code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Preamble" data-toc-modified-id="Preamble-0"><span class="toc-item-num">0 </span>Preamble</a></span><ul class="toc-item"><li><span><a href="#General-parameters" data-toc-modified-id="General-parameters-0.1"><span class="toc-item-num">0.1 </span>General parameters</a></span></li><li><span><a href="#Functions" data-toc-modified-id="Functions-0.2"><span class="toc-item-num">0.2 </span>Functions</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Midpoint-normalizer" data-toc-modified-id="Midpoint-normalizer-0.2.0.1"><span class="toc-item-num">0.2.0.1 </span>Midpoint normalizer</a></span></li></ul></li></ul></li></ul></li><li><span><a href="#A-little-exploration" data-toc-modified-id="A-little-exploration-1"><span class="toc-item-num">1 </span>A little exploration</a></span><ul class="toc-item"><li><span><a href="#Artificial" data-toc-modified-id="Artificial-1.1"><span class="toc-item-num">1.1 </span>Artificial</a></span></li><li><span><a href="#Training" data-toc-modified-id="Training-1.2"><span class="toc-item-num">1.2 </span>Training</a></span></li></ul></li><li><span><a href="#Disambiguation" data-toc-modified-id="Disambiguation-2"><span class="toc-item-num">2 </span>Disambiguation</a></span><ul class="toc-item"><li><span><a href="#Artificial" data-toc-modified-id="Artificial-2.1"><span class="toc-item-num">2.1 </span>Artificial</a></span></li><li><span><a href="#Pre-synaptic-rule" data-toc-modified-id="Pre-synaptic-rule-2.2"><span class="toc-item-num">2.2 </span>Pre-synaptic rule</a></span></li><li><span><a href="#Post-synaptic-rule" data-toc-modified-id="Post-synaptic-rule-2.3"><span class="toc-item-num">2.3 </span>Post-synaptic rule</a></span></li></ul></li></ul></div>
# -
# Thoughts about so far in why this does not work.
#
# Consider the post-synaptic case.
# 1 2 3 6 7 8
# 5
# 10 11 12 13 14 15
#
# In the post-synaptic case, the problem is that five is activated sometimes when 3 is not and 5 is activated sometimes when 12 is not (in the pre-synaptic case the problem is that five sometimes is activated when 6 or 13 are not).
#
# If we consider the problem of creating a stable connection between 3 and 5 this is what happens.
# When 3 and 5 are activated consecutively you have the the influence of the hebbian terms z_pre * z_post, but at the same time you have some anti-hebbian influence at the same time which makes the connection not so strong. Then, when 12 and 5 are activated, the connection between 3 and 5 gets the influence from the anti-hebbian terms which makes it weaker.
#
# This is the crux, if we make the hebbian term stronger so the effect of the hebbian learning beats the anti-hebbian effects in 3-5 and 12-5 then we make the connections the self-connections so strong than the inhibition is not able to suppress the pattern after.
# # Preamble
# +
import sys
sys.path.append('../')
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from mpl_toolkits.axes_grid1 import make_axes_locatable
from network import run_network_recall, train_network, run_network_recall_limit
from connectivity import designed_matrix_sequences, designed_matrix_sequences_local
from analysis import get_recall_duration_for_pattern, get_recall_duration_sequence, create_sequence_chain
from analysis import time_t1, time_t2, time_t1_local, time_t2_local, time_t2_complicated
# -
# ## General parameters
# +
# %matplotlib inline
np.set_printoptions(suppress=True, precision=5)
plt.rcParams["figure.figsize"] = [16,9]
sns.set(font_scale=3.0)
sns.set(font_scale=3.5)
sns.set_style("whitegrid", {'axes.grid': False})
lw = 8
ms = 18
alpha_graph = 0.3
color_palette = sns.color_palette()
# -
dt = 0.001
# ## Functions
# #### Midpoint normalizer
# +
class MidpointNormalize(matplotlib.colors.Normalize):
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
matplotlib.colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y))
norm = MidpointNormalize(midpoint=0)
cmap_show = matplotlib.cm.RdBu_r
# -
# # A little exploration
# ## Artificial
# +
N = 10
tau_m = 0.010
tau_z = 0.100
G = 100.0
threshold = 0.5
self_excitation = 3.0
inhibition = 1
transition = 3.0
sequences = [[i for i in range(N)]]
w = designed_matrix_sequences(N, sequences, self_excitation=self_excitation,
transition=transition, inhibition=inhibition)
plt.matshow(w, cmap='seismic')
plt.colorbar();
# -
index_from = 2
index_to = 3
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
# +
I_cue = 0
I_end = 5.0
T_cue = 3 * tau_m
t1 = time_t1(I=inhibition, T=transition, tau_z=tau_z, threshold=threshold)
t2 = time_t2(tau_z, A=self_excitation, T=transition, I=inhibition, threshold=threshold)
T_per = t1 + t2
print(t1)
print(t2)
print(T_per)
T = T_per * len(sequences[0]) + T_cue
T = 2.0
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history = np.copy(dic['x'])
z_history = dic['z']
plt.imshow(x_history, aspect='auto');
# -
pattern = 2
duration = get_recall_duration_for_pattern(x_history, pattern, dt)
print(duration)
print(t1 + t2)
start = np.where(x_history[:, 2] > 0)[0][0]
length = int(T_per / dt)
plt.plot(x_history[:, 2])
plt.axvline(start, color='red')
plt.axvline(start + length, color='red');
get_recall_duration_sequence(x_history, dt)
# ## Training
# +
N = 10
sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9]]
#sequences = [[5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
#sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
sequences =[[i for i in range(N)]]
training_time = 0.100
inter_sequence_time = 1.0
max_w = 20.0
min_w = -10.0
gh = 2.0
gah = 1.0
threshold = 0.1
tau_z = 0.050
tau_z_post = 0.005
tau_w = 1.0
epochs = 1
dic = train_network(N, dt, training_time, inter_sequence_time, sequences, tau_z,
tau_z_post, tau_w, epochs=epochs, max_w=max_w, min_w=min_w, gh=gh, gah=gah,
pre_rule=True, save_w_history=True)
w = dic['w']
plt.matshow(w, cmap=cmap_show, norm=norm)
plt.colorbar();
# -
index_from = 2
index_to = 3
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
# +
#time = np.arange(0, T, dt)
z = dic['z']
z_post = dic['z_post']
w_hist = dic['w_history']
normal = dic['normal']
negative = dic['negative']
fig = plt.figure()
ax = fig.add_subplot(211)
ax.plot(z[:, 2], label='2 pre')
ax.plot(z_post[:, 3], label='3 post')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(w_hist[:, 3, 2], label='w32')
ax.plot(normal[:, 3, 2], label='hebian')
ax.plot(negative[:, 3, 2], label='anti heb')
ax.legend(frameon=False);
# +
T = 1.0
I_end = 5.0
T_cue = 0.100
I_cue = sequences[0][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history = np.copy(dic['x'])
z_history = np.copy(dic['z'])
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.imshow(x_history, aspect='auto')
# -
# # Disambiguation
# ## Artificial
# +
N = 10
sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9]]
sequences = [[5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
#sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
N = 10
tau_m = 0.010
tau_z = 0.250
G = 100.0
threshold = 0.5
self_excitation = 2.0
inhibition = 10.0
transition = 1.0
w = designed_matrix_sequences(N, sequences, self_excitation=self_excitation,
transition=transition, inhibition=inhibition)
plt.matshow(w, cmap='seismic')
plt.colorbar();
# +
index_from = 2
index_to = 8
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
index_from = 2
index_to = 3
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
# -
t1 = time_t1(I=inhibition, T=transition, tau_z=tau_z, threshold=threshold)
t2 = time_t2(tau_z, A=self_excitation, T=transition, I=inhibition, threshold=threshold)
T_per = t1 + t2
print(t1)
print(t2)
print(T_per)
T = T_per * len(sequences[0]) + T_cue
I_end = 5.0
T_cue = 0.100
# +
I_cue = sequences[0][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_1 = np.copy(dic['x'])
z_history_1 = np.copy(dic['z'])
I_cue = sequences[1][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_2 = np.copy(dic['x'])
z_history_2 = np.copy(dic['z'])
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax1.imshow(x_history_1, aspect='auto')
ax2 = fig.add_subplot(222)
ax2.imshow(x_history_2, aspect='auto')
ax3 = fig.add_subplot(223)
ax3.imshow(z_history_1, aspect='auto')
ax4 = fig.add_subplot(224)
ax4.imshow(z_history_2, aspect='auto');
# -
I_cue
# ## Pre-synaptic rule
# +
N = 10
sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9]]
#sequences = [[5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
#sequences = [[0, 1, 2, 3, 4], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [5, 6, 2, 8, 9], [0, 1, 2, 3, 4]]
training_time = 0.100
inter_sequence_time = 1.0
max_w = 20.0
min_w = -10.0
gh = 10.0
gah = 3.0
threshold = 0.5
tau_z = 0.050
tau_z_post = 0.005
tau_w = 1.0
epochs = 1
# +
dic = train_network(N, dt, training_time, inter_sequence_time, sequences, tau_z,
tau_z_post, tau_w, epochs=epochs, max_w=max_w, min_w=min_w, gh=gh, gah=gah,
pre_rule=True, save_w_history=True)
w = dic['w']
plt.matshow(w, cmap=cmap_show, norm=norm)
plt.colorbar();
# +
time = np.arange(0, T, dt)
z = dic['z']
z_post = dic['z_post']
w_hist = dic['w_history']
normal = dic['normal']
negative = dic['negative']
fig = plt.figure()
ax = fig.add_subplot(211)
ax.plot(z[:, 2], label='2 pre')
ax.plot(z_post[:, 3], label='3 post')
ax.plot(z_post[:, 8], label='8 post')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(w_hist[:, 3, 2], label='w32')
ax.plot(normal[:, 3, 2], label='hebian')
ax.plot(negative[:, 3, 2], label='anti heb')
ax.legend(frameon=False);
# +
index_from = 2
index_to = 8
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
index_from = 2
index_to = 3
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
# +
T = 2.0
I_end = 5.0
T_cue = 0.100
I_cue = sequences[0][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_1 = np.copy(dic['x'])
z_history_1 = np.copy(dic['z'])
I_cue = sequences[1][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_2 = np.copy(dic['x'])
z_history_2 = np.copy(dic['z'])
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax1.imshow(x_history_1, aspect='auto')
ax2 = fig.add_subplot(222)
ax2.imshow(x_history_2, aspect='auto')
ax3 = fig.add_subplot(223)
ax3.imshow(z_history_1, aspect='auto')
ax4 = fig.add_subplot(224)
ax4.imshow(z_history_2, aspect='auto');
# -
# ## Post-synaptic rule
# +
dic = train_network(N, dt, training_time, inter_sequence_time, sequences, tau_z,
tau_z_post, tau_w, epochs=epochs, max_w=max_w, min_w=min_w, gh=gh, gah=gah,
pre_rule=False, save_w_history=True)
w = dic['w']
plt.matshow(w, cmap=cmap_show, norm=norm)
plt.colorbar();
# +
time = np.arange(0, T, dt)
z = dic['z']
z_post = dic['z_post']
w_hist = dic['w_history']
normal = dic['normal']
negative = dic['negative']
fig = plt.figure()
ax = fig.add_subplot(211)
ax.plot(z[:, 1], label='1 pre')
ax.plot(z_post[:, 2], label='2 post')
ax.plot(z_post[:, 8], label='8 post')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(w_hist[:, 2, 1], label='w21')
ax.plot(normal[:, 2, 1], label='hebian21')
ax.plot(negative[:, 2, 1], label='anti heb21')
ax.legend(frameon=False);
# +
index_from = 1
index_to = 2
time = np.arange(0, T, dt)
z = dic['z']
z_post = dic['z_post']
w_hist = dic['w_history']
normal = dic['normal']
negative = dic['negative']
fig = plt.figure()
ax = fig.add_subplot(211)
ax.plot(z[:, 2], label='2 pre')
ax.plot(z_post[:, 3], label='3 post')
ax.plot(z_post[:, 8], label='8 post')
ax.legend()
ax = fig.add_subplot(212)
ax.plot(w_hist[:, 3, 2], label='w32')
ax.plot(normal[:, 3, 2], label='hebian')
ax.plot(negative[:, 3, 2], label='anti heb')
ax.legend(frameon=False);
# +
index_from = 2
index_to = 8
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
index_from = 2
index_to = 3
exc_t = w[index_to, index_from]
exc_s = w[index_from, index_from]
inh = w[index_from, index_to]
print('exc_t', exc_t, 'exc_s', exc_s, 'inh', inh)
# +
T = 5.0
I_end = 5.0
T_cue = 0.100
I_cue = sequences[0][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_1 = np.copy(dic['x'])
z_history_1 = np.copy(dic['z'])
I_cue = sequences[1][0]
dic = run_network_recall_limit(N, w, G, threshold, tau_z, T, dt, I_cue, T_cue)
x_history_2 = np.copy(dic['x'])
z_history_2 = np.copy(dic['z'])
fig = plt.figure()
ax1 = fig.add_subplot(221)
ax1.imshow(x_history_1, aspect='auto')
ax2 = fig.add_subplot(222)
ax2.imshow(x_history_2, aspect='auto')
ax3 = fig.add_subplot(223)
ax3.imshow(z_history_1, aspect='auto')
ax4 = fig.add_subplot(224)
ax4.imshow(z_history_2, aspect='auto');
# -
| notebooks/2018-09-09(Disambiguation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ashikshafi08/Learning_Tensorflow/blob/main/Straight%20From%20Docs/TensorFlow_Image_Segmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="UXoJtUMR0VTG" outputId="24cc3e31-7afc-46ff-cc07-b70bf7dfab08"
# !nvidia-smi
# + [markdown] id="gcwws9xP0aN7"
# # Image Segmentation
#
# Suppose if we want to know where an object is located in the image, the shape of the object, which pixel belongs to which object etc. In this case, we want to segment the image, i.e **each pixel of the image is given a label**. Thus, the task of image segmentation is to train a neural network to output a pixel-wise mask of the image.
#
# This helps in understanding the image in the pixel level. The dataset we will be using is the Oxford-IIT Pet Dataset. The dataset consists of images, their corresponding labels, and pixel-wise masks. The masks are basically labels for each pixel.
#
# Each pixel is given one of the three categories:
# - Case 1: Pixel belonging to the pet.
# - Case 2: Pixel bordering the pet.
# - Case 3: None of the above/Surrounding pixel.
# + colab={"base_uri": "https://localhost:8080/"} id="-lCI1fS_jCcO" outputId="99ea4b45-b976-4fad-cbb8-434830021d4b"
# Downloading the tensorflow_examples
# !pip install -q git+https://github.com/tensorflow/examples.git
# + id="c3hvW6OvitAY"
# Importing the things we needed
import tensorflow as tf
from tensorflow_examples.models.pix2pix import pix2pix
import tensorflow_datasets as tfds
from IPython.display import clear_output
import matplotlib.pyplot as plt
# + [markdown] id="2oyeqJiDi6le"
# ### Download the Oxford-IIIT Pets Dataset
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 367, "referenced_widgets": ["48ecc6aec86f4f708118f199c976dc46", "<KEY>", "a2dbc159a73c494482cac3295f8ab444", "<KEY>", "<KEY>", "93e71f8fffe24d61a23bfba00115b7c0", "eb41593168e643ffa9c67597d7f241bc", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "8c148157a2c54aee89aa03e3de5f90a0", "<KEY>", "86776802238f4a15bb524a91a430de91", "2d1d9a1c718e48ed824267119754112f", "4074973b7c114853ac5afa61d8b41c53", "<KEY>", "6aeda61dd3e149fb910cf7c56396ec5e", "58ce3e40f32a4263b9e0ecc0e90ff4b3", "f4fd24615a1841c88404ee61e663ace1", "8d70c13c926048558569063e44522942", "b76e7932a3284b4d81dd0b01a8a7936b", "1ace9b31448d4d29bd955b6c07306a93", "c3b8ae184f754b6598319e066ec95609", "645a7af8f46d426d8ae6ddb4ee3637cd", "10d3a5838944446d8800dd6d92fe7cb8", "<KEY>", "1ceb24ffa1e14baea14264d7c8a6f811", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "24ea296a723c4338aef3a064f9089dee", "<KEY>", "<KEY>", "1065714db86143719e699ed0e8679055", "<KEY>", "0a68d5b502454ab599a022ad23389388", "fdfaa7605ba64e1ea88c1fb5085e6680", "<KEY>", "<KEY>", "<KEY>", "6262677481644fffad405107dbc454e6", "<KEY>", "5a7f493e2e004e838e01d745c9df14b7", "d0549a6bcfb64da1a63dc9ef31a5003d", "<KEY>", "<KEY>", "6952530d6fbc4ea18b75278357251e9b", "<KEY>", "7a5670cab16243749c5e5556b609e76b", "0b8a2a773caf48bf88deff92bad59323"]} id="Dv2QAB88jX-b" outputId="78fe450d-5ade-42bf-ef0b-8536669dae0b"
datasets , info = tfds.load('oxford_iiit_pet:3.*.*' , with_info = True)
# + [markdown] id="JLp0CYrqjekg"
# The below following code performs a simple augmentations of flipping an image and it's normalized to [0,1]. The pixels in the segmentation masks are labelled either {1, 2, 3} for the sake of convenience let's subtract 1 from the segmentation mask, resulting in labels that are : {0 , 1 ,2}
#
#
#
# + id="_QMtpYeam_TA"
# Defining the normalizing the function
def normalize(input_image , input_mask):
# Converting uint8 to float32 and normalizing it
input_image = tf.cast(input_image, tf.float32) / 255.0
input_mask -= 1
return input_image , input_mask
# + id="HB7BKEV8n3KB"
# Funtion to load image and apply some preprocessing (train set)
@tf.function
def load_image_train(datapoint):
# Resizingt the image and mask to 128 pixels
input_image = tf.image.resize(datapoint['image'] , (128 , 128))
input_mask = tf.image.resize(datapoint['segmentation_mask'] , (128 , 128))
# Randomly flip the image left and right
if tf.random.uniform(()) > 0.5:
input_image = tf.image.flip_left_right(input_image)
input_mask = tf.image.flip_left_right(input_mask)
# Applying the normalization function we created just above
input_image , input_mask = normalize(input_image , input_mask)
return input_image , input_mask
# + id="0ClIi7C-naKn"
# Funtion to load image and apply some preprocessing (test set)
def load_image_test(datapoint):
input_image = tf.image.resize(datapoint['image'] , (128 , 128))
input_mask = tf.image.resize(datapoint['segmentation_mask'] , (128 , 128))
input_image , input_mask = normalize(input_image , input_mask)
return input_image , input_mask
# + colab={"base_uri": "https://localhost:8080/"} id="LDiyAUcgoWUg" outputId="981e4def-0fdd-41e4-f877-b3c38b88c6d1"
info
# + colab={"base_uri": "https://localhost:8080/"} id="RWXi7dcfpXwj" outputId="a1b8177c-d628-4eba-d408-fcd5beb7c7d8"
# How many splits are there?
info.splits
# + id="tSX8MjQ8rjAv"
# Setting up some hyperparameters for our model
TRAIN_LENGTH = info.splits['train'].num_examples
BATCH_SIZE = 64
BUFFER_SIZE = 1000
STEPS_PER_EPOCH = TRAIN_LENGTH // BATCH_SIZE
# + [markdown] id="ugazn25SrtJm"
# Now splitting train and test into two variables and apply preprocessing methods on it.
# + colab={"base_uri": "https://localhost:8080/"} id="eDByKn06sGer" outputId="fc0d8875-b864-4fc0-f403-0c862eb36e96"
train = datasets['train'].map(load_image_train , num_parallel_calls = tf.data.AUTOTUNE)
test = datasets['test'].map(load_image_test)
train , test
# + [markdown] id="HS1NAziHsXvn"
# Inside the dataset
# - (128, 128, 3): This will be our image
# - (128, 128, 1): Mask of the image
# + colab={"base_uri": "https://localhost:8080/"} id="LhRg4b4UssYv" outputId="684cc949-c088-4d04-a760-283bb9221566"
# Applying prefetch and performance optimization methods on our data
train_dataset = train.cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE).repeat()
train_dataset = train_dataset.prefetch(buffer_size = tf.data.AUTOTUNE)
test_dataset = test.batch(BATCH_SIZE)
train_dataset , test_dataset
# + [markdown] id="JXvWb78utP6M"
# Let's take a look at an image example and it's correponding mask from the dataset.
# + id="9ARzzV__tXKG"
def display(display_list):
plt.figure(figsize = (15 , 15))
title = ['Input Image' , 'True Mask' , 'Predicted Mask']
for i in range(len(display_list)):
#print(display_list)
plt.subplot(1 , len(display_list) , i+1)
plt.title(title[i])
plt.imshow(tf.keras.preprocessing.image.array_to_img(display_list[i]))
plt.axis(False)
plt.show
# + colab={"base_uri": "https://localhost:8080/"} id="2rq8etW3txvT" outputId="1d02d8c4-118b-4d2f-e520-44363726da4c"
for image , mask in train.take(3):
sample_image , sample_mask = image , mask
sample_mask
# + colab={"base_uri": "https://localhost:8080/", "height": 320} id="T7KZXQTit6Vb" outputId="2064480d-1e64-4ba1-9943-ae5d4b19697a"
# Using the display function to plot both the input and masked image
display([sample_image , sample_mask])
# + id="Xz4pdh1CuUaV"
# + id="U69YTrH5uw8M"
| Straight From Docs/TensorFlow_Image_Segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="iU56TiIo0tbi" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629490453525, "user_tz": 240, "elapsed": 202, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="22239712-d2bb-466c-911b-b66004f2e777"
import pandas as pd
import numpy as np
import math
import matplotlib.pyplot as plt
import os
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="e867j_QnZBsZ"
# 512 HRV data per (future) CSV
# Lower threshold for outliers of 500
# Higher threshold for outliers of 2000
# + id="rnJFVl39ZIRn"
data_per_csv = 512
l_th = 500
h_th = 2000
# + [markdown] id="sK3HqQtn-XXB"
# Access data path
# + id="WhYhoh9t1alG"
def dataPath(c):
if c=='d':
return 'gdrive/My Drive/Summer Research/HRV/Diabetes/'
elif c=='h':
return 'gdrive/My Drive/Summer Research/HRV/Healthy/'
else:
raise ValueError('Parameter must be h or d')
train_data_path = 'gdrive/My Drive/Summer Research/HRV/Outlier Free'
# + [markdown] id="ZqVOAvl8Zkzn"
# New data path (to store the new CSVs)
# + id="XJP1qs0iZnzz"
def newDataPath(c):
if c=='d':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Diabetes/'
elif c=='h':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Healthy/'
else:
raise ValueError('Parameter must be h or d')
# + [markdown] id="yNJIwi7b-Zxp"
# Returns array of DataFrames for each of the CSVs for each individual
# + id="LZl4lLaM4lEj"
def HRV_DataFrame(c,num):
data_path = dataPath(c)+str(f'{num:03d}')+'/'
df = []
for files in os.listdir(data_path):
if files.endswith('.csv'):
s = pd.read_csv(data_path+files)
if s.empty == False:
s = pd.read_csv(data_path+files,usecols=[1],skiprows=[0])
df.append(s)
return df
# + [markdown] id="HUO0plE8w135"
# Plots HRV values for each individual
# + id="HYzWDSREf25i"
def plotHRV(c,num):
df = HRV_DataFrame(c,num)
plt.figure(figsize=(16,9))
for i in range(len(df)):
plt.plot(df[i], label=str(i+1), linewidth=2)
plt.title('CGM Prediction')
plt.xlabel('HRV Interval')
plt.ylabel('R to R (ms)')
plt.legend()
# + [markdown] id="7dFBC-Z4vxB_"
# Returns array of lists; detectOutliers[i][j] returns the jth value in the ith list
# + id="e-ebihTMkBgy"
def detectOutliers(c,num,lower_threshold,higher_threshold):
dfs = HRV_DataFrame(c,num)
o = []
for csv_num in range(len(dfs)):
df = dfs[csv_num]
p = list()
for i in range(len(df)):
if df.iloc[i][0] < lower_threshold or df.iloc[i][0] > higher_threshold:
p.append(i)
o.append(p)
return o
# + [markdown] id="YgLEuZ6s1ItM"
# Returns the start and length of outlier-free intervals
# + id="Kpsr6VPYMRI0"
def outlierFreeIntervalLengths(c,num,lower_threshold,higher_threshold):
dfs = detectOutliers(c,num,lower_threshold,higher_threshold)
p = list()
for csv_num in range(len(dfs)):
df = dfs[csv_num]
o = np.zeros((len(df)-1,2))
for i in range(len(df)-1):
o[i,0] = int(df[i]+1)
o[i,1] = int((df[i+1]-df[i]))
p.append(o)
return p
# + [markdown] id="7ha6N8rzjugP"
# Create folder structure
# + id="MUwCDpOVjwr6"
def folderStructure():
for i in range(9):
try:
s = newDataPath('d')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
for i in range(20):
try:
s = ''
if i == 11:
s = newDataPath('h')+f'{(i+1):03d}'+'_diabetes/'
else:
s = newDataPath('h')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
# + id="TVZjeksekPcp"
folderStructure()
# + [markdown] id="cIM70gkYXOsb"
# Put the outlier-free intervals into equal-length CSVs
# + id="zb8d1XI4WHgj"
def outlierFreeToCSV(c,num,lower_threshold,higher_threshold):
interval_length = outlierFreeIntervalLengths(c,num,lower_threshold,higher_threshold)
csvs = HRV_DataFrame(c,num)
dir = ''
if c == 'h' and num == 12:
dir = f'{num:03d}' + '_diabetes/'
else:
dir = f'{num:03d}' + '/'
count = 0
for i in range(len(interval_length)):
#access each array in the list
a = interval_length[i]
for j in range(len(a)):
if a[j,1] >= data_per_csv:
length = int(a[j,1] / data_per_csv)
for k in range(length):
count += 1
location = newDataPath(c)+dir+f'{count:03d}'+'.csv'
start = int(a[j,0]) + k*data_per_csv
end = int(a[j,0]) + (k+1)*data_per_csv
new_csv = csvs[i].loc[range(start,end),:]
new_csv.to_csv(location, index=False, header=False)
# + id="AznM_71aow23"
for i in range(9):
outlierFreeToCSV('d',i+1,l_th,h_th)
# + id="T_eI2EX_8DIx"
for i in range(20):
outlierFreeToCSV('h',i+1,l_th,h_th)
# + [markdown] id="UO3v8yrQUflB"
# Wavelet Transformed data path (to store the new new CSVs)
# + id="iEbMTl4MUjs4"
def WTDataPath(c):
if c=='d':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Wavelet Transformed Data/Diabetes/'
elif c=='h':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Wavelet Transformed Data/Healthy/'
else:
raise ValueError('Parameter must be h or d')
# + id="Z3FljkZxMcxy"
def WTDenoisedDataPath(c):
if c=='d':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Wavelet Transform Denoised Data/Diabetes/'
elif c=='h':
return 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Wavelet Transform Denoised Data/Healthy/'
else:
raise ValueError('Parameter must be h or d')
# + [markdown] id="8zWtp-RtP65g"
# Wavelet Transform
# + id="TE_y2lMgQECq"
def fourWTM(n):
#Filter banks
h0 = np.array([0.2697890,0.3947890,0.5197890,0.6447890,0.2302110,0.1052110,-0.0197890,-0.1447890])
h1 = np.array([-0.2825435,0.5553379,0.2385187,-0.0783004, -0.5834819,-0.2666627,0.0501564,0.3669755])
h2 = np.array([0.4125840,-0.6279376,0.3727824,0.1487574, -0.4125840,-0.1885590,0.0354659,0.2594909])
h3 = np.array([0.2382055,0.1088646,-0.7275830,0.5572896, -0.2382055,-0.1088646,0.0204763,0.1498171])
#Matrix of filter banks created for convenience
h = np.array([h0,h1,h2,h3])
k = int(n/4)
T = np.zeros((n,n))
for j in range(4):
for i in range(k):
if 4*i+8 > 4*k:
T[k*j+i,range((4*i),(4*i+4))] = h[j,range(4)]
T[k*j+i,range(4)] = h[j,range(4,8)]
else:
T[k*j+i,range((4*i),(4*i+8))] = h[j,range(8)]
return T
# + id="y-Chiq7lQIiR"
def four_Wavelet_Transform_Decomp(c,num):
n = data_per_csv
hrv = New_HRV_Arrays(c,num)
t = fourWTM(n)
#S tilda pseudoimage
s = np.zeros((len(hrv),data_per_csv,4))
for j in range(len(hrv)):
ts = np.matmul(t,hrv[j])
a1, d1, d2, d3 = ts[0:int(n/4)], ts[int(n/4):int(n/2)], ts[int(n/2):int(3*n/4)], ts[int(3*n/4):n]
dim = int(n/4)
A1 = np.zeros((4*dim,1))
D1 = np.zeros((4*dim,1))
D2 = np.zeros((4*dim,1))
D3 = np.zeros((4*dim,1))
for i in range(dim):
A1 = A1 + a1[i]*np.transpose(t[i:(i+1),:])
for i in range(dim):
D1 = D1 + d1[i]*np.transpose(t[i+dim:(i+dim+1),:])
for i in range(dim):
D2 = D2 + d2[i]*np.transpose(t[i+2*dim:(i+2*dim+1),:])
for i in range(dim):
D3 = D3 + d3[i]*np.transpose(t[i+3*dim:(i+3*dim+1),:])
s[j,:,:] = np.concatenate((A1,D1,D2,D3),axis=1)
return s
# + id="zPBnlphwxyCT"
def four_Wavelet_Transform_Decomp_Denoised(c,num):
n = data_per_csv
dim = int(n/4)
t = fourWTM(n)
tt = np.transpose(t)
d = np.zeros((dim,3))
D = np.zeros((n,3))
dir = WTDataPath(c)+f'{num:03d}'+'/'
num_of_csv = len([name for name in os.listdir(dir) if os.path.isfile(os.path.join(dir, name))])
#S tilda pseudoimage
s = np.zeros((num_of_csv,data_per_csv,4))
signal = np.zeros((data_per_csv,4))
for k in range(num_of_csv):
sig = np.loadtxt(dir+f'{(k+1):03d}'+'.csv', delimiter=',')
A1 = sig[:,0].reshape(-1,1)
for j in range(3):
d[:,j] = np.matmul(t[(j+1)*dim:(j+2)*dim,:],sig[:,j+1])
#Denoise details
lbda = np.std(d[:,j])*math.sqrt(2*math.log(n/4))
for i in range(dim):
if abs(d[i,j]) < lbda:
d[i,j] = 0
D[:,j] = np.matmul(tt[:, (j+1)*dim:(j+2)*dim], d[:,j])
s[k,:,:] = np.concatenate((A1,D),axis=1)
return s
# + id="60MPnqGWKJ6i"
for i in range(9):
s = four_Wavelet_Transform_Decomp_Denoised('d',i+1)
for j in range(len(s)):
np.savetxt(WTDenoisedDataPath('d')+f'{(i+1):03d}'+'/'+f'{(j+1):03d}'+'.csv', s[j], delimiter=',', fmt='%f')
# + id="xIgGFRgJMLkK"
for i in range(20):
if i != 11:
s = four_Wavelet_Transform_Decomp_Denoised('h',i+1)
for j in range(len(s)):
np.savetxt(WTDenoisedDataPath('h')+f'{(i+1):03d}'+'/'+f'{(j+1):03d}'+'.csv', s[j], delimiter=',', fmt='%f')
# + [markdown] id="PiQqDWiXWu-V"
# Returns array of DataFrames for each of the CSVs for each individual
# + id="Xb2KnPcSXE2F"
def New_HRV_Arrays(c,num):
data_path = newDataPath(c)+str(f'{num:03d}')+'/'
df = []
count = 0
for files in os.listdir(data_path):
if files.endswith('.csv'):
count += 1
s = np.loadtxt(data_path+str(f'{count:03d}')+'.csv', delimiter=',').reshape(data_per_csv,1)
df.append(s)
return df
# + [markdown] id="xlNrJwWPZx02"
# Create folder structure
# + id="y6S_GSOWZzTE"
def WT_FolderStructure():
for i in range(9):
try:
s = WTDataPath('d')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
for i in range(20):
try:
if i != 11:
s = WTDataPath('h')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
# + id="nRbzFVv0aXZW"
WT_FolderStructure()
# + id="KXdiNDpXM4Xe"
def WTDenoised_FolderStructure():
for i in range(9):
try:
s = WTDenoisedDataPath('d')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
for i in range(20):
try:
if i != 11:
s = WTDenoisedDataPath('h')+f'{(i+1):03d}'+'/'
os.mkdir(s)
except OSError as error:
continue
# + id="aWPZcAHbM9r3"
WTDenoised_FolderStructure()
# + [markdown] id="A-RCufCxgABy"
# Put the Wavelet Transformed Pseudo-images into CSVs
# + id="6HKMgVTSktQk"
for i in range(9):
s = four_Wavelet_Transform_Decomp('d',i+1)
for j in range(len(s)):
np.savetxt(WTDataPath('d')+f'{(i+1):03d}'+'/'+f'{(j+1):03d}'+'.csv', s[j], delimiter=',', fmt='%f')
# + id="NJg053ExUBIe"
for i in range(20):
if i != 11:
s = four_Wavelet_Transform_Decomp('h',i+1)
for j in range(len(s)):
np.savetxt(WTDataPath('h')+f'{(i+1):03d}'+'/'+f'{(j+1):03d}'+'.csv', s[j], delimiter=',', fmt='%f')
# + [markdown] id="E-5b4MRp1LQP"
# Graph some HRV data
# + id="RFfqFXH91Ouy"
def WTPlotHRV(c,num,subnum):
n = np.loadtxt(newDataPath(c)+str(f'{num:03d}')+'/'+str(f'{subnum:03d}')+'.csv', delimiter=',').reshape(data_per_csv,1)
wt = np.loadtxt(WTDataPath(c)+str(f'{num:03d}')+'/'+str(f'{subnum:03d}')+'.csv', delimiter=',')
wt_denoised = np.loadtxt(WTDenoisedDataPath(c)+str(f'{num:03d}')+'/'+str(f'{subnum:03d}')+'.csv', delimiter=',')
plt.figure(figsize=(32,18))
plt.plot(n, label=str(num)+':'+str(subnum)+' (Without WT)', linewidth=2)
wt_dict = {
0: ['A1',1],
1: ['D1',0],
2: ['D2',2],
3: ['D3',4]
}
for i in range(4):
plt.plot(wt[:,i]-250*(wt_dict[i][1]), label=str(num)+':'+str(subnum)+' ('+wt_dict[i][0]+')', linewidth=1)
if i != 0:
plt.plot(wt_denoised[:,i]-250*(wt_dict[i][1]), label=str(num)+':'+str(subnum)+' ('+wt_dict[i][0]+' denoised)', linewidth=2)
plt.title('HRV Data')
plt.xlabel('HRV Interval')
plt.ylabel('R to R (ms)')
plt.legend()
# + id="fN2Wb0pT1YIp" colab={"base_uri": "https://localhost:8080/", "height": 810} executionInfo={"status": "ok", "timestamp": 1629160696482, "user_tz": 240, "elapsed": 2578, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="c3f89c79-928c-4965-c547-64e82f46cfc7"
WTPlotHRV('h',8,1)
# + [markdown] id="S3B_H63hBMt2"
# Check how much data there is for diabetes, healthy patients
# + colab={"base_uri": "https://localhost:8080/"} id="S2XVODUnBQiW" executionInfo={"status": "ok", "timestamp": 1628974793368, "user_tz": 240, "elapsed": 230, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="ebe878a4-765b-4565-9008-154834ee99ff"
diabetes_count = 0
healthy_count = 0
for i in range(9):
dir = WTDataPath('d')+f'{i+1:03d}'+'/'
for files in os.listdir(dir):
if files.endswith('.csv'):
diabetes_count += 1
for i in range(20):
if i != 11:
dir = WTDataPath('h')+f'{i+1:03d}'+'/'
for files in os.listdir(dir):
if files.endswith('.csv'):
healthy_count += 1
print('Diabetes: '+str(diabetes_count)+'\nHealthy: '+str(healthy_count))
# + [markdown] id="SVSZxF8_WcEP"
# Convert time series to image
# + id="a0VeTEPuWdx4"
def HRVImage(c,num,subnum):
n = np.loadtxt(newDataPath(c)+str(f'{num:03d}')+'/'+str(f'{subnum:03d}')+'.csv', delimiter=',').reshape(data_per_csv,1)
hd = {
'd':'Diabetes/',
'h':'Healthy/'
}
plt.figure(figsize=(4,3))
plt.box(False)
plt.axis('off')
bbox_inches=0
plt.plot(n, linewidth=1, color='black')
plt.margins(0,0)
plt.savefig('gdrive/My Drive/Summer Research/HRV/Outlier Free/Images/All/'+c+str(num)+'_'+str(subnum),bbox_inches='tight',pad_inches = 0,format='png')
plt.cla()
plt.clf()
plt.close('all')
# + id="gRl7FUY1XMF0"
for i in range(9):
count = 0
dir = WTDataPath('d')+f'{i+1:03d}'+'/'
for files in os.listdir(dir):
if files.endswith('.csv'):
count += 1
HRVImage('d',i+1,count)
# + id="hg8fuDp_ILoX" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629493429662, "user_tz": 240, "elapsed": 1076127, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="edbfbb9f-9afb-4a2d-e792-76468fb787e8"
for i in range(20):
print(i+1)
count = 0
if i != 11:
dir = WTDataPath('h')+f'{i+1:03d}'+'/'
for files in os.listdir(dir):
if files.endswith('.csv'):
count += 1
HRVImage('h',i+1,count)
# + [markdown] id="DqqUoZOjN6Mi"
# Make sure all images are the same size
# + id="iHEkW1iDN8F8"
from PIL import Image
def sameSize(dir):
check_size = list()
for files in os.listdir(dir):
img = np.array(Image.open(dir+files))
if img.shape != (163, 223, 4):
check_size.append(files)
return check_size
# + colab={"base_uri": "https://localhost:8080/"} id="oYI0FgNvO4Dd" executionInfo={"status": "ok", "timestamp": 1629493565613, "user_tz": 240, "elapsed": 9504, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="f2296755-c231-41a4-abbc-4e237db0ed6c"
sameSize('gdrive/My Drive/Summer Research/HRV/Outlier Free/Images/All/')
# + [markdown] id="qdlxqdwyN5-B"
#
# + colab={"base_uri": "https://localhost:8080/"} id="FXXqAMR7i5RK" executionInfo={"status": "ok", "timestamp": 1628984994886, "user_tz": 240, "elapsed": 161, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13953621006807715822"}} outputId="1fa6ab97-f43b-4b42-a810-9fd7f4c86adb"
diabetes_count = 0
count = 0
dir = 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Images/Diabetes/'
for files in os.listdir(dir):
if files.endswith('.png'):
diabetes_count += 1
count += 1
print(diabetes_count)
# + id="J6SHAfRqE0B_"
healthy_count = 0
count = np.zeros(20)
dir = 'gdrive/My Drive/Summer Research/HRV/Outlier Free/Images/Healthy/'
for files in os.listdir(dir):
healthy_count += 1
count[int(files[0])-1] += 1
print(healthy_count)
print(count)
| Code/Preprocessing/Preprocess HRV Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Accessing zarr-formatted Daymet data on Azure
#
# The [Daymet](https://daymet.ornl.gov/) dataset contains daily minimum temperature, maximum temperature, precipitation, shortwave radiation, vapor pressure, snow water equivalent, and day length at 1km resolution for North America. The dataset covers the period from January 1, 1980 to December 31, 2019.
#
# Daymet is available in both NetCDF and Zarr format on Azure; this notebook demonstrates access to the Zarr data, which can be read into an [xarray](http://xarray.pydata.org/en/stable/) [Dataset](http://xarray.pydata.org/en/stable/data-structures.html#dataset). If you just need a subset of the data, we recommend using xarray and Zarr to avoid downloading the full dataset unnecessarily.
#
# This dataset is stored in the West Europe Azure region, so this notebook will run most efficiently on Azure compute located in the same region. If you are using this data for environmental science applications, consider applying for an [AI for Earth grant](http://aka.ms/ai4egrants) to support your compute requirements.
#
# The datasets are available in the `daymeteuwest` storage account, in the `daymet-zarr` container. Files are named according to `daymet-zarr/{frequency}/{region}.zarr`, where frequency is one of `{daily, monthly, annual}` and region is one of `{hi, na, pr}` (for Hawaii, CONUS, and Puerto Rico, respectively). For example, `daymet-zarr/daily/hi.zarr`.
#
# More complete documentation is available at [aka.ms/ai4edata-daymet](http://aka.ms/ai4edata-daymet).
# ### Environment setup
# +
import warnings
import matplotlib.pyplot as plt
import xarray as xr
import fsspec
# Neither of these are accessed directly, but both need to be
# installed; they're used via fsspec
import adlfs
import zarr
# -
# ### Constants
storage_account_name = 'daymeteuwest'
container_name = 'daymet-zarr'
frequency = 'daily' # daily, monthly, annual
region = 'hi' # hi (Hawaii), na (North America), pr (Puerto Rico)
# ### Load data into an xarray Dataset
#
# We can lazily load the data into an `xarray.Dataset` by creating a zarr store with [fsspec](https://filesystem-spec.readthedocs.io/en/latest/) and then reading it in with xarray. This only reads the metadata, so it's safe to call on a dataset that's larger than memory.
url = 'az://' + container_name + '/' + frequency + '/' + region + '.zarr'
container_name = 'daymet-zarr'
storage_account_name = 'daymeteuwest'
url = 'az://' + container_name + '/' + frequency + '/' + region + '.zarr'
store = fsspec.get_mapper(url, account_name=storage_account_name)
ds = xr.open_zarr(store, consolidated=True)
ds
# ### Working with the data
#
# Using xarray, we can quickly select subsets of the data, perform an aggregation, and plot the result. For example, we'll plot the average of the maximum temperature for the year 2009.
warnings.simplefilter('ignore', RuntimeWarning)
tmax = ds.sel(time='2009')['tmax'].mean(dim='time')
w = tmax.shape[1]; h = tmax.shape[0]; dpi = 100
fig = plt.figure(frameon=False,figsize=(w/dpi,h/dpi),dpi=dpi);
ax = plt.Axes(fig,[0., 0., 1., 1.]); ax.set_axis_off(); fig.add_axes(ax)
tmax.plot(ax=ax,cmap='inferno')
plt.show();
# Or we can visualize the timeseries of the minimum temperature over the past decade.
fig, ax = plt.subplots(figsize=(12, 4))
ds.sel(time=slice('2010', '2019'))['tmin'].mean(dim=['x', 'y']).plot(ax=ax);
# ### Chunking
#
# Each of the datasets is chunked to allow for parallel and out-of-core or distributed processing with [Dask](https://dask.org/). The different frequencies (daily, monthly, annual) are chunked so that each year is in a single chunk. The different regions in the `x` and `y` coordinates so that no single chunk is larger than about 250 MB, which is primarily important for the `na` region.
ds['prcp']
# So our `prcp` array has a shape `(14600, 584, 284)` where each chunk is `(365, 584, 284)`. Examining the store for monthly North America, we see the chunks each have size `(12, 1250, 1250)`.
na_store = fsspec.get_mapper('az://' + container_name + '/monthly/na.zarr',
account_name=storage_account_name)
na = xr.open_zarr(na_store, consolidated=True)
na['prcp']
# See http://xarray.pydata.org/en/stable/dask.html for more on how xarray uses Dask for parallel computing.
| data/daymet-zarr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Improving a model with Grid Search
#
# In this mini-lab, we'll fit a decision tree model to some sample data. This initial model will overfit heavily. Then we'll use Grid Search to find better parameters for this model, to reduce the overfitting.
#
# First, some imports.
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ### 1. Reading and plotting the data
# Now, a function that will help us read the csv file, and plot the data.
# +
def load_pts(csv_name):
data = np.asarray(pd.read_csv(csv_name, header=None))
X = data[:,0:2]
y = data[:,2]
plt.scatter(X[np.argwhere(y==0).flatten(),0], X[np.argwhere(y==0).flatten(),1],s = 50, color = 'blue', edgecolor = 'k')
plt.scatter(X[np.argwhere(y==1).flatten(),0], X[np.argwhere(y==1).flatten(),1],s = 50, color = 'red', edgecolor = 'k')
plt.xlim(-2.05,2.05)
plt.ylim(-2.05,2.05)
plt.grid(False)
plt.tick_params(
axis='x',
which='both',
bottom='off',
top='off')
return X,y
X, y = load_pts('data.csv')
plt.show()
# -
# ### 2. Splitting our data into training and testing sets
# +
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score, make_scorer
#Fixing a random seed
import random
random.seed(42)
# Split the data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
# ### 3. Fitting a Decision Tree model
# +
from sklearn.tree import DecisionTreeClassifier
# Define the model (with default hyperparameters)
clf = DecisionTreeClassifier(random_state=42)
# Fit the model
clf.fit(X_train, y_train)
# Make predictions
train_predictions = clf.predict(X_train)
test_predictions = clf.predict(X_test)
# -
# Now let's plot the model, and find the testing f1_score, to see how we did.
# The following function will help us plot the model.
def plot_model(X, y, clf):
plt.scatter(X[np.argwhere(y==0).flatten(),0],X[np.argwhere(y==0).flatten(),1],s = 50, color = 'blue', edgecolor = 'k')
plt.scatter(X[np.argwhere(y==1).flatten(),0],X[np.argwhere(y==1).flatten(),1],s = 50, color = 'red', edgecolor = 'k')
plt.xlim(-2.05,2.05)
plt.ylim(-2.05,2.05)
plt.grid(False)
plt.tick_params(
axis='x',
which='both',
bottom='off',
top='off')
r = np.linspace(-2.1,2.1,300)
s,t = np.meshgrid(r,r)
s = np.reshape(s,(np.size(s),1))
t = np.reshape(t,(np.size(t),1))
h = np.concatenate((s,t),1)
z = clf.predict(h)
s = s.reshape((np.size(r),np.size(r)))
t = t.reshape((np.size(r),np.size(r)))
z = z.reshape((np.size(r),np.size(r)))
plt.contourf(s,t,z,colors = ['blue','red'],alpha = 0.2,levels = range(-1,2))
if len(np.unique(z)) > 1:
plt.contour(s,t,z,colors = 'k', linewidths = 2)
plt.show()
plot_model(X, y, clf)
print('The Training F1 Score is', f1_score(train_predictions, y_train))
print('The Testing F1 Score is', f1_score(test_predictions, y_test))
# Woah! Some heavy overfitting there. Not just from looking at the graph, but also from looking at the difference between the high training score (1.0) and the low testing score (0.7).Let's see if we can find better hyperparameters for this model to do better. We'll use grid search for this.
#
# ### 4. (TODO) Use grid search to improve this model.
#
# In here, we'll do the following steps:
# 1. First define some parameters to perform grid search on. We suggest to play with `max_depth`, `min_samples_leaf`, and `min_samples_split`.
# 2. Make a scorer for the model using `f1_score`.
# 3. Perform grid search on the classifier, using the parameters and the scorer.
# 4. Fit the data to the new classifier.
# 5. Plot the model and find the f1_score.
# 6. If the model is not much better, try changing the ranges for the parameters and fit it again.
#
# **_Hint:_ If you're stuck and would like to see a working solution, check the solutions notebook in this same folder.**
# +
from sklearn.metrics import make_scorer
from sklearn.model_selection import GridSearchCV
clf = DecisionTreeClassifier(random_state=42)
# TODO: Create the parameters list you wish to tune.
parameters = {'max_depth':[3,4,5,6,7,8,9,10],'min_samples_leaf':[2,3,4,5,6,7,8,9,10],'min_samples_split':[2,3,4,5,6,7,8,9,10]}
# TODO: Make an fbeta_score scoring object.
scorer = make_scorer(f1_score)
# TODO: Perform grid search on the classifier using 'scorer' as the scoring method.
grid_obj = GridSearchCV(clf, parameters, scoring=scorer)
# TODO: Fit the grid search object to the training data and find the optimal parameters.
grid_fit = grid_obj.fit(X,y)
# TODO: Get the estimator.
best_clf = grid_fit.best_estimator_
# Fit the new model.
best_clf.fit(X_train, y_train)
# Make predictions using the new model.
best_train_predictions = best_clf.predict(X_train)
best_test_predictions = best_clf.predict(X_test)
# Calculate the f1_score of the new model.
print('The training F1 Score is', f1_score(best_train_predictions, y_train))
print('The testing F1 Score is', f1_score(best_test_predictions, y_test))
# Plot the new model.
plot_model(X, y, best_clf)
# Let's also explore what parameters ended up being used in the new model.
best_clf
# -
| 2. Model Evaluation and Validation/Grid_Search_Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### <span style='color:blue'>10.</span>
#
# <!--
# HW 7: Chap 12: #10, 11, 12, 14, 19
# HW 8: Chapter 12: #25, 37*, 39, 40**
# Chapter 11: #1, 2, 17 (Hint: see equation on p. 408), 20, 22, 25
# -->
#
#
# **HW Review:**
# <p><strong>Diffraction and crystallography</strong></p>
#
#
# <p><b>11.2 Describe the “phase problem” in X-ray crystallography, and at least one way the problem can be addressed (or at least circumvented to solve X-ray structures).</b></p>
#
# <p><u>See Page 420 for phase problem, See Page 421 for the way the problem can be addressed.</u></p>
#
#
#
# <p><b>11.20 Draw a set of points as a rectangular array based on unit cells of side a and b, and mark the planes with Miller indices (1,0,0), (0,1,0), (1,1,0), (1,2,0), (2,3,0), (4,1,0)<!--, and (4,$\bar{1}$,0)-->.</b>
# </p>
#
# Here's an example...
#
# $$(1,2,0) = (k,h,l) \implies (\frac{a}{h},\frac{b}{k},0) = (\frac{a}{1},\frac{b}{2},0) \\ \implies 2\times(\frac{a}{1},\frac{b}{2},0) = (2a,b,0)$$
#
# <hr>
#
# <img src="images/11.20.pdf" width="700" height="700" align="center">
#
# <hr>
#
# <!--
# <p><b>11.20 (continued) The crystal unit cell of MDM2 has dimensions <em>a</em> = 43.4 Å, <em>b</em> = 100.5 Å, <em>c</em> = 54.8 Å. What is the spacing <em>d</em> of diffraction planes with Miller indices (2,1,0)?</b></p>
# <p></p>
#
#
# \begin{equation}
# \frac{1}{d^{2}}=\frac{h^{2}}{a^{2}}+\frac{k^{2}}{b^{2}}+\frac{l^{2}}{c^{2}}
# \end{equation}
#
# <p><u>See Page 418 for more information.</u></p>
# -->
#
# **Chapter 12**
#
# **12.12 A swimmer enters a gloomier world (in one sense) on diving to greater depths. Given that the mean molar absorption coefficient of seawater in the visible region is $6.2x10^{−5}$ $dm^{3}$ $mol^{−1}$ $cm^{−1}$, calculate the depth at which a diver will experience (a) half the surface intensity of light and (b) one-tenth that intensity.**
#
#
# ### Derivation of Beer's Law:
#
# Here is an image of the situation we wish to model:
#
# <img src="http://life.nthu.edu.tw/~labcjw/BioPhyChem/Spectroscopy/pics/beersla1.gif" height="300" width="300" align="center" >
#
# The density of particles, $\rho$ and the absorption coefficient, $\alpha$ multiplied by the intensity, I shown in the 1st order differential equation:
# $$ -\frac{\partial{I}}{\partial{x}} = I \alpha \rho $$
#
# Combine like-terms to each side of the equation:
#
# $$\int_{I_{0}}^{I} \frac{\partial{I}}{I} = -\int_{0}^{x} \alpha \rho \partial{x} $$
#
#
# We know that $\int \frac{1}{x}dx = ln(x)$, so
#
# $$ln(\frac{I}{I_{0}}) = - \alpha \rho x, $$
#
#
# -----------------
#
# To get the general solution of the D.E we can take the exponential of both sides
#
# $$\frac{I}{I_{0}} = e^{-\alpha \rho x} $$
#
# **General Solution to the D.E**:
#
# $$I (x) = I_{0} e^{-\alpha \rho x}$$
#
# --------------------------
#
# Otherwise, to continue deriving Beer's Law we can use the property of logarithms:
#
# $$-ln(\frac{I}{I_{0}}) = ln(\frac{I_{0}}{I}) = \alpha \rho x, $$
#
#
# and since we know the following
#
# $$log_{10}(x) = \frac{ln(x)}{ln(10)},$$
#
# then we can say
#
# $$ log_{10}(\frac{I_{0}}{I}) = \frac{\alpha \rho x}{ln(10)}$$
#
#
# Finally, we can say that $\rho \propto c$. We can also simplify further by saying $\epsilon =\frac{\alpha}{ln(10)}$, which has units of $M^{-1}cm^{-1}$ and $x = b$, where b is in cm.
#
# $$ A = log(\frac{I_{0}}{I}) = \epsilon b c$$
#
# Now, solving for the path length $b$ gives the following expression with $c_{H_{2}O} = \rho/MW$ and $I = 0.5I_{0}$.
#
#
# $$ b = \frac{log(\frac{I_{0}}{0.5I_{0}})}{\epsilon (\rho/MW)} = \frac{0.301}{(6.2 x 10^{-5} dm^{3}. mol^{-1}.cm^{-1}) (55.5 mol.dm^{-3})} = 87 cm $$
#
# **Note**, since the information regarding salt water concentration is not provided in the question we approximated the concentration by with values for $H_{2}O$.
#
#
# <!--
# **12.14 The molar absorption coefficients of tryptophan and tyrosine at 240 nm are 2.00 × 103 dm3 mol−1 cm−1 and 1.12 × 104 dm3 mol−1 cm−1, respectively, and at 280 nm they are 5.40 × 103 dm3 mol−1 cm−1 and 1.50 × 103 dm3 mol−1 cm−1. The absorbance of a sample obtained by hydrolysis of a protein was measured in a cell of thickness 1.00 cm and was found to be 0.660 at 240 nm and 0.221 at 280 nm. What are the concentrations of the two amino acids?**
# -->
#
# **12.25 How many normal modes of vibration are there for (a) $NO_{2}$, (b) $N_{2}O$, (c) cyclohexane, and (d) hexane?**
#
# There are $3N-6$ and $3N-5$ vibrational modes (in which N is the number of atoms in molecule) for non-linear and linear molecules; respectively.
#
#
# **(a)** $NO_{2}$, Non-linear; $3N-6 = 3(3)-6 = 3$
#
# **(b)** $N_{2}O$, linear; $3N-5 = 3(3)-5 = 4$
#
# **(c)** cyclohexane, non-linear; $3N-6 = 3(18)-6 = 48$
#
# **(d)** hexane, non-linear; $3N-6 = 3(20)-6 = 54$
#
#
#
# -----------------------------------------
#
# **SIDE NOTES:**
#
# ### Rates of various processes
#
# | $\text{Process}$ | $\text{Timescales (s)}$ | $\text{Radiative}$ | $\text{Transition}$ |
# | :--: | :--: | :--: | :--: |
# | IC | $10^{-14}-10^{-11}$ | N | $S_{n} \to S_{1}$ |
# | Vib Relax | $10^{-14}-10^{-11}$ | N | ${S_{n}}^{*} \to S_{n}$ |
# | Abs | $10^{-15}$ | Y | $S_{0} \to S_{n}$ |
# | Fluor | $10^{-9}-10^{-7}$ | Y | $S_{1} \to S_{0}$ |
# | ISC | $10^{-8}-10^{-3}$ | N | $S_{1} \to T_{1}$ |
# | Phos | $10^{-4}-10^{0}$ | Y | $T_{1} \to S_{0}$ |
#
#
# - timescale of FRET are typically in ns
#
#
# -----------------------------------------
#
#
#
# **12.37 When benzophenone is illuminated with ultraviolet radiation, it is excited into a singlet state. This singlet changes rapidly into a triplet, which phosphoresces. Triethylamine acts as a quencher for the triplet. In an experiment in methanol as solvent, the phosphorescence intensity Iphos varied with amine concentration as shown below. A time-resolved laser spectroscopy experiment had also shown that the half-life of the fluorescence in the absence of quencher is 29 ms. What is the value of $k_{Q}$?**
#
#
# | $Species$ | $\text{}$ | $\text{}$ | $\text{}$ |
# | :--: | :--: | :--: | :--: |
# | $[Q]/(mol\space dm^{−3})$ | 0.0010 | 0.0050 | 0.0100 |
# | $I_{phos}/(A.U.)$ | 0.41 | 0.25 | 0.16|
#
#
# First, we need to write out the mechanism that is given in the question:
#
# >When benzophenone is illuminated with ultraviolet radiation, it is excited into a singlet state.
#
# $$ M + h\nu_{i} \rightarrow M^{*} \tag{1}$$
#
# >This singlet changes rapidly into a triplet, which phosphoresces.
#
# $$ M^{*} \rightarrow M + h\nu_{phos} \tag{2}$$
#
# >Triethylamine acts as a quencher for the triplet.
#
# $$ M^{*} + Q \rightarrow M + Q \tag{3}$$
#
#
# <hr>
#
# To model this process, we apply the steady state approximation on $[M^{*}]$ to obtain $I_{phos}$... (Do this to get your own "stern-volmer" equation that models what the questions provides).
#
# **Steady State** is an assumption that the rate of (production/destruction) is equal to zero i.e., at equilibrium.
#
# $$\frac{d[M^{*}]}{dt} = I_{abs} - k_{Q}[Q][M^{*}]-k_{phos}[M^{*}]=0$$
#
# $$ \implies (-k_{Q}[Q]-k_{phos})[M^{*}] = -I_{abs} \implies [M^{*}] = \frac{I_{abs}}{k_{Q}[Q]+k_{phos}},$$
#
# and we know that $I_{phos} = k_{phos}[M^{*}]$, so
#
# $$ I_{phos} = k_{phos} \frac{I_{abs}}{k_{Q}[Q]+k_{phos}}$$
#
# We can take the inverse of $I_{phos}$ to get the equation in the form of a line:
#
# $$ \frac{1}{I_{phos}} = \frac{1}{I_{abs}} + \frac{k_{Q}[Q]}{k_{phos}I_{abs}}$$
#
# Now, we plot the data that was given and extract the slope...
# %matplotlib inline
import plot as p
import numpy as np
Q = np.array([0.0010,0.0050, 0.0100])
Iphos = np.array([0.41, 0.25, 0.16])
x,y = Q,1/Iphos
p.simple_plot(x,y,xlabel=r'$[Q]$',ylabel=r'${I_{phos}}^{-1}$',Type='scatter',color=False,fig_size=(8,4),
fit=True, order=1, annotate_text=r"$slope=k_{Q}/(k_{phos}I_{abs})$",annotate_x=-0.005, annotate_y=5.5)
# Therefore, the linear fit gives:
# $$I_{phos}^{-1}=(424.5302 dm^{3} mol )[Q]+(1.966), $$
#
# where $\frac{k_{Q}}{k_{phos}I_{abs}} = 424.5302 dm^{3} mol $.
#
# Therefore,
#
# $$k_{Q} = \frac{(24.5302 dm^{3} mol)(2.39x10^{4} s^{-1})}{1.97} = 5.2x10^{6} dm^{3} mol^{-1} s^{-1} $$
#
# <!--
# The quantum efficiency of fluorescence is $\phi_{F}$, where
# $$\phi_{F} = \frac{\text{rate of fluor}}{I_{abs}} = \frac{k_{F} [M^{*}]}{I_{abs}} $$
# $$\phi_{F} = \frac{\text{rate of fluor}}{I_{abs}} $$
# Observed fluorescence lifetime $\tau_{0}$ is defined as
# $$\tau_{0} = \frac{\phi_{F}}{k_{F}} $$
# The question gives data for $I_{phos}$, and you need to extract the slope to obtain $k_{Q}$.
# -->
#
#
#
#
#
#
#
#
#
#
# #### [Jump to table of contents.](#Table-of-Contents:)
# <hr>
#
#
#
# <p><b>What kind of information can be obtained using FRET spectroscopy? What is the distance dependence of the FRET effect?</b></p>
#
# <p>Förster resonance energy transfer (FRET) spectroscopy is useful for studying processes involving inter and intra-molecular energy transfer and can be used to measure distances (ranging from 1 to 9 nm) in biological systems. Furthermore, conformational changes can be studied, and also good for studying bulk distances. Single molecule FRET —create histograms of binned FRET distances, ultimately revealing states.<u>See Pages 500,501 for more information.</u></p>
#
# <hr>
#
# **12.39 The Förster theory of resonance energy transfer and the basis for the FRET technique can be tested by performing fluorescence measurements on a series of compounds in which an energy donor and an energy acceptor are covalently linked by a rigid molecular linker of variable and known length. <NAME> and <NAME>, Proc. Natl. Acad. Sci. USA 58, 719 (1967), collected the following data on a family of compounds with the general composition dansyl-(l-prolyl)n-naphthyl, in which the distance R between the naphthyl donor and the dansyl acceptor was varied by increasing the number of prolyl units in the linker:**
#
#
# | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ | $\text{}$ |
# | :--: | :--: | :--: | :--: | :--: | :--: | :--: | :--: | :--: | :--: | :--: |
# | $R/nm$ | 1.2 | 1.5 | 1.8 | 2.8 | 3.1 | 3.4 | 3.7 | 4.0 | 4.3 | 4.6 |
# | $\eta_{T}$ | 0.99 | 0.94 | 0.97 | 0.82 | 0.74 | 0.65 | 0.40 | 0.28 | 0.24 | 0.16 |
#
#
#
# **Are the data described adequately by the Förster theory (eqns 12.26 and 12.27)? If so, what is the value of $R_{0}$ for the naphthyl–dansyl pair?**
#
#
# <h5> Förster theory:</h5>
#
# States that the efficiency of resonance energy transfer is related to the distance $R$ between donor-acceptor pairs by
#
# $$\eta_{T} = \frac{{R_{0}}^{6}}{{R_{0}}^{6} + {R}^{6}}, $$
#
# where $R_{0}$ is the distance at which $50 \%$ of the energy is transfered from donor to acceptor, and $R$ is the distance between donor and acceptor.
#
# First, we need to rearrange the Förster theory equation into a linearized form.
#
# $$ \frac{1}{\eta_{T}} = \frac{{R_{0}}^{6} + {R}^{6}}{{R_{0}}^{6}} = 1 + (\frac{R}{R_{0}})^{6}$$
#
# Now, we are able to plot the data:
#
#
# %matplotlib inline
import plot as p
import numpy as np
R = np.array([1.2, 1.5, 1.8, 2.8, 3.1, 3.4, 3.7, 4.0, 4.3, 4.6])
nT = np.array([0.99, 0.94, 0.97, 0.82, 0.74, 0.65, 0.40, 0.28, 0.24, 0.16])
x,y = R**6,1/nT
p.simple_plot(x,y,xlabel=r'$(R/(nm))^{6}$',ylabel=r'${\eta_{T}}^{-1}$',Type='scatter',
color=False,fig_size=(8,4),fit=True, order=1)
# Using the slope of the line $y=0.000550*x+(0.971320)$, where the slope is $0.000550 = (\frac{1}{R_{0}})^{6}$.
#
# $$R_{0} = (\frac{1}{0.000550 nm^{-6}})^{1/6} = 3.5 nm$$
| CHEM3405_Physical_Chemistry_Bio/HW_Review_03-25-20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Trading Strategy Back Testing System
# This system will be composed of the following components
# - Trading Strategy/Signals
# - Performance calculation
# - Portfolio
#
# This code is influenced and based on the book "Successful Algorithmic Trading" by <NAME>. For further detaisl go to:https://www.quantstart.com/successful-algorithmic-trading-ebook
#
import pandas as pd
import numpy as np
# ### Some Helper functions
def getDictFrom2ColDataFrame(df, index_col, other_col):
direction = df[[index_col, other_col]]
direction_dict = {k: list(v)[0] for k, v in direction.groupby(index_col)[other_col]}
return direction_dict
# ### Strategy Interface
# +
from abc import ABCMeta, abstractmethod
from enum import Enum
class TradeSignal(Enum):
"""
Trade Signal
"""
Long = 1
Short = 2
Hold = 3
class Strategy(object):
"""
Strategy is an abstract base class providing an interface for
all subsequent (inherited) strategy handling objects.
The goal of a (derived) Strategy object is to generate Signal
objects for particular symbols based on the inputs of Bars
(OHLCV) generated by a DataHandler object.
This is designed to work both with historic and live data as
the Strategy object is agnostic to where the data came from,
since it obtains the bar tuples from a queue object.
"""
__metaclass__ = ABCMeta
@abstractmethod
def calculateSignals(self):
"""
Provides the mechanisms to calculate the list of signals.
"""
raise NotImplementedError("Should implement calculate_signals()")
@abstractmethod
def calculatePnl(self):
"""
Computes the PnL after taking a long position
"""
raise NotImplementedError("Should implement calculatePnl()")
@abstractmethod
def calculateReturns(self):
"""
Computes the PnL after taking a long position
"""
raise NotImplementedError("Should implement calculateReturns()")
class BuyAndHoldStrategy(Strategy):
"""
Buy and Hold strategy: Take a long position and keep hold of the stock for a duration
"""
def __init__(self, test_feature_data):
self._signals = {}
self._signals_dict = {}
self._test_feature_data = test_feature_data
self._pnl = {}
self._return = {}
def calculateSignals(self, directions, hold_period=None, index_col='date', other_col='signal'):
self._signals = directions.copy()
self._signals['signal'] = np.where((self._signals.direction == 1.0), TradeSignal.Long, TradeSignal.Hold)
#print(self._signals.head(5))
self._signals_dict = getDictFrom2ColDataFrame(self._signals, index_col, other_col)
return self._signals, self._signals_dict
def calculatePnl(self, open_column, close_column, date_column):
for k in self._signals_dict.keys():
open_price = self._test_feature_data[self._test_feature_data[date_column] == k][open_column].tolist()[0]
close_price = self._test_feature_data[self._test_feature_data[date_column] == k][close_column].tolist()[0]
if self._signals_dict[k] == TradeSignal.Long:
num_signal = 1.0
else:
num_signal = 0.0
self._pnl[k] = float(close_price - open_price)*num_signal
#print("close_price = {0} open_price = {1} date = {2} num_signal = {3}".format(close_price, open_price, k, num_signal))
return self._pnl
def calculateReturns(self, return_column, date_column):
for k in self._signals_dict.keys():
return_price = self._test_feature_data[self._test_feature_data[date_column] == k][return_column].tolist()[0]
if self._signals_dict[k] == TradeSignal.Long:
num_signal = 1.0
else:
num_signal = 0.0
self._return[k] = float(return_price)*num_signal
return self._return
# -
# ### Test Trading Strategy
# +
data_path = ".\Data\GOOGL.csv"
test_data = pd.read_csv(data_path)
test_data = test_data[test_data.date > '2017-12-30']
test_data['direction'] = np.sign(test_data['4. close'].pct_change())
test_data['return'] = test_data['4. close'].pct_change()
test_data = test_data.dropna()
test_data.head(2)
def getDirectionDict(test_data):
direction_dict = getDictFrom2ColDataFrame(test_data, index_col='date', other_col='direction')
return direction_dict
def testStrategy():
strategy = BuyAndHoldStrategy(test_data)
#directions = getDirectionDict(test_data)
directions = test_data[['date', 'direction']]
signals, signals_dict = strategy.calculateSignals(directions)
pnl = strategy.calculatePnl(open_column = '1. open', close_column = '4. close', date_column = 'date')
returns = strategy.calculateReturns(return_column='return', date_column = 'date')
print("Number the firtst 5 signals, pnl and returns are:")
dates = list(signals_dict.keys())
for i,k in enumerate(dates[:5]):
print("Signals[{0}] = {1}".format(k, signals_dict[k]))
print("pnl[{0}] = {1}".format(k, pnl[k]))
print("returns[{0}] = {1}".format(k, returns[k]))
print("\n")
testStrategy()
# -
# ### Trading Strategy Performance estimation
# +
import numpy as np
import pandas as pd
from __future__ import print_function
class PerformanceEstimation(object):
"""
"""
@staticmethod
def computeSharpeRatio(returns, periods=252):
"""
Create the Sharpe ratio for the strategy, based on a
benchmark of zero (i.e. no risk-free rate information).
Parameters:
returns - A pandas Series representing period percentage returns.
periods - Daily (252), Hourly (252*6.5), Minutely(252*6.5*60) etc.
"""
return np.sqrt(periods) * (np.mean(returns)) / np.std(returns)
@staticmethod
def computeDrawdowns(pnl):
"""
Calculate the largest peak-to-trough drawdown of the PnL curve
as well as the duration of the drawdown. Requires that the
pnl_returns is a pandas Series.
Parameters:
pnl - A pandas Series representing period percentage returns.
Returns:
drawdown, duration - Highest peak-to-trough drawdown and duration.
"""
# Calculate the cumulative returns curve
# and set up the High Water Mark
hwm = [0]
# Create the drawdown and duration series
idx = pnl.index
drawdown = pd.Series(index = idx)
duration = pd.Series(index = idx)
# Loop over the index range
for t in range(1, len(idx)):
hwm.append(max(hwm[t-1], pnl.iloc[t]))
drawdown[t]= (hwm[t]-pnl.iloc[t])
duration[t]= (0 if drawdown[t] == 0 else duration[t-1]+1)
return drawdown, drawdown.max(), duration.max()
# -
# ### Create a simple portfolio which can generate orders and manages the profit and loss
# +
# %matplotlib inline
import matplotlib.pyplot as plt
class Portfolio(object):
def __init__(self, initial_capital, signals, test_data, num_buy_shares, ticker,close_col, signal_col):
self._initial_capital = initial_capital
self._signals = signals
self._test_data = test_data
self._num_buy_shares = num_buy_shares
self._ticker = ticker
self._close_col = close_col
self._signal_col = signal_col
self._portfolio = None
self._equity_curve = None
self._stats = None
def generatePortfolioPnl(self):
# Create a DataFrame for the traded positions
positions = pd.DataFrame(index=self._signals.index).fillna(0.0)
# Buy a 100 shares
positions[self._ticker] = self._num_buy_shares*self._signals[self._signal_col]
# Initialize the portfolio with value owned
self._portfolio = positions.multiply(test_data[self._close_col], axis=0)
self._portfolio.reindex(index=test_data.date)
self._portfolio['date'] = pd.to_datetime(test_data['date'])
# Store the difference in shares owned
pos_diff = positions.diff()
# Add `holdings` to portfolio
self._portfolio['holdings'] = (positions.multiply(test_data[self._close_col], axis=0)).sum(axis=1)
# Add `cash` to portfolio
self._portfolio['cash'] = self._initial_capital - (pos_diff.multiply(test_data[self._close_col], axis=0)).sum(axis=1).cumsum()
# Add `total` to portfolio
self._portfolio['total'] = self._portfolio['cash'] + self._portfolio['holdings']
# Add `returns` to portfolio
self._portfolio['returns'] = self._portfolio['total'].pct_change()
# Remove Nan colums
self._portfolio = self._portfolio.dropna()
# Print the first lines of `portfolio`
print("Portfolio Details (some rows):\n\n{}\n".format(self._portfolio.head()))
def visualizePortfolio(self):
# Create a figure
fig = plt.figure()
ax1 = fig.add_subplot(111, ylabel='Portfolio value in $')
# Plot the equity curve in dollars
self._portfolio['total'].plot(ax=ax1, lw=2.)
ax1.plot(self._portfolio.loc[self._signals.direction == 1.0].index,
self._portfolio.total[self._signals.direction == 1.0],
'^', markersize=10, color='m')
ax1.plot(self._portfolio.loc[self._signals.direction == -1.0].index,
self._portfolio.total[self._signals.direction == -1.0],
'v', markersize=10, color='k')
plt.title("Equity Curve")
# Show the plot
plt.show()
def visualizeEquityCurve(self):
# Create a figure
fig = plt.figure()
ax1 = fig.add_subplot(111, ylabel='Equity Curve $')
# Plot the equity curve in dollars
self._equity_curve['equity_curve'].plot(ax=ax1, lw=2.)
ax1.plot(self._equity_curve.index,
self._equity_curve['equity_curve'], color='b')
# Show the plot
plt.show()
def generateEquityCurve(self):
"""
"""
"""
Creates a pandas DataFrame from the all_holdings
list of dictionaries.
"""
curve = self._portfolio.copy()
#curve.set_index('datetime', inplace=True)
curve['returns'] = curve['total'].pct_change()
curve['equity_curve'] = (1.0+curve['returns']).cumprod()
self._equity_curve = curve.dropna()
print("Equity Curve Details (some rows):\n\n{}\n".format(self._equity_curve.head()))
return self._equity_curve
def outputSummaryStats(self):
"""
Creates a list of summary statistics for the portfolio.
"""
total_return = self._equity_curve['equity_curve'].iloc[-1]
returns = self._equity_curve['returns']
pnl = self._equity_curve['equity_curve']
sharpe_ratio = PerformanceEstimation.computeSharpeRatio(returns, periods=252)
drawdown, max_dd, dd_duration = PerformanceEstimation.computeDrawdowns(pnl)
self._equity_curve['drawdown'] = drawdown
self._stats = [("Total Return", "%0.2f%%" % ((total_return - 1.0) * 100.0)),
("Sharpe Ratio", "%0.2f" % sharpe_ratio),
("Max Drawdown", "%0.2f%%" % (max_dd * 100.0)),
("Drawdown Duration", "%d" % dd_duration)]
print("The Output Stats for thsi back test are:\n\n")
for x in self._stats:
print(x)
print("\n")
# -
# ### Test Created Trading Portfolio
# +
strategy = BuyAndHoldStrategy(test_data)
directions = test_data[['date', 'direction']]
signals, signals_dict = strategy.calculateSignals(directions)
portfolio = Portfolio(
initial_capital = 100000.0,
signals = signals,
test_data = test_data,
num_buy_shares = 100,
ticker = 'GOOGL',
close_col = '4. close',
signal_col = 'direction')
portfolio.generatePortfolioPnl()
portfolio.visualizePortfolio()
equity_curve = portfolio.generateEquityCurve()
portfolio.outputSummaryStats()
# -
| SRC/Backtesting/TradingStrategyBackTesting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Watch Me Code 1: Say My Name
#
# This program will say your name a number of times.
#
# +
name = input("What is your name? ")
times = int(input("How many times would you like me to say your name %s? " % name))
i = 1
while i <=times:
print(name)
i = i + 1
# -
| content/lessons/05/Watch-Me-Code/WMC1-Say-My-Name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# # TensorQTL QTL association testing
# + [markdown] kernel="SoS"
# This pipeline conduct QTL association tests using tensorQTL.
# + [markdown] kernel="SoS"
# 
# + [markdown] kernel="SoS"
# ## Input
# - `--molecular-pheno`, The bed.gz file containing the table describing the molecular phenotype. It shall also have a tbi index accompaning it.
#
#
# - `genotype_list` a list of whole genome plink file for each chromosome
#
#
# - `grm_list` is a file containing list of grm matrixs that generated by the GRM module of this pipeline.
#
# - `covariate` is a file with #id + samples name as colnames and each row a covariate: fixed and known covariates as well as hidden covariates recovered from factor analysis.
#
# ## Output
#
# A sets of summary statistics files for each chromosome, for both nomial significance for each test, as well as region (gene) level association evidence.
#
#
# **FIXME: please fix the statement below**
#
#
#
#
#
# + [markdown] kernel="SoS"
# # Command interface
# + kernel="Bash"
sos run TensorQTL.ipynb -h
# + [markdown] kernel="SoS"
# ## Example
# **FIXME: add it**
# + [markdown] kernel="SoS" tags=[]
# ## Global parameter settings
#
# The section outlined the parameters that can be set in the command interface.
#
# **FIXME: same comments as in APEX.ipynb**
# + kernel="SoS"
[global]
# Path to the input molecular phenotype file, per chrm, in bed.gz format.
parameter: molecular_pheno_list = path
# Covariate file, in similar format as the molecular_pheno
parameter: covariate = path
# Genotype file in plink trio format, per chrm
parameter: genotype_file_list = path
# An optional subset of region list containing a column of ENSG gene_id to limit the analysis
parameter: region_list = path("./")
# Path to the work directory of the analysis.
parameter: cwd = path('./')
# Specify the number of jobs per run.
parameter: job_size = 2
# Container option for software to run the analysis: docker or singularity
parameter: container = ''
# Prefix for the analysis output
parameter: name = 'ROSMAP'
# Specify the scanning window for the up and downstream radius to analyze around the region of interest, in units of bp
parameter: window = ['1000000']
import pandas as pd
molecular_pheno_chr_inv = pd.read_csv(molecular_pheno_list,sep = "\t")
geno_chr_inv = pd.read_csv(genotype_file_list,sep = "\t")
input_inv = molecular_pheno_chr_inv.merge(geno_chr_inv, on = "#id")
input_inv = input_inv.values.tolist()
# + [markdown] kernel="SoS"
# ## QTL Sumstat generation
# This step generate the cis-QTL summary statistics and vcov (covariate-adjusted LD) files for downstream analysis from summary statistics. The analysis is done per chromosome to reduce running time.
# + [markdown] kernel="Bash"
# ## Cis QTL Sumstat generation via tensorQTL
#
#
# + kernel="SoS"
[TensorQTL_cis_1]
input: for_each = "input_inv"
output: f'{cwd:a}/{path(_input_inv[1]):bnnn}.cis_qtl_pairs.{_input_inv[0]}.parquet',
f'{cwd:a}/{path(_input_inv[1]):bnnn}.emprical.cis_sumstats.txt',
long_table = f'{cwd:a}/{path(_input_inv[1]):bnnn}.norminal.cis_long_table.txt'
task: trunk_workers = 1, trunk_size = 1, walltime = '12h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'
bash: expand= "$[ ]", stderr = f'{_output[0]}.stderr', container = container,stdout = f'{_output[0]}.stdout'
touch $[_output[0]].time_stamp
python: expand= "$[ ]", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout' , container = container
import pandas as pd
import numpy as np
import tensorqtl
from tensorqtl import genotypeio, cis, trans
## Defineing parameter
plink_prefix_path = $[path(_input_inv[2]):nr]
expression_bed = $[path(_input_inv[1]):r]
covariates_file = "$[covariate]"
Prefix = "$[_output[0]:nnn]"
## Loading Data
phenotype_df, phenotype_pos_df = tensorqtl.read_phenotype_bed(expression_bed)
### Filter by the optional keep gene
##if $[region_list].is_file():
## region = pd.read_csv("$[region_list]","\t")
## keep_gene = region["gene_ID"].to_list()
## phenotype_df = phenotype_df.query('gene_ID in keep_gene')
## phenotype_pos_df = phenotype_pos_df.query('gene_ID in keep_gene')
covariates_df = pd.read_csv(covariates_file, sep='\t', index_col=0).T
pr = genotypeio.PlinkReader(plink_prefix_path)
genotype_df = pr.load_genotypes()
variant_df = pr.bim.set_index('snp')[['chrom', 'pos']]
## Retaining only common samples
phenotype_df = phenotype_df[np.intersect1d(phenotype_df.columns, covariates_df.index)]
phenotype_df = phenotype_df[np.intersect1d(phenotype_df.columns, genotype_df.columns)]
covariates_df = covariates_df.transpose()[np.intersect1d(phenotype_df.columns, covariates_df.index)].transpose()
if "chr" in variant_df.chrom[0]:
phenotype_pos_df.chr = [x.replace("chr","") for x in phenotype_pos_df.chr]
## cis-QTL mapping: nominal associations for all variant-phenotype pairs
cis.map_nominal(genotype_df, variant_df,
phenotype_df,
phenotype_pos_df,
Prefix, covariates_df=covariates_df)
## Load the parquet and save it as txt
pairs_df = pd.read_parquet("$[_output[0]]")
pairs_df.columns.values[6] = "pval"
pairs_df.columns.values[7] = "beta"
pairs_df.columns.values[8] = "se"
pairs_df = pairs_df.assign(
alt = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[-1])).assign(
ref = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[-2])).assign(
pos = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[0].split(":")[1])).assign(
chrom = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split(":")[0]))
pairs_df.to_csv("$[_output[2]]", sep='\t',index = None)
cis_df = cis.map_cis(genotype_df, variant_df,
phenotype_df,
phenotype_pos_df,
covariates_df=covariates_df, seed=999)
cis_df.index.name = "gene_id"
cis_df.to_csv("$[_output[1]]", sep='\t')
# + [markdown] kernel="SoS"
# ## Trans QTL Sumstat generation via tensorQTL
#
# + kernel="SoS"
[TensorQTL_trans_1]
input: for_each = "input_inv"
output: f'{cwd:a}/{path(_input_inv[1]):bnnn}.trans_sumstats.txt'
parameter: batch_size = 10000
parameter: pval_threshold = 1e-5
parameter: maf_threshold = 0.05
task: trunk_workers = 1, trunk_size = 1, walltime = '12h', mem = '40G', tags = f'{step_name}_{_output[0]:bn}'
bash: expand= "$[ ]", stderr = f'{_output[0]}.stderr', container = container,stdout = f'{_output[0]}.stdout'
touch $[_output[0]].time_stamp
python: expand= "$[ ]", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout',container =container
import pandas as pd
import numpy as np
import tensorqtl
from tensorqtl import genotypeio, cis, trans
## Defineing parameter
plink_prefix_path = $[path(_input_inv[2]):nr]
expression_bed = $[path(_input_inv[1]):r]
covariates_file = "$[covariate]"
Prefix = "$[_output[0]:nnn]"
## Loading Data
phenotype_df, phenotype_pos_df = tensorqtl.read_phenotype_bed(expression_bed)
##### Filter by the optional keep gene
##if $[region_list].is_file():
## region = pd.read_csv("$[region_list]","\t")
## keep_gene = region["gene_ID"].to_list()
## phenotype_df = phenotype_df.query('gene_ID in keep_gene')
## phenotype_pos_df = phenotype_pos_df.query('gene_ID in keep_gene')
covariates_df = pd.read_csv(covariates_file, sep='\t', index_col=0).T
pr = genotypeio.PlinkReader(plink_prefix_path)
genotype_df = pr.load_genotypes()
variant_df = pr.bim.set_index('snp')[['chrom', 'pos']]
## Retaining only common samples
phenotype_df = phenotype_df[np.intersect1d(phenotype_df.columns, covariates_df.index)]
covariates_df.transpose()[np.intersect1d(phenotype_df.columns, covariates_df.index)].transpose()
## Trans analysis
trans_df = trans.map_trans(genotype_df, phenotype_df, covariates_df, batch_size=$[batch_size],
return_sparse=True, pval_threshold=$[pval_threshold], maf_threshold=$[maf_threshold])
## Filter out cis signal
trans_df = trans.filter_cis(trans_df, phenotype_pos_df.T.to_dict(), variant_df, window=$[window])
## Output
trans_df.columns.values[1] = "gene_ID"
trans_df.columns.values[6] = "pval"
trans_df.columns.values[7] = "beta"
trans_df.columns.values[8] = "se"
trans_df = pairs_df.assign(
alt = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[1])).assign(
ref = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[2])).assign(
pos = lambda dataframe: dataframe['variant_id'].map(lambda variant_id:variant_id.split("_")[0]))
trans_df.to_csv("$[_output[0]]", sep='\t')
# + [markdown] kernel="SoS"
# **FIXME: we can consolidate these steps. I'll take a look myself after we have the MWE test**
# + kernel="SoS"
[TensorQTL_cis_2]
input: output_from("TensorQTL_cis_1")["long_table"], group_by = "all"
output: f'{cwd:a}/{name}.TensorQTL_recipe.tsv',f'{cwd:a}/{name}.TensorQTL_column_info.txt'
python: expand = "$[ ]", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'
import csv
import pandas as pd
data_tempt = pd.DataFrame({
"#chr" : [int(x.split(".")[-4].replace("chr","")) for x in [$[_input:r,]]],
"sumstat_dir" : [$[_input:r,]],
"column_info" : $[_output[1]:r]
})
column_info_df = pd.DataFrame( pd.Series( {"ID": "GENE,CHR,POS,A0,A1",
"CHR": "chrom",
"POS": "pos",
"A0": "ref",
"A1": "alt",
"SNP": "variant_id",
"STAT": "beta",
"SE": "se",
"P": "pval",
"TSS_D": "tss_distance",
"AF": "af",
"MA_SAMPLES": "ma_samples",
"MA_COUNT": "ma_count",
"GENE": "phenotype_id"}), columns = ["TensorQTL"] )
data_tempt.to_csv("$[_output[0]]",index = False,sep = "\t" )
column_info_df.to_csv("$[_output[1]]",index = True,sep = "\t" )
# + kernel="SoS"
[TensorQTL_trans_2]
input: group_by = "all"
output: f'{cwd:a}/{name}.TensorQTL_recipe.tsv',f'{cwd:a}/{name}.TensorQTL_column_info.txt'
python: expand = "$[ ]", stderr = f'{_output[0]}.stderr', stdout = f'{_output[0]}.stdout'
import csv
import pandas as pd
data_tempt = pd.DataFrame({
"#chr" : [int(x[0].split(".")[-5].replace("chr","")) for x in [$[_input:r,]]],
"sumstat_dir" : [$[_input:r,]],
"column_info" : $[_output[1]:r]
})
column_info_df = pd.DataFrame( pd.Series( {"ID": "GENE,CHR,POS,A0,A1",
"CHR": "chrom",
"POS": "pos",
"A0": "ref",
"A1": "alt",
"SNP": "variant_id",
"STAT": "beta",
"SE": "se",
"P": "pval",
"TSS_D": "tss_distance",
"AF": "af",
"MA_SAMPLES": "ma_samples",
"MA_COUNT": "ma_count",
"GENE": "phenotype_id"}), columns = ["TensorQTL"] )
data_tempt.to_csv("$[_output[0]]",index = False,sep = "\t" )
column_info_df.to_csv("$[_output[1]]",index = True,sep = "\t" )
# + kernel="SoS"
tss_distance af ma_samples ma_count
# + kernel="SoS"
Info TensorQTL
GENE,CHR,POS,A0,A1
chrom
pos
ref
alt
variant_id
beta
se
tss_distance
af
ma_samples
ma_count
phenotype_id
| code/association_scan/TensorQTL/TensorQTL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Defining a Global Variable within a Class
class NameofClass:
CLASS_ATTR_NAME = attr_value
# The main use of Class Attributes:
# - Making use of global constants related to the class
# +
#defining a class "Player" that has a class attribute MAX_POSITION with value 10
class Player:
MAX_POSITION = 10
def __init__(self):
self.position = 0
#the __init__() method that sets the position instance attribute to 0
p = Player() #creating a player p object and printing its maximum position
print(p.MAX_POSITION)
| OOP-Fundamentals/4_OOP-Instance-and-Class-Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clean dataset
#
# In the previous notebook, we used Web Scraping to collect information from Wikipedia webpages and combine them together in a .CSV file named 'Dataset'. However, this dataset is not available in the most useful format and must be cleaned before any analysis could be performed. Such a scenario is expected to occur as the web pages aren't always designed to make extracting data easy.
#
# Here, we'll understand the dataset columns and try to remove any unnecessary strings and values that bring no value to the dataset.
# ## Import libraries and read CSV file
#
# We'll first import the Pandas library which enables us to manipulate and work with the dataset. Then, we'll simply read the dataset file we created.
# +
import re
import numpy as np
import pandas as pd
dataset = pd.read_csv("Dataset.csv")
dataset.head(5)
# -
# ## Replace the headings
#
# We first begin by replacing the headings of the columns such that they reflect the data in them better. We change the text for the first column to 'Country', the third column to 'Percentage of World Population' and fourth column to 'Total Area (km2)'.
dataset.rename(columns={'Country(or dependent territory)': 'Country'}, inplace = True)
dataset.rename(columns={'% of worldpopulation': 'Percentage of World Population'}, inplace = True)
dataset.rename(columns={'Total Area': 'Total Area (km2)'}, inplace = True)
dataset.head(5)
# ## Analysing the dataset
#
# We see that almost all columns have cells which have data inside parentheses and square brackets which is not required. Thus, we can first remove all paranthesis, square brackets and the content inside them.
# +
for column in dataset.columns:
dataset[column] = dataset[column].str.replace(r"\(.*\)", "")
dataset[column] = dataset[column].str.replace(r"\[.*\]", "")
dataset.head(5)
# -
# Next, we do not need '%' sign in either column 3 or 5, thus, we can strip the cells of it.
# +
dataset['Percentage of World Population'] = dataset['Percentage of World Population'].str.strip('%')
dataset['Percentage Water'] = dataset['Percentage Water'].str.strip('%')
dataset['Percentage Water'] = dataset['Percentage Water'].str.strip()
dataset.sample(5)
# -
# Next, we remove commas from Population column.
# +
dataset['Population'] = dataset['Population'].str.replace(',', '')
dataset.head(10)
# -
# Now, we will explore the area column. Initially, we see that the information is represented in two units: sq mi and km2. We need to convert all values to km2.
#
# The formula to convert 'sq mi' to km2 is to multiply the value by 2.58999.
#
# First, we check if the cell has the units as 'sq mi', then we multiply it with 2.589999, convert it to integer and save it back to the cell else we simply convert it into integer. Before this, on taking a closer look at the values, some cells have range of areas and as a result we need to split the data at '-' and then take the first value to continue further.
# +
dataset['Total Area (km2)'] = dataset['Total Area (km2)'].str.replace(',', '')
for x in range(len(dataset['Total Area (km2)'])):
area = dataset.iloc[x]['Total Area (km2)']
if ('sq\xa0mi' in area):
area = area.split('-')[0]
area = re.sub(r'[^0-9.]+', '', area)
area = int(float(area) * 2.58999)
else:
area = area.split('-')[0]
area = re.sub(r'[^0-9.]+', '', area)
area = int(float(area))
dataset.iloc[x]['Total Area (km2)'] = area
dataset.head(5)
# -
# Let's analyse the 'Percentage Water' column further.
# For Algeria, Afghanistan, and some other countries, the value is negligible. Hence, in order to retain data and not drop these rows, we will mark these cells with 0.0.
# Chile has the character 'b' in the end which needs to be removed.
# For the columns where the value is more than 100, the actual values were missing and other content has been read instead. Thus, we must remove such rows due to lack of information.
# +
dataset['Percentage Water'] = dataset['Percentage Water'].replace('negligible', '0.0')
dataset['Percentage Water'] = dataset['Percentage Water'].replace('Negligible', '0.0')
dataset['Percentage Water'] = dataset['Percentage Water'].str.replace(r'[^0-9.]+', '')
dataset = dataset[dataset['Percentage Water'].astype(float) <= 100]
dataset.head(5)
# -
# Total GDP includes the values in the form of trillions, billions and millions. We can remove '$' and convert the words to numbers.
# +
dataset['Total Nominal GDP'] = dataset['Total Nominal GDP'].str.replace('$', '')
for x in range(len(dataset['Total Nominal GDP'])):
gdp = dataset.iloc[x]['Total Nominal GDP']
if ('trillion' in dataset.iloc[x]['Total Nominal GDP']):
gdp = re.sub(r'[^0-9.]+', '', gdp)
gdp = int(float(gdp) * 1000000000000)
elif ('billion' in dataset.iloc[x]['Total Nominal GDP']):
gdp = re.sub(r'[^0-9.]+', '', gdp)
gdp = int(float(gdp) * 1000000000)
elif ('million' in dataset.iloc[x]['Total Nominal GDP']):
gdp = re.sub(r'[^0-9.]+', '', gdp)
gdp = int(float(gdp) * 1000000)
else:
gdp = int(re.sub(r'[^0-9.]+', '', gdp))
dataset.iloc[x]['Total Nominal GDP'] = gdp
# -
# We can remove the '$' sign as well as comma from the Per Capita GDP as well.
# +
dataset['Per Capita GDP'] = dataset['Per Capita GDP'].str.replace(r'[^0-9.]+', '')
dataset.head(10)
# -
dataset.to_csv("Final_dataset.csv", index = False)
# ## Conclusion
#
# We have iterated through all columns of the dataset and removed unnecesaary characters, and unified all data into a common format. Each column presented its own set of difficulties that had to be tackled with.
# The final dataset is ready which we can use for further analysis.
| Clean dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1><span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# -
import torch
from transformers import GPT2LMHeadModel
model = GPT2LMHeadModel.from_pretrained('distilgpt2')
opt = torch.optim.Adam(model.parameters())
for epoch in range(total_epochs):
for train_step, (lm_data, edit_example) in enumerate(data_loader):
inner_opt = torch.optim.SGD(model.transformer.h[-3:].parameters(), lr=0.01)
with higher.innerloop_ctx(model, inner_opt, copy_initial_weights=False) as (fmodel, diffopt):
for edit_step in range(n_edit_steps):
loss = fmodel(edit_example).loss
diffopt.step(loss)
# Now we have fmodel which is \hat{theta}
# as well as model (the original model), which is just theta
l_base = model(lm_data).loss
l_edit = fmodel(edit_example).loss
l_loc = kl(model(lm_data).logits, fmodel(lm_data).logits)
total_loss = lambda1 * l_base + lambda2 * l_edit + lambda3 * l_loc
total_loss.backward()
| notebooks/higher.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/madhurapg/madhurapg.github.io/blob/master/gif.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="17wLKj_lQ3wo" colab_type="code" colab={}
# %%capture
# !pip install geopandas
# !pip install carto2gpd
# !pip install tornado==5.1.1
import tornado
# !pip install holoviews
# + id="whIACndEP-AS" colab_type="code" colab={}
import matplotlib.pyplot as plt
import geopandas as gpd
import numpy as np
import pandas as pd
import dask.array as da
import dask
import dask.dataframe as dd
#import carto2gpd
# + id="kHtLVLDPSHB0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="81691a5c-685e-4858-b1ca-d797ae129eeb"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="hupjbUZLnZKR" colab_type="text"
# 311 requests dataset: https://www.opendataphilly.org/dataset/311-service-and-information-requests
# + id="ni2dauTXanxn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 430} outputId="d63552d5-4a7d-4ab8-accd-4345a91c8120"
# Import the dataset
file = open("/content/drive/My Drive/Data/public_cases_fc.csv")
req = pd.read_csv(file)
req.head()
# + id="yJLSa9ayKfjG" colab_type="code" colab={}
# %%capture
# !pip install datashader
# + id="-yDvf8iQLIYx" colab_type="code" colab={}
# conversion to web mercator coordinate system
from datashader.utils import lnglat_to_meters
# create new columns 'x' (longitude) and 'y' (latitude)
req['x'], req['y'] = lnglat_to_meters(req['lon'], req['lat'])
# keep a clean dataframe
req = req.drop(['lon', 'lat'], axis=1)
# + id="CynZfIicd1bv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 395} outputId="9c352e0f-3d0f-431b-8406-a6c4b576f267"
# Create date and time columns
req['requested_datetime'] = pd.to_datetime(req['requested_datetime'])
req['requested_datetime'].head()
req['Month'] = req['requested_datetime'].dt.month
req['Year'] = req['requested_datetime'].dt.year
req['Day'] = req['requested_datetime'].dt.dayofweek
req['Day_name'] = req['requested_datetime'].dt.day_name()
req.head()
# + id="xXuIEWjiQ6kj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4f3a9987-2748-43c7-b163-1adda078535c"
req.Day.unique()
# + id="ptaEEvPz4Tfh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="45592a2b-7217-4785-d515-d4505041696c"
# Get counts of requests by department
pd.options.display.max_rows = 4000
req.agency_responsible.unique()
counts = req.groupby('agency_responsible')['objectid'].nunique()
counts = counts.sort_values()
print(counts)
# + id="Kw7sKTloe_fI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="e7f9e1ef-e6b6-4201-e752-b7459692ab7f"
# Convert to dask dataframe
data = dd.from_pandas(req, npartitions=1)
print('%s Rows' % len(data))
print('Columns:', list(data.columns))
# + id="lwHfETiBgCLv" colab_type="code" colab={}
import holoviews as hv
# + id="sVrzmhIZfIgt" colab_type="code" colab={}
points = hv.Points(data, kdims=['x', 'y'])
from holoviews.operation.datashader import datashade
import datashader as ds
import datashader.transfer_functions as tf
# Color map imports
from datashader.colors import Greys9, viridis, inferno
from colorcet import fire, bmy, bmw, bgy, cwr
datashade(points, cmap=fire).opts(width=800, height=600, bgcolor="black")
plot_width = 900
plot_height = int(plot_width*1)
# + id="URhjx9gLId_M" colab_type="code" colab={}
# %%capture
# !pip install esri2gpd
# + [markdown] id="9sf78ysxn9nZ" colab_type="text"
# Data source for Philly zipcodes: https://www.opendataphilly.org/dataset/zip-codes
# + id="oY2krf4wHMwA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="97317c39-865e-47fe-a3ab-8518827a5544"
# add Philly zipcode boundaries (coords must be in 3857)
url = "https://services.arcgis.com/fLeGjb7u4uXqeF9q/arcgis/rest/services/Zipcodes_Poly/FeatureServer/0/"
import esri2gpd
zip_codes = esri2gpd.get(url)
zip_codes = zip_codes.to_crs(epsg=3857)
# + id="G59tqbMVzZv4" colab_type="code" colab={}
def create_image(df, x_range, y_range, Cmap, w=plot_width, h=plot_height):
# create the canvas
cvs = ds.Canvas(plot_width=w, plot_height=h, x_range=x_range, y_range=y_range)
# plot 311 requests positions
agg = cvs.points(df, 'x', 'y', ds.count())
# shade
img = tf.shade(agg, cmap=Cmap, how='eq_hist')
# return an PIL image
return tf.set_background(img, "black").to_pil()
# + id="M8cNHE5bzvQ_" colab_type="code" colab={}
def plot_violations_by_day(fig, ax, data_all_days, day, zip_codes, x_range, y_range):
"""
Plot the violations for particular day of the week
"""
#some_values = ['Fire Department'] #this can be used to filter data by agency responsible
#fire_data = data_all_days.loc[data_all_days["agency_responsible"].isin(some_values)]
# trim to the specific day of the week
df_this_day = data_all_days.loc[data_all_days["Day"] == day]
day_name = list(df_this_day["Day_name"])[0]
# create the datashaded image
img = create_image(df_this_day, x_range, y_range, cwr)
# plot the image on a matplotlib axes
plt.clf()
ax = fig.gca()
ax.imshow(img, extent=[x_range[0], x_range[1], y_range[0], y_range[1]])
ax.set_axis_off()
# plot the zipcode boundaries
zip_codes.to_crs(epsg=3857).plot(ax=ax, facecolor="none", edgecolor="white")
# add a text label for the day
ax.text(
0.05,
0.9,
"311 Service and Information \n Requests in Philadelphia",
color="white",
fontsize=20,
ha="left",
transform=ax.transAxes,
)
# add a title
ax.text(
0.6,
0.1,
day_name,
color="white",
fontsize=40,
ha="left",
transform=ax.transAxes,
)
# draw the figure and return the image
fig.canvas.draw()
image = np.frombuffer(fig.canvas.tostring_rgb(), dtype="uint8")
image = image.reshape(fig.canvas.get_width_height()[::-1] + (3,))
return image
# + id="v3-4OmVizzlP" colab_type="code" colab={}
Philly = ((-75.28, -74.96), (39.86, 40.14))
x_range, y_range = [list(r) for r in lnglat_to_meters(Philly[0], Philly[1])]
# + id="S8RYIqV0z3g9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 575} outputId="87e8eb7d-7d40-4f9a-91c0-9a4364132fc6"
import imageio
# create a figure
fig, ax = plt.subplots(figsize=(10,10), facecolor='black')
# Create an image for each month
imgs = []
for day in range(7):
img = plot_violations_by_day(fig, ax, data, day, zip_codes, x_range=x_range, y_range=y_range)
imgs.append(img)
# + id="SyYlhFXf1dIE" colab_type="code" colab={}
# Combing the images for each month into a single GIF
imageio.mimsave('/content/drive/My Drive/Data/311_days.gif', imgs, fps=1);
| gif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class MyBook (Book):
def __init__(self, title, author, price):
super().__init__(title, author)
self.price = price
def display(self):
print("Title: "+ title)
print("Author: "+ author)
print("Price: "+ str(price))
| hacker-rank/30 Days of Code/13. Abstract Classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Zonal Mean Temperature
#
# Calculating the zonal mean of a quantity
# +
# %matplotlib inline
import cosima_cookbook as cc
import matplotlib.pyplot as plt
# +
from joblib import Memory
memory = Memory(cachedir='/g/data1/v45/cosima-cookbook/', verbose=0)
# -
cc.start_cluster(diagnostics_port=8787)
expts = cc.get_experiments('mom01v5')[:4]
expts
# Zonal mean allow us to look at the ocean in cross-section. In this diagnostics, we calculate the time averaged, zonal mean temperature field.
@memory.cache
def calc_zonal_mean_temp(expt, n=1):
print('Calculating {} zonal_mean_temp'.format(expt))
if expt == 'mom01v5/KDS75':
ncfile = 'ocean_month.nc'
else:
ncfile = 'ocean.nc'
zonal_temp = cc.get_nc_variable(expt, ncfile, 'temp',
chunks={'st_ocean': None},
n=n)
zonal_mean_temp = zonal_temp.mean('xt_ocean').mean('time')
zonal_mean_temp.load()
return zonal_mean_temp
# In this example, we assume that the cells are all of the same thickness in the $x$ direction. It would be accurate to perform a weighted average using the `dzt` field.
def plot_zonal_mean_temp(zonal_mean_temp):
zonal_mean_temp.plot()
plt.gca().invert_yaxis()
plt.title('{}: Zonal Mean Temp'.format(expt))
plt.figure(figsize=(16,8))
for n, expt in enumerate(expts):
plt.subplot(2,2,n+1)
zonal_mean_temp = calc_zonal_mean_temp(expt)
plot_zonal_mean_temp(zonal_mean_temp)
| DocumentedExamples/zonal_mean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#IMPORTACION DE LIBRERIAS#
import instaloader
import instabot
# +
#LOGIN INSTABOT#
bot = instabot.Bot()
bot.login(username = "patagonian_review", password = "<PASSWORD>")
#LOGIN INSTALOADER#
L = instaloader.Instaloader()
L.login("patagonian_review","Com<PASSWORD>")
# -
| Santi/Instabot testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Denzel-Loyd-Macasero/OOP58003/blob/main/%3CMacasero%2C_Denzel_Loyd_M__FinalExam%3E.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 130} id="sNNJrJQByEu0" outputId="956e703c-f10a-4a3a-d891-7635b000c988"
from tkinter import *
window = Tk()
window.title("GUI Form")
window.geometry("400x300+20+10")
lbl1 = Label(window, text = "Find the smallest number")
lbl1.grid(row=0, column=1, columnspan=3, sticy=EW)
lbl2 = Label(window, text ="Enter the first number:")
lbl2.grid(row=0, column = 0)
ent2 = Entry(window, bd=3, textvariable=con0fent2)
ent2.grid(row=1, column =1)
con0fent2 = StringVar()
lbl3 = Label(window, text = "Enter the second number: ")
lbl3.grid(row=2, colum=1)
ent3 = Entry(window,bd=3, textvarible=con0fent3)
ent3.grid(row=2,column=3)
con0fent3=Stringvar()
lbl4 = Label(window, text="Enter the third number: ")
lbl4.grid(row=3,column =0)
ent4 = Entry(window, bd=3, textvariable=con0fent4)
ent4.grid(row=3, column=1)
btn1 = Button(window, text = "find the smallest no.")
btn1.grid(row=4, column = 1)
lbl5 = Label(window, text="The smallest number:")
lbl5.grid(row=5, column=0)
con0fLargest = StringVar()
ent5 = Entry(window, bd=3, state="readonly")
ent5.grid(row=5, column=1)
def findSmallest():
L = []
L.append(eval(con0fent2.get()))
L.append(eval(con0fent3.get()))
L.append(eval(con0fent4.get()))
con0Largest.set(max(L))
window.mainloop()
mainloop()
| <Macasero,_Denzel_Loyd_M__FinalExam>.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# three types of print
# pprint
# debug print
# pydotprint
# +
# logistic regression
import numpy
import theano
import theano.tensor as T
rng = numpy.random
N = 400
feats = 784
D = (rng.randn(N, feats), rng.randint(size = N, low = 0, high = 2))
# サンプル数400で、変数が784個ある(全てが標準正規分布に従って出力される)
# 各サンプルに対して結果変数として0,1のどちらかが報告されている
# この二つの情報をまとめたものがD(tuple)
training_steps = 10000
x = T.matrix("x") # 変数を入れる箱
y = T.vector("y") # 結果を入れる箱
w = theano.shared(rng.randn(feats))
b = theano.shared(0., name = "b") # bは定数項?
print("Initial Value:")
print(w.get_value())
print(b.get_value())
# wは係数
p_1 = 1 / (1 + T.exp(-T.dot(x, w) - b)) # dot is inner product
prediction = p_1 > 0.5 # 論理値として、p_1が0.5よりも大きいと1を出力する(振り分け)
xent = -y * T.log(p_1) - (1 - y) * T.log(1 - p_1) # 尤度関数の逆
# 下のcost functionはgenralized logistic regressionのコスト関数
cost = xent.mean() + 0.01 * (w ** 2).sum()
gw, gb = T.grad(cost, [w,b]) # gradient descentをやるよ
# compile
train = theano.function(inputs = [x, y], outputs = [prediction, xent]
,updates = ((w, w - 0.1 * gw), (b, b - 0.1 * gb))) # 0.1 is a step
predict = theano.function(inputs = [x], outputs = prediction)
# pprint
theano.printing.pprint(prediction)
# -
# pre-compilation
theano.printing.debugprint(prediction)
# post-compilation
theano.printing.debugprint(predict)
# pre-compilation graph
theano.printing.pydotprint(prediction, outfile="pic/logreg_pydotprint_prediction.png", var_with_name_simple=True)
# optimized graph
theano.printing.pydotprint(train, outfile="pic/logreg_pydotprint_train.png", var_with_name_simple=True)
| theano tutorial 4 __drawing theano graphs__.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
dist = tf.distributions.Normal(loc=0., scale=3.)
dist.cdf(1.)
# Evaluate the cdf at 1, returning a scalar.
#sess = tf.Session()
#sess.run(dist.cdf(1.))
# -
tf.__version__
# +
# import quandl
# q_api_key = "bB4wp5--7XrkpGZ7-gxJ"
# quandl.ApiConfig.api_key = q_api_key
# MinPercentileDays = 100
# QuandlAuthToken = "" # not necessary, but can be used if desired
# Stocks=['GE', 'AMD', 'F', 'AAPL', 'AIG', 'CHK', 'MU', 'MSFT', 'CSCO', 'T']
# df = quandl.get_table('WIKI/PRICES', ticker=Stocks, qopts = { 'columns': ['ticker', 'volume','adj_close'] },
# date = { 'gte': '2009-01-01', 'lte': '2015-1-1' }, paginate=True)
# df.to_csv("/Users/andrewplaate/mlp3/10Stocks.csv")
# +
from importlib import reload
import policy_gradient
reload(policy_gradient)
#reload(trading_env)
# +
import gym
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
from matplotlib import interactive
interactive(True)
import gym_trading
env = gym.make('trading-v0')
#env.time_cost_bps = 0 #
env = env.unwrapped
import tensorflow as tf
import policy_gradient
# create the tf session
# Input
num_actions=10 # same as # of stocks
Variables=3
obs_dim=num_actions*Variables
NumOfHiddLayers=2
architecture = "LSTM" # right now, valid inputs are LSTM and FFNN
LR="GD"
actFunc="relu"
regulizer="l2"
regulizerScale=0.0001
#avgfilename="/home/s1793158/mlp3/FILE_Name.p"
actFuncs=["lrelu"]
name=["RNN_GD_relu_l2","RNN_GD_elu_l2","RNN_GD_lrelu_l2","RNN_GD_selu_l2","RNN_GD_selu_l2","RNN_GD_sig_l2"]
name=["RNN_GD_sig_l2"]
# from tensorflow.python.framework import ops
#ops.reset.default_graph()
for i in range(1):
tf.reset_default_graph()
sess = tf.InteractiveSession()
#tf.reset_default_graph()
# tf.set_random_seed(0)
# with tf.Session() as sess:
tf.set_random_seed(0)
#avgfilename="/afs/inf.ed.ac.uk/user/s17/s1793158/mlp3/Saving/"+name[i]+".p"
#Modelfilename="/afs/inf.ed.ac.uk/user/s17/s1793158/mlp3/SavedModels/"+name[i]
avgfilename="/Users/andrewplaate/mlp3/Saving/"+name[i]+".p"
Modelfilename="/Users/andrewplaate/mlp3/"+name[i]
sess = tf.InteractiveSession()
pg = policy_gradient.PolicyGradient(sess, obs_dim=obs_dim,
num_actions=num_actions,
NumOfLayers=NumOfHiddLayers,
Num_Of_variables=Variables,
LR=LR,
architecture=architecture,
actFunc=actFuncs[i],
learning_rate=1e-4,
regulizer =regulizer,
regulizerScale=regulizerScale,
avgfilename=avgfilename,
Modelfilename=Modelfilename,
num_hiddenRNN=24,
DropoutMemoryStates= True,
DropoutVariational_recurrent=True,
output_keep_prob=0.8,
state_keep_prob=0.8
)
# and now let's train it and evaluate its progress. NB: this could take some time...
direc="aa"
load_model=False
#grads, grads_clipped = pg.get_grads_and_clipping()
'''with tf.Session() as sess:
sess.run(init) # actually running the initialization op
_grads_clipped, _grads = sess.run(
[grads_clipped, grads],
feed_dict={pg.X: epX, pg._tf_epr: epr, pg._tf_y: epy, pg._tf_x: epx})'''
df,sf = pg.train_model(env, episodes=2, log_freq=1, load_model=False,model_dir = direc)#, load_model=True)
sess.close()
# +
# import matplotlib as mpl
# import numpy as np
# import pandas as pd
# import matplotlib.pyplot as plt
# from matplotlib import interactive
# interactive(True)
# import pickle as pkl
# loca="/Users/andrewplaate/mlp3/TestRNN/"
# # adam3 = pkl.load( open( loca+"FFN_Adam3_lrelu_l2.p", "rb" ) )
# # adam4 = pkl.load( open( loca+"FFN_Adam4_lrelu_l2.p", "rb" ) )
# # rms3 = pkl.load( open( loca+"FFN_RMS3_lrelu_l2.p", "rb" ) )
# # rms4 = pkl.load( open( loca+"FFN_RMS4_lrelu_l2.p", "rb" ) )
# # gd3 = pkl.load( open( loca+"FFN_GD3_lrelu_l2.p", "rb" ) )
# # adam3 = pkl.load( open( loca+"RNN_Adam2_lrelu_l2.p", "rb" ) )
# # adam4 = pkl.load( open( loca+"RNN_Adam3_lrelu_l2.p", "rb" ) )
# # rms3 = pkl.load( open( loca+"RNN_RMS2_lrelu_l2.p", "rb" ) )
# # rms4 = pkl.load( open( loca+"RNN_RMS3_lrelu_l2.p", "rb" ) )
# # gd3 = pkl.load( open( loca+"RNN_GD2_lrelu_l2.p", "rb" ) )
# adam3 = pkl.load( open( loca+"RNN_Adam_lrelu_l2_e3.p", "rb" ) )
# adam4 = pkl.load( open( loca+"RNN_Adam_lrelu_l2_e4.p", "rb" ) )
# rms3 = pkl.load( open( loca+"RNN_RMSProp_lrelu_l2_e3.p", "rb" ) )
# rms4 = pkl.load( open( loca+"RNN_RMSProp_lrelu_l2_e4.p", "rb" ) )
# gd3 = pkl.load( open( loca+"RNN_GD_lrelu_l2_e3.p", "rb" ) )
# gd4 = pkl.load( open( loca+"RNN_GD_lrelu_l2_e4.p", "rb" ) )
# plt.figure(figsize=(12,12))
# plt.plot(adam3[0,:],label="adam3")
# plt.plot(adam4[0,:],label="adam4")
# plt.plot(rms3[0,:],label="rms3")
# plt.plot(rms4[0,:],label="rms4")
# plt.plot(gd3[0,:],label="gd3")
# plt.legend()
# +
# meanss=pd.DataFrame(np.array([np.mean(adam3,axis=1),np.mean(adam4,axis=1),np.mean(rms3,axis=1),np.mean(rms4,axis=1),
# np.mean(gd3,axis=1),np.mean(gd4,axis=1)]))
# meanss.index= [name1,name2,name3,name4,name5,name6]
# meanss.columns=["Mean Sort","Mean Ret"]
# meanss
# std=pd.DataFrame(np.array([np.std(adam3,axis=1),np.std(adam4,axis=1),np.std(rms3,axis=1),np.std(rms4,axis=1),
# np.std(gd3,axis=1),np.std(gd4,axis=1)]))
# std.index= [name1,name2,name3,name4,name5,name6]
# std.columns=["Std Sort","Std Ret"]
# std
# DF=meanss.join(std)
# DF=DF.loc[:,["Mean Sort","Std Sort","Mean Ret","Std Ret"]]
# print(DF.round(4).to_latex())
# +
# name1="Adam(1e-3)"
# name2="Adam(1e-4)"
# name3="RMSP(1e-3)"
# name4="RMSP(1e-4)"
# name5="SGD(1e-3)"
# name6="SGD(1e-4)"
# aa=np.arange(-0.05,0.05,0.001)
# aa=100
# alphaa=0.3
# SortRet=0
# #plt.title("Total Returns")
# plt.figure(figsize=(12,6))
# plt.title("Sortino Ratio", size=18)
# #plt.title("Total Returns",size=18)
# plt.hist(adam3[SortRet,:],label= name1,alpha = alphaa,bins=aa)
# plt.hist(adam4[SortRet,:],label=name2,alpha = alphaa,bins=aa)
# plt.hist(rms3[SortRet,:],label=name3, alpha = alphaa,bins=aa)
# plt.hist(rms4[SortRet,:],label=name4, alpha =alphaa,bins=aa)
# plt.hist(gd3[SortRet,:],label=name5, alpha = alphaa,bins=aa)
# plt.hist(gd4[SortRet,:],label=name6, alpha = alphaa,bins=aa)
# plt.xticks(size=20)
# plt.yticks(size=20)
# plt.ylabel("Count",size=20)
# plt.legend( fontsize=20)
# plt.savefig('/Users/andrewplaate/mlp3/gym_trading/envs/'+"RNN_Sort_LR")
| gym_trading/envs/CW4 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Capstone project
#
# ### Project8: Traffic Light Classification
# +
import os
import cv2
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import sys
import glob
import yaml
from sklearn.utils import shuffle
from sklearn.model_selection import train_test_split
from keras.models import load_model
from keras.models import Sequential
from keras.layers import Flatten, Dense, Lambda
from keras.layers.core import Dense, Activation, Flatten, Dropout
from keras.layers import Cropping2D
from keras.layers.convolutional import Conv2D
from keras.layers.pooling import MaxPooling2D
from keras import backend as K
tf.__version__
# +
# Grab test images
img_dir = "classification_images" # Enter Directory of all images
data_path = os.path.join(img_dir,'*g')
images = glob.glob(data_path)
classification_images = []
test_images = []
# Create the list of images to be classified
for image in images:
classification_images.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images.append(np.expand_dims(img, axis=0))
# Model path
model_path = "ssd_mobilenet_v1_coco_11_06_2017/frozen_inference_graph.pb"
# Load the Tensorflow model into memory.
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(model_path, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name ='')
sess = tf.Session(graph = detection_graph)
print(len(classification_images))
# +
# Define input and output tensors (i.e. data) for the object detection classifier
# Input tensor is the image
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Output tensors are the detection boxes, scores, and classes
# Each box represents a part of the image where a particular object was detected
detection_boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Each score represents level of confidence for each of the objects.
# The score is shown on the result image, together with the class label.
detection_scores = detection_graph.get_tensor_by_name('detection_scores:0')
detection_classes = detection_graph.get_tensor_by_name('detection_classes:0')
# Number of objects detected
num_detections = detection_graph.get_tensor_by_name('num_detections:0')
# +
# Perform the actual detection by running the model with the image as input
images_with_boxes = []
# Detect traffic light boxes, scores for detection and classes
def detection(expand_image):
(boxes, scores, classes, num) = sess.run(
[detection_boxes, detection_scores, detection_classes, num_detections],
feed_dict ={image_tensor: expand_image})
return boxes, scores, classes
# Draw boxes on image
def draw_boxes(image, boxes, scores, classes):
for parameter in zip(boxes[0], classes[0], scores[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * image.shape[0])
x_max = int(box[2] * image.shape[0])
y_min = int(box[1] * image.shape[1])
y_max = int(box[3] * image.shape[1])
image = cv2.rectangle(image, (y_min,x_min),(y_max,x_max), (0,255,0), 5)
images_with_boxes = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
return image
# +
boxes = []
scores = []
classes = []
for i in range (len(classification_images)):
boxes, scores, classes = detection(test_images[i])
images_with_boxes.append(draw_boxes(classification_images[i], boxes, scores, classes))
# +
# Load train data
train_data = yaml.load(open("data_train/train.yaml"))
labels = []
train_images = []
# Cut the image looking at the boxes in the yaml
def cut_boxes(train_data):
for train_datum in train_data:
if len(train_datum['boxes']) != 0:
image_path = train_datum['path']
image = cv2.cvtColor(cv2.imread(image_path), cv2.COLOR_RGB2BGR)
for box in train_datum['boxes']:
label = box['label']
x_min = int(box['x_min'])
x_max = int(box['x_max'])
y_min = int(box['y_min'])
y_max = int(box['y_max'])
if (x_max - x_min) > 10:
train_image = image[y_min:y_max,x_min:x_max,:]
if train_image.shape[0] != 0:
try:
train_image = cv2.resize(train_image,(14,32))
train_images.append(train_image)
labels.append(label)
len(labels)
except:
pass
cut_boxes(train_data)
# -
plt.figure()
plt.imshow(train_images[6])
plt.savefig("train_images.png")
# +
# Load simulation data
# Red
red_path = "data_train_sim/tl_data_red" # Enter Directory of all images
data_red_sim_path = os.path.join(red_path,'*g')
images_sim_red = glob.glob(data_red_sim_path)
images_sim_red.sort()
# Green
green_path = "data_train_sim/tl_data_green" # Enter Directory of all images
data_green_sim_path = os.path.join(green_path,'*g')
images_sim_green = glob.glob(data_green_sim_path)
images_sim_green.sort()
# Yellow
yellow_path = "data_train_sim/tl_data_yellow" # Enter Directory of all images
data_yellow_sim_path = os.path.join(yellow_path,'*g')
images_sim_yellow = glob.glob(data_yellow_sim_path)
images_sim_yellow.sort()
# -
print(images_sim_red[0])
print(images_sim_green[0])
print(images_sim_yellow[0])
# +
# Simulation image labeling
labels_sim = []
image_sim_paths = []
for image in images_sim_red:
labels_sim.append('R')
image_sim_paths.append(image)
for image in images_sim_green:
labels_sim.append('G')
image_sim_paths.append(image)
for image in images_sim_yellow:
labels_sim.append('Y')
image_sim_paths.append(image)
# +
# # Shuffle simlation data (images and labels)
# data_list = list(zip(labels_sim, image_sim_paths))
# random.shuffle(data_list)
# labels_sim, image_sim_paths = zip(*data_list)
# +
# Perform traffic light detection on simulation data
classification_images_red = []
test_images_red = []
classification_images_green = []
test_images_green = []
classification_images_yellow = []
test_images_yellow = []
boxes_sim = []
classes_sim = []
scores_sim = []
sim_images_box = []
labels_sim = []
# RED CASE
# Create the list of images to be classified
for image in images_sim_red:
classification_images_red.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_red:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_red.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_sim_red)):
# for i in range(1):
boxes_sim, scores_sim, classes_sim = detection(test_images_red[i])
for parameter in zip(boxes_sim[0], classes_sim[0], scores_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_red[i].shape[0])
x_max = int(box[2] * classification_images_red[i].shape[0])
y_min = int(box[1] * classification_images_red[i].shape[1])
y_max = int(box[3] * classification_images_red[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_red[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
sim_image_box = cv2.resize(image,(14,32))
sim_image_box = cv2.cvtColor(sim_image_box, cv2.COLOR_BGR2RGB)
sim_images_box.append(sim_image_box)
labels_sim.append('R')
except:
pass
# GREEN CASE
# Create the list of images to be classified
for image in images_sim_green:
classification_images_green.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_green:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_green.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_sim_green)):
# for i in range(1):
boxes_sim, scores_sim, classes_sim = detection(test_images_green[i])
for parameter in zip(boxes_sim[0], classes_sim[0], scores_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_green[i].shape[0])
x_max = int(box[2] * classification_images_green[i].shape[0])
y_min = int(box[1] * classification_images_green[i].shape[1])
y_max = int(box[3] * classification_images_green[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_green[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
sim_image_box = cv2.resize(image,(14,32))
sim_image_box = cv2.cvtColor(sim_image_box, cv2.COLOR_BGR2RGB)
sim_images_box.append(sim_image_box)
labels_sim.append('G')
except:
pass
# YELLOW CASE
# Create the list of images to be classified
for image in images_sim_yellow:
classification_images_yellow.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_yellow:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_yellow.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_sim_yellow)):
# for i in range(1):
boxes_sim, scores_sim, classes_sim = detection(test_images_yellow[i])
for parameter in zip(boxes_sim[0], classes_sim[0], scores_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_yellow[i].shape[0])
x_max = int(box[2] * classification_images_yellow[i].shape[0])
y_min = int(box[1] * classification_images_yellow[i].shape[1])
y_max = int(box[3] * classification_images_yellow[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_yellow[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
sim_image_box = cv2.resize(image,(14,32))
sim_image_box = cv2.cvtColor(sim_image_box, cv2.COLOR_BGR2RGB)
sim_images_box.append(sim_image_box)
labels_sim.append('Y')
except:
pass
# +
print(len(sim_images_box))
print(len(labels_sim))
plt.figure()
plt.imshow(sim_images_box[0])
plt.savefig("sim_images_box0.png")
# -
# Label refactor: squeexe classes into the 4 we are interested in
for i in range(len(labels)):
if labels[i] == 'Yellow':
labels[i] = 'Y'
elif labels[i] == 'Red':
labels[i] = 'R'
elif labels[i] == 'RedLeft':
labels[i] = 'R'
elif labels[i] == 'RedRight':
labels[i] = 'R'
elif labels[i] == 'Green':
labels[i] = 'G'
elif labels[i] == 'GreenLeft':
labels[i] = 'G'
elif labels[i] == 'GreenRight':
labels[i] = 'G'
else:
labels[i] = 'O'
# +
# Creating a Dictionary with dict() method
label_dict = dict({'R' : 0, 'Y' : 1, 'G' : 2, 'O' : 3})
# Train lists
x_real = []
y_real = []
# Use dict to store labels
y_real = [label_dict[i] for i in labels]
# -
# Transform them for the model
x_real = np.array(train_images)
y_real = np.array(y_real)
# +
# Train with real data
model = Sequential()
model.add(Lambda(lambda x: (x / 255.0) - 0.5, input_shape=(32,14,3)))
model.add(Conv2D(filters=8, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=16, kernel_size=(3, 3),strides=(1, 1), activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=32, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=64, kernel_size=(3, 3) ,activation='relu'))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(units=100, activation='relu'))
model.add(Dense(units=20, activation='relu'))
model.add(Dense(units=4, activation='softmax'))
print(model.summary())
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(x_real,y_real,validation_split=0.2,shuffle=True,nb_epoch=16)
model.save('model_real.h5')
# +
x_sim = []
y_sim = []
# Use dict to store labels
y_sim= [label_dict[i] for i in labels_sim]
# Transform them for the model
x_sim = np.array(sim_images_box)
y_sim = np.array(y_sim)
# -
# Evaluate the model on the test data using `evaluate`
print('\n# Evaluate on test data')
results = model.evaluate(x_sim, y_sim, batch_size=128)
print('test loss, test acc:', results)
# +
# Train with simulation data
model = Sequential()
model.add(Lambda(lambda x: (x / 255.0) - 0.5, input_shape=(32,14,3)))
model.add(Conv2D(filters=8, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=16, kernel_size=(3, 3),strides=(1, 1), activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=32, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=64, kernel_size=(3, 3) ,activation='relu'))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(units=100, activation='relu'))
model.add(Dense(units=20, activation='relu'))
model.add(Dense(units=4, activation='softmax'))
print(model.summary())
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(x_sim,y_sim,validation_split=0.2,shuffle=True,nb_epoch=16)
model.save('model_sim.h5')
# -
# Evaluate the model on the test data using `evaluate`
print('\n# Evaluate on test data')
results = model.evaluate(x_real, y_real, batch_size=128)
print('test loss, test acc:', results)
# +
# Create miscellaneous data set
x_data_set = []
y_data_set = []
x_data_set = np.concatenate((x_real, x_sim))
y_data_set = np.concatenate((y_real, y_sim))
x_data_set, y_data_set = shuffle(x_data_set, y_data_set)
x_train = []
y_train = []
x_test = []
y_test = []
# Create training and testing vars
x_train, x_test, y_train, y_test = train_test_split(x_data_set, y_data_set, test_size=0.2)
# +
# Train with miscellaneous data
model = Sequential()
model.add(Lambda(lambda x: (x / 255.0) - 0.5, input_shape=(32,14,3)))
model.add(Conv2D(filters=8, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=16, kernel_size=(3, 3),strides=(1, 1), activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=32, kernel_size=(3, 3), strides=(1, 1),activation='relu'))
model.add(Dropout(0.5))
model.add(Conv2D(filters=64, kernel_size=(3, 3) ,activation='relu'))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(units=100, activation='relu'))
model.add(Dense(units=20, activation='relu'))
model.add(Dense(units=4, activation='softmax'))
print(model.summary())
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
model.fit(x_train,y_train,validation_split=0.2,shuffle=True,nb_epoch=16)
model.save('model.h5')
# -
# Evaluate the model on the test data using `evaluate`
print('\n# Evaluate on test data')
results = model.evaluate(x_test, y_test, batch_size=128)
print('test loss, test acc:', results)
# +
# load the model we saved
model = load_model('model.h5')
classification = model.predict_classes(x_train[0].reshape(1, 32, 14, 3), batch_size=10)
print(classification)
# -
plt.figure()
plt.imshow(x_train[0])
plt.savefig("x_train0.png")
# +
# Perform traffic light detection on udacity record data
classification_images_red = []
test_images_red = []
classification_images_green = []
test_images_green = []
classification_images_yellow = []
test_images_yellow = []
classification_images_none = []
test_images_none = []
boxes_record = []
classes_record = []
scores_record = []
record_images_box = []
labels_record = []
# +
# Load record data
# Red
red_path = "data_train_record/tl_data_red" # Enter Directory of all images
data_red_record_path = os.path.join(red_path,'*g')
images_record_red = glob.glob(data_red_record_path)
images_record_red.sort()
# Green
green_path = "data_train_record/tl_data_green" # Enter Directory of all images
data_green_record_path = os.path.join(green_path,'*g')
images_record_green = glob.glob(data_green_record_path)
images_record_green.sort()
# Yellow
yellow_path = "data_train_record/tl_data_yellow" # Enter Directory of all images
data_yellow_record_path = os.path.join(yellow_path,'*g')
images_record_yellow = glob.glob(data_yellow_record_path)
images_record_yellow.sort()
#None
none_path = "data_train_record/tl_data_none" # Enter Directory of all images
data_none_record_path = os.path.join(none_path,'*g')
images_record_none = glob.glob(data_none_record_path)
images_record_none.sort()
# +
# RED CASE
# Create the list of images to be classified
for image in images_record_red:
classification_images_red.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_red:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_red.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_record_red)):
# for i in range(1):
boxes_record, scores_record, classes_record = detection(test_images_red[i])
for parameter in zip(boxes_record[0], classes_record[0], scores_record[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_red[i].shape[0])
x_max = int(box[2] * classification_images_red[i].shape[0])
y_min = int(box[1] * classification_images_red[i].shape[1])
y_max = int(box[3] * classification_images_red[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_red[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
record_image_box = cv2.resize(image,(14,32))
record_image_box = cv2.cvtColor(record_image_box, cv2.COLOR_BGR2RGB)
record_images_box.append(record_image_box)
labels_record.append('R')
except:
pass
# GREEN CASE
# Create the list of images to be classified
for image in images_record_green:
classification_images_green.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_green:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_green.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_record_green)):
# for i in range(1):
boxes_record, scores_record, classes_record = detection(test_images_green[i])
for parameter in zip(boxes_record[0], classes_record[0], scores_record[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_green[i].shape[0])
x_max = int(box[2] * classification_images_green[i].shape[0])
y_min = int(box[1] * classification_images_green[i].shape[1])
y_max = int(box[3] * classification_images_green[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_green[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
record_image_box = cv2.resize(image,(14,32))
record_image_box = cv2.cvtColor(record_image_box, cv2.COLOR_BGR2RGB)
record_images_box.append(record_image_box)
labels_record.append('G')
except:
pass
# YELLOW CASE
# Create the list of images to be classified
for image in images_record_yellow:
classification_images_yellow.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_yellow:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_yellow.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_record_yellow)):
# for i in range(1):
boxes_record, scores_record, classes_record = detection(test_images_yellow[i])
for parameter in zip(boxes_record[0], classes_record[0], scores_record[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_yellow[i].shape[0])
x_max = int(box[2] * classification_images_yellow[i].shape[0])
y_min = int(box[1] * classification_images_yellow[i].shape[1])
y_max = int(box[3] * classification_images_yellow[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_yellow[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
record_image_box = cv2.resize(image,(14,32))
record_image_box = cv2.cvtColor(record_image_box, cv2.COLOR_BGR2RGB)
record_images_box.append(record_image_box)
labels_record.append('Y')
except:
pass
# NONE CASE
# Create the list of images to be classified
for image in images_record_none:
classification_images_none.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_none:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_none.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
# for i in range (len(image_record_none)):
for i in range(1):
boxes_record, scores_record, classes_record = detection(test_images_none[i])
for parameter in zip(boxes_record[0], classes_record[0], scores_record[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_none[i].shape[0])
x_max = int(box[2] * classification_images_none[i].shape[0])
y_min = int(box[1] * classification_images_none[i].shape[1])
y_max = int(box[3] * classification_images_none[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_none[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
record_image_box = cv2.resize(image,(14,32))
record_image_box = cv2.cvtColor(record_image_box, cv2.COLOR_BGR2RGB)
record_images_box.append(record_image_box)
labels_record.append('O')
except:
pass
# +
x_record = []
y_record = []
# Use dict to store labels
y_record = [label_dict[i] for i in labels_record]
# Transform them for the model
x_record = np.array(record_images_box)
y_record = np.array(y_record)
# -
# Evaluate the model on the test data using `evaluate`
print('\n# Evaluate on record data')
results = model.evaluate(x_record, y_record, batch_size=128)
print('test loss, test acc:', results)
# +
# Perform traffic light detection on new sim data
classification_images_red = []
test_images_red = []
classification_images_green = []
test_images_green = []
classification_images_yellow = []
test_images_yellow = []
classification_images_none = []
test_images_none = []
boxes_new_sim = []
classes_new_sim = []
scores_new_sim = []
new_sim_images_box = []
labels_new_sim = []
# +
# Load new_sim data
# Red
red_path = "data_train_new_sim/red" # Enter Directory of all images
data_red_new_sim_path = os.path.join(red_path,'*g')
images_new_sim_red = glob.glob(data_red_new_sim_path)
images_new_sim_red.sort()
# Green
green_path = "data_train_new_sim/green" # Enter Directory of all images
data_green_new_sim_path = os.path.join(green_path,'*g')
images_new_sim_green = glob.glob(data_green_new_sim_path)
images_new_sim_green.sort()
# Yellow
yellow_path = "data_train_new_sim/yellow" # Enter Directory of all images
data_yellow_new_sim_path = os.path.join(yellow_path,'*g')
images_new_sim_yellow = glob.glob(data_yellow_new_sim_path)
images_new_sim_yellow.sort()
# +
# RED CASE
# Create the list of images to be classified
for image in images_new_sim_red:
classification_images_red.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_red:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_red.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_new_sim_red)):
# for i in range(1):
boxes_new_sim, scores_new_sim, classes_new_sim = detection(test_images_red[i])
for parameter in zip(boxes_new_sim[0], classes_new_sim[0], scores_new_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_red[i].shape[0])
x_max = int(box[2] * classification_images_red[i].shape[0])
y_min = int(box[1] * classification_images_red[i].shape[1])
y_max = int(box[3] * classification_images_red[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_red[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
new_sim_image_box = cv2.resize(image,(14,32))
new_sim_image_box = cv2.cvtColor(new_sim_image_box, cv2.COLOR_BGR2RGB)
new_sim_images_box.append(new_sim_image_box)
labels_new_sim.append('R')
except:
pass
# GREEN CASE
# Create the list of images to be classified
for image in images_new_sim_green:
classification_images_green.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_green:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_green.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_new_sim_green)):
# for i in range(1):
boxes_new_sim, scores_new_sim, classes_new_sim = detection(test_images_green[i])
for parameter in zip(boxes_new_sim[0], classes_new_sim[0], scores_new_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_green[i].shape[0])
x_max = int(box[2] * classification_images_green[i].shape[0])
y_min = int(box[1] * classification_images_green[i].shape[1])
y_max = int(box[3] * classification_images_green[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_green[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
new_sim_image_box = cv2.resize(image,(14,32))
new_sim_image_box = cv2.cvtColor(new_sim_image_box, cv2.COLOR_BGR2RGB)
new_sim_images_box.append(new_sim_image_box)
labels_new_sim.append('G')
except:
pass
# YELLOW CASE
# Create the list of images to be classified
for image in images_new_sim_yellow:
classification_images_yellow.append(cv2.imread(image))
# Squeeze them in 1-D and convert them for the model
for img in classification_images_yellow:
img = cv2.cvtColor(img, cv2.COLOR_RGB2BGR)
test_images_yellow.append(np.expand_dims(img, axis=0))
# Cut the image looking at boxes detected
for i in range (len(images_new_sim_yellow)):
# for i in range(1):
boxes_new_sim, scores_new_sim, classes_new_sim = detection(test_images_yellow[i])
for parameter in zip(boxes_new_sim[0], classes_new_sim[0], scores_new_sim[0]):
if parameter[1] == 10 and parameter[2] >= .5:
box = parameter[0]
x_min = int(box[0] * classification_images_yellow[i].shape[0])
x_max = int(box[2] * classification_images_yellow[i].shape[0])
y_min = int(box[1] * classification_images_yellow[i].shape[1])
y_max = int(box[3] * classification_images_yellow[i].shape[1])
if (x_max - x_min) > 20:
image = classification_images_yellow[i][x_min:x_max,y_min:y_max,:]
if image.shape[0] != 0:
try:
new_sim_image_box = cv2.resize(image,(14,32))
new_sim_image_box = cv2.cvtColor(new_sim_image_box, cv2.COLOR_BGR2RGB)
new_sim_images_box.append(new_sim_image_box)
labels_new_sim.append('Y')
except:
pass
# +
x_new_sim = []
y_new_sim = []
# Use dict to store labels
y_new_sim = [label_dict[i] for i in labels_new_sim]
# Transform them for the model
x_new_sim = np.array(new_sim_images_box)
y_new_sim = np.array(y_new_sim)
# -
i = 143
plt.figure()
plt.imshow(x_new_sim[i])
print(y_new_sim[i])
# +
# load the model we saved
model = load_model('model.h5')
classification = model.predict_classes(x_new_sim[i].reshape(1, 32, 14, 3), batch_size=10)
print(classification)
# -
| ros/src/tl_detector/light_classification/Traffic_light_detector/Traffic_Light_detector.ipynb |
# ---
# jupyter:
# jupytext:
# formats: md,ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# In this exercise, you'll work on building some intuition around feature extraction. First, we'll walk through the example we did in the tutorial again, but this time, with a kernel you choose yourself. We've mostly been working with images in this course, but what's behind all of the operations we're learning about is mathematics. So, we'll also take a look at how these feature maps can be represented instead as arrays of numbers and what effect convolution with a kernel will have on them.
#
# Run the cell below to get started!
# Setup feedback system
from learntools.core import binder
binder.bind(globals())
from learntools.computer_vision.ex2 import *
# # Apply Transformations #
#
# The next few exercises walk through feature extraction just like the example in the tutorial. Run the following cell to load an image we'll use for the next few exercises.
# +
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
plt.rc('figure', autolayout=True)
plt.rc('axes', labelweight='bold', labelsize='large',
titleweight='bold', titlesize=18, titlepad=10)
plt.rc('image', cmap='magma')
image_path = '../input/computer-vision-resources/car_illus.jpg'
image = tf.io.read_file(image_path)
image = tf.io.decode_jpeg(image, channels=1)
image = tf.image.resize(image, size=[400, 400])
plt.figure(figsize=(6, 6))
plt.imshow(tf.squeeze(image), cmap='gray')
plt.axis('off')
plt.show();
# -
# You can run this cell to see some standard kernels used in image processing.
# +
import learntools.computer_vision.visiontools as visiontools
from learntools.computer_vision.visiontools import edge, bottom_sobel, emboss, sharpen
kernels = [edge, bottom_sobel, emboss, sharpen]
names = ["Edge Detect", "Bottom Sobel", "Emboss", "Sharpen"]
plt.figure(figsize=(12, 12))
for i, (kernel, name) in enumerate(zip(kernels, names)):
plt.subplot(1, 4, i+1)
visiontools.show_kernel(kernel)
plt.title(name)
plt.tight_layout()
# -
# # 1) Define Kernel #
#
# Use the next code cell to define a kernel. You have your choice of what kind of kernel to apply. One thing to keep in mind is that the *sum* of the numbers in the kernel determines how bright the final image is. Generally, you should try to keep the sum of the numbers between 0 and 1 (though that's not required for a correct answer).
#
# In general, a kernel can have any number of rows and columns. For this exercise, let's use a $3 \times 3$ kernel, which often gives the best results. Define a kernel with `tf.constant`.
# +
# YOUR CODE HERE: Define a kernel with 3 rows and 3 columns.
kernel = tf.constant([
#____,
])
# Uncomment to view kernel
# visiontools.show_kernel(kernel)
# Check your answer
q_1.check()
# -
# #%%RM_IF(PROD)%%
kernel = np.array([
[-2, -1, 0],
[-1, 1, 1],
[0, 1, 2],
])
q_1.assert_check_failed()
# #%%RM_IF(PROD)%%
kernel = tf.constant([
'abc'
])
q_1.assert_check_failed()
# #%%RM_IF(PROD)%%
kernel = tf.constant([0, 1, 2])
q_1.assert_check_failed()
# #%%RM_IF(PROD)%%
kernel = tf.constant([
[-2, -1, 0],
[-1, 1, 1],
[0, 1, 2],
])
visiontools.show_kernel(kernel)
q_1.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
q_1.hint()
#_COMMENT_IF(PROD)_
q_1.solution()
# Now we'll do the first step of feature extraction, the filtering step. First run this cell to do some reformatting for TensorFlow.
# Reformat for batch compatibility.
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
image = tf.expand_dims(image, axis=0)
kernel = tf.reshape(kernel, [*kernel.shape, 1, 1])
kernel = tf.cast(kernel, dtype=tf.float32)
# # 2) Apply Convolution #
#
# Now we'll apply the kernel to the image by a convolution. The *layer* in Keras that does this is `layers.Conv2D`. What is the *backend function* in TensorFlow that performs the same operation?
# +
# YOUR CODE HERE: Give the TensorFlow convolution function (without arguments)
conv_fn = ____
# Check your answer
q_2.check()
# -
# #%%RM_IF(PROD)%%
conv_fn = 'abc'
q_2.assert_check_failed()
# #%%RM_IF(PROD)%%
conv_fn = tf.nn.conv2d(
input=image,
filters=kernel,
strides=1, # or (1, 1)
padding='SAME',
)
q_2.assert_check_failed()
# #%%RM_IF(PROD)%%
conv_fn = tf.nn.conv2d
q_2.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
q_2.hint()
#_COMMENT_IF(PROD)_
q_2.solution()
# Once you've got the correct answer, run this next cell to execute the convolution and see the result!
# +
image_filter = conv_fn(
input=image,
filters=kernel,
strides=1, # or (1, 1)
padding='SAME',
)
plt.imshow(
# Reformat for plotting
tf.squeeze(image_filter)
)
plt.axis('off')
plt.show();
# -
# Can you see how the kernel you chose relates to the feature map it produced?
#
# # 3) Apply ReLU #
#
# Now detect the feature with the ReLU function. In Keras, you'll usually use this as the activation function in a `Conv2D` layer. What is the *backend function* in TensorFlow that does the same thing?
# +
# YOUR CODE HERE: Give the TensorFlow ReLU function (without arguments)
relu_fn = ____
# Check your answer
q_3.check()
# -
# #%%RM_IF(PROD)%%
relu_fn = 'abc'
q_3.assert_check_failed()
# #%%RM_IF(PROD)%%
relu_fn = tf.nn.relu(image_filter)
q_3.assert_check_failed()
# #%%RM_IF(PROD)%%
relu_fn = tf.nn.relu
q_3.assert_check_passed()
# Lines below will give you a hint or solution code
#_COMMENT_IF(PROD)_
q_3.hint()
#_COMMENT_IF(PROD)_
q_3.solution()
# Once you've got the solution, run this cell to detect the feature with ReLU and see the result!
#
# The image you see below is the feature map produced by the kernel you chose. If you like, experiment with some of the other suggested kernels above, or, try to invent one that will extract a certain kind of feature.
#
# +
image_detect = relu_fn(image_filter)
plt.imshow(
# Reformat for plotting
tf.squeeze(image_detect)
)
plt.axis('off')
plt.show();
# -
# In the tutorial, our discussion of kernels and feature maps was mainly visual. We saw the effect of `Conv2D` and `ReLU` by observing how they transformed some example images.
#
# But the operations in a convolutional network (like in all neural networks) are usually defined through mathematical functions, through a computation on numbers. In the next exercise, we'll take a moment to explore this point of view.
#
# Let's start by defining a simple array to act as an image, and another array to act as the kernel. Run the following cell to see these arrays.
# +
# Sympy is a python library for symbolic mathematics. It has a nice
# pretty printer for matrices, which is all we'll use it for.
import sympy
sympy.init_printing()
from IPython.display import display
image = np.array([
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 1, 1, 1],
[0, 1, 0, 0, 0, 0],
])
kernel = np.array([
[1, -1],
[1, -1],
])
display(sympy.Matrix(image))
display(sympy.Matrix(kernel))
# Reformat for Tensorflow
image = tf.cast(image, dtype=tf.float32)
image = tf.reshape(image, [1, *image.shape, 1])
kernel = tf.reshape(kernel, [*kernel.shape, 1, 1])
kernel = tf.cast(kernel, dtype=tf.float32)
# -
# # 4) Observe Convolution on a Numerical Matrix #
#
#
# What do you see? The image is simply a long vertical line on the left and a short horizontal line on the lower right. What about the kernel? What effect do you think it will have on this image? After you've thought about it, run the next cell for the answer.
# View the solution (Run this code cell to receive credit!)
q_4.check()
# Now let's try it out. Run the next cell to apply convolution and ReLU to the image and display the result.
# +
image_filter = tf.nn.conv2d(
input=image,
filters=kernel,
strides=1,
padding='VALID',
)
image_detect = tf.nn.relu(image_filter)
# The first matrix is the image after convolution, and the second is
# the image after ReLU.
display(sympy.Matrix(tf.squeeze(image_filter).numpy()))
display(sympy.Matrix(tf.squeeze(image_detect).numpy()))
# -
# Is the result what you expected?
#
# # Conclusion #
#
# In this lesson, you learned about the first two operations a convolutional classifier uses for feature extraction: **filtering** an image with a **convolution** and **detecting** the feature with the **rectified linear unit**.
#
# # Keep Going #
#
# Move on to [**Lesson 3**](#$NEXT_NOTEBOOK_URL$) to learn the final operation: **condensing** the feature map with **maximum pooling**!
| notebooks/computer_vision/raw/ex2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cryptolytic
# language: python
# name: cryptolytic
# ---
# # Imports
# !pip install scikit-learn==0.21.3
# !pip install ta==0.4.7
# !pip install pandas==0.25.1
# +
import glob
import os
import pickle
import json
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import datetime as dt
from ta import add_all_ta_features
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
# -
# #### Background on Arbitrage Models
# Arbitrage models were created with the goal of predicting arbitrage 10 min before it happens in an active crypto market. The models are generated by getting all of the combinations of 2 exchanges that support the same trading pair, engineering technical analysis features, merging that data on 'closing_time', engineering more features, and creating a target that signals an arbitrage opportunity. A valid arbitrage signal is when the arbitrage lasts >30 mins because it takes time to move coins from one exchange to the other in order to successfully complete the arbitrage trades.
#
# The models predict whether there will be an arbitrage opportunity that starts 10 mins after the prediction time and lasts for at least 30 mins, giving a user enough times to execute trades.
#
# More than 6000+ iterations of models were generated in this notebook and the best ones were selected from each possible arbitrage combination based on model selection criteria outlined later in this section. The models were Random Forest Classifier and the best model parameters varied for each dataset. The data was obtained from the respective exchanges via their api, and we did a 70/30 train/test split on 5 min candlestick data that fell anywhere in the range from Jun 2015 - Oct 2019. There was a 2 week gap left between the train and test sets to prevent data leakage. The models return 0 (no arbitrage), 1 (arbitrage from exchange 1 to exchange 2) and -1 (arbitrage from exchange 2 to exchange 1).
#
# The profit calculation incorporated fees like in the real world. We used mean percent profit as the profitability metric which represented the average percent profit per arbitrage trade if one were to act on all trades predicted by the model in the testing period, whether those predictions were correct or not.
#
# From the 6000+ iterations of models trained, the best models were narrowed down based on the following criteria:
# - How often the models predicted arbitrage when it didn't exist (False positives)
# - How many times the models predicted arbitrage correctly (True positives)
# - How profitable the model was in the real world over the period of the test set.
#
# There were 21 models that met the thresholds for model selection critera (details of these models can be found at the end of this nb). The final models were all profitable with gains anywhere from 0.2% - 2.3% within the varied testing time periods (Note: the model with >9% mean percent profit was an outlier). Visualizations for how these models performed can be viewed at https://github.com/Lambda-School-Labs/cryptolytic-ds/blob/master/finalized_notebooks/visualization/arb_performance_visualization.ipynb
#
# \* It is HIGHLY recommended to run this on sagemaker and split the training work onto 4 notebooks. These functions will take over a day to run if not split up. There are 95 total options for models, 75 of those options have enough data to train models, and with different options for parameters around ~7K models will be trained. After selecting for the best models, there were 21 good ones that were included in this project.
#
# ** Feature engineering takes a LONG time. We export data as csvs along each step to not have to re-engineer features everytime the runtime restarts: export after technical analysis features are added, and export after datasets are merged. When iterating on this work, you should first settle on any new features you want to add, create those datasets, and then move onto modeling. Try not to go back and forth with feature engineering and modeling, it will be too time consuming and inefficient.
#
# \*** There has been some feature selection done in this process where we removed highly correlated features, but not enough. There should be more exploration into whether removing features improves accuracy.
#
# \**** We haven't tried normalizing the dataset to see if it will improve accuracy, but that should be a top priority to anyone continuing this project
# ##### Folder organization:
# ├── arbitrage/ <-- The top-level directory for all arbitrage work
# │ ├── arbitrage_models.ipynb <-- notebook for arbitrage models
# │ ├── all_data/ <-- Directory with subdirectories containing 5 min candle data
# │ │ ├──bitfinex_300/
# │ │ │ └── data.csv
# │ │ ├──coinbase_pro_300/
# │ │ │ └── data.csv
# │ │ ├──gemini_300/
# │ │ │ └── data.csv
# │ │ ├──hitbtc_300/
# │ │ │ └── data.csv
# │ │ └──kraken_300/
# │ │ └── data.csv
# │ ├── data/ <-- Directory for csv files of all 5 min candle data
# │ │ └── data.csv
# │ ├── ta_data/ <-- Directory for csv files of data after ta features engineered
# │ │ └── data.csv
# │ ├── arb_data/ <-- Directory for csv files of final arbitrage training data
# │ │ └── data.csv
# │ ├── pickles/ <-- Directory for all pickle models
# │ │ └── models.pkl
# │ ├── arbitrage_pickles <-- Directory for final models after model selection
# │ │ └── models.pkl
# │ │
# │ ├── cm/ <-- Directory for confusion matrices after training models
# │ │
# │ ├── model_perf/ <-- Directory for performance csvs after training models
#
# Note: The folders /all_data and /data technically have the same exact data, the only difference is that one is structured. It doesn't make sense that there are two folders with the same exact data, but a function was written to get the combinations for arbitrage with the subdirectories so it's necessary for now until the function is rewritten ¯\\\_(ツ)_/¯
# # Data
# Get all of the 5 min candle data filepaths into a variable
csv_filepaths = glob.glob('data/*.csv')
len(csv_filepaths) #80
# # Functions
# #### Function to get all combinations of exchanges with the same trading pair
#
# Follow instructions on folder organization above in order for this to run correctly.
# +
# five supported exchanges
exchanges = ['bitfinex', 'coinbase_pro', 'gemini', 'hitbtc', 'kraken']
# function to create pairs for arbitrage datasets
def get_file_pairs(exchanges):
"""This function takes in a list of exchanges and looks through data
directories to find all possible combinations for 2 exchanges
with the same trading pair. Returns a list of all lists that
include the file pairs"""
# list for filenames of ohlcv csvs
filenames = []
for directory in os.listdir('all_data'):
# .DS_Store files can mess things up, since they aren't directories
if directory != '.DS_Store':
# for each of the files in the subdirectory
for filename in os.listdir('all_data/' + directory):
# add to list of filenames if the file is a csv
if filename.endswith('300.csv'):
filenames.append(filename)
# list for pairs of csvs
file_pairs = []
# compare filenames to eachother and append them in a list
for filename_1 in filenames:
# filenames we haven't looped through yet
remaining_filenames = filenames[filenames.index(filename_1)+1:]
# iterate through remaining filenames
for filename_2 in remaining_filenames:
# iterate through exchanges
for exchange in exchanges:
# drop the exchange from the first filename and see if the
# remaining string is contained in the second filename
if filename_1.replace(exchange, '') in filename_2:
# add the pair of filenames to the list of pairs
file_pairs.append([filename_1, filename_2])
return file_pairs
# -
pairs = get_file_pairs(exchanges)
print(len(pairs)) # 95
pairs
# #### OHLCV Data Resampling
def resample_ohlcv(df, period):
""" Changes the time period on cryptocurrency ohlcv data.
Period is a string denoted by '{time_in_minutes}T'(ex: '1T', '5T', '60T')."""
# Set date as the index. This is needed for the function to run
df = df.set_index(['date'])
# Aggregation function
ohlc_dict = {'open':'first',
'high':'max',
'low':'min',
'close': 'last',
'base_volume': 'sum'}
# Apply resampling
df = df.resample(period, how=ohlc_dict, closed='left', label='left')
return df
# #### Filling NaNs
# +
# resample_ohlcv function will create NaNs in df where there were gaps in the data.
# The gaps could be caused by exchanges being down, errors from cryptowatch or the
# exchanges themselves
def fill_nan(df):
"""Iterates through a dataframe and fills NaNs with appropriate
open, high, low, close values."""
# Forward fill close column.
df['close'] = df['close'].ffill()
# Backward fill the open, high, low rows with the close value.
df = df.bfill(axis=1)
return df
# -
# #### Feature engineering - before merge
def engineer_features(df, period='5T'):
"""Takes a df, engineers ta features, and returns a df
default period=['5T']"""
# convert unix closing_time to datetime
df['date'] = pd.to_datetime(df['closing_time'], unit='s')
# time resampling to fill gaps in data
df = resample_ohlcv(df, period)
# move date off the index
df = df.reset_index()
# create closing_time
closing_time = df.date.values
df.drop(columns='date', inplace=True)
# create feature to indicate where rows were gaps in data
df['nan_ohlcv'] = df['close'].apply(lambda x: 1 if pd.isnull(x) else 0)
# fill gaps in data
df = fill_nan(df)
# adding all the technical analysis features...
df = add_all_ta_features(df, 'open', 'high', 'low', 'close','base_volume', fillna=True)
# add closing time column
df['closing_time'] = closing_time
return df
# #### Feature Engineering - after merge
# +
def get_higher_closing_price(df):
"""returns the exchange with the higher closing price"""
# exchange 1 has higher closing price
if (df['close_exchange_1'] - df['close_exchange_2']) > 0:
return 1
# exchange 2 has higher closing price
elif (df['close_exchange_1'] - df['close_exchange_2']) < 0:
return 2
# closing prices are equivalent
else:
return 0
def get_pct_higher(df):
"""returns the percentage of the difference between ex1/ex2
closing prices"""
# if exchange 1 has a higher closing price than exchange 2
if df['higher_closing_price'] == 1:
# % difference
return ((df['close_exchange_1'] /
df['close_exchange_2'])-1)*100
# if exchange 2 has a higher closing price than exchange 1
elif df['higher_closing_price'] == 2:
# % difference
return ((df['close_exchange_2'] /
df['close_exchange_1'])-1)*100
# if closing prices are equivalent
else:
return 0
def get_arbitrage_opportunity(df):
"""function to create column showing available arbitrage opportunities"""
# assuming the total fees are 0.55%, if the higher closing price is less
# than 0.55% higher than the lower closing price...
if df['pct_higher'] < .55:
return 0 # no arbitrage
# if exchange 1 closing price is more than 0.55% higher
# than the exchange 2 closing price
elif df['higher_closing_price'] == 1:
return -1 # arbitrage from exchange 2 to exchange 1
# if exchange 2 closing price is more than 0.55% higher
# than the exchange 1 closing price
elif df['higher_closing_price'] == 2:
return 1 # arbitrage from exchange 1 to exchange 2
def get_window_length(df):
"""function to create column showing how long arbitrage opportunity has lasted"""
# convert arbitrage_opportunity column to a list
target_list = df['arbitrage_opportunity'].to_list()
# set initial window length
window_length = 5 # time in minutes
# list for window_lengths
window_lengths = []
# iterate through arbitrage_opportunity column
for i in range(len(target_list)):
# check if a value in the arbitrage_opportunity column is equal to the
# previous value in the arbitrage_opportunity column and increase
# window length
if target_list[i] == target_list[i-1]:
window_length += 5
window_lengths.append(window_length)
# if a value in the arbitrage_opportunity column is
# not equal to the previous value in the arbitrage_opportunity column
# reset the window length to five minutes
else:
window_length = 5
window_lengths.append(window_length)
# create window length column showing how long an arbitrage opportunity has lasted
df['window_length'] = window_lengths
return df
def merge_dfs(df1, df2):
"""function to merge dataframes and create final features for arbitrage data"""
# merging two modified ohlcv dfs on closing time to create arbitrage df
df = pd.merge(df1, df2, on='closing_time',
suffixes=('_exchange_1', '_exchange_2'))
# convert closing_time to datetime
df['closing_time'] = pd.to_datetime(df['closing_time'])
# Create additional date features.
df['year'] = df['closing_time'].dt.year
df['month'] = df['closing_time'].dt.month
df['day'] = df['closing_time'].dt.day
# get higher_closing_price feature to create pct_higher feature
df['higher_closing_price'] = df.apply(get_higher_closing_price, axis=1)
# get pct_higher feature to create arbitrage_opportunity feature
df['pct_higher'] = df.apply(get_pct_higher, axis=1)
# create arbitrage_opportunity feature
df['arbitrage_opportunity'] = df.apply(get_arbitrage_opportunity, axis=1)
# create window_length feature
df = get_window_length(df)
return df
# -
# #### Creating the target
# +
# specifying arbitrage window length to target, in minutes
interval = 30
def get_target_value(df, interval=30):
"""function to get target values; takes df and window length to target"""
# if the coming arbitrage window is as long as the targeted interval
if df['window_length_shift'] >= interval:
# if that window is for exchange 1 to 2
if df['arbitrage_opportunity_shift'] == 1:
return 1 # arbitrage from exchange 1 to 2
# if that window is for exchange 2 to 1
elif df['arbitrage_opportunity_shift'] == -1:
return -1 # arbitrage from exchange 2 to 1
# if no arbitrage opportunity
elif df['arbitrage_opportunity_shift'] == 0:
return 0 # no arbitrage opportunity
# if the coming window is less than our targeted interval
else:
return 0 # no arbitrage opportunity
def get_target(df, interval=interval):
"""function to create target column"""
# used to shift rows
# assumes candle length is five minutes, interval is 30 mins
rows_to_shift = int(-1*(interval/5)) # -7
# arbitrage_opportunity feature, shifted by length of targeted interval
# minus one to predict ten minutes in advance rather than five
df['arbitrage_opportunity_shift'] = df['arbitrage_opportunity'].shift(
rows_to_shift - 1)
# window_length feature, shifted by length of targeted interval minus one
# to predict ten minutes
df['window_length_shift'] = df['window_length'].shift(rows_to_shift - 1)
# creating target column; this will indicate if an arbitrage opportunity
# that lasts as long as the targeted interval is forthcoming
df['target'] = df.apply(get_target_value, axis=1)
# dropping rows where target could not be calculated due to shift
df = df[:rows_to_shift - 1] # -7
return df
def get_close_shift(df, interval=interval):
rows_to_shift = int(-1*(interval/5))
df['close_exchange_1_shift'] = df['close_exchange_1'].shift(
rows_to_shift - 2)
df['close_exchange_2_shift'] = df['close_exchange_2'].shift(
rows_to_shift - 2)
return df
# function to create profit feature
def get_profit(df):
"""function to create profit feature"""
# if exchange 1 has the higher closing price
if df['higher_closing_price'] == 1:
# return how much money you would make if you bought on exchange 2, sold
# on exchange 1, and took account of 0.55% fees
return (((df['close_exchange_1_shift'] /
df['close_exchange_2'])-1)*100)-.55
# if exchange 2 has the higher closing price
elif df['higher_closing_price'] == 2:
# return how much money you would make if you bought on exchange 1, sold
# on exchange 2, and took account of 0.55% fees
return (((df['close_exchange_2_shift'] /
df['close_exchange_1'])-1)*100)-.55
# if the closing prices are the same
else:
return 0 # no arbitrage
# -
# #### Split names when in the format exchange_trading_pair
# coinbase_pro has an extra underscore so we need a function to split it differently
def get_exchange_trading_pair(ex_tp):
# coinbase_pro
if len(ex_tp.split('_')) == 4:
exchange = ex_tp.split('_')[0] + '_' + ex_tp.split('_')[1]
trading_pair = ex_tp.split('_')[2] + '_' + ex_tp.split('_')[3]
# all other exchanges
else:
exchange = ex_tp.split('_')[0]
trading_pair = ex_tp.split('_')[1] + '_' + ex_tp.split('_')[2]
return exchange, trading_pair
# ### Generate all individual csv's with ta data (~1-2 hours)
# create /ta_data directory before running this function
def create_ta_csvs(csv_filepaths):
"""Takes a csv filename, creates a dataframe, engineers features,
and saves it as a new csv in /ta_data."""
# counter
n = 1
for file in csv_filepaths:
# create df
df = pd.read_csv(file, index_col=0)
# define period
period = '5T'
# engineer features
df = engineer_features(df, period)
print('features engineered')
# generate new filename
filename = 'ta_data/' + file.split('/')[1][:-4] + '_ta.csv'
# export csv
df.to_csv(filename)
# print progress
print(f'csv #{n} saved :)')
# update counter
n += 1
create_ta_csvs(csv_filepaths)
# ### Generate all arbitrage training data csv's (~9 hrs)
# Notes:
# - create a /arb_data directory before running this function
# - this function takes a really long time to run so it's recommended to run in sagemaker and divide the pairs in to 4 notebooks so you're running about 20 pairs in each notebook. Should take ~2-3 hours if split up on 4 notebooks.
def create_arb_csvs(pairs):
"""Takes a list of possible arbitrage combinations, finds the
appropriate datasets in /ta_data, loads datasets, merges them,
engineers more features, creates a target and exports the new
dataset as a csv"""
# counter
n = 0
# iterate through arbitrage combinations
for pair in pairs:
# define paths for the csv
csv_1, csv_2 = 'ta_data/' + pair[0][:-4] + '_ta.csv', 'ta_data/' + pair[1][:-4] + '_ta.csv'
# define exchanges and trading_pairs
ex_tp_1, ex_tp_2 = pair[0][:-8], pair[1][:-8]
exchange_1, trading_pair_1 = get_exchange_trading_pair(ex_tp_1)
exchange_2, trading_pair_2 = get_exchange_trading_pair(ex_tp_2)
print(exchange_1, trading_pair_1, exchange_2, trading_pair_2)
# define model_name for the filename
model_name = exchange_1 + '_' + ex_tp_2
print(model_name)
# create dfs from csv's that already include ta features
df1, df2 = pd.read_csv(csv_1, index_col=0), pd.read_csv(csv_2, index_col=0)
print('df 1 shape: ', df1.shape, 'df 2 shape: ', df2.shape)
# merge dfs
df = merge_dfs(df1, df2)
print('dfs merged')
print('merged df shape:' , df.shape)
# create target
df = get_target(df)
print(model_name, ' ', df.shape)
# export csv
path = 'arb_data/'
csv_filename = path + model_name + '.csv'
df.to_csv(csv_filename)
# print progress
print(f'csv #{n} saved :)')
# update counter
n += 1
create_arb_csvs(pairs)
# #### Get arbitrage data csvs into a variable
arb_data_paths = glob.glob('arb_data/*.csv')
print(len(arb_data_paths))
# ## Train models
# ##### Notes:
# - create /pickles and /arbitrage_pickles directories before running this function
# - test that this function will run to completion before running fully using just one option for each model parameter and one dataset
# - this function takes a really long time to run so it's recommended to run in sagemaker and divide the pairs in to 4 notebooks so you're running about 20 pairs in each notebook. Should take ~4 hours if split up on 4 notebooks.
# - this function was written to pick up where it left off in case something goes wrong. It will first check if a specific model exists in the /pickles directory and if not, it will then train the model. If the model does exist, it will just get the performance stats
# +
def create_models(arb_data_paths):
"""This function takes in a list of all the arbitrage data paths,
does train/test split, feature selection, trains models,
saves the pickle file, gets performance stats for the model,
and returns a dataframe of performance stats and a dictionary
of confusion matrices for each model"""
counter = 0
line = '---------------'
performance_list = []
confusion_dict = {}
# this is in case the function stops running you can pick up where you left off
# get all model paths into a variable
model_paths = glob.glob('pickles/*.pkl')
# iterate through the arbitrage csvs
for file in arb_data_paths:
# define model name
name = file.split('/')[1][:-8]
# read csv
df = pd.read_csv(file, index_col=0)
# convert str closing_time to datetime
df['closing_time'] = pd.to_datetime(df['closing_time'])
print('\n' + line*5 + '\n' + line*2 + name.upper() + line*2 + '\n' + line*5)
# 70/30 train/test split
test_train_split_row = round(len(df)*.7)
# get closing_time for t/t split
test_train_split_time = df['closing_time'][test_train_split_row]
# remove 1 week from each end of the t/t datasets to create a
# two week gap between the data - prevents data leakage
train_cutoff_time = test_train_split_time - dt.timedelta(days=7)
test_cutoff_time = test_train_split_time + dt.timedelta(days=7)
print('cutoff time:', train_cutoff_time, test_cutoff_time)
# train and test subsets
train = df[df['closing_time'] < train_cutoff_time]
test = df[df['closing_time'] > test_cutoff_time]
# printing shapes to track progress
print('train and test shape: ', train.shape, test.shape)
# pick features
# not using open, high, or low, which are highly correlated with close
# and do not improve model performance
features = ['close_exchange_1','base_volume_exchange_1',
'nan_ohlcv_exchange_1','volume_adi_exchange_1', 'volume_obv_exchange_1',
'volume_cmf_exchange_1', 'volume_fi_exchange_1','volume_em_exchange_1',
'volume_vpt_exchange_1','volume_nvi_exchange_1', 'volatility_atr_exchange_1',
'volatility_bbhi_exchange_1','volatility_bbli_exchange_1',
'volatility_kchi_exchange_1', 'volatility_kcli_exchange_1',
'volatility_dchi_exchange_1','volatility_dcli_exchange_1',
'trend_macd_signal_exchange_1', 'trend_macd_diff_exchange_1',
'trend_adx_exchange_1', 'trend_adx_pos_exchange_1',
'trend_adx_neg_exchange_1', 'trend_vortex_ind_pos_exchange_1',
'trend_vortex_ind_neg_exchange_1', 'trend_vortex_diff_exchange_1',
'trend_trix_exchange_1', 'trend_mass_index_exchange_1',
'trend_cci_exchange_1', 'trend_dpo_exchange_1', 'trend_kst_sig_exchange_1',
'trend_kst_diff_exchange_1', 'trend_aroon_up_exchange_1',
'trend_aroon_down_exchange_1', 'trend_aroon_ind_exchange_1',
'momentum_rsi_exchange_1', 'momentum_mfi_exchange_1',
'momentum_tsi_exchange_1', 'momentum_uo_exchange_1',
'momentum_stoch_signal_exchange_1', 'momentum_wr_exchange_1',
'momentum_ao_exchange_1', 'others_dr_exchange_1', 'close_exchange_2',
'base_volume_exchange_2', 'nan_ohlcv_exchange_2',
'volume_adi_exchange_2', 'volume_obv_exchange_2',
'volume_cmf_exchange_2', 'volume_fi_exchange_2',
'volume_em_exchange_2', 'volume_vpt_exchange_2',
'volume_nvi_exchange_2', 'volatility_atr_exchange_2',
'volatility_bbhi_exchange_2', 'volatility_bbli_exchange_2',
'volatility_kchi_exchange_2', 'volatility_kcli_exchange_2',
'volatility_dchi_exchange_2', 'volatility_dcli_exchange_2',
'trend_macd_signal_exchange_2',
'trend_macd_diff_exchange_2', 'trend_adx_exchange_2',
'trend_adx_pos_exchange_2', 'trend_adx_neg_exchange_2',
'trend_vortex_ind_pos_exchange_2',
'trend_vortex_ind_neg_exchange_2',
'trend_vortex_diff_exchange_2', 'trend_trix_exchange_2',
'trend_mass_index_exchange_2', 'trend_cci_exchange_2',
'trend_dpo_exchange_2', 'trend_kst_sig_exchange_2',
'trend_kst_diff_exchange_2', 'trend_aroon_up_exchange_2',
'trend_aroon_down_exchange_2',
'trend_aroon_ind_exchange_2',
'momentum_rsi_exchange_2', 'momentum_mfi_exchange_2',
'momentum_tsi_exchange_2', 'momentum_uo_exchange_2',
'momentum_stoch_signal_exchange_2',
'momentum_wr_exchange_2', 'momentum_ao_exchange_2',
'others_dr_exchange_2', 'year', 'month', 'day',
'higher_closing_price', 'pct_higher',
'arbitrage_opportunity', 'window_length']
# pick target
target = 'target'
# X, y matrix
X_train = train[features]
X_test = test[features]
y_train = train[target]
y_test = test[target]
print('train test shapes:', X_train.shape, X_test.shape, y_train.shape, y_test.shape)
# filter out datasets that are too small
if (X_train.shape[0] > 1000) and (X_test.shape[0] > 100):
# max_depth_list = [14] # just for testing to see if function completes
max_depth_list = [14, 15, 17, 18, 21, 25]
for max_depth in max_depth_list:
# max_features_list = [50] # just for testing to see if function completes
max_features_list = [50, 55, 60, 65, 70, 75, 80]
for max_features in max_features_list:
# n_estimator_list = [100] # just for testing to see if function completes
n_estimator_list = [100, 150]
for n_estimators in n_estimator_list:
# define model
model_name = name + '_' + str(max_features) + '_' + str(max_depth) + '_' + str(n_estimators)
print(line + model_name + line)
# define model filename to check if it exists
model_path = f'pickles/{model_name}.pkl'
# if the model does not exist
if model_path not in model_paths:
# instantiate model
model = RandomForestClassifier(max_features=max_features,
max_depth=max_depth,
n_estimators=n_estimators,
n_jobs=-1,
random_state=42)
# there was a weird error caused by two of the datasets which
# is why this try/except is needed to keep the function running
try:
# fit model
model = model.fit(X_train, y_train)
print('model fitted!')
# train accuracy
train_score = model.score(X_train, y_train)
print('train accuracy:', train_score)
# make predictions
y_preds = model.predict(X_test)
print('predictions made!')
# test accuracy
score = accuracy_score(y_test, y_preds)
print('test accuracy:', score)
# save model
pickle.dump(model, open('pickles/{model_name}.pkl'.format(
model_name=model_name), 'wb'))
print('pickle saved!'.format(model_name=model_name))
except:
print(line*3 + '\n' + line + 'ERROR' + line + '\n' + line*3)
break # break out of for loop if there is an error with modeling
# if the model exists
else:
# load model
model = pickle.load(open(model_path, 'rb'))
print('model loaded')
# train accuracy
train_score = model.score(X_train, y_train)
print('train accuracy:', train_score)
# make predictions
y_preds = model.predict(X_test)
print('predictions made!')
# test accuracy
score = accuracy_score(y_test, y_preds)
print('test accuracy:', score)
############## Performance metrics ###############
# TODO: put this all in a function and just return the
# metrics we want
# labels for confusion matrix
unique_y_test = y_test.unique().tolist()
unique_y_preds = list(set(y_preds))
labels = list(set(unique_y_test + unique_y_preds))
labels.sort()
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
# create confusion matrix
confusion = pd.DataFrame(confusion_matrix(y_test, y_preds),
columns=columns, index=index)
print(model_name + ' confusion matrix:')
print(confusion, '\n')
# append to confusion list
confusion_dict[model_name] = confusion
# creating dataframe from test set to calculate profitability
test_with_preds = X_test.copy()
# add column with higher closing price
test_with_preds['higher_closing_price'] = test_with_preds.apply(
get_higher_closing_price, axis=1)
# add column with shifted closing price
test_with_preds = get_close_shift(test_with_preds)
# adding column with predictions
test_with_preds['pred'] = y_preds
# adding column with profitability of predictions
test_with_preds['pct_profit'] = test_with_preds.apply(
get_profit, axis=1).shift(-2)
# filtering out rows where no arbitrage is predicted
test_with_preds = test_with_preds[test_with_preds['pred'] != 0]
# calculating mean profit where arbitrage predicted...
pct_profit_mean = test_with_preds['pct_profit'].mean()
# calculating median profit where arbitrage predicted...
pct_profit_median = test_with_preds['pct_profit'].median()
print('percent profit mean:', pct_profit_mean)
print('percent profit median:', pct_profit_median, '\n\n')
# save net performance to list
performance_list.append([name, max_features, max_depth, n_estimators,
pct_profit_mean, pct_profit_median])
######################## END OF TODO ###########################
# if there is not enough data
else:
print('{model_name}: not enough data!'.format(model_name=name))
# update count
# TODO: make a better counter that is actually useful in
# showing how much is left
counter += 1
print(counter, '\n')
# create a dataframe for performace of all models
df = pd.DataFrame(performance_list, columns = ['ex_tp', 'max_features', 'max_depth',
'n_estimators', 'pct_profit_mean','pct_profit_median'])
return df, confusion_dict
# -
df, confusion_dict = create_models(arb_data_paths)
# #### Export model performance data into csvs and JSON
# You will need /model_perf and /cm directories to store the performance csv's and JSON if you split up running the models on several notebooks. You will also have to change the name of each of the files being exported in the other notebooks otherwise you will overwrite everything.
# +
# exporting model performance to csv
df.to_csv('model_perf/perf1.csv', index=False) # change the name of this file
# exporting confusion matrices to json
class JSONEncoder(json.JSONEncoder):
def default(self, obj):
if hasattr(obj, 'to_json'):
return obj.to_json(orient='records')
return json.JSONEncoder.default(self, obj)
with open('cm/confusion1.json', 'w') as fp: # change the name of this file
json.dump(confusion_dict, fp, cls=JSONEncoder)
# -
# #### Concatenate performance and confusion matrices
# +
def concat_perf_dfs(filepaths):
df_list = []
for path in filepaths:
df = pd.read_csv(path)
df_list.append(df)
df = pd.concat(df_list)
df = df.sort_values(by='pct_profit_mean', ascending=False)
return df
def concat_dicts(filepaths):
confusion_dict = {}
for path in filepaths:
confusion = json.load(open(path))
confusion_dict.update(confusion)
return confusion_dict
# +
# check the number of files you have in each folder
# if you ran on 4 notebooks you should have 4
perf_csv_paths = glob.glob('model_perf/*.csv')
confusion_paths = glob.glob('cm/*.json')
print(len(perf_csv_paths))
print(len(confusion_paths))
# +
# concatenate all confusion matrices
confusion_dict = concat_dicts(confusion_paths)
# concatenate all performance dataframes
perf_df = concat_perf_dfs(perf_csv_paths)
# check the number of entries in performance data
# should be the number of models trained
# lengths of confusion_dict and perf_df should be same
print(len(confusion_dict.keys()))
print(len(perf_df))
# -
# #### look at performance dataframe
perf_df
# #### look at confusion dict
confusion_dict
# ## Model Selection
#
# Models were evaluated on several metrics to select the best models that predict arbitrage. The following were taken into consideration:
# - How often the models predicted arbitrage when it didn't exist (False positives)
# - How many times the models predicted arbitrage correctly (True positives)
# - How profitable the model was in the real world over the period of the test set.
#
# Model accuracy scores are not a good evaluation of model performance alone, and neither is profitability alone.
#
# **Why is accuracy score alone a bad measure?**
#
# The model can be making predictions of 0 (no arbitrage) 100% of the time and still achieve high accuracy because the dataset may not have that many arbitrage opportunities. So simply guessing 0 all the time could be more "accurate" than a model that makes arbitrage predictions. You want to make sure that the model is actually predicting arbitrage.
#
# **Why is profitability alone a bad measure?**
#
# Models that are profitable can be making predictions of arbitrage where arbitrage doesn't actually exist. Those predictions may be profitable by chance, but the model is not predicting what you want it to.
# ### Create a Dataframe with PNL and Confusion Matrix Features
def model_confusion(df, confusion_dict):
"""This function takes in the dataframe of performance stats
for all the models, their respective confusion matrices,
creates new features from the confusion matrices,
and returns a dataframe with all of the performance stats"""
line = '-------'
feature_dict = {}
model_name_list = []
# create a copy of df to not overwrite original
df = df.copy()
# iterate through all models
for i in range(len(df)):
# define model name
model_name = (df.ex_tp.iloc[i] + '_' + str(df.max_features.iloc[i])
+ '_' + str(df.max_depth.iloc[i]) + '_' + str(df.n_estimators.iloc[i]))
model_name_list.append(model_name)
# get confusion matrix for specific model
conf_mat = pd.read_json(confusion_dict[model_name])
#########################################################
############## create confusion features ################
#########################################################
# Some models never predicted -1, some never predicted 1, and
# some never predicted 1 or -1, meaning that they never predicted
# arbitrage at all. Each case needs to be handled with a conditional.
# confusion matrix has -1, 0, 1 predictions
if 'Predicted 1' in conf_mat.columns and 'Predicted -1' in conf_mat.columns:
# % incorrect predictions for 0, 1, -1
pct_wrong_0 = (conf_mat['Predicted 0'].loc[0] +
conf_mat['Predicted 0'].loc[2])/conf_mat['Predicted 0'].sum()
pct_wrong_1 = (conf_mat['Predicted 1'].loc[0] +
conf_mat['Predicted 1'].loc[1])/conf_mat['Predicted 1'].sum()
pct_wrong_neg1 = (conf_mat['Predicted -1'].loc[1] +
conf_mat['Predicted -1'].loc[2])/conf_mat['Predicted -1'].sum()
# total number correct arbitrage preds (-1)
correct_arb_neg1 = conf_mat['Predicted -1'].loc[0]
# total number correct arbitrage preds (1)
correct_arb_1 = conf_mat['Predicted 1'].loc[2]
# total number correct arbitrage preds (-1) + (1)
correct_arb = correct_arb_neg1 + correct_arb_1
# total number correct no arbitrage preds (0)
correct_arb_0 = conf_mat['Predicted 0'].loc[1]
# confusion matrix has 0, 1 predictions
elif 'Predicted 1' in conf_mat.columns:
pct_wrong_0 = conf_mat['Predicted 0'].loc[1] / conf_mat['Predicted 0'].sum()
pct_wrong_1 = conf_mat['Predicted 1'].loc[0] / conf_mat['Predicted 1'].sum()
pct_wrong_neg1 = np.nan
# total number correct arbitrage preds (-1)
correct_arb_neg1 = 0
# total number correct arbitrage preds (1)
correct_arb_1 = conf_mat['Predicted 1'].loc[1]
# total number correct arbitrage preds (-1) + (1)
correct_arb = correct_arb_neg1 + correct_arb_1
# total number correct no arbitrage preds (0)
correct_arb_0 = conf_mat['Predicted 0'].loc[0]
# confusion matrix has -1, 0 predictions
elif 'Predicted -1' in conf_mat.columns:
pct_wrong_0 = conf_mat['Predicted 0'].loc[0] / conf_mat['Predicted 0'].sum()
pct_wrong_1 = np.nan
pct_wrong_neg1 = conf_mat['Predicted -1'].loc[1] / conf_mat['Predicted -1'].sum()
# total number correct arbitrage preds (-1)
correct_arb_neg1 = conf_mat['Predicted -1'].loc[0]
# total number correct arbitrage preds (1)
correct_arb_1 = 0
# total number correct arbitrage preds (-1) + (1)
correct_arb = correct_arb_neg1 + correct_arb_1
# total number correct no arbitrage preds (0)
correct_arb_0 = conf_mat['Predicted 0'].loc[1]
# confusion matrix has only 0
else:
pct_wrong_0 = 0
pct_wrong_1 = 0
pct_wrong_neg1 = 0
correct_arb = 0
correct_arb_neg1 = 0
correct_arb_1 = 0
correct_arb_0 = 0
# add confusion features to dict
feature_list = [correct_arb, pct_wrong_0, pct_wrong_1, pct_wrong_neg1,
correct_arb_neg1, correct_arb_1, correct_arb_0]
feature_dict[model_name] = feature_list
# create a df from the new features
columns = ['correct_arb', 'pct_wrong_0', 'pct_wrong_1', 'pct_wrong_neg1',
'correct_arb_neg1', 'correct_arb_1', 'correct_arb_0']
df2 = pd.DataFrame(feature_dict).transpose().reset_index()
df2 = df2.rename(columns = {'index': 'model_name', 0: 'correct_arb', 1:'pct_wrong_0',
2: 'pct_wrong_1', 3: 'pct_wrong_neg1',
4: 'correct_arb_neg1', 5: 'correct_arb_1',
6: 'correct_arb_0'})
# merge new features with performance df
df['model_name'] = model_name_list
print(df.shape, df2.shape)
df = df.merge(df2, on='model_name').drop(columns = 'model_name')
print('shape after merge:', df.shape)
return df
df = model_confusion(perf_df, confusion_dict)
df
# #### Filter for best models
# +
# filter for models that are predicting arb when its not happening < 30% of the time
df2 = df[df['pct_wrong_0'] < 0.30]
print('shape after filetering pct_wrong_0:', df2.shape)
# filter for models that predict > 25 correct arb
df2 = df2[df2['correct_arb'] > 25]
print('shape after filtering correct_arb:', df2.shape)
# filter for models that make > 0.20% profit
df2 = df2[df2['pct_profit_mean'] > 0.2]
print('shape after filtering pct_profit_mean:', df2.shape)
# sort values to have the best model from each option listed first
# and drop the rest of the duplicates
df2 = df2.sort_values(by=['correct_arb'], ascending=False)
df2 = df2.drop_duplicates(subset='ex_tp')
print('shape after droping duplicates:', df2.shape)
# -
# #### Inspect best model stats
df2
# #### Move best models into a new folder
# Since there were >6K models trained, it is difficult to find the best ones in the folder with all of the models. Moving the models to a new folder makes them easier to find and download.
#
# Be super careful when running this next cell because it will rename the models with it's general name without the parameters - that is to abide by naming conventions in lambda functions. If you want to revert this change you will have to manually find the model parameters and rename each one. So do not run this unless you're absolutely sure that the way you filtered the models is final.
# +
# info for model names
models = df2['ex_tp'].values
max_features = df2['max_features'].values
max_depth = df2['max_depth'].values
n_estimators = df2['n_estimators'].values
# check that the list lengths are what you expect
# the length should be the number of rows in the filtered df
print(len(models), len(max_features), len(max_depth), len(n_estimators))
#### commented out to prevent accidental run ####
# for i in range(len(models)):
# # define model name
# model_name = models[i] + '_' + str(max_features[i]) + '_' + str(max_depth[i]) + '_' + str(n_estimators[i])
# # rename the filepath to move
# os.rename(f'pickles/{model_name}.pkl', f'arb_pickles/{models[i]}.pkl')
# +
# confirm that the correct number of models were moved
model_paths = glob.glob('arbitrage_pickles/*.pkl')
print(len(model_paths))
# -
# ## Limitations
#
# The data sets for all models were split by a 70/30 train/test split, meaning that the size of the test data varies greatly for each model. The size of the test set is larger or smaller depending on the starting size of the merged dataset for the two exchanges. The % profit mean is the average percentage gained if one were to act on all arbitrage opportunities predicted by that specific model during the period of the test set. With a larger test set, there is a better chance of seeing more arbitrage predictions, and the % profit mean number will be more accurate as it's averaged over more observations. Right now, it's possible that the % profit mean is skewed due to different testing timeframes where more or less arbitrage predictions are observed.
#
# When more data is available, it may be more accurate to apply evaluation metrics to the last 100-200 arbitrage predictions by the model so that you can make valid comparisons between models.
# ## What's next?
#
# - More hyperparameter tuning
# - Neural Networks
# - Implement auto model retraining
# - create a trigger that checks for model decay
# - retrain models in cloud if decayed, select best performers, save new version to S3 Buckets
# - save train/test data of newest version for reference
# - Create a bot that will act on arbitrage predictions from models
| cryptolytic/finalized_notebooks/arbitrage_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
import numpy as np
import pandas as pd
import matchzoo as mz
train_pack = mz.datasets.wiki_qa.load_data('train', task='ranking')
valid_pack = mz.datasets.wiki_qa.load_data('dev', task='ranking', filter=True)
predict_pack = mz.datasets.wiki_qa.load_data('test', task='ranking', filter=True)
preprocessor = mz.preprocessors.DSSMPreprocessor()
train_pack_processed = preprocessor.fit_transform(train_pack)
valid_pack_processed = preprocessor.transform(valid_pack)
predict_pack_processed = preprocessor.transform(predict_pack)
ranking_task = mz.tasks.Ranking(loss=mz.losses.RankCrossEntropyLoss(num_neg=4))
ranking_task.metrics = [
mz.metrics.NormalizedDiscountedCumulativeGain(k=3),
mz.metrics.NormalizedDiscountedCumulativeGain(k=5),
mz.metrics.MeanAveragePrecision()
]
model = mz.models.DSSM()
model.params['input_shapes'] = preprocessor.context['input_shapes']
model.params['task'] = ranking_task
model.params['mlp_num_layers'] = 3
model.params['mlp_num_units'] = 300
model.params['mlp_num_fan_out'] = 128
model.params['mlp_activation_func'] = 'relu'
model.guess_and_fill_missing_params()
model.build()
model.compile()
model.backend.summary()
pred_x, pred_y = predict_pack_processed[:].unpack()
evaluate = mz.callbacks.EvaluateAllMetrics(model, x=pred_x, y=pred_y, batch_size=len(pred_x))
train_generator = mz.PairDataGenerator(train_pack_processed, num_dup=1, num_neg=4, batch_size=64, shuffle=True)
len(train_generator)
history = model.fit_generator(train_generator, epochs=20, callbacks=[evaluate], workers=5, use_multiprocessing=False)
| tutorials/wikiqa/dssm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import os
import sys
import numpy as np
sys.path.append('../')
from dcase_models.data.datasets import URBAN_SED
# Features
from dcase_models.data.features import MelSpectrogram
from attprotos.layers import PrototypeReconstruction
from attprotos.losses import prototype_loss
from attprotos.model import AttProtos
from dcase_models.data.data_generator import DataGenerator
from dcase_models.data.scaler import Scaler
from dcase_models.util.files import load_json, load_pickle
from dcase_models.util.files import mkdir_if_not_exists, save_pickle
from dcase_models.util.data import evaluation_setup
from dcase_models.model.container import KerasModelContainer
from keras.layers import Layer
import matplotlib.pyplot as plt
#os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
# +
dataset_name = "URBAN_SED"
models_path = '../experiments'
dataset_path = './'
fold_name = "test"
model_folder = os.path.join(models_path, dataset_name, "AttProtos")
parameters_file = os.path.join(model_folder, 'config.json')
params = load_json(parameters_file)
params_features = params['features']["MelSpectrogram"]
if params_features['pad_mode'] == 'none':
params_features['pad_mode'] = None
params_dataset = params['datasets'][dataset_name]
params_model = params['models']["AttProtos"]
# Get and init dataset class
dataset_path = os.path.join(dataset_path, params_dataset['dataset_path'])
dataset = URBAN_SED(dataset_path)
# Get and init feature class
features = MelSpectrogram(**params_features)
print('Features shape: ', features.get_shape(10.0))
if not features.check_if_extracted(dataset):
print('Extracting features ...')
features.extract(dataset)
print('Done!')
# +
exp_folder = os.path.join(model_folder, fold_name)
# Load scaler
scaler_file = os.path.join(exp_folder, 'scaler.pickle')
scaler = load_pickle(scaler_file)
# Init data generator
data_gen_test = DataGenerator(
dataset, features, folds=[fold_name],
batch_size=params['train']['batch_size'],
shuffle=False, train=False, scaler=scaler
)
metrics = ['sed']
features_shape = features.get_shape()
n_frames_cnn = features_shape[1]
n_freq_cnn = features_shape[2]
n_classes = len(dataset.label_list)
model_container = AttProtos(
model=None, model_path=None, n_classes=n_classes,
n_frames_cnn=n_frames_cnn, n_freq_cnn=n_freq_cnn,
metrics=metrics,
**params_model['model_arguments']
)
model_container.load_model_weights(exp_folder)
# -
model_container.model.summary()
X, Y = data_gen_test.get_data()
# +
instance = 2
pred = model_container.model.predict(X[instance])[0]
h, mask1, mask2 = model_container.model_encoder.predict(X[instance])
alpha = model_container.model_encoder_mask.predict(X[instance])
H_hat, _ = model_container.model_h_hat().predict(X[instance])
X_hat = model_container.model_decoder.predict([H_hat, mask1, mask2])
print(X_hat.shape, pred.shape)
plt.figure(figsize=(12,8))
plt.subplot(1,2,1)
plt.imshow(np.concatenate(X[instance], axis=0).T, origin='lower')
plt.subplot(1,2,2)
plt.imshow(np.concatenate(X_hat, axis=0).T, origin='lower')
plt.show()
plt.figure(figsize=(12,8))
plt.subplot(1,2,1)
#plt.imshow((pred*(pred>0.5)).T, origin='lower')
plt.imshow((pred*(pred>0.5)).T, origin='lower')
plt.colorbar()
plt.subplot(1,2,2)
plt.imshow(Y[instance].T, origin='lower')
plt.show()
# +
import matplotlib.gridspec as grd
class_ix = 5
print(dataset.label_list[class_ix])
time_hops = np.argwhere(pred[:,class_ix]*(pred[:,class_ix]>0.5)!=0)
logits = pred
mask_logits = np.zeros_like(logits)
mask_logits[time_hops, class_ix] = 1
logits = logits*mask_logits
w = model_container.model.get_layer('dense').get_weights()[0]
grad = logits.dot(w.T)
grad = np.reshape(grad, (10, 32, 15))
alpha2 = np.sum(alpha, axis=1)
alpha_grad = grad*alpha2
alpha_grad = alpha_grad*(alpha_grad>0)
data = np.concatenate(X[instance], axis=0)
saliency = np.zeros_like(data)
for th in time_hops:
energy = np.sum(alpha_grad[th[0]]**2, axis=0)
profile = alpha_grad[th[0], :, np.argmax(energy)]
profile_extend = np.interp(np.arange(128), np.arange(32)*4, profile)
profile_extend = np.convolve(profile_extend, [1/32]*32, mode='same')
#plt.plot(profile_extend)
#plt.show()
saliency[th[0]*20:(th[0] + 1)*20] = profile_extend
plt.figure(figsize=(12,8))
#fig, ax = plt.subplots(nrows=2, ncols=1, sharex=True,figsize=(12,8))
#saliency = np.ma.masked_where((0.0 < saliency) &
# (saliency < 10), saliency)
#saliency = saliency*(saliency>0.1)
plt.imshow(data.T, origin='lower')
plt.imshow(saliency.T, origin='lower', cmap ='gray', alpha=0.5)
#ax = plt.subplot(gs[1])
#ax[1].imshow((pred*(pred>0.5)).T, origin='lower', interpolation='nearest', extent=[0, 200, 0, 10])
plt.show()
plt.figure(figsize=(12,8))
plt.imshow(saliency.T, origin='lower', cmap ='gist_yarg')
plt.axis('off')
plt.show()
plt.figure(figsize=(12,8))
masked_data = 2*((data+1)*saliency/2) - 1
#masked_data[np.abs(masked_data)<0.01] = -1
plt.imshow(masked_data.T, origin='lower')
plt.colorbar()
plt.axis('off')
plt.show()
#for j in range(11):
# plt.subplot(1,2,1)
# img = grad[:,:,j]*alpha2[:,:,j]
# plt.imshow((img*(img>np.mean(img))).T, origin='lower', vmin=0, vmax=np.amax(grad*alpha2))
#plt.imshow((grad[:,:,j]).T, origin='lower', vmin=np.amin(grad), vmax=np.amax(grad))
#plt.imshow((alpha2[:,:,j]).T, origin='lower', vmin=np.amin(alpha2), vmax=np.amax(alpha2))
# plt.colorbar()
# plt.subplot(1,2,2)
# plt.imshow(X_train[argmin][j].T, origin='lower')
#plt.title(dataset.label_list[j])
# plt.show()
# +
import matplotlib.gridspec as grd
images = []
classes_ix = [5, 7, 8]
for class_ix in classes_ix:
print(dataset.label_list[class_ix])
time_hops = np.argwhere(pred[:,class_ix]*(pred[:,class_ix]>0.5)!=0)
logits = pred
mask_logits = np.zeros_like(logits)
mask_logits[time_hops, class_ix] = 1
logits = logits*mask_logits
w = model_container.model.get_layer('dense').get_weights()[0]
grad = logits.dot(w.T)
grad = np.reshape(grad, (10, 32, 15))
alpha2 = np.sum(alpha, axis=1)
alpha_grad = grad*alpha2
alpha_grad = alpha_grad*(alpha_grad>0)
data = np.concatenate(X[instance], axis=0)
saliency = np.zeros_like(data)
for th in time_hops:
energy = np.sum(alpha_grad[th[0]]**2, axis=0)
profile = alpha_grad[th[0], :, np.argmax(energy)]
profile_extend = np.interp(np.arange(128), np.arange(32)*4, profile)
profile_extend = np.convolve(profile_extend, [1/32]*32, mode='same')
#plt.plot(profile_extend)
#plt.show()
saliency[th[0]*20:(th[0] + 1)*20] = profile_extend
masked_data = 2*((data+1)*saliency/2) - 1
images.append(np.expand_dims(masked_data, 0))
images = np.concatenate(images, axis=0)
data = np.concatenate(X[instance], axis=0)
plt.rcParams.update({'font.size': 12, 'font.family': 'serif'})
#plt.figure(figsize=(16,4))
plt.figure(figsize=(8,6))
plt.subplot(2,2,1)
plt.imshow(data.T, origin='lower')
plt.title('data instance')
plt.ylabel('mel filter index')
#plt.xlabel('hop time')
plt.subplot(2,2,2)
plt.imshow(images[0].T, origin='lower')
plt.title(dataset.label_list[classes_ix[0]])
#plt.xlabel('hop time')
plt.subplot(2,2,3)
plt.imshow(images[1].T, origin='lower')
plt.title(dataset.label_list[classes_ix[1]])
plt.xlabel('hop time')
plt.ylabel('mel filter index')
plt.subplot(2,2,4)
plt.imshow(images[2].T, origin='lower')
plt.title(dataset.label_list[classes_ix[2]])
plt.xlabel('hop time')
plt.savefig('attention_maps.png', dpi=300, bbox_inches = 'tight',pad_inches = 0)
plt.show()
# +
# Init data generator
data_gen_train = DataGenerator(
dataset, features, folds=['train'],
batch_size=params['train']['batch_size'],
shuffle=False, train=True, scaler=scaler
)
X_train, _ = data_gen_train.get_data()
print(X_train.shape)
_, distances = model_container.model_h_hat().predict(X_train)
print(distances.shape)
# -
argmin = np.argmin(distances, axis=0)
print(argmin.shape, X_train[argmin].shape)
plt.figure(figsize=(12,8))
for k in range(1, 11):
plt.subplot(2, 10, k)
plt.imshow(X_train[argmin][k-1].T, origin='lower')
plt.subplot(2, 10, k+10)
plt.imshow(alpha2[:,:,k-1].T, origin='lower', vmin=np.amin(alpha2), vmax=np.amax(alpha2))
# +
latent_closest, mask1, mask2 = model_container.model_encoder.predict(X_train[argmin])
prototypes = model_container.model.get_layer('lc').get_weights()[0]
distances_ = np.sum((prototypes - latent_closest)**2, axis=(1,3))
ixs = np.argmin(distances_, axis=1)
#prototypes = np.concatenate([prototypes]*5, axis=1)
#prototypes = np.concatenate([prototypes]*32, axis=2)
prototypes_zeros = np.zeros_like(latent_closest)
mask1_zeros = np.zeros_like(mask1)
mask2_zeros = np.zeros_like(mask2)
x_l = 0
for j, ix in enumerate(ixs):
prototypes_zeros[j, x_l, ix, :] = prototypes[j,0,0,:]
start = max(0, ix*2-8)
end = min(64, ix*2+8) if start !=0 else 16
start = start if end != 64 else 64 - 16
mask2_zeros[j, :, start:end] = 1
start = max(0, ix*4-16)
end = min(128, ix*4+16) if start !=0 else 32
start = start if end != 128 else 128 - 32
mask1_zeros[j, :, start:end] = 1
mask1 = mask1*mask1_zeros
mask2 = mask2*mask2_zeros
print(prototypes.shape)
prototypes_mel = model_container.model_decoder.predict([prototypes_zeros, mask1, mask2])
print(prototypes_mel.shape)
print(mask1.shape, mask2.shape)
# +
import matplotlib.patches as patches
print(ixs)
fig, axs = plt.subplots(2, 7, figsize=(16,6), sharex=True, subplot_kw={'aspect': 1})
for k in range(15):
#plt.subplot(1, 15, k)
start = max(0, ixs[k]*4-16)
end = min(ixs[k]*4+16, 128)
print(start, end, prototypes_mel[k][:, start:end].shape)
local_prototype = prototypes_mel[k][:, start:end]
if start == 0:
#local_prototype = np.concatenate((-np.ones((20, 32-end)), local_prototype), axis=1)
end = 32
if end == 128:
start = 128 - 32
#local_prototype = np.concatenate((local_prototype, -np.ones((20, 32-local_prototype.shape[1])), ), axis=1)
local_prototype = prototypes_mel[k][:, start:end]
#axs[k%2, k%7].imshow(local_prototype.T, origin='lower', extent=[0,20,start,end])
axs[k%2, k%7].imshow(prototypes_mel[k].T, origin='lower')
axs[k%2, k%7].set_title(k)
#
#axs[k%3, k%5]
print(k%2, k%7)
if k%2 == 1:
axs[k%2, k%7].set_xlabel('hop time')
if k%7 == 0:
axs[k%2, k%7].set_ylabel('mel filter index')
# Create a Rectangle patch
#rect = patches.Rectangle((0, ixs[k]*4-14), 20, 32, linewidth=1, edgecolor='r', facecolor='none')
# Add the patch to the Axes
#axs[k].add_patch(rect)
plt.savefig('local_prototypes.png', dpi=300, bbox_inches = 'tight',pad_inches = 0)
plt.show()
# +
plt.rcParams.update({'font.size': 12, 'font.family': 'serif'})
fig, axs = plt.subplots(2, 3, figsize=(7,7), sharex=True, subplot_kw={'aspect': 1})
for j, k in enumerate([5, 11, 9, 2, 6, 4]):
#plt.subplot(1, 15, k)
start = max(0, ixs[k]*4-16)
end = min(ixs[k]*4+16, 128)
print(start, end, prototypes_mel[k][:, start:end].shape)
local_prototype = prototypes_mel[k][:, start:end]
if start == 0:
#local_prototype = np.concatenate((-np.ones((20, 32-end)), local_prototype), axis=1)
end = 32
if end == 128:
start = 128 - 32
#local_prototype = np.concatenate((local_prototype, -np.ones((20, 32-local_prototype.shape[1])), ), axis=1)
local_prototype = prototypes_mel[k][:, start:end]
axs[j%2, j%3].imshow(local_prototype.T, origin='lower', extent=[0,20,start,end])
#axs[k%3, k%5]
print(j%3, j%2)
if j%2 == 1:
axs[j%2, j%3].set_xlabel('hop time')
if j%3 == 0:
axs[j%2, j%3].set_ylabel('mel filter index')
# Create a Rectangle patch
#rect = patches.Rectangle((0, ixs[k]*4-14), 20, 32, linewidth=1, edgecolor='r', facecolor='none')
# Add the patch to the Axes
#axs[k].add_patch(rect)
plt.savefig('local_prototypes.png', dpi=300, bbox_inches = 'tight',pad_inches = 0)
plt.show()
| notebooks/AttProtos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intermediate Pandas
# 
# **ToC**
#
# - [Navigating multilevel index](#Navigating-multilevel-index)
# - [Accessing rows and columns](#Accessing-rows-and-columns)
# - [Naming indices](#Naming-indices)
# - [Accessing rows and columns using cross section](#Accessing-rows-and-columns-using-cross-section)
# - [Missing data](#Missing-data)
# - [dropna](#dropna)
# - [fillna](#fillna)
# - [Data aggregation](#Data-aggregation)
# - [groupby](#groupby)
# - [mean min max](#mean-min-max)
# - [describe](#describe)
# - [transpose](#transpose)
# - [Combining DataFrames](#Combining-DataFrames)
# - [concat](#concat)
# - [merge](#merge)
# - [inner merge](#inner-merge)
# - [merge on multiple columns](#merge-on-multiple-columns)
# - [outer merge](#outer-merge)
# - [Sorting](#Sorting)
# - [left merge](#left-merge)
# - [right merge](#right-merge)
# - [join](#join)
#
# ## Navigating multilevel index
# + jupyter={"outputs_hidden": true}
import pandas as pd
import numpy as np
# + jupyter={"outputs_hidden": false}
# Index Levels
outside = ['G1','G1','G1','G2','G2','G2']
inside = [1,2,3,1,2,3]
hier_index = list(zip(outside,inside)) #create a list of tuples
hier_index
# + jupyter={"outputs_hidden": false}
#create a multiindex
hier_index = pd.MultiIndex.from_tuples(hier_index)
hier_index
# + jupyter={"outputs_hidden": false}
# Create a dataframe (6,2) with multi level index
df = pd.DataFrame(np.random.randn(6,2),index=hier_index,columns=['A','B'])
df
# -
# ### Accessing rows and columns
# You can use `loc` and `iloc` as a chain to access the elements. Go from outer index to inner index
# + jupyter={"outputs_hidden": false}
#access columns as usual
df['A']
# + jupyter={"outputs_hidden": false}
#access rows
df.loc['G1']
# + jupyter={"outputs_hidden": false}
#acess a single row form inner
df.loc['G1'].loc[1]
# + jupyter={"outputs_hidden": false}
#access a single cell
df.loc['G2'].loc[3]['B']
# -
# ### Naming indices
# Indices can have names (appear similar to column names)
# + jupyter={"outputs_hidden": false}
df.index.names
# + jupyter={"outputs_hidden": false}
df.index.names = ['Group', 'Serial']
df
# -
# ### Accessing rows and columns using cross section
# The `xs` method allows to get a cross section. The advantage is it can penetrate a multilevel index in a single step. Now that we have named the indices, we can use cross section effectively
# + jupyter={"outputs_hidden": false}
# Get all rows with Serial 1
df.xs(1, level='Serial')
# + jupyter={"outputs_hidden": false}
# Get rows with serial 2 in group 1
df.xs(['G1',2])
# -
# ## Missing data
# You can either drop rows/cols with missing values using `dropna()` or fill those cells with values using the `fillna()` methods.
#
# ### dropna
# Use `dropna(axis, thresh...)` where axis is 0 for rows, 1 for cols and thresh represents how many occurrences of nan before dropping happens
# + jupyter={"outputs_hidden": false}
d = {'a':[1,2,np.nan], 'b':[np.nan, 5, np.nan], 'c':[6,7,8]}
dfna = pd.DataFrame(d)
dfna
# + jupyter={"outputs_hidden": false}
# dropping rows with one or more na values
dfna.dropna()
# + jupyter={"outputs_hidden": false}
# dropping cols with one or more na values
dfna.dropna(axis=1)
# + jupyter={"outputs_hidden": false}
# Dropping rows only if 2 or more cols have na values
dfna.dropna(axis=0, thresh=2)
# -
# ### fillna
# + jupyter={"outputs_hidden": false}
dfna.fillna(value=999)
# + jupyter={"outputs_hidden": false}
# filling with mean value of entire dataframe
dfna.fillna(value = dfna.mean())
# + jupyter={"outputs_hidden": false}
# fill with mean value row by row
dfna['a'].fillna(value = dfna['a'].mean())
# -
# ## Data aggregation
# Pandas allows sql like control on the dataframes. You can treat each DF as a table and perform sql aggregation.
#
# ### groupby
# Format is: `df.groupby('col_name').aggregation()`
# + jupyter={"outputs_hidden": false}
comp_data = {'Company':['GOOG','GOOG','MSFT','MSFT','FB','FB'],
'Person':['Sam','Charlie','Amy','Vanessa','Carl','Sarah'],
'Sales':[200,120,340,124,243,350]}
comp_df = pd.DataFrame(comp_data)
comp_df
# -
# #### mean min max
# + jupyter={"outputs_hidden": false}
# mean sales by company - automatically only applies mean on numerical columns
comp_df.groupby('Company').mean()
# + jupyter={"outputs_hidden": false}
# standard deviation in sales by company
comp_df.groupby('Company').std()
# -
# You can run other aggregation functions like `mean, min, max, std, count` etc. Lets look at `describe` which does all of it.
# #### describe
# + jupyter={"outputs_hidden": false}
comp_df.groupby('Company').describe()
# -
# ### transpose
# Long over due, you can tile a DF by calling the `transpose()` method.
# + jupyter={"outputs_hidden": false}
comp_df.groupby('Company').describe().transpose()
# + jupyter={"outputs_hidden": false}
comp_df.groupby('Company').describe().index
# -
# ## Combining DataFrames
# You can concatenate, merge and join data frames.
#
# Lets take a look at 3 DataFrames
# + jupyter={"outputs_hidden": true}
df1 = pd.DataFrame({'A': ['A0', 'A1', 'A2', 'A3'],'B': ['B0', 'B1', 'B2', 'B3'],
'C': ['C0', 'C1', 'C2', 'C3'],'D': ['D0', 'D1', 'D2', 'D3']}, index=[0, 1, 2, 3])
df2 = pd.DataFrame({'A': ['A4', 'A5', 'A6', 'A7'], 'B': ['B4', 'B5', 'B6', 'B7'],
'C': ['C4', 'C5', 'C6', 'C7'],'D': ['D4', 'D5', 'D6', 'D7']}, index=[4, 5, 6, 7])
# + jupyter={"outputs_hidden": false}
df1
# + jupyter={"outputs_hidden": false}
df2
# -
# ### concat
# `pd.concat([list_of_df], axis=0)` will extend a dataframe either along rows or columns. All DF in the list should be of same dimension.
# + jupyter={"outputs_hidden": false}
# extend along rows
pd.concat([df1, df2]) #flows well because index is sequential and colmns match
# + jupyter={"outputs_hidden": false}
#extend along columns
pd.concat([df1, df2], axis=1) #fills NaN when index dont match
# -
# ### merge
# merge lets you do a sql merge with `inner, outer, right` and `left` joins.
# `pd.merge(left, right, how='outer', on='key')` where, `left` and `right` are your two DataFrames (tables) and `on` refers to the `foreign key`
# + jupyter={"outputs_hidden": false}
left = pd.DataFrame({'key1': ['K0', 'K1', 'K2', 'K3'],'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3']})
right = pd.DataFrame({'key1': ['K0', 'K1', 'K2', 'K3'],'B': ['C0', 'C1', 'C2', 'C3'],
'C': ['D0', 'D1', 'D2', 'D3']})
left
# + jupyter={"outputs_hidden": false}
right
# -
# #### inner merge
# Inner join keeps only the intersection.
# + jupyter={"outputs_hidden": false}
#merge along key1
pd.merge(left, right, how='inner', on='key1')
# -
# When both tables have same column names that are not used for merging (`on`) then pandas appends `x` and `y` to their names to differentiate
# #### merge on multiple columns
# Sometimes, your foreign key is composite. Then you can merge on multiple keys by passing a list to the `on` argument.
# Now lets add a key2 column to both the tables.
# + jupyter={"outputs_hidden": false}
left['key2'] = ['K0', 'K1', 'K0', 'K1']
left
# + jupyter={"outputs_hidden": false}
right['key2'] = ['K0', 'K0', 'K0', 'K0']
right
# + jupyter={"outputs_hidden": false}
pd.merge(left, right, how='inner', on=['key1', 'key2'])
# -
# `inner` merge will only keep the intersection, thus only 2 rows.
# #### outer merge
# Use `how='outer'` to keep the union of both the tables. pandas fills `NaN` when a cell has no values.
# + jupyter={"outputs_hidden": false}
om = pd.merge(left, right, how='outer', on=['key1', 'key2'])
om
# -
# ### Sorting
# Use `DataFrame.sort_values(by=columns, inplace=False, ascending=True)` to sort the table.
# + jupyter={"outputs_hidden": false}
om.sort_values(by=['key1', 'key2']) #now you got the merge sorted by columns.
# -
# #### right merge
# `how='right'` will keep all the rows of right table and drop the rows of left table that dont have a matching keys.
# + jupyter={"outputs_hidden": false}
pd.merge(left, right, how='right', on=['key1', 'key2']).sort_values(by='key1')
# -
# #### left merge
# `how='left'` will similarly keep all rows of left and those rows of right that has a matching foreign key.
# + jupyter={"outputs_hidden": false}
pd.merge(left, right, how='left', on=['key1', 'key2']).sort_values(by='key1')
# -
# ### join
# Joins are like merges but work on index instead of columns. Further, they are by default either `left` or `right` with `inner` as mode of joins. See example below:
# + jupyter={"outputs_hidden": false}
df_a = pd.DataFrame({'A': ['A0', 'A1', 'A2'],
'B': ['B0', 'B1', 'B2']},
index=['K0', 'K1', 'K2'])
df_b = pd.DataFrame({'C': ['C0', 'C2', 'C3'],
'D': ['D0', 'D2', 'D3']},
index=['K0', 'K2', 'K3'])
df_a
# + jupyter={"outputs_hidden": false}
df_b
# + jupyter={"outputs_hidden": false}
#join b to a, default mode = keep all rows of a and matching rows of b (left join)
df_a.join(df_b)
# -
# Thus all rows of df_a and those in df_b. If df_b did not have that index, then NaN for values.
# + jupyter={"outputs_hidden": false}
#join b to a
df_b.join(df_a)
# + jupyter={"outputs_hidden": false}
#outer join - union of outputs
df_b.join(df_a, how='outer')
| python_crash_course/pandas_cheat_sheet_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] cell_id="00001-5ba44c48-d6b5-40a5-a1e5-5986b1eb70da" deepnote_cell_type="markdown" deletable=false
# # Problem Set 0 - Python & Jupyter Notebooks
#
# In this problem set, you will learn how to:
#
# * navigate jupyter notebooks (like this one).
# * write and evaluate some basic *expressions* in Python, the computer language of the course.
# * learn data manipulation using the `pandas` package.
#
# For reference, you might find it useful to read [Chapter 3 of the Data 8 textbook](http://www.inferentialthinking.com/chapters/03/programming-in-python.html), [Chapter 1](https://www.inferentialthinking.com/chapters/01/what-is-data-science.html) and [Chapter 2](https://www.inferentialthinking.com/chapters/02/causality-and-experiments.html) are worth skimming as well.
# + [markdown] cell_id="00002-e3698e92-65c5-412d-9123-7c4b240ac2b2" deepnote_cell_type="markdown"
# ## 1. Jupyter Notebooks
#
# This webpage is called a Jupyter notebook. A notebook is an interactive computing environment to write programs and view their results, and also to write text. A notebook is thus an editable computer document in which you can write computer programs; view their results; and comment, annotate, and explain what is going on.
# + [markdown] cell_id="00004-c2326993-e03c-4c91-a1f8-a0a778556c43" deepnote_cell_type="markdown"
# ### 1.1. Text cells
#
# In a notebook, each rectangle containing text or code is called a *cell*.
#
# Text cells (like this one) can be edited by double-clicking on them. They're written in a simple format called [markdown](http://daringfireball.net/projects/markdown/syntax) to add formatting and section headings.
# You don't need to learn too much about markdown.
#
# After you edit a text cell, click the "run cell" button at the top that looks like `▶` in the toolbar at the top of this window, or hold down `shift` + press `return`, to confirm any changes to the text and formatting.
# + [markdown] cell_id="00005-fe58f6f7-ce48-4a4d-88ec-0ff0a62117fd" deepnote_cell_type="markdown" deletable=false
# **Question 1.1.1.** This paragraph is in its own text cell. Try editing it so that this sentence is the last sentence in the paragraph, and then click the "run cell" ▶| button or hold down `shift` + `return`. This sentence, for example, should be deleted. So should this one.
# + [markdown] cell_id="00006-ec8c2887-95f3-4b5c-bcfa-1da3cbc26f23" deepnote_cell_type="markdown"
# ### 1.2. Code cells
#
# Other cells contain code in the Python 3 language. Running a code cell will execute all of the code it contains.
#
# To run the code in a code cell, first click on that cell to activate it. It'll be highlighted with a little green or blue rectangle. Next, either press `▶` or hold down `shift` + press `return`.
#
# Try running this cell:
# + cell_id="00007-ae472ab8-54dc-43a7-be16-a6c5328fc3fc" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786118508 source_hash="a8a44a2d"
print("Hello, world!")
# + [markdown] cell_id="00010-c9132ae7-7621-4d3a-b0ae-b823e0984327" deepnote_cell_type="markdown"
# The fundamental building block of Python code is an expression. Cells can contain multiple lines with multiple expressions. When you run a cell, the lines of code are executed in the order in which they appear. Every `print` expression prints a line. Run the next cell and notice the order of the output.
# + cell_id="00011-41446c4e-9422-4753-9a79-e274f05b5650" deepnote_cell_type="code" deepnote_to_be_reexecuted=false deletable=false execution_millis=2 execution_start=1610786119046 source_hash="c21a1a0b"
print("First this line is printed,")
print("and then this one.")
# + [markdown] cell_id="00012-1f4e4107-c775-4250-8591-c7cacf0b2855" deepnote_cell_type="markdown"
# ### 1.3. Writing notebooks
#
# You can use Jupyter notebooks for your own projects or documents. When you make your own notebook, you'll need to create your own cells for text and code.
#
# To add a cell, click the + button in the menu bar. It'll start out as a text cell. You can change it to a code cell by clicking inside it so it's highlighted, clicking the drop-down box next to the restart (⟳) button in the menu bar, and choosing "Code".
# + [markdown] cell_id="00013-f36b1724-6714-4eb5-98f9-8bb5964156a0" deepnote_cell_type="markdown" tags=[]
# **Question 1.3.1.** Add a code cell below this one. Write code in it that prints out:
#
# Econometrics is what econometricians do.
#
# Run your cell to verify that it works.
# + [markdown] cell_id="00019-7bcb08be-3668-4bab-b7e2-45f73e6b6d9d" deepnote_cell_type="markdown" tags=[]
# ### 1.4. The kernel
#
# The kernel is a program that executes the code inside your notebook and outputs the results. In the top right of your window, you can see a circle that indicates the status of your kernel. If the circle is empty (⚪), the kernel is idle and ready to execute code. If the circle is filled in (⚫), the kernel is busy running some code.
#
# Next to every code cell, you'll see some text that says `In [...]`. Before you run the cell, you'll see `In [ ]`. When the cell is running, you'll see `In [*]`. If you see an asterisk (\*) next to a cell that doesn't go away, it's likely that the code inside the cell is taking too long to run, and it might be a good time to interrupt the kernel (discussed below). When a cell is finished running, you'll see a number inside the brackets, like so: `In [1]`. The number corresponds to the order in which you run the cells; so, the first cell you run will show a 1 when it's finished running, the second will show a 2, and so on.
#
# You may run into problems where your kernel is stuck for an excessive amount of time, your notebook is very slow and unresponsive, or your kernel loses its connection. If this happens, try the following steps:
# 1. At the top of your screen, click **Kernel**, then **Interrupt**.
# 2. If that doesn't help, click **Kernel**, then **Restart**. If you do this, you will have to run your code cells from the start of your notebook up until where you paused your work.
# 3. If that doesn't help, restart your server. First, save your work by clicking **File** at the top left of your screen, then **Save and Checkpoint**. Next, click **Control Panel** at the top right. Choose **Stop My Server** to shut it down, then **Start My Server** to start it back up. Then, navigate back to the notebook you were working on. You'll still have to run your code cells again.
#
# **IMPORTANT:** If you leave your notebook alone for a while, the server will "clear" the code you've run and you'll have to run your notebook from the very top again. If you've been away from your computer and after coming back you get a `<something> not defined` error this is almost certainly what happened.
# + [markdown] cell_id="00020-8dd1af80-14dd-4318-a2e2-a2083e3d7286" deepnote_cell_type="markdown"
# ### 1.5. Libraries
#
# There are many add-ons and extensions to the core of Python that are useful to using it to get work done. They are contained in what are called libraries. Let's tell Python to install some of them. Run the code cell below to do so.
#
# The following lines import `numpy` and `pandas`.
# Note that we imported `pandas` as `pd`, `numpy` as `np`.
# This means that, for example, when we call functions in `pandas`, we will always reference them with `pd` first. `pd` is like a shortcut.
# + cell_id="00021-477baf2d-9aee-42b5-9126-87c7c15c7432" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4813 execution_start=1610786265948 source_hash="1ef69260"
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
# + [markdown] cell_id="00022-af76bee0-0669-49a7-8ed4-f2b0296f9d2a" deepnote_cell_type="markdown"
# ## 2. Python: Numbers & Variables
#
# In addition to representing commands to print out lines, expressions can represent numbers and methods of combining numbers. The expression `3.2500` evaluates to the number 3.25. (Run the cell and see.)
# + cell_id="00023-4ab2cb54-482e-4098-b61c-cdece0d25c4d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1610786125030 source_hash="6b5a05b0" tags=[]
3.2500
# + [markdown] cell_id="00024-9a336f6d-d3e4-47bc-b1da-5dd9734bb562" deepnote_cell_type="markdown"
# Notice that we didn't have to `print`. When you run a notebook cell, if the last line has a value, then Jupyter helpfully prints out that value for you. However, it won't print out prior lines automatically.
# + cell_id="00025-9496fc5f-9066-4189-b031-bdade66fefbd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=5 execution_start=1610786125035 source_hash="200996a9"
print(2)
3
4
# + [markdown] cell_id="00026-952d384d-7672-4024-bba6-435a93602e00" deepnote_cell_type="markdown"
# Above, you should see that 4 is the value of the last expression, 2 is printed, but 3 is not because it was neither printed nor last.
# + [markdown] cell_id="00027-9b23cc47-9145-46f3-8c2b-685b5b47a350" deepnote_cell_type="markdown"
# ### 2.1. Arithmetic
#
# The line in the next cell subtracts. Its value is what you'd expect. Run it.
# + cell_id="00028-57401a7b-9f50-4998-8df3-c6985d9c8db0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=6 execution_start=1610786125043 source_hash="e40f2e60"
3.25 - 1.5
# + [markdown] cell_id="00029-65d182c9-e9b1-4f1a-bda3-efb03d1f8a78" deepnote_cell_type="markdown"
# Many basic arithmetic operations are built into Python. The Data 8 textbook section on [Expressions](http://www.inferentialthinking.com/chapters/03/1/expressions.html) describes all the arithmetic operators used in the course.
# The common operator that differs from typical math notation is `**`, which raises one number to the power of the other. So, `2**3` stands for $2^3$ and evaluates to 8.
#
# The order of operations is the same as what you learned in elementary school, and Python also has parentheses. For example, compare the outputs of the cells below. The second cell uses parentheses for a happy new year!
# + cell_id="00030-5a276350-c699-4459-aa56-8068c8ff3208" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786125088 source_hash="e8257ac"
5+6*5-6*3**2*2**3/4*7
# + cell_id="00031-dd95116a-b021-4225-83b6-a1b77572a5c6" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786125089 source_hash="197a43b6"
5+(6*5-(6*3))**2*((2**3)/4*7)
# + [markdown] cell_id="00032-5e18eac2-67c4-4c58-a4e4-c0875a3de5c6" deepnote_cell_type="markdown"
# In standard math notation, the first expression is
#
# $$5 + 6 \times 5 - 6 \times 3^2 \times \frac{2^3}{4} \times 7,$$
#
# while the second expression is
#
# $$5 + (6 \times 5 - (6 \times 3))^2 \times (\frac{(2^3)}{4} \times 7).$$
# + [markdown] cell_id="00033-e2d2cbaa-4530-4dd8-b7ef-52e7ddd4f62d" deepnote_cell_type="markdown" tags=[]
# **Question 2.1.1.** Write a Python expression in this next cell that's equal to $5 \times (3 \frac{10}{11}) - 50 \frac{1}{3} + 2^{.5 \times 22} - \frac{7}{33} + 4$.
#
# By "$3 \frac{10}{11}$" we mean $3+\frac{10}{11}$, not $3 \times \frac{10}{11}$.
#
# Replace the ellipses (`...`) with your expression. Try to use parentheses only when necessary.
#
# *Hint:* The correct output should start with a familiar number.
# + cell_id="00034-629156ca-cb2a-4575-8322-0a028813bb1b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false deletable=false execution_millis=2 execution_start=1610786127399 source_hash="1f85637f"
...
# + [markdown] cell_id="00035-a7dac102-1cde-4aa3-a775-c7b216ddfbd5" deepnote_cell_type="markdown"
# ### 2.2. Variables
#
# In natural language, we have terminology that lets us quickly reference complicated concepts.
#
# In Python, we do this with *assignment statements*. An assignment statement has a name on the left side of an `=` sign and an expression to be evaluated on the right.
# + cell_id="00036-2fe6d498-06b5-4665-b860-ab911b87da10" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786127420 source_hash="42fbeeaf"
ten = 3 * 2 + 4
# + [markdown] cell_id="00037-1ac7687a-b971-45f0-b84b-f287e24b89af" deepnote_cell_type="markdown"
# When you run that cell, Python first computes the value of the expression on the right-hand side, `3 * 2 + 4`, which is the number 10. Then it assigns that value to the name `ten`. At that point, the code in the cell is done running.
#
# After you run that cell, the value 10 is bound to the name `ten`:
# + cell_id="00038-52b7ddec-2d9b-4313-9f1f-bb0ae713668e" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786127460 source_hash="d8b71231"
ten
# + [markdown] cell_id="00039-97d394b6-42fe-4237-92e0-d882a4f9c7d3" deepnote_cell_type="markdown"
# The statement `ten = 3 * 2 + 4` is not asserting that `ten` is already equal to `3 * 2 + 4`, as we might expect by analogy with math notation. Rather, that line of code changes what `ten` means; it now refers to the value 10, whereas before it meant nothing at all.
#
# **IMPORTANT:** The order you run the cells matters. Even though the cell that says `ten` happens after the cell that says `ten = 3 * 2 + 4` in this notebook, you still have to run the cell that says `ten = 3 * 2 + 4` before you run the cell that says `ten`. If you start up this notebook and the first cell you run is the `ten` cell, you'll get an error that says something like `ten not defined`. That's because you haven't defined the variable `ten` above.
# + [markdown] cell_id="00040-0948d67c-7dd0-4ac5-b934-70c361446df3" deepnote_cell_type="markdown" tags=[]
# **Question 2.2.1.** Try writing code that uses a name (like `eleven`) that hasn't been assigned to anything. You'll see an error!
# + cell_id="00041-72cd5a01-c795-4af5-aa8f-645d2eaa9b25" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786127461 source_hash="1f85637f"
...
# + [markdown] cell_id="00042-50b9b262-9898-4365-a2c4-e794bd67ca2d" deepnote_cell_type="markdown"
# A common pattern in Jupyter notebooks is to assign a value to a name and then immediately evaluate the name in the last line in the cell so that the value is displayed as output.
# + cell_id="00043-422c0b69-ce93-4998-85d1-7c07cbbef544" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1610786127509 source_hash="84e41d80"
close_to_pi = 355/113
close_to_pi
# + [markdown] cell_id="00044-dca25950-ecd1-40c7-8097-278fe49c2e9b" deepnote_cell_type="markdown"
# Another common pattern is that a series of lines in a single cell will build up a complex computation in stages, naming the intermediate results.
# + cell_id="00045-0c223c8e-e4e3-4e0c-a744-0552799eebec" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786127556 source_hash="f49d2e00"
semimonthly_salary = 841.25
monthly_salary = 2 * semimonthly_salary
number_of_months_in_a_year = 12
yearly_salary = number_of_months_in_a_year * monthly_salary
yearly_salary
# + [markdown] cell_id="00046-25c0743d-9215-45c9-a77c-f3c54019a642" deepnote_cell_type="markdown"
# Names in Python can have letters (upper- and lower-case letters are both okay and count as different letters), underscores, and numbers. The first character can't be a number (otherwise a name might look like a number). And names can't contain spaces, since spaces are used to separate pieces of code from each other.
#
# Other than those rules, what you name something doesn't matter *to Python*. For example, this cell does the same thing as the above cell, except everything has a different name:
# + cell_id="00047-33a38e69-b70d-489b-b0cb-796d8ce4b9f5" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786127557 source_hash="fa245a47"
a = 841.25
b = 2 * a
c = 12
d = c * b
d
# + [markdown] cell_id="00048-97f0be3a-0928-4c7e-978e-16530d4a69c5" deepnote_cell_type="markdown"
# **However**, names are very important for making your code *readable* to yourself and others. The cell above is shorter, but it's totally useless without an explanation of what it does.
# + [markdown] cell_id="00049-1a1787ca-aadf-4cb8-bbbc-7f46f73221e7" deepnote_cell_type="markdown"
# ### 2.3. Checking your code
#
# Our notebooks include built-in *tests* to check whether your work is correct. Sometimes, there are multiple tests for a single question, and passing all of them is required to receive credit for the question. Please don't change the contents of the test cells.
#
# Run the next code cell to initialize the tests:
# + cell_id="00050-2b42e499-9ad0-4520-b13b-026f82bfce02" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1686 execution_start=1610786127557 source_hash="fd524aa"
# Using the pound symbol tells Python to ignore the rest of this line
# This was we can make comments in code
# Run this code cell to initialize the autograder
import otter
grader = otter.Notebook()
# + [markdown] cell_id="00051-6f3169db-5880-4f62-a612-af5f7ca718ca" deepnote_cell_type="markdown"
# Go ahead and attempt Question 2.3.1. Running the cell directly after it will test whether you have assigned `seconds_in_a_decade` correctly.
# If you haven't, this test will tell you the correct answer. Resist the urge to just copy it, and instead try to adjust your expression.
# + [markdown] cell_id="00052-c6984302-d81f-4812-813c-0ad9b0561d0e" deepnote_cell_type="markdown" deletable=false editable=false
# **Question 2.3.1.** Assign the name `seconds_in_a_decade` to the number of seconds between midnight January 1, 2010 and midnight January 1, 2020. Note that there are two leap years in this span of a decade. A non-leap year has 365 days and a leap year has 366 days.
#
# <!--
# BEGIN QUESTION
# name: q2_3_1
# -->
# + cell_id="00053-beaf12a3-67e5-4d78-9e29-0f507be6031b" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=2 execution_start=1610786129248 source_hash="a6086c3b" tags=[]
# Using the pound symbol tells Python to ignore the rest of this line.
# This way we can make comments in code.
# Change the next line so that it computes the number of seconds in a decade
# and assigns that number the name, seconds_in_a_decade.
seconds_in_a_decade = ...
# We've put this line in this cell so that it will print the value you've given
# to seconds_in_a_decade when you run it. You don't need to change this.
seconds_in_a_decade
# + deletable=false editable=false
grader.check("q2_3_1")
# + [markdown] cell_id="00055-052b2f15-283c-4581-ae59-52f2c9caa60a" deepnote_cell_type="markdown" tags=[]
# If the autograder found that you had set the right variable(s) to the proper value(s) that it expected, well and good: you are probably not far off track. If you failed any of the tests, go back and try again.
# + [markdown] cell_id="00123-23433c3d-aa14-409e-a776-bc230cc0de91" deepnote_cell_type="markdown"
# ## 3. Data
#
# We will be using `pandas` in this course for data manipulation and analysis. `pandas` stores data in the form of something called a `DataFrame`, which is really just another word for table.
#
# Recall that we typically use `pd` as the shortcut for `pandas`.
# + [markdown] cell_id="00124-c243a68b-fec5-4610-82fe-7c54e68b383e" deepnote_cell_type="markdown" tags=[]
# ### 3.1. Reading in a dataset
#
# Most of the time, the data we want to analyze will be in a separate file, typically as a `.csv` file.
# In this case, we want to read the files in and convert them into a tabular format.
#
# `pandas` has a specific function to read in csv files called `pd.read_csv(<file_path>)`, with the same relative file path as its argument. Your dataset will take the form of a variable, as shown below where we call our dataset `baby_df`.
#
# The `<dataframe>.head(...)` function will display the first 5 rows of the data frame by default.
# If you want to specify the number of rows displayed, you can use`dataframe.head(<num_rows>)`.
# Similarly, if you want to see the last few rows of the data frame, you can use `dataframe.tail(<num_rows>)`.
# + cell_id="00125-89701414-7a69-4d73-837f-47efff6f528f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=34 execution_start=1610786141720 source_hash="c73a235d" tags=[]
# We will call our dataset baby_df
baby_df = pd.read_csv("baby.csv") # df is short for dataframe
baby_df.head(5)
# + [markdown] cell_id="00126-85c69fc5-33b1-46dd-8844-7abb75fce106" deepnote_cell_type="markdown" tags=[]
# ### 3.2. Columns
#
# You can access the values of a particular column by using `<dataframe>['<column_name>']`.
# Notice that you have to write the column name in quotes. Single or double quotes both work fine, but here we use single quotes because it's faster to type.
#
#
# + cell_id="00127-46a06813-3191-42e0-98f2-4a3752487799" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786141755 source_hash="8c56e4ba" tags=[]
baby_df['Birth.Weight']
# + [markdown] cell_id="00130-0417633e-acf1-43bc-b379-60e7715f4d13" deepnote_cell_type="markdown" tags=[]
# ### 3.3. Getting the shape of a table
#
# The number of rows and columns in a `DataFrame` can be accessed together using the `.shape` attribute.
# Notice that the index is not counted as a column.
# + cell_id="00131-3bf05c79-f4e5-48d8-ad05-31e1ce904e22" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1 execution_start=1610786141755 source_hash="6655aad2" tags=[]
baby_df.shape
# + [markdown] cell_id="00144-632df95f-06fb-47fe-8ee1-5b6f87f19013" deepnote_cell_type="markdown" tags=[]
# ### 3.4. Selecting columns
#
# Sometimes the entire dataset contains too many columns, and we are only interested in some of the columns.
# In these situations, we would want to be able to select and display a subset of the columns from the original table.
#
# We can do this using the following syntax: `<dataframe>[[<list of columns we want to select>]]`.
#
# Note that there must be two sets of square brackets.
# + cell_id="00145-74cdf44d-d1ed-497b-b799-dcb802cbecaf" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=37 execution_start=1610786141756 source_hash="fa2bfe08" tags=[]
# Selects the columns "Birth.Weight" and "Maternal.Age" with all rows
baby_df[['Birth.Weight', 'Maternal.Age']]
# + [markdown] cell_id="00057-0b288994-33a7-4e02-8766-0c1646236d46" deepnote_cell_type="markdown" tags=[]
# We haven't actually changed anything about the dataset by doing this.
# + cell_id="00058-cec4c2cd-b646-40ea-9f37-89a944ad500f" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=21 execution_start=1610786141793 source_hash="59ff2a8a" tags=[]
baby_df.head()
# + [markdown] cell_id="00059-1369f2c8-6c7d-4430-95d3-f737dafa1c6f" deepnote_cell_type="markdown" tags=[]
# To save the dataset with fewer columns, define a new variable (or re-define the same dataset variable).
# + cell_id="00060-e76c1805-cacf-4166-8f2e-55e49f764c36" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=0 execution_start=1610786141814 source_hash="fd3fd1c" tags=[]
smaller_baby_df = baby_df[['Birth.Weight', 'Maternal.Age']]
smaller_baby_df.head()
# + [markdown] cell_id="00150-1e1b6b51-9715-4343-bdd7-44b49f09c448" deepnote_cell_type="markdown" tags=[]
# ## 4. Techniques in `pandas`
# + [markdown] cell_id="00151-361d8f9b-6258-4454-b585-495d305c8341" deepnote_cell_type="markdown" tags=[]
# ### 4.1. Filtering and boolean indexing
#
# Sometimes, we would like to filter a table by only returning rows that satisfy a specific condition.
# We can do this in `pandas` by "boolean indexing".
# The expression below returns a boolean column where an entry is `True` if the row satisfies the condition and `False` if it doesn't.
# + cell_id="00152-7bbdc810-a2c0-4e51-85ca-c2511326fdd7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=14 execution_start=1610786141815 source_hash="c1aaf83f" tags=[]
baby_df['Birth.Weight'] > 120
# + [markdown] cell_id="00153-b88a305e-2fc5-4b20-9efa-acb36cc943c3" deepnote_cell_type="markdown" tags=[]
# If we want to filter our data for all rows that satisfy `'Birth.Weight'` > 120, we can use the above expression as if we were selecting a column, as below.
# The idea is that we only want the rows where the "boolean column" is `True`.
# + cell_id="00154-efa7ca40-a0ae-4e20-82bc-b33c653c4e72" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=45 execution_start=1610786141829 source_hash="bb40d8f2" tags=[]
# Select all rows that are True in the boolean series baby_df['Birth.Weight'] > 120
baby_df[baby_df['Birth.Weight'] > 120]
# + [markdown] cell_id="00155-46430a22-366f-4593-b8fe-7d8b120fa87b" deepnote_cell_type="markdown" tags=[]
# You can tell we're missing some rows by the fact that the index on the very left is missing some numbers.
#
# Here are a few more examples:
# + cell_id="00156-42ad92f6-12db-4f2f-9c60-b666c1b78465" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=28 execution_start=1610786141875 source_hash="8dd3ab0b" tags=[]
# Return all rows where Maternal.Height is greater than or equal to 63
baby_df[baby_df['Maternal.Height'] >= 63]
# + [markdown] cell_id="00064-3738271c-7160-4ad2-9fc2-de450dacfd6f" deepnote_cell_type="markdown" tags=[]
# Notice below that to ask if something is equal to something else we use `==`, which is different from defining a variable. That uses `=`.
# + cell_id="00157-279898a5-44fd-41b5-ab88-6632d022e373" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=27 execution_start=1610786141904 source_hash="2f93444c" tags=[]
# Return all rows where Maternal.Smoker is True
baby_df[baby_df['Maternal.Smoker'] == True]
# + [markdown] cell_id="00158-5749a37f-3923-4e06-8eaa-99cca1825e93" deepnote_cell_type="markdown" tags=[]
# ### 4.2. Filtering on multiple conditions
#
# We can also filter on multiple conditions.
# If we want rows where each condition is true, we separate our criterion by the `&` symbol, where `&` represents *and*.
#
# `df[(boolean condition 1) & (boolean condition 2) & (boolean condition 2)]`
#
# If we just want one of the conditions to be true, we separate our criterion by `|` symbols, where `|` represents *or*.
#
# `df[(boolean condition 1) | (boolean condition 2) | (boolean condition 2)]`
# + cell_id="00159-bdf8bf7c-0598-443d-b23a-79b35daf1343" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=18 execution_start=1610786141931 source_hash="b13ae7f1" tags=[]
# Select all rows where Gestational.Days are above or equal to 270, but less than 280
baby_df[(baby_df['Gestational.Days'] >= 270) & (baby_df['Gestational.Days'] < 280)]
# + [markdown] cell_id="00072-51e3d449-81ee-4141-9667-61396f8e5d85" deepnote_cell_type="markdown" tags=[]
# Again though, none of this changed the original dataset.
# + cell_id="00073-4386c08f-f6a8-4d7e-9169-c1675565aa49" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=20 execution_start=1610786141950 source_hash="59ff2a8a" tags=[]
baby_df.head()
# + [markdown] cell_id="00074-76d47ce5-895b-47ca-b7b9-6c411f6106df" deepnote_cell_type="markdown" tags=[]
# To save a new dataset with your desired filtering, you should define new variables.
# + cell_id="00075-eb6dc1f2-d9d5-474c-af75-b0fe353229b2" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=23 execution_start=1610786141970 source_hash="fb1f2afe" tags=[]
gestational_range_baby_df = baby_df[(baby_df['Gestational.Days'] >= 270) & (baby_df['Gestational.Days'] < 280)]
gestational_range_baby_df.head()
# + [markdown] cell_id="00162-804f6946-457c-493f-9a44-845c19b1f007" deepnote_cell_type="markdown" output_cleared=false tags=[]
# ### 4.3. Inserting new columns
#
# Suppose we know that the device used to measure mother height was not calibrated correctly and underestimated by 2 units for each mother. To correct this, we want to add 2 to each observation of mother height. The code below shows how to do this. The syntax is similar to defining a new variable. Notice we can do math with columns as if they were numbers, and the math is appplied to each element in the column.
# + cell_id="00165-067a1f3b-64c4-48d5-8f64-e930ec68d3ed" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=43 execution_start=1610786141993 output_cleared=false source_hash="bb0cd2cc" tags=[]
baby_df['Adjusted Height'] = baby_df['Maternal.Height'] + 2
baby_df.head()
# + [markdown] cell_id="00178-08172d4e-bb6c-47f8-add8-e4e602468c9c" deepnote_cell_type="markdown" tags=[]
# ### 4.4. An exercise in `pandas`
#
# In this exercise, you will use some common functions in `pandas` that are featured above.
# + [markdown] cell_id="00179-833693b1-6828-425b-8fcd-49904ace1af5" deepnote_cell_type="markdown" deletable=false editable=false tags=[]
# **Question 4.4.1.** Read in the table `gdp.csv`, storing into the variable `gdp`.
#
# <!--
# BEGIN QUESTION
# name: q4_4_1
# -->
# + cell_id="00180-8a399cd2-cf7e-4d8c-bbea-99f319ad0b79" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=161 execution_start=1610786142037 source_hash="ce5d72c7" tags=[]
gdp = ...
gdp
# + deletable=false editable=false
grader.check("q4_4_1")
# + [markdown] cell_id="00182-11ef23d2-9c2e-45fb-bdcd-dd18ba952875" deepnote_cell_type="markdown" tags=[]
# The three variables in `gdp` that we are interested in are the following:
#
# 1. `cn` $\Rightarrow$ Capital Stock in millions of USD
# 2. `cgdpe` $\Rightarrow$ Expenditure-side Real GDP in millions of USD
# 3. `emp` $\Rightarrow$ Number of Persons employed in millions
# + [markdown] cell_id="00183-84673f29-160f-4a9b-a45e-95ea472691f1" deepnote_cell_type="markdown" deletable=false editable=false tags=[]
# **Question 4.4.2.**
# Select the columns `country`, `year`, `cn`, `cgdpe`, `emp`, from the dataframe called `gdp`. Call the new table `gdp2` and display its first five rows.
#
# <!--
# BEGIN QUESTION
# name: q4_4_2
# -->
# + cell_id="00184-4b4ab0e4-1a71-4d7b-b4fa-796ac49bfe76" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=22 execution_start=1610786142205 source_hash="d686982" tags=[]
gdp2 = ...
gdp2.head()
# + deletable=false editable=false
grader.check("q4_4_2")
# + [markdown] cell_id="00189-e9841d53-092a-48a9-bdc8-b87b3051c3ca" deepnote_cell_type="markdown" deletable=false editable=false tags=[]
# **Question 4.4.3.**
# Notice that there are a lot of -1 values. This dataset uses -1 to indicate missing data for a given country-year combination, so we don't really care about these rows.
# Filter out all the rows in which the GDP, employment level, or capital stock was recorded as -1 and store the corresponding table to the variable `cleaned_gdp`. Use the `gdp2` table you defined above.
#
# <!--
# BEGIN QUESTION
# name: q4_4_3
# -->
# + cell_id="00190-fe06e70b-f84f-4891-8023-709dce7dc377" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=4 execution_start=1610786142277 source_hash="5877bd4a" tags=[]
cleaned_gdp = ...
cleaned_gdp
# + deletable=false editable=false
grader.check("q4_4_3")
# + [markdown] cell_id="00206-36a765ee-0e6b-4715-bbf2-964e9add3729" deepnote_cell_type="markdown" deletable=false editable=false tags=[]
# **Question 4.4.4.**
# Compute the GDP per employed-person and add that as a column called `gdp_pc` to `cleaned_gdp`.
#
# *Hint*: Remember you can divide a column by another column, and this will do element-wise division.
#
# <!--
# BEGIN QUESTION
# name: q4_4_4
# -->
# + cell_id="00207-5a5abcc3-1dfc-4abc-a3ad-5a3b4be5fb23" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=52 execution_start=1610786281315 source_hash="ebaf8e4c" tags=[]
...
cleaned_gdp
# + deletable=false editable=false
grader.check("q4_4_4")
# + [markdown] deletable=false editable=false
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false
grader.check_all()
| ps0/ps0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
import sys, os
if 'google.colab' in sys.modules and not os.path.exists('.setup_complete'):
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/setup_colab.sh -O- | bash
# !pip install --upgrade https://github.com/Theano/Theano/archive/master.zip
# !pip install --upgrade https://github.com/Lasagne/Lasagne/archive/master.zip
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week05_explore/bayes.py
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week05_explore/action_rewards.npy
# !wget -q https://raw.githubusercontent.com/yandexdataschool/Practical_RL/spring20/week05_explore/all_states.npy
# !touch .setup_complete
# This code creates a virtual display to draw game images on.
# It will have no effect if your machine has a monitor.
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# !bash ../xvfb start
os.environ['DISPLAY'] = ':1'
# +
from abc import ABCMeta, abstractmethod, abstractproperty
import enum
import numpy as np
np.set_printoptions(precision=3)
np.set_printoptions(suppress=True)
import pandas
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Contents
# * [1. Bernoulli Bandit](#Part-1.-Bernoulli-Bandit)
# * [Bonus 1.1. Gittins index (5 points)](#Bonus-1.1.-Gittins-index-%285-points%29.)
# * [HW 1.1. Nonstationary Bernoulli bandit](#HW-1.1.-Nonstationary-Bernoulli-bandit)
# * [2. Contextual bandit](#Part-2.-Contextual-bandit)
# * [2.1 Bulding a BNN agent](#2.1-Bulding-a-BNN-agent)
# * [2.2 Training the agent](#2.2-Training-the-agent)
# * [HW 2.1 Better exploration](#HW-2.1-Better-exploration)
# * [3. Exploration in MDP](#Part-3.-Exploration-in-MDP)
# * [Bonus 3.1 Posterior sampling RL (3 points)](#Bonus-3.1-Posterior-sampling-RL-%283-points%29)
# * [Bonus 3.2 Bootstrapped DQN (10 points)](#Bonus-3.2-Bootstrapped-DQN-%2810-points%29)
#
# ## Part 1. Bernoulli Bandit
#
# We are going to implement several exploration strategies for simplest problem - bernoulli bandit.
#
# The bandit has $K$ actions. Action produce 1.0 reward $r$ with probability $0 \le \theta_k \le 1$ which is unknown to agent, but fixed over time. Agent's objective is to minimize regret over fixed number $T$ of action selections:
#
# $$\rho = T\theta^* - \sum_{t=1}^T r_t$$
#
# Where $\theta^* = \max_k\{\theta_k\}$
#
# **Real-world analogy:**
#
# Clinical trials - we have $K$ pills and $T$ ill patient. After taking pill, patient is cured with probability $\theta_k$. Task is to find most efficient pill.
#
# A research on clinical trials - https://arxiv.org/pdf/1507.08025.pdf
class BernoulliBandit:
def __init__(self, n_actions=5):
self._probs = np.random.random(n_actions)
@property
def action_count(self):
return len(self._probs)
def pull(self, action):
if np.any(np.random.random() > self._probs[action]):
return 0.0
return 1.0
def optimal_reward(self):
""" Used for regret calculation
"""
return np.max(self._probs)
def step(self):
""" Used in nonstationary version
"""
pass
def reset(self):
""" Used in nonstationary version
"""
# +
class AbstractAgent(metaclass=ABCMeta):
def init_actions(self, n_actions):
self._successes = np.zeros(n_actions)
self._failures = np.zeros(n_actions)
self._total_pulls = 0
@abstractmethod
def get_action(self):
"""
Get current best action
:rtype: int
"""
pass
def update(self, action, reward):
"""
Observe reward from action and update agent's internal parameters
:type action: int
:type reward: int
"""
self._total_pulls += 1
if reward == 1:
self._successes[action] += 1
else:
self._failures[action] += 1
@property
def name(self):
return self.__class__.__name__
class RandomAgent(AbstractAgent):
def get_action(self):
return np.random.randint(0, len(self._successes))
# -
# ### Epsilon-greedy agent
#
# **for** $t = 1,2,...$ **do**
#
# **for** $k = 1,...,K$ **do**
#
# $\hat\theta_k \leftarrow \alpha_k / (\alpha_k + \beta_k)$
#
# **end for**
#
# $x_t \leftarrow argmax_{k}\hat\theta$ with probability $1 - \epsilon$ or random action with probability $\epsilon$
#
# Apply $x_t$ and observe $r_t$
#
# $(\alpha_{x_t}, \beta_{x_t}) \leftarrow (\alpha_{x_t}, \beta_{x_t}) + (r_t, 1-r_t)$
#
# **end for**
#
# Implement the algorithm above in the cell below:
class EpsilonGreedyAgent(AbstractAgent):
def __init__(self, epsilon=0.01):
self._epsilon = epsilon
def get_action(self):
<YOUR CODE>
@property
def name(self):
return self.__class__.__name__ + "(epsilon={})".format(self._epsilon)
# ### UCB Agent
# Epsilon-greedy strategy heve no preference for actions. It would be better to select among actions that are uncertain or have potential to be optimal. One can come up with idea of index for each action that represents otimality and uncertainty at the same time. One efficient way to do it is to use UCB1 algorithm:
#
# **for** $t = 1,2,...$ **do**
#
# **for** $k = 1,...,K$ **do**
#
# $w_k \leftarrow \alpha_k / (\alpha_k + \beta_k) + \sqrt{2log\ t \ / \ (\alpha_k + \beta_k)}$
#
# **end for**
#
# **end for**
# $x_t \leftarrow argmax_{k}w$
#
# Apply $x_t$ and observe $r_t$
#
# $(\alpha_{x_t}, \beta_{x_t}) \leftarrow (\alpha_{x_t}, \beta_{x_t}) + (r_t, 1-r_t)$
#
# **end for**
#
# __Note:__ in practice, one can multiply $\sqrt{2log\ t \ / \ (\alpha_k + \beta_k)}$ by some tunable parameter to regulate agent's optimism and wilingness to abandon non-promising actions.
#
# More versions and optimality analysis - https://homes.di.unimi.it/~cesabian/Pubblicazioni/ml-02.pdf
class UCBAgent(AbstractAgent):
def get_action(self):
<YOUR CODE>
# ### Thompson sampling
#
# UCB1 algorithm does not take into account actual distribution of rewards. If we know the distribution - we can do much better by using Thompson sampling:
#
# **for** $t = 1,2,...$ **do**
#
# **for** $k = 1,...,K$ **do**
#
# Sample $\hat\theta_k \sim beta(\alpha_k, \beta_k)$
#
# **end for**
#
# $x_t \leftarrow argmax_{k}\hat\theta$
#
# Apply $x_t$ and observe $r_t$
#
# $(\alpha_{x_t}, \beta_{x_t}) \leftarrow (\alpha_{x_t}, \beta_{x_t}) + (r_t, 1-r_t)$
#
# **end for**
#
#
# More on Thompson Sampling:
# https://web.stanford.edu/~bvr/pubs/TS_Tutorial.pdf
class ThompsonSamplingAgent(AbstractAgent):
def get_action(self):
<YOUR CODE>
# +
from collections import OrderedDict
def get_regret(env, agents, n_steps=5000, n_trials=50):
scores = OrderedDict({
agent.name: [0.0 for step in range(n_steps)] for agent in agents
})
for trial in range(n_trials):
env.reset()
for a in agents:
a.init_actions(env.action_count)
for i in range(n_steps):
optimal_reward = env.optimal_reward()
for agent in agents:
action = agent.get_action()
reward = env.pull(action)
agent.update(action, reward)
scores[agent.name][i] += optimal_reward - reward
env.step() # change bandit's state if it is unstationary
for agent in agents:
scores[agent.name] = np.cumsum(scores[agent.name]) / n_trials
return scores
def plot_regret(agents, scores):
for agent in agents:
plt.plot(scores[agent.name])
plt.legend([agent.name for agent in agents])
plt.ylabel("regret")
plt.xlabel("steps")
plt.show()
# +
# Uncomment agents
agents = [
# EpsilonGreedyAgent(),
# UCBAgent(),
# ThompsonSamplingAgent()
]
regret = get_regret(BernoulliBandit(), agents, n_steps=10000, n_trials=10)
plot_regret(agents, regret)
# -
# # Bonus 1.1. Gittins index (5 points).
#
# Bernoulli bandit problem has an optimal solution - Gittins index algorithm. Implement finite horizon version of the algorithm and demonstrate it's performance with experiments. some articles:
# - Wikipedia article - https://en.wikipedia.org/wiki/Gittins_index
# - Different algorithms for index computation - http://www.ece.mcgill.ca/~amahaj1/projects/bandits/book/2013-bandit-computations.pdf (see "Bernoulli" section)
#
# # HW 1.1. Nonstationary Bernoulli bandit
#
# What if success probabilities change over time? Here is an example of such bandit:
class DriftingBandit(BernoulliBandit):
def __init__(self, n_actions=5, gamma=0.01):
"""
Idea from https://github.com/iosband/ts_tutorial
"""
super().__init__(n_actions)
self._gamma = gamma
self._successes = None
self._failures = None
self._steps = 0
self.reset()
def reset(self):
self._successes = np.zeros(self.action_count) + 1.0
self._failures = np.zeros(self.action_count) + 1.0
self._steps = 0
def step(self):
action = np.random.randint(self.action_count)
reward = self.pull(action)
self._step(action, reward)
def _step(self, action, reward):
self._successes = self._successes * (1 - self._gamma) + self._gamma
self._failures = self._failures * (1 - self._gamma) + self._gamma
self._steps += 1
self._successes[action] += reward
self._failures[action] += 1.0 - reward
self._probs = np.random.beta(self._successes, self._failures)
# And a picture how it's reward probabilities change over time
# +
drifting_env = DriftingBandit(n_actions=5)
drifting_probs = []
for i in range(20000):
drifting_env.step()
drifting_probs.append(drifting_env._probs)
plt.figure(figsize=(17, 8))
plt.plot(pandas.DataFrame(drifting_probs).rolling(window=20).mean())
plt.xlabel("steps")
plt.ylabel("Success probability")
plt.title("Reward probabilities over time")
plt.legend(["Action {}".format(i) for i in range(drifting_env.action_count)])
plt.show()
# -
# Your task is to invent an agent that will have better regret than stationary agents from above.
# +
# YOUR AGENT HERE SECTION
# +
drifting_agents = [
ThompsonSamplingAgent(),
EpsilonGreedyAgent(),
UCBAgent(),
YourAgent()
]
plot_regret(DriftingBandit(), drifting_agents, n_steps=20000, n_trials=10)
# -
# ## Part 2. Contextual bandit
#
# Now we will solve much more complex problem - reward will depend on bandit's state.
#
# **Real-word analogy:**
#
# > Contextual advertising. We have a lot of banners and a lot of different users. Users can have different features: age, gender, search requests. We want to show banner with highest click probability.
#
# If we want use strategies from above, we need some how store reward distributions conditioned both on actions and bandit's state.
# One way to do this - use bayesian neural networks. Instead of giving pointwise estimates of target, they maintain probability distributions
#
# <img src="bnn.png">
# Picture from https://arxiv.org/pdf/1505.05424.pdf
#
#
# More material:
# * A post on the matter - [url](http://twiecki.github.io/blog/2016/07/05/bayesian-deep-learning/)
# * Theano+PyMC3 for more serious stuff - [url](http://pymc-devs.github.io/pymc3/notebooks/bayesian_neural_network_advi.html)
# * Same stuff in tensorflow - [url](http://edwardlib.org/tutorials/bayesian-neural-network)
#
# Let's load our dataset:
# +
all_states = np.load("all_states.npy")
action_rewards = np.load("action_rewards.npy")
state_size = all_states.shape[1]
n_actions = action_rewards.shape[1]
print("State size: %i, actions: %i" % (state_size, n_actions))
# +
import theano
import theano.tensor as T
import lasagne
from lasagne import init
from lasagne.layers import *
import bayes
as_bayesian = bayes.bbpwrap(bayes.NormalApproximation(std=0.1))
BayesDenseLayer = as_bayesian(DenseLayer)
# -
# ## 2.1 Bulding a BNN agent
#
# Let's implement epsilon-greedy BNN agent
class BNNAgent:
"""a bandit with bayesian neural net"""
def __init__(self, state_size, n_actions):
input_states = T.matrix("states")
target_actions = T.ivector("actions taken")
target_rewards = T.vector("rewards")
self.total_samples_seen = theano.shared(
np.int32(0), "number of training samples seen so far")
batch_size = target_actions.shape[0] # por que?
# Network
inp = InputLayer((None, state_size), name='input')
# YOUR NETWORK HERE
out = <YOUR CODE: Your network>
# Prediction
prediction_all_actions = get_output(out, inputs=input_states)
self.predict_sample_rewards = theano.function(
[input_states], prediction_all_actions)
# Training
# select prediction for target action
prediction_target_actions = prediction_all_actions[T.arange(
batch_size), target_actions]
# loss = negative log-likelihood (mse) + KL
negative_llh = T.sum((prediction_target_actions - target_rewards)**2)
kl = bayes.get_var_cost(out) / (self.total_samples_seen+batch_size)
loss = (negative_llh + kl)/batch_size
self.weights = get_all_params(out, trainable=True)
self.out = out
# gradient descent
updates = lasagne.updates.adam(loss, self.weights)
# update counts
updates[self.total_samples_seen] = self.total_samples_seen + \
batch_size.astype('int32')
self.train_step = theano.function([input_states, target_actions, target_rewards],
[negative_llh, kl],
updates=updates,
allow_input_downcast=True)
def sample_prediction(self, states, n_samples=1):
"""Samples n_samples predictions for rewards,
:returns: tensor [n_samples, state_i, action_i]
"""
assert states.ndim == 2, "states must be 2-dimensional"
return np.stack([self.predict_sample_rewards(states) for _ in range(n_samples)])
epsilon = 0.25
def get_action(self, states):
"""
Picks action by
- with p=1-epsilon, taking argmax of average rewards
- with p=epsilon, taking random action
This is exactly e-greedy policy.
"""
reward_samples = self.sample_prediction(states, n_samples=100)
# ^-- samples for rewards, shape = [n_samples,n_states,n_actions]
best_actions = reward_samples.mean(axis=0).argmax(axis=-1)
# ^-- we take mean over samples to compute expectation, then pick best action with argmax
<YOUR CODE>
chosen_actions = <YOUR CODE: implement epsilon-greedy strategy>
return chosen_actions
def train(self, states, actions, rewards, n_iters=10):
"""
trains to predict rewards for chosen actions in given states
"""
loss_sum = kl_sum = 0
for _ in range(n_iters):
loss, kl = self.train_step(states, actions, rewards)
loss_sum += loss
kl_sum += kl
return loss_sum / n_iters, kl_sum / n_iters
@property
def name(self):
return self.__class__.__name__
# ## 2.2 Training the agent
N_ITERS = 100
def get_new_samples(states, action_rewards, batch_size=10):
"""samples random minibatch, emulating new users"""
batch_ix = np.random.randint(0, len(states), batch_size)
return states[batch_ix], action_rewards[batch_ix]
# +
from IPython.display import clear_output
from pandas import DataFrame
moving_average = lambda x, **kw: DataFrame(
{'x': np.asarray(x)}).x.ewm(**kw).mean().values
def train_contextual_agent(agent, batch_size=10, n_iters=100):
rewards_history = []
for i in range(n_iters):
b_states, b_action_rewards = get_new_samples(
all_states, action_rewards, batch_size)
b_actions = agent.get_action(b_states)
b_rewards = b_action_rewards[
np.arange(batch_size), b_actions
]
mse, kl = agent.train(b_states, b_actions, b_rewards, n_iters=100)
rewards_history.append(b_rewards.mean())
if i % 10 == 0:
clear_output(True)
print("iteration #%i\tmean reward=%.3f\tmse=%.3f\tkl=%.3f" %
(i, np.mean(rewards_history[-10:]), mse, kl))
plt.plot(rewards_history)
plt.plot(moving_average(np.array(rewards_history), alpha=0.1))
plt.title("Reward per epesode")
plt.xlabel("Episode")
plt.ylabel("Reward")
plt.show()
samples = agent.sample_prediction(
b_states[:1], n_samples=100).T[:, 0, :]
for i in range(len(samples)):
plt.hist(samples[i], alpha=0.25, label=str(i))
plt.legend(loc='best')
print('Q(s,a) std:', ';'.join(
list(map('{:.3f}'.format, np.std(samples, axis=1)))))
print('correct', b_action_rewards[0].argmax())
plt.title("p(Q(s, a))")
plt.show()
return moving_average(np.array(rewards_history), alpha=0.1)
# -
bnn_agent = BNNAgent(state_size=state_size, n_actions=n_actions)
greedy_agent_rewards = train_contextual_agent(
bnn_agent, batch_size=10, n_iters=N_ITERS)
# ## HW 2.1 Better exploration
#
# Use strategies from first part to gain more reward in contextual setting
class ThompsonBNNAgent(BNNAgent):
def get_action(self, states):
"""
picks action based by taking _one_ sample from BNN and taking action with highest sampled reward (yes, that simple)
This is exactly thompson sampling.
"""
<YOUR CODE>
thompson_agent_rewards = train_contextual_agent(ThompsonBNNAgent(state_size=state_size, n_actions=n_actions),
batch_size=10, n_iters=N_ITERS)
class BayesUCBBNNAgent(BNNAgent):
q = 90
def get_action(self, states):
"""
Compute q-th percentile of rewards P(r|s,a) for all actions
Take actions that have highest percentiles.
This implements bayesian UCB strategy
"""
<YOUR CODE>
ucb_agent_rewards = train_contextual_agent(BayesUCBBNNAgent(state_size=state_size, n_actions=n_actions),
batch_size=10, n_iters=N_ITERS)
# +
plt.figure(figsize=(17, 8))
plt.plot(greedy_agent_rewards)
plt.plot(thompson_agent_rewards)
plt.plot(ucb_agent_rewards)
plt.legend([
"Greedy BNN",
"Thompson sampling BNN",
"UCB BNN"
])
plt.show()
# -
# ## Part 3. Exploration in MDP
#
# The following problem, called "river swim", illustrates importance of exploration in context of mdp's.
# <img src="river_swim.png">
#
# Picture from https://arxiv.org/abs/1306.0940
# Rewards and transition probabilities are unknown to an agent. Optimal policy is to swim against current, while easiest way to gain reward is to go left.
class RiverSwimEnv:
LEFT_REWARD = 5.0 / 1000
RIGHT_REWARD = 1.0
def __init__(self, intermediate_states_count=4, max_steps=16):
self._max_steps = max_steps
self._current_state = None
self._steps = None
self._interm_states = intermediate_states_count
self.reset()
def reset(self):
self._steps = 0
self._current_state = 1
return self._current_state, 0.0, False
@property
def n_actions(self):
return 2
@property
def n_states(self):
return 2 + self._interm_states
def _get_transition_probs(self, action):
if action == 0:
if self._current_state == 0:
return [0, 1.0, 0]
else:
return [1.0, 0, 0]
elif action == 1:
if self._current_state == 0:
return [0, .4, .6]
if self._current_state == self.n_states - 1:
return [.4, .6, 0]
else:
return [.05, .6, .35]
else:
raise RuntumeError(
"Unknown action {}. Max action is {}".format(action, self.n_actions))
def step(self, action):
"""
:param action:
:type action: int
:return: observation, reward, is_done
:rtype: (int, float, bool)
"""
reward = 0.0
if self._steps >= self._max_steps:
return self._current_state, reward, True
transition = np.random.choice(
range(3), p=self._get_transition_probs(action))
if transition == 0:
self._current_state -= 1
elif transition == 1:
pass
else:
self._current_state += 1
if self._current_state == 0:
reward = self.LEFT_REWARD
elif self._current_state == self.n_states - 1:
reward = self.RIGHT_REWARD
self._steps += 1
return self._current_state, reward, False
# Let's implement q-learning agent with epsilon-greedy exploration strategy and see how it performs.
class QLearningAgent:
def __init__(self, n_states, n_actions, lr=0.2, gamma=0.95, epsilon=0.1):
self._gamma = gamma
self._epsilon = epsilon
self._q_matrix = np.zeros((n_states, n_actions))
self._lr = lr
def get_action(self, state):
if np.random.random() < self._epsilon:
return np.random.randint(0, self._q_matrix.shape[1])
else:
return np.argmax(self._q_matrix[state])
def get_q_matrix(self):
""" Used for policy visualization
"""
return self._q_matrix
def start_episode(self):
""" Used in PSRL agent
"""
pass
def update(self, state, action, reward, next_state):
<YOUR CODE>
# Finish implementation of q-learnig agent
def train_mdp_agent(agent, env, n_episodes):
episode_rewards = []
for ep in range(n_episodes):
state, ep_reward, is_done = env.reset()
agent.start_episode()
while not is_done:
action = agent.get_action(state)
next_state, reward, is_done = env.step(action)
agent.update(state, action, reward, next_state)
state = next_state
ep_reward += reward
episode_rewards.append(ep_reward)
return episode_rewards
# +
env = RiverSwimEnv()
agent = QLearningAgent(env.n_states, env.n_actions)
rews = train_mdp_agent(agent, env, 1000)
plt.figure(figsize=(15, 8))
plt.plot(moving_average(np.array(rews), alpha=.1))
plt.xlabel("Episode count")
plt.ylabel("Reward")
plt.show()
# -
# Let's visualize our policy:
def plot_policy(agent):
fig = plt.figure(figsize=(15, 8))
ax = fig.add_subplot(111)
ax.matshow(agent.get_q_matrix().T)
ax.set_yticklabels(['', 'left', 'right'])
plt.xlabel("State")
plt.ylabel("Action")
plt.title("Values of state-action pairs")
plt.show()
plot_policy(agent)
# As your see, agent uses suboptimal policy of going left and does not explore the right state.
# ## Bonus 3.1 Posterior sampling RL (3 points)
# Now we will implement Thompson Sampling for MDP!
#
# General algorithm:
#
# >**for** episode $k = 1,2,...$ **do**
# >> sample $M_k \sim f(\bullet\ |\ H_k)$
#
# >> compute policy $\mu_k$ for $M_k$
#
# >> **for** time $t = 1, 2,...$ **do**
#
# >>> take action $a_t$ from $\mu_k$
#
# >>> observe $r_t$ and $s_{t+1}$
# >>> update $H_k$
#
# >> **end for**
#
# >**end for**
#
# In our case we will model $M_k$ with two matricies: transition and reward. Transition matrix is sampled from dirichlet distribution. Reward matrix is sampled from normal-gamma distribution.
#
# Distributions are updated with bayes rule - see continious distribution section at https://en.wikipedia.org/wiki/Conjugate_prior
#
# Article on PSRL - https://arxiv.org/abs/1306.0940
# +
def sample_normal_gamma(mu, lmbd, alpha, beta):
""" https://en.wikipedia.org/wiki/Normal-gamma_distribution
"""
tau = np.random.gamma(alpha, beta)
mu = np.random.normal(mu, 1.0 / np.sqrt(lmbd * tau))
return mu, tau
class PsrlAgent:
def __init__(self, n_states, n_actions, horizon=10):
self._n_states = n_states
self._n_actions = n_actions
self._horizon = horizon
# params for transition sampling - Dirichlet distribution
self._transition_counts = np.zeros(
(n_states, n_states, n_actions)) + 1.0
# params for reward sampling - Normal-gamma distribution
self._mu_matrix = np.zeros((n_states, n_actions)) + 1.0
self._state_action_counts = np.zeros(
(n_states, n_actions)) + 1.0 # lambda
self._alpha_matrix = np.zeros((n_states, n_actions)) + 1.0
self._beta_matrix = np.zeros((n_states, n_actions)) + 1.0
def _value_iteration(self, transitions, rewards):
# YOU CODE HERE
state_values = <YOUR CODE: find action values with value iteration>
return state_values
def start_episode(self):
# sample new mdp
self._sampled_transitions = np.apply_along_axis(
np.random.dirichlet, 1, self._transition_counts)
sampled_reward_mus, sampled_reward_stds = sample_normal_gamma(
self._mu_matrix,
self._state_action_counts,
self._alpha_matrix,
self._beta_matrix
)
self._sampled_rewards = sampled_reward_mus
self._current_value_function = self._value_iteration(
self._sampled_transitions, self._sampled_rewards)
def get_action(self, state):
return np.argmax(self._sampled_rewards[state] +
self._current_value_function.dot(self._sampled_transitions[state]))
def update(self, state, action, reward, next_state):
<YOUR CODE>
# update rules - https://en.wikipedia.org/wiki/Conjugate_prior
def get_q_matrix(self):
return self._sampled_rewards + self._current_value_function.dot(self._sampled_transitions)
# +
from pandas import DataFrame
moving_average = lambda x, **kw: DataFrame(
{'x': np.asarray(x)}).x.ewm(**kw).mean().values
horizon = 20
env = RiverSwimEnv(max_steps=horizon)
agent = PsrlAgent(env.n_states, env.n_actions, horizon=horizon)
rews = train_mdp_agent(agent, env, 1000)
plt.figure(figsize=(15, 8))
plt.plot(moving_average(np.array(rews), alpha=0.1))
plt.xlabel("Episode count")
plt.ylabel("Reward")
plt.show()
# -
plot_policy(agent)
# ## Bonus 3.2 Bootstrapped DQN (10 points)
#
# Implement Bootstrapped DQN algorithm and compare it's performance with ordinary DQN on BeamRider Atari game. Links:
# - https://arxiv.org/abs/1602.04621
| week05_explore/week5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Automação de Processos com Selenium
# ## Funcionalidades do ChromeDriver
#
# O Webdriver tem por objetivo simular de forma automática todos os passos que o ser humano faria no navegador a fim de realizar determinada tarefa. Podemos localizar os elementos presentes no HTML de um site por meio de seus atributos (XPath, nome da classe, ID, CSS Selector etc.) e interagir com eles. Essas interações incluem:
#
# > Interações diretas, como clicar com o método `.click()`, enviar teclas com o método `send_keys()`
#
# > Extrair o código HTML que constrói aquele objeto na página web, com destaque para a extração de tabelas com o get_attribute("outerHTML") e a função `pd.read_html`
#
# Ao fazer a busca por um elemento na página web, o Selenium retorna um objeto do tipo "WebElement". Ele funciona de forma semelhante a uma lista do Python, contendo cada elemento encontrado que possui o atributo pesquisado, um em cada posição. Podemos acessar o elemento que desejamos pelo índice da lista (Ex: html[0]), para realizar as operações em cima dele.
# ## Processos comuns para utilização do Webdriver:
# #### Importar Bibliotecas
#
# > **from** selenium **import** webdriver
# >
# > **from** selenium.webdriver.common.keys **import** Keys # permite especificar qual tecla pressionar (Ex: Keys.RETURN)
#
# #### Abrir o ChromeDriver
#
# Antes de tudo, devemos instalar o `chromedriver.exe` e requerir o seu PATH (dada sua versão do Google Chrome), para abrir o aplicativo de teste automatizado.
#
# driver = webdriver.Chrome("chromedriver.exe")
#
# #### Abrir o navegador automatizado
#
# driver.get("link")
#
# #### Incluir pausas
#
# Importante para pausar a execução do código por determinado período, e aguardar o navegador carregar a ação feita anteriormente. Incluímos pausas com o módulo `time`
#
# > **from** time **import** sleep
#
# sleep(segundos)
#
# #### Fechar o navegador automatizado
#
# driver.quit()
# # Exemplos Práticos
# ### Importação dos títulos das notícias da Reuters
from selenium import webdriver
# +
driver = webdriver.Chrome("./chromedriver.exe")
driver.get("https://www.reuters.com/places/brazil")
titulos = driver.find_elements_by_class_name("story-title")
for i in titulos:
print(i.text)
driver.quit()
# -
# ### Importação de dados da B3
from selenium import webdriver
import pandas as pd
import matplotlib.pyplot as plt
# +
driver = webdriver.Chrome("./chromedriver.exe")
driver.get("http://www2.bmf.com.br/pages/portal/bmfbovespa/lumis/lum-taxas-referenciais-bmf-ptBR.asp")
html = driver.find_element_by_xpath('//*[@id="tb_principal1"]').get_attribute("outerHTML")
# Alternativamente: html = driver.find_element_by_class_name('responsive').get_attribute("outerHTML")
df = pd.read_html(html, decimal = ",", thousands = ".")[0]
driver.quit()
# +
df.columns = ["Dias Corridos", "Dólar Ano Útil", "Dólar Ano Corrente"]
df.head()
# -
plt.plot(df["Dias Corridos"], df["Ano Corrente"], label = "Dólar Futuro", color = "green", linewidth = 2)
plt.xlabel("Dias Corridos", fontsize = 12)
plt.title("Dólar Futuro")
plt.show()
# ### Baixando arquivos da web clicando em botão de "Download"
from selenium import webdriver
import pandas as pd
from time import sleep
# +
driver = webdriver.Chrome("./chromedriver.exe")
driver.get("https://sistemaswebb3-listados.b3.com.br/investorProfilePage/range?language=pt-br")
sleep(3)
driver.find_element_by_link_text('Download').click() # Clicar em download
sleep(3)
driver.quit()
# +
df = pd.read_csv("D:\\Downloads\\Perfil Investidores Faixa Etária_Jun2021.csv", sep = "|", encoding = "latin1", skiprows = 2)
df
| selenium/.ipynb_checkpoints/web scraping-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os #### FOR MULTIPLE FEATURES X, SINGLE Y ####
import sys
import tensorflow as tf
W = tf.Variable(tf.zeros([2,1]), name="weights")
b = tf.Variable(0., name="bias")
# +
# ------- define the training loop operations -------
def inference(X): # compute inference model over data X and return the result
return tf.matmul(X, W) + b
def loss(X, Y): # compute loss over training data X and expected outputs Y
Y_predicted = inference(X)
return tf.reduce_sum(tf.squared_difference(Y, Y_predicted))
def inputs(): # read/generate input training data X and expected outputs Y
# Data from http://people.sc.fsu.edu/~jburkardt/datasets/regression/x09.txt
weight_age = [[84, 46], [73, 20], [65, 52], [70, 30], [76, 57], [69, 25], [63, 28], [72, 36], [79, 57],
[75, 44], [27, 24], [89, 31], [65, 52], [57, 23], [59, 60], [69, 48], [60, 34], [79, 51],
[75, 50], [82, 34], [59, 46], [67, 23], [85, 37], [55, 40], [63, 30]]
blood_fat_content = [354, 190, 405, 263, 451, 302, 288, 385, 402, 365, 209, 290, 346, 254, 395, 434, 220,
374, 308, 220, 311, 181, 274, 303, 244]
return tf.to_float(weight_age), tf.to_float(blood_fat_content) # X, Y
def train(total_loss): # train/adjust model parameters according to computed total loss
learning_rate = 0.0000001
return tf.train.GradientDescentOptimizer(learning_rate).minimize(total_loss)
def evaluate(sess, X, Y): # evaluate the resulting trained model
print(sess.run(inference([[80., 25.]]))) # 303
print(sess.run(inference([[65., 25.]]))) # 256
# ---------------------------------------------------
# +
#saver = tf.train.Saver() # by default the saver will keep only the most recent 5 files and delete the rest
#currentDir = 'C:\\Users\\<NAME>\\Desktop\\Tensorflow learning\\my-model'
with tf.Session() as sess:
tf.global_variables_initializer().run()
X, Y = inputs()
#print(sess.run(X)) # USE sess TO PRINT VALUES TO SCREEN
total_loss = loss(X, Y)
train_op = train(total_loss)
coord = tf.train.Coordinator()
threads = tf.train.start_queue_runners(sess=sess, coord=coord)
initial_step = 0
# verify if we don't have a checkpoint saved already
#check_point = tf.train.get_checkpoint_state(currentDir)
#if check_point and check_point.model_checkpoint_path:
# saver.restore(sess, check_point.model_checkpoint_path)
# initial_step = int(check_point.model_checkpoint_path.rsplit('-', 1)[1])
# actual training loop
training_steps = 1000
for step in range(initial_step, training_steps):
sess.run([train_op])
if step % 100 == 0: # for debug purpose, see how the loss gets decremented thru training steps
print("loss: ", sess.run([total_loss]))
#saver.save(sess, currentDir, global_step=step) # creates a checkpoint file with the name template
# my-model-{step} like my-model-1000, my-model-2000...
evaluate(sess, X, Y)
#saver.save(sess, currentDir, global_step=training_steps)
coord.request_stop()
coord.join(threads)
sess.close()
# -
| 03_linearRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import numpy as np
import pandas as pd
import json
import glob
pd.set_option('display.max_rows', 300)
pd.set_option('display.max_columns', 50)
input_filespecs = [
"../../../mnt/lab9-isilon/shared/eaglemonk/examples-data-project-pvc-*/test-logs/*_longevity-test-*.log",
# "../../../tmp/test-longevity-camera-claudio-04.log",
]
records = []
for input_filespec in input_filespecs:
for input_filename in sorted(glob.glob(input_filespec)):
print(input_filename)
with open(input_filename) as file:
for line in file:
m = re.match('{.*', line)
if m:
rec1 = json.loads(line)
rec = rec1['fields']
rec['level'] = rec1['level']
rec['target'] = rec1['target']
records += [rec]
df = pd.DataFrame(records)
if "pts" not in df:
df["pts"] = np.nan
if "prev_pts" not in df:
df["prev_pts"] = np.nan
if "time_delta" not in df:
df["time_delta"] = np.nan
df["min_pts"] = pd.to_datetime(df["min_pts"], errors="coerce")
df["max_pts"] = pd.to_datetime(df["max_pts"], errors="coerce")
df["pts"] = pd.to_datetime(df["pts"], errors="coerce")
df["prev_pts"] = pd.to_datetime(df["prev_pts"], errors="coerce")
df["ms_from_prev"] = df.pts.diff() / pd.Timedelta(1, "millisecond")
df.info()
df.head(4)
stats = df[df.description.isin(["statistics"])].groupby(["stream", "probe_name"]).last()
pd.Timestamp.utcnow() - stats[["min_pts", "max_pts"]]
stats[(stats.pts_missing_count>0) |
(stats.pts_decreasing_count>0) |
(stats.corrupted_count>0) |
(stats.discontinuity_count>100) |
(stats.idle_count>5)
].T
df[df.description.isin(["statistics"])].groupby(["stream", "probe_name"]).last().T
df.groupby(["stream", "description"]).size()
df[df.level.isin(["WARN","DEBUG"])].groupby(["stream", "description", "probe_name"]).size()
df[df.description.isin(["Gap in PTS is too large"])][["time_delta", "probe_name"]].value_counts().sort_index()
df[df.description.isin(["PTS is missing"])][["pts", "time_delta", "probe_name"]]
df[df.description.isin(["PTS is decreasing"])][["pts", "time_delta", "probe_name"]]
df.iloc[300:400][df.probe_name.isin(["1-pravegasrc"])][["pts", "ms_from_prev", "prev_pts", "time_delta", "probe_name", "flags", "size", "description"]]
df2 = df[df.description.isin(["buffer"]) & df.probe_name.isin(["1-pravegasrc"])].copy()
df2["ms_from_prev"] = df2.pts.diff() / pd.Timedelta(1, "millisecond")
df2[["pts", "ms_from_prev", "element", "pad", "flags", "size", "description"]].head(200)
df2.ms_from_prev.value_counts()
df2 = df[df.description.isin(["buffer"]) & ((df.element=="pravegasrc") | ((df.element=="h264parse") & (df["pad"]=="sink")))].copy()
df2["ms_from_prev"] = df2.pts.diff() / pd.Timedelta(1, "millisecond")
df2[["pts", "ms_from_prev", "element", "pad", "flags", "size", "description"]].head(200)
df2[df2.element=="h264parse"].ms_from_prev.value_counts()
df2.ms_from_prev.value_counts()
| jupyter/notebooks/longevity_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yukinaga/ai_programming/blob/main/lecture_06/01_ml_libraries.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="6gpLdgYqPxp5"
# # 機械学習ライブラリ
# 機械学習ライブラリ、KerasとPyTorchのコードを紹介します。
# 今回はコードの詳しい解説は行いませんが、実装の大まかな流れを把握しましょう。
#
# + [markdown] id="WguhKCRZdgV_"
# ## ● Kerasのコード
# 以下のコードは、Kerasによるシンプルなニューラルネットワークの実装です。
# Irisの各花を、SetosaとVersicolorに分類します。
# 以下のコードでは、`Sequential`でモデルを作り、層や活性化関数を追加しています。
# + id="7uN-2AkcPCa5"
import numpy as np
from sklearn import datasets
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
iris = datasets.load_iris()
iris_data = iris.data
sl_data = iris_data[:100, 0] # SetosaとVersicolor、Sepal length
sw_data = iris_data[:100, 1] # SetosaとVersicolor、Sepal width
# 平均値を0に
sl_ave = np.average(sl_data) # 平均値
sl_data -= sl_ave # 平均値を引く
sw_ave = np.average(sw_data)
sw_data -= sw_ave
# 入力をリストに格納
input_data = []
correct_data = []
for i in range(100):
input_data.append([sl_data[i], sw_data[i]])
correct_data.append([iris.target[i]])
# 訓練データとテストデータに分割
input_data = np.array(input_data) # NumPyの配列に変換
correct_data = np.array(correct_data)
x_train, x_test, t_train, t_test = train_test_split(input_data, correct_data)
# ------ ここからKerasのコード ------
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras.optimizers import SGD
model = Sequential()
model.add(Dense(2, input_dim=2)) # 入力:2、中間層のニューロン数:2
model.add(Activation("sigmoid")) # シグモイド関数
model.add(Dense(1)) # 出力層のニューロン数:1
model.add(Activation("sigmoid")) # シグモイド関数
model.compile(optimizer=SGD(lr=0.3), loss="mean_squared_error", metrics=["accuracy"])
model.fit(x_train, t_train, epochs=32, batch_size=1) # 訓練
loss, accuracy = model.evaluate(x_test, t_test)
print("正解率: " + str(accuracy*100) + "%")
# + [markdown] id="J-gMuglhk1k9"
# ## ● PyTorchのコード
# 以下のコードは、PyTorchよるシンプルなニューラルネットワークの実装です。
# Irisの各花を、SetosaとVersicolorに分類します。
# 以下のコードでは、Kerasと同様に`Sequential`でモデルを作り、層や活性化関数を並べています。
# PyTorchでは、入力や正解をTensor形式のデータに変換する必要があります。
# + id="djZH-6V39iq8"
import numpy as np
from sklearn import datasets
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
iris = datasets.load_iris()
iris_data = iris.data
sl_data = iris_data[:100, 0] # SetosaとVersicolor、Sepal length
sw_data = iris_data[:100, 1] # SetosaとVersicolor、Sepal width
# 平均値を0に
sl_ave = np.average(sl_data) # 平均値
sl_data -= sl_ave # 平均値を引く
sw_ave = np.average(sw_data)
sw_data -= sw_ave
# 入力をリストに格納
input_data = []
correct_data = []
for i in range(100):
input_data.append([sl_data[i], sw_data[i]])
correct_data.append([iris.target[i]])
# 訓練データとテストデータに分割
input_data = np.array(input_data) # NumPyの配列に変換
correct_data = np.array(correct_data)
x_train, x_test, t_train, t_test = train_test_split(input_data, correct_data)
# ------ ここからPyTorchのコード ------
import torch
from torch import nn
from torch import optim
# Tensorに変換
x_train = torch.tensor(x_train, dtype=torch.float32)
t_train = torch.tensor(t_train, dtype=torch.float32)
x_test = torch.tensor(x_test, dtype=torch.float32)
t_test = torch.tensor(t_test, dtype=torch.float32)
net = nn.Sequential(
nn.Linear(2, 2), # 入力:2、中間層のニューロン数:2
nn.Sigmoid(), # シグモイド関数
nn.Linear(2, 1), # 出力層のニューロン数:1
nn.Sigmoid() # シグモイド関数
)
loss_fnc = nn.MSELoss()
optimizer = optim.SGD(net.parameters(), lr=0.3)
# 1000エポック学習
for i in range(1000):
# 勾配を0に
optimizer.zero_grad()
# 順伝播
y_train = net(x_train)
y_test = net(x_test)
# 誤差を求める
loss_train = loss_fnc(y_train, t_train)
loss_test = loss_fnc(y_test, t_test)
# 逆伝播(勾配を求める)
loss_train.backward()
# パラメータの更新
optimizer.step()
if i%100 == 0:
print("Epoch:", i, "Loss_Train:", loss_train.item(), "Loss_Test:", loss_test.item())
y_test = net(x_test)
count = ((y_test.detach().numpy()>0.5) == (t_test.detach().numpy()==1.0)).sum().item()
print("正解率: " + str(count/len(y_test)*100) + "%")
| lecture_06/01_ml_libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://medium.com/@anilbak69/5-python-pandas-tricks-to-make-data-analysis-7151b8fa0968
import pandas as pd
# part 1, generate and save the chart you'll use in the report
chart_fig = ("test_scatter.png")
df = pd.util.testing.makeDataFrame()
fig = df.plot(x='A', y='B', kind='scatter').get_figure()
fig.savefig(chart_fig)
# +
# part 2: set up a page template with desired properties
import os
from reportlab.platypus import SimpleDocTemplate, Paragraph, Spacer, PageBreak, Image
from reportlab.lib.styles import ParagraphStyle
from reportlab.rl_config import defaultPageSize
from reportlab.lib.units import inch
def page_template(canvas, doc):
'''
End Goal document:
8.5 x 11" sheet
1" margins all around
Content on page:
Justified paragraph describing the project
Centered chart
Another justified paragraph describing the chart
Left-flushed footer with author's name and report title
right-flushed footer with page number
'''
# save the current state before this function makes its changes state changes (e.g. change
# font, line styles, etc.)
canvas.saveState()
# define pg dimensions
pg_height = 11 * inch
pg_width = 8.5 * inch
# set up new doc object
out_pdf_name = "Pg_w_chart.pdf"
doc = SimpleDocTemplate(out_pdf_name)
Story = []
title_txt = "Example PDF Chart Report!"
# Paragraph(text, style, bulletText=None)
# alignment params: 0=left flush, 1=centered, 2=right flush
title_style = ParagraphStyle("Title", fontSize=18, spaceAfter=1 * inch, alignment=1)
title = Paragraph(title_txt, title_style)
Story.append(title)
# Story.append(Spacer(1, 1 * inch))
img_obj = Image(chart_fig)
Story.append(img_obj)
doc.build(Story, onFirstPage=page_template, onLaterPages=page_template)
print("created file {}".format(os.path.join(os.getcwd(), out_pdf_name)))
# -
import reportlab
print(img_obj.hAlign)
| MakePDFReport/sample_plots_to_PDF_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: yggJLNE
# language: python
# name: yggjlne
# ---
# # Example - Matplotlib Integration
# ## Import libraries and setup node_editor
# %matplotlib inline
import jupyterlab_nodeeditor as jlne
from yggdrasil import yamlfile
import matplotlib.pyplot as plt
import ipywidgets as widgets
import numpy as np
# +
# Setting up the JLNE instance
filename = "X:\College\Grad School\Research\gc-Xyzic\jupyterlab_nodeeditor\examples\model_example.yml"
model_set = yamlfile.parse_yaml(filename, model_only=True)
model_sets = jlne.parse_yggdrasil_yaml(filename)
schema = yamlfile.get_schema()
socket_types = tuple(schema.form_schema['definitions']['schema']['definitions']['simpleTypes']['enum'])
ne = jlne.NodeEditor(socket_types = socket_types)
# -
ne.socket_types = ('Temperature', 'Rainfall', 'Delta Time', 'Results')
ne.add_component(
{"inputs": [
{'title': 'Temperature Morning', 'key': 'temp1', 'socket_type': 'Temperature'},
{'title': 'Temperature Afternoon', 'key': 'temp2', 'socket_type': 'Temperature'},
{'title': 'Temperature Evening', 'key': 'temp3', 'socket_type': 'Temperature'}
],
"outputs": [
{'title': 'Results', 'key': 'results', 'socket_type': 'Temperature'}
],
"title": "Temperature Averaging"
})
# ## Display the node editor
# #### *Optional - Right click the cell and select "Create new view for output" to see it in a different tab*
# Make sure to add in a node BEFORE continuing
ne
# Setup and add the Slope and Y-Intercept Sliders
m = widgets.FloatSlider(value = 0.0, min = -2.0, max = 2.0, step = 0.1, description = "Slope")
b = widgets.FloatSlider(value = 5.0, min = 0.0, max = 10.0, step = 0.1, description = "Intercept")
ne.node_editor.nodes[0].display_element.children += (m,)
ne.node_editor.nodes[0].display_element.children += (b,)
# Now you can see the sliders on the node editor instance.
#
# This next cell will add in the graph that you can play around with.
# +
# Open initial sample plot png and add it to the sidebar
with open("test.png", "rb") as f:
image = f.read()
plotimg = widgets.Image(value = image, format = "png")
ne.node_editor.nodes[0].display_element.children += (plotimg,)
# -
# The sliders won't work until this next cell is run.
#
# The comments explain what each portion does with ipywidgets.
# +
# Use widgets to let the sliders adjust the plot
def plot_change(change):
# Create the graph
fig, ax = plt.subplots()
ax.set(xlabel='x', ylabel='y', title = "Simple Test Graph")
ax.grid()
x = np.linspace(-10, 10, num=1000)
ax.set_ylim(0, 10)
# Plot the line
ax.plot(x, m.value * x + b.value)
# Save it as an image
fig.savefig("slope.png")
# Display the image
with open("slope.png", "rb") as f:
new_plot = f.read()
plotimg.value = new_plot
# Now have the sliders change the plot
m.observe(plot_change, "value")
b.observe(plot_change, "value")
# -
# For now we have to save/load a png due to %matplotlib widget having some issues.
#
# It will probably be resolved and the example will be updated accordingly in the future.
#
| examples/example_matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Re-Purposing a Pretrained Network
#
# Since a large CNN is very time-consuming to train (even on a GPU), and requires huge amounts of data, is there any way to use a pre-calculated one instead of retraining the whole thing from scratch?
#
# This notebook shows how this can be done. And it works surprisingly well.
#
#
# ## How do we classify images with untrained classes?
#
# This notebook extracts a vector representation of a set of images using the GoogLeNet CNN pretrained on ImageNet. It then builds a 'simple SVM classifier', allowing new images can be classified directly. No retraining of the original CNN is required.
# +
import tensorflow as tf
import numpy as np
import scipy
import matplotlib.pyplot as plt
# %matplotlib inline
import time
from urllib.request import urlopen # Python 3+ version (instead of urllib2)
CLASS_DIR='./images/cars'
#CLASS_DIR='./images/seefood' # for HotDog vs NotHotDog
# -
# ### Add TensorFlow Slim Model Zoo to path
# +
import os, sys
slim_models_dir = './models/tensorflow_zoo'
if not os.path.exists(slim_models_dir):
print("Creating model/tensorflow_zoo directory")
os.makedirs(slim_models_dir)
if not os.path.isfile( os.path.join(slim_models_dir, 'models', 'README.md') ):
print("Cloning tensorflow model zoo under %s" % (slim_models_dir, ))
# !cd {slim_models_dir}; git clone https://github.com/tensorflow/models.git
sys.path.append(slim_models_dir + "/models/slim")
print("Model Zoo model code installed")
# -
# ### The Inception v1 (GoogLeNet) Architecture|
#
# 
# ### Download the Inception V1 checkpoint¶
#
# Functions for building the GoogLeNet model with TensorFlow / slim and preprocessing the images are defined in ```model.inception_v1_tf``` - which was downloaded from the TensorFlow / slim [Model Zoo](https://github.com/tensorflow/models/tree/master/slim).
#
# The actual code for the ```slim``` model will be <a href="model/tensorflow_zoo/models/slim/nets/inception_v1.py" target=_blank>here</a>.
# +
from datasets import dataset_utils
targz = "inception_v1_2016_08_28.tar.gz"
url = "http://download.tensorflow.org/models/"+targz
checkpoints_dir = './data/tensorflow_zoo/checkpoints'
if not os.path.exists(checkpoints_dir):
os.makedirs(checkpoints_dir)
if not os.path.isfile( os.path.join(checkpoints_dir, 'inception_v1.ckpt') ):
tarfilepath = os.path.join(checkpoints_dir, targz)
if os.path.isfile(tarfilepath):
import tarfile
tarfile.open(tarfilepath, 'r:gz').extractall(checkpoints_dir)
else:
dataset_utils.download_and_uncompress_tarball(url, checkpoints_dir)
# Get rid of tarfile source (the checkpoint itself will remain)
os.unlink(tarfilepath)
print("Checkpoint available locally")
# -
# Build the model and select layers we need - the features are taken from the final network layer, before the softmax nonlinearity.
# +
slim = tf.contrib.slim
from nets import inception
from preprocessing import inception_preprocessing
image_size = inception.inception_v1.default_image_size
image_size
# +
imagenet_labels_file = './data/imagenet_synset_words.txt'
if os.path.isfile(imagenet_labels_file):
print("Loading ImageNet synset data locally")
with open(imagenet_labels_file, 'r') as f:
imagenet_labels = {0: 'background'}
for i, line in enumerate(f.readlines()):
# n01440764 tench, Tinca tinca
synset,human = line.strip().split(' ', 1)
imagenet_labels[ i+1 ] = human
else:
print("Downloading ImageNet synset data from repo")
from datasets import imagenet
imagenet_labels = imagenet.create_readable_names_for_imagenet_labels()
print("ImageNet synset labels available")
# +
tf.reset_default_graph()
# This creates an image 'placeholder'
# input_image = tf.image.decode_jpeg(image_string, channels=3)
input_image = tf.placeholder(tf.uint8, shape=[None, None, 3], name='input_image')
# Define the pre-processing chain within the graph - based on the input 'image' above
processed_image = inception_preprocessing.preprocess_image(input_image, image_size, image_size, is_training=False)
processed_images = tf.expand_dims(processed_image, 0)
# Reverse out some of the transforms, so we can see the area/scaling of the inception input
numpyish_image = tf.multiply(processed_image, 0.5)
numpyish_image = tf.add(numpyish_image, 0.5)
numpyish_image = tf.multiply(numpyish_image, 255.0)
# Create the model - which uses the above pre-processing on image
# it also uses the default arg scope to configure the batch norm parameters.
print("Model builder starting")
# Here is the actual model zoo model being instantiated :
with slim.arg_scope(inception.inception_v1_arg_scope()):
logits, _ = inception.inception_v1(processed_images, num_classes=1001, is_training=False)
probabilities = tf.nn.softmax(logits)
# Create an operation that loads the pre-trained model from the checkpoint
init_fn = slim.assign_from_checkpoint_fn(
os.path.join(checkpoints_dir, 'inception_v1.ckpt'),
slim.get_model_variables('InceptionV1')
)
print("Model defined")
# -
# ### Display the network layout graph on TensorBoard
#
# This isn't very informative, since the inception graph is pretty complex...
# +
#writer = tf.summary.FileWriter(logdir='../tensorflow.logdir/', graph=tf.get_default_graph())
#writer.flush()
# -
# ### Load an Example Image
#
# Pull in an image into a numpy object :
if False:
# Read from the Web
from io import BytesIO
url = 'https://upload.wikimedia.org/wikipedia/commons/7/70/EnglishCockerSpaniel_simon.jpg'
image_string = urlopen(url).read()
im = plt.imread(BytesIO(image_string), format='jpg')
if False:
# Read from a file via a queue ==> brain damage in jupyter
#filename_queue = tf.train.string_input_producer( tf.train.match_filenames_once("./images/*.jpg") )
filename_queue = tf.train.string_input_producer( ['./images/cat-with-tongue_224x224.jpg'] )
#_ = filename_queue.dequeue() # Ditch the first value
image_reader = tf.WholeFileReader()
_, image_string = image_reader.read(filename_queue)
# Read from a file
im = plt.imread("./images/cat-with-tongue_224x224.jpg")
print(im.shape, im[0,0]) # (height, width, channels), (uint8, uint8, uint8)
def crop_middle_square_area(np_image):
h, w, _ = np_image.shape
h = int(h/2)
w = int(w/2)
if h>w:
return np_image[ h-w:h+w, : ]
return np_image[ :, w-h:w+h ]
im_sq = crop_middle_square_area(im)
im_sq.shape
# ### Run using the Example Image
#
# Let's verify that GoogLeNet / Inception-v1 and our preprocessing are functioning properly :
# +
# Now let's run the pre-trained model
with tf.Session() as sess:
# This is the loader 'op' we defined above
init_fn(sess)
# This is two ops : one merely loads the image from numpy,
# the other runs the network to get the class probabilities
np_image, np_probs = sess.run([numpyish_image, probabilities], feed_dict={input_image:im_sq})
# These are regular numpy operations
probs = np_probs[0, :]
sorted_inds = [i[0] for i in sorted(enumerate(-probs), key=lambda x:x[1])]
# And now plot out the results
plt.figure()
plt.imshow(np_image.astype(np.uint8))
plt.axis('off')
plt.show()
for i in range(5):
index = sorted_inds[i]
print('Probability %0.2f%% => [%s]' % (probs[index], imagenet_labels[index]))
# -
# ----------
# ## Use the Network to create 'features' for the training images
#
# Now go through the input images and feature-ize them at the 'logit level' according to the pretrained network.
#
# <!-- [Logits vs the softmax probabilities](images/presentation/softmax-layer-generic_676x327.png) !-->
#
# 
#
# NB: The pretraining was done on ImageNet - there wasn't anything specific to the recognition task we're doing here.
import os
classes = sorted( [ d for d in os.listdir(CLASS_DIR) if os.path.isdir("%s/%s" % (CLASS_DIR, d)) ] )
classes # Sorted for for consistency
# +
train = dict(filepath=[], features=[], target=[])
with tf.Session() as sess:
# This is the loader 'op' we defined above
init_fn(sess)
print("Loaded pre-trained model")
t0 = time.time()
for class_i, directory in enumerate(classes):
for filename in os.listdir("%s/%s" % (CLASS_DIR, directory, )):
filepath = '%s/%s/%s' % (CLASS_DIR, directory, filename, )
if os.path.isdir(filepath): continue
im = plt.imread(filepath)
im_sq = crop_middle_square_area(im)
# This is two ops : one merely loads the image from numpy,
# the other runs the network to get the 'logit features'
rawim, np_logits = sess.run([numpyish_image, logits], feed_dict={input_image:im_sq})
train['filepath'].append(filepath)
train['features'].append(np_logits[0])
train['target'].append( class_i )
plt.figure()
plt.imshow(rawim.astype('uint8'))
plt.axis('off')
plt.text(320, 50, '{}'.format(filename), fontsize=14)
plt.text(320, 80, 'Train as class "{}"'.format(directory), fontsize=12)
print("DONE : %6.2f seconds each" %(float(time.time() - t0)/len(train),))
# -
# ### Build an SVM model over the features
# +
#train['features'][0]
# -
from sklearn import svm
classifier = svm.LinearSVC()
classifier.fit(train['features'], train['target']) # learn from the data
# ### Use the SVM model to classify the test set
# +
test_image_files = [f for f in os.listdir(CLASS_DIR) if not os.path.isdir("%s/%s" % (CLASS_DIR, f))]
with tf.Session() as sess:
# This is the loader 'op' we defined above
init_fn(sess)
print("Loaded pre-trained model")
t0 = time.time()
for filename in sorted(test_image_files):
im = plt.imread('%s/%s' % (CLASS_DIR,filename,))
im_sq = crop_middle_square_area(im)
# This is two ops : one merely loads the image from numpy,
# the other runs the network to get the class probabilities
rawim, np_logits = sess.run([numpyish_image, logits], feed_dict={input_image:im_sq})
prediction_i = classifier.predict([ np_logits[0] ])
decision = classifier.decision_function([ np_logits[0] ])
plt.figure()
plt.imshow(rawim.astype('uint8'))
plt.axis('off')
prediction = classes[ prediction_i[0] ]
plt.text(350, 50, '{} : Distance from boundary = {:5.2f}'.format(prediction, decision[0]), fontsize=20)
plt.text(350, 75, '{}'.format(filename), fontsize=14)
print("DONE : %6.2f seconds each" %(float(time.time() - t0)/len(test_image_files),))
# -
# ----------------
# ## Exercise : Try your own ideas
#
# The whole training regime here is based on the way the image directories are structured. So building your own example shouldn't be very difficult.
#
# Suppose you wanted to classify pianos into Upright and Grand :
#
# * Create a ```pianos``` directory and point the ```CLASS_DIR``` variable at it
# * Within the ```pianos``` directory, create subdirectories for each of the classes (i.e. ```Upright``` and ```Grand```). The directory names will be used as the class labels
# * Inside the class directories, put a 'bunch' of positive examples of the respective classes - these can be images in any reasonable format, of any size (no smaller than 224x224).
# + The images will be automatically resized so that their smallest dimension is 224, and then a square 'crop' area taken from their centers (since ImageNet networks are typically tuned to answering on 224x224 images)
# * Test images should be put in the ```pianos``` directory itelf (which is logical, since we don't *know* their classes yet)
#
# Finally, re-run everything - checking that the training images are read in correctly, that there are no errors along the way, and that (finally) the class predictions on the test set come out as expected.
#
# If/when it works - please let everyone know : We can add that as an example for next time...
| notebooks/2-CNN/5-TransferLearning/4-ImageClassifier-inception_tf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/schaftler/DESY_FS-TUXS_Project/blob/main/DESY_GS_Optimized_v2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hfNx3g82kfhI"
# # Import libraries
# + id="Gw-S6uzrkfhM"
import numpy as np
from qutip import *
from numpy.linalg import multi_dot
import matplotlib.pyplot as plt
# + id="ld3AjTQM4axm"
#pip install qutip
# + id="0pd4Ob3RkzKr"
# + [markdown] id="k1V_siQNkfhP"
# # Initialize lists of pauli operators
# + id="G_DUtt0CkfhQ"
def initialize(N):
si = qeye(2)
#sm = np.dot(zero,one.T);
#sp = np.dot(one,zero.T);
sm=sigmap() #Qutip uses inverse notation, please check the end of the notebook
sp=sigmam()
sz=sigmaz()
sz_list = []
sp_list = []
sm_list = []
spsm_list = []
for n in range(N):
op_list = []
for m in range(N):
op_list.append(si)
op_list[n]=sz
sz_list.append(tensor(op_list))
#Create lists of sp,sm operators for KroneckerProduct(Id2,Id2,...Sp,Id2, Sm) operation
op_list[n] = sp
sp_list.append(tensor(op_list))
op_list[n] = Qobj(sm)
sm_list.append(tensor(op_list))
#Create a list of spsm operators like KroneckerProduct(Sp.Sm,Id2), yes there is
op_list[n] = Qobj(np.dot(sp,sm))
spsm_list.append(tensor(op_list))
return sp_list,sm_list,spsm_list,sz_list
# + [markdown] id="qRcj0wXpkfhR"
# # Define interaction rates
# + id="-viqGUP0kfhS"
# Specify an argument such that it is easy to select between "Dicke","General",etc
def interaction_rates(N,general=False,dicke=False,simp=True):
if simp==True:
#When all interaction rates are equal
gamma_i=np.ones(N)
gamma_j=np.ones(N)
#For the Dicke case
if dicke==True:
gamma_i=np.asarray(0.5*np.ones(N))
gamma_j=np.asarray(0.2*np.ones(N))
#For generalized case. Initially using a set of random rates between (0,1),
#but can be done using interatomic distance dependence
if general==True:
gamma_i=np.random.random(N)
gamma_j=np.random.random(N)
return gamma_i,gamma_j
# + [markdown] id="SlTrgxHakfhT"
# # Initialize the state for evaluation, here $|ee..\rangle \langle ee..|$
# + id="y5MfnDh9kfhT"
def init_state(N):
op_list = []
for i in range(N):
op_list.append(basis(2,1)) #Creating a list of states for excited atoms
#op_list[1]=basis(2,0)
ket = tensor(op_list) #Creating a total tensor product state for initally excited atoms
state = ket2dm(ket) #Creating a density matrix for the initial state
return state
# + [markdown] id="Uc3EWCzmkfhV"
# # Calculate orthogonal vectors
# + id="PITmhB_BkfhW"
#Calculate orthogonal vectors
def ovec(state):
temp=state
if type(temp)==qutip.qobj.Qobj:
temp = temp.full()
ovec=temp/np.sqrt(np.tensordot(temp,temp))
return ovec
# + [markdown] id="vcPZ4l1BkfhX"
# # Calculate absolute norm of residue
#
# + id="xtjfrRMLkfhY"
#Calculate absolute norm of residue
def abs_norm(p):
if type(p)==qutip.qobj.Qobj:
p = p.full()
e=np.ndarray.flatten(p)
for i in range(len(e)):
temp = e[i]
if temp < 0:
e[i]=-1*temp
res=np.real(sum(e))
return res
# + id="KP6OysjykfhY"
#Alternate function using inbuilt np.absolute()
def abs_norm(p):
if type(p)==qutip.qobj.Qobj:
p = p.full()
e=np.ndarray.flatten(p)
res=np.real(sum(np.absolute(e)))
return res
# + [markdown] id="h7TdYFRRkfhZ"
# # Calculate residue
# + id="bRHdVT5XkfhZ"
#Calculating the residue
def fn(e,et,state):
if type(state)==qutip.qobj.Qobj:
state = state.full()
for i in range(len(et)):
state = state-np.tensordot(state,et[i])*et[i]
if abs_norm(state)==0:
res=0.0
else:
ovec = state/np.sqrt(np.tensordot(state,state))
e.append(state)
et.append(ovec)
res = abs_norm(state)
state=(Qobj(state).unit()).full()
#print('res=',res)
#print('e=',e)
#print('et=',et)
return res,state
# + [markdown] id="KRscBg4Mkfha"
# # Calculate lindbladian for given state, interaction rates
# + id="hwQS5GkSkfha"
def liouville_gen(N,state,et,e_lis,L_list=[],gamma_i=[],gamma_j=[],num=[],dicke=False,general=False,simp=False):
if simp==True:
gamma_i,gamma_j = interaction_rates(N) #define interatcion rates depending on the model needed
if dicke==True:
gamma_i,gamma_j = interaction_rates(N,dicke=True) #define interatcion rates depending on the model needed
if general==True:
gamma_i,gamma_j = interaction_rates(N,general=True) #define interatcion rates depending on the model needed
#gamma_i=[0.2,0.3,0.4]
#gamma_j=[0.1,0.12,0.221]
sp_list,sm_list,spsm_list,sz_list=initialize(N) #initialize all the operator lists
L_list=[]
L_list.append(state)
L=[] #storing result of Lindbladian operation
num_states=[np.count_nonzero(state)] #count states
#L_list.append(state) #start with the initial state
L_tot=0
for i in range(N):
for j in range(N):
#Solve the Liouville
if i == j:
L.append(gamma_i[i]*(Qobj(multi_dot([sm_list[i],state,sp_list[i]])- 0.5*multi_dot([spsm_list[i],state]) - 0.5*multi_dot([state,spsm_list[i]]))))
#L.append(0.6*(Qobj(multi_dot([sz_list[i],state,sz_list[i]])- 0.5*multi_dot([sz_list[i],sz_list[i],state]) - 0.5*multi_dot([state,sz_list[i],sz_list[i]]))))
else:
L.append(gamma_j[i]*(Qobj(multi_dot([sm_list[j],state,sp_list[i]])-0.5*multi_dot([sp_list[i],sm_list[j],state]) - 0.5*multi_dot([state,sp_list[i],sm_list[j]]))))
#L.append(0.3*(Qobj(multi_dot([sz_list[j],state,sz_list[i]])- 0.5*multi_dot([sz_list[i],sz_list[i],state]) - 0.5*multi_dot([state,sz_list[i],sz_list[i]]))))
#if i == j:
# L.append(gamma_i[i]*(Qobj(2*multi_dot([sm_list[i],state,sp_list[i]])- multi_dot([spsm_list[i],state]) - multi_dot([state,spsm_list[i]]))).unit())
#else:
# L.append(gamma_j[i]*(Qobj(2*multi_dot([sm_list[j],state,sp_list[i]])-multi_dot([sp_list[i],sm_list[j],state]) - multi_dot([state,sp_list[i],sm_list[j]]))).unit())
size=len(L)
for n in range(size):
L_tot=L_tot+L[n] #Summing all the terms in Lindbladian
L_list.append(L_tot)
num_states.append(np.count_nonzero(L_tot))
size2=len(L_list)
for m in range(0,size2):
num_states.append(np.count_nonzero(L_list[m]))
num.append(num_states[1])
#np.real(np.asarray(L_list).full())
res,state = fn(e_lis,et,L_tot)
#np.set_printoptions(precision=3)
#dim=to_super(Qobj(state))
#residue can be used directly for next Lindbladian operation (orthogonal part) and normalize it!
#Also keep max.inter. rate = 1
#self-interactions (check literature)
#setting precision: comparing norms of resulting state and residue n_state/n_norm~1e-9
n_state = abs_norm(state)
precision=res/abs_norm(state)
if precision<1e-5:
res = np.round(res,4)
if res!=0.0:
state = state
#evec = ovec(state)
#et_lis.append(evec)
#et.append(evec)
if simp==True:
liouville_gen(N,state,et,e_lis,num,simp=True)
if dicke==True:
liouville_gen(N,state,et,e_lis,num,dicke=True)
if general==True:
liouville_gen(N,state,et,e_lis,num,general=True)
else:
#print("total no. of states =", np.count_nonzero(L_tot))
print("All states found!")
print("state=",L_tot)
print("res=",res)
print("number of states involved = ",len(num)+1)
print("states=",num)
#print('dimns=',dim)
#print('inner =',inner)
#print('et=',et)
return
# + [markdown] id="eZfGq7UEkfhb"
# # All pieces together
# + id="40lH2tUwkfhb"
def main(N,num=[],et=[],e_lis=[],dicke=False,general=False,simp=False):
initialize(N)
state=init_state(N)
e_lis.append(state)
evec = ovec(state)
#et_lis.append(evec)
et.append(evec)
if simp==True:
liouville_gen(N,state,et,e_lis,L_list=[],gamma_i=[],gamma_j=[],num=[],simp=True)
if dicke==True:
liouville_gen(N,state,et,e_lis,L_list=[],gamma_i=[],gamma_j=[],num=[],dicke=True)
if general==True:
liouville_gen(N,state,et,e_lis,L_list=[],gamma_i=[],gamma_j=[],num=[],general=True)
# + [markdown] id="wp4wmDx0kfhc"
# # Execute
# + id="jsShayHBkfhc" colab={"base_uri": "https://localhost:8080/"} outputId="f5963d74-92c4-4702-ba9d-595fba2dee25"
main(5,general=True)
# + [markdown] id="q3LJlBgDkfhc"
# # Results
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="ABfu6aYukfhd" outputId="ae6dc1f7-056d-43ac-9708-a2fbe1723949"
#Simple case (all_gamma==1)
n_atoms = [2,3,4,5,6,7]
states = [3,4,5,6,7,8]
#Dicke Case (only collective)
n_atoms = [2,3,4,5,6,7]
states = [3,5,7,10,15,18]
#Earlier, when the self interaction rate is low as compared to interatomic int. rate by 1/10 times, the discrepancy in two atoms arises,
#but if we keep it to be \gamma_{ii}=0.1,\gamma_{ij}=0.5, the # states for 3 atoms is fixed but not for 6 atoms.
#but as the rate is fixed to \gamma_{ii}=0.2,\gamma_{ij}=0.5,, the original states are retained.
#states = [4,6,9,12,16,20]
#Dicke Case (local+collective)
n_atoms = [2,3,4,5,6,7]
states = [3,6,9,12,15,20]
#Case with local dephasing, etc.
n_atoms = [2,3,4,5,6,7]
states = [4,6,9,12,16,20]
#Case with collective dephasing, etc.
n_atoms = [2,3,4,5,6,7]
states = [4,6,9,12,16,20]
#Case with (local+collective) dephasing, etc.
n_atoms = [2,3,4,5,6,7]
states = [4,6,9,12,16,20]
#General Case ()
n_atoms = [2,3,4,5,6]
states = [6,20,70,251,930]#excluding the excited state
plt.plot(n_atoms,states)
plt.plot(n_atoms,np.exp(n_atoms),'r')
# + colab={"base_uri": "https://localhost:8080/", "height": 54} id="xW7noHMcoARg" outputId="431cf003-0215-4aab-ad39-6f812f34dbb3"
to_super(Qobj(init_state(4)))
# + [markdown] id="H9mBlMfukfhd"
# # Code Structure
# + [markdown] id="25d5K-hHkfhe"
# ### 1. In the main(*args) function, specify the $\textbf{N}:$ no. of atoms, specify the kind of model that is to be evaluated, ex. for dicke case, just set, $\texttt{dicke=True}$, empty lists for counting the states, and storing the orthogonalized vectors.
#
# ### 2. The main function then initializes the pauli operators required for Lindblad calculation for given $\texttt{N}$, using the $\texttt{initialize(N)}$ function.
#
# ### 3. The $\texttt{init_state(N)}$ function is used to generate the inital state of the system.
#
# ### 4. The $\texttt{e_lis=[]}$ stores the residue at each step, it starts with storing the inital state.
#
# ### 5. $\texttt{et}$ stores the orthogonal vectors calculated at each step using the $\texttt{ovec(state)}$ function. Orthogonal vector corresponding to initial state is stored first.
#
# ### 6. Depending upon the argument provided for the type of model, (ex. for Dicke case, set $\texttt{dicke=True}$), it evaluates the all the involved states until the norm of residue is $0.0$ using the function $\texttt{liouville_gen()}$
#
# + id="qN5FYIp2hFJO"
# + [markdown] id="SeiDh5oAhLPE"
# # Dicke Model
# Ignoring the local processes and check the evolution of number of states.
# $\Gamma_{ii}=\gamma_{c}=0$, and $\gamma_{ij}=\gamma$
# + id="OipAMjO4hBAz"
# + [markdown] id="h32ebENCdrtI"
# # Generating Interaction Rates for general case
# 1. Line Shape:
#
# If only nearest neighbor interactions are considered, then $\gamma_{i,j}, |i-j|=1$
# + colab={"base_uri": "https://localhost:8080/"} id="JMUK4VkzifnQ" outputId="ff02aecf-e986-4952-e9e4-9075b34b6336"
N=4
g_self=[]
g_int=[]
g_s=0.5
g_i=0.2
for i in range(N):
for j in range(N):
if i==j:
g_self.append(g_s)
else:
if np.abs(i-j)==1:
g_int.append(g_i)
else:
g_int.append(0)
print(g_self,g_int)
# + [markdown] id="BOtv7J-MaKIH"
# # For long range interactions the interaction rate should drop off somehow
# + [markdown] id="1_fplcmxnS5u"
# ## 2. Partially filled square
# 
#
# + colab={"base_uri": "https://localhost:8080/"} id="fnqKcs1Fkq0k" outputId="0baed64f-72b4-4666-f674-b6756be483fa"
N=4
if N%2==0:
a=np.zeros(N)
sq=a@a.T
sq
# + id="R87E1Sarvv2K"
| DESY_GS_Optimized_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Stationarity
# We’re handling here a time series. As such, it’s very important the concept of [Stationarity](https://www.investopedia.com/articles/trading/07/stationary.asp). Much of the analysis carried out on financial time series data involves identifying if the series we want to predict is stationary, and if it is not, finding ways to transform it such that it is stationary. A time series is said to be stationary if its mean and variance don’t change over time, this means, it doesn’t show trends or seasonal effects.
import numpy as np
import pandas as pd
# Let's check stationarity by using [Dickey-Fuller test](https://en.wikipedia.org/wiki/Dickey%E2%80%93Fuller_test).
#
# The Dickey Fuller test is one of the most popular statistical tests. It can be used to determine the presence of unit root in the series, and hence help us understand if the series is stationary or not. The null and alternate hypothesis of this test are:
# - **Null Hypothesis**: The series has a unit root (value of a =1)
# - **Alternate Hypothesis**: The series has no unit root.
#
# If we fail to reject the null hypothesis, we can say that the series is non-stationary. This means that the series can be linear or difference stationary.
#
from utils import get_apple_stock, adf_test
apple_stock = get_apple_stock()
adf_test(apple_stock['Close'])
# Let's try differencing
from utils import difference
diffed_series, _ = difference(apple_stock['Close'])
adf_test(diffed_series)
# We can see that our statistic value of -16 is less than the value of -3.431 at 1%. This suggests that we can reject the null hypothesis with a significance level of less than 1% (i.e. a low probability that the result is a statistical fluke). Rejecting the null hypothesis means that the process has no unit root, and in turn that the time series is stationary or does not have time-dependent structure.
from utils import plot_series
plot_series(diffed_series, 'Stationary, diff(n=1), Close price')
# Let's use [Box-Cox test](http://scipy.github.io/devdocs/generated/scipy.stats.boxcox.html) and see if this transformation helps to make the series stationary
from scipy.stats import boxcox
boxcox_series, lmbda = boxcox(apple_stock['Close'])
print('Best lmbda for Box-Cox test:', lmbda)
plot_series(boxcox_series, 'After applying Box-Cox')
adf_test(boxcox_series)
# ### Trend and Seasonality
# A trend can be removed from your time series data (and data in the future) as a data preparation and cleaning exercise. This is common when using statistical methods for time series forecasting, but does not always improve results when using machine learning models. Alternately, a trend can be added, either directly or as a summary, as a new input variable to the supervised learning problem to predict the output variable.
import matplotlib.pyplot as plt
# %matplotlib inline
from statsmodels.tsa.seasonal import seasonal_decompose
from sklearn.linear_model import LinearRegression
X = np.array([i for i in range(0, len(apple_stock))]).reshape(-1, 1)
y = np.array(apple_stock['Close']).reshape(-1, 1)
linear_regressor = LinearRegression()
linear_regressor.fit(X, y)
trend = linear_regressor.predict(X)
fig, ax = plt.subplots(figsize=(15, 6))
plt.plot(y)
plt.plot(trend)
ax.set_title('Apple Close price trend')
ax.legend(['Close price', 'Trend'])
detrended_price = [y[i]-trend[i] for i in range(0, len(apple_stock))]
plot_series(detrended_price)
# There seems to be a parabola in the residuals, suggesting that perhaps a polynomial fit may have done a better job
from numpy import polyfit
degree = 4
X1d = X.flatten()
y1d = y.flatten()
coef = polyfit(X1d, y1d, degree)
print('Coefficients: %s' % coef)
# +
# create curve
curve = list()
for i in range(len(X)):
value = coef[-1]
for d in range(degree):
value += X[i]**(degree-d) * coef[d]
curve.append(value)
# plot curve over original data
plt.subplots(figsize=(15, 6))
plt.plot(y)
plt.plot(curve, color='red', linewidth=2)
# -
detrended_price = np.array([y[i]-curve[i] for i in range(0, len(apple_stock))]).flatten()
plot_series(detrended_price)
adf_test(detrended_price)
# Let's have a look at other plots that might gives as a hint about its trend and seasonality:
from pandas.plotting import lag_plot, autocorrelation_plot
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
fig, ax = plt.subplots(figsize=(15, 6))
lag_plot(apple_stock['Close'], lag=1, ax=ax)
# Let's have a look at the correlation at lag=10
fig, ax = plt.subplots(figsize=(15, 6))
lag_plot(apple_stock['Close'], lag=10, ax=ax)
# We can see a linear pattern that indicates the data is not random, with a positive and strong correlation (autocorrelation is present). Let's remove trend and confirm we get [white noise](https://www.quantstart.com/articles/White-Noise-and-Random-Walks-in-Time-Series-Analysis)
fig, ax = plt.subplots(figsize=(15, 6))
lag_plot(pd.Series(diffed_series), ax=ax)
fig, ax = plt.subplots(figsize=(15, 6))
autocorrelation_plot(apple_stock['Close'][:500], ax=ax)
fig, ax = plt.subplots(figsize=(15, 6))
autocorrelation_plot(pd.Series(diffed_series[:500]), ax=ax)
fig, ax = plt.subplots(figsize=(15, 6))
_ = plot_acf(apple_stock['Close'][:500], ax=ax)
fig, ax = plt.subplots(figsize=(15, 6))
_ = plot_acf(pd.Series(diffed_series[:500]), ax=ax)
# Decomposing the time series
from statsmodels.tsa.seasonal import seasonal_decompose
result = seasonal_decompose(apple_stock['Close'], freq=1)
plot_series(result.observed, title='Observed', figsize=(15, 2))
plot_series(result.trend, title='Trend', figsize=(15, 2))
plot_series(result.seasonal, title='Seasonal', figsize=(15, 2))
# We can confirm this timeseries is not seasonal
plot_series(result.resid, title='Residual', figsize=(15, 2))
| notebooks/stationarity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
#import BurstCube
import matplotlib as mpl
import matplotlib.pyplot as plt
import astropy.units as u
from astropy.time import Time
from gbm.data.phaii import TTE
from gbm.binning.unbinned import bin_by_time
# -
# ## Methods and classes
class Hit(object):
"""
Single interaction with a SiPM
Attributes
----------
time Time of the hits
detector Detector ID
energy Deposited energy
"""
def __init__(self, detector, time, energy):
self.time = time
self.detector = detector
self.energy = energy
def __str__(self):
return "Detector: {} Time: {:.6f} Energy: {:.2e}".format(self.detector, self.time, self.energy)
def __lt__(self, other):
'''
Less than overload for time sorting
'''
return self.time < other.time
# +
from gbm.data.phaii import TTE
from gbm.data.primitives import EventList
class CosimaSim(object):
'''
Contains info associated with a Cosima output file (.sim)
Attributes
----------
tstart Start of simulated time
tstop Stop of simulated time
nthrown Total number of simulated particles
ntrigger Total number of triggered events
hits Collection of all hits (Hit object array)
'''
def __init__(self):
self.tstart = None
self.tstop = None
self.nthrown = None
self.hits = []
@classmethod
def open(cls, filename, detector_dict, time_shift = 0*u.second):
'''
Parameters
----------
filename: str
Path to .sim file. Will be closed on destruction
detector_dict: dict
Convert from a string of x,y location to a given detector ID
Use x and y value exactly as they appear in the input file
time_shift: time Quantity, _optional_, default: 0s
Apply a time shift to the timing in the file
Return
------
obj: CosimaSim object
Warnings
--------
Concatenating files with NF and IN keywords is not supported
'''
obj = cls()
file = open(filename)
in_events = False
while True:
line = file.readline()
if not line:
break
key = line[0:2] # =\n for empty lines
if in_events:
if key == 'EN':
in_events = False
continue
# Time of hit
if key == "TI":
time = float(line.split()[1]) * u.second + time_shift
# Interaction with sensitive detectors
if key != "HT":
continue
# Format
x,y,energy = np.array(line.split(';'))[[1,2,4]]
detector = detector_dict[x.strip()+','+y.strip()]
energy = float(energy)*u.keV
# Add
obj.hits = np.append(obj.hits, Hit(detector, time, energy))
if key == 'TB':
obj.tstart = float(line.split()[1])*u.second + time_shift
elif key == 'SE':
in_events = True
elif key == 'TE':
obj.tstop = float(line.split()[1])*u.second + time_shift
elif key == 'TS':
obj.nthrown = int(line.split()[1])
elif key == 'NF' or key == 'IN':
raise InputError("NF/IN keys found in {} not supported".format(filename))
file.close()
assert obj.tstart is not None, "TB key not found in {}".format(filename)
assert obj.tstop is not None, "TE key not found in {}".format(filename)
assert obj.nthrown is not None, "TS key not found in {}".format(filename)
return obj
def merge(self, sim2, shift = 0*u.second):
'''
Combine CosimaSim object with a shift
Parameters
----------
sim2: CosimaSim
Hits to be merged
shift: time Quantity, _optional_, default: 0s
Apply a time shift to obj2
'''
self.tstart = min(self.tstart, sim2.tstart + shift)
self.tstop = max(self.tstop, sim2.tstop + shift)
self.nthrown += sim2.nthrown
for hit in sim2.hits:
hit.time = hit.time + shift
self.hits = np.append(self.hits, hit)
hits_sort = np.argsort(self.hits)
self.hits = [self.hits[n] for n in hits_sort]
def to_TTE(self, energy_channel_edges, *args, **kwargs):
'''
Convert the hits into a mocked Time-Tagged Event object
Parameters
----------
energy_channel_edges: array
Energy channel definition for discreatization
Additional *args, **kwargs are passed to TTE.from_data()
Return:
-------
tte: TTE object
'''
times = []
energy_channels = []
energy_channel_edges = [e.to(u.keV).value for e in energy_channel_edges]
nchannels = len(energy_channel_edges)-1
for hit in self.hits:
times += [hit.time.to(u.second).value]
energy_channel = np.digitize(hit.energy.to(u.keV).value, energy_channel_edges) - 1
energy_channel = min(nchannels-1, max(0, energy_channel)) #Overflow/underflow
energy_channels += [energy_channel]
evtlist = EventList.from_lists(times_list = times,
pha_list = energy_channels,
chan_lo = energy_channel_edges[:-1],
chan_hi = energy_channel_edges[1:])
tte = TTE.from_data(data = evtlist, *args, **kwargs)
return tte
# -
# ## .sim to TTE
# ### Create object
# +
from gbm.binning.unbinned import bin_by_time
import matplotlib.pyplot as plt
from gbm.plot.lightcurve import Lightcurve
detector_dict = {"4.88500,4.88500":'q0',
"-4.88500,4.88500":'q1',
"-4.88500,-4.88500":'q2',
"4.88500,-4.88500":'q3'}
t0 = 599529605.000 * u.second
sim = CosimaSim.open("sim/signal.inc1.id1.sim", detector_dict)
bkg = CosimaSim.open("sim/bkg.inc1.id1.sim", detector_dict)
sim.merge(bkg, shift=-5*u.second)
n_energy_bins = 128
energy_edges = [e*u.keV for e in np.geomspace(4.5,2000,n_energy_bins+1)]
tte = sim.to_TTE(energy_edges, trigtime = t0.to(u.second).value)
# -
# ### IO
tte.write("./")
tte = TTE.open("glg_tte_all_bn200101000_v00.fit")
# ## Analisis
# ### Lightcurve
# +
phaii = tte.to_phaii(bin_by_time, .2, time_ref=0.0)
lcplot = Lightcurve(data=phaii.to_lightcurve())
plt.show()
# -
# ### Spectrum
# +
from gbm.plot.spectrum import Spectrum
fig,ax = plt.subplots(dpi=150)
spectrum = tte.to_spectrum(time_range=(0.0, 2.0))
specplot = Spectrum(data=spectrum, axis=ax)
E = np.geomspace(30,200,100)
ax.plot(E, 2.5*(E/100)**-1.5, label=r"$\sim E^{-1.5}$")
ax.legend()
plt.show()
# -
| cosima2TTE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Creating and manipulating monte carlo spectra using MITK-MSI
#
# In this tutorial we will learn how to
# 1. create reflectance spectra from examplary tissues
# 2. how to analyse and visualize the created spectra
# 3. how to manipulate them
#
# The MITK-MSI software provides a wrapper to the popular MCML approach to simulate how light travels through tissue. This wrapper can be found in mc/sim.py.
# In this tutorial we will utilize our tissue model which uses this wrapper to create the reflectance spectra.
#
# As a prerequisit, you need a MCML monte carlo simulation which uses the format specified [here](http://omlc.org/software/mc/).
# I tested this software with the GPU accelerated version which can be found [here](https://code.google.com/archive/p/gpumcml/).
# +
# 1.1 create spectra - setup simulation environment
# some necessary imports
import logging
import numpy as np
import os
# everything related to the simulation wrapper
from mc import sim
# the factories create batches (tissue samples) and suited tissue models
from mc import factories
# function which runs simulations for each wavelength
from mc.create_spectrum import create_spectrum
# Where does your monte carlo simulation executable resides in?
MCML_EXECUTABLE = "/home/wirkert/workspace/monteCarlo/gpumcml/fast-gpumcml/gpumcml.sm_20"
# The MCML needs a simulation input file, where shall it be created?
MCI_FILENAME = "./temp.mci"
# filename of the file with the simulation results. Due to a bug in GPUMCML will always
# be created in the same folder as the MCML executable
MCO_FILENAME = "temp.mco"
# The wavelengths for which we want to run our simulation
WAVELENGTHS = np.arange(450, 720, 2) * 10 ** -9
# we want to create standard colonic tissue as specified in the IPCAI 2016 publication
# "Robust Near Real-Time Estimation of Physiological Parameters from Megapixel
# Multispectral Images with Inverse Monte Carlo and Random Forest Regression"
factory = factories.ColonMuscleMeanScatteringFactory()
# if you want to create data from the generic tissue mentioned in the paper, choose:
#factory = factories.GenericMeanScatteringFactory()
# create a simulation wrapper
# the simulation wrapper wraps the mcml executable in python code
sim_wrapper = sim.SimWrapper()
# our simulation needs to know where the input file for the simulation
# shall resign (will be automatically created)
sim_wrapper.set_mci_filename(MCI_FILENAME)
# also it needs to know where the simulation executable shall lie in
sim_wrapper.set_mcml_executable(MCML_EXECUTABLE)
# create the tissue model
# it is responsible for writing the simulation input file
tissue_model = factory.create_tissue_model()
# tell it where the input file shall lie in
tissue_model.set_mci_filename(sim_wrapper.mci_filename)
# also set the output filename
tissue_model.set_mco_filename(MCO_FILENAME)
# tell it how much photons shall be simulated. Will be set to 10**6 by standard,
# this is just an example
tissue_model.set_nr_photons(10**6)
# +
# 1.2 create spectra - create tissue samples for simulation
# setup batch with tissue instances which should be simulated
batch = factory.create_batch_to_simulate()
# we want to simulate ten tissue instances in this example
nr_samples = 10
df = batch.create_parameters(10)
# lets have a look at the dataframe. Each row corresponds to one tissue instance,
# each tissue instance is defined by various layers, which all have certain parameters
# like e.g. oxygenation (here sao2)
df
# +
# 1.3 create spectra - run simulation
# add reflectance column to dataframe
for w in WAVELENGTHS:
df["reflectances", w] = np.NAN # the reflectances have not been calculated yet, thus set no nan
# for each instance in our batch
for i in range(df.shape[0]):
# set the desired element in the dataframe to be simulated
tissue_model.set_dataframe_row(df.loc[i, :])
logging.info("running simulation " + str(i) + " for\n" +
str(tissue_model))
reflectances = create_spectrum(tissue_model, sim_wrapper, WAVELENGTHS)
# store in dataframe
for r, w in zip(reflectances, WAVELENGTHS):
df["reflectances", w][i] = r
# clean up temporarily created files
os.remove(MCI_FILENAME)
created_mco_file = os.path.join(os.path.split(MCML_EXECUTABLE)[0], MCO_FILENAME)
if os.path.isfile(created_mco_file):
os.remove(created_mco_file)
# Hooray, finished,
# now our dataframe also contains reflectances for each tissue instance:
df["reflectances"]
# +
# 2.1 analyse spectra - plot reflectances
# the usual settings for plotting in ipython notebooks
import matplotlib.pylab as plt
# %matplotlib inline
# let's have a look at our reflectances
df["reflectances"].T.plot(kind="line")
plt.ylabel("reflectance")
plt.xlabel("wavelengths [m]")
# put legend outside of plot
plt.gca().legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid()
# +
# 2.1 analyse spectra - show distribution of blood volume fraction (vhb) and sao2
# now we need some special pandas functions
import pandas as pd
# we're interested in the distribution of vhb and sao2 in the first layer (layer0)
df_vhb_sao2 = df["layer0"][["vhb", "sao2"]]
# plot a scatter matrix showing the distribution of vhb and sao2.
# of course, with this little data this does not really make sense,
# however it is a useful tool for analysis if much data is available
pd.tools.plotting.scatter_matrix(df_vhb_sao2, alpha=0.75, figsize=(6, 6))
plt.show()
# +
# 3.1 manipulate spectra - apply sliding average
# in 3.1 and 3.2 we will adapt the generated spectra to an imaginary imaging system
# This system has filters with 20nm bandwith (taken care of in 3.1)
# and takes multispectral images in 10nm steps (taken care of in 3.2)
# the module mc.dfmanipulations was written to provide some basic,
# often needed manipulations of the calculated spectra
# all dmfmanipulations are performed inplace, however, the df is also returned.
import mc.dfmanipulations as dfmani
# first copy to not lose our original data
df2 = df.copy()
# We apply a sliding average to our data. This is usefull if
# we want to see e.g. how the reflectance was recorded by bands with a certain width
# a sliding average of 11 will take the five left and five right of the current reflectance
# and average. Because we take 2nm steps of reflectance in our simulation this means
# a 20nm window.
dfmani.fold_by_sliding_average(df2, 11)
# lets again plot the reflectances
df2["reflectances"].T.plot(kind="line")
plt.ylabel("reflectance")
plt.xlabel("wavelengths [m]")
# put legend outside of plot
plt.gca().legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid()
# we can see that the bump at 560nm is "smoother"
# +
# 3.2 manipulate spectra - select certain wavelenghts
# our imaginary imaging system takes images in 10nm steps from 470 to 660nm
imaging_system_wavelengths = np.arange(470, 670, 10) * 10**-9
df3 = df2.copy()
dfmani.interpolate_wavelengths(df3, imaging_system_wavelengths)
# let's look at the newly created reflectances
df3["reflectances"].T.plot(kind="line", marker='o')
plt.ylabel("reflectance")
plt.xlabel("wavelengths [m]")
# put legend outside of plot
plt.gca().legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.grid()
# -
# that's it, folks. If you want, you can save the created dataframe easily to csv:
df.to_csv("results.csv", index=False)
| Modules/Biophotonics/python/iMC/tutorials/Monte Carlo Spectra Generation - Basic tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# tgb - 4/18/2019
# - We will train unconstrained (U), loss-constrained (L) and architecture-constrained (A) neural networks trained on the 8-column +0K experiment and validated on the same experiment. The loss-constrained networks will have varying architectures to see how MSE and energy conservation performances vary with the importance given to each in the loss function.
#
# Notebook 009 follows the notebook 005 that predicts:
# ***
# [PHQ, PHCLDLIQ, PHCLDICE, TPHYSTND, QRL, QRS, DTVKE, FSNT, FSNS, FLNT, FLNS, PRECT, PRECTEND, PRECST, PRECSTEN] as a function of:
# [QBP, QCBP, QIBP, TBP, VBP, Qdt_adiabatic, QCdt_adiabatic, QIdt_adiabatic, Tdt_adiabatic, Vdt_adiabatic, PS, SOLIN, SHFLX, LHFLX]
# # 1) Load modules and create training/validation data generators
from cbrain.imports import *
from cbrain.data_generator import *
from cbrain.cam_constants import *
from cbrain.losses import *
from cbrain.utils import limit_mem
from cbrain.layers import *
import tensorflow as tf
import tensorflow.math as tfm
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
import xarray as xr
import numpy as np
from cbrain.model_diagnostics import ModelDiagnostics
# Otherwise tensorflow will use ALL your GPU RAM for no reason
limit_mem()
TRAINDIR = '/local/Tom.Beucler/SPCAM_PHYS/'
DATADIR = '/project/meteo/w2w/A6/S.Rasp/SP-CAM/fluxbypass_aqua/'
PREFIX = '8col009_01_'
#PREFIXDS = '8col009_ds1_'
PREFIXDS = PREFIX
# %cd /filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM
# tgb - 4/18/2019 - Used preprocessed data calculated in notebook 009 @ https://github.com/tbeucler/CBRAIN-CAM/blob/master/notebooks/tbeucler_devlog/009_Generalization_Climate_Change_8col.ipynb
# +
scale_dict = load_pickle('./nn_config/scale_dicts/009_Wm2_scaling.pkl')
in_vars = load_pickle('./nn_config/scale_dicts/009_Wm2_in_vars.pkl')
out_vars = load_pickle('./nn_config/scale_dicts/009_Wm2_out_vars.pkl')
dP = load_pickle('./nn_config/scale_dicts/009_Wm2_dP.pkl')
train_gen = DataGenerator(
data_fn = TRAINDIR+PREFIXDS+'train_shuffle.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = TRAINDIR+PREFIX+'norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=True
)
valid_gen = DataGenerator(
data_fn = TRAINDIR+PREFIX+'valid.nc',
input_vars = in_vars,
output_vars = out_vars,
norm_fn = TRAINDIR+PREFIX+'norm.nc',
input_transform = ('mean', 'maxrs'),
output_transform = scale_dict,
batch_size=1024,
shuffle=False
)
# -
# # 2) Build neural networks in a loop
# tgb - 4/18/2019 - https://stackoverflow.com/questions/52320059/creating-a-new-sequential-model-inside-a-for-loop-using-keras
# tgb - 4/24/2019 - Edited for data-scarce network
# +
hsav = {}
hsav[ep] = hist.history
# 5) Save model
path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'DS1_'+str(ep)+'.h5'
NN.save(path)
print('NN saved in ',path)
# +
#alpha_array = [0,0.01,0.25,0.5,0.75,0.99,1] # Loop over weight given to MSE and conservation constraints
alpha_array = [0]
Nep = 100; hsav = {};
for alpha in alpha_array:
NN = {}
print('alpha = ',str(alpha),' and NN is ',NN)
graph = tf.Graph()
with tf.Session(graph=graph):
# 1) Create model
# Unconstrained model with 5 dense layers (Notebook 009)
inpU = Input(shape=(304,))
densout = Dense(512, activation='linear')(inpU)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(218, activation='linear')(densout)
out_layer = LeakyReLU(alpha=0.3)(densout)
NN = tf.keras.models.Model(inpU, out_layer)
print('NN is ',NN.summary())
# 2) Define loss
al = alpha/4 # Weight given to each residual
Loss = WeakLoss(inpU, inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi, name='loss',
alpha_mass=al, alpha_ent=al,
alpha_lw=al, alpha_sw=al)
# 3) Compile model
NN.compile(tf.keras.optimizers.RMSprop(), loss=Loss, metrics=[mse])
# 4) Train model
# Save history following https://stackoverflow.com/questions/49969006/save-and-load-keras-callbacks-history
# for ep in range(20):
hist = NN.fit_generator(train_gen, epochs=Nep)
hsav = hist.history
# 5) Save model
path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'DS1.h5'
NN.save(path)
print('NN saved in ',path)
# -
# tgb - 5/1/2019 - Modified to create NNA network with 0.01=alpha loss function
Nep = 20
# Repeat for architecture-constrained network
graph = tf.Graph()
with tf.Session(graph=graph):
# 1) Create model
# Unconstrained model with 5 dense layers (Notebook 009)
inpC = Input(shape=(304,))
densout = Dense(512, activation='linear')(inpC)
densout = LeakyReLU(alpha=0.3)(densout)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
densout = Dense(214, activation='linear')(densout)
densout = LeakyReLU(alpha=0.3)(densout)
surfout = SurRadLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, densout])
massout = MassConsLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, surfout])
enthout = EntConsLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, massout])
NNA = tf.keras.models.Model(inpC, enthout)
print(NNA.summary())
al = 0.01/4
Loss = WeakLoss(inpC, inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi, name='loss',
alpha_mass=al, alpha_ent=al,
alpha_lw=al, alpha_sw=al)
# 2) Compile model
NNA.compile(tf.keras.optimizers.RMSprop(), loss=Loss, metrics=[mse])
# 3) Train model
NNA.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen)
# 4) Save model
path = TRAINDIR+'HDF5_DATA/NNA0.01.h5'
NNA.save(path)
print('NN saved in ',path)
# tgb - 4/23/2019 - Repeat unconstrained and architecture-constrained examples in multiple linear regression mode to compare performances.
# To make the models linear, simply eliminate the leaky reLU layers.
# +
Nep = 20
alpha = 0
NN = {}
print('alpha = ',str(alpha),' and NN is ',NN)
graph = tf.Graph()
with tf.Session(graph=graph):
# 1) Create model
# Unconstrained model with 5 dense layers (Notebook 009)
inpU = Input(shape=(304,))
densout = Dense(512, activation='linear')(inpU)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
out_layer = Dense(218, activation='linear')(densout)
NN = tf.keras.models.Model(inpU, out_layer)
print('NN is ',NN.summary())
# 2) Define loss
al = alpha/4 # Weight given to each residual
Loss = WeakLoss(inpU, inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi, name='loss',
alpha_mass=al, alpha_ent=al,
alpha_lw=al, alpha_sw=al)
# 3) Compile model
NN.compile(tf.keras.optimizers.RMSprop(), loss=Loss, metrics=[mse])
# 4) Train model
NN.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen)
# 5) Save model
path = TRAINDIR+'HDF5_DATA/MLRL'+str(alpha)+'.h5'
NN.save(path)
print('MLR saved in ',path)
# -
# tgb - 4/23/2019 - Repeat the operation of linearing the neural network in the architecture-constrained case
# Repeat for architecture-constrained network
graph = tf.Graph()
with tf.Session(graph=graph):
# 1) Create model
# Unconstrained model with 5 dense layers (Notebook 009)
inpC = Input(shape=(304,))
densout = Dense(512, activation='linear')(inpC)
for i in range (4):
densout = Dense(512, activation='linear')(densout)
densout = Dense(214, activation='linear')(densout)
surfout = SurRadLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, densout])
massout = MassConsLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, surfout])
enthout = EntConsLayer(
inp_div=train_gen.input_transform.div,
inp_sub=train_gen.input_transform.sub,
norm_q=scale_dict['PHQ'],
hyai=hyai, hybi=hybi
)([inpC, massout])
NNA = tf.keras.models.Model(inpC, enthout)
print(NNA.summary())
# 2) Compile model
NNA.compile(tf.keras.optimizers.RMSprop(), loss=mse, metrics=[mse])
# 3) Train model
NNA.fit_generator(train_gen, epochs=Nep, validation_data=valid_gen)
# 4) Save model
path = TRAINDIR+'HDF5_DATA/MLRA.h5'
NNA.save(path)
print('MLR saved in ',path)
# ## 3) Calculate statistics and residuals in a loop
# ## 3.1) Load models to test statistics
# tgb - 4/19/2019 - Debugging
# +
# config_fn = '/filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM/pp_config/8col_rad_tbeucler_local_PostProc.yml'
# data_fn = '/local/Tom.Beucler/SPCAM_PHYS/8col009_01_valid.nc'
# dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
# # %cd $TRAINDIR/HDF5_DATA
# # !ls
# dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
# alpha = 0
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'.h5'
# NN = load_model(path,custom_objects=dict_lay)
# md = ModelDiagnostics(NN,config_fn,data_fn)
# # 3) Calculate statistics and save in pickle file
# md.compute_stats(niter=5)
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'md.pkl'
# pickle.dump(md.stats,open(path,'wb'))
# print('Stats are saved in ',path)
# # 4) Calculate budget residuals and save in pickle file
# md.compute_res(niter=5)
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'res.pkl'
# pickle.dump(md.res,open(path,'wb'))
# print('Budget residuals are saved in ',path)
# -
# ## 3.2) Statistics
# +
# config_fn = '/filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM/pp_config/8col_rad_tbeucler_local_PostProc.yml'
# data_fn = '/local/Tom.Beucler/SPCAM_PHYS/8col009_01_valid.nc'
# dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
# for alpha in alpha_array:
# # 1) Load model
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'.h5'
# NN = load_model(path,custom_objects=dict_lay)
# # 2) Define model diagnostics object
# md = ModelDiagnostics(NN,config_fn,data_fn)
# # 3) Calculate statistics and save in pickle file
# md.compute_stats()
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'md.pkl'
# pickle.dump(md.stats,open(path,'wb'))
# print('Stats are saved in ',path)
# # 4) Calculate budget residuals and save in pickle file
# md.compute_res()
# path = TRAINDIR+'HDF5_DATA/NNL'+str(alpha)+'res.pkl'
# pickle.dump(md.res,open(path,'wb'))
# print('Budget residuals are saved in ',path)
# -
# ### Statistics of architecture-constrained neural network
# +
# config_fn = '/filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM/pp_config/8col_rad_tbeucler_local_PostProc.yml'
# data_fn = '/local/Tom.Beucler/SPCAM_PHYS/8col009_01_valid.nc'
# dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
# NN = {}; MD = {};
# # 1) Load model
# path = TRAINDIR+'HDF5_DATA/NNA.h5'
# NN = load_model(path,custom_objects=dict_lay)
# # 2) Define model diagnostics object
# md = ModelDiagnostics(NN,config_fn,data_fn)
# # 3) Calculate statistics and save in pickle file
# # md.compute_stats()
# # path = TRAINDIR+'HDF5_DATA/NNAmd.pkl'
# # pickle.dump(md.stats,open(path,'wb'))
# # print('Stats are saved in ',path)
# # 4) Calculate budget residuals and save in pickle file
# md.compute_res()
# path = TRAINDIR+'HDF5_DATA/NNAres.pkl'
# pickle.dump(md.res,open(path,'wb'))
# print('Budget residuals are saved in ',path)
# -
# ### Statistics of multiple linear regressions
# tgb - 4/23/2019
# +
config_fn = '/filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM/pp_config/8col_rad_tbeucler_local_PostProc.yml'
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/8col009_01_valid.nc'
dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
# 1) Load model
path = TRAINDIR+'HDF5_DATA/MLRL0.h5'
NN = load_model(path,custom_objects=dict_lay)
# 2) Define model diagnostics object
md = ModelDiagnostics(NN,config_fn,data_fn)
# 3) Calculate statistics and save in pickle file
# md.compute_stats()
# path = TRAINDIR+'HDF5_DATA/MLRL0md.pkl'
# pickle.dump(md.stats,open(path,'wb'))
# print('Stats are saved in ',path)
# 4) Calculate budget residuals and save in pickle file
md.compute_res()
path = TRAINDIR+'HDF5_DATA/MLRL0res.pkl'
pickle.dump(md.res,open(path,'wb'))
print('Budget residuals are saved in ',path)
NN = {}; md = {};
# 1) Load model
path = TRAINDIR+'HDF5_DATA/MLRA.h5'
NN = load_model(path,custom_objects=dict_lay)
# 2) Define model diagnostics object
md = ModelDiagnostics(NN,config_fn,data_fn)
# 3) Calculate statistics and save in pickle file
md.compute_stats()
path = TRAINDIR+'HDF5_DATA/MLRAmd.pkl'
pickle.dump(md.stats,open(path,'wb'))
print('Stats are saved in ',path)
# 4) Calculate budget residuals and save in pickle file
md.compute_res()
path = TRAINDIR+'HDF5_DATA/MLRAres.pkl'
pickle.dump(md.res,open(path,'wb'))
print('Budget residuals are saved in ',path)
# -
| notebooks/tbeucler_devlog/010_Conserving_Network_Paper_Runs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Neural Networks with Keras
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# %pylab inline
import matplotlib.pylab as plt
import numpy as np
from distutils.version import StrictVersion
# +
import sklearn
print(sklearn.__version__)
assert StrictVersion(sklearn.__version__ ) >= StrictVersion('0.18.1')
# +
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
print(tf.__version__)
assert StrictVersion(tf.__version__) >= StrictVersion('1.1.0')
# +
import keras
print(keras.__version__)
assert StrictVersion(keras.__version__) >= StrictVersion('2.0.0')
# +
import pandas as pd
print(pd.__version__)
assert StrictVersion(pd.__version__) >= StrictVersion('0.20.0')
# -
# ## Solving Iris with Neural Networks
from sklearn.datasets import load_iris
iris = load_iris()
iris.data[0]
print(iris.DESCR)
# +
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
iris_df = pd.DataFrame(iris.data, columns=iris.feature_names)
CMAP = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
pd.plotting.scatter_matrix(iris_df, c=iris.target, edgecolor='black', figsize=(15, 15), cmap=CMAP)
plt.show()
# -
# ## The artificial Neuron
# 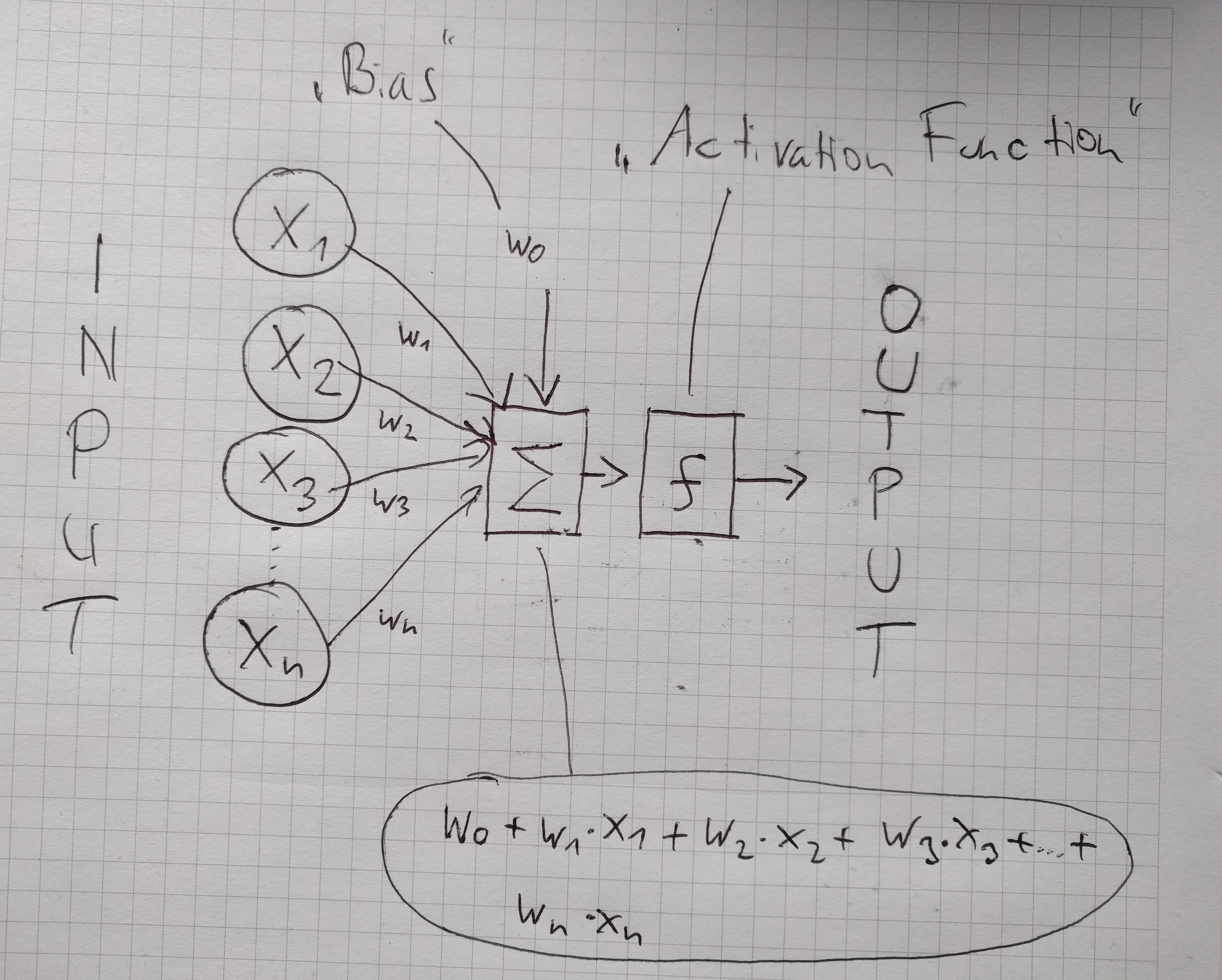
# ## Our first Neural Network with Keras
# 
# +
# # keras.layers.Input?
# -
from keras.layers import Input
inputs = Input(shape=(4, ))
# +
# # keras.layers.Dense?
# -
from keras.layers import Dense
# just linear activation (like no activation function at all)
fc = Dense(3)(inputs)
from keras.models import Model
model = Model(input=inputs, output=fc)
model.summary()
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
# this is just random stuff, no training has taken place so far
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
# ### This is the output of all 3 hidden neurons, but what we really want is a category for iris category
# * Softmax activation turns each output to a percantage between 0 and 1 all adding up to 1
# * interpretation is likelyhood of category
# 
inputs = Input(shape=(4, ))
fc = Dense(3)(inputs)
predictions = Dense(3, activation='softmax')(fc)
model = Model(input=inputs, output=predictions)
model.summary()
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
# ## Now we have likelyhoods for categories, but still our model is totally random
# # Training
# * training is performed using Backpropagation
# * each pair of ground truth input and output is passed through network
# * difference between expected output (ground truth) and actual result is summed up and forms loss function
# * loss function is to be minimized
# * optimizer defines strategy to minimize loss
# ### Optimizers: Adam and RMSprop seem nice
# 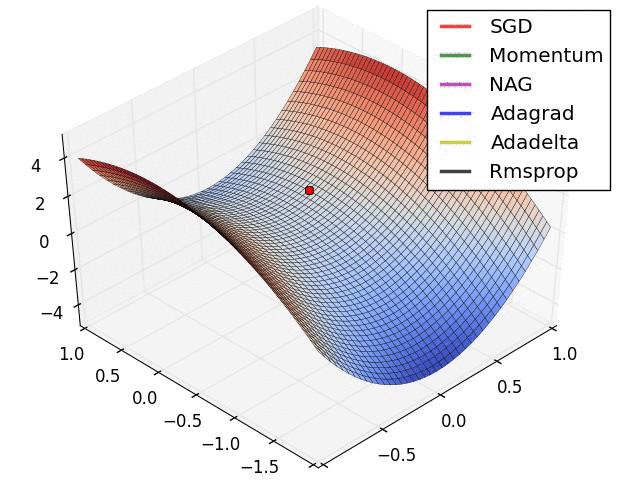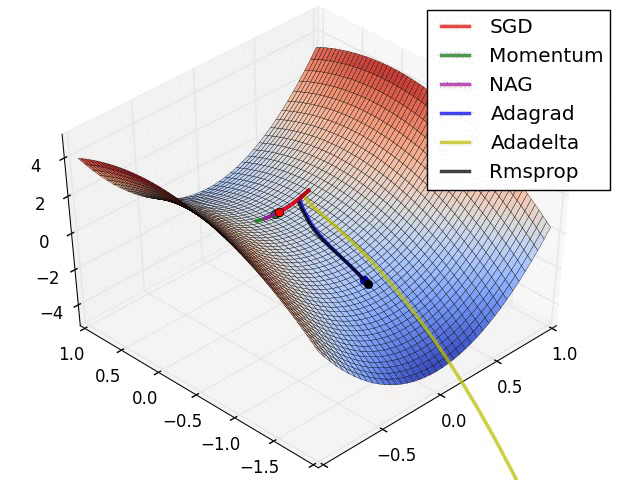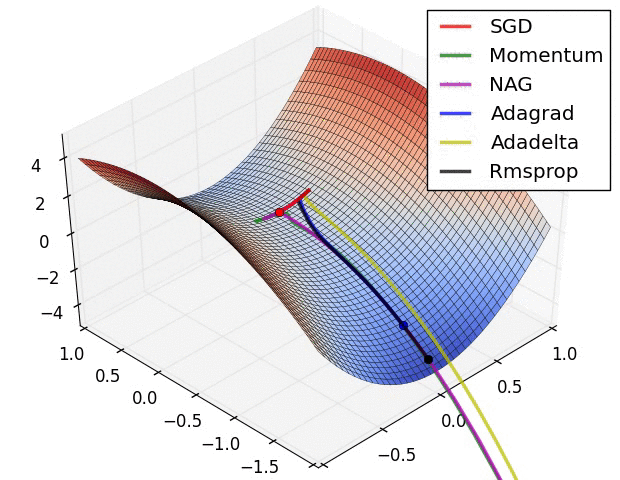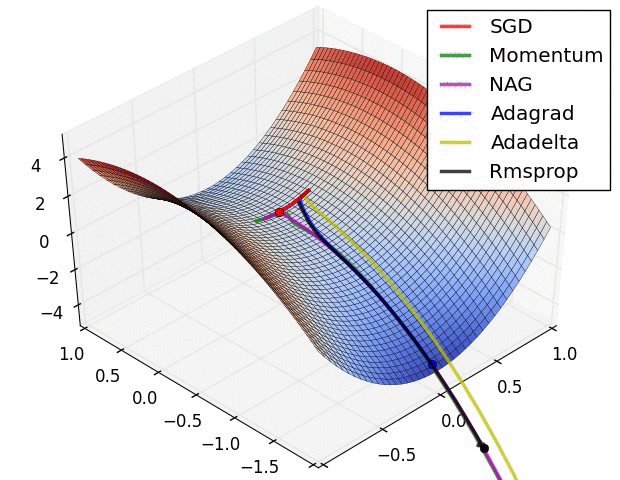
# 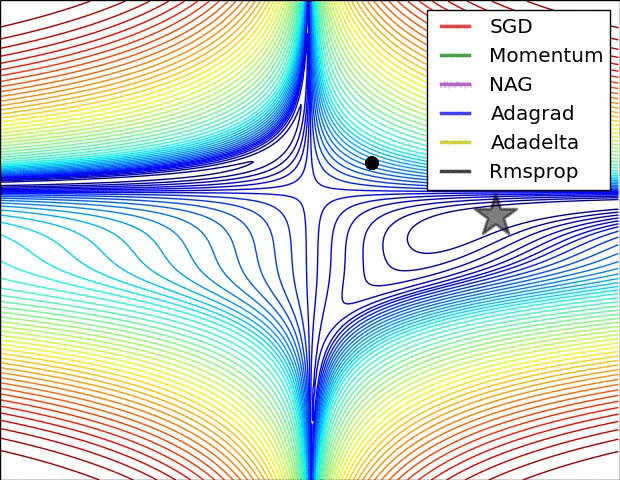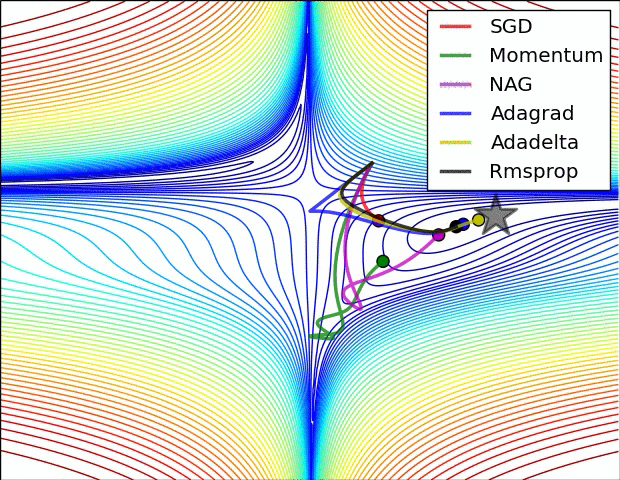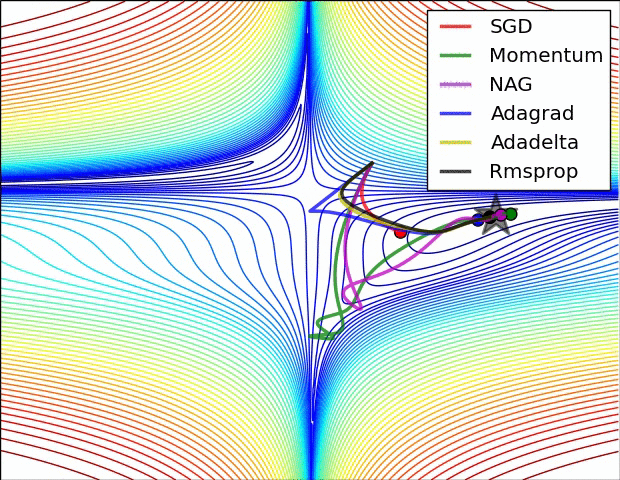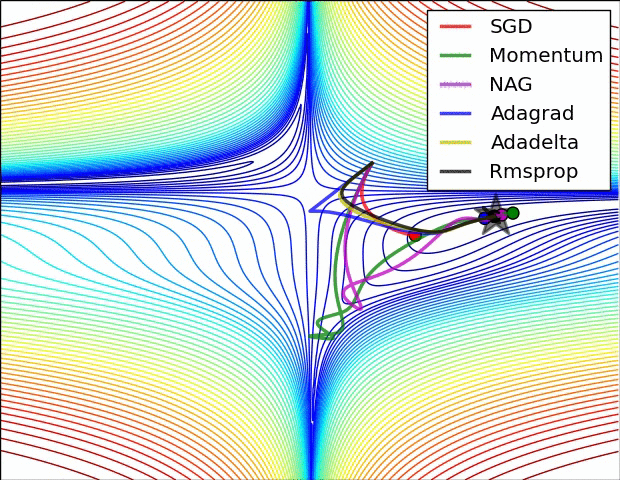
# http://cs231n.github.io/neural-networks-3/#ada
X = np.array(iris.data)
y = np.array(iris.target)
X.shape, y.shape
y[100]
# +
# tiny little pieces of feature engeneering
from keras.utils.np_utils import to_categorical
num_categories = 3
y = to_categorical(y, num_categories)
# -
y[100]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42, stratify=y)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# +
# !rm -r tf_log
tb_callback = keras.callbacks.TensorBoard(log_dir='./tf_log')
# https://keras.io/callbacks/#tensorboard
# To start tensorboard
# tensorboard --logdir=/mnt/c/Users/olive/Development/ml/tf_log
# open http://localhost:6006
# -
# %time model.fit(X_train, y_train, epochs=500, validation_split=0.2, callbacks=[tb_callback])
# # %time model.fit(X_train, y_train, epochs=500, validation_split=0.2)
# # Evaluation
model.predict(np.array([[ 5.1, 3.5, 1.4, 0.2]]))
X[0], y[0]
train_loss, train_accuracy = model.evaluate(X_train, y_train)
train_loss, train_accuracy
test_loss, test_accuracy = model.evaluate(X_test, y_test)
test_loss, test_accuracy
# ---
#
# # Hands-On
# ## Execute this notebook and improve the training results
# * Are the results you see good?
# * What kind of decision boundary can our neurons draw?
# * Can you improve the results? Can you at least match Knn?
# * What is the total mimimum of neurons you need for this problem?
# * Play around with the number of neurons in the hidden layer
# * Add another hidden layer
# * More than one hidden layer is what people call deep neural networks
# * hidden layers close to the input hopefully do the feature engeneering and extraction for us
# * Increase the number of epochs for training
# ---
# # Stop Here
# ## Save Model in Keras und TensorFlow Format
# Keras format
model.save('nn-iris.hdf5')
# ## Export as raw tf model
#
# * https://tensorflow.github.io/serving/serving_basic.html
# * https://github.com/tensorflow/serving/blob/master/tensorflow_serving/example/mnist_saved_model.py
import os
from keras import backend as K
K.set_learning_phase(0)
sess = K.get_session()
# !rm -r tf
tf.app.flags.DEFINE_integer('model_version', 1, 'version number of the model.')
tf.app.flags.DEFINE_string('work_dir', '/tmp', 'Working directory.')
FLAGS = tf.app.flags.FLAGS
export_path_base = 'tf'
export_path = os.path.join(
tf.compat.as_bytes(export_path_base),
tf.compat.as_bytes(str(FLAGS.model_version)))
classification_inputs = tf.saved_model.utils.build_tensor_info(model.input)
classification_outputs_scores = tf.saved_model.utils.build_tensor_info(model.output)
from tensorflow.python.saved_model.signature_def_utils_impl import build_signature_def, predict_signature_def
signature = predict_signature_def(inputs={'inputs': model.input},
outputs={'scores': model.output})
builder = tf.saved_model.builder.SavedModelBuilder(export_path)
builder.add_meta_graph_and_variables(
sess,
tags=[tf.saved_model.tag_constants.SERVING],
signature_def_map={
tf.saved_model.signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY: signature
})
builder.save()
# !ls -lhR tf
# ## This TensorFlow Modell can be uploaded to Google Cloud ML and called via REST interface
# +
# # cd tf
# gsutil cp -R 1 gs://irisnn
# create model and version at https://console.cloud.google.com/mlengine
# gcloud ml-engine predict --model=irisnn --json-instances=./sample_iris.json
# SCORES
# [0.9954029321670532, 0.004596732556819916, 3.3544753819114703e-07]
# -
| notebooks/workshops/tss/nn-intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from qiskit_aqua import run_algorithm
from qiskit_qcgpu_provider import QCGPUProvider
sat_cnf = """
c Example DIMACS 3-sat
p cnf 3 5
-1 -2 -3 0
1 -2 3 0
1 2 -3 0
1 -2 -3 0
-1 2 3 0
"""
params = {
"problem": { "name": "search" },
"algorithm": { "name": "Grover" },
"oracle": { "name": "SAT", "cnf": sat_cnf },
"backend": { "name": "qasm_simulator" }
}
# -
backend = QCGPUProvider().get_backend('qasm_simulator')
# %time result_qiskit = run_algorithm(params)
# %time result = run_algorithm(params, backend=backend)
print(result["result"])
| examples/aqua/Grovers Algorithm for SAT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from math import pi, atan, sin, cos
import cv2
import matplotlib.pyplot as plt
import numpy as np
from scipy import spatial
from tqdm import tqdm
from shapely import affinity
from shapely.geometry import LineString, Point
from labvision import images
import filehandling
from particletracking import dataframes, statistics
# -
def get_cgw(df, k):
tree = spatial.cKDTree(df[['x', 'y']].values)
dists, _ = tree.query(tree.data, 2)
cgw = np.mean(dists[:, 1])
return cgw * k
def coarse_order_field(df, cgw, x, y, no_of_neighbours=20):
"""
Calculate the coarse-grained field characterising local orientation order
"""
order = df.order.values
# Generate the lattice nodes to query
# x, y = np.meshgrid(x, y)
r = np.dstack((x, y))
# Get the positions of all the particles
particles = df[['x', 'y']].values
# Generate the tree from the particles
tree = spatial.cKDTree(particles)
# Query the tree at all the lattice nodes to find the nearest n particles
# Set n_jobs=-1 to use all cores
dists, indices = tree.query(r, no_of_neighbours, n_jobs=-1)
# Calculate all the coarse-grained delta functions (Katira ArXiv eqn 3
cg_deltas = np.exp(-dists ** 2 / (2 * cgw ** 2)) / (2 * pi * cgw ** 2)
# Multiply by the orders to get the summands
summands = cg_deltas * order[indices]
# Sum along axis 2 to calculate the field
field = np.sum(summands, axis=2)
return field
def get_field_threshold(fields, ls, im):
# Draw a box around an always ordered region of the image to
# calculate the phi_o
fields = np.dstack(fields)
line_selector = LineSelector(im)
op1, op2 = line_selector.points
phi_o = np.mean(
fields[op1[1] // ls:op2[1] // ls, op1[0] // ls:op2[0] // ls, :])
# Repeat for disordered
line_selector = LineSelector(im)
dp1, dp2 = line_selector.points
phi_d = np.mean(
fields[dp1[1] // ls:dp2[1] // ls, dp1[0] // ls:dp2[0] // ls, :])
field_threshold = (phi_o + phi_d) / 2
return field_threshold
class LineSelector:
def __init__(self, im):
cv2.namedWindow('line', cv2.WINDOW_NORMAL)
cv2.resizeWindow('line', 960, 540)
cv2.setMouseCallback('line', self.record)
self.points = []
while True:
cv2.imshow('line', im)
key = cv2.waitKey(1) & 0xFF
if len(self.points) == 2:
break
cv2.destroyAllWindows()
def record(self, event, x, y, flags, param):
if event == cv2.EVENT_LBUTTONDOWN:
self.points.append([x, y])
def get_extended_centre_line(p1, p2):
dx = p2[0] - p1[0]
dy = p2[1] - p1[1]
p0 = (p1[0] - dx, p1[1] - dy)
p3 = (p2[0] + dx, p2[1] + dy)
return LineString(((p0[0], p0[1]), (p3[0], p3[1])))
def find_contours(f, t):
t_low = t - 0.02 * t
t_high = t + 0.02 * 5
new_f = (f < t_high) * (f > t_low)
new_f = np.uint8(new_f)
contours = images.find_contours(new_f)
contours = images.sort_contours(contours)
try:
return contours[-1]
except IndexError as e:
print("Only one contour")
return contours
def close_contour(c):
c = np.vstack((c, c[0, :]))
return c
def get_dists(x, y, c, l):
dists = []
crosses = []
for (xp, yp) in zip(x, y):
p = Point((xp, yp))
l_rot = affinity.rotate(l, 90, p)
cross = c.intersection(l_rot)
if cross.geom_type == 'Point':
dist = cross.distance(p)
cross = cross.x, cross.y
elif cross.geom_type == 'MultiPoint':
ds = [c.distance(p) for c in cross]
dist = np.min(ds)
cross = cross[np.argmin(ds)]
cross = cross.x, cross.y
else:
dist = 0
cross = xp, yp
dists.append(dist)
crosses.append(cross)
return dists, crosses
def get_angle(im):
ls = LineSelector(im)
p1, p2 = ls.points
m = (p2[1] - p1[1]) / (p2[0] - p1[0])
a = -atan(m)
c = np.array([i // 2 for i in np.shape(im)])[::-1]
return a, c, p1, p2
# +
def plot_fft(dists, dL, pix_2_mm):
dL *= pix_2_mm
sp = [np.abs(np.fft.fft(np.array(h)*pix_2_mm))**2 for h in dists]
N = len(dists[0])
freq = np.fft.fftfreq(N, dL)[1:N//2]
y = (np.stack(sp)* dL * N)[1:N//2]
y_mean = np.mean(y, axis=0).squeeze()
y_err = np.std(y, axis=0, ddof=1).squeeze()
xplot = freq*2*np.pi
L_x = 2 * np.pi / (dL * N)
r_x = 2 * np.pi / (data.df.loc[0].r.mean() * pix_2_mm)
cgw_x = 2 * np.pi / (cgw * pix_2_mm)
xmin = 0
xmax = sum(xplot < cgw_x)
xplot = np.log10(xplot[xmin:xmax])
yplot = np.log10(y_mean[xmin:xmax])
yplot_err = 0.434 * y_err[xmin:xmax] / y_mean[xmin:xmax]
coeffs, cov = np.polyfit(xplot, yplot, 1, w=yplot_err, cov=True)
fit_func = np.poly1d(coeffs)
yfit = fit_func(xplot)
m = coeffs[0]
dm = np.sqrt(cov[0, 0])
# m, c, sm, sc = get_fit(xplot, yplot, yplot_err)
# yfit = m*xplot + c
plt.figure()
plt.errorbar(xplot, yplot, yerr=yplot_err, fmt='o')
plt.plot(xplot, yfit, label=f'Fit with gradient {m:.3f} +/- {dm:.3f}')
plt.axvline(np.log10(L_x), label='L' , c='r')
plt.axvline(np.log10(cgw_x), label='cgw', c='b')
# plt.axvline(np.log10(r_x), label='r', c='g')
plt.xlabel('log$_{10}(k = 2\pi m/L)$ [mm$^{-1}$]')
plt.ylabel('log$_{10}(<|\delta h_k|^2>L)$ [mm$^3$]')
plt.legend()
# -
dL
PIX_2_mm
def get_pix_2_mm():
_, _, p1, p2 = get_angle(ims[0])
L_pix = np.sqrt((p1[0]-p2[0])**2+(p1[1]-p2[1])**2)
L_mm = 200.0
return L_mm / L_pix
PIX_2_mm = get_pix_2_mm()
plot_fft(dists, dL, PIX_2_mm)
N = len(dists[0])
f = np.fft.fftfreq(N, dL)
f[(N-1)//2]
N
dL
# +
direc = "/media/data/Data/FirstOrder/Interfaces/RecordFluctuatingInterfaceJanuary2020/Quick/first_frames"
savename = f"{direc}/data_new.hdf5"
files = filehandling.get_directory_filenames(direc+'/*.png')
ims = [images.load(f, 0) for f in tqdm(files, 'Loading images')]
ims = [images.bgr_to_gray(im) for im in ims]
circles = [images.find_circles(im, 27, 200, 7, 16, 16)
for im in tqdm(ims, 'Finding Circles')]
data = dataframes.DataStore(savename, load=False)
for f, info in tqdm(enumerate(circles), 'Adding Circles'):
data.add_tracking_data(f, info, ['x', 'y', 'r'])
calc = statistics.PropertyCalculator(data)
calc.order()
lattice_spacing = 10
x = np.arange(0, ims[0].shape[1], lattice_spacing)
y = np.arange(0, ims[0].shape[0], lattice_spacing)
x, y = np.meshgrid(x, y)
cgw = get_cgw(data.df.loc[0], 1.85)
fields = [coarse_order_field(data.df.loc[f], cgw, x, y)
for f in tqdm(range(len(ims)), 'Calculating Fields')]
field_threshold = get_field_threshold(fields, lattice_spacing, ims[0])
contours = [find_contours(f, field_threshold)
for f in tqdm(fields, 'Calculating contours')]
# Multiply the contours by the lattice spacing and squeeze
contours = [c.squeeze()*lattice_spacing for c in contours]
# Close contours
contours = [close_contour(c) for c in contours]
# Convert to LineString
contours = [LineString(c) for c in contours]
# +
# Find the line along the centre of the tray
a, c, p1, p2 = get_angle(ims[0])
centre_line = get_extended_centre_line(p1, p2)
plt.plot(p1[0], p1[1], 'x')
plt.plot(p2[0], p2[1], '.')
plt.plot(centre_line.coords.xy[0], centre_line.coords.xy[1])
# -
# Distance between query points that determines one end of the frequency
dL = data.df.loc[0].r.mean() / 10
L = np.sqrt((p1[0]-p2[0])**2+(p1[1]-p2[1])**2)
N_query = int(L / dL)
# query points
xq, yq = np.linspace(p1[0], p2[0], N_query), np.linspace(p1[1], p2[1], N_query)
dL = np.sqrt((xq[1]-xq[0])**2 + (yq[1]-yq[0])**2)
print(len(xq), len(yq))
dists, crosses = zip(*[get_dists(xq, yq, c, centre_line) for c in tqdm(contours)])
plot_fft(dists, dL)
def draw_contour(im, c, i):
plt.figure()
c = c.coords.xy
plt.imshow(im)
plt.plot(c[0], c[1], 'r')
plt.axis('off')
plt.savefig(direc+f'/annotated/{i}_line.jpg', dpi=900)
for i in range(len(ims)):
draw_contour(ims[i], contours[i], i)
crosses_arr = np.array(crosses)
crosses_arr.shape
mean_cross = np.mean(crosses_arr, axis=0)
# %matplotlib auto
plt.imshow(ims[0], cmap='gray')
plt.plot(mean_cross[:, 0], mean_cross[:, 1], 'r-', label='mean')
plt.plot(crosses_arr[0, :, 0], crosses_arr[0, :, 1], 'y-', label='current')
plt.legend()
plt.close('all')
dists_arr = np.array(dists)
dists_arr.shape
dists_mean = np.mean(dists_arr, axis=0)
dists_sub = [(np.array(d) - dists_mean).tolist() for d in dists]
plot_fft(dists_sub, dL, PIX_2_mm)
# %matplotlib inline
plot_fft(dists, dL, PIX_2_mm)
mean_cross
from scipy import signal
y = signal.savgol_filter(mean_cross[:, 1], 901, 3)
plt.imshow(ims[0], cmap='gray')
plt.plot(*mean_cross.T)
plt.plot(mean_cross[:, 0], y)
from labvision import video
vid0filename = "/media/data/Data/FirstOrder/Interfaces/RecordFluctuatingInterfaceJanuary2020/Quick/16750001.MP4"
vid0 = video.ReadVideo(vid0filename)
ims = [vid0.read_next_frame() for f in range(vid0.num_frames)]
ims = [images.bgr_to_gray(im) for im in ims]
circles = [images.find_circles(im, 27, 200, 7, 16, 16)
for im in tqdm(ims, 'Finding Circles')]
# +
data = dataframes.DataStore(savename, load=False)
for f, info in tqdm(enumerate(circles), 'Adding Circles'):
data.add_tracking_data(f, info, ['x', 'y', 'r'])
calc = statistics.PropertyCalculator(data)
calc.order()
lattice_spacing = 10
x = np.arange(0, ims[0].shape[1], lattice_spacing)
y = np.arange(0, ims[0].shape[0], lattice_spacing)
x, y = np.meshgrid(x, y)
cgw = get_cgw(data.df.loc[0], 1.85)
fields = [coarse_order_field(data.df.loc[f], cgw, x, y)
for f in tqdm(range(len(ims)), 'Calculating Fields')]
field_threshold = get_field_threshold(fields, lattice_spacing, ims[0])
contours = [find_contours(f, field_threshold)
for f in tqdm(fields, 'Calculating contours')]
# Multiply the contours by the lattice spacing and squeeze
contours = [c.squeeze()*lattice_spacing for c in contours]
# Close contours
contours = [close_contour(c) for c in contours]
# Convert to LineString
contours = [LineString(c) for c in contours]
# -
dists, crosses = zip(*[get_dists(xq, yq, c, centre_line) for c in tqdm(contours)])
plot_fft(dists, dL, PIX_2_mm)
| first_order/interface/average_capillary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import numpy as np
class Vector:
def __init__(self, x, y):
self.x = x
self.y = y
#all the properties of vectors and the tools we can use with them. We could use numpy, but this made more intuitive sense to make our own
#class, and rewrite our own operators. It is nice to stay consistent with the class and object style from before with the balls.
def len(self):
return math.sqrt(self.x*self.x + self.y*self.y)
def __add__(self, other):
return Vector(self.x + other.x, self.y + other.y)
def __sub__(self, other):
return Vector(self.x - other.x, self.y - other.y)
def __mul__(self, other):
return Vector(self.x * other, self.y * other)
def __rmul__(self, other):
return Vector(self.x * other, self.y * other)
def __truediv__(self, other):
return Vector(self.x / other, self.y / other)
def angle(self):
return math.atan2(self.y, self.x)
def norm(self):
if self.x == 0 and self.y == 0:
return Vector(0, 0)
return self / self.len()
def dot(self, other):
return self.x*other.x + self.y*other.y
# +
Position1 = Vector(2,3)
Position2 = Vector(3,6)
Position1.dot(Position2)
# -
Position1.angle()
Total = Position1 + Position2
Total.x
Total.y
# +
class Planet:
def __init__(self, position, velocity, rotation):
#Position and Velocity are vector objects from the vector class
self.position = position
self.velocity = velocity
self.rotation = rotation
# -
Mercury = Planet(Vector(2,3), Vector(0,0), 2*np.pi)
type(Mercury)
# +
class Human:
def __init__(self, age, height):
self.age = age
self.height = height
def grow(self):
if self.height < 72:
self.height += 1
self.age += 1
else:
self.age += 1
James = Human(1,20)
# -
type(James)
# +
year = 0
while year < 100:
print("At the age of {0:0.0f} James was {1:0.0f} inches tall".format(James.age,James.height))
year += 1
# -
def spawn_objects(N):
planets = []
initial_x_coords = np.arange(1,1000, N)
initial_y_coords = np.zeros(N)
initial_z_coords = np.zeros(N)
for i in range(N):
planet_i = Planet(Vector(initial_x_coords[i], initial_y_coords[i], initial_z_coords[i]))
planets.append(planet_i)
return planets
| Demos/OOP/Classes Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.0 64-bit
# language: python
# name: python3
# ---
# %matplotlib qt
import matplotlib.pyplot as plt
from scipy.io import wavfile
from scipy import signal
import numpy as np
from glob import glob
from scipy.ndimage.filters import gaussian_filter1d
from scipy import signal
# !make -f stages/test/makefile
# %time !./stages_test
new = {}
stock = {}
for fn in glob("stages_*wav"):
try:
n = wavfile.read(fn)[1]
except Exception as e:
print(fn)
print(e)
continue
new[fn[:-4]] = n
try:
s = wavfile.read(f"stages_test_reference/{fn}")[1]
stock[fn[:-4]] = s
except:
print(f"stock file not found for {fn}; skipping")
continue
if (s==n).all():
print(f"✅ {fn}")
else:
print(f"❌ {fn}")
for i, label in enumerate(["gate", "value", "segment", "phase"]):
if (s.T[i] == n.T[i]).all():
print(f"\t✅ {label}")
else:
ixs = np.where(s.T[i] != n.T[i])[0]
print(f"\t❌ {label}")
plt.figure()
plt.plot(s.T[i], label="old")
plt.plot(n.T[i], label="new")
plt.title(f"{fn} - {label}")
plt.xlim(ixs[0], ixs[-1])
#plt.legend()
rand_tap = new["stages_random_tap_lfo"].T
((np.diff(rand_tap[0]) > 0) == (np.diff(rand_tap[1]) != 0)).all()
plt.figure()
plt.loglog(*signal.welch(new["stages_random_white_noise"].T[1], scaling="spectrum"))
plt.loglog(*signal.welch(new["stages_random_brown_noise"].T[1], scaling="spectrum"))
phase = new["stages_tap_lfo_audio_rate"].T[3]
gate = new["stages_tap_lfo_audio_rate"].T[0]
phase_peaks = signal.find_peaks(phase)[0]
gate_peaks = signal.find_peaks(gate)[0]
32000 / np.diff(phase_peaks).mean(), 32000 / np.diff(gate_peaks).mean()
plt.figure()
plt.plot(gate)
plt.plot(phase)
| stages-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# # Propiedades.
#
# En Python es posible hacer que ciertos métodos se comporten de como si fueran atributos mediante el uso de *property*.
#
# Las propiedades en Python corresponden a métodos que se comportan como "getters" y "setters" de un atributo.
#
# Un "getter" regresa el contenido de un atributo.
# Un "setter" crea o modifica el contenido de un atributo.
#
# La sintaxis para la definición de propiedades es la siguiente:
#
# ```
# class <nombre de la clase>:
# ...
# ...
# @property
# def <nombre de la propiedad>(self):
# ...
# ...
# return <objeto>
# @<nombre de la propiedad>.setter
# def<nombre de la propiedad>(self, <parametro>):
# ...
# ...
# ```
#
# La primera función definida es un getter regresará un valor de la propiedad cuando se invoque y la segunda asignará el valor a la propiedad cuando se utilice un operador de asignación.
#
# Si no se define un setter, el valor de la propiedad no podrá ser cambiado.
#
# **Ejemplo:**
# En este caso se crearan dos propiedades: _clave_ y _nombre_.
#
# * El método *\_\_init\_\_()* crea una cadena de texto basada en una estampa de tiempo y lo liga al atributo *\_\_clave*.
#
# * La propiedad _clave_ sólo cuenta con un getter.
#
# * El getter de _clave_ despliega la cadena de caracteres del atributo *\_\_clave*.
#
# * El contenido de _nombre_ es guardado en el atributo *lista_nombre*, el cual debe de ser un objeto de tipo _list_ que debe de estar conformado por un número de 2 a 3 elementos.
#
# * El setter de _nombre_ valida que se ingrese una lista o una tupla que contenga entre 2 y 3 objetos. De lo contrario, levantará una excepción de tipo *ValueError*.
#
# * El getter de _nombre_ arma y regresa un objeto de tipo _str_ a partir del atributo *lista_nombre*.
from time import time
class Persona:
'''Clase base para creación de datos personales.'''
def __init__(self):
'''Genera una clave única a partir de una estampa de tiempo y la relaciona con el atributo __clave.'''
self.__clave = str(int(time() / 0.017))[1:]
@property
def clave(self):
'''Regresa el valor del atributo "escondido" __clave.'''
return self.__clave
@property
def nombre(self):
'''Regresa una cadena de caracteres a partir de la lista contenida en lista_nombre.'''
return " ".join(self.lista_nombre)
@nombre.setter
def nombre(self, nombre):
'''Debe ingresarse una lista o tupla con entre 2 y 3 elementos.'''
if len(nombre) < 2 or len(nombre) > 3 or type(nombre) not in (list, tuple):
raise ValueError("Formato incorrecto.")
else:
self.lista_nombre = nombre
sujeto = Persona()
sujeto.nombre
sujeto.nombre = ["Jorge", "Sánchez", "Pérez"]
sujeto.nombre
sujeto.lista_nombre
sujeto.clave
sujeto.clave = 12
dir(sujeto)
getattr(sujeto, '_Persona__clave')
setattr(sujeto,"_Persona__clave", "te juanquié")
sujeto.clave
| C05_propiedades.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Random Sampling
# =============
#
# Copyright 2016 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](http://creativecommons.org/licenses/by/4.0/)
# +
from __future__ import print_function, division
import numpy
import scipy.stats
import matplotlib.pyplot as pyplot
from ipywidgets import interact, interactive, fixed
import ipywidgets as widgets
# seed the random number generator so we all get the same results
numpy.random.seed(18)
# some nicer colors from http://colorbrewer2.org/
COLOR1 = '#7fc97f'
COLOR2 = '#beaed4'
COLOR3 = '#fdc086'
COLOR4 = '#ffff99'
COLOR5 = '#386cb0'
# %matplotlib inline
# -
# Part One
# ========
#
# Suppose we want to estimate the average weight of men and women in the U.S.
#
# And we want to quantify the uncertainty of the estimate.
#
# One approach is to simulate many experiments and see how much the results vary from one experiment to the next.
#
# I'll start with the unrealistic assumption that we know the actual distribution of weights in the population. Then I'll show how to solve the problem without that assumption.
#
# Based on data from the [BRFSS](http://www.cdc.gov/brfss/), I found that the distribution of weight in kg for women in the U.S. is well modeled by a lognormal distribution with the following parameters:
weight = scipy.stats.lognorm(0.23, 0, 70.8)
weight.mean(), weight.std()
# Here's what that distribution looks like:
xs = numpy.linspace(20, 160, 100)
ys = weight.pdf(xs)
pyplot.plot(xs, ys, linewidth=4, color=COLOR1)
pyplot.xlabel('weight (kg)')
pyplot.ylabel('PDF')
None
# `make_sample` draws a random sample from this distribution. The result is a NumPy array.
def make_sample(n=100):
sample = weight.rvs(n)
return sample
# Here's an example with `n=100`. The mean and std of the sample are close to the mean and std of the population, but not exact.
sample = make_sample(n=100)
sample.mean(), sample.std()
# We want to estimate the average weight in the population, so the "sample statistic" we'll use is the mean:
def sample_stat(sample):
return sample.mean()
# One iteration of "the experiment" is to collect a sample of 100 women and compute their average weight.
#
# We can simulate running this experiment many times, and collect a list of sample statistics. The result is a NumPy array.
def compute_sample_statistics(n=100, iters=1000):
stats = [sample_stat(make_sample(n)) for i in range(iters)]
return numpy.array(stats)
# The next line runs the simulation 1000 times and puts the results in
# `sample_means`:
sample_means = compute_sample_statistics(n=100, iters=1000)
# Let's look at the distribution of the sample means. This distribution shows how much the results vary from one experiment to the next.
#
# Remember that this distribution is not the same as the distribution of weight in the population. This is the distribution of results across repeated imaginary experiments.
pyplot.hist(sample_means, color=COLOR5)
pyplot.xlabel('sample mean (n=100)')
pyplot.ylabel('count')
None
# The mean of the sample means is close to the actual population mean, which is nice, but not actually the important part.
sample_means.mean()
# The standard deviation of the sample means quantifies the variability from one experiment to the next, and reflects the precision of the estimate.
#
# This quantity is called the "standard error".
std_err = sample_means.std()
std_err
# We can also use the distribution of sample means to compute a "90% confidence interval", which contains 90% of the experimental results:
conf_int = numpy.percentile(sample_means, [5, 95])
conf_int
# The following function takes an array of sample statistics and prints the SE and CI:
def summarize_sampling_distribution(sample_stats):
print('SE', sample_stats.std())
print('90% CI', numpy.percentile(sample_stats, [5, 95]))
# And here's what that looks like:
summarize_sampling_distribution(sample_means)
# Now we'd like to see what happens as we vary the sample size, `n`. The following function takes `n`, runs 1000 simulated experiments, and summarizes the results.
def plot_sample_stats(n, xlim=None):
sample_stats = compute_sample_statistics(n, iters=1000)
summarize_sampling_distribution(sample_stats)
pyplot.hist(sample_stats, color=COLOR2)
pyplot.xlabel('sample statistic')
pyplot.xlim(xlim)
# Here's a test run with `n=100`:
plot_sample_stats(100)
# Now we can use `interact` to run `plot_sample_stats` with different values of `n`. Note: `xlim` sets the limits of the x-axis so the figure doesn't get rescaled as we vary `n`.
# +
def sample_stat(sample):
return sample.mean()
slider = widgets.IntSlider(min=10, max=1000, value=100)
interact(plot_sample_stats, n=slider, xlim=fixed([55, 95]))
None
# -
# ### Other sample statistics
#
# This framework works with any other quantity we want to estimate. By changing `sample_stat`, you can compute the SE and CI for any sample statistic.
#
# **Exercise 1**: Fill in `sample_stat` below with any of these statistics:
#
# * Standard deviation of the sample.
# * Coefficient of variation, which is the sample standard deviation divided by the sample standard mean.
# * Min or Max
# * Median (which is the 50th percentile)
# * 10th or 90th percentile.
# * Interquartile range (IQR), which is the difference between the 75th and 25th percentiles.
#
# NumPy array methods you might find useful include `std`, `min`, `max`, and `percentile`.
# Depending on the results, you might want to adjust `xlim`.
# +
def sample_stat(sample):
# TODO: replace the following line with another sample statistic
return sample.mean()
slider = widgets.IntSlider(min=10, max=1000, value=100)
interact(plot_sample_stats, n=slider, xlim=fixed([0, 100]))
None
# -
# STOP HERE
# ---------
#
# We will regroup and discuss before going on.
# Part Two
# ========
#
# So far we have shown that if we know the actual distribution of the population, we can compute the sampling distribution for any sample statistic, and from that we can compute SE and CI.
#
# But in real life we don't know the actual distribution of the population. If we did, we wouldn't need to estimate it!
#
# In real life, we use the sample to build a model of the population distribution, then use the model to generate the sampling distribution. A simple and popular way to do that is "resampling," which means we use the sample itself as a model of the population distribution and draw samples from it.
#
# Before we go on, I want to collect some of the code from Part One and organize it as a class. This class represents a framework for computing sampling distributions.
class Resampler(object):
"""Represents a framework for computing sampling distributions."""
def __init__(self, sample, xlim=None):
"""Stores the actual sample."""
self.sample = sample
self.n = len(sample)
self.xlim = xlim
def resample(self):
"""Generates a new sample by choosing from the original
sample with replacement.
"""
new_sample = numpy.random.choice(self.sample, self.n, replace=True)
return new_sample
def sample_stat(self, sample):
"""Computes a sample statistic using the original sample or a
simulated sample.
"""
return sample.mean()
def compute_sample_statistics(self, iters=1000):
"""Simulates many experiments and collects the resulting sample
statistics.
"""
stats = [self.sample_stat(self.resample()) for i in range(iters)]
return numpy.array(stats)
def plot_sample_stats(self):
"""Runs simulated experiments and summarizes the results.
"""
sample_stats = self.compute_sample_statistics()
summarize_sampling_distribution(sample_stats)
pyplot.hist(sample_stats, color=COLOR2)
pyplot.xlabel('sample statistic')
pyplot.xlim(self.xlim)
# The following function instantiates a `Resampler` and runs it.
def plot_resampled_stats(n=100):
sample = weight.rvs(n)
resampler = Resampler(sample, xlim=[55, 95])
resampler.plot_sample_stats()
# Here's a test run with `n=100`
plot_resampled_stats(100)
# Now we can use `plot_resampled_stats` in an interaction:
slider = widgets.IntSlider(min=10, max=1000, value=100)
interact(plot_resampled_stats, n=slider, xlim=fixed([1, 15]))
None
# **Exercise 2**: write a new class called `StdResampler` that inherits from `Resampler` and overrides `sample_stat` so it computes the standard deviation of the resampled data.
# +
# Solution goes here
# -
# Test your code using the cell below:
# +
def plot_resampled_stats(n=100):
sample = weight.rvs(n)
resampler = StdResampler(sample, xlim=[0, 100])
resampler.plot_sample_stats()
plot_resampled_stats()
# -
# When your `StdResampler` is working, you should be able to interact with it:
slider = widgets.IntSlider(min=10, max=1000, value=100)
interact(plot_resampled_stats, n=slider)
None
# STOP HERE
# ---------
#
# We will regroup and discuss before going on.
# Part Three
# ==========
#
# We can extend this framework to compute SE and CI for a difference in means.
#
# For example, men are heavier than women on average. Here's the women's distribution again (from BRFSS data):
female_weight = scipy.stats.lognorm(0.23, 0, 70.8)
female_weight.mean(), female_weight.std()
# And here's the men's distribution:
male_weight = scipy.stats.lognorm(0.20, 0, 87.3)
male_weight.mean(), male_weight.std()
# I'll simulate a sample of 100 men and 100 women:
female_sample = female_weight.rvs(100)
male_sample = male_weight.rvs(100)
# The difference in means should be about 17 kg, but will vary from one random sample to the next:
male_sample.mean() - female_sample.mean()
# Here's the function that computes Cohen's effect size again:
def CohenEffectSize(group1, group2):
"""Compute Cohen's d.
group1: Series or NumPy array
group2: Series or NumPy array
returns: float
"""
diff = group1.mean() - group2.mean()
n1, n2 = len(group1), len(group2)
var1 = group1.var()
var2 = group2.var()
pooled_var = (n1 * var1 + n2 * var2) / (n1 + n2)
d = diff / numpy.sqrt(pooled_var)
return d
# The difference in weight between men and women is about 1 standard deviation:
CohenEffectSize(male_sample, female_sample)
# Now we can write a version of the `Resampler` that computes the sampling distribution of $d$.
class CohenResampler(Resampler):
def __init__(self, group1, group2, xlim=None):
self.group1 = group1
self.group2 = group2
self.xlim = xlim
def resample(self):
group1 = numpy.random.choice(self.group1, len(self.group1), replace=True)
group2 = numpy.random.choice(self.group2, len(self.group2), replace=True)
return group1, group2
def sample_stat(self, groups):
group1, group2 = groups
return CohenEffectSize(group1, group2)
# NOTE: The following functions are the same as the ones in Resampler,
# so I could just inherit them, but I'm including them for readability
def compute_sample_statistics(self, iters=1000):
stats = [self.sample_stat(self.resample()) for i in range(iters)]
return numpy.array(stats)
def plot_sample_stats(self):
sample_stats = self.compute_sample_statistics()
summarize_sampling_distribution(sample_stats)
pyplot.hist(sample_stats, color=COLOR2)
pyplot.xlabel('sample statistic')
pyplot.xlim(self.xlim)
# Now we can instantiate a `CohenResampler` and plot the sampling distribution.
resampler = CohenResampler(male_sample, female_sample)
resampler.plot_sample_stats()
# This example demonstrates an advantage of the computational framework over mathematical analysis. Statistics like Cohen's $d$, which is the ratio of other statistics, are relatively difficult to analyze. But with a computational approach, all sample statistics are equally "easy".
#
# One note on vocabulary: what I am calling "resampling" here is a specific kind of resampling called "bootstrapping". Other techniques that are also considering resampling include permutation tests, which we'll see in the next section, and "jackknife" resampling. You can read more at <http://en.wikipedia.org/wiki/Resampling_(statistics)>.
| sampling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from tqdm.auto import tqdm
# -
tqdm.pandas()
pd.set_option("display.max_rows", 1000)
data = pd.read_csv("uiks-utf8.csv")
data['pct_turnout'] = (data.found_ballots / data.registered_voters) * 100
data['pct_yes'] = (data.yes_votes / data.found_ballots) * 100
data.head(2)
tol = 1e-2
indicators = {
'CloseDivisable@5': lambda x: int(np.isclose([x.pct_yes % 5.0],[0], atol=tol) or np.isclose([x.pct_turnout % 5.0], [0], atol=tol)),
'CloseDivisable@1': lambda x: int((np.isclose([x.pct_yes % 1.0],[0], atol=tol) or np.isclose([x.pct_turnout % 1.0], [0], atol=tol)) and (x.used_ballots % 10.0 == 0 or x.yes_votes % 10.0 == 0))
}
for key, fn in indicators.items():
data[key] = data.progress_apply(fn, axis=1)
data['AnyDivisable'] = data.progress_apply(lambda x: np.max([x['CloseDivisable@5'], x['CloseDivisable@1']]), axis=1)
fraud_full = data[data['CloseDivisable@5'] == 1]
print(f"Всего участков в ровными процентами: {len(fraud_full)}")
print(f"Сумма проголосовавших на таких участках: {fraud_full['found_ballots'].sum()}")
print(f"{(fraud_full['found_ballots'].sum() / data['found_ballots'].sum()) * 100:.2f}% от общего числа")
print(f"Процентное соотношение голосов за\против: " +
f"{fraud_full['pct_yes'].mean():.2f}% ЗА, " +
f"{((fraud_full.no_votes / data.found_ballots) * 100).mean():.2f}% Против, " +
f"{((fraud_full.incorrect_ballots / fraud_full.found_ballots) * 100).mean():.2f}% Испорченных")
len(fraud_full[fraud_full.region_name == 'Нижегородская область']) / len(data[data.region_name == "Нижегородская область"])
len(fraud_full[fraud_full.region_name == 'Территория за пределами РФ']) / len(data[data.region_name == "Территория за пределами РФ"])
fraud_full.region_name.value_counts()
fraud_full[fraud_full.tik_name == "39 Нижний Новгород, Нижегородская"]['found_ballots'].value_counts()
fraud = data[data.registered_voters > 500][data['CloseDivisable@5'] == 1]
print(f"Всего участков с процентом явки\голосов за, делящимся на 5: {len(fraud)}")
fraud.region_name.value_counts()
fraud[fraud.region_name == "город Москва"]
| FraudScoring.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
#
#
# [This repository](https://github.com/tamura70/taocp-sat/) provides Linux binaries and PDF files produced from the CWEB programs written by [<NAME>](https://www-cs-faculty.stanford.edu/~knuth/) for "Satisfiability", Volume 4 Fascicle 6 of [The Art of Computer Programming](https://www-cs-faculty.stanford.edu/~knuth/taocp.html) (TAOCP). The original CWEB programs were obtained on March 19, 2019 from:
#
# - [https://www-cs-faculty.stanford.edu/~knuth/programs.html](https://www-cs-faculty.stanford.edu/~knuth/programs.html)
#
# They were compiled on Ubuntu 18.04 with TeX Live 2018 by using the following Perl script.
#
# - [http://bach.istc.kobe-u.ac.jp/lect/taocp-sat/knuth/install.pl](http://bach.istc.kobe-u.ac.jp/lect/taocp-sat/knuth/install.pl)
#
# There were some errors during the compilation (see the following log file). Therefore, it is not guaranteed the compiled programs and PDFs are correct.
#
# - [http://bach.istc.kobe-u.ac.jp/lect/taocp-sat/knuth/install.log](http://bach.istc.kobe-u.ac.jp/lect/taocp-sat/knuth/install.log)
#
# These programs can be executed in [Jupyter](http://jupyter.org) notebooks on [Binder](https://mybinder.org) with [Almond](https://almond.sh) Scala kernel.
#
# - [https://mybinder.org/v2/gh/tamura70/taocp-sat/master?urlpath=lab/tree/notebooks/index.ipynb>](https://mybinder.org/v2/gh/tamura70/taocp-sat/master?urlpath=lab/tree/notebooks/index.ipynb>)(in Japanese)
#
# See the following web page for more details of how to run Knuth's programs.
#
# - [Knuth先生の『TAOCP 7.2.2.2 Satisfiability』を読む](http://bach.istc.kobe-u.ac.jp/lect/taocp-sat/) (in Japanese)
#
#
# ## Credits/Links
#
#
# Binder setup scripts are obtained from [Almond Examples](https://github.com/almond-sh/examples).
#
# - [Almond](https://almond.sh)
# - [Jupyter](http://jupyter.org)
# - [Binder](https://mybinder.org)
#
#
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multibin Coupled NormSys
# %pylab inline
# +
import pyhf
import logging
logging.basicConfig(level=logging.INFO)
from pyhf import Model
def prep_data(sourcedata):
spec = {
'signal': {
'signal': {
'data': sourcedata['signal']['bindata']['sig'],
'mods': [{'name': 'mu', 'type': 'normfactor', 'data': None}],
},
'bkg1': {
'data': sourcedata['signal']['bindata']['bkg1'],
'mods': [
{
'name': 'coupled_normsys',
'type': 'normsys',
'data': {'lo': 0.9, 'hi': 1.1},
}
],
},
'bkg2': {
'data': sourcedata['signal']['bindata']['bkg2'],
'mods': [
{
'name': 'coupled_normsys',
'type': 'normsys',
'data': {'lo': 0.5, 'hi': 1.5},
}
],
},
},
'control': {
'background': {
'data': sourcedata['control']['bindata']['bkg1'],
'mods': [
{
'name': 'coupled_normsys',
'type': 'normsys',
'data': {'lo': 0.9, 'hi': 1.1},
}
],
}
},
}
pdf = Model(spec)
data = []
for c in pdf.config.channel_order:
data += sourcedata[c]['bindata']['data']
data = data + pdf.config.auxdata
return data, pdf
# +
source = {
"channels": {
"signal": {
"binning": [2, -0.5, 1.5],
"bindata": {
"data": [105.0, 220.0],
"bkg1": [100.0, 100.0],
"bkg2": [50.0, 100.0],
"sig": [10.0, 35.0],
},
},
"control": {
"binning": [2, -0.5, 1.5],
"bindata": {"data": [110.0, 105.0], "bkg1": [100.0, 100.0]},
},
}
}
d, pdf = prep_data(source['channels'])
print(d)
init_pars = pdf.config.suggested_init()
par_bounds = pdf.config.suggested_bounds()
unconpars = pyhf.unconstrained_bestfit(d, pdf, init_pars, par_bounds)
print('UNCON', unconpars)
conpars = pyhf.constrained_bestfit(0.0, d, pdf, init_pars, par_bounds)
print('CONS', conpars)
pdf.expected_data(conpars)
# +
def plot_results(testmus, cls_obs, cls_exp, test_size=0.05):
plt.plot(mutests, cls_obs, c='k')
for i, c in zip(range(5), ['grey', 'grey', 'grey', 'grey', 'grey']):
plt.plot(mutests, cls_exp[i], c=c)
plt.plot(testmus, [test_size] * len(testmus), c='r')
plt.ylim(0, 1)
def invert_interval(testmus, cls_obs, cls_exp, test_size=0.05):
point05cross = {'exp': [], 'obs': None}
for cls_exp_sigma in cls_exp:
yvals = [x for x in cls_exp_sigma]
point05cross['exp'].append(
np.interp(test_size, list(reversed(yvals)), list(reversed(testmus)))
)
yvals = cls_obs
point05cross['obs'] = np.interp(
test_size, list(reversed(yvals)), list(reversed(testmus))
)
return point05cross
pyhf.runOnePoint(1.0, d, pdf, init_pars, par_bounds)[-2:]
mutests = np.linspace(0, 5, 61)
tests = [
pyhf.runOnePoint(muTest, d, pdf, init_pars, par_bounds)[-2:] for muTest in mutests
]
cls_obs = [test[0] for test in tests]
cls_exp = [[test[1][i] for test in tests] for i in range(5)]
plot_results(mutests, cls_obs, cls_exp)
invert_interval(mutests, cls_obs, cls_exp)
| docs/examples/notebooks/multichannel-coupled-normsys.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
First insights:
* If an input is trained, it will stay stable, even if it receives less inputs in a second run
* If an input was trained less in a first run, it will strenghten its connections later if its stimuated more often then
* But in this case it will take much more time, since only few spikes are available which takes some time to build up
* If both inputs are trained strongly, further activating patterns have not really any impact anymore
-> An input aquires a specific amount of neurons and stably saturates
-> If it's relatively weak in the beginning it can still become saturates, but then it takes longer
Next steps:
* What happens when the network is bigger?
* What happens when the input it smaller/bigger?
* What happens when a third input comes into play?
"""
# +
# Allow reload of objects
# %load_ext autoreload
# %autoreload
from pelenet.utils import Utils
from pelenet.experiments.assemblies import AssemblyExperiment
# Official modules
import numpy as np
import nxsdk.api.n2a as nx
import matplotlib
import matplotlib.pyplot as plt
from time import time
from copy import deepcopy
# -
# Overwrite default parameters (pelenet/parameters/ and pelenet/experiments/random.py)
parameters = {
# Experiment
'seed': 1, # Random seed
'trials': 20, # Number of trials
'stepsPerTrial': 50, # Number of simulation steps for every trial
'isReset': True, # Activate reset after every trial
# Network
'reservoirExSize': 400, # Number of excitatory neurons
'reservoirConnPerNeuron': 35, #35 # Number of connections per neuron
# Neurons
'refractoryDelay': 2, # Refactory period
'voltageTau': 100, #100 # Voltage time constant
'currentTau': 6, #5 # Current time constant
'thresholdMant': 1500, #1200 # Spiking threshold for membrane potential
# Plasticity
'isLearningRule': True,
'learningRule': '2^-2*x1*y0 - 2^-2*y1*x0 + 2^-4*x1*y1*y0 - 2^-3*y0*w*w',
# Input
'inputIsVary': True,
'inputVaryProbs': [0.1, 0.9],
'inputGenSpikeProb': 0.8, # Probability of spikes for the spike generators
'inputNumTargetNeurons': 100, # Number of neurons targeted by the spike generators
'inputSteps': 30, # Number of steps the input is active
# Probes
'isExSpikeProbe': True, # Probe excitatory spikes
'isInSpikeProbe': True, # Probe inhibitory spikes
'isWeightProbe': True # Probe weight matrix at the end of the simulation
}
# Initilizes the experiment and utils
exp = AssemblyExperiment(name='assemblies', parameters=parameters)
utils = Utils.instance()
# +
# Build network
exp.build()
# Get spectral radius
utils.getSpectralRadius(exp.net.initialWeights)
# -
# Run network
exp.run()
# Plot spike trains of the excitatory (red) and inhibitory (blue) neurons
exp.net.plot.reservoirSpikeTrain(figsize=(12,6))
# Weight matrix before learning
exp.net.plot.initialExWeightMatrix()
# Weight matrix after learning
exp.net.plot.trainedExWeightMatrix()
# Sorted weight matrix after learning
expSupportMask = utils.getSupportWeightsMask(exp.net.trainedWeightsExex)
exp.net.plot.weightsSortedBySupport(expSupportMask)
# New parameters
newParameters = {
**parameters,
'trials': 20,
'inputVaryProbs': [0.9, 0.1]
}
# Initilizes a second experiment
exp2 = AssemblyExperiment(name='assemblies-retrain', parameters=newParameters)
# +
#np.array_equal(exp.initialMasks.exex.toarray(), exp2.initialMasks.exex.toarray())
# -
# Build network
weights = deepcopy(exp.net.initialWeights)
weights.exex = exp.net.trainedWeightsExex
exp2.buildWithGivenMaskAndWeights(exp.net.initialMasks, weights)
# Run network
exp2.run()
# Plot spike trains of the excitatory (red) and inhibitory (blue) neurons
exp2.net.plot.reservoirSpikeTrain(figsize=(12,6))
# Weight matrix before learning
exp2.net.plot.initialExWeightMatrix()
# Weight matrix after learning
exp2.net.plot.trainedExWeightMatrix()
# Sorted weight matrix after learning
exp2SupportMask = utils.getSupportWeightsMask(exp2.net.trainedWeightsExex)
exp2.net.plot.weightsSortedBySupport(exp2SupportMask)
| assemblies-bigger-input.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='Top'></a>
#
# # Choice of output intervals<a class='tocSkip'></a>
#
# Length and number of output intervals does not influence accuracy.
# + code_folding=[]
# %load_ext autoreload
# %autoreload 2
# %load_ext watermark
import sys
import os
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
import torch
from lifelines import KaplanMeierFitter
# Make modules in "src" dir visible`
project_dir = os.path.split(os.getcwd())[0]
if project_dir not in sys.path:
sys.path.append(os.path.join(project_dir, 'src'))
import dataset
from model import Model
import utils
matplotlib.style.use('multisurv.mplstyle')
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#DataLoader" data-toc-modified-id="DataLoader-1"><span class="toc-item-num">1 </span><code>DataLoader</code></a></span></li><li><span><a href="#Model" data-toc-modified-id="Model-2"><span class="toc-item-num">2 </span>Model</a></span><ul class="toc-item"><li><span><a href="#Evaluate" data-toc-modified-id="Evaluate-2.1"><span class="toc-item-num">2.1 </span>Evaluate</a></span></li></ul></li><li><span><a href="#Plot" data-toc-modified-id="Plot-3"><span class="toc-item-num">3 </span>Plot</a></span><ul class="toc-item"><li><span><a href="#Save-to-file" data-toc-modified-id="Save-to-file-3.1"><span class="toc-item-num">3.1 </span>Save to file</a></span></li></ul></li></ul></div>
# +
DATA = utils.INPUT_DATA_DIR
MODELS = utils.TRAINED_MODEL_DIR
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# -
# # `DataLoader`
dataloaders = utils.get_dataloaders(
data_location=DATA,
labels_file='../data/labels.tsv',
modalities=['clinical', 'mRNA'],
# wsi_patch_size=299,
# n_wsi_patches=5,
# exclude_patients=exclude_cancers,
return_patient_id=True
)
# # Model
#
#
# Try several different discretization schemes of the continuous time scale with two different approaches: equidistant times (several variations) or equidistant survival probabilities (propsed by [<NAME> Borgan, 2019](https://arxiv.org/abs/1910.06724)).
#
# The survival probability scheme is based on the distribution of the event times. Estimate the survival function using the Kaplan-Meier estimator and make a grid of equidistant estimates (corresponding to quantiles of the input times). The result is a more dense grid in intervals with more events.
# +
labels = [(t, e) for t, e in dataloaders['train'].dataset.label_map.values()]
durations = [t for t, _ in labels]
events = [e for _, e in labels]
survival_prob_interval_cuts = utils.discretize_time_by_duration_quantiles(durations, events, 20)
survival_prob_interval_cuts = torch.from_numpy(survival_prob_interval_cuts)
# -
# Interval - saved weight pairs
int_and_weights = {
'30 one-year intervals':
{'intervals': torch.arange(0., 365 * 31, 365),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_epoch43_acc0.81.pth')},
'20 one-year intervals':
{'intervals': torch.arange(0., 365 * 21, 365),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_20_1year_intervals_epoch35_acc0.80.pth')},
'10 one-year intervals':
{'intervals': torch.arange(0., 365 * 11, 365),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_10_1year_intervals_epoch37_acc0.80.pth')},
'20 half-year intervals':
{'intervals': torch.arange(0., 365 * 10.1, 365 / 2),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_20_half-year_intervals_epoch47_acc0.80.pth')},
'5 one-year intervals':
{'intervals': torch.arange(0., 365 * 6, 365),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_5_1year_intervals_epoch39_acc0.80.pth')},
'10 half-year intervals':
{'intervals': torch.arange(0., 365 * 5.1, 365 / 2),
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_10_half-year_intervals_epoch35_acc0.79.pth')},
'20 variable intervals':
{'intervals': survival_prob_interval_cuts,
'weights': os.path.join(MODELS, 'clinical_mRNA_lr0.005_discretized_by_duration_quantiles_epoch50_acc0.79.pth')}
}
# +
models = {}
for x in int_and_weights:
models[x] = Model(
dataloaders=dataloaders,
output_intervals=int_and_weights[x]['intervals'],
device=device)
models[x].load_weights(int_and_weights[x]['weights'])
print()
# -
# ## Evaluate
#
# $\text{C}^{\text{td}}$ values were obtained using the Jupyter notebook used to evaluate trained models ([table-multisurv_cindices](table-multisurv_evaluation.ipynb)).
c_td = {
'30 one-year intervals': '0.822 (0.806-0.839)',
'20 one-year intervals': '0.809 (0.792-0.826)',
'10 one-year intervals': '0.810 (0.794-0.828)',
'20 half-year intervals': '0.816 (0.800-0.832)',
'5 one-year intervals': '0.816 (0.800-0.833)',
'10 half-year intervals': '0.817 (0.801-0.834)',
'20 variable intervals': '0.810 (0.793-0.828)',
}
# +
# %%time
results = {}
for model in models:
print(f' {model} -', end=' ')
results[model] = models[model].predict_dataset(dataloaders['test'].dataset)
print()
print()
# -
# # Plot
utils.plot.show_default_colors()
def predictions_to_pandas(data):
labels = pd.read_csv('../data/labels.tsv', sep='\t')
predictions = np.stack([x[0] for x in data['patient_data'].values()])
# Add prediction 1 at time 0 (patients still alive)
ones = np.ones((predictions.shape[0], 1))
predictions = np.concatenate((ones, predictions), axis=1)
predictions = pd.DataFrame(predictions, columns=[str(i) for i in range(predictions.shape[-1])])
patient_ids = list(data['patient_data'].keys())
predictions['submitter_id'] = patient_ids
predictions = predictions.merge(labels.iloc[:, :-1])
predictions.set_index('submitter_id', inplace=True)
return predictions
def get_KM_estimates(data, return_kmfitter=False):
kmf = KaplanMeierFitter()
kmf.fit(durations=data['time'].values,
event_observed=data['event'].values)
if return_kmfitter:
return kmf
x = kmf.survival_function_.index.values
y = kmf.survival_function_
return x, y
# Select and order six models
models_to_use = ['30 one-year intervals', '10 one-year intervals', '5 one-year intervals',
'20 half-year intervals', '10 half-year intervals', '20 variable intervals']
# +
default_colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
fig, axs = plt.subplots(2, 3, figsize=(18, 6))
# fig.suptitle('Pan-cancer patient survival')
axs = axs.ravel()
for i, model in enumerate(models_to_use):
predictions = predictions_to_pandas(data=results[model])
x, y = get_KM_estimates(predictions)
axs[i].plot(x, y, '--', color=default_colors[-2], label='Kaplan-Meier')
time_points = utils.plot.format_output_intervals(models[model].output_intervals)
len(time_points)
mean_probabilities = predictions[[str(i) for i in range(len(time_points))]].mean(axis=0).values
axs[i].plot(time_points, mean_probabilities, 'o', color=default_colors[0],
markersize=5, label='MultiSurv')
axs[i].set_title(model + '\n' + r'$\mathrm{C}^{\mathrm{td}}$' + f': {c_td[model]}')
axs[i].set_xlim(None, 31)
if i == len(models_to_use) - 1:
axs[i].legend()
for ax in axs.flat:
ax.set(xlabel='Time (years)', ylabel='Survival probability')
ax.label_outer()
plt.subplots_adjust(top=0.82, hspace=0.6, wspace=0.1)
# -
# ## Save to file
# + language="javascript" active=""
# IPython.notebook.kernel.execute('nb_name = "' + IPython.notebook.notebook_name + '"')
# + active=""
# pdf_file = nb_name.split('.ipynb')[0]
# utils.plot.save_plot_for_figure(figure=fig, file_name=pdf_file)
# -
# # Watermark<a class='tocSkip'></a>
# %watermark --iversions
# %watermark -v
print()
# %watermark -u -n
# [Top of the page](#Top)
| figures_and_tables/figure-choice_of_output_intervals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NAVIGATION-->
# < [Cruise Spatio-Temporal Bounds](CruiseBounds.ipynb) | [Index](Index.ipynb) | [Cruise Variables](CruiseVariables.ipynb) >
#
# <a href="https://colab.research.google.com/github/simonscmap/pycmap/blob/master/docs/CruiseTrajectory.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# <a href="https://mybinder.org/v2/gh/simonscmap/pycmap/master?filepath=docs%2FCruiseTrajectory.ipynb"><img align="right" src="https://mybinder.org/badge_logo.svg" alt="Open in Colab" title="Open and Execute in Binder"></a>
# ## *cruise_trajectory(cruiseName)*
#
# Returns a dataframe containing the trajectory of the specified cruise.
# > **Parameters:**
# >> **cruiseName: string**
# >> <br />The official cruise name. If applicable, you may also use cruise "nickname" ('Diel', 'Gradients_1' ...). <br />A full list of cruise names can be retrieved using [cruise](Cruises.ipynb) method.
#
#
# >**Returns:**
# >> Pandas dataframe.
# ### Example
# +
# #!pip install pycmap -q #uncomment to install pycmap, if necessary
import pycmap
api = pycmap.API(token='<YOUR_API_KEY>')
api.cruise_trajectory('KM1513')
# -
# <img src="figures/sql.png" alt="SQL" align="left" width="40"/>
# <br/>
# ### SQL Statement
# Here is how to achieve the same results using a direct SQL statement. Please refere to [Query](Query.ipynb) for more information.
# <code>EXEC uspCruiseTrajectoryByName 'Cruise Official Name'</code>
#
#
# **Example:**<br/>
# <code>EXEC uspCruiseTrajectoryByName 'KM1513'</code>
| docs/CruiseTrajectory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datetime variables in Python and Pandas
#
# Date and time features are important for data science problems in various industries, from sales, marketing, and finance to HR, e-commerce, retail, and many more. Predicting how the stock markets will behave tomorrow, how many products will be sold in the upcoming week, when is the best time to launch a new product, how long before a position at the company gets filled, etc. are some of the problems that we can find answers to using date and time data.
#
# This incredible amount of insight that you can unravel from the data is what makes date and time components so fun to work with! So let’s get down to the business of mastering date-time manipulation in Python.
#
# But in many cases dates and time are not in the right format and the information in those variables can't be worked with until it is transformed.
#
# In the following notebook you will learn how to work with dates and time in `Python` and `pandas`.
import pandas as pd
from datetime import datetime, date, time, timedelta
# ### Python Datetime Module
#
# First, we will have a look at the datetime module of Python. Let's start simple. We will look at the basics of the module and how to define dates and time.
# +
# We create a date variable for a random date with the datetime module and as a string…
d1 = date(2021, 5, 14)
d2 = '2021-05-14'
# …and print out the variables. Do you see any differences?
print(d1)
print(d2)
# -
# If we have a look at the types of those two variables we can see that there is a difference.
print(type(d1))
print(type(d2))
# +
# You can also easily create a variable with the date of today.
d3 = date.today()
print('Date of today: ',d3)
# You can print the day…
print('Day: ', d3.day)
# …or month…
print('Month: ', d3.month)
# …or year.
print('Year: ', d3.year)
# -
# The same applies to the time method.
t1 = time(13, 20, 13, 40)
t2 = '13:20:13.000040'
print(t1)
print(t2)
print(type(t1))
print(type(t2))
# +
# You can print the hour…
print('Hour :',t1.hour)
# …or minute…
print('Minute :',t1.minute)
# …or second…
print('Second :',t1.second)
# …and even the microseconds.
print('Microsecond :',t1.microsecond)
# -
# Above we looked at the date and time seperatly with the `date` and `time` methods. But we can also combine both with the `datetime` method.
# +
# We create a variable with a random date and time with the datetime method…
dt1 = datetime(2021, 5, 14, 11, 20, 30, 40)
# …and print it.
print(dt1)
print(type(dt1))
# -
# You can also create a variable with your local date-time.
dt2 = datetime.now()
dt2
# One really cool thing you can do with the DateTime function is to extract the day of the week! This is especially helpful in feature engineering because the value of the target variable can be dependent on the day of the week, like sales of a product are generally higher on a weekend or traffic on StackOverflow could be higher on a weekday when people are working, etc.
#
# >Note: `.weekday` returns the day of the week starting with 0 for Monday and ending with 6 for Sunday. `.isoweekday` on the other hand returns the day of the week where Monday is 1 and Sunday is 7.
# +
# We can define the date of today…
dt2 = datetime.now()
# …and print the weekday where Monday is 0…
print(dt2.weekday()) # outputs e.g. 3 for Thurday
# …or where Monday is 1.
print(dt2.isoweekday()) # outputs 4 in ISO format
# -
# Another very important feature that you can generate from the given date in a dataset is the week of the year.
# +
# We can define the date of today…
dt2 = datetime.now()
# …and retun the year, week and weekday.
print('Date:', dt2.isocalendar())
print('Week :', dt2.isocalendar()[1])
# -
# ### Formatting the dates with `strptime` and `strftime`
#
# As we saw in the first example, you have to pay attention to the type of the data. The date in datasets often is of type string. So how can we change that?
#
# - `strptime`: creates a `datetime` object from a string representing date and time. It takes two arguments: the date and the format in which your date is present.
# - `strftime`: can be used to convert the `datetime` object into a string representing date and time.
#
# The following table shows several format codes that can be used to specify a date or time format.
#
#
# | Directive | Meaning | Example |
# |: ------------- :|:-------------:| -----:|
# | %a | Abbreviated weekday name. | Sun, Mon, ...|
# | %A | Full weekday name. | Sunday, Monday, ... |
# | %w | Weekday as a decimal number. | 0, 1, ..., 6 |
# | %d | Day of the month as a zero-padded decimal. | 01, 02, ..., 31 |
# | %-d | Day of the month as a decimal number.| 1, 2, ..., 30|
# | %b | Abbreviated month name.| Jan, Feb, ..., Dec|
# | %B | Full month name.| January, February, ...|
# | %m | Month as a zero-padded decimal number.| 01, 02, ..., 12|
# | %-m | Month as a decimal number.| 1, 2, ..., 12|
# | %y | Year without century as a zero-padded decimal number.| 00, 01, ..., 99|
# | %-y | Year without century as a decimal number.| 0, 1, ..., 99|
# | %Y | Year with century as a decimal number.| 2013, 2019 etc.|
# | %H | Hour (24-hour clock) as a zero-padded decimal number.| 00, 01, ..., 23|
# | %-H | Hour (24-hour clock) as a decimal number.| 0, 1, ..., 23|
# | %I | Hour (12-hour clock) as a zero-padded decimal number.| 01, 02, ..., 12|
# | %-I | Hour (12-hour clock) as a decimal number.| 1, 2, ... 12|
# | %p| Locale’s AM or PM.| AM, PM|
# | %M| Minute as a zero-padded decimal number.| 00, 01, ..., 59|
# | %-M| Minute as a decimal number.| 0, 1, ..., 59|
# | %S| Second as a zero-padded decimal number.| 00, 01, ..., 59|
# | %-S| Second as a decimal number.| 0, 1, ..., 59|
# | %f| Microsecond as a decimal number, zero-padded on the left.| 000000 - 999999|
# | %z| UTC offset in the form +HHMM or -HHMM.| |
# | %Z| Time zone name.| |
# | %j| Day of the year as a zero-padded decimal number.| 001, 002, ..., 366|
# | %-j| Day of the year as a decimal number.| 1, 2, ..., 366|
# | %U| Week number of the year (Sunday as the first day of the week). All days in a new year preceding the first Sunday are considered to be in week 0.| 00, 01, ..., 53|
# | %W| Week number of the year (Monday as the first day of the week). All days in a new year preceding the first Monday are considered to be in week 0.| 00, 01, ..., 53|
# | %c| Locale’s appropriate date and time representation.| Mon Sep 30 07:06:05 2013|
# | %x| Locale’s appropriate date representation.| 09/30/13|
# | %X| Locale’s appropriate time representation.| 07:06:05|
# | %%| A literal '%' character.| %|
# #### Strptime
#
# With `strptime` and the right combination of the format codes (see list above) you can convert a string into a datetime type:
# +
# Create a variable with a datetime stored as string…
date = '22 April, 2020 13:20:13'
# …and convert the string to datetime type.
d1 = datetime.strptime(date,'%d %B, %Y %H:%M:%S')
print(d1)
print(type(d1))
# +
# We can create a date stored as string…
date = '2021-05-14'
print(date)
print(type(date))
# …and convert the string to datetime…
d2 = datetime.strptime(date, '%Y-%m-%d')
print(d2)
print(type(d2))
# …or onvert the string to date format.
d2 = datetime.strptime(date, '%Y-%m-%d').date()
print(d2)
print(type(d2))
# -
# #### Strftime
#
# If we want to convert a `datetime` variable into a string we can use `.strftime` and the right combination of format codes:
# +
# Create a datetime variable…
d1 = datetime.now()
print('Datetime object :',d1)
# …and convert it into a string.
new_date = d1.strftime('%d/%m/%Y %H:%M')
print('Formatted date :',new_date)
print(type(new_date))
# -
# ### Calculating with the datetime module
#
# Sometimes, you might have to find the time span between two dates, which can be another very useful feature that you can derive from a dataset. By substracting one date from another you will get a `timedelta`.
# timedelta : duration between dates
d1 = datetime(2020,4,23,11,13,10)
d2 = datetime(2021,4,23,12,13,10)
duration = d2 - d1
print(type(duration))
duration
# As you can see, the duration is returned as the number of days for the date and seconds for the time between the dates. So for your features you can also retrieve those values separately:
print('Duration:', duration)
print('Days:', duration.days) # 365
print('Seconds:' ,duration.seconds) # 3600
# But what if you actually need the duration in hours or minutes? Well, there is a simple solution for that.
#
# `timedelta` is also a class in the `datetime` module. So, you can use it to convert your duration into hours and minutes as the following code shows:
# +
# duration in hours
print('Duration in hours :', duration / timedelta(hours = 1))
# duration in minutes
print('Duration in minutes :', duration / timedelta(minutes = 1))
# duration in seconds
print('Duration in seconds :', duration / timedelta(seconds = 1))
# -
# `timedelta` also makes it possible to add and subtract integers from a `datetime` object.
# +
# Create datetime variable for current date…
d1 = datetime.now().date()
print("Today's date :",d1)
# …and add two days…
d2 = d1+timedelta(days=2)
print("Date 2 days from today :",d2)
# …or add two weeks.
d3 = d1+timedelta(weeks=2)
print("Date 2 weeks from today :",d3)
# -
# ## DateTime in Pandas
#
# We already know that Pandas is a great library for doing data analysis tasks. And so it goes without saying that Pandas also supports Python DateTime objects. It has some great methods for handling dates and times, such as `to_datetime()` and `to_timedelta()`.
# Define a date using .to_datetime()
date = pd.to_datetime('24th of April, 2020')
print(date)
print(type(date))
# You might have noticed something strange here. The type of the object returned by `.to_datetime()` is not DateTime but Timestamp. Well, don’t worry, it is just the Pandas equivalent of Python’s DateTime.
#
# We already know that timedelta gives differences in times. The Pandas `.to_timedelta()` method does the same:
# +
# Define and print a variable with the current date and time…
date = datetime.now()
print(date)
# …and print the date of tomorrow…
print(date + pd.to_timedelta(1, unit = 'D'))
# …and next week.
print(date + pd.to_timedelta(1, unit = 'W'))
# -
# ### Date Range in Pandas
#
# To make the creation of date sequences a convenient task, Pandas provides the `.date_range()` method. It accepts a start date, an end date, and an optional frequency code:
# Create a range of dates for 1 month
pd.date_range(start='24/4/2020', end='24/5/2020', freq='D')
# Instead of defining the end date, you could also define the period or number of time periods you want to generate: [See here for a list of frequency aliases.]( https://pandas.pydata.org/docs/user_guide/timeseries.html#timeseries-offset-aliases)
# +
# Define variable with date of today…
start_date = datetime.today()
# …and create sequence with consecutive minutes…
dates_start = pd.date_range(start=start_date, periods=10, freq='T')
dates_start
# -
# …or consecutive days.
dates_end = pd.date_range(start=start_date, periods=10, freq='D')
dates_end
# Let's have a look at how to create new features out of `datetime/timestamp` columns in a Pandas DataFrame. Therefore, we will create a little DataFrame which contains the `start_date` and `end_date` (see cells above) as features and a target column with three different classes.
# +
# Create target column with random classes (1, 2 or 3)
import random
randomList = []
for i in range(10):
randomList.append(random.randint(1,3))
# Create DataFrame out of previously defined variables
df = pd.DataFrame()
df['Start_date'] = dates_start
df['End_date'] = dates_end
df['Target'] = randomList
# Show first 5 rows
df.head()
# -
# The elemnts in the date columns are still timestamps
print(df.Start_date[0])
print(type(df.Start_date[0]))
# We can create multiple new features from the date column, like the day, month, year, hour, minute, etc. using the `dt` attribute. For example, let's create a new column containing the information about the day of our `end_date` column.
# Create new column with day of end_date column
df['Day_of_end_date'] = df['End_date'].dt.day
df.head(10)
# Create new column for year of start_date column
df['Year_of_start_date'] = df['Start_date'].dt.year
df.head()
# ## Exercise:
#
# Create at least two more features for example `month`, `hour` or `minute` etc. from the start and end date columns.
df['Month_of_start_date'] = df['Start_date'].dt.month
df.head()
df['Hour_of_start_date'] = df['Start_date'].dt.hour
df.head()
df['Minute_of_start_date'] = df['Start_date'].dt.minute
df.head()
# ## Working with dates on a real dataset
#
# At the end of this notebook we will have a quick look at a real dataset with weather information for the city of Seattle. You will see that the first column contains information about the date of the measurements. While importing the dataset this column is loaded as a object/string type column and we want to convert it before we start our analysis.
# Loading the dataset
df = pd.read_csv("data/seattle-weather.csv")
# Check if the data is loaded correctly
df.head()
# Check column types
df.info()
print(df.date[0])
print(type(df.date[0]))
# From the output of `df.info()` and the two print statements above we can see that the date column is loaded as object/string.
# In order to facilitate working with the date column we can change the type using `.to_datetime()`:
# Replace original column with converted one
df.date = pd.to_datetime(df.date, format='%Y/%m/%d')
df.head()
df.info()
print(df.date[0])
print(type(df.date[0]))
# And now the date column is correctly defined as datetime.
#
# ## Summary
#
# In addition to strings, floats and integers, there is another data type specific to dates and time (datetime).
# If you are dealing with time-dependent data, it is worth to convert them into the correct datetime format.
# This ensures that
# - it's easy to choose the desired format of the date (e.g. YY-MM-DD)
# - durations are calculated correctly
# - durations can be displayed in different time units easily (e.g. in hours or days)
# - additional features can be easily created (e.g. what day of the week it is)
# - visualization tools can recognize variables correctly and display them well (axis labeling).
#
#
| 11_datetime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [HR-Employee-Attrition](https://www.kaggle.com/pavansubhasht/ibm-hr-analytics-attrition-dataset)
# <a href="https://www.indusnet.co.in/what-are-predictive-metrics-and-how-can-it-help-you-predict-employee-turnover/"><img src="../images/hr_attrition.jpg"></a>
# Uncover the factors that lead to employee attrition and explore important questions such as ‘show me a breakdown of distance from home by job role and attrition’ or ‘compare average monthly income by education and attrition’. This is a fictional data set created by IBM data scientists.
# - `Education`: 1 'Below College' 2 'College' 3 'Bachelor' 4 'Master' 5 'Doctor'
# - `EnvironmentSatisfaction`: 1 'Low' 2 'Medium' 3 'High' 4 'Very High'
# - `JobInvolvement`: 1 'Low' 2 'Medium' 3 'High' 4 'Very High'
# - `JobSatisfaction`: 1 'Low' 2 'Medium' 3 'High' 4 'Very High'
# - `PerformanceRating`: 1 'Low' 2 'Good' 3 'Excellent' 4 'Outstanding'
# - `RelationshipSatisfaction`: 1 'Low' 2 'Medium' 3 'High' 4 'Very High'
# - `WorkLifeBalance`: 1 'Bad' 2 'Good' 3 'Better' 4 'Best'
# # Exploración de los datos
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
pd.options.display.max_columns = None
# Load data from a csv file
df = pd.read_csv('../datasets/WA_Fn-UseC_-HR-Employee-Attrition.csv')
df.head()
# - Aquí podemos ver que los campos `EmployeeCount`, `Over18`, y `StandardHours` tienen los mismos valores para todas las filas, así que podemos eliminarlos.
# - Además, el campo `EmployeeNumber` es un identificador y también podemos eliminarlo
df.drop(['EmployeeCount', 'Over18', 'StandardHours', 'EmployeeNumber'], 1, inplace=True)
df.head()
df.info()
# ### Desbalanceo de las clases
sns.countplot(x='Attrition', data=df)
# ### Visualización de los datos
for column in df.select_dtypes(include='int64'):
fare = sns.FacetGrid(df, hue="Attrition",aspect=2)
fare.map(sns.kdeplot, column, shade= True)
fare.set(xlim=(0, df[column].max()))
fare.add_legend()
plt.show()
# ## Preprocesamiento
# ### Valores nulos
total = df.isnull().sum().sort_values(ascending=False)
percent_1 = df.isnull().sum()/df.isnull().count()*100
percent_2 = (round(percent_1, 1)).sort_values(ascending=False)
missing_data = pd.concat([total, percent_2], axis=1, keys=['Total', '%'])
missing_data.head(5)
# ### Detección de outliers
for i, column in enumerate(df.columns):
if type(df.iat[0, i]) == np.int64:
sns.boxplot(x=column, data=df)
plt.show()
# ### Datos categóricos
df.head()
df.info()
# +
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
df['Attrition'] = le.fit_transform(df['Attrition'])
df.head()
# -
str_columns = df.select_dtypes(include='object').columns
str_columns
dummy_busnstrvl = pd.get_dummies(df['BusinessTravel'], prefix='busns_trvl')
dummy_dept = pd.get_dummies(df['Department'], prefix='dept')
dummy_edufield = pd.get_dummies(df['EducationField'], prefix='edufield')
dummy_gender = pd.get_dummies(df['Gender'], prefix='gend')
dummy_jobrole = pd.get_dummies(df['JobRole'], prefix='jobrole')
dummy_maritstat = pd.get_dummies(df['MaritalStatus'], prefix='maritalstat')
dummy_overtime = pd.get_dummies(df['OverTime'], prefix='overtime')
continuous_columns = df.select_dtypes(include='int64').columns
hrattr_continuous = df[continuous_columns]
continuous_columns
df = pd.concat(
[dummy_busnstrvl, dummy_dept, dummy_edufield, dummy_gender, dummy_jobrole, dummy_maritstat,
dummy_overtime, hrattr_continuous], axis=1)
df.head()
df.info()
# ## Selección de variables
df.head()
# +
# sns.heatmap(df.corr(), annot=True, cbar=True)
# -
# # Entrenando los modelos
# +
from sklearn.model_selection import train_test_split
X = df.drop("Attrition", axis=1)
y = df["Attrition"]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# -
y.shape
# ## Logistic Regression
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, confusion_matrix
logreg = LogisticRegression(solver='liblinear')
logreg.fit(X_train, y_train)
y_pred = logreg.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
logreg = LogisticRegression(solver='liblinear')
scores = cross_val_score(logreg, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
# -
# ## Naïve Bayes
# +
from sklearn.naive_bayes import GaussianNB
from sklearn.metrics import accuracy_score, confusion_matrix
gaussian = GaussianNB()
gaussian.fit(X_train, y_train)
y_pred = gaussian.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
gaussian = GaussianNB()
scores = cross_val_score(gaussian, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
# -
# ## Decision tree
# +
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
decision_tree = DecisionTreeClassifier()
decision_tree.fit(X_train, y_train)
y_pred = decision_tree.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
decision_tree = DecisionTreeClassifier()
scores = cross_val_score(decision_tree, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
# -
# ## Random forest
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
random_forest = RandomForestClassifier(n_estimators=100)
random_forest.fit(X_train, y_train)
y_pred = random_forest.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
rf = RandomForestClassifier(n_estimators=100)
scores = cross_val_score(rf, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
# -
importances = pd.DataFrame({'feature':X_train.columns,'importance':np.round(random_forest.feature_importances_,3)})
importances = importances.sort_values('importance',ascending=False).set_index('feature')
importances.head(15)
importances[:20].plot.bar()
# ## Support Vector Machine
# +
from sklearn.svm import SVC, LinearSVC
from sklearn.metrics import accuracy_score, confusion_matrix
linear_svc = LinearSVC(max_iter=10000)
linear_svc.fit(X_train, y_train)
y_pred = linear_svc.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
linear_svc = LinearSVC()
scores = cross_val_score(linear_svc, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
# -
# ### K Nearest Neighbors
# +
from sklearn.neighbors import KNeighborsClassifier
# experimenting with different n values
k_range = list(range(1,26))
scores = []
for k in k_range:
knn = KNeighborsClassifier(n_neighbors=k)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
scores.append(accuracy_score(y_test, y_pred))
plt.plot(k_range, scores)
plt.xlabel('Value of k for KNN')
plt.ylabel('Accuracy Score')
plt.title('Accuracy Scores for Values of k of k-Nearest-Neighbors')
plt.show()
# +
from sklearn.metrics import accuracy_score, confusion_matrix
knn = KNeighborsClassifier(n_neighbors =22)
knn.fit(X_train, y_train)
y_pred = knn.predict(X_test)
print(accuracy_score(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
# +
from sklearn.model_selection import cross_val_score
knn = KNeighborsClassifier(n_neighbors =22)
scores = cross_val_score(knn, X_train, y_train, cv=10, scoring = "accuracy")
print("Scores:", scores)
print("Mean:", scores.mean())
print("Standard Deviation:", scores.std())
| soluciones/ht_employee_attrition-solucion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Has Seattle seen a reduction in collisions from reduced speed limits?
# This notebook examines the impact of reduced speed limits in central Seattle on the number of collisions and collision related injuries.
#
# This notebook will leverage a Python package called CausalImpact, which uses a bayesian structural time series model, to investigate whether there was statistically significant drop in collisions or reduction in injuries after speed limits in Central Seattle were reduced in October of 2016.
#
# https://google.github.io/CausalImpact/CausalImpact.html
#
# **Please note that we've identified a bug in this python implementation of the CausalImpact package where an Key Error shows on some machines when the .run() command is executed. We are unable to track down the root cause to specific depency but found an issue was already opened in the repo for a similar error. This is a lesson learned for future technology reviews.
# #### Step 0: Import packages
#First we'll add another directory to paths list so we can import our module render_stats
import sys
import os
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
import pandas as pd
from causalimpact import CausalImpact
#Render_stats will be used to perform most of the data cleaning in this project.
import wa_collisions.render_stats as render_stats
# #### Step 1: Import data
# First, let's read in the Seattle collisions data and map the data to neighbhorhoods.
#
# Make sure the file_path variable is set the location of the Collisions data from you are using. The default below is the location the data will write to if you follow our instructions in Example - Prepare Data.
file_path = "../wa_collisions/data/Collisions.csv"
data = render_stats.read_collision_with_neighborhoods(file_path, contains_neighborhood=False, geo_path_root='../')
# #### Step 2: Data clean-up and visualization
# First, we'll identify neighborhoods where speed limit changed and those where it didn't. Based on looking at the boundaries of these neighborhoods we see that the boundaries are similar those shown in this report: http://www.seattle.gov/Documents/Departments/beSuperSafe/VZ_2017_Progress_Report.pdf
# We will assume all neighborhoods outside of this core group did not have their speed limits changed in October 2016.
#
# After determining this list, we will call a function which pivots the data by the number of collisions per day in neighbhorhoods where speed limits changed and those where they didn't. These can be thought of as "control" and "treatment" groups.
speed_limit_nbrhood = ['Atlantic','Pike-Market', 'Belltown', 'International District',
'Central Business District', 'First Hill', 'Yesler Terrace',
'Pioneer Square', 'Interbay','Mann','Minor']
daily_collision_count = render_stats.pivot_by_treatment(input_frame=data, treatment_list=speed_limit_nbrhood, control_list=None
, neighborhood_path='../wa_collisions/data/Neighborhoods/Neighborhoods.json'
, agg_by=None, resample_by='D')
daily_collision_count.plot()
daily_collision_count.corr()
# Plotting this data, we see that it is quite noisy and events are not highly correlated. This violates our assumption that control and treatment are similar except for the change in speed limits.
#
# What happens if we aggregate the data by month?
monthly_collision_count = render_stats.pivot_by_treatment(input_frame=data, treatment_list=speed_limit_nbrhood, control_list=None
, neighborhood_path='../wa_collisions/data/Neighborhoods/Neighborhoods.json'
, agg_by=None, resample_by='M')
monthly_collision_count.plot()
monthly_collision_count.corr()
# That's better!
#
# Looking at the plot, we see that there appears to be a slight drop in the number of collisions in 2017, but it's pretty hard to tell. It also looks like there is a similar drop in neigborhoods where speed limits are not changed. Let's run the data through Causal Impact for a closer look.
# #### Step 3: Analysis of collision counts
# First, we need to find the date range before and after the speed limit change.
date_ranges = render_stats.find_period_ranges(monthly_collision_count, transition_date="2016-10-01")
print('pre:', date_ranges[0])
print('post:', date_ranges[1])
# Now, let's perform the analysis
impact_collision_count = CausalImpact(monthly_collision_count, date_ranges[0], date_ranges[1])
impact_collision_count.run()
impact_collision_count.summary()
# Interesting. The results indicate there were 5 fewer collisions than would have been predicted based on a counterfactual. This is a -0.1% drop.
#
# This result is definitely not statistically significant.
#
# Let's plot the data for a closer look.
impact_collision_count.plot()
# #### Step 4: Analysis of only speeding related collisions
# The total count of collision doesn't appear to have been impacted by a change in speed limits in central Seattle, but what about only speeding related collisions? Surely those were affected.
#
# With lower limits, more drivers might be speeding in general. From this, we could hypothesize that more collisions could be linked to speeding.
# First we'll re-pull the filtered data and group by month. We'll plot the data and check the correlation between control and treatment again.
monthly_speeding_count = render_stats.pivot_by_treatment(input_frame=data[data['ind_speeding']==True], treatment_list=speed_limit_nbrhood, control_list=None
, neighborhood_path='../wa_collisions/data/Neighborhoods/Neighborhoods.json'
, agg_by=None, resample_by='M')
monthly_speeding_count.plot()
monthly_speeding_count.corr()
monthly_speeding_count.ix['2011-10-01':'2016-10-01'].corr()
# Interesting. We are not seeing a strong correlation here, and it is basically the same whether we look at all months or just the 5 years before the speed limit change.
#
# What does Causal Impact show?
# First, let's make sure our date range are still valid.
date_ranges = render_stats.find_period_ranges(monthly_speeding_count, transition_date="2016-10-01")
print('pre:', date_ranges[0])
print('post:', date_ranges[1])
impact_speeding_count = CausalImpact(monthly_speeding_count, date_ranges[0], date_ranges[1])
impact_speeding_count.run()
impact_speeding_count.summary()
# We are seeing a drop in the number of speeding collisions predicted, contrary to my hypothesis. Interesting.
#
# Unfortunately, due to the high variability in month collisions in general, we are not able to detect statistical significance.
#
# Plotting the data below, there is an apparent downward trend in cumulative impact though.
impact_speeding_count.plot()
# #### Step 5: Analysis of injuries
# So far, we see almost no notable impact of the speed limit changes on collision counts in Seattle, but what about injuries? Perhaps there are just as many fender benders as before but because people are driving slower, fewer people were injured.
# First, let's repull the data, plot it and check control/treatment correlation like we did above.
monthly_injuries = render_stats.pivot_by_treatment(input_frame=data, treatment_list=speed_limit_nbrhood, control_list=None
, neighborhood_path='../wa_collisions/data/Neighborhoods/Neighborhoods.json'
, agg_by='injuries', resample_by='M')
monthly_injuries.plot()
monthly_injuries.corr()
# As before, we are seeing a reasonably high correlation between treatment and control, however the plot does not suggest the number of injuries has fallen.
#
# Let's run it through Causal Impact.
date_ranges = render_stats.find_period_ranges(monthly_injuries, transition_date="2016-10-01")
print('pre:', date_ranges[0])
print('post:', date_ranges[1])
impact_injuries = CausalImpact(monthly_injuries, date_ranges[0], date_ranges[1])
impact_injuries.run()
impact_injuries.summary()
# We are seeing fewer total injuries than might have been predicted, but it's not statistically significant. Let's see how appararent this is in the plot.
impact_injuries.plot()
# #### Summary
# In this notebook, we have used Causal Impact to analyze collisions in Seattle before and after speed limits were dropped in October of 2016.
#
# While the speed limit change does not appear to have affected the total number of collisions, it's possible that it had a small impact on reducing speeding related collisions and the number of injury related collisions.
#
# Unfortunately, due to small sizes and an inherently noisy dataset, we are unable to come to any firm conclusions. However, we do see some positive signals.
| examples/Example - CausalImpact SpeedLimits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AliAqdas-repo/FallDetection/blob/main/FD_DataOrganization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="sLaem-TFSwJ4"
# #Data Organization and Structuring
# The goal of this notebook is to download the dataset and organize necessary data so that it could be used to train a Deep Learning Model for Fall Detection
# + [markdown] id="sg68eTBDS7ln"
# ##Importing Dependencies
# + id="FJ1UpxzlS4ft"
import os
import numpy as np
import matplotlib.pyplot as plt
import re
import csv
# + [markdown] id="HRoa0nBYTMZZ"
# ##Downloading Dataset from Drive
# The dataset downloaded in the first block is copied directly to Google Drive to avoid downloading again and again and can be just downloaded over Google Servers from **Drive**
# + id="peQRR0SoTKHm"
#Downloading Dataset from Google Drive
# !gdown --id 1-J5TBYvi---DW68UvVrUMhyF2u-QZLdj
# + id="ZUWZI2K3Tyzw"
#Unzipping Dataset
if not os.path.exists('/content/Dataset/'):
os.mkdir('/content/Dataset/')
# !unzip /content/dataset.zip -d /content/Dataset/
# + [markdown] id="ps622PFUUNno"
# ## Loading the Dataset in Colab
# + id="OIVASTS4T2B7"
#Function that Enlists Full Path of Data Files
def list_full_path(dir):
return [os.path.join(dir,os.path.splitext(fi)[0]) for fi in os.listdir(dir)]
# + id="F9ifSLVTUD7q"
data_files=list_full_path('/content/Dataset/FallAllD')
# + id="DaxuoQpHUQxk"
data_files[0]
# + id="td7u7gJKX004"
#Length of Data
len(data_files)
# + id="8ZKnVKXVX59R"
#Extracts the Activity Code
int(data_files[0][34:37])
# + [markdown] id="5Uaxqr1NYg6G"
# ##Splitting Data
# Splitting entire dataset into Fall and Activities of Daily Life(ADLs)
# + id="zkhdkoNAYAl4"
fall_files=[]
not_fall_files=[]
for datfile in data_files:
if not (datfile[-1]=='B' or datfile[-1]=='M'): #Rejecting Barometer and Magnetometer Data
#Splitting into Fall and Not Fall based on ActivtyID2Str.m file
if 100<int(datfile[34:37])<136:
fall_files.append(str(datfile))
else:
not_fall_files.append(str(datfile))
# + id="mUQSfCUPYd5b"
#Length of All Files
len(not_fall_files)
#Number of Sets of Sensor Measurements. We divide by 4 because we have data from 4 sensors per subject/measurement.
len(not_fall_files)/4
# + [markdown] id="VDC5pCjqZa1Z"
# ## Fusing Sensor Data
# We are fusing sensor data from different sensors with different sampling rates to create a single file that can be used as input to a Neural Network
# + id="LIrhjIUXZEFK"
#Sorting the Data
fall_files=sorted(fall_files)
not_fall_files=sorted(not_fall_files)
# + id="r4QhV9KHZNRh"
def read_into_array(filedir):
#reads data into array. If file extension isn't displayed in filedir then its default to .dat
ext=os.path.splitext(filedir)[1]
if ext=='':
ext='.dat'
with open(os.path.splitext(filedir)[0]+ext,'r') as datfile :
file_data=csv.reader(datfile,delimiter=',')
data=list()
for row in file_data:
data.append([float(val) for val in row])
return data
# + id="Q9W_fbifaA9O"
#Order of Files: Acc Bar Gyro Magn
import cv2
def save_combined_data(input_files,output_dir=None,AG_ONLY=True):
############### INPUT PARAMETERS ##################
#--------------------------------------------------
#input_files = Input File Paths
#output_dir = Directory to Output Combined Data
#AG_ONLY = Accelration and Gyro Data Only(True)
#--------------------------------------------------
OUTPUT_SAMPLES=952
TMP_MAG_SAMPLES=4800
TMP_BAR_SAMPLES=1000
if AG_ONLY:
GAP=2
TOTAL_CHANNELS=6;
#Channel Assignment Order
#-------------------------------------
# Channel 1-3 - Accelerometer
# Channel 4-6 - Gyroscope
#-------------------------------------
else:
GAP=4
TOTAL_CHANNELS=10;
dataset=np.empty((len(input_files)//GAP,OUTPUT_SAMPLES,TOTAL_CHANNELS))
#Channel Assignment Order
#-------------------------------------
# Channel 1-3 - Accelerometer
# Channel 4 - Barometer
# Channel 5-7 - Gyroscope
# Channel 8-10 - Magnetometer
#-------------------------------------
for k in range(0,len(input_files),GAP):
data=np.empty((OUTPUT_SAMPLES,TOTAL_CHANNELS))
co_channel=0 #Current Output Channel, Used to Iterate Data in Loop
for ip_file in input_files[k+0:k+GAP]:
datpts=read_into_array(ip_file)
ip_datpts=[[None]*3]*OUTPUT_SAMPLES
if len(datpts)==200:
CHANNELS=1
tmp_bar_pts=np.asarray([[0.0]*CHANNELS]*TMP_BAR_SAMPLES)
#Upsample by a factor of 5 and dropping last 48 samples
for i in range(0,5):
tmp_bar_pts[i::5,0]=np.asarray(datpts)[:,0]
ip_datpts=tmp_bar_pts[0:OUTPUT_SAMPLES]
elif len(datpts)==1600:
CHANNELS=3
tmp_mag_pts=[[None]*CHANNELS]*TMP_MAG_SAMPLES
#Zero Order Hold to Upsample Data by a Factor of 3
tmp_mag_pts[0::3]=datpts
tmp_mag_pts[1::3]=datpts
tmp_mag_pts[2::3]=datpts
#Dropping last 40 Samples and Downsampling data by factor of 5
ip_datpts=tmp_mag_pts[0:4760:5]
elif len(datpts)==4760:
CHANNELS=3
ip_datpts=datpts[::5]
else:
print('Error')
break
print(ip_file)
data[:,co_channel:co_channel+CHANNELS]=np.asarray(ip_datpts) #assigning channels to data from files
co_channel=co_channel+CHANNELS
if not output_dir==None:
print('Saving Data')
if not output_dir[-1]=='/':
if not os.path.exists(output_dir):
os.makedirs(output_dir)
np.savetxt(f'{output_dir}/{ip_file[26:-2]}.csv',data,delimiter=",")
else:
if not os.path.exists(output_dir):
os.makedirs(output_dir[:-1])
np.savetxt(f'{output_dir}/{ip_file[26:-2]}.csv',data,delimiter=",")
else:
dataset[k//GAP]=data
if output_dir==None:
return dataset
# + id="LPH9gWTXZ-sL"
save_combined_data(fall_files,'/content/data_proc_AG/fall_files/',True)
save_combined_data(not_fall_files,'/content/data_proc_AG/not_fall_files/',True)
# + [markdown] id="axuXEPR1aUtH"
# The data can now be zipped and uploaded to Google Drive to save the effort for the next time you work on this project.
# + id="E6VeHyE4akd_"
# !zip -r /content/data_proc_AG.zip /content/data_proc_AG/
# !cp /content/data_proc_AG.zip /content/drive/MyDrive/Datasets/FallDetect/data_proc_AG.zip
| FD_DataOrganization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.insert(0, "../src/")
sys.path.insert(0, "../../../src/")
sys.path.insert(0, "../../../projects/pyskip_bench/src/")
sys.path.insert(0, "../../../projects/pyskip_blox/")
# +
import random
import numpy as np
import pyskip
import pyskip_3d
from pyskip_blox import minecraft, colors
from pyskip import functional as F
import ipywebrtc
SHOW_RUNS_MODE = False
# -
def show_runs(t, runs_value=2):
q = t.reshape(len(t))
ret = pyskip.Tensor(len(t), val=q.to(bool).to(int)._tensor.array())
run_points = runs_value * (
(
(q[:(len(q) - 1)] == q[1:])
)
& (q[1:] != 0)
).to(int)
ret[1:] = ret[1:].max(run_points)
ret[0:(len(ret) - 1)] = ret[0:(len(ret) - 1)].max(runs_value * (ret[1:] == 2).to(int))
return ret.reshape(t.shape)
# %%time
# level = minecraft.PySkipMinecraftLevel.load("../data/sample_minecraft_world.pickle.gz")
level = minecraft.PySkipMinecraftLevel.load("../data/Hogwarts1.0.pickle.gz")
# %%time
t = level.megatensor()
# t = level.chunk_list[0].tensor
if SHOW_RUNS_MODE:
t = show_runs(t)
for i in range(20):
print(f"{i}=>{F.sum((t == i).to(int))}")
config = pyskip_3d.VoxelConfig()
config[0] = pyskip_3d.EmptyVoxel()
if SHOW_RUNS_MODE:
config[1] = pyskip_3d.ColorVoxel(255, 255, 255)
config[2] = pyskip_3d.ColorVoxel(255, 105, 180)
else:
for i in range(1, 256):
r, g, b = colors.color_for_id(i)
config[i] = pyskip_3d.ColorVoxel(r, g, b)
# %%time
mesh = pyskip_3d.generate_mesh(config, t._tensor)
(len(mesh.triangles) * 4 + len(mesh.positions) * 4) / (level.num_runs() * 2 * 4)
from pythreejs import *
import numpy as np
from IPython.display import display
# +
# %%time
geometry = BufferGeometry(
attributes={
"position": BufferAttribute(
np.array(mesh.positions, dtype='float32').reshape(-1, 3),
normalized=False,
),
"normal": BufferAttribute(
np.array(mesh.normals, dtype='float32').reshape(-1, 3),
normalized=False,
),
"color": BufferAttribute(
np.array(mesh.colors, dtype='uint8').reshape(-1, 4)
),
"index": BufferAttribute(
np.array(mesh.triangles, dtype='uint32'),
normalized=False,
),
},
)
# -
js_mesh = Mesh(
geometry=geometry,
material=MeshLambertMaterial(vertexColors="VertexColors"),
position=[0, 0, 0],
)
"""
camera_pos = level.xyz_to_pyskip_col(level.dense_dimensions())
camera = PerspectiveCamera(position=camera_pos, lookAt=(0,0,0), fov=20)
up = [0, 0, 0]
up[level.column_order[1]] = 1
camera.up = tuple(up)
"""
# +
light_position = level.xyz_to_pyskip_col((0, 256, 0))
point_light = PointLight(color='#ffffff', position=light_position)
global_light = AmbientLight(color='#333333')
#normals_helper = VertexNormalsHelper(js_mesh, size=0.5, color="#ffcc00")
scene = Scene(children=[js_mesh, camera, point_light, global_light], background="black")
# -
# +
renderer = Renderer(
camera=camera,
background="#b0b0b0",
background_opacity=1,
scene=scene,
# controls=[OrbitControls(controlling=camera)],
width=600,
height=500,
)
display(renderer)
# -
stream = ipywebrtc.WidgetStream(widget=renderer, max_fps=30)
recorder = ipywebrtc.ImageRecorder(filename='snapshot', format='png', stream=stream)
recorder
| projects/pyskip_3d/notebooks/render_minecraft.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/04_Python_Functions/tree/main/002_Python_Functions_Built_in)**
# </i></small></small>
# # Python `abs()`
#
# The **`abs()`** method returns the absolute value of the given number. If the number is a complex number, **`abs()`** returns its magnitude.
#
# **Syntax**:
#
# ```python
# abs(num)
# ```
# ## `abs()` Parameters
#
# **`abs()`** method takes a single argument:
#
# * **num** - a number whose absolute value is to be returned. The number can be:
# * integer
# * floating number
# * complex number
# ## Return value from `abs()`
#
# **`abs()`** method returns the absolute value of the given number.
#
# * For integers - integer absolute value is returned
# * For floating numbers - floating absolute value is returned
# * For complex numbers - magnitude of the number is returned
#
# +
# Example 1: Get absolute value of a number
# random integer
integer = -20
print('Absolute value of -20 is:', abs(integer))
#random floating number
floating = -30.33
print('Absolute value of -30.33 is:', abs(floating))
# +
# Example 2: Get magnitude of a complex number
# random complex number
complex = (3 - 4j)
print('Magnitude of 3 - 4j is:', abs(complex))
# -
| 002_Python_Functions_Built_in/001_Python_abs().ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from scipy import spatial
import matplotlib.pyplot as plt
import logging
LOGGER = logging.getLogger(__name__)
from pynhhd import create_logger
create_logger(logging.INFO)
from utils import drawing, fields
d = 1
n = 101
# -----------------------------------------------
# create points
points = d*np.indices((n,n))
points = [points[d] for d in range(2)]
points = np.moveaxis(points, 0, -1)
points = points.reshape(-1, points.shape[-1])
npoints = points.shape[0]
# -----------------------------------------------
# create a Delaunay triangulation
simplices = spatial.Delaunay(points).simplices
nfaces = simplices.shape[0]
simplices2 = simplices[np.random.permutation(nfaces)]
simplices = simplices2
# -----------------------------------------------
# create PC vector field
centroids = np.zeros((nfaces, 2))
for i in range(simplices.shape[0]):
centroids[i] = 1.0/3.0 * (points[simplices[i][0]] + points[simplices[i][1]] + points[simplices[i][2]])
c = np.array([(n-1)/2,(n-1)/2])
vf = fields.create_criticalPoint2D(centroids, np.array([(n-1)/2,(n-1)/2]), 1, 1, 0,1)
mvf = np.linalg.norm(vf, axis=1)
LOGGER.info('vf = {}, {}, {}'.format(mvf.shape, mvf.min(), mvf.max()))
# -----------------------------------------------
plt.figure()
#plt.scatter(points[:,0],points[:,1], c=p, s=60, cmap=plt.cm.jet) #discretize_colormap(plt.cm.jet,3))
plt.triplot(points[:,0],points[:,1], simplices, '-', alpha=0.2)
plt.gca().set_aspect('equal', 'box')
vrng = (0, 196.1)
k = 50
plt.figure()
drawing.draw_quivers(centroids, vf, vrng, k)
plt.gca().set_aspect('equal', 'box')
#plt.xlim([-0.1,n-1+0.1])
#plt.ylim([-0.1,n-1+0.1])
plt.show()
# +
import sys
from pynhhd import nHHD
hhd = nHHD(points = points, simplices = simplices)
hhd.decompose(vf)
# +
plt.figure()
plt.imshow(hhd.div.reshape(n,n),origin='lower',cmap=plt.cm.PiYG) #,vmax=numpy.abs(n).max(), vmin=-numpy.abs(n).max())
plt.xlim([-0.5,n-1+0.5])
plt.ylim([-0.5,n-1+0.5])
plt.colorbar()
plt.figure()
plt.imshow(hhd.curlw.reshape(n,n),origin='lower',cmap=plt.cm.PiYG) #,vmax=numpy.abs(n).max(), vmin=-numpy.abs(n).max())
plt.xlim([-0.5,n-1+0.5])
plt.ylim([-0.5,n-1+0.5])
plt.colorbar()
plt.show()
# +
plt.figure()
plt.imshow(hhd.nD.reshape(n,n),origin='lower',cmap=plt.cm.PiYG) #,vmax=numpy.abs(n).max(), vmin=-numpy.abs(n).max())
plt.xlim([-0.5,n-1+0.5])
plt.ylim([-0.5,n-1+0.5])
plt.colorbar()
plt.figure()
plt.imshow(hhd.nRu.reshape(n,n),origin='lower',cmap=plt.cm.PiYG) #,vmax=numpy.abs(n).max(), vmin=-numpy.abs(n).max())
plt.xlim([-0.5,n-1+0.5])
plt.ylim([-0.5,n-1+0.5])
plt.colorbar()
plt.show()
# +
# ----------------------------------------------------------------
mvf = np.linalg.norm(vf, axis=1)
LOGGER.info('vf = {}, {}, {}'.format(mvf.shape, mvf.min(), mvf.max()))
mr = np.linalg.norm(hhd.r, axis=1)
md = np.linalg.norm(hhd.d, axis=1)
mh = np.linalg.norm(hhd.h, axis=1)
LOGGER.info('d = {}, {}'.format(md.min(), md.max())) #, numpy.linalg.norm(md)
LOGGER.info('r = {}, {}'.format(mr.min(), mr.max())) #, numpy.linalg.norm(mr)
LOGGER.info('h = {}, {}'.format(mh.min(), mh.max())) #, numpy.linalg.norm(mh)
#vrng = (0, 196.1)
vrng = (0, 196.1)
k = 50
# ----------------------------------------------------------------
plt.figure()
#plt.scatter(points[:,0],points[:,1], c=p, s=60, cmap=plt.cm.jet) #discretize_colormap(plt.cm.jet,3))
#plt.triplot(points[:,0],points[:,1], simplices, '-', alpha=0.2)
drawing.draw_quivers(centroids, vf, vrng, k)
plt.xlim([-0.1,n-1+0.1])
plt.ylim([-0.1,n-1+0.1])
#plt.savefig('fig1.png')
plt.gca().set_aspect('equal', 'box')
# ----------------------------------------------------------------
plt.figure()
drawing.draw_quivers(centroids, hhd.r, vrng, k)
#plt.imshow(hhd.nD.reshape(n,n),origin='lower',cmap=plt.cm.PiYG) #,vmax=numpy.abs(n).max(), vmin=-numpy.abs(n).max())
plt.xlim([-0.1,n-1+0.1])
plt.ylim([-0.1,n-1+0.1])
plt.gca().set_aspect('equal', 'box')
#plt.savefig('fig2.png')
plt.figure()
drawing.draw_quivers(centroids, hhd.h, vrng, k)
plt.xlim([-0.1,n-1+0.1])
plt.ylim([-0.1,n-1+0.1])
plt.gca().set_aspect('equal', 'box')
#plt.savefig('fig3.png')
plt.show()
# -
| pynhhd-v1.1/examples/ex-nhhd2-unstructgrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/datarobot-community/mlops-examples/blob/master/MLOps_Agent/Main_Script.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# ## MLOps Agent - Python End to End
# **Author**: <NAME>
#
# #### Scope
# The scope of this Notebook is to provide instructions on how to use DataRobot's MLOps Agents.
#
# #### Requirements
#
# - Python 3.7.0
# - MLOps Agent 6.3.3
#
# Your version might be different but the below procedure should remain the same.
#Clone the repository
# !git clone https://github.com/datarobot-community/mlops-examples
#Install needed packages
# !pip install -r /content/mlops-examples/MLOps_Agent/requirements.txt
# ### Configuring the Agent
#
# To configure the agent, we just need to define the DataRobot MLOps location and our API token. By default, the agent expects the data to be spooled on the local file system. Make sure that default location (/tmp/ta) exists.
#
# The `token` needs to be your personal token found under Developer Tools in your DataRobot instance. The endpoint specified below is the DataRobot trial endpoint but you should change it if needed.
#
import datarobot as dr
import os
# +
token = "YOUR_API_TOKEN"
endpoint = "https://app2.datarobot.com"
## connect to DataRobot platform with python client.
client = dr.Client(token, "{}/api/v2".format(endpoint))
mlops_agents_tb = client.get("mlopsInstaller")
with open("/content/mlops-examples/MLOps_Agent/mlops-agent.tar.gz", "wb") as f:
f.write(mlops_agents_tb.content)
# -
# #### Once it is downloaded....and saved to your local filesystem, open/uncompress the file
# +
# !tar -xf /content/mlops-examples/MLOps_Agent/mlops-agent.tar.gz
#Save the folder where the whl file is saved
with os.popen("ls /content") as pipe:
for line in pipe:
if line.startswith('datarobot_mlops_package'):
mlops_package = line.strip()
version = line.strip()[-5:]
print(mlops_package)
print(version)
# -
#Execute command and install mlops-agent
os.system('pip install /content/{}/lib/datarobot_mlops-{}-py2.py3-none-any.whl'.format(mlops_package, version))
# ### Open Quick Start
# As noted in comment code from the Deployment Integrations tab above, open to get started with the agent software configuration steps:
#
# .../{agent install dir}/docs/html/quickstart.html
#
# Edit .../{agent install dir}/conf/mlops.agent.conf.yaml to have this (everything else can stay as default if you want)
# This file is contains the properties used by the MLOps service. Namely, the DataRobpt host url, your authentication token, the spool to use queue data to send to MLOps.
"""
# Set your DR host:
mlopsURL: "https://app2.datarobot.com"
# Set your API token
apiToken: "<KEY>"
# Create the spool directory on your system that you want MLOps to use, eg /tmp/ta
channelConfigs:
- type: "FS_SPOOL"
details: {name: "bench", spoolDirectoryPath: "/tmp/ta"}
"""
# !mkdir /tmp/taqw
# #### Commands to get you started
#
# This will allow you to start, get status, and stop the MLOps agent service. You will only need to run start for now. Run status if you want to check on the service.
# !bash /content/datarobot_mlops_package-6.3.3/bin/start-agent.sh #Change version based on the downloaded file
# +
# Shutdown - DON'T RUN THIS CELL, IT'S JUST SHOWING YOU HOW TO SHUTDOWN
# #!bash datarobot_mlops_package-6.3.3/bin/stop-agent.sh
# -
# ## Create an MLOps Model Package for a model and deploy it
# #### Train a simple RandomForestClassifier model to use for this example
# +
import pandas as pd
import numpy as np
import time
import csv
import pytz
import json
import yaml
import datetime
from sklearn.ensemble import RandomForestClassifier
TRAINING_DATA = '/content/{}/examples/data/surgical-dataset.csv'.format(mlops_package)
df = pd.read_csv(TRAINING_DATA)
columns = list(df.columns)
arr = df.to_numpy()
np.random.shuffle(arr)
split_ratio = 0.8
prediction_threshold = 0.5
train_data_len = int(arr.shape[0] * split_ratio)
train_data = arr[:train_data_len, :-1]
label = arr[:train_data_len, -1]
test_data = arr[train_data_len:, :-1]
test_df = df[train_data_len:]
# train the model
clf = RandomForestClassifier(n_estimators=10, max_depth=2, random_state=0)
clf.fit(train_data, label)
# -
# ### Create empty deployment in DataRobot MLOps
#
# Using the MLOps client, create a new model package to represent the random forest model we just created. This includes uploading the traning data and enabling data drift.
# +
from datarobot.mlops.mlops import MLOps
from datarobot.mlops.common.enums import OutputType
from datarobot.mlops.connected.client import MLOpsClient
from datarobot.mlops.common.exception import DRConnectedException
from datarobot.mlops.constants import Constants
# Read the model configuration info from the example. This is used to create the model package.
with open('/content/{}/examples/model_config/surgical_binary_classification.json'.format(mlops_package), "r") as f:
model_info = json.loads(f.read())
model_info
# Read the mlops connection info from the provided example
with open('/content/{}/conf/mlops.agent.conf.yaml'.format(mlops_package)) as file:
# The FullLoader parameter handles the conversion from YAML
# scalar values to Python the dictionary format
agent_yaml_dict = yaml.load(file, Loader=yaml.FullLoader)
MLOPS_URL = agent_yaml_dict['mlopsUrl']
API_TOKEN = agent_yaml_dict['apiToken']
# Create connected client
mlops_connected_client = MLOpsClient(MLOPS_URL, API_TOKEN)
# Add training_data to model configuration
print("Uploading training data - {}. This may take some time...".format(TRAINING_DATA))
dataset_id = mlops_connected_client.upload_dataset(TRAINING_DATA)
print("Training dataset uploaded. Catalog ID {}.".format(dataset_id))
model_info["datasets"] = {"trainingDataCatalogId": dataset_id}
# Create the model package
print('Create model package')
model_pkg_id = mlops_connected_client.create_model_package(model_info)
model_pkg = mlops_connected_client.get_model_package(model_pkg_id)
model_id = model_pkg["modelId"]
# Deploy the model package
print('Deploy model package')
# Give the deployment a name:
DEPLOYMENT_NAME="Python binary classification remote model " + str(datetime.datetime.now())
deployment_id = mlops_connected_client.deploy_model_package(model_pkg["id"],
DEPLOYMENT_NAME)
# Enable data drift tracking
print('Enable feature drift')
enable_feature_drift = TRAINING_DATA is not None
mlops_connected_client.update_deployment_settings(deployment_id, target_drift=True,
feature_drift=enable_feature_drift)
_ = mlops_connected_client.get_deployment_settings(deployment_id)
print("\nDone.")
print("DEPLOYMENT_ID=%s, MODEL_ID=%s" % (deployment_id, model_id))
DEPLOYMENT_ID = deployment_id
MODEL_ID = model_id
# -
# ## Run Model Predictions
#
# #### Call the external model's predict fuction and send prediction data to MLOps
#
# You can find Deployment and Model ID under `Deployments` --> `Monitoring` Tab. The rest of the code can stay as it is.
# +
import sys
import time
import random
import pandas as pd
from datarobot.mlops.mlops import MLOps
from datarobot.mlops.common.enums import OutputType
DEPLOYMENT_ID = 'YOUR_DEPLOYMENT_ID'
MODEL_ID = 'YOUR_MODEL_ID'
CLASS_NAMES = ["1", "0"]
# Spool directory path must match the Monitoring Agent path configured by admin.
SPOOL_DIR = "/tmp/ta"
"""
This sample code demonstrates usage of the MLOps library.
It does not have real data (or even a real model) and should not be run against a real MLOps
service.
"""
ACTUALS_OUTPUT_FILE = 'actuals.csv'
def main(deployment_id, model_id, spool_dir, class_names):
"""
This is a binary classification algorithm example.
User can call the DataRobot MLOps library functions to report statistics.
"""
# MLOPS: initialize the MLOps instance
mlops = MLOps() \
.set_deployment_id(deployment_id) \
.set_model_id(model_id) \
.set_filesystem_spooler(spool_dir) \
.init()
# Get predictions
start_time = time.time()
predictions = clf.predict_proba(test_data).tolist()
num_predictions = len(predictions)
end_time = time.time()
# Get assocation id's for the predictions so we can track them with the actuals
def _generate_unique_association_ids(num_samples):
ts = time.time()
return ["x_{}_{}".format(ts, i) for i in range(num_samples)]
association_ids = _generate_unique_association_ids(len(test_data))
# MLOPS: report the number of predictions in the request and the execution time.
mlops.report_deployment_stats(num_predictions, end_time - start_time)
# MLOPS: report the predictions data: features, predictions, class_names
mlops.report_predictions_data(features_df=test_df,
predictions=predictions,
class_names=class_names,
association_ids=association_ids)
target_column_name = columns[len(columns) - 1]
target_values = []
orig_labels = test_df[target_column_name].tolist()
print("Wrote actuals file: %s" % ACTUALS_OUTPUT_FILE)
def write_actuals_file(out_filename, test_data_labels, association_ids):
"""
Generate a CSV file with the association ids and labels, this example
uses a dataset that has labels already.
In a real use case actuals (labels) will show after prediction is done.
:param out_filename: name of csv file
:param test_data_labels: actual values (labels)
:param association_ids: association id list used for predictions
"""
with open(out_filename, mode="w") as actuals_csv_file:
writer = csv.writer(actuals_csv_file, delimiter=",")
writer.writerow(
[
Constants.ACTUALS_ASSOCIATION_ID_KEY,
Constants.ACTUALS_VALUE_KEY,
Constants.ACTUALS_TIMESTAMP_KEY
]
)
tz = pytz.timezone("America/Los_Angeles")
for (association_id, label) in zip(association_ids, test_data_labels):
actual_timestamp = datetime.datetime.now().replace(tzinfo=tz).isoformat()
writer.writerow([association_id, "1" if label else "0", actual_timestamp])
# Write csv file with labels and association Id, when output file is provided
write_actuals_file(ACTUALS_OUTPUT_FILE, orig_labels, association_ids)
# MLOPS: release MLOps resources when finished.
mlops.shutdown()
main(DEPLOYMENT_ID, MODEL_ID, SPOOL_DIR, CLASS_NAMES)
# -
# ### Upload actuals back to MLOps
# +
def _get_correct_actual_value(deployment_type, value):
if deployment_type == "Regression":
return float(value)
return str(value)
def _get_correct_flag_value(value_str):
if value_str == "True":
return True
return False
def upload_actuals():
print("Connect MLOps client")
mlops_connected_client = MLOpsClient(MLOPS_URL, API_TOKEN)
deployment_type = mlops_connected_client.get_deployment_type(DEPLOYMENT_ID)
actuals = []
with open(ACTUALS_OUTPUT_FILE, mode="r") as actuals_csv_file:
reader = csv.DictReader(actuals_csv_file)
for row in reader:
actual = {}
for key, value in row.items():
if key == Constants.ACTUALS_WAS_ACTED_ON_KEY:
value = _get_correct_flag_value(value)
if key == Constants.ACTUALS_VALUE_KEY:
value = _get_correct_actual_value(deployment_type, value)
actual[key] = value
actuals.append(actual)
if len(actuals) == 10000:
mlops_connected_client.submit_actuals(DEPLOYMENT_ID, actuals)
actuals = []
# Submit the actuals
print("Submit actuals")
mlops_connected_client.submit_actuals(DEPLOYMENT_ID, actuals)
print("Done.")
upload_actuals()
# -
# ### Stop the mlops service
# !bash /content/datarobot_mlops_package-6.3.3/bin/stop-agent.sh #Change version based on the downloaded file
| MLOps_Agent/Main_Script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.4 ('inthemoment')
# language: python
# name: python3
# ---
# +
# example.ipynb
# Authors: <NAME>
# Example how to use ITM
# -
import sys
import matplotlib.pyplot as plt
import numpy as np
sys.path.append("../")
from inthemoment import ITM, config
# Some settings
config['general']["random state seed"] = 44
itm = ITM()
# Generating the sample
itm.generate()
count, bins, ignored = plt.hist(itm.sample, 30, density=True)
plt.plot(bins, itm._sample_pdf(bins, config["sample"]["pdf"]["mean"], config["sample"]["pdf"]["sd"]),
linewidth=2, color='r')
plt.show()
# The subset
count, bins, ignored = plt.hist(itm.subset, 30, density=True)
plt.plot(bins, itm._sample_pdf(bins, config["sample"]["pdf"]["mean"], config["sample"]["pdf"]["sd"]),
linewidth=2, color='r')
plt.show()
# Fitting
fit_res = itm.fit()
fig, ax = plt.subplots()
count, bins, ignored = plt.hist(itm.subset, 30, density=True)
ax.plot(bins, itm._sample_pdf(bins, config["sample"]["pdf"]["mean"], config["sample"]["pdf"]["sd"]),
linewidth=2, color='r', label='Truth')
ax.plot(bins, itm._pdf(bins, *fit_res[0]), label="Sample",
linewidth=2, color='b')
ax.plot(bins, itm._pdf(bins, *fit_res[2]), label='ITM',
linewidth=2, color='g')
ax.plot(bins, itm._pdf(bins, *fit_res[3]), label='Scipy',
linewidth=2, color='y', ls='--')
ax.legend()
ax.set_ylim(0., 4)
plt.show()
fig.savefig("fit_res.png", facecolor='white', dpi=500)
# Example for the rebinning
count, bins = np.histogram(itm.sample, np.linspace(5., 15., 20), density=False)
old_grid = (bins[1:] + bins[:-1]) / 2.
new_grid_edges = np.linspace(5., 15., 40)
new_counts, new_grid, new_widths, new_edges = itm.rebin(
count, old_grid, new_grid_edges,
binning_scheme="Lin", negatives=True
)
plt.step(old_grid, count)
plt.step(new_grid, new_counts, ls='--')
old_grid_edges = np.linspace(80., 120., 100)
old_grid = (old_grid_edges[1:] + old_grid_edges[:-1]) / 2.
v = np.random.normal(size=10000) + 100
count, bins = np.histogram(v, bins=old_grid_edges)
old_grid_edges = np.linspace(80., 120., 100)
old_grid = (old_grid_edges[1:] + old_grid_edges[:-1]) / 2.
v = np.random.normal(size=10000) + 100
count, bins = np.histogram(v, bins=old_grid_edges)
new_grid_edges = np.linspace(80., 120., 200)
new_counts, new_grid, new_widths, new_edges = itm.rebin(
count, old_grid, new_grid_edges,
binning_scheme="Lin", negatives=True
)
plt.step(old_grid, count)
plt.step(new_grid, new_counts, ls='--')
plt.step(old_grid, count)
plt.step(new_grid, new_counts, ls='--')
plt.xlim(95., 105)
| examples/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import time
import seaborn as sns
import urllib.request
from tqdm import tqdm
import pandas as pd
# %matplotlib inline
# -
# # Quadratic Assignment Problem
# ## 1. Read data
# Popular QAP with loss function minimums: <br>
# - Nug12 12 578 (OPT) (12,7,9,3,4,8,11,1,5,6,10,2)
# - Nug14 14 1014 (OPT) (9,8,13,2,1,11,7,14,3,4,12,5,6,10)
# - Nug15 15 1150 (OPT) (1,2,13,8,9,4,3,14,7,11,10,15,6,5,12)
# - Nug16a 16 1610 (OPT) (9,14,2,15,16,3,10,12,8,11,6,5,7,1,4,13)
# - Nug16b 16 1240 (OPT) (16,12,13,8,4,2,9,11,15,10,7,3,14,6,1,5)
# - Nug17 17 1732 (OPT) (16,15,2,14,9,11,8,12,10,3,4,1,7,6,13,17,5)
# - Nug18 18 1930 (OPT) (10,3,14,2,18,6,7,12,15,4,5,1,11,8,17,13,9,16)
# - Nug20 20 2570 (OPT) (18,14,10,3,9,4,2,12,11,16,19,15,20,8,13,17,5,7,1,6)
# - Nug21 21 2438 (OPT) (4,21,3,9,13,2,5,14,18,11,16,10,6,15,20,19,8,7,1,12,17)
# - Nug22 22 3596 (OPT) (2,21,9,10,7,3,1,19,8,20,17,5,13,6,12,16,11,22,18,14,15)
# - Nug24 24 3488 (OPT) (17,8,11,23,4,20,15,19,22,18,3,14,1,10,7,9,16,21,24,12,6,13,5,2)
# - Nug25 25 3744 (OPT) (5,11,20,15,22,2,25,8,9,1,18,16,3,6,19,24,21,14,7,10,17,12,4,23,13) *
# - Nug27 27 5234 (OPT) (23,18,3,1,27,17,5,12,7,15,4,26,8,19,20,2,24,21,14,10,9,13,22,25,6,16,11) *
# - Nug28 28 5166 (OPT) (18,21,9,1,28,20,11,3,13,12,10,19,14,22,15,2,25,16,4,23,7,17,24,26,5,27,8,6) *
# - Nug30 30 6124 (OPT) (5 12 6 13 2 21 26 24 10 9 29 28 17 1 8 7 19 25 23 22 11 16 30 4 15 18 27 3 14 20)
def get_nug(no):
QAP_INSTANCE_URL = f'http://anjos.mgi.polymtl.ca/qaplib/data.d/nug{no}.dat'
qap_instance_file = urllib.request.urlopen(QAP_INSTANCE_URL)
line = qap_instance_file.readline()
n = int(line.decode()[:-1].split()[0])
A = np.empty((n, n))
qap_instance_file.readline()
for i in range(n):
line = qap_instance_file.readline()
A[i, :] = list(map(int, line.decode()[:-1].split()))
B = np.empty((n, n))
qap_instance_file.readline()
for i in range(n):
line = qap_instance_file.readline()
B[i, :] = list(map(int, line.decode()[:-1].split()))
return n, A, B
# +
n, A, B = get_nug(12)
print('Problem size: %d' % n)
print('Flow matrix:\n', A)
print('Distance matrix:\n', B)
# -
# ## 2. Objective function
def qap_objective_function(p):
s = 0.0
for i in range(n):
s += (A[i, :] * B[p, p[i]]).sum()
return s
p = [11, 6, 8, 2, 3, 7, 10, 0, 4, 5, 9, 1]
print(qap_objective_function(p), p)
# # 3. Random Sampling
# +
# %%time
T = 1000000
permutations = np.empty((T, n), dtype=np.int64)
costs = np.zeros(T)
for i in tqdm(range(T)):
permutations[i, :] = np.random.permutation(n)
costs[i] = qap_objective_function(permutations[i, :])
p = permutations[costs.argmin(), :]
print(qap_objective_function(p), p)
# +
plt.figure()
plt.hist(costs, bins=100, edgecolor='black')
plt.show()
print(costs.mean(), costs.std())
# -
# ## 4. Simulated Annealing
def qap_objective_function(p):
s = 0.0
for i in range(n):
s += (A[i, :] * B[p, p[i]]).sum()
return s
# +
q = p.copy()
i, j = 4, 8
q[i], q[j] = q[j], q[i]
# -
qap_objective_function(q)
qap_objective_function(p)
z = q.copy()
i, j = 2, 3
z[i], z[j] = z[j], z[i]
qap_objective_function(z)
def delta(p, r, s, A, B):
n = len(p)
ans = 0
q = p.copy()
q[r], q[s] = q[s], q[r]
for i in range(n):
for j in range(n):
ans += A[i][j] * B[p[i], p[j]] - A[i][j] * B[q[i], q[j]]
return -ans
def delta2(p, r, s, A, B):
n = len(p)
ans = 0
for k in range(n):
if k == r or k == s:
continue
ans += (A[s, k] - A[r, k]) * (B[p[s], p[k]] - B[p[r], p[k]])
return -2 * ans
# +
r, s = 4, 8
u, v = 2, 6
print(delta2(p, r, s, A, B))
print(delta2(q, u, v, A, B))
# -
for u in range(n):
for v in range(n):
a = delta2(p, r, s, A, B)
b = delta2(q, u, v, A, B)
c = delta2(p, u, v, A, B) + 2 * (A[r, u] - A[r, v] + A[s, v] - A[s, u]) * (B[q[s], q[u]] - B[q[s], q[v]] + B[q[r], q[v]] - B[q[r], q[u]])
if u in [r, s] or v in [r, s]:
2 + 2
else:
if b != c:
print(u, v)
print(a)
print(b)
print(c)
print()
# +
class SA:
def __init__(self, flow_m, dist_m, QAP_name='', T=10000, radius=1, alpha=1.0, cost_dist=False):
self.T = T
self.radius = radius
self.alpha = alpha
n1, n2 = flow_m.shape
n3, n4 = dist_m.shape
assert(n1 == n2 == n3 == n4)
self.FLOW = flow_m
self.DIST = dist_m
self.n = n1
self.last_perm = np.random.permutation(n1)
self.act_perm = self.last_perm.copy()
self.r, self.s = 0, 1
self.act_perm[0], self.act_perm[1] = self.act_perm[1], self.act_perm[0]
self.p_cost = self.qap_objective_function(self.act_perm)
self.costs = np.zeros(T)
self.all_perms = [self.act_perm]
self.all_jumps = []
self.QAP_name = QAP_name
self.cost_and_perms_dist = []
self.cost_dist = cost_dist
self.last_swap_costs = {}
self.act_swap_costs = {}
for u in range(self.n):
for v in range(u, self.n):
self.last_swap_costs[(u, v)] = self.delta(self.last_perm, u, v)
self.act_swap_costs[(u, v)] = self.delta(self.act_perm, u, v)
def delta(self, p, r, s):
'''
Change of Cost after swap(r, s)
'''
n = self.n
A, B = self.FLOW, self.DIST
ans = 0
for k in range(n):
if k == r or k == s:
continue
ans += (A[s, k] - A[r, k]) * (B[p[s], p[k]] - B[p[r], p[k]])
return -2 * ans
def qap_objective_function(self, p):
s = 0.0
for i in range(self.n):
s += (self.FLOW[i, :] * self.DIST[p, p[i]]).sum()
return s
def random_neighbor(self):
q = self.act_perm.copy()
for r in range(self.radius):
i, j = np.random.choice(self.n, 2, replace=False)
q[i], q[j] = q[j], q[i]
return q
def run(self):
for t in tqdm(range(self.T), desc='Simulated Annealing', position=0):
good_jump, random_jump = 0, 0
q = self.random_neighbor()
q_cost = self.qap_objective_function(q)
if(q_cost < self.p_cost):
if self.cost_dist:
self.cost_and_perms_dist.append((self.p_cost - q_cost))
self.act_perm, self.p_cost = q, q_cost
self.all_perms.append(self.act_perm)
good_jump = 1
elif(np.random.rand() < np.exp(-self.alpha * (q_cost - self.p_cost) * t / self.T)):
self.act_perm, self.p_cost = q, q_cost
self.all_perms.append(self.act_perm)
random_jump = 1
self.costs[t] = self.p_cost
self.all_jumps.append((good_jump, random_jump))
def update_swap_costs(self):
cost = 0
r, s = self.r, self.s
A, B = self.FLOW, self.DIST
q = self.act_perm
self.last_swap_costs = self.act_swap_costs
self.act_swap_costs = {}
for u in range(self.n):
for v in range(u, self.n):
if u in [r, s] or v in [r, s]:
# calculate (u, v) swap cost in O(N)
self.act_swap_costs[(u, v)] = self.delta(self.act_perm, u, v)
else:
# swap in O(1)
self.act_swap_costs[(u, v)] = self.last_swap_costs[u, v] \
+ 2 * (A[r, u] - A[r, v] + A[s, v] - A[s, u]) \
* (B[q[s], q[u]] - B[q[s], q[v]] + B[q[r], q[v]] - B[q[r], q[u]])
def run_faster(self):
'''
We make only single swaps per iter.
Insted of calculating qap_objective_function in every iteration,
we calculate cost of swapping every pair every time
we change a state.
qap_objective_function -> O(n^2) every itereation
new approach -> O(n^2) only when we change a state
'''
for t in tqdm(range(self.T), desc='Simulated Annealing', position=0):
good_jump, random_jump = 0, 0
u, v = np.random.choice(self.n, 2, replace=False)
q = self.act_perm.copy()
q[u], q[v] = q[v], q[u]
q_cost = self.act_swap_costs[(min(u, v), max(u, v))] + self.p_cost
# if q_cost != self.qap_objective_function(q):
# print('IMPLEMENTATION DOES NOT WORK!!!')
if(q_cost < self.p_cost):
self.act_perm, self.p_cost = q, q_cost
self.all_perms.append(self.act_perm)
self.r, self.s = u, v
self.last_perm = self.act_perm
self.update_swap_costs()
good_jump = 1
elif(np.random.rand() < np.exp(-self.alpha * (q_cost - self.p_cost) * t / self.T)):
self.act_perm, self.p_cost = q, q_cost
self.all_perms.append(self.act_perm)
self.r, self.s = u, v
self.last_perm = self.act_perm
self.update_swap_costs()
random_jump = 1
self.costs[t] = self.p_cost
self.all_jumps.append((good_jump, random_jump))
def plot_cost(self):
plt.figure(figsize=(15, 5))
plt.plot(self.costs)
plt.title('Cost function ' + self.QAP_name)
plt.show()
def plot_hist(self, bins):
plt.figure(figsize=(15, 5))
plt.hist(self.costs, bins=bins, edgecolor='black')
plt.title('Cost function histogram ' + self.QAP_name)
plt.show()
def plot_jumps(self):
x = np.array(self.all_jumps).reshape(-1, 50)
x = x.sum(axis=1)
f, ax = plt.subplots(2,1, figsize=(15,10))
ax[0].bar(range(x.shape[0]), x, color='green')
ax[1].bar(range(x.shape[0]), x, color='red')
ax[0].set_title('Successes')
ax[1].set_title('Accepted failures')
plt.show()
def plot_all(self, bins):
self.plot_cost()
self.plot_hist(bins=bins)
self.plot_jumps()
# -
# ### 4.1 Basic implementation
# +
# %%time
n, A, B = get_nug(12)
simulation = SA(flow_m=A, dist_m=B, QAP_name='nug12', T=500000, cost_dist=True)
simulation.run()
# -
simulation.costs.min()
simulation.plot_all(bins=50)
# ### 4.2 Improved implementation based on
# https://arxiv.org/pdf/1111.1353.pdf
# +
# %%time
n, A, B = get_nug(12)
simulation = SA(flow_m=A, dist_m=B, QAP_name='nug12', T=500000)
simulation.run_faster()
# -
simulation.costs.min()
# ## 5. Parameters tunning
# ### 5.1 Radius
# +
# %%time
scores_r = []
for r in range(1, 20):
print(f'r: {r}')
n, A, B = get_nug(12)
simulation = SA(flow_m=A, dist_m=B, QAP_name='nug12', T=200000, radius=r, cost_dist=True)
simulation.run()
scores_r.append((r, simulation.costs, simulation.cost_and_perms_dist))
# +
scores_r = np.array(scores_r)
plt.plot(scores_r[:, 0], list(map(lambda x: min(x), scores_r[:, 1])))
plt.xlabel('Distance between permutations')
plt.ylabel('Min cost')
# save scores
# pd.DataFrame(scores_r).to_csv('scores_for_different_radius.csv')
# +
# scores_r = pd.read_csv('scores_for_different_radius.csv', names=['r', 'cost'],
# header=None).reset_index(drop=True).iloc[1:]
# +
# distances between cost_f(perms) in successes
f, ax = plt.subplots(4, 5, figsize=(17,15))
for r in range(19):
ax[r // 5, r % 5].plot(scores_r[:, 2][r])
ax[r // 5, r % 5].set_title(f'r: {r}')
# -
# ### 5.2 alpha
# +
# %%time
scores_a = []
alphas = [0.01, 0.02, 0.03, 0.05, 0.1, 0.2, 0.3, 0.5, 0.7, 1.01, 1.03, 1.05, 1.1, 1.2, 1.4, 1.5, 1.7, 2, 4, 5, 10]
for a in alphas:
print(f'alpha: {a}')
n, A, B = get_nug(12)
simulation = SA(flow_m=A, dist_m=B, QAP_name='nug12', T=200000, alpha=a)
simulation.run_faster()
scores_a.append(simulation.costs)
# +
# save scores
# pd.DataFrame(scores_a).to_csv('scores_for_different_alpha.csv')
# -
scores_a = np.array(scores_a)
plt.figure(figsize=(15, 5))
plt.xticks(np.linspace(0, 10, 20))
plt.plot(alphas, scores_a.min(axis=1))
for i, a in enumerate(alphas):
print(a, ' -> ', scores_a.min(axis=1)[i])
# ## 6. SA on different QAP
# +
from urllib.error import HTTPError
nugs = []
for i in range(12, 30):
try:
x = get_nug(i)
nugs.append(x)
print(f'Nug {i} appended!')
except HTTPError:
print(f'Nug {i} wasn t found :c')
# +
# %%time
SCORES = []
for n, A, B in nugs:
print(f'Running nug: {n}')
simulation = SA(flow_m=A, dist_m=B, QAP_name=f'Nug {n}', T=1000000)
simulation.run_faster()
SCORES.append(simulation.costs)
save_costs = pd.DataFrame(simulation.costs)
save_costs.to_csv(f'scores_for_nug{n}.csv')
# +
SCORES = np.array(SCORES)
n, m = SCORES.shape
n, m
# -
f, ax = plt.subplots(4, 3, figsize=(17,15))
for i in range(n):
ax[i // 3, i % 3].plot(SCORES[i])
ax[i // 3, i % 3].set_title(f'nug{nugs[i][0]}')
print('Min cost:')
for i, s in enumerate(SCORES.min(axis=1)):
print(f'nug{nugs[i][0]}: {s}')
#
| Evo/Assignment1/SimulatedAnnealing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Residual Analysis
#
# By <NAME> and <NAME>
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
#
# ---
#
#
#
# ## Linear Regression
#
# Linear regression is one of our most fundamental modeling techniques. We use it to estimate a linear relationship between a set of independent variables $X_i$ and a dependent outcome variable $y$. Our model takes the form of:
#
# $$ y_i = \beta_{0} 1 + \beta_{i, 1} x_{i, 1} + \dots + \beta_{i, p} x_{i, p} + \epsilon_i = x_i'\beta + \epsilon_i $$
#
# For $i \in \{1, \dots, n\}$, where $n$ is the number of observations. We write this in vector form as:
#
# $$ y = X\beta + \epsilon $$
#
# Where $y$ is a $n \times 1$ vector, $X$ is a $n \times p$ matrix, $\beta$ is a $p \times 1$ vector of coefficients, and $\epsilon$ is a standard normal error term. Typically we call a model with $p = 1$ a simple linear regression and a model with $p > 1$ a multiple linear regression. More background information on regressions can be found in the lectures on [simple linear regression](https://www.quantopian.com/lectures#Linear-Regression) and [multiple linear regression](https://www.quantopian.com/lectures#Multiple-Linear-Regression).
#
# Whenever we build a model, there will be gaps between what a model predicts and what is observed in the sample. The differences between these values are known as the residuals of the model and can be used to check for some of the basic assumptions that go into the model. The key assumptions to check for are:
#
# * **Linear Fit:** The underlying relationship should be linear
# * **Homoscedastic:** The data should have no trend in the variance
# * **Independent and Identically Distributed:** The residuals of the regression should be independent and identically distributed (i.i.d.) and show no signs of serial correlation
#
# We can use the residuals to help diagnose whether the relationship we have estimated is real or spurious.
#
# Statistical error is a similar metric associated with regression analysis with one important difference: While residuals quantify the gap between a regression model predictions and the observed sample, statistical error is the difference between a regression model and the unobservable expected value. We use residuals in an attempt to estimate this error.
# Import libraries
import numpy as np
import pandas as pd
from statsmodels import regression
import statsmodels.api as sm
import statsmodels.stats.diagnostic as smd
import scipy.stats as stats
import matplotlib.pyplot as plt
import math
# # Simple Linear Regression
#
# First we'll define a function that performs linear regression and plots the results.
def linreg(X,Y):
# Running the linear regression
X = sm.add_constant(X)
model = sm.OLS(Y, X).fit()
B0 = model.params[0]
B1 = model.params[1]
X = X[:, 1]
# Return summary of the regression and plot results
X2 = np.linspace(X.min(), X.max(), 100)
Y_hat = X2 * B1 + B0
plt.scatter(X, Y, alpha=1) # Plot the raw data
plt.plot(X2, Y_hat, 'r', alpha=1); # Add the regression line, colored in red
plt.xlabel('X Value')
plt.ylabel('Y Value')
return model, B0, B1
# Let's define a toy relationship between $X$ and $Y$ that we can model with a linear regression. Here we define the relationship and construct a model on it, drawing the determined line of best fit with the regression parameters.
# +
n = 50
X = np.random.randint(0, 100, n)
epsilon = np.random.normal(0, 1, n)
Y = 10 + 0.5 * X + epsilon
linreg(X,Y)[0];
print "Line of best fit: Y = {0} + {1}*X".format(linreg(X, Y)[1], linreg(X, Y)[2])
# -
# This toy example has some generated noise, but all real data will also have noise. This is inherent in sampling from any sort of wild data-generating process. As a result, our line of best fit will never exactly fit the data (which is why it is only "best", not "perfect"). Having a model that fits every single observation that you have is a sure sign of [overfitting](https://www.quantopian.com/lectures/the-dangers-of-overfitting).
#
# For all fit models, there will be a difference between what the regression model predicts and what was observed, which is where residuals come in.
# ## Residuals
#
# The definition of a residual is the difference between what is observed in the sample and what is predicted by the regression. For any residual $r_i$, we express this as
#
# $$r_i = Y_i - \hat{Y_i}$$
#
# Where $Y_i$ is the observed $Y$-value and $\hat{Y}_i$ is the predicted Y-value. We plot these differences on the following graph:
# +
model, B0, B1 = linreg(X,Y)
residuals = model.resid
plt.errorbar(X,Y,xerr=0,yerr=[residuals,0*residuals],linestyle="None",color='Green');
# -
# We can pull the residuals directly out of the fit model.
residuals = model.resid
print residuals
# # Diagnosing Residuals
#
# Many of the assumptions that are necessary to have a valid linear regression model can be checked by identifying patterns in the residuals of that model. We can make a quick visual check by looking at the residual plot of a given model.
#
# With a residual plot, we look at the predicted values of the model versus the residuals themselves. What we want to see is just a cloud of unrelated points, like so:
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
plt.xlim([1,50]);
# What we want is a fairly random distribution of residuals. The points should form no discernible pattern. This would indicate that a plain linear model is likely a good fit. If we see any sort of trend, this might indicate the presence of autocorrelation or heteroscedasticity in the model.
# ## Appropriateness of a Linear Model
#
# By looking for patterns in residual plots we can determine whether a linear model is appropriate in the first place. A plain linear regression would not be appropriate for an underlying relationship of the form:
#
# $$Y = \beta_0 + \beta_1 X^2$$
#
# as a linear function would not be able to fully explain the relationship between $X$ and $Y$.
#
# If the relationship is not a good fit for a linear model, the residual plot will show a distinct pattern. In general, a residual plot of a linear regression on a non-linear relationship will show bias and be asymmetrical with respect to residual = 0 line while a residual plot of a linear regression on a linear relationship will be generally symmetrical over the residual = 0 axis.
#
# As an example, let's consider a new relationship between the variables $X$ and $Y$ that incorporates a quadratic term.
# +
n = 50
X = np.random.randint(0, 50, n)
epsilon = np.random.normal(0, 1, n)
Y_nonlinear = 10 - X**1.2 + epsilon
model = sm.OLS(Y_nonlinear, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
print 'beta_0: ', B0
print 'beta_1: ', B1
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
# The "inverted-U" shape shown by the residuals is a sign that a non-linear model might be a better fit than a linear one.
# ## Heteroscedasticity
#
# One of the main assumptions behind a linear regression is that the underlying data has a constant variance. If there are some parts of the data with a variance different from another part the data is not appropriate for a linear regression. **Heteroscedasticity** is a term that refers to data with non-constant variance, as opposed to homoscedasticity, when data has constant variance.
#
# Significant heteroscedasticity invalidates linear regression results by biasing the standard error of the model. As a result, we can't trust the outcomes of significance tests and confidence intervals generated from the model and its parameters.
#
# To avoid these consequences it is important to use residual plots to check for heteroscedasticity and adjust if necessary.
#
# As an example of detecting and correcting heteroscedasticity, let's consider yet another relationship between $X$ and $Y$:
# +
n = 50
X = np.random.randint(0, 100, n)
epsilon = np.random.normal(0, 1, n)
Y_heteroscedastic = 100 + 2*X + epsilon*X
model = sm.OLS(Y_heteroscedastic, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
# Heteroscedasticity often manifests as this spread, giving us a tapered cloud in one direction or another. As we move along in the $x$-axis, the magnitudes of the residuals are clearly increasing. A linear regression is unable to explain this varying variability and the regression standard errors will be biased.
#
# ### Statistical Methods for Detecting Heteroscedasticity
#
# Generally, we want to back up qualitative observations on a residual plot with a quantitative method. The residual plot led us to believe that the data might be heteroscedastic. Let's confirm that result with a statistical test.
#
# A common way to test for the presence of heteroscedasticity is the Breusch-Pagan hypothesis test. It's good to combine the qualitative analysis of a residual plot with the quantitative analysis of at least one hypothesis test. We can add the White test as well, but for now we will use only Breush-Pagan to test our relationship above. A function exists in the `statsmodels` package called `het_breushpagan` that simplifies the computation:
breusch_pagan_p = smd.het_breushpagan(model.resid, model.model.exog)[1]
print breusch_pagan_p
if breusch_pagan_p > 0.05:
print "The relationship is not heteroscedastic."
if breusch_pagan_p < 0.05:
print "The relationship is heteroscedastic."
# We set our confidence level at $\alpha = 0.05$, so a Breusch-Pagan p-value below $0.05$ tells us that the relationship is heteroscedastic. For more on hypothesis tests and interpreting p-values, refer to the [lecture on hypothesis testing.](https://www.quantopian.com/research/notebooks/Cloned%20from%20%22Quantopian%20Lecture%20Series%3A%20Hypothesis%20Testing%22%201.ipynb). Using a hypothesis test bears the risk of a false positive or a false negative, which is why it can be good to confirm with additional tests if we are skeptical.
# ### Adjusting for Heteroscedasticity
#
# If, after creating a residual plot and conducting tests, you believe you have heteroscedasticity, there are a number of methods you can use to attempt to adjust for it. The three we will focus on are differences analysis, log transformations, and Box-Cox transformations.
# #### Differences Analysis
#
# A differences analysis involves looking at the first-order differences between adjacent values. With this, we are looking at the changes from period to period of an independent variable rather than looking directly at its values. Often, by looking at the differences instead of the raw values, we can remove heteroscedasticity. We correct for it and can use the ensuing model on the differences.
# Finding first-order differences in Y_heteroscedastic
Y_heteroscedastic_diff = np.diff(Y_heteroscedastic)
# Now that we have stored the first-order differences of `Y_heteroscedastic` in `Y_heteroscedastic_diff` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
# +
model = sm.OLS(Y_heteroscedastic_diff, sm.add_constant(X[1:])).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
breusch_pagan_p = smd.het_breushpagan(residuals, model.model.exog)[1]
print breusch_pagan_p
if breusch_pagan_p > 0.05:
print "The relationship is not heteroscedastic."
if breusch_pagan_p < 0.05:
print "The relationship is heteroscedastic."
# *Note: This new regression was conducted on the differences between data, and therefore the regression output must be back-transformed to reach a prediction in the original scale. Since we regressed the differences, we can add our predicted difference onto the original data to get our estimate:*
#
# $$\hat{Y_i} = Y_{i-1} + \hat{Y}_{diff}$$
# #### Logarithmic Transformation
#
# Next, we apply a log transformation to the underlying data. A log transformation will bring residuals closer together and ideally remove heteroscedasticity. In many (though not all) cases, a log transformation is sufficient in stabilizing the variance of a relationship.
# Taking the log of the previous data Y_heteroscedastic and saving it in Y_heteroscedastic_log
Y_heteroscedastic_log = np.log(Y_heteroscedastic)
# Now that we have stored the log transformed version of `Y_heteroscedastic` in `Y_heteroscedastic_log` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
# +
model = sm.OLS(Y_heteroscedastic_log, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
# Running and interpreting a Breusch-Pagan test
breusch_pagan_p = smd.het_breushpagan(residuals, model.model.exog)[1]
print breusch_pagan_p
if breusch_pagan_p > 0.05:
print "The relationship is not heteroscedastic."
if breusch_pagan_p < 0.05:
print "The relationship is heteroscedastic."
# *Note: This new regression was conducted on the log of the original data. This means the scale has been altered and the regression estimates will lie on this transformed scale. To bring the estimates back to the original scale, you must back-transform the values using the inverse of the log:*
#
# $$\hat{Y} = e^{\log(\hat{Y})}$$
# #### Box-Cox Transformation
#
# Finally, we examine the Box-Cox transformation. The Box-Cox transformation is a powerful method that will work on many types of heteroscedastic relationships. The process works by testing all values of $\lambda$ within the range $[-5, 5]$ to see which makes the output of the following equation closest to being normally distributed:
# $$
# Y^{(\lambda)} = \begin{cases}
# \frac{Y^{\lambda}-1}{\lambda} & : \lambda \neq 0\\ \log{Y} & : \lambda = 0
# \end{cases}
# $$
#
# The "best" $\lambda$ will be used to transform the series along the above function. Instead of having to do all of this manually, we can simply use the `scipy` function `boxcox`. We use this to adjust $Y$ and hopefully remove heteroscedasticity.
#
# *Note: The Box-Cox transformation can only be used if all the data is positive*
# Finding a power transformation adjusted Y_heteroscedastic
Y_heteroscedastic_box_cox = stats.boxcox(Y_heteroscedastic)[0]
# Now that we have stored the power transformed version of `Y_heteroscedastic` in `Y_heteroscedastic_prime` let's repeat the regression and residual plot to see if the heteroscedasticity is still present:
# +
model = sm.OLS(Y_heteroscedastic_box_cox, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
# Running and interpreting a Breusch-Pagan test
breusch_pagan_p = smd.het_breushpagan(residuals, model.model.exog)[1]
print breusch_pagan_p
if breusch_pagan_p > 0.05:
print "The relationship is not heteroscedastic."
if breusch_pagan_p < 0.05:
print "The relationship is heteroscedastic."
# *Note: Now that the relationship is not heteroscedastic, a linear regression is appropriate. However, because the data was power transformed, the regression estimates will be on a different scale than the original data. This is why it is important to remember to back-transform results using the inverse of the Box-Cox function:*
#
# $$\hat{Y} = (Y^{(\lambda)}\lambda + 1)^{1/\lambda}$$
#
# ### GARCH Modeling
#
# Another approach to dealing with heteroscadasticity is through a GARCH (generalized autoregressive conditional heteroscedasticity) model. More information can be found in the [lecture on GARCH modeling](https://www.quantopian.com/lectures#ARCH,-GARCH,-and-GMM).
# ## Residuals and Autocorrelation
#
# Another assumption behind linear regressions is that the residuals are not autocorrelated. A series is autocorrelated when it is correlated with a delayed version of itself. An example of a potentially autocorrelated time series series would be daily high temperatures. Today's temperature gives you information on tomorrow's temperature with reasonable confidence (i.e. if it is 90 °F today, you can be very confident that it will not be below freezing tomorrow). A series of fair die rolls, however, would not be autocorrelated as seeing one roll gives you no information on what the next might be. Each roll is independent of the last.
#
# In finance, stock prices are usually autocorrelated while stock returns are independent from one day to the next. We represent a time dependency on previous values like so:
#
# $$Y_i = Y_{i-1} + \epsilon$$
#
# If the residuals of a model are autocorrelated, you will be able to make predictions about adjacent residuals. In the case of $Y$, we know the data will be autocorrelated because we can make predictions based on adjacent residuals being close to one another.
# +
n = 50
X = np.linspace(0, n, n)
Y_autocorrelated = np.zeros(n)
Y_autocorrelated[0] = 50
for t in range(1, n):
Y_autocorrelated[t] = Y_autocorrelated[t-1] + np.random.normal(0, 1)
# Regressing X and Y_autocorrelated
model = sm.OLS(Y_autocorrelated, sm.add_constant(X)).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# -
# Autocorrelation in the residuals in this example is not explicitly obvious, so our check is more to make absolutely certain.
#
# ### Statistical Methods for Detecting Autocorrelation
#
# As with all statistical properties, we require a statistical test to ultimately decide whether there is autocorrelation in our residuals or not. To this end, we use a Ljung-Box test.
#
# A Ljung-Box test is used to detect autocorrelation in a time series. The Ljung-Box test examines autocorrelation at all lag intervals below a specified maximum and returns arrays containing the outputs for every tested lag interval.
#
# Let's use the `acorr_ljungbox` function in `statsmodels` to test for autocorrelation in the residuals of our above model. We use a max lag interval of $10$, and see if any of the lags have significant autocorrelation:
# +
ljung_box = smd.acorr_ljungbox(residuals, lags = 10)
print "Lagrange Multiplier Statistics:", ljung_box[0]
print "\nP-values:", ljung_box[1], "\n"
if any(ljung_box[1] < 0.05):
print "The residuals are autocorrelated."
else:
print "The residuals are not autocorrelated."
# -
# Because the Ljung-Box test yielded a p-value below $0.05$ for at least one lag interval, we can conclude that the residuals of our model are autocorrelated.
# ## Adjusting for Autocorrelation
#
# We can adjust for autocorrelation in many of the same ways that we adjust for heteroscedasticity. Let's see if a model on the first-order differences of $Y$ has autocorrelated residuals:
# Finding first-order differences in Y_autocorrelated
Y_autocorrelated_diff = np.diff(Y_autocorrelated)
# +
model = sm.OLS(Y_autocorrelated_diff, sm.add_constant(X[1:])).fit()
B0, B1 = model.params
residuals = model.resid
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('Predicted Values');
plt.ylabel('Residuals');
# Running and interpreting a Ljung-Box test
ljung_box = smd.acorr_ljungbox(residuals, lags = 10)
print "P-values:", ljung_box[1], "\n"
if any(ljung_box[1] < 0.05):
print "The residuals are autocorrelated."
else:
print "The residuals are not autocorrelated."
# -
# *Note: This new regression was conducted on the differences between data, and therefore the regression output must be back-transformed to reach a prediction in the original scale. Since we regressed the differences, we can add our predicted difference onto the original data to get our estimate:*
#
# $$\hat{Y_i} = Y_{i-1} + \hat{Y_{diff}}$$
# We can also perform a log transformation, if we so choose. This process is identical to the one we performed on the heteroscedastic data up above, so we will leave it out this time.
# # Example: Market Beta Calculation
#
# Let's calculate the market beta between TSLA and SPY using a simple linear regression, and then conduct a residual analysis on the regression to ensure the validity of our results. To regress TSLA and SPY, we will focus on their returns, not their price, and set SPY returns as our independent variable and TSLA returns as our outcome variable. The regression will give us a line of best fit:
#
# $$\hat{r_{TSLA}} = \hat{\beta_0} + \hat{\beta_1}r_{SPY}$$
#
# The slope of the regression line $\hat{\beta_1}$ will represent our market beta, as for every $r$ percent change in the returns of SPY, the predicted returns of TSLA will change by $\hat{\beta_1}$.
#
# Let's start by conducting the regression the returns of the two assets.
# +
start = '2014-01-01'
end = '2015-01-01'
asset = get_pricing('TSLA', fields='price', start_date=start, end_date=end)
benchmark = get_pricing('SPY', fields='price', start_date=start, end_date=end)
# We have to take the percent changes to get to returns
# Get rid of the first (0th) element because it is NAN
r_a = asset.pct_change()[1:].values
r_b = benchmark.pct_change()[1:].values
# Regressing the benchmark b and asset a
r_b = sm.add_constant(r_b)
model = sm.OLS(r_a, r_b).fit()
r_b = r_b[:, 1]
B0, B1 = model.params
# Plotting the regression
A_hat = (B1*r_b + B0)
plt.scatter(r_b, r_a, alpha=1) # Plot the raw data
plt.plot(r_b, A_hat, 'r', alpha=1); # Add the regression line, colored in red
plt.xlabel('TSLA Returns')
plt.ylabel('SPY Returns')
# Print our result
print "Estimated TSLA Beta:", B1
# Calculating the residuals
residuals = model.resid
# -
# Our regression yielded an estimated market beta of 1.9253; according to the regression, for every 1% in return we see from the SPY, we should see 1.92% from TSLA.
#
# Now that we have the regression results and residuals, we can conduct our residual analysis. Our first step will be to plot the residuals and look for any red flags:
plt.scatter(model.predict(), residuals);
plt.axhline(0, color='red')
plt.xlabel('TSLA Returns');
plt.ylabel('Residuals');
# By simply observing the distribution of residuals, it does not seem as if there are any abnormalities. The distribution is relatively random and no patterns can be observed (the clustering around the origin is a result of the nature of returns to cluster around 0 and is not a red flag). Our qualitative conclusion is that the data is homoscedastic and not autocorrelated and therefore satisfies the assumptions for linear regression.
#
# ###Breusch-Pagan Heteroscedasticity Test
#
# Our qualitative assessment of the residual plot is nicely supplemented with a couple statistical tests. Let's begin by testing for heteroscedasticity using a Breusch-Pagan test. Using the `het_breuschpagan` function from the statsmodels package:
# +
bp_test = smd.het_breushpagan(residuals, model.model.exog)
print "Lagrange Multiplier Statistic:", bp_test[0]
print "P-value:", bp_test[1]
print "f-value:", bp_test[2]
print "f_p-value:", bp_test[3], "\n"
if bp_test[1] > 0.05:
print "The relationship is not heteroscedastic."
if bp_test[1] < 0.05:
print "The relationship is heteroscedastic."
# -
# Because the P-value is greater than 0.05, we do not have enough evidence to reject the null hypothesis that the relationship is homoscedastic. This result matches up with our qualitative conclusion.
# ###Ljung-Box Autocorrelation Test
#
# Let's also check for autocorrelation quantitatively using a Ljung-Box test. Using the `acorr_ljungbox` function from the statsmodels package and the default maximum lag:
ljung_box = smd.acorr_ljungbox(r_a)
print "P-Values:", ljung_box[1], "\n"
if any(ljung_box[1] < 0.05):
print "The residuals are autocorrelated."
else:
print "The residuals are not autocorrelated."
# Because the Ljung-Box test yielded p-values above 0.05 for all lags, we can conclude that the residuals are not autocorrelated. This result matches up with our qualitative conclusion.
#
# After having visually assessed the residual plot of the regression and then backing it up using statistical tests, we can conclude that the data satisfies the main assumptions and the linear model is valid.
# ## References
# * "Analysis of Financial Time Series", by <NAME>
# *This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| quantopian/lectures/Residuals_Analysis/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Notebook to implement path searches in graph with -
# 1. Networkx algorithms
# 2. Discovery patterns
# 3. SPARQL queries
#
# * Author: <NAME>
# * Created: 2021-09-24
# * Last edited: 2021-09-24
#
# We combine the PheKnowLator and machine reading graphs (currently separate for each NP) using Networkx and search paths in the combined graph.
import os
import os.path
import networkx as nx
import json
import urllib
import traceback
from itertools import islice
from rdflib import Graph, URIRef, BNode, Namespace, Literal
from rdflib.namespace import RDF, OWL
from tqdm import tqdm
import json
import hashlib
import pickle
import pandas as pd
import numpy as np
#import pheknowlator kg_utils
import sys
sys.path.append('../')
from pkt_kg.utils import *
KG_PATH = '/home/sanya/PheKnowLatorv2/resources/knowledge_graphs/'
MR_PATH = '/home/sanya/PheKnowLatorv2/machine_read/output_graphs/'
KG_NAME = 'PheKnowLator_v3.0.0_full_instance_inverseRelations_OWLNETS_NetworkxMultiDiGraph.gpickle'
#MR_GRAPH_NAME = 'machineread_greentea_version1.gpickle'
MR_GRAPH_NAME_GT = 'machineread_greentea_version2.gpickle'
MR_GRAPH_NAME_KT = 'machineread_kratom_version1.gpickle'
NodeLabelsFilePL = 'PheKnowLator_v3.0.0_full_instance_inverseRelations_OWLNETS_NodeLabels.txt'
NodeLabelsFileMR_gt = 'machineread_greentea_version2_NodeLabels.tsv'
NodeLabelsFileMR_kt = 'machineread_kratom_version1_NodeLabels.tsv'
#create dictionary for node labels from node labels files
df1 = pd.read_csv(KG_PATH+NodeLabelsFilePL, sep='\t')
df2 = pd.read_csv(MR_PATH+NodeLabelsFileMR_gt, sep='\t')
df3 = pd.read_csv(MR_PATH+NodeLabelsFileMR_kt, sep='\t')
df1.head()
df2.head()
df3.head()
nodeLabels = {}
for i in range(len(df1.index)):
uri = df1.at[i, 'entity_uri']
if isinstance(uri, str):
uri = uri.replace('<', '')
uri = uri.replace('>', '')
if uri not in nodeLabels:
nodeLabels[uri] = df1.at[i, 'label']
len(nodeLabels)
#N(nodeLabels) = 753217 (both PL and MR nodes combined)
for i in range(len(df2.index)):
uri = df2.at[i, 'entity_uri']
if isinstance(uri, str):
uri = uri.replace('<', '')
uri = uri.replace('>', '')
if uri not in nodeLabels:
nodeLabels[uri] = df2.at[i, 'label']
len(nodeLabels)
for i in range(len(df3.index)):
uri = df3.at[i, 'entity_uri']
if isinstance(uri, str):
uri = uri.replace('<', '')
uri = uri.replace('>', '')
if uri not in nodeLabels:
nodeLabels[uri] = df3.at[i, 'label']
len(nodeLabels)
#save to pickle file
#with open(KG_PATH+'nodeLabels_20211005.pickle', 'wb') as filep:
#pickle.dump(nodeLabels, filep)
with open(KG_PATH+'nodeLabels_20211014.pickle', 'rb') as filep:
nodeLabels = pickle.load(filep)
pl_kg = nx.read_gpickle(KG_PATH+KG_NAME)
# +
# get the number of nodes, edges, and self-loops
nodes = nx.number_of_nodes(pl_kg)
edges = nx.number_of_edges(pl_kg)
self_loops = nx.number_of_selfloops(pl_kg)
print('There are {} nodes, {} edges, and {} self-loop(s)'.format(nodes, edges, self_loops))
# get degree information
avg_degree = float(edges) / nodes
print('The Average Degree is {}'.format(avg_degree))
# +
# get 5 nodes with the highest degress
n_deg = sorted([(str(x[0]), x[1]) for x in pl_kg.degree], key=lambda x: x[1], reverse=1)[:6]
for x in n_deg:
print('Label: {}'.format(nodeLabels[x[0]]))
print('{} (degree={})'.format(x[0], x[1]))
# get network density
density = nx.density(pl_kg)
print('The density of the graph is: {}'.format(density))
# -
mr_kg = nx.read_gpickle(MR_PATH+MR_GRAPH_NAME_GT)
mr_kg2 = nx.read_gpickle(MR_PATH+MR_GRAPH_NAME_KT)
# +
# get the number of nodes, edges, and self-loops
print('Green Tea Machine Read: ')
nodes = nx.number_of_nodes(mr_kg)
edges = nx.number_of_edges(mr_kg)
self_loops = nx.number_of_selfloops(mr_kg)
print('There are {} nodes, {} edges, and {} self-loop(s)'.format(nodes, edges, self_loops))
# get degree information
avg_degree = float(edges) / nodes
print('The Average Degree is {}'.format(avg_degree))
print('Kratom Machine Read: ')
nodes = nx.number_of_nodes(mr_kg2)
edges = nx.number_of_edges(mr_kg2)
self_loops = nx.number_of_selfloops(mr_kg2)
print('There are {} nodes, {} edges, and {} self-loop(s)'.format(nodes, edges, self_loops))
# get degree information
avg_degree = float(edges) / nodes
print('The Average Degree is {}'.format(avg_degree))
# +
# get 5 nodes with the highest degress
print('Green Tea')
n_deg = sorted([(str(x[0]), x[1]) for x in mr_kg.degree], key=lambda x: x[1], reverse=1)[:6]
for x in n_deg:
print('Label: {}'.format(nodeLabels[x[0]]))
print('{} (degree={})'.format(x[0], x[1]))
# get network density
density = nx.density(mr_kg)
print('The density of the graph is: {}'.format(density))
print('Kratom')
# get 5 nodes with the highest degress
n_deg = sorted([(str(x[0]), x[1]) for x in mr_kg2.degree], key=lambda x: x[1], reverse=1)[:6]
for x in n_deg:
print('Label: {}'.format(nodeLabels[x[0]]))
print('{} (degree={})'.format(x[0], x[1]))
# get network density
density = nx.density(mr_kg2)
print('The density of the graph is: {}'.format(density))
# -
#combine graphs - PL and MR
nx_graph = nx.compose_all([pl_kg, mr_kg, mr_kg2])
print(type(nx_graph))
# +
# get the number of nodes, edges, and self-loops
nodes = nx.number_of_nodes(nx_graph)
edges = nx.number_of_edges(nx_graph)
self_loops = nx.number_of_selfloops(nx_graph)
print('There are {} nodes, {} edges, and {} self-loop(s)'.format(nodes, edges, self_loops))
# get degree information
avg_degree = float(edges) / nodes
print('The Average Degree is {}'.format(avg_degree))
# +
# get 5 nodes with the highest degress
n_deg = sorted([(str(x[0]), x[1]) for x in nx_graph.degree], key=lambda x: x[1], reverse=1)[:6]
for x in n_deg:
print('{} (degree={})'.format(x[0], x[1]))
# get network density
density = nx.density(nx_graph)
print('The density of the graph is: {}'.format(density))
# +
mr_kg_comb = nx.compose(mr_kg, mr_kg2)
# get the number of nodes, edges, and self-loops
nodes = nx.number_of_nodes(mr_kg_comb)
edges = nx.number_of_edges(mr_kg_comb)
self_loops = nx.number_of_selfloops(mr_kg_comb)
print('There are {} nodes, {} edges, and {} self-loop(s)'.format(nodes, edges, self_loops))
# get degree information
avg_degree = float(edges) / nodes
print('The Average Degree is {}'.format(avg_degree))
# +
# get 5 nodes with the highest degress
n_deg = sorted([(str(x[0]), x[1]) for x in mr_kg_comb.degree], key=lambda x: x[1], reverse=1)[:6]
for x in n_deg:
print('{} (degree={})'.format(x[0], x[1]))
# get network density
density = nx.density(mr_kg_comb)
print('The density of the graph is: {}'.format(density))
# -
#nodes and edges examples
nodes = list(nx_graph.nodes(data=True))
for x in nodes:
print(x)
break
i = 0
for u, v, keys in nx_graph.edges(keys=True):
i = i+1
print('Edge', i)
print(u, nodeLabels[str(u)])
print(keys, nodeLabels[str(keys)])
print(v, nodeLabels[str(v)])
if i==10:
break
#Useful functions
#nx_graph.get_edge_data(u, v, key=None, default=None])
#Returns the attribute dictionary associated with edge (u, v).
#key = hashable identifier, optional (default=None), Return data only for the edge with specified key.
node1 = URIRef('http://purl.obolibrary.org/obo/CHEBI_47495')
node2 = URIRef('http://purl.obolibrary.org/obo/GO_0031325')
edge_keys = list(nx_graph.get_edge_data(node1, node2).keys())
for item in edge_keys:
print(str(item))
print(nodeLabels[str(item)])
node1
node2
nx_graph[node1][node2]
nx_graph[node1][node2]
ewt = [e['weight'] for e in nx_graph[node1][node2].values()]
ewt
nx_graph.get_edge_data(node1, node2, default=0)
#nx_graph.edges[node1, node2, 'key'] = key is hashable of triple -- how do we decipher/unhash?
nx_graph.edges[node1, node2]
# ## Path Searches
# 1. Single source shortest path (saved)
# 2. k-simple paths (saved for cyp3a4, midazolam)
# 3. Bidirectional shortest paths (in nb)
# 4. Shortest paths - do
DIR_OUT = '/home/sanya/PheKnowLatorv2/output_files/'
obo = Namespace('http://purl.obolibrary.org/obo/')
napdi = Namespace('http://napdi.org/napdi_srs_imports:')
# Functions. Create function for -
# 1. Get path narrative given path or list of paths
# 2. Get path URIs given path or list of paths
# 3. Get path with machine reading output from 2017 and prior
# 4. Save path with labels to file
#
def get_path_labels(path):
path_labels = []
if len(path) < 1:
print('Path length 1, skipping')
return
for edge in zip(path, path[1:]):
data = nx_graph.get_edge_data(*edge)
pred = list(data.keys())[0]
node1_lab = str(edge[0])
node2_lab = str(edge[1])
if node1_lab in nodeLabels:
node1_lab = nodeLabels[node1_lab]
if node2_lab in nodeLabels:
node2_lab = nodeLabels[node2_lab]
pred_lab = nodeLabels[str(pred)]
if list(data.values())[0]:
if 'source_graph' in list(data.values())[0]:
source_graph = 'machine_read'
else:
source_graph = ''
else:
source_graph = ''
labels = [node1_lab, pred_lab, node2_lab, source_graph]
path_labels.append(labels)
return path_labels
def get_path_uri(path):
path_uri = []
if len(path) < 1:
print('Path length 1, skipping')
return
for edge in zip(path, path[1:]):
data = nx_graph.get_edge_data(*edge)
pred = list(data.keys())[0]
attribute = list(data.values())
uri = [str(edge[0]), pred, str(edge[1]), attribute]
path_uri.append(uri)
return path_uri
#get shortest path from green tea leaf
greentea_path = nx.single_source_shortest_path(nx_graph, napdi.camellia_sinensis_leaf)
type(greentea_path)
save1 = 'greentea_single_source_shortest_path_50.txt'
#get 20 paths from green tea single source shortest path
#if returned paths are dictionary
count = 0
for target, node_list in greentea_path.items():
count += 1
if target != napdi.camellia_sinensis_leaf:
if str(target) not in nodeLabels:
target_label = str(target).split('/')[-1]
else:
target_label = nodeLabels[str(target)]
print('\n{} - {} Path:'.format(str(napdi.camellia_sinensis_leaf).split('/')[-1], target_label))
path_labels = get_path_labels(node_list)
print(path_labels)
if count == 20:
break
#save 100 paths from green tea single source shortest path to file
#if returned paths are dictionary
count = 0
file_save = open(DIR_OUT+save1, 'w')
for target, node_list in greentea_path.items():
count += 1
if target != napdi.camellia_sinensis_leaf:
if str(target) not in nodeLabels:
target_label = str(target).split('/')[-1]
else:
target_label = nodeLabels[str(target)]
file_save.write('\n{} - {} Path:\n'.format(str(napdi.camellia_sinensis_leaf).split('/')[-1], target_label))
path_labels = get_path_labels(node_list)
for triples in path_labels:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
if count == 100:
break
file_save.close()
# +
#obo.CHEBI_83161 - St. Johns Wort extract (to test graph)
# -
#green tea and warfarin
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_10033)
pathx
#scratch try
for edge in zip(pathx,pathx[1:]):
data = nx_graph.get_edge_data(*edge)
print('Edge info: ')
print(data.values())
print('source_graph' in list(data.values())[0])
path_labels
epicatechin = obo.CHEBI_90
catechin = obo.CHEBI_23053
egcg = obo.CHEBI_4806
greentea = napdi.camellia_sinensis_leaf
edge_path_test = nx.all_simple_edge_paths(nx_graph, catechin, obo.PR_P08684, 10)
i = 0
for x in edge_path_test:
print(x)
i += 1
if i==3:
break
#returns simple paths nodes and edges,
def k_simple_edge_paths(G, source, target, k, shortestLen):
paths = nx.all_simple_edge_paths(G, source, target, cutoff=shortestLen+20)
path_l = []
path_n = []
i = 0
while i<k:
try:
print('[info] applying next operator to search for a simple path of max length {}'.format(shortestLen+20))
path = next(paths)
except StopIteration:
break
print('[info] Simple path found of length {}'.format(len(path)))
if len(path) > shortestLen:
print('[info] Simple path length greater than shortest path length ({}) so adding to results'.format(shortestLen))
path_l.append(path)
i += 1
for path in path_l:
triple_list = []
for triple in path:
subj_lab = ''
pred_lab = ''
obj_lab = ''
subj = str(triple[0])
pred = str(triple[2])
obj = str(triple[1])
if subj in nodeLabels:
subj_lab = nodeLabels[subj]
if obj in nodeLabels:
obj_lab = nodeLabels[obj]
if pred in nodeLabels:
pred_lab = nodeLabels[pred]
triple_labels = (subj_lab, pred_lab, obj_lab)
triple_list.append(triple_labels)
path_n.append(triple_list)
return path_l, path_n
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, greentea, obo.PR_P08684, 10, 0)
source = str(greentea)
target = str(obo.PR_P08684)
save2 = 'greentea_cyp3a4_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
pathx = nx.bidirectional_shortest_path(nx_graph, obo.CHEBI_23053, obo.PR_P08684)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
source = str(obo.CHEBI_23053)
target = str(obo.PR_P08684)
save2 = 'catechin_cyp3a4_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, obo.CHEBI_23053, obo.PR_P08684, 20, 2)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
source = str(obo.CHEBI_4806)
target = str(obo.CHEBI_41879)
save2 = 'EGCG_dexamethasone_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, obo.CHEBI_4806, obo.CHEBI_41879, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
source = str(obo.CHEBI_4806)
target = str(obo.UBERON_0000468)
save2 = 'EGCG_bodyweight_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, obo.CHEBI_4806, obo.UBERON_0000468, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
for item in zip(cyp3a4_edge_path_labs[0], cyp3a4_edge_paths[0]):
print(item)
source = str(obo.CHEBI_23053)
target = str(obo.PR_P08684)
save2 = 'catechin_cyp3a4_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, obo.CHEBI_23053, obo.PR_P08684, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
source = str(obo.CHEBI_23053)
target = str(obo.HP_0003074)
save2 = 'catechin_hyperglycemia_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, obo.CHEBI_23053, obo.HP_0003074, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
source = str(napdi.camellia_sinensis_leaf)
target = str(obo.PR_O08684)
save2 = 'greentea_cyp3a4_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, napdi.camellia_sinensis_leaf, obo.PR_P08684, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
cyp3a4_edge_path_labs[0]
cyp3a4_edge_paths[0]
def k_shortest_paths(G, source, target, k, weight='weight'):
return list(islice(nx.all_shortest_paths(G, source, target, weight=weight), k))
#only returns node list, use get_path_labels to generate edges and get labels
def k_simple_paths(G, source, target, k, shortestLen):
paths = nx.all_simple_paths(G, source, target, cutoff=shortestLen+20)
path_l = []
i = 0
while i < k:
try:
print('[info] applying next operator to search for a simple path of max length {}'.format(shortestLen+20))
path = next(paths)
except StopIteration:
break
print('[info] Simple path found of length {}'.format(len(path)))
if len(path) > shortestLen:
print('[info] Simple path length greater than shortest path length ({}) so adding to results'.format(shortestLen))
path_l.append(path)
i += 1
return path_l
cyp3a4_paths = k_simple_paths(nx_graph, napdi.camellia_sinensis_leaf, obo.PR_P08684, 10, 4)
str(obo.PR_P08684).split('/')[-1]
#if returned paths are list
#simple paths with max length 25
save2 = 'greentea_cyp3a4_simple_paths_10.txt'
file_save = open(DIR_OUT+save2, 'w')
source = str(napdi.camellia_sinensis_leaf)
target = str(obo.PR_P08684)
source_label = source
target_label = target
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=24):\n'.format(source_label, target_label))
i = 0
for node_list in cyp3a4_paths:
file_save.write('\nPATH: '+str(i)+'\n')
path_labels = get_path_labels(node_list)
for triples in path_labels:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
source = str(napdi.camellia_sinensis_leaf)
target = str(obo.CHEBI_6931)
save2 = 'greentea_midazolam_simple_paths_20.txt'
file_save = open(DIR_OUT+save2, 'w')
cyp3a4_edge_paths, cyp3a4_edge_path_labs = k_simple_edge_paths(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_6931, 20, 0)
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for path_list in cyp3a4_edge_path_labs:
file_save.write('\nPATH: '+str(i)+'\n')
for triples in path_list:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
#if returned paths are list
#simple paths with max length 25
save3 = 'greentea_midazolam_simple_paths_10.txt'
file_save = open(DIR_OUT+save3, 'w')
source = str(napdi.camellia_sinensis_leaf)
target = str(obo.CHEBI_6931)
source_label = source
target_label = target
if source in nodeLabels:
source_label = nodeLabels[source]
if target in nodeLabels:
target_label = nodeLabels[target]
file_save.write('\n{} - {} Simple Path (cutoff=20):\n'.format(source_label, target_label))
i = 0
for node_list in midazolam_paths:
file_save.write('\nPATH: '+str(i)+'\n')
path_labels = get_path_labels(node_list)
for triples in path_labels:
for item in triples:
file_save.write(str(item)+' ')
file_save.write('\n')
i += 1
file_save.close()
##Bidirectional shortest paths
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_10033)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.PR_P08684)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_6931)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.HP_0003418)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_9150)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
pathx = nx.bidirectional_shortest_path(nx_graph, napdi.camellia_sinensis_leaf, obo.CHEBI_7444)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
# KRATOM
kratom = napdi.mitragyna_speciosa
mitragynine = obo.CHEBI_6956
hydroxy_mitragynine = napdi['7_hydroxy_mitragynine']
hydroxy_mitragynine
pathx = nx.bidirectional_shortest_path(nx_graph, kratom, obo.PR_P08684)
path_labels = get_path_labels(pathx)
for triples in path_labels:
print(triples)
for key in nodeLabels:
print(key)
print(nodeLabels[key])
break
# ### Path searches with MR nodes as end points - predications with highest belief scores
# 1. Get MR predications with belief scores > 0.65
# 2. Use subject and object nodes as start and end points for simple path searches (shortest path would just be direct link between the nodes)
df = pd.read_csv('../machine_read/greentea_pmid_all_predicates_umls_processed.tsv', sep='\t')
df.head()
df.info()
df = df.loc[df['belief'] > 0.8]
df.info()
df = df.reset_index(drop=True)
df.head()
df = df.sort_values(by=['belief'], ascending=False)
df = df.reset_index(drop=True)
df.head()
df.to_csv('../machine_read/MR_triples_searchpath.tsv', sep='\t', index=False)
#Subject-Object pairs (testing paths) for triples from machine reading with belief scores > 0.8
'''
1. catechin (CHEBI_90) -> ABCB1 (), biosynthetic process (), transport (), apoptotic process (), coronary disease (),
cholesterol (), myocardial eschemia, cisplatin, heart disease, glucose, glucose import, glucose metabolic process,
hyperglycemia, intestinal absorption
2. epigallocatechin gallate (CHEBI_4806) -> quinone, paracetamol, Endoplasmic Reticulum Stress, ATP, ATPase, autophagy, bile acid,
transport, cell death, cholesterol, cisplatin, dexamethasone, diclofenac, digoxin, dopamine, drug metabolic process,
erythromycin, glutathione, heart failure, hemolysis, angiotensin-2, cortisol, insulin secretion, insulin resistance,
liver failure, nadolol, obesity, quercetin, tamoxifen, verapamil
3. greentea -> atorvastatin, rosuvastatin, benzo[a]pyrene, cardiovascular disease, stroke, cholesterol,
Myocardial Ischemia, Coronary Disease, Diabetes Mellitus, diclofenac, digoxin, doxorubicin, hypertension, liver disease,
nadolol, obesity, warfarin, glucose import, glutathione
EXTENDED LISTS BELOW
'''
catechin_list = ['ABCB1_gene', 'Anabolism', 'Biological_Transport', 'Apoptosis', 'Cell_Proliferation',
'Coronary_Arteriosclerosis', 'Cholesterol', 'Cytochrome_P-450_CYP1A1', 'Cytochrome_P-450_CYP1A2',
'Cytochrome_P-450_CYP3A4', 'Insulin_Secretion', 'Cisplatin', 'Heart_Diseases', 'Glucose',
'glucose_uptake', 'glucose_transport', 'Hyperglycemia', 'Obesity', 'P-Glycoprotein',
'UGT1A1_gene', 'Weight_decreased']
egcg_list = ['1,4-benzoquinone', 'ABCA1_gene', 'Acetaminophen', 'Adenosine_Triphosphatases', 'Autophagy',
'Bile_Acids', 'Bilirubin', 'Biological_Transport', 'Body_Weight', 'BRCA1_protein,_human',
'Cell_Death', 'Cell_Proliferation', 'Cholesterol', 'Cisplatin', 'Collagen', 'Coronary_Arteriosclerosis',
'Cytochrome_P-450_CYP1A1', 'Cytochrome_P-450_CYP1A2', 'Cytochrome_P-450_CYP3A4', 'Cytochrome_P-450_CYP2D6',
'Cytochrome_P-450_CYP2C19', 'drug_metabolism', 'Dexamethasone', 'Diclofenac', 'Digoxin', 'Dopamine',
'GA-Binding_Protein_Transcription_Factor', 'Gluconeogenesis', 'Glucose_Transporter',
'glucose_transport', 'glucose_uptake', 'Glutathione', 'Glycogen', 'Erythromycin', 'Heart_failure',
'Hemolysis_(disorder)', 'Inflammation', 'Hydrocortisone',
'Interleukin-1', 'Interleukin-6', 'Intestinal_Absorption', 'rosoxacin', 'UGT1A1_gene',
'Insulin_Secretion', 'Insulin_Resistance', 'Liver_Failure',
'Nadolol', 'Obesity', 'Quercetin', 'Tamoxifen', 'Verapamil']
greentea_list = ['ABCB1_gene', 'ABCG2_gene', 'Acetaminophen', 'Biological_Transport', 'Cardiovascular_Diseases',
'Cerebrovascular_accident', 'Coronary_Arteriosclerosis', 'atorvastatin', 'Benzopyrenes', 'Cholesterol',
'Cytochrome_P450', 'Cytochromes', 'Cytochrome_P-450_CYP1A1', 'Cytochrome_P-450_CYP1A2',
'Cytochrome_P-450_CYP3A4', 'Diabetes_Mellitus', 'Diclofenac', 'Digoxin', 'Doxorubicin', 'glucose_transport',
'Hypertensive_disease', 'Hay_fever', 'Interleukin-10', 'Lipid_Metabolism', 'Liver_diseases',
'Low-Density_Lipoproteins', 'Nadolol', 'Obesity', 'glucose_uptake', 'Glutathione',
'SLC2A1_protein,_human', 'SLC5A1_gene', 'SLCO1A2_gene', 'SLCO2B1_gene', 'Warfarin',
'rosuvastatin', 'rosoxacin', 'TNFSF11_protein,_human', 'TRPA1_gene', 'TRPV1_gene']
#get OBO identifiers from dataframe
node_dict = {}
for item in catechin_list:
if item not in node_dict:
print(item)
obo_id = df.loc[df['object_name'] == item]['object_obo'].values[0]
node_dict[item] = obo_id.split('/')[-1]
for item in egcg_list:
if item not in node_dict:
print(item)
obo_id = df.loc[df['object_name'] == item]['object_obo'].values[0]
node_dict[item] = obo_id.split('/')[-1]
for item in tea_list:
if item not in node_dict:
print(item)
obo_id = df.loc[df['object_name'] == item]['object_obo'].values[0]
node_dict[item] = obo_id.split('/')[-1]
len(node_dict)
node_dict['Hyperglycemia']
x = zip(cyp3a4_edge_paths, cyp3a4_edge_path_labs)
for item in x:
print(item)
print(type(item))
break
# +
from typing import Dict, List, Optional, Set, Tuple, Union
def n3(node: Union[URIRef, BNode, Literal]) -> str:
"""Method takes an RDFLib node of type BNode, URIRef, or Literal and serializes it to meet the RDF 1.1 NTriples
format.
Src: https://github.com/RDFLib/rdflib/blob/c11f7b503b50b7c3cdeec0f36261fa09b0615380/rdflib/plugins/serializers/nt.py
Args:
node: An RDFLib
Returns:
serialized_node: A string containing the serialized
"""
if isinstance(node, Literal): serialized_node = "%s" % _quoteLiteral(node)
else: serialized_node = "%s" % node.n3()
return serialized_node
# -
s = URIRef('http://napdi.org/napdi_srs_imports:camellia_sinensis_leaf')
p = URIRef('http://purl.obolibrary.org/obo/RO_0002180')
o = URIRef('http://purl.obolibrary.org/obo/CHEBI_90')
pred_key = hashlib.md5('{}{}{}'.format(n3(s), n3(p), n3(o)).encode()).hexdigest()
pred_key
nx_graph[s][o][p]
# ### Fix nodeLabels in file from build 3.0.0
import json
import pickle
import re
len(nodeLabels)
file1 = open(KG_PATH + 'nodeLabels_20211014.pickle', 'rb')
nods = pickle.load(file1)
nods['http://purl.obolibrary.org/obo/PR_Q9H9S0']
nods['http://purl.obolibrary.org/obo/PR_Q9H9S0']
nods['http://purl.obolibrary.org/obo/RO_0000057'] = 'has participant'
fileo = open(KG_PATH + 'nodeLabels_20211021.pickle', 'wb')
for key in correctLabels:
print(key)
print(correctLabels[key])
break
for key in correctLabels:
node = key.strip('<')
node = node.strip('>')
newLabel = correctLabels[key]['label']
if node in nods:
if newLabel != 'N/A':
nods[node] = newLabel
len(nods)
nods['http://purl.obolibrary.org/obo/SO_0000704']
nods['http://purl.obolibrary.org/obo/PR_Q9H9S0']
nas[:20]
nas = []
for key in correctLabels:
label = correctLabels[key]['label']
if label == 'N/A':
nas.append(key)
len(nas)
count = 1
for item in nas:
if 'PR' in item:
count += 1
count
pickle.dump(nods, fileo)
# ## Reweighting the KG
# 1. Fix subclassof chemical entity
# 2. maybe downweight subclassof
# 3. use belief scores of MR to weight?
# 4. centrality measures - node degree centrality
# 5. Fix mapping to TEA in REACH/SemRep (maps to triethylamine CHEBI_35026)
# +
#INDRA pathfinding module also searches in nx multidigraph
#https://indra.readthedocs.io/en/latest/modules/explanation/pathfinding.html
#uses belief in metadata (I think)
| PheKnowLator_notebooks/notebooks/Path_Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''venv'': venv)'
# name: python3
# ---
# # Introduction to Regression with Neural Networks with TensorFlow
# + id="A2m9e2WZC40B"
import tensorflow as tf
print(tf.__version__)
# -
# ## Creating data to view and fit
# +
import numpy as np
import matplotlib.pyplot as plt
#Create features
X = np.array([-7.0,-4.0,-1.0,2.0,5.0,8.0,11.0,14.0])
#Create labels
y = np.array([3.0,6.0,9.0,12.0,15.0,18.0,21.0,24.0])
#Visualize it
plt.scatter(X,y)
# + pycharm={"name": "#%%\n"}
y == X+10
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Input and output shapes
# + pycharm={"name": "#%%\n"}
#Create a demo tensor for our housing price prediction problem
house_info = tf.constant(['bedroom','bathroom','garage'])
house_price = tf.constant([939700])
house_info,house_price
# -
# Turn our numpy arrays into tensors
X = tf.constant(X)
y = tf.constant(y)
X,y
input_shape = X[0].shape
output_shape = y[0].shape
input_shape,output_shape
# ## Steps in modellig with TF
#
# 1. Create a model - define input and output layers and hidden layers
# 2. Compile model - loss func, optimizer and evaluation metrics
# 3. Fitting a model - Let the model try to identify the patterns
# +
# Set random seed
tf.random.set_seed(42)
# 1. Create a model using the Sequential API
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# 2. Compile model
model.compile(loss= tf.keras.losses.mae #Mean absolute error
,optimizer = tf.keras.optimizers.SGD() # Stochastic gradient descent
, metrics=['mae']
)
# 3. Fit the model
model.fit(X,y,epochs=5)
# -
# Check out X and y
X,y
# +
# Try to predict with the trained model
y_pred = model.predict([17.0]) #Expected 27
y_pred
# -
# ## Improving our model
#
# We can improve our model by altering the steps we took to create a model
#
# 1. Creating a model - add more layers, increase number of neurons, change activation functions
# 2. Compiling a model - change the optimization functions or the learning rate of the optimization function
# 3. Fitting a model - more epochs or give more data
# +
# Rebuilding our model
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss = tf.keras.losses.mae,optimizer = tf.optimizers.SGD(), metrics=['MAE'])
#3. Fit the model
model.fit(X,y,epochs= 100)
# -
# Reminding the data
X,y
# +
#Check if the prediction has improved
model.predict([17.0])# Expected 27
# +
# Rebuilding our model with further changes
# 1. Create the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(100,activation='relu'),
tf.keras.layers.Dense(1),
])
# 2. Compile the model
model.compile(loss = tf.keras.losses.mae,optimizer = tf.optimizers.SGD(), metrics=['MAE'])
#3. Fit the model
model.fit(X,y,epochs= 100)
# +
# Testing a new prediction
model.predict([17.0]) # Expected 27
#Signs of overfitting --> Better loss but worse performance with outside data
# -
# ## Evaluating a model
#
# Typical workflow when working with neural networks
#
# Build -> fit -> evaluate -> tweak -> fit -> evaluate -> tweak ->...
#
# When evaluating remeber:
# > Visualize,Visualize,Visualize
# * The data
# * The model
# * The training
# * The predictions
# +
# Make a bigger dataset
X = tf.range(-100,100,4)
X
# +
#Make labels for the dataset
y= X+10
y
# -
# Visualize the data
plt.scatter(X,y)
# ### The 3 sets...
#
# * Training (70-80%)
# * Validate - used for tunning/tweaking (10-15%)
# * Test - Evaluation/final testing (10-15%)
# +
# Check the lenght of the samples
len(X)
# +
#Because of the diminished value of samples validation set is going to be ignored for now
#Split the data into train and test
X_train = X[:40]#First 40 (80%)
X_test = X[40:]# Last 10 (20%)
y_train = y[:40]
y_test = y[40:]
len(X_test),len(y_test),len(X_train),len(y_train)
# -
# ## Visualizing the data after split
plt.figure(figsize=(10,7))
#Plot training in blue
plt.scatter(X_train,y_train,c='b',label = 'Training data')
plt.scatter(X_test,y_test,c='g',label = 'Testing data')
plt.legend()
# +
#Building a neural network for our data
model = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
model.compile(loss = tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=['mae'])
#model.fit(X_train,y_train,epochs=100)
# -
X[0].shape
# +
# Creating a model that builds automatically by defining the input_shape on the first layer
tf.random.set_seed(42)
model = tf.keras.Sequential([
tf.keras.layers.Dense(1,input_shape = [1])
])
model.compile(loss = tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=['mae'])
#model.fit(X_train,y_train,epochs=100)
# -
model.summary()
#Every time we run fit without reinstatiate the model it adds epochs to the training
#Watch out with this to not cause overfitting
model.fit(X_train,y_train,epochs=100,verbose = 0)
# * Total params - number of parameters in the model
# * Trainable params - parameters that the model can update as it trains
# * Non-trainable params - frozen parameters trained previously (imported models generally)
# +
from keras.utils.vis_utils import plot_model
plot_model(model)
# -
# ## Visualizing the model predictions
#
# Plot the predictions against the real values
# > Model copied from video notebook
# +
# Let's see if we can make another to improve our model
# 1. Create the model (this time with an extra hidden layer with 100 hidden units)
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation=None),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model.compile(loss="mae",
optimizer=tf.keras.optimizers.Adam(learning_rate=0.01),
metrics=["mae"])
# 3. Fit the model
model.fit(X, y, epochs=100)
# -
# Make some predictions
y_pred = model.predict(X_test)
y_pred
y_test
# Creating plot function
def plot_predictions(train_data=X_train,
train_labels=y_train,
test_data=X_test,
test_labels=y_test,
predictions=y_pred):
"""
Plots training data, test data and compares predictions to ground truth labels.
"""
plt.figure(figsize=(10, 7))
# Plot training data in blue
plt.scatter(train_data, train_labels, c="b", label="Training data")
# Plot testing data in green
plt.scatter(test_data, test_labels, c="g", label="Testing data")
# Plot model's predictions in red
plt.scatter(test_data, predictions, c="r", label="Predictions")
# Show the legend
plt.legend();
plot_predictions(X_train,y_train,X_test,y_test,y_pred)
# ### Evaluation metrics
#
# Depending on the problem there will be different metrics
#
# - Regression -> MAE and MSE (mainly)
# - MSE punishes more the bigger errors
#
#model.evaluate returns loss and metrics set while compiling the model
model.evaluate(X_test,y_test)
# +
#These return a list, instead of a single value
MAE = tf.metrics.mean_absolute_error(y_test,y_pred)
MSE = tf.metrics.mean_squared_error(y_test,y_pred)
MAE,MSE
# -
# Comparing different tensor must have same shape
#y_test -> (10,)
#y_pred -> (10,1)
y_test,tf.constant(y_pred)
#Adjusting y_pred
y_pred = tf.squeeze(tf.constant(y_pred))
y_pred
# +
# Now the errors are calculated correctly
MAE = tf.metrics.mean_absolute_error(y_test,y_pred)
MSE = tf.metrics.mean_squared_error(y_test,y_pred)
MAE,MSE
# +
# Make some functions to reuse MAE and MSE
def mae(y_true, y_pred):
return tf.metrics.mean_absolute_error(y_true=y_true,
y_pred=tf.squeeze(y_pred))
def mse(y_true, y_pred):
return tf.metrics.mean_squared_error(y_true=y_true,
y_pred=tf.squeeze(y_pred))
# -
# ### Running experiments to improve the model
#
# 3 Experiments
# 1. `model_1` - same model, 1 layer, trained for longer (100 epochs)
# 2. `model_2` - 2 layers, trained for longer (100 epochs)
# 3. `model_3` - 2 layers, 500 epochs
X_train,y_train
# + colab={"base_uri": "https://localhost:8080/"} id="0cLyuLdP-SIw" outputId="f6654942-3d40-4dd9-a904-38c13afadfc4"
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_1 = tf.keras.Sequential([
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model_1.compile(loss=tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=["mae"])
# 3. Fit the model
model_1.fit(X_train, y_train, epochs=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 465} id="9Wfj4dqK-z6H" outputId="08cde110-5b47-4b66-a693-36cf7014656a"
# Make and plot predictions for model_1
y_preds_1 = model_1.predict(X_test)
plot_predictions(predictions=y_preds_1)
# +
#Calculate model_1 evaluation metrics
mae_1 = mae(y_test,y_preds_1)
mse_1 = mse(y_test,y_preds_1)
mae_1,mse_1
# -
# ### Building model 2
# +
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_2 = tf.keras.Sequential([
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model_2.compile(loss=tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=["mse"])
# 3. Fit the model
model_2.fit(X_train, y_train, epochs=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 465} id="9Wfj4dqK-z6H" outputId="08cde110-5b47-4b66-a693-36cf7014656a"
# Make and plot predictions for model_1
y_preds_2 = model_2.predict(X_test)
plot_predictions(predictions=y_preds_2)
# +
#Calculate model_1 evaluation metrics
mae_2 = mae(y_test,y_preds_2)
mse_2 = mse(y_test,y_preds_2)
mae_2,mse_2
# -
# ### Build model 3
# +
# Set random seed
tf.random.set_seed(42)
# 1. Create the model
model_3 = tf.keras.Sequential([
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
model_3.compile(loss=tf.keras.losses.mae,
optimizer=tf.keras.optimizers.SGD(),
metrics=["mse"])
# 3. Fit the model
model_3.fit(X_train, y_train, epochs=500)
# + colab={"base_uri": "https://localhost:8080/", "height": 465} id="9Wfj4dqK-z6H" outputId="08cde110-5b47-4b66-a693-36cf7014656a"
# Make and plot predictions for model_1
y_preds_3 = model_3.predict(X_test)
plot_predictions(predictions=y_preds_3)
# +
#Calculate model_1 evaluation metrics
mae_3 = mae(y_test,y_preds_3)
mse_3 = mse(y_test,y_preds_3)
mae_3,mse_3
# -
# ## Comparing the results of our experiments
# +
# Let's compare our model's results using a pandas Dataframe
import pandas as pd
model_results = [['model_1',mae_1.numpy(),mse_1.numpy()],
['model_2',mae_2.numpy(),mse_2.numpy()],
['model_3',mae_3.numpy(),mse_3.numpy()]]
all_results = pd.DataFrame(model_results, columns = ['model','mae','mse'])
all_results
# -
model_2.summary()
# Looks like `model_2` performs best
# ## Tracking your experiments
#
# Can be tedious when running lots of experiments
#
# Tools to help ease the task:
#
# * TensorBoard - Helps tracking ML experiments
# * Weights and Biases - Tool for tracking all kinds of ML experiments (integrates into TensorBoard)
# ## Saving our models
#
# There are two main formats to save models:
#
# 1. SavedModel
# 2. HDF5
# Saving in SavedModel format
model_2.save('.\\saved_models\\01_saved_bestmodel')
# Saving in HDF5 format
model_2.save('.\\saved_models\\01_saved_bestmodel_HDF5.h5')
# ## Loading a saved model
# Loading models to check if they saved correctly
#SavedModel format
loaded_SavedModel_Format = tf.keras.models.load_model('.\\saved_models\\01_saved_bestmodel')
loaded_SavedModel_Format.summary()
# Compare model_2 predictions with the loaded model predictions
model_2_preds = model_2.predict(X_test)
loaded_SavedModel_format_preds = loaded_SavedModel_Format.predict(X_test)
model_2_preds == loaded_SavedModel_format_preds
# +
# Load in the .h5 model
loaded_h5_model = tf.keras.models.load_model('.\\saved_models\\01_saved_bestmodel_HDF5.h5')
loaded_h5_model.summary()
# -
# Compare model_2 predictions with the loaded model predictions
model_2_preds = model_2.predict(X_test)
loaded_h5_format_preds = loaded_h5_model.predict(X_test)
model_2_preds == loaded_h5_format_preds
# # A larger example
# +
# Read in the insurance dataset
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# -
# One-hot encoding our dataframe
insurance_one_hot = pd.get_dummies(insurance,drop_first=True)
insurance_one_hot
# +
# Create X and y values
X = insurance_one_hot.drop('charges',axis = 1)
y = insurance_one_hot['charges']
X.head()
# -
y.head()
# +
# Creating training and test sets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train,y_test = train_test_split(X,y,train_size=0.8,random_state=42)
len(X),len(X_train),len(X_test)
# +
# Build a neural network
tf.random.set_seed(42)
insurance_model = tf.keras.Sequential([
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)])
insurance_model.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.SGD(),
metrics=['mae'])
insurance_model.fit(X_train,y_train,epochs = 100)
# +
# Check the results of the model on the test data
insurance_model.evaluate(X_test,y_test)
# -
# The model results are not great, let's improve it
#
# First try -> add an extra layer with more hidden units and use Adam optimizer
# Second try -> train for longer
# +
# Build a neural network
tf.random.set_seed(42)
insurance_model_2 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)])
insurance_model_2.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(),
metrics=['mae'])
insurance_model_2.fit(X_train,y_train,epochs = 100)
# +
# Check the results of the larger model on the test data
insurance_model_2.evaluate(X_test,y_test)
# +
# Build a neural network
tf.random.set_seed(42)
insurance_model_3 = tf.keras.Sequential([
tf.keras.layers.Dense(100),
tf.keras.layers.Dense(10),
tf.keras.layers.Dense(1)])
insurance_model_3.compile(loss = tf.keras.losses.mae,
optimizer = tf.keras.optimizers.Adam(),
metrics=['mae'])
history = insurance_model_3.fit(X_train,y_train,epochs = 200)
# +
# Check the results of the larger model on the test data
insurance_model_3.evaluate(X_test,y_test)
# +
# Plot history, also known as loss or training curve
pd.DataFrame(history.history).plot()
plt.ylabel('loss')
plt.xlabel('epochs')
# -
# ## Preprocessing data (Normalization and Standardization)
#
# Neural networks tend to prefer normalization
# - Possible to test both and compare
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="A9EZSNF_3MaY" outputId="fb545d41-a657-4d70-a546-04b0250efe62"
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
# Read in the insurance dataframe
insurance = pd.read_csv("https://raw.githubusercontent.com/stedy/Machine-Learning-with-R-datasets/master/insurance.csv")
insurance
# +
from sklearn.compose import make_column_transformer
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.model_selection import train_test_split
# Create a column transformer
ct = make_column_transformer((MinMaxScaler(),['age','bmi','children']),
(OneHotEncoder(handle_unknown='ignore'),['sex','smoker','region']))
# Create X and y
X = insurance.drop('charges',axis = 1)
y = insurance['charges']
# Create train and test sets
X_train, X_test, y_train,y_test = train_test_split(X,y,test_size=0.2,random_state=42)
# Fit the column transformer to our training data
ct.fit(X_train)
# Transform training and test data with scaler anf OHencoder
X_train_normal = ct.transform(X_train)
X_test_normal = ct.transform(X_test)
# -
# Visualizing new data
X_train.loc[0],X_train_normal[0]
X_train.shape, X_train_normal.shape #More columns added by OneHotEncoder
# Test the new normalized data on a new model
# + colab={"base_uri": "https://localhost:8080/"} id="KDF9TU0_6W2r" outputId="7fd4fee5-425e-43b5-d192-787a119983eb"
# Build a neural network model to fit on our normalized data
tf.random.set_seed(42)
# 1. Create the model
insurance_model_4 = tf.keras.Sequential([
tf.keras.layers.Dense(50),
tf.keras.layers.Dense(50),
tf.keras.layers.Dense(50),
tf.keras.layers.Dense(1)
])
# 2. Compile the model
insurance_model_4.compile(loss=tf.keras.losses.mae,
optimizer=tf.keras.optimizers.Adam(),
metrics=["mae"])
# 3. Fit the model
insurance_model_4.fit(X_train_normal, y_train, epochs=100)
# + colab={"base_uri": "https://localhost:8080/"} id="wt84g6C790Y0" outputId="033111e9-a711-43de-c12d-8b816c66ca38"
# Evalaute our insurance model trained on normalized data
insurance_model_4.evaluate(X_test_normal, y_test)
| 01_neural_network_regression_in_tensorflow_edit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import Libraries
#Load Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# # Load Dataset into drugset
#Load Data From Drug Dataset
drugset = pd.read_csv('./drugdataset.csv')
drugset.head()
# # Finding the Key Statistics of the Dataset
#Key Statistics (Numerical Columns only)
drugset.describe()
# # Finding the different classes inside the dataset
#Identify number of Classes (i.e. Drugs)
drugset.Drug.unique()
# Count of Males and Females where 0 represents males and 1 represents female
drugset.groupby(['Sex']).size().reset_index(name='count')
# Count of Blood Pressure Level where 0 represents normal BP, 1 for low BP and 2 for High BP
drugset.groupby(['BP']).size().reset_index(name='count')
# Count of Cholesterol Level 0 represents normal cholesterol, 1 for high cholestrol
drugset.groupby(['Cholesterol']).size().reset_index(name='count')
# Box-Plot
drugset.boxplot(column ='Na_to_K')
# # Creating x and y variables, testing and scaling the data
# +
#Create x and y variables
X = drugset.drop('Drug',axis=1).to_numpy()
y = drugset['Drug'].to_numpy()
#Create Train and Test datasets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y,test_size = 0.2,random_state=100)
#Scale the data
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
x_train2 = sc.fit_transform(X_train)
x_test2 = sc.transform(X_test)
# -
# # Scripting for neural network to get the evaluation report and also finding confusion matrix
# +
#Script for Neural Network
from sklearn.neural_network import MLPClassifier
mlp = MLPClassifier(hidden_layer_sizes=(5,4,5),
activation='relu',solver='adam',
max_iter=10000,random_state=100)
mlp.fit(x_train2, y_train)
predictions = mlp.predict(x_test2)
#Evaluation Report and Matrix
from sklearn.metrics import classification_report, confusion_matrix
target_names=['drugY', 'drugC', 'drugX', 'drugA', 'drugB']
print(confusion_matrix(y_test,predictions))
print(classification_report(y_test,predictions,target_names=target_names))
# +
#Script for Decision Tree
from sklearn.tree import DecisionTreeClassifier
for name,method in [('DT', DecisionTreeClassifier(random_state=100))]:
method.fit(x_train2,y_train)
predict = method.predict(x_test2)
target_names=['drugY', 'drugC', 'drugX', 'drugA', 'drugB']
print('\nEstimator: {}'.format(name))
print(confusion_matrix(y_test,predict))
print(classification_report(y_test,predict,target_names=target_names))
| Assignment 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Requires a neo4j databases to be running at `localhost:7687`
# Tell socialgene where it can talk to the neo4j database
import os
os.environ["NEO4J_URI"] = "bolt://localhost:7687"
# Import required and helper modules
# +
from rich import inspect
from rich.progress import Progress
import pandas as pd
import logging
from socialgene.classes.findmybgc import FindMyBGC
from socialgene.base.socialgene import SocialGene
logging.getLogger("neo4j").setLevel(logging.WARNING)
logging.getLogger().setLevel(logging.INFO)
# -
# Setting paths here so analysis can be easily repeated with other data
genbank_file="/home/chase/Documents/data/wmmb235/john/NZ_MDRX01000001.region001_B235_keyicin/NZ_MDRX01000001.region001.gbk"
hmm_file="/home/chase/Documents/socialgene_data/micromonospora_commit_6efde564f307c5582cee1b8d7e15559dd71a7ab9/long_cache/HMM_HASH/socialgene_nr_hmms_file_1_of_1.hmm"
# Create a new FindMyBGC() object
findmybgc_object = FindMyBGC()
# Read the genbank file for keycin
findmybgc_object.parse(genbank_file)
# Run hmmsearch (with pyHMMER) to annotate the proteins with domain info
findmybgc_object.annotate_with_pyhmmer(
hmm_filepath=hmm_file,
use_neo4j_precalc=False,
cpus=0,
)
# Search NEO4J for similar proteins and their location info
findmybgc_object.query_neo4j()
#
# +
df = pd.DataFrame(
list(findmybgc_object.neo4j_results.result[0]["result_list"]),
columns=["assembly", "locus", "query_protein", "subject_protein", "start", "end", "strand"],
)
# -
# Taking a look at the number of input proteins per result assembly
#
# The top result is GCF_001855515.1 which is WMMB235
# Number of input proteins matched by result assembly
df.groupby("assembly")["query_protein"].nunique().sort_values(ascending=False)
# Same thing but splitting out by locus (contig/scaffold)
# Number of input proteins matched by result assembly and locus
df.groupby(["assembly", "locus"])["query_protein"].nunique().sort_values(ascending=False)
# Same thing as above (number of input proteins per result locus) but looking at one of results towards the bottom
z=df.groupby(["assembly", "locus"])["query_protein"].nunique().sort_values(ascending=False).to_frame()
z=z.reset_index()
z.loc[z['assembly'] == 'GCF_003667455.1']
z=df.groupby("assembly")["query_protein"].nunique().sort_values(ascending=False).to_frame()
z=z.reset_index()
z.loc[z['assembly'] == 'GCF_003667455.1']
| old/2020_01_21_keyicin.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8f01aa40-5e13-4792-b407-6e5e5fb34d2c", "showTitle": false, "title": ""}
# Creating Databases
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3b06face-e061-4253-a5a9-f85d4dc3c00b", "showTitle": false, "title": ""}
# %sql
create database Praveen
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2c8bef4e-ebc2-42df-9628-811b4afa57bd", "showTitle": false, "title": ""}
# %python
spark.sql('Create database Praveenpy')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4324f22c-bb35-4d44-9ce4-09cf82bb4cb6", "showTitle": false, "title": ""}
# %scala
spark.sql("create database Praveenscala")
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f4e79aaa-6397-47bd-a642-86ff3f58e08f", "showTitle": false, "title": ""}
# Describing Databases
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a3b6d76d-9b87-4cd1-859b-a48e6e437b83", "showTitle": false, "title": ""}
# %sql
describe database Praveen
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f58c795c-9619-4c41-82d6-e56e76ba1f6e", "showTitle": false, "title": ""}
# %python
spark.sql('describe database Praveenpy').show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2c8a84a2-f1da-4a7d-90a8-22856a3880d6", "showTitle": false, "title": ""}
# %scala
spark.sql("describe database Praveenscala").show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4a1fae6b-ea41-4ab0-95c1-b999b2f4ca04", "showTitle": false, "title": ""}
# Drop databases
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "58b2dc12-ee5b-4895-81f3-091fbb50fe67", "showTitle": false, "title": ""}
# %sql
drop database Praveen
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2078acbd-da37-4c0c-8f7f-5d5651bad2af", "showTitle": false, "title": ""}
# %python
spark.sql("Drop database praveenpy")
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7b9b4937-878b-4e89-a153-626e91a694c3", "showTitle": false, "title": ""}
# %scala
spark.sql("drop database PraveenScala")
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "10e656d9-5a0b-4a1a-b49d-946f53c40c3f", "showTitle": false, "title": ""}
# drop if tables exists use CASCADE
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fcdbec21-f3dc-4bbe-a2c2-934ad2bbc01c", "showTitle": false, "title": ""}
loc = '/FileStore/tables/data.csv'
df = spark.read.csv(loc,header=True)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e8847054-6ed1-4432-8b33-1f7e90d66c28", "showTitle": false, "title": ""}
df.write.format('parquet').saveAsTable('Praveen.mytable')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "83ff3686-b530-4f7b-b1c7-cb4ecb24f8fb", "showTitle": false, "title": ""}
# %sql
drop database Praveen cascade
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b6301a14-8075-454e-a02a-2db7c99823b6", "showTitle": false, "title": ""}
# Creating table in notebook
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "aa377780-be90-4e21-8db4-5b6e91c986e9", "showTitle": false, "title": ""}
# %sql
create table Praveenpy.first(cid int,name string,numb int)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "88718531-ce70-4ad7-99b0-9ceb4a44798c", "showTitle": false, "title": ""}
df = spark.createDataFrame([(1,'one',1),(2,'two',2)],['cid','name','numb'])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "299704bd-614a-4e2b-aaa3-7030cd5c117b", "showTitle": false, "title": ""}
df.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e4bb5973-4b47-4d56-8b1a-68e42c979513", "showTitle": false, "title": ""}
from pyspark.sql.functions import col
df = df.withColumn('cid',col('cid').cast('int'))
df = df.withColumn('numb',col('numb').cast('int'))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d60e51f7-641a-4d52-a95c-4511efeae25d", "showTitle": false, "title": ""}
df.write.mode('overwrite').saveAsTable('Praveenpy.first')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "58f1d19b-6171-4a5b-9aea-43b6099e2c64", "showTitle": false, "title": ""}
df.write.mode('overwrite').partitionBy('cid').saveAsTable('Praveenpy.two')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ff2718a6-c6f5-4c01-bee7-8c4b5390462b", "showTitle": false, "title": ""}
# %sql
select * from praveenpy.two
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "568a8f02-d2c3-4bb8-b8cf-33fb6e4601d4", "showTitle": false, "title": ""}
| Data Extraction/2. Creating databases in databricks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import math
import matplotlib.pyplot as plt
import numpy as np
from astropy import units as u
from mpl_toolkits.mplot3d import Axes3D
from plasmapy import formulary, particles
# -
# # Physics of the ExB drift
# Consider a single particle of mass $m$ and charge $q$ in a constant, uniform magnetic field $\mathbf{B}=B\ \hat{\mathbf{z}}$. In the absence of external forces, it travels with velocity $\mathbf{v}$ governed by the equation of motion
#
# $$m\frac{d\mathbf{v}}{dt} = q\mathbf{v}\times\mathbf{B}$$
#
# which simply equates the net force on the particle to the corresponding Lorentz force. Assuming the particle initially (at time $t=0$) has $\mathbf{v}$ in the $x,z$ plane (with $v_y=0$), solving reveals
#
# $$v_x = v_\perp\cos\omega_c t \quad\mathrm{;}\quad v_y = -\frac{q}{\lvert q \rvert}v_\perp\sin \omega_c t$$
#
# while the parallel velocity $v_z$ is constant. This indicates that the particle gyrates in a circular orbit in the $x,y$ plane with constant speed $v_\perp$, angular frequency $\omega_c = \frac{\lvert q\rvert B}{m}$, and Larmor radius $r_L=\frac{v_\perp}{\omega_c}$.
#
# As an example, take one proton `p+` moving with velocity $1\ m/s$ in the $x$-direction at $t=0$:
# Setup proton in uniform B field
B = 5 * u.T
proton = particles.Particle("p+")
omega_c = formulary.frequencies.gyrofrequency(B, proton)
v_perp = 1 * u.m / u.s
r_L = formulary.lengths.gyroradius(B, proton, Vperp=v_perp)
# We can define a function that evolves the particle's position according to the relations above describing $v_x,v_y$, and $v_z$. The option to add a constant drift velocity $v_d$ to the solution is included as an argument, though this drift velocity is zero by default:
def single_particle_trajectory(v_d=np.array([0, 0, 0])):
# Set time resolution & velocity such that proton goes 1 meter along B per rotation
T = 2 * math.pi / omega_c.value # rotation period
v_parallel = 1 / T * u.m / u.s
dt = T / 1e2 * u.s
# Set initial particle position
x = []
y = []
xt = 0 * u.m
yt = -r_L
# Evolve motion
timesteps = np.arange(0, 10 * T, dt.value)
for t in list(timesteps):
v_x = v_perp * math.cos(omega_c.value * t) + v_d[0]
v_y = v_perp * math.sin(omega_c.value * t) + v_d[1]
xt += +v_x * dt
yt += +v_y * dt
x.append(xt.value)
y.append(yt.value)
x = np.array(x)
y = np.array(y)
z = v_parallel.value * timesteps
return x, y, z
# Executing with the default argument and plotting the particle trajectory gives the expected helical motion, with a radius equal to the Larmor radius:
x, y, z = single_particle_trajectory()
fig = plt.figure(figsize=(6, 6))
ax = fig.add_subplot(111, projection="3d")
ax.plot(x, y, z, label="$\mathbf{F}=0$")
ax.legend()
bound = 3 * r_L.value
ax.set_xlim([-bound, bound])
ax.set_ylim([-bound, bound])
ax.set_zlim([0, 10])
ax.set_xlabel("x [m]")
ax.set_ylabel("y [m]")
ax.set_zlabel("z [m]")
plt.show()
print(f"r_L = {r_L.value:.2e} m")
print(f"omega_c = {omega_c.value:.2e} rads/s")
# How does this motion change when a constant external force $\mathbf{F}$ is added? The new equation of motion is
#
# $$m\frac{d\mathbf{v}}{dt} = q\mathbf{v}\times\mathbf{B} + \mathbf{F}$$
#
# and we can find a solution by considering velocities of the form $\mathbf{v}=\mathbf{v}_\parallel + \mathbf{v}_L + \mathbf{v}_d$. Here, $\mathbf{v}_\parallel$ is the velocity parallel to the magnetic field, $\mathbf{v}_L$ is the Larmor gyration velocity in the absence of $\mathbf{F}$ found previously, and $\mathbf{v}_d$ is some constant drift velocity perpendicular to the magnetic field. Then, we find that
#
# $$F_\parallel = m\frac{dv_\parallel}{dt} \quad\mathrm{and}\quad \mathbf{F}_\perp = q\mathbf{B}\times \mathbf{v}_d$$
#
# and applying the vector triple product yields
#
#
# $$\mathbf{v}_d = \frac{1}{q}\frac{\mathbf{F}_\perp\times\mathbf{B}}{B^2}$$
#
# In the case where the external force $\mathbf{F} = q\mathbf{E}$ is due to a constant electric field, this is the constant $\mathbf{E}\times\mathbf{B}$ drift velocity:
#
# $$\boxed{
# \mathbf{v}_d = \frac{\mathbf{E}\times\mathbf{B}}{B^2}
# }$$
#
# Built in drift functions allow you to account for the new force added to the system in two different ways:
# +
E = 0.2 * u.V / u.m # E-field magnitude
ey = np.array([0, 1, 0])
ez = np.array([0, 0, 1])
F = proton.charge * E # force due to E-field
v_d = formulary.drifts.force_drift(F * ey, B * ez, proton.charge)
print("F drift velocity: ", v_d)
v_d = formulary.drifts.ExB_drift(E * ey, B * ez)
print("ExB drift velocity: ", v_d)
# -
# The resulting particle trajectory can be compared to the case without drifts by calling our previously defined function with the drift velocity now as an argument. As expected, there is a constant drift in the direction of $\mathbf{E}\times\mathbf{B}$:
x_d, y_d, z_d = single_particle_trajectory(v_d=v_d)
# + nbsphinx-thumbnail={"output-index": 0}
fig = plt.figure(figsize=(6, 6))
ax = fig.add_subplot(111, projection="3d")
ax.plot(x, y, z, label="$\mathbf{F}=0$")
ax.plot(x_d, y_d, z_d, label="$\mathbf{F}=q\mathbf{E}$")
bound = 3 * r_L.value
ax.set_xlim([-bound, bound])
ax.set_ylim([-bound, bound])
ax.set_zlim([0, 10])
ax.set_xlabel("x [m]")
ax.set_ylabel("y [m]")
ax.set_zlabel("z [m]")
ax.legend()
plt.show()
# -
print(f"r_L = {r_L.value:.2e} m")
print(f"omega_c = {omega_c.value:.2e} rads/s")
# Of course, the implementation in our `single_particle_trajectory()` function requires the analytical solution for the velocity $\mathbf{v}_d$. This solution can be compared with that implemented in the [particle stepper notebook](simulation/particle_stepper.ipynb). It uses the Boris algorithm to evolve the particle along its trajectory in prescribed $\mathbf{E}$ and $\mathbf{B}$ fields, and thus does not require the analytical solution.
| docs/notebooks/ExB_drift.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="hddpB7Iu8ynD"
# # Implementation of Resnet
# > In this notebook I have implemented ResNet from scratch using Pytorch
# + id="pYlM3FI-8mZK"
#importing libraries
import torch
import torch.nn as nn
import torch.nn.functional as F
# + id="XwerA9Bw1BZb"
# an essential block of layers which forms resnets
class ResBlock(nn.Module):
#in_channels -> input channels,int_channels->intermediate channels
def __init__(self,in_channels,int_channels,identity_downsample=None,stride=1):
super(ResBlock,self).__init__()
self.expansion = 4
self.conv1 = nn.Conv2d(in_channels,int_channels,kernel_size=1,stride=1,padding=0)
self.bn1 = nn.BatchNorm2d(int_channels)
self.conv2 = nn.Conv2d(int_channels,int_channels,kernel_size=3,stride=stride,padding=1)
self.bn2 = nn.BatchNorm2d(int_channels)
self.conv3 = nn.Conv2d(int_channels,int_channels*self.expansion,kernel_size=1,stride=1,padding=0)
self.bn3 = nn.BatchNorm2d(int_channels*self.expansion)
self.relu = nn.ReLU()
self.identity_downsample = identity_downsample
self.stride = stride
def forward(self,x):
identity = x.clone()
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.conv2(x)
x = self.bn2(x)
x = self.relu(x)
x = self.conv3(x)
x = self.bn3(x)
#the so called skip connections
if self.identity_downsample is not None:
identity = self.identity_downsample(identity)
x += identity
x = self.relu(x)
return x
class ResNet(nn.Module):
def __init__(self,block,layers,image_channels,num_classes):
super(ResNet,self).__init__()
self.in_channels = 64
self.conv1 = nn.Conv2d(image_channels,64,kernel_size=7,stride=2,padding=3)
self.bn1 = nn.BatchNorm2d(64)
self.relu = nn.ReLU()
self.maxpool = nn.MaxPool2d(kernel_size=3,stride=2,padding=1)
#the resnet layers
self.layer1 = self._make_layer(block,layers[0],int_channels=64,stride=1)
self.layer2 = self._make_layer(block,layers[1],int_channels=128,stride=2)
self.layer3 = self._make_layer(block,layers[2],int_channels=256,stride=2)
self.layer4 = self._make_layer(block,layers[3],int_channels=512,stride=2)
self.avgpool = nn.AdaptiveAvgPool2d((1,1))
self.fc1 = nn.Linear(512*4,num_classes)
def forward(self,x):
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = self.avgpool(x)
x = x.reshape(x.shape[0],-1)
x = self.fc1(x)
return x
def _make_layer(self,block,num_res_blocks,int_channels,stride):
identity_downsample = None
layers = []
if stride!=1 or self.in_channels != int_channels*4:
identity_downsample = nn.Sequential(nn.Conv2d(self.in_channels,int_channels*4,
kernel_size=1,stride=stride),
nn.BatchNorm2d(int_channels*4))
layers.append(ResBlock(self.in_channels,int_channels,identity_downsample,stride))
#this expansion size will always be 4 for all the types of ResNets
self.in_channels = int_channels*4
for i in range(num_res_blocks-1):
layers.append(ResBlock(self.in_channels,int_channels))
return nn.Sequential(*layers)
def ResNet18(img_channel=3,num_classes=1000):
return ResNet(ResBlock,[2,2,2,2],img_channel,num_classes)
def ResNet34(img_channel=3,num_classes=1000):
return ResNet(ResBlock,[3,4,6,3],img_channel,num_classes)
def ResNet50(img_channel=3,num_classes=1000):
return ResNet(ResBlock,[3,4,6,3],img_channel,num_classes)
def ResNet101(img_channel=3,num_classes=1000):
return ResNet(ResBlock,[3,4,23,3],img_channel,num_classes)
def ResNet152(img_channel=3,num_classes=1000):
return ResNet(ResBlock,[3,8,36,3],img_channel,num_classes)
# + id="KAyfJzPP0lRS" outputId="2b5c3f62-6c73-4681-8182-42ae7996c847" colab={"base_uri": "https://localhost:8080/"}
def test():
net = ResNet101(img_channel=3,num_classes=1000)
x = torch.randn(4,3,224,224)
y = net(x).to("cuda")
print(y.size())
test()
# + id="TNlDFB7xdQ-G"
| Resnet/.ipynb_checkpoints/Resnet-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Phase transition
#
# This notebook illustrates the phase transition for perfect recovery.
#
# Observations:
#
# - it's the SBM generation that takes the most time, not so much the inference.
# Keeping 10 iterations of cavi is fine.
# - `delta_inf` works best close to $(K - 1) / K$.
# I set it at 0.799
import collabclass
import numpy as np
import matplotlib.pyplot as plt
import pickle
# +
n = 1000
k = 5
s = 5
alpha = 0.5
deltas = np.array([0.03, 0.1, 0.2])
ms = np.linspace(100, 20000, num=10, dtype=int)
n_draws = 100
# +
# %%time
np.random.seed(0)
samples = np.zeros((len(ms), n_draws), dtype=object)
for i, m in enumerate(ms):
for j in range(n_draws):
# Tuple `(us, vs, graph)`.
samples[i, j] = collabclass.sbm(m=m, n=n, k=k, s=s, alpha=alpha)
# -
def experiment(predict):
res = np.zeros((len(deltas), len(ms), 2*n_draws))
for z, delta in enumerate(deltas):
for i, m in enumerate(ms):
print(".", end="", flush=True)
for j in range(n_draws):
us, vs, graph = samples[i, j]
for o in range(2):
vs_hat = collabclass.symmetric_channel(vs, k, delta)
# Run the predictor.
vs_bar = predict(graph, vs_hat)
# Evaluate accuracy.
#fraction_connected = np.count_nonzero(graph.item_idx[:,1] > 0) / n
res[z, i, 2*j + o] = np.count_nonzero(vs_bar == vs) / n
print()
return res
# ### CAVI
# +
# %%time
np.random.seed(0)
delta_inf = 0.799
def cavi(graph, vs_hat):
alpha_prior = alpha * np.ones((len(graph.user_idx), k))
beta_prior = collabclass.init_beta(k, vs_hat, delta_inf)
_, beta = collabclass.cavi(graph, alpha_prior, beta_prior, n_iters=10)
return np.argmax(beta, axis=1)
res = experiment(cavi)
# -
# ### wvRN
# %%time
np.random.seed(0)
res2 = experiment(collabclass.wvrn)
# ### Plot data
# +
fig, ax = plt.subplots(figsize=(12, 7))
for z, delta in enumerate(deltas):
cavi = np.mean(res[z] == 1, axis=1)
wvrn = np.mean(res2[z] == 1, axis=1)
ax.plot(ms, cavi, marker="o", ms=5, color=plt.cm.tab10(z),
label=f"CAVI δ = {delta:.2f}")
ax.plot(ms, wvrn, marker="o", ms=5, color=plt.cm.tab10(z), linestyle=":",
label=f"wvRN δ = {delta:.2f}")
ax.legend()
| notebooks/synthetic-phasetrans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Notebook written by [<NAME>](https://github.com/zhedongzheng)
import tensorflow as tf
import os
from tqdm import tqdm
# +
MAX_LEN = 300
BATCH_SIZE = 32
VOCAB_SIZE = 20000
TF_RECORD_PATH = './imdb_train_fixed300.tfrecord'
if not os.path.isfile(TF_RECORD_PATH):
(X_train, y_train), (_, _) = tf.keras.datasets.imdb.load_data(num_words=VOCAB_SIZE)
X_train = tf.keras.preprocessing.sequence.pad_sequences(X_train,
MAX_LEN,
padding='post',
truncating='post')
# -
if not os.path.isfile(TF_RECORD_PATH):
writer = tf.python_io.TFRecordWriter(TF_RECORD_PATH)
for sent, label in tqdm(zip(X_train, y_train), total=len(X_train), ncols=70):
example = tf.train.Example(
features = tf.train.Features(
feature = {
'sent': tf.train.Feature(
int64_list=tf.train.Int64List(value=sent)),
'label': tf.train.Feature(
int64_list=tf.train.Int64List(value=[label])),
}))
serialized = example.SerializeToString()
writer.write(serialized)
writer.close()
# +
def _parse_fn(example_proto):
parsed_feats = tf.parse_single_example(
example_proto,
features={
'sent': tf.FixedLenFeature([MAX_LEN], tf.int64),
'label': tf.FixedLenFeature([], tf.int64)
})
return parsed_feats['sent'], parsed_feats['label']
dataset = tf.data.TFRecordDataset([TF_RECORD_PATH])
dataset = dataset.map(_parse_fn)
dataset = dataset.batch(BATCH_SIZE)
iterator = dataset.make_one_shot_iterator()
X_batch, y_batch = iterator.get_next()
print(X_batch.get_shape(), y_batch.get_shape())
# -
sess = tf.Session()
x, y = sess.run([X_batch, y_batch])
print(x.shape, y.shape)
| src_tf/data_io/tfrecord_imdb_fixed_len.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="5OMjVdD5k3H9" colab={"base_uri": "https://localhost:8080/"} outputId="29cf05f9-f780-48c5-d299-019b7e7715f3"
import pandas as pd
from sklearn.ensemble import IsolationForest
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
from sklearn.neighbors import LocalOutlierFactor
from sklearn.svm import OneClassSVM
#--------------------------------------------------
def encode(cat):
# integer encode
label_encoder = LabelEncoder()
integer_encoded = label_encoder.fit_transform(cat)
# binary encode
onehot_encoder = OneHotEncoder(sparse=False)
integer_encoded = integer_encoded.reshape(len(integer_encoded), 1)
onehot_encoded = onehot_encoder.fit_transform(integer_encoded)
return onehot_encoded
def remove_outlier(data, technique=0):
if technique == 0:
#contamination is a hyperparameter for the expected percentage of outliers in the dataset
iso = IsolationForest(contamination=0.1)
yhat = iso.fit_predict(data)
# select all rows that are not outliers
mask = yhat != -1
return mask
elif technique == 1:
# identify outliers in the training dataset
lof = LocalOutlierFactor()
yhat = lof.fit_predict(data)
# select all rows that are not outliers
mask = yhat != -1
return mask
elif technique == 2:
ee = OneClassSVM(nu=0.01)
yhat = ee.fit_predict(x)
# select all rows that are not outliers
mask = yhat != -1
return mask
else:
print('Invalid Input')
return
def main():
x = pd.read_csv('features.csv')
x = x.to_numpy()
AOD_tech = 0 # set this variable to 0 for IsolationForest, 1 for LocalOutlierFactor, and 2 for OneClassSVM
# Automatic outlier detection require all the features to be numeric
print('Removing outliers.....')
mask = remove_outlier(x, AOD_tech)
x = x[mask,:]
print('Outlier detection completed!')
return x
if __name__ == '__main__':
clean_data = main()
# + id="plYbRhrpBzpb" colab={"base_uri": "https://localhost:8080/"} outputId="10b04d94-f28a-4b1d-9e9d-18f0c7298d99"
clean_data.shape
| Outlier_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Compare DQN improvements
# - Double DQN
# - Dueling DQN
# - Prioritized experience replay
# +
from pathlib import Path
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
# -
# ## Load scores
experiment_dirs = list(Path('../experiments').glob('*'))
# +
experiments = [
('_'.join(experiment.name.split('_')[2:]), pd.read_csv(experiment / 'scores.csv'))
for experiment in experiment_dirs
if (experiment / 'scores.csv').exists()
if experiment.name.startswith('final_comparison')]
[e[0] for e in experiments]
# -
# ## Comparison graphs
# +
plt.figure(figsize=(10, 10))
plt.xlabel('Episode #')
plt.ylabel('Score')
for ([name, scores], color) in zip(experiments, mcolors.TABLEAU_COLORS):
ylim = (-5, 20)
smoothened_scores = scores['score'].rolling(window=100, min_periods=0).mean()
plt.plot(scores['episode'], smoothened_scores, label=name, color=color)
solution_episodes = scores['episode'][smoothened_scores >= 13.0]
if len(solution_episodes) > 0:
first_solution_episode = solution_episodes.iloc[0]
plt.vlines(first_solution_episode, *ylim, linestyle='--', linewidth=1, color=color)
plt.legend()
plt.savefig('../artifacts/dqn_improvements_comparison.png')
plt.show()
| notebooks/compare_dqn_improvements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Example 5.1 Modeling CSV data with Multilayer Perceptron Networks
# The first example from the Deep Learning book is modeling CSV data with a multilayer perceptron network (Patterson and Gibson. 175). This is entended to be a gentile introduction to the DL4J API using a simple model. My plan was to implement this exact model in TensorFlow using modern toolsets like Pandas, for loading data and Keras for creating, training and testing the model. I thought this would be the simpilest model to translate into TensorFlow and Keras, but I was wrong.
#
# The largest stumbling block in this transformation was the use of the Negative Log-Likelihood as the loss function. The log-likelihood is a function that is used in traditional pattern recognition to estimate parmaeters.
#
# The likelihood is the product of all of the data given the model parameters; e.g.,
#
# $$L = \prod_{k=1}^{N} p(x_k | \Theta) $$
#
# Applying the negative log to the likehood, we get
#
# $$NLL = \sum_{k=1}^{N} -\ln p(x_k | \Theta) $$
#
# where, $$p(x_k | \Theta)$$ is the Gaussian probability of $$x_k$$ given the model parameters $$\Theta$$
#
# The equation for the Gaussian probability is $$ p(x) = \frac{1}{\sqrt{2\pi\sigma^2}} e^{-1/2 ((x - \mu)^2/\sigma^2)}$$
#
# Applying the natural logarithm into the negative log likehood function, we have
#
# $$ NLL = \sum_{k=1}^{N} \frac{ln(2\pi\sigma^2)}{2} + \frac{(x_k - \mu)^2}{2\sigma^2} $$
#
# If we assume that the observed values are samples from a Gaussian distribution with a predicted mean and variance, we can minimize the loss using the negative log-likehood criterion in place of the mean-squared error, with the following loss function, where $$y_k$$ is the true value and $$x_k$$ is the predicted value
#
#
#
# ## Configure imports
# +
import tensorflow.python.platform
import tensorflow as tf
import numpy as np
import os
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Input, Dense
import wget # pip3 install wget
import importlib
matplotlib_loader = importlib.find_loader("matplotlib")
PLT_FOUND = matplotlib_loader is not None
if PLT_FOUND:
import matplotlib as pyplot
# -
# The data used in this example is artifical, two parameter data of two different labels.
#
# We are going to read a few lines from one of the data files to determine how the data is organized.
# +
path_prefix = os.path.join("data", "example1")
filenameTrain = "saturn_data_train.csv"
filenameTest = "saturn_data_eval.csv"
localFilenameTrain = os.path.join(path_prefix, filenameTrain)
localFilenameTest = os.path.join(path_prefix, filenameTest)
# Data by Dr. <NAME> (http://www.jasonbaldridge.com) to test neural network frameworks.
# Read "https://github.com/jasonbaldridge/try-tf/tree/master/simdata" and copy
# to data/example1
if (
not os.path.isdir(path_prefix)
or not os.path.exists(localFilenameTrain)
or not os.path.exists(localFilenameTest)
):
# The actual URL for the raw data is:
URL = "https://raw.githubusercontent.com/jasonbaldridge/try-tf/master/simdata/"
print("Missing Saturn simulation data!")
print("Downloading from", URL)
os.mkdir(path_prefix)
wget.download(URL + "/" + filenameTrain, localFilenameTrain)
wget.download(URL + "/" + filenameTest, localFilenameTest)
print("\n\nExample 5.1 with TensorFlow version: {}".format(tf.__version__))
print("Eager execution: {}".format(tf.executing_eagerly()))
print("\nThe first five lines from the training data file:")
fd = open(localFilenameTrain)
for i in range(5):
sys.stdout.write(fd.readline())
fd.close()
# -
# Here, we can see tha the file is arranged into three columns. The first column is the label of the two different groups of data (group 0 and group 1). The second column is are the two features. We will assume that these two features are simply the coordinates of the point that is in the labeled group; i.e., x and y.
#
# +
NUM_LABELS = 2
def pack_features_vector(features, labels):
"""Pack the features into a single array."""
features = tf.stack(list(features.values()), axis=1)
return features, labels
def get_dataset(file_path, **kwargs):
"""Extract tf.data.Dataset representations of labels and features in CSV files given data in the format of label, feat[0], feat[1]. feat[2], etc..
Args:
file_path (string): The path to one or more CSV files to load.
Returns:
tf.data.Dataset : A object that holds the (fetures, labels) data from the CSV file in batches.
"""
# Use the 'experimental' make_csv_dataset to load the input data from the CSV file
dataset = tf.data.experimental.make_csv_dataset(file_path, num_epochs=1, **kwargs)
# Pack the features from a map of tensorflow data itnoa single feature vector.
dataset = dataset.map(pack_features_vector)
# Convert the integer lables in the dataset to one-hot encoded values.
dataset = dataset.map(lambda x, y: (x, tf.one_hot(y, depth=NUM_LABELS)))
if PLT_FOUND:
pyplot.figure()
# There are only two labels in this dataset 0 or 1
idx = labels > 0.5
pyplot.scatter(feat[idx, 0], feat[idx, 1], marker="+", c="#ff0000")
idx = labels <= 0.5
pyplot.scatter(feat[idx, 0], feat[idx, 1], marker="o", c="#00ff00")
pyplot.show()
return dataset
# +
BATCH_SIZE = 50
NUM_EPOCHS = 40 # Number of epochs, full passes of the data
NUM_INPUTS = 2
NUM_OUTPUTS = 2
NUM_HIDDEN_NODES = 20
MY_SEED = 123
# Constants that specify the data to load from the .csv files.
COLUMN_NAMES = ["label", "x", "y"]
LABEL_NAME = COLUMN_NAMES[0]
LABELS = [0, 1]
# Load the training data set and test data set into batches and suffle the input data before use.
training_batches = get_dataset(
localFilenameTrain,
batch_size=BATCH_SIZE,
column_names=COLUMN_NAMES,
label_name=LABEL_NAME,
shuffle=True,
shuffle_seed=MY_SEED,
)
print("\nDataset element defintion:\n\t", training_batches.element_spec)
testing_batches = get_dataset(
localFilenameTest,
batch_size=BATCH_SIZE,
column_names=COLUMN_NAMES,
label_name=LABEL_NAME,
shuffle=True,
shuffle_seed=MY_SEED,
)
# -
# ## Models
# Next, make the regression model to predict the label. For this example, the model has two layers. The input layer is an multilayer perceptron network with an RELU activation function and the output layer is is a softmax activation function with a negative log likelihood loss function.
#
# The weight initializer from the Deep Learning book is Xavier.
#
#
# ## Loss functions
# Let's examine the negative log likelihood function again.
#
# $$ NLL = \sum_{k=1}^{N} \frac{ln(2\pi\sigma^2)}{2} + \frac{(x_k - \mu)^2}{2\sigma^2} $$
#
# If we assume that the mean is 0.0 and the variance is 1.0, the negative log likelihood function simplifies to,
#
# $$ NLL = \sum_{k=1}^{N} \frac{ln(2\pi)}{2} + \frac{(x_k - \mu)^2}{2} $$
#
# $$ NLL = \frac{N ln(2\pi)}{2} + \sum_{k=1}^{N} \frac{(x_k - \mu)^2}{2} $$
#
#
# +
class MeanSquaredError(tf.keras.losses.Loss):
"""Custom loss function for calculating the loss as the mean-sequared error between the true output and the predicted output"""
def call(self, y_true, y_pred):
y_true = tf.cast(y_true, y_pred.dtype)
return tf.reduce_mean(tf.square(y_pred - y_true), axis=-1)
class NegativeLogLikelihood(tf.keras.losses.Loss):
"""Custom loss function for calculating the loss as negative log likelihood between the true output and the predicted output"""
def call(self, y_true, y_pred):
y_true = tf.cast(y_true, y_pred.dtype)
return tf.reduce_mean(tf.square(y_pred - y_true), axis=-1)
# +
# Build the model. For this example, the model has two layers. The input layer is
# an multilayer perceptron network with an RELU activation function and the output
# layer is is a softmax activation function with a negative log likelihood loss function.
#
# The weight initializer in the Deep Learning book is Xavier and it is seeded with MY_SEED (123)
initializer = tf.keras.initializers.GlorotNormal(seed=MY_SEED)
model = Sequential(
[
tf.keras.layers.Dense(
NUM_HIDDEN_NODES, activation="relu", kernel_initializer=initializer
),
tf.keras.layers.Dense(
NUM_OUTPUTS, activation="softmax", kernel_initializer=initializer
),
]
)
# Optimizer is Adam, loss function is mean squared error
model.compile(
loss=MeanSquaredError(),
optimizer=tf.optimizers.Adam(),
metrics=["accuracy"],
)
print("\n\nFit the training data.")
history = model.fit(training_batches, epochs=NUM_EPOCHS, verbose=1)
model.summary()
if PLT_FOUND:
# plot history
pyplot.plot(history.history["loss"], label="loss")
pyplot.plot(history.history["accuracy"], label="accuracy")
pyplot.title("Training loss and accuracy (MSE loss)")
pyplot.legend()
pyplot.show()
# Run against the test set. Final evaluation of the model
scores = model.evaluate(testing_batches, verbose=0)
print("Test set analysis accuracy: %.2f%%" % (scores[1] * 100))
| example5-1_nll.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Scientist Nanodegree
#
# ### Author: <NAME>
#
# ## Supervised Learning
# ## Project: Finding Donors for *CharityML*
# Welcome to the first project of the Data Scientist Nanodegree! In this notebook, some template code has already been provided for you, and it will be your job to implement the additional functionality necessary to successfully complete this project. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `'TODO'` statement. Please be sure to read the instructions carefully!
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
# ## Getting Started
#
# In this project, you will employ several supervised algorithms of your choice to accurately model individuals' income using data collected from the 1994 U.S. Census. You will then choose the best candidate algorithm from preliminary results and further optimize this algorithm to best model the data. Your goal with this implementation is to construct a model that accurately predicts whether an individual makes more than $50,000. This sort of task can arise in a non-profit setting, where organizations survive on donations. Understanding an individual's income can help a non-profit better understand how large of a donation to request, or whether or not they should reach out to begin with. While it can be difficult to determine an individual's general income bracket directly from public sources, we can (as we will see) infer this value from other publically available features.
#
# The dataset for this project originates from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Census+Income). The datset was donated by <NAME> and <NAME>, after being published in the article _"Scaling Up the Accuracy of Naive-Bayes Classifiers: A Decision-Tree Hybrid"_. You can find the article by <NAME> [online](https://www.aaai.org/Papers/KDD/1996/KDD96-033.pdf). The data we investigate here consists of small changes to the original dataset, such as removing the `'fnlwgt'` feature and records with missing or ill-formatted entries.
# ----
# ## Exploring the Data
# Run the code cell below to load necessary Python libraries and load the census data. Note that the last column from this dataset, `'income'`, will be our target label (whether an individual makes more than, or at most, $50,000 annually). All other columns are features about each individual in the census database.
# +
# USING PYTHON 3
# Import libraries necessary for this project
import numpy as np
import pandas as pd
from time import time
from IPython.display import display # Allows the use of display() for DataFrames
import matplotlib as mpl
import matplotlib.pyplot as plt
# Import supplementary visualization code visuals.py
import visuals as vs
# Pretty display for notebooks
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# Load the Census dataset
data = pd.read_csv("census.csv")
# Success - Display the first record
display(data.head(n=1))
# -
# Plot styles
plt.style.use('fivethirtyeight')
# ### Implementation: Data Exploration
# A cursory investigation of the dataset will determine how many individuals fit into either group, and will tell us about the percentage of these individuals making more than \$50,000. In the code cell below, you will need to compute the following:
# - The total number of records, `'n_records'`
# - The number of individuals making more than \$50,000 annually, `'n_greater_50k'`.
# - The number of individuals making at most \$50,000 annually, `'n_at_most_50k'`.
# - The percentage of individuals making more than \$50,000 annually, `'greater_percent'`.
#
# **HINT:** You may need to look at the table above to understand how the `'income'` entries are formatted.
# See what the income values are
data.income.unique()
# +
# Total number of records
n_records = data.shape[0]
# Number of records where individual's income is more than $50,000
n_greater_50k = data.query('income == ">50K"').shape[0]
# Number of records where individual's income is at most $50,000
n_at_most_50k = data.query('income == "<=50K"').shape[0]
# Percentage of individuals whose income is more than $50,000
greater_percent = (n_greater_50k / n_records) * 100
# Print the results
print("Total number of records: {:,}".format(n_records))
print("Individuals making more than $50,000: {:,}".format(n_greater_50k))
print("Individuals making at most $50,000: {:,}".format(n_at_most_50k))
print("Percentage of individuals making more than $50,000: {:.2f}%".format(greater_percent))
# -
# **Featureset Exploration**
#
# * **age**: continuous.
# * **workclass**: Private, Self-emp-not-inc, Self-emp-inc, Federal-gov, Local-gov, State-gov, Without-pay, Never-worked.
# * **education**: Bachelors, Some-college, 11th, HS-grad, Prof-school, Assoc-acdm, Assoc-voc, 9th, 7th-8th, 12th, Masters, 1st-4th, 10th, Doctorate, 5th-6th, Preschool.
# * **education-num**: continuous.
# * **marital-status**: Married-civ-spouse, Divorced, Never-married, Separated, Widowed, Married-spouse-absent, Married-AF-spouse.
# * **occupation**: Tech-support, Craft-repair, Other-service, Sales, Exec-managerial, Prof-specialty, Handlers-cleaners, Machine-op-inspct, Adm-clerical, Farming-fishing, Transport-moving, Priv-house-serv, Protective-serv, Armed-Forces.
# * **relationship**: Wife, Own-child, Husband, Not-in-family, Other-relative, Unmarried.
# * **race**: Black, White, Asian-Pac-Islander, Amer-Indian-Eskimo, Other.
# * **sex**: Female, Male.
# * **capital-gain**: continuous.
# * **capital-loss**: continuous.
# * **hours-per-week**: continuous.
# * **native-country**: United-States, Cambodia, England, Puerto-Rico, Canada, Germany, Outlying-US(Guam-USVI-etc), India, Japan, Greece, South, China, Cuba, Iran, Honduras, Philippines, Italy, Poland, Jamaica, Vietnam, Mexico, Portugal, Ireland, France, Dominican-Republic, Laos, Ecuador, Taiwan, Haiti, Columbia, Hungary, Guatemala, Nicaragua, Scotland, Thailand, Yugoslavia, El-Salvador, Trinadad&Tobago, Peru, Hong, Holand-Netherlands.
# ----
# ## Preparing the Data
# Before data can be used as input for machine learning algorithms, it often must be cleaned, formatted, and restructured — this is typically known as **preprocessing**. Fortunately, for this dataset, there are no invalid or missing entries we must deal with, however, there are some qualities about certain features that must be adjusted. This preprocessing can help tremendously with the outcome and predictive power of nearly all learning algorithms.
# ### Transforming Skewed Continuous Features
# A dataset may sometimes contain at least one feature whose values tend to lie near a single number, but will also have a non-trivial number of vastly larger or smaller values than that single number. Algorithms can be sensitive to such distributions of values and can underperform if the range is not properly normalized. With the census dataset two features fit this description: '`capital-gain'` and `'capital-loss'`.
#
# Run the code cell below to plot a histogram of these two features. Note the range of the values present and how they are distributed.
# +
# Split the data into features and target label
income_raw = data['income']
features_raw = data.drop('income', axis = 1)
# Visualize skewed continuous features of original data
vs.distribution(data)
# -
# For highly-skewed feature distributions such as `'capital-gain'` and `'capital-loss'`, it is common practice to apply a <a href="https://en.wikipedia.org/wiki/Data_transformation_(statistics)">logarithmic transformation</a> on the data so that the very large and very small values do not negatively affect the performance of a learning algorithm. Using a logarithmic transformation significantly reduces the range of values caused by outliers. Care must be taken when applying this transformation however: The logarithm of `0` is undefined, so we must translate the values by a small amount above `0` to apply the the logarithm successfully.
#
# Run the code cell below to perform a transformation on the data and visualize the results. Again, note the range of values and how they are distributed.
# +
# Log-transform the skewed features
skewed = ['capital-gain', 'capital-loss']
features_log_transformed = pd.DataFrame(data = features_raw)
features_log_transformed[skewed] = features_raw[skewed].apply(lambda x: np.log(x + 1))
# Visualize the new log distributions
vs.distribution(features_log_transformed, transformed = True)
# -
# ### Normalizing Numerical Features
# In addition to performing transformations on features that are highly skewed, it is often good practice to perform some type of scaling on numerical features. Applying a scaling to the data does not change the shape of each feature's distribution (such as `'capital-gain'` or `'capital-loss'` above); however, normalization ensures that each feature is treated equally when applying supervised learners. Note that once scaling is applied, observing the data in its raw form will no longer have the same original meaning, as exampled below.
#
# Run the code cell below to normalize each numerical feature. We will use [`sklearn.preprocessing.MinMaxScaler`](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html) for this.
# +
from sklearn.preprocessing import MinMaxScaler
# Initialize a scaler, then apply it to the features
scaler = MinMaxScaler() # default=(0, 1)
numerical = [
'age', 'education-num', 'capital-gain', 'capital-loss', 'hours-per-week'
]
features_log_minmax_transform = pd.DataFrame(data=features_log_transformed)
features_log_minmax_transform[numerical] = scaler.fit_transform(
features_log_transformed[numerical])
# Show an example of a record with scaling applied
display(features_log_minmax_transform.head(n=5))
# -
# ### Implementation: Data Preprocessing
#
# From the table in **Exploring the Data** above, we can see there are several features for each record that are non-numeric. Typically, learning algorithms expect input to be numeric, which requires that non-numeric features (called *categorical variables*) be converted. One popular way to convert categorical variables is by using the **one-hot encoding** scheme. One-hot encoding creates a _"dummy"_ variable for each possible category of each non-numeric feature. For example, assume `someFeature` has three possible entries: `A`, `B`, or `C`. We then encode this feature into `someFeature_A`, `someFeature_B` and `someFeature_C`.
#
# | | someFeature | | someFeature_A | someFeature_B | someFeature_C |
# | :-: | :-: | :-: | :-: | :-: | :-: |
# | 0 | B | | 0 | 1 | 0 |
# | 1 | C | ----> one-hot encode ----> | 0 | 0 | 1 |
# | 2 | A | | 1 | 0 | 0 |
#
# Additionally, as with the non-numeric features, we need to convert the non-numeric target label, `'income'` to numerical values for the learning algorithm to work. Since there are only two possible categories for this label ("<=50K" and ">50K"), we can avoid using one-hot encoding and simply encode these two categories as `0` and `1`, respectively. In code cell below, you will need to implement the following:
# - Use [`pandas.get_dummies()`](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html?highlight=get_dummies#pandas.get_dummies) to perform one-hot encoding on the `'features_log_minmax_transform'` data.
# - Convert the target label `'income_raw'` to numerical entries.
# - Set records with "<=50K" to `0` and records with ">50K" to `1`.
# +
# One-hot encode the 'features_log_minmax_transform' data using pandas.get_dummies()
features_final = pd.get_dummies(features_log_minmax_transform)
# Encode the 'income_raw' data to numerical values
income = income_raw.apply(lambda x: 1 if x == '>50K' else 0)
# Print the number of features after one-hot encoding
encoded = list(features_final.columns)
print("{} total features after one-hot encoding.".format(len(encoded)))
# Uncomment the following line to see the encoded feature names
print(encoded)
# -
# ### Shuffle and Split Data
# Now all _categorical variables_ have been converted into numerical features, and all numerical features have been normalized. As always, we will now split the data (both features and their labels) into training and test sets. 80% of the data will be used for training and 20% for testing.
#
# Run the code cell below to perform this split.
# +
# Import train_test_split
from sklearn.model_selection import train_test_split
# Split the 'features' and 'income' data into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(features_final,
income,
test_size=0.2,
random_state=42)
# Show the results of the split
print("Training set has {} samples.".format(X_train.shape[0]))
print("Testing set has {} samples.".format(X_test.shape[0]))
# -
# ----
# ## Evaluating Model Performance
# In this section, we will investigate four different algorithms, and determine which is best at modeling the data. Three of these algorithms will be supervised learners of your choice, and the fourth algorithm is known as a *naive predictor*.
# ### Metrics and the Naive Predictor
# *CharityML*, equipped with their research, knows individuals that make more than \\$50,000 are most likely to donate to their charity. Because of this, *CharityML* is particularly interested in predicting who makes more than \\$50,000 accurately. It would seem that using **accuracy** as a metric for evaluating a particular model's performace would be appropriate. Additionally, identifying someone that *does not* make more than \\$50,000 as someone who does would be detrimental to *CharityML*, since they are looking to find individuals willing to donate. Therefore, a model's ability to precisely predict those that make more than \\$50,000 is *more important* than the model's ability to **recall** those individuals. We can use **F-beta score** as a metric that considers both precision and recall:
#
# $$ F_{\beta} = (1 + \beta^2) \cdot \frac{precision \cdot recall}{\left( \beta^2 \cdot precision \right) + recall} $$
#
# In particular, when $\beta = 0.5$, more emphasis is placed on precision. This is called the **F$_{0.5}$ score** (or F-score for simplicity).
#
# Looking at the distribution of classes (those who make at most \\$50,000, and those who make more), it's clear most individuals do not make more than \\$50,000. This can greatly affect **accuracy**, since we could simply say *"this person does not make more than \\$50,000"* and generally be right, without ever looking at the data! Making such a statement would be called **naive**, since we have not considered any information to substantiate the claim. It is always important to consider the *naive prediction* for your data, to help establish a benchmark for whether a model is performing well. That been said, using that prediction would be pointless: If we predicted all people made less than \\$50,000, *CharityML* would identify no one as donors.
#
#
# #### Note: Recap of accuracy, precision, recall
#
# **Accuracy** measures how often the classifier makes the correct prediction. It’s the ratio of the number of correct predictions to the total number of predictions (the number of test data points).
#
# **Precision** tells us what proportion of messages we classified as spam, actually were spam.
# It is a ratio of true positives(words classified as spam, and which are actually spam) to all positives(all words classified as spam, irrespective of whether that was the correct classificatio), in other words it is the ratio of
#
# `[True Positives/(True Positives + False Positives)]`
#
# **Recall(sensitivity)** tells us what proportion of messages that actually were spam were classified by us as spam.
# It is a ratio of true positives(words classified as spam, and which are actually spam) to all the words that were actually spam, in other words it is the ratio of
#
# `[True Positives/(True Positives + False Negatives)]`
#
# For classification problems that are skewed in their classification distributions like in our case, for example if we had a 100 text messages and only 2 were spam and the rest 98 weren't, accuracy by itself is not a very good metric. We could classify 90 messages as not spam(including the 2 that were spam but we classify them as not spam, hence they would be false negatives) and 10 as spam(all 10 false positives) and still get a reasonably good accuracy score. For such cases, precision and recall come in very handy. These two metrics can be combined to get the F1 score, which is weighted average(harmonic mean) of the precision and recall scores. This score can range from 0 to 1, with 1 being the best possible F1 score(we take the harmonic mean as we are dealing with ratios).
# ### Question 1 - Naive Predictor Performace
# * If we chose a model that always predicted an individual made more than $50,000, what would that model's accuracy and F-score be on this dataset? You must use the code cell below and assign your results to `'accuracy'` and `'fscore'` to be used later.
#
# **Please note** that the the purpose of generating a naive predictor is simply to show what a base model without any intelligence would look like. In the real world, ideally your base model would be either the results of a previous model or could be based on a research paper upon which you are looking to improve. When there is no benchmark model set, getting a result better than random choice is a place you could start from.
#
# **HINT:**
#
# * When we have a model that always predicts '1' (i.e. the individual makes more than 50k) then our model will have no True Negatives(TN) or False Negatives(FN) as we are not making any negative('0' value) predictions. Therefore our Accuracy in this case becomes the same as our Precision(True Positives/(True Positives + False Positives)) as every prediction that we have made with value '1' that should have '0' becomes a False Positive; therefore our denominator in this case is the total number of records we have in total.
# * Our Recall score(True Positives/(True Positives + False Negatives)) in this setting becomes 1 as we have no False Negatives.
# +
TP = np.sum(income)
FP = income.count() - TP
TN = 0 # No predicted negatives in the naive case
FN = 0 # No predicted negatives in the naive case
# Calculate accuracy, precision and recall
accuracy = TP / (TP + FP)
recall = TP / (TP + FN)
precision = TP / (TP + FP)
# Calculate F-score using the formula above for beta = 0.5 and correct values for precision and recall.
beta = 0.5
fscore = (1 + beta**2) * (precision * recall) / (beta**2* precision + recall)
# Print the results
print("Naive Predictor: [Accuracy score: {:.4f}, F-score: {:.4f}]".format(
accuracy, fscore))
# -
# ### Supervised Learning Models
# **The following are some of the supervised learning models that are currently available in** [`scikit-learn`](http://scikit-learn.org/stable/supervised_learning.html) **that you may choose from:**
# - Gaussian Naive Bayes (GaussianNB)
# - Decision Trees
# - Ensemble Methods (Bagging, AdaBoost, Random Forest, Gradient Boosting)
# - K-Nearest Neighbors (KNeighbors)
# - Stochastic Gradient Descent Classifier (SGDC)
# - Support Vector Machines (SVM)
# - Logistic Regression
# ### Question 2 - Model Application
# List three of the supervised learning models above that are appropriate for this problem that you will test on the census data. For each model chosen
#
# - Describe one real-world application in industry where the model can be applied.
# - What are the strengths of the model; when does it perform well?
# - What are the weaknesses of the model; when does it perform poorly?
# - What makes this model a good candidate for the problem, given what you know about the data?
#
# **HINT:**
#
# Structure your answer in the same format as above^, with 4 parts for each of the three models you pick. Please include references with your answer.
# **Answer:**
#
# Using [Scikit-Learn's handy decision-making cheatsheet](https://scikit-learn.org/stable/tutorial/machine_learning_map/), I determine that since we're predicting a category (whether someone has >50K income), and have labeled data, our best models are going to be **classification models**.
#
# Following the path through the cheatsheet, I begin with Linear SVC (an SVM model), then continue to KNeighbors Classifier (a K-Nearest Neighbors model), and eventually to Ensemble Classifiers. As my ensemble model I selected Random Forest.
#
# I selected: **Support Vector Machines (SVM)**, **K-Nearest Neighbors (KNeighbors)**, and **Random Forest**.
#
# Here's an analysis of each:
#
# * Describe one real-world application in industry where the model can be applied.
# * Support Vector Machines (SVM): predicting whether an individual is a user of a certain mobile app based on a number of demographic factors
# * K-Nearest Neighbors (KNeighbors): according to Scikit, "nearest neighbors has been successful in a large number of classification and regression problems, including handwritten digits and satellite image scenes. Being a non-parametric method, it is often successful in classification situations where the decision boundary is very irregular." (Source: https://scikit-learn.org/stable/modules/neighbors.html)
# * Random Forest: predicting crime from lots of different signals
# * What are the strengths of the model; when does it perform well?
# * Support Vector Machines (SVM): it is effective in high-dimensionsal spaces, it is memory efficient, and because it uses the kernel trick, we benefit from some built-in knowledge (Source: https://scikit-learn.org/stable/modules/svm.html#classification)
# * K-Nearest Neighbors (KNeighbors): can be robust when our training data is noisy, and effective when we have a lot of training data (Source: https://people.revoledu.com/kardi/tutorial/KNN/Strength%20and%20Weakness.htm)
# * Random Forest: this is an ensemble model, built from a number of Decision Trees. For that reason, Random Forest often outperforms Decision Trees alone. They can also be faster than other models.
# * What are the weaknesses of the model; when does it perform poorly?
# * Support Vector Machines (SVM): choosing the right kernel function can be difficult, and it can be difficult to tune the different parameters. (Source: https://statinfer.com/204-6-8-svm-advantages-disadvantages-applications/).
# * K-Nearest Neighbors (KNeighbors): the model can be difficult to tune, specifically finding the optimal value of K (number of nearest neighbors). The computational cost of this model is also high. (Source: https://people.revoledu.com/kardi/tutorial/KNN/Strength%20and%20Weakness.htm)
# * Random Forest: can be prone to overfitting, especially when training data is noisy. Random forests are also unable to predict outside the ranges present in the training data. (Source: http://blog.citizennet.com/blog/2012/11/10/random-forests-ensembles-and-performance-metrics)
# * What makes this model a good candidate for the problem, given what you know about the data?
# * Support Vector Machines (SVM): according to the decision chart on Scikit's website, SVM is a good first choice because we are predicting a category, have labeled data, and fewer than 100K samples. Linear SVC (and SVM model) is our first stop.
# * K-Nearest Neighbors (KNeighbors): this model is a good second choice after SVM according to the decision chart when we have non-textual data.
# * Random Forest: after SVM and KNeighbors, Ensemble Classifiers are our next option. Random Forests is one of the most popular and well-known ensemble methods to try.
#
# ### Implementation - Creating a Training and Predicting Pipeline
# To properly evaluate the performance of each model you've chosen, it's important that you create a training and predicting pipeline that allows you to quickly and effectively train models using various sizes of training data and perform predictions on the testing data. Your implementation here will be used in the following section.
# In the code block below, you will need to implement the following:
# - Import `fbeta_score` and `accuracy_score` from [`sklearn.metrics`](http://scikit-learn.org/stable/modules/classes.html#sklearn-metrics-metrics).
# - Fit the learner to the sampled training data and record the training time.
# - Perform predictions on the test data `X_test`, and also on the first 300 training points `X_train[:300]`.
# - Record the total prediction time.
# - Calculate the accuracy score for both the training subset and testing set.
# - Calculate the F-score for both the training subset and testing set.
# - Make sure that you set the `beta` parameter!
# +
# Import two metrics from sklearn - fbeta_score and accuracy_score
from sklearn.metrics import fbeta_score, accuracy_score
def train_predict(learner, sample_size, X_train, y_train, X_test, y_test):
'''
inputs:
- learner: the learning algorithm to be trained and predicted on
- sample_size: the size of samples (number) to be drawn from training set
- X_train: features training set
- y_train: income training set
- X_test: features testing set
- y_test: income testing set
'''
results = {}
# Fit the learner to the training data using slicing with 'sample_size' using .fit(training_features[:], training_labels[:])
start = time() # Get start time
learner = learner.fit(X_train[:sample_size], y_train[:sample_size])
end = time() # Get end time
# Calculate the training time
results['train_time'] = end - start
# Get the predictions on the test set(X_test),
# then get predictions on the first 300 training samples(X_train) using .predict()
start = time() # Get start time
predictions_test = learner.predict(X_test)
predictions_train = learner.predict(X_train[:300])
end = time() # Get end time
# Calculate the total prediction time
results['pred_time'] = end - start
# Compute accuracy on the first 300 training samples which is y_train[:300]
results['acc_train'] = accuracy_score(y_train[:300], predictions_train)
# Compute accuracy on test set using accuracy_score()
results['acc_test'] = accuracy_score(y_test, predictions_test)
# Compute F-score on the the first 300 training samples using fbeta_score()
results['f_train'] = fbeta_score(y_train[:300],
predictions_train[:300],
beta=0.5)
# Compute F-score on the test set which is y_test
results['f_test'] = fbeta_score(y_test, predictions_test, beta=0.5)
# Success
print("{} trained on {} samples.".format(learner.__class__.__name__,
sample_size))
# Return the results
return results
# -
# ### Implementation: Initial Model Evaluation
# In the code cell, you will need to implement the following:
# - Import the three supervised learning models you've discussed in the previous section.
# - Initialize the three models and store them in `'clf_A'`, `'clf_B'`, and `'clf_C'`.
# - Use a `'random_state'` for each model you use, if provided.
# - **Note:** Use the default settings for each model — you will tune one specific model in a later section.
# - Calculate the number of records equal to 1%, 10%, and 100% of the training data.
# - Store those values in `'samples_1'`, `'samples_10'`, and `'samples_100'` respectively.
#
# **Note:** Depending on which algorithms you chose, the following implementation may take some time to run!
# +
# Import the three supervised learning models from sklearn
from sklearn.svm import LinearSVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
# Initialize the three models
clf_A = LinearSVC(random_state=42)
clf_B = KNeighborsClassifier(n_neighbors=3)
clf_C = RandomForestClassifier(random_state=42, n_estimators=10)
# Calculate the number of samples for 1%, 10%, and 100% of the training data
samples_100 = len(y_train)
samples_10 = int(samples_100 * 0.1)
samples_1 = int(samples_100 * 0.01)
# Collect results on the learners
results = {}
for clf in [clf_A, clf_B, clf_C]:
clf_name = clf.__class__.__name__
results[clf_name] = {}
for i, samples in enumerate([samples_1, samples_10, samples_100]):
results[clf_name][i] = \
train_predict(clf, samples, X_train, y_train, X_test, y_test)
# Run metrics visualization for the three supervised learning models chosen
vs.evaluate(results, accuracy, fscore)
# -
# ----
# ## Improving Results
# In this final section, you will choose from the three supervised learning models the *best* model to use on the student data. You will then perform a grid search optimization for the model over the entire training set (`X_train` and `y_train`) by tuning at least one parameter to improve upon the untuned model's F-score.
# ### Question 3 - Choosing the Best Model
#
# * Based on the evaluation you performed earlier, in one to two paragraphs, explain to *CharityML* which of the three models you believe to be most appropriate for the task of identifying individuals that make more than \\$50,000.
#
# **HINT:**
# Look at the graph at the bottom left from the cell above(the visualization created by `vs.evaluate(results, accuracy, fscore)`) and check the F score for the testing set when 100% of the training set is used. Which model has the highest score? Your answer should include discussion of the:
# * metrics - F score on the testing when 100% of the training data is used,
# * prediction/training time
# * the algorithm's suitability for the data.
# **Answer:**
#
# Linear SVC and Random Forest performed almost the same in both accuracy and f-score. Both were above 0.8 in accuracy and just under 0.7 in f-score. The difference is quite small. Since we're going to be tuning the model next, I'm selecting Random Forest since it is a much more tuneable model with many more parameters available to optimize.
#
# (It's also notable how much slower KNeighbors was than the other two models.)
# ### Question 4 - Describing the Model in Layman's Terms
#
# * In one to two paragraphs, explain to *CharityML*, in layman's terms, how the final model chosen is supposed to work. Be sure that you are describing the major qualities of the model, such as how the model is trained and how the model makes a prediction. Avoid using advanced mathematical jargon, such as describing equations.
#
# **HINT:**
#
# When explaining your model, if using external resources please include all citations.
# **Answer:**
#
# Decision Trees are like the if-then flowcharts we made in school. For example, if you wanted to separate dogs from cats, you might first classify them based on their size, then the sounds they make, then other factors (such as whether they're panting or purring!). Decision trees are great ways to split data into categories.
#
# Random Forests are made of lots of Decision Trees (just like real forests). Random Forests are what's known as an ensemble model. That means it's an aggregated model composed of the outputs of several individual sub-models, in this case decision trees. Ensemble models often outperform individual models for the same reason that crowd-sourced predictions often outperform individual predictions.
#
# A Random Forest is an ensemble of Decision Trees, but with a bit of randomness introduced when each tree is grown. Each tree is asked to search for the best path among a random _subset_ of features, not all of the features. In our example of dogs and cats, one tree might be asked to look at only tail length and color, whereas another might be asked to contemplate purring sounds and sleep patterns. That generates a lot of diversity in the trees, and that diversity can result in a better model when all of those decision trees are brought together into an ensemble.
#
# (Source: _Hands-On Machine Learning with Scikit-Learn & TensorFlow_, Geron, p. 183, 191).
# ### Implementation: Model Tuning
# Fine tune the chosen model. Use grid search (`GridSearchCV`) with at least one important parameter tuned with at least 3 different values. You will need to use the entire training set for this. In the code cell below, you will need to implement the following:
# - Import [`sklearn.grid_search.GridSearchCV`](http://scikit-learn.org/0.17/modules/generated/sklearn.grid_search.GridSearchCV.html) and [`sklearn.metrics.make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html).
# - Initialize the classifier you've chosen and store it in `clf`.
# - Set a `random_state` if one is available to the same state you set before.
# - Create a dictionary of parameters you wish to tune for the chosen model.
# - Example: `parameters = {'parameter' : [list of values]}`.
# - **Note:** Avoid tuning the `max_features` parameter of your learner if that parameter is available!
# - Use `make_scorer` to create an `fbeta_score` scoring object (with $\beta = 0.5$).
# - Perform grid search on the classifier `clf` using the `'scorer'`, and store it in `grid_obj`.
# - Fit the grid search object to the training data (`X_train`, `y_train`), and store it in `grid_fit`.
#
# **Note:** Depending on the algorithm chosen and the parameter list, the following implementation may take some time to run!
# +
# Suppress chatty warnings, thanks StackOverflow:
# https://stackoverflow.com/questions/32612180/eliminating-warnings-from-scikit-learn
def warn(*args, **kwargs):
pass
import warnings
warnings.warn = warn
# Import 'GridSearchCV', 'make_scorer', and any other necessary libraries
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
# Initialize the classifier
clf = RandomForestClassifier(random_state=42)
# Create the parameters list you wish to tune, using a dictionary if needed.
# HINT: parameters = {'parameter_1': [value1, value2], 'parameter_2': [value1, value2]}
'''
Here are the parameters I'm choosing to tune, from the dictionary on the Scikit-Learn
website: https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html
criterion : string, optional (default=”gini”)
The function to measure the quality of a split. Supported criteria are “gini” for
the Gini impurity and “entropy” for the information gain. Note: this parameter is
tree-specific.
n_estimators : integer, optional (default=10)
The number of trees in the forest.
max_depth : integer or None, optional (default=None)
The maximum depth of the tree. If None, then nodes are expanded until all leaves
are pure or until all leaves contain less than min_samples_split samples.
min_samples_split : int, float, optional (default=2)
The minimum number of samples required to split an internal node:
'''
parameters = {
'criterion': ['gini', 'entropy'],
'n_estimators': [10, 100, 1000],
'max_depth': [1, 32],
'min_samples_split': [0.1, 1.0, 10]
}
# Make an fbeta_score scoring object using make_scorer()
scorer = make_scorer(fbeta_score, beta=0.5)
# Perform grid search on the classifier using 'scorer' as the scoring method using GridSearchCV()
grid_obj = GridSearchCV(estimator=clf,
param_grid=parameters,
scoring=scorer,
cv=3)
# Fit the grid search object to the training data and find the optimal parameters using fit()
grid_fit = grid_obj.fit(X_train, y_train)
# Get the estimator
best_clf = grid_fit.best_estimator_
# Make predictions using the unoptimized and model
predictions = (clf.fit(X_train, y_train)).predict(X_test)
best_predictions = best_clf.predict(X_test)
# Report the before-and-afterscores
print("Unoptimized model\n------")
print("Accuracy score on testing data: {:.4f}".format(
accuracy_score(y_test, predictions)))
print("F-score on testing data: {:.4f}".format(
fbeta_score(y_test, predictions, beta=0.5)))
print("\nOptimized Model\n------")
print("Final accuracy score on the testing data: {:.4f}".format(
accuracy_score(y_test, best_predictions)))
print("Final F-score on the testing data: {:.4f}".format(
fbeta_score(y_test, best_predictions, beta=0.5)))
# -
# See what the winning parameters were
best_clf
# ### Question 5 - Final Model Evaluation
#
# * What is your optimized model's accuracy and F-score on the testing data?
# * Are these scores better or worse than the unoptimized model?
# * How do the results from your optimized model compare to the naive predictor benchmarks you found earlier in **Question 1**?
#
# **Note:** Fill in the table below with your results, and then provide discussion in the **Answer** box.
# #### Results:
#
# | Metric | Unoptimized Model | Optimized Model | Naive Benchmarks |
# | :------------: | :---------------: | :-------------: | :-------------: |
# | Accuracy Score | 0.8400 | 0.8621 | 0.2478 |
# | F-score | 0.6927 | 0.7514 | 0.2917 |
#
# **Answer:**
#
# Our optimized model performs better than our unoptimized model in both accuracy and f-score. Both unoptimized and optimized models significantly outperform the naive benchmarks we calculated in Question 1.
# ----
# ## Feature Importance
#
# An important task when performing supervised learning on a dataset like the census data we study here is determining which features provide the most predictive power. By focusing on the relationship between only a few crucial features and the target label we simplify our understanding of the phenomenon, which is most always a useful thing to do. In the case of this project, that means we wish to identify a small number of features that most strongly predict whether an individual makes at most or more than \\$50,000.
#
# Choose a scikit-learn classifier (e.g., adaboost, random forests) that has a `feature_importance_` attribute, which is a function that ranks the importance of features according to the chosen classifier. In the next python cell fit this classifier to training set and use this attribute to determine the top 5 most important features for the census dataset.
# ### Question 6 - Feature Relevance Observation
# When **Exploring the Data**, it was shown there are thirteen available features for each individual on record in the census data. Of these thirteen records, which five features do you believe to be most important for prediction, and in what order would you rank them and why?
# **Answer:**
#
# The five features I predict will be most important, in order, are:
#
# * **capital-gain**: I predict that higher capital gains must correlate with higher incomes. This one is probably the most predictive!
# * **workclass (Private, Federal-gov, Local-gov, State-gov)**: I predict that people employed by private companies or the government are more likely to have incomes >\\$50K than those who have never worked or are self-employed
# * **hours-per-week**: I predict that the more hours one works, the higher their income must be. This might not hold true at very income income levels, but I expect it to be predictive of incomes >\\$50K
# * **education (Bachelors, Masters, Doctorate)**: I predict these degrees to be predictive of incomes >\\$50K
# * **occupation (Sales, Exec-managerial, Prof-specialty)**: I predict these occupational subspecialties I expect to be most predictive of incomes >\\$50K as they tend to be higher-paying jobs than the other subspecialties listed
#
# All of these seem correlated with income to me, and I expect they will help predict income >\\$50K.
# ### Implementation - Extracting Feature Importance
# Choose a `scikit-learn` supervised learning algorithm that has a `feature_importance_` attribute availble for it. This attribute is a function that ranks the importance of each feature when making predictions based on the chosen algorithm.
#
# In the code cell below, you will need to implement the following:
# - Import a supervised learning model from sklearn if it is different from the three used earlier.
# - Train the supervised model on the entire training set.
# - Extract the feature importances using `'.feature_importances_'`.
# +
# Import a supervised learning model that has 'feature_importances_'
# Train the supervised model on the training set using .fit(X_train, y_train)
# Note: I'm using the exact model from best_clf
model = RandomForestClassifier(bootstrap=True,
class_weight=None,
criterion='entropy',
max_depth=32,
max_features='auto',
max_leaf_nodes=None,
min_impurity_decrease=0.0,
min_impurity_split=None,
min_samples_leaf=1,
min_samples_split=10,
min_weight_fraction_leaf=0.0,
n_estimators=1000,
n_jobs=None,
oob_score=False,
random_state=42,
verbose=0,
warm_start=False).fit(X_train, y_train)
# Extract the feature importances using .feature_importances_
importances = model.feature_importances_
# Plot
vs.feature_plot(importances, X_train, y_train)
# -
# ### Question 7 - Extracting Feature Importance
#
# Observe the visualization created above which displays the five most relevant features for predicting if an individual makes at most or above \$50,000.
# * How do these five features compare to the five features you discussed in **Question 6**?
# * If you were close to the same answer, how does this visualization confirm your thoughts?
# * If you were not close, why do you think these features are more relevant?
# **Answer:**
#
# The five features most important to my model are:
#
# * **age**
# * **capital-gain**
# * **marital-status** (Married-civ-spouse)
# * **hours-per-week**
# * **education-num**
#
# Only two of these were in my expected list: capital-gain and hours-per-week. I completely overlooked age, which now that I am able to consider is of course correlated with income! I was surprised to find marital-status in the top predictors. And although I speculated that education would be an important feature, I was wrong in selecting degree category: instead number of years of education was important.
#
# This is a perfect illustration of why machine learning outperforms traditional programming.
# ### Feature Selection
# How does a model perform if we only use a subset of all the available features in the data? With less features required to train, the expectation is that training and prediction time is much lower — at the cost of performance metrics. From the visualization above, we see that the top five most important features contribute more than half of the importance of **all** features present in the data. This hints that we can attempt to *reduce the feature space* and simplify the information required for the model to learn. The code cell below will use the same optimized model you found earlier, and train it on the same training set *with only the top five important features*.
# +
# Import functionality for cloning a model
from sklearn.base import clone
# Reduce the feature space
X_train_reduced = X_train[X_train.columns.values[(np.argsort(importances)[::-1])[:5]]]
X_test_reduced = X_test[X_test.columns.values[(np.argsort(importances)[::-1])[:5]]]
# Train on the "best" model found from grid search earlier
clf = (clone(best_clf)).fit(X_train_reduced, y_train)
# Make new predictions
reduced_predictions = clf.predict(X_test_reduced)
# Report scores from the final model using both versions of data
print("Final Model trained on full data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, best_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, best_predictions, beta = 0.5)))
print("\nFinal Model trained on reduced data\n------")
print("Accuracy on testing data: {:.4f}".format(accuracy_score(y_test, reduced_predictions)))
print("F-score on testing data: {:.4f}".format(fbeta_score(y_test, reduced_predictions, beta = 0.5)))
# -
# ### Question 8 - Effects of Feature Selection
#
# * How does the final model's F-score and accuracy score on the reduced data using only five features compare to those same scores when all features are used?
# * If training time was a factor, would you consider using the reduced data as your training set?
# **Answer:**
#
# The final model's accuracy and f-score are better than the scores with reduced data, however not by much:
#
# | Metric | Unoptimized Model | Optimized Model | Optimized Model <br>w/ Reduced Data | Diff |
# | :------------: | :---------------: | :-------------: | :-------------: | :---------: |
# | Accuracy Score | 0.8400 | 0.8621 | 0.8384 | -0.0237 |
# | F-score | 0.6927 | 0.7514 | 0.6926 | -0.0588 |
#
# The optimized model with reduced data performs quite closely to our unoptimized model. If training time were my _only_ consideration, I would consider using the reduced data as my training set. I might happily trade off the slight decline in performance for speed.
#
# However, reducing the model to only those factors could increase bias. Age, marital status, and education are signals that might correlate with individuals from specific ethnic, racial, or socioeconomic backgrounds. Eliminating the other signals could exacerbate such bias. For equity and fairness reasons, I would avoid using the reduced data model. Since our goal is to identify potential donors from the most diverse pool of individuals, we're willing to take the longer training time.
# ### Appendix: Trying Gradient Boosting and AdaBoost
#
# After marking my project completed, the reviewer said: "FYI, the highest scores I've seen from students used either Gradient Boosting or AdaBoost." That made me interested in trying those two models to see if I can improve upon my Random Forest results.
# +
from sklearn.ensemble import GradientBoostingClassifier, AdaBoostClassifier
# Initialize the two models
clf_A = GradientBoostingClassifier(random_state=42)
clf_B = AdaBoostClassifier(random_state=42)
# Calculate the number of samples for 1%, 10%, and 100% of the training data
samples_100 = len(y_train)
samples_10 = int(samples_100 * 0.1)
samples_1 = int(samples_100 * 0.01)
# Collect results on the learners
results = {}
for clf in [clf_A, clf_B]:
clf_name = clf.__class__.__name__
results[clf_name] = {}
for i, samples in enumerate([samples_1, samples_10, samples_100]):
results[clf_name][i] = \
train_predict(clf, samples, X_train, y_train, X_test, y_test)
# Run metrics visualization for the three supervised learning models chosen
vs.evaluate(results, accuracy, fscore)
# -
results['GradientBoostingClassifier']
results['AdaBoostClassifier']
# Sure enough, both models do perform better out of the box than Random Forest.
# Performance of the three optimized models:
#
# | Metric | Random Forest<br>Optimized | Gradient Boosting<br>(Unoptimized) | AdaBoost<br>(Unoptimized) |
# | :------------: | :-------------: | :-------------: | :---------: |
# | Accuracy Score | 0.8621 | 0.8638 | 0.8607 |
# | F-score | 0.7514 | 0.7580| 0.7491 |
# Gradient Boosting, unoptimized, has a higher accuracy and f-score than my optimized Random Forest model.
| p1_finding_donors/finding_donors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas lecture 2
# In the previous lecture, spent time to understand the data types `Series` and `DataFrame` and some basic operations using them. Among those operations, we learned how to add new columns and filter rows using the operators that `pandas` has implemented that operate on the `Series` and `DataFrame` types.
#
# But, since we are working with a general purpose language in Python, we should be allowed to define any sort of complex conditions and formulas for doing the same operations without limiting ourselves to just the operators implemented by `pandas`.
#
# The next few examples should make it clear what we are talking about.
import pandas as pd
import json
movies_df = pd.read_csv('movies.csv')
movies_df.head(5)
len(movies_df)
# ## Filtering by boolean indexing
#
# ### apply() method on Series
#
# This dataset contains information about the movies present in the Internet Movies database. A lot of the information, like the genres of the movies in the list is in the **JSON** format.
#
# Let's say we wanted to create a filtered `DataFrame` containing only Comedy movies. The approach we have already learnt to filetering will not apply in this case, as there is no `pandas` operator or built in function that will check for `name == Comedy` in a custom JSON expression.
#
# The approach that we will use allows us to write _any custom function_ which looks at the value of a _one column_ in a row, and returns True if we want to keep this row, and False otherwise. We have defined such a function below - which takes in the genre JSON string, and the actual genre to filter, and returns True if the movie genres list contains that genre.
def is_genre(json_str, genre):
genres = []
json_list = json.loads(json_str.replace("'", '"'))
for genre_json in json_list:
genres.append(genre_json['name'])
return genre in genres
# The next step is to create a new boolean column, which contains `True` if that row should be kept, and `False` otherwise. Given that we already have a function that returns that boolean value, we just need a way to create a new column that is filled with the return value of that function.
#
# In `pandas`, the `Series` type has a method called `apply()` - which takes as input a function with one argument. When called on the `Series`, it returns a new `Series` that consists of the output of the function _applied_ to each value in the original `Series`.
#
# We create a new column called `is_comedy` using this method.
movies_df['is_comedy'] = movies_df['genres'].apply(lambda x: is_genre(x, 'Comedy'))
movies_df['is_comedy'].head(5)
# **Aside:** For those not familiar with the `lamda` syntax, this is a cool way in Python to write small functions without using the full function syntax. The syntax is `lambda <input variables>: <expression for return value>`. This is extremely useful for data analysis.
#
# As you can see in the code above, we call the `apply()` method to the `genres` column in the `DataFrame`, pass in a function which checks if the movie is a Comedy movie, and assign the resultant `Series` to a new column called `is_comedy`.
#
# Now, we can simply use a regular filter and filter rows by checking if the value of column `is_comedy` is `True`. Note that in the code below, we omit `== True`. Why?
comedy_movies = movies_df[movies_df['is_comedy']]
comedy_movies.head(10)
len(comedy_movies)
# Just to recap, this method teaches us two things:
#
# - How to use `apply()` method in a series so that we can create a new column using any custom function. This is the crux of why this works.
# - Using the new column to filter the rows.
#
# This approach to filtering is called **Filtering by Boolean Indexing**.
# **Exercise:** Find all successful movies made by `<NAME>`. A successful movie is one whose revenue is greater than the budget.
# ### apply() method on DataFrame
#
# If you've completed the exercise above, you will have noticed that you needed another filtering step to judge the success of the movie, after you've applied the boolean indexing filter. This was required because you couldn't include the values of other columns in your function.
#
# In general, we would like to be able to get all the columns in a row to be able to compute the value of a new column that we want to create. Fortunately, the way to do that is similar to what we've already done: there is an `apply()` on the `DataFrame` class as well. Let's learn a bit more about it.
df = pd.DataFrame([[1, 2], [3, 4], [5, 6]], columns=['A', 'B'])
df
# Series apply(). We get a new Series with the application of the function.
df['A'].apply(lambda x: x**2)
# DataFrame apply(). The function is applied on each cell of the DataFrame
df.apply(lambda x: x**2)
# But, to create a new column, this is not what we are looking for. We want that function to be applied to each row, and we want a `Series` as a return value.
#
# To do that, the `apply()` method on `DataFrame` exposes another argument called `axis`. Like in Coordinate Geometry, axis defines the direction of movement. For example, in a 2-D plane, we have two axes which are perpendicular to each other.
#
# 
#
# Their meaning is very similar in `pandas`. Our dataset is also 2 dimensional - it has rows and columns. The value of axis can be 0 or 1, where
# - 0 is used to describe the direction including all rows, and
# - 1 is used to describe the direction including all columns.
#
# Here is a diagram to show that:
#
# 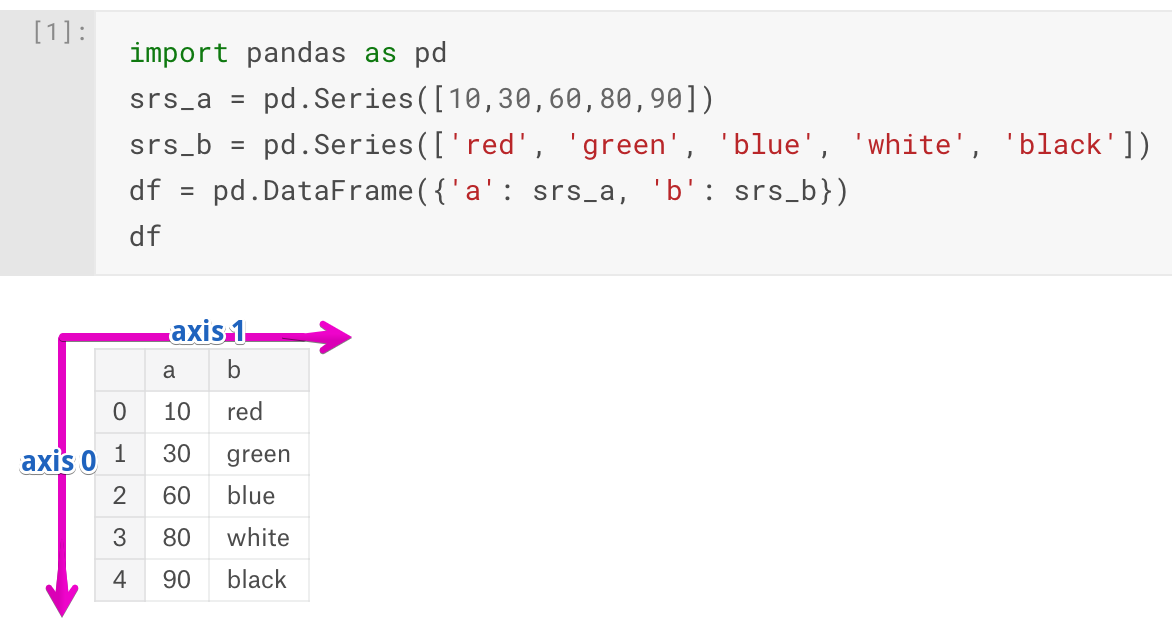
#
# Let's look at our small dataframe again. Let's say we want to print the sum of all elements in each row. What we can do is:
df['sum'] = df.apply(sum, axis=1)
df
# With axis=1, our function (`sum` in this case), is called once for each row. The argument to our function is a `Series` representing the values of that row.
#
# Just for completion sake, let's also see what happens when axis = 0. This case can be useful to create some sort of summary of the data.
df.loc[3] = df.apply(sum, axis=0)
df
# As you can see, with axis=0, the function is applied once for each column. The input to our function is a `Series` representing all the values in the column.
# Now let's do our exercise again, using the `apply()` method on `DataFrame`. Let's also twist it a bit, and look at failures instead of successes.
def studio_failure(row, studio):
studios = []
try:
json_list = json.loads(row['production_companies'].replace("'", '"'))
except:
return False
if not isinstance(json_list, list):
return False
for studio_json in json_list:
studios.append(studio_json['name'])
if row['revenue'] == '0' or row['budget'] == 0:
return False
failure = float(row['revenue']) < float(row['budget'])
return (studio in studios) and failure
columbia_failure_df = movies_df[movies_df.apply(lambda x: studio_failure(x, 'Columbia Pictures'), axis=1)]
columbia_failure_df.head(10)
# Note that
#
# - Because the data is not completely clean, the `studio_failure()` function has a bunch of conditions to eliminate certain rows.
# - We don't really need to create a new column for the output of `df.apply()`. The filter step just needs a boolean `Series` of the same length as the number of rows in the dataframe and that's what `df.apply()` returns when the `axis=1`.
# ## Exercises
#
# In this set of exercises, we are going to use a data set that contains a large list of wines, their description and reviews. The dataset is in the file [wine_data.csv](https://raw.githubusercontent.com/amangup/data-analysis-bootcamp/master/07-Pandas2/wine_data.csv).
#
# Let's start with a few exercises to get to know the data better:
#
# 1. How many wines are there?
# 2. How many US wines are there?
# 3. What is the average rating of a wine?
# 4. Let's look at the distribution of ratings. First, add the following import statement in your notebook:
#
# ```python
# import matplotlib.pyplot as plt
# ```
#
# and then run the following code in a cell:
#
# ```python
# raw_df['points'].hist()
# plt.show()
# ```
#
# 5. Do exercises 3 and 4 only for wines from the US, and only for wines from France.
# 6. Do exercises 3, 4 and 5, but this time let's look at the price of wines, instead of their rating. Note that by default the histogram is not as useful. Create a histogram of wines priced upto $100 instead.
# 7. Find if there what is the pearson correlation coefficient between the price of the wine and its rating. Hint: checkout the `corr()` method in the `DataFrame` class.
#
# ### Wine search
#
# Now that we are familiar the data, let's work on a mini project: Create a wine search which can highlight the top 5 wines that whose description matches the search keywords.
#
# We will use a well known method for keyword based ranking called the **tf-idf algorithm**.
#
# - Let's say we want to search wines which mention `oak` in their description.
# - We would like to highlight wines whose description mentions the work `oak` the maximum number of times.
# - This count of mentions is called **Term Frequency**, or **tf**.
# - Higher the **tf** of a wine, higher it's ranking.
# - Let's say the search term is `oak wine`.
# - `wine` is a common term and ideally we would not like to give that term much weight.
# - We will count the _number of descriptions_ in which the word `wine` appears. This count represents the **Inverse Document Frequency**, or **idf**.
# - If a word appears in most of the documents, that means that it's is a common word, and thus we should more or less ignore this word.
# - The precise definition of **idf** is `idf(word) = log(N_docs / N_docs_with_word)`, where `N_docs` is the total number of descriptions, and `N_docs_with_word` is the number of descriptions which contains the specific word. You can see that `idf` decreases as `N_docs_with_word` increases.
#
# Let's understand this using an example. Here is a sample description:
#
# "There's plenty of oak to this solid, peppery Merlot that's also a touch green. The cassis and cherry fruit that drives the palate is healthy and sturdy, while the finish features some tight-grained oak and firm enough tannins. Maybe too much oak given the fruit quality."
#
# - This sentence contains the word `oak` three times. Thus, `tf = 3`
# - The word 'oak' appears in 8,721 descriptions out of a total of 129,971 descriptions. Thus, `idf = log(129971/8721) = 2.7`. I've used base = `e` here.
# - The ranking score of this wine would be `tf * idf = 3 * 2.7 = 8.1` given the search term `oak`.
#
# Let's build our wine search engine by following these steps:
#
# 1. Write a function which takes a text and returns a dictionary with the word count for each word.
# 2. Add a column to your dataframe called `tf` that stores the word count dictionary corresponding to it's description.
# 3. Create another dictionary called `idf_dict` by going over all the descriptions for all wines, and map each word to its **idf** score.
# 4. Write a function which takes as argument the search keywords (as a list of strings) that:
# - calculates the score of each wine as: `tf(word_1) * idf(word_1) + ... + tf(word_k) * idf(word_k)`
# - sorts the wines in descending order of their score (Hint: use `sort_values()` method on `DataFrame`)
# - Returns a dataframe with details of the top 5 scoring wines.
#
# 5. Implement your own search ranking scheme that gives a certain priority to the rating of the wine as an augmentation of the ranking algorithm above. You can also implement a max price filter - to only search among wines priced lower than the max price.
| 07-Pandas2/.ipynb_checkpoints/pandas2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rizkiar00/HMMPosTaggingIndonesia/blob/main/PosTagging.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="HiK721jLhjVX" outputId="13c32849-9fd4-4aac-dc95-f10ebf4c1ec8"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="0YdbhjdwmXi-"
# ## Mengambil dan Mengekstrak Dataset
#
# https://github.com/kmkurn/id-pos-tagging/blob/master/data
# + colab={"base_uri": "https://localhost:8080/"} id="WJqwXN4Eh9Fd" outputId="819a385a-5c8d-47be-b9f9-edd7c8284345"
# !wget https://github.com/kmkurn/id-pos-tagging/blob/master/data/dataset.tar.gz?raw=true
# open file
import tarfile
file = tarfile.open('dataset.tar.gz?raw=true')
# extracting file
file.extractall('dataset')
file.close()
# + id="1BDYmz1Xh_q0"
# # Importing libraries
# import nltk
# import numpy as np
# import pandas as pd
# import random
# from sklearn.model_selection import train_test_split
# import pprint, time
# #download the treebank corpus from nltk
# nltk.download('treebank')
# #download the universal tagset from nltk
# nltk.download('universal_tagset')
# # reading the Treebank tagged sentences
# nltk_data = list(nltk.corpus.treebank.tagged_sents(tagset='universal'))
# #print the first two sentences along with tags
# print(nltk_data[:2])
# + [markdown] id="ZnW4R3M7mpeF"
# ## Mengambil Data Training
# + colab={"base_uri": "https://localhost:8080/"} id="LIiKVKWpldZz" outputId="7add2305-7404-48bf-c59d-86e3bd7e5995"
import csv
with open('/content/dataset/train.01.tsv', newline='') as f:
reader = csv.reader(f,delimiter='\t',quoting=csv.QUOTE_NONE)
train_set = list(reader)
print(train_set[0:40])
print(len(train_set))
# + [markdown] id="_ldqmsCEm2Nj"
# ## Mengambil Data Testing
# + colab={"base_uri": "https://localhost:8080/"} id="UiT7CyCsnHuw" outputId="53bba883-9bd6-42ff-b1b8-77fb2b11a629"
with open('/content/dataset/test.01.tsv', newline='') as f:
reader = csv.reader(f,delimiter='\t')
test_set = list(reader)
print(test_set[0:30])
print(len(test_set))
# + [markdown] id="PHXRiKJEm5qK"
# ## Preprocessing
#
#
# 1. Semua huruf jadi huruf kecil cth: LAsd -> lasd
# 2. Semua angka menjadi 0 cth: dsaq123 -> dsaq0
#
#
# + id="kROWchh61VGj"
def transform(w, lower=True, replace_digits=True):
if lower:
w = w.lower()
if replace_digits:
w = re.sub(r'\d+', '0', w)
return w
# + [markdown] id="cYrQyl9DnTOb"
# ## Membersihkan data yang kosong dan menyesuaikan format
# + colab={"base_uri": "https://localhost:8080/"} id="881vZCdwnfW-" outputId="e99072e5-c39e-4aa2-d1be-36f0292ca61c"
# create list of train and test tagged words
# train_tagged_words = [ tup for sent in train_set for tup in sent ]
# test_tagged_words = [ tup for sent in test_set for tup in sent ]
# print(len(train_tagged_words))
# print(len(test_tagged_words))
import regex as re
def Remove(tuples):
tuples = [t for t in tuples if t]
return tuples
train_set = Remove(train_set)
test_set = Remove(test_set)
train_tagged_words = []
for alist in train_set:
alist[0] = transform(alist[0])
train_tagged_words.append(tuple(alist))
test_tagged_words = []
for alist in test_set:
alist[0] = transform(alist[0])
test_tagged_words.append(tuple(alist))
print(train_tagged_words[:5])
print(test_tagged_words[:5])
# + [markdown] id="uK9t3L3_ndTs"
# ## Mengecek jenis kata dan vocabulary
# + colab={"base_uri": "https://localhost:8080/"} id="PtyjG8wcoq6z" outputId="40286d90-52c7-4b9f-db3f-f4b76a94c76a"
tags = {tag for word,tag in train_tagged_words}
print(len(tags))
print(tags)
# check total words in vocabulary
vocab = {word for word,tag in train_tagged_words}
print(len(vocab))
# + [markdown] id="DK8lLrNNnuHK"
# ## Fungsi untuk mengecek Peluang
# + id="DIUKUZIuwARY"
# compute Emission Probability
def word_given_tag(word, tag, train_bag = train_tagged_words):
tag_list = [pair for pair in train_bag if pair[1]==tag]
count_tag = len(tag_list)#total number of times the passed tag occurred in train_bag
w_given_tag_list = [pair[0] for pair in tag_list if pair[0]==word]
#now calculate the total number of times the passed word occurred as the passed tag.
count_w_given_tag = len(w_given_tag_list)
return (count_w_given_tag, count_tag)
# compute Transition Probability
def t2_given_t1(t2, t1, train_bag = train_tagged_words):
tags = [pair[1] for pair in train_bag]
count_t1 = len([t for t in tags if t==t1])
count_t2_t1 = 0
for index in range(len(tags)-1):
if tags[index]==t1 and tags[index+1] == t2:
count_t2_t1 += 1
return (count_t2_t1, count_t1)
# + [markdown] id="10IqIeLOn52z"
# #Membuat Model Peluang HMM
# + colab={"base_uri": "https://localhost:8080/", "height": 762} id="tZ_a4NIYuMYm" outputId="9d6bad56-45e5-49af-d449-eae3b9a943e6"
import numpy as np
import pandas as pd
tags_matrix = np.zeros((len(tags), len(tags)), dtype='float32')
for i, t1 in enumerate(list(tags)):
for j, t2 in enumerate(list(tags)):
tags_matrix[i, j] = t2_given_t1(t2, t1)[0]/t2_given_t1(t2, t1)[1]
tags_df = pd.DataFrame(tags_matrix, columns = list(tags), index=list(tags))
display(tags_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 920} id="GDkCyS0-xvb5" outputId="32b79164-f788-43b0-9d26-350f894ea553"
import seaborn as sns
import matplotlib.pyplot as plt
plt.figure(figsize = (25,16))
ax = sns.heatmap(tags_df, linewidths=.1,annot=True)
ax.xaxis.tick_top()
ax.tick_params(length=0)
# + [markdown] id="6cSncynXn-wT"
# ## Membuat Fungsi Viterbi
# + id="vfmTVG8Jxxnv"
def Viterbi(words, train_bag = train_tagged_words):
state = []
T = list(set([pair[1] for pair in train_bag]))
for key, word in enumerate(words):
#initialise list of probability column for a given observation
p = []
for tag in T:
if key == 0:
transition_p = tags_df.loc['Z', tag]
else:
transition_p = tags_df.loc[state[-1], tag]
# compute emission and state probabilities
emission_p = word_given_tag(words[key], tag)[0]/word_given_tag(words[key], tag)[1]
state_probability = emission_p * transition_p
p.append(state_probability)
pmax = max(p)
# getting state for which probability is maximum
state_max = T[p.index(pmax)]
state.append(state_max)
return list(zip(words, state))
# + [markdown] id="sv3fMVtqoDV7"
# ## Testing untuk 10 kata pertama pada data test
# + id="3dPEdu1RItQx" colab={"base_uri": "https://localhost:8080/"} outputId="6de028a8-ae14-4a25-b182-0f8766a4ff23"
import time
# list of 10 sents on which we test the model
test_run = test_set[0:10]
# list of tagged words
test_run_base = [tuple(sent) for sent in test_run]
# list of untagged words
test_tagged_words = [sent[0] for sent in test_run]
#Here We will only test 10 sentences to check the accuracy
#as testing the whole training set takes huge amount of time
start = time.time()
tagged_seq = Viterbi(test_tagged_words)
end = time.time()
difference = end-start
print("Time taken in seconds For The First 10th Word: ", difference)
# accuracy
check = [i for i, j in zip(tagged_seq, test_run_base) if i == j]
accuracy = len(check)/len(tagged_seq)
print('Viterbi Algorithm Accuracy For The First 10th Word: ',accuracy*100)
# + [markdown] id="bSRx4GXhoODc"
# ## Testing untuk semua data test
#
# (Dikarenakan terbatasnya google colab, maka testing dilakukan secara perlahan)
# + colab={"base_uri": "https://localhost:8080/"} id="1cVYEw-C2jRd" outputId="c2fafb45-fe29-4880-c6df-292446d1dc6d"
#Code to test all the test sentences
#(takes alot of time to run s0 we wont run it here)
# tagging the test sentences()
from pathlib import Path
import csv
test_tagged_words = [tuple(sent) for sent in test_set]
test_untagged_words = [sent[0] for sent in test_set]
#test_untagged_words
interval = 2000
filename = '/content/drive/MyDrive/output.csv'
if Path(filename).is_file():
with open(filename, newline='') as f:
reader = csv.reader(f)
listtagged_seq = list(reader)
prevtagged_seq = []
for i in listtagged_seq:
prevtagged_seq.append(tuple(i))
startidx = len(prevtagged_seq)
else:
startidx = 0
prevtagged_seq = []
endinterval = startidx + interval #(titik pada kalimat terakhir)
if endinterval > len(test_untagged_words):
endinterval = len(test_untagged_words)-1
for i in range(endinterval,len(test_untagged_words)):
if test_untagged_words[i] == '.':
break
endinterval = i+1
if len(prevtagged_seq) < len(test_tagged_words):
start = time.time()
current = start
tagged_seq = Viterbi(test_untagged_words[startidx:endinterval])
end = time.time()
difference = end-start
wordcount = endinterval - startidx
print("Finish at index: ", endinterval)
print("Number of Word Prcessed: ", wordcount)
print("Time taken in seconds: ", difference)
# accuracy
check = [i for i, j in zip(tagged_seq, test_tagged_words[startidx:endinterval]) if i == j]
accuracy = len(check)/len(tagged_seq)
print('Viterbi Algorithm Accuracy: ',accuracy*100)
prevtagged_seq.extend(tagged_seq)
listouput = []
for tuples in prevtagged_seq:
listouput.append(list(tuples))
with open(filename, "w") as f:
writer = csv.writer(f)
writer.writerows(listouput)
if len(prevtagged_seq) >= len(test_tagged_words):
print("Part Of Speech Tagging has Been Finished")
# + [markdown] id="5ulrV8oUpjnt"
# ## Hasil Akurasi dari prediksi POS Tagging pada semua data test
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="RqVX2WzQkhS1" outputId="9d1ad6f1-67d9-4ed3-a00a-344fa0595170"
with open(filename, newline='') as f:
reader = csv.reader(f)
listtagged_seq = list(reader)
prevtagged_seq = []
for i in listtagged_seq:
prevtagged_seq.append(tuple(i))
check = [i for i, j in zip(prevtagged_seq, test_tagged_words[0:len(prevtagged_seq)]) if i == j]
accuracy = len(check)/len(test_tagged_words[0:len(prevtagged_seq)])
print('Viterbi Algorithm Accuracy: ',accuracy*100)
# + [markdown] id="Rvb3JhOxpczE"
# ## NLTK untuk Word Tokenize
# + id="lB2zAn5LpXoz"
import nltk
nltk.download('punkt')
# + [markdown] id="a6BF1VxrpkrC"
# ## Testing untuk kalimat Inputan
# + colab={"base_uri": "https://localhost:8080/"} id="yw3CGY1xw-tA" outputId="836d8494-5562-4b4e-a282-70a6e4b0b7ca"
print('Masukkan Kalimat yang akan di Post Tagging: ')
sentence = input()
sentence = nltk.word_tokenize(sentence)
sentence = list(map(transform, sentence))
taggedsentence = Viterbi(sentence)
print(taggedsentence)
| PosTagging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_pdy0t0f"
# ### Problem Statement
#
# Write a function that takes an input array (or Python list) consisting of only `0`s, `1`s, and `2`s, and sorts that array in a single traversal.
#
# Note that if you can get the function to put the `0`s and `2`s in the correct positions, this will aotumatically cause the `1`s to be in the correct positions as well.
# + graffitiCellId="id_vsgisrd"
def sort_012(input_list):
pass
# + [markdown] graffitiCellId="id_rrxcwca"
# <span class="graffiti-highlight graffiti-id_rrxcwca-id_1f2p5yd"><i></i><button>Show Solution</button></span>
# + graffitiCellId="id_2sqv48t"
def test_function(test_case):
sort_012(test_case)
print(test_case)
if test_case == sorted(test_case):
print("Pass")
else:
print("Fail")
# + graffitiCellId="id_x3ai5yy"
test_case = [0, 0, 2, 2, 2, 1, 1, 1, 2, 0, 2]
test_function(test_case)
# + graffitiCellId="id_t8sucox"
test_case = [2, 1, 2, 0, 0, 2, 1, 0, 1, 0, 0, 2, 2, 2, 1, 2, 0, 0, 0, 2, 1, 0, 2, 0, 0, 1]
test_function(test_case)
# + graffitiCellId="id_mk5p9ba"
test_case = [2, 2, 0, 0, 2, 1, 0, 2, 2, 1, 1, 1, 0, 1, 2, 0, 2, 0, 1]
test_function(test_case)
| Basic Algorithms/Sort Algorithms/Sort-0-1-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Cartpole optimal control problem
#
# 
#
# A cartpole is another classical example of control. In this system, an underactuated pole is attached on top of a 1D actuacted cart. The game is to raise the pole to a standing position.
#
# The model is here:
# https://en.wikipedia.org/wiki/Inverted_pendulum
#
# We denote by $m_1$ the cart mass, $m_2$ the pole mass ($m=m_1+m_2$), $l$ the pole length, $\theta$ the pole angle w.r.t. the vertical axis, $p$ the cart position, and $g=$9.81 the gravity.
#
# The system acceleration can be rewritten as:
#
# $$\ddot{\theta} = \frac{1}{\mu(\theta)} \big( \frac{\cos \theta}{l} f + \frac{mg}{l} \sin(\theta) - m_2 \cos(\theta) \sin(\theta) \dot{\theta}^2\big),$$
#
# $$\ddot{p} = \frac{1}{\mu(\theta)} \big( f + m_2 \cos(\theta) \sin(\theta) g -m_2 l \sin(\theta) \dot{\theta} \big),$$
#
# $\hspace{12em}$with $$\mu(\theta) = m_1+m_2 \sin(\theta)^2,$$
#
# where $f$ represents the input command (i.e $f=u$) and $m=m_1+m_2$ is the total mass.
#
# ## I. Differential Action Model
#
# A Differential Action Model (DAM) describes the action (control/dynamics) in continous-time. In this exercise, we ask you to write the equation of motions for the cartpole.
#
# For more details, see the instructions inside the DifferentialActionModelCartpole class.
# +
import crocoddyl
import pinocchio
import numpy as np
class DifferentialActionModelCartpole(crocoddyl.DifferentialActionModelAbstract):
def __init__(self):
crocoddyl.DifferentialActionModelAbstract.__init__(self, crocoddyl.StateVector(4), 1, 6) # nu = 1; nr = 6
self.unone = np.zeros(self.nu)
self.m1 = 1.
self.m2 = .1
self.l = .5
self.g = 9.81
self.costWeights = [ 1., 1., 0.1, 0.001, 0.001, 1. ] # sin, 1-cos, x, xdot, thdot, f
def calc(self, data, x, u=None):
if u is None: u=model.unone
# Getting the state and control variables
x, th, xdot, thdot = np.asscalar(x[0]), np.asscalar(x[1]), np.asscalar(x[2]), np.asscalar(x[3])
f = np.asscalar(u[0])
# Shortname for system parameters
m1, m2, l, g = self.m1, self.m2, self.l, self.g
s, c = np.sin(th), np.cos(th)
###########################################################################
############ TODO: Write the dynamics equation of your system #############
###########################################################################
# Hint:
# You don't need to implement integration rules for your dynamic system.
# Remember that DAM implemented action models in continuous-time.
m = m1 + m2
mu = m1 + m2 * s**2
xddot, thddot = cartpole_dynamics(self, data, x, u) ### Write the cartpole dynamics here
data.xout = np.matrix([ xddot, thddot ]).T
# Computing the cost residual and value
data.r = np.matrix(self.costWeights * np.array([ s, 1-c, x, xdot, thdot, f ])).T
data.cost = .5* np.asscalar(sum(np.asarray(data.r)**2))
def calcDiff(model,data,x,u=None,recalc=True):
# Advance user might implement the derivatives
pass
# -
# You can get the solution by uncommenting the following line:
# +
# # %load solutions/cartpole_dyn.py
# -
# You may want to check your computation. Here is how to create the model and run the calc method.
cartpoleDAM = DifferentialActionModelCartpole()
cartpoleData = cartpoleDAM.createData()
x = cartpoleDAM.State.rand()
u = np.zeros(1)
cartpoleDAM.calc(cartpoleData,x,u)
# ## II. Write the derivatives with DAMNumDiff
#
# In the previous exercise, we didn't define the derivatives of the cartpole system. In crocoddyl, we can compute them without any additional code thanks to the DifferentialActionModelNumDiff class. This class computes the derivatives through numerical differentiation.
#
# In the following cell, you need to create a cartpole DAM that computes the derivates using NumDiff.
# Creating the carpole DAM using NumDiff for computing the derivatives.
# We specify the withGaussApprox=True to have approximation of the
# Hessian based on the Jacobian of the cost residuals.
cartpoleND = crocoddyl.DifferentialActionModelNumDiff(cartpoleDAM, True)
# After creating your cartpole DAM with NumDiff. We would like that you answer the follows:
#
# - 2 columns of Fx are null. Wich ones? Why?
#
# - can you double check the values of Fu?
#
# +
# # %load solutions/cartpole_fxfu.py
# -
# ## III. Integrate the model
#
# After creating DAM for the cartpole system. We need to create an Integrated Action Model (IAM). Remenber that an IAM converts the continuos-time action model into a discrete-time action model. For this exercise we'll use a simpletic Euler integrator.
# +
# # %load solutions/cartpole_integration.py
###########################################################################
################## TODO: Create an IAM with from the DAM ##################
###########################################################################
# Hint:
# Use IntegratedActionModelEuler
# -
# ## IV. Write the problem, create the solver
#
# First, you need to describe your shooting problem. For that, you have to indicate the number of knots and their time step. For this exercise we want to use 50 knots with $dt=$5e-2.
#
# Here is how we create the problem.
# Fill the number of knots (T) and the time step (dt)
x0 = np.matrix([ 0., 3.14, 0., 0. ]).T
T = 50
problem = crocoddyl.ShootingProblem(x0, [ cartpoleIAM ]*T, cartpoleIAM)
# Problem can not solve, just integrate:
us = [ pinocchio.utils.zero(cartpoleIAM.differential.nu) ]*T
xs = problem.rollout(us)
# In cartpole_utils, we provite a plotCartpole and a animateCartpole methods.
# +
# %%capture
# %matplotlib inline
from cartpole_utils import animateCartpole
anim = animateCartpole(xs)
# If you encounter problems probably you need to install ffmpeg/libav-tools:
# sudo apt-get install ffmpeg
# or
# sudo apt-get install libav-tools
# -
# And let's display this rollout!
#
# Note that to_jshtml spawns the video control commands.
from IPython.display import HTML
# HTML(anim.to_jshtml())
HTML(anim.to_html5_video())
# Now we want to create the solver (SolverDDP class) and run it. Display the result. **Do you like it?**
# +
# # %load solutions/cartpole_ddp.py
##########################################################################
################## TODO: Create the DDP solver and run it ###############
###########################################################################
# +
# %%capture
# %matplotlib inline
# Create animation
anim = animateCartpole(xs)
# -
# HTML(anim.to_jshtml())
HTML(anim.to_html5_video())
# ## Tune the problem, solve it
#
# Give some indication about what should be tried for solving the problem.
#
#
# - Without a terminal model, we can see some swings but we cannot stabilize. What should we do?
#
# - The most important is to reach the standing position. Can we also nullify the velocity?
#
# - Increasing all the weights is not working. How to slowly increase the penalty?
#
#
# +
# # %load solutions/cartpole_tuning.py
###########################################################################
################## TODO: Tune the weights for each cost ###################
###########################################################################
terminalCartpole = DifferentialActionModelCartpole()
terminalCartpoleDAM = crocoddyl.DifferentialActionModelNumDiff(terminalCartpole, True)
terminalCartpoleIAM = crocoddyl.IntegratedActionModelEuler(terminalCartpoleDAM)
terminalCartpole.costWeights[0] = 0 # fix me :)
terminalCartpole.costWeights[1] = 0 # fix me :)
terminalCartpole.costWeights[2] = 0 # fix me :)
terminalCartpole.costWeights[3] = 0 # fix me :)
terminalCartpole.costWeights[4] = 0 # fix me :)
terminalCartpole.costWeights[5] = 0 # fix me :)
problem = crocoddyl.ShootingProblem(x0, [ cartpoleIAM ]*T, terminalCartpoleIAM)
# +
# Creating the DDP solver
ddp = crocoddyl.SolverDDP(problem)
ddp.setCallbacks([ crocoddyl.CallbackVerbose() ])
# Solving this problem
done = ddp.solve([], [], 300)
print done
# +
# %%capture
# %matplotlib inline
# Create animation
anim = animateCartpole(xs)
# -
# HTML(anim.to_jshtml())
HTML(anim.to_html5_video())
| examples/notebooks/cartpole_swing_up.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// ## Loading Data into Tables - HDFS
//
// Let us understand how we can load data from HDFS location into Spark Metastore table.
// + tags=["remove-cell"]
// %%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/QLl0xvnQsTg?rel=0&controls=1&showinfo=0" frameborder="0" allowfullscreen></iframe>
// -
// Let us start spark context for this Notebook so that we can execute the code provided. You can sign up for our [10 node state of the art cluster/labs](https://labs.itversity.com/plans) to learn Spark SQL using our unique integrated LMS.
val username = System.getProperty("user.name")
// +
import org.apache.spark.sql.SparkSession
val username = System.getProperty("user.name")
val spark = SparkSession.
builder.
config("spark.ui.port", "0").
config("spark.sql.warehouse.dir", s"/user/${username}/warehouse").
enableHiveSupport.
appName(s"${username} | Spark SQL - Managing Tables - Basic DDL and DML").
master("yarn").
getOrCreate
// -
// If you are going to use CLIs, you can use Spark SQL using one of the 3 approaches.
//
// **Using Spark SQL**
//
// ```
// spark2-sql \
// --master yarn \
// --conf spark.ui.port=0 \
// --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
// ```
//
// **Using Scala**
//
// ```
// spark2-shell \
// --master yarn \
// --conf spark.ui.port=0 \
// --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
// ```
//
// **Using Pyspark**
//
// ```
// pyspark2 \
// --master yarn \
// --conf spark.ui.port=0 \
// --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
// ```
// * We can use load command with out **LOCAL** to get data from HDFS location into Spark Metastore Table.
// * User running load command from HDFS location need to have write permissions on the source location as data will be moved (deleted on source and copied to Spark Metastore table)
// * Make sure user have write permissions on the source location.
// * First we need to copy the data into HDFS location where user have write permissions.
import sys.process._
val username = System.getProperty("user.name")
s"hadoop fs -rm -R /user/${username}/retail_db/orders" !
s"hadoop fs -mkdir /user/${username}/retail_db" !
s"hadoop fs -put -f /data/retail_db/orders /user/${username}/retail_db" !
s"hadoop fs -ls /user/${username}/retail_db/orders" !
// * Here is the script which will truncate the table and then load the data from HDFS location to Hive table.
// + language="sql"
//
// USE itversity_retail
// + language="sql"
//
// TRUNCATE TABLE orders
// + language="sql"
//
// LOAD DATA INPATH '/user/itversity/retail_db/orders'
// INTO TABLE orders
// -
s"hadoop fs -ls /user/${username}/warehouse/${username}_retail.db/orders" !
s"hadoop fs -ls /user/${username}/retail_db/orders" !
// + language="sql"
//
// SELECT * FROM orders LIMIT 10
// + language="sql"
//
// SELECT count(1) FROM orders
// -
// * Using Spark SQL with Python or Scala
spark.sql("USE itversity_retail")
spark.sql("TRUNCATE TABLE orders")
spark.sql("""
LOAD DATA INPATH '/user/itversity/retail_db/orders'
INTO TABLE orders""")
s"hadoop fs -ls /user/${username}/retail_db/orders" !
spark.sql("SELECT * FROM orders LIMIT 10")
spark.sql("SELECT count(1) FROM orders")
// * If you look at **/user/training/retail_db** orders directory would have been deleted.
// * Move is much faster compared to copying the files by moving blocks around, hence Hive load command from HDFS location will always try to move files.
| 04_basic_ddl_and_dml/06_loading_data_into_tables_hdfs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('data.csv')
df['hN'].mean()
df
# ได้ชูไม้
# JUMP OF N
df[(df['hN']/df['hN'].mean()) >= 1.22]
# JUMP OF S
len(df[(df['hS']/df['hS'].mean()) >= 1.22])
# DOWN OF N
len(df[(df['hN']/df['hN'].mean()) <= 0.63])
# DOWN OF S
len(df[(df['hS']/df['hS'].mean()) <= 0.76])
| .ipynb_checkpoints/ExploreData-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Basic Subplot Demo
#
#
# Demo with two subplots.
# For more options, see
# :doc:`/gallery/subplots_axes_and_figures/subplots_demo`
#
# +
import numpy as np
import matplotlib.pyplot as plt
# Data for plotting
x1 = np.linspace(0.0, 5.0)
x2 = np.linspace(0.0, 2.0)
y1 = np.cos(2 * np.pi * x1) * np.exp(-x1)
y2 = np.cos(2 * np.pi * x2)
# Create two subplots sharing y axis
fig, (ax1, ax2) = plt.subplots(2, sharey=True)
ax1.plot(x1, y1, 'ko-')
ax1.set(title='A tale of 2 subplots', ylabel='Damped oscillation')
ax2.plot(x2, y2, 'r.-')
ax2.set(xlabel='time (s)', ylabel='Undamped')
plt.show()
| matplotlib/gallery_jupyter/subplots_axes_and_figures/subplot_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook-7: Lists
# ### Notebook Content
#
# In previous code camp notebooks we looked at numeric (**integers** and **floats**) and textual (**strings**) data, but it's probably been quite difficult to imagine how you'd assemble these simple data into something useful. The new data types (**lists** and **dictionaries**) will _begin_ to show you how that can happen, and they will allow you to 'express' much more complex concepts with ease.
# ### What's the Difference?
#
# Up until now, our variables have only held _one_ things: a number, or a string.
#
# In other words, we've had things like, `myNumber = 5` or `myString = "Hello world!"`. And that's it.
#
# Now, with **lists** and **dictionaries** we can store _multiple_ things: several numbers, several strings, or some mish-mash of both. That is, lists and dictionaries are data _structures_ that can contain multiple data _types_.
#
# Here's a brief summary of these data structures, highlighting their main difference:
# - A **list** is an *ordered* collection of 'items' (numbers, strings, etc.). You can ask for the first item in a list, the 3rd, or the 1,000th, it doesn't really matter because your list has an order and it keeps that order.
# - A **dictionary** is an *unordered* collection of 'items' (numbers, strings, etc.). You access items in a dictionary in a similar way to how you access a real dictionary: you have a 'key' (i.e. the word for which you want the definition) and you use this to look up the 'value' (i.e. the definition of the word).
#
# There's obviously a lot more to lists and dictionaries than this, but it's a good starting point.
#
# Let's start with **lists** in this notebook and we'll continue with **dictionaries** in the following.
#
# You can go back to our short video that talks about lists:
#
# [](https://youtu.be/UDNtW3sy-og?t=452)
# ### In this Notebook
#
# - Indexing & Slicing
# - List operations
# - Addition and Mulitplication
# - You're (not) in the list!
# - List functions
# - insert
# - append
# - index
# - len
# ----
# # Lists
#
# So a **list** is an ordered collection of items that we access by position (what Python calls the **index**) within that collection. So the first item in a list is always the first item in the list. End of (well, sort of... more on this later).
#
# Because lists contain multiple data items, we create and use them differently from the simple (single item) variables we've seen so far. You can always spot a list because it is a series of items separated by commas and grouped together between a pair of square brackets ( *[A, B, C, ..., n]* ).
#
# Here is a list of 4 items assigned to a variable called `myList`:
myList = [1,2,4,5]
print(myList)
# Lists are pretty versatile: they don't really care what kind of items you ask them to store. So you can use a list to hold items of all the other data types that we have seen so far!
#
# Below we assign a new list to a variable called `myList` and then print it out so that you can see that it 'holds' all of these different types of data:
myList = ['hi I am', 2312, 'mixing', 6.6, 90, 'strings, integers and floats']
print(myList)
# We don't want to get too technical here, but it's important to note one thing: `myList` is still just **one** thing – a list – and not six things (`'hi I am', 2312, 'mixing', 6.6, 90, 'strings, integers and floats'`). So we can only assign one list to a variable called `myList`. It's like the difference between a person and a crowd: a crowd is one thing that holds many people inside it...
#
# ## Indexing
#
# To access an item in a list you use an **index**. This is a new and important concept and it's a term that we're going to use a _lot_ in the future, so try to really get this term into your head before moving forward in the notebook.
#
# ### Accessing Elements in a List using Indexes
#
# The index is just the location of the item in the list that you want to access. So let's say that we want to fetch the second item, we access it via the *index notation* like so:
myList = ['hi', 2312, 'mixing', 6.6, 90, 'strings, integers and floats' ]
print(myList[1])
# See what has happened there? We have:
#
# 1. Assigned a list with 6 elements to the variable `myList`.
# 2. Accessed the second element in the list using that element's _index_ between a pair of square brackets (next to the list's name).
#
# Wait a sec – didn't we say *second* element? Why then is the index `1`???
#
# ### Zero-Indexing
#
# Good catch! That's because list indexes are *zero-based*: this is a fancy way to say that the count starts from 0 instead of that from 1. So the first element has index 0, and the last element has index _n-1_ (i.e. the count of the number of items in the list [_n_] minus one). Zero indexing is a bit how like the ground floor in the UK is often shown as floor 0 in a lift.
#
# To recap:
myNewList = ['first', 'second', 'third']
print("The first element is: " + myNewList[0])
print("The third element is: " + myNewList[2])
# ### Negative Indexing
#
# Since programmers are lazy, they also have a short-cut for accessing the _end_ of a list. Since positive numbers count from the start of a list, negative numbers count from the end:
print(myNewList[-1])
print(myNewList[-2])
# The _last_ element has index `-1`. So you count forwards from `0`, but backwards from `-1`.
#
# You can remember it this way: the last item in the list is at _n-1_ (where _n_ is the number of items in the list), so `...[-1]` is a sensible way to access the last item.
# #### A challenge for you!
#
# Edit the code so that it prints the 'second' element in the list
# + solution2="hidden" solution2_first=true
print("The second element is :" + myNewList[???])
# + solution2="hidden"
print("The second element is :" + myNewList[1])
# -
# ### Index Out of Range
#
# What happens when you try to access an element that doesn't exist?
#
# We know that `myList` has 3 elements, so what if we try to access the 200th element in the list? In that case Python, as usual, will inform us of the problem using an *error message* pointing to the problem:
print(myNewList[200])
# #### A challenge for you!
# + [markdown] solution2="hidden" solution2_first=true
# Do you remember the past lesson on *syntax errors* and *exceptions*? What is the error message displayed in the code above? Is it an *exception* or a *syntax error*? Can you find the explanation for what's going in the [Official Documentation](https://www.google.ie/url?sa=t&rct=j&q=&esrc=s&source=web&cd=3&ved=0ahUKEwiN3s-0qr7OAhVGIcAKHYBLAE4QFggoMAI&url=https%3A%2F%2Fdocs.python.org%2F2%2Ftutorial%2Ferrors.html&usg=AFQjCNG6q1juN8ZVXOEqOYWxE18Cv5X_qw&sig2=o92WLjkV1PNNfgpW1w9n0g&cad=rja)?
# + [markdown] solution2="hidden"
# Even if a statement or expression is syntactically correct, it may cause an error when an attempt is made to execute it. Errors detected during execution are called ***exceptions*** and are not unconditionally fatal. The last line of the error message indicates what happened.
#
# You can see from the last line to you have an error related to your index - where your list index is out of range.
# -
# ## A String is a List?
#
# Even if you didn't realise it, you have already been working with lists in a _sense_ because *strings* are _basically_ lists! Think about it this way: strings are an ordered sequence of characters because 'hello' and 'olhel' are very different words! It turns out that characters in a string can be accessed the same way we'd access a generic list.
myString = "ABCDEF"
print(myString[0])
print(myString[-1])
# ## Slicing
#
# If you want to access more than one item at a time, then you can specify a _range_ using two index values instead of just one.
#
# If you provide two numbers, Python will assume you are indicating the start and end of a _group_ of items. This operation is called *list slicing*, but keep in mind that indexes start from 0!
#
# _Note_: remember too that the error above when we tried to get the 200th element was _index out of range_. So 'range' is how Python talks about more than one list element.
shortSentence = "Now I'll just print THIS word, between the 20th and the 25th character: "
print(shortSentence[20:25])
# #### A challenge for you!
#
# Using the previous code as a guide, edit the code below so that it prints from the second to the fourth (inclusive) characters from the string:
# + solution2="shown" solution2_first=true
shortSentence2 = "A12B34c7.0"
print(shortSentence2[???:???])
# + solution2="shown"
shortSentence2 = "A12B34c7.0"
print(shortSentence2[1:4])
# -
# To print the entirety of a list from any starting position onwards, just drop the second index value while leaving the `:` in your code:
stringToPrint = "I will print from HERE onwards"
print("Starting from the 17th position: " + stringToPrint[17:])
# Notice that there are _two_ spaces between "position:" and "HERE" in the printout above? That's because the 17th character is a space. Let's make it a little more obvious:
print("Starting from the 17th position: '" + stringToPrint[17:] + "'")
# Got it?
#
# #### A challenge for you!
#
# Now, combining what we've seen above, how do you think you would print everything _up to the eighth character from the end_ (which is the space between "HERE" and "onwards")?
#
# You'll need to combine:
# 1. Negative indexing
# 2. List slicing
#
# There are two ways to do it, one way uses only one number, the other uses two. Both are correct. Why don't you try to figure them both out? For 'Way 2' below the `???` is a placeholder for a full slicing operation since if we gave you more of a hint it would make it too obvious.
# + solution2="hidden" solution2_first=true
print("Up to the 18th position (Way 1): '" + stringToPrint[???:???] + "'")
print("Up to the 18th position (Way 2): '" + ??? + "'")
# + solution2="hidden"
print("Up to the 18th position (Way 1): '" + stringToPrint[-8:] + "'")
print("Up to the 18th position (Way 2): '" + stringToPrint[22:30] + "'")
# -
# Strings have also plenty of methods that might prove to be quite useful in the future; for a fuller overview check out [this reference](https://en.wikibooks.org/wiki/Python_Programming/Variables_and_Strings).
#
# ## List operations
#
# So far, we've only created a list, but just like a real to-do list, most lists don't usually stay the same throughout the day (or the execution of your application). Their _real_ value comes when we start to change them: adding and removing items, updating an existing item, concatenating several lists (i.e. sticking them together), etc.
#
# ### Replacing an item
#
# Here's how we replace an item in a list:
# +
myNewList = ['first', 'second', 'third']
print(myNewList)
# This replaces the item in the 2nd position
myNewList[1] = 'new element'
print(myNewList)
# -
# This shouldn't surprise you too much: it's just an assignment (via "=") after all!
#
# So if you see `list[1]` on the right side of the assignment (the "=") then we are _reading from_ a list, but if you see `list[1]` on the left side of the assignment then we are _writing to_ a list.
#
# Here's an example for a (small) party at a friend's place:
# +
theParty = ['Bob','Doug','Louise','Nancy','Sarah','Jane']
print(theParty)
theParty[1] = 'Phil' # Doug is replaced at the party by Phil
print(theParty)
theParty[0] = theParty[1]
print(theParty) # Phil is an evil genius and manages to also replace Doug with a Phil clone
# -
# Got it?
#
# ### Addition and Multiplication
#
# You can also operate on entire lists at one time, rather than just on their elements individually. For instance, given two lists you might want to add them together like so:
# +
britishProgrammers = ["Babbage", "Lovelace"]
nonBritishProgrammers = ["Torvald", "Knuth"]
famousProgrammers = britishProgrammers + nonBritishProgrammers
print(famousProgrammers)
# -
# You can even multiply them, although in this particular instance it is kind of pointless:
print(britishProgrammers * 2)
# #### A challenge for you!
#
# Correct the syntax in the following code to properly define a new list:
# + solution2="hidden" solution2_first=true
otherNonBritishProgrammers = ["Wozniak" ??? "<NAME>"]
# + solution2="hidden"
otherNonBritishProgrammers = ["Wozniak","<NAME>"]
otherNonBritishProgrammers
# -
# Edit the following code to print out all the non british programmers:
# + solution2="hidden" solution2_first=true
print nonBritishProgrammers ??? otherNonBritishProgrammers
# + solution2="hidden"
print(nonBritishProgrammers + otherNonBritishProgrammers)
# -
# ### You're (not) in the list!
#
# Ever stood outside a club or event and been told: "You're not on/in the list"? Well, Python is like that too. In fact, Python tries as hard as possible to be _like_ English – this isn't by accident, it's by design – and once you've done a _bit_ of programming in Python you can start to guess how to do something by thinking about how you might say it in English.
#
# So if you want to check if an item exists in a list you can use the **in** operator:
#
# ```python
# element in list
# ```
#
# The **in** operator will return `True` if the item is present, and `False` otherwise. This is a new data type (called a Boolean) that we'll see a lot more of in two notebooks' time.
# +
print ('Lovelace' in britishProgrammers)
print ('Lovelace' in nonBritishProgrammers)
letters = ['a','b','c','d','e','f','g','h','i']
print ('e' in letters)
print ('z' in letters)
# -
# _Note:_ You might also have spotted that this time there are parentheses ("(...)") after `print`. In general, as you become more experienced you'll _always_ want to put parentheses after a `print` statement (because that's how Python3 works) but the intricacies of _why_ this is the case are a bit out of the scope of an introductory set of lessons.
#
# Anyway, if you want to check if an item does not exist in a list then you can use the **not in ** operator. Let's go back to our party:
print(theParty)
print('Bob' not in theParty)
print('Jane' not in theParty)
# So here, that 'Boolean' gives us `True` on the first `not in` because it's "true that Bob isn't at the party" and `False` on the second one because it's "not true that Jane isn't at the party"! Double-negatives aren't supposed to exist in English, but they certainly do in programming!
# #### A challenge for you!
#
# Complete the missing bits of the following code so that we print out Ada Lovelace, her full name:
# + solution2="hidden" solution2_first=true
firstProgrammerSurnames = ["Babbage", "Lovelace"]
firstProgrammerNames = ["Charles", "Ada"]
firstProgrammerSurnames[1] = firstProgrammerNames[1] + " " + firstProgrammerSurnames[1]
print("Lady "+ ???[1] +" is considered to be the first programmer.")
# + solution2="hidden"
firstProgrammerSurnames = ["Babbage", "Lovelace"]
firstProgrammerNames = ["Charles", "Ada"]
firstProgrammerSurnames[1] = firstProgrammerNames[1] + " " + firstProgrammerSurnames[1]
print("Lady "+ firstProgrammerSurnames[1] +" is considered to be the first programmer.")
# -
# _Note_ : Actually, Lady Ada Lovelace is a [fascinating person](https://en.wikipedia.org/wiki/Ada_Lovelace): she isn't just the first female programmer, she was the first programmer full-stop. For many years <NAME> got all the credit for inventing computers simply because he was a guy and designed the clever mechanical adding machine. However, lately, we've realised that Ada was the one who actually saw that Babbage hadn't just invented a better abacus, he'd invented a general-purpose computing device!
#
# She was so far ahead of her time that the code she wrote down couldn't even run on Babbage's 'simple' (i.e. remarkably complex for the time) computer, but it is now recognised as the first computer algorithm. As a result, there is now a day in her honour every year that is celebrated around the world at places like Google and Facebook, as well as at King's and MIT, because we want to recognise the fundamental contribution to computing made by women programmers.
#
# This contribution was long overlooked by the men who thought that the hard part was the machine, not the programming. Rather entertainingly (given the kudos attached to the people who created applications like Google and Facebook), most men thought that programming was just like typing and was, therefore 'women's work'. So why not take a few minutes to [recognise the important contributions of women to the history of computing](http://www.npr.org/blogs/alltechconsidered/2014/10/06/345799830/the-forgotten-female-programmers-who-created-modern-tech).
#
# 
# ## Extending Lists
#
# We've already seen that we can combine two lists using the `+` operator, but if you wanted to constantly add new items to your list then having to do something like this would be annoying:
# ```python
# myList = [] # An empty list
#
# myList = myList + ['New item']
# myList = myList + ['Another new item']
# myList = myList + ['And another one!']
#
# print(myList)
# ```
# Not just annoying, but also hard to read! As with most things, because programmers are lazy there's an easier way to write this in Python:
# ```python
# myList = [] # An empty list
#
# myList.append('New item')
# myList.append('Another new item')
# myList.append('And another one!')
#
# print(myList)
# ```
#
# Why don't you try typing it all in the coding area below? You'll get the same answer either way, but one is faster to write and easier to read!
# +
#your code here
# -
# Appending to a list using `append(...)` is actually using something called a _function_. We've not really seen this concept before and we're not going to go into it in enormous detail here (there's a whole other notebook to introduce you to functions). The things that you need to notice at _this_ point in time are:
#
# 1. That square brackets ('[' and ']') are used for list indexing.
# 2. That parentheses ('(' and ')') are (normally) used for function calls.
#
# The best thing about functions is that they are like little packages of code that can do a whole bunch of things at once (e.g. add an item to a list by modifying the list directly), but you only need to remember to write `append(...)`.
#
# What did I mean just now by 'modifying the list directly'?
#
# Notice that in the first example above we had to write:
# ```python
# myList = myList + ['New item']
# ```
# because we had to write the result of concatenating the two lists together back to the variable. The list isn't really growing, we're just overwriting what was _already_ in `myList` with the results of the list addition operation. What do you think you would get if you wrote the following code:
# ```python
# myList = [] # An empty list
#
# myList + ['New item']
# myList + ['Another new item']
# myList + ['And another one!']
#
# print(myList)
# ```
# If you aren't sure, why don't you try typing this into the coding area above and try to figure out why the answer is: ''.
#
# Meanwhile, in the second example we could just write:
# ```python
# myList.append('New item')
# ```
# and the change was made to `myList` directly! So this is easier to read _and_ it is more like what we'd expect to happen: the list grows without us having to overwrite the old variable.
# ## Other List Functions
#
# `append()` is a function, and there are many other functions that can be applied to lists such as: `len`, `insert` and `index`. You tell Python to *execute a function* (sometimes also termed *calling a function*) by writing the function's name, followed by a set of parentheses. The parentheses will contain any optional inputs necessary for the function to do its job. Appending to a list wouldn't be very useful if you couldn't tell Python _what_ to append in the first place!
#
# The `len` function is also a good example of this:
# ```python
# len(theParty)
# ```
# Here, the function `len` (lazy short-hand for _length_ ) is _passed_ `theParty` list as an input in order to do its magic.
#
# The functions `append`, `insert` and `index` work a _little_ bit differently. They have to be _called_ using `theParty` list. We're at risk of joining Alice down the rabbit-hole here, so let's just leave it at: the second set of functions are of a particular _type_ known as *methods* of the *list class*. We'll stop there.
#
# <img src="https://www.washingtonpost.com/blogs/answer-sheet/files/2013/01/alice-falling-down-rabbit-hole1.jpg" width="250" />
# In order to use a _list method_ you always need to 'prepend' (i.e. lead with) the name of list you want to work with, like so:
#
# ```python
# theParty.append("<NAME>")
# theParty.insert(2, "Anastasia")
# theParty.index("Sarah")
# ```
#
# The idea here is that methods are associated with specific types of things such as lists, whereas generic functions are kind of free-floating. Think about it this way: you can't append something to a number, because that _makes no sense_. Appending is something that only makes sense in the context of a list. In contrast, something like `len` works on several different types of variables: lists and string both!
# ### Append
#
# Reminder: here's appending...
britishProgrammers = ['Lovelace']
britishProgrammers.append("Turing")
print(britishProgrammers)
# ### Insert
#
# That's cool, but as you noticed `append` only ever inserts the new item as the last element in the list. What if you want it to go somewhere else?
#
# With `insert` you can also specify a position
print(nonBritishProgrammers)
nonBritishProgrammers.insert(1, "Swartz")
print(nonBritishProgrammers)
# ### Index
#
# Lastly, with the `index` method you can easily ask Python to find the position (index) of a given item:
# Say you want to know in where "Knuth" is
# in the list of non-British programmers...
print(nonBritishProgrammers.index("Knuth"))
# #### A challenge for you!
# Add the famous [Grace Hopper](https://en.wikipedia.org/wiki/Grace_Hopper) (inventress of the first compiler!) to the list of British programmers, and then print her index position:
# + solution2="hidden" solution2_first=true
nonBritishProgrammers.???("Hopper")
print(nonBritishProgrammers.???("Hopper"))
# + solution2="hidden"
nonBritishProgrammers.append("Hopper")
# let's check
print(nonBritishProgrammers)
print(nonBritishProgrammers.index("Hopper"))
# -
# ### Length
#
# Cool, so those were some of the *methods you can invoke on* a list. Let's focus now on some *functions* that take lists as an _input_.
#
# With the function `len` you can immediately know the `len`-gth of a given list:
print( len(britishProgrammers) )
length_of_list = len(nonBritishProgrammers)
print("There are " + str(length_of_list) + " elements in the list of non-British Programmers")
# + [markdown] solution2="hidden" solution2_first=true
# Did you see the `str(length_of_list)`? There's another function! We didn't draw attention to it before, we just told you to use `str(5.0)` to convert the float `5.0` to the string `"5.0"`. We can tell it's a function because it uses the format `functionName(...some input...)`. So the function name is `str` (as in _convert to string_) and the input is a number (in this case it's the length of the list `nonBritishProgrammers`). So now we can easily convert between different types of data: we took an integer (you can check this by adding the following line to the code above:
# ```python
# print length_of_list + 1
# ```
# And then print it out as a string. So `length_of_list` is a number, and by calling `str(length_of_list)` we changed it to a string that we could print out. Given that programmers are lazy, can you guess how you'd convert the string "3" to the _integer_ 3?
# + solution2="hidden"
print(int(length_of_list) + 1)
# + solution2="hidden"
three = "3"
# Let's check the data type
type(three)
# + solution2="hidden"
# Change to Integer
three = int(three)
type(three)
# -
# #### A challenge for you!
#
# Complete the missing bits of the following code:
# + solution2="hidden" solution2_first=true
length_of_brits = ???(britishProgrammers)
print "There are " ??? " British programmers."
# + solution2="hidden"
length_of_brits = len(britishProgrammers)
print("There are " + str(length_of_brits) + " British programmers.")
# -
# To check if the output of `range()` is a list we can use the `type()` function:
# # Code (Applied Geo-example)
# Let's have a little play with some geographical coordinates. Based on what we've just done in this notebook, what do you think would be a good data type for storing lat/long coordinates?
#
# Go on, I'll bet you can't guess!
#
# Let's use what we know to jump to a particular point on the planet...
# +
# We'll see more of this package import
# later... for now, just know that it
# provides access to a function called IFrame
from IPython.display import IFrame
# We want to view an OpenStreetMap map
siteName = "http://www.openlinkmap.org/small.php"
# Specify the location and zoom level
latitude = 63.6314
longitude = -19.6083
zoomLevel = 10
mapURL = ['?lat=', str(latitude), '&lon=', str(longitude), '&zoom=', str(zoomLevel)]
# Show the ULR
print(siteName + ''.join(mapURL))
# And now show us an inline map!
IFrame(siteName + ''.join(mapURL), width='100%', height=400)
# -
# And now let's try somewhere closer to home. Try using the `KCLMapCoordinates` list below (which tells you the x/y and zoom-level) in combination with the code above. I'd suggest copying what you need from above, pasting it into the code below, and then using what you know about accessing list information so that the map shows the location of King's College London...
# + solution2="shown" solution2_first=true
# King's College coordinates
# What format are they in? Does it seem appropriate?
KCLMapCoordinates = [51.51130657591914, -0.11596798896789551, 15]
# + solution2="shown"
# We want to view an OpenStreetMap map
siteName = "http://www.openlinkmap.org/small.php"
# Specify the location and zoom level
latitude = KCLMapCoordinates[0]
longitude = KCLMapCoordinates[1]
zoomLevel = 15
mapURL = ['?lat=', str(latitude), '&lon=', str(longitude), '&zoom=', str(zoomLevel)]
# Show the ULR
print(siteName + ''.join(mapURL))
# And now show us an inline map!
IFrame(siteName + ''.join(mapURL), width='100%', height=400)
# -
#
# ### Further references:
#
# General list or resources
# - [Awesome list of resources](https://github.com/vinta/awesome-python)
# - [Python Docs](https://docs.python.org/2.7/tutorial/introduction.html)
# - [HitchHiker's guide to Python](http://docs.python-guide.org/en/latest/intro/learning/)
# - [Python for Informatics](http://www.pythonlearn.com/book_007.pdf)
# - [Learn Python the Hard Way - Lists](http://learnpythonthehardway.org/book/ex32.html)
# - [Learn Python the Hard Way - Dictionaries](http://learnpythonthehardway.org/book/ex39.html)
# - [CodeAcademy](https://www.codecademy.com/courses/python-beginner-en-pwmb1/0/1)
#
#
# ### Credits!
#
# #### Contributors:
# The following individuals have contributed to these teaching materials:
# - [<NAME>](https://github.com/jamesdamillington)
# - [<NAME>](https://github.com/jreades)
# - [<NAME>](https://github.com/miccferr)
# - [<NAME>](https://github.com/zarashabrina)
#
# #### License
# The content and structure of this teaching project itself is licensed under the [Creative Commons Attribution-NonCommercial-ShareAlike 4.0 license](https://creativecommons.org/licenses/by-nc-sa/4.0/), and the contributing source code is licensed under [The MIT License](https://opensource.org/licenses/mit-license.php).
#
# #### Acknowledgements:
# Supported by the [Royal Geographical Society](https://www.rgs.org/HomePage.htm) (with the Institute of British Geographers) with a Ray Y Gildea Jr Award.
#
# #### Potential Dependencies:
# This notebook may depend on the following libraries: None
| notebook-07-lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import jhu2016_labs.plotting as plotting
import jhu2016_labs.models as models
# %matplotlib inline
# -
# ## Training data
#
# In real scenario, we observed a set of values, often called features, and we try to learn some model that can "explain" the observed features. Generally, we do not know the true distribution of the features. For this work we will simulate a set of single dimensional features sampled from a GMM. We will call this GMM the "true model" and in this notebook we will see different strategies to approximate the true model from the simulated training data.
true_model = models.GMM([-4, 0, 2], [1, 2, .7], [0.2, 0.2, 0.6])
X = true_model.sampleData(50)
Y = np.zeros_like(X) # The random variable has a single dimension !
x_min = -8
x_max = 7
y_min = -0.01
y_max = .4
fig, ax = plotting.plotGMM(true_model, x_min=x_min, x_max=x_max, y_min=y_min, y_max=y_max,
label='True model')
ax.plot(X, Y, '+', color='b', label='data')
ax.legend(loc='upper left')
# ## Gaussian density
#
# We can first try to model the observed data with a Gaussian density function. The Gaussian density function is defined as:
# $$
# \DeclareMathOperator{\Norm}{\mathcal{N}}
# \DeclareMathOperator{\Gam}{Gam}
# \DeclareMathOperator{\e}{exp}
# p(x \mid \mu, \sigma^2) = \Norm(x \mid \mu, \sigma^2) = \frac{1}{\sqrt{2\pi\sigma^2}} \e\{ \frac{-(x - \mu)^2}{2\sigma^2} \}
# $$
# * $\mu$ is the **mean** of the Gaussian density
# * $\sigma^2$ is the **variance** of the Gaussian density
#
#
# ##### TODO
# Try to play with the mean and the variance of the Gaussian. How each parameter change the density ?
gaussian = models.Gaussian(0., 1.) # <- 1st parameter is mean, second variance0
fig, ax = plotting.plotGaussian(gaussian)
# Alternatively, the Gaussian density function can be parameterized with a precision $\lambda$ which is the inverse of the variance:
# $$
# \DeclareMathOperator{\e}{exp}
# p(x | \mu, \lambda) = \Norm(x \mid \mu, \lambda^{-1}) = \frac{\sqrt{\lambda}}{\sqrt{2\pi}} \e\{ \frac{-\lambda(x - \mu)^2}{2} \}
# $$
# * $\mu$ is the **mean** of the Gaussian density
# * $\lambda = \frac{1}{\sigma^2}$ is the **precision** of the Gaussian density
#
# This alternative parameterization will prove to be useful when dealing with Bayesian inference for the Gaussian/GMM density.
# ### Maximum Likelihood estimation
#
# We will first try to model our data with a simple Gaussian. We need to find the parameters (mean and variance) that fit the best the data.
#
# $$
# \begin{align}
# \mu &= \frac{1}{N} \sum_i x_i \\
# \sigma^2 &= \frac{1}{N} \sum_i x_i^2
# \end{align}
# $$
#
# ##### TODO
# From the above equations, implement the maximum-likelihood solution for the Gaussian. Does the Gaussian seems to be a good model for the true density ?
# +
##########################
# Write your code here.
mean_ml =
var_ml =
##########################
# Check your solutions uncomment the following code
test_gaussian_ml = models.Gaussian.maximumLikelihood(X)
assert np.isclose(test_gaussian_ml.mean, mean_ml), 'incorrect mean'
assert np.isclose(test_gaussian_ml.var, var_ml), 'incorrect variance'
print('correct solution !')
gaussian_ml = models.Gaussian(mean_ml, var_ml)
llh = gaussian_ml.logLikelihood(X)
print('log-likelihood:', llh)
y_min = -.01
y_max = .4
fig, ax = plotting.plotGMM(true_model, y_min=y_min, y_max=y_max, label='True model')
plotting.plotGaussian(gaussian_ml, fig=fig, ax=ax, color='red', label='Gaussian ML')
ax.plot(X, Y, '+', color='b', label='data')
ax.legend(loc='upper left')
# -
# ### Bayesian Inference
#
# We can work within the Bayesian framework with the Gaussian density by putting a prior over the mean and precision. A common an convenient choice of prior for the Gaussian is the **Normal-Gamma** prior:
#
# $$
# p(\mu, \lambda \mid m_0, \kappa_0, a_0, b_0) = \Norm(\mu \mid m_0, (\kappa_0 \lambda)^{-1}) \Gam(\lambda \mid a_0, b_0)
# $$
#
# where:
#
# $$
# \Gam(\lambda \mid a_0, b_0) = \frac{1}{\Gamma(a_0)} b_0^{a_0} \lambda^{a_0 - 1} \e \{ -b_0 \lambda\}
# $$
#
# $m_0$, $\kappa_0$, $a_0$ and $b_0$ are called **hyper-parameters**. They are the parameters of the prior distribution.
#
# ##### TODO
# Play with the parameters of the Normal-Gamma. How each parameter affects the density ?
ng_prior = models.NormalGamma(0, 2, 5, 6)
x_min =-2.
x_max = 2.
y_min = 0.
y_max = 2
plotting.plotNormalGamma(ng_prior, x_min=x_min, x_max=x_max, y_min=y_min, y_max=y_max)
# Because the Normal-Gamma is the **conjugate prior** of the Normal density, the posterior distribution $p(\mu, \lambda \mid \mathbf{x})$ has a closed form solution:
#
# $$
# p(\mu, \lambda \mid \mathbf{x}) = \Norm(\mu \mid m_n, (\kappa_n \lambda)^{-1}) \Gam(\lambda \mid a_n, b_n)
# $$
#
# where:
#
# $$
# \begin{align}
# m_n &= \frac{\kappa_0 m_0 + N \bar{x}} {\kappa_0 + N} \\
# \kappa_n &= \kappa_0 + N \\
# a_n &= a_0 + \frac{N}{2} \\
# b_n &= b_0 + \frac{N}{2} ( s + \frac{\kappa_0 (\bar{x} - m_0)^2}{\kappa_0 + N} ) \\
# \bar{x} &= \frac{1}{N} \sum_i x_i \\
# s &= \frac{1}{N} \sum_i (x_i - \bar{x})^2
# \end{align}
# $$
#
# $N$ is the total number of point in the the training data and $m_n$, $\kappa_n$, $a_n$ and $b_n$ are the parameters of the posterior. Note that they are different from the hyper-parameters !!
#
# ##### TODO
# Compute the posterior distribution with 1, 5, 10, 20 and 50 data points from the training data. What do you observe ?
ng_prior = models.NormalGamma(0, 2, 5, 6)
ng_posterior = ng_prior.posterior(X[:1]) # <- Change '1' to the number of sample you want to use
x_min =-2
x_max = 2
y_min = 0.
y_max = 2
plotting.plotNormalGamma(ng_posterior, x_min=x_min, x_max=x_max, y_min=y_min, y_max=y_max)
# ### Predictive probability
#
# Now that we have our posterior distibution we can predict the probability of a new data point given the training data:
#
# $$
# p(x' \mid \mathbf{x}) = \int p(x' \mid \theta) p(\theta \mid \mathbf{x}) d \theta
# $$
#
# For the Gaussian with Normal-Gamma prior the marginal predictive distribution is the Student's t distribution:
#
# $$
# \newcommand{\diff}{\mathop{}\!d}
# \DeclareMathOperator{\St}{\mathcal{St}}
# p(x' \mid \mathbf{x}) = \int_{-\infty}^{\infty} p(x|\mu, \lambda) p(\mu, \lambda \mid \mathbf{x}) \diff \mu \diff \lambda = \St(x' \mid \mu_n, \nu, \gamma)
# $$
#
# where:
#
# $$
# \begin{align}
# \nu &= 2a_n \\
# \gamma &= \frac{a_n \kappa_n}{b_n(\kappa_n + 1)} \\
# \St(x' \mid \mu_n, \nu, \gamma) &= \frac{\Gamma(\frac{\nu}{2} + \frac{1}{2})}{\Gamma(\frac{\nu}{2})} \Big( \frac{\gamma}{\pi \nu} \Big)^{\frac{1}{2}} \Big[ 1 + \frac{\gamma (x - \mu_n)^2}{\nu} \Big]^{-\frac{\nu}{2} - \frac{1}{2}}
# \end{align}
# $$
#
# ##### TODO
# Compute the log-likelihood of the predictive distribution derived from a posterior trained with 1, 5, 10, 20 and 50 data points. How does it compare to the log-likelihood of the Gaussian trained with maximum likelihood ?
ng_posterior = ng_prior.posterior(X[:1]) # <- Change '1' by the number of sample you want to use.
predict_pdf = ng_posterior.predictiveDensity()
llh = predict_pdf.logLikelihood(X)
print('log-likelihood:', llh)
# ## Gaussian Mixture Model (GMM)
#
# The Gaussian density is a very simple function, however, in most cases of interest the density we try to model has a complex shape that cannot be expressed with a simple formula. A solution is to assume that our complex density is made of $K$ Gaussian densities. This is called a Gaussian Mixture Model (GMM) and it is defined as:
# $$
# p(x|\boldsymbol{\mu}, \boldsymbol{\lambda}, \boldsymbol{\pi}) = \sum_{k=1}^{K} \pi_k \Norm(x|\mu_k, (\lambda_k)^{-1})
# $$
# * $\boldsymbol{\mu}$ is the vector of $K$ means
# * $\boldsymbol{\lambda}$ is the vector of $K$ precisions
# * $\boldsymbol{\pi}$ is the vector of $K$ weights such that $\sum_{k=1}^K \pi_k = 1$
#
# ##### TODO
# Observe the influence of each parameters on the density and add/remove some components. Does this model suffer of the same drawbacks of the Gaussian density ?
# +
# Change the initial values of the parameters.
# you can add a component by removing a mean, a variance
# and a weight. All arrays shall always have the same number
# of elements ! Be careful, make sure that the sum of weight
# always sum up to one.
means = [-4.0, 0.0, 4.0, 5]
variances = [1.0, 2.0, 1.4, 1]
weights = [0.1, 0.4, 0.2, 0.3]
gmm = models.GMM(means, variances, weights)
y_min = 0.
y_max = .3
plotting.plotGMM(gmm, show_components=True, y_min=y_min, y_max=y_max)
# -
# ## Maximum Likelihood estimation
#
# The GMM parameters can be estimated with the **Expectation-Maximization** (EM) algorithm. The EM algorithm is an iterative algorithm that converges toward a (local) maximum of the log-likelihood of the data given the model. The EM training is as follows:
# * initialize the parameters of the GMM
# * iterate until convergence:
# * Expectation (E-step): compute the probability of the latent variable for each data point
# * Maximization (M-step): update the parameters from the statistics of the E-step.
#
# ##### TODO
# * We have implemented a simple version of the EM algorithm that iterates 10 times. Change the following code to iterates until convergence of the log-likelihood. We will consider that the log-liklihood has converged if $ \log p(\mathbf{x} | \theta^{new}) - \log p(\mathbf{x} | \theta^{old}) \le 0.01$.
# * Run the EM algorithm with different initialization: is the final log-likelihood always the same ? What can you conclude ? Is the log-likelihood greater than the one from the Gaussian estimated with Maximum-Likelihood ?
# +
# Initialization of the model
means = [-4.0, 0.0, 4.0]
variances = [1.0, 1.0, 1.]
weights = [0.3, 0.4, 0.3]
gmm = models.GMM(means, variances, weights)
## EM algorithm
print('initial log-likelihood:', gmm.logLikelihood(X))
############################################
# Change this loop to check the convergence of the EM .
for i in range(10):
# E-step
Z = gmm.EStep(X)
# M-step
gmm.MStep(X, Z)
llh = gmm.logLikelihood(X)
print('log-likelihood:', llh)
##############################################
y_min = -0.01
y_max = .4
fig, ax = plotting.plotGMM(true_model, y_min=y_min, y_max=y_max, label='True model')
plotting.plotGMM(gmm, fig=fig, ax=ax, color='r', label='GMM EM')
ax.plot(X, Y, '+', color='b', label='data')
ax.legend(loc='upper left')
# -
# ## Bayesian GMM
#
# We can also work the Bayesian inference for GMM by putting a prior over the GMM parameters. Let $\Theta$ be the set of parameters of the GMM:
# $$
# \Theta = \{ \boldsymbol{\mu}, \boldsymbol{\lambda}, \boldsymbol{\pi} \}
# $$
#
# The prior over the weights $\boldsymbol{\pi}$ will be a Dirichlet distribution:
#
# $$
# \DeclareMathOperator{\Dir}{Dir}
# p(\boldsymbol{\pi}) = \Dir(\boldsymbol{\pi} \mid \boldsymbol{\alpha}_0)
# $$
#
# and the prior of the mean and precision of the component $k$ of the mixture will be a Normal-Gamma distribution:
#
# $$
# p(\mu_k, \lambda_k) = \Norm(\mu_k \mid m_0, (\kappa_0 \lambda_k)^{-1}) \Gam(\lambda_k \mid a_0, b_0)
# $$
#
# The joint distribution of the data, the latent variables and the parameters can be written as:
#
# $$
# \begin{align}
# p(\mathbf{x}, \mathbf{z}, \Theta) &= p(\mathbf{x}, \mathbf{z} \mid \Theta)p(\Theta) \\
# &= \Bigg[ \prod_{i=0}^{N} p(x_i \mid \boldsymbol{\mu}, \boldsymbol{\lambda}, \boldsymbol{\pi}) \Bigg] \Bigg[ \Dir(\boldsymbol{\pi} \mid \boldsymbol{\alpha}_0) \prod_{k=0}^{K} \Norm(\mu_k \mid m_0, (\kappa_0 \lambda_k)^{-1}) \Gam(\lambda_k \mid a_0, b_0) \Bigg]
# \end{align}
# $$
#
# ### Gibbs Sampling:
#
# The EM algorithm cannot be applied directly to train the Bayesian GMM. Instead, we will use Gibbs Sampling (GS) to learn the posterior distribution of the parameters. Gibbs Sampling is a simple way to sample values from a complex distribution. Let's say that we want to sample values for the random variable $A$ and $B$ conditioned on $C$. Unfortunately, $p(A, B \mid C)$ is too complex to be sampled from directly. Alternatively we can sample in turn $a$ from $p(A | B = b, C)$ and $b$ from $p(B | A=a, C)$. It can be proven that if we keep sampling long enough, the set of $a$ and $b$ will be distributed according to $p(A, B | C)$. This is the Gibbs Sampling algorithm.
#
# NOTE: Gibbs Sampling is not always applicable as we cannot always sample from the conditional distributions (it may be intractable).
#
# In the following we just show the 3 conditional distributions we need for the Gibbs Sampling of the Bayesian GMM.
#
#
# ##### latent variable $z_i$
# $$
# \begin{align}
# p(z_i \mid \mathbf{x}, \Theta) &= p(z_i \mid x_i, \boldsymbol{\pi}, \boldsymbol{\mu}, \boldsymbol{\lambda}) \\
# p(z_i = k \mid x_i, \boldsymbol{\pi}, \boldsymbol{\mu}, \boldsymbol{\lambda}) &= \frac{\pi_k \Norm(x_i \mid \mu_k, \lambda_k)} {\sum_{j=0}^{K} \pi_j \Norm(x_i \mid \mu_j, \lambda_j)}
# \end{align}
# $$
#
# ###### mean and variance
# We define the set of all data point $x_i$ that are assigned to the component $k$ of the mixture as follows:
# $$
# \mathbf{x}_{(k)} = \{ x_i : z_i = k, \forall i \in \{1,... , N \} \}
# $$
# and similarly for the latent variables $\mathbf{z}$:
# $$
# \mathbf{z}_{(k)} = \{ z_i : z_i = k, \forall i \in \{1,... , N \} \}
# $$
#
# $$
# \begin{align}
# p(\mu_k, \lambda_k \mid \mathbf{x}, \mathbf{z}, \Theta_{\smallsetminus \{ \mu_k, \lambda_k \} } ) &= p(\mu_k, \lambda_k \mid \mathbf{x}_{(k)}, \mathbf{z}_{(k)}, \Theta_{\smallsetminus \{ \mu_k, \lambda_k \} } ) \\
# &= \Norm(\mu_k \mid m_{n,k}, (\kappa_{n,k} \lambda_k)^{-1}) \Gam(\lambda_k \mid a_{n,k}, b_{n,k})
# \end{align}
# $$
#
# where:
#
# $$
# \begin{align}
# m_{n,k} &= \frac{\kappa_0 m_0 + N_k \bar{x}_k} {\kappa_0 + N_k} \\
# \kappa_{n,k} &= \kappa_0 + N_k \\
# a_{n,k} &= a_0 + \frac{N_k}{2} \\
# b_{n,k} &= b_0 + \frac{N_k}{2} ( s + \frac{\kappa_0 (\bar{x}_k - m_0)^2}{\kappa_0 + N_k} ) \\
# N_k &= \left\vert \mathbf{x}_{(k)} \right\vert \\
# \bar{x}_k &= \frac{1}{N_k} \sum_{\forall x \in \mathbf{x}_{(k)}} x \\
# s_n &= \frac{1}{N} \sum_{\forall x \in \mathbf{x}_{(k)}} (x_i - \bar{x})^2
# \end{align}
# $$
#
# NOTE: these equations are very similar to the Bayesian Gaussian estimate. However, it remains some difference
#
# ##### weights
#
# $$
# \begin{align}
# p( \boldsymbol{\pi} \mid \mathbf{x}, \mathbf{z}, \Theta_{\smallsetminus \{ \boldsymbol{\pi} \} } ) &= p( \boldsymbol{\pi} \mid \mathbf{z}) \\
# &= \Dir(\boldsymbol{\pi} \mid \boldsymbol{\alpha})
# \end{align}
# $$
# where:
# $$
# \alpha_{n,k} = \alpha_{0,k} + N_k \; ; \; \forall \, k = 1\dots K
# $$
#
# ##### TODO
# * Compare the following implementation of the Gibbs Sampling to the EM algorithm. What are their differences/similarities ?
# * Run the Gibbs Sampling algorithm for 1, 2, 5, 10, 20, ... iterations. How many iterations do we need to get a reasonable estimate of the true density ?
# * Change the hyper-parameters and run the Gibbs Sampling algorithm for 3 iterations. Are the initial sampled GMM very similar ? Run more iterations, does the Gibbs Sampling converges to the same solution regardless of the initialization ?
# +
# hyper-parameters [pi_1, pi_2, ...], m, kappa, a, b
# you may try to change this.
bgmm = models.BayesianGMM([1, 1, 1], 0, 1, 2, 1)
y_min = -0.01
y_max = .4
fig, ax = plotting.plotGMM(true_model, y_min=y_min, y_max=y_max, label='True model')
for i in range(5):
# Sample the latent variables
Z = bgmm.sampleLatentVariables(X)
# Update the parameters
bgmm.sampleMeansVariances(X, Z)
bgmm.sampleWeights(Z)
# Just for plotting, this is not part of the Gibbs Sampling algorithm.
plotting.plotGMM(bgmm.gmm, fig=fig, ax=ax, color='b', lw=.5, label='sampled GMM')
gmm_avg = bgmm.averageGMM()
plotting.plotGMM(gmm_avg, fig=fig, ax=ax, color='r', lw=3, label='avg GMM')
ax.plot(X, Y, '+', color='b', label='data')
handles, labels = ax.get_legend_handles_labels()
labels, ids = np.unique(labels, return_index=True)
handles = [handles[i] for i in ids]
ax.legend(handles, labels, loc='upper left')
# -
| IntroductionToBayesianInference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # ML for Time Series
#
# ## A Unified Framework for Structured Graph Learning via Spectral Constraints
#
# <NAME> - <NAME>
#
# ## Experiments
# ### Setup
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn.datasets as skd
import sklearn.metrics as skm
from learnGraphTopology import *
# ## Denoising a multi-component graph
# First, we use the spectralGraphTopology to denoise laplacian matrices polluted with noise. We generate a 4 components adjacency matrix.
n_class_feats = 5
n_classes = 4
n_feats = n_classes * n_class_feats
prob_intra = 1
prob_extra = 0.3
max_weight_intra = 1
max_weight_extra = 0.3
adj = np.zeros((n_feats, n_feats))
for i in range(n_classes):
i_start = i * n_class_feats
i_end = i_start + n_class_feats
for ii in range(i_start, i_end):
for jj in range(ii+1, i_end):
adj[ii, jj] = np.random.binomial(n=1, p=prob_intra) * np.random.uniform(high=max_weight_intra)
plt.matshow(adj+adj.T)
plt.title("Original matrix")
plt.show()
# We add noise to the matrix
for i in range(n_feats):
for j in range(i+1, n_feats):
adj[i, j] += np.random.binomial(n=1, p=prob_extra) * np.random.uniform(high=max_weight_extra)
adj = adj + adj.T
plt.matshow(adj)
plt.title("Noisy matrix")
plt.show()
# We generate the data according to the noisy laplacian and learn a denoise laplacian corresponding to a 4 components matrix
n_samples = 100 * n_feats
# compute the laplacian and correlation matrices
lap = np.diag(adj.sum(axis=0)) - adj
theta = np.linalg.pinv(lap)
# generate samples
x = np.random.multivariate_normal(np.zeros(n_feats), theta, size=n_samples).T
# learn the laplacian and adjacency matrices
res_denoising = learn_k_component_graph(x, k=4, maxiter=10000)
if res_denoising['convergence']: print("The optimization converged!")
plt.matshow(res_denoising["Adjacency"])
plt.title("recovered adjacency matrix")
plt.show()
print('relative error: ', np.linalg.norm(res_denoising["Laplacian"] - lap)/np.linalg.norm(lap))
print('naive relative error: ', np.linalg.norm(np.linalg.pinv(np.cov(x)) - lap)/np.linalg.norm(lap))
# We show the computed laplacian: the components are well retrieved, but the coefficients have been modified during the process.
# # Denoising a bipartite graph
# Now, we test it on a bipartite graph structure.
# +
halfnodes=20
nodes=2*halfnodes
A = np.zeros([halfnodes*2,halfnodes*2])
A[halfnodes:,:halfnodes] = 1
plt.matshow(A+A.T)
plt.title("original")
plt.show()
A+=np.random.rand(nodes,nodes)*0.1
for k in range(nodes):
A[k][k]=0
A+=A.T
L=A_to_L(A)
plt.matshow(A)
plt.title("noisy")
# -
di=learn_bipartite_graph(np.linalg.pinv(L),maxiter=10000)
plt.matshow(di["Adjacency"])
plt.title("recovered adjacency matrix")
plt.show()
print(di["Adjacency"])
# This time, not only the learned matrix has the desired structure, but it also has similar coefficients to the original one.
# ## Popular synthetic datasets
def dist_mat_exp_kernel(x, sigma=1, eps=1, k=None):
n, d = x.shape
dist_mat = np.zeros((n, n))
for i in range(n):
dist_mat[i] = np.exp(-np.linalg.norm(x[i] - x, axis=1)/eps)
if not k is None:
for i in range(n):
k_th_nearest_dist = np.sort(dist_mat[i])[-k]
dist_mat[i][dist_mat[i] < k_th_nearest_dist] = 0
dist_mat = 0.5*(dist_mat + dist_mat.T)
else:
dist_mat[dist_mat<eps] = 0
return dist_mat
# We load the "two moons" dataset
n_samples = 100
moons_data, moons_labels = skd.make_moons(n_samples=n_samples)
plt.scatter(moons_data.T[0], moons_data.T[1], c=moons_labels)
plt.show()
C_moons = dist_mat_exp_kernel(moons_data, sigma=1, eps=0.1)
# We learn the graph with two components in order to find the clusters.
res_moons = learn_k_component_graph(C_moons, k=2, maxiter=2000) #2000 or more in reality
# Then, we use the first eigenvalue as hte classying metric and compute the AUC score.
score_moons = np.linalg.eigh(res_moons["Laplacian"])[1][:, 1]
print("AUC score of the spectral classifier", skm.roc_auc_score(moons_labels, score_moons))
# We display the adjacency matrix, which is block diagonal and contains the two expected components.
id_moons = np.argsort(moons_labels)
fig = plt.figure(figsize=(20, 20))
plt.matshow(res_moons["Adjacency"][id_moons][:, id_moons], fignum=0)
plt.show()
# Then, we load the "two circles" dataset:
def at_least_k_component(L,k):
la,M=np.linalg.eigh(L)
la[:k] = 0
return M@np.diag(la)@M.T
n_samples = 100
circles_data, circles_labels = skd.make_circles(n_samples=n_samples,noise=0.05,factor=0.5)
plt.scatter(circles_data.T[0], circles_data.T[1], c=circles_labels)
plt.show()
# We learn the clusters with the spectralGraphTopology package
C_circles = dist_mat_exp_kernel(circles_data, k=7,sigma=0.1)
res_circles = learn_k_component_graph(C_circles, k=2, maxiter=10000) #2000 or more in reality
print(res_circles["convergence"])
# +
#res_circles["Laplacian"]=at_least_k_component(res_circles["Laplacian"],k=2)
# -
print(nb_connected_component(res_circles["Laplacian"]))
print(np.array([[0.3,0.02],[0.02,0.3]]).round(1))
np.linalg.eigh(res_circles["Laplacian"])[0][1]
# We display the AUC of the second eigenvalue classifier and the distribution of the corresponding scores according to the true label.
score_circles = np.linalg.eigh(res_circles["Laplacian"])[1][:, 1]
print("AUC score of the spectral classifier", skm.roc_auc_score(circles_labels, score_circles))
plt.scatter(score_circles, circles_labels)
# And here is the learned adjacency matrix:
id_circles = np.argsort(circles_labels)
fig = plt.figure(figsize=(20, 20))
plt.matshow(res_circles["Adjacency"][id_circles][:, id_circles], fignum=0)
plt.show()
A=res_circles["Adjacency"][id_circles][:, id_circles]
print(np.max(A[60:,:40]),np.max(A[:40,60:]))
print(np.max(A))
# Here are unused functions to generate the helix_3D and the three_circles datasets:
def helix_3D(n_samples, h=20, angle=np.pi/2):
labels=np.array([0]*(n_samples//2)+[1]*((n_samples+1)//2))
np.random.shuffle(labels)
ha = np.random.rand(n_samples)*h
return np.array([[np.cos(a+b),np.sin(a+b),a] for a,b in zip(ha,labels)]),labels
def three_circles(n_samples,factor=0.5):
labels=np.zeros([n_samples])
labels[-n_samples//3:]=2
labels[n_samples//3:-n_samples//3]=1
np.random.shuffle(labels)
angles= np.random.rand(n_samples)*2*np.pi
print(angles,labels)
return np.array([[np.cos(a)*factor**b,np.sin(a)*factor**b] for a,b in zip(angles,labels)]),labels
print(three_circles(6))
# ## RNA-Seq dataset
#
# This dataset come from https://archive.ics.uci.edu/ml/datasets/gene+expression+cancer+RNA-Seq#
#
# It represents 801 patients suffering from cancers of 5 different types.
#
# The features of the data are RNA-Seq gene expression levels, and the label is the tumor type.
#
# To make the code work, download the dataset, extract it and save data and labels in the folder ./data/genes/
gene_df = pd.read_csv("data/genes/data.csv")
gene_df = gene_df[gene_df.columns[1:]]
gene_df
gene_labels = pd.read_csv("data/genes/labels.csv")
gene_labels = gene_labels[gene_labels.columns[1:]]
gene_labels
# The goal is to model the distribution with a $k$ component graph, in order to cluster the data into $k$ classes. As there are $k=5$ classes in reality, we try the algorithm with this value.
#
# We also normalize the expression of the genes before computing the covariance matrix.
l=gene_df.values
print(np.max(l))
to_keep = l[]
l-=np.expand_dims(np.mean(l,axis=0),axis=0)
l/=np.expand_dims(np.std(l,axis=0)+10**-9,axis=0)
C = np.cov(l)
print(np.max(l))
res_genes = learn_k_component_graph(C, k=9, maxiter=2000) #2000 or more in reality
# Then, we reorganise the matrix according to their labels to see visually if the components that should be connected are effectively connected.
id_genes = np.argsort(gene_labels.values.squeeze())
fig = plt.figure(figsize=(20, 20))
l=(res_genes["Adjacency"]>10**-3)
plt.matshow(l[id_genes][:, id_genes], fignum=0)
plt.show()
# On this representation of the matrix $\mathbb{1}_{A_{ij}>\epsilon}$ with $A$ the adjacency matrix and $\epsilon=10^{-3}$ reorganised according to the true labels, we see that the algorithm learns somehow the main components.
print(np.sort(np.linalg.eigh(res_genes["Laplacian"])[0]))
def connected_component(A):
n=A.shape[0]
cc=[-1]*n
nbcc=0
def parc(u,i,n):
cc[u]=i
for v in range(n):
if A[u][v]>10**-7 and cc[v]==-1:
parc(v,i,n)
for u in range(n):
if cc[u]==-1:
parc(u,nbcc,n)
nbcc+=1
return cc
cc=connected_component(l)
print(np.array(cc))
D=np.diagonal(res_genes["Laplacian"])
print(np.sort(D))
# Unfortunately, as the sorted values of the laplacian diagonal show, these components correspond to single nodes, connected to no other node in the dataset.
# ## Realistic sensor displacement benchmark dataset
# This part contains the experiments about the multivariate times series measuring body movements.
#
# The data is available at https://archive.ics.uci.edu/ml/machine-learning-databases/00305/
#
# To make the code work, download the dataset, extract it and save data and labels in the folder ./data/realistic_sensor_displacement/
from os import listdir
location = "data/realistic_sensor_displacement"
for filename in listdir(location)[:1]:
file=(pd.read_csv(location+'/'+filename, skiprows=0, sep='\s+'))
# +
l=file.values
print(l.shape)
# -
l=l[:,2:-1]
# We try to learn a laplacian while enforcing a $k$ components structure for its associated graph.
di=learn_k_component_graph(l.T,k=5,maxiter=1000)
# Then, we plot the adjecency matrix, reorganised according to the coefficients of the second eigenvector of the laplacian, which could give a (very) rough estimation of the learned classes, according to spectral graph theory.
V=np.linalg.eigh(di["Laplacian"])[1]
ordre=np.argsort(V.T[1,:])
fig = plt.figure(figsize=(20, 20))
plt.matshow(di["Adjacency"][ordre][:,ordre], fignum=0)
plt.show()
# We do not see any meaningful pattern in the matrix, showing that the graph learning has probably failed.
print(di["convergence"])
print(np.sort(np.linalg.eigh(di["Laplacian"])[0])[:10])
# We see that indeed, the algorithm didn't converge and the laplacian has only one zero eigenvalue.
#
# Then, we try to use thresholding to enforce the smalest coefficients to go to zero and increase the number of components of the graph.
def A_to_L(A):
D=np.diag(np.sum(A,axis=0))
return D-A
A=(di["Adjacency"])*(abs(di["Adjacency"])>10**-3)
L=A_to_L(A)
print(np.sort(np.linalg.eigh(L)[0])[:10])
# Unfortunately, the trick didn't work. We think that one explanation of this is because the data was not adapted to $k$-components graph learning
| .ipynb_checkpoints/experiment-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
import string
# # Idea
#
# Get data
# - Calculate the name length
# - Calculate the chr set
# - Calculate the chr set length
# - Calculate the ratio for the chr set length and the name length
# - Remove the duplicate letter sets
# - Create dataframe with index=names, columns=alphabet
# - Calculate the letter distribution
# - Choose argmin(letter sum); The optimum set must have atleast one of these
# - Iterate through all argmin(letter sum) names:
# - Recursion starts here
# - Mark all name letters to False
# - Update the letter distribution
# - Choose argmin(letter sum); The optimum set must have atleast one of these, but due to n cutoff not all combinations are tested.
# - Calculate the effective set length
# - Calculate the effective ratio
# - Choose the n first names with {the highest effective ratio / shortest length}
# - Iterate through the chosen names
# - The recursion ends here
# ## Read data and calculate some properties
names_df = pd.read_csv("./IMA_mineral_names.txt", sep=',', header=None, names=['names'])
names_df['names'] = names_df['names'].str.strip().str.lower()
names_df['len'] = names_df['names'].str.len()
names_df['tuple'] = names_df['names'].apply(lambda x: tuple(sorted(set(x))))
names_df['setlen'] = names_df['tuple'].apply(lambda x: len(x))
names_df['set_per_len'] = names_df['setlen']/names_df['len']
names_df.head(5)
len(names_df)
# ## Remove duplicates
def sort_and_return_smallest(df):
if len(df) == 1:
return df
df = df.sort_values(by=['len', 'names'])
return df.iloc[:1, :]
# %time names_set = names_df.groupby(by='tuple', as_index=False).apply(sort_and_return_smallest)
len(names_set)
def sort_and_return_smallest_duplicates(df):
if len(df) == 1:
return list(df['names'])
df = df.sort_values(by=['len', 'names'])
names = df.loc[df['len'] == df['len'].iloc[0], 'names']
return list(names)
# %time names_duplicates = names_df.groupby(by='tuple', as_index=False).apply(sort_and_return_smallest_duplicates)
len(names_duplicates)
# In case some of these are in the chosen set
duplicate_name_dict = {}
for value in names_duplicates:
if len(value) > 1:
duplicate_name_dict[value[0]] = value[1:]
names_set.set_index(['names'], inplace=True)
names_set.head()
# ## Create letter table
letter_df = pd.DataFrame(index=names_set.index, columns=list(string.ascii_lowercase), dtype=bool)
letter_df.loc[:] = False
# %%time
for name, set_ in zip(names_set.index, names_set['tuple']):
for letter in set_:
letter_df.loc[name, letter] = True
# ## Find argmin in the letter distribution
lowest_count_letter = letter_df.sum(0).argmin()
lowest_count_letter
# +
# Get subset based on the chosen letter
subsetlen = letter_df[letter_df[lowest_count_letter]].sum(1)
name_len = subsetlen.index.str.len()
setlen = pd.DataFrame({'set_per_len' : subsetlen/name_len, 'len' : name_len})
setlen.head()
# -
# ## Recursion
def get_min_set(df, current_items, m=46, sort_by_len=False, n_search=20):
# Gather results
results = []
# Get letter with lowest number of options
letter = df.sum(0)
letter = letter[letter > 0].argmin()
# Get subset based on the chosen letter
subsetlen = df.loc[df[letter], :].sum(1)
name_len = subsetlen.index.str.len()
setlen = pd.DataFrame({'set_per_len' : subsetlen/name_len, 'len' : name_len})
if sort_by_len:
order_of_operations = setlen.sort_values(by=['len', 'set_per_len'], ascending=True).index
else:
order_of_operations = setlen.sort_values(by=['set_per_len', 'len'], ascending=False).index
# Loop over the mineral names with chosen letter
# Ordered based on the (setlen / len)
for i, (name, letter_bool) in enumerate(df.loc[order_of_operations, :].iterrows()):
if i > n_search:
break
if sum(map(len, current_items))+len(name) >= m:
continue
# Get df containing rest of the letters
df_ = df.copy()
df_.loc[:, letter_bool] = False
# If letters are exhausted there is one result
# Check if the result is less than chosen limit m
if df_.sum(0).sum() == 0 and sum(map(len, current_items))+len(name) < m:
# This result is "the most optimal" under these names
current_items_ = current_items + [name]
len_current_items_ = sum(map(len, current_items_))
len_unique = len(set("".join(current_items_)))
results.append((len_current_items_, current_items_))
if len_current_items_ < 41:
print("len", len_current_items_, "len_unique", len_unique, current_items_, "place 1", flush=True)
continue
# Remove mineral names without new letters
df_ = df_.loc[df_.sum(1) != 0, :]
if df_.sum(0).sum() == 0:
if sum(map(len, current_items))+len(name) < m:
unique_letters = sum(map(len, map(set, current_items + [name])))
if unique_letters == len(string.ascii_lowercase):
# Here is one result (?)
current_items_ = current_items + [name]
len_current_items_ = sum(map(len, current_items_))
len_unique = len(set("".join(current_items_)))
results.append((len_current_items_, current_items_))
if len_current_items_ < 41:
print("len", len_current_items_, "len_unique", len_unique, current_items_, "place 1", flush=True)
continue
current_items_ = current_items + [name]
optimal_result = get_min_set(df_, current_items_, m=m, sort_by_len=sort_by_len, n_search=n_search)
if len(optimal_result):
results.extend(optimal_result)
return results
# ## The effective ratio criteria
# +
# %%time
res_list = []
order_of_oparations = setlen.loc[letter_df.loc[:, lowest_count_letter], :].sort_values(by=['set_per_len', 'len'], ascending=False).index
for i, (name, letter_bool) in enumerate(letter_df.ix[order_of_oparations].iterrows()):
print(name, i+1, "/", len(order_of_oparations), flush=True)
df_ = letter_df.copy()
df_.loc[:, letter_bool] = False
res = get_min_set(df_, [name], m=45, sort_by_len=False, n_search=20)
res_list.extend(res)
# -
res_df = pd.DataFrame([[item[0]] + item[1] for item in res_list]).sort_values(by=0)
res_df.head()
# ## The shortest name length criteria
# +
# %%time
res_list_ = []
order_of_oparations = setlen.loc[letter_df.loc[:, lowest_count_letter], :].sort_values(by=['set_per_len', 'len'], ascending=False).index
for i, (name, letter_bool) in enumerate(letter_df.ix[order_of_oparations].iterrows()):
print(name, i+1, "/", len(order_of_oparations), flush=True)
df_ = letter_df.copy()
df_.loc[:, letter_bool] = False
res_ = get_min_set(df_, [name], m=45, sort_by_len=True, n_search=20)
res_list_.extend(res_)
# +
#res_df_ = pd.DataFrame([[item[0]] + item[1] for item in res_list_]).sort_values(by=0)
# -
res_df.shape #, res_df_.shape
# ## Save the results
# %time res_df.to_csv("./example_but_not_optimum_no_duplicates.csv")
optimum = res_df[res_df[0] == res_df.iloc[0, 0]]
# ## Check for duplicates
optimum.iloc[:, 1:].applymap(lambda x: duplicate_name_dict.get(x, None))
optimum
# ## Validate results
optimum.apply(lambda x: "".join(sorted(set("".join(x.iloc[1:6].values)))) == string.ascii_lowercase, axis=1)
| RiddleMeThis-Mineral_name_pangrams-dynamic_programming_or_not_really.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# +
import numpy as np
import pandas as pd
import tensorflow as tf
import cv2
import matplotlib.pyplot as plt
from tensorflow.python.util import deprecation
deprecation._PRINT_DEPRECATION_WARNINGS = False
tf.logging.set_verbosity(tf.logging.INFO)
# +
import tensorflow as tf
NUM_CLASSES = 228
STARTER_LEARNING_RATE = 0.005
CUT_OFF = 0.184
DECAY_STEPS = 400000
DECAY_RATE = 0.5
def alexnet_model_fn(features, labels, mode):
"""Model function for Alexnet."""
# Input Layer
# Reshape X to 4-D tensor: [batch_size, width, height, channels]
input_layer = tf.convert_to_tensor(features["x"])
#print("input_layer: {}".format(input_layer.shape))
conv1 = tf.layers.conv2d(inputs=input_layer,filters=96,kernel_size=[11, 11],strides=4,padding="valid",activation=tf.nn.relu)
#print("conv1: {}".format(conv1.shape))
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[3, 3], strides=2, padding='valid')
#print("pool1: {}".format(pool1.shape))
conv2 = tf.layers.conv2d(inputs= pool1,filters=256,kernel_size=[5, 5],padding="same",activation=tf.nn.relu)
#print("conv2: {}".format(conv2.shape))
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[3, 3], strides=2, padding='valid')
#print("pool2: {}".format(pool2.shape))
conv3 = tf.layers.conv2d(inputs=pool2,filters=384,kernel_size=[3, 3],padding="same",activation=tf.nn.relu)
#print("conv3: {}".format(conv3.shape))
conv4 = tf.layers.conv2d(inputs=conv3,filters=384,kernel_size=[3, 3],padding="same",activation=tf.nn.relu)
#print("conv4: {}".format(conv4.shape))
conv5 = tf.layers.conv2d(inputs=conv4,filters=256,kernel_size=[3, 3],padding="same",activation=tf.nn.relu)
#print("conv5: {}".format(conv5.shape))
pool5 = tf.layers.max_pooling2d(inputs=conv5, pool_size=[3, 3], strides=2,padding='valid')
#print("pool5: {}".format(pool2.shape))
pool5_flat = tf.reshape(conv5, [-1, 12*12*256])
#print("pool5_flat: {}".format(pool5_flat.shape))
fc6 = tf.layers.dense(inputs=pool5_flat, units=4096, activation=tf.nn.relu)
#print("dense1: {}".format(fc6.shape))
dropout6 = tf.layers.dropout(inputs=fc6, rate=0.2, training=mode == tf.estimator.ModeKeys.TRAIN)
#print("dropout6: {}".format(dropout6.shape))
fc7 = tf.layers.dense(inputs=dropout6, units=4096, activation=tf.nn.relu)
#print("fc7: {}".format(fc7.shape))
dropout7 = tf.layers.dropout(inputs=fc7, rate=0.2, training=mode == tf.estimator.ModeKeys.TRAIN)
#print("dropout7: {}".format(dropout7.shape))
# Logits Layer
# Input Tensor Shape: [batch_size, 4096]
# Output Tensor Shape: [batch_size, 228]
logits = tf.layers.dense(inputs=dropout7, units=NUM_CLASSES)
#print("logits: {}".format(logits.shape))
# Generate Predictions
predictions = {
# Generate predictions (for PREDICT and EVAL mode)
"classes": tf.cast(tf.sigmoid(logits) >= CUT_OFF, tf.int8, name="class_tensor"),
# Add `sigmoid_tensor` to the graph. It is used for PREDICT and by the
# `logging_hook`.
"probabilities": tf.nn.sigmoid(logits, name="prob_tensor")
}
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions)
# Calculate Loss (for both TRAIN and EVAL modes)
#w_tensor = tf.convert_to_tensor(w)
#w_tensor = tf.reshape(w_tensor, [-1,228])
loss = tf.losses.sigmoid_cross_entropy(multi_class_labels=labels, logits=logits)#, weights=w_tensor)
#loss = tf.nn.sigmoid_cross_entropy_with_logits(labels=labels, logits=logits)
# Configure the Training Op (for TRAIN mode)
if mode == tf.estimator.ModeKeys.TRAIN:
global_step = tf.train.get_global_step()
learning_rate = tf.train.exponential_decay(
learning_rate=STARTER_LEARNING_RATE, global_step=global_step,
decay_steps=DECAY_STEPS, decay_rate=DECAY_RATE
)
if global_step % DECAY_STEPS == 0:
tf.logging.info('Learning rate at global step '+str(global_step)+': '+str(learning_rate))
optimizer = tf.train.AdagradOptimizer(learning_rate=learning_rate)
train_op = optimizer.minimize(
loss=loss,
global_step=global_step)
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
# Customize evaluation metric
def meanfscore(predictions, labels):
predictions = tf.reshape(tf.transpose(predictions), [-1])
labels = tf.convert_to_tensor(labels)
labels = tf.reshape(tf.transpose(labels), [-1])
precision_micro, update_op_p = tf.metrics.precision(labels, predictions)
recall_micro, update_op_r = tf.metrics.recall(labels, predictions)
f1_mircro = tf.div(tf.multiply(2., tf.multiply(precision_micro, recall_micro)), tf.add(precision_micro, recall_micro), name="eval_tensor")
return f1_mircro, tf.group(update_op_p, update_op_r)
def precision_micro(predictions, labels):
predictions = tf.reshape(tf.transpose(predictions), [-1])
labels = tf.convert_to_tensor(labels)
labels = tf.reshape(tf.transpose(labels), [-1])
precision_micro, update_op_p = tf.metrics.precision(labels, predictions)
return precision_micro, update_op_p
def recall_micro(predictions, labels):
predictions = tf.reshape(tf.transpose(predictions), [-1])
labels = tf.convert_to_tensor(labels)
labels = tf.reshape(tf.transpose(labels), [-1])
recall_micro, update_op_r = tf.metrics.recall(labels, predictions)
return recall_micro, update_op_r
# Add evaluation metrics (for EVAL mode)
eval_metric_ops = {
"meanfscore": meanfscore(predictions["classes"], labels),
"precision_micro": precision_micro(predictions["classes"], labels),
"recall_micro": recall_micro(predictions["classes"], labels)}
return tf.estimator.EstimatorSpec(
mode=mode, loss=loss, eval_metric_ops=eval_metric_ops)
# +
import numpy as np
import pandas as pd
import cv2
NUM_CLASSES = 228
IMAGE_WIDTH = 224
IMAGE_HEIGHT = 224
def load_images(addrs_list):
images = np.empty((len(addrs_list), IMAGE_WIDTH, IMAGE_HEIGHT, 3), dtype=np.float32)
for i, fpath in enumerate(addrs_list):
img = cv2.imread(fpath, cv2.IMREAD_COLOR)
img = cv2.resize(img, (224, 224))
images[i, ...] = img#.transpose(2, 0, 1)
if i % 1000 == 0:
print('Loading images: {}'.format(i))
return images
def get_multi_hot_labels(df, index_list):
label_id = [df['labelId'][i] for i in index_list]
labels_matrix = np.zeros([len(index_list), NUM_CLASSES], dtype=np.uint8())
for i in range(len(label_id)):
for j in range(len(label_id[i].split(' '))):
row, col = i, int(label_id[i].split(' ')[j]) - 1
labels_matrix[row][col] = 1
return labels_matrix
# +
validation_df = pd.read_csv('/home/ec2-user/SageMaker/imat/train.csv')
eval_path_list = validation_df['imagePath']
eval_data = load_images(eval_path_list)
eval_labels = get_multi_hot_labels(validation_df, list(range(validation_df.shape[0])))
# -
# Evaluate
multilabel_classifier = tf.estimator.Estimator(
model_fn=alexnet_model_fn, model_dir="/home/ec2-user/SageMaker/imat/model/multilabel_alexnet_model")
eval_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": eval_data},
y=eval_labels,
shuffle=False)
eval_results = multilabel_classifier.evaluate(input_fn=eval_input_fn)
print(eval_results)
| notebooks/Alexnet_val.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Throughputs on Ultra96
#
# This jupyter notebook reflects the throughput values on the final (Maximum AUs deployed) implementation of
# - FIFO-ONLY
# - HASH-AMAP
# - FULL-AMAP
#
# The results correspond to the Fig.7c in the paper
#
#
import pynq
from au_hardware_fifo_only import *
from au_hardware_full import *
from au_hardware_hash import *
# ## FIFO-ONLY
au = Au_fifo(Height=180, Width=240, bitfile="bitfile/fifo_1024_5_13.bit", au_number=13, fifo_depth=2048)
T = au.test_random(32768, 100) #(number of events, number of tests)
# # HASH-AMAP
au = Au_hash(Height=180, Width=240, bitfile="bitfile/hash_1024_5_18.bit", au_number=16, fifo_depth=1024)
au.test_random(32768, 100) #(number of events, number of tests)
# ## FULL-AMAP
au = Au_full(Height=180, Width=240, bitfile="bitfile/full_1024_5_4.bit", au_number=4, fifo_depth=1024)
au.test_random(32768, 100)
| hardware/drive/ultra96/test_speed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''qstcgan'': conda)'
# name: python3
# ---
# # iMLE performance
#
# Reconstructing the `cat` state from measurements of the Husimi Q function using
# iterative Maximum Likelihood Estimation (iMLE).
#
# The cat state is defined as:
#
# $$|\psi_{\text{cat}} \rangle = \frac{1}{\mathcal N} ( |\alpha \rangle + |-\alpha \rangle \big ) $$
#
# with $\alpha=2$ and normalization $\mathcal N$.
#
# ## Husimi Q function measurements
#
# The Husimi Q function can be obtained by calculating the expectation value of measuring the following operator:
#
# $$\mathcal O_i = \frac{1}{\pi}|\beta_i \rangle \langle \beta_i|$$
#
# where $|\beta_i \rangle $ are coherent states written in the Fock basis.
#
# +
import numpy as np
from qutip import coherent, coherent_dm, expect, Qobj, fidelity, rand_dm
from qutip.wigner import wigner, qfunc
import tensorflow as tf
from tqdm.auto import tqdm
import matplotlib
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
# +
hilbert_size = 32
alpha = 2
psi = coherent(hilbert_size, alpha) + coherent(hilbert_size, -alpha)
psi = psi.unit() # The .unit() function normalizes the state to have unit trace
grid = 32
xvec = np.linspace(-3, 3, grid)
yvec = np.linspace(-3, 3, grid)
q = qfunc(psi, xvec, yvec, g=2)
cmap = "Blues"
im = plt.pcolor(xvec, yvec, q, vmin=0, vmax=np.max(q), cmap=cmap, shading='auto')
plt.colorbar(im)
plt.xlabel(r"Re($\beta$)")
plt.ylabel(r"Im($\beta$)")
plt.title("Husimi Q function")
plt.show()
# -
# # Construct the measurement operators and simulated data (without any noise)
# +
X, Y = np.meshgrid(xvec, yvec)
betas = (X + 1j*Y).ravel()
m_ops = [coherent_dm(hilbert_size, beta) for beta in betas]
ops_numpy = [op.full() for op in m_ops]
data = expect(m_ops, psi)
rho_true = psi*psi.dag()
# -
# # Iterative maximum likelihood estimation
# +
fidelities = []
max_iterations = 10000
rho = rand_dm(hilbert_size, 0.8)
fidelities.append(fidelity(rho_true, rho))
pbar = tqdm(range(max_iterations))
for i in range(max_iterations):
guessed_val = expect(m_ops, rho)
ratio = data / guessed_val
R = Qobj(np.einsum("aij,a->ij", ops_numpy, ratio))
rho = R * rho * R
rho = rho / rho.tr()
f = fidelity(rho, rho_true)
fidelities.append(f)
pbar.set_description("Fidelity iMLE {:.4f}".format(f))
pbar.update()
# -
iterations = np.arange(len(fidelities))
plt.plot(iterations, fidelities, color="blue", label="iMLE")
plt.legend()
plt.xlabel("Iterations")
plt.ylabel("Fidelity")
plt.ylim(0, 1.02)
plt.grid(which='minor', alpha=0.2)
plt.grid(which='major', alpha=0.2)
plt.xscale('log')
plt.show()
| paper-figures/fig3a-imle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # General Purpose Visualization Notebook
# ## Visualize Uncertainty Quantifications
# Assumes `run_experiments` has generated results in the `[EXP_FOLDER]/results` folder.
# %load_ext autoreload
# %autoreload 2
EXP_FOLDER = 'MNIST'
# +
import sys
import os
sys.path.append(os.path.abspath(EXP_FOLDER))
import config # imported from EXP_FOLDER
import numpy as np
import glob
import torch
import matplotlib
matplotlib.rc('xtick', labelsize=8)
matplotlib.rc('ytick', labelsize=8)
import matplotlib.pyplot as plt
from ipywidgets import interact
from nn_ood.utils.viz import plot_histogram, plot_scatter, plot_rocs_by_error, plot_times, summarize_ood_results, summarize_ood_results_by_error, plot_perf_vs_runtime, plot_rocs_by_dist, generate_table_block
from nn_ood.utils.viz import plot_transform_sweep
from nn_ood.utils.inspect import norm_by_param
# -
results_dict = {}
folder_path = os.path.join(EXP_FOLDER, "results","*")
for filename in glob.glob(folder_path):
name = os.path.basename(filename)
results_dict[name] = torch.load(filename)
# ## Load Models
# +
# load base model
print("Loading models")
models = []
for i in range(config.N_MODELS):
print("loading model %d" % i)
filename = os.path.join(EXP_FOLDER, 'models', config.FILENAME + "_%d" % i)
state_dict = torch.load(filename)
model = config.make_model()
model.load_state_dict(state_dict)
model.to(config.device)
model.eval()
models.append(model)
plt.show()
model = models[0]
## Ask which uncertainty model to use for viz
import ipywidgets as widgets
names = list(config.test_unc_models.keys())
name_widget = widgets.Dropdown(
options=names,
value=names[0],
description='Uncertainty Model:',
disabled=False,
)
name_widget
# +
# load unc model
name = name_widget.value
info = config.test_unc_models[name]
print(info)
config.unfreeze_model(model)
if 'freeze' in info:
if type(info['freeze']) is bool:
freeze_frac = None
else:
freeze_frac = info['freeze']
config.freeze_model(model, freeze_frac=freeze_frac)
if 'apply_fn' is info:
model.apply(info['apply_fn'])
if 'multi_model' in info:
unc_model = info['class'](models, config.dist_fam, info['kwargs'])
else:
unc_model = info['class'](model, config.dist_fam, info['kwargs'])
if info['load_name'] is not None:
filename = os.path.join(EXP_FOLDER, "models", info['load_name']+"_"+config.FILENAME)
unc_model.load_state_dict(torch.load(filename), strict=False)
# -
# ## Visualize Samples from Dataset
config.viz_datasets(idx=5, unc_model=unc_model)
config.viz_transforms(idx=5, unc_model=unc_model)
# ## Generate plots from results
for filename, info in config.plots_to_generate.items():
summarized_results = info['summary_fn'](results_dict, *info['summary_fn_args'], **info['summary_fn_kwargs'])
info['plot_fn'](summarized_results, *info['plot_fn_args'], **info['plot_fn_kwargs'])
if 'legend' in info:
plt.legend(**info['legend'])
if 'title' in info:
plt.title(info['title'])
plt.tight_layout()
plt.savefig(os.path.join(EXP_FOLDER,filename))
plt.show()
# # Other visualization tools
# ### Histograms by technique
# +
n_techniques = len(config.keys_to_compare)
max_cols = 4
n_rows = int(np.ceil(n_techniques/max_cols))
n_cols = min(max_cols, n_techniques)
fig, axes = plt.subplots(n_rows,n_cols,figsize=[5*n_cols, 5*n_rows])
axes = axes.flatten()
for i, name in enumerate(config.keys_to_compare):
if name not in results_dict:
continue
results = results_dict[name]
plot_histogram(axes[i], results)
axes[i].set_title(name)
plt.show()
# -
# ### Scatterplots of Uncertainty vs Error
# +
n_techniques = len(config.keys_to_compare)
n_datasets = len(config.test_dataset_args)
fig, axes = plt.subplots(n_techniques,n_datasets,figsize=[2*n_datasets, 2*n_techniques], sharey='row')
for i, name in enumerate(config.keys_to_compare):
if name not in results_dict:
continue
results = results_dict[name]
plot_scatter(axes[i], results)
axes[i][n_datasets//2 + n_datasets%2 - 1].set_title(name)
plt.tight_layout()
# -
# ### Compare OoD performance vs Runtime
# +
summary = summarize_ood_results(results_dict, config.in_dist_splits, config.out_dist_splits,
keys_to_compare = config.keys_to_compare)
plot_perf_vs_runtime(summary,
colors=config.colors, figsize=[4,2.5], dpi=150, normalize_x=True)
plt.title(EXP_FOLDER)
plt.legend()
plt.tight_layout()
# -
@interact(x=(0.0,config.err_thresh*5, config.err_thresh/10.))
def g(x=config.err_thresh):
summary = summarize_ood_results_by_error(results_dict, config.splits_to_use, x,
keys_to_compare = config.keys_to_compare)
plot_perf_vs_runtime(summary,
colors=config.colors, figsize=[4,2.5], dpi=150, normalize_x=True)
plt.legend()
# ### Noise analysis
# +
if "transforms" not in dir(config):
raise NameError("No transforms to visualize for this experiment")
transforms_results_dict = {}
folder_path = os.path.join(EXP_FOLDER, "results_transforms","*")
for filename in glob.glob(folder_path):
name = os.path.basename(filename)
transforms_results_dict[name] = torch.load(filename)
plot_transform_sweep(transforms_results_dict)
# -
# ### Visualize Runtime Performance
plot_times(results_dict, keys_to_compare=config.keys_to_compare, colors=config.colors)
| experiments/visualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from scipy import stats
import itertools
from sklearn import linear_model
from sklearn.model_selection import train_test_split
from numpy import ones,vstack
from numpy.linalg import lstsq
import matplotlib.pyplot as plt
import statsmodels.api as sm;
pd.options.display.max_columns = 150;
Ture = True;
# -
typedict = {#'PID' : 'nominal',
'SalePrice' : 'Continuous',
#Matt
'LotFrontage' : 'Continuous',
'LotArea' : 'Continuous',
'maybe_LotShape' : 'Nominal',
'LandSlope' : 'Nominal',
'LandContour' : 'Nominal',
'maybe_MSZoning' : 'Nominal',
'Street_paved' : 'Nominal',
'Alley' : 'Nominal',
'Neighborhood' : 'Nominal',
'Foundation' : 'Nominal',
'Utilities' : 'Nominal',
'Heating' : 'Nominal',
'HeatingQC_nom' : 'Ordinal',
'CentralAir' : 'Nominal',
'Electrical' : 'Nominal',
'HeatingQC_ord' : 'Ordinal',
'LotShape_com' : 'Nominal',
'MSZoning_com' : 'Nominal',
'LF_Near_NS_RR' : 'Nominal',
'LF_Near_Positive_Feature' : 'Nominal',
'LF_Adjacent_Arterial_St' : 'Nominal',
'LF_Near_EW_RR' : 'Nominal',
'LF_Adjacent_Feeder_St' : 'Nominal',
'LF_Near_Postive_Feature' : 'Nominal',
'Heating_com' : 'Nominal',
'Electrical_com' : 'Nominal',
'LotConfig_com' : 'Nominal',
'LotFrontage_log' : 'Continuous',
'LotArea_log' : 'Continuous',
#Oren
'MiscFeature': 'Nominal',
'Fireplaces': 'Discrete',
'FireplaceQu': 'Ordinal',
'PoolQC': 'Ordinal',
'PoolArea': 'Continuous',
'PavedDrive': 'Nominal',
'ExterQual': 'Ordinal',
'OverallQual': 'Ordinal',
'maybe_OverallCond': 'Ordinal',
'MiscVal': 'Continuous',
'YearBuilt': 'Discrete',
'YearRemodAdd': 'Discrete',
'KitchenQual': 'Ordinal',
'Fence': 'Ordinal',
'RoofStyle': 'Nominal',
'RoofMatl': 'Nominal',
'maybe_ExterCond': 'Ordinal',
'maybe_MasVnrType': 'Nominal',
'MasVnrArea': 'Continuous',
#Mo
'BsmtQual_ord': 'Ordinal',
'BsmtCond_ord': 'Ordinal',
'BsmtExposure_ord': 'Ordinal',
'TotalBsmtSF': 'Continuous',
'BSMT_GLQ':'Continuous',
'BSMT_Rec':'Continuous',
'maybe_BsmtUnfSF': 'Continuous',
'maybe_BSMT_ALQ':'Continuous',
'maybe_BSMT_BLQ':'Continuous',
'maybe_BSMT_LwQ':'Continuous',
#Deck
'WoodDeckSF':'Continuous',
'OpenPorchSF':'Continuous',
'ScreenPorch':'Continuous',
'maybe_EnclosedPorch':'Continuous',
'maybe_3SsnPorch':'Continuous',
#Garage
'GarageFinish':'Nominal',
'GarageYrBlt':'Continuous',
'GarageCars':'Ordinal',
'GarageArea':'Continuous',
'GarageType_con':'Nominal',
'maybe_GarageQual':'Nominal',
'maybe_GarageCond':'Nominal',
# Hao-Wei
"SaleType": "Nominal",
"BldgType": "Nominal",
"Functional_ord": "Ordinal", # Changed from "Functional"
"1stFlrSF": "Continuous",
"2ndFlrSF": "Continuous",
"maybe_LowQualFinSF": "Continuous", # Rejectable p-value
"GrLivArea": "Cbontinuous",
"BsmtFullBath": "Discrete",
"maybe_BsmtHalfBath": "Discrete", # Rejectable p-value
"FullBath": "Discrete",
"maybe_HalfBath": "Discrete",
"BedroomAbvGr": "Discrete",
"KitchenAbvGr": "Discrete",
"TotRmsAbvGrd": "Discrete",
"MoSold": "Discrete", # Rejectable p-value
"YrSold": "Discrete", # Rejectable p-value
"1stFlrSF_log": "Continuous",
"2ndFlrSF_log": "Continuous",
"GrLivArea_log": "Continuous",
"number_floors": "Discrete",
"attic": "Ordinal",
"PUD": "Nominal",
#### Whose?
"SaleCondition": "Nominal",
"SalePrice_log": "Continuous",
"MS_coded": "Nominal",
"sold_datetime": "Discrete",
# Used locally in this notebook
"Months_Elapsed": "Discrete"
}
attic_dict = {"No attic": 0, "Finished": 2, "Unfinished": 1};
fence_dict = {"No Fence": 0, "Minimum Privacy": 3, "Good Privacy": 4, "Good Wood": 2 , "Minimum Wood/Wire": 1};
PoolQC_dict = {0:0, "0":0, "Fa": 1, "TA":2, "Gd":3, "Ex":4};
housing = pd.read_csv('../data/ames_housing_price_data_v2.csv', index_col = 0);
# +
y = housing["SalePrice"];
ylog = housing["SalePrice_log"];
x = housing.drop(["SalePrice", "SalePrice_log"], axis = 1);
# -
np.sum(x.isnull(), axis = 0)[:50]
| Hao-Wei/Data Cleaning (phase 2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# slow down a bit when hacking something together, e.g. I forgot to add a simple function call
# tuple unpacking is nice, but cannot be done in a nested list comprehension
# don't forget .items in for k,v in dict.items()
# use hashlib for md5 encodings
# multiline list comprehensions don't need extra parentheses, but multiline if statements do
# np.clip min and max can be omitted by specifying None
# try except looks nice untill it obscures your real error
# parsing ints to ints instead of strings is really important
# checking whether someting is an int should be done with isinstance, not with isalpha() (fails on int)
# removing from a list while iterating can be done safely by iterating over a slice(?)
# with re make sure to use r'' literal strings
# read assignment before tinkering with networkx and discovering its not necessary
# sometimes a simple for loop works better then a list comprehension when parsing the input, and just add to concept variables
# for incrementing a string, you can use chr(ord(inp)+1)
# find repeating characters re.findall(r'([a-z])\1', password)
# regex: modify operator to nongreedy by appending ?
# ok so sometime you can bruteforce a problem....
# +
from aoc_utils import *
import itertools
lines = data('input', parser=int, sep='\n')
lines.sort(reverse=True)
lines
# -
# the easy way, should have tried this probably, but only found out afterwards
total = 0
fill=150
for i in range(len(lines)):
total += sum(1 for comb in itertools.combinations(lines,i) if sum(comb)==fill)
print(f'combinations of {i} buckets, {total}')
# uncomment for part 2:
# if total > 0: break
# the hard way, was struggling with a good algo. was making for loops which exploded the result
def countnum(freebottles, tofill=150):
if not lines or tofill < 0:
return 0
if sum(freebottles) < tofill:
return 0
newtofill = tofill - freebottles[0]
count = 0
if newtofill == 0:
count+= 1
count += countnum(freebottles[1:], tofill)
return count
if newtofill < 0:
count += countnum(freebottles[1:], tofill)
return count
count += countnum(freebottles[1:], newtofill)
count += countnum(freebottles[1:], tofill)
return count
countnum(lines)
# part 2
fit = set()
def countnum(freebottles, tofill=150, used =0):
global fit
if not lines or tofill < 0:
return 0
if sum(freebottles) < tofill:
return 0
newtofill = tofill - freebottles[0]
count = 0
if newtofill == 0:
fit.add(used+1)
if used+1==4:
count+= 1
count += countnum(freebottles[1:], tofill, used)
return count
if newtofill < 0:
count += countnum(freebottles[1:], tofill, used)
return count
count += countnum(freebottles[1:], newtofill, used+1)
count += countnum(freebottles[1:], tofill, used)
return count
countnum(lines)
| advent_of_code_2015/day 17/solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ACbjNjyO4f_8"
# ##### Copyright 2019 The TensorFlow Hub Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + id="MCM50vaM4jiK"
# Copyright 2018 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] id="9qOVy-_vmuUP"
# # Approximate Nearest Neighbor(ANN) 및 텍스트 임베딩을 사용한 의미론적 검색
#
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/hub/tutorials/semantic_approximate_nearest_neighbors"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png"> GitHub에서 소스 보기</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드</a></td>
# <td><a href="https://tfhub.dev/google/universal-sentence-encoder/2"><img src="https://www.tensorflow.org/images/hub_logo_32px.png">TF Hub 모델보기</a></td>
# </table>
# + [markdown] id="7Hks9F5qq6m2"
# 이 튜토리얼에서는 입력 데이터가 제공된 [TensorFlow Hub](https://tfhub.dev)(TF-Hub) 모듈에서 임베딩을 생성하고 추출된 임베딩을 사용하여 approximate nearest neighbour(ANN) 인덱스를 빌드하는 방법을 보여줍니다. 그런 다음 이 인덱스를 실시간 유사성 일치 및 검색에 사용할 수 있습니다.
#
# 많은 양의 데이터를 처리할 때 전체 리포지토리를 스캔하여 주어진 쿼리와 가장 유사한 항목을 실시간으로 찾는 식으로 정확한 일치 작업을 수행하는 것은 효율적이지 않습니다. 따라서 속도를 크게 높이기 위해 정확한 nearest neighbor(NN) 일치를 찾을 때 약간의 정확성을 절충할 수 있는 근사 유사성 일치 알고리즘을 사용합니다.
#
# 이 튜토리얼에서는 쿼리와 가장 유사한 헤드라인을 찾기 위해 뉴스 헤드라인 자료의 텍스트를 실시간으로 검색하는 예를 보여줍니다. 키워드 검색과 달리 이 검색으로 텍스트 임베딩에 인코딩된 의미론적 유사성이 포착됩니다.
#
# 이 튜토리얼의 단계는 다음과 같습니다.
#
# 1. 샘플 데이터를 다운로드합니다.
# 2. TF-Hub 모듈을 사용하여 데이터에 대한 임베딩을 생성합니다.
# 3. 임베딩에 대한 ANN 인덱스를 빌드합니다.
# 4. 유사성 일치에 인덱스를 사용합니다.
#
# [TensorFlow Transform](https://beam.apache.org/documentation/programming-guide/)(TF-Transform)과 함께 [Apache Beam](https://www.tensorflow.org/tfx/tutorials/transform/simple)을 사용하여 TF-Hub 모듈에서 임베딩을 생성합니다. 또한 Spotify의 [ANNOY](https://github.com/spotify/annoy) 라이브러리를 사용하여 nearest neighbour(NN) 인덱스를 빌드합니다. 이 [Github 리포지토리](https://github.com/erikbern/ann-benchmarks)에서 ANN 프레임워크의 벤치마킹을 찾을 수 있습니다.
#
# 이 튜토리얼에서는 TensorFlow 1.0을 사용하며 TF-Hub의 TF1 [Hub 모듈](https://www.tensorflow.org/hub/tf1_hub_module)에서만 동작합니다. [본 튜토리얼의 TF2 업데이트 버전](https://github.com/tensorflow/hub/blob/master/examples/colab/tf2_semantic_approximate_nearest_neighbors.ipynb)을 참조하세요.
# + [markdown] id="Q0jr0QK9qO5P"
# ## 설정
# + [markdown] id="whMRj9qeqed4"
# 필요한 라이브러리를 설치합니다.
# + id="qmXkLPoaqS--"
# !pip install -q apache_beam
# !pip install -q 'scikit_learn~=0.23.0' # For gaussian_random_matrix.
# !pip install -q annoy
# + [markdown] id="A-vBZiCCqld0"
# 필요한 라이브러리를 가져옵니다.
# + id="6NTYbdWcseuK"
import os
import sys
import pathlib
import pickle
from collections import namedtuple
from datetime import datetime
import numpy as np
import apache_beam as beam
import annoy
from sklearn.random_projection import gaussian_random_matrix
import tensorflow.compat.v1 as tf
import tensorflow_hub as hub
# + id="_GF0GnLqGdPQ"
# TFT needs to be installed afterwards
# !pip install -q tensorflow_transform==0.24
import tensorflow_transform as tft
import tensorflow_transform.beam as tft_beam
# + id="tx0SZa6-7b-f"
print('TF version: {}'.format(tf.__version__))
print('TF-Hub version: {}'.format(hub.__version__))
print('TF-Transform version: {}'.format(tft.__version__))
print('Apache Beam version: {}'.format(beam.__version__))
# + [markdown] id="P6Imq876rLWx"
# ## 1. 샘플 데이터 다운로드하기
#
# [A Million News Headlines](https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/SYBGZL#) 데이터세트에는 평판이 좋은 Australian Broadcasting Corp. (ABC)에서 공급한 15년치의 뉴스 헤드라인이 수록되어 있습니다. 이 뉴스 데이터세트에는 호주에 보다 세분화된 초점을 두고 2003년 초부터 2017년 말까지 전 세계적으로 일어난 주목할만한 사건에 대한 역사적 기록이 요약되어 있습니다.
#
# **형식**: 탭으로 구분된 2열 데이터: 1) 발행일 및 2) 헤드라인 텍스트. 여기서는 헤드라인 텍스트에만 관심이 있습니다.
#
# + id="OpF57n8e5C9D"
# !wget 'https://dataverse.harvard.edu/api/access/datafile/3450625?format=tab&gbrecs=true' -O raw.tsv
# !wc -l raw.tsv
# !head raw.tsv
# + [markdown] id="Reeoc9z0zTxJ"
# 단순화를 위해 헤드라인 텍스트만 유지하고 발행일은 제거합니다.
# + id="INPWa4upv_yJ"
# !rm -r corpus
# !mkdir corpus
with open('corpus/text.txt', 'w') as out_file:
with open('raw.tsv', 'r') as in_file:
for line in in_file:
headline = line.split('\t')[1].strip().strip('"')
out_file.write(headline+"\n")
# + id="5-oedX40z6o2"
# !tail corpus/text.txt
# + [markdown] id="ls0Zh7kYz3PM"
# ## TF-Hub 모듈을 로드하는 도우미 함수
# + id="vSt_jmyKz3Xp"
def load_module(module_url):
embed_module = hub.Module(module_url)
placeholder = tf.placeholder(dtype=tf.string)
embed = embed_module(placeholder)
session = tf.Session()
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
print('TF-Hub module is loaded.')
def _embeddings_fn(sentences):
computed_embeddings = session.run(
embed, feed_dict={placeholder: sentences})
return computed_embeddings
return _embeddings_fn
# + [markdown] id="2AngMtH50jNb"
# ## 2. 데이터에 대한 임베딩 생성하기
#
# 이 튜토리얼에서는 [Universal Sentence Encoder](https://tfhub.dev/google/universal-sentence-encoder/2)를 사용하여 헤드라인 데이터에 대한 임베딩을 생성합니다. 그런 다음 문장 임베딩을 사용하여 문장 수준의 의미 유사성을 쉽게 계산할 수 있습니다. Apache Beam과 TF-Transform을 사용하여 임베딩 생성 프로세스를 실행합니다.
# + [markdown] id="F_DvXnDB1pEX"
# ### 임베딩 추출 메서드
# + id="yL7OEY1E0A35"
encoder = None
def embed_text(text, module_url, random_projection_matrix):
# Beam will run this function in different processes that need to
# import hub and load embed_fn (if not previously loaded)
global encoder
if not encoder:
encoder = hub.Module(module_url)
embedding = encoder(text)
if random_projection_matrix is not None:
# Perform random projection for the embedding
embedding = tf.matmul(
embedding, tf.cast(random_projection_matrix, embedding.dtype))
return embedding
# + [markdown] id="_don5gXy9D59"
# ### TFT preprocess_fn 메서드 만들기
# + id="fwYlrzzK9ECE"
def make_preprocess_fn(module_url, random_projection_matrix=None):
'''Makes a tft preprocess_fn'''
def _preprocess_fn(input_features):
'''tft preprocess_fn'''
text = input_features['text']
# Generate the embedding for the input text
embedding = embed_text(text, module_url, random_projection_matrix)
output_features = {
'text': text,
'embedding': embedding
}
return output_features
return _preprocess_fn
# + [markdown] id="SQ492LN7A-NZ"
# ### 데이터세트 메타데이터 만들기
# + id="d2D4332VA-2V"
def create_metadata():
'''Creates metadata for the raw data'''
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import schema_utils
feature_spec = {'text': tf.FixedLenFeature([], dtype=tf.string)}
schema = schema_utils.schema_from_feature_spec(feature_spec)
metadata = dataset_metadata.DatasetMetadata(schema)
return metadata
# + [markdown] id="5zlSLPzRBm6H"
# ### Beam 파이프라인
# + id="jCGUIB172m2G"
def run_hub2emb(args):
'''Runs the embedding generation pipeline'''
options = beam.options.pipeline_options.PipelineOptions(**args)
args = namedtuple("options", args.keys())(*args.values())
raw_metadata = create_metadata()
converter = tft.coders.CsvCoder(
column_names=['text'], schema=raw_metadata.schema)
with beam.Pipeline(args.runner, options=options) as pipeline:
with tft_beam.Context(args.temporary_dir):
# Read the sentences from the input file
sentences = (
pipeline
| 'Read sentences from files' >> beam.io.ReadFromText(
file_pattern=args.data_dir)
| 'Convert to dictionary' >> beam.Map(converter.decode)
)
sentences_dataset = (sentences, raw_metadata)
preprocess_fn = make_preprocess_fn(args.module_url, args.random_projection_matrix)
# Generate the embeddings for the sentence using the TF-Hub module
embeddings_dataset, _ = (
sentences_dataset
| 'Extract embeddings' >> tft_beam.AnalyzeAndTransformDataset(preprocess_fn)
)
embeddings, transformed_metadata = embeddings_dataset
# Write the embeddings to TFRecords files
embeddings | 'Write embeddings to TFRecords' >> beam.io.tfrecordio.WriteToTFRecord(
file_path_prefix='{}/emb'.format(args.output_dir),
file_name_suffix='.tfrecords',
coder=tft.coders.ExampleProtoCoder(transformed_metadata.schema))
# + [markdown] id="uHbq4t2gCDAG"
# ### 무작위 투영 가중치 행렬 생성하기
#
# [무작위 투영](https://en.wikipedia.org/wiki/Random_projection)은 유클리드 공간에 있는 점 집합의 차원을 줄이는 데 사용되는 간단하지만 강력한 기술입니다. 이론적 배경은 [Johnson-Lindenstrauss 보조 정리](https://en.wikipedia.org/wiki/Johnson%E2%80%93Lindenstrauss_lemma)를 참조하세요.
#
# 무작위 투영으로 임베딩의 차원을 줄이면 ANN 인덱스를 빌드하고 쿼리하는 데 필요한 시간이 줄어듭니다.
#
# 이 튜토리얼에서는 [Scikit-learn](https://scikit-learn.org/stable/modules/random_projection.html#gaussian-random-projection) 라이브러리의 [가우스 무작위 투영](https://en.wikipedia.org/wiki/Random_projection#Gaussian_random_projection)을 사용합니다.
# + id="T1aYPeOUCDIP"
def generate_random_projection_weights(original_dim, projected_dim):
random_projection_matrix = None
if projected_dim and original_dim > projected_dim:
random_projection_matrix = gaussian_random_matrix(
n_components=projected_dim, n_features=original_dim).T
print("A Gaussian random weight matrix was creates with shape of {}".format(random_projection_matrix.shape))
print('Storing random projection matrix to disk...')
with open('random_projection_matrix', 'wb') as handle:
pickle.dump(random_projection_matrix,
handle, protocol=pickle.HIGHEST_PROTOCOL)
return random_projection_matrix
# + [markdown] id="CHxZX2Z3Nk64"
# ### 매개변수 설정하기
#
# 무작위 투영 없이 원래 임베딩 공간을 사용하여 인덱스를 빌드하려면 `projected_dim` 매개변수를 `None`으로 설정합니다. 그러면 높은 차원의 임베딩에 대한 인덱싱 스텝이 느려집니다.
# + cellView="form" id="feMVXFL0NlIM"
module_url = 'https://tfhub.dev/google/universal-sentence-encoder/2' #@param {type:"string"}
projected_dim = 64 #@param {type:"number"}
# + [markdown] id="On-MbzD922kb"
# ### 파이프라인 실행하기
# + id="Y3I1Wv4i21yY"
import tempfile
output_dir = pathlib.Path(tempfile.mkdtemp())
temporary_dir = pathlib.Path(tempfile.mkdtemp())
g = tf.Graph()
with g.as_default():
original_dim = load_module(module_url)(['']).shape[1]
random_projection_matrix = None
if projected_dim:
random_projection_matrix = generate_random_projection_weights(
original_dim, projected_dim)
args = {
'job_name': 'hub2emb-{}'.format(datetime.utcnow().strftime('%y%m%d-%H%M%S')),
'runner': 'DirectRunner',
'batch_size': 1024,
'data_dir': 'corpus/*.txt',
'output_dir': output_dir,
'temporary_dir': temporary_dir,
'module_url': module_url,
'random_projection_matrix': random_projection_matrix,
}
print("Pipeline args are set.")
args
# + id="iS9obmeP4ZOA"
# !rm -r {output_dir}
# !rm -r {temporary_dir}
print("Running pipeline...")
# %time run_hub2emb(args)
print("Pipeline is done.")
# + id="JAwOo7gQWvVd"
# !ls {output_dir}
# + [markdown] id="HVnee4e6U90u"
# 생성된 임베딩의 일부를 읽습니다.
# + id="-K7pGXlXOj1N"
import itertools
embed_file = os.path.join(output_dir, 'emb-00000-of-00001.tfrecords')
sample = 5
record_iterator = tf.io.tf_record_iterator(path=embed_file)
for string_record in itertools.islice(record_iterator, sample):
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value
embedding = np.array(example.features.feature['embedding'].float_list.value)
print("Embedding dimensions: {}".format(embedding.shape[0]))
print("{}: {}".format(text, embedding[:10]))
# + [markdown] id="agGoaMSgY8wN"
# ## 3. 임베딩을 위한 ANN 인덱스 빌드하기
#
# [Approximate Nearest Neighbors Oh Yeah](https://github.com/spotify/annoy)(ANNOY)는 주어진 쿼리 포인트에 가까운 공간에서 포인트를 검색하기 위한 Python 바인딩이 있는 C++ 라이브러리입니다. 또한 ANNOY는 메모리에 매핑되는 대규모 읽기 전용 파일 기반 데이터 구조를 만들며, [Spotify](https://www.spotify.com)에서 음악 추천을 위해 빌드하고 사용합니다.
# + id="UcPDspU3WjgH"
def build_index(embedding_files_pattern, index_filename, vector_length,
metric='angular', num_trees=100):
'''Builds an ANNOY index'''
annoy_index = annoy.AnnoyIndex(vector_length, metric=metric)
# Mapping between the item and its identifier in the index
mapping = {}
embed_files = tf.gfile.Glob(embedding_files_pattern)
print('Found {} embedding file(s).'.format(len(embed_files)))
item_counter = 0
for f, embed_file in enumerate(embed_files):
print('Loading embeddings in file {} of {}...'.format(
f+1, len(embed_files)))
record_iterator = tf.io.tf_record_iterator(
path=embed_file)
for string_record in record_iterator:
example = tf.train.Example()
example.ParseFromString(string_record)
text = example.features.feature['text'].bytes_list.value[0].decode("utf-8")
mapping[item_counter] = text
embedding = np.array(
example.features.feature['embedding'].float_list.value)
annoy_index.add_item(item_counter, embedding)
item_counter += 1
if item_counter % 100000 == 0:
print('{} items loaded to the index'.format(item_counter))
print('A total of {} items added to the index'.format(item_counter))
print('Building the index with {} trees...'.format(num_trees))
annoy_index.build(n_trees=num_trees)
print('Index is successfully built.')
print('Saving index to disk...')
annoy_index.save(index_filename)
print('Index is saved to disk.')
print("Index file size: {} GB".format(
round(os.path.getsize(index_filename) / float(1024 ** 3), 2)))
annoy_index.unload()
print('Saving mapping to disk...')
with open(index_filename + '.mapping', 'wb') as handle:
pickle.dump(mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
print('Mapping is saved to disk.')
print("Mapping file size: {} MB".format(
round(os.path.getsize(index_filename + '.mapping') / float(1024 ** 2), 2)))
# + id="AgyOQhUq6FNE"
embedding_files = "{}/emb-*.tfrecords".format(output_dir)
embedding_dimension = projected_dim
index_filename = "index"
# !rm {index_filename}
# !rm {index_filename}.mapping
# %time build_index(embedding_files, index_filename, embedding_dimension)
# + id="Ic31Tm5cgAd5"
# !ls
# + [markdown] id="maGxDl8ufP-p"
# ## 4. 유사성 일치에 인덱스 사용하기
#
# 이제 ANN 인덱스를 사용하여 의미상 입력 쿼리에 가까운 뉴스 헤드라인을 찾을 수 있습니다.
# + [markdown] id="_dIs8W78fYPp"
# ### 인덱스 및 매핑 파일 로드하기
# + id="jlTTrbQHayvb"
index = annoy.AnnoyIndex(embedding_dimension)
index.load(index_filename, prefault=True)
print('Annoy index is loaded.')
with open(index_filename + '.mapping', 'rb') as handle:
mapping = pickle.load(handle)
print('Mapping file is loaded.')
# + [markdown] id="y6liFMSUh08J"
# ### 유사성 일치 메서드
# + id="mUxjTag8hc16"
def find_similar_items(embedding, num_matches=5):
'''Finds similar items to a given embedding in the ANN index'''
ids = index.get_nns_by_vector(
embedding, num_matches, search_k=-1, include_distances=False)
items = [mapping[i] for i in ids]
return items
# + [markdown] id="hjerNpmZja0A"
# ### 주어진 쿼리에서 임베딩 추출하기
# + id="a0IIXzfBjZ19"
# Load the TF-Hub module
print("Loading the TF-Hub module...")
g = tf.Graph()
with g.as_default():
embed_fn = load_module(module_url)
print("TF-Hub module is loaded.")
random_projection_matrix = None
if os.path.exists('random_projection_matrix'):
print("Loading random projection matrix...")
with open('random_projection_matrix', 'rb') as handle:
random_projection_matrix = pickle.load(handle)
print('random projection matrix is loaded.')
def extract_embeddings(query):
'''Generates the embedding for the query'''
query_embedding = embed_fn([query])[0]
if random_projection_matrix is not None:
query_embedding = query_embedding.dot(random_projection_matrix)
return query_embedding
# + id="kCoCNROujEIO"
extract_embeddings("Hello Machine Learning!")[:10]
# + [markdown] id="nE_Q60nCk_ZB"
# ### 가장 유사한 항목을 찾기 위한 쿼리 입력하기
# + cellView="form" id="wC0uLjvfk5nB"
#@title { run: "auto" }
query = "confronting global challenges" #@param {type:"string"}
print("Generating embedding for the query...")
# %time query_embedding = extract_embeddings(query)
print("")
print("Finding relevant items in the index...")
# %time items = find_similar_items(query_embedding, 10)
print("")
print("Results:")
print("=========")
for item in items:
print(item)
# + [markdown] id="wwtMtyOeDKwt"
# ## 더 자세히 알고 싶나요?
#
# [tensorflow.org](https://www.tensorflow.org/)에서 TensorFlow에 대해 자세히 알아보고 [tensorflow.org/hub](https://www.tensorflow.org/hub/)에서 TF-Hub API 설명서를 확인할 수 있습니다. 추가적인 텍스트 임베딩 모듈 및 이미지 특성 벡터 모듈을 포함해 [tfhub.dev](https://tfhub.dev/)에서 사용 가능한 TensorFlow Hub 모듈을 찾아보세요.
#
# 빠르게 진행되는 Google의 머신러닝 실무 개요 과정인 [머신러닝 집중 과정](https://developers.google.com/machine-learning/crash-course/)도 확인해 보세요.
| site/ko/hub/tutorials/semantic_approximate_nearest_neighbors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="WFHFZ9hCbwh1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 442} outputId="0b9f4f0d-2076-4d24-9faf-9cfea5403518" executionInfo={"status": "ok", "timestamp": 1583440939187, "user_tz": -60, "elapsed": 12453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
# !pip install --upgrade tables
# !pip install eli5
# + id="7U3j4_AZPG6k" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
# + id="6zidFUiCb572" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="319a6a06-605f-451b-86b2-ae6adbbad315" executionInfo={"status": "ok", "timestamp": 1583441149062, "user_tz": -60, "elapsed": 982, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + id="79YRNkgzcppm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3c784d1d-be33-4ead-9070-5a5b5c4d7855" executionInfo={"status": "ok", "timestamp": 1583441162214, "user_tz": -60, "elapsed": 2030, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
# ls
# + [markdown] id="KGtf7iD1cW4t" colab_type="text"
# # Wczytywanie danych
# + id="FJ56j8wAcacB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f1252085-3d80-497c-b536-e6eedd203f25" executionInfo={"status": "ok", "timestamp": 1583441182139, "user_tz": -60, "elapsed": 4398, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="Wm8LkBpRcuq2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="d23afc88-363c-40da-cb44-807edf0c74a9" executionInfo={"status": "ok", "timestamp": 1583441193658, "user_tz": -60, "elapsed": 805, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df.columns
# + id="3RMatp13cyWf" colab_type="code" colab={}
# + [markdown] id="dMLCm2w0dKY7" colab_type="text"
# #Dummy Model
# + id="SUEe-JGsdMet" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ef74aeb-64d5-47f8-b5e6-e08390bb5db4" executionInfo={"status": "ok", "timestamp": 1583441342293, "user_tz": -60, "elapsed": 1073, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df.select_dtypes(np.number).columns
# + id="5MYto8OadWkL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85a2c69a-90d2-491f-e361-849da1134e16" executionInfo={"status": "ok", "timestamp": 1583441745531, "user_tz": -60, "elapsed": 924, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
feats = ['car_id']
X = df[feats].values
y = df['price_value'].values
model = DummyRegressor()
model.fit(X,y)
y_pred = model.predict(X)
mae(y, y_pred)
# + id="u2EA_84pd6FI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8fcfedcb-d825-4ca2-a8da-20b04ce91583" executionInfo={"status": "ok", "timestamp": 1583441801574, "user_tz": -60, "elapsed": 618, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
[x for x in df.columns if 'price' in x]
# + id="g_HdOI7WfGzH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="efcafeef-abb3-49bd-d7da-7eff05788fbb" executionInfo={"status": "ok", "timestamp": 1583443323672, "user_tz": -60, "elapsed": 1114, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df['price_currency'].value_counts()
# + id="H-_IgWx-fM14" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f19d7584-947c-471f-dae6-9b111cd92561" executionInfo={"status": "ok", "timestamp": 1583443306854, "user_tz": -60, "elapsed": 1194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df = df [ df['price_currency'] != 'EUR']
df.shape
# + id="C1NXqeV3fxKw" colab_type="code" colab={}
# + [markdown] id="BcSZ7vr2fzlP" colab_type="text"
# ## Features
# + id="gKFrulQDf1gt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 678} outputId="a8a8e729-0d5c-468f-ddaf-bfa5f3da9298" executionInfo={"status": "ok", "timestamp": 1583442007257, "user_tz": -60, "elapsed": 844, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df.head()
# + id="7p8_YGUdf3g3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="933b4a27-9bbb-4e74-9c6d-d956c4c014e4" executionInfo={"status": "ok", "timestamp": 1583442045191, "user_tz": -60, "elapsed": 874, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
for feat in df.columns:
print(feat)
# + id="v2mC13y6gCGq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="204448a7-9e6a-475b-8a55-6a5549060287" executionInfo={"status": "ok", "timestamp": 1583442118566, "user_tz": -60, "elapsed": 568, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
df['param_color'].factorize()[0]
# + id="irJH3yrSgMZO" colab_type="code" colab={}
SUFFIX_CAT= '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list):continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="od6NrpAph4iI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b03b5f99-1cb8-480c-8ae8-4dc8ac723dff" executionInfo={"status": "ok", "timestamp": 1583442709752, "user_tz": -60, "elapsed": 522, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="1C_4noc5iU9U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="52d59999-2513-47cb-d226-61cd7077e366" executionInfo={"status": "ok", "timestamp": 1583443051085, "user_tz": -60, "elapsed": 5131, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores)
# + id="Ao3UuSIijJFE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="c21d75ed-ba1f-421f-bd91-db20d312ad2c" executionInfo={"status": "ok", "timestamp": 1583443237680, "user_tz": -60, "elapsed": 45722, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GikLv61bRiGYsmKocZbXiUI0qGst0kyU2l-F7j9Tg=s64", "userId": "14751556941823709984"}}
m = DecisionTreeRegressor(max_depth=5)
m.fit(X,y)
imp = PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="TULu85XhkLZt" colab_type="code" colab={}
| day3_simple_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ATACseq_clustering]
# language: python
# name: conda-env-ATACseq_clustering-py
# ---
import pandas as pd
import numpy as np
import scanpy as sc
import os
from sklearn.cluster import KMeans
from sklearn.cluster import AgglomerativeClustering
from sklearn.metrics.cluster import adjusted_rand_score
from sklearn.metrics.cluster import adjusted_mutual_info_score
from sklearn.metrics.cluster import homogeneity_score
import rpy2.robjects as robjects
from rpy2.robjects import pandas2ri
df_metrics = pd.DataFrame(columns=['ARI_Louvain','ARI_kmeans','ARI_HC',
'AMI_Louvain','AMI_kmeans','AMI_HC',
'Homogeneity_Louvain','Homogeneity_kmeans','Homogeneity_HC'])
workdir = './peaks_intensity_results/'
path_fm = os.path.join(workdir,'feature_matrices/')
path_clusters = os.path.join(workdir,'clusters/')
path_metrics = os.path.join(workdir,'metrics/')
os.system('mkdir -p '+path_clusters)
os.system('mkdir -p '+path_metrics)
metadata = pd.read_csv('../../input/metadata.tsv',sep='\t',index_col=0)
num_clusters = len(np.unique(metadata['label']))
print(num_clusters)
files = [x for x in os.listdir(path_fm) if x.startswith('FM')]
len(files)
files
def getNClusters(adata,n_cluster,range_min=0,range_max=3,max_steps=20):
this_step = 0
this_min = float(range_min)
this_max = float(range_max)
while this_step < max_steps:
print('step ' + str(this_step))
this_resolution = this_min + ((this_max-this_min)/2)
sc.tl.louvain(adata,resolution=this_resolution)
this_clusters = adata.obs['louvain'].nunique()
print('got ' + str(this_clusters) + ' at resolution ' + str(this_resolution))
if this_clusters > n_cluster:
this_max = this_resolution
elif this_clusters < n_cluster:
this_min = this_resolution
else:
return(this_resolution, adata)
this_step += 1
print('Cannot find the number of clusters')
print('Clustering solution from last iteration is used:' + str(this_clusters) + ' at resolution ' + str(this_resolution))
for file in files:
file_split = file[:-4].split('_')
method = file_split[1]
print(method)
pandas2ri.activate()
readRDS = robjects.r['readRDS']
df_rds = readRDS(os.path.join(path_fm,file))
fm_mat = pandas2ri.ri2py(robjects.r['data.frame'](robjects.r['as.matrix'](df_rds)))
fm_mat.fillna(0,inplace=True)
fm_mat.columns = metadata.index
adata = sc.AnnData(fm_mat.T)
adata.var_names_make_unique()
adata.obs = metadata.loc[adata.obs.index,]
df_metrics.loc[method,] = ""
#Louvain
sc.pp.neighbors(adata, n_neighbors=15,use_rep='X')
# sc.tl.louvain(adata)
getNClusters(adata,n_cluster=num_clusters)
#kmeans
kmeans = KMeans(n_clusters=num_clusters, random_state=2019).fit(adata.X)
adata.obs['kmeans'] = pd.Series(kmeans.labels_,index=adata.obs.index).astype('category')
#hierachical clustering
hc = AgglomerativeClustering(n_clusters=num_clusters).fit(adata.X)
adata.obs['hc'] = pd.Series(hc.labels_,index=adata.obs.index).astype('category')
#clustering metrics
#adjusted rank index
ari_louvain = adjusted_rand_score(adata.obs['label'], adata.obs['louvain'])
ari_kmeans = adjusted_rand_score(adata.obs['label'], adata.obs['kmeans'])
ari_hc = adjusted_rand_score(adata.obs['label'], adata.obs['hc'])
#adjusted mutual information
ami_louvain = adjusted_mutual_info_score(adata.obs['label'], adata.obs['louvain'],average_method='arithmetic')
ami_kmeans = adjusted_mutual_info_score(adata.obs['label'], adata.obs['kmeans'],average_method='arithmetic')
ami_hc = adjusted_mutual_info_score(adata.obs['label'], adata.obs['hc'],average_method='arithmetic')
#homogeneity
homo_louvain = homogeneity_score(adata.obs['label'], adata.obs['louvain'])
homo_kmeans = homogeneity_score(adata.obs['label'], adata.obs['kmeans'])
homo_hc = homogeneity_score(adata.obs['label'], adata.obs['hc'])
df_metrics.loc[method,['ARI_Louvain','ARI_kmeans','ARI_HC']] = [ari_louvain,ari_kmeans,ari_hc]
df_metrics.loc[method,['AMI_Louvain','AMI_kmeans','AMI_HC']] = [ami_louvain,ami_kmeans,ami_hc]
df_metrics.loc[method,['Homogeneity_Louvain','Homogeneity_kmeans','Homogeneity_HC']] = [homo_louvain,homo_kmeans,homo_hc]
adata.obs[['louvain','kmeans','hc']].to_csv(os.path.join(path_clusters ,method + '_clusters.tsv'),sep='\t')
df_metrics.to_csv(path_metrics+'clustering_scores.csv')
df_metrics
| Extra/BoneMarrow_noisy_p2/test_peaks/Cusanovich2018/run_clustering_intensity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # $\nu$-electron scattering
#
# Testing the neutrino-electron scattering rate calculation for Xenon. We use the free electron approximation but also include a stepping process to approximate the impact of the atomic ionisation energy levels which suppress the rate at very low recoil energies.
#
# First we look at the energy levels in Xenon. The approximation works by applying a heaviside function to the free electron neutrino-electron cross section de-weight those electrons that can't be ionised by the incoming neutrino with a defined energy. i.e.
#
# \begin{equation}
# \frac{\textrm{d}\sigma^i}{\textrm{d} E_R} = \sum\limits_{k=1}^{Z} \Theta(E_R-E^k_B) \frac{\textrm{d}\sigma^i_0}{\textrm{d}E_R}
# \end{equation}
#
# where $i$ refers to neutrino flavour, $\sigma_0$ is the cross section for a free electron, and $E^k_B$ is the ionisation energy of the $k$th level.
# ### First we take a look at the xenon energy levels:
# +
import sys
sys.path.append('../erec')
from numpy import *
import matplotlib.pyplot as plt
from Params import *
from LabFuncs import *
from DMFuncs import *
from PlotFuncs import *
from NeutrinoFuncs import *
from AtomicFuncs import *
# Function for outputting the sum of the heaviside functions
def XeStep(E_r): # E_r is always in keV
Z = 54
# Xenon energy levels (in eV) from arXiv:1610.04177
E_B_vals = array([34759.3]*2\
+[5509.8]*2+[5161.5]*2+[4835.6]*4\
+[1170.5]*2+[1024.8]*2+[961.2]*4+[708.1]*2+[694.9]*8\
+[229.4]*2+[175.6]*2+[162.8]*4+[73.8]*2+[71.7]*8\
+[27.5]*2+[13.4]*4+[12.0]*2)
TH = zeros(shape=shape(E_r))
for i in range(0,Z):
TH += 1.0*(E_r>E_B_vals[i]/1000.0)
# Neutrino rate later multiplies by Z so we divide by it here to not overcount
TH = TH/(1.0*Z)
return TH,E_B_vals
# Set up plot
fig,ax = MySquarePlot(r'$E_R$ [keV]',r'$\frac{1}{Z}\sum\limits_{k=1}^{Z} k \Theta(E_R-E^k_B)$',lfs=40,tfs=30)
# Plot step functions
ne = 2000
E_r = logspace(-1,3,ne)
st,E_B_vals = XeStep(E_r)
plt.step(E_r,st,'-',lw=5,color='firebrick')
E_prev = 0.0
for i in range(0,54):
if (E_B_vals[i]!=E_prev)&(E_B_vals[i]/1000.0>E_r[0]):
plt.plot(E_B_vals[i]*array([1,1])/1000.0,[0.0,1.1],'k-',lw=3,alpha=0.1)
E_prev = 1.0*E_B_vals[i]
# Add shell labels
plt.text(34759.3/1000.0*0.9,1.11,'K',fontsize=30)
plt.text(5509.8/1000.0*0.75,1.11,'L$_{1-3}$',fontsize=30)
plt.text(1170.5/1000.0*0.6,1.11,'M$_{1-5}$',fontsize=30)
plt.text(229.4/1000.0*0.65,1.11,'N$_{1-3}$',fontsize=30)
# Plot tweaking
#plt.yscale('log')
plt.xlim([E_r[0],E_r[-1]])
plt.ylim([0.0,1.1])
plt.xscale('log')
plt.gcf().text(0.64,0.17,r'{\bf Xenon} $Z = 54$',fontsize=35)
# Show and save
plt.show()
pltname = "XenonStep"
fig.savefig('../plots/'+pltname+'.pdf',bbox_inches='tight')
fig.savefig('../plots/plots_png/'+pltname+'.png',bbox_inches='tight')
# -
# # Neutrino-electron recoil rates for the two approximations
# +
import cmocean
# Load neutrino fluxes
NuBG = GetNuFluxes(0.0,Xe131)
E_nu_all = NuBG.Energy
Flux_all = NuBG.Flux
n_nu = NuBG.NumberOfNeutrinos
Errs = NuBG.Uncertainties
solar = NuBG.SolarLabel
fig,ax = MySquarePlot(r'$E_R$ [keV]',r'$\textrm{d}R/\textrm{d}E_R$ [ton$^{-1}$ year$^{-1}$ keV$^{-1}$]',lfs=40,tfs=30)
ne = 5000
E_r = logspace(-4.0,3.2,ne)
TH,E_B_vals = XeStep(E_r)
ymax = 5.0
ymin = 3.0e-3
col = cmocean.cm.phase(linspace(0.1,0.8,3))
# pp
dR1 = dRdEe_nu(E_r,0.0,solar[0],E_nu_all[:,0],Flux_all[:,0],Xe131,flav_Solar)
plt.plot(E_r,dR1,'--',lw=3,color=col[0,:])
plt.plot(E_r,dR1*TH,'-',lw=3,color=col[0,:])
# 7Be
dR2 = dRdEe_nu(E_r,0.0,solar[3],E_nu_all[:,3],Flux_all[:,3],Xe131,flav_Solar)\
+dRdEe_nu(E_r,0.0,solar[4],E_nu_all[:,4],Flux_all[:,4],Xe131,flav_Solar)
plt.plot(E_r,dR2,'--',lw=3,color=col[1,:])
plt.plot(E_r,dR2*TH,'-',lw=3,color=col[1,:])
# CNO
dR3 = dRdEe_nu(E_r,0.0,solar[6],E_nu_all[:,6],Flux_all[:,6],Xe131,flav_Solar)\
+dRdEe_nu(E_r,0.0,solar[7],E_nu_all[:,7],Flux_all[:,7],Xe131,flav_Solar)\
+dRdEe_nu(E_r,0.0,solar[8],E_nu_all[:,8],Flux_all[:,8],Xe131,flav_Solar)
plt.plot(E_r,dR3,'--',lw=3,color=col[2,:])
plt.plot(E_r,dR3*TH,'-',lw=3,color=col[2,:])
# pep
# dR4 = dRdEe_nu(E_r,0.0,solar[1],E_nu_all[:,1],Flux_all[:,1],Xe131,flav_Solar)
# plt.plot(E_r,dR4,'--',lw=3,color=col[3,:])
# plt.plot(E_r,dR4*TH,'-',lw=3,color=col[3,:])
plt.plot(-E_r,dR1+dR2+dR3,'k--',lw=3,label='Free electron approx.')
plt.plot(-E_r,(dR1+dR2+dR3),'k-',lw=3,label='Free electron approx. with energy levels')
plt.legend(frameon=False,fontsize=30,borderpad=0.1,handlelength=1)
plt.gcf().text(0.15,0.43,'CNO',color=col[2,:])
plt.gcf().text(0.15,0.6,'$^7$Be',color=col[1,:])
plt.gcf().text(0.15,0.81,'$pp$',color=col[0,:])
plt.gcf().text(0.75,0.82,r'{\bf Xenon}',fontsize=35)
plt.xscale('log')
plt.yscale('log')
plt.ylim([5.0e-4,10.0])
plt.xlim([0.1,2000.0])
plt.show()
pltname = "NuRate_Xe_electrons"
fig.savefig('../plots/'+pltname+'.pdf',bbox_inches='tight')
fig.savefig('../plots/plots_png/'+pltname+'.png',bbox_inches='tight')
# -
# # Does a finite energy resolution remove these features?
#
# Energy resolution in xenon detectors looks something like,
# \begin{equation}
# \frac{\sigma}{E_R} = c_1\sqrt{\frac{1 {\rm keV}}{E_R}} + c_2
# \end{equation}
# From arXiv:1610.02076 the fit to known peaks in LUX's ER background has $c_1 = 0.33\pm0.01$ and $c_2\simeq0.0$. For results to come we should be using the 2d (S1,S2) pdf for ERs but for now we just want to see how it looks.
# +
fig,ax = MySquarePlot(r'$E_R$ [keV]',r'$\textrm{d}R/\textrm{d}E_R$ [ton$^{-1}$ year$^{-1}$ keV$^{-1}$]',lfs=40,tfs=30)
n_c1_vals = 10
c1_vals = linspace(0.05,0.5,n_c1_vals)
dR0 = TH*(dR1+dR2+dR3)
col = cmocean.cm.balance(linspace(0.1,1.0,n_c1_vals))
for i in flipud(range(0,n_c1_vals)):
c1 = c1_vals[i]
c2 = 0.00
sig_E = c1*sqrt(E_r)+c2*E_r
dR = SmearE(E_r,dR0,sig_E)
plt.fill_between(E_r,dR,y2=dR0,color=col[i,:],alpha=0.5)
plt.plot(E_r,dR,'-',color=col[i,:])
plt.plot(E_r,dR0,'k-',lw=3)
plt.plot(E_r,(dR1+dR2+dR3),'k--',lw=3)
plt.gcf().text(0.15,0.15,r'$\frac{\sigma}{E_R} = c_1\sqrt{\frac{1 {\rm keV}}{E_R}} + c_2$',fontsize=50)
plt.gcf().text(0.15,0.82,r'$pp$+$^7$Be+CNO',fontsize=35)
plt.gcf().text(0.75,0.82,r'{\bf Xenon}',fontsize=35)
plt.gcf().text(0.15,0.65,r'Free electron',fontsize=35)
plt.xscale('log')
plt.yscale('log')
plt.xlim([0.1,40.0])
plt.ylim([5.0e-1,8.0])
# Custom colorbar
import matplotlib as mpl
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
cbaxes = inset_axes(ax, width="15%", height="20%", bbox_to_anchor=[560, -650, 200, 1000])
norm = mpl.colors.Normalize(vmin=c1_vals[0],vmax=c1_vals[-1])
sm = plt.cm.ScalarMappable(cmap=cmocean.cm.balance, norm=norm)
sm.set_array([])
plt.colorbar(sm,cax=cbaxes,ticks=arange(c1_vals[0],c1_vals[-1],0.1),boundaries=c1_vals,orientation='vertical')
f = plt.gcf().get_children()
cbar = f[2]
cbar.tick_params(labelsize=25)
cbar.tick_params(which='major',direction='out',width=2,length=10,right=True,top=False,pad=7)
cbar.tick_params(which='minor',direction='out',width=2,length=7,right=True,top=False)
plt.gcf().text(0.73,0.26,r'$c_1$',fontsize=40,color='k')
plt.show()
pltname = "NuRate_Xe_electrons_vs_resolution"
fig.savefig('../plots/'+pltname+'.pdf',bbox_inches='tight')
fig.savefig('../plots/plots_png/'+pltname+'.png',bbox_inches='tight')
# -
| notebooks/NeutrinoRates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multigrid and MgNet
# +
from IPython.display import IFrame
IFrame(src="https://cdnapisec.kaltura.com/p/2356971/sp/235697100/embedIframeJs/uiconf_id/41416911/partner_id/2356971?iframeembed=true&playerId=kaltura_player&entry_id=0_j0az3ah8&flashvars[streamerType]=auto&flashvars[localizationCode]=en&flashvars[leadWithHTML5]=true&flashvars[sideBarContainer.plugin]=true&flashvars[sideBarContainer.position]=left&flashvars[sideBarContainer.clickToClose]=true&flashvars[chapters.plugin]=true&flashvars[chapters.layout]=vertical&flashvars[chapters.thumbnailRotator]=false&flashvars[streamSelector.plugin]=true&flashvars[EmbedPlayer.SpinnerTarget]=videoHolder&flashvars[dualScreen.plugin]=true&flashvars[hotspots.plugin]=1&flashvars[Kaltura.addCrossoriginToIframe]=true&&wid=1_df2css9s",width='800', height='500')
# -
# This lecture includes:
#
# * Multigrid
# * MgNet
#
# ## 1. A framework of both Multigrid and MgNet
#
# * Comment out the code for MgNet, you will obtain the multigrid code.
# * Comment out the code for MG, you will obtain the MgNet code.
#
# +
import torch
import numpy as np
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
from timeit import default_timer as timer
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
use_cuda = torch.cuda.is_available()
print('Use GPU?', use_cuda)
##### For MG: inilization of A, S, Pi, R, RT #####
def get_mg_init(A=None, S=None, Pi=None, R=None, RT=None):
A_kernel = torch.tensor([[[[0,-1,0],[-1,4,-1],[0,-1,0]]]],dtype=torch.float32)
S_kernel = torch.tensor([[[[0,1/64,0],[1/64,12/64,1/64],[0,1/64,0]]]],dtype=torch.float32)
Pi_kernel = torch.tensor([[[[0,0,0],[0,0,0],[0,0,0]]]],dtype=torch.float32)
R_kernel = torch.tensor([[[[0,0.5,0.5],[0.5,1,0.5],[0.5,0.5,0]]]],dtype=torch.float32)
RT_kernel = torch.tensor([[[[0,0.5,0.5],[0.5,1,0.5],[0.5,0.5,0]]]],dtype=torch.float32)
if A is not None:
A.weight = torch.nn.Parameter(A_kernel)
if S is not None:
S.weight = torch.nn.Parameter(S_kernel)
if Pi is not None:
Pi.weight = torch.nn.Parameter(Pi_kernel)
if R is not None:
R.weight = torch.nn.Parameter(R_kernel)
if RT is not None:
RT.weight = torch.nn.Parameter(RT_kernel)
return
##### For MG: setup for prolongation and error calculation #####
RT = nn.ConvTranspose2d(1, 1, kernel_size=3, stride=2, padding=0, bias=False)
get_mg_init(None,None,None,None,RT)
A = nn.Conv2d(1, 1, kernel_size=3,stride=1, padding=1, bias=False)
get_mg_init(A,None,None,None,None)
class MgIte(nn.Module):
def __init__(self, A, S):
super().__init__()
get_mg_init(A=A,S=S) ##### For MG: inilization of A, S #####
self.A = A
self.S = S
self.bn1 =nn.BatchNorm2d(A.weight.size(0)) ##### For MgNet: BN #####
self.bn2 =nn.BatchNorm2d(S.weight.size(0)) ##### For MgNet: BN #####
def forward(self, out):
u, f = out
u = u + (self.S(((f-self.A(u))))) ##### For MG: u = u + S*(f-A*u) #####
u = u + F.relu(self.bn2(self.S(F.relu(self.bn1((f-self.A(u))))))) ##### For MgNet: add BN and ReLU #####
out = (u, f)
return out
class MgRestriction(nn.Module):
def __init__(self, A_old, A, Pi, R):
super().__init__()
get_mg_init(A=A,Pi=Pi,R=R) ##### For MG: inilization of A, Pi, R #####
self.A_old = A_old
self.A = A
self.Pi = Pi
self.R = R
self.bn1 = nn.BatchNorm2d(Pi.weight.size(0)) ##### For MgNet: BN #####
self.bn2 = nn.BatchNorm2d(R.weight.size(0)) ##### For MgNet: BN #####
def forward(self, out):
u_old, f_old = out
u = self.Pi(u_old) ##### For MG: u = Pi*u_old #####
f = self.R(f_old-self.A_old(u_old)) + self.A(u) ##### For MG: f = R*(f_old-A_old*u_old) + A*u #####
u = F.relu(self.bn1(self.Pi(u_old))) ##### For MgNet: add BN and ReLU #####
f = F.relu(self.bn2(self.R(f_old-self.A_old(u_old)))) + self.A(u) ##### For MgNet: add BN and ReLU #####
out = (u,f)
return out
class MG(nn.Module):
def __init__(self, num_channel_input, num_iteration, num_channel_u, num_channel_f, num_classes):
super().__init__()
self.num_iteration = num_iteration
self.num_channel_u = num_channel_u
##### For MgNet: Initialization layer #####
self.conv1 = nn.Conv2d(num_channel_input, num_channel_f, kernel_size=3, stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(num_channel_f)
A = nn.Conv2d(num_channel_u, num_channel_f, kernel_size=3, stride=1, padding=1, bias=False)
S = nn.Conv2d(num_channel_f, num_channel_u, kernel_size=3,stride=1, padding=1, bias=False)
layers = []
for l, num_iteration_l in enumerate(num_iteration): #l: l-th layer. num_iteration_l: the number of iterations of l-th layer
for i in range(num_iteration_l):
layers.append(MgIte(A, S))
setattr(self, 'layer'+str(l), nn.Sequential(*layers))
# set attribute. This is equivalent to define
# self.layer1 = nn.Sequential(*layers)
# self.layer2 = nn.Sequential(*layers)
# ...
# self.layerJ = nn.Sequential(*layers)
if l < len(num_iteration)-1:
A_old = A
A = nn.Conv2d(num_channel_u, num_channel_f, kernel_size=3,stride=1, padding=1, bias=False)
S = nn.Conv2d(num_channel_f, num_channel_u, kernel_size=3,stride=1, padding=1, bias=False)
##### For MG: padding=0 #####
Pi = nn.Conv2d(num_channel_u, num_channel_u, kernel_size=3,stride=2, padding=0, bias=False)
R = nn.Conv2d(num_channel_f, num_channel_f, kernel_size=3, stride=2, padding=0, bias=False)
##### For MgNet: padding=1 #####
Pi = nn.Conv2d(num_channel_u, num_channel_u, kernel_size=3,stride=2, padding=1, bias=False)
R = nn.Conv2d(num_channel_f, num_channel_f, kernel_size=3, stride=2, padding=1, bias=False)
layers= [MgRestriction(A_old, A, Pi, R)]
##### For MgNet: average pooling and fully connected layer for classification #####
self.pooling = nn.AdaptiveAvgPool2d(1) # pooling the data in each channel to size=1
self.fc = nn.Linear(num_channel_u ,num_classes)
def forward(self, u, f):
f = F.relu(self.bn1(self.conv1(f))) ##### For MgNet: initialization of f #####
if use_cuda: ##### For MgNet: initialization of u #####
u = torch.zeros(f.size(0),self.num_channel_u,f.size(2),f.size(3), device=torch.device('cuda'))
else:
u = torch.zeros(f.size(0),self.num_channel_u,f.size(2),f.size(3))
out = (u, f)
u_list.append(u) ##### For MG: save u^j, j=1,2,...,J #####
for l in range(len(self.num_iteration)):
out = getattr(self, 'layer'+str(l))(out)
u, f = out ##### For MG: save u^j, j=1,2,...,J #####
u_list.append(u) ##### For MG: save u^j, j=1,2,...,J #####
##### For MgNet: average pooling and fully connected layer for classification #####
u, f = out
u = self.pooling(u)
u = u.view(u.shape[0], -1)
u = self.fc(u)
return u
# -
# ## 2. Apply Multigrid to solve the solving the following system
#
# \begin{equation}\label{matrix}
# A\ast u =f,
# \end{equation}
# where $A\ast$ is a convolution for one channel with stride 1 and zero padding $1$
# $$
# A=\begin{bmatrix} 0 & -1 & 0 \\ -1 & 4 & -1 \\ 0 & 1 & 0 \end{bmatrix},~~
# $$
# and
# $ u \in \mathbb{R}^{n\times n} $, $ f\in \mathbb{R}^{n\times n}$ and $f_{i,j}=\dfrac{1}{(n+1)^2}$
#
#
# ### Mulrigrid code includes: (a) comment out the code in 1 for MgNet; (b) the setup and postprocessing code below
# +
def plot_solution(J,u,label_name):
N = 2 ** J -1
h = 1/2**J
X = np.arange(h, 1, h)
Y = np.arange(h, 1, h)
X, Y = np.meshgrid(X,Y) # create a mesh
a = torch.reshape(u, (N, N))
fig1 = plt.figure()
ax = Axes3D(fig1) # plot a 3D surface, (X,Y,u(X,Y))
ax.plot_surface(X, Y, np.array(a.data), rstride=1, cstride=1, cmap=plt.cm.coolwarm)
ax.set_title(label_name)
def plot_error(M,error,label_name):
#print(np.linalg.norm((f-self.A(u)).reshape(-1).detach().numpy()))
plt.figure()
plt.title('Error vs number of iterations using '+label_name)
plot = plt.plot(error)
plt.xlabel('Number of iterations')
plt.yscale('log')
plt.ylabel('Error')
plt.show()
# -
def MG1(u,f,J,num_iteration):
u_list.clear() # Save u^0,u^1,u^2,u^3...,u^J
u = MG0(u,f)
for j in range(J-1,0,-1):
u_list[j] += RT(u_list[j+1])
u = u_list[1]
return u
# +
# Model setup
num_channel_input = 1
num_channel_u = 1
num_channel_f = 1
num_classes = 1
J = 4
num_iteration = [2,2,2,2]
MG0=MG(num_channel_input, num_iteration, num_channel_u, num_channel_f, num_classes)
##### For MG: PDE setup u=sin(2*pi*x)*sin(2*pi*y) #####
N = 2 ** J -1
h = 1/2**J
u_exact = torch.ones(1,1,N,N)
f = torch.ones(1,1,N,N) / (N+1) **2
##### For MG: Muligrid iteration #####
M = 100
u = torch.randn(1,1,N,N)
error = [np.linalg.norm((A(u)-f).detach().numpy())] # calculate the Frobenius Norm of (A*u-f)
u_list =[] # Save u^0,u^1,u^2,u^3...,u^J
for m in range(M):
u = MG1(u,f,J,num_iteration)
error.append(np.linalg.norm((A(u)-f).detach().numpy())) # calculate the Frobenius Norm of (A*u-f)
##### Lian added for MG: Plot results #####
plot_error(M,error,'Multigrid')
plot_solution(J,u,'Numerical solution')
# -
# ## 3. Build and training MgNet on Cifar10
# ### MgNet code includes: (a) comment out the code in 1 for Multigrid; (b) the setup, training and test code below
# +
def adjust_learning_rate(optimizer, epoch, init_lr):
#lr = 1.0 / (epoch + 1)
lr = init_lr * 0.1 ** (epoch // 30)
for param_group in optimizer.param_groups:
param_group['lr'] = lr
return lr
minibatch_size = 128
num_epochs = 120
lr = 0.1
num_channel_input = 3
num_channel_u = 64
num_channel_f = 64
num_classes = 10
num_iteration = [1,1,1,1]
# Step 1: Define a model
my_model = MgNet(num_channel_input, num_iteration, num_channel_u, num_channel_f, num_classes)
if use_cuda:
my_model = my_model.cuda()
# Step 2: Define a loss function and training algorithm
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(my_model.parameters(), lr=lr, momentum=0.9, weight_decay = 0.0005)
# Step 3: load dataset
normalize = torchvision.transforms.Normalize(mean=(0.4914, 0.4822, 0.4465), std=(0.2023, 0.1994, 0.2010))
transform_train = torchvision.transforms.Compose([torchvision.transforms.RandomCrop(32, padding=4),
torchvision.transforms.RandomHorizontalFlip(),
torchvision.transforms.ToTensor(),
normalize])
transform_test = torchvision.transforms.Compose([torchvision.transforms.ToTensor(),normalize])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform_train)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=minibatch_size, shuffle=True)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform_test)
testloader = torch.utils.data.DataLoader(testset, batch_size=minibatch_size, shuffle=False)
# classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
start = timer()
#Step 4: Train the NNs
# One epoch is when an entire dataset is passed through the neural network only once.
for epoch in range(num_epochs):
start_epoch = timer()
current_lr = adjust_learning_rate(optimizer, epoch, lr)
start_training = timer()
my_model.train()
for i, (images, labels) in enumerate(trainloader):
if use_cuda:
images = images.cuda()
labels = labels.cuda()
# Forward pass to get the loss
outputs = my_model(0,images) # We need additional 0 input for u in MgNet
loss = criterion(outputs, labels)
# Backward and compute the gradient
optimizer.zero_grad()
loss.backward() #backpropragation
optimizer.step() #update the weights/parameters
end_training = timer()
print('Computation Time for training:',end_training - start_training)
# Training accuracy
start_training_acc = timer()
my_model.eval()
correct = 0
total = 0
for i, (images, labels) in enumerate(trainloader):
with torch.no_grad():
if use_cuda:
images = images.cuda()
labels = labels.cuda()
outputs = my_model(0,images) # We need additional 0 input for u in MgNet
p_max, predicted = torch.max(outputs, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
training_accuracy = float(correct)/total
end_training_acc = timer()
print('Computation Time for training accuracy:',end_training_acc - start_training_acc)
# Test accuracy
start_test_acc = timer()
correct = 0
total = 0
for i, (images, labels) in enumerate(testloader):
with torch.no_grad():
if use_cuda:
images = images.cuda()
labels = labels.cuda()
outputs = my_model(0,images) # We need additional 0 input for u in MgNet
p_max, predicted = torch.max(outputs, 1)
total += labels.size(0)
correct += (predicted == labels).sum()
test_accuracy = float(correct)/total
end_test_acc = timer()
print('Computation Time for test accuracy:',end_test_acc - start_test_acc)
print('Epoch: {}, learning rate: {}, the training accuracy: {}, the test accuracy: {}' .format(epoch+1,current_lr,training_accuracy,test_accuracy))
end_epoch = timer()
print('Computation Time for one epoch:',end_epoch - start_epoch)
end = timer()
print('Total Computation Time:',end - start)
# -
| _build/jupyter_execute/Module6/m6_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sentencepiece as spm
import pandas as pd
import MeCab
# ## 조사 제거해서 문장 반환하는 함수
dict_list
# +
m = MeCab.Tagger()
delete_tag = ['BOS/EOS', 'JKS', 'JKC', 'JKG', 'JKO', 'JKB', 'JKV', 'JKQ', 'JX', 'JC']
'''
"JKS": "주격 조사",
"JKC": "보격 조사",
"JKG": "관형격 조사",
"JKO": "목적격 조사",
"JKB": "부사격 조사",
"JKV": "호격 조사",
"JKQ": "인용격 조사",
"JX": "보조사",
"JC": "접속 조사",
'''
def remove_josa(sentence):
sentence_split = sentence.split() # 원본 문장 띄어쓰기로 분리
dict_list = []
for token in sentence_split: # 띄어쓰기로 분리된 각 토큰 {'단어':'형태소 태그'} 와 같이 딕셔너리 생성
m.parse('')
node = m.parseToNode(token)
word_list = []
pos_list = []
while node:
morphs = node.feature.split(',')
word_list.append(node.surface)
pos_list.append(morphs[0])
node = node.next
dict_list.append(dict(zip(word_list, pos_list)))
for dic in dict_list: # delete_tag에 해당하는 단어 쌍 지우기 (조사에 해당하는 단어 지우기)
for key in list(dic.keys()):
if dic[key] in delete_tag:
del dic[key]
combine_word = [''.join(list(dic.keys())) for dic in dict_list] # 형태소로 분리된 각 단어 합치기
result = ' '.join(combine_word) # 띄어쓰기로 분리된 각 토큰 합치기
return result # 온전한 문장을 반환
# -
# 예시 ) 너는
#
# --node--
# morphs: ['BOS/EOS', '*', '*', '*', '*', '*', '*', '*']
# word_list : ['']
# pos_list : ['BOS/EOS']
#
# --node--
# morphs: ['NP', '*', 'F', '너', '*', '*', '*', '*']
# word_list : ['', '너']
# pos_list : ['BOS/EOS', 'NP']
#
# --node--
# morphs: ['JX', '*', 'T', '는', '*', '*', '*', '*']
# word_list : ['', '너', '는']
# pos_list : ['BOS/EOS', 'NP', 'JX']
#
# --node--
# morphs: ['BOS/EOS', '*', '*', '*', '*', '*', '*', '*']
# word_list : ['', '너', '는', '']
# pos_list : ['BOS/EOS', 'NP', 'JX', 'BOS/EOS']
# dict_list, [{'': 'BOS/EOS', '너': 'NP', '는': 'JX'}]
# ## sentencepiece 적용 (original과 조사 제거 버전 비교)
KOR_data = pd.read_csv("C:/Users/<NAME>/Desktop/NMT Project/dataset/datalist_striped.csv", encoding = 'utf-8-sig')
KOR_data = KOR_data['Korean']
# +
f1 = open("original.txt", "w", encoding = 'utf-8')
f2 = open("no_josa.txt", "w", encoding = 'utf-8')
for row in KOR_data[:100000]:
f1.write(row) # 조사 제거 안한 원본 문장 저장
f1.write('\n')
f2.write(remove_josa(row)) # 조사 제거한 문장 저장
f2.write('\n')
#f.close()
#f2.close()
# +
# original 모델 생성 (조사 제거안한 문장으로 학습)
spm.SentencePieceTrainer.Train('--input=original.txt \
--model_prefix=original \
--vocab_size=100000 \
--hard_vocab_limit=false')
# 조사 제거한 문장으로 모델 생성
spm.SentencePieceTrainer.Train('--input=no_josa.txt \
--model_prefix=revise \
--vocab_size=100000 \
--hard_vocab_limit=false')
# +
#모델 불러오기
sp1 = spm.SentencePieceProcessor()
sp1.Load('original.model')
sp2 = spm.SentencePieceProcessor()
sp2.Load('revise.model')
# +
# input setence
sentence = "시 관계자는 선별진료소, 의료기관 등에 개인보호구인 덧신, 장갑·고글, 마스크 800개씩을 배포했지만 물량이 턱없이 부족하다며 정부에 추가 지원을 요청했다."
# 0. 원본 문장
print("<원본 문장>")
print(sentence+'\n')
# 1. original 모델 (조사 제거 안한 문장으로 학습)
print("<original 모델>")
print(sp1.EncodeAsPieces(sentence))
# 1. 조사 제거 모델 (조사 제거 한 문장으로 학습)
print("\n<조사 제거 모델>")
print(sp2.EncodeAsPieces(sentence))
| pre-processing codes/.ipynb_checkpoints/조사 제거, sentencepiece 적용-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Combining Dataframes using merge()
import pandas as pd
# Read the dataset
data=pd.read_csv('bigmart_data.csv')
# Counting the number of rows and columns of the dataframe.
data_rows = len(data.index)
data_columns = len(data.columns)
data_rows, data_columns
data.head()
# Splitting the data
data_part1 = data[:100]
data_part2 = data[100:200]
data_part1.head()
data_part2.head()
data_part1_rows = len(data_part1.index)
data_part1_columns = len(data_part1.columns)
data_part1_rows, data_part1_columns
data_part2_rows = len(data_part2.index)
data_part2_columns = len(data_part2.columns)
data_part2_rows, data_part2_columns
# Merging dataframes with merge() and join='outer'
combined_data1 = pd.merge(data_part1, data_part2, on='Item_Fat_Content', how='outer')
combined_data1
combined_data1_rows = len(combined_data1.index)
combined_data1_columns = len(combined_data1.columns)
combined_data1_rows, combined_data1_columns
# Merging dataframes with merge() and join='inner'
combined_data2 = pd.merge(data_part1, data_part2, on='Item_Fat_Content', how='inner')
combined_data2
combined_data2_rows = len(combined_data2.index)
combined_data2_columns = len(combined_data2.columns)
combined_data2_rows, combined_data2_columns
# Merging dataframes with merge() and join='outer'
combined_data3 = pd.merge(data_part1, data_part2, on='Item_Type', how='outer')
combined_data3
combined_data3_rows = len(combined_data3.index)
combined_data3_columns = len(combined_data3.columns)
combined_data3_rows, combined_data3_columns
# Merging dataframes with merge() and join='inner'
combined_data4 = pd.merge(data_part1, data_part2, on='Item_Type', how='inner')
combined_data4
combined_data4_rows = len(combined_data4.index)
combined_data4_columns = len(combined_data4.columns)
combined_data4_rows, combined_data4_columns
# Note that data_part1 and data_part2 have the same shared keys on 'Item_Fat_Content'.
# But this is not the case on 'Item_Type'.
# So joining 'outer' and 'inner' causes differnt effects.
# There are some keys that either of the data_part1 or data_par2 don't have.
shared_keys1 = combined_data1_rows - combined_data2_rows
shared_keys2 = combined_data3_rows - combined_data4_rows
shared_keys1, shared_keys2
# Finding these different keys:
item_type_keys_of_data_part1 = data_part1['Item_Type'].unique()
item_type_keys_of_data_part1
len(item_type_keys_of_data_part1)
item_type_keys_of_data_part2 = data_part2['Item_Type'].unique()
item_type_keys_of_data_part2
len(item_type_keys_of_data_part2)
for item1 in item_type_keys_of_data_part1:
for item2 in item_type_keys_of_data_part2:
if item1 == item2:
print(item1)
for item2 in item_type_keys_of_data_part2:
for item1 in item_type_keys_of_data_part1:
if item2 == item1:
print(item2)
for item1 in item_type_keys_of_data_part1:
if item1 not in item_type_keys_of_data_part2:
print('!!!>>> "{}" is not present in data_part2<<<!!!'.format(item1))
else: print('"{}" is present in data_part2.'.format(item1))
# 'Others' is present in data_part2, but absent in data_part1.
for item2 in item_type_keys_of_data_part2:
if item2 not in item_type_keys_of_data_part1:
print('!!!>>> "{}" is not present in data_part1 <<<!!!'.format(item2))
else: print('"{}" is present in data_part1.'.format(item2))
# So 4 samples with 'Item_Type'='Others' are present in data_part2 that are absent in data_part1.
# These 4 samples point to the difference between join innerly and outerly, i.e. shared_keys2=4.
data_part2[data_part2['Item_Type'].str.match('Others')]
# Merging dataframes with merge() and join='inner'
# In this case, data_part1 and data_part2 only have one shared key, namely 'Item_MRP' = 222.5456.
combined_data5 = pd.merge(data_part1, data_part2, on='Item_MRP', how='inner')
combined_data5
combined_data5_rows = len(combined_data5.index)
combined_data5_columns = len(combined_data5.columns)
combined_data5_rows, combined_data5_columns
# And when we join the data parts outerly, we don't miss any data.
combined_data6 = pd.merge(data_part1, data_part2, on='Item_MRP', how='outer')
combined_data6
# 199 = 99 different 'Item_MRP's of data_part1 + 99 different 'Item_MRP's of data_part2 + 1 shared key (Item_MRP' = 222.5456)
combined_data6_rows = len(combined_data6.index)
combined_data6_columns = len(combined_data6.columns)
combined_data6_rows, combined_data6_columns
# Merging dataframes with merge() and join='left'
combined_data7 = pd.merge(data_part1, data_part2, on='Item_MRP', how='left')
combined_data7
# Merging dataframes with merge() and join='right'
combined_data8 = pd.merge(data_part1, data_part2, on='Item_MRP', how='right')
combined_data8
| Pandas_Data_Manipulation4.ipynb |