code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
# -*- coding: utf-8 -*-
"""
Created on Fri Jun 11 11:02:03 2021
@author: Christian
"""
import numpy as np
import pandas as pd
from mip import Model, xsum, maximize, BINARY, CBC
from Co2_price import df_data
from electricity_demand import electricity_demand
from Electricity_Price0 import Electricity_price_pivot0
from Electricity_Price1 import Electricity_price_pivot1
from gas_price import gas_p
from heat_demand import heat_demand_pivot
from heat_price import heat_p
from elec_effciency import elec_eff
from electricity_capacity import elec_capacity
from heat_capacity import heat_capacity
from heat_boiler_capacity import heat_cap
from heat_boiler_efficiency1 import heat_eff
elec_demand = electricity_demand.values.tolist()
elec_price_wholesale = Electricity_price_pivot0
elec_price_household_32_05 = Electricity_price_pivot1
co2_price = df_data
gas_price = gas_p
heat_demand_1 = heat_demand_pivot
heat_demand_1 = heat_demand_1.interpolate(method='linear', limit_direction='forward')
heat_price_1 = heat_p
el_price1 = elec_price_wholesale.interpolate(method='linear', limit_direction='forward')
el_price = el_price1.stack().tolist() # Electricity price
del_t = 1 # duration of time step
el_demand = electricity_demand.stack().tolist() # Electicity demand
gas_pp = gas_price["Preis"].values.tolist() # Gas price for different plants
em_fc = 0.2 # emission factor
co2_p = co2_price["mean_CO2_tax"].values.tolist() # CO2 Price
capacity_el = elec_capacity.stack().tolist() # maximum capacity of electricity for the power plants
capacity_ht = heat_capacity.stack().tolist() # maximum capacity of heat for the power plants
heat_demand = heat_demand_1.stack().tolist() # heat demand
heat_price = heat_price_1.stack().tolist() # heat price
heat_ratio = 400/385 # heat ratio
eff_plants = elec_eff.stack().tolist() # the efficiency of the power plants
capacity_ht_boiler = heat_cap.stack().tolist()
eff_boiler = heat_eff.stack().tolist()
T = range(len(8784))
I = range(len(1))
J = range(len(1)) #J: boiler in set of boilers J
K = range(len(1)) #k: CHP in set of CHP plants K
m = Model("Maximizing profit", sense=maximize, solver_name=CBC)
#variable
y_t = [[m.add_var(lb = 0) for i in I] for t in T] # Electricity generation
el_sold = [[m.add_var(lb = 0) for i in I] for t in T] # electricity sold
el_bought = [m.add_var(lb=0) for t in T] # electricity bought
x_t = [[m.add_var(lb = 0) for i in I] for t in T] # Fuel consumption
z_t = [[m.add_var(lb = 0) for i in I] for t in T] # Heat generation
# objective function
# Revenue = Electricity price * demand/generation
# Cost = Fuel price = gas price + emission_factor * co2 price
# Max Proft = Revenue - Cost
heat_demand_norm = [((element / max(heat_demand)) * 385) for element in heat_demand]
heat_demand = heat_demand_norm
print(len(co2_price))
print(len(el_demand))
print(len(gas_pp))
print(len(co2_p))
print(len(capacity_el))
print(len(capacity_ht))
print(len(heat_demand))
print(len(heat_price))
print(len(eff_plants))
print(len(capacity_ht_boiler))
print(len(eff_boiler))
m.objective = xsum(el_price[t] * del_t * (el_sold[t]-el_bought[t]) - (x_t[t][i] * (gas_pp[t] * del_t + (em_fc * co2_p[t] * del_t)))
for t in T for i in I) + xsum((heat_demand[t] * heat_price[t]) for t in T)
# constraints
for t in T:
for j in J:
m += z_t[t][j] <= capacity_ht_boiler[j] # heat generation <= maximum capacity of heat of the plant
m += x_t[t][j] == z_t[t][j]/eff_boiler[j] # fuel consumption = Heat generation / efficiency of the plants
for k in K:
m += y_t[t][k] <= capacity_el[k] # electricity generation <= maximum capacity of electricity of the plant
m += z_t[t][k] <= capacity_ht[k] # heat generation <= maximum capacity of heat of the plant
m += x_t[t][k] == y_t[t][i]/eff_plants[k] # fuel consumption = Electricity generation / efficiency of the plants
m += y_t[t][k] == heat_ratio * z_t[t][k] # electricity generation >= heat ratio * heat generation
for i in I:
m += heat_demand[t] <= z_t[t][i] # heat demand >= Heat generation
m += y_t[t][i] + el_bought[t] == el_sold[t] + el_demand[t] # electricity generation + bought electricity = sold electricity + electricity demand
status = m.optimize()
obj= m.objective_value
status
| Optimization_new2_boiler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import mp_parser
import matplotlib.pyplot as plt
# Mesh Generation
a = mp_parser.mesh(.5)
a.setMesh(25, 10)
filled = a.nodes
# print(filled)
x = [i[0] for i in filled]
y = [i[1] for i in filled]
plt.plot(x,y, '.')
plt.axis('scaled')
plt.show()
a.printFile(digits = 3, folderName = "SlopeFailureAnalysis")
# +
# Soil Geometry
points = [
(0 , 0),
(24.875, 0),
(24.875,9.375),
(19, 9.375),
(9.875, 4.875),
(0, 4.875)
]
b = mp_parser.particle(points, delta = 0.125)
b.translate(0.0625, 0.0625)
filled1 = b.nodes
x1 = [i[0] for i in filled1]
y1 = [i[1] for i in filled1]
plt.plot(x1,y1, '.')
plt.axis('scaled')
plt.show()
b.printFile(fileName = 'soil', folderName = "SlopeFailureAnalysis", digits=4)
# -
# Side Boundaries
nset = []
for key, item in enumerate(a.nodes):
if item[0] == 0 or item[0] == 25:
# print(key, item)
nset.append(key)
print(nset)
# Bottom Boundaries
nset = []
for key, item in enumerate(a.nodes):
if item[1] == 0:
# print(key, item)
nset.append(key)
print(nset)
# +
# Particle Loading
particles = b.nodes
pset = []
tmp = []
for key, item in enumerate(particles):
# print(key, item)
if item[1] > 9.437 and item[0] < 22.5 and item[0] > 19.5:
print(key, item)
pset.append(key)
print(pset)
# -
import pandas as pd
df = pd.read_hdf('particles09800.h5', 'table')
print(list(df))
particles_stresses = df[['stress_xx', 'stress_yy', 'stress_zz', 'tau_xy', 'tau_yz', 'tau_xz']].values.tolist()
# +
for key, item in enumerate(particles_stresses):
print(key, item)
with open(f'result/SlopeFailureAnalysis/particles_stresses.txt', 'w') as f:
f.write(f'! {len(particles_stresses)} particle(s) \n')
f.write(f'{len(particles_stresses)}\n')
for key, item in enumerate(particles_stresses):
f.write(' '.join([f'{j:.0f}' for j in item]) + '\n')
# -
| SlopeStabilityAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch]
# language: python
# name: conda-env-pytorch-py
# ---
# +
# import torch and other libraries
import torch
import torch.nn as nn
import torchvision.models as models
import torchvision.transforms as transforms
from torch.autograd import Variable
from PIL import Image
from IPython.display import display # to display images
print(torch.__version__)
# -
# other imports
import time
import numpy as np
import pickle
# Load the pretrained model
resnet_model = models.resnet18(pretrained=True)
alexnet_model = models.alexnet(pretrained=True)
# Use the model object to select the desired layer
resnet_layer = resnet_model._modules.get('avgpool')
alexnet_layer = alexnet_model._modules.get('classifier')
resnet_model.eval()
# Set model to evaluation mode
resnet_model.eval()
alexnet_model.eval()
# image scaler to 224 x 224 pixels
scaler = transforms.Scale((224, 224))
normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
to_tensor = transforms.ToTensor()
def get_vector(image_name, model):
# 1. Load the image with Pillow library
img = Image.open(image_name)
# 2. Create a PyTorch Variable with the transformed image
t_img = Variable(normalize(to_tensor(scaler(img))).unsqueeze(0))
if model == 'resnet':
# 3. Create a vector of zeros that will hold our feature vector
# The 'avgpool' layer has an output size of 512
my_embedding = torch.zeros(512)
# 4. Define a function that will copy the output of a layer
def copy_data(m, i, o):
my_embedding.copy_(o.data)
# 5. Attach that function to our selected layer
h = resnet_layer.register_forward_hook(copy_data)
# 6. Run the model on our transformed image
resnet_model(t_img)
h.remove()
elif model == 'alexnet':
print('using alexnet...')
# 3. Create a vector of zeros that will hold our feature vector
# The 'classifier' layer has an output size of 1000
my_embedding = torch.zeros(1000)
def copy_data(m, i, o):
my_embedding.copy_(o.data)
# 5. Attach that function to our selected layer
h = alexnet_layer.register_forward_hook(copy_data)
# 6. Run the model on our transformed image
alexnet_model(t_img)
h.remove()
# 8. Return the feature vector
return my_embedding
DATA_PATH = 'data/images/'
img1 = DATA_PATH + 'golden1.jpg'
# img2 = DATA_PATH + 'golden2.jpg'
img2 = DATA_PATH + 'cat1.jpg'
# +
# get feature vectors from resnet18
n_trials = 10
img1_resnet_times = []
img2_resnet_times = []
# get feature vectors n_trial times each and average
for i in range(n_trials):
# image 1
t_start_img1 = time.time()
img1_vector = get_vector(img1, 'resnet')
img1_resnet_times.append(time.time() - t_start_img1)
# image 2
t_start_img2 = time.time()
img2_vector = get_vector(img2, 'resnet')
img2_resnet_times.append(time.time() - t_start_img2)
# output results
t_mean_img1_resnet = round(np.mean(img1_resnet_times)/n_trials, 3)
t_mean_img2_resnet = round(np.mean(img2_resnet_times)/n_trials, 3)
print('Number of Trials: {}'.format(n_trials))
print('Image 1 Feature Vector Genration Time: {}s'.format(t_mean_img1_resnet))
print('Image 2 Feature Vector Genration Time: {}s'.format(t_mean_img2_resnet))
n_frames = 10
n_corpus = 3e6
t_corpus = round(t_mean_img1_resnet*n_frames*n_corpus/3600, 3)
print('\nCorpus Size={}, Frames per Video={}'.format(int(n_corpus), n_frames))
print('Corpus Processing Time: {} hrs, or {} days'.format(t_corpus, round(t_corpus/24, 3)))
# -
# Using PyTorch Cosine Similarity
cos = nn.CosineSimilarity(dim=1, eps=1e-6)
cos_sim = cos(img1_vector.unsqueeze(0),
img2_vector.unsqueeze(0))
print('\nCosine similarity: {0}\n'.format(cos_sim))
features_test = [np.array(img1_vector), np.array(img2_vector)]
features_test = np.array(features_test)
features_test
# Resnet 18 experiments show that feature vector image generation takes aobut 1/100th of a second. This means over 45 days to process 3 million videos with 100 sampled images each.
# +
# get feature vectors from alexnet
# img1_vector_a = get_vector(img1, 'alexnet')
# img2_vector_a = get_vector(img2, 'alexnet')
# -
img1_vec = np.array(img1_vector)
img2_vec = np.array(img2_vector)
vec = np.array([img1_vec, img2_vec])
print(vec.shape)
# ## Frames extraction from videos
import os
import argparse
import FFMPEGFrames
import shutil
import importlib
from ffprobe3 import FFProbe
importlib.reload(FFMPEGFrames)
# +
videos_path = 'data/videos'
delete = True
features = {}
# loop through all the videos
i = 0
for path, subdirs, files in os.walk(videos_path):
for name in files:
if i < 2:
# extract frames from videos
video_length = int(float(FFProbe(os.path.join(path, name)).video[0].duration)) + 1
n_frames = 10
fps = n_frames/video_length
video_path = os.path.join(path, name)
print(os.path.join(path, name) + ' ' + str(video_length) + 's ' + '(' + str(fps) + ')')
f = FFMPEGFrames.FFMPEGFrames("data/video_frames/")
f.extract_frames(os.path.join(path, name), fps)
i += 1
# get feature vectors for each frame image
frames_path = f.full_output
frames = os.listdir(frames_path)
model = 'resnet'
features[video_path] = [get_vector(os.path.join(frames_path, frame), model) for frame in frames]
# delete rest of the files after extracting image features
if delete:
shutil.rmtree(f.full_output)
# -
test_dict = {'data/videos/0.mp4':np.array([1, 9, 1])}
test_dict
video_path = 'data/videos/0.mp4'
data = np.array([1, 1, 1])
f = h5py.File("test.hdf5", "w")
f.create_dataset(video_path, data)
list(f.keys())
# ## Check the extracted features
import h5py
f1 = h5py.File('test_features.hdf5', 'r+')
# +
# List all groups
print("Keys: %s" % f1.keys())
a_group_key = list(f1.keys())[0]
# # Get the data
data = list(f1[a_group_key])
data
# -
data_names = f1['mnt']['e'].keys()
data_names
f1['mnt']['e']['ucf_101_sample']['v_ApplyEyeMakeup_g01_c01.avi']['output000001.png']
f1['mnt/e/ucf_101_sample/v_ApplyEyeMakeup_g01_c01.avi'].keys()
f1.close()
| feature_extraction_experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TOOLS
#
# This notebook contains different sections that help to:
# - Generate different video variations
# - Compute metrics using external software like ffmpeg or libav
#
# It makes use of different scripts located in the /scripts folder that will help to both generate different renditions from original (1080p) input videos as well as distortions of them (attacks).
#
# It also provides means to execute metric extractions from those videos (MS-SSIM, VMAF, SSIM and PSNR) by means of bash shell. These metrics are also utilized in other notebooks.
# # Input path configuration
#
# The cell below must be executed. It prepares the notebook to point to the local repo structure where the original 1080p renditions are.
#
# The expected input structure is:
#
# ```
# data
# ├── 1080p
# │ └── 01.mp4
# ├── 720p
# │ └── 01.mp4
# ├── 480p
# │ └── 01.mp4
# ├── 360p
# │ └── 01.mp4
# └── 240p
# | └── 01.mp4
# └── 144p
# └── 01.mp4
# ```
# +
import os.path
from pathlib import Path
current_path = Path(os.path.abspath(''))
input_path = (current_path / "../data/1080p").resolve()
# -
# # 1.-Rendition creation
# ## 1.0.- Scaling and bitrate reduction
#
# In first place, a dataset with original videos is required. We propose the collection provided in the YT8M notebook available [here](https://github.com/epiclabs-io/YT8M/blob/master/yt8m-crawler.ipynb). Once CSV file with the videos metadata is generated, use [downloader](../../YT8M_downloader/downloader.py) script to download videos. Consider using various format filter values to get e.g. 60 fps videos.
#
# ## 1.1.- Watermarks
#
# There is a python script in order to insert watermarks in the videos. This script receives 4 parameters:
# - The input path (-i or --input) which is the folder containing 1080p.
# - The output path (-o or --output) which is the folder where the videos with watermark are going to be stored.
# - The metadata (-m or --metadata) which is the file containing data about the videos, the most important is the needed bitrate to enconde the video.
# - The watermark file (-w --watermark) which is the file containing the image to be applied to the video.
#
# The output looks like
#
# ```
# ├── 1080p_watermark
# │ ├── 01.mp4
# ├── 720p_watermark
# │ ├── 01.mp4
# ├── 480p_watermark
# │ ├── 01.mp4
# ├── 360p_watermark
# │ ├── 01.mp4
# ├── 240p_watermark
# │ ├── 01.mp4
# ├── 144p_watermark
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
metadata_file = (current_path / "/scripts/yt8m_data.csv").resolve()
watermark_file = (current_path / "/scripts/watermark/livepeer.png").resolve()
# %run -i '/scripts/watermark.py' -i $input_path -o $output_path -m $metadata_file -w $watermark_file
# ## 1.2.- Flips / rotations
# There is a python script in order to flip / rotate the videos. This script receives 3 parameters:
# - The input path (-i or --input) which is the folder containing 1080p.
# - The output path (-o or --output) which is the folder where the flipped videos are going to be stored.
# - The desired flip or rotation:
# - -vf or --vflip for the vertical flip
# - -hf or --hflip for the horizontal flip
# - -cf or for the 90 degrees clockwise rotation
# - -ccf for the 90 degrees counterclockwise rotation
#
# There are implemented the following ways to flip / rotate a video:
# ## 1.3.- Vertical flip
#
# The output looks like
#
# ```
# ├── 1080p_flip_vertical
# │ ├── 01.mp4
# ├── 720p_flip_vertical
# │ ├── 01.mp4
# ├── 480p_flip_vertical
# │ ├── 01.mp4
# ├── 360p_flip_vertical
# │ ├── 01.mp4
# ├── 240p_flip_vertical
# │ ├── 01.mp4
# ├── 144p_flip_vertical
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/flip.py' -i $input_path -o $output_path -vf
# ## 1.4.- Horizontal flip
#
# The output looks like
#
# ```
# ├── 1080p_flip_horizontal
# │ ├── 01.mp4
# ├── 720p_flip_horizontal
# │ ├── 01.mp4
# ├── 480p_flip_horizontal
# │ ├── 01.mp4
# ├── 360p_flip_horizontal
# │ ├── 01.mp4
# ├── 240p_flip_horizontal
# │ ├── 01.mp4
# ├── 144p_flip_horizontal
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/flip.py' -i $input_path -o $output_path -hf
# ## 1.5.- Rotate 90 degrees clockwise
#
# The output looks like
#
# ```
# ├── 1080p_rotate_90_clockwise
# │ ├── 01.mp4
# ├── 720p_rotate_90_clockwise
# │ ├── 01.mp4
# ├── 480p_rotate_90_clockwise
# │ ├── 01.mp4
# ├── 360p_rotate_90_clockwise
# │ ├── 01.mp4
# ├── 240p_rotate_90_clockwise
# │ ├── 01.mp4
# ├── 144p_rotate_90_clockwise
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/flip.py' -i $input_path -o $output_path -cf
# ## 1.6.- Rotate 90 degrees counterclockwise
#
# The output looks like
#
# ```
# ├── 1080p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ├── 720p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ├── 480p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ├── 360p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ├── 240p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ├── 144p_rotate_90_counterclockwise
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/flip.py' -i $input_path -o $output_path -ccf
# ## 1.7.- Black and white
#
# There is a python script in order to convert color videos to black and white. This script receives 2 parameters:
# - The input path (-i or --input) which is the folder containing the renditions.
# - The output path (-o or --output) which is the folder where the black and white videos are going to be stored.
#
# The output looks like
#
# ```
# ├── 1080p_black_and_white
# │ ├── 01.mp4
# ├── 720p_black_and_white
# │ ├── 01.mp4
# ├── 480p_black_and_white
# │ ├── 01.mp4
# ├── 360p_black_and_white
# │ ├── 01.mp4
# ├── 240p_black_and_white
# │ ├── 01.mp4
# ├── 144p_black_and_white
# │ ├── 01.mp4
# ```
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/black_and_white.py' -i $input_path -o $output_path
# ## 1.8.- Chroma subsampling
#
# There is a python script in order to change the chroma subsampling of the videos. This script receives 4 parameters:
# - The input path (-i or --input) which is the folder containing the renditions.
# - The output path (-o or --output) which is the folder where the subsmapled videos are going to be stored.
# - The metadata (-m or --metadata) which is the file containing data about the videos, the most important is the needed bitrate to enconde the video.
# - The chosen subsampling (-s --subsampling) which is the chroma subsampling to be applied to the video.
#
# The output looks like
#
# ```
# ├── 720p_chroma_subsampling_chosen_subsampling
# │ ├── 01.mp4
# ├── 480p_chroma_subsampling_chosen_subsampling
# │ ├── 01.mp4
# ├── 360p_chroma_subsampling_chosen_subsampling
# │ ├── 01.mp4
# ├── 240p_chroma_subsampling_chosen_subsampling
# │ ├── 01.mp4
# ├── 144p_chroma_subsampling_chosen_subsampling
# │ ├── 01.mp4
# ```
#
# Where chosen_subsampling is one of the [ffmpeg valid subsampling](https://trac.ffmpeg.org/wiki/Chroma%20Subsampling), for example:
#
# `yuv420p, yuv422p, yuv444p, yuv420p10le, yuv422p10le, yuv444p10le`
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/chroma_subsampling.py' -i $input_path -o $output_path -m $metadata_file -s yuv422p
# ## 1.9.- low bitrate
#
# There is a python script in order to change the bitrate of the videos. This script receives 4 parameters:
# - The input path (-i or --input) which is the folder containing the renditions.
# - The output path (-o or --output) which is the folder where the videos with low bitrate are going to be stored.
# - The metadata (-m or --metadata) which is the file containing data about the videos, the most important is the needed bitrate to enconde the video.
# - The chosen divisor for the bitrate (-d, --divisor) which is the divisot to be applied to the video bitrate.
#
# The output looks like
#
# ```
# ├── 1080p_low_bitrate_divisor
# │ ├── 01.mp4
# ├── 720p_low_bitrate_divisor
# │ ├── 01.mp4
# ├── 480p_low_bitrate_divisor
# │ ├── 01.mp4
# ├── 360p_low_bitrate_divisor
# │ ├── 01.mp4
# ├── 240p_low_bitrate_divisor
# │ ├── 01.mp4
# ├── 144p_low_bitrate_divisor
# │ ├── 01.mp4
# ```
#
# Where divisor is an integer greater than 0 that is going to divide the current bitrate
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/low_bitrate.py' -i $input_path -o $output_path -m $metadata_file -d 4
# ## 1.10.- vignette
#
# There is a python script in order to apply a vignette filter to the videos. This script receives 3 parameters (One is optional):
# - The input path (-i or --input) which is the folder containing the renditions.
# - The output path (-o or --output) which is the folder where the vignetted videos are going to be stored.
# - The angle (-a or --angle) which is the angle of the vignette filter to be applied to the video. This param is optional and by default is [PI/5](https://ffmpeg.org/ffmpeg-filters.html#vignette-1)
#
# The output looks like
#
# ```
# ├── 1080p_vignette_angle
# │ ├── 01.mp4
# ├── 720p_vignette_angle
# │ ├── 01.mp4
# ├── 480p_vignette_angle
# │ ├── 01.mp4
# ├── 360p_vignette_angle
# │ ├── 01.mp4
# ├── 240p_vignette_angle
# │ ├── 01.mp4
# ├── 144p_vignette_angle
# │ ├── 01.mp4
# ```
#
# Where angle is a valid angle for ffmpeg (in the [0,PI/2] range)
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/vignette.py' -i $input_path -o $output_path -m $metadata_file -a PI/4
# ## 1.11.- FPS converter
#
# This script converts source videos to common frame rates, and puts results into corresponding folders. It can be applied after all other renditions are created.
# - The input path (-i or --input) which is the folder containing the renditions.
# - The output path (-o or --output) which is the folder where the renditions are going to be stored.
# - GPU flag (-g or --gpu). Whether to use GPU codecs. You need to have Ffmpeg built with Nvidia Codecs for it to work. Default is false.
#
# The output folder names will have source-target FPS suffix appended.
#
# ```
# ├── 1080p_watermark_60-24fps
# │ ├── <filename>.mp4
# ├── 720p_watermark_60-24fps
# │ ├── <filename>.mp4
# ├── 480p_watermark_60-24fps
# │ ├── <filename>.mp4
# ├── 360p_watermark_60-24fps
# │ ├── <filename>.mp4
# ├── 240p_watermark_60-24fps
# │ ├── <filename>.mp4
# ├── 144p_watermark_60-24fps
# │ ├── <filename>.mp4
# ```
# + pycharm={"name": "#%%\n"}
output_path = (current_path / "../data").resolve()
# %run -i '/scripts/fps_converter.py' -i $input_path -o $output_path
# -
# # 2 Metric extraction
#
# In order to extract metrics from the videos above and being able to compare them, a set of scripts are made available below.
#
# ## 2.1 Compute ms-ssim
#
# There is a bash script `evaluate-ms-ssim.sh` in order to calculate the multiscale ssim. This script receives two parameters which are the path where the videos are and the path where the output should be placed. Note that the `../output` folder is the folder where other notebooks are expecting the output.
#
#
# The output structuture is inside the output folder
#
# ```
# mssim
# ├── 240
# │ ├── 01
# │ │ └── 01_240.log
# ├── 360
# │ ├── 01
# │ │ └── 01_360.log
# ├── 480
# │ ├── 01
# │ │ └── 01_480.log
# └── 720
# ├── 01
# └── 01_720.log
# ```
#
# Where the folder indicates the rendition we are using to compare against the original (1080p).
# A subfolder of this folder contains the name of the asset and finally the file containing the log.
#
# The log is a csv file, with the following structure:
#
# ```
# ms-ssim, psnr-y, psnr-u, psnr-v
# 0.986889, 32.866684, 43.274622, 42.429359
# 0.985558, 32.394349, 43.344157, 42.658971
# 0.985460, 32.521368, 43.338460, 42.580399
# 0.985896, 32.670122, 43.325404, 42.529248
# ```
# +
shell_path = Path(os.path.abspath(''))
shell_input_path = (current_path / "../data").resolve()
shell_output_path = (current_path / "../output").resolve()
# !bash '/scripts/shell/evaluate-ms-ssim.sh' $shell_input_path $shell_output_path
# -
# ## 2.1.- Compute VMAF
#
# There is a bash script `evaluate-vmaf.sh` in order to calculate the vmaf score. This script receives two parameters which are the path where the videos are and the path where the output should be placed. Note that the `../output` folder is the folder where other notebooks are expecting the output.
#
# The script will produce the following folder structure:
#
# ```
# output/vmaf
# ├── 240
# │ ├── 01
# │ │ ├── 01_240.log
# │ │ └── 01_240.log.out
# ├── 360
# │ ├── 01
# │ │ ├── 01_360.log
# │ │ └── 01_360.log.out
# ├── 480
# │ ├── 01
# │ │ ├── 01_480.log
# │ │ └── 01_480.log.out
# └── 720
# ├── 01
# ├── 01_720.log
# └── 01_720.log.out
# ```
#
# Where the folder indicates the rendition we are using to compare against the original (1080p).
# A subfolder of this folder contains the name of the asset and finally two files: One containing the result
# (videoname_rendition_resolution.log) and other containing the output from the ffmpeg (videoname_rendition_resolution.log.out).
#
# The log file contains the following information:
#
# ```
# Start calculating VMAF score...
# Exec FPS: 158.922597
# VMAF score = 90.566873
# ```
#
# The interesting line is the third one, containing the vmaf score.
#
# The .out file is not worth analyzing as it is the standard ffmpeg output
# +
shell_path = Path(os.path.abspath(''))
shell_input_path = (current_path / "../data").resolve()
shell_output_path = (current_path / "../output").resolve()
# !bash '/scripts/shell/evaluate-vmaf.sh' $shell_input_path $shell_output_path
# -
# ## 2.3.- Compute ssim and psnr
#
# There is a bash script `evaluate-psnr-ssim.sh` in order to calculate the ssim and psnr metrics. This script receives two parameters which are the path where the videos are and the path where the output should be placed. Note that the `../output` folder is the folder where other notebooks are expecting the output.
#
# The script will produce the following folder structure:
#
# The output structuture is inside the output folder for the psnr
#
# ```
# psnr
# ├── 240
# │ ├── 01
# │ │ └── 01_240.log
# ├── 360
# │ ├── 01
# │ │ └── 01_360.log
# ├── 480
# │ ├── 01
# │ │ └── 01_480.log
# └── 720
# ├── 01
# └── 01_720.log
# ```
#
# Where the folder indicates the rendition we are using to compare against the original (1080p).
# A subfolder of this folder contains the name of the asset and finally the file containing the log.
#
# The log has the following structure:
#
# n:1 mse_avg:0.60 mse_y:0.73 mse_u:0.34 mse_v:0.32 psnr_avg:50.37 psnr_y:49.50 psnr_u:52.76 psnr_v:53.01
# n:2 mse_avg:0.83 mse_y:1.01 mse_u:0.48 mse_v:0.45 psnr_avg:48.95 psnr_y:48.09 psnr_u:51.29 psnr_v:51.62
# n:3 mse_avg:0.77 mse_y:0.94 mse_u:0.45 mse_v:0.44 psnr_avg:49.25 psnr_y:48.42 psnr_u:51.59 psnr_v:51.71
# n:4 mse_avg:0.76 mse_y:0.92 mse_u:0.45 mse_v:0.43 psnr_avg:49.32 psnr_y:48.50 psnr_u:51.55 psnr_v:51.76
# n:5 mse_avg:0.65 mse_y:0.79 mse_u:0.39 mse_v:0.36 psnr_avg:50.01 psnr_y:49.18 psnr_u:52.22 psnr_v:52.58
#
#
# The output structuture is inside the output folder for the ssim
#
# ```
# ssim
# ├── 240
# │ ├── 01
# │ │ └── 01_240.log
# ├── 360
# │ ├── 01
# │ │ └── 01_360.log
# ├── 480
# │ ├── 01
# │ │ └── 01_480.log
# └── 720
# ├── 01
# └── 01_720.log
# ```
#
# Where the folder indicates the rendition we are using to compare against the original (1080p).
# A subfolder of this folder contains the name of the asset and finally the file containing the log.
#
#
# +
shell_path = Path(os.path.abspath(''))
shell_input_path = (current_path / "../data").resolve()
shell_output_path = (current_path / "../output").resolve()
# !bash '/scripts/shell/evaluate-psnr-ssim.sh' $shell_input_path $shell_output_path
| feature_engineering/notebooks/Tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
plt.rcParams.update({ # setup matplotlib to use latex for output
"pgf.texsystem": "pdflatex", # change this if using xetex or lautex
"text.usetex": True, # use LaTeX to write all text
"font.family": "serif",
"font.serif": [], # blank entries should cause plots to inherit fonts from the document
"font.sans-serif": [],
"font.monospace": [],
"axes.labelsize": 10, # LaTeX default is 10pt font.
"font.size": 10,
"legend.fontsize": 8, # Make the legend/label fonts a little smaller
"xtick.labelsize": 8,
"ytick.labelsize": 8,
"pgf.preamble": [
r"\usepackage[utf8x]{inputenc}", # use utf8 fonts becasue your computer can handle it :)
r"\usepackage[T1]{fontenc}", # plots will be generated using this preamble
]
})
# --------------------------------
#
# # Results
#
#
# ### PINN:
#
# Network: [1, 50, 50, 50, 50, 51]
#
# Num Param: 10.35k
#
# Error lambda_1 (Clean Data): 0.016499%
#
# Error lambda_2 (Clean Data): 0.011111%
#
# Error lambda_1 (Noisy Data): 0.153911%
#
# Error lambda_2 (Noisy Data): 0.045160%
#
#
# ### QRes-PINN:
# Network: [1, 20, 20, 20, 20, 51]
#
# Num Param: 4.61k
#
# Error lambda_1 (Clean Data): 0.008750%
#
# Error lambda_2 (Clean Data): 0.009676%
#
# Error lambda_1 (Noisy Data): 0.183016%
#
# Error lambda_2 (Noisy Data): 0.008587%
#
# ## Count Number of Parameters
l = [1, 50, 50, 50, 50, 51]
num_param = 0
for i in range(len(l)-1):
num_param += l[i] * l[i+1] + l[i+1]
print("PINN Num params: %.2fk" % np.round(num_param / 1000, 2))
l = [1, 20, 20, 20, 20, 51]
num_param = 0
for i in range(len(l)-1):
num_param += l[i] * l[i+1] * 2 + l[i+1]
print("QRes Num params: %.2fk" % np.round(num_param / 1000, 2))
#
# --------------------------------
#
# # Training Loss
#
pinn_clean = np.load("loss/loss_clean_PINNs.npy")
qres_clean = np.load("loss/loss_clean_QRes.npy")
pinn_noisy = np.load("loss/loss_noisy_PINNs.npy")
qres_noisy = np.load("loss/loss_noisy_QRes.npy")
# +
fig, axes = plt.subplots(1, 2, dpi=200, figsize=(4.5, 1.2))
# clean data
x_pinn = np.concatenate((
np.arange(0, 50000, 10),
np.arange(len(pinn_clean) - 5000) + 50000
))
x_qres = np.concatenate((
np.arange(0, 10000, 10),
np.arange(len(qres_clean) - 1000) + 10000
))
ax = axes[0]
sns.lineplot(x=x_pinn, y=pinn_clean, label='PINN', lw=0.5, alpha=1, ax=ax, color='#DE5521')
sns.lineplot(x=x_qres, y=qres_clean, label='QRes', lw=0.5, alpha=1, ax=ax, color='#21AADE')
ax.set_xlabel('Training Epoch', fontsize=9)
ax.set_ylabel('Training Loss (log)', fontsize=10)
ax.set_title('KdV (Clean Data)', fontsize=9)
ax.set_yscale('log')
ax.legend(loc='upper center', frameon=False, bbox_to_anchor=(0.5, 1.00))
# sns.despine()
# noisy data
x_pinn = np.concatenate((
np.arange(0, 50000, 10),
np.arange(len(pinn_noisy) - 5000) + 50000
))
x_qres = np.concatenate((
np.arange(0, 10000, 10),
np.arange(len(qres_noisy) - 1000) + 10000
))
ax = axes[1]
sns.lineplot(x=x_pinn, y=pinn_noisy, label='PINN', lw=0.5, alpha=1, ax=ax, color='#DE5521')
sns.lineplot(x=x_qres, y=qres_noisy, label='QRes', lw=0.5, alpha=1, ax=ax, color='#21AADE')
ax.set_xlabel('Training Epoch')
ax.set_title('KdV (1\% noise)', fontsize=9)
ax.set_yscale('log')
ax.legend(loc='upper center', frameon=False, bbox_to_anchor=(0.5, 1.00))
fig.savefig('figures/loss_curves.pdf', dpi=300, bbox_inches='tight')
plt.show()
# -
| QRes/main/discrete_time_identification (KdV)/eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import os
import scipy.stats as st
import iqplot
import bokeh.io
import bokeh.plotting
from bokeh.models import Legend
import numba
import bebi103
bokeh.io.output_notebook()
# -
# # Aim 1: Data Validation for Microtubule Experiments #
# ### Comparing Labeled and Unlabeled tubulin performance ###
# In the experiment conducted by Gardner et al., microtubules were labeled with fluorescent markers. We investigate whether or not these fluorescent markers influence tubulin performance, determined by time to catastrophe (s). We look at data gathered from unlabeled and labeled tubulin, and focus on three different comparisons:
# 1. ECDF of labeled vs unlabeled tubulin
# 2. Mean time to catastrophe of labeled vs unlabeled tubulin
# 3. Hypothesis testing assuming identical distributions
# Each of these strategies checks whether or not the labeled or unlabeled tubulin datasets are different in some way. If a significant difference does exist, this means that the fluorescent markers have some sort of impact on microtubule performance. This would also mean that the labeled tubulin used in the subsequent experiments do not accurately depict microtubule behavior. In this way, we hope to validate the data collected by confirming that the fluorescent markers do not influence microtubule performance. <br />
#
# To start our investigation, we read in the dataset and save the values in a tidy data frame.
# +
data_path = '../datasets'
file_path = os.path.join(data_path, 'gardner_time_to_catastrophe_dic_tidy.csv')
# get rid of the index column when loading data
df = pd.read_csv(file_path).iloc[:, 1:]
# replace True/False with labeled vs unlabeleed
df['labeled'] = df['labeled'].apply(lambda x: 'labeled tubulin' if x else 'unlabeled tubulin')
df
# -
# ### 1. ECDF comparison ###
# To determine whether or not microtubule performance is different between labeled and unlabeled tubulin, we first look at the cumulative distributions of the empirical data. If the cumulative distributions occupy the same areas, then the fluorescent markers probably do not have a strong effect on microtubule performance since unlabeled/labeled times to catastrophe are indistinguishable from each other. We use `iqplot` to display the respective ECDFs below and observe whether or not the unlabeled and labeled datasets are identically distributed.
# +
p = iqplot.ecdf(
data=df,
q='time to catastrophe (s)',
cats='labeled',
style='staircase',
conf_int = True
)
bokeh.io.show(p)
# -
# By a quick, visual inspection of the plot, it looks like the catastrophe times for microtubules with labeled and unlabeled tubulin could be identically distributed. The confidence interval for the unlabeled tubulin almost always overlaps with the labeled tubulin confidence intervals. <br /> <br />
# Since we are using the confidence intervals to check whether or not the datasets overlap, further investigation of confidence interval generation is worth exploring. The confidence intervals above were calculated with bootstrapping, but we can also use Dvoretzky-Kiefer-Wolfowitz Inequality (DKW) to compute confidence intervals for the ECDF. To start, we define an ecdf function that can compute the ecdf at an arbitrary x value.
def ecdf(x, data):
"""
This function computes the value of the ECDF built from a 1D array, data,
at arbitrary points, x, which can also be an array.
x can be an integer or float, an array of ints, or a list of ints
"""
data_sorted = np.sort(data)
ecdf_li = []
if type(x) == int or type(x) == float:
index_tup = np.where(data_sorted <= x)[0]
if index_tup.size == 0:
ecdf = 0
else:
ecdf = (index_tup[-1] + 1) / len(data)
ecdf_li.append(ecdf)
else:
for value in x:
index_tup = np.where(data_sorted <= value)[0]
if index_tup.size == 0:
ecdf = 0
else:
ecdf = (index_tup[-1] + 1) / len(data)
ecdf_li.append(ecdf)
return np.array(ecdf_li)
# The DKW inequality states that for any $\epsilon > 0$,
#
# \begin{align}
# P\left(\mathrm{sup}_x \left|F(x) - \hat{F}(x)\right| > \epsilon\right) \le 2\mathrm{e}^{-2 n \epsilon^2},
# \end{align}
#
# To calculate the DKW inequality for the microtubule catastrophe data then, we first calculate $\alpha$ which while be used to calculate $\epsilon$. For the 95% confidence interval: <br />
# \begin{align}
# 100*(1-\alpha) & = 95 \\
# 1-\alpha & = 0.95 \\
# 0.05 &= \alpha \\
# \end{align} <br />
# Now we create a function to get $\epsilon$ for a given dataset (since $n$ might vary), which I will later use to calculate the upper and lower bounds. I use the calculated $\alpha$ value and the expression: <br />
# \begin{align}
# \epsilon &= \sqrt{\frac{1}{2n} \log{\frac{2}{\alpha}}} \\
# \end{align}
# +
alpha = 0.05
def calc_epsilon(data):
n = len(data)
x = 1/(2*n)
l = np.log(2/alpha)
return np.sqrt(x*l)
# -
# Next we create a function that returns the lower bound, given by the expression: <br />
# \begin{align}
# L(x) = \max\left(0, \hat{F}(x) - \epsilon\right),
# \end{align}
def lower_bound(data):
"""
for a given array of experimental data,
this function returns a list of the DKW lower bound values
corresponding to the given data points
"""
ep = calc_epsilon(data)
l_bounds = []
for x in data:
lower = np.maximum(0, x-ep)
l_bounds.append(lower)
return l_bounds
# Now we create a function that returns the upper bound, given by the expression: <br />
# \begin{align}
# U(x) = \min\left(1, \hat{F}(x) + \epsilon\right).
# \end{align}
def upper_bound(data):
"""
for a given array of experimental data,
this function returns a list of the DKW upper bound values
corresponding to the given data points
"""
ep = calc_epsilon(data)
u_bounds = []
for x in data:
upper = np.minimum(1, x+ep)
u_bounds.append(upper)
return u_bounds
# Now I want to plot the confidence intervals for the ECDFs of the labeled and unlabeled tubulin times to catastrophe. I use the previously made `unlabeled_tubulin` and `labeled tubulin` arrays, along with the `ecdf` method made in part e to obtain the ecdf values of the two arrays. I use the `lower_bound` and `upper_bound` methods just created to also store the lower and upper bounds for each dataset. <br /><br />
# For the **unlabeled** tubulin dataset I create a dataframe that holds all the values needed for plotting. I sort the dataset by the value so I can plot in order.
# +
values_u = []
ecdf_u = []
for x in unlabeled_tubulin:
values_u.append(x)
a = ecdf([x], unlabeled_tubulin)
ecdf_u.append(a[0])
df_unlabeled = pd.DataFrame(data = {"value":values_u,
"ecdf":ecdf_u,
}
)
df_unlabeled = df_unlabeled.sort_values(by = ["value"])
e = df_unlabeled.loc[:, "ecdf"]
lower_u = lower_bound(e)
upper_u = upper_bound(e)
df_unlabeled["lower_bound"] = lower_u
df_unlabeled["upper_bound"] = upper_u
df_unlabeled.head()
# -
# Now for the **labeled** tubulin dataset I repeat this procedure of creating a dataframe and then plotting.
# +
values_l = []
ecdf_l = []
for x in labeled_tubulin:
values_l.append(x)
a = ecdf([x], labeled_tubulin)
ecdf_l.append(a[0])
df_labeled = pd.DataFrame(data = {"value":values_l,
"ecdf":ecdf_l,
}
)
df_labeled = df_labeled.sort_values(by = ["value"])
e_l = df_labeled.loc[:, "ecdf"]
lower_l = lower_bound(e_l)
upper_l = upper_bound(e_l)
df_labeled["lower_bound"] = lower_l
df_labeled["upper_bound"] = upper_l
df_labeled.head()
# -
# Finally, I create the final plot which overlays the two ecdf's with their respective confidence intervals, calculated with the DKW lower/upper bound expressions.
# +
p2 = bokeh.plotting.figure(
width=800,
height=400,
x_axis_label="time to catastrophe",
y_axis_label="ecdf",
title = "unlabeled vs. labeled tubulin",
)
e = p2.line(source = df_unlabeled, x = "value", y = "ecdf",
color = "#b2abd2", line_width = 3, alpha = 0.7)
l = p2.circle(source = df_unlabeled, x = "value", y = "lower_bound",
color = "#5e3c99", alpha = 0.5)
u = p2.circle(source = df_unlabeled, x = "value", y = "upper_bound",
color = "#5e3c99", alpha = 0.5)
e_l = p2.line(source = df_labeled, x = "value", y = "ecdf",
color = "#fdb863", line_width = 3, alpha = 0.7)
l_l = p2.circle(source = df_labeled, x = "value", y = "lower_bound",
color = "#e66101", alpha = 0.3)
u_l = p2.circle(source = df_labeled, x = "value", y = "upper_bound",
color = "#e66101", alpha = 0.3)
legend = Legend(items=[("unlabeled ecdf" , [e]),
("unlabeled lower bound" , [l]),
("unlabeled upper bound" , [u]),
("labeled ecdf" , [e_l]),
("labeled lower bound" , [l_l]),
("labeled upper bound" , [u_l]),], location="center")
p2.add_layout(legend, 'right')
p2.legend.click_policy = "hide"
bokeh.io.show(p2)
# -
# The purple dots here show the unlabeled tubulin bounds while the orange dots show the labeled tubulin bounds, and the lines show the ecdf values. <br />
# Comparing the lower bounds, the orange and purple dots seem to follow the same trajectory. This means that the unlabeled and labeled tubulin lower bound values are very similar. <br />
# Comparing the upper bounds, the labeled values the purple dots are noticeably above the orange dots. This means that the unlabeled upper bound values are slightly higher than the labeled upper bound values, though the shape of the bounds is the same. <br />
# Though the upper bound values are not as aligned as the lower bound values, it is still reasonable to conclude that these confidence intervals are very similar. Therefore, this quick visual check supports the hypothesis that microtubule times to catastrophe are identically distributed between unlabeled and labeled tubulin. <br /><br />
# This conclusion matches what we found from the iqplot calculations.
# ### 2. Mean time to catastrophe comparison ###
# Next, we compare the mean times to catastrophe between labeled and unlabeled tubulin to detect any possible differences in performance. If the mean times are close to each other, there is more reason to believe that the fluorescent markers do not affect microtubule performance. To check this, we use nonparametric bootstrapping to compute confidence intervals for the plug-in estimate for the mean time to catastrophe for each of the two conditions. First we define some functions to calculate our bootstrap replicate for each bootstrap sample.
# +
rg = np.random.default_rng()
# set up numpy arrays with values for the labeled and unlabeled tubulin
unlabeled_tubulin = df.loc[df['labeled'] == 'unlabeled tubulin', 'time to catastrophe (s)'].values
labeled_tubulin = df.loc[df['labeled'] == 'labeled tubulin', 'time to catastrophe (s)'].values
def generate_bootstrap_samples(data):
"""Draw N bootstrap samples from a 1D data set."""
return rg.choice(data, size=len(data))
def bootstrap_reps_mean(data, N=1):
"""Draw boostrap replicates of the mean from 1D data set."""
means = np.empty(N)
for i in range(N):
means[i] = np.mean(generate_bootstrap_samples(data))
return means
# -
# Now we can generate 100,000 bootstrap samples for both the labeled and unlabeled and calculate the plug-in estimate for the mean.
# +
unlabeled_means = bootstrap_reps_mean(unlabeled_tubulin, N=100000)
labeled_means = bootstrap_reps_mean(labeled_tubulin, N=100000)
unlabeled_mean_conf_int = np.percentile(unlabeled_means, [2.5, 97.5])
labeled_mean_conf_int = np.percentile(labeled_means, [2.5, 97.5])
print(f"Unlabeled tubulin time to catastrophe(s) confidence interval: [{unlabeled_mean_conf_int[0]:.2f}, {unlabeled_mean_conf_int[1]:.2f}]")
print(f"Labeled tubulin time to catastrophe(s) confidence interval: [{labeled_mean_conf_int[0]:.2f}, {labeled_mean_conf_int[1]:.2f}]")
# -
# We use the `bebi103` package to visually display these confidence intervals.
# +
labeled_mean = labeled_tubulin.mean()
unlabeled_mean = unlabeled_tubulin.mean()
summaries = [
dict(label = "unlabeled tubulin", estimate = unlabeled_mean,
conf_int = unlabeled_mean_conf_int),
dict(label = "labeled tubulin", estimate = labeled_mean,
conf_int = labeled_mean_conf_int)
]
bokeh.io.show(
bebi103.viz.confints(summaries)
)
# -
# The confidence intervals of the two categories have significant overlap. This calculation supports the previous conclusion from the ecdf since there is not a clear difference in microtubule performance between labeled and unlabeled samples. <br /> <br />
# Again, since we are using the confidence intervals to check value overlaps it is worth double checking that our confidence interval generation is appropriate. In this case, we can double check our confidence intervals with a theoretical distribution rather than the empirical distribution. Specifically, we can use the normal distribution by the central limit theorem:
# \begin{align}
# &\mu = \bar{x},\\[1em]
# &\sigma^2 = \frac{1}{n(n-1)}\sum_{i=1}^n (x_i - \bar{x})^2,
# \end{align}
# We define a function to calculate the variance of the data set using this theoretical equation.
def calc_variance(data_array):
"""This function calculates the variance of a 1D data array"""
n = data_array.size
mean = data_array.mean()
numer = 0
for i in range(n):
numer += (data_array[i] - mean) ** 2
denom = n * (n-1)
return numer/denom
# Now we perform the calculation and visualize the confidence intervals.
# +
unlabeled_variance = calc_variance(unlabeled_tubulin)
labeled_variance = calc_variance(labeled_tubulin)
labeled_conf1 = st.norm.ppf(0.025, loc=labeled_mean, scale=np.sqrt(labeled_variance))
labeled_conf2 = st.norm.ppf(0.975, loc=labeled_mean, scale=np.sqrt(labeled_variance))
unlabeled_conf1 = st.norm.ppf(0.025, loc=unlabeled_mean, scale=np.sqrt(unlabeled_variance))
unlabeled_conf2 = st.norm.ppf(0.975, loc=unlabeled_mean, scale=np.sqrt(unlabeled_variance))
print(f"Unlabeled tubulin time to catastrophe(s) confidence interval: [{unlabeled_conf1:.2f}, {unlabeled_conf2:.2f}]")
print(f"Labeled tubulin time to catastrophe(s) confidence interval: [{labeled_conf1:.2f}, {labeled_conf2:.2f}]")
summaries = [
dict(label = "unlabeled tubulin", estimate = unlabeled_mean,
conf_int = [unlabeled_conf1, unlabeled_conf2]),
dict(label = "labeled tubulin", estimate = labeled_mean,
conf_int = [labeled_conf1, labeled_conf2])
]
bokeh.io.show(
bebi103.viz.confints(summaries)
)
# -
# When comparing the confidence interval calculation from the theoretical distribution to the confidence interval derived from the empirical distribution, we can see that the confidence intervals are very similar. Again, there does not seem to be a significant difference between the times to catastrophe between the unlabeled and labeled tubulin.
# ### 3. Hypothesis testing assuming identical distributions ###
# Next, we use a permutation hypothesis test to test the hypothesis that the distribution of catastrophe times for microtubules with labeled tubulin is the same as that for unlabeled tubulin.
# + [markdown] tags=[]
# #### Step 1: State the null hypothesis.
# > The null hypothesis is that the time to catastrophe for labeled and unlabeled tubulin are identically distributed.
# -
# #### Step 2: Define a test statistic.
# > For our first NHST experiment, the test statistic that will be used is difference in means. This test statistic will offer a good comparison to results acquired in part a and b, when we compared the confidence intervals of the means of the two categories. <br /> <br />
# > For our second NHST experiment, the test statistic that will be used is difference in variance. Time to catastrophe can be modeled by a combination of exponential processes (as shown in HW6), and we know that we can get a normal approximation of the distribution using the equating moments or taylor expansion method. Since a normal distribution can be described by the mean and variance, we decided to also compare the difference of variance as a test statistic since we are interested in the question of if the labeled and unlabeled tubulin time to catastrophe come from the same distribution. <br /> <br />
# > For our third NHST experiment, the test statistic that will be used is difference in medians. We decided to conduct this test to compare a different parameter that can describe a distribution to offer more information about if the two categories come from the same distribution. This was mainly done out of curiosity.
# #### Step 3: Simulate data acquisition for the scenario where the null hypothesis is true, many many times.
# > We will concatenate the two data sets, randomly shuffle them, designate the first entries in the shuffled array to be a “labeled” data set and the rest to be a “unlabeled” data set. Our null hypothesis posits that both the labeled and unlabeled tubulin catastrophe times come from the same distribution, so our concatenated data set will include all points from both categories.
@numba.njit
def generate_perm_sample(x, y):
"""Generate a permutation sample."""
combined_data = np.concatenate((x, y))
np.random.shuffle(combined_data)
half_index = int(combined_data.size / 2)
return combined_data[:half_index], combined_data[half_index:]
# #### Step 4a: (NHST Experiment 1) Compute the p-value (the fraction of simulations for which the test statistic (diff in mean) is at least as extreme as the test statistic computed from the measured data). Do this 10 million times.
# +
@numba.njit
def generate_perm_reps_diff_mean(x, y, N):
"""Generate array of permuation replicates."""
out = np.empty(N)
for i in range(N):
x_perm, y_perm = generate_perm_sample(x, y)
out[i] = np.mean(x_perm) - np.mean(y_perm)
return out
# Compute test statistic for original data set
diff_mean = np.mean(labeled_tubulin) - np.mean(unlabeled_tubulin)
# Draw replicates
perm_reps = generate_perm_reps_diff_mean(labeled_tubulin, unlabeled_tubulin, 10000000)
# Compute p-value
p_val = np.sum(perm_reps >= diff_mean) / len(perm_reps)
print('NHST experiment 1: Difference in mean p-value =', p_val)
# -
# #### Step 4b: (NHST Experiment 2) Compute the p-value (the fraction of simulations for which the test statistic (diff in variance) is at least as extreme as the test statistic computed from the measured data). Do this 10 million times.
# +
@numba.njit
def generate_perm_reps_diff_variance(x, y, N):
"""Generate array of permuation replicates."""
out = np.empty(N)
for i in range(N):
x_perm, y_perm = generate_perm_sample(x, y)
out[i] = np.var(x_perm) - np.var(y_perm)
return out
# Compute test statistic for original data set
diff_variance = np.var(labeled_tubulin) - np.var(unlabeled_tubulin)
# Draw replicates
perm_reps = generate_perm_reps_diff_variance(labeled_tubulin, unlabeled_tubulin, 10000000)
# Compute p-value
p_val = np.sum(perm_reps >= diff_variance) / len(perm_reps)
print('NHST experiment 2: Difference in variance p-value =', p_val)
# -
# #### Step 4c: (NHST Experiment 3) Compute the p-value (the fraction of simulations for which the test statistic (diff in median) is at least as extreme as the test statistic computed from the measured data). Do this 10 million times.
# +
@numba.njit
def generate_perm_reps_diff_median(x, y, N):
"""Generate array of permuation replicates."""
out = np.empty(N)
for i in range(N):
x_perm, y_perm = generate_perm_sample(x, y)
out[i] = np.median(x_perm) - np.median(y_perm)
return out
# Compute test statistic for original data set
diff_median = np.median(labeled_tubulin) - np.median(unlabeled_tubulin)
# Draw replicates
perm_reps = generate_perm_reps_diff_median(labeled_tubulin, unlabeled_tubulin, 10000000)
# Compute p-value
p_val = np.sum(perm_reps >= diff_median) / len(perm_reps)
print('NHST experiment 3: Difference in median p-value =', p_val)
# -
# The p-value is 0.21 where the test statistic is the difference in means. <br />
# The p-value is 0.59 where the test statistic is the difference in variance. <br />
# The p-value is 0.48 where the test statistic is the difference in medians. <br />
# This means that the probability of getting a difference of means, variances, and medians, respectively, as extreme as was observed under the null hypothesis is relatively high (the null hypothesis being that the control and test samples were drawn from identical distribution). Although this result does not confirm that we can reject the null hypothesis (p-value does not represent the probability that a given hypothesis is "true"), it supports the findings in the previous sections where we do not observe a strong influence of fluorescent markers on microtubule performance.
| software/labeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # SLU12: Feature Engineering (aka Real World Data)
# ---
#
# In this notebook we will cover the following topics:
# 1. [Tidy data principles](#tidy-data)
# 2. [Types of data in Pandas](#types-data)
# 3. [Types of statistical data](#types-statistical)
# 1. [Dealing with numerical features](#deal-num)
# 2. [Dealing with categorical features](#deal-cat)
# > *Happy datasets are all alike; every unhappy dataset is unhappy in its own way.*
#
# (Shamelessly adapted from [Tolstoy's Anna Karenina](https://en.wikipedia.org/wiki/Anna_Karenina_principle).)
#
# # 1. Tidy data principles <a class="anchor" id="tidy-data"></a>
#
# At the beginning of any project, it is critical to structure datasets in a way that facilitates work.
#
# Most datasets are dataframes made up of rows and columns, containing values that belong to a variable and an observation:
# * **Variables** contain all values that measure the same thing across observations;
# * **Observations** contain all values measured on the same unit (e.g., same person) across variables.
#
# The ideas of ***tidy data*** ([Wickham, 2014](http://vita.had.co.nz/papers/tidy-data.html)) provide a standardized framework to organize and structure datasets, making them easy to manipulate, model and visualize.
# 1. Each variable forms a column;
# 2. Each observation forms a row;
# 3. Each type of observational unit forms a table (or dataframe).
#
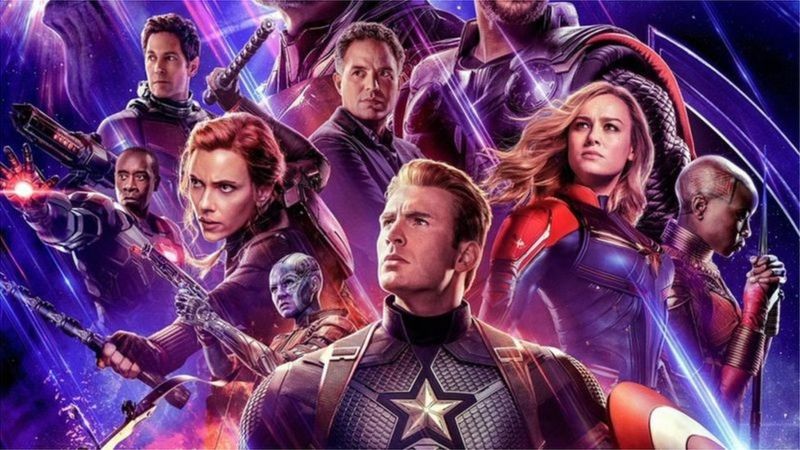
# We will be using a preprocessed version of the `avengers` dataset, by [FiveThirtyEight](https://github.com/fivethirtyeight/data/tree/master/avengers).
#
# 
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# -
avengers = pd.read_csv('data/avengers.csv')
avengers.head()
# # 2. Types of data in Pandas <a class="anchor" id="types-data"></a>
#
# ## 2.1. Numerical and object dtypes
#
# As stated above, a dataset is a collection of values, usually either numbers (quantitative) or strings (qualitative).
avengers.dtypes
# Pandas main data types are:
# * Numeric (`int`, `float`);
# * Datetime (`datetime`, `timedelta`);
# * String (`object`).
#
# The convenient `select_dtypes` method allows us to select variables (columns in our dataframe) by data type.
(avengers.select_dtypes(include='object')
.head(3))
# ## 2.2. Category dtype
#
# Pandas provide us with a `category` dtype for categorical data:
# * It allows to easily identify categorical columns, which are recognized by other Python libraries;
# * Converting a string variable with a few different values to a categorical variable saves memory;
# * By converting to a categorical we can specify an order on the categories.
#
# Let's consider a categorical feature: `Universe`.
# +
avengers_cat = avengers.copy()
avengers_cat = avengers_cat.assign(Universe=avengers['Universe'].astype('category'))
avengers_cat.describe(include='category')
# -
# Categorical data has a `categories` and an `ordered` property:
# * `Series.cat.categories` prints the different values (or levels) the variable can take on;
# * `Series.cat.ordered` prints whether the categorical variable has a natural order or not (hint: if it has, it's not purely categorical).
avengers_cat['Universe'].cat.categories
avengers_cat['Universe'].cat.ordered
# ### Ordinal data
#
# Ordinal statistical data refers to categories that have a natural order, but the distance between them is not known.
#
# We will use the `Membership` variable as an example since it appears to be an order in the degree of commitment of our avengers.
#
# We can also use the `category` dtype.
# +
avengers_ord = avengers.copy()
avengers_ord = avengers_ord.assign(Membership=avengers['Membership'].astype('category'))
avengers_ord['Membership'].cat.categories
# -
# However, this time we need to set the order for our categories, since there is one! The `category` datatype is flexible enough to accommodate this.
# +
ordered_cats = ['Honorary', 'Academy', 'Probationary', 'Full']
avengers_ord = avengers_ord.assign(Membership=avengers_ord['Membership'].cat.set_categories(ordered_cats, ordered=True))
avengers_ord['Membership'].min(), avengers_ord['Membership'].max()
# -
# Again, remember that our models need variables in numeric form, in order to be able to make sense of them.
#
# The `category` datatypes deals with this gracefully for us.
(avengers_ord.assign(Membership=avengers_ord['Membership'].cat.codes)
.sample(n=5))
# However, and as usual, there is a trade-off here:
# * If we assign integer values to our ordinal categories we are *imposing the assumption* that they are equally spaced;
# * If we convert them to dummy variables, we will lose the constraint with their order.
#
# Later in this notebook, we will see a better way to encode categorical variables into numerical ones.
# # 3. Types of statistical data <a class="anchor" id="types-statistical"></a>
#
# There are two main types of statistical data:
# 1. Numerical;
# 2. Categorical.
# ## 3.1. Dealing with numerical data <a class="anchor" id="deal-num"></a>
#
# Numerical data is information that is measurable. It's always collected in number form, although not all data in number form is numerical.
#
# Some of the things we can do with numerical data:
# * Mathematical operations (e.g., addition, distances and the normalization above);
# * Sort it in ascending or descending order.
#
# **Discrete data**
#
# Discrete data take on certain values, although the list of values may be finite or not.
#
# `TotalDeaths` and `TotalReturns` in our `avengers` data are discrete variable.
#
# **Continuous data**
#
# Continuous data can take any value within a range: `Appearances` is an example in our data.
# ### 3.1.1. Introducing sklearn-like transformers
#
# In a Data Science project, you will deal with a dataset, on which you will apply transformations (*feature engineering!*) and, after that, you will train your Machine Learning model. You have already encountered some of these transformations: for example, imputing the missing values on a given column (using pandas `fillna` method). After your model is trained, you are going to use it in a separate dataset to make predictions (either to evaluate the performance of your model on a test set or to make predictions on new, unlabeled data).
#
# If, for example, you are filling the missing values of a numerical feature with a fixed value (say `0`), then it is trivial to apply the same transformation on new, unseen data. But imagine that instead you are filling the missing values with the median value of that feature. How do you apply that transformation in practice?
#
# You have to do it in two steps:
# * You compute the median of that feature for the *training data* and fill the missing values with that value;
# * For new unseen data (e.g. the test set), you just fill the missing value with the median you calculated for the *training data*.
#
# This process is greatly facilitated by using `sklearn`-like **transformers**, which have two standard methods:
# * The `.fit()` method goes through the data and creates a mapping;
# * Then`.transform()` transforms the data using this mapping, failing gracefully you when strange things happen (e.g., unseen values or categories).
#
# Also, they can used in very convenient ways with other `sklearn` utilities and a typical workflow.
#
# For numerical data, we will introduce the following transformers, from `sklearn.preprocessing`:
from sklearn.preprocessing import KBinsDiscretizer
from sklearn.preprocessing import Binarizer
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import Normalizer
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import RobustScaler
# ### 3.1.2. Discretization of numerical data
#
# #### Binning
#
# Discretization is the transformation of continuous data into *intervals* or *bins*. Despite its name, it can be applied on discrete data to put them on a fixed number of bins.
#
# Let's try it in the `Appearances` field, which has the following distribution:
avengers['Appearances'].plot.hist(bins=10, figsize=(10, 6));
plt.xlim(0);
plt.xlabel('Number of appearances');
plt.title('Histogram of number of appearances');
# The histogram above shows the distribution of this field by splitting the instances into 10 bins, for the purpose of the plot. The [`KBinsDiscretizer` transformer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.KBinsDiscretizer.html#sklearn.preprocessing.KBinsDiscretizer) will do the same thing, at the level of the data.
#
# In the following, we will create a new feature called `Appearances_bins` containing 10 bins, from 0 to 9, with the instances uniformly distributed.
# +
# save column as a dataframe, as required by the transformer
X = avengers[['Appearances']]
# initialize transformer with desired options
binner = KBinsDiscretizer(n_bins=10, encode='ordinal', strategy='uniform')
# fit transformer to data
binner.fit(X)
# create new feature by transforming the data
avengers['Appearances_bins'] = binner.transform(X)
# -
# The histogram will look the same as above, but the feature values will range from 0 to 9 instead.
avengers['Appearances_bins'].plot.hist(figsize=(10, 6));
plt.xlim(0);
plt.xlabel('Number of appearances');
plt.title('Number of appearances after discretization');
# If you want to know the original values for the edges of each bin, call:
binner.bin_edges_
# You can play with the options of the transformer to obtain different results.
# +
# initialize transformer with desired options
binner = KBinsDiscretizer(n_bins=5, encode='ordinal', strategy='quantile')
# save data to bin
X = avengers[['Appearances']]
# fit transformer to data
binner.fit(X)
# create new feature by transforming the data
avengers['Appearances_bins_quartiles'] = binner.transform(X)
# plot histogram
avengers['Appearances_bins_quartiles'].plot.hist(figsize=(10, 6));
plt.xlim(0,4);
plt.xlabel('Number of appearances');
plt.title('Number of appearances after discretization');
# -
# #### Binarization
#
# You may just want to consider 2 bins, say you want all the Avengers with fewer than 1000 appearances to be in bin 0 and all the Avengers with more than 1000 appearances to be in bin 1. For that, you can use the [`Binarizer` transformer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.Binarizer.html#sklearn.preprocessing.Binarizer):
# +
# initialize transformer with desired options
binarizer = Binarizer(threshold = 1000)
# save data to binarize
X = avengers[['Appearances']]
# fit transformer to data
binarizer.fit(X)
# create new feature by transforming the data
avengers['Appearances_binary'] = binarizer.transform(X)
# plot histogram
avengers['Appearances_binary'].plot.hist(figsize=(4, 5));
plt.xlim(0,1);
plt.xlabel('Number of appearances');
plt.title('Number of appearances after binarization');
# -
# ### 3.1.3. Scaling of numerical data
#
# Often, the numeric variables in our dataset have very different scales, that is, take on different ranges of values.
#
# It's usually a good practice to **scale** them during the preprocessing of our data. These transformations change the data itself, but not the distribution. Why it is important to scale the data?
# * When predictor values have different ranges, particular features can dominate the algorithm (e.g., think [Euclidean distance](https://en.wikipedia.org/wiki/Euclidean_distance));
# * Different scales can make estimators unable to learn correctly from certain features in smaller ranges;
# * You don't want your feature to rely on the scale of the measurement involved;
# * Optimization methods (e.g. gradient descent) will converge faster, and otherwise they may not converge at all.
#
# A notable exception are *decision tree-based* estimators that are robust to arbitrary scaling of the data.
#
# Here, we present 4 different examples of scaling transformers.
#
# #### [MinMaxScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html#sklearn.preprocessing.MinMaxScaler)
#
# This transforms all variables so that the minimum and the maximum of the transformed data take certain values $[a,b]$, say, [0, 1]. The general formula to scale $x_i$ from a range $[x_{\min}, x_{\max}] \to [a,b]$ is:
#
# $$ x_i' = \frac{x_i - x_{\min}}{x_{\max} - x_{\min}} \cdot (b-a) + a $$
# +
# initialize transformer with desired options
minmaxscaler = MinMaxScaler(feature_range=(0,1))
# save data to scale
X = avengers[['Appearances']]
# fit transformer to data
minmaxscaler.fit(X)
# create new feature by transforming the data
avengers['Appearances_minmax'] = minmaxscaler.transform(X)
# plot histogram
avengers['Appearances_minmax'].plot.hist(figsize=(8, 5));
plt.xlim(0, 1);
plt.xlabel('Number of appearances');
plt.title('Number of appearances after min-max scaling');
# -
# #### [Normalizer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.Normalizer.html)
#
# This normalizes each ***row*** of the dataset *individually* to unit norm, independently of the other rows. You can choose between $\ell_2$ (default), $\ell_1$ and $\max$ norms.
# +
# initialize transformer with desired options
normalizer = Normalizer(norm='l2')
# save numerical columns to normalize
X = avengers[['Appearances', 'TotalDeaths', 'TotalReturns']]
# fit transformer to data
normalizer.fit(X)
# create new features by transforming the data
X_normalized = normalizer.transform(X) # recall that output is a numpy array
avengers['Appearances_normalized'] = X_normalized[:, 0]
avengers['TotalDeaths_normalized'] = X_normalized[:, 1]
avengers['TotalReturns_normalized'] = X_normalized[:, 2]
# plot histogram of normalized appearances
avengers['Appearances_normalized'].plot.hist(figsize=(8, 5));
plt.xlim(0.88, 1);
plt.xlabel('Number of appearances');
plt.title('Number of appearances after normalization');
# -
# #### [StandardScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler)
#
# Standardization means both centering the data around 0 (by removing the mean) and scaling it to unit variance:
#
# $$ z_i = \frac{x_i - \mu}{\sigma}$$
# +
# initialize transformer with desired options
standardscaler = StandardScaler()
# save data to scale
X = avengers[['Appearances']]
# fit transformer to data
standardscaler.fit(X)
# create new feature by transforming the data
avengers['Appearances_standard_scaled'] = standardscaler.transform(X)
# plot histogram
avengers['Appearances_standard_scaled'].plot.hist(figsize=(8, 5));
plt.xlabel('Number of appearances');
plt.title('Number of appearances after standard scaling');
# -
# #### [RobustScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html#sklearn.preprocessing.RobustScaler)
#
# Scalers differ from each other in the way to estimate the parameters used to shift and scale each feature. In the presence of some very large ***outliers***, using the scaler above leads to the compression of inliers. Since outliers have an influence on the minimum, maximum, mean and standard deviation, these scalers will shrink the range of the feature values.
#
# The alternative is to scale the features in a way that is robust to outliers: using the *median* (instead of the *mean*) and the Interquartile Range *('member SLU04?)*.
# +
# initialize transformer with desired options
robustscaler = RobustScaler()
# save data to scale
X = avengers[['Appearances']]
# fit transformer to data
robustscaler.fit(X)
# create new feature by transforming the data
avengers['Appearances_robust_scaled'] = robustscaler.transform(X)
# plot histogram
avengers['Appearances_robust_scaled'].plot.hist(figsize=(8, 5));
plt.xlabel('Number of appearances');
plt.title('Number of appearances after robust scaling');
# -
# ## 3.2. Dealing with categorical data <a class="anchor" id="deal-cat"></a>
#
# Categorical data represents categories (e.g., gender, marital status, hometown).
#
# Categorical variables can take on a limited, and usually fixed, number of possible values.
#
# The categories can also take on numerical values (e.g., ids), but those numbers have no mathematical meaning:
# * You can't do mathematical operations, even if [the computer says yes](https://www.youtube.com/watch?v=Lk1yoobIMUY);
# * Nor sort them in ascending or descending order.
#
# A limitation of categorical data in the form of strings is that estimators, in general, don't know how to deal with it.
# ### 3.2.1. Binary data
#
# A binary variable is a variable with only two possible values: like `Active` and `Gender` in our `avengers` dataset.
#
# Since our algorithms can't deal with data in the form of strings, we need to transform such variables to a numerical form.
#
# The method `Series.map` allows us to easily deal with this cases, mapping inputs to outputs. Let's use it convert both columns to either 0 or 1.
# +
avengers = pd.read_csv('data/avengers.csv')
(avengers.assign(Active_mapped = avengers['Active'].map({'YES': 1, 'NO': 0}),
Gender_mapped = avengers['Gender'].map({'MALE': 1, 'FEMALE': 0}))
.sample(5))
# -
# (See below for a transformer way to achieve the same thing.)
# ### 3.2.2. Enconding categorical features
#
# There are also transformers to deal with categorical features. Even though `sklearn` contains some of such transformers, there is a (much) better library which you shoud (must!) use instead, [`category_encoders`](http://contrib.scikit-learn.org/category_encoders/):
import category_encoders as ce
# In particular, we are going to explore the following encoders.
# #### Ordinal encoding
#
# The [`OrdinalEncoder` transformer](http://contrib.scikit-learn.org/category_encoders/ordinal.html) encodes categorical features as ordinal, integer feature. It uses a single column of integers, chosen randomly by default, to represent the classes.
#
# For example, the `Universe` field has 7 possible values.
avengers.sample(5, random_state=9)
avengers['Universe'].unique()
# +
# initialize transformer with desired options
ordinalencoder = ce.ordinal.OrdinalEncoder()
# save data to scale (no need to reshape)
X = avengers[['Universe']]
# fit transformer to data
ordinalencoder.fit(X)
# create new feature by transforming the data
X_encoded = ordinalencoder.transform(X)
X_encoded.sample(5, random_state=9)
# -
# If you want to see the mapping which was created:
ordinalencoder.category_mapping
# We applied the transformation on the same data we fit the transformer. What would happen if we tried to transform an instance with an ***unseen category***, say `Earth-123` or `Earth-999`?
ordinalencoder.transform(pd.DataFrame({'Universe': ['Earth-6311', 'Earth-123', 'Earth-999']}))
# It automatically imputed the value `-1`. This is because, by default, the option `handle_unknown` is set to `'value'`. The transformers in `category_encoders` take care of unseen categories by themselves!
# #### One-hot encoding
#
# Ordinal enconding, as described above, creates one new feature with integers for each categorical feature. Even though the values of the integers generally have no meaning, a Machine Learning algorithm might think otherwise and decide that, for example, a category whose encoding is `8` is more important than one with `2`!
#
# To avoid this, we can consider **one-hot encoding**, which, for each categorical feature, creates one new feature per *each category*. This can be implemented using the [`OneHotEncoder` transformer](http://contrib.scikit-learn.org/category_encoders/onehot.html).
# +
# initialize transformer with desired options
ohe = ce.one_hot.OneHotEncoder(use_cat_names=True, handle_unknown='indicator')
# save data to scale (no need to reshape)
X = avengers[['Universe']]
# fit transformer to data
ohe.fit(X)
# create new feature by transforming the data
X_ohe = ohe.transform(X)
X_ohe.sample(5, random_state=9)
# -
# Note how the names of the new features were automatically created (by using the option `use_cat_names`) and how an extra feature (`Universe_-1`) was created for unseen categories (due to the option `handle_unknown='indicator'` -- note that this is different from the Ordinal Encoder!).
# # Conclusion
#
# You now know the types of data that you will find in the real world and the options you have to manipulate data it in order to get the most of it.
#
# Make sure to review this notebook well, and when you're ready, go solve the exercises. Good luck!
| S01 - Bootcamp and Binary Classification/SLU12 - Feature Engineering/Learning notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv("housepricedata.csv")
df.head()
df.tail()
df.columns
df.shape
df.describe()
dataset = df.values
dataset
x = dataset[:,0:10]
y = dataset[:,10]
from sklearn import preprocessing
min_max_scaler = preprocessing.MinMaxScaler()
x_scale = min_max_scaler.fit_transform(x)
x_scale
from sklearn.model_selection import train_test_split
x_train, x_val_and_test, y_train, y_val_and_test = train_test_split(x_scale,y, test_size= 0.3)
x_val, x_test, y_val, y_test = train_test_split(x_val_and_test, y_val_and_test, test_size=0.5)
print(x_train.shape, x_val.shape, x_test.shape)
print(y_train.shape, y_val.shape, y_test.shape)
import tensorflow as tf
from tensorflow.keras import models,layers
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(32, activation='relu', input_shape=(10,)),
tf.keras.layers.Dense(32, activation='relu'),
tf.keras.layers.Dense(1, activation= 'sigmoid')])
model.compile(optimizer='sgd',
loss='binary_crossentropy',
metrics=['accuracy'])
hist = model.fit(x_train, y_train,
batch_size=32, epochs=100,
validation_data=(x_val, y_val))
model.evaluate(x_test, y_test)[1]
# +
plt.plot(hist.history['acc'])
plt.plot(hist.history['val_acc'])
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='lower right')
plt.show()
# -
| House Price Prediction using Deep Learning/.ipynb_checkpoints/House Price Prediction Using CNN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# name: python3
# ---
# +
import json
import pprint
import matplotlib.pyplot as plt
import pandas as pd
import datetime
# -
gamerName = input("소환사명을 입력하세요.")
gamerName
userId = None
with open(f"../data/{gamerName}_userId.txt", "r") as userId_file:
userId = int(userId_file.read().strip())
userId
# +
def call_log(gamerName):
with open(f"../data/{gamerName}_gameInfo.json", "r") as json_log:
gameInfo = json.load(json_log)
with open(f"../data/{gamerName}_timeline.json", "r") as json_log:
timeline = json.load(json_log)
return gameInfo, timeline
gameInfo, timeline = call_log(gamerName)
# -
def find_me(userId, gameIndex):
for player in gameInfo[gameIndex]['participantIdentities']:
if player['player']['accountId'] == userId:
return player['participantId']
win_rate = []
for gameIndex in range(len(gameInfo)):
for player in gameInfo[gameIndex]['participants']:
if player['participantId'] == find_me(userId, gameIndex):
win_rate.append(player['stats']['win'])
# 승률 구하는 부분
sum(win_rate) / len(win_rate) * 100
# # 매치 승률 구하기
# ## 시간대별 게임 수 시각화
gameDay = []
for i in range(len(gameInfo)):
gameDay.append(int(datetime.datetime.fromtimestamp(gameInfo[i]['gameCreation'] // 1000).strftime("%H")))
# Prepare Data
df = pd.Series(gameDay)
df = df.value_counts()
for i in range(24):
if i not in df.index:
df[i] = 0
# +
# Draw plot
fig, ax = plt.subplots(figsize=(8,5), dpi= 80)
ax.vlines(x=df.index, ymin=0, ymax=df.values, color='firebrick', alpha=0.7, linewidth=2)
ax.scatter(x=df.index, y=df.values, s=25, color='firebrick', alpha=0.7)
# Title, Label, Ticks and Ylim
ax.set_title('match per day', fontdict={'size':22})
ax.set_ylabel('match')
ax.set_xticks(df.index)
ax.set_xticklabels(df.index, rotation=0, fontdict={'horizontalalignment': 'center', 'size':10})
ax.set_ylim(0, df.values.max() + 10)
# Annotate
for index, value in df.iteritems():
ax.text(index, value+.5, s=round(value, 2), horizontalalignment= 'center', verticalalignment='bottom', fontsize=10)
plt.show()
# -
# ## 매치승률 구하기
WinPerDayDf = pd.DataFrame()
for i in range(len(gameInfo)):
WinPerDay = pd.Series()
WinPerDay['datetime'], WinPerDay['Hour'] = list(map(int, datetime.datetime.fromtimestamp(gameInfo[i]['gameCreation'] // 1000).strftime("%Y%m%d,%H").split(',')))
# print(WinPerDay['datetime'], WinPerDay['Hour'])
for player in gameInfo[i]['participants']:
if player['participantId'] == find_me(userId, i):
WinPerDay['win'] = player['stats']['win']
break
WinPerDayDf = WinPerDayDf.append(WinPerDay, ignore_index=True)
WinPerDayDf['Hour'] = WinPerDayDf['Hour'].astype(int)
WinPerDayDf['datetime'] = WinPerDayDf['datetime'].astype(int)
WinPerDayDf['win'] = WinPerDayDf['win'].astype(bool)
sortedWinPerDayDf = WinPerDayDf.sort_values(by=['datetime', 'Hour'])
sortedWinPerDayDf
totalGame = []
oneGame = []
prevDate = [sortedWinPerDayDf.iloc[0]['datetime'], sortedWinPerDayDf.iloc[0]['Hour']]
currDate = None
for index, row in sortedWinPerDayDf.iterrows():
currDate = [row['datetime'], row['Hour']]
if prevDate[0] == currDate[0] or (prevDate[0] + 1 == currDate[0] and currDate[1] < 6):
oneGame.append(row['win'])
else:
prevDate = currDate
totalGame.append(oneGame)
oneGame = [row['win']]
matchWinRate = []
for i in totalGame:
win_or_defeat = 0
for j in i:
win_or_defeat += float(j) - 0.5
if win_or_defeat > 0:
# print("승리")
matchWinRate.append(1)
# elif win_or_defeat == 0:
# # print("무승부")
# matchWinRate.append(0)
else :
# print("패배")
matchWinRate.append(0)
sum(matchWinRate) / len(matchWinRate)
| code/data_analytics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: airbnb
# language: python
# name: airbnb
# ---
# # -- Objetivo do problema
#
# --1.0. Previsão do primeiro destino que um novo usuário irá escolher.
#
# --Por que?
# --Quak tipo de modelo de negócio do Airbnb
# --Marketplace( Conecta pessoas que oferecem acamodação, com pessoas que estão procurando acomodação )
# --Oferta ( pessoas oferecendo acomodação )
# -- Tamanho do portfolio
# -- Diversidade/densidade de portfolio.
# -- Preço Médio
#
# -- Demanda ( pessoas procurando acomodação )
# -- Numero de Usuários
# -- LTV ( Lifetime Value )
# -- CAC ( Client acquisition cost )
#
# -- Gross Revenue = (phi*Numero cliente) - CAC
#
# -- Demanda ( pessoas procurando acomodação )
#
# --Proposta da solução
# - Modelo de previsão do primeiro destino de um novo usuário.
# - 1.0. Predições e salvar em uma tabela do banco de dados.
# - 2.0. API
# - Input: usuário e suas caracteristicas
# - output: usuario e suas caracteristicas com a coluna prediction
# # 0.0. Imports
# +
import random
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split, StratifiedKFold
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler, RobustScaler, StandardScaler
from sklearn.metrics import classification_report, accuracy_score, balanced_accuracy_score, cohen_kappa_score
from scikitplot.metrics import plot_confusion_matrix
from imblearn import under_sampling, over_sampling
from imblearn.over_sampling import SMOTENC
from pandas_profiling import ProfileReport
from scipy.stats import skewnorm
from scipy import stats
from tensorflow.keras import models
from tensorflow.keras import layers
from category_encoders import TargetEncoder
from IPython.core.display import display, HTML
# +
# %matplotlib inline
plt.rcParams['figure.figsize'] = [13, 8]
sns.set_style('darkgrid')
pd.set_option('display.float_format', lambda x: '%.3f' % x)
pd.set_option('display.max_columns', None )
display(HTML("<style>.container { width:80% !important; }</style>"))
# -
# ## 0.1. Helper Functions
# +
def numeric_statistics( df ):
dic = {
"type": df.dtypes.values,
"Unique_Values": df.nunique().values,
"Mean": df.mean(),
"Median": df.median(),
"Std": df.std(),
"Min": df.min(),
"Max": df.max(),
"Range": df.max() - df.min(),
"Skew": df.skew(),
"Kurtosis": df.kurtosis()
}
return pd.DataFrame( dic, index=df.columns )
def cramer_v( x, y ):
cm = pd.crosstab( x, y ).values
n = cm.sum()
r, k = cm.shape
chi2 = stats.chi2_contingency( cm )[0]
chi2corr = max( 0, chi2 - (k-1)*(r-1)/(n-1) )
kcorr = k - (k-1)**2/(n-1)
rcorr = r - (r-1)**2/(n-1)
return np.sqrt( (chi2corr/n) / ( min(kcorr-1,rcorr-1) ) )
# -
# # 1.0. Data Description
# +
# df users
df_raw = pd.read_csv( 'data/train_users_2.csv', low_memory=True )
# df sessions
df_sessions = pd.read_csv( 'data/sessions.csv', low_memory=True )
# -
#
# ## Users
df_raw.head()
#
# ## Sessions
df_sessions.head()
# ## 1.1. Data Dimension
#
# ## Users
print(f'Number of Rows: {df_raw.shape[0]}')
print(f'Number of Columns: {df_raw.shape[1]}')
#
# ## Sessions
print(f'Number of Rows: {df_sessions.shape[0]}')
print(f'Number of Columns: {df_sessions.shape[1]}')
# ## 1.2. Data types
#
# ## Users
df_raw.dtypes
#
# ## Sessions
df_sessions.dtypes
# ## 1.3. Check NA
#
# ## Users
df_raw.isnull().sum() / len(df_raw)
#
# ## Sessions
df_sessions.isnull().sum() / len(df_sessions)
# ## 1.4. FillNA
# ### 1.4.1 Users
#
# ## Costumers
# If the customer didn't have a country destination, they didn't book an apartment to travel to, my solution to this variable is going to be to exchange the NA values for the max date from a first booking.
#
# ## Age
# for this variable I decided to fill the null values with a distribution with the same patterns as the variable age.
# +
# date_first_booking
aux = df_raw[df_raw['date_first_booking'].isna()]
aux['country_destination'].value_counts( normalize=True )
# + cell_style="center"
# age
aux = df_raw[df_raw['age'].isna()]
print( aux['country_destination'].value_counts( normalize=True ), '\n\n')
aux_02 = df_raw[( df_raw['age'] > 15 ) & ( df_raw['age'] <120 )]['age']
skew = aux_02.skew()
print(f'Skew: {skew}')
std = aux_02.std()
print(f'std: {std}')
mean = aux_02.mean()
print(f'mean: {mean}')
sns.histplot( aux_02, kde=True );
# +
# date_first_booking
date_first_booking_max = pd.to_datetime( df_raw['date_first_booking'] ).max().strftime( '%Y-%m-%d' )
df_raw['date_first_booking'] = df_raw['date_first_booking'].fillna( date_first_booking_max )
# age
df_raw['age'] = df_raw['age'].apply(lambda x: skewnorm.rvs( skew, loc=mean, scale=std ).astype( 'int64' ) if pd.isna(x) else x )
# first_affiliate_tracked
df_raw = df_raw[~df_raw['first_affiliate_tracked'].isna()]
# + cell_style="center"
sns.histplot( df_raw[(df_raw['age'] > 15) & (df_raw['age'] <120)]['age'], kde=True );
# -
# ### 1.4.2 df_sessions
# +
# user_id 0.3%
df_sessions = df_sessions[~df_sessions['user_id'].isna()]
# action - 0.7%
df_sessions = df_sessions[~df_sessions['action'].isna()]
#action_type - 11%
df_sessions = df_sessions[~df_sessions['action_type'].isna()]
# action_details - 11%
df_sessions = df_sessions[~df_sessions['action_detail'].isna()]
# secs_elapsed - 1.2%
df_sessions = df_sessions[~df_sessions['secs_elapsed'].isna()]
# -
# ## 1.5. Change Data Types
# +
aux_01 = df_raw.sample().T
aux_02 = df_raw.sample().dtypes
pd.concat( [aux_01,aux_02], axis=1 )
# +
# df_raw
# date_account_created
df_raw['date_account_created'] = pd.to_datetime( df_raw['date_account_created'] )
# timestamp_first_active
df_raw['timestamp_first_active'] = pd.to_datetime( df_raw['timestamp_first_active'], format='%Y%m%d%H%M%S')
# date first_booking
df_raw['date_first_booking'] = pd.to_datetime( df_raw['date_first_booking'] )
# age
df_raw['age'] = df_raw['age'].apply( lambda x: x if pd.isna(x) else int(x) )
# -
# ## 1.6. Check Blanced Data
df_raw['country_destination'].value_counts( )
# ## 1.7. Numerical data
# ### 1.7.1 Users
# +
# df_raw
num_attributes = df_raw.select_dtypes( include=['int64','float64'] )
cat_attributes = df_raw.select_dtypes( include='object')
time_attributes = df_raw.select_dtypes( include='datetime64[ns]')
numeric_statistics( num_attributes )
# -
# ### 1.7.2 Sessions
# +
# df_sessions
num_attributes_sessions = df_sessions.select_dtypes( include=['int64','float64'] )
cat_attributes_sessions = df_sessions.select_dtypes( include='object')
time_attributes_sessions = df_sessions.select_dtypes( include='datetime64[ns]')
numeric_statistics( num_attributes_sessions )
# -
# ## 1.8. Categorical data
# ### 1.8.1 Users
# df2
cat_attributes.drop('id', axis=1 ).describe()
# + code_folding=[]
cat_attributes_list = cat_attributes.drop( 'id', axis=1 ).columns.tolist()
corr_dict = {}
for i in range( len ( cat_attributes_list ) ):
corr_list = []
for j in range( len( cat_attributes_list ) ):
ref = cat_attributes_list[i]
feat = cat_attributes_list[j]
# correlation
corr = cramer_v( cat_attributes[ ref ], cat_attributes[ feat ] )
# append a list
corr_list.append( corr )
# appende a correlation list for each ref attributs
corr_dict[ ref ] = corr_list
d = pd.DataFrame( corr_dict )
d = d.set_index( d.columns)
sns.heatmap( d, annot=True )
# -
# ### 1.8.2 Sessions
# df_sessions
cat_attributes_sessions.drop( 'user_id', axis=1 ).describe()
# + code_folding=[]
cat_attributes_list = cat_attributes_sessions.drop( 'user_id', axis=1 ).columns.tolist()
corr_dict = {}
for i in range( len ( cat_attributes_list ) ):
corr_list = []
for j in range( len( cat_attributes_list ) ):
ref = cat_attributes_list[i]
feat = cat_attributes_list[j]
# correlation
corr = cramer_v( cat_attributes_sessions[ ref ], cat_attributes_sessions[ feat ] )
# append a list
corr_list.append( corr )
# appende a correlation list for each ref attributs
corr_dict[ ref ] = corr_list
d = pd.DataFrame( corr_dict )
d = d.set_index( d.columns)
sns.heatmap( d, annot=True )
# -
# # 2.0. Featue Engineeging
df2 = df_raw.copy()
df2.shape
df2 = df_raw.copy()
df2.dtypes
# ## 2.1. Create New Features
# +
# days from first active uo to first booking
df2['first_active'] = pd.to_datetime( df2['timestamp_first_active'].dt.strftime('%Y-%m-%d' ) )
df2['days_from_first_active_until_booking'] = ( df2['date_first_booking'] - df2['first_active'] ).apply( lambda x: x.days )
# days from first active up to account created
df2['days_from_first_active_until_account_created'] = ( df2['date_account_created'] - df2['first_active'] ).apply( lambda x: x.days )
# days from account created up to first booking
df2['days_from_account_created_until_first_booking'] = ( df2['date_first_booking'] - df2['date_account_created'] ).apply( lambda x: x.days )
# ============== active ==============
# year of first active
df2['year_first_active'] = df2['first_active'].dt.year
# month of first active
df2['month_first_active'] = df2['first_active'].dt.month
# day of first active
df2['day_first_active'] = df2['first_active'].dt.day
# day of week
df2['day_of_week_first_active'] = df2['first_active'].dt.dayofweek
# week of year of first active
df2['week_first_active'] = df2['first_active'].dt.isocalendar().week.astype( 'int64' )
# ============== booking ==============
# year of first booking
df2['year_first_booking'] = df2['date_first_booking'].dt.year
# month of first booking
df2['month_first_booking'] = df2['date_first_booking'].dt.month
# day of first booking
df2['day_first_booking'] = df2['date_first_booking'].dt.day
# day of week
df2['day_of_week_first_booking'] = df2['date_first_booking'].dt.dayofweek
# week of year of first booking
df2['week_first_booking'] = df2['date_first_booking'].dt.isocalendar().week.astype('int64')
# ============== Account created ==============
# year of account_created
df2['year_account_created'] = df2['date_account_created'].dt.year
# month of account_created
df2['month_account_created'] = df2['date_account_created'].dt.month
# day of account_created
df2['day_account_created'] = df2['date_account_created'].dt.day
# day of week
df2['day_of_week_account_created'] = df2['date_account_created'].dt.dayofweek
# week of year of account_created
df2['week_account_created'] = df2['date_account_created'].dt.isocalendar().week.astype('int64')
# -
df2.shape
# # 3.0. Data Filtering
df3 = df2.copy()
# ## 3.1. Rows
# +
# there are few people with these ages and inside the dataset there is lots of weird values od age.
df3 = df3[( df3['age'] > 15 ) & ( df3['age'] <120 )]
# df_sessions = df_sessions[df_sessions['secs_elapsed'] > 0]
# -
# ## 3.2. Columns
# dropping columns which was used to create new columns
cols = ['date_account_created','timestamp_first_active','date_first_booking','first_active','date_account_created']
df3.drop( cols, axis=1, inplace=True )
# + [markdown] heading_collapsed=true
# # 4.0. EDA
# + hidden=true
df4 = df3.copy()
# + [markdown] hidden=true
# ## 4.1. Hypotesis
# + [markdown] hidden=true
# H01. Em todos os destinos, os usuários levam 15 dias, em média, para fazer a primeira reserva no airbnb, desde sua primeira ativação
#
# H02. Em todos os destinos, os usuários levam 3 dias, em médiam para fazer o cadastro no site.
#
# H03. O volume de reservas anuais feitas durante o verão aumentaram em 20% para destinos dentro dos USA.
#
# H04. Usuários do sexo masculino fazem 10% mais reservas para países fora do USA
#
# H05. O canal de Marketing Google representa 40% das reservas para países fora dos USA
#
# H06. O destino dos USA representam mais de 20% em todos os canais.
#
# H07. A idade média das pessoas é até de 35 anos em todos os destinos.
#
# H08. A porcentagem de usuários que usam o site na língua inglês-americano para reservar a acomodação em qualquer destino é maior que 90%
#
# H09. O número de reservas do AIbnb é crescente ao longo dos anos.
# + [markdown] hidden=true
# ## 4.1. Univariate Analysis - Feature Behaviour
# + hidden=true
columns = ['id','gender', 'age', 'signup_method', 'signup_flow', 'language',
'affiliate_channel', 'affiliate_provider', 'first_affiliate_tracked',
'signup_app', 'first_device_type', 'first_browser',
'country_destination', 'days_from_first_active_until_booking',
'days_from_first_active_until_account_created',
'days_from_account_created_until_first_booking','year_first_active',
'month_first_active', 'day_first_active', 'day_of_week_first_active',
'week_first_active', 'year_first_booking', 'month_first_booking',
'day_first_booking', 'day_of_week_first_booking', 'week_first_booking',
'month_account_created', 'day_account_created',
'day_of_week_account_created', 'week_account_created']# 'year_account_created']
# + hidden=true
proof = ProfileReport(df4[columns])
#proof.to_notebook_iframe()
proof.to_file( output_file='AirBnB.html' )
# + [markdown] hidden=true
# ## 4.2. Bivariate Analysis - Hypothesis Validation
# + [markdown] hidden=true
# ### H01. Em todos os destinos, os usuários levam 15 dias, em média, para fazer a primeira reserva no airbnb, desde sua primeira ativação
#
# **FALSE**
# + hidden=true
df4[['country_destination','days_from_first_active_until_booking']].groupby( 'country_destination' ).mean()
# + [markdown] hidden=true
# ### H02. Em todos os destinos, os usuários levam 3 dias, em médiam para fazer o cadastro no site.
#
# **FALSE**
# + hidden=true
df4[['country_destination','days_from_first_active_until_account_created']].groupby( 'country_destination' ).mean()
# + [markdown] hidden=true
# ### H03. O volume de reservas anuais feitas durante o verão aumentaram em 20% para destinos dentro dos USA.
#
# **FALSE**
# + hidden=true
aux_01 = df4[['year_first_booking','month_first_booking','country_destination']].groupby( ['year_first_booking', 'month_first_booking', 'country_destination' ]).size().reset_index().rename( columns={0:'count'})
aux_01
# + hidden=true
# only summer
summer = [6,7,8,9]
aux_02 = aux_01[( aux_01['month_first_booking'].isin( summer ) ) & ( aux_01['country_destination'] == 'US' ) ]
# + hidden=true
aux_02
# + hidden=true
aux_02.columns
# + hidden=true
aux_02 = aux_02[['year_first_booking','count']].groupby( 'year_first_booking' ).sum().reset_index()
sns.barplot( data=aux_02, x='year_first_booking', y='count');
# + [markdown] hidden=true
# percentage of growing
# + hidden=true
aux_02['count'].pct_change() * 100
# + [markdown] hidden=true
# ### H04. Usuários do sexo masculino fazem 10% mais reservas para países fora do USA
#
# **FALSE**
# + hidden=true
aux_01 = df4.loc[ ~df4['country_destination'].isin( ['US','NDF'] ), ['gender','country_destination']]
aux_01 = aux_01.groupby( 'gender' ).count().reset_index()
aux_01['pct-gender'] = aux_01['country_destination'].pct_change()
aux_01
# + [markdown] hidden=true
# ### H05. O canal de Marketing Google representa 40% das reservas para países fora dos USA
#
# **FALSE**
# + hidden=true
aux_01 = df4.loc[ ~df4['country_destination'].isin(['US','NDF']), ['affiliate_provider','country_destination']]
aux_01['affiliate_provider'].value_counts( normalize=True )
# + [markdown] hidden=true
# ### H06. O destino dos USA representam mais de 20% em todos os canais.
#
# **TRUE**
# + hidden=true
aux_01 = df4[['affiliate_channel','country_destination']]
aux_01['country_destination'].value_counts( normalize=True )
# + [markdown] hidden=true
# ### H07. A idade média das pessoas é até de 35 anos em todos os destinos.
#
# **FALSE**
# + hidden=true
aux_01 = df4[df4['country_destination'] != 'NDF'][['age','country_destination']]
aux_01.groupby( 'country_destination' ).mean().reset_index()
# + [markdown] hidden=true
# ### H08. A porcentagem de usuários que usam o site na língua inglês-americano para reservar a acomodação em qualquer destino é maior que 90%
#
# **TRUE**
# + hidden=true
aux_01 = df4[df4['country_destination'] != 'NDF'][['language','country_destination']]
aux_01['language'].value_counts( normalize=True )
# + [markdown] hidden=true
# ### H09. O número de reservas do AIbnb é crescente ao longo dos anos.
#
# **FALSE**
# + hidden=true
aux_01 = df4.loc[df4['country_destination'] != 'NDF', ['year_first_booking']].value_counts().reset_index()
aux_01.rename( columns={0:'count'}, inplace=True )
sns.barplot( data=aux_01, x='year_first_booking', y='count' );
# -
# # 5.0. Data Preparation
df5 = df4.copy()
# +
# filter columns with high correlation
cols = ['days_from_first_active_until_booking', 'year_first_active', 'month_first_active', 'day_first_active','day_of_week_first_active', 'week_first_active', 'month_first_booking', 'month_account_created','year_first_booking', 'week_first_booking', 'affiliate_provider','first_browser', 'first_device_type', 'language'] # high correlation
df5 = df5.drop( cols, axis=1 )
# -
df5.dtypes
# ## 5.1. Rescaling
ss = StandardScaler()
rs = RobustScaler()
mms = MinMaxScaler()
# +
# age - Standardization
df5['age'] = ss.fit_transform( df5[['age']].values )
# signup_flow - Robust Scaler
df5['signup_flow'] = rs.fit_transform( df5[['signup_flow']].values )
# days_from_first_active_until_account_created - Robust Scaler
df5['days_from_first_active_until_account_created'] = rs.fit_transform( df5[['days_from_first_active_until_account_created']].values )
# days_from_account_created_until_first_booking - Robust Scaler
df5['days_from_account_created_until_first_booking'] = rs.fit_transform( df5[['days_from_account_created_until_first_booking']].values )
# year_account_created - MinMax Scaler
df5['year_account_created'] = mms.fit_transform( df5[['year_account_created']].values )
# -
# ## 5.2 Encoding
te = TargetEncoder()
# +
# gender - One Hot Encoder
df5 = pd.get_dummies( df5, prefix=['gender'], columns=['gender'] )
# signup_method - One Hot Encoder
df5 = pd.get_dummies( df5, prefix=['signup_method'], columns=['signup_method'] )
# signup_app - One Hot Encoder
df5 = pd.get_dummies( df5, prefix=['signup_app'], columns=['signup_app'] )
# affiliate_channel - Target Encoder - trasnforma pra number pois o targetencoder nao aceita categorica
# c = {'NDF':0, 'US':1, 'other':2, 'CA':3, 'FR':4, 'IT':5, 'ES':6, 'GB':7, 'NL':8, 'DE':9, 'AU':10, 'PT':11}
c = dict( zip( df5['country_destination'].drop_duplicates().tolist(), np.arange( 1, 12 ) ) )
df5['affiliate_channel'] = te.fit_transform( df5[['affiliate_channel']].values, df5['country_destination'].map( c ) )
# first_affiliaTarget_tracked - Target Encoder
df5['first_affiliate_tracked'] = te.fit_transform( df5[['first_affiliate_tracked']].values, df5['country_destination'].map( c ) )
# -
# ## 5.3. Transformation
# +
# week_of_year_account_created
df5['week_of_year_account_created_sin'] = df5['week_account_created'].apply( lambda x: np.sin( x * (2*np.pi/52 ) ) )
df5['week_of_year_account_created_cos'] = df5['week_account_created'].apply( lambda x: np.cos( x * (2*np.pi/52 ) ) )
# day_of_week_first_booking
df5['day_of_week_first_booking_sin'] = df5['day_of_week_first_booking'].apply( lambda x: np.sin( x * (2*np.pi/7 ) ) )
df5['day_of_week_first_booking_cos'] = df5['day_of_week_first_booking'].apply( lambda x: np.cos( x * (2*np.pi/7 ) ) )
# day_account_created
df5['day_account_created_sin'] = df5['day_account_created'].apply( lambda x: np.sin( x * (2*np.pi/31 ) ) )
df5['day_account_created_cos'] = df5['day_account_created'].apply( lambda x: np.cos( x * (2*np.pi/31 ) ) )
# day_of_week_account_created
df5['day_of_week_account_created_sin'] = df5['day_of_week_account_created'].apply( lambda x: np.sin( x * (2*np.pi/7 ) ) )
df5['day_of_week_account_created_cos'] = df5['day_of_week_account_created'].apply( lambda x: np.cos( x * (2*np.pi/7 ) ) )
# -
# ## 5.3. Balanced Dataset
# ### 5.3.1 Random Undersampling
# +
# ratio balanded (it's need to set all the variable to run well)
#ratio_balanced = {'NDF': 1000}
#undersampling = under_sampling.RandomUnderSampler( sampling_strategy=ratio_balanced, random_state=32 )
# difine sample
#undersampling = under_sampling.RandomUnderSampler( sampling_strategy='auto', random_state=32 )
# apply sampler
#x_under, y_under = undersampling.fit_resample( df5, df5['country_destination'] )
# -
# ### 5.3.2 Random Oversampling
# +
# difine sample
#oversampling = over_sampling.RandomOverSampler( sampling_strategy='auto', random_state=32 )
# apply sampler
#x_over, y_over = oversampling.fit_resample( df5, df5['country_destination'] )
# +
#df5['country_destination'].value_counts()
# +
#y_over.value_counts()
# -
# ### 5.3.3 SMOTE
X = df5.drop( ['id','country_destination'], axis=1 )
y = df5['country_destination'].copy()
y.value_counts()
X.columns
sm = SMOTENC( [1,2,3], sampling_strategy='minority', n_jobs=-1 )
X, y = sm.fit_resample( X, y )
y.value_counts()
df5 = pd.concat( [X,y], axis=1 )
# # 6.0. Feature Selection
df6 = df5.copy()
df6.columns
X = df5.drop( 'country_destination', axis=1 )
y = df5['country_destination'].copy()
# # 7.0. Machine Learning
X_train, X_val, y_train, y_val = train_test_split( X, y, test_size=0.2, random_state=32 )
# ## 7.1. Baseline model
index = df5['country_destination'].value_counts().index
values = df5['country_destination'].value_counts( normalize=True ).values
yhat_random = random.choices( population=index, weights=values, k=y_val.shape[0] )
# prediction prepare
y_val_random = y_val.to_numpy()
# +
# balanced accuracy, accuracy, kappa score
dic = {
'model': 'NN',
'Accuracy': accuracy_score( y_val_random, yhat_random ),
'BalancedAccuracy': balanced_accuracy_score( y_val_random, yhat_random ),
'KappaScore': cohen_kappa_score( y_val_random, yhat_random )
}
print( pd.DataFrame( dic, index=[0] ) )
print( classification_report( y_val_random, yhat_random ) )
# confusion matrix
plot_confusion_matrix(y_val_random, yhat_random, figsize=(18,7) );
# -
# ## 7.2. NN
on = OneHotEncoder()
y_train_nn = on.fit_transform( y_train.values.reshape(-1,1) ).toarray()
print( 'Number of Rows: {}'.format( X_train.shape[0] ) )
print( 'Number of Features: {}'.format( X_train.shape[1] ) )
print( 'Number of Classes: {}'.format( y_train.nunique() ) )
# +
# model definition
model = models.Sequential()
model.add( layers.Dense( 256, input_dim=X_train.shape[1], activation='relu' ) )
model.add( layers.Dense( 12, activation='softmax' ) )
# model compile
model.compile( loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'] )
# train model
model.fit( X_train, y_train_nn, epochs=5 )
# -
# # 8.0. Model Performance
# +
# prediction
pred_nn = model.predict( X_val )
# invert prediction
yhat_nn = on.inverse_transform( pred_nn )
# prediction prepare
y_val_nn = y_val.to_numpy()
yhat_nn = yhat_nn.reshape( 1,-1 )[0]
# +
# balanced accuracy, accuracy, kappa score
dic = {
'model': 'NN',
'Accuracy': accuracy_score( y_val_nn, yhat_nn ),
'BalancedAccuracy': balanced_accuracy_score( y_val_nn, yhat_nn ),
'KappaScore': cohen_kappa_score( y_val_nn, yhat_nn )
}
print( pd.DataFrame( dic, index=[0] ) )
print( classification_report( y_val_nn, yhat_nn ) )
# confusion matrix
plot_confusion_matrix(y_val_nn, yhat_nn, figsize=(18,7) );
# -
# ## <font color=red> 8.1. Cross Validation </font>
# +
# generate k-fold
num_folds = 5
kfold = StratifiedKFold( n_splits=num_folds, shuffle=True, random_state=32 )
balanced_acc_list = []
kappa_acc_list = []
i = 1
for train_ix, val_ix in kfold.split( X_train, y_train ):
print( 'Fold Number: {}/{}'.format( i, num_folds ) )
# get fold
X_train_fold = X_train.iloc[train_ix]
y_train_fold = y_train.iloc[train_ix]
x_val_fold = X_train.iloc[val_ix]
y_val_fold = y_train.iloc[val_ix]
# target hot-encoding
ohe = OneHotEncoder()
y_train_fold_nn = ohe.fit_transform( y_train_fold.values.reshape( -1, 1 ) ).toarray()
# model definition
model = models.Sequential()
model.add( layers.Dense( 256, input_dim=X_train.shape[1], activation='relu' ) )
model.add( layers.Dense( 11, activation='softmax') )
# compile model
model.compile( loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'] )
# training model
model.fit( X_train_fold, y_train_fold_nn, epochs=100, batch_size=32, verbose=0 )
# prediction
pred_nn = model.predict( x_val_fold )
yhat_nn = ohe.inverse_transform( pred_nn )
# prepare data
y_test_nn = y_val_fold.to_numpy()
yhat_nn = yhat_nn.reshape( 1, -1 )[0]
# metrics
## Balanced Accuracy
balanced_acc_nn = balanced_accuracy_score( y_test_nn, yhat_nn )
balanced_acc_list.append( balanced_acc_nn )
## Kappa Metrics
kappa_acc_nn = cohen_kappa_score( y_test_nn, yhat_nn )
kappa_acc_list.append( kappa_acc_nn )
i += 1
print(f'Avg Balanced Accuracy: {np.mean( balanced_acc_list )} +/- {np.std( balanced_acc_list )}' )
print(f'Avg Kappa: {np.mean( kappa_acc_list )} +/- {np.std( kappa_acc_list )}' )
# -
| Airbnb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# -
df = pd.read_csv("../data/used_mobile_phone.csv")
print(df.info())
df.head()
# +
# create_date로부터 '월'을 의미하는 month 정보를 피처로 추출합니다.
df['month'] = df['create_date'].apply(lambda x: x[:7])
# 월별 거래 횟수를 계산하여 출력합니다.
df['month'].value_counts()
# -
# 일별 거래 횟수를 계산하여 그래프로 출력합니다.
# to_datetime() : 문자를 날짜형식(timestamp)으로 변환
df_day = pd.to_datetime(df['create_date'].apply(lambda x: x[:10])).value_counts()
df_day.plot()
plt.show()
# 가격의 분포를 그래프로 탐색합니다.
df['price'].hist(bins="auto")
# +
# 휴대폰 기종(phone_model)별 가격의 평균과 표춘편차를 계산합니다.
df_price_model_mean = df.groupby('phone_model')['price'].transform(lambda x: np.mean(x))
df_price_model_std = df.groupby('phone_model')['price'].transform(lambda x: np.std(x))
# 이를 바탕으로 모든 데이터의 z-score(표준화 점수)를 계산합니다.
# z-score는 평균값과 얼마나 거리가 먼지 계산해주는 통계적인 예측값이다.
df_price_model_z_score = (df['price'] - df_price_model_mean) / df_price_model_std
df_price_model_z_score.hist(bins="auto")
# +
# factory_price 피처의 분포를 탐색합니다.
df['factory_price'].hist(bins="auto")
# factory_price와 price 피처를 산점도 그래프로 출력하여 상관 관계를 살펴봅니다.
df.plot.scatter(x='factory_price', y='price')
# +
# 기종별 총 거래 데이터 개수를 집계합니다.
model_counts = df['phone_model'].value_counts()
print(model_counts.describe())
# 기종별 총 거래 데이터 개수를 상자 그림으로 살펴봅니다.
plt.boxplot(model_counts)
# -
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction import DictVectorizer
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
# +
# 데이터를 학습/테스트용 데이터로 분리합니다.
df = df[['price', 'phone_model', 'factory_price', 'maker', 'price_index', 'month']]
df = pd.get_dummies(df, columns=['phone_model', 'maker', 'month'])
X = df.loc[:, df.columns != 'price']
y = df['price']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# 랜덤 포레스트 모델을 학습합니다.
forest = RandomForestRegressor(n_estimators=1000, criterion='mse')
forest.fit(X_train, y_train)
y_train_pred = forest.predict(X_train)
y_test_pred = forest.predict(X_test)
# 학습한 모델을 평가합니다.
print('MSE train: %.3f, test: %.3f' % (mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# +
# 학습한 모델의 피처 중요도를 그래프로 살펴봅니다.
importances = forest.feature_importances_
indices = np.argsort(importances)[::-1]
plt.bar(range(X.shape[1]), importances[indices])
# 학습한 모델의 피처 중요도를 출력합니다.
feat_labels = X.columns.tolist()
# zip() 은 동일한 개수로 이루어진 자료형을 묶어 주는 역할을 하는 함수이다.
feature = list(zip(feat_labels, forest.feature_importances_))
# sorted라는 정렬함수는 시퀀스 자료형 뿐만 아니라 순서에 구애받지 않는 자료형에도 적용할 수 있습니다. 정렬된 결과는 list로 반환됩니다.
sorted(feature, key=lambda tup: tup[1], reverse=True)[:10]
# -
# month 피처 중, 영향력이 높은순으로 정렬하여 출력합니다.
for sorted_feature in sorted(feature, key=lambda tup: tup[1], reverse=True):
if "month" in sorted_feature[0]:
print(sorted_feature)
# 데이터를 다시 불러옵니다.
df = pd.read_csv("../data/used_mobile_phone.csv")
from datetime import datetime
import time
# +
# create_date 피처를 수치적으로 계산하기 위해 unixtime으로 변환하는 함수를 정의합니다.
# unixtime : 시간을 정수로 표현한 것이며 값이 클수록 '최근'에 가까워집니다.
def date_to_unixtime(date_str):
timestamp = time.mktime(datetime.strptime(date_str, '%Y-%m-%d').timetuple())
return timestamp
# create_date 피처를 '현재와 얼마나 가까운 데이터인지' 판단하기 위한 점수를 생성합니다. 먼저 unixtime으로 데이터를 변환합니다.
df['create_unixtime'] = df['create_date'].apply(lambda x: date_to_unixtime(x[:10]))
# 변환된 unixtime에 min-max 스케일링을 적용합니다.
df['create_time_score'] = (df['create_unixtime'] - df['create_unixtime'].min()) / \
(df['create_unixtime'].max() - df['create_unixtime'].min())
df[['create_date', 'create_unixtime', 'create_time_score']].head()
# +
# phone_model 피처에서 저장 용량(phone_model_storage) 피처를 추출합니다.
# split함수는 문자열을 공백 혹은 어떠한 기준으로 나눌때 사용하는 함수이다.
df['phone_model_storage'] = df['phone_model'].apply(lambda x: x.split(" ")[-1])
# phone_model 피처에서 기종 세부명(phone_model_detail) 피처를 추출합니다.
df['phone_model_detail'] = df['phone_model'].apply(lambda x: ' '.join(x.split(" ")[:-1]))
df[['phone_model_storage', 'phone_model_detail']].head()
# +
# phone_model 피처의 기종별 거래 데이터 개수를 집계합니다.
model_counts = df['phone_model'].value_counts()
# phone_model_detail 피처의 기종별 거래 데이터 개수를 집계합니다.
model_detail_counts = df['phone_model_detail'].value_counts()
data = [model_counts, model_detail_counts]
# 두 피처 간의 기종별 거래 데이터 개수를 비교합니다.
mpl_fig = plt.figure()
ax = mpl_fig.add_subplot(111)
ax.boxplot(data)
# -
# 기종명 + 용량으로 출고가를 찾기 위한 딕셔너리를 생성합니다.
model_to_factory_price_dict = {}
for index, row in df.iterrows():
model_concat = (row['phone_model_detail'], row['phone_model_storage'])
if model_concat in model_to_factory_price_dict:
pass
else:
model_to_factory_price_dict[model_concat] = row['factory_price']
print(str(model_to_factory_price_dict)[:40], "...")
# +
# 거래 가격(price)의; z-score를 계산합니다. 이는 해당 데이터의 가격이 기종의 평균에 비해 어느정도로 높거나 낮은지를 알 수 있게 하는 점수입니다.
# std() : 표준편차
df['price_by_group'] = df.groupby('phone_model_detail')['price'].transform(lambda x: (x - x.mean()) / x.std())
# 거래 가격의 z-score(price_by_group)의 분포를 그래프로 출력합니다.
ax = df['price_by_group'].hist(bins="auto")
# z-score(price_by_group) 기준으로 하위 5%, 상위 5%에 해당하는 점수를 lower_bound, upper_bound라고 지정합니다.
lower_bound = df['price_by_group'].quantile(0.05)
upper_bound = df['price_by_group'].quantile(0.95)
# lower_bound, upper_bound 그래프에 추가합니다.
ax.axvline(x=lower_bound, color='r', linestyle='dashed', linewidth=2)
ax.axvline(x=upper_bound, color='r', linestyle='dashed', linewidth=2)
# lower_bound, upper_bound를 출력합니다.
print(lower_bound)
print(upper_bound)
# +
# lower_bound보다 낮으면 0, upper_bound보다 높으면 2, 그 중간이면 1로 가격의 상태를 분류하는 함수를 정의합니다.
def get_price_level(price, lower, upper):
if price <= lower:
return "0"
elif price >= upper:
return "2"
else:
return "1"
# lower_bound보다 낮으면 0, upper_bound보다 높으면 2, 그 중간이면 1로 가격의 상태를 분류합니다.
df['price_lower'] = df.groupby('phone_model_detail')['price'].transform(lambda x: x.quantile(0.05))
df['price_upper'] = df.groupby('phone_model_detail')['price'].transform(lambda x: x.quantile(0.95))
df['price_level'] = df.apply(lambda row: get_price_level(row['price'], row['price_lower'],
row['price_upper']), axis=1)
df[['price', 'price_lower', 'price_upper', 'price_level', 'text']].head()
# -
import pickle
import re
# 중고나라 불용어 사전을 불러옵니다.
with open('../data/used_mobile_phone_stopwords.pkl', 'rb') as f:
stopwords = pickle.load(f)
# 불용어 사전에 등록된 단어 10개를 출력합니다.
print(stopwords[:10])
from konlpy.tag import Okt
# +
# ‘+’를 제외한 특수문자를 제거하고, 숫자형태의 문자를 제거합니다.
def text_cleaning(text):
text = ''.join(c for c in text if c.isalnum() or c in '+, ')
text = ''.join([i for i in text if not i.isdigit()])
return text
# 불용어에 등장하지 않는 형태소만을 추출하여 반환하는 함수입니다.
def get_pos(x):
tagger = Okt()
poses = tagger.pos(x)
return [pos[0] for pos in poses if pos[0] not in stopwords]
# 위 함수들을 적용한 형태소 추출을 테스트합니다.
df['text'] = df['text'].apply(lambda x: text_cleaning(x))
result = get_pos(df['text'][0])
print(result)
# -
from collections import Counter
# +
# get_pos() 함수를 모든 텍스트 데이터에 적용하여 형태소 말뭉치를 추출합니다.
corpus = sum(df['text'].apply(lambda x: get_pos(x)).tolist(), [])
# 추출된 형태소 말뭉치에서 가장 많이 등장한 형태소 2500개를 추출합니다.
counter = Counter(corpus)
# most_common(2500) : 데이터의 개수가 많은 순으로 정렬된 배열을 리턴
common_wrods = [key for key, _ in counter.most_common(2500)]
common_wrods
# -
# TF-IDF 벡터 생성
df['price_level'].value_counts()
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
# +
# 반출 형태소를 제외한 모든 형태소를 제거하는 함수를 정의합니다.
def get_common_pos(x):
tagger = Okt()
poses = tagger.pos(x)
return [pos[0] for pos in poses if pos[0] in common_wrods]
# 1:3:1 비율로 랜덤 샘플링을 수행합니다.
negative_random = df[df['price_level']=='0'].sample(321, random_state=30)
neutral_random = df[df['price_level']=='1'].sample(321*3, random_state=30)
positive_random = df[df['price_level']=='2'].sample(321, random_state=30)
# 샘플링 완료된 데이터셋을 정의합니다.
df_sample = negative_random.append(neutral_random).append(positive_random)
# TF-IDF를 수행하여 피쳐를 반환합니다.
# TF-IDF(Term Frequency - Inverse Document Frequency)는 정보 검색과 텍스트 마이닝에서 이용하는 가중치로,
# 여러 문서로 이루어진 문서군이 있을 때 어떤 단어가 특정 문서 내에서 얼마나 중요한 것인지를 나타내는 통계적 수치이다
index_vectorizer = CountVectorizer(tokenizer= lambda x: get_common_pos(x))
# fit_transform() 메소드는 말 그대로 fit()한 다음에 transform() 하는 것입니다
X = index_vectorizer.fit_transform(df_sample['text'].tolist())
tfidf_vectorizer = TfidfTransformer()
X = tfidf_vectorizer.fit_transform(X)
# 감성 분류를 위한 학습 데이터셋을 정의합니다.
y = df_sample['price_level']
# train_test_split 함수는 전체 데이터셋 배열을 받아서 랜덤하게 test/train 데이터 셋으로 분리해주는 함수이다.
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=30)
# NumPy 배열에서 배열의 형태를 알아보는 shape()
print(x_train.shape)
print(x_test.shape)
# -
from sklearn.metrics import accuracy_score
from sklearn.svm import SVC
from sklearn.metrics import confusion_matrix
# +
# 비선형 SVM 분류 모델을 학습하고 평가합니다.
# 서포트 벡터 머신(이하 SVM)은 결정 경계(Decision Boundary), 즉 분류를 위한 기준 선을 정의하는 모델이다.
svm = SVC(kernel='rbf', C=10.0, random_state=0, gamma=0.10)
svm.fit(x_train, y_train)
y_pred_ksvc = svm.predict(x_test)
print('Accuracy: %.2f' % accuracy_score(y_test, y_pred_ksvc))
# Confusion Matrix를 출력합니다.
# Confusion Matrix(혼동행렬) 예측값이 실제 관측값을 얼마나 정확히 예측했는지 보여주는 행렬
confmat = confusion_matrix(y_true=y_test, y_pred=y_pred_ksvc)
print(confmat)
# +
# text 피처로부터 '상품 상태 피처(product_status)'를 생성합니다.
# transform() 메서드는 입력된 객체와 동일하게 인덱스된 객체를 반환한다.
X = index_vectorizer.transform(df['text'].tolist())
X = tfidf_vectorizer.transform(X)
df['product_status'] = pd.Series(svm.predict(X))
# 랜덤 포레스트 모델 학습을 위한 데이터를 준비합니다.
df = df[['price', 'factory_price', 'maker', 'price_index', 'create_time_score', 'phone_model_storage',
'phone_model_detail', 'product_status']]
df = pd.get_dummies(df, columns=['maker', 'phone_model_storage', 'phone_model_detail', 'product_status'])
# 학습/테스트 데이터를 분리합니다.
X = df.loc[:, df.columns != 'price']
y = df['price']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# 랜덤 포레스트 모델을 학습하고 평가합니다.
forest = RandomForestRegressor(n_estimators=1000, criterion='mse')
forest.fit(X_train, y_train)
y_train_pred = forest.predict(X_train)
y_test_pred = forest.predict(X_test)
# mean_squared_error : RMSE를 구하는 함수
print('MSE train: %.3f, test: %.3f' % (mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# 피처 중요도 plot을 출력합니다.
importances = forest.feature_importances_
plt.plot(importances, "o")
# 피처 중요도를 print로 출력합니다.
feat_labels = X.columns.tolist()
feature = list(zip(feat_labels, forest.feature_importances_))
sorted(feature, key=lambda tup: tup[1], reverse=True)[:10]
# -
# 테스트 데이터의 y값과 예측된 y값을 산점도 그래프를 활용하여 상관 관계를 살펴봅시다.
plt.scatter(y_test.values, y_test_pred)
from sklearn.model_selection import RandomizedSearchCV
# +
# Randomized Search 로 찾아낼 파라미터 후보군을 각각 리스트로 선정합니다.
n_estimators = [int(x) for x in np.linspace(start = 200, stop = 2000, num = 10)]
max_features = ['auto', 'sqrt']
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
bootstrap = [True, False]
# RandomizedSearchCV 오브젝트를 생성하여 모델을 정의합니다.
# n_estimators : 생성할 tree의 개수와
# max_features : 최대 선택할 특성의 수입니다.
# max_depth : 트리의 최대 깊이, 깊이가 깊어지면 과적합될 수 있으므로 적절히 제어 필요
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'bootstrap': bootstrap}
forest = RandomForestRegressor()
# RandomizedSearchCV 반복적인 실행으로 최적의 파라미터를 찾기 위한 방법
optimal_forest = RandomizedSearchCV(estimator = forest,
param_distributions = random_grid,
n_iter = 100,
cv = 3,
verbose=2,
random_state=42,
n_jobs = -1)
# RandomizedSearchCV 모델을 학습합니다.
X = df.loc[:, df.columns != 'price']
y = df['price']
optimal_forest.fit(X,y)
# -
# 앞서 선정한 파라미터 후보군 중에서 가장 좋은 결과를 보인 파리미터의 조합을 출력합니다.
print(optimal_forest.best_params_)
# +
# 최적의 파라미터를 적용한 모델로 중고 휴대폰의 가격을 예측하고 평가합니다.
y_train_pred = optimal_forest.predict(X_train)
y_test_pred = optimal_forest.predict(X_test)
print('MSE train: %.3f, test: %.3f' % (
mean_squared_error(y_train, y_train_pred),
mean_squared_error(y_test, y_test_pred)))
print('R^2 train: %.3f, test: %.3f' % (
r2_score(y_train, y_train_pred),
r2_score(y_test, y_test_pred)))
# 가격 예측 모델의 피처 중요도 plot을 출력합니다.
importances = optimal_forest.best_estimator_.feature_importances_
indices = np.argsort(importances)[::-1]
plt.bar(range(X.shape[1]), importances[indices])
# 가격 예측 모델의 피처 중요도를 출력합니다.
feat_labels = X.columns.tolist()
feature = list(zip(feat_labels, optimal_forest.best_estimator_.feature_importances_))
sorted(feature, key=lambda tup: tup[1], reverse=True)[:10]
# -
# 테스트 데이터의 y값과 예측된 y값을 산점도 그래프를 활용하여 상관 관계를 살펴봅시다.
plt.scatter(y_test.values, y_test_pred)
import joblib
# +
# 프로그램에서 입력값에 대한 피처를 찾기 위한 딕셔너리를 생성합니다.
col_to_index_dict = {}
# enumerate는 "열거하다"라는 뜻이다.
# 이 함수는 순서가 있는 자료형(리스트, 튜플, 문자열)을 입력으로 받아 인덱스 값을 포함하는 enumerate 객체를 돌려준다.
for idx, col in enumerate(df.columns[1:]):
col_to_index_dict[col] = idx
print(str(col_to_index_dict)[:40], "...")
# +
# 모델 예측을 위한 중간 오브젝트들을 각각 파일로 저장합니다.
with open('../data/used_mobile_pickles/model_to_factory_price_dict.pickle', 'wb') as f:
pickle.dump(model_to_factory_price_dict, f, pickle.HIGHEST_PROTOCOL)
with open('../data/used_mobile_pickles/col_to_index_dict.pickle', 'wb') as f:
pickle.dump(col_to_index_dict, f, pickle.HIGHEST_PROTOCOL)
with open('../data/used_mobile_pickles/common_words.pickle', 'wb') as f:
pickle.dump(common_wrods, f, pickle.HIGHEST_PROTOCOL)
with open('../data/used_mobile_pickles/init_vectorizer_vocabulary.pickle', 'wb') as f:
pickle.dump(index_vectorizer.vocabulary_, f, pickle.HIGHEST_PROTOCOL)
with open('../data/used_mobile_pickles/tfidf_vectorizer.pickle', 'wb') as f:
pickle.dump(tfidf_vectorizer, f, pickle.HIGHEST_PROTOCOL)
with open('../data/used_mobile_pickles/svm_classifier.pickle', 'wb') as f:
pickle.dump(svm, f, pickle.HIGHEST_PROTOCOL)
joblib.dump(optimal_forest.best_estimator_, '../data/used_mobile_pickles/rf_regressor.pickle', compress = 1)
# +
# 예측에 필요한 파일 로드
with open('../data/used_mobile_pickles/model_to_factory_price_dict.pickle', 'rb') as f:
model_to_factory_price_dict = pickle.load(f)
with open('../data/used_mobile_pickles/col_to_index_dict.pickle', 'rb') as f:
col_to_index_dict = pickle.load(f)
with open('../data/used_mobile_pickles/common_words.pickle', 'rb') as f:
common_words = pickle.load(f)
with open('../data/used_mobile_pickles/init_vectorizer_vocabulary.pickle', 'rb') as f:
init_vectorizer_vocabulary = pickle.load(f)
with open('../data/used_mobile_pickles/tfidf_vectorizer.pickle', 'rb') as f:
tfidf_vectorizer = pickle.load(f)
with open('../data/used_mobile_pickles/svm_classifier.pickle', 'rb') as f:
svm_classifier = pickle.load(f)
rf_regressor = joblib.load('../data/used_mobile_pickles/rf_regressor.pickle')
# +
import numpy as np
from konlpy.tag import Okt
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
# -
class Almhago():
def __init__(self, model_to_factory_price_dict, col_to_index_dict,
common_words, init_vectorizer_vocabulary, tfidf_vectorizer,
prd_status_classifier, price_regressor):
self._model_to_factory_price_dict = model_to_factory_price_dict
self._col_to_index_dict = col_to_index_dict
self._common_words = common_words
self._init_vectorizer_vocabulary = init_vectorizer_vocabulary
self._index_vectorizer = self._init_index_vectorizer()
self._tfidf_vectorizer = tfidf_vectorizer
self._prd_status_classifier = prd_status_classifier
self._price_regressor = price_regressor
def _get_common_pos(self, x):
tagger = Okt()
poses = tagger.pos(x)
return [pos[0] for pos in poses if pos[0] in self._common_words]
def _text_cleaning(self, text):
text = ''.join(c for c in text if c.isalnum() or c in '+, ')
text = ''.join([i for i in text if not i.isdigit()])
return text
def _init_index_vectorizer(self):
word_index_vectorizer = CountVectorizer(tokenizer = lambda x: self._get_common_pos(x))
word_index_vectorizer.vocabulary_ = self._init_vectorizer_vocabulary
return word_index_vectorizer
def _get_ftr_price(self, model_name, storage):
return self._model_to_factory_price_dict[(model_name, storage)]
def _get_prd_status(self, text):
X = self._index_vectorizer.transform([self._text_cleaning(program_test_dict['text'])])
X = self._tfidf_vectorizer.transform(X)
return self._prd_status_classifier.predict(X)[0]
def _print_almhago(self, model, price, prd_status):
status = ""
if prd_status == "0":
status = "불량한"
elif prd_status == "1":
status = "양호한"
else:
status = "좋은"
print("선택하신", model, "모델은", status, "상태입니다. Almhago 예상 가격은", str(int(price[0])), "원 입니다.")
def predict(self, input_dict):
feature = np.zeros(64)
feature[self._col_to_index_dict['factory_price']] = self._get_ftr_price(input_dict['phone_model_detail'],
input_dict['phone_model_storage'])
feature[self._col_to_index_dict['price_index']] = input_dict['price_index']
feature[self._col_to_index_dict['create_time_score']] = input_dict['create_time_score']
feature[self._col_to_index_dict["_".join(["maker", input_dict['maker']])]] = 1
feature[self._col_to_index_dict["_".join(["phone_model_detail", input_dict['phone_model_detail']])]] = 1
feature[self._col_to_index_dict["_".join(["phone_model_storage", input_dict['phone_model_storage']])]] = 1
feature[self._col_to_index_dict["_".join(["product_status", self._get_prd_status(input_dict['text'])])]] = 1
# predict result
predicted_price = self._price_regressor.predict([feature])
self._print_almhago(input_dict['phone_model_detail'],
predicted_price,
self._get_prd_status(input_dict['text']))
# +
# 데이터의 가장 최근 시점인 2017년 4월을 기준으로 하기 위한 두 개(price_index, create_time_score)의 피처를 정의합니다.
recent_price_index = 95.96
recent_create_time_score = 1.0
# 판매 가격을 예측하고자 하는 메이커, 기종, 용량을 입력하고, 상품의 상태를 나타내는 게시글을 입력합니다.
program_test_dict = {"maker": "apple",
"phone_model_detail": "iphone 6",
"phone_model_storage": "16gb",
"text": '아이폰6 스페이스그레이 16기가 10만원에 팔아요~ 직접거래 \
메인보드 세척 완료 한 침수 폰 입니다폰 안켜지는 상테이구요 고쳐서 쓰실분 가져가세요10만원에 팔아요 \
리퍼한지 얼마안되서 기스 이런거 하나도 없습니당~서울 강남 근처 직거래 가능하며택배거래도 가능해요',
"price_index": recent_price_index,
"create_time_score": recent_create_time_score}
# 정의한 피처를 파라미터로 하여 almhago 오브젝트를 생성합니다.
almhago = Almhago(model_to_factory_price_dict, col_to_index_dict,
common_wrods, init_vectorizer_vocabulary, tfidf_vectorizer,
svm_classifier, rf_regressor)
# 입력한 데이터로 판매 가격을 예측합니다.
almhago.predict(program_test_dict)
# -
| chapter5/.ipynb_checkpoints/01-used-phone-price-prediction-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Script launched with a break in the logging loop just to test if it works well
# !python train.py -i 3 -li 3 \
# --train-dataset-path ../data/coco_data \
# --style-image-path ../data/styles/sketch.jpg \
# --trained-models-dir ../outputs/trained_transfer_models
| src/test_train_transfer_net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import pandas as pd
import io
from pandas.tseries.offsets import MonthEnd
# +
# Building blocks for the URL
entrypoint = 'https://sdw-wsrest.ecb.europa.eu/service/' # Using protocol 'https'
resource = 'data' # The resource for data queries is always'data'
flowRef ='ICP' # Dataflow describing the data that needs to be returned, exchange rates in this case
key = 'M.U2.N.000000.4.ANR' # Defining the dimension values, explained below
# Define the parameters
'''parameters = {
'startPeriod': '2000-01-01', # Start date of the time series
'endPeriod': '2018-10-01' # End of the time series
}'''
request_url = entrypoint + resource + '/'+ flowRef + '/' + key
response = requests.get(request_url, headers={'Accept': 'text/csv'})
inflation_europe = pd.read_csv(io.StringIO(response.text))
inflation_europe = inflation_europe[['TIME_PERIOD', 'OBS_VALUE']]
inflation_europe.rename(columns={
'TIME_PERIOD': 'Date',
'OBS_VALUE': 'Value'
}, inplace=True)
inflation_europe.set_index('Date', inplace=True)
inflation_europe.index = pd.to_datetime(inflation_europe.index, format='%Y-%m')
inflation_europe.index = pd.to_datetime(inflation_europe.index, format='%Y') + MonthEnd(1)
inflation_europe
# -
inflation_europe.to_pickle(r'..\02-Data\09-Inflation_Europe\01-Inflation_percentage_1997-01-31_2021-06-30_monthly_ecb.pkl')
| 03-Notebooks/09-Inflation_Europe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: distortion function
#
# The distortion can be assigned to the optics of the telescope. The distortion function should receive an array with a shape of $(2, N)$. The first and second elements are the x- and y-positions on the focal plane, respectively. An array with the same shape is returned, which contains the coordinates with distortion.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
import warpfield as w
from warpfield.DUMMY import get_jasmine
from astropy.coordinates import SkyCoord, Longitude, Latitude, Angle
from astropy.time import Time
import astropy.units as u
# An artificial source set is used in this notebook. Define a grid in a ±0.3° region around the origin of the ICRS frame.
arr = np.linspace(-0.3, 0.3, 30)
xx,yy = np.meshgrid(arr,arr)
ra = [x*u.degree for x in xx.flat]
dec = [y*u.degree for y in yy.flat]
src = SkyCoord(ra, dec, frame='icrs')
# The telescope is pointed toward the origin of the ICRS frame. The position angle is 0.0°.
pointing = SkyCoord(0.0*u.deg,0.0*u.deg, frame='icrs')
position_angle = Angle(0.0, unit='degree')
jasmine = get_jasmine(pointing, position_angle)
# The figure below shows the on-sky distribution of the artifical sources. A well-organized grid pattern is recognized.
ax = w.display_sources(pointing, src, title='grid w/o distortion')
# The `identity_transformation` is assigned to the optics if not specified.
jasmine.optics.distortion
# This function does not change the given position at all. Thus, the source distribution on the plane is the same as on the sky.
jasmine.display_focal_plane(src)
# As long as the input and output formats are valid, any function can be assigned to the distortion function of the optics. The module provides the `distortion_generator` function to genrate a common distortion function.
#
#
# The `distortion_generator` receives three parameters. The first argument $K$ is the radial component. The second and third arguments, $S$ and $T$, define the tangential component.
# The distortion function is defined as follows:
#
# $$
# \begin{cases}
# r = \sqrt{x^2 + y^2}, ~~ r_6 = r/10^6, \\
# x' = x \left(1 + \sum_n{K_n {r_6}^{2n}} \right)
# + \left(S_1\left(r^2 + 2x^2\right) + 2S_2xy \right)
# \left(1 + \sum_n{T_n {r_6}^{2n}} \right)\\
# y' = y \left(1 + \sum_n{K_n {r_6}^{2n}} \right)
# + \left(2S_1xy + S_2\left(r^2 + 2y^2\right) \right)
# \left(1 + \sum_n{T_n {r_6}^{2n}} \right)\\
# \end{cases}
# $$
#
# The $K$ and $T$ can be given as vectors. The $S$ should be an array with two elements.
# Here is an example of a radial distortion function. A barrel-type distortion pattern can be recognized. Different distortion patterns are obtained with different parameters.
from warpfield.distortion import distortion_generator
distortion = distortion_generator(K=-5e1)
jasmine.set_distortion(distortion)
jasmine.display_focal_plane(src)
# The $S$ parameter is specified in the followng example. The distortion pattern is completely different from the previous one.
distortion = distortion_generator(S=[-8,4])
jasmine.set_distortion(distortion)
jasmine.display_focal_plane(src)
| notebook/tutorial_07_simple_distortion_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finish Installation
# I would strongly reccomend recloning the repositort in our project directory. Unlike the home directory you can store over 1TB of data and your contents are safe there until 90 days after project expires in 2022:
# cd /expanse/lustre/projects/sio134/hmangipu/
git clone https://github.com/gmooers96/CBRAIN-CAM.git
# In the repository, you will need to complete the directories
# cd CBRAIN-CAM/MAPS/
# mkdir models
# mkdir model_graphs
# cd model_graphs
# mkdir latent_space
# mkdir losses
# mkdir model_diagrams
# mkdir reconstructions
# cd ../
# mkdir Bash_Scripts
# cd Bash_Scripts
# mkdir outputs
# cd ../
# At this point you will need to bring over the preprocessed files and change the path files to train/test data. I would reccomend:
# cd /expanse/lustre/projects/sio134/hmangipu/CBRAIN-CAM/MAPS/
# mkdir Preprocessed_Data
# mv {all of your data}/ /expanse/lustre/projects/sio134/hmangipu/CBRAIN-CAM/MAPS/Preprocessed_Data/
# Go to the config file and change all paths in the config file you care about to:
# cd model_config
#new file path
#Preprocessed_Data/Centered_50_50/Space_Time_W_Training.npy
#ect....
"data": {
"training_data_path": "Preprocessed_Data/Centered_50_50/Space_Time_W_Training.npy",
"test_data_path": "Preprocessed_Data/Centered_50_50/Space_Time_W_Test.npy",
"train_labels": "/fast/gmooers/Preprocessed_Data/Centered_50_50/Y_Train.npy",
"test_labels": "/fast/gmooers/Preprocessed_Data/Centered_50_50/Improved_Y_Test.npy",
"max_scalar": "Preprocessed_Data/Centered_50_50/Space_Time_Max_Scalar.npy",
"min_scalar": "Preprocessed_Data/Centered_50_50/Space_Time_Min_Scalar.npy"
},
# I would call in an interactive gpu node and test the script to see if it runs. If run is successful go to your Bash_Scripts/ Directory and make a bash script called {your bash scripts name here}.sh. I believe your script should be the same as mine but just swap out your email for mine so you get a notification when it finishes.
# +
# #!/bin/bash
#SBATCH -A sio134
#SBATCH --job-name="conv_vae"
#SBATCH --output="conv_vae.%j.%N.out"
#SBATCH --partition=gpu
#SBATCH --nodes=1
#SBATCH --gpus=4
#SBATCH --ntasks-per-node=40
#SBATCH --export=ALL
#SBATCH -t 24:00:00
#SBATCH --mem=374G
#SBATCH --no-requeue
#SBATCH --mail-user={<EMAIL>
#SBATCH --mail-type=end # email me when the job finishes
module purge
module load gpu
module load slurm
module load openmpi
module load amber
# cd ../
source activate {name of your gpu environment}
python3 train_fully_conv.py --id 83
# -
# At this point you should be able to submit a full job to train. Feel free to submit multiple jobs if you are interested:
sbatch {your bash scripts name here}.sh
# Mine takes ~37 to ~40 seconds per epoch to train. Let me know if yours is significantly faster or slower. You can monitor your job and finds its job number by:
squeue -u {your username}
# If you realize you made a mistake you can cancel a job by:
scancel {your job number}
# If everything is working at point I would go back to your home directory and remove the git clone there to clear up space (can only have a max of 100G in the home directory)
# cd /home/{$USER}
# #rm -r CBRAIN-CAM/
# The last thing I would reccomend doing is editing your bashrm script so you can move around Expanse easier (e.g. make shortcuts of commands you will make often). You can access it by:
vi $HOME/.bashrc
# on the vim screen, under the lines:
# User specific aliases and functions
module load gcc
module load slurm
# You can add in any shortcut you want. Edit in vim by pressing {i}, end your edits by hitting {the escape key}, save your changes by ":wq" or ":q" if you want to quit and not make changes. Below are the ones I think are helpful to add:
alias PROJECT="cd /expanse/lustre/projects/sio134/gmooers/CBRAIN-CAM/MAPS/"
alias TEMP="cd /expanse/lustre/scratch/gmooers/temp_project/"
alias QUE="squeue -u gmooers"
alias GPUENV="conda activate GPU2"
alias CPUENV="conda activate CPU"
alias SMALL_GPU="srun --partition=gpu-debug --pty --account=sio134 --nodes=1 --ntasks-per-node=40 --mem=374G --gpus=4 -t 00:30:00 --wait=0 --export=ALL /bin/bash"
alias SMALL_CPU="srun --partition=debug --pty --account=sio134 --nodes=1 --ntasks-per-node=128 --mem=248G -t 00:30:00 --wait=0 --export=ALL /bin/bash"
alias CPU_JOB="srun --partition=compute --pty --account=sio134 --nodes=2 --ntasks-per-node=128 --mem=248G -t 03:00:00 --wait=0 --export=ALL /bin/bash"
alias GPU_JOB="srun --partition=gpu --pty --account=sio134 --nodes=1 --ntasks-per-node=40 --mem=374G --gpus=4 -t 03:00:00 --wait=0 --export=ALL /bin/bash"
alias S="screen"
alias SR="screen -r"
alias JN="jupyter notebook --port=8888 --no-browser"
alias J="jupyter notebook --port="
alias N=" --no-browser"
alias P="module purge"
alias DE="conda deactivate"
alias BASH="vi $HOME/.bashrc"
# to see a list of oyu alias's, just type "alias" in the command line
| MAPS/Expanse_Set_Up_Guide.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Author: <NAME>
# Date: 03/12/2020
# # Imports
import pandas as pd
import urllib.request
import selenium
from selenium import webdriver
import webbrowser
import requests
import os
import pynput
import time
# # Things To Know
cwd = os.getcwd()
files = os.listdir(cwd)
f = open(cwd+"/credentials.txt", "r")
username = f.readline()
password = <PASSWORD>()
login_pos = (823,560)
password_pos = (596,508)
posts_pos = (65,410)
add_new_pos = (40,370)
byname_pos = (400,416)
# # Methods
# +
def type_word(name,keyboard):
for i in name:
keyboard.press(i)
keyboard.release(i)
def press_button(pos,mouse):
mouse.position = pos
mouse.click(pynput.mouse.Button.left)
def post_article(article,mouse,keyboard):
wait(2)
press_button(posts_pos,mouse)
wait(3)
press_button(add_new_pos,mouse)
wait(3)
type_word(article.title,keyboard)
add_byname(article.author,mouse,keyboard)
def add_byname(author,mouse,keyboard):
wait(2)
mouse.position=(383,423)
wait(1)
press_button(byname_pos, mouse)
byname = "By " #+ author
wait(2)
type_word(byname, keyboard)
mouse.position=(611,375)
click()
mouse.position = (611,564)
click()
hyperlink = generate_hyperlink(author,mouse,keyboard)
type_word(hyperlink, keyboard)
def click():
mouse.press(pynput.mouse.Button.left)
mouse.release(pynput.mouse.Button.left)
def generate_hyperlink(author,mouse,keyboard):
name = ""
for i in author:
if i==' ':
name = name + "-"
else:
name = name + i
html_hyperlink = '<a href="http://mxanvil.net/category/">'+name+'</a>'
return html_hyperlink
def click_menu():
mouse.position = (611,375)
mouse.click(pynput.mouse.Button.left)
def click_html():
mouse.position = (611,565)
mouse.click(pynput.mouse.Button.left)
def highlight_author_name(author,mouse,keyboard):
mouse.position = (393,416)
mouse.press(pynput.mouse.Button.left)
a=len(author)
mouse.move(11*a,0)
mouse.release(pynput.mouse.Button.left)
def wait(i):
time.sleep(i)
def login(keyboard, mouse):
wait(5)
type_word(username,keyboard)
press_button(password_pos,mouse)
type_word(password,keyboard)
press_button(login_pos,mouse)
# -
# # Classes
class Article:
def __init__(self, title, author, teaser, issue_num, txt):
self.title = title
self.author = author
self.teaser = teaser
self.issue_number = issue_num
self.text = txt
# # Opening Site
t0=time.time()
url = 'http://mxanvil.net/wp-admin'
webbrowser.open(url)
keyboard = pynput.keyboard.Controller()
mouse = pynput.mouse.Controller()
wait(5)
type_word(username,keyboard)
press_button(password_pos,mouse)
type_word(password,keyboard)
press_button(login_pos,mouse)
wait(5)
article = Article("Sample","<NAME>","Read now","9","great article plz read")
wait(2)
press_button(posts_pos,mouse)
wait(3)
mouse.position = add_new_pos
mouse.press(pynput.mouse.Button.left)
mouse.release(pynput.mouse.Button.left)
wait(3)
type_word(article.title,keyboard)
add_byname(article.author,mouse,keyboard)
t1=time.time()
print(str(t1-t0))
#
| Posting_Script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Parent class and sub class have a same method with same number of arguments and when you call that method from sub class then which method will be called?
# +
class Vehicle:
def __init__(self, color, maxSpeed):
self.color = color
self.__maxSpeed = maxSpeed #Making maxSpeed as private using '__' before the memeber
def getMaxSpeed(self): #get method is a public function
return self.__maxSpeed
def setMaxSpeed(self, maxSpeed): #set method is a public function
self.__maxSpeed = maxSpeed
def print(self): #Another way of accessing printing private members --> printing within a class
print("Color : ",self.color)
print("MaxSpeed :", self.__maxSpeed)
class Car(Vehicle):
def __init__(self, color, maxSpeed, numGears, isConvertible):
super().__init__(color, maxSpeed) #Inheriting from vehicle class
self.numGears = numGears #Passing via arguments in car class
self.isConvertible = isConvertible
def print(self):
print("NumGears :", self.numGears)
print("IsConvertible :", self.isConvertible)
c = Car("red", 35, 5, False)
c.print() #Car print() is printed
# -
# ** Eventhough the print() is available in both Vehicle and Car class, print() of Car (derived class) got executed because it's called from derived class
# ### Ans) First it checks the child class in which the print() is called if it's not there then the control goes to its parent class if it's not there then the control goes to it's parent class and so on and so forth
# +
class Vehicle:
def __init__(self, color, maxSpeed):
self.color = color
self.__maxSpeed = maxSpeed #Making maxSpeed as private using '__' before the memeber
def getMaxSpeed(self): #get method is a public function
return self.__maxSpeed
def setMaxSpeed(self, maxSpeed): #set method is a public function
self.__maxSpeed = maxSpeed
def print(self): #Another way of accessing printing private members --> printing within a class
print("Color : ",self.color)
print("MaxSpeed :", self.__maxSpeed)
class Car(Vehicle):
def __init__(self, color, maxSpeed, numGears, isConvertible):
super().__init__(color, maxSpeed) #Inheriting from vehicle class
self.numGears = numGears #Passing via arguments in car class
self.isConvertible = isConvertible
c = Car("red", 35, 5, False)
c.print() #Car print() is printed
# -
# ** Here in the above example there is no print() in derived class so the control goes to its parent class and the print() got executed
| 04 OOPS-2/4.3 Inheritance cont.....ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="2na19Y5CDOqn"
# <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/horizontal-primary-light.png" alt="he-black-box" width="600"/>
#
#
# # Homomorphic Encryption using Duet: Data Owner
# ## Tutorial 0: Basic operations
#
#
# Welcome!
# This tutorial will show you how to use Duet with homomorphic encryption and some use cases. This notebook illustrates the Data Owner view on the operations.
#
# We will focus on Duet's integration with [TenSEAL](https://github.com/OpenMined/TenSEAL).
# TenSEAL is a Python library for doing homomorphic encryption operations on tensors. It's built on top of [Microsoft SEAL](https://github.com/Microsoft/SEAL), a C++ library implementing the BFV and CKKS homomorphic encryption schemes.
#
#
# If you want to learn more about TenSEAL, we recommend the following tutorials:
# - ['Tutorial 0 - Getting Started'](https://github.com/OpenMined/TenSEAL/blob/master/tutorials/Tutorial%200%20-%20Getting%20Started.ipynb).
# - ['Tutorial 1: Training and Evaluation of Logistic Regression on Encrypted Data'](https://github.com/OpenMined/TenSEAL/blob/master/tutorials/Tutorial%201%20-%20Training%20and%20Evaluation%20of%20Logistic%20Regression%20on%20Encrypted%20Data.ipynb).
# - ['Tutorial 2: Working with Approximate Numbers'](https://github.com/OpenMined/TenSEAL/blob/master/tutorials/Tutorial%202%20-%20Working%20with%20Approximate%20Numbers.ipynb).
#
#
# Let's now start the tutorial with a brief review of what homomorphic encryption is, but keep in mind that you don't need to be a crypto expert to use these features.
# + [markdown] id="xkPW4dGzDOqv"
# ## Homomorphic Encryption
#
# __Definition__ : Homomorphic encryption (HE) is a technique that allows computations to be made on ciphertexts and generates results that when decrypted, corresponds to the result of the same computations made on plaintexts.
#
# <img src="https://github.com/OpenMined/TenSEAL/raw/master/tutorials/assets/he-black-box.png" alt="he-black-box" width="600"/>
#
# This means that an HE scheme lets you encrypt two numbers *X* and *Y*, add their encrypted versions so that it gets decrypted to *X + Y*, the addition could have been a multiplication as well.
# + [markdown] id="WgCmGQ2oDas5"
# COLAB
# + id="dJ5_SLqpDdQ8"
# %%capture
# This only runs in colab and clones the code sets it up and fixes a few issues,
# you can skip this if you are running Jupyter Notebooks
import sys
if "google.colab" in sys.modules:
branch = "grace_dev" # change to the branch you want
# ! git clone --single-branch --branch $branch https://github.com/godormad/PySyft.git
# ! cd PySyft && ./scripts/colab.sh # fixes some colab python issues
sys.path.append("/content/PySyft/src") # prevents needing restart
# + [markdown] id="lFWq7Mg8DOqy"
# ### Setup
#
# All modules are imported here, make sure everything is installed by running the cell below.
# + id="1oy7SegeDOq1" outputId="05e1332b-cbec-4f06-bd70-c2a8d1804f39" colab={"base_uri": "https://localhost:8080/"}
# !pip install tenseal
# !pip show tenseal
import syft as sy
import tenseal as ts
import pytest
sy.load_lib("tenseal")
# + [markdown] id="VmfGmx7SDOq3"
# ### Start Duet Data Owner instance
# + id="hfEyQ8k5DOq7" outputId="60e8e314-6436-446f-8057-<KEY>" colab={"base_uri": "https://localhost:8080/", "height": 728}
# Start Duet local instance
duet = sy.launch_duet(loopback=False)
# + [markdown] id="ZmXTNum-DOq_"
# ### Theory: Homomorphic encryption schemes
#
# __TenSEAL__ supports two encryption schemes:
# - __BFV__, a scheme for operations on integers.
# - __CKKS__, a scheme for operations on approximate numbers. This scheme is much better suited for ML applications and we will focus more on it.
#
# There are a few major steps for each scheme:
# 1. __Keys Generation__: in this step, we generate public and private keys that will be used for encryption/decryption.
# 2. __Encryption__: this is the process of converting a plaintext into an encrypted ciphertext. This step requires an encryption key(or a public key).
# 3. __Decryption__: this is the process of converting a ciphertext back into a plaintext. This step requires a decryption key(or a secret key). This step cannot be done on the Data Scientist endpoint.
# + [markdown] id="mfI6O6naDOrB"
# ### Theory: Homomorphic encryption parameters
#
# __TenSEAL__ requires a few parameters to set the keys up:
# - __The polynomial modulus degree(poly_modulus_degree).__ This parameter directly affects the size of the ciphertext, the security of the scheme(bigger is better), but also the computational performance of the scheme(bigger is worse)
# - __The coefficient modulus sizes(coeff_mod_bit_sizes).__ This parameter is an array of bit sizes and directly affects the size of the ciphertext, the security of the scheme(bigger is worse), and the depth of computation allowed in the encrypted space(longer is better).
# - __The scaling factor(global_scale).__ This parameter is only used for the approximate schemes(CKKS) and directly affects the precision of computation and decryption.
# + [markdown] id="twQh9VaDDOrE"
# ### Theory: Homomorphic encryption keys
#
# __TenSEAL__ generates a few keys internally, each with another use case:
# - __The Private Key(or the secret/decryption key)__. This key is used for decrypting ciphertexts, and it is used to derive the other keys. __DO NOT SHARE IT OUTSIDE THE DATA OWNER PROCESS__.
# - __The Public key(or the encryption key)__. This key is used for encrypting the plain data to a ciphertext. You can safely share it with the Data Scientist.
# - __The Relinearization Keys(optional)__. This key is used for controlling the quality of the ciphertexts after encrypted multiplications. Generate it only if you are doing encrypted multiplications. You can safely share it with the Data Scientist.
# - __The Galois Keys(optional)__. This key is needed to perform encrypted vector rotation operations on ciphertexts. Generate it only if you are evaluating convolutions on encrypted data. You can safely share it with the Data Scientist.
# + [markdown] id="QmC6LUOdDOrG"
# ### TenSEAL Context
#
# Now that we had a short introduction, let's get to work.
#
# The first step to do for a Data Owner is to generate a security context containing security parameters and encryption keys.
# + id="KeMSz1tZDOrH" outputId="dce97b03-ece8-4758-ee42-c77a982b9f0d" colab={"base_uri": "https://localhost:8080/"}
context = ts.Context(
ts.SCHEME_TYPE.CKKS,
poly_modulus_degree=8192,
coeff_mod_bit_sizes=[60, 40, 40, 60]
)
context.global_scale = 2**40
context
# + [markdown] id="VqMCywESDOrL"
# ### Encrypt the data
#
# + id="xw_GE4jxDOrM" outputId="6a5ec68c-8166-4f28-e6e3-8bbe1c9dcccd" colab={"base_uri": "https://localhost:8080/"}
v1 = [0, 1, 2, 3, 4]
v2 = [4, 3, 2, 1, 0]
enc_v1 = ts.ckks_vector(context, v1)
enc_v2 = ts.ckks_vector(context, v2)
(enc_v1, enc_v2)
# + [markdown] id="TvVzqehaDOrN"
# ### Make Context and Encrypted Vectors Referenceable over Duet
# + id="JPmSMv5lDOrO"
# tag them so our partner can easily reference it
ctx_ptr = context.send(duet, searchable=True, tags=["context"])
enc_v1_ptr = enc_v1.send(duet, searchable=True, tags=["enc_v1"])
enc_v2_ptr = enc_v2.send(duet, searchable=True, tags=["enc_v2"])
# + id="_EeqtFeUDOrP" outputId="275c6c54-d7ba-45d7-830e-c973bf9d2a6e" colab={"base_uri": "https://localhost:8080/", "height": 143}
# we can see that our three objects are now inside the store we control
duet.store.pandas
# + [markdown] id="SgfSXkwADOrQ"
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 1 : Now STOP and run the Data Scientist notebook until the same checkpoint.
# + id="rGSx98mhDOrR" outputId="ba90fece-baf2-4135-9c77-ef8b39b5d8b0" colab={"base_uri": "https://localhost:8080/", "height": 213}
# We can see our duet partner has requested the two encrypted vectors and the public context
duet.requests.pandas
# + [markdown] id="OaNM37xpDOrS"
# ### Approve the requests
# + id="ByrS-0IODOrS"
duet.requests[0].accept()
duet.requests[0].accept()
duet.requests[0].accept()
# + id="iLAXCwHVDOrU" outputId="f6c43bd9-922d-445b-e601-b76999084c53" colab={"base_uri": "https://localhost:8080/", "height": 32}
# The requests should have been handled
duet.requests.pandas
# + [markdown] id="lQ3QvtyQDOrU"
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 2 : Now STOP and run the Data Scientist notebook until the same checkpoint.
# + [markdown] id="jJWGsOCmDOrU"
# ### Get the computation results from store and decrypt them locally
# + id="205C9PmQDOrV" outputId="a0b46a18-c041-45e6-d696-87b17e5bcd42" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted add
result_add = duet.store["result_add"].get(delete_obj=False)
result_add.link_context(context)
result_add
# + id="qqjb29hoDOrX" outputId="96373593-600c-4543-997f-43713c4a8727" colab={"base_uri": "https://localhost:8080/"}
decrypted_result = result_add.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [v1 + v2 for v1, v2 in zip(v1, v2)]
decrypted_result
# + id="yfTP_cGlDOrZ" outputId="6cf5ef44-b152-45ba-8066-d5da91f888d6" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted - plain add
result_iadd = duet.store["result_iadd"].get(delete_obj=False)
result_iadd.link_context(context)
decrypted_result = result_iadd.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [v1 + v2 for v1, v2 in zip(v1, [10, 10, 10, 10, 10])]
decrypted_result
# + id="5PdeTmkaDOra" outputId="49f23078-94ed-488c-ac99-1ba5dd9dd891" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted subtraction
result_sub = duet.store["result_sub"].get(delete_obj=False)
result_sub.link_context(context)
decrypted_result = result_sub.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [v1 - v2 for v1, v2 in zip(v1, v2)]
decrypted_result
# + id="2trb1jlgDOrb" outputId="4827e4c0-55d5-46f6-9005-2a3744400782" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted multiplication
result_mul = duet.store["result_mul"].get(delete_obj=False)
result_mul.link_context(context)
decrypted_result = result_mul.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [v1 * v2 for v1, v2 in zip(v1, v2)]
decrypted_result
# + id="8TeLnhaaDOrc" outputId="63c30e22-f935-4743-8e54-4ed01245ecba" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted power
result_pow = duet.store["result_pow"].get(delete_obj=False)
result_pow.link_context(context)
decrypted_result = result_pow.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [v ** 3 for v in v1]
decrypted_result
# + id="J2rLNJVGDOrc" outputId="9baef6d9-b0d1-42ab-94f3-2447ae672437" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted negation
result_neg = duet.store["result_neg"].get(delete_obj=False)
result_neg.link_context(context)
decrypted_result = result_neg.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [-v for v in v1]
decrypted_result
# + id="JY3VukJDDOrd" outputId="800f6090-10dd-4333-8f82-9aaaf4151449" colab={"base_uri": "https://localhost:8080/"}
# Validate the encrypted polynomial evaluation for 1 + X^2 + X^3
result_poly = duet.store["result_poly"].get(delete_obj=False)
result_poly.link_context(context)
decrypted_result = result_poly.decrypt()
assert pytest.approx(decrypted_result, abs=10**-3) == [1 + v**2 + v**3 for v in v1]
decrypted_result
# + [markdown] id="r7BNH_OlDOrd"
# ### <img src="https://github.com/OpenMined/design-assets/raw/master/logos/OM/mark-primary-light.png" alt="he-black-box" width="100"/> Checkpoint 3 : Well done!
# + [markdown] id="jFDfj1FfDOre"
# # Congratulations!!! - Time to Join the Community!
#
# Congratulations on completing this notebook tutorial! If you enjoyed this and would like to join the movement toward privacy preserving, decentralized ownership of AI and the AI supply chain (data), you can do so in the following ways!
#
# ### Star PySyft and TenSEAL on GitHub
#
# The easiest way to help our community is just by starring the Repos! This helps raise awareness of the cool tools we're building.
#
# - [Star PySyft](https://github.com/OpenMined/PySyft)
# - [Star TenSEAL](https://github.com/OpenMined/TenSEAL)
#
# ### Join our Slack!
#
# The best way to keep up to date on the latest advancements is to join our community! You can do so by filling out the form at [http://slack.openmined.org](http://slack.openmined.org). #lib_tenseal and #code_tenseal are the main channels for the TenSEAL project.
#
# ### Donate
#
# If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
#
# [OpenMined's Open Collective Page](https://opencollective.com/openmined)
| examples/homomorphic-encryption/Tutorial_0_TenSEAL_Syft_Data_Owner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hubble Source Catalog API Notebook: SMC Color-Magnitude Diagram
# ### August 2019, <NAME>
#
# A [new MAST interface](https://catalogs.mast.stsci.edu/hsc) supports queries to the current and previous versions of the [Hubble Source Catalog](https://archive.stsci.edu/hst/hsc). It allows searches of the summary table (with multi-filter mean photometry) and the detailed table (with all the multi-epoch measurements). It also has an associated [API](https://catalogs.mast.stsci.edu/docs/hsc.html), which is used in this notebook.
#
# This is based on part of [HSC Use Case #2](https://archive.stsci.edu/hst/hsc/help/use_case_2_v3.html).
# * It searches the HSC for point-like objects in the Small Magellanic Cloud (SMC) with ACS/WFC V and I band measurements,
# * selects a subset of those objects in a V-I color range,
# * plots the positions of the objects on the sky, and
# * plots the color-magnitude diagram for the selected objects.
#
# The whole process takes only about 2 minutes to complete.
#
# This notebook is available for [download](hscv3_smc_api.ipynb). Another [simple notebook](hscv3_api.ipynb) demonstrates other search capabilities of the API to find variable objects and plot their light curves. A more complex notebook that shows how to access the proper motion tables using the HSC API is also [available](sweeps_hscv3p1_api.html).
# # Instructions:
# * Complete the initialization steps [described below](#Initialization).
# * Run the notebook.
#
# Running the notebook from top to bottom takes about 2 minutes.
#
# # Table of Contents
# * [Initialization](#Initialization)
# * [Find variable objects the SMC](#smc)
# * [Use MAST name resolver](#resolver)
# * [Search HSC summary table](#summary)
# * [Show object positions on the sky](#positions)
# * [Plot the color-magnitude diagram](#cmd)
# # Initialization <a class="anchor" id="Initialization"></a>
# ### Install Python modules
#
# _This notebook requires the use of **Python 3**._
#
# This needs the `requests` and `fastkde` modules in addition to the common requirements of `astropy`, `numpy` and `scipy`. For anaconda versions of Python the installation commands are:
#
# <pre>
# conda install requests
# pip install fastkde
# </pre>
# +
# %matplotlib inline
import astropy, pylab, time, sys, os, requests, json
import numpy as np
from astropy.table import Table
from astropy.io import ascii
from fastkde import fastKDE
from scipy.interpolate import RectBivariateSpline
from astropy.modeling import models, fitting
# Set page width to fill browser for longer output lines
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# set width for pprint
astropy.conf.max_width = 150
# -
# ## Useful functions
#
# Execute HSC searches and resolve names using [MAST query](https://mast.stsci.edu/api/v0/MastApiTutorial.html).
# +
hscapiurl = "https://catalogs.mast.stsci.edu/api/v0.1/hsc"
def hsccone(ra,dec,radius,table="summary",release="v3",format="csv",magtype="magaper2",
columns=None, baseurl=hscapiurl, verbose=False,
**kw):
"""Do a cone search of the HSC catalog
Parameters
----------
ra (float): (degrees) J2000 Right Ascension
dec (float): (degrees) J2000 Declination
radius (float): (degrees) Search radius (<= 0.5 degrees)
table (string): summary, detailed, propermotions, or sourcepositions
release (string): v3 or v2
magtype (string): magaper2 or magauto (only applies to summary table)
format: csv, votable, json
columns: list of column names to include (None means use defaults)
baseurl: base URL for the request
verbose: print info about request
**kw: other parameters (e.g., 'numimages.gte':2)
"""
data = kw.copy()
data['ra'] = ra
data['dec'] = dec
data['radius'] = radius
return hscsearch(table=table,release=release,format=format,magtype=magtype,
columns=columns,baseurl=baseurl,verbose=verbose,**data)
def hscsearch(table="summary",release="v3",magtype="magaper2",format="csv",
columns=None, baseurl=hscapiurl, verbose=False,
**kw):
"""Do a general search of the HSC catalog (possibly without ra/dec/radius)
Parameters
----------
table (string): summary, detailed, propermotions, or sourcepositions
release (string): v3 or v2
magtype (string): magaper2 or magauto (only applies to summary table)
format: csv, votable, json
columns: list of column names to include (None means use defaults)
baseurl: base URL for the request
verbose: print info about request
**kw: other parameters (e.g., 'numimages.gte':2). Note this is required!
"""
data = kw.copy()
if not data:
raise ValueError("You must specify some parameters for search")
if format not in ("csv","votable","json"):
raise ValueError("Bad value for format")
url = "{}.{}".format(cat2url(table,release,magtype,baseurl=baseurl),format)
if columns:
# check that column values are legal
# create a dictionary to speed this up
dcols = {}
for col in hscmetadata(table,release,magtype)['name']:
dcols[col.lower()] = 1
badcols = []
for col in columns:
if col.lower().strip() not in dcols:
badcols.append(col)
if badcols:
raise ValueError('Some columns not found in table: {}'.format(', '.join(badcols)))
# two different ways to specify a list of column values in the API
# data['columns'] = columns
data['columns'] = '[{}]'.format(','.join(columns))
# either get or post works
# r = requests.post(url, data=data)
r = requests.get(url, params=data)
if verbose:
print(r.url)
r.raise_for_status()
if format == "json":
return r.json()
else:
return r.text
def hscmetadata(table="summary",release="v3",magtype="magaper2",baseurl=hscapiurl):
"""Return metadata for the specified catalog and table
Parameters
----------
table (string): summary, detailed, propermotions, or sourcepositions
release (string): v3 or v2
magtype (string): magaper2 or magauto (only applies to summary table)
baseurl: base URL for the request
Returns an astropy table with columns name, type, description
"""
url = "{}/metadata".format(cat2url(table,release,magtype,baseurl=baseurl))
r = requests.get(url)
r.raise_for_status()
v = r.json()
# convert to astropy table
tab = Table(rows=[(x['name'],x['type'],x['description']) for x in v],
names=('name','type','description'))
return tab
def cat2url(table="summary",release="v3",magtype="magaper2",baseurl=hscapiurl):
"""Return URL for the specified catalog and table
Parameters
----------
table (string): summary, detailed, propermotions, or sourcepositions
release (string): v3 or v2
magtype (string): magaper2 or magauto (only applies to summary table)
baseurl: base URL for the request
Returns a string with the base URL for this request
"""
checklegal(table,release,magtype)
if table == "summary":
url = "{baseurl}/{release}/{table}/{magtype}".format(**locals())
else:
url = "{baseurl}/{release}/{table}".format(**locals())
return url
def checklegal(table,release,magtype):
"""Checks if this combination of table, release and magtype is acceptable
Raises a ValueError exception if there is problem
"""
releaselist = ("v2", "v3")
if release not in releaselist:
raise ValueError("Bad value for release (must be one of {})".format(
', '.join(releaselist)))
if release=="v2":
tablelist = ("summary", "detailed")
else:
tablelist = ("summary", "detailed", "propermotions", "sourcepositions")
if table not in tablelist:
raise ValueError("Bad value for table (for {} must be one of {})".format(
release, ", ".join(tablelist)))
if table == "summary":
magtypelist = ("magaper2", "magauto")
if magtype not in magtypelist:
raise ValueError("Bad value for magtype (must be one of {})".format(
", ".join(magtypelist)))
def mastQuery(request, url='https://mast.stsci.edu/api/v0/invoke'):
"""Perform a MAST query.
Parameters
----------
request (dictionary): The MAST request json object
url (string): The service URL
Returns the returned data content
"""
# Encoding the request as a json string
requestString = json.dumps(request)
r = requests.post(url, data={'request': requestString})
r.raise_for_status()
return r.text
def resolve(name):
"""Get the RA and Dec for an object using the MAST name resolver
Parameters
----------
name (str): Name of object
Returns RA, Dec tuple with position
"""
resolverRequest = {'service':'Mast.Name.Lookup',
'params':{'input':name,
'format':'json'
},
}
resolvedObjectString = mastQuery(resolverRequest)
resolvedObject = json.loads(resolvedObjectString)
# The resolver returns a variety of information about the resolved object,
# however for our purposes all we need are the RA and Dec
try:
objRa = resolvedObject['resolvedCoordinate'][0]['ra']
objDec = resolvedObject['resolvedCoordinate'][0]['decl']
except IndexError as e:
raise ValueError("Unknown object '{}'".format(name))
return (objRa, objDec)
# -
# ## Find objects in the SMC <a name="smc"></a>
#
# This is based on [HSC Use Case #2](https://archive.stsci.edu/hst/hsc/help/use_case_2_v3.html), which includes an example of creating a color-magnitude diagram for the SMC using <a href="https://mastweb.stsci.edu/hcasjobs">MAST CasJobs</a>. This is simple to do using the HSC API.
# ### Use MAST name resolver to get position of the SMC <a name="resolver"></a>
target = 'SMC'
ra, dec = resolve(target)
print(target,ra,dec)
# ### Select objects with the desired magnitudes and colors near the SMC <a name="summary"></a>
#
# This searches the summary table for objects in a 3x3 degree box centered on the galaxy that have measurements in both ACS F555W and F814W. It computes the V-I color and selects only objects in the range -1.5 < V-I < 1.5. This large query ultimately returns more than 700,000 objects and takes about a minute to complete.
# +
# save typing a quoted list of columns
columns = """MatchID,MatchRA,MatchDec,CI,A_F555W,A_F814W""".split(",")
columns = [x.strip() for x in columns]
columns = [x for x in columns if x and not x.startswith('#')]
# select objects with at least one ACS F555W and ACS F814W measurement
# and with concentration index 0.9 < CI < 1.6, consistent with point sources
# search a large 3x3 degree box in RA and Dec centered on the SMC
ddec = 1.5
dra = ddec/np.cos(np.radians(dec))
constraints = {'A_F555W_N.gte': 1, 'A_F814W_N.gte': 1, 'CI.gt':0.9, 'CI.lt':1.6,
'MatchDec.gt': dec-ddec, 'MatchDec.lt': dec+ddec,
'MatchRA.gt': ra-dra, 'MatchRA.lt': ra+dra}
# do a search with a large number of rows allowed
t0 = time.time()
tab = ascii.read(hscsearch(table="summary",release='v3',
columns=columns,verbose=True,pagesize=2000000,**constraints))
print("{:.1f} s: retrieved data and converted to {}-row astropy table".format(time.time()-t0, len(tab)))
# compute color column and select for objects in more limited color range
tab['V-I'] = tab['A_F555W'] - tab['A_F814W']
tab = tab[(tab['V-I'] < 1.5) & (tab['V-I'] > -1.5)]
print("{:.1f} s: selected {} objects with -1.5 < V-I < 1.5".format(time.time()-t0, len(tab)))
# clean up the output format
tab['A_F555W'].format = "{:.3f}"
tab['A_F814W'].format = "{:.3f}"
tab['V-I'].format = "{:.3f}"
tab['CI'].format = "{:.3f}"
tab['MatchRA'].format = "{:.6f}"
tab['MatchDec'].format = "{:.6f}"
tab
# -
# ### Plot object positions on the sky <a name="positions"></a>
#
# We mark the galaxy center as well. These fields are sprinkled all over the galaxy (as determined by the HST proposals).
pylab.rcParams.update({'font.size': 16})
pylab.figure(1,(10,10))
pylab.plot(tab['MatchRA'], tab['MatchDec'], 'bo', markersize=1,
label='{} HSC measurements'.format(len(tab)))
pylab.plot(ra,dec,'rx',label=target,markersize=10)
pylab.gca().invert_xaxis()
pylab.gca().set_aspect(1.0/np.cos(np.radians(dec)))
pylab.xlabel('RA [deg]')
pylab.ylabel('Dec [deg]')
pylab.legend(loc='best')
# ### Plot the color-magnitude diagram <a name="cmd"></a>
#
# This uses the `fastkde` module to get a kernel density estimate in order to plot a dense scatterplot.
# +
# Calculate the point density
t0 = time.time()
x = tab['V-I']
y = tab['A_F555W']
myPDF,axes = fastKDE.pdf(x,y,numPoints=2**9+1)
print("kde took {:.1f} sec".format(time.time()-t0))
# interpolate to get z values at points
finterp = RectBivariateSpline(axes[1],axes[0],myPDF)
z = finterp(y,x,grid=False)
# Sort the points by density, so that the densest points are plotted last
idx = z.argsort()
xs, ys, zs = x[idx], y[idx], z[idx]
# select a random subset of points in the most crowded regions to speed up plotting
wran = np.where(np.random.random(len(zs))*zs<0.05)[0]
print("Plotting {} of {} points".format(len(wran),len(zs)))
xs = xs[wran]
ys = ys[wran]
zs = zs[wran]
pylab.rcParams.update({'font.size': 16})
pylab.figure(1,(12,10))
pylab.scatter(xs, ys, c=zs, s=2, edgecolor='', cmap='plasma')
pylab.ylabel('V [mag]')
pylab.xlabel('V - I [mag]')
pylab.xlim(-1.5,1.5)
pylab.ylim(14,27)
pylab.gca().invert_yaxis()
pylab.title('{:,} stars in the Small Magellanic Cloud'.format(len(tab)))
pylab.colorbar()
pylab.tight_layout()
pylab.savefig("smc_cmd.png")
# -
| hscv3_smc_api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
pd.__version__ # 0.24.2
# - Getting Started: http://pandas.pydata.org/pandas-docs/stable/getting_started/index.html
# - 10 Minutes to pandas: http://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html
# - Pandas Cheat Sheet: http://pandas.pydata.org/Pandas_Cheat_Sheet.pdf
# # Design
#
# http://pandas.pydata.org/pandas-docs/stable/getting_started/overview.html
# ## Primary Data Structures
# | Dimensions | Name | Description |
# | ---------- | ---- | ----------- |
# | 1 | Series | **homogeneously**-typed array |
# | 2 | DataFrame | size-mutable tabular structure with potentially **heterogeneously**-typed column |
#
# - Pandas objects (Index, Series, DataFrame) can be thought of as **containers for arrays**, which hold the actual data and do the actual computation. For many types, **the underlying array is a numpy.ndarray**
# - **DataFrame is a container for Series, and Series is a container for scalars**
# - insert and remove objects from these containers in a dictionary-like fashion
# - the axes are intended to lend more semantic meaning to the data
#
# ```python
# for col in df.columns:
# series = df[col]
# # do something with series
# ```
# ## Mutability and copying of data
#
# - All pandas data structures are **value-mutable** (the values they contain can be altered) but **not always size-mutable**.The length of a Series cannot be changed, but, for example, columns can be inserted into a DataFrame.
#
# - However, the vast majority of methods produce new objects and leave the input data untouched. In general we like to **favor immutability where sensible**.
# ## Index
#
# https://pandas.pydata.org/pandas-docs/version/0.24.2/reference/indexing.html#index
#
# `pd.Index`: Immutable ndarray implementing an ordered, sliceable set.
display(pd.Index([1,2,3]))
display(pd.Index([1,3,2]))
display(pd.Index([1,2,3])[:2])
display(pd.Index([1,2,'a']))
display(pd.Index(['a','b','c']))
display(pd.Index(['a','b','b']))
display(pd.DatetimeIndex(['2000-01','2000-03','2001-01']))
arrays = [[1, 1, 2, 2], ['red', 'blue', 'red', 'blue']]
display(pd.MultiIndex.from_arrays(arrays, names=('number', 'color')))
# ## Series
#
# https://pandas.pydata.org/pandas-docs/stable/reference/series.html
#
# index + single column data
sr = pd.Series(data=[1,2], index=['row_1', 'row_2'])
display(sr)
display(type(sr))
# ## DataFrame
#
# https://pandas.pydata.org/pandas-docs/stable/reference/frame.html
#
# index + column + tabular data
# +
df = pd.DataFrame(data=[1,2], index=['row_1', 'row_2'], columns=['col_1'])
display(df)
df = pd.DataFrame(data=['foo',2], index=[('a',1), ('b',2)], columns=['col_1'])
display(df)
df = pd.DataFrame(data=[[1,2],[2,2]], index=[('a',1), ('b',2)], columns=['col_1', 'col_2'])
display(df)
# -
# ### Conversion: Dict
# +
# dict to dataframe
df = pd.DataFrame({'col_1': [10, 'aa', (1,'e'), 30, 45],
'col_2': [13, 'cc', (3,'f'), 33, 48],
'col_3': [17, 'dd', (5,'g'), 37, 52]})
display(df)
data = {'col_1': [3, 2, 1, 0], 'col_2': ['a', 'b', 'c', 'd']}
df = pd.DataFrame.from_dict(data, orient='columns')
display(df)
data = {'row_1': [3, 2, 1, 0], 'row_2': ['a', 'b', 'c', 'd']}
df = pd.DataFrame.from_dict(data, orient='index')
display(df)
# +
# dataframe to dict
df = pd.DataFrame({'col_1': [10, 'aa', (1,'e'), 30, 45],
'col_2': [13, 'cc', (3,'f'), 33, 48],
'col_3': [17, 'dd', (5,'g'), 37, 52]})
display(df)
print('----------\norient=dict:')
display( df.to_dict(orient='dict')) # default
print('orient=list:')
display(df.to_dict(orient='list'))
print('----------\norient=records:')
display(df.to_dict(orient='records'))
print('orient=index:')
display(df.to_dict(orient='index'))
print('----------\norient=split:')
display(df.to_dict(orient='split'))
print('----------\norient=series:')
display(df.to_dict(orient='series'))
# -
# ### Conversion: Numpy Array
# array to dataframe
display(pd.DataFrame(np.array([3,2,1,0])))
display(pd.DataFrame(np.array([[3,2],[1,0]])))
# dataframe to array
df = pd.DataFrame({'col_1': [10, 'aa', (1,'e'), 30, 45],
'col_2': [13, 'cc', (3,'f'), 33, 48],
'col_3': [17, 'dd', (5,'g'), 37, 52]})
display(df)
display(df.to_numpy()) # Depreciated: .values / as_matrix()
display(df.to_numpy().T)
display(df.T.to_numpy())
# ### View
# +
# reset_index
df = pd.DataFrame(data=[1,2], index=[('a',1), ('b',2)], columns=['col_1'])
display(df)
display(df.reset_index())
display(df.reset_index(drop=True))
# -
# rename column
display(df.rename(index=str, columns={"col_1": "x", "col_2": "y"}))
# Transpose
display(df.T)
# # File IO
# ## Input
#
# - `pd.read_json`
# - `pd.read_excel`
# - `pd.read_csv`
# - `pd.read_pickle` # for pandas objects or other objects such as python dict
#
# More: https://pandas.pydata.org/pandas-docs/stable/reference/io.html
# ## Output
#
# - `df.to_csv`
# - `df.to_dict(orient=)`
# - `df.to_excel`
# - `df.to_pickle`
#
# More: https://pandas.pydata.org/pandas-docs/stable/reference/frame.html#serialization-io-conversion
# ### Excel with Style
#
# StyleFrame: https://styleframe.readthedocs.io/en/0.2/#
# + deletable=false editable=false run_control={"frozen": true}
# # Excel with style
# from StyleFrame import StyleFrame, Styler
#
# sf = StyleFrame.read_excel('xxx.xlsx', sheet_name='Sheet1') #, read_style=True / StyleFrame support only .xlsx
#
# font_blue = Styler(font_color='blue')
# font_red = Styler(font_color='red')
#
# for col_name in sf.columns:
# sf.apply_style_by_indexes(indexes_to_style=sf[sf[col_name].isin(some_list)], # decide rows
# styler_obj=font_blue,
# cols_to_style=col_name) # decide cols
# sf.apply_style_by_indexes(indexes_to_style=sf[sf[col_name].isin(another_list)],
# styler_obj=font_red,
# cols_to_style=col_name)
#
# sf.to_excel('xxx_styled.xlsx').save()
# -
# # Indexing / Selecting / Slicing
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html
#
# index的label和integer position
# ## [ ]
#
# - series: select row
# - dataframe: select column
sr = pd.Series(data=['aa','bb','cc', 'dd'], index=['a',1,2,3])
display(sr)
display(sr[pd.Index([1,3,2])])
display(sr[[1,3,2]])
display(sr[[1,'a',2]])
df = pd.DataFrame(data=[[1,2],[3,4]], index=['row_1', 'row_2'], columns=['col_1','col_2'])
display(df)
display(df[['col_1']]) # column
# df[['row_1', 'row_2']] Error
# ## loc / iloc
# ### single index
sr = pd.Series(data=['aa','bb','cc', 'dd'], index=['a',1,2,3])
display(sr)
df = pd.DataFrame(data=[1,2], index=['row_1', 'row_2'], columns=['col_1'])
display(df)
display(df.loc[['row_1', 'row_2']])
try:
df.loc[('row_1', 'row_2')] # for multiindex: https://pandas.pydata.org/pandas-docs/stable/user_guide/advanced.html
except Exception as e:
print(type(e), e)
# ### tuple index
df = pd.DataFrame(data=[1,2], index=[('a',1), ('b',2)], columns=['col_1'])
display(df)
# + deletable=false editable=false run_control={"frozen": true}
# df.loc[('a',1)] # KeyError: 'a'
# df.loc[('a',1),] # KeyError: "None of [('a', 1)] are in the [index]"
# -
display(df.loc[[('a',1)],])
display(df.loc[[('a',1),],])
display(df.loc[[('a',1), ('b',2)],])
display(df.loc[[('a',1),], 'col_1'])
# ### hierarchical index (MultiIndex)
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/advanced.html
arrays = [np.array(['bar', 'bar', 'baz', 'baz', 'foo', 'foo', 'qux', 'qux']),
np.array(['one', 'two', 'one', 'two', 'one', 'two', 'one', 'two'])]
df = pd.DataFrame(np.random.randn(8, 4), index=arrays)
display(df)
display(df.loc[('baz', 'two')]) # series
display(df.loc[[('baz', 'two')]]) # dataframe
display(df.loc[[('baz', 'two'), ('foo', 'one')]]) # dataframe
# ### iloc
# +
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
display(df.iloc[0]) # series
display(df.iloc[-1])
display(df.iloc[0:2]) # dataframe
display(df.iloc[1:])
display(df.iloc[-1:])
display(df.iloc[::2])
# -
# ## Boolean Filtering
df = pd.DataFrame(data=[[1,2],[2,1]], index=[('a',1), ('b',2)], columns=['col_1', 'col_2'])
display(df)
display(type(df['col_1'] == 1))
display(df['col_1'] == 1)
display(df[df['col_1'] == 1])
# +
# combine
bool_1 = df['col_1'] == 2
bool_2 = df['col_2'] == 1
display(bool_1)
display(bool_2)
display(bool_1 & bool_2)
display(df[bool_1 & bool_2])
# +
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
display(df[(df.AAA <= 6) & (df.index.isin([0, 2, 4]))])
display(df.loc[(df['BBB'] > 25) | (df['CCC'] >= -40), 'AAA'])
display(df)
df.loc[(df['BBB'] > 25) | (df['CCC'] >= -40), 'AAA'] = 1
display(df)
# +
# query: write filtering rules
display(df.query('BBB > CCC'))
display(df[df.BBB > df.CCC]) # same result as the previous expression
# -
# ## Sampling
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
display(df.sample(n=2))
display(df.sample(n=2, axis=1))
# # Merge / Join / Concat
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/merging.html
#
# - `.add()`
# - `.merge()`
# - `.join()`
# # Math
#
# https://pandas.pydata.org/pandas-docs/stable/reference/frame.html#computations-descriptive-stats
#
# ## Unary Operation
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
# ### Series
display(sum(df['AAA'])) # == df['AAA'].sum()
display(max(df['AAA'])) # == df['AAA'].max()
display(df['AAA'].mean())
display(df['AAA'].value_counts()) # series
# ### DataFrame
display(df.sum()) # series
display(df * 2) # dataframe
display(df.describe()) # dataframe
# ## Binary Operation
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
# ### Series
# +
display(df['AAA'] + df['BBB'])
display(df['AAA'] * df['BBB'])
import scipy.spatial.distance as distance
display(1 - distance.cosine(df['AAA'], df['BBB']))
# -
# ### DataFrame
display(df + df)
display(df / df)
display(df.T @ df) # Matrix dot product
# # Iteration
#
#
# do not have to convert to list/dict, Pandas can do iteration - performance
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
# ## Along axis
#
# - `.iteritems()`
# - `.iterrows()`
# - `.itertuples()`
for col in df:
print(col)
for columns, series in df.iteritems():
print(columns)
print(series)
# +
for index, series in df.iterrows():
print(index)
print(series)
for row in df.itertuples():
print(row)
# -
# ## Apply / Map
# https://stackoverflow.com/questions/19798153/difference-between-map-applymap-and-apply-methods-in-pandas
#
# - `map` works element-wise on a Series.
# - `applymap` works element-wise on a DataFrame
# - `apply` works on a row / column basis of a DataFrame (`df.apply()`), also works on series(`sr.apply()`)
# - If func returns a scalar or a list, `apply(func)` returns a Series.
# - If func returns a Series, `apply(func)` returns a DataFrame.
#
#
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.map.html
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.applymap.html
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.apply.html
# - https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.apply.html
#
# +
import math
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
display(df['AAA'].apply(math.sqrt))
display(df.apply(np.sqrt, axis=0))
# -
# apply + lambda
display(df)
display(df['AAA'].apply(lambda x: pd.Series([x] * 5)))
display(df.apply(lambda x: x.loc[0] + 1, axis=0)) # index (row), axis=0
display(df.apply(lambda x: x, axis=1)) # column, axis=1
display(df.apply(lambda x: x.loc['AAA'] + 1, axis=1)) # column
display(df.apply(lambda x: x.loc['AAA'] + x.loc['BBB'], axis=1)) # multi-columns, same as: df['AAA'] + df['BBB']
display(df.apply(lambda x: max([x['BBB'], x['CCC']]), axis=1))
# +
# Normalize
df = pd.DataFrame({'AAA': [4, 5, 6, 7],
'BBB': [10, 20, 30, 40],
'CCC': [100, 50, -30, -50]})
display(df)
df['CCC'] = df['CCC'].apply(lambda x: x/max(df['CCC']))
display(df)
# -
# ### Apply with Progress Bar
# +
from tqdm import tqdm
tqdm.pandas()
df.progress_apply(lambda x: max([x['BBB'], x['CCC']]), axis=1)
# +
from tqdm.autonotebook import tqdm
tqdm.pandas()
df.progress_apply(lambda x: max([x['BBB'], x['CCC']]), axis=1)
# -
# ## Group
#
# 1. Splitting the data into groups based on some criteria
# 2. Applying a function to each group independently
# 3. Combining the results into a data structure
#
#
# - https://pandas.pydata.org/pandas-docs/stable/reference/groupby.html
# - https://pandas.pydata.org/pandas-docs/stable/user_guide/groupby.html
# ### Structure
# +
df = pd.DataFrame({'AAA': [1, 2, 8, 2],
'BBB': [1, 20, 30, 40],
'CCC': [0, 50, -30, -50]})
display(df)
grouped = df.groupby('AAA')
print(type(grouped))
print(grouped)
print('------------')
print(grouped.groups)
print(grouped.groups[8]) # indexing with group's name (dict-like)
# -
# ### Iteration
#
for name, data in grouped: # iterated as tuple(name, data)
print(name)
print(type(data), data)
# ### Grouped by Time Period
# +
dates = pd.date_range('1/10/2000', periods=60)
df = pd.DataFrame(np.random.randn(60, 4),
index=dates, columns=['A', 'B', 'C', 'D'])
display(df)
# -
for name, data in df.groupby(pd.Grouper(freq='M')): # or '1M'
print('\n', name)
print(data)
for name, data in df.groupby(pd.Grouper(freq='30d')):
print('\n', name)
print(data)
# ### Group + Apply
# +
dates = pd.date_range('1/10/2000', periods=60)
df = pd.DataFrame(np.random.randn(60, 4),
index=dates, columns=['A', 'B', 'C', 'D'])
grouped = df.groupby(pd.Grouper(freq='1M'))
sr = grouped.apply(lambda x: sum(x['B']))
display(sr)
# -
sr.index = sr.index.map(lambda x: str(x)[:7])
display(sr)
# # Missing Data
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html
#
# - `pd.NaT`
# - `np.nan`
# - `.isna()` / `notna()`
# - `.fillna()`
# # Performance
#
# http://pandas.pydata.org/pandas-docs/stable/user_guide/enhancingperf.html
#
# pandas is fast. Many of the low-level algorithmic bits have been extensively tweaked in Cython code. However, as with anything else generalization usually sacrifices performance. So if you focus on one feature for your application you may be able to create a faster specialized tool.
# ## Time
# ### Dependencies
#
# https://pandas.pydata.org/pandas-docs/stable/install.html#recommended-dependencies
#
# - numexpr
# - bottleneck
#
# ### Use Wisely
#
# https://engineering.upside.com/a-beginners-guide-to-optimizing-pandas-code-for-speed-c09ef2c6a4d6
#
# - Avoid loops; they’re slow and, in most common use cases, unnecessary.
# - If you must loop, use `.apply()`, not iteration functions.
# - Vectorization is usually better than scalar operations. Most common operations in Pandas can be vectorized.
# - **Vector operations on NumPy arrays are more efficient than on native Pandas series.**
#
# https://realpython.com/fast-flexible-pandas/
#
# 1. **Use vectorized operations: Pandas methods and functions with no for-loops.**
# 2. **Use the `.apply()` method with a callable.**
# 3. **Use `.itertuples()`: iterate over DataFrame rows as namedtuples from Python’s collections module.**
# 4. **Use `.iterrows()`: iterate over DataFrame rows as (index, pd.Series) pairs.** While a Pandas Series is a flexible data structure, it can be costly to construct each row into a Series and then access it.
# 5. Use “element-by-element” for loops, updating each cell or row one at a time with `df.loc` or `df.iloc`. (Or, `.at`/`.iat` for fast scalar access.)
#
# ---
#
# 1. Try to use vectorized operations where possible rather than approaching problems with the `for x in df`... mentality. If your code is home to a lot of for-loops, it might be better suited to working with native Python data structures, because Pandas otherwise comes with a lot of overhead.
# 2. If you have more complex operations where vectorization is simply impossible or too difficult to work out efficiently, use the `.apply()` method.
# 3. If you do have to loop over your array (which does happen), use `.iterrows()` or `.itertuples()` to improve speed and syntax.
# 4. **Pandas has a lot of optionality**, and there are almost always several ways to get from A to B. Be mindful of this, compare how different routes perform, and choose the one that works best in the context of your project.
# 5. Once you’ve got a data cleaning script built, avoid reprocessing by storing your intermediate results with HDFStore.
# 6. Integrating NumPy into Pandas operations can often improve speed and simplify syntax.
#
# https://www.dataquest.io/blog/pandas-big-data/
#
# 1. **Downcasting numeric columns to more efficient types**.
# 2. **Converting string columns to the categorical type**.
# ### Parallelize
#
# http://blog.adeel.io/2016/11/06/parallelize-pandas-map-or-apply/
# ### Cython / Numba / pandas.eval()
#
# https://pandas.pydata.org/pandas-docs/stable/user_guide/enhancingperf.html
# ## Space
#
# https://towardsdatascience.com/why-and-how-to-use-pandas-with-large-data-9594dda2ea4c
#
# 1. chunking
# 2. drop useless columns
| 02_Pandas/pandas_data_tips.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="u2UXutvEvpUj"
# # Question Answering with BERT and HuggingFace 🤗 (Fine-tuning)
#
# In the previous Hugging Face ungraded lab, you saw how to use the pipeline objects to use transformer models for NLP tasks. I showed you that the model didn't output the desired answers to a series of precise questions for a context related to the history of comic books.
#
# In this lab, you will fine-tune the model from that lab to give better answers for that type of context. To do that, you'll be using the [TyDi QA dataset](https://ai.google.com/research/tydiqa) but on a filtered version with only English examples. Additionally, you will use a lot of the tools that Hugging Face has to offer.
#
# You have to note that, in general, you will fine-tune general-purpose transformer models to work for specific tasks. However, fine-tuning a general-purpose model can take a lot of time. That's why you will be using the model from the question answering pipeline in this lab.
#
# First, let's install some packages that you will use during the lab.
# + id="7rW5HyNyv3YC" outputId="2a06419a-8890-47fc-80f2-9c071a1b2015" colab={"base_uri": "https://localhost:8080/"}
# !pip install transformers datasets torch;
# + [markdown] id="FrEglXPmvpUr"
# ## Fine-tuning a BERT model
#
# As you saw in the previous lab, you can use these pipelines as they are. But sometimes, you'll need something more specific to your problem, or maybe you need it to perform better on your production data. In these cases, you'll need to fine-tune a model.
#
# Here, you'll fine-tune a pre-trained DistilBERT model on the TyDi QA dataset.
#
# To fine-tune your model, you will leverage three components provided by Hugging Face:
#
# * Datasets: Library that contains some datasets and different metrics to evaluate the performance of your models.
# * Tokenizer: Object in charge of preprocessing your text to be given as input for the transformer models.
# * Transformers: Library with the pre-trained model checkpoints and the trainer object.
#
#
# + [markdown] id="g0Rg-e4jBFFs"
# ### Datasets
#
# To get the dataset to fine-tune your model, you will use [🤗 Datasets](https://huggingface.co/docs/datasets/), a lightweight and extensible library to share and access datasets and evaluation metrics for NLP easily. You can download Hugging Face datasets directly using the `load_dataset` function from the `datasets` library. Although the most common approach is to use `load_dataset`, for this lab you will use a filtered version containing only the English examples. You can read them from a public GCP bucket and use the `load_from_disk` function.
#
# Hugging Face `datasets` allows to load data in several formats, such as CSV, JSON, text files and even parquet. You can see more about the supported formats in the [documentation](https://huggingface.co/docs/datasets/loading.html)
#
# We already prepared the dataset for you, so you don't need to uncomment the code from the cell below if you don't want to load all the data and then filter the English examples. If you want to download the dataset by yourself, you can uncomment the following cell and then jump to the [cell](#datasets_type) in which you can see the type of object you get after loading the dataset.
# + id="NKRe_dD0J4E0"
# You can download the dataset and process it to obtain the same dataset we are loading from disk
# Uncomment the following lines to download the dataset directly
# from datasets import load_dataset
# train_data = load_dataset('tydiqa', 'primary_task')
# tydiqa_data = train_data.filter(lambda example: example['language'] == 'english')
# + [markdown] id="oS-ZAEylgg5e"
# If you want to use the dataset provided by us, please run the following cells. First, we will download the dataset from the GCP bucket.
# + id="3ceaiHQyPFv_" outputId="226274fc-96bf-485b-bf67-0bd8862b0056" colab={"base_uri": "https://localhost:8080/"}
# Download dataset from bucket.
# !wget https://storage.googleapis.com/nlprefresh-public/tydiqa_data.zip
# + id="NtfXWozHZ6oZ"
# Uncomment if you want to check the size of the file. It should be around 319M.
# #!ls -alh tydiqa_data.zip
# + [markdown] id="t0f3Q5V0lZh5"
# Now, let's unzip the dataset
# + id="mfB5xT-DZjbn" outputId="24c690f8-89ed-4ba9-dee9-53f5e60595ad" colab={"base_uri": "https://localhost:8080/"}
# Unzip inside the dataset folder
# !unzip tydiqa_data
# + [markdown] id="ufqLptl1S_a6"
# Given that we used Apache Arrow format to save the dataset, you have to use the `load_from_disk` function from the `datasets` library to load it. To access the preprocessed dataset we created, you should execute the following commands.
# + id="x68dqaoXg5Ra" outputId="b5b8ecaa-9546-4e07-d3b6-c3368c6fea4a" colab={"base_uri": "https://localhost:8080/"}
# Execute this cell if you will use the data we processed instead of downloading it.
from datasets import load_from_disk
#The path where the dataset is stored
path = '/content/tydiqa_data/'
#Load Dataset
tydiqa_data = load_from_disk(path)
tydiqa_data
# + [markdown] id="1hfzBZU3T47O"
# <a id='datasets_type'></a>
# You can check below that the type of the loaded dataset is a `datasets.arrow_dataset.Dataset`. This object type corresponds to an Apache Arrow Table that allows creating a hash table that contains the position in memory where data is stored instead of loading the complete dataset into memory. But you don't have to worry too much about that. It is just an efficient way to work with lots of data.
# + id="gkeppC3GQiW6" outputId="3e3ace6a-43db-4c1c-99d8-a2e7e7aa0de9" colab={"base_uri": "https://localhost:8080/"}
# Checking the object type for one of the elements in the dataset
type(tydiqa_data['train'])
# + [markdown] id="q_HLaNtQaFlR"
# You can also check the structure of the dataset:
# + id="2l9ANJTrbP-U" outputId="f54a46ba-952d-4578-9b15-06b0bd28e3eb" colab={"base_uri": "https://localhost:8080/"}
tydiqa_data['train']
# + [markdown] id="2xRO1yIkvpUt"
# You can see that each example is like a dictionary object. This dataset consists of questions, contexts, and indices that point to the start and end position of the answer inside the context. You can access the index using the `annotations` key, which is a kind of dictionary.
# + id="KNVpW6lADk92" outputId="c639d790-2a04-4d7c-c396-2ec403985172" colab={"base_uri": "https://localhost:8080/"}
idx = 600
# start index
start_index = tydiqa_data['train'][idx]['annotations']['minimal_answers_start_byte'][0]
# end index
end_index = tydiqa_data['train'][idx]['annotations']['minimal_answers_end_byte'][0]
print("Question: " + tydiqa_data['train'][idx]['question_text'])
print("\nContext (truncated): "+ tydiqa_data['train'][idx]['document_plaintext'][0:512] + '...')
print("\nAnswer: " + tydiqa_data['train'][idx]['document_plaintext'][start_index:end_index])
# + [markdown] id="Z-lZgDTEYm74"
# The question answering model predicts a start and endpoint in the context to extract as the answer. That's why this NLP task is known as extractive question answering.
#
# To train your model, you need to pass start and endpoints as labels. So, you need to implement a function that extracts the start and end positions from the dataset.
#
# The dataset contains unanswerable questions. For these, the start and end indices for the answer are equal to `-1`.
# + id="Ty_QDcdKYw9a" outputId="e4392bc0-2583-48e0-95f6-f3ee1a33d740" colab={"base_uri": "https://localhost:8080/"}
tydiqa_data['train'][0]['annotations']
# + [markdown] id="lHWcNMudcAuO"
# Now, you have to flatten the dataset to work with an object with a table structure instead of a dictionary structure. This step facilitates the pre-processing steps.
# + id="xDCAQQtoCs_r"
# Flattening the datasets
flattened_train_data = tydiqa_data['train'].flatten()
flattened_test_data = tydiqa_data['validation'].flatten()
# + [markdown] id="q5wUa5xED0fK"
# Also, to make the training more straightforward and faster, we will extract a subset of the train and test datasets. For that purpose, we will use the Hugging Face Dataset object's method called `select()`. This method allows you to take some data points by their index. Here, you will select the first 3000 rows; you can play with the number of data points but consider that this will increase the training time.
# + id="BkcIhpEnDHSJ"
# Selecting a subset of the train dataset
flattened_train_data = flattened_train_data.select(range(3000))
# Selecting a subset of the test dataset
flattened_test_data = flattened_test_data.select(range(1000))
# + [markdown] id="fBXrmwXhc13M"
# ### Tokenizers
#
# Now, you will use the [tokenizer](https://huggingface.co/transformers/main_classes/tokenizer.html) object from Hugging Face. You can load a tokenizer using different methods. Here, you will retrieve it from the pipeline object you created in the previous Hugging Face lab. With this tokenizer, you can ensure that the tokens you get for the dataset will match the tokens used in the original DistilBERT implementation.
#
# When loading a tokenizer with any method, you must pass the model checkpoint that you want to fine-tune. Here, you are using the`'distilbert-base-cased-distilled-squad'` checkpoint.
#
# You can uncomment the cell below to load the tokenizer without creating a pipeline object.
#
# + id="LInV3b_HyAIF" outputId="2bacf5aa-35cb-41b8-92fd-8dd4970f422a" colab={"base_uri": "https://localhost:8080/", "height": 145, "referenced_widgets": ["3f36586ca9614a64ab1f6f9378df1490", "130ef5be7835403c820002b4600bb8ee", "fc41328fbf804b3ab9ced0dc6607138f", "bd48a989578d435e8c7ff28e69e2ff9d", "<KEY>", "d9a30e49a30a4445920167c0d888ee91", "<KEY>", "f9b472ff9aae41238ad1d25ef0425f63", "<KEY>", "e138d0c1afe145b1a505b30a70a2f745", "6565fefb972d40adb529bec74a3f9102", "b260ef614be44f52934e5441fde24946", "d1318b1ec5254beba3067efdd2d8a0e2", "ac25e836ccd147728d58aa5db5e9f6de", "f238f6e5004e4da89bacbf1aab8650e7", "<KEY>", "05f745caf23a4ab6ab10600ce00770d7", "<KEY>", "603ae16b56db484ea45a1cb2ed1ddb18", "<KEY>", "<KEY>", "<KEY>", "51e3d37b14664465a2f5b292a947c372", "<KEY>", "3e12f946c064418bb7e6569008bea439", "<KEY>", "<KEY>", "879329dc8a2d488cb1a9cbce2ff077ec", "442e0fd5084c4ed18dcc526184b28a82", "<KEY>", "<KEY>", "<KEY>", "e113a903a6ee496dbafb190d2f5129a3", "2f84ba11280542a699ff89dfdab7e194", "f6283192f62749f9a5514f21f8b8cb4e", "07f39708d928453587fe4dc7f66c35a5", "<KEY>", "3a28250b9b014a909048dac6719ebdd4", "6e2e403aa36347b886ed5a46735b5794", "ffd7f80a1070440fa1e324719573638a", "1efb5b733cc24cad8e3ac4756385115f", "a7c9b2f0087842e8867aebad450f1ad7", "<KEY>", "<KEY>"]}
# Import the AutoTokenizer from the transformers library
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained("distilbert-base-cased-distilled-squad")
# + [markdown] id="qz6YtVcOh3qP"
# Given the characteristics of the dataset and the question-answering task, you will need to add some steps to pre-process the data after the tokenization:
#
# 1. When there is no answer to a question given a context, you will use the `CLS` token, a unique token used to represent the start of the sequence.
#
# 2. Tokenizers can split a given string into substrings, resulting in a subtoken for each substring, creating misalignment between the list of dataset tags and the labels generated by the tokenizer. Therefore, you will need to align the start and end indices with the tokens associated with the target answer word.
#
# 3. Finally, a tokenizer can truncate a very long sequence. So, if the start/end position of an answer is `None`, you will assume that it was truncated and assign the maximum length of the tokenizer to those positions.
#
# Those three steps are done within the `process_samples` function defined below.
# + id="3l-r4wI06LU7"
# Processing samples using the 3 steps described.
def process_samples(sample):
tokenized_data = tokenizer(sample['document_plaintext'], sample['question_text'], truncation="only_first", padding="max_length")
input_ids = tokenized_data["input_ids"]
# We will label impossible answers with the index of the CLS token.
cls_index = input_ids.index(tokenizer.cls_token_id)
# If no answers are given, set the cls_index as answer.
if sample["annotations.minimal_answers_start_byte"][0] == -1:
start_position = cls_index
end_position = cls_index
else:
# Start/end character index of the answer in the text.
gold_text = sample["document_plaintext"][sample['annotations.minimal_answers_start_byte'][0]:sample['annotations.minimal_answers_end_byte'][0]]
start_char = sample["annotations.minimal_answers_start_byte"][0]
end_char = sample['annotations.minimal_answers_end_byte'][0] #start_char + len(gold_text)
# sometimes answers are off by a character or two – fix this
if sample['document_plaintext'][start_char-1:end_char-1] == gold_text:
start_char = start_char - 1
end_char = end_char - 1 # When the gold label is off by one character
elif sample['document_plaintext'][start_char-2:end_char-2] == gold_text:
start_char = start_char - 2
end_char = end_char - 2 # When the gold label is off by two characters
start_token = tokenized_data.char_to_token(start_char)
end_token = tokenized_data.char_to_token(end_char - 1)
# if start position is None, the answer passage has been truncated
if start_token is None:
start_token = tokenizer.model_max_length
if end_token is None:
end_token = tokenizer.model_max_length
start_position = start_token
end_position = end_token
return {'input_ids': tokenized_data['input_ids'],
'attention_mask': tokenized_data['attention_mask'],
'start_positions': start_position,
'end_positions': end_position}
# + [markdown] id="Q3LAsWSyk_Rm"
# To apply the `process_samples` function defined above to the whole dataset, you can use the `map` method as follows:
# + id="rGbYd7QnFetG" outputId="6ef24640-b756-41f5-d630-7362b0cb7139" colab={"base_uri": "https://localhost:8080/", "height": 81, "referenced_widgets": ["7ac95ce0c2df4f2d81e0bd517fbc47b3", "<KEY>", "c53ec539e27c44829816ad67725b7821", "<KEY>", "9e8707b4dffd41e3bef7b1077fe16a17", "63afeafe09bc4ebd87a4ac7e8b92f0a8", "<KEY>", "<KEY>", "<KEY>", "5716150d89a64f92a7b605a726a02901", "ec883dffb2154853b5f621a3a4853127", "a79d1eafe6834530af9ec5a05f151701", "aa64c556d13040d293c2c8e716ed9deb", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "7ac172b52cd1429eb84e19394d858fba", "<KEY>", "3f442de4bfe949c1ba91d94da00ced55", "625f1571d4fe421798d0517277c785e4"]}
# Tokenizing and processing the flattened dataset
processed_train_data = flattened_train_data.map(process_samples)
processed_test_data = flattened_test_data.map(process_samples)
# + [markdown] id="wCpPhYKJluMA"
# # Transformers
#
# The last component of Hugging Face that is useful for fine-tuning a transformer corresponds to the pre-trained models you can access in multiple ways.
#
# For this lab, you will use the same model from the question-answering pipeline that you loaded before. But, if you want, you can uncomment the cell below to load the pre-trained model without accessing the pipeline parameters.
#
# You will set the base model layers fixed so the training process doesn't take too long. To do so, you have to set the `requires_grad` parameter for the `base_model` layers to `False`.
# + id="jR3VqjNc1Vb3" outputId="2a6d95eb-25b7-4342-f465-f92f9e06d0b9" colab={"base_uri": "https://localhost:8080/", "height": 49, "referenced_widgets": ["211026dc0da342198f23c9e819197cbd", "695ade9aa4684836ae4a6484d14a5a0b", "5ab01b509894478682291a70cd81a9f8", "c9730a2c9f124e039e61ea7fe98ad9c0", "<KEY>", "983eeb857cde4dcab5a7a7d809ab108a", "b2f300d0e1ca41e289841f0161405d76", "7c7a165a213e4547a36f86e869bc837e", "<KEY>", "34d2cde86ea1452782e226700e3261ec", "ff37fc5115f644bd8b93ae38b11c9f24"]}
# Import the AutoModelForQuestionAnswering for the pre-trained model. We will only fine tune the head of the model
from transformers import AutoModelForQuestionAnswering
model = AutoModelForQuestionAnswering.from_pretrained("distilbert-base-cased-distilled-squad")
# + [markdown] id="K29BYtnsm1yH"
# Now, you can take the necessary columns from the datasets to train/test and return them as Pytorch Tensors.
# + id="0X14G89noLfW"
columns_to_return = ['input_ids','attention_mask', 'start_positions', 'end_positions']
processed_train_data.set_format(type='pt', columns=columns_to_return)
processed_test_data.set_format(type='pt', columns=columns_to_return)
# + [markdown] id="yjoUFWu_nLRq"
# Here, we give you the F1 score as a metric to evaluate your model's performance. We will use this metric for simplicity, although it is based on the start and end values predicted by the model. If you want to dig deeper on other metrics that can be used for a question and answering task, you can also check [this colab notebook resource](https://colab.research.google.com/github/huggingface/notebooks/blob/master/examples/question_answering.ipynb) from the Hugging Face team.
# + id="xcW2wPnirsJk"
from sklearn.metrics import f1_score
def compute_f1_metrics(pred):
start_labels = pred.label_ids[0]
start_preds = pred.predictions[0].argmax(-1)
end_labels = pred.label_ids[1]
end_preds = pred.predictions[1].argmax(-1)
f1_start = f1_score(start_labels, start_preds, average='macro')
f1_end = f1_score(end_labels, end_preds, average='macro')
return {
'f1_start': f1_start,
'f1_end': f1_end,
}
# + [markdown] id="KuhASU4evpUu"
# Now, you will use the Hugging Face [Trainer](https://huggingface.co/transformers/main_classes/trainer.html) to fine-tune your model.
# + id="nxyOwf5utXAt" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="af5ee882-9e13-4e53-b196-72d0eae71d42"
# Training the model may take around 15 minutes.
from transformers import Trainer, TrainingArguments
training_args = TrainingArguments(
output_dir='model_results5', # output directory
overwrite_output_dir=True,
num_train_epochs=3, # total number of training epochs
per_device_train_batch_size=8, # batch size per device during training
per_device_eval_batch_size=8, # batch size for evaluation
warmup_steps=20, # number of warmup steps for learning rate scheduler
weight_decay=0.01, # strength of weight decay
logging_dir=None, # directory for storing logs
logging_steps=50
)
trainer = Trainer(
model=model, # the instantiated 🤗 Transformers model to be trained
args=training_args, # training arguments, defined above
train_dataset=processed_train_data, # training dataset
eval_dataset=processed_test_data, # evaluation dataset
compute_metrics=compute_f1_metrics
)
trainer.train()
# + [markdown] id="Ic_wNlBHCRMn"
# And, in the next cell, you can evaluate the fine-tuned model's performance on the test set.
# + id="92N11A076wRA" outputId="20dd23dc-f0d1-40c9-902f-bb27a65023bd" colab={"base_uri": "https://localhost:8080/", "height": 248}
# The evaluation may take around 30 seconds
trainer.evaluate(processed_test_data)
# + [markdown] id="_HubPkRbnzh_"
# ### Using your Fine-Tuned Model
#
# After training and evaluating your fine-tuned model, you can check its results for the same questions from the previous lab.
#
# For that, you will tell Pytorch to use your GPU or your CPU to run the model. Additionally, you will need to tokenize your input context and questions. Finally, you need to post-process the output results to transform them from tokens to human-readable strings using the `tokenizer`.
# + id="yxZGsmIOBCZ6" outputId="42b95d9e-240e-4a68-cba7-33cc8f65c04c" colab={"base_uri": "https://localhost:8080/"}
import torch
text = r"""
The Golden Age of Comic Books describes an era of American comic books from the
late 1930s to circa 1950. During this time, modern comic books were first published
and rapidly increased in popularity. The superhero archetype was created and many
well-known characters were introduced, including Superman, Batman, Captain Marvel
(later known as SHAZAM!), Captain America, and Wonder Woman.
Between 1939 and 1941 Detective Comics and its sister company, All-American Publications,
introduced popular superheroes such as Batman and Robin, Wonder Woman, the Flash,
Green Lantern, Doctor Fate, the Atom, Hawkman, Green Arrow and Aquaman.[7] Timely Comics,
the 1940s predecessor of Marvel Comics, had million-selling titles featuring the Human Torch,
the Sub-Mariner, and Captain America.[8]
As comic books grew in popularity, publishers began launching titles that expanded
into a variety of genres. Dell Comics' non-superhero characters (particularly the
licensed Walt Disney animated-character comics) outsold the superhero comics of the day.[12]
The publisher featured licensed movie and literary characters such as Mickey Mouse, Donald Duck,
Roy Rogers and Tarzan.[13] It was during this era that noted Donald Duck writer-artist
<NAME> rose to prominence.[14] Additionally, MLJ's introduction of Archie Andrews
in Pep Comics #22 (December 1941) gave rise to teen humor comics,[15] with the Archie
Andrews character remaining in print well into the 21st century.[16]
At the same time in Canada, American comic books were prohibited importation under
the War Exchange Conservation Act[17] which restricted the importation of non-essential
goods. As a result, a domestic publishing industry flourished during the duration
of the war which were collectively informally called the Canadian Whites.
The educational comic book Dagwood Splits the Atom used characters from the comic
strip Blondie.[18] According to historian <NAME>, appealing comic-book
characters helped ease young readers' fear of nuclear war and neutralize anxiety
about the questions posed by atomic power.[19] It was during this period that long-running
humor comics debuted, including EC's Mad and Carl Barks' Uncle Scrooge in Dell's Four
Color Comics (both in 1952).[20][21]
"""
questions = ["What superheroes were introduced between 1939 and 1941 by Detective Comics and its sister company?",
"What comic book characters were created between 1939 and 1941?",
"What well-known characters were created between 1939 and 1941?",
"What well-known superheroes were introduced between 1939 and 1941 by Detective Comics?"]
for question in questions:
inputs = tokenizer.encode_plus(question, text, return_tensors="pt")
#print("inputs", inputs)
#print("inputs", type(inputs))
input_ids = inputs["input_ids"].tolist()[0]
inputs.to("cuda")
text_tokens = tokenizer.convert_ids_to_tokens(input_ids)
answer_model = model(**inputs)
answer_start = torch.argmax(
answer_model['start_logits']
) # Get the most likely beginning of answer with the argmax of the score
answer_end = torch.argmax(answer_model['end_logits']) + 1 # Get the most likely end of answer with the argmax of the score
answer = tokenizer.convert_tokens_to_string(tokenizer.convert_ids_to_tokens(input_ids[answer_start:answer_end]))
print(f"Question: {question}")
print(f"Answer: {answer}\n")
# + [markdown] id="_yTDQ6kn6pWS"
# You can compare those results with those obtained using the pipeline, as you did in the previous lab. As a reminder, here are those results:
#
# ```
# What popular superheroes were introduced between 1939 and 1941?
# >> teen humor comics
# What superheroes were introduced between 1939 and 1941 by Detective Comics and its sister company?
# >> <NAME>
# What comic book characters were created between 1939 and 1941?
# >> Archie
# Andrews
# What well-known characters were created between 1939 and 1941?
# >> Archie
# Andrews
# What well-known superheroes were introduced between 1939 and 1941 by Detective Comics?
# >> <NAME>
# ```
# + [markdown] id="uf-v8mUSLqXN"
# **Congratulations!**
#
# You have finished this series of ungraded labs. You were able to:
#
# * Explore the Hugging Face Pipelines, which can be used right out of the bat.
#
# * Fine-tune a model for the Extractive Question & Answering task.
#
# I recommend you go through the free [Hugging Face course](https://huggingface.co/course/chapter1) to explore their ecosystem in more detail and find different ways to use the `transformers` library.
# + id="5ElDBNE2SppH"
| Week-3/Ungraded-Assignments/C4_W3_2_Question_Answering_with_BERT_and_HuggingFace_Pytorch_tydiqa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# MODIFY!
# use Robust!
model_name = 'ela-net-rb-wo'
# # Import Libraries & Data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df = pd.read_csv('./data/d-wo-ns.csv')
X = df.drop('throughput',axis=1)
y = df['throughput']
# ---
# # Scale Data
# +
# Split the data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=42)
# Scale the data
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
scaler.fit(X_train)
# -
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
# # Determine Hyperparameters
# MODIFY!
from sklearn.linear_model import ElasticNetCV
model = ElasticNetCV(l1_ratio=[.1, .5, .7,.8,.9, .95, .99, 1],max_iter=1000000,cv=10)
model
model.fit(X_train, y_train)
model.l1_ratio_
model.alpha_
hp = pd.Series(name=f'{model_name} HP', data=[model.l1_ratio_, model.alpha_],index=['l1_ratio','alpha'])
hp
hp.to_csv(f'./hyperparameters/{model_name}.csv')
# # Score Model
# +
# MODIFY!
from sklearn.linear_model import ElasticNet
score_model = ElasticNet(
alpha = model.alpha_,
l1_ratio = model.l1_ratio_
)
score_model
# +
from sklearn.model_selection import cross_validate
scores = cross_validate(
score_model,
X_train,
y_train,
scoring=[
'neg_mean_absolute_error',
'neg_mean_squared_error',
'neg_root_mean_squared_error',
],
cv=10
)
# -
scores = pd.DataFrame(scores)
scores
mean_scores = scores.mean()
mean_scores
# # Export Model Score
mean_scores = mean_scores.rename(f'{model_name}')
mean_scores[[2,3,4]] = mean_scores[[2,3,4]].apply(abs)
# +
# mean_scores
# -
mean_scores = mean_scores.rename({
'fit_time':'Fit Time',
'score_time':'Score Time',
'test_neg_mean_absolute_error':'MAE',
'test_neg_mean_squared_error':'MSE',
'test_neg_root_mean_squared_error':'RMSE'
})
mean_scores
mean_scores['STD FT'] = scores.fit_time.std()
mean_scores['STD ST'] = scores.score_time.std()
mean_scores['STD MAE'] = scores.test_neg_mean_absolute_error.std()
mean_scores['STD MSE'] = scores.test_neg_mean_squared_error.std()
mean_scores['STD RMSE'] = scores.test_neg_root_mean_squared_error.std()
mean_scores
mean_scores.to_csv(f'./scores/{model_name}.csv')
# # Holdout Test
score_model.fit(X_train,y_train)
y_holdout = score_model.predict(X_test)
sns.histplot(y_holdout,bins=40,kde=True)
# +
fig,ax = plt.subplots()
sns.kdeplot(y_test, ax=ax, shade=True, label='Observations')
sns.kdeplot(y_holdout, ax=ax,shade=True,label='Predictions')
ax.legend(loc='best')
# +
fig,ax = plt.subplots()
sns.ecdfplot(y_test, ax=ax, label='Observations')
sns.ecdfplot(y_holdout, ax=ax,label='Predictions')
plt.axvline(x=y_test.mean(),color='grey',linestyle='--')
ax.legend(loc='best')
# +
from sklearn.metrics import mean_absolute_error, mean_squared_error
mae = mean_absolute_error(y_test,y_holdout)
mse = mean_squared_error(y_test,y_holdout)
rmse = np.sqrt(mse)
# -
err_df = pd.DataFrame(data=[mae, mse, rmse],index=['MAE','MSE','RMSE'],columns=[f'{model_name}'])
err_df
err_df.to_csv(f'./holdout-test/{model_name}.csv')
# mean_scores.to_csv(f'./opt-model-err/{model_name}-err.csv')
res = y_test - y_holdout
res.describe().drop('count')
sns.histplot(data=res, kde=True,bins=40)
# +
ax = sns.scatterplot(x=y_test, y=res)
ax.set(ylabel='Residuals', xlabel='Test Label')
plt.axhline(y=0,color='red',linestyle='--')
# there should be no clear pattern / curve in the plot
# we see a positive correlation between Test Label and Residuals -> later models should avoid this pattern
# +
import scipy as sp
fig, ax = plt.subplots()
sp.stats.probplot(res,plot=ax);
# -
# # Export Optimized Model
from sklearn.preprocessing import RobustScaler
scaler = RobustScaler()
scaler.fit(X)
# +
# MODIFY!
X = scaler.transform(X)
op_model = score_model
op_model.fit(X,y)
# -
y_pred = op_model.predict(X)
sns.histplot(y_pred,bins=40,kde=True)
# +
fig,ax = plt.subplots()
sns.kdeplot(y, ax=ax, shade=True, label='Observations')
sns.kdeplot(y_pred, ax=ax,shade=True,label='Predictions')
ax.legend(loc='best')
# +
fig,ax = plt.subplots()
sns.ecdfplot(y, ax=ax, label='Observations')
sns.ecdfplot(y_pred, ax=ax,label='Predictions')
plt.axvline(x=y.mean(),color='grey',linestyle='--')
ax.legend(loc='best')
# -
mae = mean_absolute_error(y,y_pred)
mse = mean_squared_error(y,y_pred)
rmse = np.sqrt(mse)
err_df = pd.DataFrame(data=[mae, mse, rmse],index=['MAE','MSE','RMSE'],columns=[f'{model_name}'])
err_df
err_df.to_csv(f'./model-err/{model_name}.csv')
# mean_scores.to_csv(f'./opt-model-err/{model_name}-err.csv')
from joblib import dump, load
dump(op_model, f'./opt-models/{model_name}.joblib')
res = y - y_pred
res.describe().drop('count')
sns.histplot(data=res, kde=True,bins=40)
# +
ax = sns.scatterplot(x=y_test, y=res)
ax.set(ylabel='Residuals', xlabel='Test Label')
plt.axhline(y=0,color='red',linestyle='--')
# there should be no clear pattern / curve in the plot
# we see a positive correlation between Test Label and Residuals -> later models should avoid this pattern
# +
import scipy as sp
fig, ax = plt.subplots()
sp.stats.probplot(res,plot=ax);
# -
# DONE!
| 1-dl-project/dl-4-ela-net-opt-rb-wo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
def off_win(row, games_df):
score = games_df.iloc[row.loc['gid'] - 1]
if row['off'] == score['h']:
if score.ptsh > score.ptsv:
return 1
else:
return 0
elif row['off'] == score['v']:
if score.ptsv > score.ptsh:
return 1
else:
return 0
else:
raise Exception('Teams do not match: %s vs. %s; %s vs. %s' % (
row['off'], row['def'], score['v'], score['h']))
plays_df = pd.read_csv('../data/raw/PLAY.csv')
games_df = pd.read_csv('../data/raw/GAME.csv')
plays_df = plays_df.loc[:, [
'gid',
'off',
'def',
'qtr',
'min',
'sec',
'ptso',
'ptsd',
'timo',
'timd',
'dwn',
'ytg',
'yfog'
]]
plays_df['y'] = plays_df.apply(lambda row: off_win(row, games_df), axis=1)
plays_df.to_csv('../data/Xy.csv')
| notebooks/format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (PythonData)
# language: python
# name: pythondata
# ---
# +
# Dependencies
import json
import pandas as pd
import csv
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
analyzer = SentimentIntensityAnalyzer()
from textblob import TextBlob
# # Twitter API Keys
# consumer_key = "yUHOWwWG9Pj3C8f21tnm4rWOb"
# consumer_secret = "<KEY>"
# access_token = "<KEY>"
# access_token_secret = "<KEY>"
# +
# # Setup Tweepy API Authentication
# auth = tweepy.OAuthHandler(consumer_key, consumer_secret)
# auth.set_access_token(access_token, access_token_secret)
# api = tweepy.API(auth, parser=tweepy.parsers.JSONParser())
# # Get all tweets from home feed
# public_tweets = api.home_timeline()
# Import CSV file into Data Frame
tweeter_data = pd.read_csv("Output_tweeter_bitcoin.csv", dtype=str)
tweeter_data.head()
# len(df_tweeter_data)
# -
df_tweeter_data = pd.DataFrame(tweeter_data)
df_tweeter_data.head()
columns = ['date','text','username','keyword']
df1_tweeter_data = pd.DataFrame(df_tweeter_data, columns=columns)
df1_tweeter_data.head()
len(df1_tweeter_data)
df2_tweeter_data = df1_tweeter_data.drop_duplicates(["text","username"])
len(df2_tweeter_data)
df2_tweeter_data=df2_tweeter_data.fillna('')
# +
# Search for all tweets
tweets_text = df2_tweeter_data["text"]
# counter = 0
compound_list = []
positive_list = []
negative_list = []
neutral_list = []
all_tweets = []
for tweet in tweets_text:
if type(tweet) is str:
# all_tweets.append(tweet)
# tx= TextBlob(t)
# counter+=1
# print(counter)
# print(tweet)
compound = analyzer.polarity_scores(tweet)["compound"]
# print(compound)
pos = analyzer.polarity_scores(tweet)["pos"]
neu = analyzer.polarity_scores(tweet)["neu"]
neg = analyzer.polarity_scores(tweet)["neg"]
# # # Add each value to the appropriate list
compound_list.append(compound)
positive_list.append(pos)
negative_list.append(neg)
neutral_list.append(neu)
# # # Print the Averages
# print(compound_list)
print(len(compound_list))
# print(*************)
# print("+ve" + str(positive_list))
print(np.mean(positive_list))
print(len(negative_list))
print(len(neutral_list))
# print("-ve" + str(negative_list))
# print("neutral" + str(neutral_list))
# # len(tweets_text)
# -
import numpy as np
test = np.array(compound_list)
df2_tweeter_data["Compound_score"] = compound_list
df2_tweeter_data.head()
# +
# wordlist=pd.DataFrame();
# polarity=[]
# subj=[]
# def sentiment_calc(text):
# try:
# return TextBlob(text).sentiment
# except:
# return None
# df2_tweeter_data['sentiment'] = df2_tweeter_data['text'].apply(sentiment_calc)
# # for t in df2_tweeter_data["text"]:
# # tx= TextBlob(t)
# # polarity.append(tx.sentiment.polarity)
# # subj.append(tx.sentiment.subjectivity)
# # poltweet= pd.DataFrame({'polarity':polarity,'subjectivity':subj})
# # poltweet.polarity.plot(title='Polarity')
# # plt.show()
# -
df2_tweeter_data.head(5)
| For review/Project_Crypto_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Package imports
# %matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import random
import time
import math
import pdb
import pickle
import pandas as pd
from sklearn import preprocessing
import seaborn as sns
import os
import time
#from datasets import dataset_utils
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# Main slim library
slim = tf.contrib.slim
# +
# Model Architecture Defs
def mlp_model(inputs, net_arch, is_training, scope="mlp", net_type='mlpnet'):
"""Creates the regression model.
Args:
inputs: A node that yields a `Tensor` of size [batch_size, dimensions].
net_arch: A model architecture with node_sizes, dropout, and other hyper-parmas
is_training: Whether or not we're currently training the model.
scope: An optional variable_op scope for the model.
Returns:
predictions: 1-D `Tensor` of shape [batch_size] of responses.
end_points: A dict of end points representing the hidden layers.
"""
end_points = {}
if net_type == 'mlpnet':
with tf.variable_scope(scope, 'mlp', [inputs]):
# Set the default weight _regularizer and acvitation for each fully_connected layer.
with slim.arg_scope([slim.fully_connected], activation_fn=tf.nn.relu,
weights_regularizer=slim.l2_regularizer(net_arch['reg'])):
# Creates a fully connected layer from the inputs
net = slim.fully_connected(inputs, net_arch['l1'], normalizer_fn=slim.batch_norm, scope='fc1')
end_points['fc1'] = net
net = slim.dropout(net, net_arch['keep_p'], is_training=is_training,scope='drop1')
# Adds another fully connected layer
net = slim.fully_connected(net, net_arch['l2'], normalizer_fn=slim.batch_norm, scope='fc2')
end_points['fc2'] = net
# net = slim.dropout(net, net_arch['keep_p'], is_training=is_training)
#Adds another fully connected layer
# net = slim.fully_connected(net, net_arch['l3'], normalizer_fn=slim.batch_norm, scope='fc3')
# end_points['fc3'] = net
#net = slim.dropout(net, net_arch['keep_p'], is_training=is_training)
# Creates a fully-connected layer with a single hidden unit. Note that the
# layer is made linear by setting activation_fn=None.
predictions = slim.fully_connected(net, net_arch['output'], activation_fn=None,
normalizer_fn=slim.batch_norm, scope='prediction')
end_points['out'] = predictions
elif net_type == 'mult_mod':
with tf.variable_scope(scope, 'mult_mod', [inputs]):
d = inputs['d']
cs = inputs['cs']
attr = inputs['attr']
if net_arch['dist_mod'] == 'embed':
attr_mod = slim.fully_connected(attr, net_arch['aux_embed'], activation_fn=tf.nn.relu,
weights_regularizer=slim.l2_regularizer(net_arch['reg']),
normalizer_fn=slim.batch_norm, scope='attr_mod')
d_mod = tf.multiply(d,attr_mod)
print('shapes: d:{}, attr:{}, attr_mod {}, dmod {}'.format(d.shape,attr.shape,
attr_mod.shape,d_mod.shape))
predictions = slim.fully_connected(tf.concat([d_mod,cs],1), net_arch['aux_out'], activation_fn=None,
normalizer_fn=slim.batch_norm,
weights_regularizer=slim.l2_regularizer(net_arch['reg']),
scope='aux_prediction')
elif net_arch['dist_mod'] == 'mult':
d_mod = tf.multiply(d,attr)
print('shapes: d:{}, attr:{}, dmod {}'.format(d.shape,attr.shape,d_mod.shape))
predictions = slim.fully_connected(d_mod, net_arch['aux_out'], activation_fn=None,
normalizer_fn=slim.batch_norm,
weights_regularizer=slim.l2_regularizer(net_arch['reg']),
scope='aux_prediction')
else:
d_mod = d
predictions = slim.fully_connected(tf.concat([d_mod,cs],1), net_arch['aux_out'], activation_fn=None,
normalizer_fn=slim.batch_norm,
weights_regularizer=slim.l2_regularizer(net_arch['reg']),
scope='aux_prediction')
end_points['aux_out'] = predictions
if d_mod.shape[1] > 1:
end_points['d_mod'] = tf.reduce_sum(tf.pow(d_mod,2),1,keep_dims=True)
else:
end_points['d_mod'] = d_mod
end_points['dist'] = d
elif net_type == 'dvec_mod':
with tf.variable_scope(scope, 'dvec_mod', [inputs]):
dvec = inputs['d']
cs = inputs['cs']
attr = inputs['attr']
if net_arch['dist_mod'] == 'embed':
attr_mod = slim.fully_connected(attr, net_arch['aux_embed'], activation_fn=tf.nn.relu,
normalizer_fn=slim.batch_norm,scope='attr_mod')
d_mod = tf.multiply(dvec,attr_mod,name='d_mod')
print('shapes: d:{}, attr:{}, attr_mod {}, dmod {}'.format(dvec.shape,attr.shape,
attr_mod.shape,d_mod.shape))
else:
d_mod = dvec
print('shapes: d:{}, dmod {}'.format(dvec.shape,d_mod.shape))
net = slim.fully_connected(tf.concat([d_mod,cs],1), net_arch['l3'], activation_fn=tf.nn.relu,
weights_regularizer=slim.l2_regularizer(net_arch['reg']),
normalizer_fn=slim.batch_norm, scope='fc3')
net = slim.dropout(net, net_arch['keep_p'], is_training=is_training)
predictions = slim.fully_connected(net, net_arch['aux_out'], activation_fn=None,
normalizer_fn=slim.batch_norm, scope='aux_prediction')
end_points['aux_out'] = predictions
end_points['d_mod'] = tf.reduce_sum(tf.pow(d_mod,2),1,keep_dims=True)
end_points['dist'] = tf.reduce_sum(tf.pow(dvec,2),1,keep_dims=True)
else:
print('Unknown Net Arch')
return predictions, end_points
def produce_batch(batch_size, noise=0.2):
xs = np.random.random(size=[batch_size, 1]) * 4
ys = np.round(xs + np.random.normal(size=[batch_size, 1], scale=noise))
return [xs.astype(np.float32), ys.astype(np.int32)]
def convert_data_to_tensors(x, y):
inputs = tf.constant(x)
inputs.set_shape([None, 1])
outputs = tf.constant(y)
outputs.set_shape([None, 1])
return inputs, outputs
# +
# Load data
baseline_dir = '/projects/nbhagwat/Traj_prediction/ADNI/exp_setup/'
demo_cols = ['AGE','APOE4']
cohort = 'ALL_ADNI'
clinical_scale = 'MMSE'
exp_name = 'Exp_502_{}_traj_{}_tp_var_tp'.format(cohort,clinical_scale)
use_cs = False
use_ct = True
var_tp = True
shuffle_labels = False # What's your performance with random labels.
scale_data = False
feat_sufix = ''
exp_setup_path = baseline_dir + 'KFolds/' + exp_name + '_sKF.pkl'
with open(exp_setup_path, 'rb') as f:
exp_setup = pickle.load(f, encoding='latin')
#Check if loading the correct experimental setup
if exp_name != exp_setup['exp_name']:
print('Incorrect exp_name. Selected:{}, Retrieved:{}'.format(exp_name, exp_setup['exp_name']))
else:
print('exp_name: {}'.format(exp_name))
df = exp_setup['df']
kf = exp_setup['sKF']
feat_cols = []
if use_cs:
print('using CS columns')
if var_tp:
cs_cols = ['MMSE_bl','MMSE_var_tp'] #+ demo_cols
else:
cs_cols = ['MMSE_bl'] + demo_cols
feat_cols = feat_cols + cs_cols
feat_sufix = feat_sufix + '_CS'
print(len(feat_cols))
if use_ct:
print('using CT columns')
if var_tp:
ct_cols_bl = list(df.columns[pd.Series(df.columns).str.contains('CT_bl')])
ct_cols_tp_rate = list(df.columns[pd.Series(df.columns).str.contains('CT_var_tp')])
ct_cols = ct_cols_bl + ct_cols_tp_rate
else:
ct_cols = list(df.columns[pd.Series(df.columns).str.contains('CT')])
print('# of CT cols {}'.format(len(ct_cols)))
#Exclude certain ROIs (based on QC e.g. increase CT )
# exclude_roi_list = ['OLF'] #PCL'REC'TPOmid
# roi_suffix_list = ['L_CT_bl','R_CT_bl','L_CT_var_tp','R_CT_var_tp']
# for exclude_roi in exclude_roi_list:
# for roi_suffix in roi_suffix_list:
# ct_cols.remove(exclude_roi+'.'+roi_suffix)
print('# of CT cols after exclusion {}'.format(len(ct_cols)))
feat_cols = feat_cols + ct_cols
feat_sufix = feat_sufix + '_CT'
X_raw = np.array(df[feat_cols]) #Otherwise scaling and shifting will remain the same for next model.
y = np.array(df['traj']).astype(int)
#Flip labels (sanity check to see of contrastive loss is dependent on the label values)
#y = np.square(y-1)
print('X,y shapes:{},{}'.format(X_raw.shape,y.shape))
# Create list of all the fold-subsets (needed for parallelization)
Sub_train_kf =[]
Sub_test_kf = []
X_train_kf = []
X_test_kf = []
y_train_kf = []
y_test_kf = []
delta_t_train_kf = []
delta_t_test_kf = []
age_train_kf = []
age_test_kf = []
apoe_train_kf = []
apoe_test_kf = []
cs_bl_train_kf = []
cs_bl_test_kf = []
cs_tp_train_kf = []
cs_tp_test_kf = []
for train, test in kf:
Sub_train_kf.append(df['PTID'].values[train])
Sub_test_kf.append(df['PTID'].values[test])
delta_t_train_kf.append(df['tp'].values[train]/12.0)
delta_t_test_kf.append(df['tp'].values[test]/12.0)
age_tr = df['AGE'].values[train]
age_te = df['AGE'].values[test]
apoe_tr = df['APOE4'].values[train]
apoe_te = df['APOE4'].values[test]
cs_bl_tr = df['MMSE_bl'].values[train]
cs_bl_te = df['MMSE_bl'].values[test]
cs_tp_tr = df['MMSE_var_tp'].values[train]
cs_tp_te = df['MMSE_var_tp'].values[test]
X_tr = X_raw[train]
y_tr = y[train]
X_te = X_raw[test]
y_te = y[test]
if scale_data:
scaler = preprocessing.StandardScaler().fit(X_tr)
X_tr = scaler.transform(X_tr)
X_te = scaler.transform(X_te)
scaler = preprocessing.StandardScaler().fit(age_tr.reshape(-1,1))
age_tr = scaler.transform(age_tr)
age_te = scaler.transform(age_te)
# scaler = preprocessing.StandardScaler().fit(apoe_tr.reshape(-1,1))
# apoe_tr = scaler.transform(apoe_tr)
# apoe_te = scaler.transform(apoe_te)
scaler = preprocessing.StandardScaler().fit(cs_bl_tr.reshape(-1,1))
cs_bl_tr = scaler.transform(cs_bl_tr)
cs_bl_te = scaler.transform(cs_bl_te)
scaler = preprocessing.StandardScaler().fit(cs_tp_tr.reshape(-1,1))
cs_tp_tr = scaler.transform(cs_tp_tr)
cs_tp_te = scaler.transform(cs_tp_te)
print('cs_bl_tr: {}'.format(np.mean(cs_bl_tr)))
print('cs_bl_te: {}'.format(np.mean(cs_bl_te)))
print('cs_tp_tr: {}'.format(np.mean(cs_tp_tr)))
print('cs_tp_te: {}'.format(np.mean(cs_tp_te)))
print('apoe_tr: {}'.format(np.mean(apoe_tr)))
print('apoe_te: {}'.format(np.mean(apoe_te)))
X_train_kf.append(X_tr)
y_train_kf.append(y_tr)
age_train_kf.append(age_tr)
apoe_train_kf.append(apoe_tr)
cs_bl_train_kf.append(cs_bl_tr)
cs_tp_train_kf.append(cs_tp_tr)
# No need to resample test data
X_test_kf.append(X_te)
y_test_kf.append(y_te)
age_test_kf.append(age_te)
apoe_test_kf.append(apoe_te)
cs_bl_test_kf.append(cs_bl_te)
cs_tp_test_kf.append(cs_tp_te)
# +
#Load AIBL data
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve, auc
cohort = 'AIBL'
tp_name = 'var_tp'
traj_csv = '/projects/nbhagwat/Traj_prediction/AIBL/exp_setup/AIBL_trajectory_labels_2class_{}_3cstp_from_m54_autoselect.csv'.format(clinical_scale)
traj_data = pd.read_csv(traj_csv)
input_csv = '/projects/nbhagwat/Traj_prediction/AIBL/exp_setup/{}_{}.csv'.format(cohort,tp_name)
in_data = pd.read_csv(input_csv)
input_traj_merge_data = pd.merge(in_data,traj_data[['RID','traj']],on='RID',how='inner')
print('merge subx: {}'.format(len(input_traj_merge_data)))
print('Traj distribution 0:{},1:{}'.format(np.sum(input_traj_merge_data['traj'].values==0),
np.sum(input_traj_merge_data['traj'].values==1)))
ct_cols = list(in_data.columns[pd.Series(in_data.columns).str.contains('CT')])
aibl_CT = np.array(input_traj_merge_data[ct_cols].values)
aibl_age = input_traj_merge_data['AGE'].values
aibl_apoe = input_traj_merge_data['APOE4'].values
aibl_cs_bl = input_traj_merge_data['MMSE_bl'].values
aibl_cs_tp = input_traj_merge_data['MMSE_m18'].values
aibl_y = np.array(input_traj_merge_data['traj'].values)
aibl_y = np.vstack((np.array(aibl_y==0).astype(int),np.array(aibl_y==1).astype(int))).T
print(aibl_CT.shape,aibl_y.shape)
#Create pairs
tp_offset = 78
X_L = aibl_CT[:,:tp_offset]
X_R = aibl_CT[:,tp_offset:]
aibl_pairs = np.concatenate((X_L[:,np.newaxis,:],X_R[:,np.newaxis,:]),axis=1)
print(aibl_te_pairs.shape)
# +
def contrastive_loss(y,d,loss_func,use_auxnet,class_weights):
if use_auxnet:
if loss_func == 'softmax':
loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y,logits=d))
elif loss_func == 'weighted_softmax':
losses = tf.nn.softmax_cross_entropy_with_logits(labels=y,logits=d)
loss = tf.reduce_mean(tf.losses.compute_weighted_loss(losses,weights=class_weights))
elif loss_func == 'hinge':
loss = tf.reduce_mean(tf.losses.hinge_loss(labels=y,logits=d))
else: #This is purly contrastive
loss1 = (1-y) * tf.square(d)
loss2 = y * tf.square(tf.maximum((1 - d),0))
loss = tf.reduce_mean(loss1 +loss2)/2
return loss
def compute_accuracy(prediction,labels,use_aux_net):
with tf.name_scope('Accuracy'):
if use_aux_net:
correct_prediction = tf.equal(tf.argmax(prediction,1), tf.argmax(labels,1))
else:
pred_labels = prediction.ravel() > 0.5
correct_prediction = tf.equal(pred_labels,labels.ravel())
acc = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
return acc
def next_batch(s,e,inputs,labels,tr_delta_t,aux_var_dict,use_aux_net,n_classes):
input1 = inputs[s:e,0]
input2 = inputs[s:e,1]
tr_batch_delta_t = np.reshape(tr_delta_t[s:e],(len(range(s,e)),1))
aux_var_batch_dict = {}
if use_aux_net:
y = np.reshape(labels[s:e],(len(range(s,e)),n_classes)) #softmax output
for aux_var in aux_var_dict:
aux_var_batch_dict[aux_var] = np.reshape(aux_var_dict[aux_var][s:e],(len(range(s,e)),1))
else:
y = np.reshape(labels[s:e],(len(range(s,e)),1))
return input1,input2,y,tr_batch_delta_t,aux_var_batch_dict
def pickleIt(my_data,save_path):
f = open(save_path, 'wb')
pickle.dump(my_data, f,protocol=2)
f.close()
def variable_summaries(var, name):
"""Attach a lot of summaries to a Tensor."""
with tf.name_scope('summaries'):
mean = tf.reduce_mean(var)
tf.scalar_summary('mean/' + name, mean)
with tf.name_scope('stddev'):
stddev = tf.sqrt(tf.reduce_mean(tf.square(var - mean)))
tf.scalar_summary('sttdev/' + name, stddev)
tf.scalar_summary('max/' + name, tf.reduce_max(var))
tf.scalar_summary('min/' + name, tf.reduce_min(var))
tf.histogram_summary(name, var)
# +
# Run models
batch_size = 100
n_folds = 1
lr = 0.005
ct_offset = 78
n_classes = 2
use_mod = False
use_delta_t = False
use_aux_net = True
# net_type = '#'mult_mod'#'dvec_mod''
model_choice = 'Siamese_Layer2_dist_mod_slim'
save_perf = True
#Tensorboard
logs_path = "/tmp/tf_logs/21"
#Track performance for each fold
kf_train_acc = []
kf_valid_acc = []
kf_test_acc = []
df_perf_concat = pd.DataFrame()
aibl_df_perf_concat = pd.DataFrame()
if use_cs:
tp_offset = ct_offset + 1
else:
tp_offset = ct_offset
loss_func = 'weighted_softmax'
class_weights_prior = [1,1]
# class_weights_prior = [0.15,0.60,0.25] #inverse weights don't work (sanitiy check)
net_arch_list = [
{'input':tp_offset,'l1':25,'l2':25,'output':10,'aux_embed':1,'aux_out':2,
'num_epochs':50,'keep_p':0.8,'reg':0.01,'dist_mod':'embed','net_type':'dvec_mod'},
]
#
start_time = time.time()
for fid in range(n_folds):
print('')
print('Foldx:{}'.format(fid+1))
df_perf = pd.DataFrame()
aibl_df_perf = pd.DataFrame()
hyp_tr_perf_list = []
hyp_va_perf_list = []
hyp_te_perf_list = []
hyp_dict_list = []
for hyp, net_arch in enumerate(net_arch_list):
num_epochs = net_arch['num_epochs']
net_type = net_arch['net_type']
#list keeping track of each batch perf
print('net_arch: {}'.format(net_arch))
train_loss = []
train_acc = []
valid_acc = []
test_acc = []
tr_dist_df = pd.DataFrame()
if use_cs: #Append CS columns appropriately
X_L = np.hstack((X_train_kf[fid][:,0].reshape(len(X_train_kf[fid]),1), X_train_kf[fid][:,2:tp_offset+1]))
X_R = np.hstack((X_train_kf[fid][:,1].reshape(len(X_train_kf[fid]),1), X_train_kf[fid][:,tp_offset+1:]))
else:
X_L = X_train_kf[fid][:,:tp_offset]
X_R = X_train_kf[fid][:,tp_offset:]
pairs = np.concatenate((X_L[:,np.newaxis,:],X_R[:,np.newaxis,:]),axis=1)
y = y_train_kf[fid]
#One-hot encoding for the labels
if use_aux_net:
y = np.vstack((np.array(y==0).astype(int),np.array(y==1).astype(int))).T
#y = tf.squeeze(slim.one_hot_encoding(y, n_classes)).eval()
#Create train-validation sets
tr_split = int(0.90*(len(y)))
tr_pairs = pairs[:tr_split,:,:]
va_pairs = pairs[tr_split:,:,:]
tr_y = y[:tr_split]
va_y = y[tr_split:]
tr_delta_t = delta_t_train_kf[fid][:tr_split].reshape(len(tr_y),1)
va_delta_t = delta_t_train_kf[fid][tr_split:].reshape(len(va_y),1)
#All the aux vars
tr_aux_var_dict = {'tr_age': age_train_kf[fid][:tr_split].reshape(len(tr_y),1),
'tr_apoe':apoe_train_kf[fid][:tr_split].reshape(len(tr_y),1),
'tr_cs_bl':cs_bl_train_kf[fid][:tr_split].reshape(len(tr_y),1),
'tr_cs_tp':cs_tp_train_kf[fid][:tr_split].reshape(len(tr_y),1)}
va_age = age_train_kf[fid][tr_split:].reshape(len(va_y),1)
va_apoe = apoe_train_kf[fid][tr_split:].reshape(len(va_y),1)
va_cs_bl = cs_bl_train_kf[fid][tr_split:].reshape(len(va_y),1)
va_cs_tp = cs_tp_train_kf[fid][tr_split:].reshape(len(va_y),1)
print(pairs.shape, tr_split, len(y))
if use_cs:
X_L = np.hstack((X_test_kf[fid][:,0].reshape(len(X_test_kf[fid]),1), X_test_kf[fid][:,2:tp_offset+1]))
X_R = np.hstack((X_test_kf[fid][:,1].reshape(len(X_test_kf[fid]),1), X_test_kf[fid][:,tp_offset+1:]))
else:
X_L = X_test_kf[fid][:,:tp_offset]
X_R = X_test_kf[fid][:,tp_offset:]
te_pairs = np.concatenate((X_L[:,np.newaxis,:],X_R[:,np.newaxis,:]),axis=1)
te_y = y_test_kf[fid]
#One-hot encoding for the labels
if use_aux_net:
te_y = np.vstack((np.array(te_y==0).astype(int),np.array(te_y==1).astype(int))).T
#te_y = tf.squeeze(slim.one_hot_encoding(te_y, n_classes)).eval()
te_delta_t = delta_t_test_kf[fid].reshape(len(te_y),1)
te_age = age_test_kf[fid].reshape(len(te_y),1)
te_apoe = apoe_test_kf[fid].reshape(len(te_y),1)
te_cs_bl = cs_bl_test_kf[fid].reshape(len(te_y),1)
te_cs_tp = cs_tp_test_kf[fid].reshape(len(te_y),1)
print('pair_shapes, tr:{},va:{},te:{},label shapes, tr:{},va:{},te:{}'.format(tr_pairs.shape,va_pairs.shape,
te_pairs.shape,tr_y.shape,
va_y.shape,te_y.shape))
# Define input and model tensors + optimizer
with tf.Graph().as_default():
images_L = tf.placeholder(tf.float32,shape=([None,net_arch['input']]),name='Baseline_CT')
images_R = tf.placeholder(tf.float32,shape=([None,net_arch['input']]),name='Timepoint_2_CT')
if use_aux_net:
labels = tf.placeholder(tf.float32,shape=([None,n_classes]),name='labels')
class_weights = tf.placeholder(tf.float32,shape=([None]),name='class_weights')
else:
labels = tf.placeholder(tf.float32,shape=([None,1]),name='gt')
delta_t = tf.placeholder(tf.float32,shape=([None,1]),name='dt') #Time between two scans
age = tf.placeholder(tf.float32,shape=([None,1]),name='age') #Time between two scans
apoe = tf.placeholder(tf.float32,shape=([None,1]),name='apoe') #Time between two scans
cs_bl = tf.placeholder(tf.float32,shape=([None,1]),name='cs_bl') #Time between two scans
cs_tp = tf.placeholder(tf.float32,shape=([None,1]),name='cs_tp') #Time between two scans
is_training = tf.placeholder(tf.bool, name='is_training') #for batch normalization
with tf.variable_scope("siamese") as scope:
model1, nodes = mlp_model(images_L, net_arch, is_training, net_type='mlpnet')
scope.reuse_variables()
model2, nodes = mlp_model(images_R, net_arch, is_training, net_type='mlpnet')
#Compute distance embeddings
# distance_vec = tf.div(tf.subtract(model1,model2),tf.norm(tf.subtract(model1,model2)))
distance_vec = tf.concat([model1,model2],1)
#distance_vec = tf.subtract(model1,model2)
if use_delta_t: #encoded in months
#distance = 1.0 - tf.exp(-1*(tf.div(tf.reduce_sum(tf.pow(tf.subtract(model1,model2),2),1,keep_dims=True),delta_t)))
distance = 1.0 - tf.exp(-1*tf.div(tf.reduce_sum(tf.pow(tf.subtract(model1,model2),2),1,keep_dims=True),delta_t))
else:
distance = 1.0 - tf.exp(-1*tf.reduce_sum(tf.pow(tf.subtract(model1,model2),2),1,keep_dims=True))
if use_aux_net:
#Add other nodes to incorporate age, apoe4, time, mmse etc.
cs_data = tf.concat([cs_bl,cs_tp],1)
attr_list = [tf.add(1.0,apoe)]
attr_data = tf.concat(attr_list,1)
if net_type == 'dvec_mod':
mixed_data = {'d':distance_vec, 'cs':cs_data, 'attr':attr_data}
elif net_type == 'mult_mod':
mixed_data = {'d':distance, 'cs':cs_data, 'attr':attr_data}
else:
print('Unknow auxnet type: {}'.format(net_type))
model3, aux_nodes = mlp_model(mixed_data , net_arch,is_training, net_type=net_type)
with tf.name_scope('loss'):
loss = contrastive_loss(labels,model3,loss_func,use_aux_net,class_weights)
else:
loss = contrastive_loss(labels,distance,loss_func,use_aux_net)
#Tensorboard
# create a summary for our cost and accuracy
tf.summary.scalar("loss", loss)
tf.summary.histogram("distance", distance)
tf.summary.histogram("dmod", aux_nodes['d_mod'])
tf.summary.histogram("distance_vec", distance_vec)
#if you want to look at variables
t_vars = tf.trainable_variables()
d_vars = [var for var in t_vars if 'fc' in var.name]
# for dv in t_vars:
# tf.summary.histogram(dv.name, dv)
tf.contrib.layers.summarize_collection(tf.GraphKeys.TRAINABLE_VARIABLES)
optimizer = tf.train.AdamOptimizer(learning_rate = lr).minimize(loss)
#optimizer = tf.train.RMSPropOptimizer(0.0001,momentum=0.9,epsilon=1e-6).minimize(loss)
if fid==0:
# Print name and shape of each tensor.
print("Layers")
for k, v in nodes.items():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# Print name and shape of parameter nodes (values not yet initialized)
print("Parameters")
for v in slim.get_model_variables():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
#Tensorboard
summary_op = tf.summary.merge_all()
# create log writer object
train_writer = tf.summary.FileWriter(logs_path + '/train', graph=tf.get_default_graph())
test_writer = tf.summary.FileWriter(logs_path + '/test', graph=tf.get_default_graph())
# Launch the graph
with tf.Session(config=tf.ConfigProto(log_device_placement=False)) as sess:
tf.global_variables_initializer().run()
#initialize_all_variables().run()
# Training cycle
for epoch in range(num_epochs):
avg_loss = 0.
avg_acc = 0.
total_batch = int(tr_pairs.shape[0]/batch_size)
# Loop over all batches
for i in range(total_batch):
s = i * batch_size
e = (i+1) *batch_size
# Fit training using batch data
input1,input2,y,tr_batch_delta_t,tr_aux_var_batch_dict = next_batch(s,e,tr_pairs,tr_y,
tr_delta_t,
tr_aux_var_dict,
use_aux_net,n_classes)
class_weights_array = np.ones(len(y))
for c in range(len(class_weights_prior)):
class_weights_array[np.argmax(y,axis=1)==c] = class_weights_prior[c]
if use_aux_net:
tr_batch_age = tr_aux_var_batch_dict['tr_age']
tr_batch_apoe = tr_aux_var_batch_dict['tr_apoe']
tr_batch_cs_bl = tr_aux_var_batch_dict['tr_cs_bl']
tr_batch_cs_tp = tr_aux_var_batch_dict['tr_cs_tp']
_,loss_value,predict,tr_dist_batch,summary = sess.run([optimizer,loss,model3,
distance,summary_op],
feed_dict={images_L:input1,
images_R:input2,labels:y,
delta_t: tr_batch_delta_t,age:tr_batch_age,
apoe: tr_batch_apoe,cs_bl: tr_batch_cs_bl,
cs_tp: tr_batch_cs_tp,
class_weights: class_weights_array,
is_training:True})
else:
_,loss_value,predict = sess.run([optimizer,loss,distance],
feed_dict={images_L:input1,images_R:input2,labels:y,
delta_t: tr_batch_delta_t, is_training:True})
tr_acc = compute_accuracy(predict,y,use_aux_net).eval()
avg_loss += loss_value
avg_acc +=tr_acc*100
#Tensorboard
train_writer.add_summary(summary, epoch * total_batch + i)
train_loss.append(avg_loss/(total_batch))
train_acc.append(avg_acc/total_batch)
tr_d_mod = aux_nodes['d_mod'].eval({images_L:va_pairs[:,0],images_R:va_pairs[:,1],labels:y,
delta_t:va_delta_t, age:va_age, apoe: va_apoe,
cs_bl: va_cs_bl,cs_tp: va_cs_tp, is_training:False})
tr_dist = aux_nodes['dist'].eval({images_L:va_pairs[:,0],images_R:va_pairs[:,1],labels:y,
delta_t:va_delta_t, age:va_age, apoe: va_apoe,
cs_bl: va_cs_bl,cs_tp: va_cs_tp, is_training:False})
#One-hot encoding for the labels
if use_aux_net:
y = np.reshape(va_y,(va_y.shape[0],n_classes))
else:
y = np.reshape(va_y,(va_y.shape[0],1))
class_weights_array = np.ones(len(y))
for c in range(len(class_weights_prior)):
class_weights_array[np.argmax(y,axis=1)==c] = class_weights_prior[c]
if use_aux_net:
feature3,summary=sess.run([model3,summary_op],feed_dict={images_L:va_pairs[:,0],images_R:va_pairs[:,1],labels:y,
delta_t:va_delta_t, age:va_age, apoe: va_apoe,
cs_bl: va_cs_bl,cs_tp: va_cs_tp,
class_weights: class_weights_array, is_training:False})
va_acc = compute_accuracy(feature3,y,use_aux_net).eval()*100
else:
va_acc = compute_accuracy(predict,y,use_aux_net).eval()*100
valid_acc.append(va_acc)
#Tensorboard
test_writer.add_summary(summary, epoch*total_batch)
if epoch%10 == 0:
print('Epoch:{}, Accuracy train:{:4.3f},valid:{:4.3f}'.format(epoch,avg_acc/total_batch,va_acc))
#print('Epoch:{}, Accuracy train:{:4.3f},valid:{:4.3f}'.format(epoch,tr_acc,va_acc))
print('distance:{:4.3f},{:4.3f}<->{:4.3f}, loss:{:4.3f}'.format(np.mean(tr_dist),np.min(tr_dist),
np.max(tr_dist),avg_loss/(total_batch)))
print('dmod:{:4.3f},{:4.3f}<->{:4.3f}'.format(np.mean(tr_d_mod),np.min(tr_d_mod),np.max(tr_d_mod)))
#Accuracy of the entire training set at the end of training
if use_aux_net:
y = np.reshape(tr_y,(tr_y.shape[0],n_classes))
else:
y = np.reshape(tr_y,(tr_y.shape[0],1))
if use_aux_net:
feature3,predict=sess.run([model3,distance],feed_dict={images_L:tr_pairs[:,0],images_R:tr_pairs[:,1],labels:y,
delta_t:tr_delta_t,age:tr_aux_var_dict['tr_age'],
apoe: tr_aux_var_dict['tr_apoe'],cs_bl: tr_aux_var_dict['tr_cs_bl'],
cs_tp: tr_aux_var_dict['tr_cs_tp'], is_training:False})
tr_acc = compute_accuracy(feature3,y,use_aux_net).eval()
tr_dist_df['label']= np.argmax(y,1)
else:
tr_acc = compute_accuracy(predict,y,use_aux_net).eval()
tr_dist_df['label']= np.squeeze(np.concatenate(y))
tr_dist_df['distance'] = np.squeeze(predict)
tr_dist_df['d_mod'] = aux_nodes['d_mod'].eval(feed_dict={images_L:tr_pairs[:,0],images_R:tr_pairs[:,1],labels:y,
delta_t:tr_delta_t,age:tr_aux_var_dict['tr_age'],
apoe: tr_aux_var_dict['tr_apoe'],cs_bl: tr_aux_var_dict['tr_cs_bl'],
cs_tp: tr_aux_var_dict['tr_cs_tp'], is_training:False})
print('Accuracy training set %0.2f' % (100 * tr_acc))
print('Accuracy validation set %0.2f' % (va_acc))
# Evaluate Test set after training
if use_aux_net:
y = np.reshape(te_y,(te_y.shape[0],n_classes))
else:
y = np.reshape(te_y,(te_y.shape[0],1))
if use_aux_net:
feature3, predict= sess.run([model3,distance],
feed_dict={images_L:te_pairs[:,0],
images_R:te_pairs[:,1],labels:y,
delta_t:te_delta_t, age:te_age, apoe:te_apoe,
cs_bl:te_cs_bl, cs_tp:te_cs_tp, is_training:False})
te_acc = compute_accuracy(feature3,y,use_aux_net).eval()
else:
te_acc = compute_accuracy(predict,y,use_aux_net).eval()
test_acc.append(te_acc)
print('Accuracy test set %0.2f' % (100 * te_acc))
print('')
####################################################################
# Test model on AIBL
feature3, predict= sess.run([model3,distance],
feed_dict={images_L:aibl_pairs[:,0],
images_R:aibl_pairs[:,1],labels:aibl_y,
age:aibl_age, apoe:aibl_apoe,
cs_bl:aibl_cs_bl, cs_tp:aibl_cs_tp, is_training:False})
aibl_acc = compute_accuracy(feature3,aibl_y,use_aux_net).eval()
print('Accuracy AIBL test set %0.3f' % (100 * aibl_acc))
# Save AIBL perf
aibl_df_perf['model'] = np.tile(model_choice,len(aibl_y))
aibl_df_perf['kf'] = np.tile(kf+1,len(aibl_y))
aibl_df_perf['act_label'] = np.argmax(aibl_y,1)
aibl_df_perf['pred_label'] = np.argmax(feature3,1)
aibl_df_perf['pred_prob'] = list(tf.nn.softmax(feature3).eval())
aibl_df_perf_concat = aibl_df_perf_concat.append(aibl_df_perf)
####################################################################
#prediction weight
trained_wts = [var for var in t_vars if 'mlp/aux_prediction/weights:0' in var.name]
trained_biases = [var for var in t_vars if 'mlp/aux_prediction/BatchNorm/beta' in var.name]
pred_wt = trained_wts[0].eval()
# pred_b = trained_biases[0].eval()
print('pred weight {}'.format(pred_wt))
# Model embeddings: for t-SNE
train_feature_1 = model1.eval(feed_dict={images_L:tr_pairs[:,0],is_training:False})
train_feature_2 = model2.eval(feed_dict={images_R:tr_pairs[:,1],is_training:False})
test_feature_1 = model1.eval(feed_dict={images_L:te_pairs[:,0],is_training:False})
test_feature_2 = model2.eval(feed_dict={images_R:te_pairs[:,1],is_training:False})
hyp_dict = {}
hyp_tr_perf_list.append(tr_acc)
hyp_va_perf_list.append(va_acc)
hyp_te_perf_list.append(te_acc)
hyp_dict['hyp'] = hyp
hyp_dict['va_acc'] = va_acc
hyp_dict['te_acc'] = te_acc
hyp_dict['distance'] = predict
hyp_dict['d_mod'] = aux_nodes['d_mod'].eval(feed_dict={images_L:te_pairs[:,0],
images_R:te_pairs[:,1],labels:y,
delta_t:te_delta_t, age:te_age, apoe:te_apoe,
cs_bl:te_cs_bl, cs_tp:te_cs_tp, is_training:False})
if use_aux_net:
hyp_dict['pred_label'] = np.argmax(tf.nn.softmax(feature3).eval(),1)
hyp_dict['pred_prob'] = list(tf.nn.softmax(feature3).eval())
hyp_dict['pred_wt'] = [pred_wt]
else:
hyp_dict['pred_label'] = (predict.ravel() > 0.5).astype(int)
a = np.squeeze(np.minimum(predict,np.ones((len(predict),1))))
b = np.squeeze(1-a)
y_pred_prob = np.vstack((b,a)).T
hyp_dict['pred_prob'] = list(y_pred_prob)
hyp_dict_list.append(hyp_dict)
## Pick the best hyper-param based on va_acc
opt_hyp = np.argmax(np.array(hyp_te_perf_list))
opt_hyp_dict = hyp_dict_list[opt_hyp]
print('opt_hyp:{}, hyp_va_perf:{},hyp_te_perf:{}'.format(opt_hyp,hyp_va_perf_list,hyp_te_perf_list))
#Save perf method 1: df of all subjects
df_perf['PTID'] = Sub_test_kf[fid]
df_perf['model'] = np.tile(model_choice,len(te_y))
df_perf['kf'] = np.tile(fid+1,len(te_y))
df_perf['distance'] = opt_hyp_dict['distance']
df_perf['d_mod'] = opt_hyp_dict['d_mod']
if use_aux_net:
df_perf['act_label'] = np.argmax(te_y,1)
df_perf['pred_label'] = opt_hyp_dict['pred_label'] #np.argmax(tf.nn.softmax(feature3).eval(),1)
df_perf['pred_prob'] = opt_hyp_dict['pred_prob'] #list(tf.nn.softmax(feature3).eval())
df_perf['pred_wt'] = opt_hyp_dict['pred_wt']*len(te_y)
else:
df_perf['act_label'] = te_y
df_perf['pred_label'] = opt_hyp_dict['pred_label'] #(predict.ravel() > 0.5).astype(int)
df_perf['pred_prob'] = opt_hyp_dict['pred_prob'] #list(y_pred_prob)
#df_perf['test_embed_L'] = list(test_feature_1) #commented to test out simpler hyp search routine
#df_perf['test_embed_R'] = list(test_feature_2)
df_perf_concat = df_perf_concat.append(df_perf)
kf_train_acc.append(hyp_tr_perf_list[opt_hyp])
kf_valid_acc.append(hyp_va_perf_list[opt_hyp])
kf_test_acc.append(hyp_te_perf_list[opt_hyp])
#Save perf df
if save_perf:
pickleIt(df_perf_concat, '{}TF_perf/{}_feat{}_{}_df_perf.pkl'.format(baseline_dir,exp_name,feat_sufix,model_choice))
pickleIt(aibl_df_perf_concat, '{}TF_perf/aibl_{}_feat{}_{}_df_perf.pkl'.format(baseline_dir,exp_name,feat_sufix,model_choice))
#df_perf_concat.to_pickle('{}TF_perf/{}_feat{}_{}_df_perf.pkl'.format(baseline_dir,exp_name,feat_sufix,model_choice))
print('total_time:: {:3.2f}'.format((time.time() - start_time)/60.0))
# +
#K-fold stats
print(kf_train_acc)
print(kf_valid_acc)
print(kf_test_acc)
print(np.mean(kf_train_acc),np.mean(kf_valid_acc),np.mean(kf_test_acc))
#pickleIt(df_perf_concat, '{}TF_perf/{}_feat{}_{}_df_perf.pkl'.format(baseline_dir,exp_name,feat_sufix,model_choice))
#Use delta_t, No CS, No_age_apoe4
#No delta_t, use CS, No_age_apoe4
# [0.94759828, 0.9323144, 0.92139739, 0.91703057, 0.9323144, 0.90829694, 0.92139739, 0.93246186, 0.9237473, 0.9368192]
# [82.352942228317261, 80.392158031463623, 84.313726425170898, 84.313726425170898, 86.274510622024536, 82.352942228317261, 80.392158031463623, 80.392158031463623, 84.313726425170898, 90.38461446762085]
# [0.85964912, 0.84210527, 0.84210527, 0.80701756, 0.7719298, 0.82456142, 0.85964912, 0.83928573, 0.85714287, 0.80000001]
# 0.927338 83.5482662916 0.830345
#Use delta_t, use CS, No_age_apoe4
# [0.93449783, 0.93449783, 0.93013102, 0.94759828, 0.94323146, 0.93449783, 0.94104803, 0.95424837, 0.92810458, 0.94335514]
# [82.352942228317261, 84.313726425170898, 82.352942228317261, 84.313726425170898, 80.392158031463623, 90.196079015731812, 86.274510622024536, 82.352942228317261, 86.274510622024536, 88.461536169052124]
# [0.85964912, 0.89473683, 0.80701756, 0.84210527, 0.7719298, 0.89473683, 0.82456142, 0.9285714, 0.875, 0.81818181]
# 0.939121 84.7285073996 0.851649
#Use delta_t, use CS, use_age_apoe4
# [0.84934503, 0.90611351, 0.89301312, 0.88864625, 0.89082968, 0.86899567, 0.86899567, 0.88235289, 0.8562091, 0.90196079]
# [88.235300779342651, 82.352948188781738, 80.392163991928101, 88.235300779342651, 84.313732385635376, 82.352948188781738, 84.313732385635376, 76.470595598220825, 86.274516582489014, 90.384620428085327]
# [0.84210527, 0.87719297, 0.84210527, 0.87719297, 0.85964912, 0.84210527, 0.82456142, 0.83928573, 0.91071433, 0.87272727]
# 0.880646 84.3325859308 0.858764
df_perf_concat.head(5)
# +
# Plot Acc and Loss
print(np.mean(np.array(test_acc)))
print(test_acc)
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.plot(train_loss, 'g-')
ax2.plot(train_acc, 'b-')
ax2.plot(valid_acc, 'r-')
ax1.set_xlabel('epoch')
ax1.set_ylabel('loss', color='g')
ax2.set_ylabel('acc', color='b')
ax1.grid()
# +
#TSNE
from sklearn.manifold import TSNE
tsne_df = pd.DataFrame()
tsne_df_train = pd.DataFrame()
tsne_df_train['PTID'] = Sub_train_kf[fid][:tr_split]
tsne_df_train['subset'] = np.tile('train',len(tr_y))
tsne_df_train['KF'] = np.tile(fid,len(tr_y))
tsne_df_test = pd.DataFrame()
tsne_df_test['PTID'] = Sub_test_kf[fid]
tsne_df_test['subset'] = np.tile('test',len(te_y))
tsne_df_test['KF'] = np.tile(fid,len(te_y))
if use_aux_net:
tsne_df_train['labels'] = np.argmax(tr_y,1)
tsne_df_test['labels'] = np.argmax(te_y,1)
else:
tsne_df_train['labels'] = tr_y
tsne_df_test['labels'] = te_y
tsne_train_layers = {'l1':train_feature_1,'l2':train_feature_2}
tsne_test_layers = {'l1':test_feature_1,'l2':test_feature_2}
for l in tsne_train_layers:
tsne = TSNE(n_components=2, random_state=0,init='pca')
tsne_embed = tsne.fit_transform(tsne_train_layers[l])
tsne_df_train['TSNE_{}_x'.format(l)] = tsne_embed[:,0]
tsne_df_train['TSNE_{}_y'.format(l)] = tsne_embed[:,1]
for l in tsne_test_layers:
tsne = TSNE(n_components=2, random_state=0,init='pca')
tsne_embed = tsne.fit_transform(tsne_test_layers[l])
tsne_df_test['TSNE_{}_x'.format(l)] = tsne_embed[:,0]
tsne_df_test['TSNE_{}_y'.format(l)] = tsne_embed[:,1]
tsne_df = tsne_df_train.append(tsne_df_test)
print(len(tsne_df))
tsne_df = pd.merge(tsne_df,df[['PTID','ORIGPROT']],on='PTID')
print(len(tsne_df))
# -
flatui = ["#9b59b6", "#3498db", "#e74c3c", "#34495e", "#2ecc71", "#95a5a6"]
plot_df = tsne_df[tsne_df['subset']=='train']
with sns.axes_style("whitegrid"):
g1 = sns.lmplot('TSNE_l1_x','TSNE_l1_y',fit_reg=False,hue='labels',col='subset',data=plot_df,col_wrap=2,
markers='o',size=6,sharey=False,sharex=False,palette=flatui)
g1 = sns.lmplot('TSNE_l2_x','TSNE_l2_y',fit_reg=False,hue='labels',col='subset',data=plot_df,col_wrap=2,
markers='o',size=6,sharey=False,sharex=False,palette=flatui)
# +
#Plot embeddings
f = plt.figure(figsize=(16,9))
c = ['#ff0000', '#0000ff', '#ffff00', '#00ff00', '#009999',
'#ff00ff', '#00ffff', '#990000', '#999900', '#009900' ]
mark_size= 10
plt.subplot(1,2,1)
x = train_feature_1[:,0]
y = train_feature_1[:,1]
labels = tr_y[:,0] #left_digits
for i in range(2):
plt.plot(x[labels==i],y[labels==i],'.', c=c[i], markersize=mark_size)
plt.grid()
plt.title('Left side')
plt.subplot(1,2,2)
x = train_feature_2[:,0]
y = train_feature_2[:,1]
for i in range(2):
plt.plot(x[labels==i],y[labels==i],'.', c=c[i], markersize=mark_size)
plt.grid()
plt.title('Right side')
# -
sns.factorplot(y='distance',x='label',data=tr_dist_df,kind='box')
sns.factorplot(y='d_mod',x='label',data=tr_dist_df,kind='box')
f = plt.figure(figsize=(16,9))
a = tr_dist_df[tr_dist_df['label']==0]['distance'].values
b = tr_dist_df[tr_dist_df['label']==1]['distance'].values
plt.subplot(1,2,1)
plt.hist(a,bins=50,normed=1,facecolor='red',alpha=0.5,label='Traj_0')
plt.hist(b,bins=50,normed=1,facecolor='blue',alpha=0.5,label='Traj_1')
plt.legend()
plt.subplot(1,2,2)
a = tr_dist_df[tr_dist_df['label']==0]['d_mod'].values
b = tr_dist_df[tr_dist_df['label']==1]['d_mod'].values
plt.hist(a,bins=50,normed=1,facecolor='red',alpha=0.5,label='Traj_0')
plt.hist(b,bins=50,normed=1,facecolor='blue',alpha=0.5,label='Traj_1')
plt.legend()
# +
## Test code for TF slim
# +
n_classes = 5
net_arch = {'l1':50,'l2':50,'l3':8,'output':n_classes,'keep_p':0.8}
with tf.Graph().as_default():
# Dummy placeholders for arbitrary number of 1d inputs and outputs
inputs = tf.placeholder(tf.float32, shape=(None, 1))
outputs = tf.placeholder(tf.float32, shape=(None, n_classes))
# Build model
predictions, end_points = mlp_model(inputs,net_arch=net_arch)
# Print name and shape of each tensor.
print("Layers")
for k, v in end_points.items():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# Print name and shape of parameter nodes (values not yet initialized)
print("\n")
print("Parameters")
for v in slim.get_model_variables():
print('name = {}, shape = {}'.format(v.name, v.get_shape()))
# batch data
x_train, y_train = produce_batch(200)
x_test, y_test = produce_batch(200)
print(np.shape(y_train),np.shape(y_test))
plt.scatter(x_train, y_train[:,0])
# +
# The following snippet trains the regression model using a mean_squared_error loss.
ckpt_dir = '/tmp/tfslim_model_test2/'
with tf.Graph().as_default():
tf.logging.set_verbosity(tf.logging.INFO)
print(x_train.shape,y_train.shape)
inputs, targets = convert_data_to_tensors(x_train, y_train)
# Make the model.
predictions, nodes = mlp_model(inputs, is_training=True, net_arch=net_arch)
# Add the loss function to the graph.
#loss = tf.losses.mean_squared_error(labels=targets, predictions=predictions)
one_hot_labels = tf.squeeze(slim.one_hot_encoding(y_train, n_classes))
slim.losses.softmax_cross_entropy(predictions, one_hot_labels)
# The total loss is the uers's loss plus any regularization losses.
total_loss = slim.losses.get_total_loss()
# Create some summaries to visualize the training process:
tf.summary.scalar('losses/Total_Loss', total_loss)
# Specify the optimizer and create the train op:
optimizer = tf.train.AdamOptimizer(learning_rate=0.005)
train_op = slim.learning.create_train_op(total_loss, optimizer)
# Run the training inside a session.
final_loss = slim.learning.train(
train_op,
logdir=ckpt_dir,
number_of_steps=5000,
save_summaries_secs=5,
log_every_n_steps=500)
print("Finished training. Last batch loss:", final_loss)
print("Checkpoint saved in %s" % ckpt_dir)
# +
with tf.Graph().as_default():
inputs, targets = convert_data_to_tensors(x_test, y_test)
# Create the model structure. (Parameters will be loaded below.)
predictions, end_points = mlp_model(inputs, is_training=False, net_arch=net_arch)
one_hot_labels = tf.squeeze(slim.one_hot_encoding(y_test, n_classes))
# Make a session which restores the old parameters from a checkpoint.
sv = tf.train.Supervisor(logdir=ckpt_dir)
with sv.managed_session() as sess:
inputs, predictions, targets = sess.run([inputs, predictions, one_hot_labels])
print(np.squeeze(inputs).shape,np.argmax(targets,1).shape,np.argmax(predictions,1).shape)
#plt.scatter(np.squeeze(inputs), np.argmax(targets,1), c='r',alpha=0.5);
plt.scatter(np.squeeze(inputs), np.argmax(predictions,1), c='b',alpha=0.5);
plt.title('red=true, blue=predicted')
# -
| legacy_code/TF_siamese_adni_slim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="vg-HOITYJAy0"
# #### Основы программирования в Python для социальных наук
#
# *Автор: <NAME>, НИУ ВШЭ*
#
#
# ## Cамостоятельная работа 3
#
# ## Вариант 1
#
# Самостоятельная работа по темам:
#
# * Web-scraping
#
# **Списывание и использование телефонов**
#
# Первое предупреждение, сделанное ассистентом или преподавателем: - 1 балл к вашей оценке.
#
# Второе предупреждение: работа аннулируется без права переписывания.
#
# **Все задачи, где не указано иного, должны принимать значения на ввод (решение для частного случая будет считаться неполным).**
# + [markdown] colab_type="text" id="S3g4tDtrMoxX"
# **Задание 1.**
# *5 баллов*
#
# 1. На странице в https://en.wikipedia.org/wiki/List_of_Rick_and_Morty_episodes нужно найти таблицу под названием "Season 1 (2013–14)".
# 2. С помощью поиска по тегам, нужно сохранить из таблицы следующие колонки: 'Title', 'Directed By', 'U.S. viewers (millions)'. Каждая колонка таблицы должна быть сохранена в отдельную переменную (см. ниже названия), внутри которой лежит список, где первое значение - название колонки.
#
# Обратите внимание, положение элемента в ряде с table headers (th) и в обыкновенных рядах (теги td) может не совпадать.
#
# Например, колонки 'Title' список будет выглядеть так:
# ['Title', "Pilot", ...остальные значения..., "Ricksy Business"]
#
# Значения для 'U.S. viewers (millions)' на этом этапе могут содержать ссылки (например, '1.10[7]')
#
# 3. Выведите эти три списка командой
# print(titles)
# print(directors)
# print(viewers)
# + colab={} colab_type="code" id="KRw05f_FKOXQ"
import requests
from bs4 import BeautifulSoup
website_url = requests.get('https://en.wikipedia.org/wiki/List_of_Rick_and_Morty_episodes').text
soup = BeautifulSoup(website_url,'lxml')
My_table = soup.find_all('table',{'class':'wikitable plainrowheaders wikiepisodetable'})
rows = My_table[0].find_all('tr')
Title = []
Title.append(rows[0].find_all('th')[2].get_text().strip())
for row in rows[1:]:
r = row.find_all('td')
Title.append(r[1].get_text().strip())
Directed = []
Directed.append(rows[0].find_all('th')[3].get_text().strip())
for row in rows[1:]:
r = row.find_all('td')
Directed.append(r[2].get_text().strip())
Viewers = []
Viewers.append(rows[0].find_all('th')[6].get_text().strip())
for row in rows[1:]:
r = row.find_all('td')
value = r[5].get_text().strip()
values = value.split('[')
Viewers.append(float(values[0]))
print(Title)
print(Directed)
print(Viewers)
# + [markdown] colab_type="text" id="sZFN--3fNTt2"
# **Задание 2.**
# *5 баллов (каждый шаг 1 балл)*
#
# 1. Из списка viewers удалите то, что в wiki было ссылками (номера в квадратных строках) и переведите значения во float.
# Правильный список viewers будет выглядеть так:
# ['U.S. viewers(millions)', 1.1 ...]
#
# 2. Создайте словарь season1, в котором ключ - название эпизода, а значение - список из имени режиссера, количества зрителей (в формате float), и номера эпизода (не учитывая названия колонок в нулевом элементе). Например, пара ключ-значение для первого эпизода будет выглядеть так
# `'"Pilot"': ['<NAME>', 1.1, 1]`
# 2. Напишите функцию, которая берет аргументом название эпизода и печатает следующую строку: {номер эпизода} Episode {название эпизода} (directed by {имя режиссера} was watched by {количество зрителей} million viewers in the US.
#
# 3. Запустите вашу функцию в цикле для всех эпизодов (не учитывая названия колонки в нулевом элементе).
#
# 5. Сколько всего зрителей посмотрело эпизоды, которые режиссировал <NAME>?
# +
# Запустите эту ячейку
titles = ['Title', '"Pilot"', '"Lawnmower Dog"', '"Anatomy Park"', '"M. Night Shaym-Aliens!"', '"Meeseeks and Destroy"', '"Rick Potion #9"', '"Raising Gazorpazorp"', '"Rixty Minutes"', '"Something Ricked This Way Comes"', '"Close Rick-counters of the Rick Kind"', '"Ricksy Business"']
viewers = ['U.S. viewers(millions)', '1.10[5]', '1.51[6]', '1.30[7]', '1.32[8]', '1.61[9]', '1.75[10]', '1.76[11]', '1.48[12]', '1.54[13]', '1.75[14]', '2.13[15]']
directors = ['Directed by', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>', '<NAME>']
print(titles)
print(viewers)
print(directors)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="pwLxGVugPVys" outputId="0eb3382d-eba5-4e68-8edb-2b3e15a57ac0"
# 1
new_viewers = []
new_viewers.append(viewers[0])
for i in viewers[1:]:
value = i.split('[')
new_viewers.append(float(value[0]))
new_viewers
# +
# 2
season1 = {}
for idx, value in enumerate(titles[1:]):
season1[value] = [Directed[idx+1], Viewers[idx+1], idx+1]
season1
# -
# 3
def episodes_info(episode):
for idx, value in enumerate(titles[1:]):
if value == episode:
number = idx+1
director = Directed[idx+1]
viewers = Viewers[idx+1]
print(f'{number} Episode {episode} directed by {director} was watched by {viewers} million viewers in the US')
#4
for episode in Title[1:]:
episodes_info(episode)
# +
# 5
summ = 0
for idx, value in enumerate(directors[1:]):
if value == '<NAME>':
summ += Viewers[idx+1]
summ
| web_scraping_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
import nltk
from nltk.stem import PorterStemmer
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
# import the data
df = pd.read_csv('mail_data.csv')
df.head()
# shape of the dataset
df.shape
# checking the distribution of target vriable
df['Category'].value_counts()
# checking for any missing values
df.isnull().sum()
# Cleaning the dataset
stemmer = PorterStemmer()
corpus = []
for i in range(0 , len(df)):
review = re.sub('[^a-zA-Z]', ' ', df['Message'][i])
review = review.lower()
review = review.split()
review = [stemmer.stem(word) for word in review if word not in set(stopwords.words('english'))]
review = ' '.join(review)
corpus.append(review)
# Vectorizing the dataset and creating the bag of words
vect = TfidfVectorizer()
X = vect.fit_transform(corpus).toarray()
# converting the textual target to numerical target
df.replace({'Category' : {'ham' : 0, 'spam' : 1 }}, inplace = True)
Y = df['Category']
print(X)
print(Y)
# train test split
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, stratify = Y, random_state =2)
# training model using naive bayes classifier
model = MultinomialNB()
model.fit(x_train, y_train)
y_pred = model.predict(x_test)
from sklearn.metrics import confusion_matrix
matrix = confusion_matrix(y_test, y_pred)
matrix
from sklearn.metrics import accuracy_score
accuracy = accuracy_score(y_test, y_pred)
print('ACCURACY IS :',accuracy)
| Spam classifier (with data cleaning).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="FQiOKlk03GgY" colab_type="text"
# # DCGAN: Generate the images with Deep Convolutional GAN
# Note: This notebook is created from [chainer/examples/dcgan](https://github.com/chainer/chainer/tree/master/examples/dcgan). If you want to run it as script, please refer to the above link.
#
# In this notebook, we generate images with **generative adversarial network (GAN)**.
#
# 
# + [markdown] id="UC-m7Hwh4Y7Z" colab_type="text"
# First, we execute the following cell and install "Chainer" and its GPU back end "CuPy". If the "runtime type" of Colaboratory is GPU, you can run Chainer with GPU as a backend.
# + id="vctT_8EWSEOJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} executionInfo={"status": "ok", "timestamp": 1593380371577, "user_tz": -120, "elapsed": 37670, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="41aab673-e5c8-43e8-90a9-d00f3fddc3c3"
# !curl https://colab.chainer.org/install | sh -
# + [markdown] id="cH8mST0B5IK2" colab_type="text"
# Let's import the necessary modules, then check the version of Chainer, NumPy, CuPy, Cuda and other execution environments.
# + id="v00bch6E5Gf6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} executionInfo={"status": "ok", "timestamp": 1593380376377, "user_tz": -120, "elapsed": 33286, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="1560b8c6-a34a-49bd-e4be-1e15b7a34ffe"
import os
import numpy as np
import chainer
from chainer import cuda
import chainer.functions as F
import chainer.links as L
from chainer import Variable
from chainer.training import extensions
chainer.print_runtime_info()
# + [markdown] id="kjBUS6Cr8Sye" colab_type="text"
# ## 1. Setting parameters
# Here we set the parameters for training.
#
# * `` n_epoch``: Epoch number. How many times we pass through the whole training data.
# * `` n_units``: Number of units. How many hidden state vectors each Recursive Neural Network node has.
# * `` batchsize``: Batch size. How many train data we will input as a block when updating parameters.
# * `` n_label``: Number of labels. Number of classes to be identified. Since there are 5 labels this time, `` 5``.
# * `` epoch_per_eval``: How often to perform validation.
# * `` is_test``: If `` True``, we use a small dataset.
# * `` gpu_id``: GPU ID. The ID of the GPU to use. For Colaboratory it is good to use `` 0``.
# + id="1Ha6-ramShVF" colab_type="code" colab={}
# parameters
n_epoch = 3 # number of epochs, ideally >50, but we can write down only 1 to test the notebook
n_hidden = 100 # number of hidden units
batchsize = 50 # minibatch size
snapshot_interval = 10000 # number of iterations per snapshots
display_interval = 100 # number of iterations per display the status
gpu_id = 0
out_dir = 'result'
seed = 0 # random seed
# + [markdown] id="Q735WN-T6C4J" colab_type="text"
# ## 2. Preparation of training data and iterator
# In this notebook, we will use the training data which are preprocessed by [chainer.datasets.get_cifar10](https://docs.chainer.org/en/stable/reference/generated/chainer.datasets.get_cifar10.html#chainer.datasets.get_cifar10).
#
# From [Wikipedia](https://en.wikipedia.org/wiki/CIFAR-10), it says
#
# > The CIFAR-10 dataset (Canadian Institute For Advanced Research) is a collection of images that are commonly used to train machine learning and computer vision algorithms. It is one of the most widely used datasets for machine learning research.The CIFAR-10 dataset contains 60,000 32x32 color images in 10 different classes. The 10 different classes represent airplanes, cars, birds, cats, deer, dogs, frogs, horses, ships, and trucks. There are 6,000 images of each class.
#
# Let’s retrieve the CIFAR-10 dataset by using Chainer’s dataset utility function `get_cifar10`. CIFAR-10 is a set of small natural images. Each example is an RGB color image of size 32x32. In the original images, each component of pixels is represented by one-byte unsigned integer. This function scales the components to floating point values in the interval `[0, scale]`.
# + id="nZVIeRChTmM5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593380406023, "user_tz": -120, "elapsed": 18611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="975b3eeb-bf53-490d-c9b1-8d46acff358b"
# Load the CIFAR10 dataset if args.dataset is not specified
train, _ = chainer.datasets.get_cifar10(withlabel=False, scale=255.)
# + id="DGuctCHNyE45" colab_type="code" colab={}
train_iter = chainer.iterators.SerialIterator(train, batchsize)
# + [markdown] id="GVUZPtjo8ycv" colab_type="text"
#
# ## 3. Preparation of the model
#
# Let's define the network.
# We will create the model called DCGAN(Deep Convolutional GAN).
# As shown below, it is a model using CNN(Convolutional Neural Network) as its name suggests.
#
# 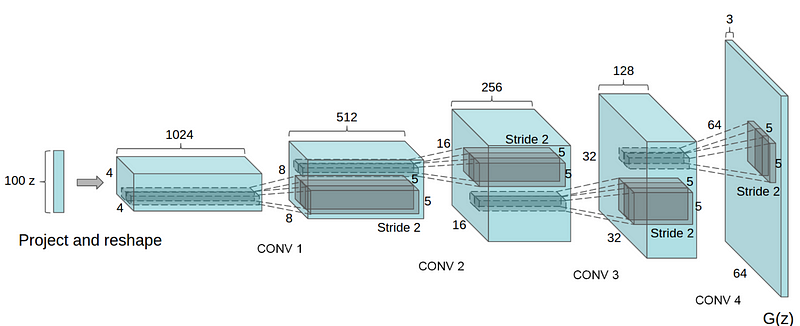 cited from [1]
#
#
# First, let’s define a network for the generator.
# + id="aGICf_k0hzW8" colab_type="code" colab={}
class Generator(chainer.Chain):
def __init__(self, n_hidden, bottom_width=4, ch=512, wscale=0.02):
super(Generator, self).__init__()
self.n_hidden = n_hidden
self.ch = ch
self.bottom_width = bottom_width
with self.init_scope():
w = chainer.initializers.Normal(wscale)
self.l0 = L.Linear(self.n_hidden, bottom_width * bottom_width * ch,
initialW=w)
self.dc1 = L.Deconvolution2D(ch, ch // 2, 4, 2, 1, initialW=w)
self.dc2 = L.Deconvolution2D(ch // 2, ch // 4, 4, 2, 1, initialW=w)
self.dc3 = L.Deconvolution2D(ch // 4, ch // 8, 4, 2, 1, initialW=w)
self.dc4 = L.Deconvolution2D(ch // 8, 3, 3, 1, 1, initialW=w)
self.bn0 = L.BatchNormalization(bottom_width * bottom_width * ch)
self.bn1 = L.BatchNormalization(ch // 2)
self.bn2 = L.BatchNormalization(ch // 4)
self.bn3 = L.BatchNormalization(ch // 8)
def make_hidden(self, batchsize):
return np.random.uniform(-1, 1, (batchsize, self.n_hidden, 1, 1)).astype(np.float32)
def __call__(self, z):
h = F.reshape(F.relu(self.bn0(self.l0(z))),
(len(z), self.ch, self.bottom_width, self.bottom_width))
h = F.relu(self.bn1(self.dc1(h)))
h = F.relu(self.bn2(self.dc2(h)))
h = F.relu(self.bn3(self.dc3(h)))
x = F.sigmoid(self.dc4(h))
return x
# + [markdown] id="nwjn5u_is1_f" colab_type="text"
# When we make a network in Chainer, we should follow some rules:
#
# 1. Define a network class which inherits `Chain`.
# 2. Make `chainer.links` ‘s instances in the `init_scope():` of the initializer `__init__`.
# 3. Concatenate `chainer.links` ‘s instances with `chainer.functions` to make the whole network.
#
# If you are not familiar with constructing a new network, you can read [this tutorial](http://localhost:8000/guides/models.html#creating-models).
#
# As we can see from the initializer `__init__`, the Generator uses the deconvolution layer `Deconvolution2D` and the batch normalization `BatchNormalization`. In `__call__`, each layer is concatenated by relu except the last layer.
#
# Because the first argument of `L.Deconvolution` is the channel size of input and the second is the channel size of output, we can find that each layer halve the channel size. When we construct `Generator` with `ch=1024`, the network is same with the image above.
#
#
# ---
#
# Note
#
# > Be careful when you concatenate a fully connected layer’s output and a convolutinal layer’s input. As we can see the 1st line of `__call__`, the output and input have to be concatenated with reshaping by `reshape`.
#
#
# ---
#
#
# + [markdown] id="kPvBD2pMw1AG" colab_type="text"
# In addtion, let’s define a network for the discriminator.
# + id="ECEfDBnK7We-" colab_type="code" colab={}
class Discriminator(chainer.Chain):
def __init__(self, bottom_width=4, ch=512, wscale=0.02):
w = chainer.initializers.Normal(wscale)
super(Discriminator, self).__init__()
with self.init_scope():
self.c0_0 = L.Convolution2D(3, ch // 8, 3, 1, 1, initialW=w)
self.c0_1 = L.Convolution2D(ch // 8, ch // 4, 4, 2, 1, initialW=w)
self.c1_0 = L.Convolution2D(ch // 4, ch // 4, 3, 1, 1, initialW=w)
self.c1_1 = L.Convolution2D(ch // 4, ch // 2, 4, 2, 1, initialW=w)
self.c2_0 = L.Convolution2D(ch // 2, ch // 2, 3, 1, 1, initialW=w)
self.c2_1 = L.Convolution2D(ch // 2, ch // 1, 4, 2, 1, initialW=w)
self.c3_0 = L.Convolution2D(ch // 1, ch // 1, 3, 1, 1, initialW=w)
self.l4 = L.Linear(bottom_width * bottom_width * ch, 1, initialW=w)
self.bn0_1 = L.BatchNormalization(ch // 4, use_gamma=False)
self.bn1_0 = L.BatchNormalization(ch // 4, use_gamma=False)
self.bn1_1 = L.BatchNormalization(ch // 2, use_gamma=False)
self.bn2_0 = L.BatchNormalization(ch // 2, use_gamma=False)
self.bn2_1 = L.BatchNormalization(ch // 1, use_gamma=False)
self.bn3_0 = L.BatchNormalization(ch // 1, use_gamma=False)
def __call__(self, x):
h = add_noise(x)
h = F.leaky_relu(add_noise(self.c0_0(h)))
h = F.leaky_relu(add_noise(self.bn0_1(self.c0_1(h))))
h = F.leaky_relu(add_noise(self.bn1_0(self.c1_0(h))))
h = F.leaky_relu(add_noise(self.bn1_1(self.c1_1(h))))
h = F.leaky_relu(add_noise(self.bn2_0(self.c2_0(h))))
h = F.leaky_relu(add_noise(self.bn2_1(self.c2_1(h))))
h = F.leaky_relu(add_noise(self.bn3_0(self.c3_0(h))))
return self.l4(h)
# + [markdown] id="2QnK8A8rw18G" colab_type="text"
# The `Discriminator` network is almost same with the transposed network of the `Generator`. However, there are minor different points:
#
# 1. Use `leaky_relu` as activation functions
# 2. Deeper than `Generator`
# 3. Add some noise when concatenating layers
# + id="eb6tMwLj0ss9" colab_type="code" colab={}
def add_noise(h, sigma=0.2):
xp = cuda.get_array_module(h.data)
if chainer.config.train:
return h + sigma * xp.random.randn(*h.shape)
else:
return h
# + [markdown] id="iWCNMuMvxQ51" colab_type="text"
# Let’s make the instances of the `Generator` and the `Discriminator`.
# + id="3KFGlbfJ7Wta" colab_type="code" colab={}
gen = Generator(n_hidden=n_hidden)
dis = Discriminator()
# + [markdown] id="lkuabIJoy3NF" colab_type="text"
# ## 4. Preparing Optimizer
# + [markdown] id="6L5PIzCYyHlm" colab_type="text"
# Next, let’s make optimizers for the models created above.
# + id="GkxLt_f7y3WM" colab_type="code" colab={}
# Setup an optimizer
def make_optimizer(model, alpha=0.0002, beta1=0.5):
optimizer = chainer.optimizers.Adam(alpha=alpha, beta1=beta1)
optimizer.setup(model)
optimizer.add_hook(
chainer.optimizer_hooks.WeightDecay(0.0001), 'hook_dec')
return optimizer
# + id="yWkvOh6CzZWZ" colab_type="code" colab={}
opt_gen = make_optimizer(gen)
opt_dis = make_optimizer(dis)
# + [markdown] id="OTq_sX298-ZR" colab_type="text"
# ## 5. Preparation and training of Updater · Trainer
#
# The GAN need the two models: the generator and the discriminator. Usually, the default updaters pre-defined in Chainer take only one model. So, we need to define a custom updater for the GAN training.
#
# The definition of `DCGANUpdate`r is a little complicated. However, it just minimize the loss of the discriminator and that of the generator alternately. We will explain the way of updating the models.
#
# As you can see in the class definiton, `DCGANUpdater` inherits `StandardUpdater`. In this case, almost all necessary functions are defined in `StandardUpdater`, we just override the functions of `__init__` and `update_core`.
#
#
#
# ---
# Note
#
# > We do not need to define `loss_dis` and `loss_gen` because the functions are called only in `update_core`. It aims at improving readability.
#
# ---
#
#
# + id="jsW8yD_jyq6y" colab_type="code" colab={}
class DCGANUpdater(chainer.training.updaters.StandardUpdater):
def __init__(self, *args, **kwargs):
self.gen, self.dis = kwargs.pop('models')
super(DCGANUpdater, self).__init__(*args, **kwargs)
def loss_dis(self, dis, y_fake, y_real):
batchsize = len(y_fake)
L1 = F.sum(F.softplus(-y_real)) / batchsize
L2 = F.sum(F.softplus(y_fake)) / batchsize
loss = L1 + L2
chainer.report({'loss': loss}, dis)
return loss
def loss_gen(self, gen, y_fake):
batchsize = len(y_fake)
loss = F.sum(F.softplus(-y_fake)) / batchsize
chainer.report({'loss': loss}, gen)
return loss
def update_core(self):
gen_optimizer = self.get_optimizer('gen')
dis_optimizer = self.get_optimizer('dis')
batch = self.get_iterator('main').next()
x_real = Variable(self.converter(batch, self.device)) / 255.
xp = chainer.backends.cuda.get_array_module(x_real.data)
gen, dis = self.gen, self.dis
batchsize = len(batch)
y_real = dis(x_real)
z = Variable(xp.asarray(gen.make_hidden(batchsize)))
x_fake = gen(z)
y_fake = dis(x_fake)
dis_optimizer.update(self.loss_dis, dis, y_fake, y_real)
gen_optimizer.update(self.loss_gen, gen, y_fake)
# + [markdown] id="btSYZF-50Yhz" colab_type="text"
# In the intializer `__init__`, an addtional key word argument `models` is required as you can see the codes below. Also, we use key word arguments `iterator`, `optimizer` and `device`. Be careful for the `optimizer`. We needs not only two models but also two optimizers. So, we should input `optimizer` as dictionary `{'gen': opt_gen, 'dis': opt_dis}`. In the `DCGANUpdater`, you can access the iterator with `self.get_iterator('main')`. Also, you can access the optimizers with `self.get_optimizer('gen')` and `self.get_optimizer('dis')`.
#
# In `update_core`, the two loss functions `loss_dis` and `loss_gen` are minimized by the optimizers. At first two lines, we access to the optimizers. Then, we generates next batch of training data by `self.get_iterator('main').next()`, and convert `batch` to `x_real` to make the training data suitable for `self.device` (e.g. GPU or CPU). After that, we minimize the loss functions with the optimizers.
#
#
# ---
# Note
#
# > When we define `update_core`, we usually want manipulate `array` with `numpy` library. Be careful that `array` should be `numpy` on CPU, but on GPU it should be `cupy`. But you do not need to write `if` condition because you can access the correct library by `xp = chainer.backends.cuda.get_array_module(array.data)`. On GPU, `xp` is `cupy`, otherwise `numpy`.
#
#
# ---
#
#
# + id="1Nl65Ivsye5-" colab_type="code" colab={}
updater = DCGANUpdater(
models=(gen, dis),
iterator=train_iter,
optimizer={
'gen': opt_gen, 'dis': opt_dis},
device=gpu_id)
trainer = chainer.training.Trainer(updater, (n_epoch, 'epoch'), out=out_dir)
# + id="e-QROwBN0JMB" colab_type="code" colab={}
from PIL import Image
import chainer.backends.cuda
def out_generated_image(gen, dis, rows, cols, seed, dst):
@chainer.training.make_extension()
def make_image(trainer):
np.random.seed(seed)
n_images = rows * cols
xp = gen.xp
z = Variable(xp.asarray(gen.make_hidden(n_images)))
with chainer.using_config('train', False):
x = gen(z)
x = chainer.backends.cuda.to_cpu(x.data)
np.random.seed()
x = np.asarray(np.clip(x * 255, 0.0, 255.0), dtype=np.uint8)
_, _, H, W = x.shape
x = x.reshape((rows, cols, 3, H, W))
x = x.transpose(0, 3, 1, 4, 2)
x = x.reshape((rows * H, cols * W, 3))
preview_dir = '{}/preview'.format(dst)
preview_path = preview_dir +\
'/image{:0>8}.png'.format(trainer.updater.iteration)
if not os.path.exists(preview_dir):
os.makedirs(preview_dir)
Image.fromarray(x).save(preview_path)
return make_image
# + id="nXAIZSy9cQdu" colab_type="code" colab={}
snapshot_interval = (snapshot_interval, 'iteration')
display_interval = (display_interval, 'iteration')
trainer.extend(
extensions.snapshot(filename='snapshot_iter_{.updater.iteration}.npz'),
trigger=snapshot_interval)
trainer.extend(extensions.snapshot_object(
gen, 'gen_iter_{.updater.iteration}.npz'), trigger=snapshot_interval)
trainer.extend(extensions.snapshot_object(
dis, 'dis_iter_{.updater.iteration}.npz'), trigger=snapshot_interval)
trainer.extend(extensions.LogReport(trigger=display_interval))
trainer.extend(extensions.PrintReport([
'epoch', 'iteration', 'gen/loss', 'dis/loss',
]), trigger=display_interval)
trainer.extend(extensions.ProgressBar(update_interval=100))
trainer.extend(
out_generated_image(
gen, dis,
10, 10, seed, out_dir),
trigger=snapshot_interval)
# + id="cHrEr46hzvuW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 884} executionInfo={"status": "ok", "timestamp": 1593381257972, "user_tz": -120, "elapsed": 264328, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="5ad2da3d-867a-455f-fe80-be5a5bc294ea"
# Run the training
trainer.run()
# + [markdown] id="N7wiomoIvjc1" colab_type="text"
# ## 6. Checking the performance with test data
# + id="pigrILri0p7b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"status": "ok", "timestamp": 1593381344588, "user_tz": -120, "elapsed": 460, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="02e90423-460a-4999-9d1d-1288fa45a84f" language="bash"
# ls result/preview
# + id="u47Am2e85xG3" colab_type="code" colab={}
from IPython.display import Image, display_png
import glob
image_files = sorted(glob.glob(out_dir + '/preview/*.png'))
# + id="gyeXhpTH7j_l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} executionInfo={"status": "error", "timestamp": 1593381359574, "user_tz": -120, "elapsed": 451, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyMjILKo4X-uSOnHqHCWcSsndp3-3-2CJW5WO4jFM=s64", "userId": "14982494247992103515"}} outputId="954da1b6-a42e-456f-af9a-6f4812befddd"
display_png(Image(image_files[0])) # first snapshot
# + id="njq0O-RZ7kHB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 336} outputId="70d8cb78-e19a-4f80-bfa6-e9ad5b8b7edf"
display_png(Image(image_files[-1])) # last snapshot
# + [markdown] id="_UBmkq3xYmSX" colab_type="text"
# ## Reference
# [1] [Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks](https://arxiv.org/abs/1511.06434)
| GAN/Intro2AI_dcgan_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise notebook :
# +
import warnings
warnings.simplefilter('ignore', FutureWarning)
import pandas as pd
from datetime import datetime
# -
london = pd.read_csv('London_2014.csv', skipinitialspace=True)
london.head()
# `Note that the right hand side of the table has been cropped to fit on the page.
# You’ll find out how to remove rogue spaces.`
# ## Every picture tells a story
#
# It can be difficult and confusing to look at a table of rows of numbers and make any
# meaningful interpretation especially if there are many rows and columns.
# Handily, pandas has a method called **plot()** which will visualise data for us by producing
# a chart.
#
# The following line of code tells Jupyter to display inside this notebook any graph that is created.
# %matplotlib inline
# The `plot()` method can make a graph of the values in a column. Gridlines are turned on by the `grid` argument.
#
# To plot `‘Max Wind SpeedKm/h ’`, it’s as simple as this code:
london['Max Wind SpeedKm/h'].plot(grid=True)
# The `grid=True` argument makes the gridlines (the dotted lines in the image above)
# appear, which make values easier to read on the chart. The chart comes out a bit small,
# so the graph can be made bigger by giving the method a `figsize=(x,y)` argument where `x` and `y` are integers that determine the length of the `x-axis` and `y-axis`.
london['Max Wind SpeedKm/h'].plot(grid=True, figsize=(10,5))
# That’s better! The argument given to the `plot()` method, `figsize=(10,5)` simply tells
# `plot()` that the `x-axis` should be 10 units wide and the `y-axis` should be 5 units high. In
# the above graph the `x-axis` (the numbers at the bottom) shows the dataframe’s index, so 0
# is 1 January and 50 is 18 February.
# The `y-axis` (the numbers on the side) shows the range of wind speed in kilometres per
# hour. It is clear that the windiest day in 2014 was somewhere in mid-February and the
# wind reached about `66 kilometers per hour`.
# By default, the `plot()` method will try to generate a line, although as you’ll see in a later
# modules, it can produce other chart types too.
#
# Multiple lines can be plotted by selecting multiple columns.
london[['Max Wind SpeedKm/h', 'Mean Wind SpeedKm/h']].plot(grid=True, figsize=(10,5))
# ### Task
#
# In the cell below, write code to plot the minimum, mean, and maximum temperature during 2014 in London.
london[['Min TemperatureC', 'Max TemperatureC','Mean TemperatureC']].plot(grid=True, figsize=(15,5))
# ## Changing a dataframe's index
# We have seen that by default every dataframe has an integer index for its rows which
# starts from `0`.
# The dataframe we’ve been using, london , has an index that goes from `0 to 364`. The
# row indexed by 0 holds data for the first day of the year and the row indexed by 364 holds
# data for the last day of the year. However, the column `'GMT' holds datetime64` values
# which would make a more intuitive index.
# Changing the index to `datetime64` values is as easy as assigning to the dataframe’s
# index attribute the contents of the `'GMT'` column, is done by assigning to the dataframe's `index` attribute the contents of the `'GMT`' column, like this:
london['GMT'] = pd.to_datetime(london['GMT'])
london.index = london['GMT']
london.head(2)
# `Notice that the 'GMT' column still remains and that the index has been labelled to show
# that it has been derived from the 'GMT' column.`
# The `iloc` attribute can still be used to get and display rows by number, but now you can now also use the `datetime64` index to get a row by date, using the dataframe's `loc` attribute, like this:
london.loc[datetime(2014, 1, 1)]
# A query such as *'Return all the rows where the date is between December 8th and December 12th'* can now be done succinctly like this:
# +
london.loc[datetime(2014,12,8) : datetime(2014,12,12)]
#The meaning of the above code is get the rows beween and including
#the indices datetime(2014,12,8) and datetime(2014,12,12)
# -
# Because the table is in date order, we can be confident that only the rows with dates
# between 8 December 2014 and 12 December 2014 (inclusive) will be returned. However if
# the table had not been in date order, we would have needed to sort it first, like this:
london = london.sort_index()
london
# Now we have a `datetime64` index, let's plot `'Max Wind SpeedKm/h'` again:
london['Max Wind SpeedKm/h'].plot(grid=True, figsize=(10,5))
# Now it is much clearer that the worst winds were in mid February.
#
# ### Task
# Use the code cell below to plot the values of `'Mean Humidity'` during spring (full months of March, April and May).
# +
london = london.sort_index()
london.loc[datetime(2014,3,1) : datetime(2014,5,31)]['Mean Humidity'].plot(grid=True, figsize=(10,5))
# -
# Your project this week is to find out what would have been the best two weeks of weather
# for a 2014 vacation in a capital of a **BRICS** country.
# I’ve written up my analysis of the best two weeks of weather in London, UK, which you can
# open in project notebook.
# The structure is very simple: besides the introduction and the conclusions, there is one
# section for each step of the analysis – obtaining, cleaning and visualising the data.
# Once you’ve worked through my analysis you should open a dataset for just one of the
# BRICS capitals: Brasilia, Moscow, Delhi, Beijing or Cape Town, the dataset has been downloade and can be found in the folder. The choice of capital is up
# to you. You should then work out the best two weeks, according to the weather, to choose
# for a two-week holiday in your chosen capital city.
#
# Once again, do not open the file with Excel , but you could take a look using a text
# editor.
# In my project, I was looking for a two
# week period that had relatively high temperatures and little rain. If you choose a capital in
# a particularly hot and dry country you will probably be looking for relatively cool weather
# and low humidity.
#
# Note that the London file has the dates in a column named ‘GMT’ whereas in the BRICS
# files they are in a column named ‘Date’. You will need to change the Python code
# accordingly. You should also change the name of the variable, London, according to the
# capital you choose.
#
# ## GOOD LUCK!
# +
import warnings
warnings.simplefilter('ignore', FutureWarning)
import pandas as pd
from datetime import datetime
Moscow = pd.read_csv('Moscow_SVO_2014.csv', skipinitialspace=True)
# -
Moscow = Moscow.rename(columns={'WindDirDegrees<br />' : 'WindDirDegrees'})
Moscow['WindDirDegrees'] = Moscow['WindDirDegrees'].str.rstrip('<br />')
Moscow['WindDirDegrees'] = Moscow['WindDirDegrees'].astype('float64')
Moscow['Date'] = pd.to_datetime(Moscow['Date'])
Moscow.dtypes
Moscow[Moscow['Date'] == datetime(2014, 6, 4)]
Moscow.index = Moscow['Date']
summer = Moscow.loc[datetime(2014,6,1) : datetime(2014,8,31)]
summer[summer['Mean TemperatureC'] >= 25]
# %matplotlib inline
summer['Mean TemperatureC'].plot(grid=True, figsize=(15,10))
summer[['Mean TemperatureC', 'Precipitationmm']].plot(grid=True, figsize=(10,5))
Aug = summer.loc[datetime(2014,8,1) : datetime(2014,8,31)]
Aug[['Mean TemperatureC', 'Precipitationmm']].plot(grid=True, figsize=(10,5))
capetown = pd.read_csv('CapeTown_CPT_2014.csv', skipinitialspace=True)
capetown.columns
capetown['Date'] = pd.to_datetime(capetown['Date'])
Moscow.dtypes
capetown[capetown['Date'] == datetime(2014, 6, 4)]
capetown.index = capetown['Date']
summer = capetown.loc[datetime(2014,1,1) : datetime(2014,2,28)]
summer[summer['Mean TemperatureC'] >= 25]
# %matplotlib inline
summer['Mean TemperatureC'].plot(grid=True, figsize=(10,5))
summer[['Mean TemperatureC', 'Precipitationmm']].plot(grid=True, figsize=(10,5))
| Florence Wangechi WT - 021-089/17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + cellView="form" colab={} colab_type="code" id="wDlWLbfkJtvu"
#@title Copyright 2020 Google LLC. Double-click here for license information.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="TL5y5fY9Jy_x"
# # Introduction to Neural Nets
#
# This Colab builds a deep neural network to perform more sophisticated linear regression than the earlier Colabs.
# + [markdown] colab_type="text" id="7RDY3EeAluPd"
# ## Learning Objectives:
#
# After doing this Colab, you'll know how to do the following:
#
# * Create a simple deep neural network.
# * Tune the hyperparameters for a simple deep neural network.
# + [markdown] colab_type="text" id="XGj0PNaJlubZ"
# ## The Dataset
#
# Like several of the previous Colabs, this Colab uses the [California Housing Dataset](https://developers.google.com/machine-learning/crash-course/california-housing-data-description).
# + [markdown] colab_type="text" id="tX_umRMMsa3z"
# ## Use the right version of TensorFlow
#
# The following hidden code cell ensures that the Colab will run on TensorFlow 2.X.
# + cellView="form" colab={} colab_type="code" id="lM75uNH-sTv2"
#@title Run on TensorFlow 2.x
# #%tensorflow_version 2.x
from __future__ import absolute_import, division, print_function, unicode_literals
# + [markdown] colab_type="text" id="xchnxAsaKKqO"
# ## Import relevant modules
#
# The following hidden code cell imports the necessary code to run the code in the rest of this Colaboratory.
# + cellView="form" colab={} colab_type="code" id="9n9_cTveKmse"
#@title Import relevant modules
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow.keras import layers
from matplotlib import pyplot as plt
import seaborn as sns
# The following lines adjust the granularity of reporting.
pd.options.display.max_rows = 10
pd.options.display.float_format = "{:.1f}".format
print("Imported modules.")
# + [markdown] colab_type="text" id="X_TaJhU4KcuY"
# ## Load the dataset
#
# Like most of the previous Colab exercises, this exercise uses the California Housing Dataset. The following code cell loads the separate .csv files and creates the following two pandas DataFrames:
#
# * `train_df`, which contains the training set
# * `test_df`, which contains the test set
#
# + colab={} colab_type="code" id="JZlvdpyYKx7V"
train_df = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv")
train_df = train_df.reindex(np.random.permutation(train_df.index)) # shuffle the examples
test_df = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_test.csv")
# + [markdown] colab_type="text" id="8ldP-5z1B2vL"
# ## Normalize values
#
# When building a model with multiple features, the values of each feature should cover roughly the same range. The following code cell normalizes datasets by converting each raw value to its Z-score. (For more information about Z-scores, see the Classification exercise.)
# + cellView="form" colab={} colab_type="code" id="g8HC-TDgB1D1"
#@title Convert raw values to their Z-scores
# Calculate the Z-scores of each column in the training set:
train_df_mean = train_df.mean()
train_df_std = train_df.std()
train_df_norm = (train_df - train_df_mean)/train_df_std
# Calculate the Z-scores of each column in the test set.
test_df_mean = test_df.mean()
test_df_std = test_df.std()
test_df_norm = (test_df - test_df_mean)/test_df_std
print("Normalized the values.")
# + [markdown] colab_type="text" id="b9ehCgIRjTxy"
# ## Represent data
#
# The following code cell creates a feature layer containing three features:
#
# * `latitude` X `longitude` (a feature cross)
# * `median_income`
# * `population`
#
# This code cell specifies the features that you'll ultimately train the model on and how each of those features will be represented. The transformations (collected in `my_feature_layer`) don't actually get applied until you pass a DataFrame to it, which will happen when we train the model.
# + colab={} colab_type="code" id="8EkNAQhnjSu-"
# Create an empty list that will eventually hold all created feature columns.
feature_columns = []
# We scaled all the columns, including latitude and longitude, into their
# Z scores. So, instead of picking a resolution in degrees, we're going
# to use resolution_in_Zs. A resolution_in_Zs of 1 corresponds to
# a full standard deviation.
resolution_in_Zs = 0.3 # 3/10 of a standard deviation.
# Create a bucket feature column for latitude.
latitude_as_a_numeric_column = tf.feature_column.numeric_column("latitude")
latitude_boundaries = list(np.arange(int(min(train_df_norm['latitude'])),
int(max(train_df_norm['latitude'])),
resolution_in_Zs))
latitude = tf.feature_column.bucketized_column(latitude_as_a_numeric_column, latitude_boundaries)
# Create a bucket feature column for longitude.
longitude_as_a_numeric_column = tf.feature_column.numeric_column("longitude")
longitude_boundaries = list(np.arange(int(min(train_df_norm['longitude'])),
int(max(train_df_norm['longitude'])),
resolution_in_Zs))
longitude = tf.feature_column.bucketized_column(longitude_as_a_numeric_column,
longitude_boundaries)
# Create a feature cross of latitude and longitude.
latitude_x_longitude = tf.feature_column.crossed_column([latitude, longitude], hash_bucket_size=100)
crossed_feature = tf.feature_column.indicator_column(latitude_x_longitude)
feature_columns.append(crossed_feature)
# Represent median_income as a floating-point value.
median_income = tf.feature_column.numeric_column("median_income")
feature_columns.append(median_income)
# Represent population as a floating-point value.
population = tf.feature_column.numeric_column("population")
feature_columns.append(population)
# Convert the list of feature columns into a layer that will later be fed into
# the model.
my_feature_layer = tf.keras.layers.DenseFeatures(feature_columns)
# + [markdown] colab_type="text" id="Ak_TMAzGOIFq"
# ## Build a linear regression model as a baseline
#
# Before creating a deep neural net, find a [baseline](https://developers.google.com/machine-learning/glossary/#baseline) loss by running a simple linear regression model that uses the feature layer you just created.
#
# + cellView="form" colab={} colab_type="code" id="QF0BFRXTOeR3"
#@title Define the plotting function.
def plot_the_loss_curve(epochs, mse):
"""Plot a curve of loss vs. epoch."""
plt.figure()
plt.xlabel("Epoch")
plt.ylabel("Mean Squared Error")
plt.plot(epochs, mse, label="Loss")
plt.legend()
plt.ylim([mse.min()*0.95, mse.max() * 1.03])
plt.show()
print("Defined the plot_the_loss_curve function.")
# + cellView="form" colab={} colab_type="code" id="RW4Qe710LgnG"
#@title Define functions to create and train a linear regression model
def create_model(my_learning_rate, feature_layer):
"""Create and compile a simple linear regression model."""
# Most simple tf.keras models are sequential.
model = tf.keras.models.Sequential()
# Add the layer containing the feature columns to the model.
model.add(feature_layer)
# Add one linear layer to the model to yield a simple linear regressor.
model.add(tf.keras.layers.Dense(units=1, input_shape=(1,)))
# Construct the layers into a model that TensorFlow can execute.
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.MeanSquaredError()])
return model
def train_model(model, dataset, epochs, batch_size, label_name):
"""Feed a dataset into the model in order to train it."""
# Split the dataset into features and label.
features = {name:np.array(value) for name, value in dataset.items()}
label = np.array(features.pop(label_name))
history = model.fit(x=features, y=label, batch_size=batch_size,
epochs=epochs, shuffle=True)
# Get details that will be useful for plotting the loss curve.
epochs = history.epoch
hist = pd.DataFrame(history.history)
rmse = hist["mean_squared_error"]
return epochs, rmse
print("Defined the create_model and train_model functions.")
# + [markdown] colab_type="text" id="f47LmxF5X_pu"
# Run the following code cell to invoke the the functions defined in the preceding two code cells. (Ignore the warning messages.)
#
# **Note:** Because we've scaled all the input data, **including the label**, the resulting loss values will be *much less* than previous models.
#
# **Note:** Depending on the version of TensorFlow, running this cell might generate WARNING messages. Please ignore these warnings.
# + colab={} colab_type="code" id="tsfE4ujDL4ju"
# The following variables are the hyperparameters.
learning_rate = 0.01
epochs = 15
batch_size = 1000
label_name = "median_house_value"
# Establish the model's topography.
my_model = create_model(learning_rate, my_feature_layer)
# Train the model on the normalized training set.
epochs, mse = train_model(my_model, train_df_norm, epochs, batch_size, label_name)
plot_the_loss_curve(epochs, mse)
test_features = {name:np.array(value) for name, value in test_df_norm.items()}
test_label = np.array(test_features.pop(label_name)) # isolate the label
print("\n Evaluate the linear regression model against the test set:")
my_model.evaluate(x = test_features, y = test_label, batch_size=batch_size)
# + [markdown] colab_type="text" id="3014ezH3C7jT"
# ## Define a deep neural net model
#
# The `create_model` function defines the topography of the deep neural net, specifying the following:
#
# * The number of [layers](https://developers.google.com/machine-learning/glossary/#layer) in the deep neural net.
# * The number of [nodes](https://developers.google.com/machine-learning/glossary/#node) in each layer.
#
# The `create_model` function also defines the [activation function](https://developers.google.com/machine-learning/glossary/#activation_function) of each layer.
# + cellView="both" colab={} colab_type="code" id="pedD5GhlDC-y"
def create_model(my_learning_rate, my_feature_layer):
"""Create and compile a simple linear regression model."""
# Most simple tf.keras models are sequential.
model = tf.keras.models.Sequential()
# Add the layer containing the feature columns to the model.
model.add(my_feature_layer)
# Describe the topography of the model by calling the tf.keras.layers.Dense
# method once for each layer. We've specified the following arguments:
# * units specifies the number of nodes in this layer.
# * activation specifies the activation function (Rectified Linear Unit).
# * name is just a string that can be useful when debugging.
# Define the first hidden layer with 20 nodes.
model.add(tf.keras.layers.Dense(units=20,
activation='relu',
name='Hidden1'))
# Define the second hidden layer with 12 nodes.
model.add(tf.keras.layers.Dense(units=12,
activation='relu',
name='Hidden2'))
# Define the output layer.
model.add(tf.keras.layers.Dense(units=1,
name='Output'))
model.compile(optimizer=tf.keras.optimizers.Adam(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.MeanSquaredError()])
return model
# + [markdown] colab_type="text" id="anH4A_yCcZx2"
# ## Define a training function
#
# The `train_model` function trains the model from the input features and labels. The [tf.keras.Model.fit](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential#fit) method performs the actual training. The `x` parameter of the `fit` method is very flexible, enabling you to pass feature data in a variety of ways. The following implementation passes a Python dictionary in which:
#
# * The *keys* are the names of each feature (for example, `longitude`, `latitude`, and so on).
# * The *value* of each key is a NumPy array containing the values of that feature.
#
# **Note:** Although you are passing *every* feature to `model.fit`, most of those values will be ignored. Only the features accessed by `my_feature_layer` will actually be used to train the model.
# + colab={} colab_type="code" id="4jv_lJYTcrEF"
def train_model(model, dataset, epochs, label_name,
batch_size=None):
"""Train the model by feeding it data."""
# Split the dataset into features and label.
features = {name:np.array(value) for name, value in dataset.items()}
label = np.array(features.pop(label_name))
history = model.fit(x=features, y=label, batch_size=batch_size,
epochs=epochs, shuffle=True)
# The list of epochs is stored separately from the rest of history.
epochs = history.epoch
# To track the progression of training, gather a snapshot
# of the model's mean squared error at each epoch.
hist = pd.DataFrame(history.history)
mse = hist["mean_squared_error"]
return epochs, mse
# + [markdown] colab_type="text" id="D-IXYVfvM4gD"
# ## Call the functions to build and train a deep neural net
#
# Okay, it is time to actually train the deep neural net. If time permits, experiment with the three hyperparameters to see if you can reduce the loss
# against the test set.
#
# + cellView="both" colab={} colab_type="code" id="nj3v5EKQFY8s"
# The following variables are the hyperparameters.
learning_rate = 0.01
epochs = 20
batch_size = 1000
# Specify the label
label_name = "median_house_value"
# Establish the model's topography.
my_model = create_model(learning_rate, my_feature_layer)
# Train the model on the normalized training set. We're passing the entire
# normalized training set, but the model will only use the features
# defined by the feature_layer.
epochs, mse = train_model(my_model, train_df_norm, epochs,
label_name, batch_size)
plot_the_loss_curve(epochs, mse)
# After building a model against the training set, test that model
# against the test set.
test_features = {name:np.array(value) for name, value in test_df_norm.items()}
test_label = np.array(test_features.pop(label_name)) # isolate the label
print("\n Evaluate the new model against the test set:")
my_model.evaluate(x = test_features, y = test_label, batch_size=batch_size)
# + [markdown] colab_type="text" id="wlPXK-SmmjQ2"
# ## Task 1: Compare the two models
#
# How did the deep neural net perform against the baseline linear regression model?
# + cellView="form" colab={} colab_type="code" id="hI7ojsL7nnBE"
#@title Double-click to view a possible answer
# Assuming that the linear model converged and
# the deep neural net model also converged, please
# compare the test set loss for each.
# In our experiments, the loss of the deep neural
# network model was consistently lower than
# that of the linear regression model, which
# suggests that the deep neural network model
# will make better predictions than the
# linear regression model.
# + [markdown] colab_type="text" id="Y5IKmk7D49_n"
# ## Task 2: Optimize the deep neural network's topography
#
# Experiment with the number of layers of the deep neural network and the number of nodes in each layer. Aim to achieve both of the following goals:
#
# * Lower the loss against the test set.
# * Minimize the overall number of nodes in the deep neural net.
#
# The two goals may be in conflict.
# + cellView="form" colab={} colab_type="code" id="wYG5qXpP5a9n"
#@title Double-click to view a possible answer
# Many answers are possible. We noticed the
# following trends:
# * Two layers outperformed one layer, but
# three layers did not perform significantly
# better than two layers; two layers
# outperformed one layer.
# In other words, two layers seemed best.
# * Setting the topography as follows produced
# reasonably good results with relatively few
# nodes:
# * 10 nodes in the first layer.
# * 6 nodes in the second layer.
# As the number of nodes in each layer dropped
# below the preceding, test loss increased.
# However, depending on your application, hardware
# constraints, and the relative pain inflicted
# by a less accurate model, a smaller network
# (for example, 6 nodes in the first layer and
# 4 nodes in the second layer) might be
# acceptable.
# + [markdown] colab_type="text" id="Pu7R_ZpDopIj"
# ## Task 3: Regularize the deep neural network (if you have enough time)
#
# Notice that the model's loss against the test set is *much higher* than the loss against the training set. In other words, the deep neural network is [overfitting](https://developers.google.com/machine-learning/glossary/#overfitting) to the data in the training set. To reduce overfitting, regularize the model. The course has suggested several different ways to regularize a model, including:
#
# * [L1 regularization](https://developers.google.com/machine-learning/glossary/#L1_regularization)
# * [L2 regularization](https://developers.google.com/machine-learning/glossary/#L2_regularization)
# * [Dropout regularization](https://developers.google.com/machine-learning/glossary/#dropout_regularization)
#
# Your task is to experiment with one or more regularization mechanisms to bring the test loss closer to the training loss (while still keeping test loss relatively low).
#
# **Note:** When you add a regularization function to a model, you might need to tweak other hyperparameters.
#
# ### Implementing L1 or L2 regularization
#
# To use L1 or L2 regularization on a hidden layer, specify the `kernel_regularizer` argument to [tf.keras.layers.Dense](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense). Assign one of the following methods to this argument:
#
# * `tf.keras.regularizers.l1` for L1 regularization
# * `tf.keras.regularizers.l2` for L2 regularization
#
# Each of the preceding methods takes an `l` parameter, which adjusts the [regularization rate](https://developers.google.com/machine-learning/glossary/#regularization_rate). Assign a decimal value between 0 and 1.0 to `l`; the higher the decimal, the greater the regularization. For example, the following applies L2 regularization at a strength of 0.05.
#
# ```
# model.add(tf.keras.layers.Dense(units=20,
# activation='relu',
# kernel_regularizer=tf.keras.regularizers.l2(l=0.01),
# name='Hidden1'))
# ```
#
# ### Implementing Dropout regularization
#
# You implement dropout regularization as a separate layer in the topography. For example, the following code demonstrates how to add a dropout regularization layer between the first hidden layer and the second hidden layer:
#
# ```
# model.add(tf.keras.layers.Dense( *define first hidden layer*)
#
# model.add(tf.keras.layers.Dropout(rate=0.25))
#
# model.add(tf.keras.layers.Dense( *define second hidden layer*)
# ```
#
# The `rate` parameter to [tf.keras.layers.Dropout](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dropout) specifies the fraction of nodes that the model should drop out during training.
#
# + cellView="form" colab={} colab_type="code" id="tflt9TZEDARW"
#@title Double-click for a possible solution
# The following "solution" uses L2 regularization to bring training loss
# and test loss closer to each other. Many, many other solutions are possible.
def create_model(my_learning_rate, my_feature_layer):
"""Create and compile a simple linear regression model."""
# Discard any pre-existing version of the model.
model = None
# Most simple tf.keras models are sequential.
model = tf.keras.models.Sequential()
# Add the layer containing the feature columns to the model.
model.add(my_feature_layer)
# Describe the topography of the model.
# Implement L2 regularization in the first hidden layer.
model.add(tf.keras.layers.Dense(units=20,
activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.04),
name='Hidden1'))
# Implement L2 regularization in the second hidden layer.
model.add(tf.keras.layers.Dense(units=12,
activation='relu',
kernel_regularizer=tf.keras.regularizers.l2(0.04),
name='Hidden2'))
# Define the output layer.
model.add(tf.keras.layers.Dense(units=1,
name='Output'))
model.compile(optimizer=tf.keras.optimizers.Adam(lr=my_learning_rate),
loss="mean_squared_error",
metrics=[tf.keras.metrics.MeanSquaredError()])
return model
# Call the new create_model function and the other (unchanged) functions.
# The following variables are the hyperparameters.
learning_rate = 0.007
epochs = 140
batch_size = 1000
label_name = "median_house_value"
# Establish the model's topography.
my_model = create_model(learning_rate, my_feature_layer)
# Train the model on the normalized training set.
epochs, mse = train_model(my_model, train_df_norm, epochs,
label_name, batch_size)
plot_the_loss_curve(epochs, mse)
test_features = {name:np.array(value) for name, value in test_df_norm.items()}
test_label = np.array(test_features.pop(label_name)) # isolate the label
print("\n Evaluate the new model against the test set:")
my_model.evaluate(x = test_features, y = test_label, batch_size=batch_size)
| mlcc-exercises_en/intro_to_neural_nets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Underfitting vs. Overfitting
# +
print(__doc__)
# Author: <NAME>
# <NAME>
# License: BSD 3 clause
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
# +
def f(x):
""" function to approximate by polynomial interpolation"""
return x * np.sin(x)
# generate points used to plot
x_plot = np.linspace(0, 10, 100)
# generate points and keep a subset of them
x = np.linspace(0, 10, 100)
print(x[:5])
rng = np.random.RandomState(0)
rng.shuffle(x)
print(x[:5])
x = np.sort(x[:50])
y = f(x)
# Add noise to targets
#y[::5] += 3 * (0.5 - rng.rand(x.shape[0] // 5))
# +
# create matrix versions of these arrays
X = x[:, np.newaxis]
X_plot = x_plot[:, np.newaxis]
colors = ['teal', 'yellowgreen', 'gold']
lw = 2
plt.plot(x_plot, f(x_plot), color='cornflowerblue', linewidth=lw, label="ground truth")
plt.scatter(x, y, color='navy', s=30, marker='o', label="training points")
plt.legend(loc='lower left')
plt.show()
# +
plt.scatter(x, y, color='navy', s=30, marker='o', label="training points")
plt.plot(x_plot, f(x_plot), color='cornflowerblue', linewidth=lw, label="ground truth")
for count, degree in enumerate([3, 4, 5]):
model = make_pipeline(PolynomialFeatures(degree), Ridge())
model.fit(X, y)
y_plot = model.predict(X_plot)
plt.plot(x_plot, y_plot, color=colors[count], linewidth=lw,
label="degree %d" % degree)
plt.legend(loc='lower left')
plt.show()
# +
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
def true_fun(X):
return np.cos(1.5 * np.pi * X)
np.random.seed(0)
n_samples = 30
degrees = [1, 4,10, 15]
X = np.sort(np.random.rand(n_samples))
y = true_fun(X) + np.random.randn(n_samples) * 0.1
plt.figure(figsize=(14, 5))
for i in range(len(degrees)):
ax = plt.subplot(1, len(degrees), i + 1)
plt.setp(ax, xticks=(), yticks=())
polynomial_features = PolynomialFeatures(degree=degrees[i],
include_bias=False)
linear_regression = LinearRegression()
pipeline = Pipeline([("polynomial_features", polynomial_features),
("linear_regression", linear_regression)])
pipeline.fit(X[:, np.newaxis], y)
# Evaluate the models using crossvalidation
scores = cross_val_score(pipeline, X[:, np.newaxis], y,
scoring="neg_mean_squared_error", cv=10)
X_test = np.linspace(0, 1, 100)
plt.plot(X_test, pipeline.predict(X_test[:, np.newaxis]), label="Model")
plt.plot(X_test, true_fun(X_test), label="True function")
plt.scatter(X, y, edgecolor='b', s=20, label="Samples")
plt.xlabel("x")
plt.ylabel("y")
plt.xlim((0, 1))
plt.ylim((-2, 2))
plt.legend(loc="best")
plt.title("Degree {}\nMSE = {:.2e}(+/- {:.2e})".format(
degrees[i], -scores.mean(), scores.std()))
plt.show()
# -
from sklearn.datasets.samples_generator import make_blobs
import matplotlib.pyplot as plt
from pandas import DataFrame
import numpy as np
# +
# generate 2d classification dataset
X, y = make_blobs(n_samples=1000, centers=3, n_features=2)
# scatter plot, dots colored by class value
df = DataFrame(dict(x=X[:,0], y=X[:,1], label=y))
colors = {0:'red', 1:'blue', 2:'green'}
fig, ax = pyplot.subplots()
grouped = df.groupby('label')
for key, group in grouped:
group.plot(ax=ax, kind='scatter', x='x', y='y', label=key, color=colors[key])
pyplot.show()
# -
X = 5 * rng.rand(10000, 1)
y = np.sin(X).ravel()
# +
rng = np.random.RandomState(0)
# Generate sample data
X = 5 * rng.rand(10000, 1)
y = np.sin(X).ravel()
# Add noise to targets
y[::5] += 3 * (0.5 - rng.rand(X.shape[0] // 5))
# -
X_plot = np.linspace(0, 5, 100000)[:, None]
train_size = 100
plt.title("Sine scatter plot with added Noise", fontsize=18)
plt.grid(True)
N=50
colors = np.random.rand(N)
plt.scatter(X[:100], y[:100], c='b', label='data', zorder=1, edgecolors=(1, 1, 1))
plt.show()
| content/Pandas Operations/.ipynb_checkpoints/Bias Variance Trade off - TODO-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="OpcQOtRw2OjO"
# # 6. Numpy basics
# + [markdown] id="0aKm_Dcq2OjY"
# The structures offered by Python are limited, especially if we deal with the multi-dimensional lists that images are. The solution to this is the Numpy package, which offers a new object called an array. On top of offering this structure, Numpy also offers a vast number of functions that operate directly on arrays, sparing us the tedious task of making sure we handle all the pixels of an image. Let's first import Numpy:
# + executionInfo={"elapsed": 934, "status": "ok", "timestamp": 1616247469278, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="rJMAvyT42OjZ"
import numpy as np
import urllib.request
urllib.request.urlretrieve('https://raw.githubusercontent.com/guiwitz/PyImageCourse_beginner/master/svg.py', 'svg.py')
from svg import numpy_to_svg
# + [markdown] id="uZQYtPhc2OjZ"
# ## 6.1 What is an array
# + [markdown] id="gVSm5qJO2OjZ"
# Now let us create arrays. The two simplest ways of doing that is to create arrays filled with 0's or 1's. For example to create a 6x4 array of zeros:
# + executionInfo={"elapsed": 611, "status": "ok", "timestamp": 1616247475383, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="kFHWasNt2Oja"
myarray = np.zeros((4,6))
# + [markdown] id="B-d6smMh2Ojb"
# An array is just a matrix, i.e. a grid of numbers with multiple dimensions. For example the array above look like this:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 550, "status": "ok", "timestamp": 1616247477314, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="klXLihdT2Ojd" outputId="a46adfea-0271-4210-c864-1f13069e4e39"
numpy_to_svg(myarray)
# + [markdown] id="JHVOzBvn2Oje"
# And the "inside" of the array looks like this:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 639, "status": "ok", "timestamp": 1616247480616, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="Vb1qBblx2Ojf" outputId="d027fa7c-6ad8-4955-db61-d37f38e8b269"
myarray
# + [markdown] id="ik2jd7VT2Ojf"
# We see that we have 6 separate list of 4 0's grouped into a larger list. This entire structure is an array of 6 rows and 4 columns. In the frame of this course you can also see it as a 6x4 pixel image.
#
# Seeing rows and columns as a system of coordinates, we can access to specific pixels. For example the pixels at row = 3 and columns = 2 is:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 581, "status": "ok", "timestamp": 1616247481275, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="r_VZfMfO2Ojg" outputId="486a7ecd-a0a1-410c-b989-87978be35e16"
myarray[3,2]
# + [markdown] id="M-xr04tC2Ojg"
# We can even modify its value:
# + executionInfo={"elapsed": 639, "status": "ok", "timestamp": 1616247483599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="ZADiI9Z42Ojg"
myarray[3,2] = 13
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 401, "status": "ok", "timestamp": 1616247483600, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="hVv0h8Nn2Ojh" outputId="e44924e7-8f6b-407a-abd6-1bf7037b50c7"
myarray
# + [markdown] id="u9tcGi4X2Ojh"
# Similarly we can crearte an array filled with 1s:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 487, "status": "ok", "timestamp": 1616247485396, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="hhmQ3MoX2Oji" outputId="4882f825-0197-4959-c607-8984d13833d0"
np.ones((3,6))
# + [markdown] id="YSMXBcZ52Oji"
# Finally we can create a one-dimensional array with evenly spaced valeus (similar to range) using:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 792, "status": "ok", "timestamp": 1616247487494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="uHZnR3vV2Oji" outputId="501d4356-3b4d-41ad-b68e-3b3accf25f21"
np.arange(0,30,3)
# + [markdown] id="PWBLj0h22Oji"
# ## 6.2 Simple calculus with arrays
# + [markdown] id="7iNYVlHw2Oji"
# The beautiful thing with arrays, is that you can consider them like an object and forget that they are composed of multiple elements. For example we can just add a value to all all pixels using:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 531, "status": "ok", "timestamp": 1616247490459, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="3gHEKK4n2Ojj" outputId="e73f56fd-18b3-40e6-d32a-ca4a88493e72"
myarray + 32
# + [markdown] id="B8mePU5I2Ojj"
# Of course as long as we don't reassign this new state to our variable it remains unchanged:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 534, "status": "ok", "timestamp": 1616247491962, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="lRBQqXX72Ojj" outputId="13941f50-e4e4-44e5-b378-dd7873e25e5d"
myarray
# + [markdown] id="zB6RdPSQ2Ojj"
# We have to write:
# + executionInfo={"elapsed": 589, "status": "ok", "timestamp": 1616247493157, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="VVyf4d8I2Ojk"
myarray = myarray + 32
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 495, "status": "ok", "timestamp": 1616247494848, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="utkPbh3G2Ojk" outputId="937c718d-89ea-4c20-f48f-68ed7f060148"
myarray
# + [markdown] id="m5-OT2Wg2Ojk"
# Note that if we didn't have those nice Numpy properties, we would have to "manually" go through all pixels to adjust their values:
# + executionInfo={"elapsed": 739, "status": "ok", "timestamp": 1616247497016, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="MWNeovif2Ojk"
for x in range(4):
for y in range(6):
myarray[x,y] = myarray[x,y] + 5
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 370, "status": "ok", "timestamp": 1616247497018, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="_b5Cryng2Ojl" outputId="901d774d-0aca-4182-e175-d9e3bd5d3821"
myarray
# + [markdown] id="F0WRZb6Y2Ojl"
# Of course we can do much more complex operations on these arrays. For example we can use the cosinus function of Numpy and apply it to the entire array:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 731, "status": "ok", "timestamp": 1616247500543, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="9S7uh7tv2Ojl" outputId="6a9c9050-48a0-466d-b50f-0dd9e98f0a08"
np.cos(myarray)
# + [markdown] id="3MspOSEn2Ojl"
# ## 6.3 Operations combining arrays
# + [markdown] id="PmY3kZVh2Ojm"
# In addition to operations that apply to entire arrays, we can do also operations combining multiple arrays. For example we can add two arrays:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 541, "status": "ok", "timestamp": 1616247503303, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="6BFc72QL2Ojm" outputId="e520fe8b-18ad-41d9-e7c6-d60b8cb7486c"
myarray1 = 2*np.ones((4,4))
myarray2 = 5*np.ones((4,4))
myarray3 = myarray1 * myarray2
myarray3
# + [markdown] id="CrLD35vm2Ojm"
# The one important constraint is of course that the two arrays used in an operation need to have the same size. Otherwise Numpy doesn't know which pairs of pixel to consider.
# + colab={"base_uri": "https://localhost:8080/", "height": 198} executionInfo={"elapsed": 583, "status": "error", "timestamp": 1616247506734, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="NwzUV6Ra2Ojm" outputId="5cc50d8a-cf58-45ee-d214-6b02a7cfbdc5"
myarray1 = 2*np.ones((3,3))
myarray2 = 5*np.ones((4,4))
myarray3 = myarray1 * myarray2
# + [markdown] id="kUWVRxsE2Ojm"
# ## 6.4 Higher dimensions
# + [markdown] id="MzC-GfYs2Ojm"
# Until now we have almost only dealt with 2D arrays that look like a simple grid:
# + colab={"base_uri": "https://localhost:8080/", "height": 171} executionInfo={"elapsed": 709, "status": "ok", "timestamp": 1616247514948, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="iy3JkQiT2Ojn" outputId="c28b7c9d-726c-4c0e-9f79-8ce1e5d1375e"
myarray = np.ones((5,10))
numpy_to_svg(myarray)
# + [markdown] id="cxXW6-hT2Ojn"
# We are not limited to create 1 or 2 dimensional arrays. We can basically create any-dimension array. For example if we do 3D imaging, we are going to have a series of planes assembled in a "cube". For example if we acquired 5 planes of a 10px by 10px image, we would have something like:
# + executionInfo={"elapsed": 668, "status": "ok", "timestamp": 1616247517864, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="Vqfed2eK2Ojn"
array3D = np.ones((10,10,5))
# + colab={"base_uri": "https://localhost:8080/", "height": 388} executionInfo={"elapsed": 797, "status": "ok", "timestamp": 1616247518247, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="9mDh5Fd72Ojn" outputId="e5dcd1d3-f808-4069-9ab2-6bae7954bc41"
numpy_to_svg(array3D)
# + [markdown] id="NGvDnyJ72Ojo"
# In higher dimensions, the array content output is not very readable. To know what we are dealing with, an **extremely** helpul property associated to arrays is shape:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 456, "status": "ok", "timestamp": 1616247520675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="xTSnEYbm2Ojo" outputId="e8fbc3fb-4765-4a6e-a686-5c22b9665bf1"
array3D.shape
# + [markdown] id="8St0sVfx2Ojo"
# This gives us the size of the array in each dimension.
#
# **Note that shape, which is a parameter of the array, is used almost like a method, however without parenthesis.**
# + [markdown] id="yCTLA5pa2Ojo"
# ## 6.5 Methods and parameters of arrays
# + [markdown] id="FO2q252b2Ojo"
# Just like variables and structures, arrays have also associated methods. As we have just seen, they also have associated parameters that one can call without ().
#
# Sometimes it is unclear whether we deal with a method or parameter. Just try and see what gives an error!
#
# These methods/parameters give a lot of information on the array and are very helpful. For example, one can ask what is the largest element:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 559, "status": "ok", "timestamp": 1616247527416, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="oku5BCrq2Ojo" outputId="a8c5af76-6317-46aa-d7e5-2a98d80a3f6e"
myarray.max
# + [markdown] id="TIg_UbpF2Ojp"
# This indicates that we actually deal with a method:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 786, "status": "ok", "timestamp": 1616247528433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="5suXfChJ2Ojp" outputId="64d18a8e-5d26-4ba6-f533-fb3e46df12c5"
myarray.max()
# + [markdown] id="p2eZkjD_2Ojp"
# We can also e.g. calculate the sum of all elements:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 564, "status": "ok", "timestamp": 1616247530226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="zOGzUcZa2Ojt" outputId="39a1fd62-592e-4ee1-cf1a-ea11e5aa5e62"
myarray.sum()
# + [markdown] id="eM1dwibj2Ojt"
# You can actively read the documentation to lear about all methods. The simplest is usually just to do a [Google Search](https://www.google.com/search?ei=5rd2XKqwN8SUsAfj7IuICw&q=sum+of+numpy+array&oq=sum+of+numpy+array&gs_l=psy-ab.3..0i7i30l7j0i203l2j0i5i30.23694.24024..24373...0.0..0.66.193.3......0....1..gws-wiz.......0i71.0RxePnmQfcg) for whatever you are looking for.
# + [markdown] id="Mys9ytER2Ojt"
# ## 6.6 Logical operations
# + [markdown] id="aBYE6AXG2Oju"
# Just like variables, arrays can be composed of booleans. Usually they are obtained by using a logical operation on a standard array:
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 617, "status": "ok", "timestamp": 1616247534190, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="5oFiTWHo2Oju" outputId="fc9d1d9d-c62b-42be-e5f4-28dcedb05e85"
myarray = np.zeros((4,4))
myarray[2,3] = 1
myarray
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 542, "status": "ok", "timestamp": 1616247535525, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="IPzALsPt2Oju" outputId="5bb61a4e-da48-4bab-f7d8-d51751a48e16"
myarray > 0
# + [markdown] id="KtniuTTI2Oju"
# Exactly as for simple variables, we can assign this boolean array to a new variable directly:
# + executionInfo={"elapsed": 378, "status": "ok", "timestamp": 1616247536793, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="Zvo048nl2Oju"
myboolean = myarray > 0
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 592, "status": "ok", "timestamp": 1616247538160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgT0K2JVYzEsjzsS5nhkUVjUrSIJ5jHzXnBoYrmVf8=s64", "userId": "16033870147214403532"}, "user_tz": -60} id="hoNitifV2Oju" outputId="67e118d0-9477-4cf6-ecc9-cddf1151deed"
myboolean
# + [markdown] id="KFXJl9q72Ojv"
# ## 6.7 Array slicing
# + [markdown] id="Do1rec-F2Ojv"
# A vast chapter is array slicing, the fact to only consider a part of an array, as when cropping an image for example. We are however going to treat that part later using actual images, to make the examples more understandable.
# + id="itR31wwx2Ojv"
| 06-Numpy_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: zipline
# language: python
# name: zipline
# ---
# # 运行
# +
from zipline.pipeline import Pipeline, STOCK_DB, Fundamentals
from zipline.pipeline.data import USEquityPricing
from zipline.pipeline.filters import StaticSids
# -
from zipline.research.run import run_pipeline
from zipline.pipeline.builtin import NDays,IsStock
def make_pipeline():
ndays = NDays()
time_to_market = Fundamentals.time_to_market.latest
return Pipeline(
columns = {
'上市天数':ndays,
'time_to_market':time_to_market
},
screen = IsStock()
)
df = run_pipeline(make_pipeline(), '2017-11-29','2017-11-30')
df.head()
import numpy as np
import pandas as pd
# # 验证
todays = df.index.get_level_values(0)
days = (todays.tz_localize(None) - pd.to_datetime(df['time_to_market'].values)).days
assert all(days == df['上市天数'])
| docs/memo/pipeline/Factors/test_ndays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Build Features
#
# As a recap, the [training data](../data/processed/train-physicists-from-1901.csv), [validation data](../data/processed/validation-physicists-from-1901.csv) and [test data](../data/processed/test-physicists-from-1901.csv) contain information on physicists who were eligible to receive a Nobel Prize in Physics. That is, they were alive on and after 10 December 1901, the date the prize was first awarded.
#
# All of the physicists in the training data are deceased and all the physicists in the validation and test data are alive. Recall that the Nobel Prize in Physics cannot be awarded posthumously and one of the goals of this project is to try to predict the next Physics Nobel Laureates. As a result, the data was purposely sampled in this way, so that the training set can be used to build models, which predict whether a living physicist is likely to be awarded the Nobel Prize in Physics.
#
# It is time to use the training, validation and test data, along with the other various pieces of data: [Nobel Physics Laureates](../data/raw/nobel-physics-prize-laureates.csv), [Nobel Chemistry Laureates](../data/raw/nobel-chemistry-prize-laureates.csv), [Places](../data/processed/places.csv) and [Countries](../data/processed/Countries-List.csv), to create features that may help in predicting Physics Nobel Laureates.
# ## Setting up the Environment
#
# An initialization step is needed to setup the environment:
# - The locale needs to be set for all categories to the user’s default setting (typically specified in the LANG environment variable) to enable correct sorting of words with accents.
# +
import locale
locale.setlocale(locale.LC_ALL, '')
# +
from datetime import datetime
import numpy as np
import pandas as pd
from pycountry_convert import country_alpha2_to_country_name
from pycountry_convert import country_name_to_country_alpha3
from pycountry_convert import country_alpha2_to_continent_code
from pycountry_convert import country_alpha3_to_country_alpha2
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.preprocessing import OneHotEncoder
from src.data.country_utils import nationality_to_alpha2_code
from src.features.features_utils import rank_hot_encode
# -
# ## Reading in the Data
#
# First let's read in the training, validation and test data and the list of Nobel Physics Laureates.
train_physicists = pd.read_csv('../data/processed/train-physicists-from-1901.csv')
train_physicists.head()
validation_physicists = pd.read_csv('../data/processed/validation-physicists-from-1901.csv')
validation_physicists.head()
test_physicists = pd.read_csv('../data/processed/test-physicists-from-1901.csv')
test_physicists.head()
nobel_physicists = pd.read_csv('../data/raw/nobel-physics-prize-laureates.csv')
nobel_physicists.head()
# There are some variants of laureate names in the training, validation and test data. As we will be searching for whether academic advisors, students, spouses, children, etc. of a physicist are physics laureates, for convenience it's useful to merge the `name` field into Nobel Physicists dataframe.
nobel_columns = ['Year', 'Laureate', 'name', 'Country', 'Rationale']
nobel_physicists = pd.merge(nobel_physicists,
train_physicists.append(validation_physicists).append(test_physicists),
how = 'left', left_on = 'Laureate',
right_on = 'fullName')[nobel_columns]
nobel_physicists.head()
# Now let's read in the list of Nobel Chemistry Laureates.
nobel_chemists = pd.read_csv('../data/raw/nobel-chemistry-prize-laureates.csv')
nobel_chemists.head()
# Again, we will be searching for whether academic advisors, students, spouses, children, etc. of a physicist are chemistry laureates. So for convenience it's useful to merge the `name` field into Nobel Chemists dataframe.
nobel_chemists = pd.merge(nobel_chemists,
train_physicists.append(validation_physicists).append(test_physicists),
how = 'left', left_on = 'Laureate',
right_on = 'fullName')[nobel_columns]
nobel_chemists.head()
# These are essentially physicists who are Chemistry Nobel Laureates. Surpringly there are quite a few of them. Of course, as noted previously, *<NAME>* is the only double laureate in Physics and Chemistry.
nobel_chemists[nobel_chemists.name.notna()].name
# It is worth noting that if there are alternative names of Chemistry Nobel Laureates in the physicists dataframe other than those above, they will *not* be found. However, we do not expect many of these, as at one point all the redirected URLS for the names were retrieved and very few were associated with laureates. In fact you can still see that some of these names are present in the [DBpedia redirects](../data/raw/dbpedia-redirects.csv) (e.g. search for "Marie_Curie"). When processing the physicist data earlier, the imputing of these redirects was removed for the names as a few of them were wrong. For instance, *<NAME>'s* children redirect back to him! (e.g. search for "Carl_Feynman" and "Michelle_Feynmann" in the DBpedia redirects or directly try http://dbpedia.org/page/Carl_Feynman or http://dbpedia.org/page/Michelle_Feynmann in your browser.
#
# Another interesting observation is that the only one in this list still alive is *<NAME>*. So we should not expect to see many, if any, physicists in the validation or test set who have Chemistry Nobel Laureate academic advisors, notable students, spouses etc. This is clearly a facet in which the training data is different from the validation and test data. Such differences can make learning difficult.
#
# Now, let's read the places and nationalities data into a dataframe. It's important at this point to turn off the default behavior of pandas which is to treat the string literal 'NA' as a missing value. In the dataset, 'NA' is both the continent code of North America and the ISO 3166 alpha-2 country code of Namibia. We then have to impute the missing values since pandas replaces them with the empty string.
places = pd.read_csv('../data/processed/places.csv', keep_default_na=False)
places = places.replace('', np.nan)
assert(all(places[places.countryAlpha3Code == 'USA']['continentCode'].values == 'NA'))
places.head()
nationalities = pd.read_csv('../data/processed/Countries-List.csv', keep_default_na=False)
nationalities = nationalities.replace('', np.nan)
assert(nationalities[nationalities.Name == 'Namibia']['ISO 3166 Code'].values == 'NA')
nationalities.head()
# Finally, with all the data read in, we can now move on to the real work, creating the features.
# ## Creating the Features
#
# It is now time to create the features from the collected data. The features we will create are listed in the table below along with their type and description. The features can be grouped into three main groups, with the bulk of features falling in the first group, then the second group and so on:
#
# 1. Features related to *professional and personal relationships* that the physicists have to *physics or chemistry laureates*, *educational institutions*, *work institutions*, *countries* and *continents*.
#
# 2. Features related to the subfield of focus of the physicist denoting whether s/he is a *experimental physicist*, *theoretical physicist* or an *astronomer*.
#
# 3. Features related to personal characteristics of the physicist, namely, *gender* and *number of years lived*.
#
# Remember that in the first group, there are people and institutions from different countries and continents that are directly involved in the [selection and voting process for the Nobel Prize in Physics](https://www.nobelprize.org/nomination/physics/) and therefore have a direct influence on those who become laureates. The second group is connected to subjective biases that may or may not exist concerning the major subfield of research of the physicist. While the third group is connected to subjective biases that may or may not exist concerning the gender and age of a physicist. Although the latter is also related to the invention or discovery "standing the test of time".
#
# | Feature | Type | Description |
# | :---: | :---: | :---: |
# | alma_mater | Categorical | List of universities attended |
# | alma_mater_continent_codes | Categorical | List of continent codes of universities attended |
# | alma_mater_country_alpha_3_codes | Categorical | List of country codes of universities attended |
# | birth_continent_codes | Categorical | List of continent codes of birth countries |
# | birth_country_alpha_3_codes | Categorical | List of country codes of birth countries |
# | citizenship_continent_codes | Categorical | List of continent codes of coutries of citizenship |
# | citizenship_country_alpha_3_codes | Categorical | List of country codes of citizenship |
# | gender | Binary | Gender of the physicist (male / female) |
# | is_astronomer | Binary | Is the physicist an astronomer? (yes / no) |
# | is_experimental_physicist | Binary | Is the physicist an experimental physicist? (yes / no) |
# | is_theoretical_physicist | Binary | Is the physicist a theoretical physicist? (yes / no) |
# | num_alma_mater | Ordinal | No. of universities attended |
# | num_alma_mater_continent_codes | Ordinal | No. of continent codes of universities attended |
# | num_alma_mater_country_alpha_3_codes | Ordinal | No. of country codes of universities attended |
# | num_birth_continent_codes | Ordinal | No. of continent codes of birth countries |
# | num_birth_country_alpha_3_codes | Ordinal | No. of birth country codes |
# | num_chemistry_laureate_academic_advisors | Ordinal | No. of chemistry laureate academic advisors |
# | num_chemistry_laureate_children | Ordinal | No. of chemistry laureate children |
# | num_chemistry_laureate_doctoral_advisors | Ordinal | No. of chemistry laureate doctoral advisors |
# | num_chemistry_laureate_doctoral_students | Ordinal | No. of chemistry laureate doctoral students |
# | num_chemistry_laureate_influenced | Ordinal | No. of chemistry laureates the physicist influenced |
# | num_chemistry_laureate_influenced_by | Ordinal | No. of chemistry laureates the physicist was influenced by |
# | num_chemistry_laureate_notable_students | Ordinal | No. of chemistry laureate notable students |
# | num_chemistry_laureate_parents | Ordinal | No. of chemistry laureate parents |
# | num_chemistry_laureate_spouses | Ordinal | No. of chemistry laureate spouses |
# | num_citizenship_continent_codes | Ordinal | No. continent codes of countries of citizenship |
# | num_citizenship_country_alpha_3_codes | Ordinal | No. of country codes of citizenship |
# | num_physics_laureate_academic_advisors | Ordinal | No. of physics laureate academic advisors |
# | num_physics_laureate_children | Ordinal | No. of physics laureate children |
# | num_physics_laureate_doctoral_advisors | Ordinal | No. of physics laureate doctoral advisors |
# | num_physics_laureate_doctoral_students | Ordinal | No. of physics laureate doctoral students |
# | num_physics_laureate_influenced | Ordinal | No. of physics laureates the physicist influenced |
# | num_physics_laureate_influenced_by | Ordinal | No. of physics laureates the physicist was influenced by |
# | num_physics_laureate_notable_students | Ordinal | No. of physics laureate notable students |
# | num_physics_laureate_parents | Ordinal | No. of physics laureate parents |
# | num_physics_laureate_spouses | Ordinal | No. of physics laureate spouses |
# | num_residence_continent_codes | Ordinal | No. of continent codes of residence countries |
# | num_residence_country_alpha_3_codes | Ordinal | No. of residence country codes |
# | num_workplaces | Ordinal | No. of workplaces |
# | num_workplaces_continent_codes | Ordinal | No. of continent codes of countries of workplaces |
# | num_workplaces_country_alpha_3_codes | Ordinal | No. of country codes of countries worked in |
# | num_years_lived_group | Ordinal | No. of years lived group (18-24, 25-34, etc.) |
# | residence_continent_codes | Categorical | List of continent codes of countries of residence |
# | residence_country_alpha_3_codes | Categorical | List of country codes of countries of residence |
# | workplaces | Categorical | List of workplaces |
# | workplaces_continent_codes | Categorical | List of continent codes of countries worked in |
# | workplaces_country_alpha_3_codes | Categorical | List of country codes of countries worked in |
#
# Some comments are also warranted with regards to the types of the feature variables. As you can see, there are three types of variables:
#
# 1. **Ordinal** variables.
#
# 2. **Categorical** variables.
#
# 3. **Binary** (**dichotomous**) variables.
#
#
# The categorical variables are all lists of varying lengths of **places** and therefore are not in the appropriate form for machine learning. Once we create them they will be encoded into binary variables and the lists will be discarded. You may ask why the encoding is done with categorical yes / no values rather than 0 / 1 values? It is because the algorithms that we will be processing the data with would treat 0 / 1 values as quantitive in nature, which clearly is not desired. Essentially, we will be left with two variable types, binary variables and ordinal variables. OK time to create the features.
# +
def build_features(physicists, nobel_physicists, nobel_chemists, places, nationalities):
"""Build features for the physicists.
Args:
physicists (pandas.DataFrame): Physicists dataframe.
nobel_physicists (pandas.DataFrame): Nobel Physics
Laureate dataframe.
nobel_chemists (pandas.DataFrame): Nobel Chemistry
Laureate dataframe.
places (pandas.DataFrame): Places dataframe.
nationality (pandas.DataFrame): Nationalies dataframe.
Returns:
pandas.DataFrame: Features dataframe.
"""
features = physicists.copy()[['fullName', 'name', 'gender']].rename(
mapper={'fullName': 'full_name'}, axis='columns')
features['num_years_lived_group'] = _build_num_years_lived_group(
physicists.birthDate, physicists.deathDate)
_build_physics_subfield_features(features, physicists)
_build_num_laureates_features(features, physicists,
nobel_physicists, nobel_chemists)
_build_citizenship_features(features, physicists, nationalities)
_build_places_features(features, physicists, places)
features = features.drop('name', axis='columns')
return features
def _build_physics_subfield_features(features, physicists):
features_to_build = {
'is_theoretical_physicist': {'categories': 'Theoretical physicists',
'others': 'theoretical physic'},
'is_experimental_physicist': {'categories': 'Experimental physicists',
'others': 'experimental physic'},
'is_astronomer': {'categories': 'astronomers',
'others': 'astronom'}
}
for feature, search_terms in features_to_build.items():
features[feature] = _build_physics_subfield(
physicists.categories, physicists.field, physicists.description,
physicists.comment, search_terms=search_terms)
def _build_num_laureates_features(features, physicists, nobel_physicists,
nobel_chemists):
features_to_build = {
'laureate_academic_advisors': 'academicAdvisor',
'laureate_doctoral_advisors': 'doctoralAdvisor',
'laureate_doctoral_students': 'doctoralStudent',
'laureate_notable_students': 'notableStudent',
'laureate_children': 'child',
'laureate_parents': 'parent',
'laureate_spouses': 'spouse',
'laureate_influenced': 'influenced',
'laureate_influenced_by': 'influencedBy'
}
for feature, relation in features_to_build.items():
features['num_physics_' + feature] = _build_num_laureates(
physicists[relation], nobel_physicists.Laureate, nobel_physicists.name)
features['num_chemistry_' + feature] = _build_num_laureates(
physicists[relation], nobel_chemists.Laureate, nobel_chemists.name)
# drop columns where the counts are all zeros
non_zero = (features != 0).any(axis='rows')
features.drop(non_zero[non_zero == False].index, axis='columns', inplace=True)
def _build_places_features(features, physicists, places):
features_to_build = {
'birth_country_alpha_3_codes': 'birthPlace',
'birth_continent_codes': 'birthPlace',
'residence_country_alpha_3_codes': 'residence',
'residence_continent_codes': 'residence',
'alma_mater': 'almaMater',
'alma_mater_country_alpha_3_codes': 'almaMater',
'alma_mater_continent_codes': 'almaMater',
'workplaces': 'workplaces',
'workplaces_country_alpha_3_codes': 'workplaces',
'workplaces_continent_codes': 'workplaces'
}
for feature, place in features_to_build.items():
code = 'countryAlpha3Code'
if 'continent' in feature:
code = 'continentCode'
if feature in ['alma_mater', 'workplaces']:
features[feature] = physicists[place].apply(
_get_alma_mater_or_workplaces)
else:
features[feature] = _build_places_codes(
physicists[place], places.fullName, places[code])
features['num_' + feature] = features[feature].apply(len)
def _build_citizenship_features(features, physicists, nationalities):
citizenship = physicists.citizenship.apply(
_get_citizenship_codes, args=(nationalities,))
nationality = physicists.nationality.apply(
_get_citizenship_codes, args=(nationalities,))
citizenship_description = physicists.description.apply(
_get_citizenship_codes, args=(nationalities,))
features['citizenship_country_alpha_3_codes'] = (
(citizenship + nationality + citizenship_description).apply(
lambda ctz: list(sorted(set(ctz)))))
features['num_citizenship_country_alpha_3_codes'] = (
features.citizenship_country_alpha_3_codes.apply(len))
features['citizenship_continent_codes'] = (
features.citizenship_country_alpha_3_codes.apply(
lambda al3: list(sorted({country_alpha2_to_continent_code(
country_alpha3_to_country_alpha2(cd)) for cd in al3}))))
features['num_citizenship_continent_codes'] = (
features.citizenship_continent_codes.apply(len))
def _build_num_years_lived_group(birth_date, death_date):
death_date_no_nan = death_date.apply(_date_no_nan)
birth_date_no_nan = birth_date.apply(_date_no_nan)
years_lived = ((death_date_no_nan - birth_date_no_nan) /
pd.to_timedelta(1, 'Y'))
years_lived = years_lived.apply(np.floor)
years_lived_group = years_lived.apply(_years_lived_group)
return years_lived_group
def _years_lived_group(years_lived):
assert(years_lived >= 18 and years_lived <= 120)
groups = {
range(18, 25): '18-24',
range(25, 35): '25-34',
range(35, 50): '35-49',
range(50, 65): '50-64',
range(65, 80): '65-79',
range(80, 95): '80-94',
range(95, 121): '95-120'
}
for range_, code in groups.items():
if years_lived in range_:
return groups[range_]
def _build_physics_subfield(categories, field, description, comment, search_terms):
cat_theoretical_physicist = categories.apply(
lambda cat: search_terms['categories'] in cat)
field_theoretical_physicist = field.apply(
lambda fld: search_terms['others'] in fld.lower() if isinstance(fld, str)
else False)
desc_theoretical_physicist = description.apply(
lambda desc: search_terms['others'] in desc.lower() if isinstance(desc, str)
else False)
comm_theoretical_physicist = description.apply(
lambda comm: search_terms['others'] in comm.lower() if isinstance(comm, str)
else False)
subfield = (cat_theoretical_physicist |
field_theoretical_physicist |
desc_theoretical_physicist |
comm_theoretical_physicist)
subfield = subfield.apply(lambda val: 'yes' if val == True else 'no')
return subfield
def _build_num_laureates(series, laureates, names):
laureate_names = series.apply(_get_nobel_laureates,
args=(laureates, names))
return laureate_names.apply(len)
def _build_places_codes(places_in_physicists, full_name_in_places, places_codes):
codes = places_in_physicists.apply(_get_places_codes,
args=(full_name_in_places, places_codes))
return codes
def _get_alma_mater_or_workplaces(cell):
if isinstance(cell, float):
return list()
places = set()
places_in_cell = cell.split('|')
for place_in_cell in places_in_cell:
# group colleges of University of Oxford and University of Cambridge
# with their respective parent university
if place_in_cell.endswith(', Cambridge'):
places.add('University of Cambridge')
elif place_in_cell.endswith(', Oxford'):
places.add('University of Oxford')
else:
places.add(place_in_cell)
places = list(places)
places.sort(key=locale.strxfrm)
return places
def _get_citizenship_codes(series, nationalities):
alpha_2_codes = nationality_to_alpha2_code(series, nationalities)
if isinstance(alpha_2_codes, float):
return list()
alpha_2_codes = alpha_2_codes.split('|')
alpha_3_codes = [country_name_to_country_alpha3(
country_alpha2_to_country_name(alpha_2_code))
for alpha_2_code in alpha_2_codes]
return alpha_3_codes
def _get_nobel_laureates(cell, laureates, names):
laureates_in_cell = set()
if isinstance(cell, str):
# assume the same name if only differs by a hyphen
# or whitespace at front or end of string
values = cell.strip().replace('-', ' ').split('|')
for value in values:
if value in laureates.values:
laureates_in_cell.add(value)
if names.str.contains(value, regex=False).sum() > 0:
laureates_in_cell.add(value)
laureates_in_cell = list(laureates_in_cell)
return laureates_in_cell
def _get_places_codes(cell, full_name_in_places, places_codes):
codes = set()
if isinstance(cell, str):
places = cell.split('|')
for place in places:
code_indices = full_name_in_places[
full_name_in_places == place].index
assert(len(code_indices) <= 1)
if len(code_indices) != 1:
continue
code_index = code_indices[0]
codes_text = places_codes[code_index]
if isinstance(codes_text, float):
continue
codes_in_cell = codes_text.split('|')
for code_in_cell in codes_in_cell:
if code_in_cell:
codes.add(code_in_cell)
codes = list(codes)
codes.sort()
return codes
def _date_no_nan(date):
if isinstance(date, str):
return datetime.strptime(date, '%Y-%m-%d').date()
return datetime(2018, 10, 24).date() # fix the date for reproducibility
# -
train_features = build_features(train_physicists, nobel_physicists, nobel_chemists, places, nationalities)
assert((len(train_features) == len(train_physicists)))
assert(len(train_features.columns) == 45)
train_features.head()
validation_features = build_features(
validation_physicists, nobel_physicists, nobel_chemists, places, nationalities)
assert((len(validation_features) == len(validation_physicists)))
assert(len(validation_features.columns) == 37)
validation_features.head()
test_features = build_features(test_physicists, nobel_physicists, nobel_chemists, places, nationalities)
assert((len(test_features) == len(test_physicists)))
assert(len(test_features.columns) == 36)
test_features.head()
# So there are more features in the training set than in the validation and test sets. So what are these extra features? These are mainly related to the relationships physicists have with chemistry and physics laureates. It seems like the data is not so rich, especially with regards to more modern physicists.
train_features.columns.difference(validation_features.columns).tolist()
train_features.columns.difference(test_features.columns).tolist()
# Any machine models that we build will have parameters chosen using the validation set and be evaluated on the test set. The tempting thing to do is to reduce the features to the common set of features between the training, validation and test sets, which have variability across all three datasets. We will do this for the training and validation sets as it seems a perfectly reasonable thing to do. However, using the test set would clearly be *data snooping* (cheating) as it is meant to be unseen data, and as such, cannot be used to make any decisions during the modeling process. So to ensure that the test set features are identical to the training set features, we will "pad" the extra features in the test set with all "0" values and remove any extra features that are not present in the training set.
feature_cols = train_features.columns.intersection(validation_features.columns)
assert(validation_features.equals(validation_features[feature_cols]))
train_features = train_features[feature_cols]
assert((len(train_features.columns) == len(validation_features.columns)))
assert(sorted(train_features.columns.tolist()) == sorted(validation_features.columns.tolist()))
train_features.head()
feature_cols = test_features.columns.intersection(train_features.columns)
test_features = test_features[feature_cols]
test_features['num_physics_laureate_influenced'] = 0
test_features['num_physics_laureate_influenced_by'] = 0
assert((len(test_features.columns) == len(train_features.columns)))
assert(sorted(test_features.columns.tolist()) == sorted(train_features.columns.tolist()))
test_features.head()
# Now we will binary encode the list features. Due to the binary encoding there will be a differing number of features in the training, test and validation sets. We will follow a methodology analagous to the above in order to ensure that the features are identical in the training, validation and test sets.
#
# The differing features occur due to the differing *country codes*, *workplaces*, *educational institutions*, etc. that the physicists are associated with. Some of the differences are due variability in the data and some are caused by the **selection** bias that we deliberately introduced in our data sampling process. The latter issue is an important one that we will return to in a later notebook.
#
# We will also be using a `presence_threshold` to group binary features that only appear in a few instances into an "other" category. This is intended to reduce the dimensionality of the feature space and help to prevent overfitting during the model building phase. Let's go ahead and "binarize" the list features now.
# +
def binarize_list_features(features, train_features=None, presence_threshold=0.0, pad_features=False):
"""Binarize list features.
Binary encode the list categorical features in the
features dataframe.
Args:
features (pandas.DataFrame): Features dataframe.
train_features (pandas.DataFrame, optional): Defaults to None.
Training features dataframe. Pass this parameter when
building features for a test or validation set so that
features not found in the training features are grouped
into the "other category" that is mentioned in
`presence_threshold` below.
presence_threshold (float, optional): Defaults to 0.0. For
each category in a categorical list feature, the
fraction of physicists for which the category is
present will be calculated. If the fraction is below
this threshold it will grouped into the "other"
category (represented by one or more "*'s'" in its
name). This is intended for "bucketing" rare
values to keep the dimensionality of the feature
space down and reduce chances of overfitting. Set
this value to zero to prevent any grouping of
values. Note that this value will be ignored when
`train_features` is provided.
pad_features (bool, optional): Defaults to False. Pad binary
features not found in the training set with all 'no'
values. This should be set to True for a test set to
ensure that the test set features will match the training
set features.
Returns:
pandas.DataFrame: Features dataframe.
"""
# union of places and citizenship (without the counts)
series_to_binarize = {
'birth_country_alpha_3_codes': 'born_in_',
'birth_continent_codes': 'born_in_',
'residence_country_alpha_3_codes': 'lived_in_',
'residence_continent_codes': 'lived_in_',
'alma_mater': 'alumnus_of_',
'alma_mater_country_alpha_3_codes': 'alumnus_in_',
'alma_mater_continent_codes': 'alumnus_in_',
'workplaces': 'worked_at_',
'workplaces_country_alpha_3_codes': 'worked_in_',
'workplaces_continent_codes': 'worked_in_',
'citizenship_country_alpha_3_codes': 'citizen_of_',
'citizenship_continent_codes': 'citizen_in_'
}
for series, prefix in series_to_binarize.items():
binarized = _binarize_list_feature(features[series], prefix,
train_features, presence_threshold)
features = features.drop(series, axis='columns').join(binarized)
# add extra features in test set to sync with training set
if pad_features:
cols_to_add = set(train_features.columns) - set(features.columns)
shape=(len(features), len(cols_to_add))
features_to_pad = pd.DataFrame(
np.full(shape, 'no'), index=features.index, columns=cols_to_add)
features = features.join(features_to_pad)
return features
def _binarize_list_feature(series, prefix, train_features=None,
presence_threshold=0.0):
mlb = MultiLabelBinarizer()
binarized = pd.DataFrame(
mlb.fit_transform(series),
columns=[prefix + class_.replace(' ', '_') for class_ in mlb.classes_],
index=series.index)
if not (presence_threshold <= 0.0) or train_features is not None:
if train_features is not None:
cols_to_group = [col for col in binarized.columns if col not in
train_features.columns]
else:
cols_to_group = binarized.mean() < presence_threshold
cols_to_group = cols_to_group[cols_to_group.values].index.tolist()
# look for at least one '1' value in the row for a physicist
if cols_to_group:
other_col = binarized[cols_to_group].applymap(
lambda val: True if val == 1 else False).any(axis='columns')
other_col.name = _series_name(series.name, prefix)
binarized = binarized.drop(cols_to_group, axis='columns').join(other_col)
binarized = binarized.applymap(lambda val: 'yes' if val == 1 else 'no')
return binarized
def _series_name(name, prefix):
if name.endswith('alpha_3_codes'):
other_name = '***'
elif name.endswith('continent_codes'):
other_name = '**'
else:
other_name = '*'
return prefix + other_name
# -
train_features = binarize_list_features(train_features, presence_threshold=0.01)
assert(len(train_features.columns) == 157)
train_features.head()
validation_features = binarize_list_features(validation_features, train_features=train_features)
assert(len(validation_features.columns) == 147)
validation_features.head()
# So there are more features in the training set than in the validation set. So what are these extra features?
train_features.columns.difference(validation_features.columns).tolist()
# Working on the Manhattan Project certainly makes sense! OK let's reduce the training and validation set to the common set of features amongst them.
feature_cols = train_features.columns.intersection(validation_features.columns)
assert(validation_features.equals(validation_features[feature_cols]))
train_features = train_features[feature_cols]
assert((len(train_features.columns) == len(validation_features.columns)))
assert(sorted(train_features.columns.tolist()) == sorted(validation_features.columns.tolist()))
train_features.head()
# There are less features in the test set so let's "pad" the remaining binary features with "no" to ensure that the features are identical between the training and test sets.
test_features = binarize_list_features(test_features, train_features=train_features, pad_features=True)
assert(sorted(test_features.columns.tolist()) == sorted(train_features.columns.tolist()))
test_features.head()
# The features almost look good now, but there is one thing that is troubling. The mix of binary and ordinal variables complicates matters when it comes to machine learning. There are issues related to the following:
# - [How to correctly scale features for machine learning algorithms](https://stats.stackexchange.com/questions/69568/whether-to-rescale-indicator-binary-dummy-predictors-for-lasso)?
# - [Difficulty of interpretability of coefficients in generalized linear models](https://andrewgelman.com/2009/07/11/when_to_standar/)
# - [The bias introduced by lower importance given towards binary variables in random forests](https://roamanalytics.com/2016/10/28/are-categorical-variables-getting-lost-in-your-random-forests/)
#
# We would like to avoid these issues altogether.
#
# Seeing as the majority of the features are binary, it makes sense to convert the ordinal variables to binary variables. However, we do not want to lose the ordinal information that is present in these variables. The [rank-hot encoder](http://scottclowe.com/2016-03-05-rank-hot-encoder/) is an encoding that converts ordinal variables to binary variables whilst maintaining the ordinal information. *<NAME>*, PhD student studying neuroinformatics at the University of Edinburgh, explains that "the **rank-hot encoder** is similar to a *one-hot encoder*, except every feature up to and including the current rank is hot." He illustrates this with the following example:
#
# <table>
# <thead>
# <tr>
# <th>Satisfaction</th>
# <th>Rank Index</th>
# <th>One-Hot Encoding</th>
# <th>Rank-Hot Encoding</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>Very bad</td>
# <td>0</td>
# <td><code class="highlighter-rouge">[1, 0, 0, 0, 0]</code></td>
# <td><code class="highlighter-rouge">[0, 0, 0, 0]</code></td>
# </tr>
# <tr>
# <td>Bad</td>
# <td>1</td>
# <td><code class="highlighter-rouge">[0, 1, 0, 0, 0]</code></td>
# <td><code class="highlighter-rouge">[1, 0, 0, 0]</code></td>
# </tr>
# <tr>
# <td>Neutral</td>
# <td>2</td>
# <td><code class="highlighter-rouge">[0, 0, 1, 0, 0]</code></td>
# <td><code class="highlighter-rouge">[1, 1, 0, 0]</code></td>
# </tr>
# <tr>
# <td>Good</td>
# <td>3</td>
# <td><code class="highlighter-rouge">[0, 0, 0, 1, 0]</code></td>
# <td><code class="highlighter-rouge">[1, 1, 1, 0]</code></td>
# </tr>
# <tr>
# <td>Very good</td>
# <td>4</td>
# <td><code class="highlighter-rouge">[0, 0, 0, 0, 1]</code></td>
# <td><code class="highlighter-rouge">[1, 1, 1, 1]</code></td>
# </tr>
# </tbody>
# </table>
#
# He goes on to say, "Instead of answering the query “Is the satisfaction x?”, the entries in a rank-hot encoder tell us “Is the satisfaction level at least x?”. This representation of the data allows a linear model to explain the effect of a high-rank as the additive composition of the effect of each rank in turn."
#
# Sounds very useful doesn't it! Plus there are some other very nice properties of this encoding scheme that are explained in the blog. The main cons of rank-hot encoding, which are shared with one-hot encoding, are:
#
# - The feature space gets larger.
# - Information is lost whenever a categorical value is observed in a new instance (i.e. in the test set) that was not observed in the training (or validation) set.
#
# However, the benefits mentioned earlier are so important that they outweigh these downsides. Plus there are ways of dealing with the increase in the size of the feature space. OK let's go ahead and **rank-hot encode** the ordinal features.
ordinal_cols = [col for col in train_features.columns if col.startswith('num_')]
enc = OneHotEncoder(categories='auto', sparse=False, dtype='int64', handle_unknown='ignore')
enc.fit(train_features[ordinal_cols].append(validation_features[ordinal_cols]))
train_features = rank_hot_encode(train_features, enc, columns=ordinal_cols)
train_features = train_features.replace({0: 'no', 1: 'yes'})
assert(len(train_features.columns) == 206)
assert(train_features.select_dtypes('int64').empty)
assert(all(train_features.notna()))
train_features.head()
validation_features = rank_hot_encode(validation_features, enc, columns=ordinal_cols)
validation_features = validation_features.replace({0: 'no', 1: 'yes'})
assert(sorted(validation_features.columns.tolist()) == sorted(train_features.columns.tolist()))
assert(validation_features.select_dtypes('int64').empty)
assert(all(validation_features.notna()))
validation_features.head()
test_features = rank_hot_encode(test_features, enc, columns=ordinal_cols)
test_features = test_features.replace({0: 'no', 1: 'yes'})
assert(sorted(test_features.columns.tolist()) == sorted(train_features.columns.tolist()))
assert(test_features.select_dtypes('int64').empty)
assert(all(test_features.notna()))
test_features.head()
# The following columns in the training features have no variation in the values. Nothing can be learnt from these features, so let's drop them.
no_variation = (train_features != 'no').any(axis='rows')
no_variation = no_variation[no_variation == False]
assert(len(no_variation) == 3)
no_variation
train_features = train_features.drop(no_variation.index, axis='columns')
assert(len(train_features) == len(train_physicists))
assert(len(train_features.columns.tolist()) == 203)
assert(len(train_features.select_dtypes('object').columns) == len(train_features.columns.tolist()))
assert(all(train_features.notna()))
train_features.head()
validation_features = validation_features.drop(no_variation.index, axis='columns')
assert(len(validation_features) == len(validation_physicists))
assert(sorted(validation_features.columns.tolist()) == sorted(train_features.columns.tolist()))
assert(len(validation_features.select_dtypes('object').columns) == len(validation_features.columns.tolist()))
assert(all(validation_features.notna()))
validation_features.head()
test_features = test_features.drop(no_variation.index, axis='columns')
assert(len(test_features) == len(test_physicists))
assert(sorted(test_features.columns.tolist()) == sorted(train_features.columns.tolist()))
assert(len(test_features.select_dtypes('object').columns) == len(test_features.columns.tolist()))
assert(all(test_features.notna()))
test_features.head()
# Let's take a quick look at the features that remain.
sorted(train_features.drop('full_name', axis='columns').columns.tolist())
# The binary encoding has increased the dimensionality of the problem. There are now 202
# features (excluding the `full_name`) for 542 observations in the training set, 192 observations in the validation set and 193 observations in the test set. A model that is fit to such data could be prone to overfitting and a dimensionality reduction on this data may be warranted.
# ## Persisting the Data
#
# Now we have the training, validation and test features dataframes, let's persist them for future use.
train_features = train_features.reindex(sorted(train_features.columns), axis='columns')
train_features.head()
validation_features = validation_features.reindex(sorted(validation_features.columns), axis='columns')
validation_features.head()
test_features = test_features.reindex(sorted(test_features.columns), axis='columns')
test_features.head()
train_features.to_csv('../data/processed/train-features.csv', index=False)
validation_features.to_csv('../data/processed/validation-features.csv', index=False)
test_features.to_csv('../data/processed/test-features.csv', index=False)
| nobel_physics_prizes/notebooks/3.0-build-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
# # Examples
using MatchingMarkets
# ## <NAME> Sotomayor (1990)
# ### Example 2.9
m, n = 5, 4;
m_prefs = [
1, 2, 3, 4, 0,
4, 2, 3, 1, 0,
4, 3, 1, 2, 0,
1, 4, 3, 2, 0,
1, 2, 4, 0, 3,
]
m_prefs = reshape(m_prefs, n+1, m)
f_prefs = [
2, 3, 1, 4, 5, 0,
3, 1, 2, 4, 5, 0,
5, 4, 1, 2, 3, 0,
1, 4, 5, 2, 3, 0,
]
f_prefs = reshape(f_prefs, m+1, n)
deferred_acceptance(m_prefs, f_prefs)
deferred_acceptance(f_prefs, m_prefs)
# ### Example 2.17
m = n = 4;
m_prefs = [
1, 2, 3, 4, 0,
2, 1, 4, 3, 0,
3, 4, 1, 2, 0,
4, 3, 2, 1, 0,
]
m_prefs = reshape(m_prefs, n+1, m)
f_prefs = [
4, 3, 2, 1, 0,
3, 4, 1, 2, 0,
2, 1, 4, 3, 0,
1, 2, 3, 4, 0,
]
f_prefs = reshape(f_prefs, m+1, n)
deferred_acceptance(m_prefs, f_prefs)
deferred_acceptance(f_prefs, m_prefs)
# ### Example 5.24
m_caps = fill(2, m)
f_caps = m_caps
deferred_acceptance(m_prefs, f_prefs, m_caps, f_caps)
deferred_acceptance(f_prefs, m_prefs, f_caps, m_caps)
# ### Page 162
m, n = 7, 5;
s_prefs = [
5, 1, 0, 2, 3, 4,
2, 5, 1, 0, 3, 4,
3, 1, 0, 2, 4, 5,
4, 1, 0, 2, 3, 5,
1, 2, 0, 3, 4, 5,
1, 3, 0, 2, 4, 5,
1, 3, 4, 0, 2, 6,
]
s_prefs = reshape(s_prefs, n+1, m)
c_prefs = [
1, 2, 3, 4, 5, 6, 7, 0,
5, 2, 0, 1, 3, 4, 6, 7,
6, 7, 3, 0, 1, 2, 4, 5,
7, 4, 0, 1, 2, 3, 5, 6,
2, 1, 0, 3, 4, 5, 6, 7,
]
c_prefs = reshape(c_prefs, m+1, n)
caps = ones(Int, n)
caps[1] = 3
caps
deferred_acceptance(s_prefs, c_prefs, caps)
deferred_acceptance(s_prefs, c_prefs, caps, SProposing)
deferred_acceptance(s_prefs, c_prefs, caps, CProposing)
# ## <NAME> (1989)
# ### Section 1.6.5
m, n = 11, 5;
s_prefs = [
3, 1, 5, 4, 0, 2,
1, 3, 4, 2, 5, 0,
4, 5, 3, 1, 2, 0,
3, 4, 1, 5, 0, 2,
1, 4, 2, 0, 3, 5,
4, 3, 2, 1, 5, 0,
2, 5, 1, 3, 0, 4,
1, 3, 2, 5, 4, 0,
4, 1, 5, 0, 2, 3,
3, 1, 5, 2, 4, 0,
5, 4, 1, 3, 2, 0,
]
s_prefs = reshape(s_prefs, n+1, m)
c_prefs = [
3, 7, 9, 11, 5, 4, 10, 8, 6, 1, 2, 0,
5, 7, 10, 6, 8, 2, 3, 11, 0, 1, 4, 9,
11, 6, 8, 3, 2, 4, 7, 1, 10, 0, 5, 9,
10, 1, 2, 11, 4, 9, 5, 3, 6, 8, 0, 7,
2, 4, 10, 7, 6, 1, 8, 3, 11, 9, 0, 5,
]
c_prefs = reshape(c_prefs, m+1, n)
caps = [4, 1, 3, 2, 1];
deferred_acceptance(s_prefs, c_prefs, caps)
deferred_acceptance(s_prefs, c_prefs, caps, SProposing)
deferred_acceptance(s_prefs, c_prefs, caps, CProposing)
# The book claims that the following matching $M_7$ is the hospital-optimal
# (college-optimal in our language) stable matching:
# $$
# \begin{array}
# ~ & r_1 & r_2 & r_3 & r_4 & r_5 & r_6 & r_7 & r_8 & r_9 & r_{10} & r_{11} \\
# M_7 & h_4 & h_4 & h_3 & h_1 & h_1 & h_3 & h_2 & h_3 & h_1 & h_5 & h_1
# \end{array}
# $$
#
# but the above matching, say $M_8$, appears to be the hospital-optimal matching:
# $$
# \begin{array}
# ~ & r_1 & r_2 & r_3 & r_4 & r_5 & r_6 & r_7 & r_8 & r_9 & r_{10} & r_{11} \\
# M_8 & h_4 & h_5 & h_3 & h_1 & h_1 & h_3 & h_2 & h_3 & h_1 & h_4 & h_1
# \end{array}
# $$
| examples/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Conclusion
#
# We covered a lot of ground in this chapter from the basic ideas of probability to fallacies in probabilistic reasoning. Still, probability is a fundamental tool in data science. Returning to our definition of data science, it says:
#
# > Data science is the application of math and computers to solve problems that stem from a lack of knowledge, constrained by the small number of people with any interest in the answers.
#
# The fundamental problem of Science is extrapolating general conclusions from specific data sets. That is, inference. Probability is our fundamental tool for dealing with the problem of inference because it allows us to model our uncertainty in a very rigorous way. Still...it's not a panacea. It does not magically make inference deductive instead of inductive.
#
# Although the types, rules and laws of probability are important, the most important part of this chapter is Bayes Rule (or Theorem). Bayes Rule allows us tells us how to update our beliefs based on evidence and forms the basis for Statistical Inference discussed in this book.
# ## Review
#
# *When asked to give an example, do not use dice, coins or cards (or any gambling device).*
#
# 1. Why is a coin flip deterministic but we still need probability to model it?
# 2. What is our working definition of probability?
# 3. If $A = {a_1, a_2, a_3}$ and $B = {b_1, b_2}$, then answer the following questions:
# 1. What does $P(A, B)$ denote?
# 2. What does $P(A)$ denote? How did we arrive at it from $P(A, B)$?
# 3. What does $P(A=a_1$) denote?
# 4. What does $P(a_1)$ denote? Why should we be careful when using a "shorthand"?
# 5. What does $P(A|B)$ denote? How many probability distributions does it represent?
# 6. What does $P(A|B)P(B)$ denote? Write it out.
# 7. Express $P(A)$ using the Total Probability.
# 4. What is the difference between an **outcome** and an **event**? Give an example not shown elsewhere.
# 5. Give an example of the **independence** of two outcomes.
# 6. Give an example of **conditional independence** of three outcomes.
# 7. What is the **Gambler's Fallacy**?
# 8. What is the **Inverse Probability Fallacy**?
# 9. What is the **Prosecutor's Fallacy**?
# 10. Joe has been randomly selected for drug testing from a population that has about 3% heroin use. Joe tests positive for heroin use ($u$). The test used correctly identifies users 95% of the time $P(+|u) = 0.95$ and correctly identifies non-users 90% of the time $P(-|c) = 0.90$ ($c$ for "clean"). What is the probability that Joe is using heroin $P(u|+)$? What are the increment and total evidences in this problem?
| fundamentals_2018.9/probability/conclusion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ! cd ../ && python setup.py develop
# +
import numpy as np
import pandas as pd
import logging
# %load_ext autoreload
# %autoreload 2
from mercari import nn_conv1d
# +
small = False
df_train, df_test = nn_conv1d.load_data(small=small)
df_train, df_test, target_scaler, embedding_params = nn_conv1d.preprocess(df_train, df_test)
train_ids, valid_ids, submission = nn_conv1d.load_valid_ids(df_train)
X = nn_conv1d.get_keras_data(df_train)
y = df_train['target']
# -
train_nonzero = (df_train['price'].iloc[train_ids] != 0).values.nonzero()[0]
X_train = nn_conv1d.slice_keras_data(X, train_ids)
X_train = nn_conv1d.slice_keras_data(X_train, train_nonzero)
y_train = y.values[train_ids][train_nonzero]
X_valid = nn_conv1d.slice_keras_data(X, valid_ids)
y_valid = y[valid_ids]
train_kwargs = dict(epochs=3, batch_size=1024)
model_rmsles, model_predictions = [], []
for i in range(1):
model, optimizer = nn_conv1d.get_model(
X_train, embedding_params, lr=0.004)
rmsle = nn_conv1d.train_model(
model, optimizer,
X_train=X_train, y_train=y_train,
X_valid=X_valid, y_valid=y_valid,
target_scaler=target_scaler,
**train_kwargs)
model_rmsles.append(rmsle)
y_pred_valid = model.predict(X_valid, batch_size=nn_conv1d.PRED_BATCH)
model_predictions.append(y_pred_valid)
logging.info(f'rmsle: {rmsle:.5f}')
submission['price'] = nn_conv1d.target_to_price(
np.mean(model_predictions, axis=0), target_scaler)
submission.to_csv(f'valid_nn_conv1d_{i}.csv', index=False)
logging.info(
f'Mean model RMSLE: {np.mean(model_rmsles)} '
f'± {1.96 * np.std(model_rmsles)}')
mean_rmsle = nn_conv1d.get_rmsle(
df_train['price'][valid_ids],
nn_conv1d.target_to_price(np.mean(model_predictions, axis=0),
target_scaler))
logging.info(f'RMSLE of prediction mean: {mean_rmsle}')
| notebooks/nn_conv1d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch_lifestream
# language: python
# name: pytorch_lifestream
# ---
# %cd ../
import random
import math
def rand_log(a, b):
la, lb = math.log(a), math.log(b)
return round(math.exp(random.random() * (lb - la) + la), 6)
def rand_int(a, b):
return round(random.random() * (b - a) + a, 6)
params = [
{
'lambd': rand_log(0.001, 0.1),
'hidden_size': int(rand_log(256, 2048)),
'prj_size': 0 if random.random() < 0.5 else int(rand_log(64, 512)),
'batch_size': int(rand_log(32, 512)),
'lr': rand_log(0.00002, 0.2),
'weight_decay': 0 if random.random() < 0.5 else rand_log(0.0001, 0.1),
'step_size': [10, 20, 30][random.randint(0, 2)],
'step_gamma': rand_int(0.4, 1.0),
}
for _ in range(30)
]
with open('bin/scen_bt_tuning_1.sh', 'wt') as f:
for p in params:
name = '-'.join([f'{k}_{v}' for k, v in p.items()])
if p['prj_size'] == 0:
prj_s = ' '
else:
prj_s = f""" "params.head_layers=[[Linear, {{in_features: {p['hidden_size']}, out_features: {p['prj_size']}, bias: false}}], [BatchNorm1d, {{num_features: {p['prj_size']}}}], [ReLU, {{ }}], [Linear, {{in_features: {p['prj_size']}, out_features: {p['prj_size']}, bias: false}}], [BatchNorm1d, {{num_features: {p['prj_size']}, affine: False}}]]" """
f.write(f"""
export SC_SUFFIX="bt_tuning_{name}"
python ../../pl_train_module.py \\
logger_name=${{SC_SUFFIX}} \\
params.train.lambd={p['lambd']} \\
params.rnn.hidden_size="{p['hidden_size']}" \\
{prj_s} \\
data_module.train.batch_size={p['batch_size']} \\
params.train.lr={p['lr']} \\
params.train.weight_decay={p['weight_decay']} \\
params.lr_scheduler.step_size={p['step_size']} \\
params.lr_scheduler.step_gamma={p['step_gamma']} \\
model_path="models/gender_mlm__$SC_SUFFIX.p" \\
--conf "conf/barlow_twins_params.hocon"
python ../../pl_inference.py \
inference_dataloader.loader.batch_size=500 \\
model_path="models/gender_mlm__$SC_SUFFIX.p" \\
output.path="data/emb__${{SC_SUFFIX}}" \\
--conf "conf/barlow_twins_params.hocon"
""")
# ```
# sh bin/scen_bt_tuning_3.sh
#
# # rm results/res_bt_tuning.txt
# # # rm -r conf/embeddings_validation.work/
# python -m embeddings_validation \
# --conf conf/embeddings_validation_short.hocon --workers 10 --total_cpu_count 20 \
# --conf_extra \
# 'report_file: "../results/res_bt_tuning.txt",
# auto_features: ["../data/emb__bt_tuning_*.pickle", "../data/barlow_twins_embeddings.pickle"]'
# less -S results/res_bt_tuning.txt
#
#
# ```
import pandas as pd
df = pd.read_fwf('results/res_bt_tuning.txt').iloc[10:95, :2]
df.columns = ['name', 'm']
df
df.iloc[0, 0]
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={0: 'k'}).set_index('k')['m'].reset_index() \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5), logx=True)
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={1: 'k'}).set_index('k')['m'].reset_index() \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5), logx=True)
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={2: 'k'}).set_index('k')['m'].reset_index() \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5))
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={3: 'k'}).set_index('k')['m'].reset_index() \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5), logx=True)
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={4: 'k'}).set_index('k')['m'].reset_index() \
.assign(k=lambda x: x['k'].str.replace('^', '-'), regex=False) \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5), logx=True)
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={5: 'k'}).set_index('k')['m'].reset_index() \
.assign(k=lambda x: x['k'].str.replace('^', '-'), regex=False) \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5))
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={6: 'k'}).set_index('k')['m'].reset_index() \
.assign(k=lambda x: x['k'].str.replace('^', '-'), regex=False) \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5))
pd.concat([
df.name.str.replace('e-', 'e^').str.split(r'-', expand=True),
df.m,
], axis=1).rename(columns={7: 'k'}).set_index('k')['m'].reset_index() \
.assign(k=lambda x: x['k'].str.replace('^', '-'), regex=False) \
.assign(n=lambda x: pd.to_numeric(x['k'].str.split('_').str[-1])) \
.set_index('n').sort_index()['m'].astype(float).plot(figsize=(12, 5))
# !cat bin/scen_bt_tuning_1.sh
| experiments/scenario_age_pred/notebooks/make_tuning_script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="1rRo8oNqZ-Rj"
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow as tf
print("TensorFlow version: {}".format(tf.__version__))
# + colab={} colab_type="code" id="p9kxxgzvzlyz"
model = tf.keras.Sequential()
model.add(tf.keras.layers.Dense(units=64, activation='relu', input_dim=100))
model.add(tf.keras.layers.Dense(units=10, activation='softmax'))
model.summary()
| Exercise01/Exercise01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 调用模型进行测试
# +
from xgboost import XGBClassifier
import pandas as pd
import numpy as np
# -
# ## 读取数据
# path to where the data lies
dpath = './data/'
test = pd.read_csv(dpath +"RentListingInquries_FE_test.csv")
test.head()
# ## 准备数据
# +
# 提供的数据中无listing_id
# test_id = test['listing_id']
# X_test = test.drop([ "listing_id"], axis=1)
X_test = test
# -
# ## load 训练好的模型
# +
#保存数据
import pickle
xgb = pickle.load(open("xgb_model.pkl", 'rb'))
# -
# ## 在测试上测试,并生成测试结果提交文件
# +
y_test_pred = xgb.predict_proba(X_test)
out_df1 = pd.DataFrame(y_test_pred)
out_df1.columns = ["high", "medium", "low"]
# out_df = pd.concat([test_id,out_df1], axis = 1)
out_df = out_df1
out_df.to_csv("xgb_Rent.csv", index=False)
# -
| 8_xgboost_Rental_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# first we need to load the data from the .mat file recorded using HEKA, which exported in .mat
# first we gonna try scipy as https://scipy-cookbook.readthedocs.io/items/Reading_mat_files.html
from scipy.io import loadmat
from tkinter.filedialog import askopenfilenames
from tkinter import Tk
import os
root1 = Tk()
filez = askopenfilenames(parent = root1, title = 'Choose file')
for fullFileName in root1.tk.splitlist(filez):
filename = fullFileName
(root, ext) =os.path.splitext(filename)
print(filename)
#x = loadmat('./')
# -
| Python_script/Data_loading.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Based on Tutorial
# https://www.analyticsvidhya.com/blog/2019/06/comprehensive-guide-text-summarization-using-deep-learning-python/
import numpy as np
import pandas as pd
import re
from bs4 import BeautifulSoup
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from nltk.corpus import stopwords
from tensorflow.keras.layers import Input, LSTM, Embedding, Dense, Concatenate, TimeDistributed, Bidirectional
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import EarlyStopping
import warnings
pd.set_option("display.max_colwidth", 200)
warnings.filterwarnings("ignore")
# Load data
data=pd.read_csv("../common-data/Reviews.csv")
# Checking data
data.head
# Checking data stats
data.info()
# Checking data and cleaning
data.drop_duplicates(subset=['Text'],inplace=True) #dropping duplicates
data.dropna(axis=0,inplace=True) #dropping na
data.info()
# +
# Data cleaning
contraction_mapping = {"ain't": "is not", "aren't": "are not","can't": "cannot", "'cause": "because", "could've": "could have", "couldn't": "could not",
"didn't": "did not", "doesn't": "does not", "don't": "do not", "hadn't": "had not", "hasn't": "has not", "haven't": "have not",
"he'd": "he would","he'll": "he will", "he's": "he is", "how'd": "how did", "how'd'y": "how do you", "how'll": "how will", "how's": "how is",
"I'd": "I would", "I'd've": "I would have", "I'll": "I will", "I'll've": "I will have","I'm": "I am", "I've": "I have", "i'd": "i would",
"i'd've": "i would have", "i'll": "i will", "i'll've": "i will have","i'm": "i am", "i've": "i have", "isn't": "is not", "it'd": "it would",
"it'd've": "it would have", "it'll": "it will", "it'll've": "it will have","it's": "it is", "let's": "let us", "ma'am": "madam",
"mayn't": "may not", "might've": "might have","mightn't": "might not","mightn't've": "might not have", "must've": "must have",
"mustn't": "must not", "mustn't've": "must not have", "needn't": "need not", "needn't've": "need not have","o'clock": "of the clock",
"oughtn't": "ought not", "oughtn't've": "ought not have", "shan't": "shall not", "sha'n't": "shall not", "shan't've": "shall not have",
"she'd": "she would", "she'd've": "she would have", "she'll": "she will", "she'll've": "she will have", "she's": "she is",
"should've": "should have", "shouldn't": "should not", "shouldn't've": "should not have", "so've": "so have","so's": "so as",
"this's": "this is","that'd": "that would", "that'd've": "that would have", "that's": "that is", "there'd": "there would",
"there'd've": "there would have", "there's": "there is", "here's": "here is","they'd": "they would", "they'd've": "they would have",
"they'll": "they will", "they'll've": "they will have", "they're": "they are", "they've": "they have", "to've": "to have",
"wasn't": "was not", "we'd": "we would", "we'd've": "we would have", "we'll": "we will", "we'll've": "we will have", "we're": "we are",
"we've": "we have", "weren't": "were not", "what'll": "what will", "what'll've": "what will have", "what're": "what are",
"what's": "what is", "what've": "what have", "when's": "when is", "when've": "when have", "where'd": "where did", "where's": "where is",
"where've": "where have", "who'll": "who will", "who'll've": "who will have", "who's": "who is", "who've": "who have",
"why's": "why is", "why've": "why have", "will've": "will have", "won't": "will not", "won't've": "will not have",
"would've": "would have", "wouldn't": "would not", "wouldn't've": "would not have", "y'all": "you all",
"y'all'd": "you all would","y'all'd've": "you all would have","y'all're": "you all are","y'all've": "you all have",
"you'd": "you would", "you'd've": "you would have", "you'll": "you will", "you'll've": "you will have",
"you're": "you are", "you've": "you have"}
"""
Convert everything to lowercase
Remove HTML tags
Contraction mapping
Remove (‘s)
Remove any text inside the parenthesis ( )
Eliminate punctuations and special characters
Remove stopwords
"""
stop_words = set(stopwords.words('english'))
def text_cleaner(text):
newString = text.lower()
newString = BeautifulSoup(newString, "lxml").text
newString = re.sub(r'\([^)]*\)', '', newString)
newString = re.sub('"','', newString)
newString = ' '.join([contraction_mapping[t] if t in contraction_mapping else t for t in newString.split(" ")])
newString = re.sub(r"'s\b","",newString)
newString = re.sub("[^a-zA-Z]", " ", newString)
tokens = [w for w in newString.split() if not w in stop_words]
return (" ".join(tokens)).strip()
# -
# Clean text
cleaned_text = []
for t in data['Text']:
cleaned_text.append(text_cleaner(t))
data['Cleaned_Text']=cleaned_text
# Clean summary text
cleaned_text = []
for t in data['Summary']:
cleaned_text.append(text_cleaner(t))
data['Cleaned_Summary']=cleaned_text
data.head()
data['Cleaned_Summary'] = data['Cleaned_Summary'].apply(lambda x : '_START_ '+ x + ' _END_')
# Printing cleaned text and summary
for i in range(5):
print("Review:",data['Cleaned_Text'][i])
print("Summary:",data['Cleaned_Summary'][i])
print("\n")
# ### Learning Summary
max_len_text=80
max_len_summary=10
from sklearn.model_selection import train_test_split
x_tr,x_val,y_tr,y_val=train_test_split(data['Cleaned_Text'],data['Cleaned_Summary'],test_size=0.1,random_state=0,shuffle=True)
# +
# Text tokenizer
x_tokenizer = Tokenizer()
x_tokenizer.fit_on_texts(list(x_tr))
#convert text sequences into integer sequences
x_tr = x_tokenizer.texts_to_sequences(x_tr)
x_val = x_tokenizer.texts_to_sequences(x_val)
#padding zero upto maximum length
x_tr = pad_sequences(x_tr, maxlen=max_len_text, padding='post')
x_val = pad_sequences(x_val, maxlen=max_len_text, padding='post')
x_voc_size = len(x_tokenizer.word_index) +1
# +
# Summary tokenizer
y_tokenizer = Tokenizer()
y_tokenizer.fit_on_texts(list(y_tr))
#convert summary sequences into integer sequences
y_tr = y_tokenizer.texts_to_sequences(y_tr)
y_val = y_tokenizer.texts_to_sequences(y_val)
#padding zero upto maximum length
y_tr = pad_sequences(y_tr, maxlen=max_len_summary, padding='post')
y_val = pad_sequences(y_val, maxlen=max_len_summary, padding='post')
y_voc_size = len(y_tokenizer.word_index) +1
# -
# Attention
from attention import AttentionLayer
# +
from keras import backend as K
K.clear_session()
latent_dim = 500
# Encoder
encoder_inputs = Input(shape=(max_len_text,))
enc_emb = Embedding(x_voc_size, latent_dim,trainable=True)(encoder_inputs)
#LSTM 1
encoder_lstm1 = LSTM(latent_dim,return_sequences=True,return_state=True)
encoder_output1, state_h1, state_c1 = encoder_lstm1(enc_emb)
#LSTM 2
encoder_lstm2 = LSTM(latent_dim,return_sequences=True,return_state=True)
encoder_output2, state_h2, state_c2 = encoder_lstm2(encoder_output1)
#LSTM 3
encoder_lstm3=LSTM(latent_dim, return_state=True, return_sequences=True)
encoder_outputs, state_h, state_c= encoder_lstm3(encoder_output2)
# Set up the decoder.
decoder_inputs = Input(shape=(None,))
dec_emb_layer = Embedding(y_voc_size, latent_dim,trainable=True)
dec_emb = dec_emb_layer(decoder_inputs)
#LSTM using encoder_states as initial state
decoder_lstm = LSTM(latent_dim, return_sequences=True, return_state=True)
decoder_outputs,decoder_fwd_state, decoder_back_state = decoder_lstm(dec_emb,initial_state=[state_h, state_c])
#Attention Layer
attn_layer = AttentionLayer(name='attention_layer')
attn_out, attn_states = attn_layer([encoder_outputs, decoder_outputs])
# Concat attention output and decoder LSTM output
decoder_concat_input = Concatenate(axis=-1, name='concat_layer')([decoder_outputs, attn_out])
#Dense layer
decoder_dense = TimeDistributed(Dense(y_voc_size, activation='softmax'))
decoder_outputs = decoder_dense(decoder_concat_input)
# Define the model
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
model.summary()
# -
# Setting up and running the model
model.compile(optimizer='rmsprop', loss='sparse_categorical_crossentropy')
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1)
history=model.fit([x_tr,y_tr[:,:-1]], y_tr.reshape(y_tr.shape[0],y_tr.shape[1], 1)[:,1:] ,epochs=5,callbacks=[es],batch_size=512, validation_data=([x_val,y_val[:,:-1]], y_val.reshape(y_val.shape[0],y_val.shape[1], 1)[:,1:]))
from matplotlib import pyplot
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
pyplot.show()
# ### Deconding results
reverse_target_word_index=y_tokenizer.index_word
reverse_source_word_index=x_tokenizer.index_word
target_word_index=y_tokenizer.word_index
print(target_word_index)
print (reverse_target_word_index)
# +
# encoder inference
encoder_model = Model(inputs=encoder_inputs,outputs=[encoder_outputs, state_h, state_c])
# decoder inference
# Below tensors will hold the states of the previous time step
decoder_state_input_h = Input(shape=(latent_dim,))
decoder_state_input_c = Input(shape=(latent_dim,))
decoder_hidden_state_input = Input(shape=(max_len_text,latent_dim))
# Get the embeddings of the decoder sequence
dec_emb2= dec_emb_layer(decoder_inputs)
# To predict the next word in the sequence, set the initial states to the states from the previous time step
decoder_outputs2, state_h2, state_c2 = decoder_lstm(dec_emb2, initial_state=[decoder_state_input_h, decoder_state_input_c])
#attention inference
attn_out_inf, attn_states_inf = attn_layer([decoder_hidden_state_input, decoder_outputs2])
decoder_inf_concat = Concatenate(axis=-1, name='concat')([decoder_outputs2, attn_out_inf])
# A dense softmax layer to generate prob dist. over the target vocabulary
decoder_outputs2 = decoder_dense(decoder_inf_concat)
# Final decoder model
decoder_model = Model(
[decoder_inputs] + [decoder_hidden_state_input,decoder_state_input_h, decoder_state_input_c],
[decoder_outputs2] + [state_h2, state_c2])
# -
def decode_sequence(input_seq):
# Encode the input as state vectors.
e_out, e_h, e_c = encoder_model.predict(input_seq)
# Generate empty target sequence of length 1.
target_seq = np.zeros((1,1))
# Choose the 'start' word as the first word of the target sequence
target_seq[0, 0] = target_word_index['start']
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + [e_out, e_h, e_c])
# Sample a token
sampled_token_index = np.argmax(output_tokens[0, -1, :])
sampled_token = reverse_target_word_index[sampled_token_index]
if(sampled_token!='end'):
decoded_sentence += ' '+sampled_token
# Exit condition: either hit max length or find stop word.
if (sampled_token == 'end' or len(decoded_sentence.split()) >= (max_len_summary-1)):
stop_condition = True
# Update the target sequence (of length 1).
target_seq = np.zeros((1,1))
target_seq[0, 0] = sampled_token_index
# Update internal states
e_h, e_c = h, c
return decoded_sentence
# +
def seq2summary(input_seq):
newString=''
for i in input_seq:
if((i!=0 and i!=target_word_index['start']) and i!=target_word_index['end']):
newString=newString+reverse_target_word_index[i]+' '
return newString
def seq2text(input_seq):
newString=''
for i in input_seq:
if(i!=0):
newString=newString+reverse_source_word_index[i]+' '
return newString
# -
import sys
for i in range(10):
# for i in range(len(x_val)):
print("Review:",seq2text(x_val[i]))
print("Original summary:",seq2summary(y_val[i]))
try:
print("Predicted summary:",decode_sequence(x_val[i].reshape(1,max_len_text)))
except: # catch *all* exceptions
print("Predicted summary: some issue.")
print("\n")
| l20-textsumm/ExploreAbstractive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install simpletransformers
from simpletransformers.classification import ClassificationModel, ClassificationArgs
import pandas as pd
import logging
import jsonlines
import matplotlib.pyplot as plt
import numpy as np
import json
import nltk
from nltk.tokenize import TweetTokenizer
from nltk.corpus import stopwords
import re
import itertools
import emoji
import sklearn
from simpletransformers.language_representation import RepresentationModel
from sklearn.feature_extraction.text import CountVectorizer
# get texts that are related to the source tweet
def get_valid_text(tweet):
all_id = []
for item in tweet:
all_id.append(item['id'])
invalid_id = []
for item in tweet:
if item['in_reply_to_status_id'] not in all_id:
invalid_id.append(item['id'])
text = tweet[0]['text']
for i in range(1, len(tweet)):
if tweet[i]['in_reply_to_status_id'] in all_id and tweet[i]['in_reply_to_status_id'] not in invalid_id:
text = text + ' ' + tweet[i]['text']
return text
# +
# load data from each json file
train_list = []
with open('train.data.jsonl', 'r') as file:
for item in jsonlines.Reader(file):
train_list.append(item)
with open ('train.label.json', 'r') as file1:
train_labels = json.loads(file1.read())
train_data_list = []
for tweet in train_list:
tweetId = tweet[0]['id_str']
text = get_valid_text(tweet)
text = text.replace('\n', '').replace('\r', '')
if train_labels[tweetId] == 'non-rumour':
train_data_list.append('1\t' + text + '\n')
elif train_labels[tweetId] == 'rumour':
train_data_list.append('0' + '\t' + text + '\n')
dev_list = []
with open('dev.data.jsonl', 'r') as file:
for item in jsonlines.Reader(file):
dev_list.append(item)
with open ('dev.label.json', 'r') as file1:
dev_labels = json.loads(file1.read())
dev_data_list = []
for tweet in dev_list:
tweetId = tweet[0]['id_str']
text = get_valid_text(tweet)
text = text.replace('\n', '').replace('\r', '')
if dev_labels[tweetId] == 'non-rumour':
dev_data_list.append('1\t' + text + '\n')
elif dev_labels[tweetId] == 'rumour':
dev_data_list.append('0' + '\t' + text + '\n')
# +
test_list = []
with open('test.data.jsonl', 'r') as file:
for item in jsonlines.Reader(file):
test_list.append(item)
test_data_list = []
for tweet in test_list:
tweetId = tweet[0]['id_str']
text = get_valid_text(tweet)
text = text.replace('\n', '').replace('\r', '')
test_data_list.append(text + '\n')
# +
covid_list = []
with open('covid.data.jsonl', 'r') as file:
for item in jsonlines.Reader(file):
covid_list.append(item)
covid_data_list = []
for tweet in covid_list:
tweetId = tweet[0]['id_str']
text = get_valid_text(tweet)
text = text.replace('\n', '').replace('\r', '')
covid_data_list.append(text + '\n')
# +
# create data frames
train_df = []
for item in train_data_list:
item = item.split('\t')
train_df.append([item[1], int(item[0])])
train_df = pd.DataFrame(train_df, columns=['text', 'label'])
dev_df = []
for item in dev_data_list:
item = item.split('\t')
dev_df.append([item[1], int(item[0])])
dev_df = pd.DataFrame(dev_df, columns=['text', 'label'])
# -
test_df = []
for item in test_data_list:
test_df.append(item)
test_df = pd.DataFrame(test_df, columns=['text'])
covid_df = []
for item in covid_data_list:
covid_df.append(item)
covid_df = pd.DataFrame(covid_df, columns=['text'])
# tokenisation and normalisation
def preprocess(text):
stopw = set(stopwords.words('english'))
text = text.lower()
tt = TweetTokenizer()
text = tt.tokenize(text)
for i in range(len(text) - 1, -1, -1):
containAlphabets = False
for j in range(len(text[i])):
if ord(text[i][j]) >= 97 and ord(text[i][j]) <= 122:
containAlphabets = True
break
if not containAlphabets:
text.pop(i)
for word in text:
if word in stopw:
text.remove(word)
return ' '.join(text)
train_df['text'] = train_df['text'].apply(preprocess)
dev_df['text'] = dev_df['text'].apply(preprocess)
test_df['text'] = test_df['text'].apply(preprocess)
covid_df['text'] = covid_df['text'].apply(preprocess)
# over-sampling of the training data
train_non_rumour = train_df.loc[train_df['label'] == 1]
train_rumour = train_df.loc[train_df['label'] == 0]
train_df = train_df.append(train_rumour)
# the hyperparameters of BERT
train_args = {
'evaluate_during_training': True,
'logging_steps': 100,
'num_train_epochs': 3,
'evaluate_during_training_steps': 100,
'save_eval_checkpoints': False,
'train_batch_size': 32,
'eval_batch_size': 64,
'overwrite_output_dir': True,
'fp16': False,
'wandb_project': "visualization-demo"
}
# create the BERT model
bert_model = ClassificationModel('bert', 'bert-base-cased', num_labels=2, use_cuda=False, args=train_args)
# train the BERT model
bert_model.train_model(train_df, eval_df=dev_df)
# evaluate using dev_data
result, model_outputs, wrong_predictions = bert_model.eval_model(dev_df, acc=sklearn.metrics.accuracy_score)
# predict labels of test data
test_sentences = []
for text in test_df['text']:
test_sentences.append(text)
predictions, raw_outputs = bert_model.predict(test_sentences)
# predict labels of dev data
dev_sentences = []
for text in dev_df['text']:
dev_sentences.append(text)
dev_predictions, dev_raw_outputs = bert_model.predict(dev_sentences)
# predict labels of task2 COVID-19 data
covid_sentences = []
for text in covid_df['text']:
covid_sentences.append(text)
covid_predictions, covid_raw_outputs = bert_model.predict(covid_sentences)
# get output and convert them to json files
output = {}
for i in range(len(test_list)):
if predictions[i] == 0:
output[test_list[i][0]['id_str']] = 'rumour'
else:
output[test_list[i][0]['id_str']] = 'non-rumour'
dev_output = {}
for i in range(len(dev_list)):
if dev_predictions[i] == 0:
dev_output[dev_list[i][0]['id_str']] = 'rumour'
else:
dev_output[dev_list[i][0]['id_str']] = 'non-rumour'
with open('test-output.json', 'w') as f:
json.dump(output, f)
with open('dev-output.json', 'w') as f:
json.dump(dev_output, f)
covid_rumours = []
covid_nonrumours = []
for i in range(len(covid_sentences)):
if covid_predictions[i] == 0:
covid_rumours.append(covid_sentences[i])
else:
covid_nonrumours.append(covid_sentences[i])
# +
with open('covid_rumours.txt', 'w', encoding='utf-8') as f:
for sentence in covid_rumours:
f.write(sentence + '\n')
with open('covid_nonrumours.txt', 'w', encoding='utf-8') as f:
for sentence in covid_nonrumours:
f.write(sentence + '\n')
| COMP90042_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from skimage.color import rgb2gray
from skimage import io
from skimage.filters import gaussian
from skimage.segmentation import active_contour
fig_size=(7, 7)
# +
I = io.imread('./images/tema10_act1a.png', as_gray=False)
# Define an outline
s = np.linspace(0, 2*np.pi, 500)
x = 145 + 125*np.cos(s)
y = 100 + 50*np.sin(s)
init = np.array([x, y]).T
# Print image + outline
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(I, cmap=plt.cm.gray)
ax.plot(init[:, 0], init[:, 1], '--r', lw=3)
plt.show()
# +
snake = active_contour(gaussian(I, 4), init, alpha=0.015, beta=100, gamma=0.001, w_line=-10)
#snake = active_contour(gaussian(I, 1), init, alpha=0.1, w_edge=2, w_line=-1)
#snake = active_contour(gaussian(I, 5), init, alpha=0.015, beta=10, gamma=0.001)
# Print image + outline
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(I, cmap=plt.cm.gray)
ax.plot(init[:, 0], init[:, 1], '--r', lw=3)
ax.plot(snake[:, 0], snake[:, 1], '-b', lw=3)
plt.show()
# +
markers = rank.gradient(gaussian(I, 2), disk(1))
mask = markers <= 10
markers2 = markers.copy()
markers2[mask] = 0
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(mask, cmap=plt.cm.nipy_spectral, interpolation='nearest')
plt.show()
# -
img = gaussian(I, 10) - gaussian(I, 1)
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(1 - I, cmap=plt.cm.gray, interpolation='nearest')
plt.show()
# +
import skimage.morphology
img = skimage.morphology.closing(I < 100, disk(5))
bw = I < 100
bw[img] = 0.0
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(bw, cmap=plt.cm.binary, interpolation='nearest')
plt.show()
# -
img
I = io.imread('./images/tema10_act1b.png', as_gray=False)
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(I, cmap=plt.cm.gray)
plt.show()
I = io.imread('./images/tema10_act1c.png', as_gray=False)
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(I, cmap=plt.cm.gray)
plt.show()
# +
from skimage.filters import rank
from skimage.morphology import watershed, disk
markers = rank.gradient(gaussian(I, 3), disk(1))
mask = markers <= 10
markers2 = markers.copy()
markers2[mask] = 0
fig, ax = plt.subplots(figsize=fig_size)
ax.imshow(mask, cmap=plt.cm.nipy_spectral, interpolation='nearest')
plt.show()
# -
np.histogram(markers)
mask
| Percepcion_Computacional/Actividades/Lab1/Manuel_Pasieka_ActiveContour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="7b073cc577c4d8466b3cd192215c96714776b87a"
# # Prediticing spot prices for AWS EC2 Instances
#
#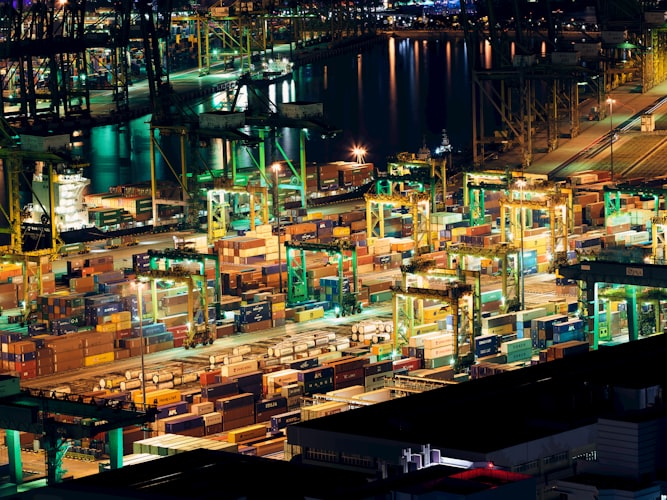 
# + [markdown] _uuid="7bad6db2cda184ce3cd4365e3c29e9b940c7cf99"
# # Table of Contents
#
# * Introduction
# * Background
# * Import libraries
# * EDA (Exploratory Data Analysis)
# * Cleaning
# * Implement Model
# * Conculsion on results
# + [markdown] _uuid="b1784ff7fe303a9161ddd9e820a331c1de9fd809"
# # Introduction
#
# The purpose of this experiment is to train a deep learning model to predict an outcome on time series data. I will be using the Fast.ai library for the model. More specifically, we will be predicting the Spot prices for specifc regions.
# + [markdown] _uuid="15face38841009cf72374099634e1c0af702c32f"
# # Background
#
# Amazon Web Services [(AWS)](https://aws.amazon.com) provides virtual computing environments via their EC2 service. You can launch instances with your favourite operating system, select pre-configured instance images or create your own. Why this is revelant to data sciensits is because generally to run deep learning models you need a machine with a good GPU. EC2 can be configured with a P2/P3 instance and can be configured with up to 8 or 16 GPUs respectively!
#
# However, you can request Spot Instance Pricing. Which basically charges you for the spot price that is in effect for the duration of your instance running time. They are adjusted based on long-term trends in supply and demand for Spot instance capacity. Spot instances can be discounted at up to 90% off compared to On-Demand pricing.
#
#
# Our goal will be to predict Spot pricing for the different global regions on offer:
#
# * US East
# * US West
# * South America (East)
# * EU (European Union) West
# * EU Central
# * Canda
# * Asia Pacific North East 1
# * Asia Pacific North East 2
# * Asia Pacific South
# * Asia Pacific Southeast 1
# * Asia Pacific Southeast 2
#
#
#
#
# + [markdown] _uuid="14ca6867f2d46f9e68eb1ec285cb0005e60b3dd2"
# # Import Libraries
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import seaborn as sns
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from IPython.display import HTML, display
from fastai.structured import *
from fastai.column_data import *
np.set_printoptions(threshold=50, edgeitems=20)
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import os
print(os.listdir("../input"))
# + [markdown] _uuid="201278f8f2c44812147c22a9a915fdc22c26e595"
# Lets import all the tables
# + _uuid="15696f1e4a1081f3a5a783d4a65759d6f0a4e4e9"
PATH = "../input/"
PATH_WRITE = "/kaggle/working/"
# + _uuid="1333ffc434045d439ccdf75599713b01524fb9d4"
# ls {PATH}
# + _uuid="d2073317a9e4d38d920b48a4b04aed4f47d9af98"
table_names = ['ap-southeast-2', 'ap-northeast-2', 'ca-central-1', 'us-east-1',
'ap-northeast-1', 'ap-south-1', 'sa-east-1', 'eu-west-1',
'ap-southeast-1', 'us-west-1', 'eu-central-1']
# + _uuid="6aa4b36a89e87d6b4e1c4535166ea8befb31a52a"
tables = [pd.read_csv(f'{PATH}{fname}.csv', low_memory=False) for fname in table_names]
# + [markdown] _uuid="5ad73f558fa28a266760e9d72fbe60b47afe89d9"
#
# + [markdown] _uuid="be2d115a8d3c888150ad7a65e0439126caf9aaf0"
#
# + [markdown] _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# # EDA
# + [markdown] _uuid="8182ec4199ee1d9a2cffb9c77a018b7b5b9b0709"
# Lets call head and take a look at what our data looks like.
# + _uuid="99cfee5cb7ae0f1821e1ebd2b484c3c280cc1d05"
for t in tables: display(t.head())
# + [markdown] _uuid="f1994a0ed99160540a45175bbd2f894b7794f15d"
# Lets call summary
# + _uuid="6f11d0dce90e036093f0b43717d877e84a83a9af"
for t in tables: display(DataFrameSummary(t).summary())
# + [markdown] _uuid="6e0269dd1faadb7b9c8845c68050ef6d805b7a0d"
# I think we need to change some of the columns names
# + _uuid="3d04b074c36921d29a32ed584e5150d7fcf9f11d"
new_labels = ['Date', 'Instance Type', 'OS', 'Region', 'Price ($)']
# + _uuid="5c4c71fe145020dfabd7664fdfad9957d9580467"
for t in tables:
t.columns = new_labels
# + _uuid="4edb090bfcb6f7da8731cb19b82b8b969a8f2a65"
for t in tables: display(t.head())
# + _uuid="0e6c6f2f9560a1ae5174e4a109e553865d7c2ce0"
for t in tables:
plt.figure(figsize=(12,8))
sns.countplot(t['Instance Type'], order=t['Instance Type'].value_counts().iloc[:20].index)
plt.xticks(rotation=90);
# + [markdown] _uuid="0927bba0acb6e575c8314a81e1ddf1b37d39593d"
# List of questions to ask:
#
# * Average price for certain instances in each region
# * Frequent instance type
# * Seasonlity of instances
# * Determine if there are any stationary variables
# * Which instance type is frequently link with what OS?
# * Need to plot instances in time-intervalse eg: between 21:00 - 22:00
#
# Also need to figure out how to give each region a table name for the graphs.
# + [markdown] _uuid="cfb5327d9a5eabb4ab81c465e907612a29e60665"
# Lets look at the tables seperately:
# + [markdown] _uuid="c97c7783e5f10166d31470fd77d4f15412d7f81c"
# # US East
# + _uuid="8f0080bd992cc111fa212f68725e7d4143d42bd7"
us_east = pd.read_csv("../input/us-east-1.csv")
PATH_USEAST = "../input/us-east-1.csv"
# + _uuid="771b9559727208d9cc4a92908ed2299783502221"
us_east.columns = new_labels
us_east.head()
# + _uuid="3a5295d73b5cd33f781389af425279e368615dff"
us_east['Date'].head()
# + [markdown] _uuid="67249332ca94a3ddef869438a140851faf220649"
# We need to parse the dates otherwise the dates will not apear on axis. The format string needs to match the format in the column EXACTLY! For more info look [here](http://strftime.org/) and [here](https://codeburst.io/dealing-with-datetimes-like-a-pro-in-pandas-b80d3d808a7f)
# + _uuid="8050f3757339a1f7d4ac6a5cf53742a822e7eb1f"
us_east['Date'] = pd.to_datetime(us_east['Date'], format='%Y-%m-%d %H:%M:%S+00:00', utc=False)
# + _uuid="3cadbe39219948fb1d426c64ffdeeb220aee53e8"
us_east.info()
# + _uuid="b669c11ac7ce0720272f1c58e726c601dbe770d6"
us_east['Date'].head(500)
# + [markdown] _uuid="890cccd166a16535c780c8fe9bfaf3f2c5970569"
# ## Instance: d2.xlarge
# + _uuid="702bd81a9aeefb67b322ecde7cb3c762c9cfb70a"
d2_= us_east[us_east['Instance Type'] == "d2.xlarge"].set_index('Date')
# + _uuid="e51c7233771be424504993b79eca75de49281a5d"
d2_Unix = us_east[us_east['OS'] == "Linux/UNIX"].set_index('Date')
# + _uuid="29d7ce1d5fa3568c6e5903cd1718b6a57901ec83"
d2_Suse = us_east[us_east['OS'] == "SUSE Linux"].set_index('Date')
# + _uuid="f804f552f9bad2a1f69d2812dd8292eef527a5d0"
d2_Win = us_east[us_east['OS'] == "Windows"].set_index('Date')
# + _uuid="31e4448882cec143bb652f0b990e4ad1437d67a8"
d2.head()
# + _uuid="fe24da3a2854fdd6a29fc25849ce5385d70e9b56"
d2.head()
# + _uuid="d95860367c735bcfdb77d03961a989892ea59eb8"
d2.head(100).plot(title="d2.xlarge Instances", figsize=(15,10))
# + _uuid="056b6b2d3ea1ed5e3555db825cd5b32f0cc262a2"
d2_Suse.head(100).plot(title="d2.xlarge Instances OS:SUSE Linux", figsize=(15,10))
# + _uuid="e15d4358545a3b1e25dd06e08844f87078f511e0"
d2_Unix.head(100).plot(title="d2.xlarge Instances OS:Linux/UNIX", figsize=(15,10))
# + _uuid="1679d1a1ef9ae4ba38c61f26a564909c329029fd"
d2_Win.head(100).plot(title="d2.xlarge Instances OS:Windows", figsize=(15,10))
# + [markdown] _uuid="9e94558673e39d3f9dae356a4eff7884780b6b39"
# Looks like windows Instances can get quite pricey with highs of around roughly `$` 7 dollars - `$` 29!🤷♀️
# + _uuid="c14e29285bc45faa5dee702b9aeb63e45ec0c528"
# + [markdown] _uuid="b9b68b8e92d857d477f2c2f1b3232dd010dfa608"
# # Cleaning
# + [markdown] _uuid="8597eae5ce3f38d3639ac8d1f7f179f93107056c"
# Lets go over the steps for checking for signs that we need to clean our dataset.
# + _uuid="7ed00f89ebaa4aca707565f3026da6845b723224"
us_east['Instance Type'].value_counts(dropna=False)
# + _uuid="dbf35b53819101f70fd2e4a7314bc6b74d769451"
us_east['OS'].value_counts(dropna=False)
# + _uuid="498cfb62d44d7ec0a8b8f3990a21a1d00771e58b"
len(us_east.isnull())
# + [markdown] _uuid="d32ba041302daf33b57c5cc8dcde6de4211c2c85"
# Out of 3 721 999 entries none have null values
# + [markdown] _uuid="26cbbc7c79e74b5138f74485f49010703ff1aeff"
# Lets train on another dataset
# + _uuid="dba41d247ed841e90f79fbf64988dd971d235576"
eu_west = pd.read_csv("../input/eu-west-1.csv")
PATH_euwest = "../input/"
# + _uuid="07068b76e9f552c529c12df4f2d9e9a897208faa"
eu_west.columns = new_labels
# + _uuid="d88b3507756b7897788f2b23d68e6da4172b17b0"
eu_west.info()
# + _uuid="8ce6ee52d87688e416ecb20d3e5f920c4563955e"
eu_west['Instance Type'].value_counts(dropna=False)
# + _uuid="208c5223a6888af8b55bf46c424cfbdab8973d38"
len(eu_west.isnull())
# + [markdown] _uuid="3f4f9f80d4da3fc49243b1b64664923adda826b0"
# # Implement Model
# + [markdown] _uuid="2752a3185ed1f147c4b74640ddd890933c57e6c7"
# Things to do:
#
# * Len the Instance type [done]
# * Add date part [done]
#
# * Create cat & continous vars [done] - do not have any other kind continous var!!
# * Process datasets [done]
# * Split Dataset - via datetime [done]
# * Create RMSE metric
# * Create model data object
# * calculate embeddings
# * Train model
# + _uuid="562f468726ea67e65218898d00f1e6ca791ed282"
add_datepart(eu_west, 'Date', drop=False)
# + _uuid="493504dfffc43d7c81e477014083c3b52068322b"
eu_west.reset_index(inplace=True)
eu_west.to_feather(f'{PATH_WRITE}eu_west')
eu_west.shape
# + _uuid="f600ad3efe637a2fb80d3b4bed29f137dbb949a1"
eu_west=pd.read_feather(f'{PATH_WRITE}eu_west')
# + _uuid="a08768c6633df34ed24bc9f102331821924929ed"
eu_west.columns
# + _uuid="8a1a3381d944815b63b87d65af47c4eef29a64b9"
joined = eu_west
joined_test = eu_west
# + _uuid="295750f97cafbeec20373223840d2bd3f4a5357b"
joined.to_feather(f'{PATH_WRITE}joined')
joined_test.to_feather(f'{PATH_WRITE}joined_test')
# + _uuid="acefae1dc369cbd1929d0840ddc6f69c3b9b7fd8"
joined = pd.read_feather(f'{PATH_WRITE}joined')
joined_test = pd.read_feather(f'{PATH_WRITE}joined_test')
# + _uuid="8d38ae66a7c1e66e55b24aac927de6c2876367d1"
joined.head()
# + _uuid="bb4a47301e8500c903ce4d24afdcb3fba3016c47"
cat_vars = [
'Instance Type',
'OS',
'Region',
'Year' ,
'Month' ,
'Week' ,
'Day',
'Dayofweek',
'Dayofyear'
]
contin_vars = ['Elapsed']
n = len(joined); n
# + _uuid="469771cb58be289167cc9677433bffa0ac7f34c3"
dep = 'Price ($)'
joined = joined[cat_vars+contin_vars+[dep,'Date']].copy()
# + _uuid="f0a1728b4e65795cb4b3cbc7df25a32b88b62194"
joined_test[dep] = 0
joined_test = joined_test[cat_vars+contin_vars+[dep,'Date',]].copy()
# + _uuid="16d7b8502a81be38017484591c2c6cf3d1930290"
for cat in cat_vars: joined[cat] = joined[cat].astype('category').cat.as_ordered()
eu_west['Price ($)'] = eu_west['Price ($)'].astype('float32')
# + _uuid="815f7b25e38ad2378bb32cf56d4885741611c5ce"
for contin in contin_vars:
joined[contin] = joined[contin].astype('float32')
joined_test[contin] = joined_test[contin].astype('float32')
# + _uuid="35e9e09bb6ecde69698ac21485e8bce8dd4ac451"
# + _uuid="f73e956e68eff54f02a1e35bc2ae5a4d199582a4"
idxs = get_cv_idxs(n, val_pct=150000/n)
joined_sample = joined.iloc[idxs].set_index("Date")
samp_size = len(joined_sample); samp_size
# + _uuid="c569c0680c96fe69c344a93705605e90b6158799"
samp_size = n
# + _uuid="bd49c1f9c1604fc90cc1645e57e084506e621a78"
joined_sample.head()
# + _uuid="ad1b07bad0f5daf6e88ca8be2b1ec7ad9d0b923d"
df_train, y, nas, mapper = proc_df(joined_sample,'Price ($)', do_scale=True)
yl = np.log(y)
# + _uuid="525f0158369d7decd8eefe1a7d06b59a52530956"
joined_test = joined_test.set_index("Date")
# + _uuid="177a699a1e1c35184a72e7a2bbb5ac54c365c278"
df_test, _, nas, mapper = proc_df(joined_test,'Price ($)', do_scale=True,mapper=mapper, na_dict=nas )
# + _uuid="750f4759b0c2df8992016d75fb028637cf540c95"
# %debug
# + _uuid="552594f719cb631cb50278d6ba792e5d5e87a49d"
df_train.info()
# + _uuid="b514e2e36c702b09bf730de2340123c296b37f9e"
# + _uuid="5e48c68defd113f99015422f789bb3d97ce1f2ef"
# + _uuid="4e0302562587c70d38a412eb2a923361f5ecdfe2"
train_val_split = 0.80
train_size = int(2383999 * train_val_split);train_size
val_idx = list(range(train_size, len(df_train)))
# + _uuid="e4e6c15c973aebd60da7eebf957893b6878a7b5b"
val_idx = np.flatnonzero(
(df_train.index<=datetime.datetime(2017,4,12)) & (df_train.index>=datetime.datetime(2017,4,12)))
# + _uuid="11d28330c430fa3cb4e5883741bee811306e3ea3"
val_idx=[0]
# + _uuid="1d6d86b449864ced33399722e294ebdd4154de7e"
len(val_idx)
# + [markdown] _uuid="e904d533e6602f21a76e2df8b448d6b983e071f3"
# We can put our Model.
# + _uuid="3c688932429fcc053032c0cee82dac3f9b791382"
def inv_y(a): return np.exp(a)
def exp_rmspe(y_pred, targ):
targ = inv_y(targ)
pct_var = (targ - inv_y(y_pred))/targ
return math.sqrt((pct_var**2).mean())
max_log_y = np.max(yl)
y_range = (0, max_log_y*1.2)
# + _uuid="d81d6e0c364ca98d50524a4198da3aa26eb195c5"
md = ColumnarModelData.from_data_frame(PATH_euwest, val_idx, df_train, yl.astype(np.float32),
cat_flds=cat_vars, bs=128, test_df=df_test)
# + _uuid="f9ea1ccef026627f1f4e03319dbfc8b3a6b011ac"
cat_sz = [(c, len(df_train[c].cat.categories)+1) for c in cat_vars]
# + _uuid="e6590bbd697a09851a263e2e5b241bf0e3c03679"
# + [markdown] _uuid="7730bf002329986d5ac5575ac0e118114c45789d"
# # Conclusion on Results
# + [markdown] _uuid="f85254622ca7a4aaac0bf5ad18771d82231a9c85"
#
| Prediticing EC2 Spot Prices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from helpers.utilities import *
# %run helpers/notebook_setup.ipynb
# + tags=["parameters"]
clinical_data_path = 'data/clean/clinical/data.csv'
output_path = 'data/clean/clinical/data_with_derived_variables.csv'
# -
clinical_data = read_csv(
clinical_data_path, index_col=0,
parse_dates=['AdmissionDate', 'Birthday', 'DateDeath']
)
# ## Adding derived clinical variables:
# ### Age
clinical_data['age_at_admission'] = clinical_data['AdmissionDate'] - clinical_data['Birthday']
clinical_data['age_at_admission_years'] = clinical_data['age_at_admission'].dt.days / 360
clinical_data['age_at_admission_years'].hist();
clinical_data['age_at_admission_years'].describe()
clinical_data['age_at_admission_years'].round().median()
# ### Survival
clinical_data['survival'] = (
clinical_data['DateDeath'] - clinical_data['AdmissionDate']
)
clinical_data['survival'].describe()
# The figures above describe only the 22 deceased patients. As the follow-up was 6 months:
follow_up = np.timedelta64(6, 'M')
clinical_data['censored_survival'] = clinical_data.survival.fillna(follow_up)
clinical_data['censored_survival'].tail()
# Convert to days:
clinical_data['survival'] = clinical_data['survival'].dt.days
clinical_data['censored_survival'] = clinical_data['censored_survival'].dt.days
clinical_data['censored_survival'].hist();
# ### Adding disease data
# Extract vectors with nicely formatted (and ordered) condition (disease) names:
from plots.annotations import conditions_names, tuberculosis_status
conditions_names
granular_conditions = clinical_data.index.str.split('.').str[1]
by_condition = granular_conditions.map(conditions_names)
clinical_data['Meningitis'] = by_condition
assert not clinical_data['Meningitis'].isnull().any()
tuberculosis_status
clinical_data['Tuberculosis'] = granular_conditions.map(tuberculosis_status).fillna('-')
clinical_data['Meningitis_with_tuberculosis_status'] = clinical_data.apply(
lambda c: (
c.Meningitis
if c.Meningitis != 'Tuberculosis' else
c.Tuberculosis + ' ' + c.Meningitis.lower()
),
axis=1
)
clinical_data['Meningitis_with_tuberculosis_status'].head()
# ### Saving expanded dataset
clinical_data.to_csv(output_path)
# TODO: split into dependent/independent variables and confounders?
# ## Correlation of the clinical variables
# TODO
# ## Which variables to use for analyses?
#
# ### Regression
#
# We may want to regress on:
# - symptoms severity (e.g. duration of lethargy, reduced consciousness, sickness, etc)
# - some of the interval clinical variables (Level of glucose in CSF, CD4 count)
# - ordinal variables (in between the regression and classification - ordinal/multinomial regression):
# - grade of TBM (1-3)
# - tuberculosis "certainty" status on the TB (+viral?) patients only
#
# Variable selection:
# - lasso: per each objective (to determine the top proteins influencing each response variable)
# - differential "expression"
#
# ### Classification
#
# - HIV status (logistic regression?)
# - Patient group (4 meningitis groups)
# - CSF appearance (4 combinations or 2x2)
#
| analyses/Clinical_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ¡A practicar!
# ### Usar un condicional para que diga si es un número par o impar
número = 7
# ### Usar un bucle para que devuelva el doble del valor de cada elemento en una lista
lista = [6,23,44,2,57]
# + active=""
# ### Usar un bucle para que muestre asteriscos del 1 al 5
# *
# **
# ***
# ****
# *****
# -
# ### Define una función que te devuelva la suma de todos los números pares entre cero y n. Donde n entra como argumento de la función
| exercises_003.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime
import h5py
import itertools
import librosa
import math
import numpy as np
import os
import pandas as pd
import soundfile as sf
import sys
import time
sys.path.append('../src')
import localmodule
# Define constants.
data_dir = localmodule.get_data_dir()
dataset_name = localmodule.get_dataset_name()
full_audio_name = "_".join([dataset_name, "full-audio"])
full_audio_dir = os.path.join(data_dir, full_audio_name)
sample_rate = localmodule.get_sample_rate()
args = ["unit01"]
unit_str = args[0]
sf_hop_length = 512 # default value for melspectrogram in librosa
sf_sr = 22050 # defaut value for sample rate in librosa
chunk_duration = 256 # in seconds
chunk_length = chunk_duration * sample_rate
# Print header.
start_time = int(time.time())
print(str(datetime.datetime.now()) + " Start.")
print("Running spectral flux on " + dataset_name + ", " + unit_str + ".")
print('h5py version: {:s}.'.format(h5py.__version__))
print('librosa version: {:s}.'.format(librosa.__version__))
print('numpy version: {:s}'.format(np.__version__))
print('pandas version: {:s}'.format(pd.__version__))
print('soundfile version: {:s}'.format(sf.__version__))
print("")
# Create HDF5 container of ODF curves.
full_spectralflux_name = "_".join([dataset_name, "spectral-flux"])
full_spectralflux_dir = os.path.join(data_dir, full_spectralflux_name)
os.makedirs(full_spectralflux_dir, exist_ok=True)
out_name = unit_str
out_path = os.path.join(full_spectralflux_dir, out_name + ".hdf5")
out_file = h5py.File(out_path)
# Load GPS coordinates.
gps_name = "_".join([dataset_name, "gps-coordinates.csv"])
gps_path = os.path.join(data_dir, gps_name)
gps_df = pd.read_csv(gps_path)
gps_row = gps_df.loc[gps_df["Unit"] == unit_str].iloc[0]
# Load UTC starting times.
utc_name = "_".join([dataset_name, "utc-start-times.csv"])
utc_path = os.path.join(data_dir, utc_name)
utc_df = pd.read_csv(utc_path)
utc_row = utc_df.loc[utc_df["Unit"] == unit_str].iloc[0]
# Copy over metadata.
out_file["dataset_name"] = dataset_name
out_file["unit"] = unit_str
out_file["sample_rate"] = sample_rate
out_file["utc_start_time"] = utc_row["UTC"]
gps_group = out_file.create_group("gps_coordinates")
gps_group["latitude"] = gps_row["Latitude"]
gps_group["longitude"] = gps_row["Longitude"]
# Open full audio file as FLAC.
recordings_name = "_".join([dataset_name, "full-audio"])
recordings_dir = os.path.join(data_dir, recordings_name)
recording_name = unit_str + ".flac"
recording_path = os.path.join(recordings_dir, recording_name)
full_audio = sf.SoundFile(recording_path)
full_audio_length = len(full_audio)
# Initialize dataset of onset detection function (ODF).
dataset_size = (1, full_audio_length)
spectralflux_dataset = out_file.create_dataset(
"spectral-flux_odf", dataset_size)
# +
# Loop over chunks.
#for chunk_id in range(n_chunks):
chunk_id = n_chunks - 1
# Load audio chunk.
chunk_start = chunk_id * chunk_length
chunk_stop = min(chunk_start + chunk_length, full_audio_length)
full_audio.seek(chunk_start)
chunk_waveform = full_audio.read(chunk_stop-chunk_start)
# Compute spectral flux.
odf = librosa.onset.onset_strength(chunk_waveform)
# Delete last sample to compensate for padding.
odf = odf[:-1]
# -
# Write to HDF5 dataset.
# hop_start is an integer because chunk_start is both a multiple
# of sample_rate and hop_length = chunk_duration.
hop_start = int((chunk_start*sf_sr) / (sample_rate*sf_hop_length))
n_hops_in_chunk = odf.shape[0]
hop_stop = min(hop_start + n_hops_in_chunk, n_hops)
spectralflux_dataset[:, hop_start:hop_stop] = odf
chunk_start + chunk_length
full_audio_length
| notebooks/run-spectral-flux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="b5e6e994"
# [](https://colab.research.google.com/github/ThomasAlbin/Astroniz-YT-Tutorials/blob/main/[ML1]-Asteroid-Spectra/13_dl_autoencoder_clustering.ipynb)
# + [markdown] id="f14c9d7a"
# # Step 13: Autoencoder Clustering
#
# In this final tutorial, we will use the latent space to cluster our asteroide spectra using Gaussian Mixture Models (GMMs). However, there are tons of ways how to cluster data in an un-supervised way. One could also apply GMMs directly on the spectra itself. Using Autoencoders, one could also train the deep learning architecture on C, S and X spectra and find "Other" spectra as outliers; considering deviations in the reconstructed spectrum.
#
# Now, today's new part starts at [cell 11](#SessionStart)
# + id="8bb6f3e5-ad7e-4a03-b940-b78f819ca0f8"
# Import standard libraries
import os
# Import installed libraries
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
# Scikit-Learn stuff
import sklearn
from sklearn import preprocessing
from sklearn.model_selection import StratifiedShuffleSplit
# Keras
import tensorflow.keras as keras
import tensorflow as tf
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# + id="6e63686b-e7ab-4d53-8542-3e33ed31769d"
# Set seeds to create reproducible experiments
np.random.seed(1)
tf.random.set_seed(1)
# + colab={"base_uri": "https://localhost:8080/"} id="e3330fcb-2536-461b-8d51-c8f8c1bb0cea" outputId="fc158338-95ad-40e7-ac71-619be4d4749c"
# Let's mount the Google Drive, where we store files and models (if applicable, otherwise work
# locally)
try:
from google.colab import drive
drive.mount('/gdrive')
core_path = "/gdrive/MyDrive/Colab/asteroid_taxonomy/"
except ModuleNotFoundError:
core_path = ""
# Load the level 2 asteroid data
asteroids_df = pd.read_pickle(os.path.join(core_path, "data/lvl2/", "asteroids.pkl"))
# Allocate the spectra to one array and the classes to another one
asteroids_X = np.array([k["Reflectance_norm550nm"].tolist() for k in asteroids_df["SpectrumDF"]])
asteroids_y = np.array(asteroids_df["Main_Group"].to_list())
asteroids_y_bus = np.array(asteroids_df["Bus_Class"].to_list())
# + [markdown] id="bc40785e-79a9-4335-b7da-6b807bb915c2"
# ## ShuffleSplit
#
# Although we do not conduct a classification ML experiment, we still consider the distribution of the classes to train our network properly.
# + id="8d56dda9-5218-448d-8fd4-b317a6caca5c"
# In this example we create a single test-training split with a ratio of 0.8 / 0.2
sss = StratifiedShuffleSplit(n_splits=1, test_size=0.2)
# Create a simple, single train / test split
for train_index, test_index in sss.split(asteroids_X, asteroids_y):
X_train, X_test = asteroids_X[train_index], asteroids_X[test_index]
y_train, y_test = asteroids_y[train_index], asteroids_y[test_index]
y_train_bus, y_test_bus = asteroids_y_bus[train_index], asteroids_y_bus[test_index]
# + [markdown] id="b9266e90-fe81-4424-8eef-4af9af622034"
# ## Scaling
#
# This time we are creating a scikit-learn scaler for our spectra data. The model's prediction signals need to be transformed inversely later on to display them correctly.
# + id="3b2d9fa8-531b-4f32-943f-08e37cb733d4"
# Import the preprocessing module
from sklearn import preprocessing
# Instantiate the StandardScaler (mean 0, standard deviation 1) and use the training data to fit
# the scaler
scaler = preprocessing.StandardScaler().fit(X_train)
# Transform now the training data
X_train_scaled = scaler.transform(X_train)
# Scale the testing data ...
X_test_scaled = scaler.transform(X_test)
# And expanding the dimensionality for our ConvNet-based Autoencoder
X_train_scaled = np.expand_dims(X_train_scaled, axis=2)
X_test_scaled = np.expand_dims(X_test_scaled, axis=2)
# + [markdown] id="d48fcbf6-e5fc-478f-ac77-8bb1087f9042"
# ## Building the Autoencoder
#
# Now we create a ConvNet Autoencoder. Note that we are not using Keras-Tuner this time. Feel free to apply Keras-Tuner as a small coding exercise.
# + colab={"base_uri": "https://localhost:8080/"} id="652dbd5c-7f7f-424b-9e28-58a91000dad0" outputId="8f785871-cb7b-4ce9-c5ea-3a950d9aa598"
# Get the number of inputs
n_inputs = asteroids_X.shape[1]
# Let's create an autoencoder with a 2-D latent space
n_bottleneck = 5
def create_model():
# Input layer, this time without a normalisation layer
input_layer = keras.Input(shape=(n_inputs, 1))
# Conv Layers (we won't be using maxpooling, since the dimensionaliy is 49 and we do not
# alter the data in our example
hidden_layer = keras.layers.Conv1D(filters=16,
activation="relu",
kernel_size=3, padding="same")(input_layer)
hidden_layer = keras.layers.Conv1D(filters=32,
activation="relu",
kernel_size=3, padding="same")(hidden_layer)
# Encoder ("Bottleneck" of the Autoencoder)
bottleneck_lay = keras.layers.Flatten()(hidden_layer)
bottleneck_lay = keras.layers.Dense(n_bottleneck)(bottleneck_lay)
# The original shape must be restored and reshaped accordingly
reset_lay = keras.layers.Dense(49*32)(bottleneck_lay)
reshape_lay = keras.layers.Reshape((49, 32))(reset_lay)
# First and second hidden decoder layers
hidden_layer = keras.layers.Conv1DTranspose(filters=32,
kernel_size=3,
strides=1,
activation="relu",
padding="same")(reshape_lay)
hidden_layer = keras.layers.Conv1DTranspose(filters=16,
kernel_size=3,
strides=1,
activation="relu",
padding="same")(hidden_layer)
# Ouput layer (same size as input layer)
output_layer = keras.layers.Conv1D(1, 1, padding="same")(hidden_layer)
# Create model
model = keras.models.Model(inputs=input_layer, outputs=output_layer)
# Create encoder model
encoder_model = keras.models.Model(inputs=input_layer, outputs=bottleneck_lay)
# We return the model and the encoder
return model, encoder_model
model, encoder_model = create_model()
# Compile the model and use a regression loss function
model.compile(optimizer='adam', loss='mse')
# Show the model summary
model.summary()
# + id="2ca843b1-e07a-4f84-9869-7cb8e9fcaa97"
# Train the model
end_epoch = 500
batch_size = 32
# Early Stopping for our final model
es_callback = keras.callbacks.EarlyStopping(monitor='val_loss', patience=5)
history = model.fit(X_train_scaled, X_train_scaled,
epochs=end_epoch,
batch_size=batch_size,
verbose=0,
validation_split=0.25,
callbacks=[es_callback])
# + [markdown] id="6f777c18-5ad6-4099-af57-5dae7a2dabd4"
# ## The loss function
#
# Let's show the loss of the training and test data. As you can see, the minimum-plateau is reached quite fast. The test data performs slightly better than the training data, since the loss results for the training data are based on an average of the batch size. The test results are based on all data.
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="6434731b-7e67-4b22-ba7a-782b56ffca56" outputId="512bba5a-7d83-44ad-9c81-b0660ab7f026"
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# plot the training and validation loss
plt.figure(figsize=(10,8))
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
# Add legend and labels
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("MSE Loss")
# ... aaaaand plot!
plt.show()
# + [markdown] id="44548c22-503d-4bfa-afde-607ce1dbf912"
# ## Signal Reconstruction
#
# ... can be done by using the entire model to predict the output spectra. The following code snippet takes a signal (change the index value in the first line to use any other signal) and predicts the Autoencoder based spectrum.
#
# As one can see, the results are "quite ok" but not perfect. The shape of the spectra can be reproduced. However, the signals are quite noisy.
# + colab={"base_uri": "https://localhost:8080/", "height": 353} id="9e2bd0ef-eaf5-4aa6-843c-6792ecb10e68" outputId="e3830fee-f7fe-46c8-f5d6-d64a158e5d0c"
# Which index shall be displayed?
index_val = 5
# Original signal
org_signal = scaler.inverse_transform(X_train_scaled[index_val].reshape(1, -1))[0]
# Reconstructed signal
rec_signal = scaler.inverse_transform(model.predict(X_train_scaled)[index_val].reshape(1, -1))[0]
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# plot the training and reconstructed data
plt.figure(figsize=(12,5))
plt.plot(org_signal, label='Original')
plt.plot(rec_signal, label='Reconstructed')
# Add legend and labels
plt.legend()
plt.xlabel("Spectra: array index")
plt.ylabel("Normalized Reflectance")
# ... aaaaand plot!
plt.show()
# + id="d33b8a3f-b653-4cc1-acde-94cab67d92eb"
# Create dataframe that contains the encoder values and the corresponding class to see whether the
# autoencoder values cluster in a way
# Encode the spectra
X_train_encoded = encoder_model.predict(X_train_scaled)
X_test_encoded = encoder_model.predict(X_test_scaled)
# Merge the data
X_encoded = np.vstack((X_train_encoded, X_test_encoded))
# Instantiate the StandardScaler (mean 0, standard deviation 1) and use the encoded data to fit
# the scaler
enc_scaler = preprocessing.StandardScaler().fit(X_encoded)
# Transform now the encoded data (later used for our scikit-learn method)
X_encoded_scaled = enc_scaler.transform(X_encoded)
# Merge the classes
y_main = np.hstack((y_train, y_test))
y_bus = np.hstack((y_train_bus, y_test_bus))
# Create a column names array for the encoded space
encoder_space = [f"enc{enc_nr+1}" for enc_nr in range(n_bottleneck)]
encoder_space_cols = encoder_space.copy()
encoder_space.extend(["Main_Group", "Bus_Class"])
# Create the dataframe
encoded_df = pd.DataFrame(np.hstack((X_encoded_scaled,
y_main[np.newaxis].transpose(),
y_bus[np.newaxis].transpose())),
columns=encoder_space)
# Change the dtype to float
encoded_df.loc[:, encoder_space[:-2]] = encoded_df.loc[:, encoder_space[:-2]].astype(float)
# + [markdown] id="rWBGad2pPRwq"
# ## Gaussian Mixture Model
# <a id='SessionStart'></a>
# With the encoded values we will now conduct a clustering experiment using Gaussian Mixture Models (GMMs). The multi-dimensional data will be used to fit a varying number of Gaussians.
#
# The question is: How many Gaussians are needed to fit the data... "properly"?
#
# For this, we will fit an increasing number of Gaussians and compute the corresponding, so-called *Bayesion Information Criterion* (BIC). This values ia a quantified version of Occam's Razor: more parameters may describe the data better, but more parameters will be punished. Goal: finding the perfect balance between "explainability" and "overfitting".
# + colab={"base_uri": "https://localhost:8080/"} id="RMiAx1ZQKiAU" outputId="2518eeda-8de1-4d9c-c4f1-c576d87070e3"
import sklearn.mixture
import tqdm
# Result dataframe that will contain the number of Gaussian components, the Bayesion Information
# Criterion (BIC) and the model itself
gmm_results_df = pd.DataFrame([], columns=["Nr_Comp", "BIC", "Model"])
# We iterate through a number of "component guesses"
max_gauss = 15
for index, gauss_components in tqdm.tqdm(enumerate(np.arange(1, max_gauss+1, 1))):
# Create and fit a temporary Gaussian Mixture Model
temp_gmm = sklearn.mixture.GaussianMixture(n_components=gauss_components,
covariance_type="full")
temp_gmm.fit(X_encoded_scaled)
# Store the number of components, the BIC and the model
gmm_results_df.loc[index, "Nr_Comp"] = gauss_components
gmm_results_df.loc[index, "BIC"] = temp_gmm.bic(X_encoded_scaled)
gmm_results_df.loc[index, "Model"] = temp_gmm
# + colab={"base_uri": "https://localhost:8080/", "height": 530} id="kt8AeR_-PkNm" outputId="c6463377-bcb8-41f6-b2f0-d14cda9aefbb"
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# Plotting the BIC vs. the number of components
plt.figure(figsize=(10, 8))
plt.plot(gmm_results_df["Nr_Comp"],
gmm_results_df["BIC"],
linestyle="dashed",
marker="o",
markersize=5,
color="w",
alpha=0.7)
# Color the minimum value
gmm_results_best = gmm_results_df.loc[gmm_results_df["BIC"] == gmm_results_df["BIC"].min()]
plt.plot(gmm_results_best["Nr_Comp"],
gmm_results_best["BIC"],
marker="o",
markersize=15,
color="tab:green",
alpha=0.7)
# Some formatting
plt.xlabel("Number of Gaussian Components")
plt.ylabel("BIC")
plt.grid(linestyle="dashed", alpha=0.3)
plt.xlim(1, max_gauss)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="lqZ6IlUESln8" outputId="7c19db99-8885-4d2a-87a9-5df25cac025b"
# Let's take the best GMM!
best_gmm = gmm_results_best["Model"].iloc[0]
# Create a new dataframe column that labels the spectra based on our GMM model:
encoded_df.loc[:, "GMM_Class"] = best_gmm.predict(encoded_df[encoder_space_cols].values)
encoded_df
# + colab={"base_uri": "https://localhost:8080/", "height": 512} id="AG52flFUVxJ6" outputId="2b6b6ec1-d455-476a-d3fc-960a074257ce"
# Groupby the Main group and the GMM classification
encoded_grouped_df = pd.crosstab(index=encoded_df["Main_Group"],
columns=encoded_df["GMM_Class"],
values=encoded_df["enc1"],
aggfunc="count")
# Extract data, column and index names for plotting purposes
encoded_grouped_values = encoded_grouped_df.values
encoded_grouped_main = encoded_grouped_df.index.values
encoded_grouped_gmm = [f"C{k}" for k in encoded_grouped_df.columns.values]
# Matplotlib settings
# Set the dark mode and the font size and style
plt.style.use('dark_background')
plt.rc('font', family='serif', size=18)
# Create a matrix-like plot of the results
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(111)
cax = ax.matshow(encoded_grouped_values, cmap="afmhot")
fig.colorbar(cax, label="Number of spectra", fraction=0.05)
# Some plotting settings
ax.set_xticks(range(len(encoded_grouped_gmm)))
ax.set_yticks(range(len(encoded_grouped_main)))
ax.set_xticklabels(encoded_grouped_gmm)
ax.set_yticklabels(encoded_grouped_main)
ax.set_xlabel("GMM based classification")
ax.set_ylabel("Main Group")
ax.xaxis.set_label_position('top')
# -
# ## Summary & Outlook
#
# Applying a GMM on our data assumes one thing: the multi-dimensional data is distributed in a Gaussian way. But is it? Well not 100 %; but here I wanted to finish this ML project with a mixture of Autoencoders and GMMs. One can
| [ML1]-Asteroid-Spectra/13_dl_autoencoder_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# HIDDEN
from datascience import *
# %matplotlib inline
path_data = '../../../data/'
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
import math
import numpy as np
# ### The SD and the Normal Curve ###
#
# We know that the mean is the balance point of the histogram. Unlike the mean, the SD is usually not easy to identify by looking at the histogram.
#
# However, there is one shape of distribution for which the SD is almost as clearly identifiable as the mean. That is the bell-shaped disribution. This section examines that shape, as it appears frequently in probability histograms and also in some histograms of data.
# ### A Roughly Bell-Shaped Histogram of Data ###
# Let us look at the distribution of heights of mothers in our familiar sample of 1,174 mother-newborn pairs. The mothers' heights have a mean of 64 inches and an SD of 2.5 inches. Unlike the heights of the basketball players, the mothers' heights are distributed fairly symmetrically about the mean in a bell-shaped curve.
baby = Table.read_table(path_data + 'baby.csv')
heights = baby.column('Maternal Height')
mean_height = np.round(np.mean(heights), 1)
mean_height
sd_height = np.round(np.std(heights), 1)
sd_height
baby.hist('Maternal Height', bins=np.arange(55.5, 72.5, 1), unit='inch')
positions = np.arange(-3, 3.1, 1)*sd_height + mean_height
plots.xticks(positions);
# The last two lines of code in the cell above change the labeling of the horizontal axis. Now, the labels correspond to "average $\pm$ $z$ SDs" for $z = 0, \pm 1, \pm 2$, and $\pm 3$. Because of the shape of the distribution, the "center" has an unambiguous meaning and is clearly visible at 64.
# ### How to Spot the SD on a Bell Shaped Curve ###
#
# To see how the SD is related to the curve, start at the top of the curve and look towards the right. Notice that there is a place where the curve changes from looking like an "upside-down cup" to a "right-way-up cup"; formally, the curve has a point of inflection. That point is one SD above average. It is the point $z=1$, which is "average plus 1 SD" = 66.5 inches.
#
# Symmetrically on the left-hand side of the mean, the point of inflection is at $z=-1$, that is, "average minus 1 SD" = 61.5 inches.
#
# In general, **for bell-shaped distributions, the SD is the distance between the mean and the points of inflection on either side.**
# ### The standard normal curve ###
#
# All the bell-shaped histograms that we have seen look essentially the same apart from the labels on the axes. Indeed, there is really just one basic curve from which all of these curves can be drawn just by relabeling the axes appropriately.
#
# To draw that basic curve, we will use the units into which we can convert every list: standard units. The resulting curve is therefore called the *standard normal curve*.
# The standard normal curve has an impressive equation. But for now, it is best to think of it as a smoothed outline of a histogram of a variable that has been measured in standard units and has a bell-shaped distribution.
#
# $$
# \phi(z) = {\frac{1}{\sqrt{2 \pi}}} e^{-\frac{1}{2}z^2}, ~~ -\infty < z < \infty
# $$
# +
# HIDDEN
# The standard normal curve
plot_normal_cdf()
# -
# As always when you examine a new histogram, start by looking at the horizontal axis. On the horizontal axis of the standard normal curve, the values are standard units.
#
# Here are some properties of the curve. Some are apparent by observation, and others require a considerable amount of mathematics to establish.
#
# - The total area under the curve is 1. So you can think of it as a histogram drawn to the density scale.
#
# - The curve is symmetric about 0. So if a variable has this distribution, its mean and median are both 0.
#
# - The points of inflection of the curve are at -1 and +1.
#
# - If a variable has this distribution, its SD is 1. The normal curve is one of the very few distributions that has an SD so clearly identifiable on the histogram.
# Since we are thinking of the curve as a smoothed histogram, we will want to represent proportions of the total amount of data by areas under the curve.
#
# Areas under smooth curves are often found by calculus, using a method called integration. It is a fact of mathematics, however, that the standard normal curve cannot be integrated in any of the usual ways of calculus.
#
# Therefore, areas under the curve have to be approximated. That is why almost all statistics textbooks carry tables of areas under the normal curve. It is also why all statistical systems, including a module of Python, include methods that provide excellent approximations to those areas.
from scipy import stats
# ### The standard normal "cdf" ###
#
# The fundamental function for finding areas under the normal curve is `stats.norm.cdf`. It takes a numerical argument and returns all the area under the curve to the left of that number. Formally, it is called the "cumulative distribution function" of the standard normal curve. That rather unwieldy mouthful is abbreviated as cdf.
#
# Let us use this function to find the area to the left of $z=1$ under the standard normal curve.
# +
# HIDDEN
# Area under the standard normal curve, below 1
plot_normal_cdf(1)
# -
# The numerical value of the shaded area can be found by calling `stats.norm.cdf`.
stats.norm.cdf(1)
# That's about 84%. We can now use the symmetry of the curve and the fact that the total area under the curve is 1 to find other areas.
#
# The area to the right of $z=1$ is about 100% - 84% = 16%.
# +
# HIDDEN
# Area under the standard normal curve, above 1
plot_normal_cdf(lbound=1)
# -
1 - stats.norm.cdf(1)
# The area between $z=-1$ and $z=1$ can be computed in several different ways. It is the gold area under the curve below.
# +
# HIDDEN
# Area under the standard normal curve, between -1 and 1
plot_normal_cdf(1, lbound=-1)
# -
# For example, we could calculate the area as "100% - two equal tails", which works out to roughly 100% - 2x16% = 68%.
#
# Or we could note that the area between $z=1$ and $z=-1$ is equal to all the area to the left of $z=1$, minus all the area to the left of $z=-1$.
stats.norm.cdf(1) - stats.norm.cdf(-1)
# By a similar calculation, we see that the area between $-2$ and 2 is about 95%.
# +
# HIDDEN
# Area under the standard normal curve, between -2 and 2
plot_normal_cdf(2, lbound=-2)
# -
stats.norm.cdf(2) - stats.norm.cdf(-2)
# In other words, if a histogram is roughly bell shaped, the proportion of data in the range "average $\pm$ 2 SDs" is about 95%.
#
# That is quite a bit more than Chebychev's lower bound of 75%. Chebychev's bound is weaker because it has to work for all distributions. If we know that a distribution is normal, we have good approximations to the proportions, not just bounds.
# The table below compares what we know about all distributions and about normal distributions. Notice that when $z=1$, Chebychev's bound is correct but not illuminating.
#
# | Percent in Range | All Distributions: Bound | Normal Distribution: Approximation |
# | :--------------- | :---------------- --| :-------------------|
# |average $\pm$ 1 SD | at least 0% | about 68% |
# |average $\pm$ 2 SDs | at least 75% | about 95% |
# |average $\pm$ 3 SDs | at least 88.888...% | about 99.73% |
| notebooks/14/3/SD_and_the_Normal_Curve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# example of a stacked model for binary classification
from numpy import hstack
from numpy import array
from sklearn.datasets import make_blobs
from sklearn.model_selection import KFold
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
# +
# create a meta dataset
def create_meta_dataset(data_x, yhat1, yhat2):
# convert to columns
yhat1 = array(yhat1).reshape((len(yhat1), 1))
yhat2 = array(yhat2).reshape((len(yhat2), 1))
# stack as separate columns
meta_X = hstack((data_x, yhat1, yhat2))
return meta_X
# make predictions with stacked model
def stack_prediction(model1, model2, meta_model, X):
# make predictions
yhat1 = model1.predict_proba(X)[:, 0]
yhat2 = model2.predict_proba(X)[:, 0]
# create input dataset
meta_X = create_meta_dataset(X, yhat1, yhat2)
# predict
return meta_model.predict(meta_X)
# +
# create the inputs and outputs
X, y = make_blobs(n_samples=1000, centers=2, n_features=100, cluster_std=20)
# split
X, X_val, y, y_val = train_test_split(X, y, test_size=0.33)
# collect out of sample predictions
data_x, data_y, knn_yhat, cart_yhat = list(), list(), list(), list()
kfold = KFold(n_splits=10, shuffle=True)
for train_ix, test_ix in kfold.split(X):
# get data
train_X, test_X = X[train_ix], X[test_ix]
train_y, test_y = y[train_ix], y[test_ix]
data_x.extend(test_X)
data_y.extend(test_y)
# fit and make predictions with cart
model1 = DecisionTreeClassifier()
model1.fit(train_X, train_y)
yhat1 = model1.predict_proba(test_X)[:, 0]
cart_yhat.extend(yhat1)
# fit and make predictions with cart
model2 = KNeighborsClassifier()
model2.fit(train_X, train_y)
yhat2 = model2.predict_proba(test_X)[:, 0]
knn_yhat.extend(yhat2)
# construct meta dataset
meta_X = create_meta_dataset(data_x, knn_yhat, cart_yhat)
# fit final submodels
model1 = DecisionTreeClassifier()
model1.fit(X, y)
model2 = KNeighborsClassifier()
model2.fit(X, y)
# construct meta classifier
meta_model = LogisticRegression(solver='liblinear')
meta_model.fit(meta_X, data_y)
# evaluate sub models on hold out dataset
acc1 = accuracy_score(y_val, model1.predict(X_val))
acc2 = accuracy_score(y_val, model2.predict(X_val))
print('Model1 Accuracy: %.3f, Model2 Accuracy: %.3f' % (acc1, acc2))
# evaluate meta model on hold out dataset
yhat = stack_prediction(model1, model2, meta_model, X_val)
acc = accuracy_score(y_val, yhat)
print('Meta Model Accuracy: %.3f' % (acc))
| notebooks/machine_learning_algorithms/8d-Ensemble-Stacking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
Image('../Images/introduction-to-machine-learning_social.png')
# > **Hello and Welcome back**
#
# > **In this module we are Building model of data categorised from clutser 3.**
#
# > **Cluster 3 was obtained from doc2vec of (ftca+Consumer compliance handbook) and Grievance description**
#
# # Table of Contents
# 1. [Step 1 - Import libraries](#ch1)
# 1. [Step 2 - Step 1: Read the dataset](#ch2)
# 1. [Step 3 - Data Exploration](#ch3)
# 1. [Step 4 - preprocessing](#ch4)
# 1. [Step 5 - Models](#ch5)
# 1. [Step 6 - Deep Learning](#ch10)
# 1. [Step 7 - SMOTE](#ch8)
# 1. [Step 8 - Model building using SMOTE](#ch9)
# 1. [Step 9 - Cross validation method of model building](#ch6)
# 1. [Step 10 - grid Search](#ch11)
# <a id="ch1"></a>
# # Import Libraries
# +
# This Python 3 environment comes with many helpful analytics libraries installed
#load packages
import sys #access to system parameters https://docs.python.org/3/library/sys.html
print("Python version: {}". format(sys.version))
import pandas as pd #collection of functions for data processing and analysis modeled after R dataframes with SQL like features
print("pandas version: {}". format(pd.__version__))
import matplotlib #collection of functions for scientific and publication-ready visualization
print("matplotlib version: {}". format(matplotlib.__version__))
import numpy as np #foundational package for scientific computing
print("NumPy version: {}". format(np.__version__))
import scipy as sp #collection of functions for scientific computing and advance mathematics
print("SciPy version: {}". format(sp.__version__))
import IPython
from IPython import display #pretty printing of dataframes in Jupyter notebook
print("IPython version: {}". format(IPython.__version__))
import sklearn #collection of machine learning algorithms
print("scikit-learn version: {}". format(sklearn.__version__))
#misc libraries
import random
import time
#ignore warnings
import warnings
warnings.filterwarnings('ignore')
# +
#Common Model Algorithms
from sklearn import svm, tree, linear_model, neighbors, naive_bayes, ensemble, discriminant_analysis, gaussian_process
from xgboost import XGBClassifier
#Common Model Helpers
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
from sklearn import feature_selection
from sklearn import model_selection
from sklearn import metrics
from sklearn.model_selection import train_test_split,GridSearchCV
from sklearn.metrics import accuracy_score
from sklearn import preprocessing
#Visualization
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
from pandas.tools.plotting import scatter_matrix
#Configure Visualization Defaults
# #%matplotlib inline = show plots in Jupyter Notebook browser
# %matplotlib inline
mpl.style.use('ggplot')
sns.set_style('white')
pylab.rcParams['figure.figsize'] = 12,8
# -
# Importing Classifier Modules
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC, LinearSVC
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import Perceptron
from sklearn.linear_model import SGDClassifier
from sklearn.ensemble import AdaBoostClassifier
from imblearn.over_sampling import SMOTE # Oversampling
from keras.layers import LSTM, Dropout
from sklearn.neural_network import MLPClassifier
# <a id="ch2"></a>
# # Read the dataset
# Reading the train and test data using pd.read_csv
train_data = pd.read_csv('../Model Building dataset/final_train_data.csv')
test_data = pd.read_csv('../Model Building dataset/final_test_data.csv')
# Shufling the data using sample(frac=1), where frac =1 would mean entire data
train_data = train_data.sample(frac=1)
test_data = test_data.sample(frac=1)
# In order to print the first 10 rows use .head()
train_data.head()
# In order to print the first 10 rows use .head()
test_data.head()
# +
#train_data.BankGrade.replace({'deficient':'0','satisfactory':'1','outstanding':'2'})
# -
# <a id="ch3"></a>
# # Data Exploration
# Creating a datafarme of BankID and GrievanceID column, which we willbe using in the end for combining the predictions.
test_data_bankID_GrievanceID = test_data[['BankID','GrievanceID']]
# Checking the data types of the data
train_data.dtypes
# Checking the data types of the data
train_data.dtypes
# Check if there are any common BankID in train and test data
set(train_data['BankID']).intersection(set(test_data['BankID']))
# Renaming the columns names of the clusters in test data with the of train data cluster column names.
test_data.rename(columns={'test_result_Cluster_using_doc2vec_pdf_normal_method': 'result_Cluster_using_doc2vec_pdf_normal_method',
'result_cluster_using_cleaned_data_test': 'result_cluster_using_cleaned_data' ,
'result_cluster_pdf_combined_test': 'result_cluster_pdf_combined',
'result_Cluster_tfidf_test':'result_Cluster_tfidf'}, inplace=True)
# Selecting the important attributes for model building.
train_X_columns_cluster_3 = ['LineOfBusiness','ResolutionComments', 'Disputed','DAYSDIFF', 'result_cluster_pdf_combined']
# We have removed GrievanceID,BankID,
#
# State - Without state column accuracy is increasing.
#
# Grievance category - It is a sub category of Line of Business.
#
# DateOfGrievance, DateOfResolution - We have taken a difference and stored them in DAYDIFF column.
train_Y_columns = ['BankGrade']
# taking only the data of the columns specidfied in "train_X_columns_cluster_3"
train_X_cluster_3 = train_data.loc[:,train_X_columns_cluster_3]
# Taking the data stored in column BankGrade
train_Y = train_data.loc[:,train_Y_columns]
# #### Test part
# taking only the data of the columns specidfied in "train_X_columns_cluster_3"
test_data_cluster_3 = test_data.loc[:, train_X_columns_cluster_3]
# <a id="ch4"></a>
# # Preprocessing
# # Cluster 3
# ## Method 1: Dummies
# # Dummies
# We are doing dummy only for columns for which the order doesn't matter. In the data we are selecting "State", "Grievance_Category", "LineOfBusiness".
# ###### Train Part
# Dummyfying the columns because they are of categorical type
train_X_cluster_3 =pd.get_dummies(train_X_cluster_3)
train_X_cluster_3.shape # To check the shape of the data
# ##### Test part
# Dummyfying the columns because they are of categorical type
test_data_cluster_3 =pd.get_dummies(test_data_cluster_3)
test_data_cluster_3.shape # To check the shape of the data
# # Train validation split
# Splitting the data into train and valid with 70% train and 30% validation
train_split_X, val_split_X, train_split_Y, val_split_Y = train_test_split(train_X_cluster_3, train_Y, test_size=0.30, random_state=42)
# <a id="ch5"></a>
# # Model buidling
# # Logistic regression
# +
# Creating an object for logistic regression
clf = LogisticRegression()
#Using the object created for logistic regression to fit on train data
clf.fit(train_split_X, train_split_Y)
##Using the object created for logistic regression to predict on validation data
y_pred_log_reg = clf.predict(val_split_X)
#Using the object created for logistic regression to predict on test data
test_y_pred_log_reg = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_log_reg_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_log_reg: ' +' '+str(acc_log_reg_train) + ' percent')
#Calculating the accuracy of validation data
acc_log_reg_validation = accuracy_score(val_split_Y, y_pred_log_reg)
print('validation_accuracy_log_reg: '+str(round((acc_log_reg_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['log_reg_pred']=test_y_pred_log_reg
#Grouping the BankID and log_reg_pred
test_predictions_log_reg=test_data.groupby(['BankID','log_reg_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_log_reg.log_reg_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_log_reg =test_predictions_log_reg.log_reg_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_log_reg.to_csv('../Submission/Cluster_3_predictions/test_predictions_log_reg.csv')
# -
# # DT
# +
# Creating an object for decision Trees
clf = DecisionTreeClassifier()
#Using the object created for decision Trees to fit on train data
clf.fit(train_split_X, train_split_Y)
#Using the object created for decision Trees to predict on validation data
y_pred_dt = clf.predict(val_split_X)
#Using the object created for decision Trees to predict on test data
test_y_pred_dt = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_dt_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_dt: ' +' '+str(acc_dt_train) + ' percent')
#Calculating the accuracy of validation data
acc_dt_validation = accuracy_score(val_split_Y, y_pred_dt)
print('validation_accuracy_dt: '+str(round((acc_dt_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['dt_pred']=test_y_pred_dt
#Grouping the BankID and dt_pred
test_predictions_dt=test_data.groupby(['BankID','dt_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_dt.dt_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_dt =test_predictions_dt.dt_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_dt.to_csv('../Submission/Cluster_3_predictions/test_predictions_dt.csv')
# -
# ## RF
# +
# Creating an object for Random Forest
clf = RandomForestClassifier(n_estimators=100)
#Using the object created for Random Forest to fit on train data
clf.fit(train_split_X, train_split_Y)
#Using the object created for Random Forest to predict on validation data
y_pred_rf = clf.predict(val_split_X)
#Using the object created for Random Forest to predict on test data
test_y_pred_rf = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_rf_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_rf: ' +' '+str(acc_rf_train) + ' percent')
#Calculating the accuracy of validation data
acc_rf_validation = accuracy_score(val_split_Y, y_pred_rf)
print('validation_accuracy_rf: '+str(round((acc_rf_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['rf_pred']=test_y_pred_rf
#Grouping the BankID and rf_pred
test_predictions_rf=test_data.groupby(['BankID','rf_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_rf.rf_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_rf =test_predictions_rf.rf_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_rf.to_csv('../Submission/Cluster_3_predictions/test_predictions_rf.csv')
# -
# ## GNB
# +
# Creating an object for Gaussian naive Bayes
clf = GaussianNB()
#Using the object created for Gaussian naive Bayes to fit on train data
clf.fit(train_split_X, train_split_Y)
#Using the object created for Gaussian naive Bayes to predict on validation data
y_pred_gnb = clf.predict(val_split_X)
#Using the object created for Gaussian naive Bayes to predict on test data
test_y_pred_gnb = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_gnb_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_gnb: ' +' '+str(acc_gnb_train) + ' percent')
#Calculating the accuracy of validation data
acc_gnb_validation = accuracy_score(val_split_Y, y_pred_gnb)
print('validation_accuracy_gnb: '+str(round((acc_gnb_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['gnb_pred']=test_y_pred_gnb
#Grouping the BankID and gnb_pred
test_predictions_gnb=test_data.groupby(['BankID','gnb_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_gnb.gnb_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_gnb =test_predictions_gnb.gnb_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_gnb.to_csv('../Submission/Cluster_3_predictions/test_predictions_gnb.csv')
# -
# # Adaboost
# +
# Creating an object for Adaboost
clf = AdaBoostClassifier()
#Using the object created for Adaboost to fit on train data
clf.fit(train_split_X, train_split_Y)
#Using the object created for Adaboost to predict on validation data
y_pred_adaboost = clf.predict(val_split_X)
#Using the object created for Adaboost to predict on test data
test_y_pred_adaboost = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_adaboost_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_adaboost: ' +' '+str(acc_adaboost_train) + ' percent')
#Calculating the accuracy of validation data
acc_adaboost_validation = accuracy_score(val_split_Y, y_pred_adaboost)
print('validation_accuracy_adaboost: '+str(round((acc_adaboost_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['ada_pred']=test_y_pred_adaboost
#Grouping the BankID and ada_pred
test_predictions_ada=test_data.groupby(['BankID','ada_pred'])
# Printing the value counts of each and their subclasses
test_predictions_ada.ada_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_ada =test_predictions_ada.ada_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_ada.to_csv('../Submission/Cluster_3_predictions/test_predictions_ada.csv')
# -
# ## SGD
# +
# Creating an object for Stocahstic gradient decsent
clf = SGDClassifier(max_iter=5, tol=None)
#Using the object created for Stocahstic gradient decsent to fit on train data
clf.fit(train_split_X, train_split_Y)
#Using the object created for Stocahstic gradient decsent to predict on validation data
y_pred_sgd = clf.predict(val_split_X)
#Using the object created for Stocahstic gradient decsent to predict on test data
test_y_pred_sgd = clf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_sgd_train = round( clf.score(train_split_X, train_split_Y) * 100, 2)
print ('train_accuracy_log_reg' +' '+str(acc_sgd_train) + ' percent')
#Calculating the accuracy of validation data
acc_sgd_validation = accuracy_score(val_split_Y, y_pred_sgd)
print('validation_accuracy_log_reg '+str(round((acc_sgd_validation)*100,2)) + 'percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['sgd_pred']=test_y_pred_sgd
#Grouping the BankID and sgd_pred
test_predictions_sgd=test_data.groupby(['BankID','sgd_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_sgd.sgd_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_sgd =test_predictions_sgd.sgd_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_sgd.to_csv('../Submission/Cluster_3_predictions/test_predictions_sgd.csv')
# +
# creating a dataframe that will print the accuarcy of train and test of all the models.
models = pd.DataFrame({
'Model': ['Logistic Regression',
'Decision Tree',
'Random Forest',
'Naive Bayes (GNB)',
'Adaboost',
'Stochastic Gradient Decent'],
'Score_train': [acc_log_reg_train,
acc_dt_train,
acc_rf_train,
acc_gnb_train,
acc_adaboost_train,
acc_sgd_train],
'Score_valid': [acc_log_reg_validation,
acc_dt_validation,
acc_rf_validation,
acc_gnb_validation,
acc_adaboost_validation,
acc_sgd_validation]
})
models.sort_values(by='Score_train', ascending=False) # sorting by score_train
# -
models.sort_values(by='Score_valid', ascending=False)# sorting by score_valid
# <a id="ch10"></a>
# # Deep learning
#importing models for deeplearning
from keras.models import Sequential
from keras.layers import Dense
from keras.utils import to_categorical
from keras.utils import np_utils, normalize
#Converting the target column in to values 0,1 and 2.
train_Y_values=train_data.BankGrade.replace({'deficient':'0','satisfactory':'1','outstanding':'2'})
train_Y.shape
train_Y_values= pd.DataFrame(train_Y_values)
train_Y_values =train_Y_values.BankGrade.astype('int')
train_Y_values.dtypes
train_Y_values= pd.DataFrame(train_Y_values)
labels_np_utils = np_utils.to_categorical(train_Y_values["BankGrade"]-1,num_classes =3)
labels_np_utils.shape
# +
mlp_model = Sequential()#creating a sequential model
mlp_model.add(Dense(128, input_dim=12, activation='relu', kernel_initializer='normal'))
mlp_model.add(Dense(64, activation='relu', kernel_initializer='normal'))
mlp_model.add(Dense(32, activation='relu', kernel_initializer='normal'))
mlp_model.add(Dense(16, activation='relu', kernel_initializer='normal'))
mlp_model.add(Dense(3, activation='softmax', kernel_initializer='normal'))
# -
mlp_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
mlp_model.fit(train_X_cluster_3, labels_np_utils, epochs=30, batch_size=64)
# +
test_pred_mlp_DL =mlp_model.predict_classes(test_data_cluster_3)
train_pred_mlp_DL =mlp_model.predict_classes(train_X_cluster_3)
# -
#printing the tarin accuracy
acc_train_mlp_DL = round( clf.score(train_X_cluster_3, train_Y) * 100, 2)
print ('train_accuracy_log_reg' +' '+str(acc_train_mlp_DL) + ' percent')
# +
# In the below method we are copying the predictions and labelling them.
test_data['mlp_DL_pred']=test_pred_mlp_DL
#Replacing the numeric prediction by their category names.
test_data['mlp_DL_pred']=test_data['mlp_DL_pred'].replace({0:'deficient',1:'satisfactory',2:'outstanding'})
#test_data['mlp_DL_pred']=test_pred_mlp_DL
#Grouping the BankID and mlp_DL_pred
test_predictions_mlp_DL=test_data.groupby(['BankID','mlp_DL_pred'])
# Printing the value counts of each and their subclasses.
test_predictions_mlp_DL.mlp_DL_pred.value_counts()
# + active=""
# # Storing the value counts in another variable
# test_predictions_mlp_DL =test_predictions_mlp_DL.mlp_DL_pred.value_counts()
#
# # Transferring the values to a csv file.
# test_predictions_mlp_DL.to_csv('../Submission trials/cluster3/test_predictions_mlp_DL.csv')
# -
# <a id="ch8"></a>
# # SMOTE
train_Y.head(1)
#Creating a SMOTE object
sm = SMOTE(random_state=2)
#Smoting the data into smoteX and smoteY
X_train_smote, y_train_smote = sm.fit_sample(train_X_cluster_3, train_Y)
# checking the number of rows in X_train after smote
X_train_smote.shape
# checking the number of rows in original train_X_cluster_3
train_X_cluster_3.shape
# checking the number of rows in original train_Y
train_Y.shape
# checking the number of rows in Y_train after smote
y_train_smote.shape
# Counting the number of rows containing outstanding as the category after SMOTE
(y_train_smote=='outstanding').sum()
# Counting the number of rows containing deficient as the category after SMOTE
(y_train_smote=='deficient').sum()
# Counting the number of rows containing satisfactory as the category after SMOTE
(y_train_smote=='satisfactory').sum()
# Counting the number of rows containing outstanding as the category
(train_Y=='outstanding').sum()
# Counting the number of rows containing deficient as the category
(train_Y=='deficient').sum()
# Counting the number of rows containing satisfactory as the category
(train_Y=='satisfactory').sum()
# <a id="ch9"></a>
# # Model building using SMOTE
# ## Logistic regression SMOTE
# +
# Creating an object for logistic regression
lr = LogisticRegression()
#Using the object created for logistic regression to fit on train data
lr.fit(X_train_smote, y_train_smote)
#Using the object created for logistic regression to predict on test data
pred_lr_SMOTE = lr.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_log = round(lr.score(X_train_smote, y_train_smote)*100,2)
acc_log
# +
# In the below method we are copying the predictions and labelling them.
test_data['log_reg_pred_smote']=pred_lr_SMOTE
#Grouping the BankID and log_reg_pred_smote
SMOTE_predictions_log_reg=test_data.groupby(['BankID','log_reg_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_log_reg.log_reg_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_log_reg =SMOTE_predictions_log_reg.log_reg_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_log_reg.to_csv('../Submission trials/cluster3/SMOTE_predictions_log_reg.csv')
# -
# # Decision tree smote
# +
# Creating an object for decision trees
dt = DecisionTreeClassifier()
#Using the object created for decision trees to fit on train data
dt.fit(X_train_smote, y_train_smote)
#Using the object created for decision trees to predict on test data
pred_dt_SMOTE = dt.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_dt = round(dt.score(X_train_smote, y_train_smote)*100,2)
print(acc_dt)
# In the below method we are copying the predictions and labelling them.
test_data['dt_pred_smote']=pred_dt_SMOTE
#Grouping the BankID and dt_pred_smote
SMOTE_predictions_dt=test_data.groupby(['BankID','dt_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_dt.dt_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_dt=SMOTE_predictions_dt.dt_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_dt.to_csv('../Submission trials/cluster3/SMOTE_predictions_dt.csv')
# -
# ## random forest SMOTE
# +
# Creating an object for random forest
rf=RandomForestClassifier(n_estimators=100)
#Using the object created for random forest to fit on train data
rf.fit(X_train_smote, y_train_smote)
#Using the object created for random forest to predict on test data
pred_rf_SMOTE = rf.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_rf = round(rf.score(X_train_smote, y_train_smote)*100,2)
acc_rf
# +
# In the below method we are copying the predictions and labelling them.
test_data['rf_pred_smote']=pred_rf_SMOTE
#Grouping the BankID and rf_pred_smote
SMOTE_predictions_rf=test_data.groupby(['BankID','rf_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_rf.rf_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_rf =SMOTE_predictions_rf.rf_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_rf.to_csv('../Submission trials/cluster3/SMOTE_predictions_rf.csv')
# -
# # GNB SMOTE
# +
# Creating an object for Gaussian Naive Bayes
gnb = GaussianNB()
#Using the object created for Gaussian Naive Bayes to fit on train data
gnb.fit(X_train_smote, y_train_smote)
#Using the object created for Gaussian Naive Bayes to predict on test data
pred_gnb_SMOTE = gnb.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_gnb = round(gnb.score(X_train_smote, y_train_smote)*100,2)
print(acc_gnb)
# In the below method we are copying the predictions and labelling them.
test_data['gnb_pred_smote']=pred_gnb_SMOTE
#Grouping the BankID and gnb_pred_smote
SMOTE_predictions_gnb=test_data.groupby(['BankID','gnb_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_gnb.gnb_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_gnb =SMOTE_predictions_gnb.gnb_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_gnb.to_csv('../Submission trials/SMOTE_predictions_gnb.csv')
# -
# # Adaboost SMOTE
# +
# Creating an object for adaboost
ada = AdaBoostClassifier()
#Using the object created for adaboost to fit on train data
ada.fit(X_train_smote, y_train_smote)
#Using the object created for adaboost to predict on test data
pred_ada_SMOTE = ada.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_ada = round(ada.score(X_train_smote, y_train_smote)*100,2)
print(acc_ada)
# In the below method we are copying the predictions and labelling them.
test_data['ada_pred_smote']=pred_ada_SMOTE
#Grouping the BankID and ada_pred_smote
SMOTE_predictions_ada=test_data.groupby(['BankID','ada_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_ada.ada_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_ada =SMOTE_predictions_ada.ada_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_ada.to_csv('../Submission trials/SMOTE_predictions_ada.csv')
# -
# # SGD SMOTE
# +
# Creating an object for Stochastic Gradient descent
sgd = SGDClassifier(max_iter=5, tol=None)
#Using the object created for Stochastic Gradient descent to fit on train data
sgd.fit(X_train_smote, y_train_smote)
#Using the object created for Stochastic Gradient descent to predict on test data
pred_sgd_SMOTE = sgd.predict(test_data_cluster_3)
#Calculating the accuracy of train data
acc_sgd = round(sgd.score(X_train_smote, y_train_smote)*100,2)
print(acc_sgd)
# In the below method we are copying the predictions and labelling them.
test_data['sgd_pred_smote']=pred_sgd_SMOTE
#Grouping the BankID and sgd_pred_smote
SMOTE_predictions_sgd=test_data.groupby(['BankID','sgd_pred_smote'])
# Printing the value counts of each and their subclasses.
SMOTE_predictions_sgd.sgd_pred_smote.value_counts()
# + active=""
# # Storing the value counts in another variable
# SMOTE_predictions_sgd =SMOTE_predictions_sgd.sgd_pred_smote.value_counts()
#
# # Transferring the values to a csv file.
# SMOTE_predictions_sgd.to_csv('../Submission trials/SMOTE_predictions_sgd.csv')
# +
# creating a dataframe that will print the accuarcy of train and test of all the models.
models_SMOTE = pd.DataFrame({
'Model': ['Logistic Regression',
'Decision Tree',
'Random Forest',
'Naive Bayes (GNB)',
'Adaboost',
'Stochastic Gradient Decent'],
'Score_train': [acc_log,
acc_dt,
acc_rf,
acc_gnb,
acc_ada,
acc_sgd
]
})
models.sort_values(by='Score_train', ascending=False) # sorting by score_train
# -
# # Predictions selected to create csv file
# <a id="ch6"></a>
# # Cross validation method of model building
data1 = pd.concat([train_X_cluster_3, train_Y], axis=1)
data1.shape
Target = ['BankGrade']
data1_x_bin=train_X_cluster_3.columns
test_data_cluster_3_X_bin = train_X_cluster_3.columns
# +
#Machine Learning Algorithm (MLA) Selection and Initialization
MLA = [
#Ensemble Methods
#ensemble.AdaBoostClassifier(),
#ensemble.BaggingClassifier(),
#ensemble.ExtraTreesClassifier(),
#ensemble.GradientBoostingClassifier(),
ensemble.RandomForestClassifier(),
#Gaussian Processes
#gaussian_process.GaussianProcessClassifier(),
#GLM
linear_model.LogisticRegressionCV(),
#linear_model.PassiveAggressiveClassifier(),
#linear_model.RidgeClassifierCV(),
#linear_model.SGDClassifier(),
#linear_model.Perceptron(),
#Navies Bayes
#naive_bayes.BernoulliNB(),
#naive_bayes.GaussianNB(),
#Nearest Neighbor
#neighbors.KNeighborsClassifier(),
#SVM
#svm.SVC(probability=True),
#svm.NuSVC(probability=True),
#svm.LinearSVC(),
#Trees
#tree.DecisionTreeClassifier(),
#tree.ExtraTreeClassifier(),
#Discriminant Analysis
#discriminant_analysis.LinearDiscriminantAnalysis(),
#discriminant_analysis.QuadraticDiscriminantAnalysis(),
#xgboost: http://xgboost.readthedocs.io/en/latest/model.html
#XGBClassifier()
]
#split dataset in cross-validation with this splitter class: http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.ShuffleSplit.html#sklearn.model_selection.ShuffleSplit
#note: this is an alternative to train_test_split
cv_split = model_selection.ShuffleSplit(n_splits = 10, test_size = .3, train_size = .6, random_state = 0 ) # run model 10x with 60/30 split intentionally leaving out 10%
#create table to compare MLA metrics
MLA_columns = ['MLA Name', 'MLA Parameters','MLA Train Accuracy Mean', 'MLA Test Accuracy Mean' ,'MLA Time']
MLA_compare = pd.DataFrame(columns = MLA_columns)
#create table to compare MLA predictions
MLA_predict = data1[Target]
MLA_predictions=test_data[['sgd_pred_smote']]
#index through MLA and save performance to table
row_index = 0
for alg in MLA:
#set name and parameters
MLA_name = alg.__class__.__name__
MLA_compare.loc[row_index, 'MLA Name'] = MLA_name
MLA_compare.loc[row_index, 'MLA Parameters'] = str(alg.get_params())
#score model with cross validation: http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_validate.html#sklearn.model_selection.cross_validate
cv_results = model_selection.cross_validate(alg, data1[data1_x_bin], data1[Target], cv = cv_split)
MLA_compare.loc[row_index, 'MLA Time'] = cv_results['fit_time'].mean()
MLA_compare.loc[row_index, 'MLA Train Accuracy Mean'] = cv_results['train_score'].mean()
MLA_compare.loc[row_index, 'MLA Test Accuracy Mean'] = cv_results['test_score'].mean()
#if this is a non-bias random sample, then +/-3 standard deviations (std) from the mean, should statistically capture 99.7% of the subsets
# MLA_compare.loc[row_index, 'MLA Test Accuracy 3*STD'] = cv_results['test_score'].std()*3 #let's know the worst that can happen!
#save MLA predictions - see section 6 for usage
alg.fit(data1[data1_x_bin], data1[Target])
MLA_predict[MLA_name] = alg.predict(data1[data1_x_bin])
MLA_predictions[MLA_name] = alg.predict(test_data_cluster_3[test_data_cluster_3_X_bin])
row_index+=1
#print and sort table: https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.sort_values.html
MLA_compare.sort_values(by = ['MLA Test Accuracy Mean'], ascending = False, inplace = True)
print(MLA_predict)
print(MLA_predictions)
MLA_compare
# +
test_data['RandomForestClassifier']=MLA_predictions.iloc[:,1]
test_data
#Grouping the BankID and ada_pred_smote
MLA_groupby=test_data.groupby(['BankID','RandomForestClassifier'])
# Printing the value counts of each and their subclasses.
MLA_groupby.RandomForestClassifier.value_counts()
# + active=""
# # Storing the value counts in another variable
# CV_RandomForestClassifier =MLA_groupby.RandomForestClassifier.value_counts()
#
# # Transferring the values to a csv file.
# CV_RandomForestClassifier.to_csv('../Submission trials/CV_RandomForestClassifier.csv')
# + active=""
# prediction_cross_validation = {'Adaboost_train' : 0.666067,
# 'Adaboost_test' : 0.666207,
# 'Bagging_classifier_train' : 0.979213,
# 'Bagging_classifier_test' : 0.647783,
# 'GradientBoostingClassifier_train': 0.685112,
# 'GradientBoostingClassifier_test' : 0.677974,
# 'RandomForestClassifier_train' : 0.974354,
# 'RandomForestClassifier_test' : 0.627919,
# 'ExtraTreesClassifier_train': 1,
# 'ExtraTreesClassifier_test':0.613525,
# 'LogisticRegressionCV_train':0.653456,
# 'LogisticRegressionCV_test':0.654036,
# 'PassiveAggressiveClassifier_train':0.526301
# 'PassiveAggressiveClassifier_test':0.527341
# 'RidgeClassifierCV_train':0.670318
# 'RidgeClassifierCV_test':0.66699
# 'SGDClassifier_train':0.53266
# 'SGDClassifier_test':0.534389
# 'Perceptron_train':0.402776
# 'Perceptron_test':0.402819
# 'BernoulliNB_train':0.632874
# 'BernoulliNB_test':0.633371
# 'GaussianNB_train':0.644303
# 'GaussianNB_test':0.64448
#
# +
#barplot using https://seaborn.pydata.org/generated/seaborn.barplot.html
sns.barplot(x='MLA Test Accuracy Mean', y = 'MLA Name', data = MLA_compare, color = 'm')
#prettify using pyplot: https://matplotlib.org/api/pyplot_api.html
plt.title('Machine Learning Algorithm Accuracy Score \n')
plt.xlabel('Accuracy Score (%)')
plt.ylabel('Algorithm')
# -
# <a id="ch11"></a>
# ## Grid Search
# + active=""
# #Grid search on deeplearning model
# from sklearn.model_selection import GridSearchCV
# from keras.models import Sequential
# from keras.layers import Dense
# from keras.wrappers.scikit_learn import KerasClassifier
#
# #Create a function to read a model
# def create_model(optimizer='adam'):
#
# model = Sequential()
# model.add(Dense(64, input_dim = 12, activation='relu'))
# model.add(Dense(32, activation='relu'))
# model.add(Dense(16, activation='relu'))
# model.add(Dense(8, activation='relu'))
# model.add(Dense(3, activation='softmax'))
#
# model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# return model
#
# # storing train_X_cluster_3 and target values in X and Y
# X=train_X_cluster_3
# Y= labels_np_utils
#
#
# model = KerasClassifier(build_fn=create_model, epochs =100, batch_size = 64, verbose=0 )
#
# #Different optimizers to be used. You can even increase the list of optimizers by adding more optimizers
# optimizer=['SGD', "RMSprop", "Adagrad", "Adadelta", "Adam", "Adamax", "Nadam"]
#
# #Creating a parametere grid to be searched
# param_grid = dict(optimizer=optimizer)
#
# #With the help of grid search method we are finding the best paarameters.
# grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=-1)
#
# #Storing the grid result for predictions
# grid_result = grid.fit(X,Y)
#
# #Printing the best parameters and best score of GridSearchCV
# print("Best %f using %s" %(grid_result.best_score_, grid_result.best_params_))
#
# #Taking a mean of test score
# means = grid_result.cv_results_['mean_test_score']
#
# #Taking the parameters used
# params = grid_result.cv_results_['params']
#
#
# for mean, param in zip(means, params):
# print("%f with: %r" %(mean, param))
# + active=""
# # Grid search on XGB classifier model
# + active=""
# #Giving a list of parameters to be searched by GridSearchCV
# clf_tune = XGBClassifier(n_estimators= 300, n_jobs= -1)
# parameters = {'max_depth': [5,10,12,15,20],
# 'min_child_weight':[2,3,4,5,7],
# 'reg_lambda': [0.50,0.75,1,1.25,1.5]}
# + active=""
# #Performing GridSearch on the train data
# grid = GridSearchCV(clf_tune, param_grid = parameters, n_jobs= -1, cv = 3 )
# grid.fit(train_split_X, train_split_Y)
# print('Best parameters: {}'.format(grid.best_params_))
# + active=""
# #Selecting the best paramters
# MAX_DEPTH = grid.best_params_['max_depth']
# MIN_CHILD_WEIGHT = grid.best_params_['min_child_weight']
# REG_LAMDBA = grid.best_params_['reg_lambda']
# + active=""
# #After selecting the best parameters training the model on those parameters
# clf_post_tune = XGBClassifier(max_depth= MAX_DEPTH , n_estimators= 500,
# min_child_weight= MIN_CHILD_WEIGHT, reg_lambda= REG_LAMDBA)
# + active=""
# #Fitting the model on best parameters
# clf_post_tune.fit(train_split_X, train_split_Y)
# + active=""
# #Calculating the accuracy score on validation data
# cross_val_score(clf_post_tune, val_split_X,val_split_Y, cv = 5, scoring= 'accuracy') .mean()
# -
# # Best accuracy on test data was obtained from randomforest from cross validation method -> 68%
# # Model selection
from IPython.display import Image
Image('../Images/Criteria for selection.png')
# **Keeping the above three dimensions for model selection in mind, I have selected RandomForest as my final model for my predictions.**
#
# > **Accuarcy -> It gave me the best accuracy when compared to other models.**
#
# > **Interpretability -> Random Forest is easy to intrpret. Reducing the complexity of the parameter.**
#
# > **Computational complexity -> Computationally it was faster when compared to models like svm, xgBoost, Adaboost.**
# # Summary and Conclusion
Image('../Images/Summary.png')
# **In the project the task assigned to me was to cluster the Grievance Description into UFDP (Un-fair and deceptive business practice) and Non – UFDP and then along with the other parameters provided classify bank’s performance as outstanding, satisfactory or deficient.**
#
# **I have taken the Grievance description and cleaned the description by the techniques of text mining. I have converted the Grievnace descripion to small letters,removed punctuation, removed stop words, removed words that occured commonly, removed words that occured rarely, tokenized the Grievnace descripion, Lemmatized the Grievnace descripion, calculated tf-idf for the Grievnace descripion. Converted the text data into vector representation using Doc2vec technique. In doc2vec i have used the ftca document and consumer compliance document and compared the Grievnace descripionwith the doc2vec vector values of ftca and consumer compliance handbook documents and with the help of Machine learning algorithm (KMeans clustering) clustered the Grievnace descripion as UFDP and Non UFDP. In my scenario the Grievnace descripion's vector value is closer to ftca document then it is classified as UFDP otherwise as Non UFDP.**
#
# **Further, replaced the Grievance description column with the cluster values in the main data. Considering 'LineOfBusiness','ResolutionComments', 'Disputed','DAYSDIFF', 'cluster values' columns for my mdoel building. I dummyfied these columns. Used Machine learning and deep learning algorithms on this data to predict the performance of the banks. I used 'Logistic Regression','Decision Tree','Random Forest','Naive Bayes (GNB)','Adaboost','Stochastic Gradient Decent' and 'Artifitial neural network' as my list of algorithms to predict the Bankgrade (outstanding/satisfactory/deficient) of the banks provided in my test data.**
#
# **Out of all the alogorithms Random Forest gave me the best accuracy of 68% on my test data.**
#
# **I would recommend the business to use my AI agent in the risk compliance department. This AI agent will save a lot of time, money and keeping other aspects in mind it will help the business grow rapidly.**
# ### What more could i have tried?
#
# **I wanted to try LSTM or Skip thoughts approach before the classification of the Grievance description into UFDP and Non UFDP by clustering algorithm.**
#
# **A detailed description of skip thoughts can be found in this link: https://medium.com/@sanyamagarwal/my-thoughts-on-skip-thoughts-a3e773605efa **
Image('../Images/Thankyou.jpeg')
| Model Building/Model_Building_Cluster_3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#torch-specific imports
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
#torchvision
import torchvision
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import numpy as np
import time
import os
import copy
device = torch.device('cpu')
dir_data = './data/yoga_poses'
mean = np.array([0.5, 0.5, 0.5])
std = np.array([0.25, 0.25, 0.25])
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(mean, std)
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize(mean, std)
]),
}
# +
image_datasets = {x: datasets.ImageFolder(os.path.join(dir_data, x),
data_transforms[x])
for x in ['train', 'val']}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x],
batch_size=5,shuffle=True, num_workers=0)
for x in ['train', 'val']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val']}
class_names = image_datasets['train'].classes
# -
print(class_names)
def imshow(inp, title):
inp = inp.numpy().transpose((1, 2, 0))
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
plt.title(title)
plt.show()
# +
# Get a batch of training data
inputs, classes = next(iter(dataloaders['train']))
# Make a grid from batch
out = torchvision.utils.make_grid(inputs)
# -
imshow(out, title=[class_names[x] for x in classes])
def train_model(model, criterion, optimizer, scheduler, num_epochs=25):
since = time.time()
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
optimizer.zero_grad()
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
if phase == 'train':
scheduler.step()
epoch_loss = running_loss / dataset_sizes[phase]
epoch_acc = running_corrects.double() / dataset_sizes[phase]
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, epoch_loss, epoch_acc))
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(
time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
return model
# +
#### Finetuning the convnet ####
# Load a pretrained model and reset final fully connected layer.
model = models.resnet18(pretrained=True)
num_ftrs = model.fc.in_features
# +
# Here the size of each output sample is set to 2.
# Alternatively, it can be generalized to nn.Linear(num_ftrs, len(class_names)).
model.fc = nn.Linear(num_ftrs, 2)
model = model.to(device)
criterion = nn.CrossEntropyLoss()
# -
# Observe that all parameters are being optimized
optimizer = optim.SGD(model.parameters(), lr=0.001)
step_lr_scheduler = lr_scheduler.StepLR(optimizer, step_size=7, gamma=0.1)
model = train_model(model, criterion, optimizer,
step_lr_scheduler, num_epochs=25)
# +
# Freeze all but last fully connected layer
# We need to set requires_grad == False to freeze the parameters so that the gradients are not computed in backward()
model_conv = torchvision.models.resnet18(pretrained=True)
for param in model_conv.parameters():
param.requires_grad = False
# +
# Parameters of newly constructed modules have requires_grad=True by default
num_ftrs = model_conv.fc.in_features
model_conv.fc = nn.Linear(num_ftrs, 2)
# +
model_conv = model_conv.to(device)
criterion = nn.CrossEntropyLoss()
# +
# Observe that only parameters of final layer are being optimized as
# opposed to before.
optimizer_conv = optim.SGD(model_conv.fc.parameters(), lr=0.001, momentum=0.9)
# +
# Decay LR by a factor of 0.1 every 7 epochs
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_conv, step_size=7, gamma=0.1)
model_conv = train_model(model_conv, criterion, optimizer_conv,
exp_lr_scheduler, num_epochs=25)
# +
# save the model locally so you can upload it to boon.ai
torch.save(model, 'yoga_model.pt')
#model = torch.load('yoga_model.pt')
| Yoga_Model_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: standardization_env
# language: python
# name: standardization_env
# ---
# # MARINe: Sea star and Katharina count conversion
#
# MARINe conducts long-term monitoring at sites along the coast of North America approximately annually. At sites where they are sufficiently abundant, ochre sea stars (Pisaster ochraceus) are monitored either within band transects or in irregularly sized plots. Other organisms, such as other sea stars or Katharina tunicata, are often also counted if they are present within the plots. There are generally 3 replicate plots per site. These plots are permanent, but chosen to target high densities of sea stars. They are intended to track changes in density and size frequency within a site, and the resulting data should not be used for comparisons between sites.
#
# Within each plot, sea stars are counted and measured. Measurements are taken from the center of the disk to the tip of the longest ray and are performed using calipers. These measurements are recorded to the nearest 5 mm for small sea stars (< 10 mm arm length) and to the nearest 10 mm for larger sea stars. Often, sizes have to be estimated due to the orientation or inaccessibility of the sea star. Early surveys binned sea stars into size classes, and these size classes shifted once through time. If there are many sea stars within a particular plot, only a subset of them may be measured.
#
# If Katharina are present in a sea star plot, they are counted, and up to 15 of them may be measured to the nearest 10 mm.
#
# At sites where sea stars are not abundant, a timed search protocol is used to document rarity.
#
# **Resources:**
# - DataONE link: https://data.piscoweb.org/metacatui/view/doi:10.6085/AA/marine_ltm.4.8
# +
## Import packages
import pandas as pd
import numpy as np
from datetime import datetime
import pyworms
from SPARQLWrapper import SPARQLWrapper, JSON
# -
# ## Load data
# +
## Load data
data = pd.read_csv('MARINe_LTM_counts_2020.csv')
print(data.shape)
data.head()
# +
## Site table
site = pd.read_csv('MARINe_site_table_2020.csv')
print(site.shape)
site.head()
# +
## Species table
species = pd.read_csv('MARINe_species_table_2020.csv')
print(species.shape)
species.head()
# -
# ## Conversion
#
# ### Occurrence
# Here, it seems like an **event** can be a survey, uniquely defined by the site code, year, and season. Actually, it looks like sometimes there have been multiple surveys within a given site, year, and season. So maybe the best way to go is site, min date, max date.
#
# ```python
# # Instances where multiple surveys occurred within a given site, year, and season
# out = data.groupby(['site_code', 'marine_common_year', 'season_name'])['min_survey_date'].nunique()
# out[out > 1]
#
# # Examine the dates for an example:
# data[(data['site_code'] == 'BOA') & (data['marine_common_year'] == 2003)].iloc[:, 0:20]
# ```
#
# An **occurrence** can be defined as an individual organism observed during an event.
#
# Measurements only pertain to occurrences, so I don't need a separate event file.
# +
## eventID
occ = pd.DataFrame({'eventID':data['site_code'] + '_' + data['min_survey_date'] + '_' + data['max_survey_date']})
print(occ.shape)
occ.head()
# +
## eventDate
occ['eventDate'] = data['min_survey_date'] + '/' + data['max_survey_date']
occ.head()
# +
## datasetName
occ['datasetName'] = 'MARINe LTM - sea star and katharina counts and sizes'
occ.head()
# +
## Merge with site table to get locality, county, stateProvince, countryCode, decimalLat, decimalLon
# Add site code to occ
occ['site_code'] = data['site_code']
# Define columns to merge from site table
site_cols = [
'site_code',
'marine_site_name',
'county',
'state_province',
'country',
'latitude',
'longitude',
]
# Define DwC terms for these columns after merge
dwc_cols = [
'eventID',
'eventDate',
'datasetName',
'locality',
'county',
'stateProvince',
'countryCode',
'decimalLatitude',
'decimalLongitude',
]
# Merge
occ = occ.merge(site[site_cols], how='left', on='site_code')
occ.drop(columns=['site_code'], inplace=True)
occ.columns = dwc_cols
print(occ.shape)
occ.head()
# -
# **Note** that ideally we would check county and state names against the [Getty Thesaurus of Geographic Names](http://www.getty.edu/research/tools/vocabularies/tgn/).
#
# Here, I've implemented a search using GTGN's SPARQL endpoint for all county names in the US. I've then filtered the output to obtain county names only in relevant states, and checked the county names in the MARINe dataset against that list. There's probably a way to write a more nuanced SPARQL query to check each individual name, but that seems like it will take more investment than I want to put in right now.
# +
## Search GTGN for all counties in the US
# Define sql strings
counties_query = """
select distinct * {
?place skos:inScheme tgn: ;
gvp:placeTypePreferred [gvp:prefLabelGVP [xl:literalForm ?type]];
gvp:placeType|(gvp:placeType/gvp:broaderGenericExtended) [rdfs:label "counties"@en];
gvp:broaderPartitiveExtended [rdfs:label "United States"@en];
gvp:prefLabelGVP [xl:literalForm ?name];
gvp:parentString ?parents}
"""
divisions_query = """
select distinct * {
?place skos:inScheme tgn: ;
gvp:placeTypePreferred [gvp:prefLabelGVP [xl:literalForm ?type]];
gvp:placeType|(gvp:placeType/gvp:broaderGenericExtended) [rdfs:label "national divisions"@en];
gvp:broaderPartitiveExtended [rdfs:label "United States"@en];
gvp:prefLabelGVP [xl:literalForm ?name];
gvp:parentString ?parents}
"""
# Set up query
sparql = SPARQLWrapper("http://vocab.getty.edu/sparql")
sparql.setReturnFormat(JSON)
sparql.setQuery(counties_query)
# Obtain county results
try:
counties_ret = sparql.query().convert()
except e:
print(e)
# Obtain national division results (Alaska has boroughs and census districts, not counties)
sparql.setQuery(divisions_query)
try:
div_ret = sparql.query().convert()
except e:
print(e)
# -
# **Ok, so the question is: How can I search for placeType is counties OR national divisions??**
# +
## Clean result
# Extract into data frame
county_df = pd.DataFrame(counties_ret['results']['bindings'])
county_df = county_df.applymap(lambda x: x['value'])
div_df = pd.DataFrame(div_ret['results']['bindings'])
div_df = div_df.applymap(lambda x: x['value'])
# Concatenate
county_df = pd.concat([county_df, div_df])
county_df.drop_duplicates(inplace=True)
# Unpack state, country etc. that each county is located in
county_df[['state', 'country', 'continent', 'planet', 'other']] = county_df['parents'].str.split(', ', expand=True)
# Filter
county_df = county_df[(county_df['country'] == 'United States') & (county_df['state'].isin(occ['stateProvince'].unique()))].copy()
county_df.head()
# +
## Check MARINe counties
for c in occ['county'].unique():
if c not in county_df['name'].unique():
print('County {} is not listed in GTGN. Double check name'.format(c))
# -
# All county names appear to be accurate.
# +
## Clean countryCode
occ['countryCode'] = occ['countryCode'].str.replace('United States', 'US')
occ.head()
# +
## coordinatUncertaintyInMeters
occ['coordinateUncertaintyInMeters'] = 350
occ.head()
# +
## Add minimumDepthInMeters, maximumDepthInMeters, samplingProtocol and samplingEffort
# Depth
occ['minimumDepthInMeters'] = 0
occ['maximumDepthInMeters'] = 0
# Protocol
occ['samplingProtocol'] = data['method_code']
occ['samplingProtocol'] = occ['samplingProtocol'].replace({
'BT25':'Band transect 2m x 5m',
'GSES':'General search entire site',
'IP':'Irregular plot',
'TS30':'Timed search 30 minutes'
})
# Effort
occ['samplingEffort'] = data['num_plots_sampled'].astype(str) + ' plot(s)'
occ.loc[occ['samplingProtocol'] == 'Band transect 2m x tm', 'samplingEffort'] = data['num_plots_sampled'].astype(str) + ' transect(s)'
occ.loc[occ['samplingProtocol'].isin(['General search entire site', 'Timed search 30 minutes']), 'samplingEffort'] = '1 site'
occ.head()
# -
# **Note** that for the protocols 'general search entire site' and 'timed search 30 minutes' the effort is always 1. **I've set the effort to 1 site (rather than n plot(s) or n transect(s) as I've done with the irregular plot and transect surveys).**
# +
## occurrenceID
occ['occurrenceID'] = data.groupby(['site_code', 'min_survey_date', 'max_survey_date'])['species_code'].cumcount()+1
occ['occurrenceID'] = occ['eventID'] + '_' + occ['occurrenceID'].astype(str)
occ.head()
# +
## scientificName
# Get species codes
occ['scientificName'] = data['species_code']
# Create scientificName column in species table
sp = species[['species_code', 'kingdom', 'phylum', 'class', 'order', 'family', 'genus', 'species']]
sp = sp.replace('NULL', np.nan, regex=True)
sp['scientificName'] = sp['species']
sp['scientificName'] = sp['scientificName'].combine_first(sp['family'])
sp['scientificName'] = sp['scientificName'].combine_first(sp['order'])
sp['scientificName'] = sp['scientificName'].combine_first(sp['class'])
sp['scientificName'] = sp['scientificName'].combine_first(sp['phylum'])
sp['scientificName'] = sp['scientificName'].combine_first(sp['kingdom'])
# Build dictionary mapping codes to names
sp_dict = dict(zip(sp['species_code'], sp['scientificName']))
# Replace codes with names in occ
occ['scientificName'] = occ['scientificName'].replace(sp_dict)
occ.head()
# +
## Get unique names
names = occ['scientificName'].unique()
names
# +
## Check names on WoRMS
results = pyworms.aphiaRecordsByMatchNames(names.tolist())
if len(results) == len(names): print('All names found.')
# Unpack results
worms_out = pd.json_normalize(results[0])
for i in range(1, len(results)):
norm = pd.json_normalize(results[i])
worms_out = pd.concat([worms_out, norm])
worms_out
# -
# **So, there's a lot that could be done here.** One thing that comes to mind is handling the output more elegantly, and/or handling what happens when a name isn't found. Right now, an empty list (i.e. `[]`) is returned. And if I query a name that's almost right (like Pisaster gigantea instead of Pisaster giganteus), it will match, but the match_type column will say `near_2` instead of `exact`. Finally, I'm not sure what the difference is between `scientificname` and `valid_name`.
# +
## Merge to add remaining taxonomy columns
# Indicate desired columns to merge
worms_cols = [
'AphiaID',
'scientificname',
'kingdom',
'phylum',
'class',
'order',
'family',
'genus',
'lsid'
]
# Give desired dwc column names
dwc_cols = occ.columns.to_list()
dwc_cols.extend([
'taxonID',
'kingdom',
'phylum',
'class',
'order',
'family',
'genus',
'scientificNameID',
])
# Merge
occ = occ.merge(worms_out[worms_cols], how='left', left_on='scientificName', right_on='scientificname')
occ = occ.drop(columns=['scientificname'])
occ.columns = dwc_cols
print(occ.shape)
occ.head()
# +
## Change taxonID to int
occ['taxonID'] = occ['taxonID'].astype(int)
occ.head()
# +
## Add other name-related columns
occ['nameAccordingTo'] = 'WoRMS'
occ['occurrenceStatus'] = 'present'
occ['basisOfRecord'] = 'HumanObservation'
occ.head()
# +
## Add count and size-related info
# Count
occ['individualCount'] = data['total']
# Change occurrenceStatus to absent if individualCount = 0
occ.loc[occ['individualCount'] == 0, 'occurrenceStatus'] = 'absent'
# Size
occ['organismQuantity'] = data['size_bin']
occ['organismQuantityType'] = 'Length of longest arm in millimeters'
occ.head()
# +
## Update organismQuantityType to reflect size measurement for Katharina
occ.loc[occ['scientificName'] == 'Katharina tunicata', 'organismQuantityType'] = 'Anterior to posterior length in millimeters'
# -
# Since I can, I'll put the size information directly in the occurrence table using `organismQuantity` and `organismQuantityType`. I can then also include it in a MoF file.
#
# Also, there are 3010 records where `individualCount` = 0 (i.e. the species of a particular size class wasn't observed). Rani says it's accurate to label these observations as 'absent'.
#
# Finally, **note** that there are 3455 records where `size_bin` is NaN. This is in addition to 1172 records where it is 'NM', or not measured. Rani clarified that NaN has been used if the species was not measured and is never measured during surveys. NM is used when some individuals of the species have been measured, just not this particular individual. NM should only appear when there's a count but no size.
#
# To check this:
# ```python
# 0 in occ.loc[occ['organismQuantity'] == 'NM', 'individualCount'].unique()
# ```
#
# For my purposes, it's fine to treat both of these values as NaN.
# +
## Clean organismQuantity
# To integer using nullable int data type in pandas
occ['organismQuantity'] = occ['organismQuantity'].replace('NM', np.nan)
converted = [int(s) if type(s) == str else np.nan for s in occ['organismQuantity']]
occ['organismQuantity'] = pd.Series(converted, dtype='Int32')
occ.head()
# -
# Learn more about nullable integers in Pandas: https://pandas.pydata.org/pandas-docs/stable/user_guide/integer_na.html
# +
## Save
occ.to_csv('MARINe_LTM_seastarkat_size_count_occurrence_20210825.csv', index=False)
# -
# ### MoF
# +
## Assemble measurementType, measurementValue
size = occ.loc[occ['organismQuantity'].isna() == False, ['eventID', 'occurrenceID', 'organismQuantityType', 'organismQuantity']].copy()
size.rename(columns={
'organismQuantityType':'measurementType',
'organismQuantity':'measurementValue'
}, inplace=True)
size['measurementType'] = 'Length maximum of biological entity specified elsewhere' # Closest term I can find on NVS
size.insert(3, 'measurementID', 'OBSMAXLX')
size.head()
# -
# **I'm not sure** of the best place to put the measurementID, or if, indeed, that's even the appropriate field. Here's the link to the P01 vocabulary term: http://vocab.nerc.ac.uk/collection/P01/current/OBSMAXLX/
# +
## Add measurementUnit, measurementMethod
# measurementUnit
size['measurementUnit'] = 'millimeters'
# measurementMethod
kath_occIDs = occ.loc[occ['scientificName'] == 'Katharina tunicata', 'occurrenceID']
size['measurementMethod'] = \
'measured with calipers from the center of the sea star to the tip of the longest arm and rounded to the nearest 5 mm if sea star is < 10 mm in size; rounded to the nearest 10 mm otherwise'
size.loc[size['occurrenceID'].isin(kath_occIDs), 'measurementMethod'] = \
'measured with calipers from the anterior to the posterior and rounded to the nearest 10 mm'
size.head()
# +
## Check for NaN values in string columns
size.isna().sum()
# +
## Save
size.to_csv('MARINe_LTM_seastarkat_size_count_mof_20210825.csv', index=False, na_rep='')
# -
# ## Questions
#
# None.
# +
## Number of years each MPA was surveyed
test = data[['site_code', 'min_survey_date', 'max_survey_date', 'marine_common_year']]
test = test.drop_duplicates()
test = test.merge(site[['site_code', 'mpa_name']], how='left', on='site_code')
test = test.dropna()
num_surveys_per_year = test.groupby(['site_code', 'marine_common_year'], as_index=False)['min_survey_date'].nunique() # 1-2, mode=2
num_sites_per_mpa = test.groupby('mpa_name', as_index=False)['site_code'].nunique() # 1-3, mode=1
test = test.groupby('mpa_name', as_index=False)['marine_common_year'].nunique()
test = test.sort_values('mpa_name')
test.to_csv('marine_sea_star_counts_years_per_mpa.csv', index=False)
# -
test['marine_common_year'].max()
# +
## Number of plots per survey
test = data[['site_code', 'min_survey_date', 'max_survey_date', 'marine_common_year', 'num_plots_sampled']]
test = test.drop_duplicates()
plots_per_survey = test.groupby(['site_code', 'min_survey_date'], as_index=False)['num_plots_sampled'].nunique()
plots_per_survey[plots_per_survey['num_plots_sampled'] > 1]
# -
# Hmmm... I find that surprising. **Should that be happening?**
# +
## Case 1
out = data[(data['site_code'] == 'BOB') & (data['min_survey_date'] == '2005-07-20')]
out[['site_code', 'min_survey_date', 'max_survey_date', 'target_assemblage', 'species_code', 'size_bin', 'total', 'num_plots_sampled', 'method_code']]
# +
## Case 2
out = data[(data['site_code'] == 'DMN') & (data['min_survey_date'] == '2010-05-29')]
out[['site_code', 'min_survey_date', 'max_survey_date', 'target_assemblage', 'species_code', 'size_bin', 'total', 'num_plots_sampled', 'method_code']]
# -
# So I guess the question is: **Why is the num_plots_sampled column sometimes different for Katharina counts than for sea stars? In the example above, does that mean Katharina was only looked for in 1 plot as opposed to 3? Or does it mean that Katharina were only found in 1 plot?**
#
# Either way, this only happens in a few cases, so will disregard for now and calculate the general number of plots per survey.
test['num_plots_sampled'].hist() # Typically 3
| MARINe/MARINe_LTM_count_conversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# %matplotlib inline
# The exercise uses depricated functions
import warnings
warnings.filterwarnings('ignore')
# # Marathon Finishing Time Example
#
# This notebook shows the use of seaborn with the data from a marathon
# ## Data exploration
url = 'https://raw.githubusercontent.com/jakevdp/marathon-data/master/marathon-data.csv'
data = pd.read_csv(url)
data.head()
data.dtypes
import datetime
def convert_time(s):
h,m,s = map(int,s.split(':'))
return datetime.timedelta(hours=h, minutes=m, seconds=s)
#Import data with a better format
data = pd.read_csv(url, converters={'split':convert_time, 'final':convert_time})
print(data.dtypes)
data.head()
# Add columns for analysis
data['split_sec'] = data['split'] / np.timedelta64(1, 's')
data['final_sec'] = data['final'] / np.timedelta64(1, 's')
data.head()
# ## Visual Exploration
# ### Jointplot
with sns.axes_style('white'):
g = sns.jointplot("split_sec", "final_sec", data, kind='hex')
g.ax_joint.plot(np.linspace(4000, 16000),
np.linspace(8000, 32000), ':k')
# ### Bar chart
data['split_frac'] = 1 - 2 * data['split_sec'] / data['final_sec']
sns.distplot(data['split_frac'], kde=False);
plt.axvline(0, color="k", linestyle="--");
# ### Pair Plots
g = sns.PairGrid(data, vars=['age', 'split_sec', 'final_sec', 'split_frac'],
hue='gender', palette='RdBu_r')
g.map(plt.scatter, alpha=0.8)
g.add_legend();
# ### KDE
sns.kdeplot(data.split_frac[data.gender=='M'], label='men', shade=True)
sns.kdeplot(data.split_frac[data.gender=='W'], label='women', shade=True)
plt.xlabel('split_frac');
# ### Factor Plots - Violin
sns.violinplot("gender", "split_frac", data=data,
palette=["lightblue", "lightpink"]);
data['age_dec'] = data.age.map(lambda age: 10 * (age // 10))
men = (data.gender == 'M')
women = (data.gender == 'W')
with sns.axes_style(style=None):
sns.violinplot("age_dec", "split_frac", hue="gender", data=data,
split=True, inner="quartile",
palette=["lightblue", "lightpink"]);
# ### Correlations
g = sns.lmplot('final_sec', 'split_frac', col='gender', data=data,
markers=".", scatter_kws=dict(color='c'))
g.map(plt.axhline, y=0.1, color="k", ls=":");
| Python Data Science Handbook/8. SEABORN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import os
import pandas as pd
import math
import re
# +
### process an inputted formula to determine the anions and cations
# formula inputted by user (don't know A, B, X at this point; can be in any format)
CCX3 = 'PbTiO3'
# collect all the elements
els = re.findall('[A-Z][a-z]?', CCX3)
# collect all the elements with their stoichiometric amounts if provided
el_num_pairs = [[el_num_pair[idx] for idx in range(len(el_num_pair)) if el_num_pair[idx] != ''][0]
for el_num_pair in re.findall('([A-Z][a-z]\d*)|([A-Z]\d*)', CCX3)]
# anion is element with 3 equivalents
X = [el_num_pair.replace('3', '') for el_num_pair in el_num_pairs if '3' in el_num_pair][0]
# cations are the other elements
cations = [el for el in els if el != X]
print('The input formula = %s' % CCX3)
print('The anion is determined to be %s' % X)
print('The cations are determined to be %s' % cations)
# +
### define some common oxidation states
# oxidation states if the element is the anion
X_ox_dict = {'N' : -3,
'O' : -2,
'S' : -2,
'Se' : -2,
'F' : -1,
'Cl' : -1,
'Br' : -1,
'I' : -1}
# common cation oxidation states
plus_one = ['H', 'Li', 'Na', 'K', 'Rb', 'Cs', 'Fr', 'Ag']
plus_two = ['Be', 'Mg', 'Ca', 'Sr', 'Ba', 'Ra']
plus_three = ['Sc', 'Y', 'La', 'Al', 'Ga', 'In',
'Pr', 'Nd', 'Pm', 'Sm', 'Eu', 'Gd', 'Tb',
'Dy', 'Ho', 'Er', 'Tm', 'Yb', 'Lu']
# +
### make dictionary of Shannon ionic radii
# starting with table available at v.web.umkc.edu/vanhornj/Radii.xls with Sn2+ added from 10.1039/c5sc04845a
# and organic cations from 10.1039/C4SC02211D
df = pd.read_csv('Shannon_Effective_Ionic_Radii.csv')
df = df.rename(columns = {'OX. State': 'ox',
'Coord. #': 'coord',
'Crystal Radius': 'rcryst',
'Ionic Radius': 'rion',
'Spin State' : 'spin'})
df['spin'] = [spin if spin in ['HS', 'LS'] else 'only_spin' for spin in df.spin.values]
def get_el(row):
ION = row['ION']
if ' ' in ION:
return ION.split(' ')[0]
elif '+' in ION:
return ION.split('+')[0]
elif '-' in ION:
return ION.split('-')[0]
df['el'] = df.apply(lambda row: get_el(row), axis = 1)
# get allowed oxidation states for each ion
el_to_ox = {}
for el in df.el.values:
el_to_ox[el] = list(set(df.ox.get((df['el'] == el)).tolist()))
# get ionic radii as function of oxidation state -> coordination number -> spin state
ionic_radii_dict = {}
for el in el_to_ox:
# list of Shannon oxidation states for each element
oxs = el_to_ox[el]
ox_to_coord = {}
for ox in oxs:
# list of coordination numbers for each (element, oxidation state)
coords = df.coord.get((df['el'] == el) & (df['ox'] == ox)).tolist()
ox_to_coord[ox] = coords
coord_to_spin = {}
for coord in ox_to_coord[ox]:
# list of spin states for each (element, oxidation state, coordination number)
spin = df.spin.get((df['el'] == el) & (df['ox'] == ox) & (df['coord'] == coord)).tolist()
coord_to_spin[coord] = spin
spin_to_rad = {}
for spin in coord_to_spin[coord]:
# list of radiis for each (element, oxidation state, coordination number)
rad = df.rion.get((df['el'] == el) & (df['ox'] == ox) & (df['coord'] == coord) & (df['spin'] == spin)).tolist()[0]
spin_to_rad[spin] = rad
coord_to_spin[coord] = spin_to_rad
ox_to_coord[ox] = coord_to_spin
ionic_radii_dict[el] = ox_to_coord
# assign spin state for transition metals (assumes that if an ion can be high-spin, it will be)
spin_els = ['Cr', 'Mn', 'Fe', 'Co', 'Ni', 'Cu']
starting_d = [4, 5, 6, 7, 8, 9]
d_dict = dict(zip(spin_els, starting_d))
for el in spin_els:
for ox in ionic_radii_dict[el].keys():
for coord in ionic_radii_dict[el][ox].keys():
if len(ionic_radii_dict[el][ox][coord].keys()) > 1:
num_d = d_dict[el] + 2 - ox
if num_d in [4, 5, 6, 7]:
ionic_radii_dict[el][ox][coord]['only_spin'] = ionic_radii_dict[el][ox][coord]['HS']
else:
ionic_radii_dict[el][ox][coord]['only_spin'] = ionic_radii_dict[el][ox][coord]['LS']
elif 'HS' in ionic_radii_dict[el][ox][coord].keys():
ionic_radii_dict[el][ox][coord]['only_spin'] = ionic_radii_dict[el][ox][coord]['HS']
elif 'LS' in ionic_radii_dict[el][ox][coord].keys():
ionic_radii_dict[el][ox][coord]['only_spin'] = ionic_radii_dict[el][ox][coord]['LS']
print('e.g., the ionic radius of Ti4+ at CN of 6 = %.3f Angstrom' % ionic_radii_dict['Ti'][4][6]['only_spin'])
# +
### determine the allowed oxidation states for each element in the compound
allowed_ox_dict = {}
for cation in cations:
# if cation is commonly 1+, make that the only allowed oxidation state
if cation in plus_one:
allowed_ox_dict[cation] = [1]
# if cation is commonly 2+, make that the only allowed oxidation state
elif cation in plus_two:
allowed_ox_dict[cation] = [2]
else:
# otherwise, use the oxidation states that have corresponding Shannon radii
allowed_ox_dict[cation] = [val for val in list(ionic_radii_dict[cation].keys()) if val > 0]
# assign the oxidation state of X based on the allowed anion oxidation states
allowed_ox_dict[X] = [X_ox_dict[X]]
for el in els:
print('The allowed oxidation state(s) of %s = %s' % (el, allowed_ox_dict[el]))
# -
### find all charge-balanced cation oxidation state combinations
ox1s = allowed_ox_dict[cations[0]]
ox2s = allowed_ox_dict[cations[1]]
oxX = allowed_ox_dict[X][0]
bal_combos = []
for ox1 in ox1s:
for ox2 in ox2s:
if ox1 + ox2 == -3*oxX:
bal_combos.append((ox1, ox2))
print('The charge balanced combinations are %s' % bal_combos)
# +
### choose the most likely charge-balanced combination
combos = bal_combos
# generate a dictionary of {cation : electronegativity} for help with assignment
chi_dict = {}
with open('electronegativities.csv') as f:
for line in f:
line = line.split(',')
if line[0] in cations:
chi_dict[line[0]] = float(line[1][:-1])
# if only one charge-balanced combination exists, use it
if len(combos) == 1:
ox_states = dict(zip(cations, combos[0]))
# if two combos exists and they are the reverse of one another
elif (len(combos) == 2) and (combos[0] == combos[1][::-1]):
# assign the minimum oxidation state to the more electronegative cation
min_ox = np.min(combos[0])
max_ox = np.max(combos[1])
epos_el = [el for el in cations if chi_dict[el] == np.min(list(chi_dict.values()))][0]
eneg_el = [el for el in cations if el != epos_el][0]
ox_states = {epos_el : max_ox,
eneg_el : min_ox}
else:
# if one of the cations is probably 3+, let it be 3+
if (cations[0] in plus_three) or (cations[1] in plus_three):
if X == 'O':
if (3,3) in combos:
combo = (3,3)
ox_states = dict(zip(ox_states, list(combo)))
# else compare electronegativities - if 0.9 < chi1/chi2 < 1.1, minimize the oxidation state diff
elif np.min(list(chi_dict.values())) > 0.9 * np.max(list(chi_dict.values())):
diffs = [abs(combo[0] - combo[1]) for combo in combos]
mindex = [idx for idx in range(len(diffs)) if diffs[idx] == np.min(diffs)]
if len(mindex) == 1:
mindex = mindex[0]
combo = combos[mindex]
ox_states = dict(zip(cations, combo))
else:
min_ox = np.min([combos[idx] for idx in mindex])
max_ox = np.max([combos[idx] for idx in mindex])
epos_el = [el for el in cations if chi_dict[el] == np.min(list(chi_dict.values()))][0]
eneg_el = [el for el in cations if el != epos_el][0]
ox_states = {epos_el : max_ox,
eneg_el : min_ox}
else:
diffs = [abs(combo[0] - combo[1]) for combo in combos]
maxdex = [idx for idx in range(len(diffs)) if diffs[idx] == np.max(diffs)]
if len(maxdex) == 1:
maxdex = maxdex[0]
combo = combos[maxdex]
else:
min_ox = np.min([combos[idx] for idx in maxdex])
max_ox = np.max([combos[idx] for idx in maxdex])
epos_el = [el for el in cations if chi_dict[el] == np.min(list(chi_dict.values()))][0]
eneg_el = [el for el in cations if el != epos_el][0]
ox_states = {epos_el : max_ox,
eneg_el : min_ox}
print('The electronegativities of %s = %.2f and %s = %.2f'
% (cations[0], chi_dict[cations[0]], cations[1], chi_dict[cations[1]]))
print('The assigned oxidation states are therefore %s = %.2f and %s = %.2f'
% (cations[0], ox_states[cations[0]], cations[1], ox_states[cations[1]]))
# +
### we know the oxidation states, but not which cation is A or B (yet)
### produce a dictionary of each cation as A or B
radii_dict = {}
for el in cations:
tmp_dict = {}
# get the oxidation state
ox = ox_states[el]
# get the coordination numbers for that cation by Shannon
coords = list(ionic_radii_dict[el][ox].keys())
# get the B CN as the one available nearest 6
B_coords = [abs(coord - 6) for coord in coords]
mindex = [idx for idx in range(len(B_coords)) if B_coords[idx] == np.min(B_coords)][0]
B_coord = coords[mindex]
# get the A CN as the one available nearest 12
A_coords = [abs(coord - 12) for coord in coords]
mindex = [idx for idx in range(len(A_coords)) if A_coords[idx] == np.min(A_coords)][0]
A_coord = coords[mindex]
# produce the equivalent B-site and A-site radii
B_rad = ionic_radii_dict[el][ox][B_coord]['only_spin']
A_rad = ionic_radii_dict[el][ox][A_coord]['only_spin']
tmp_dict['A_rad'] = A_rad
tmp_dict['B_rad'] = B_rad
radii_dict[el] = tmp_dict
for el in cations:
print('The radius of %s on the A site would be %.2f Angstrom' % (el, radii_dict[el]['A_rad']))
print('The radius of %s on the B site would be %.2f Angstrom' % (el, radii_dict[el]['B_rad']))
# +
### determine A and B, where A is the larger cation
el1 = list(radii_dict.keys())[0]
el2 = list(radii_dict.keys())[1]
if (radii_dict[el1]['A_rad'] > radii_dict[el2]['B_rad']) and (radii_dict[el1]['B_rad'] > radii_dict[el2]['A_rad']):
pred_A = el1
elif (radii_dict[el1]['A_rad'] < radii_dict[el2]['B_rad']) and (radii_dict[el1]['B_rad'] < radii_dict[el2]['A_rad']):
pred_A = el2
elif (radii_dict[el1]['A_rad'] > radii_dict[el2]['A_rad']) and (radii_dict[el1]['B_rad'] > radii_dict[el2]['B_rad']):
pred_A = el1
elif (radii_dict[el1]['A_rad'] < radii_dict[el2]['A_rad']) and (radii_dict[el1]['B_rad'] < radii_dict[el2]['B_rad']):
pred_A = el2
elif (radii_dict[el1]['B_rad'] > radii_dict[el2]['B_rad']):
pred_A = el1
elif (radii_dict[el1]['B_rad'] < radii_dict[el2]['B_rad']):
pred_A = el2
elif (radii_dict[el1]['A_rad'] > radii_dict[el2]['A_rad']):
pred_A = el1
elif (radii_dict[el1]['A_rad'] < radii_dict[el2]['A_rad']):
pred_A = el2
else:
# if the A and B radii are the same for both elements, choose the more oxidized element
if ox_dict[el1] < ox_dict[el2]:
pred_A = el1
else:
# if the elements have the same radii and oxidation state, choose at random
pred_A = el2
pred_B = [el for el in cations if el != pred_A][0]
print('%s is predicted to be A site with oxidation state = %i and radius = %.2f'
% (pred_A, ox_states[pred_A], radii_dict[pred_A]['A_rad']))
print('%s is predicted to be B site with oxidation state = %i and radius = %.2f'
% (pred_B, ox_states[pred_B], radii_dict[pred_B]['B_rad']))
# +
### make classification using tau
nA = ox_states[pred_A]
rA = radii_dict[pred_A]['A_rad']
rB = radii_dict[pred_B]['B_rad']
rX = ionic_radii_dict[X][X_ox_dict[X]][6]['only_spin']
tau = rX/rB - nA * (nA - (rA/rB)/np.log(rA/rB))
print('tau = %.2f which is %s 4.18, so %s is predicted %s'
% (tau, '<' if tau < 4.18 else '>', CCX3, 'perovskite' if tau < 4.18 else 'nonperovskite'))
| classify_CCX3_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Freyberg USG Model PEST setup example
# Herein, we will show users how to use pyEMU to setup a groundwater model for use in pest. Except using the Unstructured Grid (usg) version of MODFLOW. We will cover the following topics:
# - setup pilot points as parameters, including 1st-order tikhonov regularization
# - setup other model inputs as parameters
# - setup simulated water levels as observations
# - setup simulated water budget components as observations (or forecasts)
# - create a pest control file and adjust observation weights to balance the objective function
#
# Note that, in addition to `pyemu`, this notebook relies on `flopy`. `flopy` can be obtained (along with installation instructions) at https://github.com/modflowpy/flopy.
#
# %matplotlib inline
import os
import shutil
import platform
import numpy as np
import pandas as pd
from matplotlib.patches import Rectangle as rect
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore",
message="ModflowDis.sr is deprecated. use Modflow.sr")
from mpl_toolkits.axes_grid1 import make_axes_locatable
import matplotlib as mpl
newparams = {'legend.fontsize':10, 'axes.labelsize':10,
'xtick.labelsize':10, 'ytick.labelsize':10,
'font.family':'Univers 57 Condensed',
'pdf.fonttype':42}
plt.rcParams.update(newparams)
import pyemu
# ## Model background
# This example is based on the synthetic classroom model of Freyberg(1988). The model is a 2-dimensional MODFLOW model with 3 layers, 40 rows, and 20 columns. The model has 2 stress periods: an initial steady-state stress period used for calibration, and a 5-year transient stress period. The calibration period uses the recharge and well flux of Freyberg(1988); the last stress period use 25% less recharge and 25% more pumping to represent future conditions for a forecast period.
#
# This model has been modified using Gridgen to include a quadtree mesh at the location of the river.
#
# Freyberg, <NAME>. "AN EXERCISE IN GROUND‐WATER MODEL CALIBRATION AND PREDICTION." Groundwater 26.3 (1988): 350-360.
# +
#load the existing model and save it in a new dir and make sure it runs
import flopy
model_ws = os.path.join("freyberg_usg")
ml = flopy.modflow.Modflow.load("freyberg.usg.nam",model_ws=model_ws,verbose=False,version='mfusg',forgive=True,check=False)
ml.exe_name = "mfusg"
# ml.model_ws = "temp"
EXE_DIR = os.path.join("..","bin")
if "window" in platform.platform().lower():
EXE_DIR = os.path.join(EXE_DIR,"win")
elif "darwin" in platform.platform().lower():
EXE_DIR = os.path.join(EXE_DIR,"mac")
else:
EXE_DIR = os.path.join(EXE_DIR,"linux")
[shutil.copy2(os.path.join(EXE_DIR,f),os.path.join("temp",f)) for f in os.listdir(EXE_DIR)]
ml.external_path = "."
ml.change_model_ws('temp',reset_external=True)
ml.write_input()
def ref2array(fn):
f = open(fn)
lines = f.readlines()
vals = lines[0].split()
vals = [float(val) for val in vals]
return np.array(vals)
def array2ref(fn, vals):
f = open(fn, 'w')
line = ''.join(f'{val}\n' for val in vals)
f.write(line)
nlay = ml.nlay
for lay in np.arange(0,nlay):
for prop in ['hk', 'vka', 'ss']:
filename_lyr = f'{prop}_Layer_{lay+1}.ref'
if prop == 'vka':
filename_lyr = f'{prop}{lay+1}.ref'
ogref_lyr = ref2array(fn=os.path.join(ml.model_ws, filename_lyr))
array2ref(fn=os.path.join(ml.model_ws, filename_lyr),vals=ogref_lyr)
ml.run_model()
# -
def hdobj2data(hdsobj):
# convert usg hdsobj to array of shape (nper, nnodes)
hds = []
kstpkpers = hdsobj.get_kstpkper()
for kstpkper in kstpkpers:
data = hdsobj.get_data(kstpkper=kstpkper)
fdata = []
for lay in range(len(data)):
fdata += data[lay].tolist()
hds.append(fdata)
return np.array(hds)
node_df = pd.read_csv(os.path.join("Freyberg","misc","obs_nodes.dat"),delim_whitespace=True)
hdsobj = flopy.utils.HeadUFile(os.path.join(ml.model_ws,"freyberg.usg.hds"))
hds = hdobj2data(hdsobj)
nper,nnodes = hds.shape
(nper,nnodes)
# +
data = []
for i, dfrow in node_df.iterrows():
name, node = dfrow['name'], dfrow['node']
r = np.random.randn(nper) #add some random noise to the observations
for sp in range(nper):
hd = hds[sp,node-1]
rhd = r[sp] + hd #add some random noise to the observations
data.append([rhd,name,node,sp])
obs_df = pd.DataFrame(data,columns=['head','name','node','sp'])
obs_df.to_csv(os.path.join('obs.csv'),index=False)
obs_df
# -
# ### Use the GSF to make a Spatial Refrence structure
gsf = pyemu.gw_utils.GsfReader(os.path.join(model_ws,"freyberg.usg.gsf"))
gsf_df = gsf.get_node_data()
gsf_df["xy"] = gsf_df.apply(lambda i: (i['x'],i['y']),axis=1)
gsf_df.head()
# # Parameters
#
# ## pilot points
#
# Here we will import pilot point locations from a csv
#
pp_df = pd.read_csv(os.path.join("Freyberg","misc","pp_usg.csv"))
pp_df['xy'] = pp_df.apply(lambda i: (i['x'],i['y']),axis=1)
pp_df.head()
# ## Lets plot the model grid and pilot points to see what we're even working with
# +
usg_shp = os.path.join(model_ws,'misc','freyberg.usg.shp')
active_shp = os.path.join(model_ws,'misc','active_extent.shp')
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1, aspect='equal')
xoff, yoff = 619741.4, 3363170.9
rotation=0
proj4 = '+proj=utm +zone=14 +ellps=GRS80 +towgs84=0,0,0,0,0,0,0 +units=m +no_defs'
ml.modelgrid.set_coord_info(xoff, yoff, rotation, proj4=proj4)
mapview = flopy.plot.PlotMapView(model=ml,ax=ax)
mapview.plot_shapefile(usg_shp, edgecolor='k', facecolor='none')
mapview.plot_shapefile(active_shp, edgecolor='k', facecolor='g',alpha=.1)
ax.scatter(pp_df['x'],pp_df['y'])
# -
# ### setup pilot point locations
#
# first specify what pilot point names we want to use for each model layer (counting from 0). Here we will setup pilot points for ``hk``, ``sy`` and ``rech``. The ``rech`` pilot points will be used as a single multiplier array for all stress periods to account for potential spatial bias in recharge.
#
# ### setup sr dict (location for each node)
# for usg, we need to do some trickery to support the unstructured by layers concept
# this is just for array-based parameters, list-based pars are g2g because they have an index
# +
nlay = ml.nlay
sr_dict_by_layer = {}
for lay in range(nlay):
df_lay = gsf_df.loc[gsf_df['layer'] == lay+1]
df_lay.loc[:,"node"] = df_lay['node'] - df_lay['node'].min()
srd = {n:xy for n,xy in zip(df_lay.node.values,df_lay.xy.values)}
sr_dict_by_layer[lay+1] = srd
# -
# ### lets also set up zones for the pilot points in the model. For ease lets just use the active cells to represent each zone
# +
ibound = ml.bas6.ibound.array
zones = {}
n = 0
for lay in range(nlay):
nl = len(sr_dict_by_layer[lay+1])
zones[lay+1] = np.zeros(nl)
zones[lay+1] = np.array([ibound[n:n+nl]])
n += nl
zones
# -
v = pyemu.geostats.ExpVario(contribution=1.0,a=1000)
gs = pyemu.geostats.GeoStruct(variograms=v)
template_ws = 'template3'
pf = pyemu.utils.PstFrom('temp',template_ws, longnames=True,remove_existing=True,
zero_based=False,spatial_reference=gsf.get_node_coordinates(zero_based=True))
# +
# for lay in range(nlay):
# pf.add_parameters(os.path.join(f"hk_Layer_{lay+1}.ref"), par_type="pilotpoints",
# par_name_base=f"hk{lay+1}_pp", pp_space=pp_df.copy(),
# geostruct=gs, spatial_reference=sr_dict_by_layer[lay+1],
# upper_bound=10.0, lower_bound=0.1,zone_array=zones[lay+1],use_pp_zones=True)
# pf.add_parameters(os.path.join(f"vka{lay+1}.ref"), par_type="pilotpoints",
# par_name_base=f"vka{lay+1}_pp",pp_space=pp_df.copy(),
# geostruct=gs,spatial_reference=sr_dict_by_layer[lay+1],
# upper_bound=10.0,lower_bound=0.1,zone_array=zones[lay+1],use_pp_zones=True)
# pf.add_parameters(os.path.join(f"ss_Layer_{lay+1}.ref"), par_type="pilotpoints",
# par_name_base=f"ss{lay+1}_pp", pp_space=pp_df.copy(),
# geostruct=gs, spatial_reference=sr_dict_by_layer[lay+1],
# upper_bound=10.0, lower_bound=0.1,zone_array=zones[lay+1],use_pp_zones=True)
for lay in range(nlay):
pf.add_parameters(os.path.join(f"hk_Layer_{lay+1}.ref"), par_type="pilotpoints",
par_name_base=f"hk{lay+1}_pp", pp_space=pp_df.copy(),
geostruct=gs, spatial_reference=sr_dict_by_layer[lay+1],
upper_bound=10.0, lower_bound=0.1,zone_array=zones[lay+1])
pf.add_parameters(os.path.join(f"vka{lay+1}.ref"), par_type="pilotpoints",
par_name_base=f"vka{lay+1}_pp",pp_space=pp_df.copy(),
geostruct=gs,spatial_reference=sr_dict_by_layer[lay+1],
upper_bound=10.0,lower_bound=0.1,zone_array=zones[lay+1])
pf.add_parameters(os.path.join(f"ss_Layer_{lay+1}.ref"), par_type="pilotpoints",
par_name_base=f"ss{lay+1}_pp", pp_space=pp_df.copy(),
geostruct=gs, spatial_reference=sr_dict_by_layer[lay+1],
upper_bound=10.0, lower_bound=0.1,zone_array=zones[lay+1])
# -
pf.add_observations('obs.csv',index_cols=['name','sp'],use_cols=['obsval'],ofile_sep=',')
pf.mod_sys_cmds.append(f'mfusg freyberg.usg.nam')
# +
pf.add_py_function('usg_sim.py','hdobj2data()',is_pre_cmd='none')
pf.add_py_function('usg_sim.py','get_sim_hds(model_ws = ".")',is_pre_cmd=False)
pf.extra_py_imports.append("flopy")
pf.extra_py_imports.append("numpy as np")
pf.extra_py_imports.append("os")
pf.extra_py_imports.append("pandas as pd")
# -
pst = pf.build_pst('freyberg.usg.pst')
pst.control_data.noptmax = 0
pst.write(os.path.join(template_ws,"freyberg.usg.pst"),version=2)
pyemu.os_utils.run("pestpp-ies freyberg.usg.pst",cwd=template_ws,verbose=True)
| examples/modflow_to_pest_like_a_boss_but_its_Unstructured.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lista 01 - Introdução e Revisão Numpy
#
# [NumPy](http://numpy.org) é um pacote incrivelmente poderoso em Python, onipresente em qualquer projeto de ciência de dados. Possui forte integração com o [Pandas](http://pandas.pydata.org), outra ferramenta que iremos abordar na matéria. NumPy adiciona suporte para matrizes multidimensionais e funções matemáticas que permitem que você execute facilmente cálculos de álgebra linear. Este notebook será uma coleção de exemplos de álgebra linear computados usando NumPy.
#
# ## Numpy
#
# Para fazer uso de Numpy precisamos importar a biblioteca
# +
# -*- coding: utf8
import numpy as np
# -
# Quando pensamos no lado prático de ciência de dados, um aspecto chave que ajuda na implementação de novos algoritmos é a vetorização. De forma simples, vetorização consiste do uso de tipos como **escalar**, **vetor** e **matriz** para realizar uma computação mais eficaz (em tempo de execução).
#
# Uma matriz é uma coleção de valores, normalmente representada por uma grade 𝑚 × 𝑛, onde 𝑚 é o número de linhas e 𝑛 é o número de colunas. Os comprimentos das arestas 𝑚 e 𝑛 não precisam ser necessariamente diferentes. Se tivermos 𝑚 = 𝑛, chamamos isso de matriz quadrada. Um caso particularmente interessante de uma matriz é quando 𝑚 = 1 ou 𝑛 = 1. Nesse caso, temos um caso especial de uma matriz que chamamos de vetor. Embora haja um objeto de matriz em NumPy, faremos tudo usando matrizes NumPy porque elas podem ter dimensões maiores que 2.
#
# 1. **Escalar:** Um vetor de zero dimensões
1
# 2. **Vetor:** Representa uma dimensão
# Abaixo vamos criar um vetor simples. Inicialmente, vamos criar uma lista.
data_list = [3.5, 5, 2, 8, 4.2]
# Observe o tipo da mesma.
type(data_list)
# Embora vetores e listas sejam parecidos, vetores Numpy são otimizados para operações de Álgebra Linear. Ciência de Dados faz bastante uso de tais operações, sendo este um dos motivos da dependência em Numpy.
#
# Abaixo criamos um vetor.
data = np.array(data_list)
print(data)
print(type(data))
# Observe como podemos somar o mesmo com um número. Não é possível fazer tal operação com listas.
data + 7
# 3. **Matrizes:** Representam duas dimensões.
X = np.array([[2, 4],
[1, 3]])
X
# Podemos indexar as matrizes e os vetores.
data[0]
X[0, 1] # aqui é primeira linha, segunda coluna
# Podemos também criar vetores/matrizes de números aleatórios
X = np.random.randn(4, 3) # Gera números aleatórios de uma normal
print(X)
# ### Indexando
#
# Pegando a primeira linha
X[0] # observe que 0 é a linha 1, compare com o X[0, 1] de antes.
X[1] # segunda
X[2] # terceira
# Observe como todos os tipos retornados são `array`. Array é o nome genérico de Numpy para vetores e matrizes.
# `X[:, c]` pega uma coluna
X[:, 0]
X[:, 1]
# `X[um_vetor]` pega as linhas da matriz. `X[:, um_vetor]` pega as colunas
X[[0, 0, 1]] # observe que pego a primeira linha, indexada por 0, duas vezes
# Abaixo pego a segunda a primeira coluna
X[:, [1, 0]]
# ### Indexação Booleana
#
# `X[vetor_booleano]` retorna as linhas (ou colunas quando X[:, vetor_booleano]) onde o vetor é true
X[[True, False, True, False]]
X[:, [False, True, True]]
# ### Reshape, Flatten e Ravel
#
# Todo vetor ou matriz pode ser redimensionado. Observe como uma matriz abaixo de 9x8=72 elementos. Podemos redimensionar os mesmos para outros arrays de tamanho 72.
X = np.random.randn(9, 8)
# Criando uma matriz de 18x4.
X.reshape((18, 4))
# Ou um vetor de 72
X.reshape(72)
# A chamada flatten e ravel faz a mesma coisa, criam uma visão de uma dimensão da matriz.
X.flatten()
X.ravel()
# As funções incorporadas ao NumPy podem ser facilmente chamadas em matrizes. A maioria das funções são aplicadas a um elemento de array (como a multiplicação escalar). Por exemplo, se chamarmos `log()` em um array, o logaritmo será obtido de cada elemento.
np.log(data)
# Mean tira a média
np.mean(data)
# Algumas funções podem ser chamadas direto no vetor, nem todas serão assim. O importante é ler a [documentação](http://numpy.org) e aprender. Com um pouco de prática você vai se acostumando.
data.mean()
# Abaixo temos a mediana,
np.median(data) # por exemplo, não existe data.median(). Faz sentido? Não. Mas é assim.
# Em matrizes as funções operam em todos os elemntos.
np.median(X)
X.mean()
np.log(X + 10)
# Porém, caso você queira a media de linhas ou colunas use `axis`. Antes, vamos ver o tamanho do vetor.
X.shape
np.mean(X, axis=0) # média das colunas. como temos 8 colunas, temos 8 elementos.
np.mean(X, axis=0).shape
np.mean(X, axis=1) # média das linhas
np.mean(X, axis=1).shape
# Lembre-se que eixo 0 é coluna. Eixo 1 é linas.
# ### Multiplicação de Matrizes
# Para transpor uma matriz fazemos uso de .T
X.shape
X.T.shape
X.T
# Para multiplicar matrizes, do ponto de visto de multiplicação matricial como definido na álgebra linear, fazemos uso de `@`.
X @ X.T
# O uso de `*` realiza uma operação ponto a ponto
X * X
# Observe a diferença de tamanhos
(X * X).shape
(X @ X.T).shape
# **Pense:** Para o nosso `X` de tamanho `(9, 8)`, qual o motivo de `X * X.T` não funcionar? Qual o motivo de `X @ X` não funcionar?
# ## Correção Automática
#
# Nossa correção automática depende das funções abaixo. Tais funções comparam valores que serão computados pelo seu código com uma saída esperada. Normalmente, vocês não fazer uso de tais funções em notebooks como este. Porém, elas são chave em ambientes de testes automáticos (fora do nosso escopo).
#
# Observe como algumas funções comparam valores e outras comparam vetores. Além do mais, temos funções para comparar dentro de algumas casas decimais.
# +
from numpy.testing import assert_almost_equal
from numpy.testing import assert_equal
from numpy.testing import assert_array_almost_equal
from numpy.testing import assert_array_equal
# +
# caso você mude um dos valores vamos receber um erro!
assert_array_equal(2, 2)
# caso você mude um dos valores vamos receber um erro!
assert_array_equal([1, 2], [1, 2])
# caso você mude um dos valores vamos receber um erro!
assert_almost_equal(3.1415, 3.14, 1)
# -
# Caso você mude um dos valores abaixo vamos receber um erro! Como o abaixo.
#
# ```
# -----------------------------------------------------------------------
# AssertionError Traceback (most recent call last)
# <ipython-input-10-396672d880f2> in <module>
# ----> 1 assert_equal(2, 3) # caso você mude um dos valores vamos receber um erro!
#
# ~/miniconda3/lib/python3.7/site-packages/numpy/testing/_private/utils.py in assert_equal(actual, desired, err_msg, verbose)
# 413 # Explicitly use __eq__ for comparison, gh-2552
# 414 if not (desired == actual):
# --> 415 raise AssertionError(msg)
# 416
# 417 except (DeprecationWarning, FutureWarning) as e:
#
# AssertionError:
# Items are not equal:
# ACTUAL: 2
# DESIRED: 3
# ```
# É essencial que todo seu código execute sem erros! Portanto, antes de submeter clique em `Kernel` no menu acima. Depois clique em `Restart & Execute All.`
#
# **Garanta que o notebook executa até o fim!** Isto é, sem erros como o acima.
# ## Funções em Python
# Para criar uma função em Python fazemos uso da palavra-chave:
# ```python
# def
# ```
#
# Todos nossos exercícios farão uso de funções. **Mantenha a assinatura das funções exatamente como requisitado, a correção automática depende disso.** Abaixo, temos um exempo de uma função que imprime algo na tela!
def print_something(txt):
print(f'Você passou o argumento: {txt}')
print_something('DCC 212')
# Podemos também dizer o tipo do argumento, porém faremos pouco uso disto em ICD.
def print_something(txt: str):
print(f'Você passou o argumento: {txt}')
print_something('DCC 212')
# Abaixo temos uma função que soma, a soma, dois vetores
def sum_of_sum_vectors(array_1, array_2):
return (array_1 + array_2).sum()
x = np.array([1, 2])
y = np.array([1, 2])
sum_of_sum_vectors(x, y)
# Abaixo temos um teste, tais testes vão avaliar o seu código. Nem todos estão aqui no notebook!
assert_equal(6, sum_of_sum_vectors(x, y))
# ## Exercício 01
#
# Inicialmente, crie uma função que recebe duas listas de numéros, converte as duas para um vetor numpy usando `np.array` e retorna o produto interno das duas listas.
#
# __Dicas:__
# 1. Tente fazer um código sem nenhum **for**! Ou seja, numpy permite operações em vetores e matrizes, onde: `np.array([1, 2]) + np.array([2, 2]) = np.array([3, 4])`.
#
# __Funções:__
# 1. `np.sum(array)` soma os elementos do array. `array.sum()` tem o mesmo efeito!
def inner(array_1, array_2):
# Seu código aqui!
# Apague o return None abaixo e mude para seu retorno
return None
x1 = np.array([2, 4, 8])
x2 = np.array([10, 100, 1000])
assert_equal(20 + 400 + 8000, inner(x1, x2))
# ## Exercício 02
#
# Implemente uma função utilizando numpy que recebe duas matrizes, multiplica as duas e retorne o valor médio das células da multiplicação. Por exemplo, ao multiplicar:
#
# ```
# [1 2]
# [3 4]
#
# com
#
# [2 1]
# [1 2]
#
# temos
#
# [4 5 ]
# [10 11]
#
# onde a média de [4, 5, 10, 11] é
#
# 7.5, sua resposta final!
# ```
#
#
# __Dicas:__
# 1. Use o operador @ para multiplicar matrizes!
def medmult(X_1, X_2):
# Seu código aqui!
# Apague o return None abaixo e mude para seu retorno
return None
X = np.array([1, 2, 3, 4]).reshape(2, 2)
Y = np.array([2, 1, 1, 2]).reshape(2, 2)
assert_equal(7.5, medmult(X, Y))
| listas/l1/.ipynb_checkpoints/dcc212l1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="5PYYaGcToEcq"
from keras.preprocessing import image
from keras.models import Model, Sequential
from keras.layers import Activation, Dense, GlobalAveragePooling2D, BatchNormalization, Dropout, Conv2D, Conv2DTranspose, AveragePooling2D, MaxPooling2D, UpSampling2D, Input, Reshape
from keras import backend as K
from keras.optimizers import Nadam, Adam, SGD
from keras.metrics import categorical_accuracy, binary_accuracy
#from keras_contrib.losses import jaccard
import tensorflow as tf
import numpy as np
import pandas as pd
import glob
import PIL
from PIL import Image
import matplotlib.pyplot as plt
# %matplotlib inline
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="B7rHGhE8uCwb"
def jaccard_distance(y_true, y_pred, smooth=100):
intersection = K.sum(K.abs(y_true * y_pred), axis=-1)
sum_ = K.sum(K.square(y_true), axis = -1) + K.sum(K.square(y_pred), axis=-1)
jac = (intersection + smooth) / (sum_ - intersection + smooth)
return (1 - jac)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="NboSwhqOvlTw"
def iou(y_true, y_pred, smooth = 100):
intersection = K.sum(K.abs(y_true * y_pred), axis=-1)
sum_ = K.sum(K.square(y_true), axis = -1) + K.sum(K.square(y_pred), axis=-1)
jac = (intersection + smooth) / (sum_ - intersection + smooth)
return jac
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="umQIcnQM-U8y"
# To read the images in numerical order
import re
numbers = re.compile(r'(\d+)')
def numericalSort(value):
parts = numbers.split(value)
parts[1::2] = map(int, parts[1::2])
return parts
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="JqPblx6quzaP"
# Initializing all the images into 4d arrays.
filelist_trainx = sorted(glob.glob('trainx/*.jpg'), key=numericalSort)
#filelist_trainx.sort()
X_train = np.array([np.array(Image.open(fname)) for fname in filelist_trainx])
filelist_trainy = sorted(glob.glob('trainy/*.jpg'), key=numericalSort)
#filelist_trainy.sort()
Y_train = np.array([np.array(Image.open(fname)) for fname in filelist_trainy])
filelist_testx = sorted(glob.glob('testx/*.jpg'), key=numericalSort)
#filelist_testx.sort()
X_test = np.array([np.array(Image.open(fname)) for fname in filelist_testx])
filelist_testy = sorted(glob.glob('testy/*.jpg'), key=numericalSort)
#filelist_testy.sort()
Y_test = np.array([np.array(Image.open(fname)) for fname in filelist_testy])
filelist_valx = sorted(glob.glob('validationx/*.jpg'), key=numericalSort)
#filelist_valx.sort()
X_val = np.array([np.array(Image.open(fname)) for fname in filelist_valx])
filelist_valy = sorted(glob.glob('validationy/*.jpg'), key=numericalSort)
#filelist_valy.sort()
Y_val = np.array([np.array(Image.open(fname)) for fname in filelist_valy])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 393} colab_type="code" executionInfo={"elapsed": 1414, "status": "ok", "timestamp": 1526788841790, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-AhklfYPJ2yU/AAAAAAAAAAI/AAAAAAAAMLU/1G-c09ybTmY/s50-c-k-no/photo.jpg", "userId": "117392830770812166897"}, "user_tz": -330} id="wfRNBcdtu3Lt" outputId="2fb029f1-c983-4242-b696-16a0c7d23541"
index = 100
plt.imshow(Y_val[index])
print ("y = " + str(np.squeeze(Y_val[:, index])))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="MQk0x9hlu5xL"
def UnPooling2x2ZeroFilled(x):
out = tf.concat([x, tf.zeros_like(x)], 3)
out = tf.concat([out, tf.zeros_like(out)], 2)
sh = x.get_shape().as_list()
if None not in sh[1:]:
out_size = [-1, sh[1] * 2, sh[2] * 2, sh[3]]
return tf.reshape(out, out_size)
else:
shv = tf.shape(x)
ret = tf.reshape(out, tf.stack([-1, shv[1] * 2, shv[2] * 2, sh[3]]))
return ret
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="gnSdLXT60fex"
# Pipe Line
(x_train, y_train), (x_test, y_test), (x_val, y_val) = (X_train, Y_train), (X_test, Y_test), (X_val, Y_val)
def model_seg():
# Convolution Layers (BatchNorm after non-linear activation)
img_input = Input(shape= (192, 256, 3))
x = Conv2D(16, (3, 3), padding='same', name='conv1')(img_input)
x = BatchNormalization(name='bn1')(x)
x = Activation('relu')(x)
x = Conv2D(32, (3, 3), padding='same', name='conv2')(x)
x = BatchNormalization(name='bn2')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(64, (3, 3), padding='same', name='conv3')(x)
x = BatchNormalization(name='bn3')(x)
x = Activation('relu')(x)
x = Conv2D(64, (3, 3), padding='same', name='conv4')(x)
x = BatchNormalization(name='bn4')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(128, (3, 3), padding='same', name='conv5')(x)
x = BatchNormalization(name='bn5')(x)
x = Activation('relu')(x)
x = Conv2D(128, (4, 4), padding='same', name='conv6')(x)
x = BatchNormalization(name='bn6')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(256, (3, 3), padding='same', name='conv7')(x)
x = BatchNormalization(name='bn7')(x)
x = Dropout(0.5)(x)
x = Activation('relu')(x)
x = Conv2D(256, (3, 3), padding='same', name='conv8')(x)
x = BatchNormalization(name='bn8')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(512, (3, 3), padding='same', name='conv9')(x)
x = BatchNormalization(name='bn9')(x)
x = Activation('relu')(x)
x = Dense(1024, activation = 'relu', name='fc1')(x)
x = Dense(1024, activation = 'relu', name='fc2')(x)
# Deconvolution Layers (BatchNorm after non-linear activation)
x = Conv2DTranspose(256, (3, 3), padding='same', name='deconv1')(x)
x = BatchNormalization(name='bn19')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(256, (3, 3), padding='same', name='deconv2')(x)
x = BatchNormalization(name='bn12')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(128, (3, 3), padding='same', name='deconv3')(x)
x = BatchNormalization(name='bn13')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(128, (4, 4), padding='same', name='deconv4')(x)
x = BatchNormalization(name='bn14')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(128, (3, 3), padding='same', name='deconv5')(x)
x = BatchNormalization(name='bn15')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(64, (3, 3), padding='same', name='deconv6')(x)
x = BatchNormalization(name='bn16')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(32, (3, 3), padding='same', name='deconv7')(x)
x = BatchNormalization(name='bn20')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(16, (3, 3), padding='same', name='deconv8')(x)
x = BatchNormalization(name='bn17')(x)
x = Dropout(0.5)(x)
x = Activation('relu')(x)
x = Conv2DTranspose(1, (3, 3), padding='same', name='deconv9')(x)
x = BatchNormalization(name='bn18')(x)
x = Activation('sigmoid')(x)
pred = Reshape((192,256))(x)
model = Model(inputs=img_input, outputs=pred)
model.compile(optimizer= Adam(lr = 0.003), loss= [jaccard_distance], metrics=[iou])
hist = model.fit(x_train, y_train, epochs= 300, batch_size= 16,validation_data=(x_test, y_test), verbose=1)
model.save("model.h5")
accuracy = model.evaluate(x=x_test,y=y_test,batch_size=16)
print("Accuracy: ",accuracy[1])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 7167} colab_type="code" executionInfo={"elapsed": 14978949, "status": "ok", "timestamp": 1526771788192, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-AhklfYPJ2yU/AAAAAAAAAAI/AAAAAAAAMLU/1G-c09ybTmY/s50-c-k-no/photo.jpg", "userId": "117392830770812166897"}, "user_tz": -330} id="p6A3nXVqejtK" outputId="fa12556c-eb27-4aa8-b2d9-3689d4d4964c"
model_seg()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 70} colab_type="code" executionInfo={"elapsed": 9438, "status": "ok", "timestamp": 1526798149909, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-AhklfYPJ2yU/AAAAAAAAAAI/AAAAAAAAMLU/1G-c09ybTmY/s50-c-k-no/photo.jpg", "userId": "117392830770812166897"}, "user_tz": -330} id="s3ZCJatJekS8" outputId="96330358-cf48-48b9-8e9d-b28ed08d927f"
(x_train, y_train), (x_test, y_test), (x_val, y_val) = (X_train, Y_train), (X_test, Y_test), (X_val, Y_val)
# Convolution Layers (BatchNorm after non-linear activation)
img_input = Input(shape= (192, 256, 3))
x = Conv2D(16, (3, 3), padding='same', name='conv1')(img_input)
x = BatchNormalization(name='bn1')(x)
x = Activation('relu')(x)
x = Conv2D(32, (3, 3), padding='same', name='conv2')(x)
x = BatchNormalization(name='bn2')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(64, (3, 3), padding='same', name='conv3')(x)
x = BatchNormalization(name='bn3')(x)
x = Activation('relu')(x)
x = Conv2D(64, (3, 3), padding='same', name='conv4')(x)
x = BatchNormalization(name='bn4')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(128, (3, 3), padding='same', name='conv5')(x)
x = BatchNormalization(name='bn5')(x)
x = Activation('relu')(x)
x = Conv2D(128, (4, 4), padding='same', name='conv6')(x)
x = BatchNormalization(name='bn6')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(256, (3, 3), padding='same', name='conv7')(x)
x = BatchNormalization(name='bn7')(x)
x = Dropout(0.5)(x)
x = Activation('relu')(x)
x = Conv2D(256, (3, 3), padding='same', name='conv8')(x)
x = BatchNormalization(name='bn8')(x)
x = Activation('relu')(x)
x = MaxPooling2D()(x)
x = Conv2D(512, (3, 3), padding='same', name='conv9')(x)
x = BatchNormalization(name='bn9')(x)
x = Activation('relu')(x)
x = Dense(1024, activation = 'relu', name='fc1')(x)
x = Dense(1024, activation = 'relu', name='fc2')(x)
# Deconvolution Layers (BatchNorm after non-linear activation)
x = Conv2DTranspose(256, (3, 3), padding='same', name='deconv1')(x)
x = BatchNormalization(name='bn19')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(256, (3, 3), padding='same', name='deconv2')(x)
x = BatchNormalization(name='bn12')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(128, (3, 3), padding='same', name='deconv3')(x)
x = BatchNormalization(name='bn13')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(128, (4, 4), padding='same', name='deconv4')(x)
x = BatchNormalization(name='bn14')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(128, (3, 3), padding='same', name='deconv5')(x)
x = BatchNormalization(name='bn15')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(64, (3, 3), padding='same', name='deconv6')(x)
x = BatchNormalization(name='bn16')(x)
x = Activation('relu')(x)
x = Conv2DTranspose(32, (3, 3), padding='same', name='deconv7')(x)
x = BatchNormalization(name='bn20')(x)
x = Activation('relu')(x)
x = UpSampling2D()(x)
x = Conv2DTranspose(16, (3, 3), padding='same', name='deconv8')(x)
x = BatchNormalization(name='bn17')(x)
x = Dropout(0.5)(x)
x = Activation('relu')(x)
x = Conv2DTranspose(1, (3, 3), padding='same', name='deconv9')(x)
x = BatchNormalization(name='bn18')(x)
x = Activation('sigmoid')(x)
pred = Reshape((192,256))(x)
model = Model(inputs=img_input, outputs=pred)
model.compile(optimizer= Adam(lr = 0.003), loss= [jaccard_distance], metrics=[iou])
model.load_weights("model.h5")
predictions_valid = model.predict(x_val, batch_size=16, verbose=1)
accuracy = model.evaluate(x=x_val,y=y_val,batch_size=16)
print("Accuracy: ",accuracy[1])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="3iO1w1gdcNXR"
index = 45
predict_input = x_val[index]
ground_truth = y_val[index]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="D1F3mjdab6nP"
predictions =model.predict(predict_input.reshape(1,192,256,3), batch_size=1)
prediction = predictions.reshape(192, 256)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 561} colab_type="code" executionInfo={"elapsed": 1307, "status": "ok", "timestamp": 1526801717813, "user": {"displayName": "<NAME>", "photoUrl": "//lh4.googleusercontent.com/-AhklfYPJ2yU/AAAAAAAAAAI/AAAAAAAAMLU/1G-c09ybTmY/s50-c-k-no/photo.jpg", "userId": "117392830770812166897"}, "user_tz": -330} id="AuEYLNmoSlYc" outputId="f55716cb-0b99-4012-b6cf-4c9c7ae3f85e"
#index = 45
plt.figure()
plt.imshow(prediction)
plt.title('Predicted')
plt.figure()
plt.imshow(ground_truth)
plt.title('Ground Turth')
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="6EFLQ9oOzKua"
| melanoma_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulation of congestion in a warehouse
#
# ### Libraries:
import numpy as np
import pandas as pd
# ## Extracting the center of distribution(warehouse) information:
# 
ots = pd.read_csv("data/ot.csv",sep=',')
ots=ots.sort_values('Order')
xlsx_file = "data/layout.xlsx"
layout = pd.read_excel(xlsx_file, sheet_name="layout")
adj=pd.read_excel(xlsx_file, sheet_name="adyacencia")
num_of_orders=len(ots['Order'].unique())
aisles=layout['aisle'].unique()
ots
layout
adj
aisles
num_of_orders
# ## Pseudocode:
# 
# ## Some Considerations:
# * Equal characteristics to all items
# * Narrow aisles
# * constant velocity for all pickers
# * constant picking time for all pickers
picking_time=10 #[s]
velocity=4 #[m/s]
# ## Initializing some variables:
# +
list1=[]
for x in range(num_of_orders):
obj=list(ots.loc[ots["Order"]==x+1]["Cod.Prod"])
list1.append(obj)
orders=np.array(list1)
l=[]
for aisle in aisles:
l.append([aisle,False, 0])
aisle_bool=pd.DataFrame(l,columns=["aisle","ocuppied","picker"])
# -
orders
aisle_bool
# ## The Code:
import code
help(code.assignment_of_orders)
help(code.next_move)
help(code.sort)
help(code.route_exist)
help(code.routing)
help(code.product_in_aisle)
help(code.ocuppied)
help(code.time)
code.time(orders,1,ots,layout,adj)
code.time(orders,2,ots,layout,adj)
code.time(orders,20,ots,layout,adj)
| Final_presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
# -
# ## Internal stability example 4
#
# ### How to use this notebook?
# Try to change the dynamic matrix $A$ of the stable linear system below in order to obtain a system with two divergent modes and then change the initial conditions in order to hide the divergent behaviour.
#
# $$
# \dot{x} = \underbrace{\begin{bmatrix}0&1\\-2&-2\end{bmatrix}}_{A}x
# $$
#
# Try to answer:
# - Is it possible to achieve this? If yes, in which particular case?
# +
# %matplotlib inline
import control as control
import numpy
import sympy as sym
from IPython.display import display, Markdown
import ipywidgets as widgets
import matplotlib.pyplot as plt
#matrixWidget is a matrix looking widget built with a VBox of HBox(es) that returns a numPy array as value !
class matrixWidget(widgets.VBox):
def updateM(self,change):
for irow in range(0,self.n):
for icol in range(0,self.m):
self.M_[irow,icol] = self.children[irow].children[icol].value
#print(self.M_[irow,icol])
self.value = self.M_
def dummychangecallback(self,change):
pass
def __init__(self,n,m):
self.n = n
self.m = m
self.M_ = numpy.matrix(numpy.zeros((self.n,self.m)))
self.value = self.M_
widgets.VBox.__init__(self,
children = [
widgets.HBox(children =
[widgets.FloatText(value=0.0, layout=widgets.Layout(width='90px')) for i in range(m)]
)
for j in range(n)
])
#fill in widgets and tell interact to call updateM each time a children changes value
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
self.children[irow].children[icol].observe(self.updateM, names='value')
#value = Unicode('<EMAIL>', help="The email value.").tag(sync=True)
self.observe(self.updateM, names='value', type= 'All')
def setM(self, newM):
#disable callbacks, change values, and reenable
self.unobserve(self.updateM, names='value', type= 'All')
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].unobserve(self.updateM, names='value')
self.M_ = newM
self.value = self.M_
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].observe(self.updateM, names='value')
self.observe(self.updateM, names='value', type= 'All')
#self.children[irow].children[icol].observe(self.updateM, names='value')
#overlaod class for state space systems that DO NOT remove "useless" states (what "professor" of automatic control would do this?)
class sss(control.StateSpace):
def __init__(self,*args):
#call base class init constructor
control.StateSpace.__init__(self,*args)
#disable function below in base class
def _remove_useless_states(self):
pass
# +
# Preparatory cell
A = numpy.matrix([[0.,1.],[-2.,-2.]])
X0 = numpy.matrix([[1.],[0.]])
Aw = matrixWidget(2,2)
Aw.setM(A)
X0w = matrixWidget(2,1)
X0w.setM(X0)
# +
# Misc
#create dummy widget
DW = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
#create button widget
START = widgets.Button(
description='Test',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Test',
icon='check'
)
def on_start_button_clicked(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW.value> 0 :
DW.value = -1
else:
DW.value = 1
pass
START.on_click(on_start_button_clicked)
# +
# Main cell
def main_callback(A, X0, DW):
sols = numpy.linalg.eig(A)
sys = sss(A,[[0],[1]],[1,0],0)
pole = control.pole(sys)
if numpy.real(pole[0]) != 0:
p1r = abs(numpy.real(pole[0]))
else:
p1r = 1
if numpy.real(pole[1]) != 0:
p2r = abs(numpy.real(pole[1]))
else:
p2r = 1
if numpy.imag(pole[0]) != 0:
p1i = abs(numpy.imag(pole[0]))
else:
p1i = 1
if numpy.imag(pole[1]) != 0:
p2i = abs(numpy.imag(pole[1]))
else:
p2i = 1
print('A\'s eigenvalues are:',round(sols[0][0],4),'and',round(sols[0][1],4))
#T = numpy.linspace(0, 60, 1000)
T, yout, xout = control.initial_response(sys,X0=X0,return_x=True)
fig = plt.figure("Free response", figsize=(16,5))
ax = fig.add_subplot(121)
plt.plot(T,xout[0])
plt.grid()
ax.set_xlabel('time [s]')
ax.set_ylabel(r'$x_1$')
ax1 = fig.add_subplot(122)
plt.plot(T,xout[1])
plt.grid()
ax1.set_xlabel('time [s]')
ax1.set_ylabel(r'$x_2$')
alltogether = widgets.VBox([widgets.HBox([widgets.Label('$A$:',border=3),
Aw,
widgets.Label(' ',border=3),
widgets.Label('$X_0$:',border=3),
X0w,
START])])
out = widgets.interactive_output(main_callback, {'A':Aw, 'X0':X0w, 'DW':DW})
out.layout.height = '350px'
display(out, alltogether)
# +
#create dummy widget 2
DW2 = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
DW2.value = -1
#create button widget
START2 = widgets.Button(
description='Show answers',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Click for view the answers',
icon='check'
)
def on_start_button_clicked2(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW2.value> 0 :
DW2.value = -1
else:
DW2.value = 1
pass
START2.on_click(on_start_button_clicked2)
def main_callback2(DW2):
if DW2 > 0:
display(Markdown(r'''>Answer: The only initial condition that hides completly the divergent modes is the state space origin.
$$ $$
Example:
$$
A = \begin{bmatrix} 1 & 0 \\ 0 & 1 \end{bmatrix}
$$'''))
else:
display(Markdown(''))
#create a graphic structure to hold all widgets
alltogether2 = widgets.VBox([START2])
out2 = widgets.interactive_output(main_callback2,{'DW2':DW2})
#out.layout.height = '300px'
display(out2,alltogether2)
# -
| ICCT_si/.ipynb_checkpoints/SS-21-Internal_stability_example_4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import datetime
import matplotlib.pyplot as plt
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import ensemble
a = pd.read_csv('USDCAD_Candlestick_1_M_BID_27.12.2020-28.12.2020.csv')
df = pd.DataFrame(a)
display(df.head())
display(df.tail())
# Set the date as datetime
df['datetime'] = pd.to_datetime(df['Gmt time'])
df = df.set_index(df['datetime'])
df.index.names = [None]
df['Close'].plot(figsize=(40,20))
# +
# Create return (open-close) normalized
# We use this value to train mode
df['return_next'] = df['Open'].shift(-1) - df['Close'].shift(-1)
return_range = df['return_next'].max() - df['return_next'].min()
df['return_next'] = df['return_next'] / return_range
df['return'] = df['Open'] - df['Close']
return_range = df['return'].max() - df['return'].min()
df['return'] = df['return'] / return_range
# If return > 0, green; otherwise, red
df['return'].plot(figsize=(40,20))
# +
# Make label, 1 as rising price, 0 as falling price - prediction for the following timestamp
df['label'] = df['return_next'].apply(lambda x: 1 if x>0.0 else 0)
#drop columns that show NAN
df.dropna(inplace=True)
df.head(10)
# -
df.dtypes
# +
# Make training dataset
n_features = 60 # number of features
train_x = np.array([]).reshape([-1,n_features])
train_y = np.array([]).reshape([-1,1])
for index, row in df.iterrows():
i = df.index.get_loc(index)
if i<n_features:
continue
_x = np.array(df[i-n_features+1:i+1]['return']).T.reshape([1, -1])
_y = df.iloc[i]['label']
train_x = np.vstack((train_x, _x))
train_y = np.vstack((train_y, _y))
train_y = train_y.reshape([-1])
print(train_x.shape)
print(train_y.shape)
print('%% of Class0 : %f' % (np.count_nonzero(train_y == 0)/float(len(train_y))))
print('%% of Class1 : %f' % (np.count_nonzero(train_y == 1)/float(len(train_y))))
# +
# Define Model and fit
# Here we use 95% of data for training, and last 5% for testing
##clf = GradientBoostingClassifier(random_state=0, learning_rate=0.01, n_estimators=10000)
clf = ensemble.GradientBoostingClassifier(verbose=3)
train_len = int(len(train_x)*0.95)
x_train = train_x[:train_len]
y_train = train_y[:train_len]
clf.fit(x_train, y_train)
# -
accuracy = clf.score(x_train, y_train)
print('Testing Accuracy: %f' % accuracy)
# +
# Predict test data
pred = clf.predict(train_x[train_len:])
# Calculate equity..
contracts = 10000.0
commission = 0.0
df_trade = pd.DataFrame(train_x[train_len:,-1], columns=['return'])
df_trade['label'] = train_y[train_len:]
df_trade['pred'] = pred
df_trade['won'] = df_trade['label'] == df_trade['pred']
df_trade['return'] = df_trade['return'].shift(-1) * return_range
df_trade.drop(df_trade.index[len(df_trade)-1], inplace=True)
def calc_profit(row):
if row['won']:
return abs(row['return'])*contracts - commission
else:
return -abs(row['return'])*contracts - commission
df_trade['profit'] = df_trade.apply(lambda row: calc_profit(row), axis=1)
df_trade['equity'] = df_trade['profit'].cumsum()
display(df_trade.tail())
df_trade.plot(y='equity', figsize=(10,4), title='Backtest with $10000 initial capital')
plt.xlabel('Trades')
plt.ylabel('Equity (USD)')
for r in df_trade.iterrows():
if r[1]['won']:
plt.axvline(x=r[0], linewidth=0.5, alpha=0.8, color='g')
else:
plt.axvline(x=r[0], linewidth=0.5, alpha=0.8, color='r')
# +
# Calculate summary of trades
n_win_trades = float(df_trade[df_trade['profit']>0.0]['profit'].count())
n_los_trades = float(df_trade[df_trade['profit']<0.0]['profit'].count())
print("Net Profit : $%.2f" % df_trade.tail(1)['equity'])
print("Number Winning Trades : %d" % n_win_trades)
print("Number Losing Trades : %d" % n_los_trades)
print("Percent Profitable : %.2f%%" % (100*n_win_trades/(n_win_trades + n_los_trades)))
print("Avg Win Trade : $%.3f" % df_trade[df_trade['profit']>0.0]['profit'].mean())
print("Avg Los Trade : $%.3f" % df_trade[df_trade['profit']<0.0]['profit'].mean())
print("Largest Win Trade : $%.3f" % df_trade[df_trade['profit']>0.0]['profit'].max())
print("Largest Los Trade : $%.3f" % df_trade[df_trade['profit']<0.0]['profit'].min())
print("Profit Factor : %.2f" % abs(df_trade[df_trade['profit']>0.0]['profit'].sum()/df_trade[df_trade['profit']<0.0]['profit'].sum()))
df_trade['profit'].hist(bins=20)
# -
| ML_Binary_Options_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Resources
# A lot of resources are mentioned here: http://mccormickml.com/2016/04/27/word2vec-resources/
#
# As we know, CBOW is learning to predict the word by the context. Or maximize the probability of the target word by looking at the context. And this happens to be a problem for rare words. For example, given the context `yesterday was a really [...] day` CBOW model will tell you that most probably the word is `beautiful` or `nice`. Words like `delightful` will get much less attention of the model, because it is designed to predict the most probable word. This word will be smoothed over a lot of examples with more frequent words.
#
# On the other hand, the skip-gram model is designed to predict the context. Given the word `delightful` it must understand it and tell us that there is a huge probability that the context is `yesterday was really [...] day`, or some other relevant context. With skip-gram the word delightful will not try to compete with the word beautiful but instead, delightful+context pairs will be treated as new observations.
# +
import torch
import torch.nn as nn
import torch.autograd as autograd
import torch.optim as optim
import torch.nn.functional as F
import operator
# Continuous Bag of Words model
class CBOW(nn.Module):
def __init__(self, context_size=2, embedding_size=100, vocab_size=None):
super(CBOW, self).__init__()
self.embeddings = nn.Embedding(vocab_size, embedding_size)
self.linear1 = nn.Linear(embedding_size, vocab_size)
def forward(self, inputs):
lookup_embeds = self.embeddings(inputs)
embeds = lookup_embeds.sum(dim=0)
out = self.linear1(embeds)
out = F.log_softmax(out)
return out
def make_context_vector(context, word_to_ix):
idxs = [word_to_ix[w] for w in context]
tensor = torch.LongTensor(idxs)
return autograd.Variable(tensor)
# +
CONTEXT_SIZE = 2 # 2 words to the left, 2 to the right
EMBEDDING_SIZE = 10
raw_text = """We are about to study the idea of a computational process.
Computational processes are abstract beings that inhabit computers.
As they evolve, processes manipulate other abstract things called data.
The evolution of a process is directed by a pattern of rules
called a program. People create programs to direct processes. In effect,
we conjure the spirits of the computer with our spells.""".lower().split()
# How could you do a better pre-processing?
# Maybe a sentence tokenizer?
# Maybe a word lemmatizer?
# Should you take a bigger corpus, Replace this small corpus with a bigger one
# Maybe you should remove stopwords
# Maybe you should just Google?
# +
# Create the vocabulary
vocab = set(raw_text)
vocab_size = len(vocab)
word_to_ix = {word: i for i, word in enumerate(vocab)}
data = []
for i in range(2, len(raw_text) - 2):
context = [raw_text[i - 2], raw_text[i - 1],
raw_text[i + 1], raw_text[i + 2]]
target = raw_text[i]
data.append((context, target))
print (data[0])
# +
loss_func = nn.CrossEntropyLoss()
net = CBOW(CONTEXT_SIZE, embedding_size=EMBEDDING_SIZE, vocab_size=vocab_size)
optimizer = optim.SGD(net.parameters(), lr=0.01)
# The training loop
for epoch in range(100):
total_loss = 0
for context, target in data:
context_var = make_context_vector(context, word_to_ix)
net.zero_grad()
## Enter code to get log_probs from model
target = autograd.Variable(torch.LongTensor([word_to_ix[target]]))
loss = loss_func(log_probs.reshape(1,-1), target)
loss.backward()
optimizer.step()
total_loss += loss.data
print("Loss for epoch ", epoch, " : ", total_loss)
# -
# Now let's find embedding for every word
vocab_to_embedding = {}
for word in vocab:
vocab_to_embedding[word] = net.embeddings.forward(make_context_vector([word], word_to_ix))
def find_k_similar_words(word, k = 5):
word = word.lower()
if word not in vocab:
print ("Not found ", word)
return []
a = vocab_to_embedding[word]
max_sim = -1
sim_here = {}
for b in vocab_to_embedding:
emb = vocab_to_embedding[b]
sim = torch.dot(a.reshape(-1),emb.reshape(-1))/(a.norm()*emb.norm())
sim_here[b] = sim.data[0]
sorted_t = sorted(sim_here.items(), key=operator.itemgetter(1))
sorted_t.reverse()
return sorted_t[:k]
find_k_similar_words('program.', 5)
# ### Could you define a Skip Gram model?
| Day9/NLP-Lab/CBOW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # scikit-learn-random forest
#environment setup with watermark
# %load_ext watermark
# %watermark -a 'Gopala KR' -u -d -v -p watermark,numpy,pandas,matplotlib,nltk,sklearn,tensorflow,theano,mxnet,chainer
# Credits: Forked from [PyCon 2015 Scikit-learn Tutorial](https://github.com/jakevdp/sklearn_pycon2015) by <NAME>
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import seaborn;
from sklearn.linear_model import LinearRegression
from scipy import stats
import pylab as pl
seaborn.set()
# -
# ## Random Forest Classifier
# Random forests are an example of an *ensemble learner* built on decision trees.
# For this reason we'll start by discussing decision trees themselves.
#
# Decision trees are extremely intuitive ways to classify or label objects: you simply ask a series of questions designed to zero-in on the classification:
import fig_code
fig_code.plot_example_decision_tree()
# The binary splitting makes this extremely efficient.
# As always, though, the trick is to *ask the right questions*.
# This is where the algorithmic process comes in: in training a decision tree classifier, the algorithm looks at the features and decides which questions (or "splits") contain the most information.
#
# ### Creating a Decision Tree
#
# Here's an example of a decision tree classifier in scikit-learn. We'll start by defining some two-dimensional labeled data:
# +
from sklearn.datasets import make_blobs
X, y = make_blobs(n_samples=300, centers=4,
random_state=0, cluster_std=1.0)
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap='rainbow');
# +
# We have some convenience functions in the repository that help
from fig_code import visualize_tree, plot_tree_interactive
# Now using IPython's ``interact`` (available in IPython 2.0+, and requires a live kernel) we can view the decision tree splits:
plot_tree_interactive(X, y);
# -
# Notice that at each increase in depth, every node is split in two **except** those nodes which contain only a single class.
# The result is a very fast **non-parametric** classification, and can be extremely useful in practice.
#
# **Question: Do you see any problems with this?**
# ### Decision Trees and over-fitting
#
# One issue with decision trees is that it is very easy to create trees which **over-fit** the data. That is, they are flexible enough that they can learn the structure of the noise in the data rather than the signal! For example, take a look at two trees built on two subsets of this dataset:
# +
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier()
plt.figure()
visualize_tree(clf, X[:200], y[:200], boundaries=False)
plt.figure()
visualize_tree(clf, X[-200:], y[-200:], boundaries=False)
# -
# The details of the classifications are completely different! That is an indication of **over-fitting**: when you predict the value for a new point, the result is more reflective of the noise in the model rather than the signal.
# ## Ensembles of Estimators: Random Forests
#
# One possible way to address over-fitting is to use an **Ensemble Method**: this is a meta-estimator which essentially averages the results of many individual estimators which over-fit the data. Somewhat surprisingly, the resulting estimates are much more robust and accurate than the individual estimates which make them up!
#
# One of the most common ensemble methods is the **Random Forest**, in which the ensemble is made up of many decision trees which are in some way perturbed.
#
# There are volumes of theory and precedent about how to randomize these trees, but as an example, let's imagine an ensemble of estimators fit on subsets of the data. We can get an idea of what these might look like as follows:
# +
def fit_randomized_tree(random_state=0):
X, y = make_blobs(n_samples=300, centers=4,
random_state=0, cluster_std=2.0)
clf = DecisionTreeClassifier(max_depth=15)
rng = np.random.RandomState(random_state)
i = np.arange(len(y))
rng.shuffle(i)
visualize_tree(clf, X[i[:250]], y[i[:250]], boundaries=False,
xlim=(X[:, 0].min(), X[:, 0].max()),
ylim=(X[:, 1].min(), X[:, 1].max()))
from IPython.html.widgets import interact
interact(fit_randomized_tree, random_state=[0, 100]);
# -
# See how the details of the model change as a function of the sample, while the larger characteristics remain the same!
# The random forest classifier will do something similar to this, but use a combined version of all these trees to arrive at a final answer:
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators=100, random_state=0, n_jobs=-1)
visualize_tree(clf, X, y, boundaries=False);
# By averaging over 100 randomly perturbed models, we end up with an overall model which is a much better fit to our data!
#
# *(Note: above we randomized the model through sub-sampling... Random Forests use more sophisticated means of randomization, which you can read about in, e.g. the [scikit-learn documentation](http://scikit-learn.org/stable/modules/ensemble.html#forest)*)
# Not good for random forest:
# lots of 0, few 1
# structured data like images, neural network might be better
# small data, might overfit
# high dimensional data, linear model might work better
# ## Random Forest Regressor
# Above we were considering random forests within the context of classification.
# Random forests can also be made to work in the case of regression (that is, continuous rather than categorical variables). The estimator to use for this is ``sklearn.ensemble.RandomForestRegressor``.
#
# Let's quickly demonstrate how this can be used:
# +
from sklearn.ensemble import RandomForestRegressor
x = 10 * np.random.rand(100)
def model(x, sigma=0.3):
fast_oscillation = np.sin(5 * x)
slow_oscillation = np.sin(0.5 * x)
noise = sigma * np.random.randn(len(x))
return slow_oscillation + fast_oscillation + noise
y = model(x)
plt.errorbar(x, y, 0.3, fmt='o');
# +
xfit = np.linspace(0, 10, 1000)
yfit = RandomForestRegressor(100).fit(x[:, None], y).predict(xfit[:, None])
ytrue = model(xfit, 0)
plt.errorbar(x, y, 0.3, fmt='o')
plt.plot(xfit, yfit, '-r');
plt.plot(xfit, ytrue, '-k', alpha=0.5);
# -
# As you can see, the non-parametric random forest model is flexible enough to fit the multi-period data, without us even specifying a multi-period model!
#
# Tradeoff between simplicity and thinking about what your data is.
#
# Feature engineering is important, need to know your domain: Fourier transform frequency distribution.
# ## Random Forest Limitations
#
# The following data scenarios are not well suited for random forests:
# * y: lots of 0, few 1
# * Structured data like images where a neural network might be better
# * Small data size which might lead to overfitting
# * High dimensional data where a linear model might work better
test complete; Gopal
| tests/scikit-learn/scikit-learn-random-forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Build Speech data files
# +
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from IPython.display import display
# %matplotlib inline
# +
df = pd.read_csv('data/audio_features.csv')
df = df[df['label'].isin([0, 1, 2, 3, 4, 5, 6, 7])]
print(df.shape)
display(df.head())
# change 7 to 2
df['label'] = df['label'].map({0: 0, 1: 1, 2: 1, 3: 2, 4: 2, 5: 3, 6: 4, 7: 5})
df.head()
# +
df.to_csv('data/no_sample_df.csv')
# oversample fear
fear_df = df[df['label']==3]
for i in range(30):
df = df.append(fear_df)
sur_df = df[df['label']==4]
for i in range(10):
df = df.append(sur_df)
df.to_csv('data/modified_df.csv')
# +
emotion_dict = {'ang': 0,
'hap': 1,
'sad': 2,
'neu': 3,}
# emotion_dict = {'ang': 0,
# 'hap': 1,
# 'exc': 2,
# 'sad': 3,
# 'fru': 4,
# 'fea': 5,
# 'sur': 6,
# 'neu': 7,
# 'xxx': 8,
# 'oth': 8}
scalar = MinMaxScaler()
df[df.columns[2:]] = scalar.fit_transform(df[df.columns[2:]])
df.head()
# +
x_train, x_test = train_test_split(df, test_size=0.20)
x_train.to_csv('data/s2e/audio_train.csv', index=False)
x_test.to_csv('data/s2e/audio_test.csv', index=False)
print(x_train.shape, x_test.shape)
# -
# ## Define preprocessing functions for text
# +
import unicodedata
def unicodeToAscii(s):
return ''.join(
c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn'
)
# Lowercase, trim, and remove non-letter characters
def normalizeString(s):
s = unicodeToAscii(s.lower().strip())
s = re.sub(r"([.!?])", r" \1", s)
s = re.sub(r"[^a-zA-Z.!?]+", r" ", s)
return s
# -
# ## Build Text data files
# +
import re
import os
import pickle
useful_regex = re.compile(r'^(\w+)', re.IGNORECASE)
file2transcriptions = {}
for sess in range(1, 6):
transcripts_path = 'data/IEMOCAP_full_release/Session{}/dialog/transcriptions/'.format(sess)
transcript_files = os.listdir(transcripts_path)
for f in transcript_files:
with open('{}{}'.format(transcripts_path, f), 'r') as f:
all_lines = f.readlines()
for l in all_lines:
audio_code = useful_regex.match(l).group()
transcription = l.split(':')[-1].strip()
# assuming that all the keys would be unique and hence no `try`
file2transcriptions[audio_code] = transcription
# save dict
with open('data/t2e/audiocode2text.pkl', 'wb') as file:
pickle.dump(file2transcriptions, file)
len(file2transcriptions)
# -
audiocode2text = pickle.load(open('data/t2e/audiocode2text.pkl', 'rb'))
# +
# Prepare text data
text_train = pd.DataFrame()
text_train['wav_file'] = x_train['wav_file']
text_train['label'] = x_train['label']
text_train['transcription'] = [normalizeString(audiocode2text[code]) for code in x_train['wav_file']]
text_test = pd.DataFrame()
text_test['wav_file'] = x_test['wav_file']
text_test['label'] = x_test['label']
text_test['transcription'] = [normalizeString(audiocode2text[code]) for code in x_test['wav_file']]
text_train.to_csv('data/t2e/text_train.csv', index=False)
text_test.to_csv('data/t2e/text_test.csv', index=False)
print(text_train.shape, text_test.shape)
| 4_prepare_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="8WMvRGjnf5rr"
# #!mkdir -p ~/.kaggle
# #!cp kaggle.json ~/.kaggle/
# + colab={"base_uri": "https://localhost:8080/"} id="AvT3UpYtf7iL" outputId="cb2b0f44-0345-4542-d8fa-c658ab37559f"
# #!pip install kaggle
# + id="MyGWXrUHf9DV"
# #!chmod 600 /root/.kaggle/kaggle.json
# + colab={"base_uri": "https://localhost:8080/"} id="0x0p4J09f-Wt" outputId="dbcac03e-58b2-49e2-8428-2b10171fac78"
# #!kaggle competitions download -c home-credit-default-risk
# + colab={"base_uri": "https://localhost:8080/"} id="BY9VjC4xiDl6" outputId="1910b447-4c2e-4624-b417-8fe2ab08a941"
# #!unzip \*.zip -d dataset
# + id="bY_gS3lSj18f"
# #!rm *zip *csv
# + id="bbZooWebfvD3"
import os
import gc
import numpy as np
import pandas as pd
import multiprocessing as mp
from scipy.stats import kurtosis
import matplotlib.pyplot as plt
import seaborn as sns
warnings.simplefilter(action='ignore', category=FutureWarning)
# + id="1SNgv8eKfvD5"
DATA_DIRECTORY = "dataset"
# + colab={"base_uri": "https://localhost:8080/"} id="0PHJ3WiVfvD7" outputId="dfec7a9f-69e1-4da0-d941-4756149f223a"
df_train = pd.read_csv(os.path.join(DATA_DIRECTORY, 'application_train.csv'))
df_test = pd.read_csv(os.path.join(DATA_DIRECTORY, 'application_test.csv'))
df = df_train.append(df_test)
del df_train, df_test; gc.collect()
# + id="lbsWdO_HfvD7"
df = df[df['AMT_INCOME_TOTAL'] < 20000000]
df = df[df['CODE_GENDER'] != 'XNA']
df['DAYS_EMPLOYED'].replace(365243, np.nan, inplace=True)
df['DAYS_LAST_PHONE_CHANGE'].replace(0, np.nan, inplace=True)
# + id="DRM56nqxfvD8"
def get_age_group(days_birth):
age_years = -days_birth / 365
if age_years < 27: return 1
elif age_years < 40: return 2
elif age_years < 50: return 3
elif age_years < 65: return 4
elif age_years < 99: return 5
else: return 0
# + id="0Y9u5rCYfvD9"
docs = [f for f in df.columns if 'FLAG_DOC' in f]
df['DOCUMENT_COUNT'] = df[docs].sum(axis=1)
df['NEW_DOC_KURT'] = df[docs].kurtosis(axis=1)
df['AGE_RANGE'] = df['DAYS_BIRTH'].apply(lambda x: get_age_group(x))
# + id="7tGXFHxRfvD9"
df['EXT_SOURCES_PROD'] = df['EXT_SOURCE_1'] * df['EXT_SOURCE_2'] * df['EXT_SOURCE_3']
df['EXT_SOURCES_WEIGHTED'] = df.EXT_SOURCE_1 * 2 + df.EXT_SOURCE_2 * 1 + df.EXT_SOURCE_3 * 3
np.warnings.filterwarnings('ignore', r'All-NaN (slice|axis) encountered')
for function_name in ['min', 'max', 'mean', 'nanmedian', 'var']:
feature_name = 'EXT_SOURCES_{}'.format(function_name.upper())
df[feature_name] = eval('np.{}'.format(function_name))(
df[['EXT_SOURCE_1', 'EXT_SOURCE_2', 'EXT_SOURCE_3']], axis=1)
# + id="l88M7zcefvD-"
df['CREDIT_TO_ANNUITY_RATIO'] = df['AMT_CREDIT'] / df['AMT_ANNUITY']
df['CREDIT_TO_GOODS_RATIO'] = df['AMT_CREDIT'] / df['AMT_GOODS_PRICE']
df['ANNUITY_TO_INCOME_RATIO'] = df['AMT_ANNUITY'] / df['AMT_INCOME_TOTAL']
df['CREDIT_TO_INCOME_RATIO'] = df['AMT_CREDIT'] / df['AMT_INCOME_TOTAL']
df['INCOME_TO_EMPLOYED_RATIO'] = df['AMT_INCOME_TOTAL'] / df['DAYS_EMPLOYED']
df['INCOME_TO_BIRTH_RATIO'] = df['AMT_INCOME_TOTAL'] / df['DAYS_BIRTH']
df['EMPLOYED_TO_BIRTH_RATIO'] = df['DAYS_EMPLOYED'] / df['DAYS_BIRTH']
df['ID_TO_BIRTH_RATIO'] = df['DAYS_ID_PUBLISH'] / df['DAYS_BIRTH']
df['CAR_TO_BIRTH_RATIO'] = df['OWN_CAR_AGE'] / df['DAYS_BIRTH']
df['CAR_TO_EMPLOYED_RATIO'] = df['OWN_CAR_AGE'] / df['DAYS_EMPLOYED']
df['PHONE_TO_BIRTH_RATIO'] = df['DAYS_LAST_PHONE_CHANGE'] / df['DAYS_BIRTH']
# + id="--oHB5mIfvD-"
def do_mean(df, group_cols, counted, agg_name):
gp = df[group_cols + [counted]].groupby(group_cols)[counted].mean().reset_index().rename(
columns={counted: agg_name})
df = df.merge(gp, on=group_cols, how='left')
del gp
gc.collect()
return df
# + id="-AjK9cklfvD_"
def do_median(df, group_cols, counted, agg_name):
gp = df[group_cols + [counted]].groupby(group_cols)[counted].median().reset_index().rename(
columns={counted: agg_name})
df = df.merge(gp, on=group_cols, how='left')
del gp
gc.collect()
return df
# + id="vRkrot_8fvD_"
def do_std(df, group_cols, counted, agg_name):
gp = df[group_cols + [counted]].groupby(group_cols)[counted].std().reset_index().rename(
columns={counted: agg_name})
df = df.merge(gp, on=group_cols, how='left')
del gp
gc.collect()
return df
# + id="URTqRsY6fvEA"
def do_sum(df, group_cols, counted, agg_name):
gp = df[group_cols + [counted]].groupby(group_cols)[counted].sum().reset_index().rename(
columns={counted: agg_name})
df = df.merge(gp, on=group_cols, how='left')
del gp
gc.collect()
return df
# + id="uH-QMvWOfvEA"
group = ['ORGANIZATION_TYPE', 'NAME_EDUCATION_TYPE', 'OCCUPATION_TYPE', 'AGE_RANGE', 'CODE_GENDER']
df = do_median(df, group, 'EXT_SOURCES_MEAN', 'GROUP_EXT_SOURCES_MEDIAN')
df = do_std(df, group, 'EXT_SOURCES_MEAN', 'GROUP_EXT_SOURCES_STD')
df = do_mean(df, group, 'AMT_INCOME_TOTAL', 'GROUP_INCOME_MEAN')
df = do_std(df, group, 'AMT_INCOME_TOTAL', 'GROUP_INCOME_STD')
df = do_mean(df, group, 'CREDIT_TO_ANNUITY_RATIO', 'GROUP_CREDIT_TO_ANNUITY_MEAN')
df = do_std(df, group, 'CREDIT_TO_ANNUITY_RATIO', 'GROUP_CREDIT_TO_ANNUITY_STD')
df = do_mean(df, group, 'AMT_CREDIT', 'GROUP_CREDIT_MEAN')
df = do_mean(df, group, 'AMT_ANNUITY', 'GROUP_ANNUITY_MEAN')
df = do_std(df, group, 'AMT_ANNUITY', 'GROUP_ANNUITY_STD')
# + id="pgJRRETefvEB"
def label_encoder(df, categorical_columns=None):
if not categorical_columns:
categorical_columns = [col for col in df.columns if df[col].dtype == 'object']
for col in categorical_columns:
df[col], uniques = pd.factorize(df[col])
return df, categorical_columns
# + id="fvOARQAgfvEB"
def drop_application_columns(df):
drop_list = [
'CNT_CHILDREN', 'CNT_FAM_MEMBERS', 'HOUR_APPR_PROCESS_START',
'FLAG_EMP_PHONE', 'FLAG_MOBIL', 'FLAG_CONT_MOBILE', 'FLAG_EMAIL', 'FLAG_PHONE',
'FLAG_OWN_REALTY', 'REG_REGION_NOT_LIVE_REGION', 'REG_REGION_NOT_WORK_REGION',
'REG_CITY_NOT_WORK_CITY', 'OBS_30_CNT_SOCIAL_CIRCLE', 'OBS_60_CNT_SOCIAL_CIRCLE',
'AMT_REQ_CREDIT_BUREAU_DAY', 'AMT_REQ_CREDIT_BUREAU_MON', 'AMT_REQ_CREDIT_BUREAU_YEAR',
'COMMONAREA_MODE', 'NONLIVINGAREA_MODE', 'ELEVATORS_MODE', 'NONLIVINGAREA_AVG',
'FLOORSMIN_MEDI', 'LANDAREA_MODE', 'NONLIVINGAREA_MEDI', 'LIVINGAPARTMENTS_MODE',
'FLOORSMIN_AVG', 'LANDAREA_AVG', 'FLOORSMIN_MODE', 'LANDAREA_MEDI',
'COMMONAREA_MEDI', 'YEARS_BUILD_AVG', 'COMMONAREA_AVG', 'BASEMENTAREA_AVG',
'BASEMENTAREA_MODE', 'NONLIVINGAPARTMENTS_MEDI', 'BASEMENTAREA_MEDI',
'LIVINGAPARTMENTS_AVG', 'ELEVATORS_AVG', 'YEARS_BUILD_MEDI', 'ENTRANCES_MODE',
'NONLIVINGAPARTMENTS_MODE', 'LIVINGAREA_MODE', 'LIVINGAPARTMENTS_MEDI',
'YEARS_BUILD_MODE', 'YEARS_BEGINEXPLUATATION_AVG', 'ELEVATORS_MEDI', 'LIVINGAREA_MEDI',
'YEARS_BEGINEXPLUATATION_MODE', 'NONLIVINGAPARTMENTS_AVG', 'HOUSETYPE_MODE',
'FONDKAPREMONT_MODE', 'EMERGENCYSTATE_MODE'
]
for doc_num in [2,4,5,6,7,9,10,11,12,13,14,15,16,17,19,20,21]:
drop_list.append('FLAG_DOCUMENT_{}'.format(doc_num))
df.drop(drop_list, axis=1, inplace=True)
return df
# + id="SYuEW0klfvEC"
df, le_encoded_cols = label_encoder(df, None)
df = drop_application_columns(df)
# + id="KCnOZUiKfvEC"
df = pd.get_dummies(df)
# + id="oKDRi8ssfvEC"
bureau = pd.read_csv(os.path.join(DATA_DIRECTORY, 'bureau.csv'))
# + id="an0Q-OdSfvEC"
bureau['CREDIT_DURATION'] = -bureau['DAYS_CREDIT'] + bureau['DAYS_CREDIT_ENDDATE']
bureau['ENDDATE_DIF'] = bureau['DAYS_CREDIT_ENDDATE'] - bureau['DAYS_ENDDATE_FACT']
bureau['DEBT_PERCENTAGE'] = bureau['AMT_CREDIT_SUM'] / bureau['AMT_CREDIT_SUM_DEBT']
bureau['DEBT_CREDIT_DIFF'] = bureau['AMT_CREDIT_SUM'] - bureau['AMT_CREDIT_SUM_DEBT']
bureau['CREDIT_TO_ANNUITY_RATIO'] = bureau['AMT_CREDIT_SUM'] / bureau['AMT_ANNUITY']
# + id="a-6yU6iUfvED"
def one_hot_encoder(df, categorical_columns=None, nan_as_category=True):
original_columns = list(df.columns)
if not categorical_columns:
categorical_columns = [col for col in df.columns if df[col].dtype == 'object']
df = pd.get_dummies(df, columns=categorical_columns, dummy_na=nan_as_category)
categorical_columns = [c for c in df.columns if c not in original_columns]
return df, categorical_columns
# + id="MLZf1YK4fvED"
def group(df_to_agg, prefix, aggregations, aggregate_by= 'SK_ID_CURR'):
agg_df = df_to_agg.groupby(aggregate_by).agg(aggregations)
agg_df.columns = pd.Index(['{}{}_{}'.format(prefix, e[0], e[1].upper())
for e in agg_df.columns.tolist()])
return agg_df.reset_index()
# + id="ZjLTbd4tfvED"
def group_and_merge(df_to_agg, df_to_merge, prefix, aggregations, aggregate_by= 'SK_ID_CURR'):
agg_df = group(df_to_agg, prefix, aggregations, aggregate_by= aggregate_by)
return df_to_merge.merge(agg_df, how='left', on= aggregate_by)
# + id="BkjdcDZPfvED"
def get_bureau_balance(path, num_rows= None):
bb = pd.read_csv(os.path.join(path, 'bureau_balance.csv'))
bb, categorical_cols = one_hot_encoder(bb, nan_as_category= False)
bb_processed = bb.groupby('SK_ID_BUREAU')[categorical_cols].mean().reset_index()
agg = {'MONTHS_BALANCE': ['min', 'max', 'mean', 'size']}
bb_processed = group_and_merge(bb, bb_processed, '', agg, 'SK_ID_BUREAU')
del bb; gc.collect()
return bb_processed
# + id="NOaYMiuifvEE"
bureau, categorical_cols = one_hot_encoder(bureau, nan_as_category= False)
bureau = bureau.merge(get_bureau_balance(DATA_DIRECTORY), how='left', on='SK_ID_BUREAU')
bureau['STATUS_12345'] = 0
for i in range(1,6):
bureau['STATUS_12345'] += bureau['STATUS_{}'.format(i)]
# + colab={"base_uri": "https://localhost:8080/"} id="Br6IzFW7fvEE" outputId="7dab293a-e607-420a-a782-83dd492ce4dc"
features = ['AMT_CREDIT_MAX_OVERDUE', 'AMT_CREDIT_SUM_OVERDUE', 'AMT_CREDIT_SUM',
'AMT_CREDIT_SUM_DEBT', 'DEBT_PERCENTAGE', 'DEBT_CREDIT_DIFF', 'STATUS_0', 'STATUS_12345']
agg_length = bureau.groupby('MONTHS_BALANCE_SIZE')[features].mean().reset_index()
agg_length.rename({feat: 'LL_' + feat for feat in features}, axis=1, inplace=True)
bureau = bureau.merge(agg_length, how='left', on='MONTHS_BALANCE_SIZE')
del agg_length; gc.collect()
# + id="EBx-5Fk_fvEE"
BUREAU_AGG = {
'SK_ID_BUREAU': ['nunique'],
'DAYS_CREDIT': ['min', 'max', 'mean'],
'DAYS_CREDIT_ENDDATE': ['min', 'max'],
'AMT_CREDIT_MAX_OVERDUE': ['max', 'mean'],
'AMT_CREDIT_SUM': ['max', 'mean', 'sum'],
'AMT_CREDIT_SUM_DEBT': ['max', 'mean', 'sum'],
'AMT_CREDIT_SUM_OVERDUE': ['max', 'mean', 'sum'],
'AMT_ANNUITY': ['mean'],
'DEBT_CREDIT_DIFF': ['mean', 'sum'],
'MONTHS_BALANCE_MEAN': ['mean', 'var'],
'MONTHS_BALANCE_SIZE': ['mean', 'sum'],
'STATUS_0': ['mean'],
'STATUS_1': ['mean'],
'STATUS_12345': ['mean'],
'STATUS_C': ['mean'],
'STATUS_X': ['mean'],
'CREDIT_ACTIVE_Active': ['mean'],
'CREDIT_ACTIVE_Closed': ['mean'],
'CREDIT_ACTIVE_Sold': ['mean'],
'CREDIT_TYPE_Consumer credit': ['mean'],
'CREDIT_TYPE_Credit card': ['mean'],
'CREDIT_TYPE_Car loan': ['mean'],
'CREDIT_TYPE_Mortgage': ['mean'],
'CREDIT_TYPE_Microloan': ['mean'],
'LL_AMT_CREDIT_SUM_OVERDUE': ['mean'],
'LL_DEBT_CREDIT_DIFF': ['mean'],
'LL_STATUS_12345': ['mean'],
}
BUREAU_ACTIVE_AGG = {
'DAYS_CREDIT': ['max', 'mean'],
'DAYS_CREDIT_ENDDATE': ['min', 'max'],
'AMT_CREDIT_MAX_OVERDUE': ['max', 'mean'],
'AMT_CREDIT_SUM': ['max', 'sum'],
'AMT_CREDIT_SUM_DEBT': ['mean', 'sum'],
'AMT_CREDIT_SUM_OVERDUE': ['max', 'mean'],
'DAYS_CREDIT_UPDATE': ['min', 'mean'],
'DEBT_PERCENTAGE': ['mean'],
'DEBT_CREDIT_DIFF': ['mean'],
'CREDIT_TO_ANNUITY_RATIO': ['mean'],
'MONTHS_BALANCE_MEAN': ['mean', 'var'],
'MONTHS_BALANCE_SIZE': ['mean', 'sum'],
}
BUREAU_CLOSED_AGG = {
'DAYS_CREDIT': ['max', 'var'],
'DAYS_CREDIT_ENDDATE': ['max'],
'AMT_CREDIT_MAX_OVERDUE': ['max', 'mean'],
'AMT_CREDIT_SUM_OVERDUE': ['mean'],
'AMT_CREDIT_SUM': ['max', 'mean', 'sum'],
'AMT_CREDIT_SUM_DEBT': ['max', 'sum'],
'DAYS_CREDIT_UPDATE': ['max'],
'ENDDATE_DIF': ['mean'],
'STATUS_12345': ['mean'],
}
BUREAU_LOAN_TYPE_AGG = {
'DAYS_CREDIT': ['mean', 'max'],
'AMT_CREDIT_MAX_OVERDUE': ['mean', 'max'],
'AMT_CREDIT_SUM': ['mean', 'max'],
'AMT_CREDIT_SUM_DEBT': ['mean', 'max'],
'DEBT_PERCENTAGE': ['mean'],
'DEBT_CREDIT_DIFF': ['mean'],
'DAYS_CREDIT_ENDDATE': ['max'],
}
BUREAU_TIME_AGG = {
'AMT_CREDIT_MAX_OVERDUE': ['max', 'mean'],
'AMT_CREDIT_SUM_OVERDUE': ['mean'],
'AMT_CREDIT_SUM': ['max', 'sum'],
'AMT_CREDIT_SUM_DEBT': ['mean', 'sum'],
'DEBT_PERCENTAGE': ['mean'],
'DEBT_CREDIT_DIFF': ['mean'],
'STATUS_0': ['mean'],
'STATUS_12345': ['mean'],
}
# + id="dun76VF7fvEF"
agg_bureau = group(bureau, 'BUREAU_', BUREAU_AGG)
active = bureau[bureau['CREDIT_ACTIVE_Active'] == 1]
agg_bureau = group_and_merge(active,agg_bureau,'BUREAU_ACTIVE_',BUREAU_ACTIVE_AGG)
closed = bureau[bureau['CREDIT_ACTIVE_Closed'] == 1]
agg_bureau = group_and_merge(closed,agg_bureau,'BUREAU_CLOSED_',BUREAU_CLOSED_AGG)
del active, closed; gc.collect()
for credit_type in ['Consumer credit', 'Credit card', 'Mortgage', 'Car loan', 'Microloan']:
type_df = bureau[bureau['CREDIT_TYPE_' + credit_type] == 1]
prefix = 'BUREAU_' + credit_type.split(' ')[0].upper() + '_'
agg_bureau = group_and_merge(type_df, agg_bureau, prefix, BUREAU_LOAN_TYPE_AGG)
del type_df; gc.collect()
for time_frame in [6, 12]:
prefix = "BUREAU_LAST{}M_".format(time_frame)
time_frame_df = bureau[bureau['DAYS_CREDIT'] >= -30*time_frame]
agg_bureau = group_and_merge(time_frame_df, agg_bureau, prefix, BUREAU_TIME_AGG)
del time_frame_df; gc.collect()
# + id="uu17NWC9fvEF"
sort_bureau = bureau.sort_values(by=['DAYS_CREDIT'])
gr = sort_bureau.groupby('SK_ID_CURR')['AMT_CREDIT_MAX_OVERDUE'].last().reset_index()
gr.rename({'AMT_CREDIT_MAX_OVERDUE': 'BUREAU_LAST_LOAN_MAX_OVERDUE'}, inplace=True)
agg_bureau = agg_bureau.merge(gr, on='SK_ID_CURR', how='left')
agg_bureau['BUREAU_DEBT_OVER_CREDIT'] = \
agg_bureau['BUREAU_AMT_CREDIT_SUM_DEBT_SUM']/agg_bureau['BUREAU_AMT_CREDIT_SUM_SUM']
agg_bureau['BUREAU_ACTIVE_DEBT_OVER_CREDIT'] = \
agg_bureau['BUREAU_ACTIVE_AMT_CREDIT_SUM_DEBT_SUM']/agg_bureau['BUREAU_ACTIVE_AMT_CREDIT_SUM_SUM']
# + colab={"base_uri": "https://localhost:8080/"} id="RJ1GVozgfvEG" outputId="01a6ce0c-410c-4c14-b8d3-58bb681a1e25"
df = pd.merge(df, agg_bureau, on='SK_ID_CURR', how='left')
del agg_bureau, bureau
gc.collect()
# + id="TK28yDE_fvEG"
prev = pd.read_csv(os.path.join(DATA_DIRECTORY, 'previous_application.csv'))
pay = pd.read_csv(os.path.join(DATA_DIRECTORY, 'installments_payments.csv'))
# + id="DimspUdZfvEG"
PREVIOUS_AGG = {
'SK_ID_PREV': ['nunique'],
'AMT_ANNUITY': ['min', 'max', 'mean'],
'AMT_DOWN_PAYMENT': ['max', 'mean'],
'HOUR_APPR_PROCESS_START': ['min', 'max', 'mean'],
'RATE_DOWN_PAYMENT': ['max', 'mean'],
'DAYS_DECISION': ['min', 'max', 'mean'],
'CNT_PAYMENT': ['max', 'mean'],
'DAYS_TERMINATION': ['max'],
# Engineered features
'CREDIT_TO_ANNUITY_RATIO': ['mean', 'max'],
'APPLICATION_CREDIT_DIFF': ['min', 'max', 'mean'],
'APPLICATION_CREDIT_RATIO': ['min', 'max', 'mean', 'var'],
'DOWN_PAYMENT_TO_CREDIT': ['mean'],
}
PREVIOUS_ACTIVE_AGG = {
'SK_ID_PREV': ['nunique'],
'SIMPLE_INTERESTS': ['mean'],
'AMT_ANNUITY': ['max', 'sum'],
'AMT_APPLICATION': ['max', 'mean'],
'AMT_CREDIT': ['sum'],
'AMT_DOWN_PAYMENT': ['max', 'mean'],
'DAYS_DECISION': ['min', 'mean'],
'CNT_PAYMENT': ['mean', 'sum'],
'DAYS_LAST_DUE_1ST_VERSION': ['min', 'max', 'mean'],
# Engineered features
'AMT_PAYMENT': ['sum'],
'INSTALMENT_PAYMENT_DIFF': ['mean', 'max'],
'REMAINING_DEBT': ['max', 'mean', 'sum'],
'REPAYMENT_RATIO': ['mean'],
}
PREVIOUS_LATE_PAYMENTS_AGG = {
'DAYS_DECISION': ['min', 'max', 'mean'],
'DAYS_LAST_DUE_1ST_VERSION': ['min', 'max', 'mean'],
# Engineered features
'APPLICATION_CREDIT_DIFF': ['min'],
'NAME_CONTRACT_TYPE_Consumer loans': ['mean'],
'NAME_CONTRACT_TYPE_Cash loans': ['mean'],
'NAME_CONTRACT_TYPE_Revolving loans': ['mean'],
}
PREVIOUS_LOAN_TYPE_AGG = {
'AMT_CREDIT': ['sum'],
'AMT_ANNUITY': ['mean', 'max'],
'SIMPLE_INTERESTS': ['min', 'mean', 'max', 'var'],
'APPLICATION_CREDIT_DIFF': ['min', 'var'],
'APPLICATION_CREDIT_RATIO': ['min', 'max', 'mean'],
'DAYS_DECISION': ['max'],
'DAYS_LAST_DUE_1ST_VERSION': ['max', 'mean'],
'CNT_PAYMENT': ['mean'],
}
PREVIOUS_TIME_AGG = {
'AMT_CREDIT': ['sum'],
'AMT_ANNUITY': ['mean', 'max'],
'SIMPLE_INTERESTS': ['mean', 'max'],
'DAYS_DECISION': ['min', 'mean'],
'DAYS_LAST_DUE_1ST_VERSION': ['min', 'max', 'mean'],
# Engineered features
'APPLICATION_CREDIT_DIFF': ['min'],
'APPLICATION_CREDIT_RATIO': ['min', 'max', 'mean'],
'NAME_CONTRACT_TYPE_Consumer loans': ['mean'],
'NAME_CONTRACT_TYPE_Cash loans': ['mean'],
'NAME_CONTRACT_TYPE_Revolving loans': ['mean'],
}
PREVIOUS_APPROVED_AGG = {
'SK_ID_PREV': ['nunique'],
'AMT_ANNUITY': ['min', 'max', 'mean'],
'AMT_CREDIT': ['min', 'max', 'mean'],
'AMT_DOWN_PAYMENT': ['max'],
'AMT_GOODS_PRICE': ['max'],
'HOUR_APPR_PROCESS_START': ['min', 'max'],
'DAYS_DECISION': ['min', 'mean'],
'CNT_PAYMENT': ['max', 'mean'],
'DAYS_TERMINATION': ['mean'],
# Engineered features
'CREDIT_TO_ANNUITY_RATIO': ['mean', 'max'],
'APPLICATION_CREDIT_DIFF': ['max'],
'APPLICATION_CREDIT_RATIO': ['min', 'max', 'mean'],
# The following features are only for approved applications
'DAYS_FIRST_DRAWING': ['max', 'mean'],
'DAYS_FIRST_DUE': ['min', 'mean'],
'DAYS_LAST_DUE_1ST_VERSION': ['min', 'max', 'mean'],
'DAYS_LAST_DUE': ['max', 'mean'],
'DAYS_LAST_DUE_DIFF': ['min', 'max', 'mean'],
'SIMPLE_INTERESTS': ['min', 'max', 'mean'],
}
PREVIOUS_REFUSED_AGG = {
'AMT_APPLICATION': ['max', 'mean'],
'AMT_CREDIT': ['min', 'max'],
'DAYS_DECISION': ['min', 'max', 'mean'],
'CNT_PAYMENT': ['max', 'mean'],
# Engineered features
'APPLICATION_CREDIT_DIFF': ['min', 'max', 'mean', 'var'],
'APPLICATION_CREDIT_RATIO': ['min', 'mean'],
'NAME_CONTRACT_TYPE_Consumer loans': ['mean'],
'NAME_CONTRACT_TYPE_Cash loans': ['mean'],
'NAME_CONTRACT_TYPE_Revolving loans': ['mean'],
}
# + id="aMtS_j-0fvEG"
ohe_columns = [
'NAME_CONTRACT_STATUS', 'NAME_CONTRACT_TYPE', 'CHANNEL_TYPE',
'NAME_TYPE_SUITE', 'NAME_YIELD_GROUP', 'PRODUCT_COMBINATION',
'NAME_PRODUCT_TYPE', 'NAME_CLIENT_TYPE']
prev, categorical_cols = one_hot_encoder(prev, ohe_columns, nan_as_category= False)
# + id="pRCjHKr_fvEH"
prev['APPLICATION_CREDIT_DIFF'] = prev['AMT_APPLICATION'] - prev['AMT_CREDIT']
prev['APPLICATION_CREDIT_RATIO'] = prev['AMT_APPLICATION'] / prev['AMT_CREDIT']
prev['CREDIT_TO_ANNUITY_RATIO'] = prev['AMT_CREDIT']/prev['AMT_ANNUITY']
prev['DOWN_PAYMENT_TO_CREDIT'] = prev['AMT_DOWN_PAYMENT'] / prev['AMT_CREDIT']
total_payment = prev['AMT_ANNUITY'] * prev['CNT_PAYMENT']
prev['SIMPLE_INTERESTS'] = (total_payment/prev['AMT_CREDIT'] - 1)/prev['CNT_PAYMENT']
# + colab={"base_uri": "https://localhost:8080/"} id="kKtpSpdvfvEH" outputId="00920193-72a9-4b1c-dbe8-6a06a6a118c9"
approved = prev[prev['NAME_CONTRACT_STATUS_Approved'] == 1]
active_df = approved[approved['DAYS_LAST_DUE'] == 365243]
active_pay = pay[pay['SK_ID_PREV'].isin(active_df['SK_ID_PREV'])]
active_pay_agg = active_pay.groupby('SK_ID_PREV')[['AMT_INSTALMENT', 'AMT_PAYMENT']].sum()
active_pay_agg.reset_index(inplace= True)
active_pay_agg['INSTALMENT_PAYMENT_DIFF'] = active_pay_agg['AMT_INSTALMENT'] - active_pay_agg['AMT_PAYMENT']
active_df = active_df.merge(active_pay_agg, on= 'SK_ID_PREV', how= 'left')
active_df['REMAINING_DEBT'] = active_df['AMT_CREDIT'] - active_df['AMT_PAYMENT']
active_df['REPAYMENT_RATIO'] = active_df['AMT_PAYMENT'] / active_df['AMT_CREDIT']
active_agg_df = group(active_df, 'PREV_ACTIVE_', PREVIOUS_ACTIVE_AGG)
active_agg_df['TOTAL_REPAYMENT_RATIO'] = active_agg_df['PREV_ACTIVE_AMT_PAYMENT_SUM']/\
active_agg_df['PREV_ACTIVE_AMT_CREDIT_SUM']
del active_pay, active_pay_agg, active_df; gc.collect()
# + id="cy_90CNPfvEH"
prev['DAYS_FIRST_DRAWING'].replace(365243, np.nan, inplace= True)
prev['DAYS_FIRST_DUE'].replace(365243, np.nan, inplace= True)
prev['DAYS_LAST_DUE_1ST_VERSION'].replace(365243, np.nan, inplace= True)
prev['DAYS_LAST_DUE'].replace(365243, np.nan, inplace= True)
prev['DAYS_TERMINATION'].replace(365243, np.nan, inplace= True)
# + colab={"base_uri": "https://localhost:8080/"} id="oKkzGTKDfvEH" outputId="71a1783b-0f44-4c95-9179-09060d74079c"
prev['DAYS_LAST_DUE_DIFF'] = prev['DAYS_LAST_DUE_1ST_VERSION'] - prev['DAYS_LAST_DUE']
approved['DAYS_LAST_DUE_DIFF'] = approved['DAYS_LAST_DUE_1ST_VERSION'] - approved['DAYS_LAST_DUE']
# + id="S7__kCNdfvEH"
categorical_agg = {key: ['mean'] for key in categorical_cols}
# + colab={"base_uri": "https://localhost:8080/"} id="gMbsbdBxfvEH" outputId="bf292c17-f8c2-4a69-bec4-11d8b5897659"
agg_prev = group(prev, 'PREV_', {**PREVIOUS_AGG, **categorical_agg})
agg_prev = agg_prev.merge(active_agg_df, how='left', on='SK_ID_CURR')
del active_agg_df; gc.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="p1RNgm09fvEI" outputId="bd2d79bb-73e0-49d5-9faa-43abb227e2f4"
agg_prev = group_and_merge(approved, agg_prev, 'APPROVED_', PREVIOUS_APPROVED_AGG)
refused = prev[prev['NAME_CONTRACT_STATUS_Refused'] == 1]
agg_prev = group_and_merge(refused, agg_prev, 'REFUSED_', PREVIOUS_REFUSED_AGG)
del approved, refused; gc.collect()
# + id="O7qD6BA0fvEI"
for loan_type in ['Consumer loans', 'Cash loans']:
type_df = prev[prev['NAME_CONTRACT_TYPE_{}'.format(loan_type)] == 1]
prefix = 'PREV_' + loan_type.split(" ")[0] + '_'
agg_prev = group_and_merge(type_df, agg_prev, prefix, PREVIOUS_LOAN_TYPE_AGG)
del type_df; gc.collect()
# + id="CY0nCa6YfvEI"
pay['LATE_PAYMENT'] = pay['DAYS_ENTRY_PAYMENT'] - pay['DAYS_INSTALMENT']
pay['LATE_PAYMENT'] = pay['LATE_PAYMENT'].apply(lambda x: 1 if x > 0 else 0)
dpd_id = pay[pay['LATE_PAYMENT'] > 0]['SK_ID_PREV'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="fJEKqzojfvEI" outputId="a2b32f2a-44bf-4a04-e196-ca3e97edce1e"
agg_dpd = group_and_merge(prev[prev['SK_ID_PREV'].isin(dpd_id)], agg_prev,
'PREV_LATE_', PREVIOUS_LATE_PAYMENTS_AGG)
del agg_dpd, dpd_id; gc.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="KtR6hzvqfvEI" outputId="9642558c-8cb3-4f03-b787-1957145f5662"
for time_frame in [12, 24]:
time_frame_df = prev[prev['DAYS_DECISION'] >= -30*time_frame]
prefix = 'PREV_LAST{}M_'.format(time_frame)
agg_prev = group_and_merge(time_frame_df, agg_prev, prefix, PREVIOUS_TIME_AGG)
del time_frame_df; gc.collect()
del prev; gc.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="xoQyR7H0fvEI" outputId="20cbcc8f-6c6c-4605-a2a0-c8906cd9b778"
df = pd.merge(df, agg_prev, on='SK_ID_CURR', how='left')
del agg_prev; gc.collect()
# + id="DRPV_MkNfvEJ"
pos = pd.read_csv(os.path.join(DATA_DIRECTORY, 'POS_CASH_balance.csv'))
pos, categorical_cols = one_hot_encoder(pos, nan_as_category= False)
# + id="sUcHMMAPfvEJ"
pos['LATE_PAYMENT'] = pos['SK_DPD'].apply(lambda x: 1 if x > 0 else 0)
# + id="oaZhWd2pfvEJ"
POS_CASH_AGG = {
'SK_ID_PREV': ['nunique'],
'MONTHS_BALANCE': ['min', 'max', 'size'],
'SK_DPD': ['max', 'mean', 'sum', 'var'],
'SK_DPD_DEF': ['max', 'mean', 'sum'],
'LATE_PAYMENT': ['mean']
}
categorical_agg = {key: ['mean'] for key in categorical_cols}
pos_agg = group(pos, 'POS_', {**POS_CASH_AGG, **categorical_agg})
# + id="2W8LswKEfvEJ"
sort_pos = pos.sort_values(by=['SK_ID_PREV', 'MONTHS_BALANCE'])
gp = sort_pos.groupby('SK_ID_PREV')
temp = pd.DataFrame()
temp['SK_ID_CURR'] = gp['SK_ID_CURR'].first()
temp['MONTHS_BALANCE_MAX'] = gp['MONTHS_BALANCE'].max()
# + id="fGb58fVgfvEJ"
temp['POS_LOAN_COMPLETED_MEAN'] = gp['NAME_CONTRACT_STATUS_Completed'].mean()
temp['POS_COMPLETED_BEFORE_MEAN'] = gp['CNT_INSTALMENT'].first() - gp['CNT_INSTALMENT'].last()
temp['POS_COMPLETED_BEFORE_MEAN'] = temp.apply(lambda x: 1 if x['POS_COMPLETED_BEFORE_MEAN'] > 0
and x['POS_LOAN_COMPLETED_MEAN'] > 0 else 0, axis=1)
# + id="OO5yJzmDfvEJ"
temp['POS_REMAINING_INSTALMENTS'] = gp['CNT_INSTALMENT_FUTURE'].last()
temp['POS_REMAINING_INSTALMENTS_RATIO'] = gp['CNT_INSTALMENT_FUTURE'].last()/gp['CNT_INSTALMENT'].last()
# + colab={"base_uri": "https://localhost:8080/"} id="MEv7JqcJfvEK" outputId="841b5604-7aa1-494c-9e9a-ede7651b10e3"
temp_gp = temp.groupby('SK_ID_CURR').sum().reset_index()
temp_gp.drop(['MONTHS_BALANCE_MAX'], axis=1, inplace= True)
pos_agg = pd.merge(pos_agg, temp_gp, on= 'SK_ID_CURR', how= 'left')
del temp, gp, temp_gp, sort_pos; gc.collect()
# + id="cvF8ll7zfvEK"
pos = do_sum(pos, ['SK_ID_PREV'], 'LATE_PAYMENT', 'LATE_PAYMENT_SUM')
# + id="Oq4_EqNnfvEK"
last_month_df = pos.groupby('SK_ID_PREV')['MONTHS_BALANCE'].idxmax()
# + id="X7wY3nANfvEK"
sort_pos = pos.sort_values(by=['SK_ID_PREV', 'MONTHS_BALANCE'])
gp = sort_pos.iloc[last_month_df].groupby('SK_ID_CURR').tail(3)
gp_mean = gp.groupby('SK_ID_CURR').mean().reset_index()
pos_agg = pd.merge(pos_agg, gp_mean[['SK_ID_CURR','LATE_PAYMENT_SUM']], on='SK_ID_CURR', how='left')
# + id="_Pu9XXz7fvEK"
drop_features = [
'POS_NAME_CONTRACT_STATUS_Canceled_MEAN', 'POS_NAME_CONTRACT_STATUS_Amortized debt_MEAN',
'POS_NAME_CONTRACT_STATUS_XNA_MEAN']
pos_agg.drop(drop_features, axis=1, inplace=True)
# + id="wEBJeLzIfvEK"
df = pd.merge(df, pos_agg, on='SK_ID_CURR', how='left')
# + id="5Uquq6bIfvEL"
pay = do_sum(pay, ['SK_ID_PREV', 'NUM_INSTALMENT_NUMBER'], 'AMT_PAYMENT', 'AMT_PAYMENT_GROUPED')
pay['PAYMENT_DIFFERENCE'] = pay['AMT_INSTALMENT'] - pay['AMT_PAYMENT_GROUPED']
pay['PAYMENT_RATIO'] = pay['AMT_INSTALMENT'] / pay['AMT_PAYMENT_GROUPED']
pay['PAID_OVER_AMOUNT'] = pay['AMT_PAYMENT'] - pay['AMT_INSTALMENT']
pay['PAID_OVER'] = (pay['PAID_OVER_AMOUNT'] > 0).astype(int)
# + id="uu06JXUBfvEL"
pay['DPD'] = pay['DAYS_ENTRY_PAYMENT'] - pay['DAYS_INSTALMENT']
pay['DPD'] = pay['DPD'].apply(lambda x: 0 if x <= 0 else x)
pay['DBD'] = pay['DAYS_INSTALMENT'] - pay['DAYS_ENTRY_PAYMENT']
pay['DBD'] = pay['DBD'].apply(lambda x: 0 if x <= 0 else x)
# + id="Fnmt7K3ofvEL"
pay['LATE_PAYMENT'] = pay['DBD'].apply(lambda x: 1 if x > 0 else 0)
# + id="2ts1QqkkfvEL"
pay['INSTALMENT_PAYMENT_RATIO'] = pay['AMT_PAYMENT'] / pay['AMT_INSTALMENT']
pay['LATE_PAYMENT_RATIO'] = pay.apply(lambda x: x['INSTALMENT_PAYMENT_RATIO'] if x['LATE_PAYMENT'] == 1 else 0, axis=1)
# + id="cX7qT6dPfvEL"
pay['SIGNIFICANT_LATE_PAYMENT'] = pay['LATE_PAYMENT_RATIO'].apply(lambda x: 1 if x > 0.05 else 0)
# + id="IWWzMR8WfvEL"
pay['DPD_7'] = pay['DPD'].apply(lambda x: 1 if x >= 7 else 0)
pay['DPD_15'] = pay['DPD'].apply(lambda x: 1 if x >= 15 else 0)
# + id="PDOerU_bfvEM"
INSTALLMENTS_AGG = {
'SK_ID_PREV': ['size', 'nunique'],
'DAYS_ENTRY_PAYMENT': ['min', 'max', 'mean'],
'AMT_INSTALMENT': ['min', 'max', 'mean', 'sum'],
'AMT_PAYMENT': ['min', 'max', 'mean', 'sum'],
'DPD': ['max', 'mean', 'var'],
'DBD': ['max', 'mean', 'var'],
'PAYMENT_DIFFERENCE': ['mean'],
'PAYMENT_RATIO': ['mean'],
'LATE_PAYMENT': ['mean', 'sum'],
'SIGNIFICANT_LATE_PAYMENT': ['mean', 'sum'],
'LATE_PAYMENT_RATIO': ['mean'],
'DPD_7': ['mean'],
'DPD_15': ['mean'],
'PAID_OVER': ['mean']
}
pay_agg = group(pay, 'INS_', INSTALLMENTS_AGG)
# + id="4cxhtjXZfvEM"
INSTALLMENTS_TIME_AGG = {
'SK_ID_PREV': ['size'],
'DAYS_ENTRY_PAYMENT': ['min', 'max', 'mean'],
'AMT_INSTALMENT': ['min', 'max', 'mean', 'sum'],
'AMT_PAYMENT': ['min', 'max', 'mean', 'sum'],
'DPD': ['max', 'mean', 'var'],
'DBD': ['max', 'mean', 'var'],
'PAYMENT_DIFFERENCE': ['mean'],
'PAYMENT_RATIO': ['mean'],
'LATE_PAYMENT': ['mean'],
'SIGNIFICANT_LATE_PAYMENT': ['mean'],
'LATE_PAYMENT_RATIO': ['mean'],
'DPD_7': ['mean'],
'DPD_15': ['mean'],
}
for months in [36, 60]:
recent_prev_id = pay[pay['DAYS_INSTALMENT'] >= -30*months]['SK_ID_PREV'].unique()
pay_recent = pay[pay['SK_ID_PREV'].isin(recent_prev_id)]
prefix = 'INS_{}M_'.format(months)
pay_agg = group_and_merge(pay_recent, pay_agg, prefix, INSTALLMENTS_TIME_AGG)
# + id="jrgz3KSWfvEM"
def add_features_in_group(features, gr_, feature_name, aggs, prefix):
for agg in aggs:
if agg == 'sum':
features['{}{}_sum'.format(prefix, feature_name)] = gr_[feature_name].sum()
elif agg == 'mean':
features['{}{}_mean'.format(prefix, feature_name)] = gr_[feature_name].mean()
elif agg == 'max':
features['{}{}_max'.format(prefix, feature_name)] = gr_[feature_name].max()
elif agg == 'min':
features['{}{}_min'.format(prefix, feature_name)] = gr_[feature_name].min()
elif agg == 'std':
features['{}{}_std'.format(prefix, feature_name)] = gr_[feature_name].std()
elif agg == 'count':
features['{}{}_count'.format(prefix, feature_name)] = gr_[feature_name].count()
elif agg == 'skew':
features['{}{}_skew'.format(prefix, feature_name)] = skew(gr_[feature_name])
elif agg == 'kurt':
features['{}{}_kurt'.format(prefix, feature_name)] = kurtosis(gr_[feature_name])
elif agg == 'iqr':
features['{}{}_iqr'.format(prefix, feature_name)] = iqr(gr_[feature_name])
elif agg == 'median':
features['{}{}_median'.format(prefix, feature_name)] = gr_[feature_name].median()
return features
# + id="SC_YdOTnfvEM"
def chunk_groups(groupby_object, chunk_size):
n_groups = groupby_object.ngroups
group_chunk, index_chunk = [], []
for i, (index, df) in enumerate(groupby_object):
group_chunk.append(df)
index_chunk.append(index)
if (i + 1) % chunk_size == 0 or i + 1 == n_groups:
group_chunk_, index_chunk_ = group_chunk.copy(), index_chunk.copy()
group_chunk, index_chunk = [], []
yield index_chunk_, group_chunk_
# + id="6Qwkz1R6fvEM"
def add_trend_feature(features, gr, feature_name, prefix):
y = gr[feature_name].values
try:
x = np.arange(0, len(y)).reshape(-1, 1)
lr = LinearRegression()
lr.fit(x, y)
trend = lr.coef_[0]
except:
trend = np.nan
features['{}{}'.format(prefix, feature_name)] = trend
return features
# + id="9IAqfPACfvEN"
def parallel_apply(groups, func, index_name='Index', num_workers=0, chunk_size=100000):
if num_workers <= 0: num_workers = 8
#n_chunks = np.ceil(1.0 * groups.ngroups / chunk_size)
indeces, features = [], []
for index_chunk, groups_chunk in chunk_groups(groups, chunk_size):
with mp.pool.Pool(num_workers) as executor:
features_chunk = executor.map(func, groups_chunk)
features.extend(features_chunk)
indeces.extend(index_chunk)
features = pd.DataFrame(features)
features.index = indeces
features.index.name = index_name
return features
# + id="xsFKZmAlfvEN"
def trend_in_last_k_instalment_features(gr, periods):
gr_ = gr.copy()
gr_.sort_values(['DAYS_INSTALMENT'], ascending=False, inplace=True)
features = {}
for period in periods:
gr_period = gr_.iloc[:period]
features = add_trend_feature(features, gr_period, 'DPD',
'{}_TREND_'.format(period))
features = add_trend_feature(features, gr_period, 'PAID_OVER_AMOUNT',
'{}_TREND_'.format(period))
return features
group_features = ['SK_ID_CURR', 'SK_ID_PREV', 'DPD', 'LATE_PAYMENT',
'PAID_OVER_AMOUNT', 'PAID_OVER', 'DAYS_INSTALMENT']
gp = pay[group_features].groupby('SK_ID_CURR')
func = partial(trend_in_last_k_instalment_features, periods=[12, 24, 60, 120])
g = parallel_apply(gp, func, index_name='SK_ID_CURR', chunk_size=10000).reset_index()
pay_agg = pay_agg.merge(g, on='SK_ID_CURR', how='left')
# + id="ZFzMYZnRfvEN"
def installments_last_loan_features(gr):
gr_ = gr.copy()
gr_.sort_values(['DAYS_INSTALMENT'], ascending=False, inplace=True)
last_installment_id = gr_['SK_ID_PREV'].iloc[0]
gr_ = gr_[gr_['SK_ID_PREV'] == last_installment_id]
features = {}
features = add_features_in_group(features, gr_, 'DPD',
['sum', 'mean', 'max', 'std'],
'LAST_LOAN_')
features = add_features_in_group(features, gr_, 'LATE_PAYMENT',
['count', 'mean'],
'LAST_LOAN_')
features = add_features_in_group(features, gr_, 'PAID_OVER_AMOUNT',
['sum', 'mean', 'max', 'min', 'std'],
'LAST_LOAN_')
features = add_features_in_group(features, gr_, 'PAID_OVER',
['count', 'mean'],
'LAST_LOAN_')
return features
g = parallel_apply(gp, installments_last_loan_features, index_name='SK_ID_CURR', chunk_size=10000).reset_index()
pay_agg = pay_agg.merge(g, on='SK_ID_CURR', how='left')
# + id="VRas5FZafvEN"
df = pd.merge(df, pay_agg, on='SK_ID_CURR', how='left')
del pay_agg, gp, pay; gc.collect()
# + id="2e95uvEZfvEN"
cc = pd.read_csv(os.path.join(DATA_DIRECTORY, 'credit_card_balance.csv'))
cc, cat_cols = one_hot_encoder(cc, nan_as_category=False)
cc.rename(columns={'AMT_RECIVABLE': 'AMT_RECEIVABLE'}, inplace=True)
# + id="Zq77SAhGfvEN"
cc['LIMIT_USE'] = cc['AMT_BALANCE'] / cc['AMT_CREDIT_LIMIT_ACTUAL']
# + id="p9owpV45fvEO"
cc['PAYMENT_DIV_MIN'] = cc['AMT_PAYMENT_CURRENT'] / cc['AMT_INST_MIN_REGULARITY']
# + id="tQQZETBofvEO"
cc['LATE_PAYMENT'] = cc['SK_DPD'].apply(lambda x: 1 if x > 0 else 0)
# + id="OSjn_Ng_fvEO"
cc['DRAWING_LIMIT_RATIO'] = cc['AMT_DRAWINGS_ATM_CURRENT'] / cc['AMT_CREDIT_LIMIT_ACTUAL']
# + id="Q3lP62brfvEO"
CREDIT_CARD_AGG = {
'MONTHS_BALANCE': ['min'],
'AMT_BALANCE': ['max'],
'AMT_CREDIT_LIMIT_ACTUAL': ['max'],
'AMT_DRAWINGS_ATM_CURRENT': ['max', 'sum'],
'AMT_DRAWINGS_CURRENT': ['max', 'sum'],
'AMT_DRAWINGS_POS_CURRENT': ['max', 'sum'],
'AMT_INST_MIN_REGULARITY': ['max', 'mean'],
'AMT_PAYMENT_TOTAL_CURRENT': ['max', 'mean', 'sum', 'var'],
'AMT_TOTAL_RECEIVABLE': ['max', 'mean'],
'CNT_DRAWINGS_ATM_CURRENT': ['max', 'mean', 'sum'],
'CNT_DRAWINGS_CURRENT': ['max', 'mean', 'sum'],
'CNT_DRAWINGS_POS_CURRENT': ['mean'],
'SK_DPD': ['mean', 'max', 'sum'],
'SK_DPD_DEF': ['max', 'sum'],
'LIMIT_USE': ['max', 'mean'],
'PAYMENT_DIV_MIN': ['min', 'mean'],
'LATE_PAYMENT': ['max', 'sum'],
}
cc_agg = cc.groupby('SK_ID_CURR').agg(CREDIT_CARD_AGG)
cc_agg.columns = pd.Index(['CC_' + e[0] + "_" + e[1].upper() for e in cc_agg.columns.tolist()])
cc_agg.reset_index(inplace= True)
# + id="sYlkVrfCfvEO"
last_ids = cc.groupby('SK_ID_PREV')['MONTHS_BALANCE'].idxmax()
last_months_df = cc[cc.index.isin(last_ids)]
cc_agg = group_and_merge(last_months_df,cc_agg,'CC_LAST_', {'AMT_BALANCE': ['mean', 'max']})
# + id="xJ4dpQtofvEO"
CREDIT_CARD_TIME_AGG = {
'CNT_DRAWINGS_ATM_CURRENT': ['mean'],
'SK_DPD': ['max', 'sum'],
'AMT_BALANCE': ['mean', 'max'],
'LIMIT_USE': ['max', 'mean']
}
for months in [12, 24, 48]:
cc_prev_id = cc[cc['MONTHS_BALANCE'] >= -months]['SK_ID_PREV'].unique()
cc_recent = cc[cc['SK_ID_PREV'].isin(cc_prev_id)]
prefix = 'INS_{}M_'.format(months)
cc_agg = group_and_merge(cc_recent, cc_agg, prefix, CREDIT_CARD_TIME_AGG)
# + id="woq5rGjYfvEP"
df = pd.merge(df, cc_agg, on='SK_ID_CURR', how='left')
del cc, cc_agg; gc.collect()
# + id="5Vf2G4VXfvEP"
def add_ratios_features(df):
df['BUREAU_INCOME_CREDIT_RATIO'] = df['BUREAU_AMT_CREDIT_SUM_MEAN'] / df['AMT_INCOME_TOTAL']
df['BUREAU_ACTIVE_CREDIT_TO_INCOME_RATIO'] = df['BUREAU_ACTIVE_AMT_CREDIT_SUM_SUM'] / df['AMT_INCOME_TOTAL']
df['CURRENT_TO_APPROVED_CREDIT_MIN_RATIO'] = df['APPROVED_AMT_CREDIT_MIN'] / df['AMT_CREDIT']
df['CURRENT_TO_APPROVED_CREDIT_MAX_RATIO'] = df['APPROVED_AMT_CREDIT_MAX'] / df['AMT_CREDIT']
df['CURRENT_TO_APPROVED_CREDIT_MEAN_RATIO'] = df['APPROVED_AMT_CREDIT_MEAN'] / df['AMT_CREDIT']
df['CURRENT_TO_APPROVED_ANNUITY_MAX_RATIO'] = df['APPROVED_AMT_ANNUITY_MAX'] / df['AMT_ANNUITY']
df['CURRENT_TO_APPROVED_ANNUITY_MEAN_RATIO'] = df['APPROVED_AMT_ANNUITY_MEAN'] / df['AMT_ANNUITY']
df['PAYMENT_MIN_TO_ANNUITY_RATIO'] = df['INS_AMT_PAYMENT_MIN'] / df['AMT_ANNUITY']
df['PAYMENT_MAX_TO_ANNUITY_RATIO'] = df['INS_AMT_PAYMENT_MAX'] / df['AMT_ANNUITY']
df['PAYMENT_MEAN_TO_ANNUITY_RATIO'] = df['INS_AMT_PAYMENT_MEAN'] / df['AMT_ANNUITY']
df['CTA_CREDIT_TO_ANNUITY_MAX_RATIO'] = df['APPROVED_CREDIT_TO_ANNUITY_RATIO_MAX'] / df[
'CREDIT_TO_ANNUITY_RATIO']
df['CTA_CREDIT_TO_ANNUITY_MEAN_RATIO'] = df['APPROVED_CREDIT_TO_ANNUITY_RATIO_MEAN'] / df[
'CREDIT_TO_ANNUITY_RATIO']
df['DAYS_DECISION_MEAN_TO_BIRTH'] = df['APPROVED_DAYS_DECISION_MEAN'] / df['DAYS_BIRTH']
df['DAYS_CREDIT_MEAN_TO_BIRTH'] = df['BUREAU_DAYS_CREDIT_MEAN'] / df['DAYS_BIRTH']
df['DAYS_DECISION_MEAN_TO_EMPLOYED'] = df['APPROVED_DAYS_DECISION_MEAN'] / df['DAYS_EMPLOYED']
df['DAYS_CREDIT_MEAN_TO_EMPLOYED'] = df['BUREAU_DAYS_CREDIT_MEAN'] / df['DAYS_EMPLOYED']
return df
# + id="2ZVxuFbffvEP"
df = add_ratios_features(df)
# + id="8PLtBIz3fvEP"
df.replace([np.inf, -np.inf], np.nan, inplace=True)
# -
df.to_csv('data.csv', index=False)
| notebooks/CSE499B/data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # “吟诗作对”机器人
# 硬件准备:虚谷号、麦克风(连接到虚谷号)、小音箱(连接到虚谷号)
#
# 注意事项:
# - 本程序有两个文件组成,其中”start.mp3“为录音提示语。
# - 请逐步运行单元格中的代码,即可查看具体效果。
# - 如果使用USB小音箱,请务必在电源口接上2A的电源。
# - 请确认虚谷号处于上网状态。
#
# 本范例的具体介绍请参考百度AI的文档。https://ai.baidu.com/docs#/ASR-Online-Python-SDK/f71dfc54
# ### 准备工作1:导入基本库
import pyaudio
import wave
import os
from aip import AipSpeech
# ### 准备工作2:设置认证信息和参数
#
# 注:这里用的是测试账号,有访问次数的限制,如果测试失败,请使用自己的账号信息。
""" 你的 APPID AK SK """
APP_ID = "15126848"
API_KEY = "BPaS8KCk1B6Io9EqEOw1pOH3"
SECRET_KEY = "<KEY>"
""" 这里是参数设置,请不要改动 """
CHUNK = 1024
FORMAT = pyaudio.paInt16
CHANNELS = 1
RATE = 16000
RECORD_SECONDS = 3
client = AipSpeech(APP_ID, API_KEY, SECRET_KEY)
# 参数介绍:
# - CHUNK:指定每个缓冲区的帧数。
# - FORMAT:采样大小和格式。我们这里是pyaudio.paInt16,即16位int型。
# - CHANNELS:声道数,这里我们设定的是单声道。
# - RATE:采样频率,录音设备在一秒钟内对声音信号的采样次数,采样频率越高声音的还原就越真实越自然。这里是16000。这里是为了匹配后期语音识别的要求设置的。常用的有8kHz, 16kHz, 32kHz, 48kHz, 11.025kHz, 22.05kHz, 44.1kHz.
# - RECORD_SECONDS:录音秒数
# ### 核心函数1:
#
# rec函数的作用是录音处理。
def rec(file_name):
p = pyaudio.PyAudio()
stream = p.open(format=FORMAT,
channels=CHANNELS,
rate=RATE,
input=True,
frames_per_buffer=CHUNK)
print("开始录音,请说话...")
frames = []
for i in range(0, int(RATE / CHUNK * RECORD_SECONDS)):
data = stream.read(CHUNK)
frames.append(data)
print("录音结束!")
stream.stop_stream()
stream.close()
p.terminate()
wf = wave.open(file_name, 'wb')
wf.setnchannels(CHANNELS)
wf.setsampwidth(p.get_sample_size(FORMAT))
wf.setframerate(RATE)
wf.writeframes(b''.join(frames))
wf.close()
return file_name
# ### 核心函数2:
#
# audio_to_text函数的作用是将录音转化为文字。
def audio_to_text(wav_file):
with open(wav_file, 'rb') as fp:
file_context = fp.read()
print("正在识别语音...")
res = client.asr(file_context, 'wav', 16000, {
'dev_pid': 1536,
})
if (res["err_msg"]=="success."):
res_str = res.get("result")[0]
else:
res_str = "错误,没有识别出任何内容!"
return res_str
# ### 核心函数3:
#
# tts函数的作用是将文字result转为语音,并且朗读出来。
def tts(txt):
result = client.synthesis(txt, 'zh', 1, {'vol': 5,})
# 识别正确返回语音二进制 错误则返回dict 参照下面错误码
if not isinstance(result, dict):
with open('auido.mp3', 'wb') as f:
f.write(result)
os.system('play auido.mp3')
# ### 开始工作
#
# 描述:当用户说“床前明月光”,虚谷号会说“疑是地上霜”,如果接上音箱,将听到女声的朗读。
#添加诗词库,用户读前一句(奇数),机器对出下一句(偶数)
shici = ['床前明月光', '疑是地上霜', '举头望明月','低头思故乡']
shici.extend(['大漠沙如雪', '燕山月似钩', '何当金络脑','快走踏清秋'])
#可以用类似VB的语法,采用+的方式
shici=shici + ['海上生明月', '天涯共此时']
#输出列表
print(shici)
#播放提示语
os.system('play start.mp3')
#开始录音
ret_s = audio_to_text(rec("input.wav"))
print(ret_s)
for i in range(len(shici)//2):
#print(shici[i*2-1])
if shici[i*2] in ret_s:
back=shici[i*2+1]
break
else:
back="在下才疏学浅,甘拜下风。"
print(back)
#输出语音
tts(back)
# ### 拓展思考:
#
# 1.请给“诗词库”(变量shici)增加新的数据,使其可以支持更多的回答。
#
# 2.修改变量shici的内容,让这个程序可以在其他的特定场合工作。
#
| 课程汇集/虚谷号内置课程目录/9.人工智能综合应用/04.吟诗作对机器人/吟诗作对机器人.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#imports
import time
start_time = time.time()
import numpy as np
from matplotlib import pyplot as plt
from keras import Sequential
from keras.layers import Dense, Dropout, Conv1D, Flatten
from keras.metrics import binary_accuracy
#from keras.utils import np_utils
print("--- %s seconds ---" % (time.time() - start_time))
# +
# import datasets with time taken!
#smoll
start_time = time.time()
smoll = np.loadtxt("/home/willett/NeutrinoData/small_CNN_input_processed.txt", comments='#')
print("--- %s seconds ---" % (time.time() - start_time))
print(smoll.shape)
""" # commented out to save computation
#and the full
start_time = time.time()
fll = np.loadtxt("/home/willett/NeutrinoData/full_CNN_input_processed.txt", comments='#')
print("--- %s seconds ---" % (time.time() - start_time))
print(fll.shape)
"""
""" # commented out to save computation
#and the full
start_time = time.time()
fll = np.loadtxt("/home/willett/NeutrinoData/test_CNN_input_processed.txt", comments='#')
print("--- %s seconds ---" % (time.time() - start_time))
print(fll.shape)
"""
# extract title
pls = open("/home/willett/NeutrinoData/small_CNN_input_processed.txt", "r")
title = pls.readline()
title = title[2:-1]
print(title)
# +
# creating a dataset switch, change what UsedData is to change CNN
UD = smoll # Used Data = <dataset>
UDLength = UD.shape[0]
print("shape: ",UD.shape,"\nsize: ", UD.size," \nlength: ", UDLength)
# dataset is expected in this format:
# FirstLayer LastLayer NHits AverageZP Thrust PID_Angle PID_Front PID_LLR_M FirstLayer LastLayer NHits_Low AverageZP Thrust_Lo PID_Angle PID_Front PID_LLR_M Energy_As Angle_Bet Distance_Bet Sig Bg
# with Sig and Bg expected as one hot vectors.
# +
# splitting X = dataset , Y = one hot vectors
X = UD[:,0:-2]
Y = UD[:,-2:1000]
print("X shape: ",X.shape,"\nY shape: ", Y.shape)
# they will be split into testing and training at compile
# +
# this is a convolutional network so data must be spacially relevant: i.e. columns must be swapped.
# Convolution kernel size = (2,)
#swapping PID angle and PID front for high energy so two charge related variables in one convolution
PIDAH = X[:,5]
PIDFH = X[:,6]
X[:,5:7] = np.column_stack((PIDFH,PIDAH))
#swapping PID angle and PID front for low energy so two charge related variables in one convolution
PIDAL = X[:,13]
PIDFL = X[:,14]
X[:,13:15] = np.column_stack((PIDFL,PIDAL))
#swapping Energy Asymetry and Distance so two geometric related variables in one convolution
# While simultaneously padding with zeros!
EAS = X[:,16]
DB = X[:,-1]
print(EAS[0],DB[0])
X2 = np.zeros((UDLength,20))
X2[:,0:-1] = X
X2[:,16] = DB
X2[:,18] = EAS
#To debug print X before and X2 after, see if they swap
# -
X2 = np.expand_dims(X2, axis=2) # i.e. reshape (569, 30) to (569, 30, 1) for convolution
# +
# inevitable bias removal...
# +
# set variables:
InDim = (X2.shape[1],X2.shape[2]) #input dimension
Fltr = 4 #dimensionality of output space
KD = 2 # kernel size
Width = 8 # width of dense layers ~ 0.75 input
DR = 0.5 # rate of dropout
# linear model with a convolutional and 3 dense layers.
Model = Sequential()
Model.add(Conv1D(Fltr, KD , input_shape=InDim , activation="sigmoid", use_bias=True )) #conv
Model.add(Flatten())
Model.add(Dense(Width, activation="sigmoid", use_bias=True)) #1
Model.add(Dropout(DR) )
Model.add(Dense(Width, activation="sigmoid", use_bias=True)) #2
Model.add(Dropout(DR) )
Model.add(Dense(Width, activation="sigmoid", use_bias=True)) #3
Model.add(Dropout(DR) )
Model.add(Dense(2, activation="softmax", use_bias=True)) # output layer
# +
# compile model:
# For a binary classification problem
Model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy', 'binary_accuracy' ])
# +
# Trainining
# Train the model, iterating on the data in batches of 32 samples
history = Model.fit(X2, # the dataset
Y, #true or false values for the dataset
epochs=100, #number of iteration over data
batch_size=32, #number of trainings between tests
verbose=1, #prints one line per epoch of progress bar
validation_split=0.2 ) #ratio of test to train
# -
#summarise history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + active=""
# from sklearn.metrics import confusion_matrix
# import itertools
#
#
# def plot_confusion_matrix(cm, classes,
# normalize=False,
# title='Confusion matrix',
# cmap=plt.cm.Blues):
# """
# This function prints and plots the confusion matrix.
# Normalization can be applied by setting `normalize=True`.
# """
# if normalize:
# cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
# print("Normalized confusion matrix")
# else:
# print('Confusion matrix, without normalization')
#
# print(cm)
#
# plt.imshow(cm, interpolation='nearest', cmap=cmap)
# plt.title(title)
# plt.colorbar()
# tick_marks = np.arange(len(classes))
# plt.xticks(tick_marks, classes, rotation=45)
# plt.yticks(tick_marks, classes)
#
# fmt = '.2f' if normalize else 'd'
# thresh = cm.max() / 2.
# for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
# plt.text(j, i, format(cm[i, j], fmt),
# horizontalalignment="center",
# color="white" if cm[i, j] > thresh else "black")
#
# plt.tight_layout()
# plt.ylabel('True label')
# plt.xlabel('Predicted label')
#
#
# y_prob=model.predict(X_test)
# y_pred = y_prob.argmax(axis=-1)
# y_test_labels = y_test.argmax(axis=-1)
# cnf_matrix=confusion_matrix(y_test_labels, y_pred)
# class_names = ['No hurricane','hurricane']
# plot_confusion_matrix(cnf_matrix, classes=class_names,
# title='Confusion matrix, without normalization')
#
# -
Y.sum(0)[0] / Y.sum(0)[1] = SNRatio
| ANN/CNN1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pywt
from matplotlib import pyplot as plt
import matplotlib.image as mpimg
from pywt._doc_utils import wavedec2_keys, draw_2d_wp_basis
x=mpimg.imread('/home/karina/images.jpeg')
shape = x.shape
max_lev = 2
label_levels = 2
fig, axes = plt.subplots(2, 3, figsize=[14, 8])
for level in range(0, max_lev + 1):
if level == 0:
axes[0, 0].set_axis_off()
axes[1, 0].imshow(x, cmap=plt.cm.gray)
axes[1, 0].set_title('Image')
axes[1, 0].set_axis_off()
continue
draw_2d_wp_basis(shape, wavedec2_keys(level), ax=axes[0, level],
label_levels=label_levels)
axes[0, level].set_title('{} level\ndecomposition'.format(level))
c = pywt.wavedec2(x, 'db2', mode='periodization', level=level)
c[0] /= np.abs(c[0]).max()
for detail_level in range(level):
c[detail_level + 1] = [d/np.abs(d).max() for d in c[detail_level + 1]]
arr, slices = pywt.coeffs_to_array(c)
axes[1, level].imshow(arr, cmap=plt.cm.gray)
axes[1, level].set_title('Coefficients\n({} level)'.format(level))
axes[1, level].set_axis_off()
plt.tight_layout()
plt.show()
# -
| Groups/Group_ID_42/wavelet/transform/transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Qw1_7TM6n0o9"
# <h1><center><font color='Red'><b>Team 12's attempt to help the Child Mind Institute</center></b></font></h1>
# + [markdown] id="jkiJNkoJpcnW"
# <font color='blue'><b> In this notebook, we will be explaining our thought process behind what we did so that we could help the Child Mind Institute. We will also mention the challenges we faced and the potential drawbacks with our approaches. We found it better to present our work through jupyter and markdown instead of an app or dashboard since our work does not involve much of data and output visualization. We did think about visualization and wherever we have some ideas about how we could visualize, we will mention those.</b></font>
# + [markdown] id="XEMvrstPe1GU"
# *We have used some html to style our markdown. It somehow doesn't show it on Github, but it should be seen when it is opened as a jupyter notebook.*
# + [markdown] id="h_jCVbhTtYye"
# <font color='purple'><b>The data provided by the Child Mind Institue *consisted* of:<br>
# 1) a folder with docx files for differnt focus groups. The contents of the docx files were mainly dialogues between Moderators and Parents where moderators asked questions based on surveys filled by the Parents pertaining to how lockdown and Technology are affecting their children.<br>
# 2) some csv files based on inputs from Prolific Academic.<br>
# 3) a crisis logger csv file.</b></font>
# + [markdown] id="mGQiLoQBvoKA"
# <font color='purple'><b>We started with the docx files because we found those to contain the maximum unstructured text from which we could draw insights. We started with reading a few documents to understand what we need from them. After thinking hard, we realized the best insights can be drawn from text summarization, but for that, we also need to provide the relevant text from the documents. We realized the dialogues from the Parents will be most useful for the Child Mind Institute to understand the Parents'and their children's Pshycology. Based on our thinking, we realized our task can be broadly broken down into:</b></font><br>
#
# <font color='purple'><b> 1) Extracting relevent text (processing the documents)</b></font><br>
# <font color='purple'><b> 2) passing the extracted text to a summarizer function.</b></font>
# + [markdown] id="U5Cq4aqQzP47"
# **The first step was to create the right environment for our tasks, which meant cloning the forking and ML-for-good repository, installing the required packages. We realized the requirement of certain packages as we progressed with out tasks. There are some packages that were not used as well in the final code Below cell shows it all**
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="sXAJvYMP0WWT" outputId="ad01654b-1a59-4744-f3f9-bff1c9f07dcc"
# #!rm -r ML-for-Good-Hackathon
# !git clone https://github.com/morganstanley/ML-for-Good-Hackathon.git
# !pip install textract
# !pip install rake-nltk
# !pip install sentence_transformers
# !pip install bert-extractive-summarizer
# !pip install python-docx
# !pip install vaderSentiment
import pandas as pd
import os
import textract
import re
import gensim
from gensim.summarization.summarizer import summarize
from gensim.summarization import keywords
from rake_nltk import Rake
import nltk
nltk.download('stopwords')
nltk.download('punkt')
from sentence_transformers import SentenceTransformer
#sbert_model = SentenceTransformer('bert-base-nli-mean-tokens')
import tensorflow as tf
import tensorflow_hub
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from summarizer import Summarizer
import docx
from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
# + [markdown] id="0wtyqqstygO0"
# <h2><center><font color='Blue'><b>Document Preprocessing and extracting relevant text</center></b></font></h2>
# + [markdown] id="zhztdFm89UPA"
# <font color='purple'><b>As mentioned already, we realized that the most relevent text is Parents'dialogues. Just extracting the Parents'dialogues involved several steps, like removing certain types of strings, collecting a list of all diaglogues and joining dialogues of individual speakers. <br>
#
# Broadly, the steps are <br>
# 1) Remove irrelevant strings like (silence, [inaudible], next line (\n), extra tabs (\t) etc.) <br>
# 2) We collect each dialogue from the speakers: Parents, Moderators, Speaker, Adminitrator.<br>
# 3) We ultimately want only the Parents'dialogues to get more insights on the impact of lockdown and tech on their children.<br>
# 4) All the dialogues of a Parent are combined for further analysis.<br>
# 5) The function returns a dictionary with keys as Parents and the values as their combined dialogue from the entire doc.
# </b></font>
#
# <font color='purple'><b>An important observation from the results we saw in doc_preprocessing is that it was not very trivial to get absolutely clean text, so it is possible that some times the resulting text has some not so relevant strings. </b></font>
# + id="AKau_ixF-7J_"
def doc_preprocessing(path):
"""
Code for preprocessing docx files under focusGroups. The steps involved are:
1) Remove irrelevant strings like (silence, [inaudible], next line (\n), extra tabs (\t) etc.)
2) We collect each dialogue from the speakers: Parents, Moderators, Speaker, Adminitrator.
3) We ultimately want only the Parents'dialogues to get more insights on the impact of lockdown and tech on their children.
4) All the dialogues of a Parent are combined for further analysis.
5) The function returns a dictionary with keys as Parents and the values as their combined dialogue from the entire doc.
"""
# reading the docx file with textract
text = textract.process(path)
#setting utf-8 for decoding the text since the file read is in bytes
text = text.decode('utf-8')
#removing certain substrings like \n, \t, (silence)
text =text.replace('\n',"")
text = text.replace('\t',"")
text = text.replace('(silence)',"")
text = text.replace('. ','.')
#splitting sentences by full stop (.) and question mark (?)
split_sent=re.split(r'[?.]', text)
#removing empty strings from the list of sentences
split_sent = list(filter(('').__ne__, split_sent))
#remove irrelevant sentences like [inaudible 00:13:35]
split_sent = [sent for sent in split_sent if not re.search("inaudible",sent)]
#the next two lines are for collecting all dialogues by each speaker
dial_characters = ('Administrator','Moderator','Parent','Speaker',' Administrator',' Moderator',' Parent',' Speaker')
dial_start=[ind for ind,l in enumerate(split_sent) if l.startswith(dial_characters)]
#next, we combine dialogues for each speaker till another speaker starts a dialogue.
list_of_dial = []
for i in range(len(dial_start)-1):
list_of_dial.append('.'.join(split_sent[dial_start[i]:dial_start[i+1]]))
#we create a list of only the dialogues of Parents
list_of_Parent_dial = [sent for sent in list_of_dial if sent.startswith('Parent')]
# we get the index for each parent present in a session
num_ind = [sent.split(" ")[1].split(':')[0] for sent in list_of_Parent_dial]
parent_num = list(set([int(word) for sent in num_ind for word in sent.split() if word.isdigit()]))
# we combine all that a Parent has said in one dialogue of that Parent
all_dial_comb = []
for i in parent_num:
all_dial_comb.append('.'.join([sent for sent in list_of_Parent_dial if sent.startswith(('Parent '+str(i)+':',' Parent '+str(i)+':'))]))
# list with slight processing of each Parent dialogue
all_d_comb_n=[]
for i in range(len(all_dial_comb)):
all_d_comb_n.append(all_dial_comb[i].replace('Parent '+str(i+1)+":","").replace(' Parent '+str(i+1)+":",""))
# dictionary with keys as Parent names (eg: Parent 1) and the values as their combined dialogue.
dial_dict = dict(zip(['Parent '+str(i) for i in parent_num],all_d_comb_n))
# list processing of the values in the dict and return the dict.
for key in list(dial_dict.keys()):
dial_dict[key] = dial_dict.get(key).replace("My name's Parent",'').replace('Parent','').replace('Moderator','').replace('Administrator','')
#dial_dict[key] = re.sub('[0-9]','',dial_dict.get(key))
return dial_dict
# + [markdown] id="HsDYZkGDBhSg"
# **A quick execution of the doc_preprocessing function above can be seen below. We just chose a single file for demo, we will see later that we automatically do it for all the files and save the output in doc files, in a loop.**
# + colab={"base_uri": "https://localhost:8080/"} id="hCpQtwhO_BZO" outputId="c7cc5548-aa49-4cd8-c02c-cde4f382c2ec"
doc_preprocessing('./ML-for-Good-Hackathon/Data/FocusGroups/Media_Group4.docx')
# + [markdown] id="k4zjBztqDpTz"
# <font color='purple'><b> After getting the relevant text from the focus group documents, we needed to get insights from this relevant text. We realized the best solution would be to perform summarization. </b></font><br>
#
# <font color='purple'><b> There are two types of text summarization in NLP: Abstractive and Extractive. We were excited to try abstractive text summarization which would summarize by understanding the language context. Abstractive summarization uses the variants of the well known Neural Network architecture 'Transformers', but we weren't getting very appropriate results using that.</b></font> <br>
#
# <font color='purple'><b> Our conclusion is that it didn't work because the language is the documents isn't very clean like how it is in well written articles, but it is in the form of dialogues.</b></font> <br>
#
# <font color='purple'><b> We finally stuck to extractive text summarization.</b></font> <br>
#
# <font color='purple'><b> We were also trying key phrase extraction, but it didn't give very meaningful results, we think that it's because the documents don't contain clean language. </b></font> <br>
#
# <font color='purple'><b> For extractive text summarization, one could either summarization by percentage of text we want from the original text or by word count or in some cases, by number of sentences. We used two extractive text summarizers, one from the Gensim package (with percentage specification and word count specification) and other using the Bert extractive summarizer(with percentage specification and number of sentences specification). We realized some summarizers did better in some cases and others did better in the rest. Therefore, the purpose to use multiple summarizers is that the people from Child Mind Institute can collectively get insights from each type of summarizer. It is also to be noted that in some documents, some parents hardly had sufficient dialogue and the Gensim summarizer particularly gave a warning that there aren't relevant sentences or enough sentences. Below is the function for extractive text summarization.</b></font>
#
# <font color='purple'><b> We start with creating a Bert summarizer object as that loads the Bert model</b></font>
# + colab={"base_uri": "https://localhost:8080/", "height": 249, "referenced_widgets": ["1ae7b065d0724868b61916765ce189bb", "<KEY>", "c4be99dd645041669d4e5bbed34a17ca", "<KEY>", "<KEY>", "<KEY>", "fd43e5c9e4cd4ad78cfeb191eaf0db59", "a0b9196b15a7439b8d9ddd556ed42e78", "bb9f8b29e8f442a897be24e8028a4360", "<KEY>", "<KEY>", "<KEY>", "2f78b9d90079423487c3ad6517ce55e6", "bb35851106e04a6dacb05d77ecc6bab0", "f397e096b8d74683990f7ea4ced93d97", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "08e02e7a295b4e46a70ed969b6374d30", "c9e5553d4950432c8492a3d466cb2ff4", "33641fa5e4184a3895641469ce58afd6", "cb929a1f7f474e1d8f9c164b5eb23a20", "<KEY>", "<KEY>", "d1136f06d8c04e5a80bb6fe8d920d79d", "648356a10c904868b02a09f1083ea3ed", "534f31ff183d4983acaa48b40e927c1a", "96e89724fced4835aa9376f7ccb53967", "<KEY>", "b589607adade45d7aa6baddc471a33da", "<KEY>", "c4751cc6f5a6486dad167974eefb888d", "65775788330841b09901beacb81b9170", "90013e2d48db4a028d34087e3a5592f0", "<KEY>", "<KEY>", "<KEY>", "71bbad3f37ee46d385d88e39e7aac437", "f2ece0dd70c546d7b8eedde6242b10dd", "7f2e0b2226354a5ead19caa44b52c977", "82abb758df2b49af9c54a90e0ff03ffa", "<KEY>", "78761a9b7b824055b291c62e8c37e951", "b2a114269d5e4855a925a5fa948dde0e", "<KEY>", "fb13ec67ca6c49eb9ac56ab240d8bd8c", "6d323d42f08c4ec18648cc546feb5074", "80995ff4ef964b8e825d83726590939e", "8077c7fab94e4eeaa93dad809b2f00eb", "f956f3e7333e4893be05037acd5930db", "<KEY>", "<KEY>", "7ac3370ca42845b0aeed2d5ca40a5fe8"]} id="VJzHMJbZLhN1" outputId="3960622b-9488-4655-ed47-5934a30fb3a4"
#Creating the Bert Summarizer object
model = Summarizer()
# + id="rFSuOBVwE-4o"
def extractive_text_summarization(path):
"""
This function uses the processed data from doc_preprocessing function. We perform an extractive text summarization for each Parents dialogues.
We have used the summarizer from gensim package and we also use the Bert summarizer. With these packages, we can extract important text based on word
count, percentage, number of sentences. We had tried key phrase extraction which didn't work well.
"""
#first run the doc_preprocessing function for the file in the path
imp_dial_parents = doc_preprocessing(path)
# Summary (0.5% of the original content): Gensim
summ_per = [summarize(dialogue.replace(".",". "), ratio = 0.1) for dialogue in list(imp_dial_parents.values())]
# Summary (200 words): Gensim
summ_words = [summarize(dialogue.replace(".",". "), word_count = 200) for dialogue in list(imp_dial_parents.values())]
#creating dictionaries with keys as Parents (eg:Parent 1) and values as their summarized dialogues
keys = list(imp_dial_parents.keys())
summ_per = dict(zip(keys,summ_per))
summ_words = dict(zip(keys,summ_words))
# Using Bert extractive summarizer with ratio specification
result1 = [model(dialogue, ratio=0.2) for dialogue in list(imp_dial_parents.values())]
# Using Bert extractive summarizer with number of sentences specification
result2 = [model(dialogue, num_sentences=6) for dialogue in list(imp_dial_parents.values())]
#creating dictionaries with keys as Parents (eg:Parent 1) and values as their summarized dialogues
bert_ratio = dict(zip(keys,result1))
bert_num_sent = dict(zip(keys,result2))
# Ingnore the below commented code. This was an attempt to extract key phrases, but we didn't get great results. We have still kept it since we tried it.
# r = Rake()
# key_ph_list = []
# for i in range(len(keys)):
# r.extract_keywords_from_text(list(imp_dial_parents.values())[i])
# key_ph_list.append(r.get_ranked_phrases()[:30])
# key_phrases = dict(zip(keys,key_ph_list))
#return all the dictionaries with keys as Parents and values as extractive summaries for different types of summarizers
return summ_per, summ_words, bert_ratio, bert_num_sent
# + [markdown] id="-Tb47pbILFHL"
# **A quick execution of the extractive_text_summarization function above can be seen below. We just chose a single file for demo, we will see later that we automatically do it for all the files along with the doc_preprocessing function and save the output in doc files, in a loop. Note that the first two summarizations is from Gensim (with word count and ratio/percentage specifications respectively) and the last two are from Bert (with word count and number of sentences specifications respectively). We take care of this detail in the final execution of the loop where we process and perform this function on all doc files.**
# + colab={"base_uri": "https://localhost:8080/"} id="j0b9DOb2LStt" outputId="fc274afd-23f2-4529-a271-f66f5afe161b"
extractive_text_summarization('./ML-for-Good-Hackathon/Data/FocusGroups/Media_Group4.docx')
# + [markdown] id="K1fc08BaNS01"
# **The below code executes doc_preprocessing and extractive text summarization for all the doc files. We also save the results in doc files.**
# + id="t2peSz8ONjKy"
list_of_docs = ['./ML-for-Good-Hackathon/Data/FocusGroups/'+file for file in os.listdir('./ML-for-Good-Hackathon/Data/FocusGroups')]
#Executing the extractive_text_summarization function and saving the outputs
para_topics = ['gensim_summ_ratio', 'gensim_summ_words', 'bert_summ_ratio', 'bert_summ_num_sent']
for doc in list_of_docs:
extract = extractive_text_summarization(doc)
mydoc = docx.Document()
for i in range(len(extract)):
mydoc.add_paragraph(para_topics[i]+':')
for key in extract[i].keys():
mydoc.add_paragraph(key+': '+extract[i].get(key))
mydoc.save('summ_outputs_'+doc.split('/')[-1])
# + [markdown] id="a-N_iHojYphC"
# <font color='purple'><b> A possible way to visualize outputs from the summarizer is by creating some kind of an app where we just upload a document and it hightlights the extracted text in the document itself. This could be very beneficial for the Child Mind Institue since they would be looking at the summarized extracted text as part of the document itself and if they want, they could look at the text around the highlighted text to get more context, this is still work in progress.</b></font><br>
# + [markdown] id="tJAlVCteOX5P"
# <h2><center><font color='Blue'><b>Processing data from Prolific Academic and drawing insights from it</center></b></font></h2>
# + [markdown] id="1joCXx9cOzjE"
# <font color='purple'><b> When we saw the Prolific Academic csv files, we realized that there are too many columns and we started wondering what insights could we possibly draw from such a file. We started looking at a group of columns and found a feature called "suspectedinfected" in the Nov 2020 and April 2021 files and felt that that could be considered a target feature in some classification type model. We also noticed several columns that had too many missing values and felt that those might not give much insights. We also found a bunch of features with sentences abd started thinking about how we could represent them if we were to create a model. After thinking hard, below are the steps we followed to draw some insights. </b></font><br>
#
# <font color='purple'><b> We followed each of these steps individually for Adult and Parent data from Nov 2020 and April 2021, and did not combine the data because we felt that a more granular analysis might also give more insights. We later received an email about an update in the Prolific academic data, so we further looked at the features to see if there were any changes. We realized that the same approach can be used even on the updated data. Our outputs are finally created for each file even from the updated Nov 2020 and April 2021 data.</b></font><br>
#
# <font color='purple'><b>
# 1) remove features with more than 50% NaNs.<br>
# 2) find the top 19 or 20 features that are most correlated to suspectedidentified among each of int and float features.<br>
# 3) replace NaNs in the string features with 'Information not available'.<br>
# 4) combine dataframes from 2) and 3).<br>
# 5) remove all rows with NaNs.<br>
# 6) set target as suspectedidentified. <br>
# 7) for the string features, get embeddings for each using the Universal sentence encoder pretrained model from tensorflow hub.<br>
# 8) For each of the string features, step 7) would give us embeddings of dimension 571. We perform PCA on each to reduce the dimension to 5 for each.<br>
# 9) In 8), the new features as embeddings are given names as feature_name followed by an index.<br>
# 10) By step 9), we are done with preprocessing. We then fit a random forest classifier to arrive at the important features. This might help Child Mind
# Institute to focus on a fewer features. In case of embeddings, the features for which some components of the embeddings are important can be focussed on.
# </b></font><br>
#
# <font color='purple'><b> An important point to note is that our RF classifier is not created here for the purpose of prediction, we mainly want to assess feature importance given the data. Having said that, if we get a data for a quarter with exactly the same set of features as in the any of the csv files above, we can try prediction as well </b></font><br>
#
# <font color='purple'><b> Below is the code demonstrating each step above. It is to be noted that above works when we can set 'suspectedinfected' as the target. We noticed that for the files from quarters or months before Nov 2020, 'suspectedinfected' was not a feature. Depending on how future turns out after Covid, the feature may or may not be there. In those situations, we would again have to look at all features and see if something can be considered a target feature. If we don't find anything that can be used as an appropriate target, we may have to look for different ways to draw insights.</b></font><br>
#
# <font color='purple'><b> We start with loading the Universal Sentence Encoder pretrained model. We set the parameter in the KerasLayer from tensorflow hub as False since we will not be using these embeddings for fine tuning or feature extraction.</b></font><br>
# + colab={"base_uri": "https://localhost:8080/"} id="MWGrqVTtVAIH" outputId="b76570cf-2f9b-48f3-a69a-da966d8cbe43"
uni_encoder = tensorflow_hub.KerasLayer('https://tfhub.dev/google/universal-sentence-encoder/4',trainable=False)
# + id="_SLaD2FwO0f0"
def preprocess_prolificacademic_and_feature_importances(path):
"""
only works for Nov 2020 and April 2021 data since the data before Nov 2020 doesn't contain the variable suspectedinfected. We set the target
feature as suspectedidentified and fit a random forest classifier on the processed dataset. Note that the purpose of the classifier is not to
predict because we realize that the features in the data need not be same always. The purpose is to just suggest the more important features in
the processed dataset that can. The preprocessing steps are the following:
1) remove features with more than 50% NaNs.
2) find the top 19 or 20 features that are most correlated to suspectedidentified among each of int and float features.
3) replace NaNs in the string features with 'Information not available'.
4) combine dataframes from 2) and 3).
5) remove all rows with NaNs.
6) set target as suspectedidentified.
7) for the string features, get embeddings for each using the Universal sentence encoder pretrained model from tensorflow hub.
8) For each of the string features, step 7) would give us embeddings of dimension 571. We perform PCA on each to reduce the dimension to 5 for each.
9) In 8), the new features as embeddings are given names as feature_name followed by an index.
10) By step 9), we are done with preprocessing. We then fit a random forest classifier to arrive at the important features. This might help Child Mind
Institute to focus on a fewer features. In case of embeddings, the features for which some components of the embeddings are important can be focussed on.
"""
data2 = pd.read_csv(path)
#remove features with >50% NaNs and also timestamps and ID columns
to_be_removed = list((data2.isnull().sum() * 100 / len(data2) >50)[data2.isnull().sum() * 100 / len(data2) >50].index)
time_col = [col for col in data2.columns if col.startswith('timestamp')]
to_be_removed = to_be_removed+['ID']+time_col
data2.drop(to_be_removed, axis=1, inplace=True)
#finding and grouping features by their dtype. There are some of type int64, some of type float64 and some string features (dtype: object)
g = data2.columns.to_series().groupby(data2.dtypes).groups
g_dic = {k.name: v for k, v in g.items()}
#for all the float features, find the top 20 most correlated with the feature 'suspectedinfected'and create a subset of the dataframe with only these features
float_f = list(g_dic.get('float64'))
f_var = data2[float_f]
f_var_cor = f_var.corr()
f_var_list = list(f_var_cor.suspectedinfected.sort_values(ascending=False)[1:20].index)
print(f_var_list)
f_var = f_var[f_var_list]
#for all the int features, find the top 20 most correlated with the feature 'suspectedinfected'and create a subset of the dataframe with only these features
int_f = list(g_dic.get('int64'))
int_var = data2[int_f+['suspectedinfected']]
i_var_corr = int_var.corr()
i_var_list = list(i_var_corr.suspectedinfected.sort_values(ascending=False)[0:20].index)
print(i_var_list)
int_var = int_var[i_var_list]
#for all string features, replace NaN by 'Information not available'
var_to_preprocess = list(g_dic.get('object'))
string_f = data2[var_to_preprocess]
string_f=string_f.fillna('information not available')
#create a combined dataframe with the 20 float, 20 int and the string features
combined_df = pd.concat([f_var, int_var,string_f], axis=1)
print(f"'% of rows usable for training':{len(combined_df.dropna())/len(combined_df)}")
#drop rows with NaNs. The above line prints the % of rows that remain
combined_df.dropna(inplace=True)
#set target as suspectedinfected
target = combined_df.pop('suspectedinfected')
print(len(combined_df))
#for each string feature in combined_df, get embeddings with the pretrained Universal Sentence Encoder model and then perform PCA to reduce dimension of embeddings of each feature to 5
for feat in var_to_preprocess:
a=uni_encoder(list(combined_df[feat]))
principal=PCA(n_components=5)
principal.fit(a)
x=principal.transform(a)
y = pd.DataFrame(x,columns=[feat+str(i) for i in range(1,6)],index=combined_df.index)
combined_df = pd.concat([combined_df,y],axis=1)
#drop the original string features
combined_df.drop(var_to_preprocess,axis=1,inplace=True)
#fit a Random Forest Classifier with X=combined_df and Y=target(suspectedinfected)
clf = RandomForestClassifier(n_estimators=500,max_depth=10,random_state=42)
clf.fit(combined_df,target)
#get feature importances as dict and then, convert to a dataframe, return the top 50 important features.
feats = {} # a dict to hold feature_name: feature_importance
for feature, importance in zip(combined_df.columns, clf.feature_importances_):
feats[feature] = importance #add the name/value pair
importances = pd.DataFrame.from_dict(feats, orient='index').rename(columns={0: 'Gini-importance'})
importances = importances.sort_values(by='Gini-importance',ascending=False)
#importances.head(50)
return importances.head(50)
# + [markdown] id="wccNRqwyT_Ti"
# **A quick execution of the preprocess_prolificacademic_and_feature_importances function above can be seen below. We just chose a single file for demo, we will see later that we automatically do it for all the files in a loop.**
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="hH_0gZmoURRp" outputId="a847ade7-f95b-4412-ce6a-103361ef3bd5"
preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/updated_data/November/CRISIS_Adult_November_2020.csv')
# + [markdown] id="Gpfq9IHsWImZ"
# **The below code executes preprocess_prolificacademic_and_feature_importances for all the prolific academic csv files (old as well as updated ones from Nov 2020 and April 2021). We also save the results in a single xlsx file with multiple sheets, each containing results from a single input csv file and named accordingly.**
# + id="U-Yjk6v0Wtw8"
April2021_adult_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/April 2021/Data/CRISIS_Adult_April_2021.csv')
Nov2020_adult_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/November 2020/Data/CRISIS_Adult_November_2020.csv')
April2021_parent_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/April 2021/Data/CRISIS_Parent_April_2021.csv')
Nov2020_parent_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/November 2020/Data/CRISIS_Parent_November_2020.csv')
updated_April2021_adult_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/updated_data/April_21/CRISIS_Adult_April_2021.csv')
updated_April2021_parent_feat_imp=preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/updated_data/April_21/CRISIS_Parent_April_2021.csv')
updated_Nov2020_adult_feat_imp = preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/updated_data/November/CRISIS_Adult_November_2020.csv')
updated_Nov2020_parent_feat_imp = preprocess_prolificacademic_and_feature_importances('./ML-for-Good-Hackathon/Data/ProlificAcademic/updated_data/November/CRISIS_Parent_November_2020.csv')
#saving results from preprocess_prolificacademic_and_feature_importances in a single xlsx file
def save_xls(list_dfs, xls_path,required_dfs):
with pd.ExcelWriter(xls_path) as writer:
for n, df in enumerate(list_dfs):
df.to_excel(writer,required_dfs[n])
writer.save()
xls_path = '/content/prolificacademic_feature_imp.xlsx'
#getting names of the dataframe created above as strings
required_dfs = [df for df in globals() if 'feat_imp' in df]
#creating the list of dataframes in the same order as their names as string are listed in required_dfs
list_dfs = [April2021_adult_feat_imp,
Nov2020_adult_feat_imp,
April2021_parent_feat_imp,
Nov2020_parent_feat_imp,
updated_April2021_adult_feat_imp,
updated_April2021_parent_feat_imp,
updated_Nov2020_adult_feat_imp,
updated_Nov2020_parent_feat_imp]
required_dfs = [df[:31] if len(df) > 31 else df for df in required_dfs]
save_xls(list_dfs,xls_path,required_dfs)
# + [markdown] id="K4u5ITCdXDKD"
# <h2><center><font color='Blue'><b>Sentiment Analysis on the crisis logger csv file</center></b></font></h2>
# + [markdown] id="NcpH4ccwXYlJ"
# <font color='purple'><b> The crisis logger csv file already looked process and the immediate thought was to perform sentiment analysis. We used Vader to get Positive, neutral and negative scores for each transcription and found the overall rating for each. We finally save the results in a xlsx file along with the other details from the crisis logger file. Below is the function, the execution and how we finally save the result </b></font><br>
# + id="XXwPXgkuYK-o"
def sentiment_scores(sentence):
"""
getting sentiment scores for every sentence in crisislogger file.
"""
sid_obj = SentimentIntensityAnalyzer()
sentiment_dict=sid_obj.polarity_scores(sentence)
Negative_score = sentiment_dict['neg']*100
Neutral_score = sentiment_dict['neu']*100
Positive_score = sentiment_dict['pos']*100
if (sentiment_dict['compound'] >= 0.05):
Overall_rate = 'Positive'
elif (sentiment_dict['compound'] <= - 0.05):
Overall_rate = 'Negative'
else :
Overall_rate = 'Neutral'
return Negative_score,Neutral_score,Positive_score,Overall_rate
#executing sentiment_scores on crisislogger data and combining the output with the crisis logger file itself, exporting the result into an xlsx file.
data = pd.read_csv(r"./ML-for-Good-Hackathon/Data/CrisisLogger/crisislogger.csv")
res = data['transcriptions'].apply(sentiment_scores)
data['Negative_score'] = [x[0] for x in res]
data['Neutral_score'] = [x[1] for x in res]
data['Positive_score'] = [x[2] for x in res]
data['Overall_rate']=[x[3] for x in res]
data.to_excel('./sentiment_scores_crisislogger.xlsx')
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="eyRbnyd2YYrM" outputId="ef13d8ce-f296-486d-b29b-c3fd076cf596"
data.head(10)
# + [markdown] id="Pq6ld0bPaEgz"
# <h2><center><font color='Blue'><b> Thank you so much for this amazing opportunity! We are extremely happy that we got to work for such a wonderful cause.</center></b></font></h2>
| Team 12 submission/Final_Notebook_and_codes/Final_ML_for_good.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# In this notebook, we compute invariant sets for the 3D system with univariate input of Example ?.? of [???].
# We need to select an semidefinite programming solver to find the invariant set. A complete list of available solvers is [available here](https://www.juliaopt.org/JuMP.jl/stable/installation/#Getting-Solvers-1).
using MosekTools
using JuMP
solver = with_optimizer(Mosek.Optimizer, QUIET=true)
# We first define the safe sets and input sets for our system using [Polyhedra](https://github.com/JuliaPolyhedra/Polyhedra.jl).
using Polyhedra
G = [ 0.8304 0.9608 -0.6607
-0.5279 0.7682 -0.2108
-0.9722 -0.5355 -0.7135
0.1539 -0.7305 0.8130
0.0238 -0.4138 -0.4418
0.2655 0.8882 -0.8859
-0.6859 0.4937 -0.0190
-0.1077 -0.8368 0.8729
-0.8887 0.0932 0.3885
-0.4192 -0.9132 0.3742
-0.1649 -0.9065 0.1010
0.2987 0.9829 0.9178
0.6361 -0.6986 -0.7053]
F = [0.0366
0.1687
0.8988
0.0233
0.3298
0.1828
0.2025
0.1244
0.2920
0.3577
0.3586
0.3896
0.5930]
safe_set = polyhedron(hrep(G, F), DefaultLibrary{Float64}(solver))
cheby_center, cheby_radius = chebyshevcenter(safe_set, solver)
using SemialgebraicSets
input_set = FullSpace()
# We new define the dynamic of our system and create it using [MathematicalSystems](https://github.com/JuliaReach/MathematicalSystems.jl/).
using MathematicalSystems
A = [ 0 1 -2
3 -4 5
-6 7 8]
B = reshape([-1; 2; 4], 3, 1)
system = ConstrainedLinearControlDiscreteSystem(A, B, safe_set, input_set)
# We now use SwitchOnSafety to compute the invariant set. We start with symmetric ellipsoid centered at the origin.
using SwitchOnSafety
sym_variable = Ellipsoid(symmetric = true)
max_vol_sym_ell = invariant_set(system, solver, sym_variable)
# This computes the maximal *volume* ellipsoid.
# We can instead maximize the integral of the quadratic form defining the ellipsoid over the hypercube.
# This corresponds to maximizing the sum of squares of its semi-axes instead or equivalently the trace of the positive definite matrix defining the quadratic form.
max_tr_sym_ell = invariant_set(system, solver, sym_variable, volume_heuristic = ell -> L1_heuristic(ell, ones(3)))
# We can see the result below.
using Plots
plot(project(safe_set, [2, 3]))
plot!(project(max_vol_sym_ell, [2, 3]), color=:orange)
plot!(project(max_tr_sym_ell, [2, 3]), color=:green)
# We can see that forcing the center of the ellipsoid to coincide with the Chebyshev center of the safe set is quite conservative.
# We can ask instead to search for any ellipsoid with the chebyshev of the safe set in its interior.
# To avoid having to solve Bilinear Matrix Inequalities, we set the S-procedure scaling to `1.0`.
using SwitchOnSafety
variable = Ellipsoid(point = SetProg.InteriorPoint(cheby_center))
max_vol_ell = invariant_set(system, solver, variable, λ = 1.0)
# Instead of maximizing the volume, we can maximize the L1 norm as above.
using SwitchOnSafety
max_tr_ell = invariant_set(system, solver, variable, λ = 1.0, volume_heuristic = ell -> L1_heuristic(ell, ones(3)))
# We can see that we get a larger ellipsoids for the volume maximization but not for the L1 norm as maximizing the L1 integral over the hypercube centered at the origin is not a very good heuristic. We should instaed maximize the L1 integral over the safe set but this is not implemented yet in [SetProg](https://github.com/blegat/SetProg.jl).
using Plots
plot(project(safe_set, [1, 2]))
plot!(project(max_vol_ell, [1, 2]), color=:orange)
plot!(project(max_tr_ell, [1, 2]), color=:green)
using Plots
plot(project(safe_set, [2, 3]))
plot!(project(max_vol_ell, [2, 3]), color=:orange)
plot!(project(max_tr_ell, [2, 3]), color=:green)
using Plots
plot(project(safe_set, [1, 3]))
plot!(project(max_vol_ell, [1, 3]), color=:orange)
plot!(project(max_tr_ell, [1, 3]), color=:green)
| examples/Controlled_Invariant_Sets/3D_System_unbounded_univariate_control_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''scraping-clacso'': conda)'
# metadata:
# interpreter:
# hash: 95bb600096007ab9f567dd294a4f24ed256a7f8cd3baa781798d9a0497e2540f
# name: python3
# ---
import pandas as pd
import numpy as np
import requests
from bs4 import BeautifulSoup
from lxml import html
import scrapy
from time import sleep
import urllib3
import json
from selenium import webdriver
import random
def mexico_puebla(link):
mexico_puebla = pd.DataFrame()
links_abiertos = []
titulos_abiertos = []
pdfs_proyectos = []
descripciones = []
driver = webdriver.Firefox()
driver.get(link)
metas = driver.find_elements_by_xpath('//h4[@class="entry-title"]/a')
for meta in metas:
link = meta.get_attribute('href')
if 'programa' not in link:
links_abiertos.append(link)
titulos_abiertos.append(meta.text)
driver.quit()
n = 0
while n < len(links_abiertos):
for link_abierto in links_abiertos:
documentos = []
desc = ''
driver = webdriver.Firefox()
driver.get(link_abierto)
docs = driver.find_elements_by_xpath('//div[@class="post-wrapper-content"]//a')
try:
desc = driver.find_element_by_xpath('//div[@class="post-wrapper-content"]').text
except:
desc = ''
descripciones.append(desc)
for doc in docs:
pdf = doc.get_attribute('href')
if 'google' not in pdf:
documentos.append(pdf)
driver.quit()
pdfs_proyecto = ''
for doc in documentos:
pdfs_proyecto = pdfs_proyecto+doc+', '
pdfs_proyectos.append(pdfs_proyecto.strip(', '))
n+=1
mexico_puebla['Título'] = titulos_abiertos
mexico_puebla['Estado'] = 'Abierto'
mexico_puebla['Descripción'] = descripciones
mexico_puebla['Link'] = links_abiertos
mexico_puebla['Pdfs'] = pdfs_proyectos
return mexico_puebla
# +
estados = ['http://www.concytep.gob.mx/convocatorias-y-programas-abiertos/', 'http://www.concytep.gob.mx/convocatorias-cerradas-y-resultados/page/1/', 'http://www.concytep.gob.mx/convocatorias-cerradas-y-resultados/page/2/', 'http://www.concytep.gob.mx/convocatorias-cerradas-y-resultados/page/3/', 'http://www.concytep.gob.mx/convocatorias-cerradas-y-resultados/page/4/']
mexico_pueblaT = pd.DataFrame()
puebla1 = mexico_puebla(estados[0])
puebla2 = mexico_puebla(estados[1])
puebla3 = mexico_puebla(estados[2])
puebla4 = mexico_puebla(estados[3])
puebla5 = mexico_puebla(estados[4])
#len(pdfs_proyectos)
# +
frames = [puebla1, puebla2, puebla3, puebla4, puebla5]
puebla = pd.concat(frames)
puebla.reset_index(drop=True, inplace=True)
# -
puebla.to_excel('Mexico_Puebla.xlsx')
| test_scrapper/Mexico_puebla.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python395jvsc74a57bd0a8e76fc1bb3a146466b516168aa5f10571582da0c13260574329d12e34a70cb3
# ---
# +
# %matplotlib inline
# %config InlineBackend.figure_formats = {'retina', 'png'}
# %load_ext autoreload
# %autoreload 2
import glob
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# -
df = pd.read_csv('2021data.csv')
df.head()
# +
df_import = df.loc[(df['Flow'] == 'Imports')]
df_tes = df.loc[(df['Flow'] == 'Total energy supply')]
test=pd.DataFrame()
df_tes = df_tes.set_index(['Time'], ['Product'])
df_import = df_import.set_index(['Time'], ['Product'])
df_import['id'] = df_import['Value'] / df_tes['Value']
# df_import.to_csv('temp.csv')
# -
df_import['Product'].unique()
df_import = df_import.query("Product == 'Biogasoline' | Product == 'Primary solid biofuels'| Product == 'Biodiesels'")
df_import['id'] *= 100
df_import.head()
# +
# sns.boxplot(
# data=df_import, x='Product', y='id', hue='Import-dependent?'
# )
plt.figure(figsize=(10, 6))
sns.cubehelix_palette(start=.5, reverse = True, as_cmap=True)
ax = sns.barplot(
data=df_import.query("TIME == 2000 | TIME == 2010 | TIME == 2019"),
x="Product", y="id", hue="TIME", palette = 'crest')
ax.set_ylabel('Import Dependency (%)', fontsize='large')
ax.set_xlabel(' ', fontsize='medium')
ax.legend(frameon=False, ncol=3, fontsize='large')
# plt.axhline(y=1, color='grey', linestyle='-.')
plt.savefig(
"figure_4_202111.pdf", dpi=600,
bbox_inches='tight', pad_inches=0.2
)
# -
| fig4_importdepend/fig_4.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 📃 Solution for Exercise M1.05
#
# The goal of this exercise is to evaluate the impact of feature preprocessing
# on a pipeline that uses a decision-tree-based classifier instead of a logistic
# regression.
#
# - The first question is to empirically evaluate whether scaling numerical
# features is helpful or not;
# - The second question is to evaluate whether it is empirically better (both
# from a computational and a statistical perspective) to use integer coded or
# one-hot encoded categories.
# +
import pandas as pd
adult_census = pd.read_csv("../datasets/adult-census.csv")
# -
target_name = "class"
target = adult_census[target_name]
data = adult_census.drop(columns=[target_name, "education-num"])
# As in the previous notebooks, we use the utility `make_column_selector`
# to select only columns with a specific data type. Besides, we list in
# advance all categories for the categorical columns.
# +
from sklearn.compose import make_column_selector as selector
numerical_columns_selector = selector(dtype_exclude=object)
categorical_columns_selector = selector(dtype_include=object)
numerical_columns = numerical_columns_selector(data)
categorical_columns = categorical_columns_selector(data)
# -
# ## Reference pipeline (no numerical scaling and integer-coded categories)
#
# First let's time the pipeline we used in the main notebook to serve as a
# reference:
# +
import time
from sklearn.model_selection import cross_validate
from sklearn.pipeline import make_pipeline
from sklearn.compose import ColumnTransformer
from sklearn.preprocessing import OrdinalEncoder
from sklearn.ensemble import HistGradientBoostingClassifier
categorical_preprocessor = OrdinalEncoder(handle_unknown="use_encoded_value",
unknown_value=-1)
preprocessor = ColumnTransformer([
('categorical', categorical_preprocessor, categorical_columns)],
remainder="passthrough")
model = make_pipeline(preprocessor, HistGradientBoostingClassifier())
start = time.time()
cv_results = cross_validate(model, data, target)
elapsed_time = time.time() - start
scores = cv_results["test_score"]
print("The mean cross-validation accuracy is: "
f"{scores.mean():.3f} +/- {scores.std():.3f} "
f"with a fitting time of {elapsed_time:.3f}")
# -
# ## Scaling numerical features
#
# Let's write a similar pipeline that also scales the numerical features using
# `StandardScaler` (or similar):
# +
# solution
import time
from sklearn.preprocessing import StandardScaler
preprocessor = ColumnTransformer([
('numerical', StandardScaler(), numerical_columns),
('categorical', OrdinalEncoder(handle_unknown="use_encoded_value",
unknown_value=-1),
categorical_columns)])
model = make_pipeline(preprocessor, HistGradientBoostingClassifier())
start = time.time()
cv_results = cross_validate(model, data, target)
elapsed_time = time.time() - start
scores = cv_results["test_score"]
print("The mean cross-validation accuracy is: "
f"{scores.mean():.3f} +/- {scores.std():.3f} "
f"with a fitting time of {elapsed_time:.3f}")
# + [markdown] tags=["solution"]
# ### Analysis
#
# We can observe that both the accuracy and the training time are approximately
# the same as the reference pipeline (any time difference you might observe is
# not significant).
#
# Scaling numerical features is indeed useless for most decision tree models in
# general and for `HistGradientBoostingClassifier` in particular.
# -
# ## One-hot encoding of categorical variables
#
# We observed that integer coding of categorical variables can be very
# detrimental for linear models. However, it does not seem to be the case for
# `HistGradientBoostingClassifier` models, as the cross-validation score
# of the reference pipeline with `OrdinalEncoder` is reasonably good.
#
# Let's see if we can get an even better accuracy with `OneHotEncoder`.
#
# Hint: `HistGradientBoostingClassifier` does not yet support sparse input
# data. You might want to use
# `OneHotEncoder(handle_unknown="ignore", sparse=False)` to force the use of a
# dense representation as a workaround.
# +
# solution
import time
from sklearn.preprocessing import OneHotEncoder
categorical_preprocessor = OneHotEncoder(handle_unknown="ignore", sparse=False)
preprocessor = ColumnTransformer([
('one-hot-encoder', categorical_preprocessor, categorical_columns)],
remainder="passthrough")
model = make_pipeline(preprocessor, HistGradientBoostingClassifier())
start = time.time()
cv_results = cross_validate(model, data, target)
elapsed_time = time.time() - start
scores = cv_results["test_score"]
print("The mean cross-validation accuracy is: "
f"{scores.mean():.3f} +/- {scores.std():.3f} "
f"with a fitting time of {elapsed_time:.3f}")
# + [markdown] tags=["solution"]
# ### Analysis
#
# From an accuracy point of view, the result is almost exactly the same.
# The reason is that `HistGradientBoostingClassifier` is expressive
# and robust enough to deal with misleading ordering of integer coded
# categories (which was not the case for linear models).
#
# However from a computation point of view, the training time is
# much longer: this is caused by the fact that `OneHotEncoder`
# generates approximately 10 times more features than `OrdinalEncoder`.
#
# Note that the current implementation `HistGradientBoostingClassifier`
# is still incomplete, and once sparse representation are handled
# correctly, training time might improve with such kinds of encodings.
#
# The main take away message is that arbitrary integer coding of
# categories is perfectly fine for `HistGradientBoostingClassifier`
# and yields fast training times.
# + [markdown] tags=["solution"]
# <div class="admonition important alert alert-info">
# <p class="first admonition-title" style="font-weight: bold;">Important</p>
# <p>Which encoder should I use?</p>
# <table border="1" class="colwidths-auto docutils">
# <thead valign="bottom">
# <tr><th class="head"></th>
# <th class="head">Meaningful order</th>
# <th class="head">Non-meaningful order</th>
# </tr>
# </thead>
# <tbody valign="top">
# <tr><td>Tree-based model</td>
# <td><tt class="docutils literal">OrdinalEncoder</tt></td>
# <td><tt class="docutils literal">OrdinalEncoder</tt></td>
# </tr>
# <tr><td>Linear model</td>
# <td><tt class="docutils literal">OrdinalEncoder</tt> with caution</td>
# <td><tt class="docutils literal">OneHotEncoder</tt></td>
# </tr>
# </tbody>
# </table>
# <ul class="last simple">
# <li><tt class="docutils literal">OneHotEncoder</tt>: will always do something meaningful, but can be
# unnecessary slow with trees.</li>
# <li><tt class="docutils literal">OrdinalEncoder</tt>: can be detrimental for linear models unless your category
# has a meaningful order and you make sure that <tt class="docutils literal">OrdinalEncoder</tt> respects this
# order. Trees can deal with <tt class="docutils literal">OrdinalEncoder</tt> fine as long as they are deep
# enough.</li>
# </ul>
# </div>
| notebooks/03_categorical_pipeline_sol_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # CS375 - Assignment 3: LN model
#
# In this notebook, we show the training and validation results of training the LN model on natural scene and white noise data.
# %matplotlib inline
import os
import numpy as np
import tensorflow as tf
import pymongo as pm
import gridfs
import cPickle
import scipy.signal as signal
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook, trange
# ### Getting data from the database
# Let's connect to the database and pull the data training and test data that is stored while training our network.
# In order to find the right experiment id, it is useful to display which experiments are stored in the database first.
# +
# connect to database
dbname = 'ln_model'
wn = 'whitenoise'
ns = 'naturalscene'
exp_id = 'trainval0'
port = 24444
conn = pm.MongoClient(port = port)
c_wn = conn[dbname][wn + '.files']
c_ns = conn[dbname][ns + '.files']
# print out saved experiments in collection coll
print(c_wn.distinct('exp_id'))
print(c_ns.distinct('exp_id'))
# -
# ### Plotting the training curve
# +
from scipy.signal import convolve
def smooth_signal(lst, smooth=5):
return convolve(lst, np.ones((smooth))/smooth, 'valid')
def get_losses(coll, exp_id):
"""
Gets all loss entries from the database and concatenates them into a vector
"""
q_train = {'exp_id' : exp_id, 'train_results' : {'$exists' : True}}
return np.array([_r['loss']
for r in coll.find(q_train, projection = ['train_results'])
for _r in r['train_results']])
def get_steps(coll, exp_id):
q_train = {'exp_id' : exp_id, 'train_results' : {'$exists' : True}}
steps = [i['step'] for i in coll.find(q_train)]
return steps
# +
loss_wn = get_losses(c_wn, exp_id)
loss_ns = get_losses(c_ns, exp_id)
# Plot the training loss
plt.figure()
plt.title('Loss over steps')
plt.xlabel('Steps')
plt.ylabel('Loss')
plt.plot(loss_wn,label='Whitenoise',color='black')
plt.plot(loss_ns,label='Natural scenes',color='lightblue')
plt.legend()
smooth_wn = smooth_signal(loss_wn, smooth=50)
smooth_ns = smooth_signal(loss_ns, smooth=50)
plt.figure()
plt.title('Smoothed loss over steps')
plt.xlabel('Steps')
plt.ylabel('Loss')
plt.plot(smooth_wn,label='Whitenoise',color='black')
plt.plot(smooth_ns,label='Natural scenes',color='lightblue')
plt.legend()
# -
# From these plots, we see that the LN model more easily fits the white noise than natural scenes, as shown by the noticeably lower loss. However, the LN model also converges at about the same rate for both stimulus types, largely bottoming out after around 1000 steps.
# ### Poisson loss
# +
def get_validation_data(coll,exp_id, key1='topn_val', key2='loss'):
"""
Gets the validation data from the database (except for gridfs data)
"""
q_val = {'exp_id' : exp_id, 'validation_results' : {'$exists' : True}, 'validates' : {'$exists' : False}}
val_steps = coll.find(q_val, projection = ['validation_results'])
return [val_steps[i]['validation_results'][key1][key2]
for i in range(val_steps.count())]
### PLOT VALIDATION RESULTS HERE
# losses = get_validation_data('mc_colorize')
losses = get_validation_data(c_wn,exp_id, 'test_loss', 'poisson_loss')
wn_min_index = np.argmin(losses[1:])
plt.plot(losses,label='Whitenoise',color='black')
losses = get_validation_data(c_ns,exp_id, 'test_loss', 'poisson_loss')
ns_min_index = np.argmin(losses[1:])
plt.plot(losses,label='Natural scenes',color='lightblue')
plt.title('Poisson loss')
plt.legend()
plt.xlabel('Validation steps')
plt.ylabel('Poisson loss')
# -
# The validation loss curves show evidence of overfitting, particularly on the natural scenes dataset, as validation loss begins to increase near the end of the training curve.
# # Validation (Pearson Correlation)
def get_correlation_val(coll):
# get validation data
val_results = {}
steps = get_steps(coll, 'trainval0')
pearson_results = {}
for step in steps:
val_results[step] = {}
val_exp_id = 'testval_step%s' % step
val_result = list(coll.find({'exp_id' : val_exp_id,
'validation_results' : {'$exists' : True}},
projection=['validation_results']))
val_results[step]['naturalscene'] = np.mean(val_result[-1]['validation_results']['naturalscene_pearson'].values())
val_results[step]['whitenoise'] = np.mean(val_result[-1]['validation_results']['whitenoise_pearson'].values())
return val_results
# +
wn_val = get_correlation_val(c_wn)
ns_val = get_correlation_val(c_ns)
f, axes = plt.subplots(1, 2, figsize=(12,6))
ax1, ax2 = axes
ax1.plot([wn_val[s]['naturalscene'] for s in sorted(wn_val.keys())], linewidth=2, label='naturalscene')
ax1.plot([wn_val[s]['whitenoise'] for s in sorted(wn_val.keys())], linewidth=2, label='whitenoise')
ax1.set_title('Model Trained on White Noise', fontsize=18, y=1.05)
ax2.plot([ns_val[s]['naturalscene'] for s in sorted(ns_val.keys())], linewidth=2, label='naturalscene')
ax2.plot([ns_val[s]['whitenoise'] for s in sorted(ns_val.keys())], linewidth=2, label='whitenoise')
ax2.set_title('Model Trained on Natural Scenes', fontsize=18, y=1.05)
for ax in [ax1, ax2]:
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.spines['bottom'].set_position(('axes', -0.05))
ax.spines['left'].set_position(('axes', -0.05))
ax.legend(fontsize=14)
ax.set_ylim(0,.8)
ax.set_xlim(0,50)
ax.set_xlabel('Step', fontsize=14)
ax.set_ylabel('Pearson Correlation', fontsize=14)
plt.subplots_adjust(wspace=.5)
# -
# As expected, the correlation coefficient is higher when the input stimulus type and the training stimulus type are the same. In addition, the correlation coefficients of the CNN validation results dominate those of the LN results for all four training-validation stimulus combinations.
| assignment3/ln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# ### Re-creating convolution intro notebook
#
# Exploring convolutions using mnist dataset
#
# Finding edges - horizontal/vertical and diagonal
#
# + deletable=true editable=true
import tensorflow as tf
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
from ipywidgets import interact, interactive, fixed
from tensorflow.examples.tutorials import mnist
# + deletable=true editable=true
data = mnist.input_data.read_data_sets("MNIST_data")
# + deletable=true editable=true
# Get the images and the labels
"""
images, labels = data.train.images, data.train.labels
images.shape
# Save the images to disk after resizing
images = images.reshape((55000,28,28))
%mkdir ./data/MNIST_data
%mkdir ./data/MNIST_data/train
np.savez_compressed("./data/MNIST_data/train", images = images, labels = labels)
"""
# + deletable=true editable=true
data = np.load("./data/MNIST_data/train.npz")
images = data['images']
labels = data['labels']
# + deletable=true editable=true
images.shape
# + deletable=true editable=true
# Plotting many images
# plt.imshow(images[0],interpolation=None?
# + deletable=true editable=true
plt.imshow(images[0],interpolation=None)
# + deletable=true editable=true
def plot(img):
plt.imshow(img, interpolation=None, cmap='gray')
# + deletable=true editable=true
# Plotting multiple images
def plots(images, titles=None):
fig = plt.figure(figsize=(12,8))
for i in range(len(images)):
sp = fig.add_subplot(1, len(images), i + 1)
if titles is not None:
sp.set_title(titles[i])
plt.imshow(images[i], cmap = 'gray')
# + deletable=true editable=true
plots(images[0:4], labels[0:4])
# + deletable=true editable=true
top = [[-1, -1, -1],
[1,1,1],
[0,0,0]]
# + deletable=true editable=true
plot(top)
# + deletable=true editable=true
# Zoom image - plotting only a specific subsection of the image
r = (0, 28)
def zoomim(x1 = 0, x2 = 28, y1 = 0, y2 = 28):
plot(images[0,y1:y2, x1:x2])
# Interactive - these are from ipython widgets
w = interactive(zoomim, x1 = r, x2 = r, y1 = r, y2 = r)
w
# + deletable=true editable=true
dims
# + deletable=true editable=true
# He is applying some sort of exponential function
# Not really - index_exp stands for index Expression and gets you a slice object
# that can later be used to get a slice of the array
# gets the parameters in a dictionary - nice!
k = w.kwargs
dims = np.index_exp[k['y1']:k['y2']:1, k['x1']:k['x2']]
images[0][dims]
# + deletable=true editable=true
corrtop = np.correlate?
# + deletable=true editable=true
from scipy.ndimage.filters import correlate, convolve
# + deletable=true editable=true
corrtop = correlate(images[0], top)
# + deletable=true editable=true
plot(corrtop)
# + deletable=true editable=true
np.rot90(top, 1)
# + deletable=true editable=true
np.rot90(top, 2)
# + deletable=true editable=true
convtop = convolve(images[0], np.rot90(top, 2))
# + deletable=true editable=true
plot(convtop)
# + deletable=true editable=true
# Convolution with the twice rotated array is the same as correlation with the unrotated one
# allclose checks if the two matrices are close within a tolerance
np.allclose(corrtop, convtop)
# + deletable=true editable=true
straights = [np.rot90(top, i) for i in range(4)]
# + deletable=true editable=true
plots(straights,titles = [0, 1, 2, 3])
# + deletable=true editable=true
br = [[0,0,1],
[0,1,-1.5],
[1,-1.5,0]]
# + deletable=true editable=true
diags = [np.rot90(br, i) for i in range(4)]
plots(diags)
# + deletable=true editable=true
rots = straights + diags
plots(rots)
# + deletable=true editable=true
corrs = [correlate(images[0], rot) for rot in rots]
plots(corrs)
# + deletable=true editable=true
from skimage.measure import block_reduce
# + deletable=true editable=true
# block_reduce is to do max pooling or averaging out a block
def pool(im):
return block_reduce(im, (7,7), np.max)
# + deletable=true editable=true
plots([pool(im) for im in corrs])
# + deletable=true editable=true
# comparing eights and ones by looking at what the correlation/convolution does for them
eights = [images[i] for i in range(len(images)) if labels[i] ==8]
ones = [images[i] for i in range(len(images)) if labels[i] ==1]
# + deletable=true editable=true
plots(eights[:5])
plots(ones[:5])
# + deletable=true editable=true
pool8 = [np.array([pool(correlate(im, rot)) for im in eights]) for rot in rots]
# + deletable=true editable=true
pool8[0].shape
# + deletable=true editable=true
plots(pool8[0][0:5])
# + deletable=true editable=true
def normalize(arr):
return (arr - arr.mean())/arr.std()
# + deletable=true editable=true
# np.mean?
# + deletable=true editable=true
filts8 = np.array([ims.mean(axis = 0) for ims in pool8])
filts8 = normalize(filts8)
# + deletable=true editable=true
plots(filts8)
# + deletable=true editable=true
#Folding code
asarts
| nbs/convolution-intro-re-creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# (Note that all units unless otherwise mentioned are SI units.)
import numpy as np
import numexpr as ne
import matplotlib.pyplot as plt
import pickle,os
from multislice import prop,prop_utils
# Importing all the required libraries.
def make_zp_from_rings(n,grid_size):
zp = np.zeros((grid_size,grid_size))
for i in range(n):
if i%2 == 1 :
locs_ = np.load('ring_locs_'+str(i)+'.npy')
locs_ = tuple((locs_[0],locs_[1]))
vals_ = np.load('ring_vals_'+str(i)+'.npy')
zp[locs_] = vals_
return zp
# *make_zp_from_rings* : make a zone plate from the rings which were created earlier.
# * *Inputs* : n - number of rings, grid_size
# * *Outputs* : a numpy array containing the zone plate
os.chdir(os.getcwd()+str('/rings'))
parameters = pickle.load(open('parameters.pickle','rb'))
for i in parameters : print(i,' : ',parameters[i])
zp_coords = parameters['zp_coords']
grid_size = parameters['grid_size']
zp = make_zp_from_rings(100,int(grid_size))
plt.imshow(zp,extent=np.array(zp_coords)*1e6)
plt.title('Zone plate pattern')
plt.xlabel('axes in um')
plt.colorbar()
plt.show()
# Importing zone plate pattern and the parameters associated with it. Plotting the zone plate after creating it from the rings. One needs to save the image with high dpi to clearly see the zone plate.
beta = parameters['beta']
delta = parameters['delta']
step_xy = parameters['step_xy']
wavel = parameters['wavelength in m']
f = parameters['focal_length']
L = step_xy*np.shape(zp)[0]
m,n = np.shape(zp)
zp_thickness = 10e-6
# *Setting up the parameters for the simulation*<br>
# * *beta and delta* : delta and beta at the relevant energy <br>
# * *step_xy* : sampling in the xy plane<br>
# * *wavel* : wavelength<br>
# * *f* : focal length<br>
# * *wavefront* : wavefront that is initialzed with magnitude of 1 and phase of 0<br>
# * *L* : support length<br>
# * *m,n* : dimensions of input zp<zp>
# * *zp_thickness* : thickness of zone plate <br>
i = 0 #Angle in degrees
theta = (i)*(np.pi/180)
slope = np.tan(theta)
x = np.linspace(zp_coords[0],zp_coords[1],n)
X,Y = np.meshgrid(x,x)
z1 = 2*np.pi*(1/wavel)*slope*X
del X,Y,x
wave_in = np.multiply(np.ones(np.shape(zp),dtype='complex64'),np.exp(1j*(z1)))
del z1
# Creating the input wave for the simulation. The input wave has a position(along one axis) dependent phase to simulate the effect of tilting the zone plate (instead of tilting the zone plate itself though tilting the zone plate might be more efficient in the long run)
number_of_steps = prop_utils.number_of_steps(step_xy,wavel,zp_thickness)*2
ne.set_vml_num_threads(12)
wave_exit,L1 = prop_utils.optic_illumination(wave_in,zp,delta,beta,zp_thickness,
step_xy,wavel,number_of_steps,0,0)
del zp,wave_in
plt.imshow(np.abs(wave_exit),extent=np.array([-L1/2,L1/2,-L1/2,L1/2])*1e6,cmap='jet')
plt.xlabel('axes in um')
plt.title('wavefront(magnitude) at zone plate exit')
plt.colorbar()
plt.show()
print(np.max(np.abs(wave_exit)))
step_z = f
p = prop_utils.decide(step_z,step_xy,L,wavel)
print('Propagation to focal plane')
print('Fresnel Number :',((L**2)/(wavel*step_z)))
wave_focus,L2 = p(wave_exit - np.ones(np.shape(wave_exit)),step_xy,L,wavel,step_z)
wave_focus = wave_focus + np.ones(np.shape(wave_exit))
plt.imshow(np.abs(wave_focus),extent=np.array([-L2/2,L2/2,-L2/2,L2/2])*1e6,cmap='jet')
plt.xlabel('axes in um')
plt.title('wavefront(magnitude) at focal plane')
plt.colorbar()
plt.show()
# The focal spot size is ~50 pixels and one can't see it when the whole wavefront (40k by 40k) is visualized !
# Another thing one can notice is the faint outline of a square, this is because the output grid dimensions are different from the input grid dimensions and the length of the side of this square is nothing but the support length of the input plane.
x = np.linspace(-1,1,grid_size)*L2/2
plt.plot(x,abs(wave_focus)[int(grid_size/2),:])
plt.xlabel('axes in um')
plt.show()
print('slice in x-direction along the focal plane')
# The above plot shows that the location of the focal spot has shifted due to the tilt of the input wave.
focal_spot_size = 25
focal_spot,x_,y_,max_val = prop_utils.get_focal_spot(np.abs(wave_focus),grid_size,focal_spot_size)
plt.imshow(np.log(focal_spot**2+1),extent = np.array([-1,1,-1,1])*(focal_spot_size)*(L2/grid_size)*(1e9),cmap='jet')
plt.title('focal spot intensity (log scale)')
plt.xlabel('axes in nm')
plt.colorbar()
plt.show()
| notebooks/finite_difference/2d/zone_plate/zp_make/simulate_zp_with_tilt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# This notebook contains the code to reproduce the results of the paper:
#
# [<NAME>](https://perso.uclouvain.be/benoit.legat), [<NAME>](https://perso.uclouvain.be/raphael.jungers/content/home). [**Parallel optimization on the Entropic Cone**](http://sites.uclouvain.be/sitb2016/Proceedings_SITB2016_preliminary.pdf). [*37rd Symposium on Information Theory in the Benelux*](http://sites.uclouvain.be/sitb2016), 2016.
using EntropicCone
# Select a solver installed. [`GLPK`](https://github.com/JuliaOpt/GLPK.jl) is not the fastest but it is free and reliable, [`Gurobi`](https://github.com/JuliaOpt/Gurobi.jl) is fast and reliable but it is commercial (although academic licenses are available). Many other solvers are available, see [here](http://www.juliaopt.org/) for a list.
using GLPKMathProgInterface
solver = GLPKSolverLP()
c = ingleton(4,1,2,3,4)
cut = nonnegative(4, 1234)
using CutPruners
n = 4
h = polymatroidcone(Float64, 4)
newcut = :AddImmediately
(sp, allnodes) = @time stochasticprogram(c, h, solver, 7, cut, newcut, AvgCutPruningAlgo.([-1,-1,-1,-1,-1,-1,-1]));
length(allnodes)
s = StructDualDynProg.SDDP(sp, 3, stopcrit=StructDualDynProg.IterLimit(1), verbose=1, K=-1);
s = StructDualDynProg.SDDP(sp, 3, stopcrit=StructDualDynProg.IterLimit(1), verbose=1, K=-1);
s = StructDualDynProg.SDDP(sp, 3, stopcrit=StructDualDynProg.IterLimit(1), verbose=1, K=-1);
s = StructDualDynProg.SDDP(sp, 3, stopcrit=StructDualDynProg.IterLimit(1), verbose=1, K=-1);
CutPruners.exactpruning!(sp.data[1].nlds.FCpruner, solver)
CutPruners.ncuts(sp.data[1].nlds.FCpruner)
s.status
s.sol
s.objval
| examples/Parallel optimization on the Entropic Cone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#--- Cargamos el paquete torch y otras utilidades
import torch
from torch.autograd import Variable # Para convertir nuestras varaibles a tensores
import numpy as np
x = np.linspace(-10, 10, 1000)
y = np.sin(x) + 0.5*np.random.rand(1000) # Calculamos una función senoidal
np.shape(x), np.shape(y)
# +
#--- Visualizamos los datos
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(7,5))
plt.plot(x,y,'*')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# +
#--- Convertimos los numpy a tensores
X = Variable(torch.from_numpy(x)).float()
X = torch.unsqueeze(X,dim=1)
Y = Variable(torch.from_numpy(y)).float()
Y = torch.unsqueeze(Y,dim=1)
# -
#--- Definimos la ANN
input_dim = 1
model = torch.nn.Sequential(
torch.nn.Linear(input_dim, 200),
torch.nn.ReLU(),
torch.nn.Linear(200, 100),
torch.nn.ReLU(),
torch.nn.Linear(100, 50),
torch.nn.ReLU(),
torch.nn.Linear(50, input_dim)
)
model
#-- Definimos el opmimizador y el score de evaluación
optimizer = torch.optim.Adam(model.parameters(), lr=0.01)
criterion = torch.nn.MSELoss() # Para una regresión es mejor usar MSE
# +
#-- Definimos el número de épocas
n_epoch = 1000
loss_list = []
# Entrenamos la red
for epoch in range(n_epoch+1):
Y_pred = model(X) # Hace una predicción basada en X
loss = criterion(Y_pred, Y)
loss_list.append(loss.item())
optimizer.zero_grad() # Limpia los gradientes para una nueva predicción
loss.backward() # Backpropagation
optimizer.step() # Aplica los gradientes
if (epoch%100 == 0):
print('Epoch: [1/{}], loss: {}'.format(epoch+1,loss.item()))
plt.scatter(X.data.numpy(), Y.data.numpy())
plt.plot(X.data.numpy(), Y_pred.data.numpy(), 'g-', lw=3)
plt.show()
# -
fig = plt.figure()
plt.plot(range(n_epoch+1), loss_list)
plt.xlabel('n_epoch')
plt.ylabel('MSE')
plt.show()
| soluciones/Entrega4_sol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy
import toyplot.color
numpy.random.seed(1234)
# +
# Random graph
vertices = ["a", "b", "c", "d", "e", "f", "g", "h", "i", "j"]
random_graph = numpy.random.choice(vertices, (40, 2))
# Random tree
#edges = toyplot.generate.prufer_tree(numpy.random.choice(4, 12))
#edges = toyplot.generate.prufer_tree([1,1])
reingold_a = numpy.array([[0,1],[0,2],[1,3],[1,4],[2,5],[2,6],[3,7],[3,8],[4,9],[4,10],[6,11],[9,12],[9,13],[10,14],[10,15],[12,16],[14,17]])
reingold_b = numpy.array([[0,1],[0,2],[0,3],[0,4],[0,5],[0,6],[1,7],[1,8],[1,9],[1,10],[1,11],[1,12],[1,13],[4,14],[6,15],[13,16],[15,17],[15,18],[15,19],[15,20],[15,21],[15,22],[15,23]])
reingold_c = numpy.array([[0,1],[0,2],[1,3],[2,4],[2,5],[3,6],[4,7],[5,8],[5,9],[6,10],[7,11],[9,12],[10,13],[12,14]])
# Ring graph
#source = numpy.arange(6)
#target = (source + 1) % 6
#edges = numpy.column_stack((source, target))
graph = reingold_c
# +
# %%time
colormap = toyplot.color.LinearMap(toyplot.color.Palette(["white", "yellow", "orange", "red"]))
canvas, axes, mark = toyplot.graph(
graph,
#layout=toyplot.layout.FruchtermanReingold(),
layout=toyplot.layout.Buchheim(),
vcolor=colormap,
vmarker="o",
#varea=40,
vsize=20,
vopacity=1,
vstyle={"stroke":"black"},
ecolor="black",
eopacity=0.2,
estyle={},
width=800,
height=800,
);
axes.show = False
axes.padding = 10
vcoordinates = numpy.copy(mark.vcoordinates)
axes.text(vcoordinates.T[0], vcoordinates.T[1], mark.vids, color="black")
| notebooks/graph-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZqmhyRHvxBnO"
# #**Sentiment Analysis based on Movie Review**
# + id="LQJxIrE3wyEk"
import numpy as np
import pandas as pd
from sklearn.neural_network import MLPClassifier
import matplotlib.pyplot as plt
import zipfile
# + id="iodPOkruxeYQ"
zip = zipfile.ZipFile('train.tsv.zip')
zip.extractall()
# + id="f8o6Y3_HxgDP"
zip = zipfile.ZipFile('test.tsv.zip')
zip.extractall()
# + colab={"base_uri": "https://localhost:8080/", "height": 407} id="iqbLHh_ZxJ8G" outputId="a5db78f2-434d-49c2-ebca-d9007eb871d9"
df_train=pd.read_csv('train.tsv',sep='\t')
df_train=df_train.dropna(how='any')
df_train=df_train.drop(columns=['PhraseId','SentenceId'])
df_train
# + colab={"base_uri": "https://localhost:8080/"} id="VYE9HUz-yhbG" outputId="4a553b01-d3ec-46df-e802-0bb3a3eed0ca"
df_train['Sentiment'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="dQm1hYurxwkr" outputId="d3ba4935-aea0-45fc-b36c-4d2cc3daa190"
import re
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
corpus = []
corpus1=[]
corpus2=[]
for i in range(0, 156060):
review = re.sub('[^a-zA-Z]', ' ', df_train['Phrase'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
all_stopwords = stopwords.words('english')
all_stopwords.remove('not')
review = [ps.stem(word) for word in review if not word in set(all_stopwords)]
review = ' '.join(review)
corpus.append(review)
# + id="5duYCoOSyUl7"
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
y=df_train.iloc[:,-1].values
# + id="XVTcAuq2yWkw"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.20, random_state = 0)
# + colab={"base_uri": "https://localhost:8080/"} id="NxuPlgIr03jU" outputId="6bfa1daf-26ef-494c-b159-a5e8f055223e"
model=MLPClassifier(alpha=0.01, batch_size=256, epsilon=1e-08, learning_rate='adaptive', max_iter=100)
model.fit(X_train,y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="49paZbD91p4s" outputId="c817b2a1-7efb-472a-f55c-7856f4ec05de"
res = model.predict(X_test)
res=np.round(res)
np.set_printoptions(precision=2)
print(res)
# + colab={"base_uri": "https://localhost:8080/"} id="iLPDpxeO1riV" outputId="a368ce19-7752-4ec6-c9a1-3c9a86dc2c4b"
from sklearn.metrics import accuracy_score
print("Accuracy Score for the algorithm=>{}%".format(round(accuracy_score(y_test,res)*100),2))
# + [markdown] id="4N3VTODOzQsI"
# #**Testing the Model**
# + colab={"base_uri": "https://localhost:8080/", "height": 407} id="Xq8xnLrJzOJ7" outputId="ae96121f-37ab-453f-a90f-0f7c3bcbd2e7"
df_test=pd.read_csv('test.tsv',sep='\t')
df_test=df_test.dropna(how='any')
df_test=df_test.drop(columns=['PhraseId','SentenceId'])
df_test
# + colab={"base_uri": "https://localhost:8080/"} id="Bekiftwg5Pmg" outputId="e065d61b-7f82-4d7c-de2c-662f41ccd111"
import re
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
corpus = []
corpus1=[]
corpus2=[]
for i in range(0, 66292):
review = re.sub('[^a-zA-Z]', ' ', df_train['Phrase'][i])
review = review.lower()
review = review.split()
ps = PorterStemmer()
all_stopwords = stopwords.words('english')
all_stopwords.remove('not')
review = [ps.stem(word) for word in review if not word in set(all_stopwords)]
review = ' '.join(review)
corpus.append(review)
# + id="fHze8EYE5Xap"
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features = 1500)
X = cv.fit_transform(corpus).toarray()
# + colab={"base_uri": "https://localhost:8080/"} id="pRSg9enG5Uwz" outputId="66410616-66db-497c-e8da-55ea289acef5"
res = model.predict(X_test)
res=np.round(res)
np.set_printoptions(precision=2)
print(res)
# + colab={"base_uri": "https://localhost:8080/"} id="8hyxxNpx5fGl" outputId="39b54d7a-cd5b-4755-e310-5ac73d8374b0"
for i in range(len(df_test)):
for j in range(len(res)):
if i==j:
print("The catgory for this phrase{0} is{1}" .format(df_test['Phrase'][i],res[i]))
| SAMR_DNLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:plantcv] *
# language: python
# name: conda-env-plantcv-py
# ---
# # Module 4: Measuring plant phenotypes with PlantCV - Multiple plants
#
# [PlantCV homepage](https://plantcv.danforthcenter.org/)
#
# [PlantCV documentation](https://plantcv.readthedocs.io/en/stable/)
# %matplotlib widget
import matplotlib
from plantcv import plantcv as pcv
import numpy as np
import cv2
matplotlib.rcParams["figure.max_open_warning"] = False
pcv.params.debug = "plot"
pcv.params.text_size = 10
pcv.params.text_thickness = 10
pcv.params.line_thickness = 10
pcv.__version__
# ## Refresher: plant segmentation
# Open image file
img, imgpath, imgname = pcv.readimage(filename="images/10.9.1.244_pos-165-002-009_2020-02-29-20-05.jpg")
# Convert the RGB image into a grayscale image by choosing one of the HSV or LAB channels
gray_img = pcv.rgb2gray_lab(rgb_img=img, channel="a")
# Instead of setting a manual threshold, try an automatic threshold method such as Otsu
bin_img = pcv.threshold.otsu(gray_img=gray_img, max_value=255, object_type="dark")
# Remove "salt" noise from the binary image
filter_bin = pcv.fill(bin_img=bin_img, size=100)
# ## Measuring the shape and color of objects in digital image
#
# At this stage we have a binary mask that labels plant pixels (white) and background pixels (black). There are multiple plants but we cannot tell which pixels belong to each plant
# Identify connected components (contours) using the binary image
cnt, cnt_str = pcv.find_objects(img=img, mask=filter_bin)
# Plot each contour to see where they are
pcv.params.color_sequence = "random"
cp = img.copy()
for i in range(0, len(cnt)):
cv2.drawContours(cp, cnt, i, pcv.color_palette(num=100, saved=False)[0], thickness=-1, hierarchy=cnt_str)
pcv.plot_image(cp)
# We have distinct contours for each plant and some (most) plants are composed of multiple contours, how do we assign these to individual plants?
# Create a region of interest (ROI) for one plant
# Filter the contours using the ROI
# Flatten contours into a single object
plant, mask = pcv.object_composition(img=img, contours=plant_cnt, hierarchy=plant_str)
# Measure the size and shape of the plant
shape_img = pcv.analyze_object(img=img, obj=plant, mask=mask)
# Output measurements
print(f"Leaf area = {pcv.outputs.observations['default']['area']['value']} pixels")
print(f"Convex hull area = {pcv.outputs.observations['default']['convex_hull_area']['value']} pixels")
print(f"Solidity = {pcv.outputs.observations['default']['solidity']['value']}")
print(f"Perimeter = {pcv.outputs.observations['default']['perimeter']['value']} pixels")
print(f"Width = {pcv.outputs.observations['default']['width']['value']} pixels")
print(f"Height = {pcv.outputs.observations['default']['height']['value']} pixels")
print(f"Center of mass = {pcv.outputs.observations['default']['center_of_mass']['value']}")
print(f"Convex hull vertices = {pcv.outputs.observations['default']['convex_hull_vertices']['value']}")
print(f"Plant in frame = {pcv.outputs.observations['default']['object_in_frame']['value']}")
print(f"Bounding ellipse center = {pcv.outputs.observations['default']['ellipse_center']['value']}")
print(f"Bounding ellipse center major axis length = {pcv.outputs.observations['default']['ellipse_major_axis']['value']} pixels")
print(f"Bounding ellipse center minor axis length = {pcv.outputs.observations['default']['ellipse_minor_axis']['value']} pixels")
print(f"Bounding ellipse angle of rotation = {pcv.outputs.observations['default']['ellipse_angle']['value']} degrees")
print(f"Bounding ellipse eccentricity = {pcv.outputs.observations['default']['ellipse_eccentricity']['value']}")
# Measure the color properties of the plant
color_hist = pcv.analyze_color(rgb_img=img, mask=mask, colorspaces="hsv")
# 
# Output measurements
print(f"Hue circular mean = {pcv.outputs.observations['default']['hue_circular_mean']['value']} degrees")
print(f"Hue circular mean standard deviation = {pcv.outputs.observations['default']['hue_circular_std']['value']} degrees")
print(f"Hue median = {pcv.outputs.observations['default']['hue_median']['value']} degrees")
| module4/part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''base'': conda)'
# name: python3
# ---
from tfConstrainedGauss import solve_me, random_cov_mat, \
random_non_zero_idx_pairs, InputsME, convert_mat_to_mat_non_zero, convert_mat_non_zero_to_mat
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import numpy as np
# # Test simple 3x3 matrix problem with MaxEnt approach
# +
n = 3
# Non-zero idxs
non_zero_idx_pairs = [(0,0),(1,1),(2,2),(1,0),(2,1)]
# Random cov mat
cov_mat = random_cov_mat(n, unit_diag=True)
prec_mat_structure = convert_mat_non_zero_to_mat(
n=n,
non_zero_idx_pairs=non_zero_idx_pairs,
mat_non_zero=np.full(len(non_zero_idx_pairs),1.0)
)
clear_cmap = ListedColormap(['white', 'none'])
gridspec = {'width_ratios': [1, 1, 0.1]}
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, gridspec_kw=gridspec, dpi=150)
fig.set_size_inches(8,3)
ax1.matshow(prec_mat_structure,cmap='Greys')
ax1.set_title("precision mat structure")
z2plot = ax2.matshow(cov_mat)
ax2.imshow(prec_mat_structure,cmap=clear_cmap)
ax2.set_title("cov mat constraints")
plt.colorbar(z2plot,cax=ax3)
# -
# ## Train
# +
target_cov_mat_non_zero = convert_mat_to_mat_non_zero(
n=n,
non_zero_idx_pairs=non_zero_idx_pairs,
mat=cov_mat
)
inputs = InputsME(
n=n,
non_zero_idx_pairs=non_zero_idx_pairs,
target_cov_mat_non_zero=target_cov_mat_non_zero,
epochs=500,
learning_rate=0.01,
use_weighted_loss=False
)
results = solve_me(inputs)
# -
results.report()
# ## Plot learned cov matrix constraints
# +
init_cov_mat_reconstructed = convert_mat_non_zero_to_mat(
n=n,
non_zero_idx_pairs=non_zero_idx_pairs,
mat_non_zero=results.init_cov_mat_reconstructed_non_zero
)
low=0.4
high=1.2
clear_cmap = ListedColormap(['white', 'none'])
gridspec = {'width_ratios': [1, 1, 1, 0.1]}
fig, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, gridspec_kw=gridspec, dpi=150)
fig.set_size_inches(15,5)
fig.suptitle("cov mat")
ax1.matshow(init_cov_mat_reconstructed,vmin=low,vmax=high)
ax1.imshow(prec_mat_structure,cmap=clear_cmap)
ax1.set_title('initial guess')
ax2.matshow(cov_mat,vmin=low,vmax=high)
ax2.imshow(prec_mat_structure,cmap=clear_cmap)
ax2.set_title('target')
z3plot = ax3.matshow(results.learned_cov_mat,vmin=low,vmax=high)
ax3.imshow(prec_mat_structure,cmap=clear_cmap)
ax3.set_title('learned')
plt.colorbar(z3plot,cax=ax4)
# -
# ## Plot precision matrix learned and it's inverse = full covariance matrix
# +
init_cov_mat_reconstructed = convert_mat_non_zero_to_mat(
n=n,
non_zero_idx_pairs=non_zero_idx_pairs,
mat_non_zero=results.init_cov_mat_reconstructed_non_zero
)
clear_cmap = ListedColormap(['white', 'none'])
gridspec = {'width_ratios': [1, 0.1, 1, 0.1]}
fig, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, gridspec_kw=gridspec, dpi=150)
fig.set_size_inches(15,5)
z1plot = ax1.matshow(results.learned_prec_mat,vmin=-2,vmax=5)
ax1.imshow(prec_mat_structure,cmap=clear_cmap)
ax1.set_title('learned precision mat')
plt.colorbar(z1plot,cax=ax2)
z3plot = ax3.matshow(results.learned_cov_mat,vmin=0.4,vmax=1.2)
ax3.set_title('learned cov mat')
plt.colorbar(z3plot,cax=ax4)
# -
| examples/example_n3_me.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/grigorjevas/Discogs-price-prediction/blob/main/Preparing_data_and_models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Xv_wGJsno5A-"
# # Modelling Discogs Marketplace price predictions
# + [markdown] id="2fIw2e7ro-NB"
# ## EDA and selecting data
# + colab={"base_uri": "https://localhost:8080/"} id="cAN3tZ58pOwh" outputId="9b32717c-9449-4200-d296-a00d56707764"
from google.colab import drive
drive.mount('/content/drive')
# + id="ut_ECGCUo4dS"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.linear_model import LinearRegression
import pickle
# + id="v2QNkNPRpeEC"
df = pd.read_csv("/content/drive/MyDrive/Data/electro_raw_data.csv", parse_dates=["release_date"], na_values="N/A")
# + colab={"base_uri": "https://localhost:8080/", "height": 598} id="vvFgXt29rm5H" outputId="7ab7cf4a-c611-4ad9-bb64-72f9cfe6c518"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="LvRzPjDGrhCk" outputId="8ef27490-686a-46ac-8e01-16aa2fc12fe9"
df.shape
# + [markdown] id="nIh_ZEnNting"
# Convert release date to year
# + id="zDcJU1JVozOa"
df["release_year"] = df['release_date'].dt.year
# + [markdown] id="_cP7meZ6n-Zu"
# Only keep records priced under 100 eur
# + id="oG0NixnbnjLK"
df = df[df["price"] < 100]
# + [markdown] id="rX4P_vXrtnGd"
# Parse item condition to numerical values
# + id="GEZ6nca3tgru"
# Parses item condition value to numerical values
def parse_item_condition_to_int(condition: str) -> int:
return {
"Poor (P)": 0,
"Fair (F)": 0,
"Good (G)": 1,
"Good Plus (G+)": 1,
"Very Good (VG)": 2,
"Very Good Plus (VG+)": 3,
"Generic": 3,
"Not Graded": 3,
"No Cover": 3,
"Near Mint (NM or M-)": 4,
"Mint (M)": 5
}[condition]
# + id="eVNJ8Wettui8"
df["media_condition"] = df["media_condition"].apply(
lambda cond: parse_item_condition_to_int(cond))
df["sleeve_condition"] = df["sleeve_condition"].apply(
lambda cond: parse_item_condition_to_int(cond))
# + [markdown] id="eMPYItJGt5ci"
# Drop N/A rows
# + id="itBExpQpt9ss"
df = df.dropna()
# + [markdown] id="xeT5KOxOuIs2"
# Drop unneeded columns
# + id="Fdq3nFSkuKiF"
df = df.drop(["artist", "title", "label", "release_date", "release_page"], axis="columns")
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="WoSqwTvlr9ok" outputId="e98846ab-4338-466d-b3ed-0d39f2e8e3a4"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="bzWPu5-WtdN0" outputId="817767b8-4cf2-4096-c4fc-6d29ffb7b973"
df.shape
# + [markdown] id="W4s1zCOVvdBl"
# ## Encoding and scaling data
# + id="akayn3ZKuDcM"
one_hot_encoder = OneHotEncoder().fit(df[["release_format"]])
# + id="hGoBON0Nyc9M"
encoded_data = one_hot_encoder.transform(df[["release_format"]]).todense()
# + id="u5VLUEjTvwKt"
scaler = StandardScaler().fit(
df.drop(["release_format", "price"], axis="columns"))
# + id="VRmyTpPqwHbT"
scaled_data = scaler.transform(
df.drop(["release_format", "price"], axis="columns"))
# + id="6MQuLLqpwTBp"
encoded_features = np.concatenate([encoded_data, scaled_data], axis=-1)
# + id="eXL4CeiPy1r5"
train_features, test_features, train_price, test_price = train_test_split(
encoded_features, df["price"]
)
# + [markdown] id="8QP8jYJ303qK"
# ## Training the model
# + colab={"base_uri": "https://localhost:8080/"} id="grKT6gMJzprv" outputId="32df43fb-0c39-43c8-915d-a838aa58cc85"
model = LinearRegression(fit_intercept=False)
model.fit(train_features, train_price)
# + [markdown] id="4-VmX3y105vt"
# ## Evaluating the model
# + colab={"base_uri": "https://localhost:8080/"} id="s3V8OUV60Ern" outputId="b3f66d92-a928-43f6-db79-8767fe4ff6f5"
score = model.score(test_features, test_price)
predicted = model.predict(test_features)
mae = metrics.mean_absolute_error(test_price, predicted)
print(f"Model score: {score}")
print(f"Mean absolute error: {mae}")
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="2rJofwNp3nE5" outputId="80da7ad0-bc24-4fd3-a7d0-305a61a096d0"
expected = test_price
plt.scatter(expected, predicted)
plt.plot([0, 50], [0, 50], '--k')
plt.axis('tight')
plt.xlabel('True price')
plt.ylabel('Predicted price')
# + [markdown] id="aj1q6k1ZUfZY"
# ## Exporting the models
# + id="edL4kNNzUmbx"
with open("model.pkl", "wb") as f:
pickle.dump(model, f)
# + id="LH8wr4KsUwBj"
with open("one_hot_encoder.pkl", "wb") as f:
pickle.dump(one_hot_encoder, f)
# + id="zJAOuRxJ3qxx"
with open("scaler.pkl", "wb") as f:
pickle.dump(scaler, f)
# + id="A0cIti2UAdzf"
| Preparing_data_and_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
with open('style.css', 'r') as file:
css = file.read()
HTML(css)
# # Solving a Fixed-Point Equation
# Assume we want to find a solution of the equation
# $$ x = \cos(x). $$
# As a first step, we would try to visualize the situation and plot both the function $y = \cos(x)$ and the line $y = x$. This is done in the following cell:
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import math
plt.rcParams['figure.figsize'] = [15, 10]
n = 1000
x = [ 0.5 * math.pi * i / 1000 for i in range(1001)]
y = [math.cos(0.5 * math.pi * i / 1000) for i in range(1001)]
plt.margins(0.00)
plt.plot(x[:640], x[:640], label="y=x")
plt.plot(x, y, label="y = cos(x)")
plt.grid()
plt.xlabel("x")
plt.ylabel("y")
plt.legend()
plt.title("Solve x = cos(x)")
plt.show()
# plt.savefig("xEqualsCosX.pdf")
# -
# The figure above clearly shows that the line $y = x$ intersects the function $y = \cos(x)$ at some point $\bar{x}$ that lies somewhere in the intervall $[0.6, 0.8]$. In order to compute $\bar{x}$, we try the following
# <em style="color:blue">fixed-point iteration</em>.
# <ol>
# <li> We initialize $x_0$ with the value $1.0$. </li>
# <li> We define $x_{n+1} := \cos(x_n)$.
# </ol>
# Our hope is that the sequence $(x_n)_{n\in\mathbb{N}}$ converges to the solution $\bar{x}$, i.e. we hope that
# $$ \lim\limits_{n\rightarrow\infty} x_n = \bar{x}. $$
# Why should this be true? Well let us assume that the limit of this sequence exists and define
# $$ \bar{x} := \lim\limits_{n\rightarrow\infty} x_n. $$
# Then we have the following:
# $$
# \begin{array}{lcl}
# \cos\bigl(\bar{x}\bigr) & = & \cos\Bigl(\lim\limits_{n\rightarrow\infty} x_n\Bigr) \\
# & = & \lim\limits_{n\rightarrow\infty} \cos(x_n) \\
# & = & \lim\limits_{n\rightarrow\infty} x_{n+1} \\
# & = & \lim\limits_{n\rightarrow\infty} x_{n} \\
# & = & \bar{x}
# \end{array}
# $$
# Therefore, if the sequence converges to some limit, ten the limit is indeed a solution of the equation $x = \cos(x)$. The question whether the sequence does indeed converge is decided <em style="color:blue">experimentally</em> as follows:
# +
import math
x = 1.0
old_x = 0.0
i = 1
while abs(x - old_x) >= 4.0E-16:
old_x = x
x = math.cos(x)
print(f'{i} : {x}')
i += 1
# -
# It seems that the sequence $(x_n)_{n\in\mathbb{N}}$ does indeed converge and the solution $\bar{x}$ to the equation $x=\cos(x)$ satisfies
# $$ \bar{x} \approx 0.739085133215161. $$
# Furthermore, up to the precision of *Python's* floating point numbers, the limit $\bar{x}$ satisfies the equation
# $\cos(\bar{x}) = \bar{x}$:
x
math.cos(x)
x - math.cos(x)
# The tiny difference between <tt>x</tt> and <tt>cos(x)</tt> is due to rounding errors.
# Let us try to generalize this approach. The function <tt>solve</tt> shown below takes two arguments:
# - The first argument `f` is a real valued function that takes a single real valued argument.
# - The second argument `x0` is the start value of the fixed-point iteration.</li>
#
# The function tries to solve the equation:
# $$x = f(x) $$
#
# This is done by defining a sequence
# - $x_1 = f(x_0)$,
# - $x_2 = f(x_1)$, and, in general,
# - $x_{n+1} = f(x_n)$.
#
# If this sequence converges, i.e. if the absolute value of the difference $x_{n+1} - x_n$ is less than
# $10^{-15}$, then the value $x_{n+1}$ is returned. If the function does not converge within $10000$ steps, the special value
# `None` is returned instead. This value is always returned if a function terminates without explicitly returning a value.
def solve(f, x0):
"""
Solve the equation f(x) = x using a fixed point iteration.
x0 is the start value.
"""
x = x0
for n in range(10000): # at most 10000 iterations
oldX = x
x = f(x)
if abs(x - oldX) < 1.0e-15:
return x
print("solution to x = cos(x): ", solve(math.cos, 0));
def f(x):
return 1/(1+x)
solve(f, 0)
# Below, the expression `lambda x: 1/(1+x)` defines the function that maps $x$ to the value $\frac{1}{1+x}$.
print("solution to x = 1/(1+x):", solve(lambda x: 1/(1+x), 0));
| Python/Fixed-Point-Iteration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# Download data from google
# +
url = 'https://www.gstatic.com/covid19/mobility/Region_Mobility_Report_CSVs.zip'
r = requests.get(url, allow_redirects=True)
open('Region_Mobility_Report_CSVs.zip', 'wb').write(r.content)
# -
# Unzip files
import zipfile
import os
if not os.path.exists("CSVs"):
os.mkdir("CSVs")
with zipfile.ZipFile('Region_Mobility_Report_CSVs.zip', 'r') as zip_ref:
zip_ref.extractall("CSVs")
import pandas as pd
data = pd.read_csv("CSVs/2020_DE_Region_Mobility_Report.csv")
import math
germany=data[data["sub_region_1"].isnull()]
germany.set_index("date")
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
import matplotlib.style as style
style.use('seaborn-poster')
style.use('ggplot')
# +
interesting_colums=[]
for label, _ in germany.items():
if "_baseline" in label:
interesting_colums.append(label)
smoothing=7
for columnname in interesting_colums:
plt.plot(germany["date"],germany[columnname].rolling(smoothing).mean(),label=columnname.split("_")[0])
plt.xticks(rotation='vertical')
plt.ylabel("percent change from baseline ({} day average)".format(smoothing))
plt.gca().xaxis.set_major_locator(plt.MultipleLocator(15))
plt.legend()
#plt.savefig("mobility.svg")
plt.show()
# +
import numpy as np
stack=[]
names=[]
for columnname in interesting_colums:
stack.append(germany[columnname].rolling(smoothing).mean())
names.append(columnname.split("_")[0])
stack=np.array(stack)
plt.stackplot(germany["date"],stack,labels=names)
plt.xticks(rotation='vertical')
plt.ylabel("percent change from baseline ({} day average)".format(smoothing))
plt.gca().xaxis.set_major_locator(plt.MultipleLocator(15))
plt.legend(loc='upper left')
plt.show()
# -
| MobilityReport.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: website
# language: python
# name: website
# ---
# # Non-Profits & Tech
# > "Leveling the playing field"
#
# - toc: true
# - branch: master
# - categories: [data, non-profits, civic tech]
# - image: images/posts/datajawn/bees.jpg
# - badges: false
# - hide: false
#hide
import pandas as pd
import numpy as np
import altair as alt
# There is a growing disparity in access to tech, and in turn, access to data. You have to have a certain income to afford a smart phone and a level of understanding to sign up for tech services, yes. But I'm particularly interested in the disparity in access to tech _across organizations_. Large, for-profit hospitals can afford more expensive electronic health records and billing tools than a small non-profit organization that serves the homeless. This disparity means that organizations serving individuals with low socioeconomic status have limited access to tools that can automate repeating processes, build basic reports, send data extracts to partners and funders, and generally spend less time entering and processing data.
# In the tech world, the folks programming and building the actual product generally don't think about costs (outside of developer time). For companies who make money from the product the technologists are building, success is determined by revenue.
#
# 
#
# A thriving charity's first success is being able to afford basic operations. A non-proft has "made it" when they can say "what else can we pay for".
#
# We're going to review the cost of different technical solutions non-profits might use, how things like grant structures prohibit them from engaging technology, how that keeps them from doing what they want, and finally what technologists can do to fill the gap and support them.
#
# # How to Read this Guide
# **This post is going to review the cost of different technical solutions non-profits might use, how things like grant structures prohibit them from engaging technology, how that keeps them from increasing efficiency, and finally what technologists can do to fill the gap and support them.**
#
# For non-profits and government agencies, there's not a lot of passive (legal) revenue to the organization, so we're gonna cover that choose-your-own-adventure style.
#
# If you are in tech, start here. If you are a non-profit, I'm going assume you're familiar with funding. Start [further down](#SaaS-Me-If-You-Can).
#
# # Making money the hard way
# ## A primer for folks who haven't hustled to avoid revenue
#
# Let's start with the assumption that all non-profits need some amount of funding. The most common methods for getting these funds are:
# 1. Donations
# 2. Grants
# 3. Services
#
# ### Donations
# For the sake of not letting myself ramble, we're going to focus on financial donations. Many non-profits exist on the model of donations of goods (like Goodwill), selling those goods, and then generating operating costs from those goods. It's a financial model. It exists. The end.
#
# Non-profit organizations normally accept financial donations, but simply relying on someone to wake up and think "you know what, I'm going to send a check to United Way today" is probably not going to result in a reliable stream of income, if any income at all. If a non-profit wants a reliable amount of funds from donations, they need a fundraising campaign. Fundraising campaigns are not free - if Susan G. Komen spends $200,000 buying fancy pink shoes for football players and raises $800,000 (in an imaginary land in which you can directly attribute all financial donations to a single campaign), they're actually raising $600,000. They spent 25 cents for every dollar raised. In order to include this money in an annual budget, the non-profit needs to hold regular fundraising campaigns. This is why most non-profits will employ a Fundraising Coordinator or Marketing Specialist. If they plan on relying on donations as a revenue stream, they need to efficiently and consistently fundraise.
#
# ### Services
# Some non-profits receive funding for their programs through direct services given to members. For example, a non-profit that offers Medication Assisted Treatment may submit insurance claims for participants who have received services through the non-profit. Most insurance companies will have set money amounts for each procedure reported on each claim.
#
# Services may also come in the form of goods, like yellow silicone bracelets that are $3 beacons of virtue...
#
# >twitter: https://twitter.com/skinnamarieke/status/1171062154499411968
#
# ### Grants
# Grants are a sum of money given to an organization for a specific purpose. Grants come in two basic flavors: a sweet but tangy public grant or an earthy, umami private grant. In my experience, private grants are a little less stringent with their requirements but here's a basic breakdown of how to receive and use grant funding.
#
# **Step 1: Funding Opportunity Announcement (FOA)**
# Grant agency puts out an FOA (it's the government, did you think there wasn't an acronym?) that specifies what the grant is for, how much you could receive, who is eligible, timeline, and how to apply. Federal FOA's are almost humorously specific, like this "[Small Grants Program for Leading English Discussion Groups in Sapporo, Japan](https://www.grants.gov/web/grants/search-grants.html)"
#
# **Step 2: Application and Award**
# Agency seeking funding writes up their grant application (which usually is about the size of a graduate thesis) including a need statement, program narrative, proposed budget (or a budget narrative), discussion of impact, blood type, head shots, and $20 worth of pennies from 1960. Some of those are a joke but grant applicants have to be really careful what they include in their applications, using specific language, as the submissions are filtered through an [automated](https://www.nsf.gov/pubs/policydocs/pappg18_1/pappg_2.jsp#IIC2) check initally.
#
# **Step 3: Administration**
# Remember that grant budget you put in your application? Now you have to stick to it. The grant will specify how much money is allotted for each component: personnel, services, materials, etc. Grant awardees are required to not only stick to these budgets, but also report weekly to the funding agency.
#
# For multi-year grants, awardees are oftentimes required to re-apply (with the understanding that they will still receive the grant). Funding agencies may adjust the amount of funds given to awardees at this time.
#
#
#
# # Identifying and Adopting Tech, An Example
# ## SaaS Me If You Can
# Back in the day, I was the data manager for two federal grants that were implemented by a state government. Our grant provided direct services for citizens, so a portion of the grant was earmarked for those costs. We were required to spend a certain percentage of grant funds on direct services.
#
# The program I worked for required a 60/40 split when it came to direct services. This means we had to spend at least 60% of our allotted money on paying for medical services for our participants. The grant also required that we report very specific data elements from those services.
#
# Because the grant was federally funded but implemented at the state level, the program was responsible for both salary and benefits for all employees. The Program was also responsible for paying an administrative fee to the state - for using state resources (internet, office space, legal services, etc).
#
# For the sake of easy math, let's say the program was granted $1,000,000.
#
# **_Charts are interactive, so you can click or hover for more info._**
# +
#hide-input
breakdown = pd.DataFrame({'Resource':['Program Administrator','Data Manager', 'Case Manager','Program Specialist','Program Coordinator', 'Administrative', 'Direct Services'],
'Budget': [95000, 85000, 45000,75000,75000,20000,600000,]
})
breakdown["Program"] = "Program"
budget = alt.Chart(breakdown).mark_bar().encode(
x=alt.X('Budget', title='Program Budget Breakdown'),
y=alt.Y('Program', axis=alt.Axis(labels=False, title ='')),
color=alt.Color('Resource', legend=None),
tooltip=['Resource', 'Budget']
)
budget
# -
# The grant requires that we find individuals in need of services, so we'll need a Program Coordinator. It also requires a Program Specialist to engage with the organizations that provide the service and receive data reports from those organizations. This person should understand what data is required, how it comes in, will probably need to enter that data, and coordinate with service providers and participants regularly. The grant requires a Data Manager for bi-annual reporting of Minimum Data Elements (MDEs) identified and verified by the funding agency and ad-hoc program evaluation. The funding agency also requires regular reporting of progress and a specific adminstration of the grant, thus requiring a Program Administrator. Because the staff for the program will need materials and space, there is an annual administrative budget. This fits exactly into the $1,000,000 allotted.
# # How important is tech?
#
# Based on this budget breakdown, there is no room for paying for developers or third party tooling.
#
# But is it necessary? How much is tech used nowadays anyways? Are there any industries that don't need technology?
# I pulled [2019 data from the Bureau of Labor Statistics](https://www.bls.gov/data/) to see what percentage of each major industry is technologists.
#hide
jobs = pd.read_csv("data/oes_all_x_tech.csv", encoding='utf-8')
jobs['Percent Tech'] = round((jobs['Technologists']/jobs['total_emp']),4)
# +
#hide-input
horizontal_stripplot = alt.Chart(jobs).mark_circle(size = 100).encode(
x=alt.X('Percent Tech:Q', axis=alt.Axis(format='.0%'), title='% Tech Employees' ),
color=alt.Color('Industry:N', legend=None, ),
tooltip=[alt.Tooltip('Industry:N'), alt.Tooltip('Percent Tech:Q', format='.2%')]
).configure_facet(
spacing=0
).configure_view(
stroke=None
).configure_axis(
labelFontSize=16,
titleFontSize=16
).configure_axisY(maxExtent=55)
horizontal_stripplot
# -
# Many of the outliers make sense here - the Information industry is just "Data Processing, Hosting, and Related Services". Professional, Scientific, and Technical Services includes "Computer Systems Design and Related Services" companies which need developers to create their revenue stream.
#
# Within these categories, there's Health Care and Social Assistance, with technologists composing 0.69% of its workforce, and Other Services, with 0.94%. _Health Care_ includes hospitals, medical providers, and emergency services. _Social Assistance_ is mostly the companies we're looking for here: Individual and Family Services, Community Food and Housing, and Emergency and Other Relief Services. _Other services_ includes Grantmaking and Giving Services, Social Advocacy Organizations, and Business, Professional, Labor, Political, and Similar Organizations.
# ### Non-Profit Industries
# If you break out the Health Care and Other Services industries to pinpoint non-profit organizations, you find Grantmaking & Giving Services (3.2% Tech), Social Assistance (0.2% Tech), and Social Advocacy Organizations (2% Tech). Even with the highest percentage of tech employees, Grantmaking & Giving Services pales in comparison to their close relative Finance & Insurance (7.3%).
#
# # The Cost of Tech
# ## Tech Debt
#hide
detail = pd.read_csv("data/nonprofits_detail.csv", encoding='utf-8')
salary = jobs[['Industry', 'annual_salary_tech', 'annual_salary_all']]
salary = salary.rename(columns={'Industry':'Industry', 'annual_salary_tech':'Tech', 'annual_salary_all':'All'})
melted = pd.melt(salary.reset_index(), id_vars=['Industry'], value_vars=['Tech', 'All'], var_name='Occupation', value_name='Average Salary')
# Clearly, tech contributes to every industry in the US. This is reasonable. Let's say you want to buy a shirt from a store but you can't get to the store. Remember the days of catalog ordering? How much more work is it to mail or call an order in? Imagine managing all of those orders without an inventory or order management system. The benefit of tech is automating mundane and repetitive activities. Every organization needs that!
#
# **So why isn't it happening with non-profits?**
#
# ### The Johari Window
#
# The first project I worked on with <NAME> was actually born from a desire to do some ad-hoc data analysis for a local non-profit I love. This non-profit has two dozen separate programs, providing medical and non-medical services to a criminally underserved population. These services are paid for by a combination of donations, grants, and direct service models. When I met with them, I was hoping that they'd be able to just zip together a de-identified extract of their data to do some work on.
#
# Once I started talking with them, I realized that they stored their data in excel spreadsheets, partner systems, and paper forms. Because they didn't have access to dynamic tech solutions, they couldn't track a participant's activities across multiple programs (or even within a program).
#
# This leads me to the **Johari Window**.
#
# 
#
#
# Generally, this is an intrapersonal principle but it applies here. When I was a data manager for a grant program, I knew what tech could do for us even if I couldn't build it myself (known-unknowns). Companies that have tech teams know what they can build and have the capacity to build it (known-knowns). Even organizations that know they need to collect data, and there are tools that do it, but may be a bit lost when it comes to implementing them are empowered to reach out for help (unknown-knowns). When tech solutions are unknown-unknowns, not only is there an inability to know what you can ask for but even knowing that it can be asked!
#
# **If you don't have someone at the table who knows what you can do with tech, your organization won't know how to leverage tech to improve their process.**
# Let's circle back to [the example I laid out earlier](#SaaS-Me-If-You-Can).
# I was the only "tech" person on the team working on those programs. My plate was full managing the data requirements of both the program and the grant, so I had no extra bandwidth (and definitely did not have the prowess) to develop a tool to manage all of our data for free.
#
# The state employed an IT department, passionate and kind folks who worked to develop and manage external and in-house solutions but who were overwhelmed with the needs of each siloed program.
#
# _What is the cheapest way to improve access to technology?_
# ## In-House Team
#
# Using the same BLS data, I looked at how much it costs to employ a technologist.
#
# On average, technologists in any industry make significantly more than an average employee in that industry.
# +
#hide-input
horizontal_stripplot = alt.Chart(melted).mark_circle(size = 100).encode(
x=alt.X('Average Salary:Q', axis=alt.Axis(title='Average Annual Salary'), scale=alt.Scale(zero=False)),
y=alt.Y('Occupation:N', axis=alt.Axis(title=None)),
color=alt.Color('Industry:N', legend=None, ),
tooltip=[alt.Tooltip('Industry:N'), alt.Tooltip('Average Salary:Q')]
).configure_view(
stroke=None
).configure_axis(
labelFontSize=16,
titleFontSize=16
).configure_view(continuousHeight=600, continuousWidth=400)
horizontal_stripplot
# -
# This is not limited to private industry - tech salaries are consistently higher across all industries!
#
# Developing a tech solution usually requires more than one technologist! If you want to build even a simple app, you need someone to develop it and someone to check the work!
# ## OOTB Solutions
#
# In the example, every state had a program funded by one of our grants and 22 states had a program funded by the other grant. Because the data we were required to report to the CDC (visit level medical information for each participant) was discrete and consistent, a third party company had actually created an Out-of-the-Box SaaS app for data collection.
#
# This solution was not free, unfortunately.
#
# In the first year, we had to pay $150,000 to have the solution set up. This included data migration from our historical systems to the new system and licensing fees for each program.
#
# After the initial setup, we had to pay $25,000 for licensing and hosting. We also had to create a slush fund ($20,000) for ad-hoc development fees.
#
# That means that the app would cost us $330,000 over 5 years! If all of the grant money is accounted for, how will the program drum up an extra $330,000?
#
# ### Customization
#
# Even though this solution was was built _specifically_ for our grants, the third party provider **still** recommended that we earmark $20,000 per year for customization. If you are a program that offers both direct services and general services, who collects data both at the time of service and may need to enter data at a later time, if you need to be able to control access to data at system, form, and program levels; you will have a lot of customization work.
#
# In this scenario, an OOTB solution may incur more costs than hiring internally. For example, a non-profit looking to use Salesforce but needing specific data fields may incur a lot of customization-related costs just to ensure they are meeting grant requirements!
# # How Tech Helps
#
# Why should you care about this? Working in tech isn't perfect, but it's a stable form of employment that pays very well, is in high demand, and allows a lot of us introverts to just solve problems all day! It's a great gig.
#
# In the interest of keeping this brief, here are three arguments for increasing non-profit and government access to up-to-date technologies.
#
# ## Better Data
# We are recognizing the economic divide in the data created by modern tech. Tech solutions are built to make _money_. There's no money to be made from populations with lower socioeconomic status. This means the user data for those solutions does not represent an entire swath of the population! If you want to build predictive tooling for an entire population, you need equal amounts of data to **represent the entire population**.
#
# ## Better Services
# If a non-profit is spending 4 hours every week cleaning and preparing data for direct service claims when they can spend 5 minutes making that extract in a BI tool, imagine what they would then do with the extra 203 hours each year. Non-profits that offer individual services but use tools like Excel cannot easily see a participant's journey within a program, nonetheless within the organization. Better tech means better service.
#
# Non-profits can operate with a similar budget! By automating simple processes, they have more capacity to do the work that makes their jobs rewarding!
#
# ## Better Knowledge
# Giving programs access to better data collection tools means we will have access to more complete data on issues we are working to resolve (with limited data!). By encouraging more standardized data collection, we are building capacity for a more complete picture!
#
# # Contribute to Civic Tech
#
# So what can you do?
#
# **As a citizen**
# You may have a local [Code for America brigade](https://brigade.codeforamerica.org/)! Reach out to them. If you don't, start one!
#
# If you're not a technologist, managing a project is absolutely a huge need for these brigades!
#
# **As a non-profit**
# See above :)
#
# **As a tech company**
# Encourage developers to work on these projects! Local companies can donate actual developer time to work on projects - these projects don't happen overnight. Increasing volunteer capacity is just as valuable (and, honestly, more valuable) than a financial donation.
#
# Also, you can do pro bono work yourself! Are there any non-profits that can benefit from your product? Can you help them set it up and then offer them access to it for very cheap or free?
#
# **As a decision maker**
# MAKE TECH-SPECIFIC GRANT FUNDING. STOP SILOING WORK FOR STATE AND CITY LEVEL PROJECTS.
| _notebooks/2020-06-15-non-profits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="https://ithaka-labs.s3.amazonaws.com/static-files/images/tdm/tdmdocs/CC_BY.png"><br />
#
# Created by [<NAME>](http://nkelber.com) and Ted Lawless for [JSTOR Labs](https://labs.jstor.org/) under [Creative Commons CC BY License](https://creativecommons.org/licenses/by/4.0/)<br />
# For questions/comments/improvements, email <EMAIL>.<br />
# ___
#
# # Finding Significant Words Using TF/IDF
#
# **Description:**
# This [notebook](https://docs.constellate.org/key-terms/#jupyter-notebook) shows how to discover significant words. The method for finding significant terms is [tf-idf](https://docs.constellate.org/key-terms/#tf-idf). The following processes are described:
#
# * An educational overview of TF-IDF, including how it is calculated
# * Using the `tdm_client` to retrieve a dataset
# * Filtering based on a pre-processed ID list
# * Filtering based on a [stop words list](https://docs.constellate.org/key-terms/#stop-words)
# * Cleaning the tokens in the dataset
# * Creating a [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary)
# * Creating a [gensim](https://docs.constellate.org/key-terms/#gensim) [bag of words](https://docs.constellate.org/key-terms/#bag-of-words) [corpus](https://docs.constellate.org/key-terms/#corpus)
# * Computing the most significant words in your [corpus](https://docs.constellate.org/key-terms/#corpus) using [gensim](https://docs.constellate.org/key-terms/#gensim) implementation of [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf)
#
# **Use Case:** For Learners (Detailed explanation, not ideal for researchers)
#
# [Take me to the **Research Version** of this notebook ->](./finding-significant-terms-for-research.ipynb)
#
# **Difficulty:** Intermediate
#
# **Completion time:** 60 minutes
#
# **Knowledge Required:**
# * Python Basics Series ([Start Python Basics I](./python-basics-1.ipynb))
#
# **Knowledge Recommended:**
# * [Exploring Metadata](./metadata.ipynb)
# * [Working with Dataset Files](./working-with-dataset-files.ipynb)
# * [Pandas I](./pandas-1.ipynb)
# * [Creating a Stopwords List](./creating-stopwords-list.ipynb)
# * A familiarity with [gensim](https://docs.constellate.org/key-terms/#gensim) is helpful but not required.
#
# **Data Format:** [JSON Lines (.jsonl)](https://docs.constellate.org/key-terms/#jsonl)
#
# **Libraries Used:**
# * `pandas` to load a preprocessing list
# * `csv` to load a custom stopwords list
# * [gensim](https://docs.constellate.org/key-terms/#gensim) to help compute the [tf-idf](https://docs.constellate.org/key-terms/#tf-idf) calculations
# * [NLTK](https://docs.constellate.org/key-terms/#nltk) to create a stopwords list (if no list is supplied)
#
# **Research Pipeline:**
#
# 1. Build a dataset
# 2. Create a "Pre-Processing CSV" with [Exploring Metadata](./exploring-metadata.ipynb) (Optional)
# 3. Create a "Custom Stopwords List" with [Creating a Stopwords List](./creating-stopwords-list.ipynb) (Optional)
# 4. Complete the TF-IDF analysis with this notebook
# ____
# ## What is "Term Frequency- Inverse Document Frequency" (TF-IDF)?
#
# [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) is used in [machine learning](https://docs.constellate.org/key-terms/#machine-learning) and [natural language processing](https://docs.constellate.org/key-terms//#nlp) for measuring the significance of particular terms for a given document. It consists of two parts that are multiplied together:
#
# 1. Term Frequency- A measure of how many times a given word appears in a document
# 2. Inverse Document Frequency- A measure of how many times the same word occurs in other documents within the corpus
#
# If we were to merely consider [word frequency](https://docs.constellate.org/key-terms/#word-frequency), the most frequent words would be common [function words](https://docs.constellate.org/key-terms/#function-words) like: "the", "and", "of". We could use a [stopwords list](https://docs.constellate.org/key-terms/#stop-words) to remove the common [function words](https://docs.constellate.org/key-terms/#function-words), but that still may not give us results that describe the unique terms in the document since the uniqueness of terms depends on the context of a larger body of documents. In other words, the same term could be significant or insignificant depending on the context. Consider these examples:
#
# * Given a set of scientific journal articles in biology, the term "lab" may not be significant since biologists often rely on and mention labs in their research. However, if the term "lab" were to occur frequently in a history or English article, then it is likely to be significant since humanities articles rarely discuss labs.
# * If we were to look at thousands of articles in literary studies, then the term "postcolonial" may be significant for any given article. However, if were to look at a few hundred articles on the topic of "the global south," then the term "postcolonial" may occur so frequently that it is not a significant way to differentiate between the articles.
#
# The [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) calculation reveals the words that are frequent in this document **yet rare in other documents**. The goal is to find out what is unique or remarkable about a document given the context (and *the given context* can change the results of the analysis).
#
# Here is how the calculation is mathematically written:
#
# $$tfidf_{t,d} = tf_{t,d} \cdot idf_{t,D}$$
#
# In plain English, this means: **The value of [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) is the product (or multiplication) of a given term's frequency multiplied by its inverse document frequency.** Let's unpack these terms one at a time.
#
# ### Term Frequency Function
#
# $$tf_{t,d}$$
# The number of times (t) a term occurs in a given document (d)
#
# ### Inverse Document Frequency Function
#
# $$idf_i = \mbox{log} \frac{N}{|{d : t_i \in d}|}$$
# The inverse document frequency can be expanded to the calculation on the right. In plain English, this means: **The log of the total number of documents (N) divided by the number of documents that contain the term**
#
# ### TF-IDF Calculation in Plain English
#
# $$(Times-the-word-occurs-in-given-document) \cdot \mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# There are variations on the [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) formula, but this is the most widely-used version.
# ### An Example Calculation of TF-IDF
#
# Let's take a look at an example to illustrate the fundamentals of [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf). First, we need several texts to compare. Our texts will be very simple.
#
# * text1 = 'The grass was green and spread out the distance like the sea.'
# * text2 = 'Green eggs and ham were spread out like the book.'
# * text3 = 'Green sailors were met like the sea met troubles.'
# * text4 = 'The grass was green.'
#
# The first step is we need to discover how many unique words are in each text.
#
# |text1|text2|text3|text4|
# | --- | ---| --- | --- |
# |the|green|green|the|
# |grass|eggs|sailors|grass|
# |was|and|were|was|
# |green|ham|met|green|
# |and|were|like| |
# |spread|spread|the| |
# |out|out|sea| |
# |into|like|met| |
# |distance|the|troubles| |
# |like|book| | |
# |sea| | | |
#
#
# Our four texts share some similar words. Next, we create a single list of unique words that occur across all three texts. (When we use the [gensim](https://docs.constellate.org/key-terms/#gensim) library later, we will call this list a [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary).)
#
# |id|Unique Words|
# |---| --- |
# |0|and|
# |1|book|
# |2|distance|
# |3|eggs|
# |4|grass|
# |5|green|
# |6|ham|
# |7|like|
# |8|met|
# |9|out|
# |10|sailors|
# |11|sea|
# |12|spread|
# |13|the|
# |14|troubles|
# |15|was|
# |16|were|
#
# Now let's count the occurences of each unique word in each sentence
#
# |id|word|text1|text2|text3|text4|
# |---|---|---|---|---|---|
# |0|and|1|1|0|0|
# |1|book|0|1|0|0|
# |2|distance|1|0|0|0|
# |3|eggs|0|1|0|0|
# |4|grass|1|0|0|1|
# |5|green|1|1|1|1|
# |6|ham|0|1|0|0|
# |7|like|1|1|1|0|
# |8|met|0|0|2|0|
# |9|out|1|1|0|0|
# |10|sailors|0|0|1|0|
# |11|sea|1|0|1|0|
# |12|spread|1|1|0|0|
# |13|the|3|1|1|1|
# |14|troubles|0|0|1|0|
# |15|was|1|0|0|1|
# |16|were|0|1|1|0|
# ### Computing TF-IDF (Example 1)
#
# We have enough information now to compute [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) for every word in our corpus. Recall the plain English formula.
#
# $$(Times-the-word-occurs-in-given-document) \cdot \mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# We can use the formula to compute [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) for the most common word in our corpus: 'the'. In total, we will compute [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) four times (once for each of our texts).
#
# |id|word|text1|text2|text3|text4|
# |---|---|---|---|---|---|
# |13|the|3|1|1|1|
#
# text1: $$ tf-idf = 3 \cdot \mbox{log} \frac{4}{(4)} = 3 \cdot \mbox{log} 1 = 3 \cdot 0 = 0$$
# text2: $$ tf-idf = 1 \cdot \mbox{log} \frac{4}{(4)} = 1 \cdot \mbox{log} 1 = 1 \cdot 0 = 0$$
# text3: $$ tf-idf = 1 \cdot \mbox{log} \frac{4}{(4)} = 1 \cdot \mbox{log} 1 = 1 \cdot 0 = 0$$
# text4: $$ tf-idf = 1 \cdot \mbox{log} \frac{4}{(4)} = 1 \cdot \mbox{log} 1 = 1 \cdot 0 = 0$$
#
# The results of our analysis suggest 'the' has a weight of 0 in every document. The word 'the' exists in all of our documents, and therefore it is not a significant term to differentiate one document from another.
#
# Given that idf is
#
# $$\mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# and
#
# $$\mbox{log} 1 = 0$$
# we can see that [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) will be 0 for any word that occurs in every document. That is, if a word occurs in every document, then it is not a significant term for any individual document.
# ### Computing TF-IDF (Example 2)
#
# Let's try a second example with the word 'out'. Recall the plain English formula.
#
# $$(Times-the-word-occurs-in-given-document) \cdot \mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# We will compute [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) four times, once for each of our texts.
#
# |id|word|text1|text2|text3|text4|
# |---|---|---|---|---|---|
# |9|out|1|1|0|0|
#
# text1: $$ tf-idf = 1 \cdot \mbox{log} \frac{4}{(2)} = 1 \cdot \mbox{log} 2 = 1 \cdot .3010 = .3010$$
# text2: $$ tf-idf = 1 \cdot \mbox{log} \frac{4}{(2)} = 1 \cdot \mbox{log} 2 = 1 \cdot .3010 = .3010$$
# text3: $$ tf-idf = 0 \cdot \mbox{log} \frac{4}{(2)} = 0 \cdot \mbox{log} 2 = 0 \cdot .3010 = 0$$
# text4: $$ tf-idf = 0 \cdot \mbox{log} \frac{4}{(2)} = 0 \cdot \mbox{log} 2 = 0 \cdot .3010 = 0$$
#
# The results of our analysis suggest 'out' has some significance in text1 and text2, but no significance for text3 and text4 where the word does not occur.
# ### Computing TF-IDF (Example 3)
#
# Let's try one last example with the word 'met'. Here's the [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) formula again:
#
# $$(Times-the-word-occurs-in-given-document) \cdot \mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# And here's how many times the word 'met' occurs in each text.
#
# |id|word|text1|text2|text3|text4|
# |---|---|---|---|---|---|
# |8|met|0|0|2|0|
#
# text1: $$ tf-idf = 0 \cdot \mbox{log} \frac{4}{(1)} = 0 \cdot \mbox{log} 4 = 1 \cdot .6021 = 0$$
# text2: $$ tf-idf = 0 \cdot \mbox{log} \frac{4}{(1)} = 0 \cdot \mbox{log} 4 = 1 \cdot .6021 = 0$$
# text3: $$ tf-idf = 2 \cdot \mbox{log} \frac{4}{(1)} = 2 \cdot \mbox{log} 4 = 2 \cdot .6021 = 1.2042$$
# text4: $$ tf-idf = 0 \cdot \mbox{log} \frac{4}{(1)} = 0 \cdot \mbox{log} 4 = 1 \cdot .6021 = 0$$
#
# As should be expected, we can see that the word 'met' is very significant in text3 but not significant in any other text since it does not occur in any other text.
# ### The Full TF-IDF Example Table
#
# Here are the original sentences for each text:
#
# * text1 = 'The grass was green and spread out the distance like the sea.'
# * text2 = 'Green eggs and ham were spread out like the book.'
# * text3 = 'Green sailors were met like the sea met troubles.'
# * text4 = 'The grass was green.'
#
# And here's the corresponding [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) scores for each word in each text:
#
# |id|word|text1|text2|text3|text4|
# |---|---|---|---|---|---|
# |0|and|.3010|.3010|0|0|
# |1|book|0|.6021|0|0|
# |2|distance|.6021|0|0|0|
# |3|eggs|0|.6021|0|0|
# |4|grass|.3010|0|0|.3010|
# |5|green|0|0|0|0|
# |6|ham|0|.6021|0|0|
# |7|like|.1249|.1249|.1249|0|
# |8|met|0|0|1.2042|0|
# |9|out|.3010|.3010|0|0|
# |10|sailors|0|0|.6021|0|
# |11|sea|.3010|0|.3010|0|
# |12|spread|.3010|.3010|0|0|
# |13|the|0|0|0|0|
# |14|troubles|0|0|.6021|0|
# |15|was|.3010|0|0|.3010|
# |16|were|0|.3010|.3010|0|
#
# There are a few noteworthy things in this data.
#
# * The [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) score for any word that does not occur in a text is 0.
# * The scores for almost every word in text4 are 0 since it is a shorter version of text1. There are no unique words in text4 since text1 contains all the same words. It is also a short text which means that there are only four words to consider. The words 'the' and 'green' occur in every text, leaving only 'was' and 'grass' which are also found in text1.
# * The words 'book', 'eggs', and 'ham' are significant in text2 since they only occur in that text.
#
# Now that you have a basic understanding of how [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) is computed at a small scale, let's try computing [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) on a [corpus](https://docs.constellate.org/key-terms/#corpus) which could contain millions of words.
#
# ---
# ## Computing TF-IDF with your Dataset
# We'll use the tdm_client library to automatically retrieve the dataset in the JSON file format.
#
# Enter a [dataset ID](https://docs.constellate.org/key-terms/#dataset-ID) in the next code cell.
#
# If you don't have a dataset ID, you can:
# * Use the sample dataset ID already in the code cell
# * [Create a new dataset](https://constellate.org/builder)
# * [Use a dataset ID from other pre-built sample datasets](https://constellate.org/dataset/dashboard)
# Default dataset is "Shakespeare Quarterly," 1950-present
dataset_id = "7e41317e-740f-e86a-4729-20dab492e925"
# Next, import the `tdm_client`, passing the `dataset_id` as an argument using the `get_dataset` method.
# Importing your dataset with a dataset ID
import tdm_client
# Pull in the dataset that matches `dataset_id`
# in the form of a gzipped JSON lines file.
dataset_file = tdm_client.get_dataset(dataset_id)
# ## Apply Pre-Processing Filters (if available)
# If you completed pre-processing with the "Exploring Metadata and Pre-processing" notebook, you can use your CSV file of dataset IDs to automatically filter the dataset. Your pre-processed CSV file must be in the root folder.
# +
# Import a pre-processed CSV file of filtered dataset IDs.
# If you do not have a pre-processed CSV file, the analysis
# will run on the full dataset and may take longer to complete.
import pandas as pd
import os
pre_processed_file_name = f'data/pre-processed_{dataset_id}.csv'
if os.path.exists(pre_processed_file_name):
df = pd.read_csv(pre_processed_file_name)
filtered_id_list = df["id"].tolist()
use_filtered_list = True
print('Pre-Processed CSV found. Successfully read in ' + str(len(df)) + ' documents.')
else:
use_filtered_list = False
print('No pre-processed CSV file found. Full dataset will be used.')
# -
# ## Load Stopwords List
#
# If you have created a stopword list in the stopwords notebook, we will import it here. (You can always modify the CSV file to add or subtract words then reload the list.) Otherwise, we'll load the NLTK [stopwords](https://docs.constellate.org/key-terms/#stop-words) list automatically.
# +
# Load a custom data/stop_words.csv if available
# Otherwise, load the nltk stopwords list in English
# Create an empty Python list to hold the stopwords
stop_words = []
# The filename of the custom data/stop_words.csv file
stopwords_list_filename = 'data/stop_words.csv'
if os.path.exists(stopwords_list_filename):
import csv
with open(stopwords_list_filename, 'r') as f:
stop_words = list(csv.reader(f))[0]
print('Custom stopwords list loaded from CSV')
else:
# Load the NLTK stopwords list
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
print('NLTK stopwords list loaded')
# -
# ## Define a Unigram Processing Function
# In this step, we gather the unigrams. If there is a Pre-Processing Filter, we will only analyze documents from the filtered ID list. We will also process each unigram, assessing them individually. We will complete the following tasks:
#
# * Lowercase all tokens
# * Remove tokens in stopwords list
# * Remove tokens with fewer than 4 characters
# * Remove tokens with non-alphabetic characters
#
# We can define this process in a function.
# +
# Define a function that will process individual tokens
# Only a token that passes through all three `if`
# statements will be returned. A `True` result for
# any `if` statement does not return the token.
def process_token(token):
token = token.lower()
if token in stop_words: # If True, do not return token
return
if len(token) < 4: # If True, do not return token
return
if not(token.isalpha()): # If True, do not return token
return
return token # If all are False, return the lowercased token
# -
# ## Collect lists of Document IDs, Titles, and Unigrams
#
# Next, we process all the unigrams into a list called `documents`. For demonstration purposes, this code runs on a limit of 500 documents, but we can change this to process all the documents. We are also collecting the document titles and ids so we can reference them later.
# +
# Collecting the unigrams and processing them into `documents`
limit = None # Change number of documents being analyzed. Set to `None` to do all documents.
n = 0
documents = []
document_ids = []
document_titles = []
for document in tdm_client.dataset_reader(dataset_file):
processed_document = []
document_id = document['id']
document_title = document['title']
if use_filtered_list is True:
# Skip documents not in our filtered_id_list
if document_id not in filtered_id_list:
continue
document_ids.append(document_id)
document_titles.append(document_title)
unigrams = document.get("unigramCount", [])
for gram, count in unigrams.items():
clean_gram = process_token(gram)
if clean_gram is None:
continue
processed_document.append(clean_gram)
if len(processed_document) > 0:
documents.append(processed_document)
n += 1
if (limit is not None) and (n >= limit):
break
print('Unigrams collected and processed.')
# -
# At this point, we have unigrams collected for all our documents insde the `documents` list variable. Each index of our list is a single document, starting with `documents[0]`. Each document is, in turn, a list with a single stringe for each unigram.
#
# **Note:** As we collect the unigrams for each document, we are simply including them in a list of strings. This is not the same as collecting them into word counts, and we are not using a Counter() object here like the Word Frequencies notebook.
#
# The next cell demonstrates the contents of each item in our `document` list. Essentially,
# +
# Show the unigrams collected for a particular document
# Change n to see another document
n=0
print('Total documents: ', len(documents))
print('Preview of a single document: ', documents[n])
# -
documents[0]
# If we wanted to see word frequencies, we could convert the lists at this point into `Counter()` objects. The next cell demonstrates that operation.
# +
# Convert a given document into a Counter object to determine
# word frequencies count
# Import counter to help count word frequencies
from collections import Counter
# Use n to represent a particular document index
# Change n to see another document
n = 0
word_freq = Counter(documents[n])
word_freq.most_common(25)
# -
# Now that we have all the cleaned unigrams for every document in a list called `documents`, we can use Gensim to compute TF/IDF.
# ---
# ## Using Gensim to Compute "Term Frequency- Inverse Document Frequency"
#
# It will be helpful to remember the basic steps we did in the explanatory [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) example:
#
# 1. Create a list of the frequency of every word in every document
# 2. Create a list of every word in the [corpus](https://docs.constellate.org/key-terms/#corpus)
# 3. Compute [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) based on that data
#
# So far, we have completed the first item by creating a list of the frequency of every word in every document. Now we need to create a list of every word in the corpus. In [gensim](https://docs.constellate.org/key-terms/#gensim), this is called a "dictionary". A [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) is similar to a [Python dictionary](https://docs.constellate.org/key-terms/#python-dictionary), but here it is called a [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) to show it is a specialized kind of dictionary.
#
# ### Creating a Gensim Dictionary
#
# Let's create our [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary). A [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) is a kind of masterlist of all the words across all the documents in our corpus. Each unique word is assigned an ID in the gensim dictionary. The result is a set of key/value pairs of unique tokens and their unique IDs.
import gensim
dictionary = gensim.corpora.Dictionary(documents)
# Now that we have a [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary), we can get a preview that displays the number of unique tokens across all of our texts.
print(dictionary)
# The [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) stores a unique identifier (starting with 0) for every unique token in the corpus. The [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) does not contain information on word frequencies; it only catalogs all the unique words in the corpus. You can see the unique ID for each token in the text using the .token2id() method.
list(dictionary.token2id.items())
# We could also look up the corresponding ID for a token using the ``.get`` method.
# +
# Get the value for the key 'people'. Return 0 if there is no token matching 'people'.
# The number returned is the gensim dictionary ID for the token.
dictionary.token2id.get('people', 0)
# -
# For the sake of example, we could also discover a particular token using just the ID number. This is not something likely to happen in practice, but it serves here as a demonstration of the connection between tokens and their ID number.
#
# Normally, [Python dictionaries](https://docs.constellate.org/key-terms/#python-dictionary) only map from keys to values (not from values to keys). However, we can write a quick for loop to go the other direction. This cell is simply to demonstrate how the [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) is connected to the list entries in the [gensim](https://docs.constellate.org/key-terms/#gensim) ``bow_corpus``.
# +
# Find the token associated with a token id number
token_id = 100
# If the token id matches, print out the associated token
for dict_id, token in dictionary.items():
if dict_id == token_id:
print(token)
# -
# ## Creating a Bag of Words Corpus
#
# The next step is to connect our word frequency data found within ``documents`` to our [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) token IDs. For every document, we want to know how many times a word (notated by its ID) occurs. We will create a [Python list](https://docs.constellate.org/key-terms/#python-list) called ``bow_corpus`` that will turn our word counts into a series of [tuples](https://docs.constellate.org/key-terms/#tuple) where the first number is the [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) token ID and the second number is the word frequency.
#
# 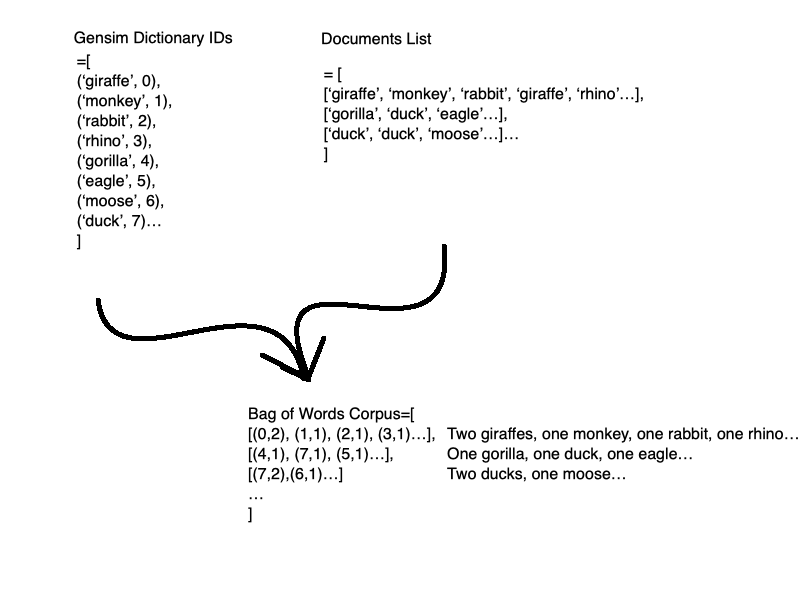
# +
# Create a bag of words corpus
bow_corpus = [dictionary.doc2bow(doc) for doc in documents]
print('Bag of words corpus created successfully.')
# +
# Examine the bag of words corpus for a specific document
# Select a particular document in our documents list
n=0
# List out a slice of the first ten items in that document
list(bow_corpus[n][:10])
# -
# Using IDs can seem a little abstract, but we can discover the word associated with a particular ID. For demonstration purposes, the following code will replace the token IDs in the last example with the actual tokens.
word_counts = [[(dictionary[id], count) for id, count in line] for line in bow_corpus]
list(word_counts[n][:10])
# ## Create the `TfidfModel`
# The next step is to create the [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) model which will set the parameters for our implementation of [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf). In our [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) example, the formula for [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) was:
#
# $$(Times-the-word-occurs-in-given-document) \cdot \mbox{log} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-word)}$$
#
# In [gensim](https://docs.constellate.org/key-terms/#gensim), the default formula for measuring [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) uses log base 2 instead of log base 10, as shown:
#
# $$(Times-the-word-occurs-in-given-document) \cdot \log_{2} \frac{(Total-number-of-documents)}{(Number-of-documents-containing-the-word)}$$
#
# If you would like to use a different formula for your [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) calculation, there is a description of [parameters you can pass](https://radimrehurek.com/gensim/models/tfidfmodel.html).
# Create our gensim TF-IDF model
model = gensim.models.TfidfModel(bow_corpus)
# Now, we apply our model to the ``bow_corpus`` to create our results in ``corpus_tfidf``. The ``corpus_tfidf`` is a python list of each document similar to ``bow_document``. Instead of listing the frequency next to the [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) ID, however, it contains the [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) score for the associated token. Below, we display the first document in ``corpus_tfidf``.
# +
# Create TF-IDF scores for the ``bow_corpus`` using our model
corpus_tfidf = model[bow_corpus]
# List out the TF-IDF scores for the first 10 tokens of the first text in the corpus
list(corpus_tfidf[0][:10])
# -
# Let's display the tokens instead of the [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary) IDs.
# +
example_tfidf_scores = [[(dictionary[id], count) for id, count in line] for line in corpus_tfidf]
# List out the TF-IDF scores for the first 10 tokens of the first text in the corpus
list(example_tfidf_scores[0][:10])
# -
# ## Find Top Terms in a Single Document
# Finally, let's sort the terms by their [TF-IDF](https://docs.constellate.org/key-terms/#tf-idf) weights to find the most significant terms in the document.
# +
# Sort the tuples in our tf-idf scores list
# Choosing a document by its index number
# Change n to see a different document
n = 0
def Sort(tfidf_tuples):
tfidf_tuples.sort(key = lambda x: x[1], reverse=True)
return tfidf_tuples
# Print the document id and title
print('Title: ', document_titles[n])
print('ID: ', document_ids[n])
#List the top twenty tokens in our example document by their TF-IDF scores
list(Sort(example_tfidf_scores[n])[:20])
# -
# We could also analyze across the entire corpus to find the most unique terms. These are terms that appear in a particular text, but rarely or never appear in other texts. (Often, these will be proper names since a particular article may mention a name often but the name may rarely appear in other articles. There's also a fairly good chance these will be typos or errors in optical character recognition.)
# +
# Define a dictionary ``td`` where each document gather
td = {
dictionary.get(_id): value for doc in corpus_tfidf
for _id, value in doc
}
# Sort the items of ``td`` into a new variable ``sorted_td``
# the ``reverse`` starts from highest to lowest
sorted_td = sorted(td.items(), key=lambda kv: kv[1], reverse=True)
# -
for term, weight in sorted_td[:25]: # Print the top 25 terms in the entire corpus
print(term, weight)
# ## Display Most Significant Term for each Document
# We can see the most significant term in every document.
# +
# For each document, print the ID, most significant/unique word, and TF/IDF score
n = 0
for n, doc in enumerate(corpus_tfidf):
if len(doc) < 1:
continue
word_id, score = max(doc, key=lambda x: x[1])
print(document_ids[n], dictionary.get(word_id), score)
if n >= 10:
break
# -
# ## Ranking documents by TF-IDF Score for a Search Word
#
from collections import defaultdict
terms_to_docs = defaultdict(list)
for doc_id, doc in enumerate(corpus_tfidf):
for term_id, value in doc:
term = dictionary.get(term_id)
terms_to_docs[term].append((doc_id, value))
if doc_id >= 500:
break
# +
# Pick a unigram to discover its score across documents
search_term = 'coriolanus'
# Display a list of documents and scores for the search term
matching = terms_to_docs.get(search_term)
for doc_id, score in sorted(matching, key=lambda x: x[1], reverse=True):
print(document_ids[doc_id], score)
# -
| finding-significant-terms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### 1. In Git, branching is the inverse process to:
# ##### Ans: merging
# #### 2. A detailed branching history can be shown by:
# ##### Ans: git show-branch
# #### 3. To examine an earlier version of a file in commit 3888bc981a, do:
# ##### Ans: git show 3888bc981a:kernel/sys.c
# #### 4. You can list all current branches on the local machine with:
# ##### Ans:
# - git branch
# - git branch --list
# - git branch -v
# #### 5. The command git checkout some_branch
# ##### Ans: switches to some_branch
| Coursera/Using Git for Distributed Development/Week-2/Quiz/Branches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/clustering/clustering_gmm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="rnmk5B1V4Wdw"
# # Clustering with Gaussian mixture models
# + id="SSmXaJG75K0A"
# Standard Python libraries
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import time
import numpy as np
import glob
import matplotlib.pyplot as plt
import PIL
import imageio
from IPython import display
import sklearn
import seaborn as sns;
sns.set(style="ticks", color_codes=True)
import pandas as pd
pd.set_option('precision', 2) # 2 decimal places
pd.set_option('display.max_rows', 20)
pd.set_option('display.max_columns', 30)
pd.set_option('display.width', 100) # wide windows
# + [markdown] id="vO95k4h84-iR"
#
#
# In this section, we show how to find clusters in an unlabeled 2d version of the Iris dataset by fitting a GMM using sklearn.
#
#
#
# + id="LiLzt46c45vL" colab={"base_uri": "https://localhost:8080/", "height": 552} outputId="2dd8c6d4-dc0e-4318-b9af-57a08c5492ff"
import seaborn as sns
from sklearn.datasets import load_iris
#from sklearn.cluster import KMeans
from sklearn.mixture import GaussianMixture
iris = load_iris()
X = iris.data
fig, ax = plt.subplots()
idx1 = 2; idx2 = 3;
ax.scatter(X[:, idx1], X[:, idx2], c="k", marker=".")
ax.set(xlabel = iris.feature_names[idx1])
ax.set(ylabel = iris.feature_names[idx2])
#save_fig("iris-2d-unlabeled")
plt.show()
K = 3
y_pred = GaussianMixture(n_components=K, random_state=42).fit(X).predict(X)
mapping = np.array([2, 0, 1])
y_pred = np.array([mapping[cluster_id] for cluster_id in y_pred])
colors = sns.color_palette()[0:K]
markers = ('s', 'x', 'o', '^', 'v')
fig, ax = plt.subplots()
for k in range(0, K):
ax.plot(X[y_pred==k, idx1], X[y_pred==k, idx2], color=colors[k], \
marker=markers[k], linestyle = 'None', label="Cluster {}".format(k))
ax.set(xlabel = iris.feature_names[idx1])
ax.set(ylabel = iris.feature_names[idx2])
plt.legend(loc="upper left", fontsize=12)
#save_fig("iris-2d-gmm")
plt.show()
| notebooks/clustering_gmm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SUM OF INTERVALS
#
# Write a function called `sum_of_intervals` that accepts an array of intervals, and returns the sum of all the interval lengths. Overlapping intervals should only be counted once.
# Intervals are represented by a pair of integers in the form of an array. The first value of the interval will always be less than the second value. Interval example: `[1, 5]` is an interval from 1 to 5. The length of this interval is 4.
#
# List containing overlapping intervals:
#
# `[[1,4], [7, 10], [3, 5]]`
#
# The sum of the lengths of these intervals is 7. Since `[1, 4]` and `[3, 5]` overlap, we can treat the interval as `[1, 5]`, which has a length of 4.
#
# Examples:
#
# `sum_of_intervals([[1,2], [6, 10], [11, 15]]) => 9`
#
# `sum_of_intervals([[1,4], [7, 10], [3, 5]]) => 7`
#
# `sum_of_intervals([[1,5], [10, 20], [1, 6], [16, 19], [5, 11]]) => 19`
# ## Function Definitions
def sum_of_intervals(intervals):
interval_lists_extended = []
for interval in intervals:
interval_lists_extended.extend(list(range(interval[0], interval[1])))
return len(set(interval_lists_extended))
print(sum_of_intervals([[1,2], [6, 10], [11, 15]]) == 9)
print(sum_of_intervals([[1,4], [7, 10], [3, 5]]) == 7)
print(sum_of_intervals([[1,6], [10, 20], [1, 5], [16, 19], [5, 11]]) == 19)
| mk018-sum_of_intervals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
from resources.workspace import *
# # The ensemble (Monte-Carlo) approach
# is an approximate method for doing Bayesian inference. Instead of computing the full posterior distributions, we instead try to generate ensembles from them.
#
# An ensemble is an *iid* sample. I.e. a set of "members" ("particles", "realizations", or "sample points") that have been drawn ("sampled") independently from the same distribution. With the EnKF, these assumptions are generally tenous, but pragmatic.
#
# Ensembles can be used to characterize uncertainty: either by reconstructing (estimating) the distribution from which it is assumed drawn, or by computing various *statistics* such as the mean, median, variance, covariance, skewness, confidence intervals, etc (any function of the ensemble can be seen as a "statistic"). This is illustrated by the code below.
# +
# Parameters
b = 0
B = 25
B12 = sqrt(B)
def true_pdf(x):
return ss.norm.pdf(x,b,sqrt(B))
# Plot true pdf
xx = 3*linspace(-B12,B12,201)
fig, ax = plt.subplots()
ax.plot(xx,true_pdf(xx),label="True");
# Sample and plot ensemble
M = 1 # length of state vector
N = 100 # ensemble size
E = b + B12*randn((N,M))
ax.plot(E, zeros(N), '|k', alpha=0.3, ms=100)
# Plot histogram
nbins = max(10,N//30)
heights, bins, _ = ax.hist(E,normed=1,bins=nbins,label="Histogram estimate")
# Plot parametric estimate
x_bar = np.mean(E)
B_bar = np.var(E)
ax.plot(xx,ss.norm.pdf(xx,x_bar,sqrt(B_bar)),label="Parametric estimate")
ax.legend();
# Uncomment AFTER Exc 4:
# dx = bins[1]-bins[0]
# c = 0.5/sqrt(2*pi*B)
# for height, x in zip(heights,bins):
# ax.add_patch(mpl.patches.Rectangle((x,0),dx,c*height/true_pdf(x+dx/2),alpha=0.3))
# Also set
# * N = 10**4
# * nbins = 50
# -
# **Exc 2:** Which approximation to the true pdf looks better: Histogram or the parametric?
# Does one approximation actually start with more information? The EnKF takes advantage of this.
# **Exc 4*:** Suppose the histogram bars get normalized (divided) by the value of the pdf at their location.
# How do you expect the resulting histogram to look?
# Test your answer by uncommenting the block in the above code.
# #### Exc 5*:
# Use the method of `gaussian_kde` from `scipy.stats` to make a "continuous histogram" and plot it above.
# `gaussian_kde`
# +
#show_answer("KDE")
# -
# **Exc 6 (Multivariate Gaussian sampling):**
# Suppose $\mathbf{z}$ is a standard Gaussian,
# i.e. $p(\mathbf{z}) = \mathcal{N}(\mathbf{z} \mid 0,\mathbf{I}_M)$,
# where $\mathbf{I}_M$ is the $M$-dimensional identity matrix.
# Let $\mathbf{x} = \mathbf{L}\mathbf{z} + \mathbf{b}$.
# Recall [Exc 3.1](T3%20-%20Univariate%20Kalman%20filtering.ipynb#Exc-3.1:),
# which yields $p(\mathbf{x}) = \mathcal{N}(\mathbf{x} \mid \mathbf{b}, \mathbf{L}^{}\mathbf{L}^T)$.
#
# * (a). $\mathbf{z}$ can be sampled using `randn((M,1))`. How (where) is `randn` defined?
# * (b). Consider the above definition of $\mathbf{x}$ and the code below.
# Complete it so as to generate a random realization of $\mathbf{x}$.
# Hint: matrix-vector multiplication can be done using the symbol `@`.
# +
M = 3 # ndim
b = 10*ones(M)
B = diag(1+arange(M))
L = np.linalg.cholesky(B) # B12
print("True mean and cov:")
print(b)
print(B)
### INSERT ANSWER (b) ###
# +
#show_answer('Gaussian sampling a')
# +
#show_answer('Gaussian sampling b')
# -
# * (c). Now sample $N = 100$ realizations of $\mathbf{x}$
# and collect them in an $M$-by-$N$ "ensemble matrix" $\mathbf{E}$.
# Avoid `for` loops (the main thing to figure out is:
# how to add a (mean) vector to a matrix).
# +
N = 100 # ensemble size
E = ### INSERT ANSWER (c) ###
# Use the code below to assess whether you got it right
x_bar = np.mean(E,axis=1)
B_bar = np.cov(E)
print("Estimated mean and cov:")
with printoptions(precision=1):
print(x_bar)
print(B_bar)
plt.matshow(B_bar,cmap="Blues"); plt.grid('off'); plt.colorbar()
# +
#show_answer('Gaussian sampling c')
# -
# **Exc 8*:** How erroneous are the ensemble estimates on average?
# +
#show_answer('Average sampling error')
# -
# **Exc 10:** Given the previous ensemble matrix $\mathbf{E}$, compute its sample mean $\overline{\mathbf{x}}$ and covariance matrix, $\overline{\mathbf{B}}$:
# $$ \overline{\mathbf{x}} = \frac{1}{N} \sum_{n=1}^N \mathbf{x}_n \\
# \overline{\mathbf{B}} = \frac{1}{N-1} \sum_{n=1}^N (\mathbf{x}_n - \overline{\mathbf{x}}) (\mathbf{x}_n - \overline{\mathbf{x}})^T $$
# +
# Don't use numpy's mean, cov
def estimate_mean_and_cov(E):
M, N = E.shape
### INSERT ANSWER ###
return x_bar, B_bar
x_bar, B_bar = estimate_mean_and_cov(E)
print(x_bar)
print(B_bar)
# +
#show_answer('ensemble moments')
# -
# **Exc 12:** Why is the normalization by $(N-1)$ for the covariance computation?
# +
#show_answer('Why (N-1)')
# -
# **Exc 14:** Like Matlab, Python (numpy) is quicker if you "vectorize" loops. This is emminently possible with computations of ensemble moments.
# * (a). Let $\mathbf{X} = \begin{bmatrix}
# \mathbf{x}_1 -\mathbf{\bar{x}}, & \ldots & \mathbf{x}_n -\mathbf{\bar{x}}, & \ldots & \mathbf{x}_N -\mathbf{\bar{x}}
# \end{bmatrix} \, .
# $
# Show that $\overline{\mathbf{B}} = \mathbf{X} \mathbf{X}^T /(N-1)$.
# * (b). Code up this formula for $\overline{\mathbf{B}}$ and insert it in `estimate_mean_and_cov(E)`
# +
#show_answer('ensemble moments vectorized')
# -
# **Exc 16:** Implement the cross-covariance estimator $\overline{Cov(\mathbf{x},\mathbf{y})} = \frac{1}{N-1} \sum_{n=1}^N (\mathbf{x}_n - \overline{\mathbf{x}}) (\mathbf{y}_n - \overline{\mathbf{y}})^T $.
# If you can, use a vectorized form similarly to Exc 14a.
def estimate_cross_cov(Ex,Ey):
### INSERT ANSWER ###
# +
#show_answer('estimate cross')
# -
# **Exc 18 (error notions)*:**
# * (a). What's the difference between error residual?
# * (b). What's the difference between error and bias?
# * (c). Show `MSE = RMSE^2 = Bias^2 + Var`
# +
#show_answer('errors')
# -
# ### Next: [Writing your own EnKF](T8 - Writing your own EnKF.ipynb)
| tutorials/T7 - Ensemble [Monte-Carlo] approach.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="KIgEaDEGw7F4" outputId="d8caf782-867d-4342-9979-5b62de14784d"
from google.colab import drive
drive.mount('/content/drive')
% cd "/content/drive/MyDrive/using-a2c"
# + id="WBBuiXbixIaA"
import optuna
from optuna.visualization import plot_contour
from optuna.visualization import plot_edf
from optuna.visualization import plot_intermediate_values
from optuna.visualization import plot_optimization_history
from optuna.visualization import plot_parallel_coordinate
from optuna.visualization import plot_param_importances
from optuna.visualization import plot_slice
# + id="yU3yps6-5_X4"
study_name = 'delta_coefficient_study'
# + id="dJdHr6P9xbIh"
study = optuna.load_study(study_name=study_name, storage='sqlite:///{}.db'.format(study_name))
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="nnAXGnn4yX2C" outputId="908622db-a79e-4527-abf9-1fa5f417493b"
plot_optimization_history(study)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="89RJvZBGyiLj" outputId="47a8de9e-bcdc-4edf-86dc-da4910d35c9f"
plot_parallel_coordinate(study)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="JWAvejWN5t1G" outputId="2429bae9-0e25-40db-da27-76e4c264fe6b"
plot_contour(study)
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="DlA24fQs54B3" outputId="a467b9d1-76df-4c73-c67f-1aef0e55ecdb"
plot_param_importances(study)
| optuna_HPO_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Probability distributions & meteor shower gazing
#
#
# **Our goals for today:**
# - Discuss some key statistics topics: samples versus populations and empirical versus theorectical distributions
# - Simulate a head/tail coin toss and well drilling i.e. Binomial distribution
# - Simulate meteors entering Earth's atmosphere i.e. Poisson distribution
# - Simulate geomagnetic polarity reversals i.e. Gamma distribution
# - Use Gutenberg-Richter to assess earthquake probability
#
# ## Setup
#
# Run this cell as it is to setup your environment.
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import scipy as scipy
import scipy.stats as sps
# ## Flipping a coin
#
# Let's pretend we are flipping a coin 10 times using ```np.random.choice([0, 1])```. How many times will be get heads? 1 is heads, 0 is tails. Let's use a for loop and get Python to simulate such a coin flip scenario for us.
#
# This code block is the first time we are using a **for loop**. For loops, result in a a chunk of code (in python the chunk that is indented) being run multiple times. In this case, the code will get looped through 10 times -- specified by ```range(0,10)```.
for flip in range(0,10):
flip_result = np.random.choice([0, 1])
print(flip_result)
# Now let's record how many times the result was heads. We will make a list called `flip_results` and have it be blank to start. Each time we go through the code we will append the result to the list:
# +
flip_results = []
for flip in range(0,10):
flip_result = np.random.choice([0, 1])
flip_results.append(...)
flip_results
# -
# We can calculate how many times were heads by taking the sum of the list:
# +
#write code here
# -
# Now let's flip the coin 10 times and do that 10 times. Each time we flip it, let's record how many heads resulted from the flip.
# +
number_heads = []
for flip_experiment in range (0,10):
flip_results = []
for flip in range(0,10):
flip_result = np.random.choice([0, 1])
flip_results.append(flip_result)
number_heads.append(...)
number_heads
# -
plt.hist(number_heads,bins=[-0.5,0.5,1.5,2.5,3.5,4.5,5.5,6.5,7.5,8.5,9.5,10.5],density=True)
plt.show()
# <font color=goldenrod>**_Code for you to write_**</font>
#
# Instead of doing 10 coin flips 10 times, do 10 coin flips 1000 times. Plot the histogram of the result.
# ## Binomial distribution:
#
# ### Theoretical
#
# A relatively straight-forward distribution is the _binomial_ distribution which describes the probability of a particular outcome when there are only two possibilities (yes or no, heads or tails, 1 or 0). For example, in a coin toss experiment (heads or tails), if we flip the coin $n$ times, what is the probability of getting $x$ 'heads'? We assume that the probability $p$ of a head for any given coin toss is 50%; put another way $p$ = 0.5.
#
# The binomial distribution can be described by an equation:
#
# $$P=f(x,p,n)= \frac{n!}{x!(n-x)!}p^x(1-p)^{n-x}$$
#
# We can look at this kind of distribution by evaluating the probability for getting $x$ 'heads' out of $n$ attempts. We'll code the equation as a function, and calculate the probability $P$ of a particular outcome (e.g., $x$ heads in $n$ attempts).
#
# Note that for a coin toss, $p$ is 0.5, but other yes/no questions can be investigated as well (e.g., chance of finding a fossil in a sedimentary layer, whether or not a landslide occurs following an earthquake).
def binomial_probability(x,p,n):
"""
This function computes the probability of getting x particular outcomes (heads) in n attempts, where p is the
probability of a particular outcome (head) for any given attempt (coin toss).
Parameters
----------
x : number of a particular outcome
p : probability of that outcome in a given attempt
n : number of attempts
Returns
---------
prob : probability of that number of the given outcome occuring in that number of attempts
"""
prob = (np.math.factorial(n)/(np.math.factorial(x)*np.math.factorial(n-x)))*(p**(x))*(1.-p)**(n-x)
return prob
# We can use this function to calculate the probability of getting 10 heads ($x=10$) when there are 10 coin tosses ($n=10$) given with the $p$ (probability) of 0.5.
binomial_probability(x=10,p=0.5,n=10)
# Let's calculate the probability of getting [ 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10] heads.
head_numbers = np.arange(0,11)
head_numbers
# +
prob_heads = 0.5
n_flips = 10
probabilities = []
for head_number in head_numbers:
prob = binomial_probability(...)
probabilities.append(prob)
probabilities
# -
# <font color=goldenrod>**_Code for you to write_**</font>
#
# Make a plot where you both plot the histogram from 1000 coin flips (using ```plt.hist()``` with ```density=True```) and you plot the results head_numbers probabilities (using ```plt.plot()```).
# +
plt.hist()
plt.plot()
plt.xlabel('Number of heads out of $n$ attempts') # add labels
plt.ylabel('Fraction of times with this number of heads')
plt.title('Coin flip results (n=10)');
# -
# Hopefully what we should see is that number of coin flips from our random samples matches the theoritical probability distribution pretty well. The more flip experiments we numerically take, the better it should match.
# ### Empirical
#
# The type of sampling we were doing above where we were flipping coins is called a _Monte Carlo simulation_. We can use simulate data from all sorts of distributions. Let's keep focusing on the binomial distribution and look at using the ```np.random.binomial``` function.
help(np.random.binomial)
# `np.random.binomial( )` requires 2 parameters, $n$ and $p$, with an optional keyword argument `size` (if `size` is not specified, it returns a single trial). We could have used this function earlier to get the number of heads that were flipped, but the way we did it also worked.
#
# Let's follow the example the is given in the `np.random.binomial( )` docstring.
#
# A company drills 9 wild-cat oil exploration wells (high risk drilling in unproven areas), each with an estimated probability of success of 0.1. All nine wells fail. What is the probability of that happening? *Note that success in this context means that liquid hydocarbons came out of the well. In reality, you may not consider this a success given that the result is that more hydrocarbons will be combusted as a result, leading to higher atmospheric carbon dioxide levels and associated global warming.*
#
# If we do ```np.random.binomial(9, 0.1, 100)``` we will get a list of 100 values that represent the number of wells that yielded oil when there is a 10% (p = 0.1) chance of each individual well yielding oil.
np.random.binomial(9, 0.1, 100)
np.random.binomial(9, 0.1, 100) == 0
np.sum()
# We can write a function that uses this process to simulate fraction of times that there no successful wells for a given number of wells, a given probability and a given number of simulations;
def wildcat_failure_rate(n_wells,prob,n_simulations):
'''
Simulate the number of times that there are no successful wells for a given number of wells and a given probability for each well.
Parameters
----------
n_wells : number of wells drilled in each simulation
prob : probability that each well will be successful
n_simulations : number of times that drilling n_wells is simulated
'''
failure_rate = sum(np.random.binomial(n_wells, prob, n_simulations) == 0)/n_simulations
return failure_rate
# <font color=goldenrod>**Put the `wildcat_failure_rate` function to use**</font>
#
# Use the function to simulate the failure rate for the above scenario (10 wells drilled, 0.1 probability of success for each well) and do it for 10 simulations
# <font color=goldenrod>**Put the `wildcat_failure_rate` function to use**</font>
#
# Use the function to simulate the failure rate for the same scenario for 1000 simulations
# <font color=goldenrod>**Put the `wildcat_failure_rate` function to use**</font>
#
# Use the function to simulate the failure rate for 100,000 simulations
# <font color=goldenrod>**Put the `binomial_probability` function to use**</font>
#
# In the examples above we are simulating the result. Instead we could use the binomial_probability distribution to calculate the probability. Go ahead and do this for this wildcat drilling example.
binomial_probability()
# **How well does the calculated binomial_probability match the simulated wildcat_failure rates? How many times do you need to simulate the problem to get a number that matches the theoretical probability?**
#
# *Write your answer here*
# ## Poisson distribution:
#
# A Poisson Distribution gives the probability of a number of events in an interval generated by a Poisson process: the average time between events is known, but the exact timing of events is random. The events must be independent and may occur only one at a time.
#
# Within Earth and Planetary Science there are many processes that approximately meet this criteria.
#
# ### Theoretical
#
# The Poisson distribution gives the probability that an event (with two possible outcomes) occurs $k$ number of times in an interval of time where $\lambda$ is the expected rate of occurance. The Poisson distribution is the limit of the binomial distribution for large $n$. So if you take the limit of the binomial distribution as $n \rightarrow \infty$ you'll get the Poisson distribution:
#
# $$P(k) = e^{-\lambda}\frac{\lambda^{k}}{k!}$$
#
def poisson_probability(k,lam):
"""
This function computes the probability of getting k particular outcomes when the expected rate is lam.
"""
# compute the poisson probability of getting k outcomes when the expected rate is lam
prob = (np.exp(-1*lam))*(lam**k)/np.math.factorial(k)
#return the output
return prob
# ## Observing meteors
#
# <img src="./images/AMS_TERMINOLOGY.png" width = 600>
#
# From https://www.amsmeteors.org/meteor-showers/meteor-faq/:
#
# > **How big are most meteoroids? How fast do they travel?** The majority of visible meteors are caused by particles ranging in size from about that of a small pebble down to a grain of sand, and generally weigh less than 1-2 grams. Those of asteroid origin can be composed of dense stony or metallic material (the minority) while those of cometary origin (the majority) have low densities and are composed of a “fluffy” conglomerate of material, frequently called a “dustball.” The brilliant flash of light from a meteor is not caused so much by the meteoroid’s mass, but by its high level of kinetic energy as it collides with the atmosphere.
#
# > Meteors enter the atmosphere at speeds ranging from 11 km/sec (25,000 mph), to 72 km/sec (160,000 mph!). When the meteoroid collides with air molecules, its high level of kinetic energy rapidly ionizes and excites a long, thin column of atmospheric atoms along the meteoroid’s path, creating a flash of light visible from the ground below. This column, or meteor trail, is usually less than 1 meter in diameter, but will be tens of kilometers long.
#
# > The wide range in meteoroid speeds is caused partly by the fact that the Earth itself is traveling at about 30 km/sec (67,000 mph) as it revolves around the sun. On the evening side, or trailing edge of the Earth, meteoroids must catch up to the earth’s atmosphere to cause a meteor, and tend to be slow. On the morning side, or leading edge of the earth, meteoroids can collide head-on with the atmosphere and tend to be fast.
#
# > **What is a meteor shower? Does a shower occur “all at once” or over a period of time?** Most meteor showers have their origins with comets. Each time a comet swings by the sun, it produces copious amounts of meteoroid sized particles which will eventually spread out along the entire orbit of the comet to form a meteoroid “stream.” If the Earth’s orbit and the comet’s orbit intersect at some point, then the Earth will pass through this stream for a few days at roughly the same time each year, encountering a meteor shower. The only major shower clearly shown to be non-cometary is the Geminid shower, which share an orbit with the asteroid (3200 Phaethon): one that comes unusually close to the sun as well as passing through the earth’s orbit. Most shower meteoroids appear to be “fluffy”, but the Geminids are much more durable as might be expected from asteroid fragments.
#
# ## Observing the Southern Taurids meteor shower
#
# Let's say you are planning to go out and try to see shooting stars tonight in a rural location. You might be in luck because there is an active shower:
#
# > **Southern Taurids**
#
# > *Active from September 28th to December 2, 2021. The peak is November 4-5, 2021*
#
# > The Southern Taurids are a long-lasting shower that reaches a barely noticeable maximum on October 9 or 10. The shower is active for more than two months but rarely produces more than five shower members per hour, even at maximum activity. The Taurids (both branches) are rich in fireballs and are often responsible for increased number of fireball reports from September through November. https://www.amsmeteors.org/meteor-showers/meteor-shower-calendar/
#
# At a rate of 5 observed meteors per hour, what is the probability of observing 6?
#
# We can use the Poisson probability function to answer this question:
lamb = 5
k = 6
prob = poisson_probability(k,lamb)
print (prob)
# So that result tells us that there is a 14.6% chance of observing exactly 6, but it would be much more helpful to be able to visualize the probability distribution. So let's go through and calculate the probability of seeing any number between 0 and 10. First, we can make an array between 0 and 11:
number_meteors_seen = np.arange(0,11)
number_meteors_seen
# +
taurid_meteor_sighting_probability = []
taurid_meteor_rate = 5
for n in number_meteors_seen:
prob = poisson_probability(number_meteors_seen[n],taurid_meteor_rate)
taurid_meteor_sighting_probability.append(prob)
# -
taurid_meteor_sighting_probability
plt.plot(number_meteors_seen,taurid_meteor_sighting_probability,label='Southern Taurids ($\lambda = 5$)')
plt.legend()
plt.show()
# When there is not an active shower the background meteor rate is about 2 an hour (although it is variable depending on time of night and season; see more here: https://www.amsmeteors.org/meteor-showers/meteor-faq/).
#
# <font color=goldenrod>**_Code for you to write_**</font>
#
# - **Calculate the probability of seeing different numbers of meteors when the background rate is 2 an hour (lambda = 2).**
# - **Plot that probability alongside the probability of seeing those same numbers during the Southern Taurids shower.**
# ## Simulate meteor observing
#
# There are many cases where it can be useful to simulate data sets. In this case, one could simulate what your experience could be in terms of the number of hours you could spend looking at the night sky and seeing 1 meteor or more on a normal night vs. a night with the Southern Taurids shower ongoing.
#
# We can use the `np.random.poisson` function to simulate 'realistic' data.
#
# `np.random.poisson( )` requires 1 parameter `lam` and an optional parameter `size`. Each call to `np.random.poisson( )` returns `size` number of draws from a Poisson distribution with $\lambda =$ `lam`.
#
# Let's try it with $\lambda = 2$ (the background rate).
# +
lam = 2
number_hours_watched = 100
number_hours_w_meteor_sighting = []
for n in np.arange(0,number_hours_watched):
number_meteors = np.random.poisson(lam)
if number_meteors >= 1:
number_hours_w_meteor_sighting.append(1)
fraction_w_sighting = len(number_hours_w_meteor_sighting)/number_hours_watched
print('percent of hours watched with a meteor sighting')
print(fraction_w_sighting*100)
# -
# <font color=goldenrod>**_Code for you to write_**</font>
#
# - **Do the same meteor watching simulation with $\lambda = 5$ (the Southern Taurids rate). Do it for 10 hours, 100 hours, 100,000 hours.**
# ### Earthquake Probability
#
# The occurrence of earthquakes is also a Poisson process, events occur randomly in time, and the average recurrence can be determined from Gutenberg-Richter. Last week we estimated the Gutenberg-Richter statistic which gives the annual rate of earthquakes above a given magnitude. Applying the Poisson distribution to this problem $\lambda = N * {\Delta}time$, where N is the annual rate. It is common to consider ${\Delta}time=30 yrs$. Why is that?
#
# Given the Gutenberg-Richter relationshipfor the San Franciso Bay Area: log10(N)= 3.266 - 0.797M, use the Poisson's distribution to find the probability of 0-9 M5 events in a 30 year period.
#
# How does the probability change with the number of events? How does that reconcile with the rate of M5 earthquakes?
#
# ***Write your answer here***
# ### Poisson Probability of 1 or more earthquakes
#
# The Poisson probability of zero events has an interesting use in characterizing earthquake hazard.
#
# $P(k=0)=e^{-\lambda}$
#
# The complement of the zero event probability is the probability of 1 or more earthquakes occuring in the period of time. It is this probability that is used in earthquake forecast reports. The probability of one or more events is written as;
#
# $P(k >= 1) = 1 - e^{-\lambda}$
#
# Determine the probability of 1 or more M4, M5, M6 and M7 in a 30 year period.
#
# ***Write answer here***
#
# How do the probabilities change if a 10 year period is considered?
#
# ***Write answer here***
# **Export the notebook as .pdf and upload to bcourses**
| week05_inclass/Week05_InClass_Probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Search Engine Based on StackOverflow Questions
# <h1>1. Business Problem </h1>
# <h2> 1.1 Description </h2>
# <p style='font-size:18px'><b> Description </b></p>
# <p>
# Stack Overflow is the largest, most trusted online community for developers to learn, share their programming knowledge, and build their careers.<br />
# <br />
# Stack Overflow is something which every programmer use one way or another. Each month, over 50 million developers come to Stack Overflow to learn, share their knowledge, and build their careers. It features questions and answers on a wide range of topics in computer programming. The website serves as a platform for users to ask and answer questions, and, through membership and active participation, to vote questions and answers up or down and edit questions and answers in a fashion similar to a wiki or Digg. As of April 2014 Stack Overflow has over 4,000,000 registered users, and it exceeded 10,000,000 questions in late August 2015. Based on the type of tags assigned to questions, the top eight most discussed topics on the site are: Java, JavaScript, C#, PHP, Android, jQuery, Python and HTML.<br />
# <br />
# </p>
# <p style='font-size:18px'><b> Problem Statemtent </b></p>
# Build a search engine based on StackOverflow questions, the search results should include the semantic meaning.
# <p style='font-size:18px'><b> Source:</b> https://archive.org/details/stackexchange </p>
#
# <h2> 1.2 Source / useful links </h2>
# Data Source :https://archive.org/details/stackexchange <br>
# Youtube : https://youtu.be/nNDqbUhtIRg <br>
# Research paper : Word2Vec https://arxiv.org/abs/1301.3781 <br>
# Research paper : Doc2Vec https://arxiv.org/abs/1405.4053 <br>
# Research paper : BERT https://arxiv.org/abs/1810.04805 <br>
# Research paper : Universal Sentance Encoder https://arxiv.org/abs/1803.11175 <br>
# word2vec Blog : https://blog.acolyer.org/2016/04/21/the-amazing-power-of-word-vectors/
#
#
# <h2> 1.3 Real World / Business Objectives and Constraints </h2>
#
# 1. the search results should include the semantic meaning.
# 2. Think of scalable architecture and try to reduce the time to return the results.
# <h1>2. Machine Learning problem </h1>
# <h2> 2.1 Data </h2>
# <h3> 2.1.1 Data Overview </h3>
# <br>
# All of the data is in 1 zip-file: stackoverflow.com-Posts.7z <br />
# inside the zip file there one xml file : Posts.xml
# <pre>
# <b>Posts.xml</b> contains 16 columns: Id,PostTypeId,AcceptedAnswerId,CreationDate,Score,ViewCount,Body,OwnerUserId,LastEditorUserId,LastEditDate,LastActivityDate,Title,Tags,AnswerCount,CommentCoun,FavoriteCount <br />
# <b>Size of Posts.xml</b> - 14.6GB<br />
# <b>Number of rows in Posts.xml</b> = 3 Million records <br />
# </pre>
# <b>Note : </b> we are not using this whole data for this case study, due to limited resources and time constraints<br/>
# <pre>
#
# <b>the dataframe that we are using has 3 columns: </b> Id,title, Body <br />
#
# <b>Number of rows in dataframe</b> = 112357 <br />
# </pre>
#
# -> The questions are randomized and contains a mix of verbose text sites as well as sites related to math and programming. The number of questions from each site may vary, and no filtering has been performed on the questions (such as closed questions).<br />
# <br />
#
# __Data Field Explaination__
#
# Dataset contains 112357 rows. The columns in the table are:<br />
# <pre>
# <b>Id</b> - Unique identifier for each question<br />
# <b>Title</b> - The question's title<br />
# <b>Body</b> - The body of the question<br />
#
# </pre>
#
# <br />
# <h3>2.1.2 Example Data point </h3>
# <h4>2.1.2.1 Posts.xml </h4>
# Id="1" <br />
# PostTypeId="1" <br />
# AcceptedAnswerId="3" <br />
# CreationDate="2016-08-02T15:39:14.947" <br />
# Score="8" <br />
# ViewCount="436" <br />
# Body="<p>What does "backprop" mean? Is the "backprop" term basically the same as "backpropagation" or does it have a different meaning?</p>
" <br />
# OwnerUserId="8" <br />
# LastEditorUserId="2444" <br />
# LastEditDate="2019-11-16T17:56:22.093" <br />
# LastActivityDate="2019-11-16T17:56:22.093" <br />
# Title="What is "backprop"?" <br />
# Tags="<neural-networks><backpropagation><terminology><definitions>" <br />
# AnswerCount="3" <br />
# CommentCount="0" <br />
# FavoriteCount="1" <br />
# <h4>2.1.2.2 file1.csv </h4>
# Id : "2214" <br/>
#
# Title : <pre> "When will the AI can replace developer or tester"</pre> <br/>
# Body : <pre> "<p>Now AI can replace call center, worker(in the factory) and going to replace court. When will the AI can replace developer or tester?</p>

<p>I want to know how long can AI replace developer. e.g. next 10 years because...</p>
"</pre><br/>
#
# <h2>2.2 Mapping the real-world problem to a Machine Learning Problem </h2>
# <h3> 2.2.1 Type of Machine Learning Problem </h3>
# <pre> <p>It is basically finding semamtic similarity between documents.Here, the terminology “document” could refer to a web-page, a word document, a paragraph of text, an essay, a sentence, or even just a single word. <br>
# Two semantically similar documents, will contain many of the same topics while two semantically different documents will not have many topics in common. Machine learning methods are described which can take as input large collections of documents and use those documents to automatically learn semantic similarity relations.</pre>
# <h3>2.2.2 Performance metric </h3>
# <b>Cosine similarity</b> :
# Cosine similarity calculates similarity by measuring the
# cosine of angle between two vectors. This is calculated as
# 
# link: https://en.wikipedia.org/wiki/Cosine_similarity
# <h1> 3. Exploratory Data Analysis </h1>
# <h2> 3.1 Data Loading </h2>
# <h3> 3.1.1 downloading data and extracting the zip file </h3>
# + colab={"base_uri": "https://localhost:8080/", "height": 243} colab_type="code" id="JhPXnS-rzyYw" outputId="940bc979-866b-4ab3-9347-c20d1d0502a5"
# !wget --header 'Host: ia800107.us.archive.org' --user-agent 'Mozilla/5.0 (Windows NT 10.0; Win64; x64; rv:76.0) Gecko/20100101 Firefox/76.0' --header 'Accept: text/html,application/xhtml+xml,application/xml;q=0.9,image/webp,*/*;q=0.8' --header 'Accept-Language: en-US,en;q=0.5' --referer 'https://archive.org/details/stackexchange' --header 'Cookie: PHPSESSID=cv4gm885ijhaelfot7md17gtd7' --header 'Upgrade-Insecure-Requests: 1' 'https://ia800107.us.archive.org/27/items/stackexchange/academia.stackexchange.com.7z' --output-document 'academia.stackexchange.com.7z'
# + colab={"base_uri": "https://localhost:8080/", "height": 131} colab_type="code" id="lRw4bMbEzzsL" outputId="5496daa8-0ae6-4685-97da-d815be13627c"
# !pip install patool #installing patoolib
import patoolib #importing
patoolib.extract_archive('/content/academia.stackexchange.com.7z')#extracting archive
# -
# <h3> 3.1.2 parsing the xml file </h3>
# + colab={} colab_type="code" id="InUjeyWjyfOM"
#https://stackoverflow.com/questions/1912434/how-do-i-parse-xml-in-python
import xml.etree.ElementTree as ET
root = ET.parse('Posts.xml').getroot()
# -
# <h3> 3.1.3 extracting the title and body from xml file </h3>
# + colab={} colab_type="code" id="rSTVr8oQyfOZ"
title = []
text = []
count = 0
count2 = 0
for child in root.iter('row'):
if child.attrib['PostTypeId'] == "1":#postTypeId ==1 means question post and postTypeId == 2 means answer post
if child.attrib['Title'] is not None:
title.append(child.attrib['Title'])
text.append(child.attrib['Body'])
elif child.attrib['Title'] is None:# in case if the post doesent have title it will just have body
text.append(child.attrib['Body'])
# + colab={"base_uri": "https://localhost:8080/", "height": 36} colab_type="code" id="HWUBo6bp1_eT" outputId="f7268eae-4094-425c-a2e8-6db78ce33f93"
len(text)
# -
# <h3> 3.1.4 saving the title and text into dataframe and then into csv </h3>
# + colab={} colab_type="code" id="Xgi78b0TyfOn"
import pandas as pd
df = pd.DataFrame({'title':title,"body":text})
# + colab={"base_uri": "https://localhost:8080/", "height": 206} colab_type="code" id="wIzDN43kyfOv" outputId="d4431ee8-d9cf-412e-c6ae-1881b8ee530d"
df.head()
# + colab={} colab_type="code" id="PbXZlCcGyfO6"
df.to_csv('file1.csv')
| questions_extract.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib
matplotlib.rcParams['figure.figsize'] = [8, 6]
import os
from darwinian_shift import DarwinianShift, BigwigLookup, hide_top_and_right_axes
# +
plot_dir = "./Transmembrane_figures/"
if not os.path.exists(plot_directory):
os.makedirs(plot_directory)
# Colours
unobserved_colour = '#BBBBBB'
observed_colour = '#EE7733'
# -
# The bigwig can be downloaded from here http://hgdownload.soe.ucsc.edu/goldenPath/hg19/phyloP100way/
# or by running
# wget http://hgdownload.soe.ucsc.edu/goldenPath/hg19/phyloP100way/hg19.100way.phyloP100way.bw
# Warning This file is approx 9Gb.
bw = BigwigLookup("hg19.100way.phyloP100way.bw", name='Phylop score')
transmembrane_section = {
'transcript_id': 'ENST00000277541',
'start': 1736, 'end': 1756
}
data = pd.read_excel("aau3879_TableS2.xlsx", skiprows=17, sheet_name='Mutations_collapsed_by_distance')
d = DarwinianShift(
data=data,
source_genome='grch37',
lookup=bw,
included_mutation_types='missense'
)
s = d.run_section(transmembrane_section)
s.plot_scatter(show_residues=True, show_legend=False, figsize=(6, 3),
unobserved_mutation_colour=unobserved_colour, missense_mutation_colour=observed_colour,
unmutated_marker_size=15, base_marker_size=15)
hide_top_and_right_axes()
plt.tight_layout()
plt.savefig(os.path.join(plot_dir, 'transmembrane_phylop_scatter.pdf'), transparent=True);
s.plot_mutation_rate_scatter(figsize=(3, 3), show_legend=False,
unobserved_mutation_colour=unobserved_colour, missense_mutation_colour=observed_colour,
unmutated_marker_size=15, base_marker_size=15)
hide_top_and_right_axes()
plt.title('')
plt.tight_layout()
plt.savefig(os.path.join(plot_dir, 'transmembrane_phylop_vs_mutation_rate.pdf'), transparent=True);
| Transmembrane_mutations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp data.tabular
# -
# # Data Tabular
#
# > Main Tabular functions used throughout the library.
#export
from tsai.imports import *
from fastai.tabular.all import *
path = untar_data(URLs.ADULT_SAMPLE)
df = pd.read_csv(path/'adult.csv')
splits = RandomSplitter()(range_of(df))
cat_names = ['workclass', 'education', 'marital-status', 'occupation', 'relationship', 'race']
cont_names = ['age', 'fnlwgt', 'education-num']
procs = [Categorify, FillMissing, Normalize]
y_names = 'salary'
y_block = CategoryBlock()
pd.options.mode.chained_assignment=None
to = TabularPandas(df, procs=procs, cat_names=cat_names, cont_names=cont_names,
y_names=y_names, y_block=y_block, splits=splits, inplace=True,
reduce_memory=False)
# +
# export
class TabularDataset():
"A `Numpy` dataset from a `TabularPandas` object"
def __init__(self, to):
self.cats = to.cats.to_numpy().astype(np.long)
self.conts = to.conts.to_numpy().astype(np.float32)
self.ys = to.ys.to_numpy()
def __getitem__(self, idx): return self.cats[idx], self.conts[idx], self.ys[idx]
def __len__(self): return len(self.cats)
@property
def c(self): return 0 if self.ys is None else 1 if isinstance(self.ys[0], float) else len(np.unique(self.ys))
class TabularDataLoader(DataLoader):
def __init__(self, dataset, bs=1, num_workers=0, device=None, train=False, **kwargs):
device = ifnone(device, default_device())
super().__init__(dataset, bs=min(bs, len(dataset)), num_workers=num_workers, shuffle=train, device=device, drop_last=train, **kwargs)
self.device, self.shuffle = device, train
def create_item(self, s): return s
def get_idxs(self):
idxs = Inf.count if self.indexed else Inf.nones
if self.n is not None: idxs = list(range(len(self.dataset)))
if self.shuffle: self.shuffle_fn()
return idxs
def create_batch(self, b):
return self.dataset[b[0]:b[0]+self.bs]
def shuffle_fn(self):
"Shuffle dataset after each epoch"
rng = np.random.permutation(len(self.dataset))
self.dataset.cats = self.dataset.cats[rng]
self.dataset.conts = self.dataset.conts[rng]
self.dataset.ys = self.dataset.ys[rng]
def to(self, device):
self.device = device
# def ds_to(self, device=None):
self.dataset.cats = tensor(self.dataset.cats).to(device=self.device)
self.dataset.conts = tensor(self.dataset.conts).to(device=self.device)
self.dataset.ys = tensor(self.dataset.ys).to(device=self.device)
# -
train_ds = TabularDataset(to.train)
valid_ds = TabularDataset(to.valid)
train_dl = TabularDataLoader(train_ds, bs=512, train=True)
valid_dl = TabularDataLoader(valid_ds, bs=512)
dls = DataLoaders(train_dl,valid_dl)
emb_szs = get_emb_sz(to)
net = TabularModel(emb_szs, 3, 2, layers=[200,100])#.cuda()
learn = Learner(dls, net, metrics=accuracy, loss_func=CrossEntropyLossFlat())
learn.fit(1, 1e-2)
#hide
create_scripts()
beep()
| nbs/005_data.tabular.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kaggle - WiDS Datathon 2022
#
# ## Modelo Nº1
#
# ### Description:
#
# - All the features
# - The entire dataset for train (not separation train/test)
# - Catboost
# - Hiperparameters by default
# - Long value of iterations: 10000
#
# - Public Score = 42.525
# # 1. Modules
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
#model
from catboost import CatBoostRegressor
# -
# # 2. Data
# +
#paths to the Github repo
train_path = 'https://raw.githubusercontent.com/ccollado7/wds-datathon-2022/main/data/raw/train.csv'
test_path = 'https://raw.githubusercontent.com/ccollado7/wds-datathon-2022/main/data/raw/test.csv'
submit_path = 'https://raw.githubusercontent.com/ccollado7/wds-datathon-2022/main/data/raw/sample_solution.csv'
# +
#train dataset
train = pd.read_csv(train_path)
train.head()
# +
#test dataset
test = pd.read_csv(test_path)
test.head()
# +
#sumbit
submit = pd.read_csv(submit_path)
submit.head()
# -
x_train = train.drop(['id','site_eui'], axis=1)
x_train.columns
y_train = train['site_eui']
cat_columns = list(train.select_dtypes(include=['object']).columns)
cat_columns
model_1_1 = CatBoostRegressor(cat_features = cat_columns,
iterations=10000,
verbose=10)
model_1_1.fit(x_train,y_train,plot=True)
x_test = test.drop('id', axis=1)
x_test.columns
submit['site_eui'] = model_1_1.predict(x_test)
submit.to_csv('modelo_1_1.csv',index=False)
| notebooks/modelos/modelo_1/modelo_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df_population = pd.read_excel("population_indian_states.xlsx")
df_population.dtypes
df_population.columns
df_population['population'] = df_population['population'].replace({',':''}, regex=True)
df_population['population'] = df_population['population'].astype(str).astype(int)
df_population
df_vaccines = pd.read_excel("vaccines_jan6.xlsx")
# +
# df = pd.concat([df_vaccines, df_population], ignore_index=True, sort=False)
# -
df = pd.merge(df_vaccines, df_population, how='outer',left_on=['state'], right_on=['state'])
df.columns
df.drop('Unnamed: 0', axis=1, inplace=True)
df.drop('sr_no', axis=1, inplace=True)
df.columns
df = df.drop(37)
df = df.drop(38)
df = df.drop(39)
df.head(3)
# +
# df.xs(0)['population'] = 380581.0
# -
df.dtypes
| population-stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COMPARISON OF DENOISING PARAMETERS (*SPARSE2D*)
# ---
# The denoising method was implemented from <a href="https://github.com/CEA-COSMIC/pysap.git" target="_blank">pySAP</a> in `pysap.extensions.sparse2d`. We tested the following combinations of parameters.
#
# **1st combination :** `tab_n_sigma = [5]`, `type_of_filtering = 3`
#
# **2nd combination :** `tab_n_sigma = [5]`, `type_of_multiresolution_transform = 2`
#
# **3rd combination :** `tab_n_sigma = [5]`, `type_of_multiresolution_transform = 24`, `type_of_filtering = 6`
#
# ## CONTENTS
# ---
#
# 1. [DATA PREPARATION](#DATA)
# 1. [METRICS OF COMPARISON](#METRICS)
# 1. [RESULTS](#RESULTS)
# 1. [Visualisation](#VISUALISATION)
# 1. [RESULTS TO DATAFRAME](#RESULTS-TO-DATAFRAME)
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
# # DATA
# ---
# +
#Importation
input_path = '/Users/alacan/Documents/Cosmostat/Codes/BlendHunter'
blended = np.load('/Users/alacan/Documents/Cosmostat/Codes/BlendHunter/bh_5/blended_noisy.npy', allow_pickle=True)
not_blended =np.load('/Users/alacan/Documents/Cosmostat/Codes/BlendHunter/bh_5/not_blended_noisy.npy', allow_pickle=True)
# +
#Import denoised images
def load_images(path= None, method=None, noise=None):
img = np.load(path+'/denoised/bh_{}_denoised{}/test_images.npy'.format(noise, method), allow_pickle=True)
return img
#Import original images
def get_images(sample):
return np.array([sample[obj]['galsim_image'][0].array for obj in range(36000,40000)])
#Import noisy images
def get_noisy(path=None, noise=None):
return np.load(path+'/bh_{}/test_images.npy'.format(noise), allow_pickle=True)
# -
#Original images
original_blended = get_images(blended)
original_not_blended = get_images(not_blended)
original = np.concatenate((original_blended, original_not_blended))
#Noisy_images
noisy5 = get_noisy(noise=5, path=input_path)
noisy14 = get_noisy(noise=14, path=input_path)
noisy18 = get_noisy(noise=18, path=input_path)
noisy40 = get_noisy(noise=40, path=input_path)
# +
#Denoised images
dn_images5_1 = load_images(path = input_path, method = 1, noise=5)
dn_images5_2 = load_images(path = input_path, method = 2, noise=5)
dn_images5_3 = load_images(path = input_path, method = 3, noise=5)
dn_images14_1 = load_images(path = input_path, method = 1, noise=14)
dn_images14_2 = load_images(path = input_path, method = 2, noise=14)
dn_images14_3 = load_images(path = input_path, method = 3, noise=14)
dn_images40_1 = load_images(path = input_path, method = 1, noise=40)
dn_images40_2 = load_images(path = input_path, method = 2, noise=40)
dn_images40_3 = load_images(path = input_path, method = 3, noise=40)
dn_images18_1 = load_images(path = input_path, method = 1, noise=18)
dn_images18_2 = load_images(path = input_path, method = 2, noise=18)
dn_images18_3 = load_images(path = input_path, method = 3, noise=18)
dn_images26_1 = load_images(path = input_path, method = 1, noise=26)
dn_images35_1 = load_images(path = input_path, method = 1, noise=35)
# +
#Denoised images for different noise realisations
dn_images51_1= load_images(path = input_path, method = 1, noise = 51)
dn_images52_1= load_images(path = input_path, method = 1, noise = 52)
dn_images53_1= load_images(path = input_path, method = 1, noise = 53)
dn_images54_1= load_images(path = input_path, method = 1, noise = 54)
dn_images141_1= load_images(path = input_path, method = 1, noise = 141)
dn_images142_1= load_images(path = input_path, method = 1, noise = 142)
dn_images143_1= load_images(path = input_path, method = 1, noise = 143)
dn_images144_1= load_images(path = input_path, method = 1, noise = 144)
dn_images181_1= load_images(path = input_path, method = 1, noise = 181)
dn_images182_1= load_images(path = input_path, method = 1, noise = 182)
dn_images183_1= load_images(path = input_path, method = 1, noise = 183)
dn_images184_1= load_images(path = input_path, method = 1, noise = 184)
# -
#Import segmentation maps
seg_map_b = np.load('/Users/alacan/Documents/Cosmostat/Codes/BlendHunter/bh_5/test_seg_map.npy', allow_pickle=True)
seg_map_nb = np.array([not_blended[val]['seg_map'][0].array for val in range(36000, 40000)])
seg_map = np.concatenate((seg_map_b, seg_map_nb))
# # METRICS
# ---
# +
#Calculation residual noise
def res_noise(images = None, mask=None):
return np.array([np.mean(abs(images[i][mask[i] == 0])) for i in range(len(images))])
#Calculation of lost information in galaxies
def lost_info(images=None, mask=None):
return np.array([np.mean(abs(images[i][mask[i] != 0] - original[i][mask[i] != 0])) for i in range(len(images))])
#Calculation of SNR
def snr_alpha(images=None, sigma=None, mask=None):
alpha = np.array([np.sum(images[i][mask[i] == 1] ** 2) for i in range(len(images))])
return np.array([np.sqrt(alpha[i] / sigma**2) for i in range(len(alpha))])
def snr_beta(images=None, sigma=None, mask=None):
beta = np.array([np.sum(images[i][mask[i] == 2] ** 2) for i in range(0, 4000)])
return np.array([np.sqrt(beta[i] / sigma**2) for i in range(len(beta))])
# Calculation of PSNR
import math
import cv2
def psnr(img1, img2):
mse = np.mean( (img1 - img2) ** 2 )
if mse == 0:
return 100
PIXEL_MAX = 255.0
return 10*math.log10((PIXEL_MAX **2) / mse)
def mean_psnr_noisy(images, round_to = None):
return round(float(np.mean((*[psnr(images[0][i], original_blended[i]) for i in range(len(images[0]))], *[psnr(images[1][i], original_not_blended[i]) for i in range(len(images[1]))]))), round_to)
def mean_psnr(denoised_images=None, round_to = None):
return round(float(np.mean([psnr(denoised_images[i], original[i]) for i in range(len(original))])), round_to)
#Original SNR
snr = np.mean(snr_alpha(images=original, sigma=5.0, mask= seg_map)) + np.mean(snr_beta(images=original, sigma=5.0, mask= seg_map))
# -
# # RESULTS
# ---
#PSNR on noisy images
psnr5 = mean_psnr_noisy(noisy5, round_to = 3)
psnr14 = mean_psnr_noisy(noisy14, round_to = 3)
psnr18 = mean_psnr_noisy(noisy18, round_to = 3)
psnr40 = mean_psnr_noisy(noisy40, round_to = 3)
# +
#Calculation of residual noise
mean5_1 = round(float(np.mean(res_noise(images=dn_images5_1, mask=seg_map))), 3)
mean5_2 = round(float(np.mean(res_noise(images=dn_images5_2, mask=seg_map))), 3)
mean5_3 = round(float(np.mean(res_noise(images=dn_images5_3, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 5 : {}'.format(mean5_1))
print('Mean residual noise with 2nd method for sigma_noise = 5 : {}'.format(mean5_2))
print('Mean residual noise with 3rd method for sigma_noise = 5 : {}'.format(mean5_3))
snr5_1 = float(np.mean(snr_alpha(images=dn_images5_1, sigma=5.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images5_1, sigma=5.0, mask= seg_map)))
snr5_2 = float(np.mean(snr_alpha(images=dn_images5_2, sigma=5.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images5_2, sigma=5.0, mask= seg_map)))
snr5_3 = float(np.mean(snr_alpha(images=dn_images5_3, sigma=5.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images5_3, sigma=5.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 5 : {}'.format(round(snr5_1/snr, 5)))
print('SNR ratio with 2nd method for sigma_noise = 5 : {}'.format(round(snr5_2/snr, 5)))
print('SNR ratio with 3rd method for sigma_noise = 5 : {}'.format(round(snr5_3/snr, 5)))
psnr5_1 = round(float(np.mean([psnr(dn_images5_1[i], original[i]) for i in range(len(original))])), 3)
psnr5_2 = round(float(np.mean([psnr(dn_images5_2[i], original[i]) for i in range(len(original))])), 3)
psnr5_3 = round(float(np.mean([psnr(dn_images5_3[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 5 : {}'.format(psnr5_1))
print('PSNR with 2nd method for sigma_noise = 5 : {}'.format(psnr5_2))
print('PSNR with 3rd method for sigma_noise = 5 : {}'.format(psnr5_3))
# -
#Calculation of residual noise
mean14_1 = round(float(np.mean(res_noise(images=dn_images14_1, mask=seg_map))), 3)
mean14_2 = round(float(np.mean(res_noise(images=dn_images14_2, mask=seg_map))), 3)
mean14_3 = round(float(np.mean(res_noise(images=dn_images14_3, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 14 : {}'.format(mean14_1))
print('Mean residual noise with 2nd method for sigma_noise = 14 : {}'.format(mean14_2))
print('Mean residual noise with 3rd method for sigma_noise = 14 : {}'.format(mean14_3))
#Calculation of lost info
snr14_1 = float(np.mean(snr_alpha(images=dn_images14_1, sigma=14.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images14_1, sigma=14.0, mask= seg_map)))
snr14_2 = float(np.mean(snr_alpha(images=dn_images14_2, sigma=14.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images14_2, sigma=14.0, mask= seg_map)))
snr14_3 = float(np.mean(snr_alpha(images=dn_images14_3, sigma=14.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images14_3, sigma=14.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 14 : {}'.format(round(snr14_1/snr, 3)))
print('SNR ratio with 2nd method for sigma_noise = 14 : {}'.format(round(snr14_2/snr, 3)))
print('SNR ratio with 3rd method for sigma_noise = 14 : {}'.format(round(snr14_3/snr, 3)))
psnr14_1 = round(float(np.mean([psnr(dn_images14_1[i], original[i]) for i in range(len(original))])), 3)
psnr14_2 = round(float(np.mean([psnr(dn_images14_2[i], original[i]) for i in range(len(original))])), 3)
psnr14_3 = round(float(np.mean([psnr(dn_images14_3[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 14 : {}'.format(psnr14_1))
print('PSNR with 2nd method for sigma_noise = 14 : {}'.format(psnr14_2))
print('PSNR with 3rd method for sigma_noise = 14 : {}'.format(psnr14_3))
#Calculation of residual noise
mean18_1 = round(float(np.mean(res_noise(images=dn_images18_1, mask=seg_map))), 3)
mean18_2 = round(float(np.mean(res_noise(images=dn_images18_2, mask=seg_map))), 3)
mean18_3 = round(float(np.mean(res_noise(images=dn_images18_3, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 18 : {}'.format(mean18_1))
print('Mean residual noise with 2nd method for sigma_noise = 18 : {}'.format(mean18_2))
print('Mean residual noise with 3rd method for sigma_noise = 18 : {}'.format(mean18_3))
#Calculation of lost info
snr18_1 = float(np.mean(snr_alpha(images=dn_images18_1, sigma=18.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images18_1, sigma=18.0, mask= seg_map)))
snr18_2 = float(np.mean(snr_alpha(images=dn_images18_2, sigma=18.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images18_2, sigma=18.0, mask= seg_map)))
snr18_3 = float(np.mean(snr_alpha(images=dn_images18_3, sigma=18.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images18_3, sigma=18.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 18 : {}'.format(round(snr18_1/snr, 3)))
print('SNR ratio with 2nd method for sigma_noise = 18 : {}'.format(round(snr18_2/snr, 3)))
print('SNR ratio with 3rd method for sigma_noise = 18 : {}'.format(round(snr18_3/snr, 3)))
psnr18_1 = round(float(np.mean([psnr(dn_images18_1[i], original[i]) for i in range(len(original))])), 3)
psnr18_2 = round(float(np.mean([psnr(dn_images18_2[i], original[i]) for i in range(len(original))])), 3)
psnr18_3 = round(float(np.mean([psnr(dn_images18_3[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 18 : {}'.format(psnr18_1))
print('PSNR with 2nd method for sigma_noise = 18 : {}'.format(psnr18_2))
print('PSNR with 3rd method for sigma_noise = 18 : {}'.format(psnr18_3))
#Calculation of residual noise
mean26_1 = round(float(np.mean(res_noise(images=dn_images26_1, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 26 : {}'.format(mean26_1))
#Calculation of lost info
snr26_1 = float(np.mean(snr_alpha(images=dn_images26_1, sigma=26.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images26_1, sigma=26.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 26 : {}'.format(round(snr26_1/snr, 3)))
psnr26_1 = round(float(np.mean([psnr(dn_images26_1[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 26 : {}'.format(psnr26_1))
#Calculation of residual noise
mean35_1 = round(float(np.mean(res_noise(images=dn_images35_1, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 35 : {}'.format(mean35_1))
#Calculation of lost info
snr35_1 = float(np.mean(snr_alpha(images=dn_images35_1, sigma=35.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images35_1, sigma=35.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 35 : {}'.format(round(snr35_1/snr, 3)))
psnr35_1 = round(float(np.mean([psnr(dn_images35_1[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 35 : {}'.format(psnr35_1))
#Calculation of residual noise
mean40_1 = round(float(np.mean(res_noise(images=dn_images40_1, mask=seg_map))), 3)
mean40_2 = round(float(np.mean(res_noise(images=dn_images40_2, mask=seg_map))), 3)
mean40_3 = round(float(np.mean(res_noise(images=dn_images40_3, mask=seg_map))), 3)
print('---RESIDUAL NOISE AND QUALITY OF DENOISING---')
print('Mean residual noise with 1st method for sigma_noise = 40 : {}'.format(mean40_1))
print('Mean residual noise with 2nd method for sigma_noise = 40 : {}'.format(mean40_2))
print('Mean residual noise with 3rd method for sigma_noise = 40 : {}'.format(mean40_3))
#Calculation of lost info
snr40_1 = float((np.mean(snr_alpha(images=dn_images40_1, sigma=40.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images40_1, sigma=40.0, mask= seg_map))))
snr40_2 = float(np.mean(snr_alpha(images=dn_images40_2, sigma=40.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images40_2, sigma=40.0, mask= seg_map)))
snr40_3 = float(np.mean(snr_alpha(images=dn_images40_3, sigma=40.0, mask= seg_map)) + np.mean(snr_beta(images=dn_images40_3, sigma=40.0, mask= seg_map)))
print('---SNR---')
print('SNR ratio with 1st method for sigma_noise = 40 : {}'.format(round(snr40_1/snr, 3)))
print('SNR ratio with 2nd method for sigma_noise = 40 : {}'.format(round(snr40_2/snr, 3)))
print('SNR ratio with 3rd method for sigma_noise = 40 : {}'.format(round(snr40_3/snr, 3)))
psnr40_1 = round(float(np.mean([psnr(dn_images40_1[i], original[i]) for i in range(len(original))])), 3)
psnr40_2 = round(float(np.mean([psnr(dn_images40_2[i], original[i]) for i in range(len(original))])), 3)
psnr40_3 = round(float(np.mean([psnr(dn_images40_3[i], original[i]) for i in range(len(original))])), 3)
print('---PSNR AND QUALITY OF RECONSTRUCTION---')
print('PSNR with 1st method for sigma_noise = 40 : {}'.format(psnr40_1))
print('PSNR with 2nd method for sigma_noise = 40 : {}'.format(psnr40_2))
print('PSNR with 3rd method for sigma_noise = 40 : {}'.format(psnr40_3))
# ## MORE NOISE REALISATIONS
#PSNR on noisy images
list_psnr_5 = [mean_psnr(denoised_images=img, round_to=3) for img in [dn_images5_1, dn_images51_1, dn_images52_1, dn_images53_1, dn_images54_1]]
list_psnr_14 = [mean_psnr(denoised_images=img, round_to=3) for img in [dn_images14_1, dn_images141_1, dn_images142_1, dn_images143_1, dn_images144_1]]
list_psnr_18 = [mean_psnr(denoised_images=img, round_to=3) for img in [dn_images18_1, dn_images181_1, dn_images182_1, dn_images183_1, dn_images184_1]]
# +
#Mean PSNR
mean_psnr5 = np.mean(list_psnr_5)
mean_psnr14 = np.mean(list_psnr_14)
mean_psnr18 = np.mean(list_psnr_18)
#Std deviation of PSNR
std_psnr5 = np.std(list_psnr_5)
std_psnr14 = np.std(list_psnr_14)
std_psnr18 = np.std(list_psnr_18)
# -
# ## VISUALISATION
#
# +
#List of sigma_noise values
sig = [5.0, 14.0, 18.0, 40.0]
#List of values for each metric per combination
list_means1 = [mean5_1, mean14_1,mean18_1, mean40_1]
list_means2 = [mean5_2, mean14_2,mean18_2, mean40_2]
list_means3 = [mean5_3, mean14_3,mean18_3, mean40_3]
list_snr1 = [round(snr5_1, 3), round(snr14_1,3),round(snr18_1 ,3), round(snr40_1 ,3)]
list_snr2 = [round(snr5_2,3), round(snr14_2,3),round(snr18_2,3), round(snr40_2,3)]
list_snr3 = [round(snr5_3,3), round(snr14_3,3),round(snr18_3,3), round(snr40_3,3)]
list_psnr1 = [psnr5_1, psnr14_1,psnr18_1, psnr40_1]
list_psnr2 = [psnr5_2, psnr14_2,psnr18_2, psnr40_2]
list_psnr3 = [psnr5_3, psnr14_3,psnr18_3, psnr40_3]
#list PSNR on noisy images
list_psnr = [psnr5, psnr14, psnr18, psnr40]
# +
sns.set(context='notebook', style='whitegrid', palette='deep')
font = {'family': 'monospace',
'color': 'k',
'weight': 'normal',
'size': 16.5}
fig, ax = plt.subplots(2,figsize=(20,19), sharey='row')
fig.suptitle('Comparing different parameters combinations in denoising method (Sparse2D)', fontdict={'family': 'monospace','color': 'k','weight': 'bold'}, fontsize=22)
#First subplot (Accuracy w.r.t. sigma_noise)
ax[0].set_title('Mean Residual Noise', fontdict=font, fontsize=20)
ax[0].plot(sig, list_means1, 'k',marker='o', linewidth=3, label='1st combination')
ax[0].plot(sig, list_means2, 'k', ls='dashdot', marker ='o', linewidth=3, label='2nd combination')
ax[0].plot(sig, list_means3, 'k:', marker ='o', linewidth=3, label='3rd combination')
ax[0].legend(loc='upper left', shadow=True, fontsize=18)
ax[0].set_ylabel('Mean residual Noise', fontdict = font)
ax[0].set_xlabel('Noise standard deviation ($\sigma_{noise}$) on images before denoising', fontdict = font)
ax[0].tick_params(axis='both', which='major', labelsize=17)
#Second subplot (Missed blends by SExtractor and BH's false negatives in a future commit)
ax[1].set_title('PSNR level', fontdict=font, fontsize=20)
ax[1].plot(sig, list_psnr1, 'k',marker='o', linewidth=3, label='1st combination')
ax[1].plot(sig, list_psnr2, 'k', ls='dashdot', marker ='o', linewidth=3, label='2nd combination')
ax[1].plot(sig, list_psnr3, 'k:', marker ='o', linewidth=3, label='3rd combination')
ax[1].plot(sig, list_psnr, 'grey', ls = '--',marker ='o', linewidth=3, label='Noisy images')
ax[1].tick_params(axis='both', which='major', labelsize=17)
ax[1].legend(loc='upper right', shadow=True, fontsize=18)
ax[1].set_ylabel('PSNR', fontdict=font)
ax[1].set_xlabel('Noise standard deviation ($\sigma_{noise}$) on images before denoising', fontdict = font)
plt.subplots_adjust(hspace=0.3)
plt.show()
# +
##Error bars
#fig = plt.figure(figsize=(20,10))
#plt.errorbar(sig[:-1], [mean_psnr5, mean_psnr14, mean_psnr18], yerr=[std_psnr5, std_psnr14, std_psnr18], label = 'sigma_noise = 5')
#plt.legend()
#plt.show()
# -
# # RESULTS TO DATAFRAME
def new_col_dataframe(dataframe=None, metric= None,method=None, data=None):
dataframe['{}_{}'.format(metric,method)] = pd.DataFrame(data)
return dataframe
# Create a DataFrame object
results= pd.DataFrame(sig, columns=['sigma_noise'])
print('The shape of our dataframe is '+str(results.shape))
results = new_col_dataframe(dataframe=results, metric = 'res_noise', method= 1, data = list_means1)
results = new_col_dataframe(dataframe=results, metric = 'res_noise', method= 2, data = list_means2)
results = new_col_dataframe(dataframe=results, metric = 'res_noise', method= 3, data = list_means3)
results = new_col_dataframe(dataframe=results, metric = 'snr', method= 1, data = list_snr1)
results = new_col_dataframe(dataframe=results, metric = 'snr', method= 2, data = list_snr2)
results = new_col_dataframe(dataframe=results, metric = 'snr', method= 3, data = list_snr3)
results = new_col_dataframe(dataframe=results, metric = 'psnr', method= 1, data = list_psnr1)
results = new_col_dataframe(dataframe=results, metric = 'psnr', method= 2, data = list_psnr2)
results = new_col_dataframe(dataframe=results, metric = 'psnr', method= 3, data = list_psnr3)
results = new_col_dataframe(dataframe=results, metric = 'psnr_noisy_img', method= '', data = list_psnr)
#Save dataframe
results.to_csv('/Users/alacan/Documents/Cosmostat/Codes/BlendHunter/results_denoising.csv')
# +
## DATAFRAME FOR ERROR BARS
# Create a DataFrame object
results_method1= pd.DataFrame(sig[:-1], columns=['sigma_noise'])
print('The shape of our dataframe is '+str(results.shape))
#List of values
list_m = [mean_psnr5, mean_psnr14, mean_psnr18]
list_std = [std_psnr5, std_psnr14, std_psnr18]
results_method1 = new_col_dataframe(dataframe=results_method1, metric = 'Mean_PSNR', method= 1, data = list_m)
results_method1 = new_col_dataframe(dataframe=results_method1, metric = 'std_PSNR', method= 1, data = list_std)
# -
#Save dataframe
results_method1.to_csv('/Users/alacan/Documents/Cosmostat/Codes/BlendHunter/results_denoising_method1.csv')
| notebooks/denoising_tests.ipynb |