code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Perhitungan Nilai Pengaturan Arduino UNO Terbaik Untuk Tracking Teleskop Ganda Zeiss
#
# Oleh : <NAME> <br>
# Tanggal : 29 Oktober 2020 <br>
# Ref : Mo<NAME>, 2020 <br>
# Versi : 1.0 <br>
# Kontak : <EMAIL> <br>
#
# ---
# 1. Import modul yang digunakan
import numpy as np
# Dari buku Astronomical Almanac 2020 halaman B9, diketahui bahwa pada tahun 2020,
#
# $$\bar t_{sidereal}\ (s) = 0.997 269 566 32\ \bar t_{solar}\ (s)$$
#
# Spesifikasi perhitungan (patokan Arduino UNO):
# - $\Delta t = 62.5$ ns (Resolusi waktu Arduino UNO dengan crystal clock 16 MHz)
# - Prescale $= 2^3 - 2^{10}$
# - Count $= 16$ bit $= 65536$ count
#
# Spesifikasi perhitungan (tambahan info dari Teleskop Zeiss dan kamera ZWO ASI 224MM):
# - $F_{teleskop} \sim 11000$ mm (Panjang fokus Teleskop Zeiss)
# - $pix_{size} = 3.75\ \mu$m (Ukuran piksel kamera)
#
# Challenge: Menghitung nilai prescale dan count terdekat serta pergeseran <i>tracking</i> dalam piksel per jam
# 2. Definisikan fungsi untuk menentukan nilai prescaler dan count yang menghasilkan nilai perhitungan terdekat dengan nilai yang diinginkan
def find_nearest_val(value, timer=1):
# Define initial values
res_t = 62.5 # Time resolution of UNO's crystal clock (nanoseconds)
f_tele = 11000 # mm
p_size = 3.75 # micron
# Define nearest prescale and count indices
near_scale, near_count = 0, 0
# Declaring a variable to store the minimum absolute difference
delta = float('inf')
if timer==0:
# Pre scaler values
pre_scale = np.array([1, 8, 64, 256, 1024])
# Acceptable count value
count_1 = np.arange(1, 257, 1)
count_2 = np.arange(1, 1025, 1)
for i in range(len(pre_scale)):
for j in range(len(count_1)):
for k in range(len(count_2)):
# Finding the value
time = res_t * pre_scale[i] * count_1[j] * count_2[k] / 1e6
# Updating the index of the answer if new absolute difference is less than delta
if abs(time - value) < delta:
delta = abs(time - value)
near_scale, near_count_1, near_count_2 = i, j, k
# Calculate pixel scale
p_scale = 206265. * p_size / f_tele / 1000. # arcseconds per pixel
# Calculate nearest value (miliseconds)
result = res_t * pre_scale[near_scale] * count_1[near_count_1] * count_2[near_count_2] / 1e6
# Calculate error of calculated over required values (percent)
error = delta / value * 100.
# Calculate error of calculated over required values (arcseconds per hour)
err_angle = 360. * 3600. * error / 100. / 24
# Calculate error of calculated over required values (pixels per hour)
err_pixel = err_angle / p_scale
param = [timer, pre_scale[near_scale], count_1[near_count_1], count_2[near_count_2], result, delta, error, err_angle, err_pixel]
print("Value to be reached = %.8f ms \n" %(value))
print("Timer = %i" %(timer))
print("\nResult!")
print("Nearest prescaler = %i" %(pre_scale[near_scale]))
print("Nearest count = %i" %(count_1[near_count_1]))
print("Looping = %i" %(count_2[near_count_2]))
print("Nearest value with 16MHz clock = %.8f ms" %(result))
print("Difference of required and calculated time = %.8f ms" %(delta))
print("Difference ratio = %.5f percent" %(error))
print("Difference rate = %.5f arcsec per hour" %(err_angle))
print("Difference rate = %.5f pixels per hour" %(err_pixel))
elif timer==1:
# Pre scaler values
pre_scale = np.array([1, 8, 64, 256, 1024])
# Acceptable count value
count = np.arange(1, 65537, 1)
for i in range(len(pre_scale)):
for j in range(len(count)):
# Finding the value
time = res_t * pre_scale[i] * count[j] / 1e6
# Updating the index of the answer if new absolute difference is less than delta
if abs(time - value) < delta:
delta = abs(time - value)
near_scale, near_count = i, j
# Calculate pixel scale
p_scale = 206265. * p_size / f_tele / 1000. # arcseconds per pixel
# Calculate nearest value (miliseconds)
result = res_t * pre_scale[near_scale] * count[near_count] / 1e6
# Calculate error of calculated over required values (percent)
error = delta / value * 100.
# Calculate error of calculated over required values (arcseconds per hour)
err_angle = 360. * 3600. * error / 100. / 24
# Calculate error of calculated over required values (pixels per hour)
err_pixel = err_angle / p_scale
param = [timer, pre_scale[near_scale], count[near_count], result, delta, error, err_angle, err_pixel]
print("Value to be reached = %.8f ms \n" %(value))
print("Timer = %i" %(timer))
print("\nResult!")
print("Nearest prescaler = %i" %(pre_scale[near_scale]))
print("Nearest count = %i" %(count[near_count]))
print("Nearest value with 16MHz clock = %.8f ms" %(result))
print("Difference of required and calculated time = %.8f ms" %(delta))
print("Difference ratio = %.5f percent" %(error))
print("Difference rate = %.5f arcsec per hour" %(err_angle))
print("Difference rate = %.5f pixels per hour" %(err_pixel))
elif timer==2:
# Pre scaler values
pre_scale = np.array([1, 8, 32, 64, 128, 256, 1024])
# Acceptable count value
count_1 = np.arange(1, 257, 1)
count_2 = np.arange(1, 1025, 1)
for i in range(len(pre_scale)):
for j in range(len(count_1)):
for k in range(len(count_2)):
# Finding the value
time = res_t * pre_scale[i] * count_1[j] * count_2[k] / 1e6
# Updating the index of the answer if new absolute difference is less than delta
if abs(time - value) < delta:
delta = abs(time - value)
near_scale, near_count_1, near_count_2 = i, j, k
# Calculate pixel scale
p_scale = 206265. * p_size / f_tele / 1000. # arcseconds per pixel
# Calculate nearest value (miliseconds)
result = res_t * pre_scale[near_scale] * count_1[near_count_1] * count_2[near_count_2] / 1e6
# Calculate error of calculated over required values (percent)
error = delta / value * 100.
# Calculate error of calculated over required values (arcseconds per hour)
err_angle = 360. * 3600. * error / 100. / 24
# Calculate error of calculated over required values (pixels per hour)
err_pixel = err_angle / p_scale
param = [timer, pre_scale[near_scale], count_1[near_count_1], count_2[near_count_2], result, delta, error, err_angle, err_pixel]
print("Value to be reached = %.8f ms \n" %(value))
print("Timer = %i" %(timer))
print("\nResult!")
print("Nearest prescaler = %i" %(pre_scale[near_scale]))
print("Nearest count = %i" %(count_1[near_count_1]))
print("Looping = %i" %(count_2[near_count_2]))
print("Nearest value with 16MHz clock = %.8f ms" %(result))
print("Difference of required and calculated time = %.8f ms" %(delta))
print("Difference ratio = %.5f percent" %(error))
print("Difference rate = %.5f arcsec per hour" %(err_angle))
print("Difference rate = %.5f pixels per hour" %(err_pixel))
else:
raise ValueError("Salah input nilai Timer!")
return param
# 3. Definisikan nilai yang ingin dicapai
sidereal = 1994.53913264/2. # miliseconds
mars = 996.7291988 # miliseconds
# 4. Menghitung nilai yang ingin dicapai (sideris)
sidereal_res = find_nearest_val(sidereal, timer=0)
sidereal_res = find_nearest_val(sidereal, timer=1)
sidereal_res = find_nearest_val(sidereal, timer=2)
# 5. Menghitung nilai yang ingin dicapai (mars)
mars_res = find_nearest_val(mars, timer=1)
| zeiss_tracking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Agent training
# In this notebook, a DQN agent is traing to solve the Navigation environment
# installing requirements
# !pip -q install ./python
# ### 1. Introducing Hyperparameters
# +
INPUT_SIZE = 37 # state size
NUM_ACTIONS = 4 # the number of possible actions
AVG_SOL_REWARD = 13.0 # over the window of 100 episode, the min average reward to stop
FC1_UNITS = 128
# I would like to urge the agent not to take too much actions without a reward
# e.g. moving left and right to avoid a blue banana will call it (idle_reward)
# for that, I penalize actions with 0 rewards
# However, that penalty shall decreas as we advance further in the episode
# Because, most likely it needs to travel further to find more yellow bananas
idle_reward_start = -0.5
idle_reward_decay = 0.8
# -
# ### 2. Setting up the environment
from unityagents import UnityEnvironment
env = UnityEnvironment(file_name="/data/Banana_Linux_NoVis/Banana.x86_64")
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 3. Instantiating the Agent
# +
from dqn_agent import Agent
from model import QNetwork
import torch
SEED = 13
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
#create the agent
agent = Agent(action_size=NUM_ACTIONS,
seed=SEED,
qnetwork_local = QNetwork(INPUT_SIZE, NUM_ACTIONS, SEED, FC1_UNITS).to(device),
qnetwork_target = QNetwork(INPUT_SIZE, NUM_ACTIONS, SEED, FC1_UNITS).to(device))
# -
# ### 3. The training loop
# +
import numpy as np
from collections import deque
import torch.optim as optim
from replay_buffer import ReplayBuffer
def dqn(n_episodes=10000, eps_start=1.0, eps_end=0.01, eps_decay=0.995):
"""Deep Q-Learning.
Params
======
n_episodes (int): maximum number of training episodes
eps_start (float): starting value of epsilon, for epsilon-greedy action selection
eps_end (float): minimum value of epsilon
eps_decay (float): multiplicative factor (per episode) for decreasing epsilon
"""
scores = [] # list containing scores from each episode
scores_window = deque(maxlen=100) # last 100 scores
eps = eps_start # initialize epsilon
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name]
state = env_info.vector_observations[0]
score = 0
idle_reward = idle_reward_start
while True:
#print(f'\ridle_reward: {idle_reward}', end="")
#print(f'idle_reward: {idle_reward}')
action = agent.act(state, eps)
env_info = env.step(action)[brain_name] # send the action to the environment
next_state = env_info.vector_observations[0] # get the next state
reward = idle_reward if env_info.rewards[0] == 0 else env_info.rewards[0] # get the reward
done = env_info.local_done[0] # see if episode has finished
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
idle_reward *= idle_reward_decay
if done:
break
scores_window.append(score) # save most recent score
scores.append(score) # save most recent score
eps = max(eps_end, eps_decay*eps) # decrease epsilon
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)), end="")
if i_episode % 100 == 0:
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_window)))
if np.mean(scores_window)>=AVG_SOL_REWARD:
print('\nEnvironment solved in {:d} episodes!\tAverage Score: {:.2f}'.format(i_episode-100, np.mean(scores_window)))
torch.save(agent.qnetwork_local.state_dict(), 'model.pt')
break
return scores
scores = dqn()
# -
# ### 4. Plot the scores
# +
import matplotlib.pyplot as plt
# %matplotlib inline
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(len(scores)), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# -
env.close()
| Navigation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="2n2UNZwpmCGm"
# # Utilisation des scripts
#
# Ce notebook vise à montrer l'utilisation des scripts du projet afin d'effectuer une augmentation de données sur une base de données.
# + [markdown] id="JVf_sKXN9H9f"
# # Installation du projet `bfc` et accès aux données
#
# Les login et mot de passe sont nécessaires pour accéder à des repos privés à partir de Google/Colab. Cela ne serait pas le cas en accès public.
# + id="CPOKLGaw9Qhx"
import os
from getpass import getpass
# + id="UK5TprNvnjMN" outputId="3f83d8b8-68b0-4322-c837-462c435b8ba3" colab={"base_uri": "https://localhost:8080/"}
# %%shell
pip uninstall opencv-python-headless
pip install opencv-python-headless==4.1.2.30
# + colab={"base_uri": "https://localhost:8080/"} id="36zlJOI5A0Go" outputId="ee84a7c8-6690-4950-fa6e-4f8917724e0d"
user = getpass('GitHub user')
password = getpass('<PASSWORD>')
os.environ['GITHUB_AUTH'] = user + ':' + password
# + colab={"base_uri": "https://localhost:8080/"} id="nF7dOnmdAc71" outputId="02b73727-86de-47ad-d0b8-bd0c509240f3"
# %%shell
git clone https://$GITHUB_AUTH@github.com/tibocour/IA.git
# + [markdown] id="pGC1hAM2nHu1"
# # Installation des dépendances
# + colab={"base_uri": "https://localhost:8080/"} id="0NA-ih6CBXLN" outputId="4381ddf8-2b8b-4113-8fd3-53b4dd9bc994"
# %%shell
pip install -r IA/requirements.txt
# + [markdown] id="d7J-CbLf5zk0"
# # Augmentation de données
#
# Téléchargement de la base de données à partir de la release GitHub (optionnel si utilisation des bases du répertoire `IA/data`)
# + id="tfVpVem7OHfC" outputId="40782ba4-6105-46c2-b9b7-22c1282073c8" colab={"base_uri": "https://localhost:8080/"}
# %%shell
wget https://github.com/tibocour/IA/releases/download/v5/project-5-at-2022-02-08-15-39-1926817e.zip
# + colab={"base_uri": "https://localhost:8080/"} id="zreMVUWh0ewv" outputId="902d74b3-63a7-4843-a926-41086dac7e4b"
# %%shell
# ls IA/data
# + [markdown] id="ZVzFfb4r2BfH"
# Pour choisir la base de données à augmenter
# + id="3akksJC8O1Q6"
# database = os.path.join("IA", "data", "megots150images.zip")
# or
database = "project-5-at-2022-02-08-15-39-1926817e.zip"
# + id="HJyT0Cn-z5gE"
os.environ.update({"database": database})
# + [markdown] id="HoMByU_X2Hqd"
# Mise en cohérence de la base issue de `label-studio` et augmentation de données
# + colab={"base_uri": "https://localhost:8080/", "height": 496} id="MgZJA6UQx2GP" outputId="14c31917-2a54-4238-850e-08fad931a985"
# %%shell
# rm -rf train_*.zip valid_*.zip
python IA/python/label_studio_voc_converter.py --zip $database
# + [markdown] id="z5cNZbbUzQjp"
# Pour Télécharger les fichiers générés (peut être long...)
# + id="0XpQLmcNPsAu"
filename = os.path.basename(database)
# + id="ZRUZSPPay4Mb"
from google.colab import files
files.download(f"train_{filename}")
files.download(f"valid_{filename}")
| notebooks/data_augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Item 12 Avoid else Block after for and while loops
# Else block runs after loop
for _ in range(1):
print(1)
else:
print("ELSE!")
for _ in range(0):
print(1)
else:
print("ELSE!")
# Using break will skip it though
for _ in range(10):
print(1)
break
else:
print("ELSE!")
# ## Things to remember
# * Else blocks after loops are executed immediately if it did not encounter a break in the loop
# * Just don't use it, since it is not intuitive and confusing
| notes/Chapter_1:_Pythonic_Thinking/Item 12 Avoid else Blocks after for and while loops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# # Imports
using Plots
using Suppressor # to make gif
using WebIO # to display gif
using Random
using StatsBase: sample # for sample/choice
import Random: shuffle!
plot([1], st=:heatmap); # speed-up
# # Chromosome
# True = dominant
@inbounds begin
struct Chromosome
alleles::Array{Bool}
end
function is_expressed(c::Chromosome) :: Bool
# Julia arrays start at 1
return c.alleles[1] || c.alleles[2]
end
function shuffle!(c1::Chromosome, c2::Chromosome)
r1, r2 = rand(1:2, 2)
l1 = [c1.alleles[r1],c2.alleles[r2]]
l2 = [c1.alleles[3-r1],c2.alleles[3-r2]] # to get the other one
copyto!(c1.alleles, l1)
copyto!(c2.alleles, l2)
end
end
# # Organism
struct Organism
genes::Array{Chromosome}
alive::Array{Bool} # Organism is immutable to prevent multiple object creations
reproduced::Array{Bool} # Organism is immutable to prevent multiple object creations
function Organism()
new([Chromosome(rand(Bool, 2)) for _ in 1:255], [true], [false])
end
end
@inbounds begin
function phenotype(o::Organism)::Int64
return count(x->is_expressed(x), (o.genes))
end
function death!(o::Organism, chance::Float64)
if rand() < chance
o.alive[1] = false
end
end
function reproduce!(o1::Organism, o2::Organism)
if o1 == o2
# In case no mate found
o1.reproduced[1] = true
return
else
if o1.alive[1] && o2.alive[1]
for i in 1:255
shuffle!.(o1.genes, o2.genes)
end
o1.reproduced[1] = o2.reproduced[1] = true
else
if o1.alive[1]
# o2 is dead
o1.reproduced[1] = true
copyto!(o2.genes, o1.genes)
o2.alive[1] = true
o2.reproduced[1] = true
else
# o1 is dead
o2.reproduced[1] = true
copyto!(o1.genes, o2.genes)
o1.alive[1] = true
o1.reproduced[1] = true
end
end
end
end
function get_chance(o::Organism, g::Int64)
@fastmath abs(phenotype(o) - g)/255 * 0.85 + 0.05
end
function virginify(o::Organism)
o.reproduced[1] = false
end
end # inbounds
# # Plate functions
@inbounds begin
function findmate(plate::Matrix{Organism}, x::Int64, y::Int64) :: Organism
xshift = [-1,0,1]
yshift = [-1,0,1]
if x == 256
deleteat!(xshift, 3)
elseif x == 1
deleteat!(xshift, 1)
end
if y == 256
deleteat!(yshift, 3)
elseif y == 1
deleteat!(yshift, 1)
end
shifts = [[xn,yn] for xn in xshift for yn in yshift if x!=0 || y!=0]
for (i, shift) in enumerate(shifts)
newx, newy = [x, y] + shift
if plate[newy, newx].reproduced[1]
deleteat!(shifts, i)
end
end
if length(shifts) == 0
push!(shifts, [0,0])
end
newx, newy = [x, y] + sample(shifts)
return plate[newy, newx]
end
function death_cycle(plate::Matrix{Organism})
for (i,o) in enumerate(plate)
x = div(i,256) # Julia is column major
death!(o, get_chance(o, x))
end
end
function reproduce_cycle(plate::Matrix{Organism})
for (i,o) in enumerate(plate)
x = div(i-1,256) + 1
y = (i-1)%256+1
if o.reproduced[1]
continue
else
reproduce!(o, findmate(plate, x, y))
end
end
end
function generation(plate::Matrix{Organism}, anim::Animation, genNum::Int64)
virginify.(plate)
death_cycle(plate)
reproduce_cycle(plate)
p = plot(phenotype.(plate), st=:heatmap, clims=(0,255), color=:greys)
#display(p)
frame(anim, p)
end
end
# # initialize
@inbounds @fastmath begin
ground = reshape([i-1 for i in 1:256 for j in 1:256], 256, 256)
plot(ground, st = :heatmap, color=:greys, clims=(0,255))
end
@inbounds @fastmath begin
plate = reshape([Organism() for _ in 1:256*256], 256,256);
end
;
@inbounds begin
anim = Animation()
p = plot(phenotype.(plate), st=:heatmap, clims=(0,255), color=:greys, title="Generation 0")
frame(anim, p)
for i in 1:100
generation(plate, anim, i)
end
end
@suppress_err begin
gif(anim, "run.gif", fps = 15)
display(Node(:img, src="run.gif?modified=$(rand())"))
end
anim = Animation()
generation(plate, anim, 0)
| Darwin Hackathon Version.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="5-7vRh7EGS30"
# ## **PySpark SQL - Getting Started Notebook**
# This notebook provides an example for installing Spark dependencies and a simple "getting started" syntax for PySpark SQL.
#
# NOTE: This notebook is designed to be used with Google's Colab notebook and the Python 3 runtime.
# + [markdown] id="aZ24BRz6FMwN"
# ## **Install Spark Dependencies**
#
#
# + id="upqpisH2IoMy"
# Install Spark dependencies
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !wget --no-cookies --no-check-certificate https://dlcdn.apache.org/spark/spark-3.2.1/spark-3.2.1-bin-hadoop3.2.tgz
# !tar zxvf spark-3.2.1-bin-hadoop3.2.tgz
# !pip install -q findspark
# !pip install pyspark
# + id="X2xvv-QnsQZs"
# !ls -al | grep spark
# + [markdown] id="3NgWpb22FchD"
# ## **Set env variables within Pyspark**
# + id="I47MHeUcyH1j"
# Set up required environment variables
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-3.2.1-bin-hadoop3.2"
# + [markdown] id="6wa2iydLFsWW"
# ## **Download Data**
# + id="rYe4U8Jnx39a"
# !wget https://raw.githubusercontent.com/zaratsian/iaa-2022/main/session_02/bikeshare_station_info.csv
# !wget https://raw.githubusercontent.com/zaratsian/iaa-2022/main/session_02/bikeshare_trips.csv
# + [markdown] id="lvMtRJwUFzie"
# ## **Import Python and PySpark Libraries**
# + id="2-SIpC_-aw0t"
import datetime
from pyspark import SparkContext
from pyspark.sql import SparkSession
from pyspark.sql.types import *
from pyspark.sql.functions import monotonically_increasing_id, col, expr, when, concat, lit, udf, split
from pyspark.ml.linalg import Vectors
from pyspark.ml.regression import GBTRegressor, LinearRegression, GeneralizedLinearRegression
from pyspark.ml.classification import GBTClassifier
from pyspark.ml.feature import VectorIndexer, VectorAssembler, StringIndexer
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml import Pipeline
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# + [markdown] id="XyT917EuF7Pv"
# ## **Initialize Spark Session**
# + id="niAz2S672M_m"
spark = SparkSession.builder.appName("Bikesharing SparkSQL").master("local[*]").getOrCreate()
# + [markdown] id="hL4NPf_ZF_OF"
# ## **Read CSV into Spark**
# + id="hC6C3wKwyBbt"
bikeshare_trips = spark.read.load('bikeshare_trips.csv', format="csv", header=True, inferSchema=True)
# + [markdown] id="Nqf2c9xOGFDb"
# ## **Display first few records**
# + id="Deym0a0pyNNu"
bikeshare_trips.show(5, truncate=False)
# + [markdown] id="N_v2cyCFGMn1"
# ## **Execute Sample SparkSQL query**
# + id="TPheTgdLyddA"
bikeshare_trips.createOrReplaceTempView("bikeshare_trips")
spark.sql("SELECT subscriber_type, count(*) as count FROM bikeshare_trips group by subscriber_type order by count desc").show(truncate=False)
# + id="nEWC_7BUzNKN"
| session_02/pyspark_sql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Load Package
import os # OS module is to interact with your operating system
# curDir = os.getcwd() #get current working directory
# print(curDir)
# os.mkdir('newDir') # creating new folder
# help(os)
root_dir ='/Users/shouzhenghuang/Desktop/finance_fraud_detect' #up direcotry
os.chdir(root_dir) #Change the current working directory to the specified path.
pwd
model_dir = './model_checkpoint'
if not os.path.exists(model_dir):
os.mkdir(model_dir)
else:
print(f"{model_dir} is already existed !")
# module will be auto-reloaded by default
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
from IPython.display import display
import copy
import pickle
pd.set_option('display.max_columns', 50)
# -
# # Load data
# +
train_path = './data/trainset-281-29.xlsx'
train_df = pd.read_excel(train_path)
predict_path = './data/testset-for-participants.xlsx'
predict_df = pd.read_excel(predict_path)
# -
# # Missing data exploratory
for df in [train_df, predict_df]:
for col in df.columns:
df[col] = df[col].apply(lambda x: None if str.lower(str(x)) in ['none', 'non', 'nan'] else x)
# +
columns = ['feature_name', 'missing_num', 'type']
train_agg = train_df.isnull().sum(axis=0).reset_index().sort_values(by=0)
train_agg.rename(columns={0:"predict_missing"}, inplace=True)
predict_agg = predict_df.isnull().sum(axis=0).reset_index().sort_values(by=0)
predict_agg.rename(columns={0:"predict_missing"}, inplace=True)
miss_agg = pd.merge(train_agg, predict_agg, on='index')
# -
train_agg
| notebook/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LHL_Bootcamp
# language: python
# name: lhl_bootcamp
# ---
# +
# import numpy
import numpy as np
# import matplotlib
import matplotlib.pyplot as plt
# set the figure size for each figure in this tutorial
plt.rcParams["figure.figsize"] = (10,6)
# -
# # Line Plot
# In this example, we will create a sine wave with x-range (0-100) generate via `Numpy`
# +
# 200 values from the interval <0,100>, equidistantly divided
x = np.linspace(0,100,200)
y = np.sin(x)
# a line plot
plt.plot(x,y,'red')
plt.show()
# -
# # Scatter Plot
# +
# 200 random values from the interval <0,10>
x = 10*np.random.rand(200,1)
# 200 random values from the interval <0,15>
y = 15*np.random.rand(200,1)
# a scatter plot
plt.scatter(x,y)
plt.show()
# -
# # Histogram
# +
# 200 random values from the interval <0,15>
y = 15*np.random.rand(200,1)
# a histogram with 20 bins
plt.hist(y,bins=20)
plt.show()
# -
# # Graphs on Common Axes
# In this example, we will plot two mathematical functions (sin(x), sin(x)*cos(x)) on one figure with shared axis.
# +
# 200 values from the interval <0,100>, equidistantly divided
x = np.linspace(0,100,200)
# sin(x) values
y1 = np.sin(x)
# sin(x)*cos(x) values
y2 =(np.sin(x))*(np.cos(x))
# a line plot of sin(x), red line
plt.plot(x,y1,'red')
# a line plot of sin(x)*cos(x), blue line
plt.plot(x,y2,'blue')
plt.show()
# -
# # Subplots
# Continuing from above, now we want a separate figure with its own axis for each function
# +
# the first figure
plt.subplot(2,1,1)
plt.plot(x,y1,'red')
plt.title('sin(x)')
# the second figure
plt.subplot(2,1,2)
plt.plot(x,y2,'blue')
plt.title('sin(x)*(cos(x))')
# automatically adjust the subplot parameters to give a specified padding
plt.tight_layout()
plt.show()
# -
# The first two parameters of plt.subplot() are the shapes of a grid with figures. In our example, we created a grid with 2 rows and one column. If we want to have our figures side by side (1 row, 2 columns) we just simply swap these parameters. The third parameter is the index of the actual figure we are plotting in. The index starts at 1 in the upper left corner of a grid and increases to the right.
# # Legends
# For this example, we will use the famous Iris dataset via Pandas and SKlearn libraries
# +
# import pandas
import pandas as pd
# import sklearn datasets
from sklearn import datasets
# +
# load iris dataset
iris = datasets.load_iris()
# create dataframe
iris_df = pd.DataFrame(iris.data, columns=iris.feature_names)
# create target
iris_df['target'] = iris.target
# map the target values to the target names
iris_df['target_name'] =iris_df.target.map(
{0: 'setosa',
1: 'versicolor',
2: 'virginica'}
)
iris_df.head()
# +
# Now, let's plot a scatter plot of sepal length and sepal width for each type of the Iris flower.
# Iris setosa
setosa = iris_df[iris_df.target_name == 'setosa']
# Iris versicolor
versicolor = iris_df[iris_df.target_name == 'versicolor']
# Iris virginica
virginica = iris_df[iris_df.target_name == 'virginica']
# plot setosa
plt.scatter(setosa['sepal length (cm)'], setosa['sepal width (cm)'],
marker ='o', color = 'red', label = 'setosa')
# plot versicolor
plt.scatter(versicolor['sepal length (cm)'], versicolor['sepal width (cm)'],
marker ='o', color = 'green', label = 'versicolor')
# plot virginica
plt.scatter(virginica['sepal length (cm)'], virginica['sepal width (cm)'],
marker ='o', color = 'blue', label = 'virginica')
# legend location
plt.legend(loc='upper right')
# plot title
plt.title('Iris flower')
# x-axis title
plt.xlabel('sepal length (cm)')
# y-axis title
plt.ylabel('sepal width (cm)')
plt.show()
# -
# By setting the label parameter in each scatter plot we set the name showed in the legend.
# # Annotations
# If we want to add annotations to the figure we created in the example above, we can do that by making the following changes in the code.
# +
# the same code as before
plt.scatter(setosa['sepal length (cm)'],setosa['sepal width (cm)'],
marker ='o', color = 'red', label = 'setosa')
plt.scatter(versicolor['sepal length (cm)'],versicolor['sepal width (cm)'],
marker ='o', color = 'green', label = 'versicolor')
plt.scatter(virginica['sepal length (cm)'],virginica['sepal width (cm)'],
marker ='o', color = 'blue', label = 'virginica')
# new lines of code
# it can be tricky to find the right coordinates for the first time
######################
plt.annotate('setosa', xy =(5.0,3.5),
xytext = (4.25,4.0), arrowprops={'color':'red'})
plt.annotate('versicolor', xy =(7.2,3.6),
xytext = (6.5,4.0), arrowprops={'color':'red'})
plt.annotate('virginica', xy =(5.05,1.95),
xytext = (5.5,1.75), arrowprops={'color':'red'})
######################
# the same code as before
plt.legend(loc='upper right')
plt.title('Iris flower')
plt.xlabel('sepal length (cm)')
plt.ylabel('sepal width (cm)')
plt.ylim(1.5,4.7)
plt.show()
# -
# To add annotations we used the `plt.annotate()` function. The xy parameter is a tuple containing the position which the arrow is pointing to. The xytext is a tuple containing the position where the text of the annotation is placed.
| Data Visualization/.ipynb_checkpoints/Matplotlib I-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Trend
# Trend is defined as the direction of growth over a period of time. There can be be upwards, downwards, and flat trends. Many times, trend is obvious by looking at a simple plot of data.
#
# There are two main reasons for trend discovery:
#
# 1. Remove a trend from nonstationary to prepare for modeling (particularly for forecasting)
# 2. Being able to charecterize the behavior by separating trend
#
# Time series models can be influenced by falsely correlated due to trends. A common example is that a rising inflation would cause an inflation for home sales prices.
# ## Setup
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import statsmodels.api as sm
import statsmodels.formula.api as smf
import statsmodels.tsa.api as smt
import random
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
# %matplotlib inline
# -
# Typically trend is shown as a linear regression which is just a straight line. But this model falls flat if there trend starts to change. There could be a global trend and more then one local trends.
# ### Linear Trend
# +
air = pd.read_csv('data/international-airline-passengers.csv', header=0, index_col=0, parse_dates=[0])
target = air.reset_index()['n_pass_thousands']
X = [i for i in range(0, len(air))]
X = np.reshape(X, (len(X), 1))
y = air['n_pass_thousands'].values
#model = LinearRegression()
#model.fit(X, y)
# calculate trend
#trend = model.predict(X)
# plot trend
from statsmodels import regression
def linreg(X,Y):
# Running the linear regression
X = sm.add_constant(X)
model = regression.linear_model.OLS(Y, X).fit()
a = model.params[0]
b = model.params[1]
X = X[:, 1]
# Return summary of the regression and plot results
X2 = np.linspace(X.min(), X.max(), 100)
Y_hat = X2 * b + a
fig = plt.figure(1, figsize=(15, 9))
ax = fig.add_subplot(111)
ax.plot(X2, Y_hat, 'r', label="Linear Trend")
ax.plot(y, label='Time Series')
ax.legend()
#X2 = np.linspace(X.min(), X.max(), 100)
#Y_hat = X2 * b + a
#plt.scatter(X, Y, alpha=0.3) # Plot the raw data
#plt.plot(X2, Y_hat, 'r', alpha=0.9); # Add the regression line, colored in red
#plt.xlabel('X Value')
plt.ylabel('Y Value')
return model.summary()
summary = linreg(X, y)
# -
# Sometimes a Linear trend doesn't show the picture. The trend could be going down, up, and flat all in one viewable period.
# ### Quadratic Trend
# +
#make a trend changing series
y = [5*np.random.normal() for j in range(50)] + [30 + 5 * np.random.normal() for j in range(50)] + [50 + 5 * np.random.normal() for j in range(50)] + [20 + 5 * np.random.normal() for j in range(50)]
X = [x for x in range(len(y))]
X = np.reshape(X, (len(X), 1))
model = LinearRegression()
model.fit(X, y)
# calculate trend
trend = model.predict(X)
#degree = 2
# experimenting with other degrees for best fit is an option
model = make_pipeline(PolynomialFeatures(3), Ridge())
model.fit(X, y)
quadratic = model.predict(X)
fig = plt.figure(1, figsize=(15, 9))
ax = fig.add_subplot(111)
ax.plot(trend, label="Linear Trend")
ax.plot(quadratic, label="Quadratic Trend")
ax.plot(X, y, label='Time Series')
ax.legend()
# -
# Here you can see the data isn't always trending up even though the Linear regression indicated a rising trend through out the entire window. Adding some polynomial regression shows a clearer picture on how the data is trending.
# ## Removing trend from a time series
# Statsmodel comes with a detrending function that can take out a trend with 3 different orders:
# * Linear
# * Quadratic
# * Constant
# +
y = np.array(y)
y = np.reshape(y, (len(y), 1))
constant = smt.detrend(y, order=0)
linear = smt.detrend(y, order=1)
quadratic = smt.detrend(y, order=2)
fig = plt.figure(1, figsize=(15, 9))
ax = fig.add_subplot(111)
ax.plot(linear, label="Linear Detrend")
ax.plot(quadratic, label="Quadratic detrend")
ax.plot(constant, label='Constant detrend')
ax.plot(X, y, label='Time Series')
ax.legend()
# -
# # Trend Estimation
#time series used
df = pd.read_csv('data/international-airline-passengers.csv', header=0, index_col=0, parse_dates=[0])['n_pass_thousands']
df.plot()
# ## Moving Average Filtering
# Moving average filtering is a linear model that isn't great for trend estimation but can provide a baseline to use to judge other models. If there is a large window size without extreme outliers, it exposes a long-term trend. It's not robust to outliers and abrupt changes, particularly for smaller window sizes.
window = 12 #the larger the window, the more robust to outliers it will be
average_signal = df.rolling(window).mean()
average_signal.plot()
df.plot()
# ## Median Filtering
# Median filter is a nonlinear model that is very robust to noise. The trend signal is most likely not to smooth. Large median windows could cause the median not to change.
window = 12
median_signal = df.rolling(window).median()
median_signal.plot()
df.plot()
# ## EWMA
# Exponentially weighted moving average is a linear model that provides different weights to values in the window. Weights are better distributed so could provide a better estimate. This model is not robust to outliers and abrupt changes. Very flexible about terms of weights and puts more emphasis on the spatial window in the signal.
window = 12
df.ewm(span=window).mean().plot() #very simple example, many more options
df.plot()
# ## Bandpass Filtering
# It filters based on frequency of the response of the series. It reduces the effect of low range (long term) and very high frequency(short-term, volatility) and exposes a trend.
# +
df = pd.read_csv('data/international-airline-passengers.csv', header=0, index_col=0, parse_dates=[0])
from scipy import signal
filter_order = 4 #need to add notes
low_cutoff_frequency = 0.001 #need to add notes
high_cutoff_frequency = 0.15 #need to add notes
b, a = signal.butter(filter_order, [low_cutoff_frequency, high_cutoff_frequency], btype='bandpass')
#plot bandpass
new_df = df
bandpass_filtered = signal.filtfilt(b, a, new_df['n_pass_thousands'])
new_df['filtered'] = new_df['n_pass_thousands'] - bandpass_filtered
new_df['n_pass_thousands'].plot()
new_df['filtered'].plot()
# -
# ## Hodrick-Prescott Filter
# This filter decomposes the time-series into a trend and cyclical component. A linear model that resembles bandpass filter and works best when noise is normally distributed.
# +
lamb = 1000
cycle, trend = sm.tsa.filters.hpfilter(df['n_pass_thousands'], lamb=lamb)
cycle.plot()
trend.plot()
df['n_pass_thousands'].plot()
# -
# ## l_1 Trend Filtering
# A nonlinear model that is generally very smooth. Yields good results when series is exponentially distributed but can be computationally expensive. Changes in trend could be used as outlier detection.
# +
#No python example
# -
| Trend.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python py3
# language: python
# name: py3
# ---
# +
import pandas as pd
import numpy as np
from tqdm import tqdm
import os
from jointSBM import jointSBM
import warnings
warnings.filterwarnings('ignore')
# -
dir_path = "data/N_100_fixed_200_alpha_20/"
edgelistsFiles = [os.path.join(dir_path,f) for f in os.listdir(dir_path) if f.find("edge")>0]
membFiles = [os.path.join(dir_path,f) for f in os.listdir(dir_path) if f.find("memb")>0]
graphs = {}
for f in edgelistsFiles:
graphName = os.path.split(f)[1].split(".")[0]
graphs[graphName] = pd.read_csv(f).values-1.
groundTruth = {}
for f in membFiles:
graphName = os.path.split(f)[1].split(".")[0]
groundTruth[graphName] = pd.read_csv(f).values[:,1]
# +
a = jointSBM(graphs, 6, groundTruth = groundTruth, n = 200, symmetric = False)
a.prepare_data()
a.fit(printLoss = True,parallel = True)
# -
print(a.measures[a.iter])
| Simple example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "26331cfec4792162c2f8fe9b562702d2", "grade": false, "grade_id": "cell-f2b2468124042cfe", "locked": true, "schema_version": 3, "solution": false, "task": false} id="5fyquvOrzzkG"
# _Lambda School Data Science, Unit 2_
#
# ---
#
# 👇 **Do not change the code in this cell.** If you're working in Google Colab, you can run this cell to install `category_encoders` and `pdpbox`.
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "6e84593629f1e735cc6423e463199480", "grade": false, "grade_id": "cell-656c869f2d287493", "locked": true, "schema_version": 3, "solution": false, "task": false} id="8vvVzINUzzkH"
# %%capture
import sys
if 'google.colab' in sys.modules:
# !pip install category_encoders
# !pip install pdpbox
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "be6f7489d8b09d515eed676f06ac2d3b", "grade": false, "grade_id": "cell-dbdc2fe26ba31738", "locked": true, "schema_version": 3, "solution": false, "task": false} id="HmDBGH3lzzkI"
# # Sprint Challenge: Predict Chicago Food Inspections 🍕
#
# In this challenge, you'll use data from the [Chicago Department of Public Health](https://www.chicago.gov/city/en/depts/cdph/provdrs/healthy_restaurants/svcs/food-protection-services.html) to build a model to predict whether a food establishment passed inspection or not.
#
# The purpose of this model is to help inspectors use their time more efficiently by identifying establishments that will likely fail inspection. In other words, this model should be able to predict whether an establishment will fail inspection *before* the inspector arrives at the establishment.
#
# # Directions
#
# This notebook contains 12 tasks, which cover the material we've learned in this sprint. Here's a summary:
#
# - **Task 1:** Importing data.
# - **Task 2:** Identifying data leakage.
# - **Task 3:** Writing a wrangle function.
# - **Task 4:** Splitting data into a feature matrix and target vector.
# - **Task 5:** Splitting data into training and validation sets.
# - **Task 6:** Establishing baseline accuracy.
# - **Task 7:** Building model with bagging predictor.
# - **Task 8:** Building model with boosting predictor.
# - **Task 9 (`stretch goal`):** Plotting ROC curves.
# - **Task 10:** Generating classification report.
# - **Task 11:** Calculating permutation importances.
# - **Task 12 (`stretch goal`):** Creating PDP interaction plot.
#
# For each task you should do the following:
#
# - Read the task instructions.
# - Write your code in the cell below the task. Delete the `raise NotImplementedError` before your start.
# - Run the testing cell below the task. If you get an error, read the error message and re-evaluate your code.
#
# **You should limit your code to the following libraries:**
#
# - `category_encoders`
# - `numpy`
# - `matplotlib`
# - `pandas`
# - `pdpbox`
# - `sklearn`
# - `xgboost`
#
# **A word of warning:** The virtual machine that will check your answers is small. So, where applicable, don't use huge values for `n_estimators` (`>100`) or `n_jobs` (keep at `-1`).
#
# If you'd like to import all your libraries at the start of your notebook, you can do so in the code block below 👇
# + deletable=false nbgrader={"cell_type": "code", "checksum": "313c53b0dc59a11bb7bfaefbf995fe2c", "grade": false, "grade_id": "cell-44be413734e30691", "locked": false, "schema_version": 3, "solution": true, "task": false} id="dajLG--uzzkI"
# YOUR CODE HERE
import pandas as pd
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import classification_report, roc_curve, plot_confusion_matrix, plot_roc_curve
import matplotlib.pyplot as plt
from category_encoders import OrdinalEncoder
from sklearn.model_selection import train_test_split
from sklearn.inspection import permutation_importance
from pdpbox.pdp import pdp_isolate, pdp_plot, pdp_interact, pdp_interact_plot
#raise NotImplementedError()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "48dd82293df0d9af1aa7efac7f7468fa", "grade": false, "grade_id": "cell-602d346d44303e87", "locked": true, "schema_version": 3, "solution": false, "task": false} id="koJcNmltzzkI"
# # I. Wrangle Data
#
# **Task 1:** Change the code below to import your dataset. Be sure to examine the columns carefully and determine if one of them should be set as the index.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "dafd956974169191567e3544c18186a5", "grade": false, "grade_id": "cell-8b9246d8d97a80ff", "locked": false, "schema_version": 3, "solution": true, "task": false} colab={"base_uri": "https://localhost:8080/", "height": 646} id="twyf9tMKzzkJ" outputId="9f3b0d1c-d136-4db8-b73a-3040ebcb8b68"
'''T1. Import data file.'''
url = 'https://drive.google.com/uc?export=download&id=1aUnQ4AJK4UtW8JL9zPyYUMtkjIgQpqKT'
df = pd.read_csv(url, parse_dates=['Inspection Date'], index_col='Inspection Date')
# YOUR CODE HERE
df.head()
#raise NotImplementedError()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "493ede874f1e2c163a74021a41d8775d", "grade": false, "grade_id": "cell-1b2eb047117d89ab", "locked": true, "schema_version": 3, "solution": false, "task": false} id="r_fXHIQ8zzkJ"
# **Task 1 Test**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "0d68298cfa23877cd61b8ba487c19dc6", "grade": true, "grade_id": "cell-e9593d4f4ed7a9bb", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="S8Orzo-DzzkJ"
'''T1 Test'''
assert isinstance(df, pd.DataFrame), 'Have you created a DataFrame named `df`?'
assert len(df) == 51916
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "1ee62609b3bf18520406b3837f7843a6", "grade": false, "grade_id": "cell-9e90dce33ddd0506", "locked": true, "schema_version": 3, "solution": false, "task": false} id="stoUh0yEzzkJ"
# **Task 2:** Given that this model is supposed to generate predictions *before* an inspection is conducted, identify the numerical feature that is an example of **data leakage.** Assign the column name to the variable `'leaky_col'`.
#
# **Remember:** Leakage is when your feature matrix includes columns that will not be available to your model at the time it make predictions.
#
#
# + deletable=false nbgrader={"cell_type": "code", "checksum": "27b878765c52a092c3c56791dde91d5d", "grade": false, "grade_id": "cell-ef24afc9168ad64f", "locked": false, "schema_version": 3, "solution": true, "task": false} id="9bw47Iz-zzkJ"
'''T2. Identify data leakage column.'''
leaky_col = 'Serious Violations Found'
# YOUR CODE HERE
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "9874ad6b513dd2c2e409aa1d6610a65e", "grade": false, "grade_id": "cell-378fd448d54e6fc0", "locked": true, "schema_version": 3, "solution": false, "task": false} id="NBtIftLizzkJ"
# **Task 2 Test**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "5a57c6a47e502a421524daf29beb7941", "grade": true, "grade_id": "cell-8429f30efb2a7bf7", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="0sTm_SM8zzkJ"
'''T2 Test'''
# This is a hidden test.
# You'll see the result when you submit to Canvas.
assert isinstance(leaky_col, str), '`leaky_col` should be type `str`.'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "b17eb5d6c465729f58b1739a11ea5b96", "grade": false, "grade_id": "cell-2f7298cea62c493e", "locked": true, "schema_version": 3, "solution": false, "task": false} id="QdrghT66zzkK"
# **Task 3:** Add to the `wrangle` function below so that it does the following:
#
# - Removes the "leaky" column.
# - Removes high-cardinality categorical columns (more than `500` categories).
# - Removes categorical columns that have only one category.
# - Removes numerical columns that are unique identifiers for each observation, not features that would affect the target.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "40e1745bb407a170e1dec8221d37fc3c", "grade": false, "grade_id": "cell-d6fc5ee398afff4f", "locked": false, "schema_version": 3, "solution": true, "task": false} id="Nv64Hwg2zzkK"
'''T3. Write wrangle function.'''
def wrangle(df):
df.drop(columns= ['State', 'Inspection ID', 'License #', leaky_col], inplace=True)
categorical_cols = df.select_dtypes('object').columns
threshold = 500
high_card_cols = [col for col in categorical_cols if df[col].nunique() > threshold]
df.drop(high_card_cols, axis = 1, inplace= True)
return df
# YOUR CODE HERE
df = wrangle(df)
df.head()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "22afb4a381b73f41b02f83ca4a0102bd", "grade": false, "grade_id": "cell-7b5d539f39db8415", "locked": true, "schema_version": 3, "solution": false, "task": false} id="oO2P9wjzzzkK"
# **Task 3 Test**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "05d5f964ad8d8bf468bd907abaa85213", "grade": true, "grade_id": "cell-49f495efb58bcd9f", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="DRmQKNdAzzkK"
'''T3 Test'''
assert df.select_dtypes('object').nunique().max() < 500, 'Have you dropped the high-cardinality columns?'
assert df.select_dtypes('object').nunique().min() > 1, 'Have you dropped the column with only one category?'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f93a157b989f3327402d7b93a31bb595", "grade": false, "grade_id": "cell-aea953fa5337fc1f", "locked": true, "schema_version": 3, "solution": false, "task": false} id="hippEpVOzzkK"
# # II. Split Data
#
# **Task 4:** Split the DataFrame `df` into the feature matrix `X` and the target vector `y`. Your target is `'Fail'`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "7bfd4151db9dd6615a624da2954138e8", "grade": false, "grade_id": "cell-b21b1c40f5478337", "locked": false, "schema_version": 3, "solution": true, "task": false} id="cS2JgwfMzzkK"
'''T4. Split feature matrix and target vector.'''
target = 'Fail'
# YOUR CODE HERE
y = df[target]
X = df.drop(target, axis=1)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "2f00ae85d10438328017da8b01b77354", "grade": false, "grade_id": "cell-d9a64e5a6bd2a37d", "locked": true, "schema_version": 3, "solution": false, "task": false} id="xj-csbySzzkK"
# **Task 4 Test**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "04db76e9023f0b61187af1e39513d377", "grade": true, "grade_id": "cell-a1d912e28c9f7522", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="ShEfdBdszzkL"
'''T4 Test'''
assert y.shape == (51916,), '`y` either has the wrong number of rows, or is two-dimentional.'
assert len(X) == 51916, '`X` has the wrong number of rows.'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "e719471298c6c418489a657a500b7d0e", "grade": false, "grade_id": "cell-b575fbda93b87f6a", "locked": true, "schema_version": 3, "solution": false, "task": false} id="gzJ2h1CLzzkL"
# **Task 5:** Split your dataset into training and validation sets.
#
# - Your training set (`X_train`, `y_train`) should contain inspections conducted before 2017.
# - Your validation set (`X_val`, `y_val`) should contain inspections conducted during or after 2017.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f92b2b9f9460a17c987c23188e3c31b1", "grade": false, "grade_id": "cell-0bb47689fd4667ed", "locked": false, "schema_version": 3, "solution": true, "task": false} id="XLGehTKZzzkL"
'''T5. Split dataset into training and validation sets.'''
# YOUR CODE HERE
train_mask = df.index.year < 2017
X_train, y_train = X.loc[train_mask], y.loc[train_mask]
val_mask = df.index.year >= 2017
X_val, y_val = X.loc[val_mask], y.loc[val_mask]
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "60e77b85e7db0d0cb2522b8caa399e77", "grade": false, "grade_id": "cell-8517b2d477256843", "locked": true, "schema_version": 3, "solution": false, "task": false} id="GQwxhLpuzzkL"
# **Task 5 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "56e9e6ef918d3a662decf3f6d67dfd01", "grade": true, "grade_id": "cell-52cf3ef1934a4278", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="eko6H4k4zzkL"
'''T5 Test'''
assert len(X_train) == len(y_train) == 41827, 'Your training set has the wrong number of observations.'
assert len(X_val) == len(y_val) == 10089, 'Your validation set has the wrong number of observations.'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "0641d242faca29af591ebea98bc88ed6", "grade": false, "grade_id": "cell-2e9a4c74f50ed0fc", "locked": true, "schema_version": 3, "solution": false, "task": false} id="VXDWnx-hzzkL"
# # III. Establish Baseline
#
# **Task 6:** Establish the baseline accuracy score for this classification problem using your training set. Save the score to the variable `baseline_acc`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b658c6901efe4fe564387be697265352", "grade": false, "grade_id": "cell-3d21cc97649be107", "locked": false, "schema_version": 3, "solution": true, "task": false} id="QFGBAjcIzzkL"
'''T6. Establish baseline accuracy.'''
# YOUR CODE HERE
baseline_acc = y_train.value_counts(normalize=True).max()
print('Baseline accuracy:', baseline_acc)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "6098c9452685d029c07b96f5295b5c1d", "grade": false, "grade_id": "cell-56d5801c8831c15b", "locked": true, "schema_version": 3, "solution": false, "task": false} id="HBuy8RgozzkL"
# **Task 6 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "8527a8f2e79c09d69519059e56c54272", "grade": true, "grade_id": "cell-abdc4cbe95e9d1da", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="70FQjyeCzzkL"
'''T6 Test'''
assert isinstance(baseline_acc, float), '`baseline_acc` should be type float. Have you defined the variable?'
assert 0.0 <= baseline_acc <= 1.0
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "8a0f7eab56c05e92a441333652ccf6cf", "grade": false, "grade_id": "cell-7d68939c4eced62c", "locked": true, "schema_version": 3, "solution": false, "task": false} id="jQ0soA0NzzkL"
# # IV. Build Model
#
# In this section, you want to answer the question: Which ensemble method performs better with this data — bagging or boosting?
#
# **Task 7:** Build a model that includes a bagging predictor (`RandomForest`). Your predictor should be part of a pipeline named `model_bag` that includes any transformers that you think are necessary.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "49a48b340c0ee7f9630c3ce57e4ca439", "grade": false, "grade_id": "cell-889285d53fdbe282", "locked": false, "schema_version": 3, "solution": true, "task": false} id="jnmm1LGvzzkL"
'''T7. Build model with bagging predictor.'''
# YOUR CODE HERE
model_bag = make_pipeline(
OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier(random_state=42, max_depth=12)
)
model_bag.fit(X_train, y_train);
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "81055fe5d87448fd8e4aff2ca4f10ea0", "grade": false, "grade_id": "cell-72dac6ede9a13038", "locked": true, "schema_version": 3, "solution": false, "task": false} id="fr3Ehm7szzkM"
# **Tast 7 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "57725e1ca8a837d8fa761271f994ad44", "grade": true, "grade_id": "cell-cddc5d7d2170877b", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="2kf5-SlKzzkM"
'''T7 Testing'''
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
assert isinstance(model_bag, Pipeline), '`model_bag` is the wrong data type. Have you assigned your pipeline to the correct variable name?'
assert isinstance(model_bag[-1], RandomForestClassifier), 'Your predictor should be a `RandomForestClassifier`.'
assert hasattr(model_bag[-1], 'feature_importances_'), 'Have you trained your model?'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "5e3c031075213c671f8f9b321585e9eb", "grade": false, "grade_id": "cell-d9750931390fe58f", "locked": true, "schema_version": 3, "solution": false, "task": false} id="Nh3r5FKfzzkM"
# **Task 8:** Build a model that includes a boosting predictor (`GradientBoostingClassifier` from `sklearn` or `XGBClassifier` from `xgboost`). Your predictor should be part of a pipeline named `model_boost` that includes any transformers that you think are necessary.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "a30d11fa6b0d4e143f4572b0baf65afb", "grade": false, "grade_id": "cell-37f16b5811ae5223", "locked": false, "schema_version": 3, "solution": true, "task": false} id="Vw0G2AjSzzkM"
'''T8. Build model with boosting predictor.'''
# YOUR CODE HERE
model_boost = make_pipeline(
OrdinalEncoder(),
SimpleImputer(),
GradientBoostingClassifier(random_state=42)
)
model_boost.fit(X_train, y_train);
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "64d1c2ff5004fe02082dc204299e0e70", "grade": false, "grade_id": "cell-3699731f62fa5db3", "locked": true, "schema_version": 3, "solution": false, "task": false} id="1nYAfwAtzzkM"
# **Task 8 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "35744289b60d1556e064bc09da544566", "grade": true, "grade_id": "cell-90deb42a1c052402", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="So08LGtdzzkM"
'''T8 Testing'''
from xgboost import XGBClassifier
from sklearn.ensemble import GradientBoostingClassifier
assert isinstance(model_boost, Pipeline), '`model_boost` is the wrong data type. Have you assigned your pipeline to the correct variable name?'
assert any([isinstance(model_boost[-1], XGBClassifier),
isinstance(model_boost[-1], GradientBoostingClassifier)]), 'Your predictor should be `XGBClassifier` or `GradientBoostingClassifier`.'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "ebed19854d947c07608d2cb6c356f7ec", "grade": false, "grade_id": "cell-dc041ac00c805cff", "locked": true, "schema_version": 3, "solution": false, "task": false} id="8khTSXZszzkM"
# # V. Check Metrics
#
# Here are the accuracy scores for your two models. Did you beat the baseline? Which of your two models appears to perform better on your validation set?
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "fc12491c98afa1dd0767422ce0a07b22", "grade": false, "grade_id": "cell-c0206a761fccab6c", "locked": true, "schema_version": 3, "solution": false, "task": false} id="T17r-tpFzzkM"
print('Bagging Model')
print('Training accuracy:', model_bag.score(X_train, y_train))
print('Validation accuracy:', model_bag.score(X_val, y_val))
print()
print('Boosting Model')
print('Training accuracy:', model_boost.score(X_train, y_train))
print('Validation accuracy:', model_boost.score(X_val, y_val))
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "044f92b1e1321a290f39d2a3d4f756ed", "grade": false, "grade_id": "cell-17e8e5433e896bc5", "locked": true, "schema_version": 3, "solution": false, "task": false} id="zMM8qc91zzkM"
# **Task 9 (`stretch_goal`):** Plot the ROC-curve for both of your models (you can plot them one-at-a-time, side-by-side, or in the same plot).
# + deletable=false nbgrader={"cell_type": "code", "checksum": "e4a418ccf53481f58016cf1828e973da", "grade": false, "grade_id": "cell-769e4a780bb22283", "locked": false, "schema_version": 3, "solution": true, "task": false} id="CXrhjwzwzzkM"
'''T9. Plot ROC-curve.'''
# YOUR CODE HERE
Boost = plot_roc_curve(model_boost, X_val, y_val)
Bag = plot_roc_curve(model_bag,
X_val,
y_val,
ax=Boost.ax_)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c2a7559346e95bf6ecd31e90dcefd3be", "grade": false, "grade_id": "cell-1b8571c3a6a034f5", "locked": true, "schema_version": 3, "solution": false, "task": false} id="g_JO0dLxzzkM"
# **Task 10:** Choose one of your models based on your validation accuracy score or your ROC curves. Then create a classification report for that model using your validation data. Save the text of the report to the variable name `model_cr`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2ef340cbc8ec53da648b929c0bab96ef", "grade": false, "grade_id": "cell-49891c4ce9bf5f37", "locked": false, "schema_version": 3, "solution": true, "task": false} id="UEBMzvjLzzkM"
'''T10. Generate classification report for one model.'''
from sklearn.metrics import classification_report
# YOUR CODE HERE
model_cr = classification_report(y_val, model_bag.predict(X_val))
print(model_cr)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "103296abc9f51aa4b883c35c418275cc", "grade": false, "grade_id": "cell-7b5374efd0e40c69", "locked": true, "schema_version": 3, "solution": false, "task": false} id="a8Zq3zEKzzkM"
# **Task 10 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "110853de9aaaba37cb2fe601091b1e7d", "grade": true, "grade_id": "cell-94e04c938f3f5f84", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="ZDnI8fYdzzkN"
assert isinstance(model_cr, str), '`model_cr` should be type `str`.'
assert all(term in model_cr for term in ['precision', 'recall', 'f1-score', 'support']), 'Is this a classification report?'
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "00449a4317e9b4d383f2ca7d58a2b0af", "grade": false, "grade_id": "cell-d2b4843352d3085a", "locked": true, "schema_version": 3, "solution": false, "task": false} id="66nPcMbmzzkN"
# **Task 11:** Using your best model, create a DataFrame `permutation_importances` with the model's permutation importances based on your validation data.
#
# - The index of the DataFrame should be your feature names.
# - The first column should be the mean importance.
# - The second column should be the importance standard deviation.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "f8e984b2a5083b74b7eb0abec46f8d63", "grade": false, "grade_id": "cell-72936eec6980072b", "locked": false, "schema_version": 3, "solution": true, "task": false} id="6ozugglWzzkN"
'''T11. Create DataFrame of permutation importances.'''
# YOUR CODE HERE
perm_importances = permutation_importance(model_bag,
X_val,
y_val,
n_repeats=5,
n_jobs=-1,
random_state=42)
data = {'mean': perm_importances['importances_mean'],
'std': perm_importances['importances_std']}
permutation_importances = pd.DataFrame(data, index=X_val.columns).sort_values(by='mean')
permutation_importances.head()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "dccb1015d0f5c2f8b23f780eb8e4abf7", "grade": false, "grade_id": "cell-9eb949d189e401bc", "locked": true, "schema_version": 3, "solution": false, "task": false} id="d-HTADpCzzkN"
# **Task 11 Testing**
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d84e55cd6e77a52c576972ab68477c90", "grade": true, "grade_id": "cell-a4d8990e7070c2dd", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} id="QL9Miip2zzkN"
'''Task 11 Test'''
assert isinstance(permutation_importances, pd.DataFrame), '`permutation_importances` should be type `DataFrame`.'
assert permutation_importances.shape == (7,2)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f005618a5eb29cde9fbc73ab717b0e1c", "grade": false, "grade_id": "cell-3d8938c1715a596d", "locked": true, "schema_version": 3, "solution": false, "task": false} id="jSyF4VmtzzkN"
# **Task 12 (`stretch goal`):** Using your best model, create a PDP interaction plot to examine how `'Latitude'` and `'Longitude'` inform predictions. Remember to user your validation data.
#
# **Note:** Because of the way that `pdp_interact` works, it will throw an error if there are `NaN` values in your validation set. To avoid this problem, be sure to set `dataset` to `X_val.dropna()`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "ea5a8422d2efafe17ee63a77c5db1e41", "grade": false, "grade_id": "cell-224d3b408f9bdd88", "locked": false, "schema_version": 3, "solution": true, "task": false} id="SVFbtxz2zzkN"
'''T12. Create PDP interaction plot for "Latitude" and "Longitude".'''
features = ['Longitude', 'Latitude']
# YOUR CODE HERE
isolate = pdp_interact(
model_bag,
dataset = X_val.dropna(),
model_features = X_val.columns,
features = features
)
pdp_interact_plot(isolate, plot_type='grid', feature_names= features)
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "ea11277dc1543cb3a51dc7cf050af8a9", "grade": false, "grade_id": "cell-60908df556f5057e", "locked": true, "schema_version": 3, "solution": false, "task": false} id="0ipFz6cazzkN"
# What do you think? Is there a relationship between location and failing a food saftey inspection?
# + [markdown] id="z-EVUa158mdE"
# #### Based off the pdp interaction plot, there is a relationship between location and failing a food safety inspection.
| DS_Unit_3_Sprint_3_Sprint_Challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
from sklearn.linear_model import LinearRegression
# -
data = pd.read_csv(r'C:\Users\Dylan\Desktop\ML\SATandGPAplusRandom.csv')
data.describe()
# dependent variables
x = data[['SAT', 'Rand 1,2,3']]
# independent variable
y = data['GPA']
reg = LinearRegression()
reg.fit(x,y)
# Coef
reg.coef_
#Intercepts
reg.intercept_
# +
# R squared
reg.score(x,y)
# +
# need adjusted R squared due to multiple variables
# Formula for Adjusted R^2
# -
# adjR^2 = 1 - ( 1 - R^2) * (n-1) / (n - p - 1)
x.shape
# +
r2 = reg.score(x,y)
n = x.shape[0]
p = x.shape[1]
adjustedR2 = 1-(1-r2)*(n-1)/(n-p-1)
# -
print(adjustedR2)
# +
# one or more of the predictors have little explanatory power
# -
from sklearn.feature_selection import f_regression
f_regression(x,y)
p_values = f_regression(x,y)[1]
print(p_values)
p_values.round(3)
print(x.columns)
# +
# SAT Useful, while RAND 1,2,3 is useless
# -
| images/Multiple Linear Regression and F-test to find relevant variables with sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: FastAi
# language: python
# name: fastai
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <center>
#
# # ⚔Fast.ai中的尝试
#
# <br>
# <br>
#
# </center>
#
# <div class="alert alert-block alert-success">
# <b>Fast.ai Version:</b> v2.0.17 <b>Draft Date:</b> 2020-06-09
# <br>
#
# <br>
# <br>
#
# 记录`fast.ai` 使用中的一些尝试 ...
# </div>
#
# -
from fastai.vision.all import *
# ## 标记
# + [markdown] hidden=true slideshow={"slide_type": "fragment"}
# <div class="alert alert-block alert-info">
# <b>提示💡:</b>
#
# 蓝色给大家一些新的提示要点,可以进行拓展操作.
#
# <br>
# <b>经验分享👣</b>
#
# 分享一些经验,踩过的坑.
# </div>
# + [markdown] hidden=true slideshow={"slide_type": "fragment"}
# <div class="alert alert-block alert-warning">
# <b>知识回顾📕:</b>
#
# 这部分会对学过的知识进行关联和回顾,便于大家进行巩固.
# </div>
# + [markdown] hidden=true slideshow={"slide_type": "fragment"}
# <div class="alert alert-block alert-success">
# 概要✅:
#
# 绿色部分为章节内容核心点,提醒关注.
# </div>
# + [markdown] hidden=true slideshow={"slide_type": "fragment"}
# <div class="alert alert-block alert-danger">
# <b>重要📌</b>
#
# 红色标识了重点注意事项,提醒大家注意.
# </div>
# + [markdown] hidden=true
# <details><summary>🤔--这是被隐藏的内容,思考之后再打开👇</summary>
# <center>
# >
# >
# 打开之后你将看到内容的详情 ...
# >
# >
# </center>
# </details>
# -
# ## 自定义Transforms
def centerSizeCrop(image, crop_size):
rows, cols = image.shape[:2]
x = round((cols - crop_size) / 2.0)
y = round((rows - crop_size) / 2.0)
img = image[y:y+crop_size, x:x+crop_size]
return img
def fixsizeCrop(img, size,crop):
img = cv2.cvtColor(np.asarray(img),cv2.COLOR_RGB2BGR)
img = cv2.resize(img, size)
img = centerSizeCrop(img, crop)
return Image.fromarray(cv2.cvtColor(img,cv2.COLOR_BGR2RGB))
class SizeTfm(Transform):
def __init__(self,size,crop):
if isinstance(size,int): size=(size,size)
self.size = size
self.crop = crop
def encodes(self, img: Image.Image):
return fixsizeCrop(img, self.size,self.crop)
path = untar_data(URLs.PETS)
files = get_image_files(path/'images')
img = PILImage.create(files[0])
img
img.size
SizeTfm((500,333),320)(img)
# ## 如何获取validation sets的一个批数据
path = untar_data(URLs.PETS)
files = get_image_files(path/"images")
def label_func(f): return f[0].isupper()
data = ImageDataLoaders.from_name_func(path, files, label_func, item_tfms=Resize(224))
with data.valid.fake_l.no_multiproc():
out = next(iter(data.valid))
out[0].shape, out[1].shape
| 3.fastai/01.EfficientNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Processor snippets
#
# Here are a bunch of Processor related snippets either used for processes that work well of some specific machine.
| notebooks/snippet_processor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: deep-learning-python3
# language: python
# name: deep-learning
# ---
# + [markdown] deletable=true editable=true run_control={"frozen": false, "read_only": false}
# # Overview
#
# This project tries to train a recurrent neural network to learn how to automatically punctuate a sentence by reading it character by character.
#
# ### Surprises
#
# use GRU.
#
# ### Todo:
# - [ ] get data
#
# ### Done:
# - [x] Add temperature to generator
# - [x] get training to work
# - [x] use optim and Adam
# - [x] add self-feeding generator
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from utils import forward_tracer, backward_tracer, Char2Vec, num_flat_features
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import numpy as np
from tqdm import tqdm
from IPython.display import clear_output
import os
from bs4 import BeautifulSoup
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def get_content(fn):
with open(fn, 'r') as f:
source = ""
for line in f:
source += line
return source
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def source_gen(path="../engadget_data/"):
for child, folders, files in os.walk(path):
for fn in files:
if fn[0] is ".":
pass
else:
src = get_content(path + fn)
soup = BeautifulSoup(src, 'html.parser')
src = soup.getText()
yield fn, src
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
for fn, text in source_gen():
print(text)
break
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
import math
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def batch_gen(seq_length, source):
s_l = len(source)
b_n = math.ceil(s_l/seq_length)
s_pad = source + " " * (b_n * seq_length - s_l)
for i in range(b_n):
yield s_pad[i*seq_length: (i+1)*seq_length]
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def get_chars():
step = 0
freq = {}
keys = []
for file_name, source in tqdm(source_gen()):
for char in source:
try:
freq[char] += 1
except KeyError:
freq[char] = 1
keys.append(char)
#if step%10000 == 9999:
# print(str(step) + ": ln: " + str(len(keys)) + str(["".join(keys)]))
return keys, freq
ks, freqs = get_chars()
print("".join(ks))
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
order = np.argsort([freqs[k] for k in ks])[::-1]
chars_ordered = "".join(np.array([k for k in ks])[order])
print(chars_ordered[:140])
plt.title('Frequency of each character')
plt.plot(np.array([math.log10(freqs[k]) for k in ks])[order])
plt.ylabel('$\log_10$ frequency')
plt.xlabel('character index')
plt.show()
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
input_chars = list(" ABCDEFGHIJKLMNOPQRSTUVWXYZabcdefghijklmnopqrstuvwxyz01234567890")
output_chars = ["<nop>", "<cap>"] + list(".,;:?!\"'")
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
class GruRNN(nn.Module):
def __init__(self, input_size, hidden_size, output_size, layers=1, bi=False):
super(GruRNN, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.output_size = output_size
self.layers = layers
self.bi_mul = 2 if bi else 1
self.encoder = nn.Linear(input_size, hidden_size)
self.gru = nn.GRU(input_size, hidden_size, self.layers, bidirectional=bi)
self.decoder = nn.Linear(hidden_size * self.bi_mul, output_size)
self.softmax = F.softmax
def forward(self, x, hidden):
#embeded = self.encoder(x)
embeded = x
#print(embeded.view(-1, 1, self.input_size).size())
#print(hidden.size())
gru_output, hidden = self.gru(embeded.view(-1, 1, self.input_size), hidden.view(self.layers * self.bi_mul, -1, self.hidden_size))
#print(gru_output.size())
output = self.decoder(gru_output.view(-1, self.hidden_size * self.bi_mul))
return output, hidden
def init_hidden(self, random=False):
if random:
return Variable(torch.randn(self.layers * self.bi_mul, self.hidden_size))
else:
return Variable(torch.zeros(self.layers * self.bi_mul, self.hidden_size))
"""
input_size = 105
hidden_size = 105
output_size = 105
layers = 2
gRNN = GruRNN(input_size, hidden_size, output_size, layers)
gRNN(Variable(torch.FloatTensor(10000, 105)),
Variable(torch.FloatTensor(layers, 105)))"""
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
class Engadget():
def __init__(self, model, char2vec=None, output_char2vec=None):
self.model = model
if char2vec is None:
self.char2vec = Char2Vec()
else:
self.char2vec = char2vec
if output_char2vec is None:
self.output_char2vec = self.char2vec
else:
self.output_char2vec = output_char2vec
self.loss = 0
self.losses = []
def init_hidden_(self, random=False):
self.hidden = model.init_hidden(random)
return self
def save(self, fn="GRU_Engadget.tar"):
torch.save({
"hidden": self.hidden,
"state_dict": model.state_dict(),
"losses": self.losses
}, fn)
def load(self, fn):
checkpoint = torch.load(fn)
self.hidden = checkpoint['hidden']
model.load_state_dict(checkpoint['state_dict'])
self.losses = checkpoint['losses']
def setup_training(self, learning_rate):
self.optimizer = optim.Adam(model.parameters(), lr=learning_rate)
self.loss_fn = nn.CrossEntropyLoss()
self.init_hidden_()
def reset_loss(self):
self.loss = 0
def forward(self, input_text, target_text):
self.hidden = self.hidden.detach()
self.optimizer.zero_grad()
self.next_(input_text)
target_vec = Variable(self.output_char2vec.char_code(target_text))
self.loss += self.loss_fn(self.output, target_vec)
def descent(self):
if self.loss is 0:
print(self.loss)
print('Warning: loss is zero.')
return
self.loss.backward()
self.optimizer.step()
self.losses.append(self.loss.cpu().data.numpy())
self.reset_loss()
def embed(self, input_data):
self.embeded = Variable(self.char2vec.one_hot(input_data))
return self.embeded
def next_(self, input_text):
self.output, self.hidden = self.model(self.embed(input_text), self.hidden)
return self
def softmax_(self, temperature=0.5):
self.softmax = self.model.softmax(self.output/temperature)
return self
def output_chars(self, start=None, end=None):
indeces = torch.multinomial(self.softmax[start:end]).view(-1)
return self.output_char2vec.vec2list(indeces)
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def apply_punc(text_input, text_output):
result = ""
for char1, char2 in zip(text_input, text_output):
if char2 == "<cap>":
result += char1.upper()
elif char2 != "<nop>":
result += char1 + char2
else:
result += char1
return result
result = apply_punc("t s", ['<cap>', '<nop>', ','])
assert(result == "T s,")
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def extract_punc(string_input, input_chars, output_chars):
input_source = []
output_source = []
for i, char in enumerate(string_input):
# print(i, char)
if char.isupper() and len(output_source) > 0:
output_source.append("<cap>")
input_source.append(char.lower())
elif char in output_chars and len(output_source) > 0:
output_source[-1] = char
elif char in input_chars:
input_source.append(char)
output_source.append("<nop>")
return input_source, output_source
i, o = extract_punc("This's a simple ATI chassis.", input_chars, output_chars)
result = apply_punc("".join(i), o)
print(result)
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
char2vec = Char2Vec(chars=input_chars, add_unknown=True)
output_char2vec = Char2Vec(chars = output_chars)
input_size = char2vec.size
output_size = output_char2vec.size
print("input_size is: " + str(input_size) + "; ouput_size is: " + str(output_size))
hidden_size = input_size
layers = 1
model = GruRNN(input_size, hidden_size, output_size, layers=layers, bi=True)
egdt = Engadget(model, char2vec, output_char2vec)
egdt.load('./Gru_Engadget_1_layer_bi.tar')
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
learning_rate = 2e-3
egdt.setup_training(learning_rate)
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
model.zero_grad()
egdt.reset_loss()
seq_length = 200
for epoch_num in range(40):
step = 0
for file_name, source in tqdm(source_gen()):
for source_ in batch_gen(seq_length, source):
step += 1
input_source, output_source = extract_punc(source_, egdt.char2vec.chars, egdt.output_char2vec.chars)
try:
egdt.forward(input_source, output_source)
if step%1 == 0:
egdt.descent()
except KeyError:
print(source)
raise KeyError
if step%400 == 399:
clear_output(wait=True)
print('Epoch {:d}'.format(epoch_num))
egdt.softmax_()
fig = plt.figure(figsize=(16, 8))
fig.subplots_adjust(hspace=0.0625)
plt.subplot(131)
plt.title("Input")
plt.imshow(egdt.embeded[:130].data.byte().numpy(), cmap="Greys_r", interpolation="none")
plt.subplot(132)
plt.title("Output")
im = plt.imshow(egdt.output[:20].data.byte().numpy(), cmap="Greys_r", interpolation="none")
cb = plt.colorbar(im, fraction=0.08); cb.outline.set_linewidth(0)
plt.subplot(133)
plt.title("Softmax Output")
im = plt.imshow(egdt.softmax[:20].cpu().data.numpy(), interpolation="none")
cb = plt.colorbar(im, fraction=0.08); cb.outline.set_linewidth(0)
plt.show()
plt.figure(figsize=(10, 3))
plt.title('Training loss')
plt.plot(egdt.losses, label="loss", linewidth=3, alpha=0.4)
plt.show()
# print(source_)
result = apply_punc(input_source, egdt.output_chars())
print(result)
# + [markdown] deletable=true editable=true run_control={"frozen": false, "read_only": false}
# ## Now use the network to generate text!
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
egdt.save('./data/Gru_Engadget_1_layer_bi.tar')
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
from ipywidgets import widgets
from IPython.display import display
# + [markdown] deletable=true editable=true run_control={"frozen": false, "read_only": false}
# **Note**: Again, using a zero initial state for the hidden state gives a much better result for the prediction.
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
def predict_next(input_text, gen_length=None, temperature=0.05):
if gen_length is None:
gen_length = len(input_text)
clear_output(wait=True)
#egdt = Engadget(model).init_hidden_(random=True)
egdt.init_hidden_()
egdt.next_(input_text)
egdt.softmax_()
output = egdt.output_chars()
#print(output)
result = apply_punc(input_text, output)
print(result)
plt.figure(figsize=(12, 9))
plt.subplot(311)
plt.title("Input")
plt.imshow(egdt.embeded[:130].data.byte().numpy().T, cmap="Greys_r", interpolation="none")
plt.subplot(312)
plt.title("Output")
plt.imshow(egdt.output[:130].data.byte().numpy().T, interpolation="none")
plt.subplot(313)
plt.title("Softmax")
plt.imshow(egdt.softmax[:130].cpu().data.numpy().T, interpolation="none")
plt.show()
predict_next(" this wont be a simple sentense it doesnt have puntuation yet the network can add", 200, 1)
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
text_input = widgets.Text()
display(text_input)
def handle_submit(sender):
#print(text_input.value)
predict_next(text_input.value, 2000, temperature=0.5)
text_input.on_submit(handle_submit)
# + [markdown] deletable=true editable=true run_control={"frozen": false, "read_only": false}
# The ge
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
# + deletable=true editable=true run_control={"frozen": false, "read_only": false}
| deep-auto-punctuation/engadget_1_layer_bi.arch/Learning Punctuations by reading Engadget.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/Rishit-dagli/TFUG-Mysuru-2020/blob/master/TFQuantum_starter.ipynb)
# # Getting started with [TensorFlow Quantum](https://www.tensorflow.org/quantum)
#
# In this notebook you will build your first hybrid quantum classical model with
# [Cirq](https://cirq.readthedocs.io/en/stable/) and TensorFlow Quantum (TFQ). We will build a very simple model to do
# binary classification in this notebook. You will then use Keras to create a wrapper for the model and simulate it to
# train and evluate the model.
# > Note: This notebook is designed to be run in Google Colab if you want to run it locally or on a Jupyter notebook you
# would skip the code cells with the `Colab only` comment.
# ## Setup
# ### Install TensorFlow 2.x (Colab only)
# Colab only
pip install -q tensorflow==2.1.0
# ### Install TensorFlow Quantum (Colab only)
# Colab only
pip install -q tensorflow-quantum
# ### Imports
# Now import TensorFlow and the module dependencies:
# +
import cirq
import random
import numpy as np
import sympy
import tensorflow as tf
import tensorflow_quantum as tfq
from matplotlib import pyplot as plt
from cirq.contrib.svg import SVGCircuit
# -
# ### Place a qubit on the grid
#
# You will then place a qubit on thee grid
qubit = cirq.GridQubit(0, 0)
# ## Prepare quantum data
#
# The first thing you would do is set up the labels and parameters for preparation of the quantum data. For simplicity
# here we have included just 2 data points `a` and `b`.
expected_labels = np.array([[1, 0], [0, 1]])
# Randonly rotate `x` and `z` axis
angle = np.random.uniform(0, 2 * np.pi)
# ## Building the quantum Circuit
#
# You will now build the quantum circuit and also convert it into a tensor
a = cirq.Circuit(cirq.ry(angle)(qubit))
b = cirq.Circuit(cirq.ry(angle + np.pi / 2)(qubit))
quantum_data = tfq.convert_to_tensor([a, b])
SVGCircuit(a)
SVGCircuit(b)
# ## Building the hybrid model
#
# This section also shows the interoperatability between TensorFlow and Cirq. With the TFQ PQC layer you can easily
# embed your quantum part of the model within a standard classical Keras model.
q_data_input = tf.keras.Input(shape = (), dtype = tf.dtypes.string)
theta = sympy.Symbol("theta")
q_model = cirq.Circuit(cirq.ry(theta)(qubit))
expectation = tfq.layers.PQC(q_model, cirq.Z(qubit))
expectation_output = expectation(q_data_input)
classifier = tf.keras.layers.Dense(2, activation = tf.keras.activations.softmax)
classifier_output = classifier(expectation_output)
# You will now define the optimizer and loss functions for your model
model = tf.keras.Model(inputs = q_data_input,
outputs = classifier_output)
model.compile(optimizer = tf.keras.optimizers.Adam(learning_rate = 0.1),
loss = tf.keras.losses.CategoricalCrossentropy())
# ## Traaining the model
#
# Training the model is just like training any other Keras model and is made easy.
history = model.fit(x = quantum_data,
y = expected_labels,
epochs = 250)
# ## Evaluating the model
plt.plot(history.history['loss'])
plt.title('model loss')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train'], loc='upper left')
plt.show()
# ## Performing inference
noise = np.random.uniform(-0.25, 0.25, 2)
test_data = tfq.convert_to_tensor([
cirq.Circuit(cirq.ry(noise[0])(qubit)),
cirq.Circuit(cirq.ry(noise[1] + np.pi/2)(qubit)),
])
# You can see in the below cell that our model does a good job with this data though it was very easy.
predictions = model.predict(test_data)
predictions
| TFQuantum_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training a Sentiment Analysis LSTM Using Noisy Crowd Labels
# In this tutorial, we'll provide a simple walkthrough of how to use Snorkel to resolve conflicts in a noisy crowdsourced dataset for a sentiment analysis task, and then use these denoised labels to train an LSTM sentiment analysis model which can be applied to new, unseen data to automatically make predictions!
#
# Specifically, we'll look at:
# 1. Loading data via SparkSQL
# 2. Creating basic Snorkel objects: `Candidates`, `Contexts`, and `Labels`
# 3. Training the `GenerativeModel` to resolve labeling conflicts
# 4. Training a simple LSTM sentiment analysis model, which can then be used on new, unseen data!
#
# Note that this is a simple tutorial meant to give an overview of the mechanics of using Snorkel-- we'll note places where more careful fine-tuning could be done!
#
# ## Installing `PySpark`
#
# Please see the [official instructions](https://spark.apache.org/docs/latest/spark-standalone.html)!
# ### Task Detail: Weather Sentiments in Tweets
#
# In this tutorial we focus on the [Weather sentiment](https://www.crowdflower.com/data/weather-sentiment/) task from [Crowdflower](https://www.crowdflower.com/).
#
# In this task, contributors were asked to grade the sentiment of a particular tweet relating to the weather. Contributors could choose among the following categories:
# 1. Positive
# 2. Negative
# 3. I can't tell
# 4. Neutral / author is just sharing information
# 5. Tweet not related to weather condition
#
# The catch is that 20 contributors graded each tweet. Thus, in many cases contributors assigned conflicting sentiment labels to the same tweet.
#
# The task comes with two data files (to be found in the `data` directory of the tutorial:
# 1. [weather-non-agg-DFE.csv](data/weather-non-agg-DFE.csv) contains the raw contributor answers for each of the 1,000 tweets.
# 2. [weather-evaluated-agg-DFE.csv](data/weather-evaluated-agg-DFE.csv) contains gold sentiment labels by trusted workers for each of the 1,000 tweets.
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
import numpy as np
from snorkel import SnorkelSession
session = SnorkelSession()
# ## Step 1: Preprocessing - Data Loading with Spark SQL and Dataframes
# First, we initialize a `SparkSession`, which manages a connection to a local Spark master which allows us to preprocess the raw data and prepare convert them to the necessary `Snorkel` format:
# +
# Initialize Spark Environment and Spark SQL
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
from pyspark.sql.types import *
from pyspark import SparkContext, SparkConf
spark = SparkSession \
.builder \
.master("local") \
.appName("Snorkel Crowdsourcing Demo") \
.config("spark.some.config.option", "some-value") \
.getOrCreate()
# -
# We can now load the raw data for our crowdsourcing task (stored in a local csv file) into a dataframe.
# +
# Load Raw Crowdsourcing Data
raw_crowd_answers = spark.read.format("csv").option("header", "true").csv("data/weather-non-agg-DFE.csv")
raw_crowd_answers.printSchema()
# Load Groundtruth Crowdsourcing Data
gold_crowd_answers = spark.read.format("csv").option("header", "true").csv("data/weather-evaluated-agg-DFE.csv")
gold_crowd_answers.createOrReplaceTempView("gold_crowd_answers")
# Filter out low-confidence answers
gold_answers = spark.sql("SELECT tweet_id, sentiment, tweet_body FROM gold_crowd_answers WHERE correct_category ='Yes' and correct_category_conf = 1").orderBy("tweet_id")
# Keep Only the Tweets with Available Groundtruth
candidate_labeled_tweets = raw_crowd_answers.join(gold_answers, raw_crowd_answers.tweet_id == gold_answers.tweet_id).select(raw_crowd_answers.tweet_id,raw_crowd_answers.tweet_body,raw_crowd_answers.worker_id,raw_crowd_answers.emotion)
# -
# As mentioned above, contributors can provide conflicting labels for the same tweet:
candidate_labeled_tweets.select("worker_id", "emotion", "tweet_body").orderBy("tweet_id").show()
# ## Step 2: Generating Snorkel Objects
#
# ### `Candidates`
#
# `Candidates` are the core objects in Snorkel representing objects to be classified. We'll use a helper function to create a custom `Candidate` sub-class, `Tweet`, with values representing the possible labels that it can be classified with:
# +
from snorkel.models import candidate_subclass
values = list(map(
lambda r: r.emotion,
candidate_labeled_tweets.select("emotion").distinct().collect()
))
Tweet = candidate_subclass('Tweet', ['tweet'], values=values)
# -
# ### `Contexts`
#
# All `Candidate` objects point to one or more `Context` objects, which represent the raw data that they are rooted in. In this case, our candidates will each point to a single `Context` object representing the raw text of the tweet.
#
# Once we have defined the `Context` for each `Candidate`, we can commit them to the database. Note that we also split into two sets while doing this:
#
# 1. **Training set (`split=0`):** The tweets for which we have noisy, conflicting crowd labels; we will resolve these conflicts using the `GenerativeModel` and then use them as training data for the LSTM
#
# 2. **Test set (`split=1`):** We will pretend that we do not have any crowd labels for this split of the data, and use these to test the LSTM's performance on unseen data
# +
from snorkel.models import Context, Candidate
from snorkel.contrib.models.text import RawText
# Make sure DB is cleared
session.query(Context).delete()
session.query(Candidate).delete()
# Now we create the candidates with a simple loop
tweet_bodies = candidate_labeled_tweets \
.select("tweet_id", "tweet_body") \
.orderBy("tweet_id") \
.distinct()
# Generate and store the tweet candidates to be classified
# Note: We split the tweets in two sets: one for which the crowd
# labels are not available to Snorkel (test, 10%) and one for which we assume
# crowd labels are obtained (to be used for training, 90%)
total_tweets = tweet_bodies.count()
test_split = total_tweets*0.1
for i, t in enumerate(tweet_bodies.collect()):
split = 1 if i <= test_split else 0
raw_text = RawText(stable_id=t.tweet_id, name=t.tweet_id, text=t.tweet_body)
tweet = Tweet(tweet=raw_text, split=split)
session.add(tweet)
session.commit()
# -
# ### `Labels`
#
# Next, we'll store the labels for each of the training candidates in a sparse matrix (which will also automatically be saved to the Snorkel database), with one row for each candidate and one column for each crowd worker:
# +
from snorkel.annotations import LabelAnnotator
from collections import defaultdict
# Extract worker votes
# Cache locally to speed up for this small set
worker_labels = candidate_labeled_tweets.select("tweet_id", "worker_id", "emotion").collect()
wls = defaultdict(list)
for row in worker_labels:
wls[row.tweet_id].append((row.worker_id, row.emotion))
# Create a label generator
def worker_label_generator(t):
"""A generator over the different (worker_id, label_id) pairs for a Tweet."""
for worker_id, label in wls[t.tweet.name]:
yield worker_id, label
labeler = LabelAnnotator(label_generator=worker_label_generator)
# %time L_train = labeler.apply(split=0)
L_train
# -
# Finally, we load the ground truth ("gold") labels for both the training and test sets, and store as numpy arrays"
# +
gold_labels = defaultdict(list)
# Get gold labels in verbose form
verbose_labels = dict([(t.tweet_id, t.sentiment)
for t in gold_answers.select("tweet_id", "sentiment").collect()])
# Iterate over splits, align with Candidate ordering
for split in range(2):
cands = session.query(Tweet).filter(Tweet.split == split).order_by(Tweet.id).all()
for c in cands:
gold_labels[split].append(values.index(verbose_labels[c.tweet.name]) + 1)
train_cand_labels = np.array(gold_labels[0])
test_cand_labels = np.array(gold_labels[1])
# -
# ## Step 3: Resolving Crowd Conflicts with the Generative Model
#
# Until now we have converted the raw crowdsourced data into a labeling matrix that can be provided as input to `Snorkel`. We will now show how to:
#
# 1. Use `Snorkel's` generative model to learn the accuracy of each crowd contributor.
# 2. Use the learned model to estimate a marginal distribution over the domain of possible labels for each task.
# 3. Use the estimated marginal distribution to obtain the maximum a posteriori probability estimate for the label that each task takes.
# +
# Imports
from snorkel.learning.gen_learning import GenerativeModel
# Initialize Snorkel's generative model for
# learning the different worker accuracies.
gen_model = GenerativeModel(lf_propensity=True)
# -
# Train the generative model
gen_model.train(
L_train,
reg_type=2,
reg_param=0.1,
epochs=30
)
# ### Infering the MAP assignment for each task
# Each task corresponds to an indipendent random variable. Thus, we can simply associate each task with the most probably label based on the estimated marginal distribution and get an accuracy score:
accuracy = gen_model.score(L_train, train_cand_labels)
print("Accuracy: {:.10f}".format(accuracy))
# ### Majority vote
#
# It seems like we did well- but how well? Given that this is a fairly simple task--we have 20 contributors per tweet (and most of them are far better than random)--**we expect majority voting to perform extremely well**, so we can check against majority vote:
# +
from collections import Counter
# Collect the majority vote answer for each tweet
mv = []
for i in range(L_train.shape[0]):
c = Counter([L_train[i,j] for j in L_train[i].nonzero()[1]])
mv.append(c.most_common(1)[0][0])
mv = np.array(mv)
# Count the number correct by majority vote
n_correct = np.sum([1 for i in range(L_train.shape[0]) if mv[i] == train_cand_labels[i]])
print("Accuracy: {:.10f}".format(n_correct / float(L_train.shape[0])))
print("Number incorrect: {:.0f}".format(L_train.shape[0] - n_correct))
# -
# We see that while majority vote makes 9 errors, the Snorkel model makes only 2! What about an average crowd worker?
# ### Average human accuracy
#
# We see that the average accuracy of a single crowd worker is in fact much lower:
accs = []
for j in range(L_train.shape[1]):
n_correct = np.sum([1 for i in range(L_train.shape[0]) if L_train[i,j] == train_cand_labels[i]])
acc = n_correct / float(L_train[:,j].nnz)
accs.append(acc)
print("Mean Accuracy: {:.10f}".format(np.mean(accs)))
# ## Step 4: Training an ML Model with Snorkel for Sentiment Analysis over Unseen Tweets
#
# In the previous step, we saw that Snorkel's generative model can help to denoise crowd labels automatically. However, what happens when we don't have noisy crowd labels for a tweet?
#
# In this step, we'll use the estimates of the generative model as _probabilistic training labels_ to train a simple LSTM sentiment analysis model, which takes as input a tweet **for which no crowd labels are available** and predicts its sentiment.
#
# First, we get the probabilistic training labels (_training marginals_) which are just the marginal estimates of the generative model:
train_marginals = gen_model.marginals(L_train)
from snorkel.annotations import save_marginals
save_marginals(session, L_train, train_marginals)
# Next, we'll train a simple LSTM:
# +
# from snorkel.learning import TextRNN - v0.6.3
from snorkel.learning.tensorflow import TextRNN # v0.7-beta
train_kwargs = {
'lr': 0.01,
'dim': 100,
'n_epochs': 200,
'dropout': 0.2,
'print_freq': 5
}
lstm = TextRNN(seed=1701, cardinality=Tweet.cardinality)
train_cands = session.query(Tweet).filter(Tweet.split == 0).order_by(Tweet.id).all()
lstm.train(train_cands, train_marginals, **train_kwargs)
# -
test_cands = session.query(Tweet).filter(Tweet.split == 1).order_by(Tweet.id).all()
accuracy = lstm.score(test_cands, test_cand_labels)
print("Accuracy: {:.10f}".format(accuracy))
# We see that we're already close to the accuracy of an average crowd worker! If we wanted to improve the score, we could tune the LSTM model using grid search (see the Intro tutorial), use [pre-trained word embeddings](https://nlp.stanford.edu/projects/glove/), or many other common techniques for getting state-of-the-art scores. Notably, we're doing this without using gold labels, but rather noisy crowd-labels!
#
# For more, checkout the other tutorials!
| tutorials/crowdsourcing/Crowdsourced_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # [Day 8: Seven Segment Search](https://adventofcode.com/2021/day/8)
import collections as cl
# ## Part 1
example_data = [
"be cfbegad cbdgef fgaecd cgeb fdcge agebfd fecdb fabcd edb | fdgacbe cefdb cefbgd gcbe",
"edbfga begcd cbg gc gcadebf fbgde acbgfd abcde gfcbed gfec | fcgedb cgb dgebacf gc",
"fgaebd cg bdaec gdafb agbcfd gdcbef bgcad gfac gcb cdgabef | cg cg fdcagb cbg",
"fbegcd cbd adcefb dageb afcb bc aefdc ecdab fgdeca fcdbega | efabcd cedba gadfec cb",
"aecbfdg fbg gf bafeg dbefa fcge gcbea fcaegb dgceab fcbdga | gecf egdcabf bgf bfgea",
"fgeab ca afcebg bdacfeg cfaedg gcfdb baec bfadeg bafgc acf | gebdcfa ecba ca fadegcb",
"dbcfg fgd bdegcaf fgec aegbdf ecdfab fbedc dacgb gdcebf gf | cefg dcbef fcge gbcadfe",
"bdfegc cbegaf gecbf dfcage bdacg ed bedf ced adcbefg gebcd | ed bcgafe cdgba cbgef",
"egadfb cdbfeg cegd fecab cgb gbdefca cg fgcdab egfdb bfceg | gbdfcae bgc cg cgb",
"gcafb gcf dcaebfg ecagb gf abcdeg gaef cafbge fdbac fegbdc | fgae cfgab fg bagce",
]
# +
segment_to_digit = {
"abcefg": 0,
"cf": 1,
"acdeg": 2,
"acdfg": 3,
"bcdf": 4,
"abdfg": 5,
"abdefg": 6,
"acf": 7,
"abcdefg": 8,
"abcdfg": 9,
}
pattern_lengths = {segment_to_digit[pattern]: len(pattern) for pattern, n in segment_to_digit.items()}
digits_by_length = cl.defaultdict(list)
for digit, length in pattern_lengths.items():
digits_by_length[length].append(digit)
unique_lengths = [digits[0] for length, digits in digits_by_length.items() if len(digits) == 1]
print(f"Unique lengths: {', '.join([str(d) for d in sorted(unique_lengths)])}")
# -
def parse_line(line):
patterns, digits = map(str.strip, line.split("|"))
return {"patterns": patterns.split(" "), "digits": digits.split(" ") }
def part1(input_data):
cnt = cl.Counter()
for line in input_data:
cnt.update(map(len, parse_line(line)["digits"]))
return sum([cnt[pattern_lengths[n]] for n in unique_lengths])
print(f"Check part 1: {part1(example_data) == 26}")
with open(r"..\data\Day 08 input.txt", "r") as fh_in:
input_data = fh_in.readlines()
print(f"Input check: {len(input_data) == 200}")
print(f"Answer part 1: {part1(input_data)}")
# ## Part 2
def decode(line, segment_to_digit=segment_to_digit):
parsed_line = parse_line(line)
# create list of patterns sorted by length
patterns_by_len = cl.defaultdict(list)
for pattern in parsed_line["patterns"]:
patterns_by_len[len(pattern)].append(set(pattern))
# combine patterns to extract segment information
segments = {}
segments["a"] = patterns_by_len[3][0] - patterns_by_len[2][0]
segments_bd = patterns_by_len[4][0] - patterns_by_len[3][0]
segments_eg = patterns_by_len[7][0] - patterns_by_len[4][0] - segments["a"]
segments_afg = sorted([five - segments_bd for five in patterns_by_len[5]], key=len)[0]
segments["f"] = segments_afg - segments["a"] - segments_eg
segments["g"] = segments_afg - segments["a"] - segments["f"]
segments["e"] = segments_eg - segments["g"]
segments["c"] = sorted([five - segments_bd - segments_afg for five in patterns_by_len[5]], key=len)[1]
segments["b"] = sorted(
[six - segments_eg - segments["f"] - segments["a"] - segments["c"] for six in patterns_by_len[6]],
key=len
)[0]
segments["d"] = segments_bd - segments["b"]
# decode by remapping and lookup
mapping = str.maketrans({list(segments[to_digit])[0]: to_digit for to_digit in segments})
decoded = [segment_to_digit["".join(sorted(digit.translate(mapping)))] for digit in parsed_line["digits"]]
return int("".join(map(str, decoded)))
example_line = "acedgfb cdfbe gcdfa fbcad dab cefabd cdfgeb eafb cagedb ab | cdfeb fcadb cdfeb cdbaf"
print(f"Check part 2 (line): {decode(example_line) == 5353}")
# +
example_answers = {
"be cfbegad cbdgef fgaecd cgeb fdcge agebfd fecdb fabcd edb | fdgacbe cefdb cefbgd gcbe": 8394,
"edbfga begcd cbg gc gcadebf fbgde acbgfd abcde gfcbed gfec | fcgedb cgb dgebacf gc": 9781,
"fgaebd cg bdaec gdafb agbcfd gdcbef bgcad gfac gcb cdgabef | cg cg fdcagb cbg": 1197,
"fbegcd cbd adcefb dageb afcb bc aefdc ecdab fgdeca fcdbega | efabcd cedba gadfec cb": 9361,
"aecbfdg fbg gf bafeg dbefa fcge gcbea fcaegb dgceab fcbdga | gecf egdcabf bgf bfgea": 4873,
"fgeab ca afcebg bdacfeg cfaedg gcfdb baec bfadeg bafgc acf | gebdcfa ecba ca fadegcb": 8418,
"dbcfg fgd bdegcaf fgec aegbdf ecdfab fbedc dacgb gdcebf gf | cefg dcbef fcge gbcadfe": 4548,
"bdfegc cbegaf gecbf dfcage bdacg ed bedf ced adcbefg gebcd | ed bcgafe cdgba cbgef": 1625,
"egadfb cdbfeg cegd fecab cgb gbdefca cg fgcdab egfdb bfceg | gbdfcae bgc cg cgb": 8717,
"gcafb gcf dcaebfg ecagb gf abcdeg gaef cafbge fdbac fegbdc | fgae cfgab fg bagce": 4315,
}
for line, answer in example_answers.items():
print(f"{line}: {decode(line) == answer}")
# -
print(f"Answer part 2: {sum([decode(line) for line in input_data])}")
| src/Day 08 Seven Segment Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="oTTHFcXOgADc"
# ! pip install visualkeras
# + id="IdDaQbrm2HWr"
import os
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm
import tensorflow as tf
from keras.layers import Conv2D, UpSampling2D, Dense, MaxPooling2D, BatchNormalization, Dropout, BatchNormalization, Conv2DTranspose
from keras.models import Sequential
import random
import visualkeras
# + id="yxajSPEo2Zm6"
dataset_path = "drive/MyDrive/tiny-imagenet-200/"
models_path = 'drive/MyDrive/Università/Magistrale/VCS/models/'
# + id="a5jmzluS21u5"
SIZE = 64
# + [markdown] id="8U2-s1l4VPtN"
# ## Dataset loading
# + id="vldArmIt23eN"
X_train = np.load(dataset_path+'x_train.npy')
Y_train = np.load(dataset_path+'y_train.npy')
X_valid = np.load(dataset_path+'x_valid.npy')
Y_valid = np.load(dataset_path+'y_valid.npy')
X_test = np.load(dataset_path+'x_test.npy')
Y_test = np.load(dataset_path+'y_test.npy')
# + id="Si5XVI0pwn4h"
X_train = tf.image.convert_image_dtype(X_train, tf.float32)
Y_train = tf.image.convert_image_dtype(Y_train, tf.float32)
X_valid = tf.image.convert_image_dtype(X_valid, tf.float32)
Y_valid = tf.image.convert_image_dtype(Y_valid, tf.float32)
X_test = tf.image.convert_image_dtype(X_test, tf.float32)
Y_test = tf.image.convert_image_dtype(Y_test, tf.float32)
# + id="jxcKIbh725M7"
print(f"{len(X_train)} training examples")
print(f"{len(X_valid)} validation examples")
print(f"{len(X_test)} test examples")
# + [markdown] id="Se7WZpabiGDq"
# ## Sample of the dataset
# + id="yxE3J25Pbtau"
from mpl_toolkits.axes_grid1 import ImageGrid
from skimage.color import lab2rgb
samples = []
for i in range(12):
samples.append(round(random.random()*len(X_train)))
images = []
for i in samples:
img = np.zeros((SIZE, SIZE, 3))
img[:,:,0] = X_train[i][:,:,0]
img[:,:,1:] = Y_train[i]*128
images.append(lab2rgb(img))
fig = plt.figure(figsize=(8., 8.))
grid = ImageGrid(fig, 111,
nrows_ncols=(4, 3),
axes_pad=0.1,
)
for ax, im in zip(grid, images):
ax.imshow(im)
ax.set_yticklabels([])
ax.set_xticklabels([])
plt.show()
# + [markdown] id="zXmk-dvm7CS3"
# ## CNN
#
# Baseline model to test the correctness of the learning framework.
# + id="q7UAz3lL3wfj"
model = Sequential(name=("CNN"))
model.add(Conv2D(32, (3, 3), activation='relu', padding='same', input_shape=(SIZE, SIZE, 1)))
model.add(BatchNormalization())
model.add(Conv2D(64, (3, 3), activation='relu', padding='same'))
model.add(BatchNormalization())
model.add(Conv2D(128, (3, 3), activation='relu', padding='same'))
model.add(BatchNormalization())
model.add(Conv2D(2, (1, 1), activation='tanh', padding='valid'))
# + [markdown] id="zJv5WlDP8Op5"
# ## Polychromify
# Main model: deep convolutional autoencoder.
# + id="BKJDPwcKsByb"
model = Sequential(name=("Polychromify"))
model.add(Conv2D(64, (3, 3), activation='relu', padding='same', strides=2, input_shape=(SIZE, SIZE, 1)))
model.add(Conv2D(128, (3, 3), activation='relu', strides=2, padding='same'))
model.add(Conv2D(256, (3, 3), activation='relu', strides=2, padding='same'))
model.add(BatchNormalization())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Conv2D(256, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(128, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(64, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
# Output layer
model.add(Conv2D(2, (1,1), activation='tanh', padding='valid'))
# + [markdown] id="dMtRoxQ9Kxab"
# ## Polychromify (overfit)
# + id="8dyjLSTrBJ5S"
# I want to test a more aggressive colorization despite having lower generalization metrics values (metrics vs qualitative results problem)
# In this model I simply removed the dropout and trained for 50 epochs, in this way we achieve a complete overfitting.
model = Sequential(name=("Polychromify-overfit"))
model.add(Conv2D(64, (3, 3), activation='relu', padding='same', strides=2, input_shape=(SIZE, SIZE, 1)))
model.add(Conv2D(128, (3, 3), activation='relu', strides=2, padding='same'))
model.add(Conv2D(256, (3, 3), activation='relu', strides=2, padding='same'))
model.add(BatchNormalization())
model.add(Dense(128, activation='relu'))
# model.add(Dropout(0.5))
model.add(Dense(64, activation='relu'))
# model.add(Dropout(0.5))
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Conv2D(256, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(128, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
model.add(Conv2D(64, (3,3), activation='relu', padding='same'))
model.add(UpSampling2D((2, 2)))
# Output layer
model.add(Conv2D(2, (1,1), activation='tanh', padding='valid'))
# + [markdown] id="MpZqg6s87LMy"
# ## Compile & train
# + id="5YoipxRal4VX"
def PSNR(y_true,y_pred):
return tf.image.psnr(y_true, y_pred, max_val=1.0)
# + id="6Bawtzbf30gx"
model.compile(optimizer="adam", loss='mse', metrics=[PSNR])
model.summary()
visualkeras.layered_view(model)
# + id="DeytlRRc31wZ"
percentage = 1
training_size = round(len(X_train)*percentage)
print(f"Using {percentage*100}% of the available training data:\ntrain: {training_size}/{len(X_train)}")
# + id="kbMokko534LR"
history = model.fit(
X_train[:training_size], Y_train[:training_size],
validation_data=(X_valid, Y_valid),
epochs=25, # Polychromify = 25 | Polychromify-overfit = 50
batch_size=64)
# + [markdown] id="Tj1GaLqV7Q6e"
# ## Plots, validation & model save
# + id="sgu1fchc350R"
def plot_history(history,model_name):
fig, axes = plt.subplots(1, 2, figsize=(15, 6))
ax = axes.ravel()
# Loss
ax[0].plot(history['loss'])
ax[0].plot(history['val_loss'])
ax[0].set_title('model loss')
ax[0].set_ylabel('loss')
ax[0].set_xlabel('epochs')
ax[0].legend(['train', 'validation'], loc='upper right')
# PSNR
ax[1].plot(history['PSNR'])
ax[1].plot(history['val_PSNR'])
ax[1].set_title('model PSNR')
ax[1].set_ylabel('PSNR')
ax[1].set_xlabel('epochs')
ax[1].legend(['train', 'validation'], loc='upper left')
fig.tight_layout()
fig.suptitle(model_name,fontsize=16)
plt.show()
# + id="GFUUi8mtABsL"
plot_history(history.history,model.name)
# + id="c8DEvm085Nto"
print("validation")
model.evaluate(X_valid, Y_valid, batch_size=64, verbose=1)
print("test")
model.evaluate(X_test, Y_test, batch_size=64, verbose=1)
# + id="FvGBQS3HNSXR"
model.save(models_path+model.name)
# + [markdown] id="wWa2CKqMzpBG"
# ## Load model
# + id="azAqhCYCOCZY"
model_name = "Polychromify" # change here to load another one
model_loaded = tf.keras.models.load_model(models_path+model_name, custom_objects={'PSNR':PSNR})
# + id="4F9HUG2tjt43"
model_loaded.summary()
visualkeras.layered_view(model_loaded)
# + id="eHspHFTLjLeg"
print("validation")
model_loaded.evaluate(X_valid, Y_valid, batch_size=64, verbose=1)
print("test")
model_loaded.evaluate(X_test, Y_test, batch_size=64, verbose=1)
# + [markdown] id="eEU_Ty2f7V3C"
# ## Visualization
# + id="jjaQ07mh5PVb"
from skimage.color import rgb2lab, lab2rgb
from skimage.transform import resize
from skimage.metrics import structural_similarity as ssim
from skimage.metrics import peak_signal_noise_ratio as psnr
from keras.preprocessing.image import ImageDataGenerator, img_to_array, load_img, array_to_img
# + id="Svs7klyZ5Rmw"
def plot_comparison(img_resized,img_recolored, figsize=(7,3),cast=False):
fig, axes = plt.subplots(1, 2, figsize=figsize)
ax = axes.ravel()
ax[0].imshow(img_resized)
ax[0].set_title("Resized")
ax[1].imshow(img_recolored)
ax[1].set_title("Recolored")
fig.tight_layout()
plt.show()
# + id="VzmvJYmy5TNp"
X_data = X_test
Y_data = Y_test
for i, x in enumerate(X_data[:10]):
img_color = np.array([x], dtype=float)
output = model.predict(img_color)
result = np.zeros((SIZE, SIZE, 3))
result[:,:,0] = x[:,:,0]
result[:,:,1:] = output[0]*128
recolored = lab2rgb(result)
original = np.zeros((SIZE, SIZE, 3))
original[:,:,0] = x[:,:,0]
original[:,:,1:] = Y_data[i]*128
original = lab2rgb(original)
ssim = tf.image.ssim(original, recolored, max_val=1.0, filter_size=11,
filter_sigma=1.5, k1=0.01, k2=0.03)
print(f"SSIM {ssim}")
print(f"PSNR {PSNR(original,recolored)}")
plot_comparison(original, recolored)
# + [markdown] id="QHJli7-xzbCy"
# ## Metric evaluation
# + id="-gLYzmcN2tSK"
import warnings
warnings.filterwarnings("ignore")
# + id="KZMaUcVLzAie"
SSIM_tot = 0
PSNR_tot = 0
size = len(X_test)
for i, x in enumerate(tqdm(X_test[:size])):
img_color = np.array([x], dtype=float)
output = model.predict(img_color)
result = np.zeros((SIZE, SIZE, 3))
result[:,:,0] = x[:,:,0]
result[:,:,1:] = output[0]*128
recolored = lab2rgb(result)
original = np.zeros((SIZE, SIZE, 3))
original[:,:,0] = x[:,:,0]
original[:,:,1:] = Y_test[i]*128
original = lab2rgb(original)
ssim = tf.image.ssim(original, recolored, max_val=1.0, filter_size=11,
filter_sigma=1.5, k1=0.01, k2=0.03)
SSIM_tot += ssim
PSNR_tot += PSNR(original,recolored)
print(f"SSIM {SSIM_tot/size} - PSNR {PSNR_tot/size}")
# + [markdown] id="F_Ip8JmwB6wH"
# ## Separate training test
# + id="hqwYhbWTAn8K"
model_A = Sequential(name=("Polychromify_A"))
model_A.add(Conv2D(64, (3, 3), activation='relu', strides=2, padding='same', input_shape=(SIZE, SIZE, 1)))
model_A.add(Conv2D(128, (3, 3), activation='relu', strides=2, padding='same'))
model_A.add(Conv2D(256, (3, 3), activation='relu', strides=2, padding='same'))
model_A.add(BatchNormalization())
model_A.add(Dense(128, activation='relu'))
model_A.add(Dropout(0.5))
model_A.add(Dense(64, activation='relu'))
model_A.add(Dropout(0.5))
model_A.add(Dense(128, activation='relu'))
model_A.add(BatchNormalization())
model_A.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model_A.add(UpSampling2D((2, 2)))
model_A.add(Conv2D(128, (3,3), activation='relu', padding='same'))
model_A.add(UpSampling2D((2, 2)))
model_A.add(Conv2D(64, (3,3), activation='relu', padding='same'))
model_A.add(UpSampling2D((2, 2)))
# Output layer
model_A.add(Conv2D(1, (1, 1), activation='tanh', padding='valid'))
# + id="ylu-M4qxEIEK"
model_B = Sequential(name=("Polychromify_B"))
model_B.add(Conv2D(64, (3, 3), activation='relu', strides=2, padding='same', input_shape=(SIZE, SIZE, 1)))
model_B.add(Conv2D(128, (3, 3), activation='relu', strides=2, padding='same'))
model_B.add(Conv2D(256, (3, 3), activation='relu', strides=2, padding='same'))
model_B.add(BatchNormalization())
model_B.add(Dense(128, activation='relu'))
model_B.add(Dropout(0.5))
model_B.add(Dense(64, activation='relu'))
model_B.add(Dropout(0.5))
model_B.add(Dense(128, activation='relu'))
model_B.add(BatchNormalization())
model_B.add(Conv2D(256, (3, 3), activation='relu', padding='same'))
model_B.add(UpSampling2D((2, 2)))
model_B.add(Conv2D(128, (3,3), activation='relu', padding='same'))
model_B.add(UpSampling2D((2, 2)))
model_B.add(Conv2D(64, (3,3), activation='relu', padding='same'))
model_B.add(UpSampling2D((2, 2)))
# Output layer
model_B.add(Conv2D(1, (1, 1), activation='tanh', padding='valid'))
# + id="n_MrFRGFEWzO"
model_A.compile(optimizer="adam", loss='mse', metrics=[PSNR])
model_B.compile(optimizer="adam", loss='mse', metrics=[PSNR])
# + id="0Ztpfs45tbqD"
model_A.summary()
visualkeras.layered_view(model_A)
# + id="DrsCyV9FtaBW"
model_B.summary()
visualkeras.layered_view(model_B)
# + id="hiax43owAy0l"
history_A = model_A.fit(
X_train[:training_size], Y_train[:,:,:,0][:training_size],
validation_data=(X_valid, Y_valid[:,:,:,0]),
epochs=30,
batch_size=64)
# + id="e6Yg7gV_CZTI"
history_B = model_B.fit(
X_train[:training_size], Y_train[:,:,:,1][:training_size],
validation_data=(X_valid, Y_valid[:,:,:,1]),
epochs=30,
batch_size=64)
# + id="rXKDUGXCiP7B"
plot_history(history_A.history,model_A.name)
plot_history(history_B.history,model_B.name)
# + id="0l79WS4vh3Hf"
model_A.save(models_path+model_A.name)
model_B.save(models_path+model_B.name)
# + [markdown] id="VM0pvk5L1E0k"
# ### Visualization
# + id="nCS1xwfvDxLM"
for i, x in enumerate(X_test[:10]):
img_color = []
img_color.append(x)
img_color = np.array(img_color, dtype=float)
output_A = model_A.predict(img_color)
output_A = output_A*128
output_B = model_B.predict(img_color)
output_B = output_B*128
result = np.zeros((SIZE, SIZE, 3))
result[:,:,0] = img_color[0][:,:,0]
result[:,:,1] = output_A[0][:,:,0]
result[:,:,2] = output_B[0][:,:,0]
recolored = lab2rgb(result)
original = np.zeros((SIZE, SIZE, 3))
original[:,:,0] = x[:,:,0]
original[:,:,1:] = Y_test[i]*128
original = lab2rgb(original)
ssim = tf.image.ssim(original, recolored, max_val=1.0, filter_size=11,
filter_sigma=1.5, k1=0.01, k2=0.03)
print(f"SSIM {ssim}")
print(f"PSNR {PSNR(original,recolored)}")
plot_comparison(original, recolored)
# + [markdown] id="SBp-0Hh71HCD"
# ### Metrics
# + id="V_kQbhq5wuSz"
SSIM_tot = 0
PSNR_tot = 0
size = len(X_test)
for i, x in enumerate(tqdm(X_test[:size])):
img_color = []
img_color.append(x)
img_color = np.array(img_color, dtype=float)
output_A = model_A.predict(img_color)
output_A = output_A*128
output_B = model_B.predict(img_color)
output_B = output_B*128
result = np.zeros((SIZE, SIZE, 3))
result[:,:,0] = img_color[0][:,:,0]
result[:,:,1] = output_A[0][:,:,0]
result[:,:,2] = output_B[0][:,:,0]
recolored = lab2rgb(result)
original = np.zeros((SIZE, SIZE, 3))
original[:,:,0] = x[:,:,0]
original[:,:,1:] = Y_test[i]*128
original = lab2rgb(original)
ssim = tf.image.ssim(original, recolored, max_val=1.0, filter_size=11,
filter_sigma=1.5, k1=0.01, k2=0.03)
SSIM_tot += ssim
PSNR_tot += PSNR(original,recolored)
print(f"SSIM {SSIM_tot/size} - PSNR {PSNR_tot/size}")
| notebooks/models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/allenpatrickargente/OOP-1-1/blob/main/OOP_Comcepts_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZRBlJwAZMK7x"
# Classes with Multiple Objects
# + id="8IKHkXTOK1nw"
class Birds:
def __init__(self,birds_name):
self.birds_name=birds_name
def flying_birds(self):
print(f"{self.birds_name} flies above the sky")
def non_flying_birds(self):
print(f"{self.birds_name} is the national bird of Australia")
vulture=Birds("Griffon Vulture")
crane=Birds("Common Crane")
emu=Birds("Emu")
vulture.flying_birds()
crane.flying_birds()
emu.non_flying_birds()
# + [markdown] id="TNPYfZHPPTWB"
# Encapsulation(mangling with double underscore
# + colab={"base_uri": "https://localhost:8080/"} id="jTxgxtlOPL0F" outputId="54f5a7c0-5fcb-4a75-bb0f-a36ca2d018c7"
class foo:
def __init__(self,a,b):
self.a=a
self.b=b
def add(self):
return self.a+self.b
object_foo=foo(3,4)
object_foo.add()
object_foo__a= 6
object_foo__b= 7
object_foo.add()
# + [markdown] id="IDhh_6gCRfBB"
#
# + colab={"base_uri": "https://localhost:8080/"} id="H6cUTMgbRfa0" outputId="8186bc09-85ce-4fb6-d161-ec0b3eae0047"
class Counter:
def __init__(self):
self.__current = 0
def increment(self):
self.__current += 1
def value(self):
return self.__current
def reset(self):
self.__current = 0
number=Counter()
number__current=1
number.increment()
number.increment()
number.increment()
print(number.value())
# + [markdown] id="AcxdGwqCU9qE"
# inheritance
# + colab={"base_uri": "https://localhost:8080/"} id="Sw-B04bzVBz3" outputId="16d161e5-8859-4967-c15e-077f0e399698"
class Person:
def __init__(self,firstname,surname):
self.firstname=firstname
self.surname=surname
def fullname(self):
print(self.firstname,self.surname)
person = Person("AllenPatrick","Argente")
person.fullname()
class Teacher(Person):
pass
person2 = Teacher("Maam","Maria")
person2.fullname()
class Student(Person):
pass
person3 = Student("AllenPAtrick","Argente")
person3.fullname()
# + [markdown] id="yYZMaeIEXjAL"
# Polymorphism
# + colab={"base_uri": "https://localhost:8080/"} id="DL10rF1GXsTZ" outputId="c4e3b393-8f29-4c10-c233-ebc6d6e6b977"
class RegularPolygon:
def __init__(self,side):
self.side = side
class Square(RegularPolygon):
def area(self):
return self.side * self.side
class EquilateralTriangle(RegularPolygon):
def area(self):
return self.side * self.side * 0.433
obj1 = Square(4)
print(obj1.area())
obj2 = EquilateralTriangle(3)
print(obj2.area())
# + [markdown] id="Ykrg88c9Y3pk"
# Application 1
# + [markdown] id="tlS_EOcHZHYL"
# 1.Create a Python program that displays the name of the three students(Student 1,Student 2, and Student 3) and their term grades
#
# 2.Create a Class name Person and attributes -std1,std2,std2,pre,mid,fin
#
# 3.Compute the average of each term grade using Grade() method
#
# 4.infomration about students grades must be hidden from others
| OOP_Comcepts_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp sparse.sparsifier
# -
# # Sparsifier
#
# > Make your neural network sparse
#hide
from nbdev.showdoc import *
from fastai.vision.all import *
#export
import torch
import torch.nn as nn
from fastcore.basics import store_attr
from fasterai.sparse.criteria import *
#hide
from torchvision.utils import make_grid
import matplotlib.pyplot as plt
# A sparse vector, as opposed to a dense one, is a vector which contains a lot of zeroes. When we speak about making a neural network sparse, we thus mean that the network's weight are mostly zeroes.
#
# With fasterai, you can do that thanks to the `Sparsifier` class.
# Let's start by creating a model
model = resnet18()
# As you probably know, weights in a convolutional neural network have 4 dimensions ($ c_{out} \times c_{in} \times k_h \times k_w$)
model.conv1.weight.ndim
#hide
def plot_kernels(layer, save=None):
kernels = layer.weight.detach().clone()
kernels = kernels - kernels.min()
kernels = kernels/kernels.max()
plt.figure(figsize=(10,10))
img = make_grid(kernels, nrow=8, padding=1, pad_value=1)
plt.axis('off')
plt.imshow(img.detach().permute(1,2,0).cpu())
if save: plt.savefig(f'{save}.pdf')
# In the case of ResNet18, the dimension of the first layer weights is $64 \times 3 \times 7 \times 7$. We thus can plot each of the $64$ filter as a $7 \times 7$ color image (because they contains $3$ channels).
plot_kernels(model.conv1)
#export
class Sparsifier():
def __init__(self, model, granularity, method, criteria):
store_attr()
self._save_weights() # Save the original weights
def prune_layer(self, m, sparsity):
weight = self.criteria(m, self.granularity)
mask = self._compute_mask(self.model, weight, sparsity)
m.register_buffer("_mask", mask) # Put the mask into a buffer
self._apply(m)
def prune_model(self, sparsity):
for m in self.model.modules():
if isinstance(m, nn.Conv2d): self.prune_layer(m, sparsity)
def _apply(self, m):
mask = getattr(m, "_mask", None)
if mask is not None: m.weight.data.mul_(mask)
if self.granularity == 'filter' and m.bias is not None:
if mask is not None: m.bias.data.mul_(mask.squeeze()) # We want to prune the bias when pruning filters
def _mask_grad(self):
for m in self.model.modules():
if isinstance(m, nn.Conv2d) and hasattr(m, '_mask'):
mask = getattr(m, "_mask")
if m.weight.grad is not None: m.weight.grad.mul_(mask)
if self.granularity == 'filter' and m.bias is not None:
if m.bias.grad is not None: m.bias.grad.mul_(mask.squeeze())
def _reset_weights(self):
for m in self.model.modules():
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
init_weights = getattr(m, "_init_weights", m.weight)
init_biases = getattr(m, "_init_biases", m.bias)
with torch.no_grad():
if m.weight is not None: m.weight.copy_(init_weights)
if m.bias is not None: m.bias.copy_(init_biases)
self._apply(m)
if isinstance(m, nn.BatchNorm2d) or isinstance(m, nn.BatchNorm1d): m.reset_parameters()
def _save_weights(self):
for m in self.model.modules():
if isinstance(m, nn.Conv2d) or isinstance(m, nn.Linear):
m.register_buffer("_init_weights", m.weight.clone())
if m.bias is not None: m.register_buffer("_init_biases", m.bias.clone())
def _clean_buffers(self):
for m in self.model.modules():
if hasattr(m, '_mask'): del m._buffers["_mask"]
if hasattr(m, '_init_weights'): del m._buffers["_init_weights"]
if hasattr(m, '_init_biases'): del m._buffers["_init_biases"]
def _compute_mask(self, model, weight, sparsity):
if self.method == 'global':
global_weight = torch.cat([self.criteria(m, self.granularity).view(-1) for m in model.modules() if isinstance(m, nn.Conv2d)])
threshold = torch.quantile(global_weight, sparsity/100) # Compute the threshold globally
elif self.method == 'local':
threshold = torch.quantile(weight.view(-1), sparsity/100) # Compute the threshold locally
else: raise NameError('Invalid Method')
if threshold > weight.max(): threshold = weight.max() # Make sure we don't remove every weight of a given layer
mask = weight.ge(threshold).to(dtype=weight.dtype)
return mask
# The `Sparsifier` class allows us to remove some (part of) the filters, that are considered to be less useful than others. This can be done by first creating an instance of the class, specifying:
#
# - The `granularity`, i.e. the part of filters that you want to remove. Typically, we usually remove weights, vectors, kernels or even complete filters.
# - The `method`, i.e. if you want to consider each layer independently (`local`), or compare the parameters to remove across the whole network (`global`).
# - The `criteria`, i.e. the way to assess the usefulness of a parameter. Common methods compare parameters using their magnitude, the lowest magnitude ones considered to be less useful.
# Once this is specified, just use the `Sparsifier.prune_model`, indicating the percentage of sparsity to you want to apply. One can also specify a single layer to prune by using the `Sparsifier.prune_layer` method.
show_doc(Sparsifier.prune_model)
# There are several ways in which we can make that first layer sparse. You will find the most important below:
# ## Pruning individual weights
# As a reminder, the first convolutional layer of ResNet18 could be represented as a $64 \times 3 \times 7 \times 7$ tensor, meaning that the whole layer posess $9408$ weights.
#
# When removing individual weights, we do not impose any constraint on where in the first layer the weight removed should come from. We only compare them all together and remove the ones that are the least important, according to our `criteria`. We thus remove $0d$ blocks from our $4d$ weight tensor.
# Here is for example how you would proceed to remove $80 \%$ of the weights of the network.
model = resnet18()
pruner = Sparsifier(model, 'weight', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
# And here are what the first layer filters would look like.
plot_kernels(model.conv1)
# ## Pruning Vectors
# When removing vectors, we don't compare $9408$ weights anymore as we now impose to remove complete vectors of weights from the first layer. We now remove $1d$ blocks from our layer and can chose to remove `column` vectors from the filters, but also `row` of `channel` vectors. In those cases, we thus compare respectively $64 \times 3 \times 7$ column vectors, or $64 \times 7 \times 3$ row vectors, or $64 \times 7 \times 7$ channel vectors, and remove the least important according to our criteria.
# The consequence is that, we now start to see some structure emerging in our remaining weights.
# > Note: If you look closely, you will see some red/green/blue vectors remaining. This means that we removed the 2 other vectors corresponding to the same spatial location but located at other color channel.
model = resnet18()
pruner = Sparsifier(model, 'column', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
model = resnet18()
pruner = Sparsifier(model, 'row', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
model = resnet18()
pruner = Sparsifier(model, 'channel', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
# A particular granularity to operate with is what we called `shared_weight`, which in this case consists of comparing $3 \times 7 \times 7$ blocks together. This results in an unstructured pruning scheme on a single filter, but this scheme is shared with all the other filters !
model = resnet18()
pruner = Sparsifier(model, 'shared_weight', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
# ## Pruning Kernels
# Having a network sparse kernel-wise means that we removed $2d$ blocks from our filter (generally from a spatial point of view). In our case, it means comparing each of the $64 \times 3$ kernels and removing the least important ones.
model = resnet18()
pruner = Sparsifier(model, 'kernel', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
# ## Pruning Filters
# Finally, we can choose to remove entire filters. This means that now, we compare the $64$ filters together, thus removing $3d$ blocks.
model = resnet18()
pruner = Sparsifier(model, 'filter', 'local', large_final)
pruner.prune_layer(model.conv1, 80)
plot_kernels(model.conv1)
| nbs/01a_sparsifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''networkx'': conda)'
# name: python3
# ---
# > **`(!)`** 2021 Written by <NAME> `<<EMAIL>>`
# > This is free and unencumbered software released into the public domain.
# +
import math
from itertools import combinations
import networkx as nx
from matplotlib import pyplot as plt
import randy
# +
def sigmoid(x):
return 1 / (1 + math.exp(-x))
def plot_goose(g):
"""Draw the graph with nodes in the common core sligthly bigger"""
node_size = [300 if core else 100 for _, core in g.nodes(data='core', default=False)]
node_color = ['red' if core else 'black' for _, core in g.nodes(data='core', default=False)]
node_color = [n for _, n in g.nodes(data='class')]
edge_color = ['red' if core else 'black' for _, _, core in g.edges(data='core', default=False)]
nx.draw(g, node_size=node_size, node_color=node_color, cmap=plt.cm.tab20, edge_color=edge_color)
plt.show()
def dump_goose(g):
"""Dump graph shuffling node id's"""
pomegranate = list(g.nodes())
randy.shuffle(pomegranate)
dump = ''
dump += f'{g.number_of_nodes()} {g.number_of_edges()}\n'
dump += ' '.join([str(g.nodes[n]['class']) for n in pomegranate]) + '\n'
edges = list()
for n1, n2 in g.edges():
edges.append((pomegranate[n1], pomegranate[n2]))
dump += '\n'.join(f'{s} {d}' for s, d in sorted(edges))
return dump
# -
def foie_gras(family_size=5, core_nodes=10, density=.5, family_divergence=.1, num_classes=1, **options):
"""Create a family of graph with a common core"""
core = nx.random_graphs.fast_gnp_random_graph(core_nodes, density)
for n in core.nodes():
core.nodes[n]['class'] = randy.choice(range(num_classes))
core.nodes[n]['core'] = True
for s, d in core.edges():
core.edges[s, d]['core'] = True
geese = list()
for f in range(family_size):
new_goose = nx.Graph(core)
# add random edges
while randy.boolean(p_true=sigmoid(family_divergence)):
s, d = randy.choice(new_goose.nodes()), randy.choice(new_goose.nodes())
new_goose.add_edge(s, d)
avg_extra_nodes = core_nodes * family_divergence
# add random nodes
for new_node in range(core_nodes, int(randy.scale_random(core_nodes, core_nodes+2*avg_extra_nodes,
core_nodes+avg_extra_nodes, 1))):
new_goose.add_node(new_node)
new_goose.nodes[new_node]['class'] = randy.choice(range(num_classes))
for d in new_goose.nodes():
if randy.boolean(p_true=density):
new_goose.add_edge(new_node, d)
geese.append(new_goose)
return geese
family = foie_gras(family_size=4, core_nodes=6, density=.5, family_divergence=1, num_classes=4)
for n, graph in enumerate(family):
print(f"# GRAPH {n}")
plot_goose(graph)
with open(f'out/family0_g{n}.txt', 'w') as f:
f.write(dump_goose(graph))
| foie-gras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import libraries
# +
# Ignore future warnings
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
from matplotlib.axes._axes import _log as matplotlib_axes_logger
matplotlib_axes_logger.setLevel('ERROR')
# +
# Pandas
import pandas as pd
# Numpy
import numpy as np
# Plotting
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from matplotlib import cm
# Preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
# +
# Learning algorithms
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
# Pipeline
from sklearn.pipeline import Pipeline
# Model Selection
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import learning_curve
from sklearn.model_selection import validation_curve
# Metrics
from sklearn.metrics import roc_curve
from sklearn.metrics import auc
from sklearn.metrics import roc_auc_score
from sklearn.metrics import make_scorer
from sklearn.metrics import silhouette_samples
# Dimensionality reduction
from sklearn.decomposition import PCA
from sklearn.decomposition import KernelPCA
from sklearn.decomposition import FastICA
from sklearn.random_projection import GaussianRandomProjection # Randomized Projections
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis # LDA
# Clustering
from sklearn.cluster import KMeans
from sklearn.mixture import GaussianMixture # Expectation Maximization
# -
# ### Load Data
# +
magic_df = pd.read_csv('./data/magic04.data', header=None, error_bad_lines=False)
# Full Data
X = magic_df.iloc[:, :-1].values
y = magic_df.iloc[:, -1].values
# Convert labels from ['h', 'g'] to [1, 0]
le = LabelEncoder()
y = le.fit_transform(y)
# Split data into train and test
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.25, random_state=42)
# -
# ### Plotting Functions
def plot_variance(var_exp, cum_var_exp, title=None, save_as=None):
plt.bar(range(1, len(var_exp)+1), var_exp, alpha=0.5, align='center',
label='individual explained variance')
plt.step(range(1, len(var_exp)+1), cum_var_exp, where='mid',
label='cumulative explained variance')
plt.ylabel('Explained Variance Ratio')
plt.xlabel('Principal Components')
plt.legend(loc='best')
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_decision_regions(X, y, classifier, resolution=0.02, title=None, save_as=None):
# Setup marker generator and color map
markers = ('s', 'x', 'o', '^', 'v')
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
cmap = ListedColormap(colors[:len(np.unique(y))])
# Plot the decision surface
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution),
np.arange(x2_min, x2_max, resolution))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.4, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
plt.title(title)
# Plot class samples
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0],
y=X[y == cl, 1],
alpha=0.6,
c=cmap(idx),
edgecolor='black',
marker=markers[idx],
label=cl)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_elbow_distortions(max_clusters, X, title=None, save_as=None):
distortions = []
for i in range(1, max_clusters+1):
km = KMeans(n_clusters=i,
init='k-means++',
n_init=10,
max_iter=300,
random_state=42)
km.fit(X)
distortions.append(km.inertia_)
plt.plot(range(1, max_clusters+1), distortions, marker='o')
plt.xlabel('Number of clusters')
plt.ylabel('Distortion')
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_elbow_bic(max_components, X, title=None, save_as=None):
bic = []
for i in range(1, max_components+1):
gmm = GaussianMixture(n_components=i,
covariance_type='full',
reg_covar=1e-06,
max_iter=100,
n_init=1,
init_params='kmeans',
random_state=42)
gmm.fit(X)
bic.append(gmm.bic(X))
plt.plot(range(1, max_components+1), bic, marker='s')
plt.xlabel('Number of Components')
plt.ylabel('Bayesian Information Criterion')
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_silhouette(km, X, title=None, save_as=None):
y_km = km.predict(X)
cluster_labels = np.unique(y_km)
n_clusters = cluster_labels.shape[0]
silhouette_vals = silhouette_samples(X, y_km, metric='euclidean')
y_ax_lower, y_ax_upper = 0, 0
yticks = []
for i, c in enumerate(cluster_labels):
c_silhouette_vals = silhouette_vals[y_km == c]
c_silhouette_vals.sort()
y_ax_upper += len(c_silhouette_vals)
color = cm.jet(float(i) / n_clusters)
plt.barh(range(y_ax_lower, y_ax_upper), c_silhouette_vals, height=1.0,
edgecolor='none', color=color)
yticks.append((y_ax_lower + y_ax_upper) / 2.)
y_ax_lower += len(c_silhouette_vals)
silhouette_avg = np.mean(silhouette_vals)
plt.axvline(silhouette_avg, color="orange", linestyle="--")
plt.yticks(yticks, cluster_labels + 1)
plt.ylabel('Cluster')
plt.xlabel('Silhouette Coefficient')
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_kmeans(km, X, xlim=None, ylim=None, title=None, save_as=None):
y_km = km.predict(X)
colors = ('lightgreen', 'gray', 'blue', 'orange', 'cyan')
markers = ('s', 'o', 'v', 'p', 'd')
for i in range(0, km.n_clusters):
plt.scatter(X[y_km == i, 0],
X[y_km == i, 1],
s=50, c=colors[i],
marker=markers[i], edgecolor='black',
label='cluster '+str(i+1))
plt.scatter(km.cluster_centers_[:, 0],
km.cluster_centers_[:, 1],
s=250, marker='*',
c='red', edgecolor='black',
label='centroids')
plt.legend(scatterpoints=1)
plt.grid()
plt.xlim(xlim)
plt.ylim(ylim)
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_gmm(gmm, X, index, xlim=None, ylim=None, title=None, save_as=None):
import itertools
from scipy import linalg
import matplotlib as mpl
y_gmm = gmm.predict(X)
color_iter = itertools.cycle(['green', 'blue', 'orange', 'purple'])
means = gmm.means_
covariances = gmm.covariances_
for i, (mean, covar, color) in enumerate(zip(
means, covariances, color_iter)):
v, w = linalg.eigh(covar)
v = 2. * np.sqrt(2.) * np.sqrt(v)
u = w[0] / linalg.norm(w[0])
# as the DP will not use every component it has access to
# unless it needs it, we shouldn't plot the redundant
# components.
if not np.any(y_gmm == i):
continue
plt.scatter(X[y_gmm == i, 0], X[y_gmm == i, 1], color=color)
# Plot an ellipse to show the Gaussian component
angle = np.arctan(u[1] / u[0])
angle = 180. * angle / np.pi # convert to degrees
ell = mpl.patches.Ellipse(mean, v[0], v[1], 180. + angle, color=color)
ell.set_alpha(0.5)
plt.grid()
plt.xlim(xlim)
plt.ylim(ylim)
plt.title(title)
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
def plot_discriminability(eigen_vals, ylim=None, save_as=None):
tot = sum(eigen_vals.real)
discr = [(i / tot) for i in sorted(eigen_vals.real, reverse=True)]
cum_discr = np.cumsum(discr)
plt.bar(range(0, len(eigen_vals)), discr, alpha=0.5, align='center',
label='individual class-discriminatory information ratio')
plt.step(range(0, len(eigen_vals)), cum_discr, where='mid',
label='cumulative class-discriminatory information ratio')
plt.ylabel('Class-discriminatory Information Ratio')
plt.xlabel('Linear Discriminants')
plt.ylim(ylim)
plt.legend(loc='best')
plt.tight_layout()
if save_as != None:
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
train_auc_list = []
test_auc_list = []
# ## Clustering Algorithms
# ### K-Means
# Elbow around k=5
plot_elbow_distortions(max_clusters=20, X=X_train, title='K-means Elbow Plot', save_as='MAGIC/kmeans_elbow_plot')
kmeans = KMeans(n_clusters=5, init='k-means++',
n_init=10, max_iter=300, verbose=0,
random_state=42, n_jobs=-1,
algorithm='auto')
# %%time
kmeans.fit(X_train)
X_train_kmeans = kmeans.predict(X_train).reshape(-1, 1)
X_test_kmeans = kmeans.predict(X_test).reshape(-1, 1)
# %%time
# Kmeans data only
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_kmeans, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_kmeans, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_kmeans, y_test))
y_pred = pipe_mlp.predict(X_train_kmeans)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_kmeans)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
X_train_kmeans_combined = np.concatenate([X_train, X_train_kmeans], axis=1)
X_test_kmeans_combined = np.concatenate([X_test, X_test_kmeans], axis=1)
# %%time
# Kmeans combined with original data
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_kmeans_combined, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_kmeans_combined, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_kmeans_combined, y_test))
y_pred = pipe_mlp.predict(X_train_kmeans_combined)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_kmeans_combined)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
plot_silhouette(km=kmeans, X=X_train, title='K-means Silhouette Plot', save_as='MAGIC/silhouette_plot')
# ### Expectation Maximization (Gaussian Mixture)
plot_elbow_bic(max_components=20, X=X_train, title='EM BIC Plot', save_as='MAGIC/EM_BIC_plot')
gaussian_mix = GaussianMixture(n_components=3, covariance_type='full',
max_iter=100, init_params='kmeans',
random_state=42, verbose=0, verbose_interval=10)
# %%time
gaussian_mix.fit(X_train)
X_train_gmm = gaussian_mix.predict(X_train).reshape(-1, 1)
X_test_gmm = gaussian_mix.predict(X_test).reshape(-1, 1)
# +
# %%time
# EM data only
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_gmm, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_gmm, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_gmm, y_test))
y_pred = pipe_mlp.predict(X_train_gmm)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_gmm)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
X_train_gmm_combined = np.concatenate([X_train, X_train_gmm], axis=1)
X_test_gmm_combined = np.concatenate([X_test, X_test_gmm], axis=1)
# +
# %%time
# EM combined with original data
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_gmm_combined, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_gmm_combined, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_gmm_combined, y_test))
y_pred = pipe_mlp.predict(X_train_gmm_combined)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_gmm_combined)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ## Dimensionality Reduction Algorithms
# ### Principal Component Analysis
pca = PCA(n_components=4, copy=True,
whiten=False, svd_solver='auto',
tol=0.0, iterated_power='auto',
random_state=42)
# %%time
X_train_pca = pca.fit_transform(X_train)
X_test_pca = pca.transform(X_test)
# +
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_pca, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_pca, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_pca, y_test))
y_pred = pipe_mlp.predict(X_train_pca)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_pca)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Independent Component Analysis
ica = FastICA(n_components=8, algorithm='parallel',
whiten=True, fun='logcosh', fun_args=None,
max_iter=200, tol=0.0001, w_init=None,
random_state=42)
# %%time
X_train_ica = ica.fit_transform(X_train)
X_test_ica = ica.transform(X_test)
# +
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_ica, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_ica, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_ica, y_test))
y_pred = pipe_mlp.predict(X_train_ica)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_ica)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Random Projection (Gaussian)
grp = GaussianRandomProjection(n_components=5,
random_state=42)
# %%time
X_train_grp = grp.fit_transform(X_train)
X_test_grp = grp.transform(X_test)
# +
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_grp, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_grp, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_grp, y_test))
y_pred = pipe_mlp.predict(X_train_grp)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_grp)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Linear Discriminant Analysis
lda = LinearDiscriminantAnalysis(n_components=1,
solver='svd',
store_covariance=False,
tol=0.0001)
# %%time
X_train_lda = lda.fit_transform(X_train, y_train)
X_test_lda = lda.transform(X_test)
# +
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train_lda, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_lda, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_lda, y_test))
y_pred = pipe_mlp.predict(X_train_lda)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_lda)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Random Forest Feature Selection
# %%time
feat_labels = magic_df.columns[:-1]
forest = RandomForestClassifier(n_estimators=500,
random_state=42)
forest.fit(X_train, y_train)
importances = forest.feature_importances_
indices = np.argsort(importances)[::-1]
X_train_rf = X_train[:, indices[:5]]
X_test_rf = X_test[:, indices[:5]]
# +
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(120,),
random_state=42))])
pipe_mlp.fit(X_train_rf, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train_rf, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test_rf, y_test))
y_pred = pipe_mlp.predict(X_train_rf)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test_rf)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Original
# %%time
pipe_mlp = Pipeline([('scl', StandardScaler()),
('clf', MLPClassifier(max_iter=1000,
activation='relu',
solver='adam',
hidden_layer_sizes=(160,),
random_state=42))])
pipe_mlp.fit(X_train, y_train)
# +
print('Train Accuracy: %.3f' % pipe_mlp.score(X_train, y_train))
print('Test Accuracy: %.3f' % pipe_mlp.score(X_test, y_test))
y_pred = pipe_mlp.predict(X_train)
print('Train AUC: %.3f' % roc_auc_score(y_train, y_pred))
train_auc_list.append(roc_auc_score(y_train, y_pred))
y_pred = pipe_mlp.predict(X_test)
print('Test AUC: %.3f' % roc_auc_score(y_test, y_pred))
test_auc_list.append(roc_auc_score(y_test, y_pred))
# -
# ### Overall Results
# +
labels_list = ['K-means Only', 'K-means Augmented', 'EM Only', 'EM Augmented', 'PCA', 'ICA', 'GRP', 'LDA', 'RF', 'Original']
training_times_list = [6.89, 39.57, 17.30, 31.00, 14.13, 31.83, 14, 3.25, 37.60, 27.40]
train_auc_list = np.array(train_auc_list).reshape(len(labels_list))
test_auc_list = np.array(test_auc_list).reshape(len(labels_list))
time_zipped = zip(labels_list, training_times_list)
time_sorted = sorted(time_zipped, key=lambda x:x[1], reverse=True)
train_zipped = zip(labels_list, train_auc_list)
train_sorted = sorted(train_zipped, key=lambda x:x[1])
test_zipped = zip(labels_list, test_auc_list)
test_sorted = sorted(test_zipped, key=lambda x:x[1])
# -
y_pos = np.arange(len(labels_list))
plt.barh(y_pos, [times for labels, times in time_sorted], align='center', alpha=0.5)
plt.yticks(y_pos, [labels for labels, times in time_sorted])
plt.xlabel('Training Time (s)')
plt.title('Training Time Comparison')
for i, v in enumerate([times for labels, times in time_sorted]):
plt.text(v-3, i, "%.1f" % v, color='black', va='center')
plt.tight_layout()
save_as = 'MAGIC/NN_time_comparison'
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
y_pos = np.arange(len(labels_list))
plt.barh(y_pos, [auc for labels, auc in train_sorted], align='center', alpha=0.5)
plt.yticks(y_pos, [labels for labels, auc in train_sorted])
plt.xlabel('Train ROC AUC')
plt.title('Train ROC AUC Comparison')
for i, v in enumerate([auc for labels, auc in train_sorted]):
plt.text(v-0.1, i, "%.3f" % v, color='black', va='center')
plt.tight_layout()
save_as = 'MAGIC/NN_train_auc_comparison'
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
y_pos = np.arange(len(labels_list))
plt.barh(y_pos, [auc for labels, auc in test_sorted], align='center', alpha=0.5)
plt.yticks(y_pos, [labels for labels, auc in test_sorted])
plt.xlabel('Test ROC AUC')
plt.title('Test ROC AUC Comparison')
for i, v in enumerate([auc for labels, auc in test_sorted]):
plt.text(v-0.1, i, "%.3f" % v, color='black', va='center')
plt.tight_layout()
save_as = 'MAGIC/NN_test_auc_comparison'
plt.savefig('./figures/' + save_as + '.png', dpi=300)
plt.show()
| unsupervised learning/MAGIC - NN with Clustering and DR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from SimPEG import *
from simpegEM1D import EM1D, EM1DSurveyTD, Utils1D, get_vertical_discretization_time, set_mesh_1d, piecewise_pulse
import numpy as np
# %pylab inline
# +
from simpegEM1D import skytem_HM_2015, skytem_LM_2015
wave_HM = skytem_HM_2015()
wave_LM = skytem_LM_2015()
time_HM = np.logspace(-5, -2, 31)
time_LM = np.logspace(-5, -2, 31)
hz = get_vertical_discretization_time(
np.unique(np.r_[time_HM, time_LM]), facter_tmax=0.5, factor_tmin=10., n_layer=30
)
mesh1D = set_mesh_1d(hz)
depth = -mesh1D.gridN[:-1]
LocSigZ = -mesh1D.gridCC
time_input_currents_HM = wave_HM.current_times[-7:]
input_currents_HM = wave_HM.currents[-7:]
time_input_currents_LM = wave_LM.current_times[-13:]
input_currents_LM = wave_LM.currents[-13:]
TDsurvey = EM1DSurveyTD(
rx_location = np.array([0., 0., 100.+30.]),
src_location = np.array([0., 0., 100.+30.]),
topo = np.r_[0., 0., 100.],
depth = depth,
rx_type = 'dBzdt',
wave_type = 'general',
src_type = 'CircularLoop',
a = 13.,
I = 1.,
time = time_HM,
time_input_currents=time_input_currents_HM,
input_currents=input_currents_HM,
n_pulse = 2,
base_frequency = 25.,
use_lowpass_filter=True,
high_cut_frequency=210*1e3,
moment_type='dual',
time_dual_moment = time_HM,
time_input_currents_dual_moment=time_input_currents_LM,
input_currents_dual_moment=input_currents_LM,
base_frequency_dual_moment=210,
)
sig_half = 1./50.
chi_half = 0.
expmap = Maps.ExpMap(mesh1D)
# -
sig = np.ones(TDsurvey.n_layer)*sig_half
eta = np.zeros(TDsurvey.n_layer)
tau = np.ones(TDsurvey.n_layer) * 0.001
c = np.ones(TDsurvey.n_layer) * 0.7
blk_ind = (-30>LocSigZ) & (-50<LocSigZ)
sig[blk_ind] = 1./10.
eta[blk_ind] = 0.5
blk_ind = (-50>LocSigZ) & (-300<LocSigZ)
sig[blk_ind] = 1./1000.
blk_ind = -300>LocSigZ
sig[blk_ind] = 1./1000
m_true = np.log(sig)
import matplotlib
matplotlib.rcParams['font.size'] = 14
fig, ax = subplots(1,1, figsize=(4, 5))
Utils1D.plotLayer(1./sig, mesh1D, showlayers=False, ax=ax)
ax.set_xlabel("Resistivity ($\Omega$m)")
ax.set_ylim(-200, 0.)
plt.tight_layout()
fig.savefig('resistivity', dpi=200)
# %%time
prob = EM1D(mesh1D, sigmaMap=expmap, verbose=False, eta=eta, tau=tau, c=c)
if prob.ispaired:
prob.unpair()
if TDsurvey.ispaired:
TDsurvey.unpair()
prob.pair(TDsurvey)
prob.chi = np.zeros(TDsurvey.n_layer)
d_true = TDsurvey.dpred(m_true)
# J = prob.getJ_sigma(m_true)
# %%time
prob = EM1D(mesh1D, sigmaMap=expmap, verbose=False)
if prob.ispaired:
prob.unpair()
if TDsurvey.ispaired:
TDsurvey.unpair()
prob.pair(TDsurvey)
prob.chi = np.zeros(TDsurvey.n_layer)
d_true_em = TDsurvey.dpred(m_true)
# J = prob.getJ_sigma(m_true)
np.random.seed(1)
TDsurvey.dtrue = d_true
std = 0.05
noise = std*abs(TDsurvey.dtrue)*np.random.randn(*TDsurvey.dtrue.shape)
floor = 0.
std = 0.1
TDsurvey.dobs = TDsurvey.dtrue+noise
uncert = abs(TDsurvey.dobs)*std+floor
dmisfit = DataMisfit.l2_DataMisfit(TDsurvey)
uncert = (abs(TDsurvey.dobs)*std+floor)
dmisfit.W = 1./ uncert
d_true_ip = d_true - d_true_em
fig, axes = subplots(1,1, figsize = (6,4))
axes.plot(TDsurvey.time*1e3, -d_true[:TDsurvey.n_time], '-', lw=2, color='k')
axes.plot(TDsurvey.time*1e3, d_true[:TDsurvey.n_time], '--', lw=2, color='k')
axes.plot(TDsurvey.time*1e3, -d_true_em[:TDsurvey.n_time], '-', lw=2, color='b')
axes.set_xscale('log');
axes.set_yscale('log');
plt.legend(("EMIP (+)", "EMIP (-)", "EM (+)"), loc=3)
plt.xlabel("Time (ms)")
plt.ylabel("SkyTEM data (V/A-m$^2$)")
plt.tight_layout()
# fig.savefig('skytem_data', dpi=200)
fig, axes = subplots(1,1, figsize = (6,4))
axes.plot(TDsurvey.time, -d_true[:TDsurvey.n_time], '-', lw=2, color='k')
axes.plot(TDsurvey.time, d_true[:TDsurvey.n_time], '--', lw=2, color='k')
axes.plot(TDsurvey.time, -d_true_em[:TDsurvey.n_time], '-', lw=2, color='b')
axes.plot(TDsurvey.time, -d_true_ip[:TDsurvey.n_time], '-', lw=2, color='r')
axes.plot(TDsurvey.time, d_true_ip[:TDsurvey.n_time], '--', lw=2, color='r')
axes.set_xscale('log');
axes.set_yscale('log');
plt.legend(("EMIP (+)", "EMIP (-)", "EM (+)","IP (+)", "IP (-)"), loc=3)
plt.xlabel("Time (ms)")
plt.ylabel("SkyTEM data (V/A-m$^2$)")
plt.tight_layout()
# fig.savefig('skytem_data', dpi=200)
| notebooks/examples/EM1D_inversion_TD_layers_dual_moment_ip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cz4071
# language: python
# name: cz4071
# ---
import requests, random, tqdm
import pandas as pd
from bs4 import BeautifulSoup
random.seed(42)
df = pd.read_excel('DataScientists.xls')
print('old', df.shape)
df = df[df.dblp.duplicated(keep='last')].sort_values(by=['name']).reset_index(drop=True)
print('new', df.shape)
df.head()
def get_links(node_url):
'''node_url: url of node person
returns: set of people linked to person
'''
soup = BeautifulSoup(requests.get(node_url).content, 'html5lib')
name = [component.text.strip() for component in soup.find('span', attrs={'class': 'name primary'})][0]
links = {para.text.strip() for para in soup.find_all('span', attrs={'itemprop': 'author'})}
links.remove(name)
links = ','.join(sorted(list(links)))
return dict(name=name, links=links)
clean_df = dict(name=[], country=[], institution=[], links=[], expertise=[])
with tqdm.tqdm(total=len(df)) as pbar:
for _, row in df.iterrows():
_, country, institution, node_url, _ = row.values
data = get_links(node_url)
clean_df['name'].append(data['name'])
clean_df['links'].append(data['links'])
clean_df['country'].append(country)
clean_df['institution'].append(institution)
clean_df['expertise'].append(random.randint(1, 10))
pbar.update(1)
clean_df = pd.DataFrame.from_dict(clean_df)
clean_df.head()
clean_df.to_csv('clean.csv', index=False)
| clean_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Conditional list comprehensions for time-stamped data
# Great, you've successfully extracted the data of interest, the time, from a pandas DataFrame! Let's tweak your work further by adding a conditional that further specifies which entries to select.
#
# In this exercise, you will be using a list comprehension to extract the time from time-stamped Twitter data. You will add a conditional expression to the list comprehension so that you only select the times in which entry[17:19] is equal to '19'. The pandas package has been imported as pd and the file 'tweets.csv' has been imported as the df DataFrame for your use.
#
# ### Instructions
#
# - Extract the column 'created_at' from df and assign the result to tweet_time.
#
# - Create a list comprehension that extracts the time from each row in tweet_time. Each row is a string that represents a timestamp, and you will access the 12th to 19th characters in the string to extract the time. Use entry as the iterator variable and assign the result to tweet_clock_time. Additionally, add a conditional expression that checks whether entry[17:19] is equal to '19'.
import pandas as pd
df=pd.read_csv('tweets.csv')
df.head()
# +
# Extract the created_at column from df: tweet_time
tweet_time = df['created_at']
# Extract the clock time: tweet_clock_time
tweet_clock_time = [entry[11:19] for entry in tweet_time if entry[17:19]==19 ]
# Print the extracted times
print(tweet_clock_time)
# -
# here entry[17:19]=='19' need to give '' cause match the string. Said that Each row is a string
# +
# Extract the created_at column from df: tweet_time
tweet_time = df['created_at']
# Extract the clock time: tweet_clock_time
tweet_clock_time = [entry[11:19] for entry in tweet_time if entry[17:19]=='19' ]
# Print the extracted times
print(tweet_clock_time)
| Python Data Science Toolbox -Part 2/List comprehensions and generators/13. Conditional list comprehensions for time-stamped data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ThabisoEDSA/MBTI-Personality-Classifier/blob/master/EDSA_Thabz_Xente.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="vaFCeHrokBZh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="b2455d70-7298-448c-a408-a2d57ff091b9"
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="DL8-yNOJkZe4" colab_type="text"
# #### Xente Fraud Detection System Classification
# >>> Author : <NAME>
# + [markdown] id="FvtqT6KUk6pE" colab_type="text"
# #### Importing packages and Data
# + id="LtR0pEBbk3j7" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.metrics import log_loss
import seaborn as sns
from scipy import stats
from scipy.stats import norm, skew #for some statistics
from scipy.special import boxcox1p
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# + id="O9D-1jYklHps" colab_type="code" colab={}
#### Let's read in our MBTI Headset with 'read_csv'
train = pd.read_csv('/content/gdrive/My Drive/Explore/Xente/training.csv' )
test = pd.read_csv('/content/gdrive/My Drive/Explore/Xente/testing.csv' )
sample = pd.read_csv('/content/gdrive/My Drive/Explore/Xente/sample_submission.csv' )
# + id="6_8nyre3QgDg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a3ad9bc2-ab5c-4914-c099-85207ac62ef6"
len(sample)
# + id="vVnxq82Xl7dq" colab_type="code" colab={}
xe_train = train.copy()
xe_test = test.copy()
# + id="TErFGH4AmGZD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ef115e64-dadd-4de9-b46c-dd4fbc7c642d"
len(xe_train)
# + id="AkTYjgndmI-T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="2f60bb58-f217-4305-d6f8-e4dcbc2c9ef6"
#### now let's check if there are any missing values in this dataset
xe_train.isnull().sum()
# + id="ztSfIhSopNPo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7687be3f-b90d-41fc-ea16-63362cdb7b5c"
##The data has no missing values
### Let's count the number of fraudulent transanctions versus non fradulent trans
print('We have : ', train[train['FraudResult']==1]['FraudResult'].count(), 'fradulent transactions')
# + id="UQ7hz6nwpzHy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dfbbcbaf-941a-4088-c18f-39ddd34c2adc"
print('We have : ', train[train['FraudResult']==0]['FraudResult'].count(), 'non-fradulent transactions')
# + id="NsonkBgcq2rj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="86370507-badb-4150-c596-a4986b1de946"
train.columns
# + [markdown] id="Jr-jRCLysgF7" colab_type="text"
# - as we can see we have a very imbalanced dataset as the number of non-fradulent transaction are way higher
# - this implies we must aim to deal with thi issue
# + id="YS5DVAA4swYn" colab_type="code" colab={}
### what categories of services do we have
# + id="SZ_bCuaCthBv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="898782b8-9835-42ce-8eed-58120eda724d"
train['ProductCategory'].unique()
# + id="qNGLE1EPwlmq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="dcbed6b4-c930-4b85-dc4b-7cd113e8e169"
#### what service providers do we have
train['ProviderId'].unique()
# + id="cKHsgl_XxQsi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="89704d87-d9ad-4a9e-ccc9-1c22bfeec894"
#### what product
train['ProductId'].unique()
# + id="JqjFsSSjxuaK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="b4787789-7330-489b-fa3f-bbc6600047de"
#### There are 23 products on
#### let's check how many channels do we have
train['ChannelId'].unique()
# + id="jjC2ZP6qyPqb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="056ae569-3b05-4ee8-b6fc-9cc595168f42"
train['TransactionStartTime'].unique()
# + id="DK87zT4iyfLL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a587e780-e518-4476-88d9-3345a948ace3"
### what pricing stratergy is used
train['PricingStrategy'].unique()
# + id="FmdgyuGDyvo7" colab_type="code" colab={}
# + [markdown] id="lilB3IGp0hQ_" colab_type="text"
# - Let's view our data as categorical and numerical column
# - First let's chech the munerical columns
# + id="AT7lT2ap0ufr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="278a36c5-ab75-41a0-aa4f-945fead0d798"
xe_train.select_dtypes(include=['int','int64','float']).columns
print(len(xe_train.select_dtypes(include=['int','int64','float']).columns))
# + id="TxpQvFui0xEY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="dfcca73b-762f-4348-ce74-de51c13c10d6"
xe_train.select_dtypes(include='object').columns
print(xe_train.select_dtypes(include='object').columns)
# + id="zaZrK-sQ2YAO" colab_type="code" colab={}
combined = pd.concat([xe_train,xe_test], sort=False)
# + id="eR8_nLcSjjYw" colab_type="code" colab={}
#### dealing with the imbalance in the data
# + id="xzyMWq4E9oA-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="b5a4cd14-3fac-4d3c-d522-783dc7b5a4db"
combined.isnull().sum()
# + id="dWsxeDYybgAZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a9e78d36-4f0b-44ad-858b-f4f186e5e11a"
len(combined)
# + id="UtUxxLJGcIMG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4a322aaa-e75c-4304-b13e-a2b1e26e92e6"
### let's loo
values_count = combined.nunique().sort_values()
np.sum(values_count == 1)
# + id="Xz7NmunIclUH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ddfe6298-04ec-4cd4-e298-1213bb53ee70"
#### now let's look at duplicates
duplicates = []
for i, ref in enumerate(combined.columns[:-1]):
for other in combined.columns[i + 1:-1]:
if other not in duplicates and np.all(combined[ref] == combined[other]):
duplicates.append(other)
len(duplicates)
# + id="9Tk8sixuc9hn" colab_type="code" colab={}
# + id="d5vYZl_Hdwlv" colab_type="code" colab={}
# + id="H0vFL4vX-p2g" colab_type="code" colab={}
### our train dataset had no missing values but the test dataset introduced some N/As
### therefore we have to fill them in with zeros
combined['FraudResult']=combined['FraudResult'].fillna(0)
# + id="4CdTleu0_NKq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="e29aef98-9bd1-4081-fe2e-e8e082725d9d"
### Let's recheck our dataset for missing values
combined.isnull().sum()
# + id="Oxj1x7tq1Vdk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5f71ff56-3855-4fa7-e83e-00990be369b0"
#### Since we have no missing values lets lable encode ou categorical variables
###Let's label encode catecorical variables above
#Label Encoding
cols = ('TransactionId', 'BatchId', 'AccountId', 'SubscriptionId', 'CustomerId',
'CurrencyCode', 'ProviderId', 'ProductId', 'ProductCategory',
'ChannelId', 'TransactionStartTime')
# process columns, apply LabelEncoder to categorical features
for c in cols:
if c in cols:
lbl = LabelEncoder()
lbl.fit(list(combined[c].values))
combined[c] = lbl.transform(list(combined[c].values))
# shape
print('Shape combined: {}'.format(combined.shape))
# + id="xQo0X9NH2Fuc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="c49da5b4-85c3-42b9-c528-d7e0c22eb40f"
combined.head(3)
# + id="QSPbQpWB2jZM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 348} outputId="ab330564-0ea5-4d18-a015-13991e88ab60"
#### let's just deal with skewed data
num_feats = new_data.dtypes[new_data.dtypes != "object"].index
# Check the skew of all numerical features
skewed_feats = new_data[num_feats].apply(lambda x: skew(x.dropna())).sort_values(ascending=False)
print("\nSkew in numerical features: \n")
skewness = pd.DataFrame({'Skew' :skewed_feats})
skewness = skewness.drop('FraudResult', 0)
skewness.head(15)
# + id="JjrmLz8y3nvM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5809c7e8-7cd2-443c-9f31-9211358fb129"
#### let's just box cox transform this thing and see what happens
#Correct for skewness by using boxcox1p
skewness = skewness[abs(skewness) > 0.75]
print("There are {} skewed numerical features to Box Cox transform".format(skewness.shape[0]))
from scipy.special import boxcox1p
skewed_features = skewness.index
lam = 0.15
for feat in skewed_features:
new_data[feat] = boxcox1p(new_data[feat], lam)
# + id="44JxvrkJ8jsW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="78ae90c3-1ef7-46e1-f3be-7b73c908b8b3"
# I didn't use getdummies
#Get Dummies
#combined = pd.get_dummies(combined)
#len_train = train.shape[0]
#combined.head(3)
combined.head()
# + id="YmhuK7FJ9aN-" colab_type="code" colab={}
###
### As we can see boxcoxing this introduces nans in the value column
### we will replace that with mean of the column
combined['Amount']=combined['Amount'].fillna(combined['Amount'].mean())
# + id="NnjqUFGP9Up2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="66c7fe86-058f-4ceb-f26c-b3ad700385d8"
#Based on data descriptions the following columns NA's will be filled with 'None'
combined.isnull().sum()
# + id="7o3VdMD7AzPf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="d1e98721-4203-4e9a-fa71-e82ad9563805"
combined.info()
# + id="FDzsP1ccB3uq" colab_type="code" colab={}
combined['FraudResult'] = combined.FraudResult.astype(int)
# + id="xxXB4DWV4Yxc" colab_type="code" colab={}
#### Drop a few columns let's see what happens
# + id="Pxkc4SFsCvhj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="107f78c2-b68d-4983-f59f-d5c7034d8d88"
combined.head()
# + id="e8L7sVPV6s0k" colab_type="code" colab={}
len_train = len(xe_train)
X_train = combined[:len_train]
y_train = X_train.FraudResult.values
X_train = X_train.drop('FraudResult', 1)
X_test = combined[len_train:]
y_test = X_test.FraudResult.values
Xtest = X_test.drop(['FraudResult'], 1)
# + id="QTRnoETZ6vTt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b282eb18-9d52-41f9-8acf-84bb485abe1e"
len(X_train)
# + id="HaDkDfzQ7OP1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="67de40c8-71e8-492c-dbd7-1b2a103eb5d9"
len(y_train)
# + id="sF9aDchr7vJd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="15e17829-382f-4048-9900-5973ee81fc11"
len(X_test)
# + id="m1pqd0gsTnim" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4af44e6e-ba48-465a-bfda-9d58e4b0135a"
len(y_test)
# + id="Z2pVu8bQUBbw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="ff32e882-54e5-48ab-c196-a12a70d0db64"
X_test.drop('FraudResult',1, inplace=True)
X_test.head(3)
# + id="naCsV3lWTp7K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="45519eb8-ab8a-4430-ecf6-701595e94561"
### great now let's do the train test split
model = LogisticRegression()
#y = combined['FraudResult']
#X = combined.drop('FraudResult',1)
#X_train,X_test,y_train ,y_test = train_test_split(X,y, test_size= 0.50,random_state=40)
#### train the model on the training data
model.fit(X_train,y_train)
y_hat = model.predict( X_test)
print('Accuracy Score:',accuracy_score(y_test, y_hat)*100)
#print(accuracy_score(y_hat,label_test1)*100)
print(log_loss(y_test,y_hat,labels=[0,1]))
#X_test.head(3)
# + id="nss3QuK_T005" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e77f426-5e3d-49aa-baa9-b79959d0fcfe"
len(y_hat)
# + id="MFjFKVnqV5Sx" colab_type="code" colab={}
#Output to CSV
output_avg = pd.DataFrame({'A': xe_test.TransactionId, 'B': y_hat})
output_avg.to_csv('/content/gdrive/My Drive/Explore/Xente/submission1_xente.csv', index=False)
# + id="dM1-OyPLWY2R" colab_type="code" colab={}
| EDSA_Thabz_Xente.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solving the problem of 3-coloring using CSP
#
# This is a small example to show how you can encode the problem of [3-coloring](https://en.wikipedia.org/wiki/Graph_coloring#Vertex_coloring) into a constraint satisfaction problem and using a CSP solver to solve the problem.
#
# We will use Google's [OR-Tools](https://developers.google.com/optimization/introduction/overview) in Python (see this [example](csp.ipynb) for details on how to set this up).
#
# Let's start by creating a CSP instance.
# +
from ortools.sat.python import cp_model
model = cp_model.CpModel();
# -
# ## 3-coloring
#
# In the problem of 3-coloring, you are given as input a finite (undirected) [graph](https://en.wikipedia.org/wiki/Graph_(discrete_mathematics)) $G = (V,E)$, consisting of a set $V$ of *vertices* (or *nodes*) and a set $E$ of *edges*. Each edge $e \in E$ consists of a set $\{v_1,v_2\}$ of exactly two vertices in $V$.
#
# A *coloring* of a graph $G = (V,E)$ is a function $\mu : V \rightarrow C$ that assigns to each vertex $v \in V$ a color $\mu(v) \in C$. The coloring $\mu$ is called *proper* if for every edge $\{v_1,v_2\} \in E$, the coloring assigns different colors to the two endpoints of the edge—that is, $\mu(v_1) \neq \mu(v_2)$.
#
# In the problem of 3-coloring, the question is to decide if there exists a proper coloring $\mu : V \rightarrow \{1,2,3\}$ of the graph that only uses three colors.
#
# ## Encoding the graph and colorings
#
# Take some graph $G = (V,E)$, that comprises an input for the 3-coloring problem.
# Suppose that the vertices $V = \{ v_1,\dotsc,v_n \}$ are numbered from $1$ to $n$.
#
# We will encode this graph in Python in the following way.
#
# To explain this, take the following example graph $G = (V,E)$, where $V = \{v_1,v_2,v_3,v_4\}$ and $E = \{ \{v_1,v_2\}, \{v_1,v_3\}, \{v_2,v_3\}, \{v_2,v_4\}, \{v_3,v_4\} \}$. We encode the set of edges as a list of pairs, where each vertex is indicated by its index.
num_vertices = 4;
edges = [(1,2),(1,3),(2,3),(2,4),(3,4)];
# Then, we will have to represent the possible 3-colorings of the graph $G$ using variables. We will do this as follows. For each vertex $v_i$, we will have a variable $x_i$ with domain $\{1,2,3\}$.
vars = dict()
for i in range(1,num_vertices+1):
vars[i] = model.NewIntVar(1, 3, "x{}".format(i));
# The different assignments of the variables $x_1,\dotsc,x_4$ to the domain $\{1,2,3\}$ correspond exactly to all possible 3-colorings of the vertices $v_1,\dotsc,v_4$.
#
# ## Ensuring that colorings are proper
#
# To make sure that only truth assignments are allowed that correspond to *proper* 3-colorings, we need to add some constraints.
#
# For each edge $\{ v_{i_1},v_{i_2} \} \in E$, and each color $c$, we will make sure that the vertices $v_{i_1}$ and $v_{i_2}$ do not both get colored with color $c$.
# For each edge $\{ v_{i_1},v_{i_2} \} \in E$, we add a constraint that expresses that the value assigned to $x_{i_1}$ is different from the value assigned to $x_{i_2}$.
for (i,j) in edges:
model.Add(vars[i] != vars[j]);
# ## Calling the CSP solver and constructing a coloring
#
# All together, the constraints in our CSP instance ensure the following. Every assignment that satisfies the constraints (if such an assignment exists) corresponds to a proper 3-coloring of the graph $G$. Therefore, we can now call the CSP solver to find a solution to the CSP instance, if it exists. And if it exists, we can use it to construct a proper 3-coloring of the graph.
#
# For our simple example, a solution to the CSP instance exists, and we get a proper 3-coloring of the graph.
# +
solver = cp_model.CpSolver();
answer = solver.Solve(model);
if answer == cp_model.FEASIBLE:
print("The graph is 3-colorable:");
for i in range(1,num_vertices+1):
print("- Vertex {} gets color {}".format(i,solver.Value(vars[i])));
else:
print("The graph is not 3-colorable!");
| examples/3coloring-csp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Kipoi python API
# ## Quick start
# There are three basic building blocks in kipoi:
#
# - **Source** - provides Models and DataLoaders.
# - **Model** - makes the prediction given the numpy arrays.
# - **Dataloader** - loads the data from raw files and transforms them into a form that is directly consumable by the Model
# 
# ## List of main commands
#
#
# Get/list sources
# - `kipoi.list_sources()`
# - `kipoi.get_source()`
#
#
# List models/dataloaders
# - `kipoi.list_models()`
# - `kipoi.list_dataloaders()`
#
# Get model/dataloader
# - `kipoi.get_model()`
# - `kipoi.get_dataloader_factory()`
#
# Load only model/dataloader description from the yaml file without loading the model
#
# - `kipoi.get_model_descr()`
# - `kipoi.get_dataloader_descr()`
#
# Install the dependencies
# - `kipoi.install_model_dependencies()`
# - `kipoi.install_dataloader_dependencies()`
import kipoi
# ### Source
#
# Available sources are specified in the config file located at: `~/.kipoi/config.yaml`. Here is an example config file:
#
# ```yaml
# model_sources:
# kipoi: # default
# type: git-lfs # git repository with large file storage (git-lfs)
# remote_url: <EMAIL>:kipoi/models.git # git remote
# local_path: ~/.kipoi/models/ # local storage path
# gl:
# type: git-lfs # custom model
# remote_url: https://i12g-gagneurweb.informatik.tu-muenchen.de/gitlab/gagneurlab/model-zoo.git
# local_path: /s/project/model-zoo
# ```
#
# There are three different model sources possible:
#
# - **`git-lfs`** - git repository with source files tracked normally by git and all the binary files like model weights (located in `files*` directories) are tracked by [git-lfs](https://git-lfs.github.com).
# - Requires `git-lfs` to be installed.
# - **`git`** - all the files including weights (not recommended)
# - **`local`** - local directory containing models defined in subdirectories
#
# For **`git-lfs`** source type, larger files tracked by `git-lfs` will be downloaded into the specified directory `local_path` only after the model has been requested (when invoking `kipoi.get_model()`).
#
# #### Note
#
# A particular model/dataloader is defined by its source (say `kipoi` or `my_git_models`) and the relative path of the desired model directory from the model source root (say `rbp/`).
#
# A directory is considered a model if it contains a `model.yaml` file.
import kipoi
# +
import warnings
warnings.filterwarnings('ignore')
import logging
logging.disable(1000)
# -
kipoi.list_sources()
s = kipoi.get_source("kipoi")
s
kipoi.list_models().head()
# ## Model
#
# Let's choose to use the `rbp_eclip/UPF1` model from kipoi
MODEL = "rbp_eclip/UPF1"
# **NOTE:** If you are using python2, use a different model like `MaxEntScan/3prime` to following this example.
# Note. Install all the dependencies for that model:
# add --gpu flag to install gpu-compatible dependencies (e.g. installs tensorflow-gpu instead of tensorflow)
# !kipoi env install {MODEL}
model = kipoi.get_model(MODEL)
# ### Available fields:
#
# #### Model
#
# - type
# - args
# - info
# - authors
# - name
# - version
# - tags
# - doc
# - schema
# - inputs
# - targets
# - default_dataloader - loaded dataloader class
#
#
# - predict_on_batch()
# - source
# - source_dir
# - pipeline
# - predict()
# - predict_example()
# - predict_generator()
#
# #### Dataloader
#
# - type
# - defined_as
# - args
# - info (same as for the model)
# - output_schema
# - inputs
# - targets
# - metadata
#
#
# - source
# - source_dir
# - example_kwargs
# - init_example()
# - batch_iter()
# - batch_train_iter()
# - batch_predict_iter()
# - load_all()
model
model.type
# ### Info
model.info
model.info.version
# ### Schema
dict(model.schema.inputs)
model.schema.targets
# ### Default dataloader
#
# Model already has the default dataloder present. To use it, specify
model.source_dir
model.default_dataloader
model.default_dataloader.info
# ### Predict_on_batch
model.predict_on_batch
# ### Others
# Model source
model.source
# model location directory
model.source_dir
# ## DataLoader
DataLoader = kipoi.get_dataloader_factory(MODEL)
# same as DataLoader = model.default_dataloader
# A dataloader will most likely require input arguments in which the input files are defined, for example input fasta files or bed files, based on which the model input is generated. There are several options where the dataloader input keyword arguments are displayed:
# Display information about the dataloader
print(DataLoader.__doc__)
# Alternatively the dataloader keyword arguments can be displayed using the function:
kipoi.print_dl_kwargs(DataLoader)
# ## Run dataloader on some examples
# each dataloader already provides example files which can be used to illustrate its use:
DataLoader.example_kwargs
import os
# # cd into the source directory
os.chdir(DataLoader.source_dir)
# !tree
dl = DataLoader(**DataLoader.example_kwargs)
# could be also done with DataLoader.init_example()
# This particular dataloader is of type Dataset
# i.e. it implements the __getitem__ method:
dl[0].keys()
dl[0]["inputs"]["seq"][:5]
dl[0]["inputs"]["seq"][:5]
len(dl)
# ### Get the whole dataset
whole_data = dl.load_all()
whole_data.keys()
whole_data["inputs"]["seq"].shape
# ### Get the iterator to run predictions
it = dl.batch_iter(batch_size=1, shuffle=False, num_workers=0, drop_last=False)
next(it)["inputs"]["seq"].shape
model.predict_on_batch(next(it)["inputs"])
# ### Pipeline
#
# Pipeline object will take the dataloader arguments and run the whole pipeline directly:
#
# ```
# dataloader arguments --Dataloader--> numpy arrays --Model--> prediction
# ```
example_kwargs = model.default_dataloader.example_kwargs
preds = model.pipeline.predict_example()
preds
model.pipeline.predict(example_kwargs)
next(model.pipeline.predict_generator(example_kwargs, batch_size=2))
from kipoi_utils.data_utils import numpy_collate
numpy_collate_concat(list(model.pipeline.predict_generator(example_kwargs)))
# ### Re-train the Keras model
#
# Keras model is stored under the `.model` attribute.
model.model.compile("adam", "binary_crossentropy")
train_it = dl.batch_train_iter(batch_size=2)
# +
# model.model.summary()
# -
model.model.fit_generator(train_it, steps_per_epoch=3, epochs=1)
| notebooks/python-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Configuration
with open('startup.py') as f:
exec(f.read())
public(ConfigParser)
help(ConfigParser.read)
import conf
highlight(conf)
help(conf.asdict)
site.addsitedir(str(Path.home() / 'Development/hw-4.2.0/hw'))
from tools import *
# hide
os.chdir('../')
# +
class Configuration():
'''Class to handle program configuration.
'''
CONF_DIR_NAMES = ['etc', 'conf', 'cfg']
def __init__(self):
'''Scan the `etc`, `cfg` and `conf` directories for compatible files and parse them.
'''
for dirname in CONF_DIR_NAMES:
# -
pp(os.confstr_names)
help(zip)
| nb/config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import string
import cv2
import matplotlib.pyplot as plt
import numpy as np
from captcha.image import ImageCaptcha
from bson import ObjectId
% matplotlib inline
% config InlineBackend.figure_format = 'retina'
characters = string.digits + string.ascii_uppercase
def batch_generate_captcha_mat(count, cnt=1, w=30, h=30):
capt = ImageCaptcha(width=cnt * 34 + 26, height=60)
mat_x = np.ndarray((count, w, h), dtype=np.uint8)
mat_y = np.zeros((count, corpus_len), dtype=np.uint8)
for i in range(0, count):
cid = random.randint(0, corpus_len - 1)
c = corpus[cid]
capt_img = np.array(capt.generate_image(c))
gray_img = cv2.cvtColor(capt_img, cv2.COLOR_RGB2GRAY)
scle_img = cv2.resize(gray_img, (w, h))
mat_x[i] = scle_img
mat_y[i][cid] = 1.0
return mat_x, mat_y
def show_img(img, zoom=4, dpi=80):
w = img.shape[0]
h = img.shape[1]
plt.figure(figsize=(w*zoom/dpi, h*zoom/dpi), dpi=dpi)
plt.axis('off')
plt.imshow(img, cmap=plt.get_cmap('gray'))
plt.show()
return
def batch_generate_captcha_to_file(corpus, batch=1, capt_len=1, w=30, h=30):
capt = ImageCaptcha(width=capt_len * 30, height=60)
for i in range(0, batch):
chars = ''
for j in range(0, capt_len):
chars += random.choice(corpus)
# image = capt.generate_image(chars)
filename = 'data/{}_{}.png'.format(str(ObjectId()), chars)
capt.write(chars, filename)
batch_generate_captcha_to_file(characters, batch=1, capt_len=4)
# -
| captcha.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Subrepresentations
#
# ## Subrepresentations
#
# A subrepresentation is a subvector space that is also a representation of $G$ under the same action.
#
# ```{admonition} Definition (Subrepresentation)
# :class: definition
#
# Let $V$ be a linear representation of a group $G$. A subspace $W$ of $V$ is a subrepresentation if $W$ is invariant under $G$ - that is, if $g.w\in W$ for all $g\in G$ and all $w\in W$.
# ```
#
# ```{admonition} Examples (Subrepresentation)
# :class: example
#
# - Every subspace is a subrepresentation of the trivial representation on any vector space since the trivial $G$ action takes every subspace back to itself
#
# - The tautological representation of $D_4$ on $\mathbb{R}^2$ has no proper non-zero subrepresentations because there is no line taken back to itself under every symmetry of the square, that is, there is no line left invariant by $D_4$.
# - The vertex permutation representation of $D_4$ on $\mathbb{R}^4$, induced by the action of $D_4$ on a set of basis elements $\{e_1,e_2,e_3,e_4\}$ indexed by the vertices of a square does have a proper non-trivial subrepresentation. For example, the one dimensional subspace spanned by $e_1+e_2+e_3+e_4$ is fixed by $D_4$ -- when $D_4$ acts, it permutes $e_i$ so their sum remains unchanged. Therefore, for all $g\in D_4$, we have
#
# $$
# g.(\lambda,\lambda,\lambda,\lambda)=(\lambda,\lambda,\lambda,\lambda)
# $$
#
# for all vectors in the one-dimensional subspace of $\mathbb{R}^4$. Then, $D_4$ acts trivially on this one-dimensional subrepresentation.
#
# - Another representation of the vertex permutation representation of $D_4$ on $\mathbb{R}^4$ is the subspace $W\subset \mathbb{R}^4$ of vectors whose coordinates sum to 0. When $D_4$ acts by permutinf the coordinates, it leaves their sum unchanged, For example, $B$ sends $(1,2,3,-6)$ to $(-6,3,2,1)$ if vectors are labelled counterclockwise from the upper right. The space $W$ is a three-dimensional representation of $D_4$ on $\mathbb{R}^4$. Note that $W$ is a non-trivial subrepresentation since the elements of $G$ do move the vectors around in the space $W$.
#
# - The standard representaion of $S_n$: Let $S_n$ acts on a vector space of dimension $n$, say $\mathbb{R}^n$, by permuting the $n$ vectors of a fixed basis. Note that the subspace spanned by the sum of the basis vectors is fixed by the action of $S_n$ -- it is a subrepresentation on which $S_n$ acts trivially. More interestingly, the $(n-1)$-dimensional subspace
#
# $$
# W=\left\{\begin{pmatrix}x_1\\x_2\\\vdots\\x_n\end{pmatrix}:\sum_{i=1}^n x_i=0\right\} \subset\mathbb{R}^n
# $$
#
# is also invariant under the permutation action. This is called the standard representation of $S_n$.
#
# ```
#
# ## Direct sum of representations
#
# For representations that have non-zero subrepresentations, it is always possible to simplify the matrix representatives in such a way that all subrepresentations are clearly visible, as blocks in these matrices. Let us take the permutation representation of $S_4$, $\rho:S_3\to GL(\mathbb{R}^3)$:
#
# $$
# \begin{align*}
# e\mapsto \begin{pmatrix}1&0&0\\0&1&0\\0&0&1\end{pmatrix}, \qquad (12)\mapsto \begin{pmatrix}0&1&0\\1&0&0\\0&0&1\end{pmatrix}, \qquad (13)\mapsto \begin{pmatrix}0&0&1\\0&1&0\\1&0&0\end{pmatrix}\\
# (23)\mapsto \begin{pmatrix}1&0&0\\0&0&1\\0&1&0\end{pmatrix}, \qquad (123)\mapsto \begin{pmatrix}0&0&1\\1&0&0\\0&1&0\end{pmatrix}, \qquad (132)\mapsto \begin{pmatrix}0&1&0\\0&0&1\\1&0&0\end{pmatrix}
# \end{align*}
# $$
#
# We know that it does have two subrepresentations:
# - a one-dimensional trivial representation
# - a two-dimensional standard representation
#
# Let us define
#
# $$
# M=\begin{pmatrix}1&-\frac{1}{2}&-\frac{1}{2}\\0&\frac{\sqrt{3}}{2}&-\frac{\sqrt{3}}{2}\\1&1&1\end{pmatrix}
# $$
#
# Then by direct calculation:
#
# $$
# \begin{align*}
# M \rho(e) M^{-1} =\begin{pmatrix}1&0&0\\0&1&0\\0&0&1\end{pmatrix}\qquad M \rho((12)) M^{-1} =\begin{pmatrix}-\frac{1}{2}&\frac{\sqrt{3}}{2}&0\\\frac{\sqrt{3}}{2}&\frac{1}{2}&0\\0&0&1\end{pmatrix}\\
# M \rho((13)) M^{-1} =\begin{pmatrix}-\frac{1}{2}&-\frac{\sqrt{3}}{2}&0\\-\frac{\sqrt{3}}{2}&\frac{1}{2}&0\\0&0&1\end{pmatrix}\qquad M \rho((23)) M^{-1} =\begin{pmatrix}1&0&0\\0&-1&0\\0&0&1\end{pmatrix}\\
# M \rho((123)) M^{-1} =\begin{pmatrix}-\frac{1}{2}&-\frac{\sqrt{3}}{2}&0\\\frac{\sqrt{3}}{2}&-\frac{1}{2}&0\\0&0&1\end{pmatrix}\qquad M \rho((132)) M^{-1} =\begin{pmatrix}-\frac{1}{2}&\frac{\sqrt{3}}{2}&0\\-\frac{\sqrt{3}}{2}&-\frac{1}{2}&0\\0&0&1\end{pmatrix}\\
# \end{align*}
# $$
#
# All these matrices are block-diagonal with a two-dimensional block and a one-dimensional block. By multiplying by $M$ from the left and $M^{-1}$ from the right we just changed the basis of the three-dimensional space. This change of basis exposed the seubrepresentations of the permutation representation of $S_3$.
#
# ```{admonition} Definition (Direct sum of representations)
# :class: definition
#
# Suppose that $G$ acts on vector spaces $V$ and $W$. We can define an action of $G$ coordinate-wise on their direct sum as:
#
# $$
# g.(v,w)=(g.v,g.w)\in V\oplus W
# $$
#
# The matrix associated to every $g$ acting on $V\oplus W$ will be a block diagonal matrix
#
# $$
# \begin{pmatrix}
# \rho_1(g)&0\\0&\rho_2(g)
# \end{pmatrix}
# $$
#
# obtained from the $n\times n$ matrix $\rho_1(g)$ describing the action of $g$ on $V$, and the $m\times m$ matrix $\rho_2(g)$ describing the action of $g$ on $W$. We call $V\oplus W$ the direct sum of representations of $G$.
#
# ```
#
# We are interested in decomposing representations in direct susm of their subrepresentations. To do it we need one more notion:
#
# ```{admonition} Definition (Isomorphic representations)
# :class: definition
#
# Two representations $V$ and $W$ of $G$ are isomorphic if there is a vector space isomorphism between them that preserves the action of $G$, that is, if there exists an isomorphism $\phi:V\to W$ such that
#
# $$
# R_V(g)=\phi^{-1}\circ R_W(g)\circ \phi
# $$
#
# for all $g\in G$, where $R_V:G\to GL(V)$ and $R_W:G\to GL(W)$ are two representations.
#
# ```
#
# ```{admonition} Example (Isomorphic representations)
# :class: example
#
# We have shown that there exists a matrix $M$ such that for all $g\in S_3$ we have
#
# $$
# M R(g) M^{-1}=\begin{pmatrix}R_1(g)&0\\0&R_2(g)\end{pmatrix}
# $$
#
# where $R$ is the permutation representation of $S_3$, $R_1$ is the standard representation of $S_3$, and $R_2$ is the trivial representation of $S_3$. This implies that
#
# $$
# R\cong R_1\oplus R_2
# $$
#
# ```
#
# ## Searching for subrepresentations
#
# Let us consider the following four-dimensional representation $V$ of $D_4$ (called the vertex permutation representation):
# - for the rotation: $A \mapsto \begin{pmatrix}0&0&0&1\\1&0&0&0\\0&1&0&0\\0&0&1&0\end{pmatrix}
# - for the horizontal reflection: $B \mapsto \begin{pmatrix}0&1&0&0\\1&0&0&0\\0&0&0&1\\0&0&1&0\end{pmatrix}
#
# The remaining matrices can be found using the fact that we have a group representation, i.e. it is a group homomorphism. For example
#
# $$
# A^2=A\cdot A\mapsto \phi(A)\cdot \phi(A)=\begin{pmatrix}0&0&0&1\\1&0&0&0\\0&1&0&0\\0&0&1&0\end{pmatrix} ^2=\begin{pmatrix}0&0&1&0\\0&0&0&1\\1&0&0&0\\0&1&0&0\end{pmatrix}
# $$
#
# We want to find all subrepresentations of $V$. To find interesting subrepresentations of $V$, we can look for non-zero vectors in $\mathbb{R}^4$ whose orbits under the action of $D_4$ span proper subspaces of $V$. One way is to find vectors with small orbits. For example, the vector $(1,1,1,1)$ is fixed by $D_4$. It spans a one-dimensional subrepresentation of $V$ where $D_4$ acts trivially.
#
# Another vector with a small orbit is $w=(-1,1,-1,1)$. Note that $A$ acts on $w$ to produce $(1,-1,1,-1)=-w$. Also the reflection $B$ acts the same way and produces $-w$. All other elements of $D_4$ act by superpositions of these two elements, and therefore the orbit of $w$ is just $\{w,-w\}$. Thus the one-dimensional subspace spanned by $w$ is a subrepresentation of $V$. This is not a trivial representation since some elements act by the multiplication by $-1$.
#
# There is also a two-dimensional subrepresentation of $V$ that can be found by considering the orbit of $u=(1,1,-1,-1)$. Its orbit is:
#
# $$
# \mathrm(Orb)(u)=\{(1,1,-1,-1),(-1,1,1,-1),(-1,-1,1,1),(1-1,-1,1)\}
# $$
#
# that are points on a two-dimensional subspace $T$ of $\mathbb{R}^4$ that has a basis $(1,1,-1,-1)$ and $(-1,1,1,-1)$. It is easy to check that the vertex representation action of $D_4$ on $T$ is identical with the tautological representation of $D_4$ on this two-plane.
#
# Finally, since the three subrepresentations described above span $V$, there is a direct sum decomposition of representations:
#
# $$
# V\cong T\oplus \mathbb{R}(1,1,1,1)\oplus \mathbb{R} (1,-1,1,-1)
# $$
#
# Importantly, none of these subrepresentations can be further decomposed.
| docs/_sources/Lectures/Lecture5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [View in Colaboratory](https://colab.research.google.com/github/Sumindar/cnn/blob/master/mnistcnn.ipynb)
# + id="F1W4WJ1xIBpV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="5f0b778f-29e7-4f74-a4dc-29b7f10db775"
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# + id="Euj_wmZRF1rB" colab_type="code" colab={}
from __future__ import print_function
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
from keras.optimizers import RMSprop
K.tensorflow_backend._get_available_gpus()
from matplotlib.pyplot import imshow
import numpy as np
batch_size = 128
num_classes = 10
epochs = 12
# + id="R9wnAWwiF5Oh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="71f766cf-ba6e-451c-d88c-63aceff940f1"
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# + id="0A0cZ4RsF9-M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3bc2135a-0dec-4158-e099-3361b6a1e0ff"
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
y_train.shape
# + id="5DsXuaMWIkzL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="92ea683d-ca9e-4e72-fd8e-4ad81290711f"
y_train[0]
# + id="IpICAsCsHmFX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="000ace85-8652-470c-afc7-b220123c2aa2"
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.summary()
# + id="hlIBt6D-I5dJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="d74f610e-5ba0-4984-88b7-d08a17b598d9"
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# + id="s6OzDGoQz27l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 364} outputId="e34330ee-b4f3-4ad0-9e7e-557f29bde6c7"
imshow(x_test[1].reshape(28,28))
# + id="b0IjB8LY0IX-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="42d1d858-e3b0-4bd5-e8cc-d8fbbb643a02"
np.where(y_test[1]==1)
| mnistcnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import glob
import tensorflow as tf
import tensorflow.keras as keras
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
import cv2
import random
import pickle
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten, Conv2D, MaxPooling2D
import warnings
warnings.filterwarnings("ignore")
warnings.simplefilter("ignore")
# To run on GPU
gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=0.7)
sess = tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
# +
# Fetch all the files from the image folder
files = glob.glob('images/**')
print(files)
dictval={}
i = 0
# Iterate over every file and try to save data to the dictval
for file in files:
print(file)
if "batches.meta" in file:
# batches.meta contains the data for the label names
# and size of the batch
with open(file,'rb') as fo:
data = pickle.load(fo, encoding='bytes')
print(data)
else:
with open(file, 'rb') as fo:
temp = pickle.load(fo, encoding='bytes')
#print(temp)
if i == 0:
dictval['data']= list(temp[b'data'])
dictval['labels']= list(temp[b'labels'])
else:
dictval['data'] = dictval['data'] + list(temp[b'data'])
dictval['labels'] = dictval['labels'] + list(temp[b'labels'])
i+=1
# -
# Convert the bytes to the normal string
print(data[b'label_names'])
labels = [x.decode('utf-8') for x in data[b'label_names']]
print(labels)
# +
alldata = dictval['data']
alldatalabels = dictval['labels']
trainingdata = []
def create_training_data():
def reshapedata(imdata, imlabel):
print(len(imdata))
# fig=plt.figure(figsize=(8, 8))
for i in range(len(imdata)):
# for i in range(1,5):
# This data is the in the format of 3072 array elements
temp = imdata[i]
#print(temp)
#print(len(temp))
# To reshape the data
img = np.reshape(temp, (3, 32,32)).T
#print(img.shape)
# Convert the numpy array into the RGB format
img = Image.fromarray(img, 'RGB')
# To see the image without correct orientation
#plt.imshow(img)
#plt.show()
# img is in rotated format, so we need to rotate the image
# to get the original orientation
img = img.rotate(270)
# Here gray conversion is done: in our application color images are not need because we
# can get the same information in the gray image.
# Benefit of using gray image : It will reduce the calculations by 3(RGB have 3 channels)
img = cv2.cvtColor(np.array(img), cv2.COLOR_RGB2BGR)
img = cv2.cvtColor(np.array(img), cv2.COLOR_BGR2RGB)
# Just to make sure that every image is 32*32
# img = cv2.resize(img, (32,32))
# To see the image in the correct orientation
# fig.add_subplot(5, 5, i)
# plt.imshow(img)
# Just to verify that every label is int
if type(imlabel[i]) != type(2):
continue
class_num = imlabel[i]
# plt.title(labels[class_num])
# print(labels[class_num])
# break
# To create training data:
# I have appened the image data and the label
# temp[0]: This is image
# temp[1]: This is label
temp = [img, class_num]
trainingdata.append(temp)
#break
reshapedata(alldata, alldatalabels)
create_training_data()
print("This is the dimension of the data")
print(len(trainingdata))
# -
# This is to make the data shuffled randomly
import random
random.shuffle(trainingdata)
# +
# This is just to check whether every data is
# append correctly or not.
for sample in trainingdata[:10]:
print("label = %d" %sample[1])
X = []
Y = []
# This is to store all the images in X
# and all the labels in Y
for features, label in trainingdata:
X.append(features)
Y.append(label)
# To reshape informat of tensorflow
X = np.array(X).reshape(-1,32, 32, 3)
print(X.shape)
# plt.imshow(X[4,:,:,:])
# plt.show()
# +
#X = pickle.load(open("X.pickle", 'rb'))
#Y = pickle.load(open("Y.pickle", 'rb'))
# to normalize the data.
X = X.astype('float32')
X /= 255.0
plt.imshow(X[4,:,:,:])
plt.show()
# print(X[8,:,:,:])
# 60% Training data
x_train = X[:30000]
y_train = Y[:30000]
# 20% Testing data
x_test = X[30000:40000]
y_test = Y[30000:40000]
# 20% Validation data
x_val = X[40000:50000]
y_val = Y[40000:50000]
# +
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', input_shape=(32, 32, 3)))
model.add(Dropout(0.3))
model.add(Conv2D(32, (3, 3), activation='relu'))
model.add(Dropout(0.3))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(Dropout(0.3))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(Dropout(0.3))
model.add(Conv2D(128, (3, 3), activation='relu'))
model.add(Dropout(0.2))
model.add(Conv2D(128, (3, 3), activation='relu'))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(20, activation='relu'))
model.add(Dense(10, activation='softmax'))
# model = Sequential()
# model.add(Conv2D(32, (3, 3), padding='same', activation='relu', input_shape=(32, 32, 3)))
# model.add(Dropout(0.2))
# # model.add(Conv2D(32,(3,3),padding='same', activation='relu'))
# # model.add(Dropout(0.2))
# model.add(Conv2D(64,(3,3),padding='same',activation='relu'))
# model.add(Dropout(0.2))
# # model.add(Conv2D(64,(3,3),padding='same',activation='relu'))
# # model.add(Dropout(0.2))
# model.add(Conv2D(128,(3,3),padding='same',activation='relu'))
# model.add(Dropout(0.2))
# # model.add(Conv2D(128,(3,3),padding='same',activation='relu'))
# # model.add(Dropout(0.2))
# model.add(Flatten())
# # model.add(Dropout(0.2))
# model.add(Dense(128,activation='relu'))
# model.add(Dropout(0.2))
# model.add(Dense(10, activation='softmax'))
# model.summary()
# -
# Model compilation is done
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy',metrics=['accuracy'])
# Here we are going to train the model
model.fit(x_train, y_train,batch_size=200, validation_data=(x_val, y_val), epochs = 100)
# To find the test accuracy
test_loss, test_acc = model.evaluate(x_test, y_test)
print(test_acc)
model.save('ourmodel.h5')
saved_model = tf.keras.models.load_model('ourmodel.h5')
y_pred = saved_model.predict(x_test)
print(y_pred[8])
predval = 105
count = 0
for i in range(len(y_pred)):
if np.argmax(y_pred[i]) == y_test[i]:
count +=1
accuracy = count/len(y_pred)
print(accuracy)
# print(np.argmax(y_pred[predval]))
# print(y_test[predval])
| source_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
#为了解决一些MacOS的系统问题。
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
os.environ['KMP_DUPLICATE_LIB_OK']='True'
# +
'''
循环神经网络的TensorFlow实践代码。
'''
from tensorflow.keras import models, layers, losses, optimizers
#设置超参数。
INPUT_UNITS = 56
TIME_STEPS = 14
HIDDEN_SIZE = 256
NUM_CLASSES = 10
EPOCHS = 5
BATCH_SIZE = 64
LEARNING_RATE = 1e-3
#初始化循环神经网络模型。
model = models.Sequential()
model.add(layers.LSTM(HIDDEN_SIZE))
model.add(layers.Dense(NUM_CLASSES))
#设定神经网络的损失函数、优化方式,以及评估方法。
model.compile(optimizer=optimizers.Adam(LEARNING_RATE),
loss=losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# +
import pandas as pd
#使用pandas,读取fashion_mnist的训练和测试数据文件。
train_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_train.csv')
test_data = pd.read_csv('../datasets/fashion_mnist/fashion_mnist_test.csv')
#从训练数据中,拆解出训练特征和类别标签。
X_train = train_data[train_data.columns[1:]]
y_train = train_data['label']
#从测试数据中,拆解出测试特征和类别标签。
X_test = test_data[train_data.columns[1:]]
y_test = test_data['label']
# +
from sklearn.preprocessing import StandardScaler
#初始化数据标准化处理器。
ss = StandardScaler()
#标准化训练数据特征。
X_train = ss.fit_transform(X_train)
#标准化测试数据特征。
X_test = ss.transform(X_test)
# +
X_train = X_train.reshape([-1, TIME_STEPS, INPUT_UNITS])
#使用fashion_mnist的训练集数据训练网络模型。
model.fit(X_train, y_train.values, batch_size=BATCH_SIZE, epochs=EPOCHS, verbose=1)
# +
X_test = X_test.reshape([-1, TIME_STEPS, INPUT_UNITS])
#使用fashion_mnist的测试集数据评估网络模型的效果。
result = model.evaluate(X_test, y_test.values, verbose=0)
print('循环神经网络(TensorFlow版本)在fashion_mnist测试集上的准确率为: %.2f%%。' %(result[1] * 100))
| Chapter_6/Section_6.4.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 12807, "status": "ok", "timestamp": 1552773429425, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="ZZG4BqkENEyd" outputId="0b8bd808-eba5-4de2-9c01-64058b721fba"
# Taken from
# https://stackoverflow.com/questions/48750199/google-colaboratory-misleading-information-about-its-gpu-only-5-ram-available
# memory footprint support libraries/code
# !ln -sf /opt/bin/nvidia-smi /usr/bin/nvidia-smi
# !pip install gputil
# !pip install psutil
# !pip install humanize
import psutil
import humanize
import os
import GPUtil as GPU
GPUs = GPU.getGPUs()
# Colab only provides one GPU and it is not always guaranteed
gpu = GPUs[0]
def printm():
process = psutil.Process(os.getpid())
print("RAM Free: " + humanize.naturalsize( psutil.virtual_memory().available ), " | Proc size: " + humanize.naturalsize( process.memory_info().rss))
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 2475, "status": "ok", "timestamp": 1552773429426, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="BBvIvBoyg68g" outputId="64e90310-ddee-4cb4-a672-41a831cc0503"
printm()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 4124, "status": "ok", "timestamp": 1552773455053, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="yV1m-9ZGuKGj" outputId="32fde251-f5a5-4b68-9b12-9e42974bffae"
# Clone repo
# !git clone https://github.com/MatchLab-Imperial/keras_triplet_descriptor
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 978, "status": "ok", "timestamp": 1552773459405, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="pyZSqhZ5LACT" outputId="ce0a5f2e-73e9-43f0-f80b-41950680731a"
# Change directory
# %cd keras_triplet_descriptor
# + colab={"base_uri": "https://localhost:8080/", "height": 462} colab_type="code" executionInfo={"elapsed": 190209, "status": "ok", "timestamp": 1552773652267, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="307CBCL-FjX4" outputId="9b10a27f-fe9c-4492-a089-784596f08ac0"
# Download data
# !wget -O hpatches_data.zip https://imperialcollegelondon.box.com/shared/static/ah40eq7cxpwq4a6l4f62efzdyt8rm3ha.zip
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 249292, "status": "ok", "timestamp": 1552773724185, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="36mBTFvPCxY9" outputId="b0b44eeb-fe9c-4c11-e2e1-8a67725abd68"
# Extract data
# !unzip -q ./hpatches_data.zip
# !rm ./hpatches_data.zip
# + colab={} colab_type="code" id="o0KYfe-at9KN"
import sys
import json
import os
import glob
import time
import tensorflow as tf
import numpy as np
import cv2
import random
import keras
from keras import backend as K
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda, Reshape
from keras.layers import Conv2D, MaxPooling2D, BatchNormalization, Conv2DTranspose
from keras.layers import Input, UpSampling2D, concatenate
from read_data import HPatches, DataGeneratorDesc, hpatches_sequence_folder, DenoiseHPatches, tps
from utils import generate_desc_csv, plot_denoise, plot_triplet
# + colab={} colab_type="code" id="NXL31ez-AT5h"
random.seed(1234)
np.random.seed(1234)
tf.set_random_seed(1234)
# + colab={} colab_type="code" id="ABKDHB9RApZk"
hpatches_dir = './hpatches'
splits_path = 'splits.json'
splits_json = json.load(open(splits_path, 'rb'))
split = splits_json['a']
train_fnames = split['train']
test_fnames = split['test']
seqs = glob.glob(hpatches_dir+'/*')
seqs = [os.path.abspath(p) for p in seqs]
# -
seqs_train = list(filter(lambda x: x.split('\\')[-1] in train_fnames, seqs))
seqs_test = list(filter(lambda x: x.split('\\')[-1] in split['test'], seqs))
len(seqs)
len(seqs_train)
len(seqs_test)
# + [markdown] colab_type="text" id="qeWik0vMEtuC"
# ## Models and loss
# + colab={} colab_type="code" id="W6QbkHnbuIUD"
def get_denoise_model(shape, do = 0, activate = 'selu'):
inputs = Input(shape)
## Encoder starts
conv1 = Conv2D(16, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(inputs)
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1)
## Bottleneck
conv2 = Conv2D(32, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(pool1)
## Now the decoder starts
up3 = Conv2D(64, 2, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(UpSampling2D(size = (2,2))(conv2))
merge3 = concatenate([conv1,up3], axis = -1)
conv3 = Conv2D(64, 3, activation = 'relu', padding = 'same', kernel_initializer = 'he_normal')(merge3)
conv4 = Conv2D(1, 3, padding = 'same')(conv3)
shallow_net = Model(inputs = inputs, outputs = conv4)
return shallow_net
def get_descriptor_model(shape, activate= 'relu'):
'''Architecture copies HardNet architecture'''
init_weights = keras.initializers.he_normal()
descriptor_model = Sequential()
descriptor_model.add(Conv2D(32, 3, padding='same', input_shape=shape, use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Conv2D(32, 3, padding='same', use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Conv2D(64, 3, padding='same', strides=2, use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Conv2D(64, 3, padding='same', use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Conv2D(128, 3, padding='same', strides=2, use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Conv2D(128, 3, padding='same', use_bias = True, kernel_initializer=init_weights))
descriptor_model.add(BatchNormalization(axis = -1))
descriptor_model.add(Activation(activate))
descriptor_model.add(Dropout(0.3))
descriptor_model.add(Conv2D(128, 8, padding='valid', use_bias = True, kernel_initializer=init_weights))
# Final descriptor reshape
descriptor_model.add(Reshape((128,)))
return descriptor_model
def triplet_loss(x):
output_dim = 128
a, p, n = x
_alpha = 1.0
positive_distance = K.mean(K.square(a - p), axis=-1)
negative_distance = K.mean(K.square(a - n), axis=-1)
return K.expand_dims(K.maximum(0.0, positive_distance - negative_distance + _alpha), axis = 1)
# + [markdown] colab_type="text" id="RlS5zcV7EJgp"
# ## Denoising Image Patches
#
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" executionInfo={"elapsed": 1293, "status": "ok", "timestamp": 1552773922597, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "15717585103977390251"}, "user_tz": 0} id="-eUSba93Dttj" outputId="9888dc14-c015-469b-f13c-d1576f667cbb"
from keras.layers import LeakyReLU
shape = (32, 32, 1)
denoise_model = keras.models.load_model('./denoise_base.h5')
# -
# ## Vary Learning Rate
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="DVmDZIRTHPDa" outputId="94bda2ad-e40e-4c44-e74f-be30d956eed6"
from keras.layers import Lambda
shape = (32, 32, 1)
xa = Input(shape=shape, name='a')
xp = Input(shape=shape, name='p')
xn = Input(shape=shape, name='n')
descriptor_model = get_descriptor_model( shape)
ea = descriptor_model(xa)
ep = descriptor_model(xp)
en = descriptor_model(xn)
loss = Lambda(triplet_loss)([ea, ep, en])
sgd1 = keras.optimizers.SGD(lr=0.00001, momentum=0.9, nesterov=True)
sgd2 = keras.optimizers.SGD(lr=0.0001, momentum=0.9, nesterov=True)
sgd3 = keras.optimizers.SGD(lr=0.001, momentum=0.9, nesterov=True)
sgd4 = keras.optimizers.SGD(lr=0.01, momentum=0.9, nesterov=True)
sgd5 = keras.optimizers.SGD(lr=0.1, momentum=0.9, nesterov=True)
descriptor_model_trip_sgd1 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd2 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd3 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd4 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd5 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd1.compile(loss='mean_absolute_error', optimizer=sgd1)
descriptor_model_trip_sgd2.compile(loss='mean_absolute_error', optimizer=sgd2)
descriptor_model_trip_sgd3.compile(loss='mean_absolute_error', optimizer=sgd3)
descriptor_model_trip_sgd4.compile(loss='mean_absolute_error', optimizer=sgd4)
descriptor_model_trip_sgd5.compile(loss='mean_absolute_error', optimizer=sgd5)
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="YIR1cH4fDwKj" outputId="b17c0b36-e90e-4df0-e969-0ab13136786d"
### Descriptor loading and training
# Loading images
hPatches = HPatches(train_fnames=train_fnames, test_fnames=test_fnames,
denoise_model=denoise_model, use_clean=False)
# Creating training generator
training_generator = DataGeneratorDesc(*hPatches.read_image_file(hpatches_dir, train=1), num_triplets=10000)
# Creating validation generator
val_generator = DataGeneratorDesc(*hPatches.read_image_file(hpatches_dir, train=0), num_triplets=10000)
# + colab={"base_uri": "https://localhost:8080/", "height": 183} colab_type="code" id="3RQmOMU92csu" outputId="648c64db-a797-4bb6-c7de-5eb45d27da25"
plot_triplet(training_generator)
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="QPyc8as42WTQ" outputId="69ea09ee-8198-42d5-a654-2bd3e339e5b2"
#epochs = 1
### As with the denoising model, we use a loop to save for each epoch
## #the weights in an external website in case colab stops.
### reset, so e.g. calling 5 times fit(epochs=1) behave as fit(epochs=5)
### If you have a model saved from a previous training session
### Load it in the next line
# descriptor_model_trip.set_weights(keras.models.load_model('./descriptor.h5').get_weights())
# descriptor_model_trip.optimizer = keras.models.load_model('./descriptor.h5').optimizer
#for e in range(epochs):
descriptor_history_sgd1 = descriptor_model_trip_sgd1.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd2 = descriptor_model_trip_sgd2.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd3 = descriptor_model_trip_sgd3.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd4 = descriptor_model_trip_sgd4.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd5 = descriptor_model_trip_sgd5.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
# -
# ## Plot Losses
# + colab={} colab_type="code" id="XFA_8uN4Eb3B"
import matplotlib.pyplot as plt
# +
def plot_history(history, history2, history3, history4, history5, metric = None):
# Plots the loss history of training and validation (if existing)
# and a given metric
if metric != None:
fig, axes = plt.subplots(2,1, figsize=(8, 10))
#axes[0].plot(history.history[metric])
#axes[0].plot(history2.history[metric])
#axes[0].plot(history3.history[metric])
#axes[0].plot(history4.history[metric])
#axes[0].plot(history5.history[metric])
#axes[0].plot(history6.history[metric])
try:
#axes[0].plot(history.history['val_'+metric])
#axes[0].plot(history2.history['val2_'+metric])
#axes[0].plot(history3.history['val3_'+metric])
axes[0].legend(['lr=1e-5', 'lr=1e-4', 'lr=1e-3', 'lr=1e-2', 'lr=1e-1'], loc='upper right')
except:
pass
axes[0].set_title('MAE Vs. No of Epochs for Various Learning Rates')
axes[0].set_ylabel('Mean Absolute Error')
axes[0].set_xlabel('Epoch')
fig.subplots_adjust(hspace=0.5)
axes[1].plot(history.history['loss'])
axes[1].plot(history2.history['loss'])
axes[1].plot(history3.history['loss'])
axes[1].plot(history4.history['loss'])
axes[1].plot(history5.history['loss'])
try:
#axes[1].plot(history.history['val_loss'])
axes[1].legend(['lr=1e-5', 'lr=1e-4', 'lr=1e-3', 'lr=1e-2', 'lr=1e-1'], loc='upper right')
except:
pass
axes[1].set_title('MAE Vs. No of Epochs for Various Learning Rates')
axes[1].set_ylabel('Mean Absolute Error')
axes[1].set_xlabel('Epoch')
else:
plt.plot(history.history['loss'])
try:
plt.plot(history.history['val_loss'])
plt.legend(['Train', 'Val'])
except:
pass
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plot_history(descriptor_history_sgd1, descriptor_history_sgd2, descriptor_history_sgd3, descriptor_history_sgd4, descriptor_history_sgd5, 'mean_absolute_error')
# +
def plot_val_history(history, history2, history3, history4, history5, metric = None):
# Plots the loss history of training and validation (if existing)
# and a given metric
if metric != None:
fig, axes = plt.subplots(2,1, figsize=(8, 10))
#axes[0].plot(history.history[metric])
#axes[0].plot(history2.history[metric])
#axes[0].plot(history3.history[metric])
try:
#axes[0].plot(history.history['val_'+metric])
#axes[0].plot(history2.history['val_'+metric])
#axes[0].plot(history3.history['val_'+metric])
#axes[0].plot(history4.history['val_'+metric])
#axes[0].plot(history5.history['val_'+metric])
#axes[0].plot(history6.history['val_'+metric])
axes[0].legend(['lr=1e-5', 'lr=1e-4', 'lr=1e-3', 'lr=1e-2', 'lr=1e-1'], loc='upper right')
except:
pass
axes[0].set_title('Validation Loss Vs. No of Epochs for Various Learning Rates')
axes[0].set_ylabel('Validation Loss')
axes[0].set_xlabel('Epoch')
fig.subplots_adjust(hspace=0.5)
#axes[1].plot(history.history['loss'])
#axes[1].plot(history2.history['loss'])
#axes[1].plot(history3.history['loss'])
try:
axes[1].plot(history.history['val_loss'])
axes[1].plot(history2.history['val_loss'])
axes[1].plot(history3.history['val_loss'])
axes[1].plot(history4.history['val_loss'])
axes[1].plot(history5.history['val_loss'])
#axes[1].plot(history6.history['val_loss'])
axes[1].legend(['lr=1e-5', 'lr=1e-4', 'lr=1e-3', 'lr=1e-2', 'lr=1e-1'], loc='upper right')
except:
pass
axes[1].set_title('Validation Loss Vs. No of Epochs for Various Learning Rates')
axes[1].set_ylabel('Validation Loss')
axes[1].set_xlabel('Epoch')
else:
plt.plot(history.history['loss'])
try:
plt.plot(history.history['val_loss'])
plt.legend(['Train', 'Val'])
except:
pass
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plot_val_history(descriptor_history_sgd1, descriptor_history_sgd2, descriptor_history_sgd3, descriptor_history_sgd4, descriptor_history_sgd5, 'mean_absolute_error')
# -
# ## <NAME>
# +
sgd1 = keras.optimizers.SGD(lr=0.1, momentum=0.9, nesterov=True)
sgd2 = keras.optimizers.SGD(lr=0.1, momentum=0.8, nesterov=True)
sgd3 = keras.optimizers.SGD(lr=0.1, momentum=0.7, nesterov=True)
sgd4 = keras.optimizers.SGD(lr=0.1, momentum=0.6, nesterov=True)
sgd5 = keras.optimizers.SGD(lr=0.1, momentum=0.5, nesterov=True)
descriptor_model_trip_sgd1 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd2 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd3 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd4 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd5 = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip_sgd1.compile(loss='mean_absolute_error', optimizer=sgd1)
descriptor_model_trip_sgd2.compile(loss='mean_absolute_error', optimizer=sgd2)
descriptor_model_trip_sgd3.compile(loss='mean_absolute_error', optimizer=sgd3)
descriptor_model_trip_sgd4.compile(loss='mean_absolute_error', optimizer=sgd4)
descriptor_model_trip_sgd5.compile(loss='mean_absolute_error', optimizer=sgd5)
# +
#epochs = 1
### As with the denoising model, we use a loop to save for each epoch
## #the weights in an external website in case colab stops.
### reset, so e.g. calling 5 times fit(epochs=1) behave as fit(epochs=5)
### If you have a model saved from a previous training session
### Load it in the next line
# descriptor_model_trip.set_weights(keras.models.load_model('./descriptor.h5').get_weights())
# descriptor_model_trip.optimizer = keras.models.load_model('./descriptor.h5').optimizer
#for e in range(epochs):
descriptor_history_sgd1 = descriptor_model_trip_sgd1.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd2 = descriptor_model_trip_sgd2.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd3 = descriptor_model_trip_sgd3.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd4 = descriptor_model_trip_sgd4.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
descriptor_history_sgd5 = descriptor_model_trip_sgd5.fit_generator(generator=training_generator, epochs=5, verbose=1, validation_data=val_generator)
### Saves optimizer and weights
#descriptor_model_trip.save('descriptor.h5')
### Uploads files to external hosting
# #!curl -F "file=@descriptor.h5" https://file.io
# -
# ## Plot Losses
# +
def plot_history(history, history2, history3, history4, history5, metric = None):
# Plots the loss history of training and validation (if existing)
# and a given metric
if metric != None:
fig, axes = plt.subplots(2,1, figsize=(8, 10))
#axes[0].plot(history.history[metric])
#axes[0].plot(history2.history[metric])
#axes[0].plot(history3.history[metric])
#axes[0].plot(history4.history[metric])
#axes[0].plot(history5.history[metric])
try:
#axes[0].plot(history.history['val_'+metric])
#axes[0].plot(history2.history['val2_'+metric])
#axes[0].plot(history3.history['val3_'+metric])
axes[0].legend(['0.9', '0.8', '0.7', '0.6', '0.5'], loc='best')
except:
pass
axes[0].set_title('MAE Vs. No of Epochs for Various Momentum Values')
axes[0].set_ylabel('Mean Absolute Error')
axes[0].set_xlabel('Epoch')
fig.subplots_adjust(hspace=0.5)
axes[1].plot(history.history['loss'])
axes[1].plot(history2.history['loss'])
axes[1].plot(history3.history['loss'])
axes[1].plot(history4.history['loss'])
axes[1].plot(history5.history['loss'])
try:
#axes[1].plot(history.history['val_loss'])
axes[1].legend(['0.9', '0.8', '0.7', '0.6', '0.5'], loc='best')
except:
pass
axes[1].set_title('MAE Vs. No of Epochs for Various Momentum Values')
axes[1].set_ylabel('Mean Absolute Error')
axes[1].set_xlabel('Epoch')
else:
plt.plot(history.history['loss'])
try:
plt.plot(history.history['val_loss'])
plt.legend(['Train', 'Val'])
except:
pass
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plot_history(descriptor_history_sgd1, descriptor_history_sgd2, descriptor_history_sgd3, descriptor_history_sgd4, descriptor_history_sgd5, 'mean_absolute_error')
# +
def plot_val_history(history, history2, history3, history4, history5, metric = None):
# Plots the loss history of training and validation (if existing)
# and a given metric
if metric != None:
fig, axes = plt.subplots(2,1, figsize=(8, 10))
#axes[0].plot(history.history[metric])
#axes[0].plot(history2.history[metric])
#axes[0].plot(history3.history[metric])
try:
#axes[0].plot(history.history['val_'+metric])
#axes[0].plot(history2.history['val_'+metric])
#axes[0].plot(history3.history['val_'+metric])
#axes[0].plot(history4.history['val_'+metric])
#axes[0].plot(history5.history['val_'+metric])
axes[0].legend(['0.9', '0.8', '0.7', '0.6', '0.5'], loc='best')
except:
pass
axes[0].set_title('Validation Loss Vs. No of Epochs for for Various Momentum Values')
axes[0].set_ylabel('Validation Loss')
axes[0].set_xlabel('Epoch')
fig.subplots_adjust(hspace=0.5)
#axes[1].plot(history.history['loss'])
#axes[1].plot(history2.history['loss'])
#axes[1].plot(history3.history['loss'])
try:
axes[1].plot(history.history['val_loss'])
axes[1].plot(history2.history['val_loss'])
axes[1].plot(history3.history['val_loss'])
axes[1].plot(history4.history['val_loss'])
axes[1].plot(history5.history['val_loss'])
axes[1].legend(['0.9', '0.8', '0.7', '0.6', '0.5'], loc='best')
except:
pass
axes[1].set_title('Validation Loss Vs. No of Epochs for Various Momentum Values')
axes[1].set_ylabel('Validation Loss')
axes[1].set_xlabel('Epoch')
else:
plt.plot(history.history['loss'])
try:
plt.plot(history.history['val_loss'])
plt.legend(['Train', 'Val'])
except:
pass
plt.title('Model Loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plot_val_history(descriptor_history_sgd1, descriptor_history_sgd2, descriptor_history_sgd3, descriptor_history_sgd4, descriptor_history_sgd5, 'mean_absolute_error')
# -
# ## Save Baseline Model
# +
sgd1 = keras.optimizers.SGD(lr=0.1, momentum=0.7, nesterov=True)
descriptor_model_trip = Model(inputs=[xa, xp, xn], outputs=loss)
descriptor_model_trip.compile(loss='mean_absolute_error', optimizer=sgd1)
# +
#epochs = 1
### As with the denoising model, we use a loop to save for each epoch
## #the weights in an external website in case colab stops.
### reset, so e.g. calling 5 times fit(epochs=1) behave as fit(epochs=5)
### If you have a model saved from a previous training session
### Load it in the next line
# descriptor_model_trip.set_weights(keras.models.load_model('./descriptor.h5').get_weights())
# descriptor_model_trip.optimizer = keras.models.load_model('./descriptor.h5').optimizer
#for e in range(epochs):
descriptor_history = descriptor_model_trip.fit_generator(generator=training_generator, epochs=20, verbose=1, validation_data=val_generator)
descriptor_model_trip.save('descriptor_base.h5')
### Saves optimizer and weights
#descriptor_model_trip.save('descriptor.h5')
### Uploads files to external hosting
# #!curl -F "file=@descriptor.h5" https://file.io
| Baseline Models/Baseline L2-Net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 重用 TF model
# 训练原模型
# +
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
mnist = input_data.read_data_sets("data/")
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_hidden3 = 50
n_hidden4 = 50
n_hidden5 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int64, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1", reuse=tf.AUTO_REUSE)
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2", reuse=tf.AUTO_REUSE)
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3", reuse=tf.AUTO_REUSE)
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4", reuse=tf.AUTO_REUSE)
hidden5 = tf.layers.dense(hidden4, n_hidden5, activation=tf.nn.relu, name="hidden5", reuse=tf.AUTO_REUSE)
logits = tf.layers.dense(hidden5, n_outputs, name="outputs", reuse=tf.AUTO_REUSE)
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
threshold = 1.0
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var)
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 10
batch_size = 100
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(n_epochs):
for iteration in range(mnist.train.num_examples // batch_size):
X_batch, y_batch = mnist.train.next_batch(batch_size)
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: mnist.test.images,
y: mnist.test.labels})
print(epoch, "Test accuracy:", accuracy_val)
save_path = saver.save(sess, "model/minist_model.ckpt")
# -
| chapter 11 - save model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 10: Data Preparation & CNN Transfer Learning (Optional)
#
# ### [<NAME>](http://staffwww.dcs.shef.ac.uk/people/H.Lu/) @ [COM4509/6509 MLAI2019](https://github.com/maalvarezl/MLAI)
#
# ### This lab is provided as *optional* materials for those interested. There will be no assessment on this material though I will be happy to answer questions in lab or discussion board.
#
# ### More materials on deep learning will be updated at my [SimplyDeep](https://github.com/haipinglu/SimplyDeep/) project in future.
# ## Objective
#
# * To load and preprocess/augment data from a non trivial dataset.
# * To make use of pretrained models for image classification
#
# **Suggested reading**:
# * [PyTorch tutorial on *Writing Custom Datasets, DataLoaders and Transforms*](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html)
# * [PyTorch tutorial on *Transfer Learning for Computer Vision Tutorial*](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html)
# * [Data preparation - Wikipedia](https://en.wikipedia.org/wiki/Data_preparation)
# * [Transfer learning - Stanford CNN course](https://cs231n.github.io/transfer-learning/)
#
# **This notebook is based on PyTorch tutorials on [*Writing Custom Datasets, DataLoaders and Transforms*](https://pytorch.org/tutorials/beginner/data_loading_tutorial.html) and [*Transfer Learning for Computer Vision Tutorial*](https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html) by `<NAME> <https://chsasank.github.io>` under the BSD license.**
# ## Why
#
# **Data preparation** is important when applying machine learning to real-world problems. Mostly, data are not in a format that can be directly analysed by machine learning algorithms, and we need to preprocess the data to such a format. Often, it is also helpful to inspect data to know it (well) before machine learning, to avoid unpleasant surprises.
#
# **Transfer learning** can help improve the performance on a particular learning problem by leveraging knowledge gained from a different but related problem. This can also alleviate the demanded resources and accelerate development.
#
#
# %matplotlib inline
# ## 1. Data Preparation
#
# PyTorch provides many tools to make data loading easy and hopefully, to make your code more readable. With conda, we should have ``scikit-image`` (for image io and transforms) and ``pandas`` (for easier csv parsing) already.
# +
from __future__ import print_function, division
import os
import torch
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
plt.ion() # interactive mode
# -
# ### 1.1 The facial pose dataset
# The dataset we are going to deal with is that of facial pose, with 68 different landmark points are annotated for each face.
#
# **Download** the dataset from [here](https://download.pytorch.org/tutorial/faces.zip) and unzip the images to a directory named `data/faces/`.
#
# **Landmarks**: the dataset comes with a csv file `face_landmarks.csv` with each row containing the image filename followed by the $x$ and $y$ coordinates of the 68 landmarks.
landmarks_frame = pd.read_csv('data/faces/face_landmarks.csv')
landmarks_frame.head()
# **Inspection**: let us take a look at the third image (index $n=2$) `10comm-decarlo.jpg`.
# +
n = 2 #The index
img_name = landmarks_frame.iloc[n, 0]
landmarks = landmarks_frame.iloc[n, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
print('Image name: {}'.format(img_name))
print('Landmarks shape: {}'.format(landmarks.shape))
print('First 4 Landmarks: {}'.format(landmarks[:4]))
# -
# Let's write a simple helper function to show an image and its landmarks
# and use it to show a sample.
#
#
#
# +
def show_landmarks(image, landmarks):
"""Show image with landmarks"""
plt.imshow(image)
plt.scatter(landmarks[:, 0], landmarks[:, 1], s=10, marker='.', c='r')
plt.pause(0.001) # pause a bit so that plots are updated
plt.figure()
show_landmarks(io.imread(os.path.join('data/faces/', img_name)),landmarks)
plt.show()
# -
# ### Define a custom torch Dataset
#
# ``torch.utils.data.Dataset`` is an abstract class representing a dataset. Our custom dataset should inherit ``Dataset`` and **override** the following methods:
#
# - ``__len__`` so that ``len(dataset)`` returns the size of the dataset.
# - ``__getitem__`` to support the indexing such that ``dataset[i]`` can
# be used to get $i$\ th sample
#
# Let's create a dataset class for our face landmarks dataset. We will read the csv in ``__init__`` but leave the reading of images to ``__getitem__``. This is memory efficient because all the images are not stored in the memory at once but read as required.
#
# Sample of our dataset will be a dict ``{'image': image, 'landmarks': landmarks}``. Our dataset will take an
# optional argument ``transform`` so that any required processing can be applied on the sample. We will see the usefulness of ``transform`` below.
class FaceLandmarksDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, csv_file, root_dir, transform=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied on a sample.
"""
self.landmarks_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.landmarks_frame)
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
img_name = os.path.join(self.root_dir, self.landmarks_frame.iloc[idx, 0])
image = io.imread(img_name)
landmarks = self.landmarks_frame.iloc[idx, 1:]
landmarks = np.array([landmarks])
landmarks = landmarks.astype('float').reshape(-1, 2)
sample = {'image': image, 'landmarks': landmarks}
if self.transform:
sample = self.transform(sample)
return sample
# ### 1.2 Test our custom Dataset
# Let's instantiate this class and iterate through the data samples. We will print the sizes of first $N$ samples and show their landmarks.
#
#
#
# +
face_dataset = FaceLandmarksDataset(csv_file='data/faces/face_landmarks.csv', root_dir='data/faces/')
fig = plt.figure()
N=3 #number to show
for i in range(len(face_dataset)):
sample = face_dataset[i]
print(i, sample['image'].shape, sample['landmarks'].shape)
ax = plt.subplot(1, N, i + 1)
plt.tight_layout()
ax.set_title('Sample #{}'.format(i))
ax.axis('off')
show_landmarks(**sample)
if i == N-1:
#plt.show()
break
# -
# ### 1.3 Transforms
#
# #### Define transforms
# One issue we can see from the above is that the samples are not of the
# same size. Most neural networks expect the images of a fixed size.
# Therefore, we will need to write some prepocessing code.
# Let's create three transforms:
#
# - ``Rescale``: to scale the image
# - ``RandomCrop``: to crop from image randomly. This is data
# augmentation.
# - ``ToTensor``: to convert the numpy images to torch images (we need to
# swap axes).
#
# We will write them as **callable classes** instead of simple functions so
# that parameters of the transform need not be passed everytime it's
# called. For this, we just need to implement ``__call__`` method and
# if required, ``__init__`` method. We can then use a transform like this:
#
# ```python
# tsfm = Transform(params)
# transformed_sample = tsfm(sample)
# ```
# Observe below how these transforms had to be applied **both** on the image and
# landmarks.
#
#
#
# +
class Rescale(object):
"""Rescale the image in a sample to a given size.
Args:
output_size (tuple or int): Desired output size. If tuple, output is matched to output_size.
If int, smaller of image edges is matched to output_size keeping aspect ratio the same.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = transform.resize(image, (new_h, new_w))
# h and w are swapped for landmarks because for images,
# x and y axes are axis 1 and 0 respectively
landmarks = landmarks * [new_w / w, new_h / h]
return {'image': img, 'landmarks': landmarks}
class RandomCrop(object):
"""Crop randomly the image in a sample.
Args:
output_size (tuple or int): Desired output size. If int, square crop is made.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
landmarks = landmarks - [left, top]
return {'image': image, 'landmarks': landmarks}
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image),
'landmarks': torch.from_numpy(landmarks)}
# -
# #### Compose transforms and apply them to a sample
#
# Now, we apply the transforms on a sample.
#
# Let's say we want to rescale the shorter side of the image to 256 and
# then randomly crop a square of size 224 from it. i.e, we want to compose
# ``Rescale`` and ``RandomCrop`` transforms.
# ``torchvision.transforms.Compose`` is a simple callable class which allows us
# to do this.
#
#
#
# +
scale = Rescale(256)
crop = RandomCrop(128)
composed = transforms.Compose([Rescale(256),
RandomCrop(224)])
n=65 #Choose image 65 to apply the transform
# Apply each of the above transforms on sample.
#fig = plt.figure()
sample = face_dataset[n]
for i, tsfrm in enumerate([scale, crop, composed]):
transformed_sample = tsfrm(sample)
print(i)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tsfrm).__name__)
show_landmarks(**transformed_sample)
#plt.show()
# -
# ### 1.4 Iterating through the dataset
# Let's put this all together to create a dataset with composed
# transforms. To summarize, every time this dataset is sampled:
#
# - An image is read from the file on the fly
# - Transforms are applied on the read image
# - Since one of the transforms is random, data is augmentated on
# sampling
#
# We can iterate over the created dataset with a ``for i in range``
# loop as before. Here, we rescale all images to the same size of 256x256 and then crop a 224x224 subimage from each.
# +
transformed_dataset = FaceLandmarksDataset(csv_file='data/faces/face_landmarks.csv',
root_dir='data/faces/',
transform=transforms.Compose([
Rescale(256),
RandomCrop(224),
ToTensor()
]))
for i in range(len(transformed_dataset)):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['landmarks'].size())
if i == 3:
break
# -
# However, we are losing a lot of features by using a simple ``for`` loop to
# iterate over the data. In particular, we are missing out on:
#
# - Batching the data
# - Shuffling the data
# - Load the data in parallel using ``multiprocessing`` workers.
#
# ``torch.utils.data.DataLoader`` is an iterator which provides all these
# features. Parameters used below should be clear. One parameter of
# interest is ``collate_fn``. You can specify how exactly the samples need
# to be batched using ``collate_fn``. However, default collate should work
# fine for most use cases. Check out the [DataLoader API here](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader).
#
# **`num_workers`** specifies the number of workers for [multi-process data loading](https://pytorch.org/docs/stable/data.html#single-and-multi-process-data-loading). On (my) Windows, if `num_workers` is set to a positive number, there will be a `BrokenPipeError`. On (my) Linux, there is no such probem, e.g., setting `num_workers=4` works fine. I am not sure about Mac.
# +
dataloader = DataLoader(transformed_dataset, batch_size=4,
shuffle=True, num_workers=0)
# Helper function to show a batch
def show_landmarks_batch(sample_batched):
"""Show image with landmarks for a batch of samples."""
images_batch, landmarks_batch = \
sample_batched['image'], sample_batched['landmarks']
batch_size = len(images_batch)
im_size = images_batch.size(2)
grid_border_size = 2
grid = utils.make_grid(images_batch)
plt.imshow(grid.numpy().transpose((1, 2, 0)))
for i in range(batch_size):
plt.scatter(landmarks_batch[i, :, 0].numpy() + i * im_size + (i + 1) * grid_border_size,
landmarks_batch[i, :, 1].numpy() + grid_border_size,
s=10, marker='.', c='r')
plt.title('Batch from dataloader')
for i_batch, sample_batched in enumerate(dataloader):
print(i_batch, sample_batched['image'].size(),
sample_batched['landmarks'].size())
# observe 4th batch and stop.
if i_batch == 3:
plt.figure()
show_landmarks_batch(sample_batched)
plt.axis('off')
plt.ioff()
plt.show()
break
# -
#
# ## 2. CNN Transfer Learning in Computer Vision
# Next, we will train a convolutional neural network (CNN) for image classification using transfer learning. As pointed out in the [cs231n notes](https://cs231n.github.io/transfer-learning)
# > In practice, very few people train an entire Convolutional Network
# from scratch (with random initialization), because it is relatively
# rare to have a dataset of sufficient size. Instead, it is common to
# pretrain a ConvNet on a very large dataset (e.g. ImageNet, which
# contains 1.2 million images with 1000 categories), and then use the
# ConvNet either as an initialization or a fixed feature extractor for
# the task of interest.
#
# These two major transfer learning scenarios look as follows:
#
# - **Finetuning the convnet**: Instead of random initializaion, we
# initialize the network with a pretrained network, like the one that is
# trained on imagenet 1000 dataset. Rest of the training looks as
# usual.
# - **ConvNet as fixed feature extractor**: Here, we will freeze the weights
# for all of the network except that of the final fully connected
# layer. This last fully connected layer is replaced with a new one
# with random weights and only this layer is trained.
#
#
#
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import torchvision
from torchvision import datasets, models
import time
import copy
# ### 2.1 Load the `ants` & `bees` Data
#
# We will train a model to classify **ants** and **bees**. We have about 120 training images each for ants and bees. There are 75 validation images for each class. Usually, this is a very small dataset to generalize upon, if trained from scratch. Since we are using transfer learning, we should be able to generalize reasonably well. This dataset is a very small subset of [imagenet](https://en.wikipedia.org/wiki/ImageNet). We will use torchvision and torch.utils.data packages for loading the data.
#
# **Download** the data from [here](https://download.pytorch.org/tutorial/hymenoptera_data.zip) to our `data` directory and extract it there.
#
# #### Compose the transforms
#
# Check out the [transforms API here](https://pytorch.org/docs/stable/torchvision/transforms.html) for avaiable transforms in PyTorch.
#
# **Note**: the numbers `[0.485, 0.456, 0.406]` and `[0.229, 0.224, 0.225]` in `Normalize` are from the [imagenet](https://en.wikipedia.org/wiki/ImageNet). See discussions [here](https://stackoverflow.com/questions/58151507/why-pytorch-officially-use-mean-0-485-0-456-0-406-and-std-0-229-0-224-0-2) and also [here ](https://github.com/jacobgil/pytorch-grad-cam/issues/6).
#
# **ImageFolder** is a method of `torchvision.datasets`. See the [`ImageFolder` API here](https://pytorch.org/docs/stable/torchvision/datasets.html#imagefolder).
# It assumes that images are organized in the following way: ::
#
# root/ants/xxx.png
# root/ants/xxy.jpeg
# root/ants/xxz.png
# .
# .
# .
# root/bees/123.jpg
# root/bees/nsdf3.png
# root/bees/asd932_.png
#
# where 'ants', 'bees' etc. are class labels.
# +
# Data augmentation and normalization for training
# Just normalization for validation
data_transforms = {
'train': transforms.Compose([
transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
}
data_dir = 'data/hymenoptera_data'
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x), data_transforms[x])
for x in ['train', 'val']}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x], batch_size=4,
shuffle=True, num_workers=4)
for x in ['train', 'val']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val']}
class_names = image_datasets['train'].classes
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# -
# #### Inspect data by visualizing a few images
#
# Let's visualize a few training images so as to understand the data augmentations.
#
#
# +
def imshow(inp, title=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0)) #Tensor to numpy format conversion
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
# Get a batch of training data
inputs, classes = next(iter(dataloaders['train']))
# Make a grid from batch
out = torchvision.utils.make_grid(inputs)
imshow(out, title=[class_names[x] for x in classes])
# -
# ### 2.2 Training the model
#
# Now, let's write a general function to train a model. Here, we will
# illustrate:
#
# - Scheduling the learning rate
# - Saving the best model
# - Recording the computational time.
#
# In the following, parameter ``scheduler`` is an LR scheduler object from
# ``torch.optim.lr_scheduler``. Check out [its API here](https://pytorch.org/docs/stable/optim.html#how-to-adjust-learning-rate)
def train_model(model, criterion, optimizer, scheduler, num_epochs=25):
since = time.time()
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
if phase == 'train':
scheduler.step()
epoch_loss = running_loss / dataset_sizes[phase]
epoch_acc = running_corrects.double() / dataset_sizes[phase]
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, epoch_loss, epoch_acc))
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
print()
time_elapsed = time.time() - since
print('Training complete in {:.0f}m {:.0f}s'.format(time_elapsed // 60, time_elapsed % 60))
print('Best val Acc: {:4f}'.format(best_acc))
# load best model weights
model.load_state_dict(best_model_wts)
return model
# #### Visualizing the model predictions
#
# Generic function to display predictions for a few images
def visualize_model(model, num_images=6):
was_training = model.training
model.eval()
images_so_far = 0
fig = plt.figure()
with torch.no_grad():
for i, (inputs, labels) in enumerate(dataloaders['val']):
inputs = inputs.to(device)
labels = labels.to(device)
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
for j in range(inputs.size()[0]):
images_so_far += 1
ax = plt.subplot(num_images//2, 2, images_so_far)
ax.axis('off')
ax.set_title('predicted: {}'.format(class_names[preds[j]]))
imshow(inputs.cpu().data[j])
if images_so_far == num_images:
model.train(mode=was_training)
return
model.train(mode=was_training)
# ### 2.3 Finetuning the convnet
#
# Load a pretrained model and reset final fully connected layer.
# +
model_ft = models.resnet18(pretrained=True)
num_ftrs = model_ft.fc.in_features
# Here the size of each output sample is set to 2.
# Alternatively, it can be generalized to nn.Linear(num_ftrs, len(class_names)).
model_ft.fc = nn.Linear(num_ftrs, 2)
model_ft = model_ft.to(device)
criterion = nn.CrossEntropyLoss()
# Observe that all parameters are being optimized
optimizer_ft = optim.SGD(model_ft.parameters(), lr=0.001, momentum=0.9)
# Decay LR by a factor of 0.1 every 7 epochs
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_ft, step_size=7, gamma=0.1)
# -
# #### Train and evaluate
#
# We set the number of epochs small to save time.
max_epochs=5;
model_ft = train_model(model_ft, criterion, optimizer_ft, exp_lr_scheduler, num_epochs=max_epochs)
visualize_model(model_ft)
# ### 2.4 ConvNet as fixed feature extractor
#
# Here, we need to freeze all the network except the final layer. We need to set ``requires_grad == False`` to freeze the parameters so that the gradients are not computed in ``backward()``.
#
# You can read more about this in the [documentation](https://pytorch.org/docs/master/notes/autograd.html#excluding-subgraphs-from-backward)
# +
model_conv = torchvision.models.resnet18(pretrained=True)
for param in model_conv.parameters():
param.requires_grad = False
# Parameters of newly constructed modules have requires_grad=True by default
num_ftrs = model_conv.fc.in_features
model_conv.fc = nn.Linear(num_ftrs, 2)
model_conv = model_conv.to(device)
criterion = nn.CrossEntropyLoss()
# Observe that only parameters of final layer are being optimized as
# opposed to before.
optimizer_conv = optim.SGD(model_conv.fc.parameters(), lr=0.001, momentum=0.9)
# Decay LR by a factor of 0.1 every 7 epochs
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_conv, step_size=7, gamma=0.1)
# -
# #### Train and evaluate
#
# On CPU this will take about half the time compared to previous scenario. This is expected as gradients don't need to be computed for most of the network. However, forward does need to be computed.
model_conv = train_model(model_conv, criterion, optimizer_conv,
exp_lr_scheduler, num_epochs=max_epochs)
# +
visualize_model(model_conv)
plt.ioff()
plt.show()
# -
# ## 3. Exercises
#
#
# * Construct image transforms not yet available in `torchvision.transforms` and use them to compose various manipulations of images.
# * Replace the `resnet18` model with some other [pretrained models](https://pytorch.org/docs/stable/torchvision/models.html) and compare their performance.
# * Take some other adventures, e.g., how about freezing all the network except the **last two** layers?
# * Explore other `torchvision` APIs.
| lab/Lab 10 - Data prep & Transfer learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.019091, "end_time": "2021-11-08T23:58:29.611505", "exception": false, "start_time": "2021-11-08T23:58:29.592414", "status": "completed"} tags=[]
# # Introduction
#
# In the last tutorial, we learned how to select relevant data out of a DataFrame or Series. Plucking the right data out of our data representation is critical to getting work done, as we demonstrated in the exercises.
#
# However, the data does not always come out of memory in the format we want it in right out of the bat. Sometimes we have to do some more work ourselves to reformat it for the task at hand. This tutorial will cover different operations we can apply to our data to get the input "just right".
#
# **To start the exercise for this topic, please click [here](https://www.kaggle.com/kernels/fork/595524).**
#
# We'll use the Wine Magazine data for demonstration.
# + _kg_hide-input=true papermill={"duration": 1.82228, "end_time": "2021-11-08T23:58:31.448860", "exception": false, "start_time": "2021-11-08T23:58:29.626580", "status": "completed"} tags=[]
import pandas as pd
pd.set_option('max_rows', 5)
import numpy as np
reviews = pd.read_csv("../input/wine-reviews/winemag-data-130k-v2.csv", index_col=0)
# + papermill={"duration": 0.046391, "end_time": "2021-11-08T23:58:31.513029", "exception": false, "start_time": "2021-11-08T23:58:31.466638", "status": "completed"} tags=[]
reviews
# + [markdown] papermill={"duration": 0.015656, "end_time": "2021-11-08T23:58:31.544911", "exception": false, "start_time": "2021-11-08T23:58:31.529255", "status": "completed"} tags=[]
# # Summary functions
#
# Pandas provides many simple "summary functions" (not an official name) which restructure the data in some useful way. For example, consider the `describe()` method:
# + papermill={"duration": 0.035287, "end_time": "2021-11-08T23:58:31.596103", "exception": false, "start_time": "2021-11-08T23:58:31.560816", "status": "completed"} tags=[]
reviews.points.describe()
# + [markdown] papermill={"duration": 0.016412, "end_time": "2021-11-08T23:58:31.629292", "exception": false, "start_time": "2021-11-08T23:58:31.612880", "status": "completed"} tags=[]
# This method generates a high-level summary of the attributes of the given column. It is type-aware, meaning that its output changes based on the data type of the input. The output above only makes sense for numerical data; for string data here's what we get:
# + papermill={"duration": 0.058779, "end_time": "2021-11-08T23:58:31.704915", "exception": false, "start_time": "2021-11-08T23:58:31.646136", "status": "completed"} tags=[]
reviews.taster_name.describe()
# + [markdown] papermill={"duration": 0.016959, "end_time": "2021-11-08T23:58:31.740729", "exception": false, "start_time": "2021-11-08T23:58:31.723770", "status": "completed"} tags=[]
# If you want to get some particular simple summary statistic about a column in a DataFrame or a Series, there is usually a helpful pandas function that makes it happen.
#
# For example, to see the mean of the points allotted (e.g. how well an averagely rated wine does), we can use the `mean()` function:
# + papermill={"duration": 0.027076, "end_time": "2021-11-08T23:58:31.785122", "exception": false, "start_time": "2021-11-08T23:58:31.758046", "status": "completed"} tags=[]
reviews.points.mean()
# + [markdown] papermill={"duration": 0.017043, "end_time": "2021-11-08T23:58:31.819891", "exception": false, "start_time": "2021-11-08T23:58:31.802848", "status": "completed"} tags=[]
# To see a list of unique values we can use the `unique()` function:
# + papermill={"duration": 0.03973, "end_time": "2021-11-08T23:58:31.877142", "exception": false, "start_time": "2021-11-08T23:58:31.837412", "status": "completed"} tags=[]
reviews.taster_name.unique()
# + [markdown] papermill={"duration": 0.017478, "end_time": "2021-11-08T23:58:31.912654", "exception": false, "start_time": "2021-11-08T23:58:31.895176", "status": "completed"} tags=[]
# To see a list of unique values _and_ how often they occur in the dataset, we can use the `value_counts()` method:
# + papermill={"duration": 0.044701, "end_time": "2021-11-08T23:58:31.975267", "exception": false, "start_time": "2021-11-08T23:58:31.930566", "status": "completed"} tags=[]
reviews.taster_name.value_counts()
# + [markdown] papermill={"duration": 0.018645, "end_time": "2021-11-08T23:58:32.012104", "exception": false, "start_time": "2021-11-08T23:58:31.993459", "status": "completed"} tags=[]
# # Maps
#
# A **map** is a term, borrowed from mathematics, for a function that takes one set of values and "maps" them to another set of values. In data science we often have a need for creating new representations from existing data, or for transforming data from the format it is in now to the format that we want it to be in later. Maps are what handle this work, making them extremely important for getting your work done!
#
# There are two mapping methods that you will use often.
#
# [`map()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.map.html) is the first, and slightly simpler one. For example, suppose that we wanted to remean the scores the wines received to 0. We can do this as follows:
# + papermill={"duration": 0.082535, "end_time": "2021-11-08T23:58:32.114121", "exception": false, "start_time": "2021-11-08T23:58:32.031586", "status": "completed"} tags=[]
review_points_mean = reviews.points.mean()
reviews.points.map(lambda p: p - review_points_mean)
# + [markdown] papermill={"duration": 0.018689, "end_time": "2021-11-08T23:58:32.152418", "exception": false, "start_time": "2021-11-08T23:58:32.133729", "status": "completed"} tags=[]
# The function you pass to `map()` should expect a single value from the Series (a point value, in the above example), and return a transformed version of that value. `map()` returns a new Series where all the values have been transformed by your function.
#
# [`apply()`](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.apply.html) is the equivalent method if we want to transform a whole DataFrame by calling a custom method on each row.
# + papermill={"duration": 14.129845, "end_time": "2021-11-08T23:58:46.301025", "exception": false, "start_time": "2021-11-08T23:58:32.171180", "status": "completed"} tags=[]
def remean_points(row):
row.points = row.points - review_points_mean
return row
reviews.apply(remean_points, axis='columns')
# + [markdown] papermill={"duration": 0.019162, "end_time": "2021-11-08T23:58:46.340299", "exception": false, "start_time": "2021-11-08T23:58:46.321137", "status": "completed"} tags=[]
# If we had called `reviews.apply()` with `axis='index'`, then instead of passing a function to transform each row, we would need to give a function to transform each *column*.
#
# Note that `map()` and `apply()` return new, transformed Series and DataFrames, respectively. They don't modify the original data they're called on. If we look at the first row of `reviews`, we can see that it still has its original `points` value.
# + papermill={"duration": 0.035886, "end_time": "2021-11-08T23:58:46.395719", "exception": false, "start_time": "2021-11-08T23:58:46.359833", "status": "completed"} tags=[]
reviews.head(1)
# + [markdown] papermill={"duration": 0.019766, "end_time": "2021-11-08T23:58:46.435659", "exception": false, "start_time": "2021-11-08T23:58:46.415893", "status": "completed"} tags=[]
# Pandas provides many common mapping operations as built-ins. For example, here's a faster way of remeaning our points column:
# + papermill={"duration": 0.031045, "end_time": "2021-11-08T23:58:46.486756", "exception": false, "start_time": "2021-11-08T23:58:46.455711", "status": "completed"} tags=[]
review_points_mean = reviews.points.mean()
reviews.points - review_points_mean
# + [markdown] papermill={"duration": 0.020448, "end_time": "2021-11-08T23:58:46.527844", "exception": false, "start_time": "2021-11-08T23:58:46.507396", "status": "completed"} tags=[]
# In this code we are performing an operation between a lot of values on the left-hand side (everything in the Series) and a single value on the right-hand side (the mean value). Pandas looks at this expression and figures out that we must mean to subtract that mean value from every value in the dataset.
#
# Pandas will also understand what to do if we perform these operations between Series of equal length. For example, an easy way of combining country and region information in the dataset would be to do the following:
# + papermill={"duration": 0.106601, "end_time": "2021-11-08T23:58:46.655438", "exception": false, "start_time": "2021-11-08T23:58:46.548837", "status": "completed"} tags=[]
reviews.country + " - " + reviews.region_1
# + [markdown] papermill={"duration": 0.020571, "end_time": "2021-11-08T23:58:46.696826", "exception": false, "start_time": "2021-11-08T23:58:46.676255", "status": "completed"} tags=[]
# These operators are faster than `map()` or `apply()` because they uses speed ups built into pandas. All of the standard Python operators (`>`, `<`, `==`, and so on) work in this manner.
#
# However, they are not as flexible as `map()` or `apply()`, which can do more advanced things, like applying conditional logic, which cannot be done with addition and subtraction alone.
#
# # Your turn
#
# If you haven't started the exercise, you can **[get started here](https://www.kaggle.com/kernels/fork/595524)**.
# + [markdown] papermill={"duration": 0.023375, "end_time": "2021-11-08T23:58:46.740966", "exception": false, "start_time": "2021-11-08T23:58:46.717591", "status": "completed"} tags=[]
# ---
#
#
#
#
# *Have questions or comments? Visit the [course discussion forum](https://www.kaggle.com/learn/pandas/discussion) to chat with other learners.*
| course/Pandas/summary-functions-and-maps.ipynb |
# ---
# title: "Visualize Neural Network Architecutre"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "How to visualize neural network architecture in Python."
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Preliminaries
# Load libraries
from keras import models
from keras import layers
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from keras.utils import plot_model
# ## Construct Neural Network Architecture
# +
# Start neural network
network = models.Sequential()
# Add fully connected layer with a ReLU activation function
network.add(layers.Dense(units=16, activation='relu', input_shape=(10,)))
# Add fully connected layer with a ReLU activation function
network.add(layers.Dense(units=16, activation='relu'))
# Add fully connected layer with a sigmoid activation function
network.add(layers.Dense(units=1, activation='sigmoid'))
# -
# ## Visualize Network Architecture
# Visualize network architecture
SVG(model_to_dot(network, show_shapes=True).create(prog='dot', format='svg'))
# ## Save To File
# Save the visualization as a file
plot_model(network, show_shapes=True, to_file='network.png')
| content/deep_learning/keras/.ipynb_checkpoints/visualize_neural_network_architecture-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import sys
sys.path.insert(1, '../../../../script/')
import math
import collections
import numpy as np
import pandas as pd
import scipy.stats as stats
import matplotlib.pyplot as plt
import seaborn as sns
#import missingno as msno
from collections import defaultdict
from scipy.stats.stats import pearsonr
from sklearn.model_selection import train_test_split
from imblearn.pipeline import Pipeline, make_pipeline
from imblearn.under_sampling import RandomUnderSampler
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import GridSearchCV
from imblearn.over_sampling import SMOTE
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score, f1_score, classification_report
from sklearn.metrics import roc_curve, auc, roc_auc_score
from sklearn.metrics import plot_confusion_matrix
import pydotplus
from sklearn import tree
from IPython.display import Image
import cleaning_class as cl
import cleaning_test as cl_test
# %load_ext autoreload
# %autoreload 2
# +
def report(results, n_top=3):
configurations = {}
c_i = 0
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean training score: {0:.3f} (std: {1:.3f})".format(
results['mean_train_score'][candidate],
results['std_train_score'][candidate]))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
configurations[c_i] = results['params'][candidate]
c_i += 1
return configurations
def report_multiple(results, n_top=3, scoring = 'roc_auc'):
configurations = {}
c_i = 0
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_' + scoring] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean training score:", end = '')
print(" roc_auc: {0:.3f} (std: {1:.3f})".format(
results['mean_train_roc_auc'][candidate],
results['std_train_roc_auc'][candidate]), end = '')
print(", Accuracy: {0:.3f} (std: {1:.3f})".format(
results['mean_train_accuracy'][candidate],
results['std_train_accuracy'][candidate]), end = '')
print(", f1: {0:.3f} (std: {1:.3f})".format(
results['mean_train_f1'][candidate],
results['std_train_f1'][candidate]))
print("Mean validation score:", end = '')
print(" roc_auc: {0:.3f} (std: {1:.3f})".format(
results['mean_test_roc_auc'][candidate],
results['std_test_roc_auc'][candidate]), end = '')
print(", Accuracy: {0:.3f} (std: {1:.3f})".format(
results['mean_test_accuracy'][candidate],
results['std_test_accuracy'][candidate]), end = '')
print(", f1: {0:.3f} (std: {1:.3f})".format(
results['mean_test_f1'][candidate],
results['std_test_f1'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
configurations[c_i] = results['params'][candidate]
c_i += 1
return configurations
# -
# # Check the feature used for classification
#
# ne ho selezionate solo alcune, speravo cambiasse qualcosa, possiamo scrivere che dopo diversi test sul decision tree ci siamo resi conto che queste hanno più importanza rispetto alle altre, le abbiamo selezionate per essere sicuri di non andare ada incorrere nella curse of dimensionality
df = pd.read_csv('../../data/training.csv')
cl.cleaning(df)
df.columns
df = pd.get_dummies(df)
df = pd.get_dummies(df, columns=['WheelTypeID'])
df.columns
# # Split in train and validation
#
# validation condiviso con le varie tecniche per il confronto, fatto con lo stratified per tenere tutto bilanciato con le classi.
# +
attributes = [col for col in df.columns if col != 'IsBadBuy']
x = df[attributes].values
y = df['IsBadBuy']
x_train, x_val, y_train, y_val = train_test_split(x, y, test_size=0.3, stratify=y)
# -
# # - Undersampling
# uso from RandomUnderSampler:
# https://imbalanced-learn.readthedocs.io/en/stable/under_sampling.html
# ### GRIDSEARCH
# +
param_list = {'criterion': ['gini', 'entropy'],
'max_depth': [None] + list(np.arange(2, 10)),
'min_samples_split': list(np.arange(2, 40)),
'min_samples_leaf': list(np.arange(1, 40)),
}
new_params = {'decisiontreeclassifier__' + key: param_list[key] for key in param_list}
skf = StratifiedKFold(n_splits=3)
clf = DecisionTreeClassifier(criterion='gini', max_depth=None, min_samples_split=2, min_samples_leaf=1)
imba_pipeline = make_pipeline(RandomUnderSampler(), clf)
scoring = ['accuracy', 'f1', 'roc_auc' ]
random_search = RandomizedSearchCV(imba_pipeline, param_distributions=new_params, n_iter=1000, cv=skf, scoring=scoring, refit = 'roc_auc', n_jobs = 4, verbose = 1, return_train_score=True)
random_search.fit(x_train, y_train)
cnfs = report_multiple(random_search.cv_results_, n_top=3, scoring = 'roc_auc')
# -
# ### Perform Clustering
rus = RandomUnderSampler()
x_u_train_resampled, y_u_train_resampled = rus.fit_resample(x_train, y_train)
models_u = []
y_pred_vals_u = []
y_pred_trains_u = []
hyper_ps = random_search.cv_results_
for cnf in cnfs.values():
criterion = cnf['decisiontreeclassifier__criterion']
max_depth = cnf['decisiontreeclassifier__max_depth']
min_samples_split = cnf['decisiontreeclassifier__min_samples_split']
min_samples_leaf = cnf['decisiontreeclassifier__min_samples_leaf']
clf = DecisionTreeClassifier(criterion=criterion, max_depth=max_depth, min_samples_split=min_samples_split, min_samples_leaf=min_samples_leaf)
clf = clf.fit(x_u_train_resampled, y_u_train_resampled)
models_u.append(clf)
y_pred = clf.predict(x_val)
y_pred_tr = clf.predict(x_u_train_resampled)
y_pred_vals_u.append(y_pred)
y_pred_trains_u.append(y_pred_tr)
# ### Analyze the classification results
roc_auc_models_u_val = []
for i in range(0,len(cnfs)):
print("model {}".format(i))
print('Train Accuracy %s' % accuracy_score(y_u_train_resampled, y_pred_trains_u[i]))
print('Train F1-score %s' % f1_score(y_u_train_resampled, y_pred_trains_u[i], average=None))
fpr, tpr, _ = roc_curve(y_u_train_resampled, y_pred_trains_u[i])
roc_auc = auc(fpr, tpr)
roc_auc = roc_auc_score(y_u_train_resampled, y_pred_trains_u[i], average=None)
print("Train roc_auc: {}".format(roc_auc))
print()
print('Test Accuracy %s' % accuracy_score(y_val, y_pred_vals_u[i]))
print('Test F1-score %s' % f1_score(y_val, y_pred_vals_u[i], average=None))
fpr, tpr, _ = roc_curve(y_val, y_pred_vals[i])
roc_auc = auc(fpr, tpr)
roc_auc = roc_auc_score(y_val, y_pred_vals[i], average=None)
print("Test roc_auc: {}".format(roc_auc))
roc_auc_models_u_val.append(roc_auc)
print(classification_report(y_val, y_pred_vals_u[i]))
print(confusion_matrix(y_val, y_pred_vals_u[i]))
for i in range(0,len(cnfs)):
print("model {} - roc_auc: {}".format(i, roc_auc_models_u_val[i]))
# ### Choose the best model
dot_data = tree.export_graphviz(models_u[0], out_file=None,
feature_names=attributes,
class_names=['BadBuy' if x == 1 else 'GoodBuy' for x in clf.classes_],
filled=True, rounded=True,
special_characters=True,
max_depth=3)
graph = pydotplus.graph_from_dot_data(dot_data)
Image(graph.create_png())
# Evaluate the performance
# +
plt.figure(figsize=(8, 5))
plt.plot(fpr, tpr, label='ROC curve (area = %0.2f)' % (roc_auc_models_u_val[0]))
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.tick_params(axis='both', which='major')
plt.legend(loc="lower right", fontsize=14, frameon=False)
plt.show()
# -
# # - Uppersampling
# rimetto insieme x_train e y_train per poter aumentare i record della classe minoritaria (lo 1) uso SMOTE:
# https://imbalanced-learn.readthedocs.io/en/stable/over_sampling.html
# ### GRIDSEARCH
# +
param_list = {
'criterion': ['gini', 'entropy'],
'max_depth': [None] + list(np.arange(2, 20)),
'min_samples_split': list(np.arange(2, 40)),
'min_samples_leaf': list(np.arange(2, 40)),
'min_impurity_decrease': [0.75e-6, 0.5e-6, 1e-6]
}
new_params = {'decisiontreeclassifier__' + key: param_list[key] for key in param_list}
skf = StratifiedKFold(n_splits=3)
clf = DecisionTreeClassifier(criterion='gini', max_depth=None, min_samples_split=2, min_samples_leaf=1)
imba_pipeline = make_pipeline(SMOTE(sampling_strategy= 1.0, random_state=42), clf)
scoring = ['accuracy', 'f1', 'roc_auc' ]
#grid_imba = RandomizedSearchCV(imba_pipeline, param_distributions=new_params, cv=skf, n_iter=400, return_train_score=True, scoring='f1', n_jobs = 4, verbose = 1)
grid_imba = RandomizedSearchCV(imba_pipeline, param_distributions=new_params, n_iter=100000, cv=skf, scoring=scoring, refit = 'roc_auc', n_jobs = 4, verbose = 1, return_train_score=True)
grid_imba.fit(x, y)
cnfs = report_multiple(grid_imba.cv_results_, n_top=3, scoring = 'roc_auc')
# -
cnfs = report_multiple(grid_imba.cv_results_, n_top=3, scoring = 'roc_auc')
# ### Perform Clustering
sm = SMOTE(sampling_strategy=1.0, random_state=42)
x_o_train_resampled, y_o_train_resampled = sm.fit_sample(x_train, y_train)
models_o = []
y_pred_vals_o = []
y_pred_trains_o = []
hyper_ps = grid_imba.cv_results_
for cnf in cnfs.values():
criterion = cnf['decisiontreeclassifier__criterion']
max_depth = cnf['decisiontreeclassifier__max_depth']
min_samples_split = cnf['decisiontreeclassifier__min_samples_split']
min_samples_leaf = cnf['decisiontreeclassifier__min_samples_leaf']
clf = DecisionTreeClassifier(criterion=criterion, max_depth=max_depth, min_samples_split=min_samples_split, min_samples_leaf=min_samples_leaf)
clf = clf.fit(x_o_train_resampled, y_o_train_resampled)
models_o.append(clf)
y_pred = clf.predict(x_val)
y_pred_tr = clf.predict(x_o_train_resampled)
y_pred_vals_o.append(y_pred)
y_pred_trains_o.append(y_pred_tr)
roc_auc_models_o_val = []
for i in range(0,len(cnfs)):
print("model {}".format(i))
print('Train Accuracy %s' % accuracy_score(y_o_train_resampled, y_pred_trains_o[i]))
print('Train F1-score %s' % f1_score(y_o_train_resampled, y_pred_trains_o[i], average=None))
fpr, tpr, _ = roc_curve(y_o_train_resampled, y_pred_trains_o[i])
roc_auc = auc(fpr, tpr)
roc_auc = roc_auc_score(y_o_train_resampled, y_pred_trains_o[i], average=None)
print("Train roc_auc: {}".format(roc_auc))
print()
print('Test Accuracy %s' % accuracy_score(y_val, y_pred_vals_o[i]))
print('Test F1-score %s' % f1_score(y_val, y_pred_vals_o[i], average=None))
fpr, tpr, _ = roc_curve(y_val, y_pred_vals_o[i])
roc_auc = auc(fpr, tpr)
roc_auc = roc_auc_score(y_val, y_pred_vals_o[i], average=None)
print("Test roc_auc: {}".format(roc_auc))
roc_auc_models_o_val.append(roc_auc)
print(classification_report(y_val, y_pred_vals_o[i]))
print(confusion_matrix(y_val, y_pred_vals_o[i]))
# ### Analyze the classification performance
for i in range(0,len(cnfs)):
print("model {} - roc_auc: {}".format(i, roc_auc_models_o_val[i]))
# ### Choose the best model
# Visualize the decision tree
dot_data = tree.export_graphviz(models_o[2], out_file=None,
feature_names=attributes,
class_names=['BadBuy' if x == 1 else 'GoodBuy' for x in clf.classes_],
filled=True, rounded=True,
special_characters=True,
max_depth=4)
graph = pydotplus.graph_from_dot_data(dot_data)
Image(graph.create_png())
# +
# %matplotlib inline
plt.figure(figsize=(8, 5))
plt.plot(fpr, tpr, label='ROC curve (area = %0.2f)' % (roc_auc_models_o_val[2]))
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.tick_params(axis='both', which='major')
plt.legend(loc="lower right", fontsize=14, frameon=False)
plt.show()
# -
# # Model evaluation on test set
# il miglior modello è quello con la tecnica dell'undersampling, valutiamo il tutto sul test set
df_test = pd.read_csv('../../data/test.csv')
cl_test.cleaning(df_test)
df_test.columns
df_test = pd.get_dummies(df_test)
df_test = pd.get_dummies(df_test, columns=['WheelTypeID'])
df_test.columns
attributes = [col for col in df_test.columns if col != 'IsBadBuy']
x_test = df_test[attributes].values
y_test = df_test['IsBadBuy']
y_pred_test = models_u[0].predict(x_test)
# +
print('Test Accuracy %s' % accuracy_score(y_test, y_pred_test))
print('Test F1-score %s' % f1_score(y_test, y_pred_test, average=None))
print(classification_report(y_test, y_pred_test))
# Plot non-normalized confusion matrix
titles_options = [("Confusion matrix, without normalization", None),
("Normalized confusion matrix", 'true')]
for title, normalize in titles_options:
disp = plot_confusion_matrix(models[0], x_test, y_test,
cmap=plt.cm.Blues,
normalize=normalize)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.show()
# -
fpr, tpr, _ = roc_curve(y_test, y_pred_test)
roc_auc = auc(fpr, tpr)
roc_auc = roc_auc_score(y_test, y_pred_test, average=None)
print("model {} - roc_auc: {}".format(0, roc_auc))
roc_auc_models.append(roc_auc)
# +
plt.figure(figsize=(8, 5))
plt.plot(fpr, tpr, label='ROC curve (area = %0.2f)' % (roc_auc))
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.tick_params(axis='both', which='major')
plt.legend(loc="lower right", fontsize=14, frameon=False)
plt.show()
| notebook/4_classification/6_CFU/old/c_report-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2020년 2월 25일 화요일
# ### 프로그래머스 스택/큐: 기능개발
# ### 문제 : https://programmers.co.kr/learn/courses/30/lessons/42586
# ### 블로그 : https://somjang.tistory.com/entry/Programmers-%EC%8A%A4%ED%83%9D%ED%81%90-%EA%B8%B0%EB%8A%A5%EA%B0%9C%EB%B0%9C-Python
# ### 첫번째 시도
def solution(progresses, speeds):
answer = []
min_key = 0
work_times_dic = {}
for i in range(len(progresses)):
work_time = (100 - progresses[i]) // speeds[i]
if (100-progresses[i]) % speeds[i] != 0:
work_time = work_time + 1
my_keys = work_times_dic.keys()
if i > 0:
min_key = min(my_keys)
if min_key < work_time and (work_time not in my_keys):
work_times_dic[work_time] = 1
elif min_key > work_time:
work_times_dic[min_key] = work_times_dic[min_key] + 1
work_times_dic = sorted(work_times_dic.items())
for i in range(len(work_times_dic)):
answer.append(work_times_dic[i][1])
return answer
solution([93, 30, 55], [1,30, 5])
# ---
# ### 두번째 시도
def solution(progresses, speeds):
answer = []
min_key = 0
work_times_dic = {}
for i in range(len(progresses)):
work_time = (100 - progresses[i]) // speeds[i]
if (100-progresses[i]) % speeds[i] != 0:
work_time = work_time + 1
my_keys = work_times_dic.keys()
if i > 0:
min_key = min(my_keys)
if min_key < work_time and (work_time not in my_keys):
work_times_dic[work_time] = 1
elif min_key > work_time:
work_times_dic[min_key] = work_times_dic[min_key] + 1
else:
work_times_dic[work_time] = work_times_dic[work_time] + 1
work_times_dic = sorted(work_times_dic.items())
for i in range(len(work_times_dic)):
answer.append(work_times_dic[i][1])
return answer
solution([93, 30, 55], [1,30, 5])
# ---
# ### 세번째 시도
def solution(progresses, speeds):
answer = []
min_key = 0
finish_time = []
finish_time_stack = []
for i in range(len(progresses)):
work_time = (100 - progresses[i]) // speeds[i]
if (100-progresses[i]) % speeds[i] != 0:
work_time = work_time + 1
finish_time.append(work_time)
finish_time_dic = {finish_time[0]:1}
finish_time_stack.append(finish_time[0])
for i in range(1, len(finish_time)):
if finish_time[i-1] > finish_time[i]:
finish_time_dic[finish_time[i-1]] = finish_time_dic[finish_time[i-1]] + 1
elif finish_time[i] not in finish_time_dic.keys():
finish_time_dic[finish_time[i]] = 1
finish_time_stack.append(finish_time[i])
else:
finish_time_dic[finish_time[i]] = finish_time_dic[finish_time[i]] + 1
for key in finish_time_dic.keys():
answer.append(finish_time_dic[key])
print(finish_time_dic)
return answer
solution([93, 30, 55], [1,30, 5])
solution([ 93 , 30 , 55 , 60 ], [ 1, 30 , 5 , 40 ])
solution([40, 93, 30, 55, 60, 65], [60, 1, 30, 5 , 10, 7])
# ### 네번째 시도
def solution(progresses, speeds):
answer = []
min_key = 0
finish_time = []
finish_time_stack = []
for i in range(len(progresses)):
work_time = (100 - progresses[i]) // speeds[i]
if (100-progresses[i]) % speeds[i] != 0:
work_time = work_time + 1
finish_time.append(work_time)
finish_time_stack.append(finish_time[0])
finish_time_dic = {finish_time[0]:1}
for i in range(1, len(finish_time)):
if finish_time[i-1] < finish_time[i] and (finish_time[i] > finish_time_stack[-1]):
finish_time_dic[finish_time[i]] = 1
finish_time_stack.append(finish_time[i])
else:
finish_time_dic[finish_time_stack[-1]] = finish_time_dic[finish_time_stack[-1]] + 1
for key in finish_time_dic.keys():
answer.append(finish_time_dic[key])
return answer
solution([40, 93, 30, 55, 60, 65], [60, 1, 30, 5 , 10, 7])
| DAY 001 ~ 100/DAY019_[Programmers] 스택 큐 기능개발 (Python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/a-dhingra/pandas_exercises/blob/master/02_Filtering_%26_Sorting/Chipotle/Exercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="e4qY_o85gh5n"
# # Ex1 - Filtering and Sorting Data
# + [markdown] id="V87zCzUxgh52"
# This time we are going to pull data directly from the internet.
# Special thanks to: https://github.com/justmarkham for sharing the dataset and materials.
#
# ### Step 1. Import the necessary libraries
# + id="rV3J1DiMgh53"
import pandas as pd
import numpy as np
# + [markdown] id="49QNIDa5gh55"
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv).
# + id="2fNxboP5g0AY" outputId="50a56a8b-722a-40ab-d969-fbfc590221e1" colab={"base_uri": "https://localhost:8080/", "height": 204}
url = r'https://raw.githubusercontent.com/justmarkham/DAT8/master/data/chipotle.tsv'
data = pd.read_csv(url, sep='\t')
data[:5]
# + [markdown] id="VSTYyf9vgh56"
# ### Step 3. Assign it to a variable called chipo.
# + id="CHfgKmSzgh56" outputId="481eb527-812c-4512-b54c-cceeea61bc3d" colab={"base_uri": "https://localhost:8080/", "height": 204}
chipo = pd.DataFrame(data)
chipo.head()
# + [markdown] id="zkZkXBQtgh57"
# ### Step 4. How many products cost more than $10.00?
# + id="BbhtHtTPjzTO" outputId="302af803-cb1a-4c17-84f0-faa5db2e41b5" colab={"base_uri": "https://localhost:8080/"}
# chipo.head()
len(chipo[chipo['item_price'].apply( lambda x: float(x[1:-1])) > 10])
# + [markdown] id="LGUN54QRgh58"
# ### Step 5. What is the price of each item?
# ###### print a data frame with only two columns item_name and item_price
# + id="YSwOgyECgh59" outputId="43afa1f0-4b11-4691-aac5-9a5c2f2a91c8" colab={"base_uri": "https://localhost:8080/", "height": 204}
df = chipo[['item_name','item_price']]
type(df)
df.head()
# + id="buCqB7bVmC1e" outputId="510db3c7-5355-4c62-daca-b63211aa8414" colab={"base_uri": "https://localhost:8080/", "height": 359}
df1 = df.drop_duplicates(subset=['item_name'], keep='first')
df1.head(10)
# without duplicate 'item_name' values
# + id="_ME7LK6vm8Bi" outputId="f5427ac1-6c33-4551-b8f8-24f6449a118e" colab={"base_uri": "https://localhost:8080/", "height": 359}
df.head(10) # with duplicate 'item_name' values
# + [markdown] id="Gnkkwcjygh59"
# ### Step 6. Sort by the name of the item
# + id="4IMXu1RomCCl" outputId="a767d293-d514-421a-d572-0448f50e3073" colab={"base_uri": "https://localhost:8080/", "height": 1000}
sorted_df1 = df1.sort_values(by='item_name')
sorted_df1
# + [markdown] id="gpWq9HLQgh5-"
# ### Step 7. What was the quantity of the most expensive item ordered?
# + id="h_BFeNdJgh5_"
price = [float(x[1:-1]) for x in chipo['item_price']]
chipo['price'] = price
# + id="hAkulsggsFr1" outputId="e9ca63eb-4bfa-454b-deda-f027745780aa" colab={"base_uri": "https://localhost:8080/"}
max_price = max(price)
max_price
# + id="9q9WhSjcctSs" outputId="fc849f2c-2dad-4d41-fe3d-8480c8903457" colab={"base_uri": "https://localhost:8080/", "height": 80}
chipo.loc[chipo['price']==max_price]
# + id="NzBP__tJFqpC"
# + [markdown] id="Glkq7aO1gh5_"
# ### Step 8. How many times was a Veggie Salad Bowl ordered?
# + id="K03FVXyigh6A" outputId="8cdd9239-ef77-40f6-817a-b71eb9131185" colab={"base_uri": "https://localhost:8080/"}
len(chipo[chipo['item_name'] =='Veggie Salad Bowl']['order_id'])
# + [markdown] id="oD4vfUpSgh6A"
# ### Step 9. How many times did someone order more than one Canned Soda?
# + id="xztu_29Jgh6B" outputId="930d7ba9-4f48-4b57-8764-226400fe6606" colab={"base_uri": "https://localhost:8080/"}
len(chipo[(chipo['item_name']=='Canned Soda') & (chipo['quantity']>1)])
# + id="0oZTU5yWdoMN"
| 02_Filtering_&_Sorting/Chipotle/Exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises
from __future__ import division, print_function, unicode_literals
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from sklearn import datasets
# %matplotlib inline
# Problem # 8
iris=datasets.load_iris()
X=iris['data'][:,(2,3)]
y=iris['target']
setosa_or_versicolor = (y == 0) | (y == 1)
X = X[setosa_or_versicolor]
y = y[setosa_or_versicolor]
from sklearn.svm import SVC,LinearSVC
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
_c=5
_loss='hinge'
_randomstate=42
lin_pipeline=Pipeline([
('imputer', SimpleImputer(strategy="median")),
('scaler',StandardScaler()),
])
X_scaled=lin_pipeline.fit_transform(X)
# Linear Classification
lin_clf=LinearSVC(loss=_loss,C=_c,random_state=_randomstate)
lin_clf.fit(X_scaled,y)
print("LinearSVC: ", lin_clf.intercept_, lin_clf.coef_)
# SVC
svc_clf=SVC(kernel='linear',C=_c)
svc_clf.fit(X_scaled,y)
print("SVC: ", svc_clf.intercept_, svc_clf.coef_)
# +
# SGDC Classifier
alpha = 1 / (_c * len(X))
from sklearn.linear_model import SGDClassifier
sgdc_clf=SGDClassifier(loss=_loss, learning_rate="constant", eta0=0.001, alpha=alpha,
max_iter=100000, tol=-np.infty, random_state=_randomstate)
sgdc_clf.fit(X_scaled,y)
print("SGDC: ", sgdc_clf.intercept_, sgdc_clf.coef_)
# +
w1 = -lin_clf.coef_[0, 0]/lin_clf.coef_[0, 1]
b1 = -lin_clf.intercept_[0]/lin_clf.coef_[0, 1]
w2 = -svc_clf.coef_[0, 0]/svc_clf.coef_[0, 1]
b2 = -svc_clf.intercept_[0]/svc_clf.coef_[0, 1]
w3 = -sgdc_clf.coef_[0, 0]/sgdc_clf.coef_[0, 1]
b3 = -sgdc_clf.intercept_[0]/sgdc_clf.coef_[0, 1]
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
lin_clf.fit(X_scaled, y)
svc_clf.fit(X_scaled, y)
sgdc_clf.fit(X_scaled, y)
# Transform the decision boundary lines back to the original scale
line1 = scaler.inverse_transform([[-10, -10 * w1 + b1], [10, 10 * w1 + b1]])
line2 = scaler.inverse_transform([[-10, -10 * w2 + b2], [10, 10 * w2 + b2]])
line3 = scaler.inverse_transform([[-10, -10 * w3 + b3], [10, 10 * w3 + b3]])
# Plot all three decision boundaries
plt.figure(figsize=(11, 4))
plt.plot(line1[:, 0], line1[:, 1], "k:", label="LinearSVC")
plt.plot(line2[:, 0], line2[:, 1], "b--", linewidth=2, label="SVC")
plt.plot(line3[:, 0], line3[:, 1], "r-", label="SGDClassifier")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs") # label="Iris-Versicolor"
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo") # label="Iris-Setosa"
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper center", fontsize=14)
plt.axis([0, 5.5, 0, 2])
plt.show()
# -
# Problem #9
try:
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1, cache=True)
except ImportError:
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original')
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test=train_test_split(mnist['data'],mnist['target'], test_size=0.2,random_state=42)
from sklearn.svm import LinearSVC
lin_clf=None
lin_clf=LinearSVC(random_state=_randomstate)
lin_clf.fit(X_train,y_train)
# +
from sklearn.metrics import accuracy_score
y_pred = lin_clf.predict(X_train)
accuracy_score(y_train, y_pred)
# -
scaler=StandardScaler()
X_train_scaled=scaler.fit_transform(X_train.astype(np.float32))
X_test_scaled=scaler.fit_transform(X_test.astype(np.float32))
lin_clf.fit(X_train_scaled,y_train)
y_pred = lin_clf.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
from sklearn.svm import SVC
svc_clf=None
svc_clf=SVC(decision_function_shape="ovr",gamma="auto")
svc_clf.fit(X_train_scaled,y_train)
y_pred = svc_clf.predict(X_train_scaled)
from sklearn.metrics import accuracy_score
accuracy_score(y_train, y_pred)
from scipy.stats import reciprocal, uniform
from sklearn.model_selection import RandomizedSearchCV
param_grid = [
# try 12 (3×4) combinations of hyperparameters
{
'C': reciprocal(1, 10),
'gamma': reciprocal(0.001, 0.1),
}
]
random_search = RandomizedSearchCV(svc_clf, param_grid,n_iter=10, verbose=2, cv=3)
random_search.fit(X_train_scaled[:1000],y_train[:1000])
random_search.best_estimator_
random_search.best_score_
random_search.best_estimator_.fit(X_train_scaled, y_train)
y_pred = random_search.best_estimator_.predict(X_train_scaled)
accuracy_score(y_train, y_pred)
# Problem #10
from sklearn.datasets import fetch_california_housing
housing=fetch_california_housing()
X=housing['data']
y=housing['target']
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
# -
lin_pipeline=Pipeline([
('imputer', SimpleImputer(strategy="median")),
('scaler',StandardScaler()),
])
X_train_scaled=lin_pipeline.fit_transform(X_train)
X_test_scaled=lin_pipeline.transform(X_test)
# +
from sklearn.svm import LinearSVR
lin_svr = LinearSVR(random_state=42)
lin_svr.fit(X_train_scaled, y_train)
# -
from sklearn.metrics import mean_squared_error
y_pred=lin_svr.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
mse
np.sqrt(mse)
# +
from sklearn.svm import SVR
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import reciprocal, uniform
param_distributions = {"gamma": reciprocal(0.001, 0.1), "C": uniform(1, 10)}
rnd_search_cv = RandomizedSearchCV(SVR(), param_distributions, n_iter=10, verbose=2, cv=3, random_state=42)
rnd_search_cv.fit(X_train_scaled, y_train)
# -
y_pred=rnd_search_cv.predict(X_train_scaled)
mse = mean_squared_error(y_train, y_pred)
mse
np.sqrt(mse)
| Hands-on-ML(Scikit-Learn&TF)/src/Labs/05_SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
1) Find the root - preorder[0]
2) Find inorder of both left and right subtree - Left sub tree followed by root and the rest will be right sub tree.
---> Search for root in inorder. (Left of root is left subtree and right of root is right subtree)
3) Find preorder of left and right subtree - Find the middle point between left and right sub tree.
4) Use recursion to build left and right subtree
5) Connect root with both
** Assume all data is unique
# -
Pre order (Root, left, right):
In order (Left, Root, right):
# +
import queue
class BinaryTreeNode:
def __init__(self, data):
self.data = data
self.left = None
self.right = None
def binary_search(a, x, si, ei):
if si > ei:
return -1
mid = (si + ei)//2
if a[mid] == x:
return mid
elif a[mid] > x:
return binary_search(a, x, si, mid-1)
else:
return binary_search(a, x, mid+1, ei)
def buildTreePreOrder(preorder, inorder):
if len(preorder) == 0:
return None
root = BinaryTreeNode(preorder[0])
rootIndex = binary_search(inorder, root.data, 0, len(inorder))
leftInorder = inorder[:rootIndex]
rightInorder = inorder[rootIndex + 1:]
x = len(leftInorder)
leftPreorder = preorder[1:1+x]
rightPreorder = preorder[1+x:]
root.left = buildTreePreOrder(leftPreorder, leftInorder) # Where to call recursion?
root.right = buildTreePreOrder(rightPreorder, rightInorder)
return root
def printLevelATNewLine(root):
if root==None:
return
inputQ = queue.Queue()
outputQ = queue.Queue()
inputQ.put(root)
while not inputQ.empty():
while not inputQ.empty():
curr = inputQ.get()
print(curr.data, end=' ')
if curr.left!=None:
outputQ.put(curr.left)
if curr.right!=None:
outputQ.put(curr.right)
print()
inputQ, outputQ = outputQ, inputQ
#n=int(input())
n = 7
#preorder = [int(i) for i in input().strip().split()]
#inorder = [int(i) for i in input().strip().split()]
#preorder = [1, 2, 3, 4, 15, 5, 6, 7, 8, 10, 9, 12]
#inorder = [4, 15, 3, 2, 5, 1, 6, 10, 8, 7, 9, 12]
inorder = [4,2,5,1,6,3,7]
preorder = [1,2,4,5,3,6,7]
root = buildTreePreOrder(preorder, inorder)
printLevelATNewLine(root)
# +
def buildTreeFromPreIn(preorder, inorder):
if len(preorder) == 0:
return None
rootData = preorder[0]
root = BinaryTreeNode(rootData)
rootIndexInInorder = -1
for i in range(0, len(inorder)):
if inorder[i] == rootData:
rootIndexInInorder = i
break
if rootIndexInInorder == -1:
return None
leftInorder = inorder[0:rootIndexInorder]
rightInorder = inorder[rootIndexInorder + 1:]
lenLeftSubtree = len(leftInorder)
leftPreorder = preorder[1:lenLeftSubtree + 1]
rightPreorder = preorder[lenLeftSubtree + 1:]
leftChild = buildTreeFromPreIn(leftPreorder, leftInorder)
rightChild = buildTreeFromPreIn(rightPreorder, rightInorder)
root.left = leftChild
root.right = rightChild
return root
inorder = [4,2,5,1,6,3,7]
preorder = [1,2,4,5,3,6,7]
root = buildTreePreOrder(preorder, inorder)
printLevelATNewLine(root)
printTreeDetailed(root)
# -
| 14 Binary Trees - 2/14.09 Construct a tree using inorder and preorder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from typing import List
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.axes import Axes
import torch
import torch.nn as nn
from torch.optim import SGD
from torch.distributions.multivariate_normal import MultivariateNormal
from sklearn.linear_model import BayesianRidge
from scipy.stats import multivariate_normal, sem
from experiments.linear_regression_posterior import (
get_features_and_targets,
compute_true_posterior,
build_model_and_callbacks,
fit_model,
)
from experiments.utils.metrics import compute_distance_between_matrices, compute_gaussian_wasserstein_distance
from swafa.fa import OnlineEMFactorAnalysis
np.set_printoptions(suppress=True) # don't use scientific notation
# %load_ext autoreload
# %autoreload 2
# -
# ### Define parameters
# +
# data
n_samples, n_features = 1000, 2
feature_covar = [
[1, 0.5],
[0.5, 1]
]
alpha = 0.01
beta = 0.1
lambda_ = alpha / beta
# callbacks
latent_dim = 1
gradient_weight_posterior_kwargs = dict(
latent_dim=latent_dim,
optimiser=SGD,
optimiser_kwargs=dict(lr=0.001),
n_warm_up_time_steps=100,
random_seed=1,
)
em_weight_posterior_kwargs = dict(
latent_dim=latent_dim,
n_warm_up_time_steps=100,
random_seed=1,
)
posterior_update_epoch_start = 1
posterior_eval_epoch_frequency = 100000
# training
batch_size = 100
pre_train_lr = 1e-3
pre_train_n_epochs = 500
swa_lr = 1e-2
swa_weight_decay = 1e-3
swa_n_epochs = 100
# -
# ### Define helper functions
# +
def run_experiment(n_samples: int, n_features: int, feature_covar: np.ndarray, alpha: float, beta: float,
latent_dim: int, gradient_weight_posterior_kwargs: dict, em_weight_posterior_kwargs: dict,
posterior_update_epoch_start: int, posterior_eval_epoch_frequency: int, batch_size: int,
pre_train_lr: float, pre_train_n_epochs: int, swa_lr: float, swa_weight_decay: float,
swa_n_epochs: int, n_trials: int) -> (List[float], List[float], List[float]):
relative_distances_from_mean = []
relative_distances_from_cov = []
wasserstein_distances = []
for n in range(n_trials):
results = run_trial(
n_samples=n_samples,
n_features=n_features,
feature_covar=feature_covar,
alpha=alpha,
beta=beta,
latent_dim=latent_dim,
gradient_weight_posterior_kwargs=gradient_weight_posterior_kwargs,
em_weight_posterior_kwargs=em_weight_posterior_kwargs,
posterior_update_epoch_start=posterior_update_epoch_start,
posterior_eval_epoch_frequency=posterior_eval_epoch_frequency,
batch_size=batch_size,
pre_train_lr=pre_train_lr,
pre_train_n_epochs=pre_train_n_epochs,
swa_lr=swa_lr,
swa_weight_decay=swa_weight_decay,
swa_n_epochs=swa_n_epochs,
random_seed=n
)
relative_distances_from_mean.append(
compute_distance_between_matrices(
results['true_posterior_mean'],
results['empirical_mean'],
) /
compute_distance_between_matrices(
results['true_posterior_mean'],
torch.zeros_like(results['true_posterior_mean']),
)
)
relative_distances_from_cov.append(
compute_distance_between_matrices(
results['true_posterior_covar'],
results['empirical_covar'],
) /
compute_distance_between_matrices(
results['true_posterior_covar'],
torch.zeros_like(results['true_posterior_covar']),
)
)
wasserstein_distances.append(
compute_gaussian_wasserstein_distance(
mean1=results['true_posterior_mean'],
covar1=results['true_posterior_covar'],
mean2=results['empirical_mean'],
covar2=results['empirical_covar'],
) / n_features
)
return relative_distances_from_mean, relative_distances_from_cov, wasserstein_distances
def run_trial(n_samples: int, n_features: int, feature_covar: np.ndarray, alpha: float, beta: float,
latent_dim: int, gradient_weight_posterior_kwargs: dict, em_weight_posterior_kwargs: dict,
posterior_update_epoch_start: int, posterior_eval_epoch_frequency: int, batch_size: int,
pre_train_lr: float, pre_train_n_epochs: int, swa_lr: float, swa_weight_decay: float,
swa_n_epochs: int, random_seed: int) -> dict:
X, y = generate_data(n_samples, n_features, feature_covar, random_seed)
true_posterior_mean, true_posterior_covar, _, _ = compute_true_posterior(X, y, alpha=alpha, beta=beta)
(
model,
gradient_posterior_update_callback,
em_posterior_update_callback,
sklearn_posterior_eval_callback,
gradient_posterior_eval_callback,
em_posterior_eval_callback,
) = build_model_and_callbacks(
X=X,
true_posterior_mean=true_posterior_mean,
true_posterior_covar=true_posterior_covar,
model_optimiser_class=SGD,
model_optimiser_kwargs=None,
posterior_latent_dim=latent_dim,
gradient_weight_posterior_kwargs=gradient_weight_posterior_kwargs,
em_weight_posterior_kwargs=em_weight_posterior_kwargs,
posterior_update_epoch_start=posterior_update_epoch_start,
posterior_eval_epoch_frequency=posterior_eval_epoch_frequency,
model_random_seed=random_seed,
)
callbacks = [
gradient_posterior_update_callback,
em_posterior_update_callback,
sklearn_posterior_eval_callback,
gradient_posterior_eval_callback,
em_posterior_eval_callback,
]
model.optimiser_kwargs = dict(lr=pre_train_lr)
fit_model(X=X, y=y, model=model, callbacks=None, n_epochs=pre_train_n_epochs, batch_size=batch_size)
w_pretrained = torch.clone(model.output_layer.weight.data).numpy().squeeze()
model.optimiser_kwargs = dict(lr=swa_lr, weight_decay=swa_weight_decay)
fit_model(X=X, y=y, model=model, callbacks=callbacks, n_epochs=swa_n_epochs, batch_size=batch_size)
empirical_mean, empirical_covar = sklearn_posterior_eval_callback.get_empirical_mean_and_covariance()
W = np.vstack(sklearn_posterior_eval_callback.weight_iterates)
return dict(
W=W,
w_pretrained=w_pretrained,
true_posterior_mean=true_posterior_mean,
true_posterior_covar=true_posterior_covar,
empirical_mean=empirical_mean,
empirical_covar=empirical_covar,
)
def generate_data(n_samples: int, n_features: int, feature_covar: np.ndarray, random_seed: int,
) -> (np.ndarray, np.ndarray):
np.random.seed(random_seed)
torch.manual_seed(random_seed)
p_x = MultivariateNormal(loc=torch.zeros(n_features), covariance_matrix=torch.Tensor(feature_covar))
X = p_x.sample((n_samples,))
p_theta = MultivariateNormal(loc=torch.zeros(n_features), covariance_matrix=torch.eye(n_features) / alpha)
theta = p_theta.sample().reshape(-1, 1)
epsilon = torch.normal(torch.zeros(n_samples), 1 / np.sqrt(beta) * torch.ones(n_samples))
y = X.mm(theta).squeeze() + epsilon
return X, y
# -
# ### Run experiment
relative_distances_from_mean, relative_distances_from_cov, wasserstein_distances = run_experiment(
n_samples=n_samples,
n_features=n_features,
feature_covar=feature_covar,
alpha=alpha,
beta=beta,
latent_dim=latent_dim,
gradient_weight_posterior_kwargs=gradient_weight_posterior_kwargs,
em_weight_posterior_kwargs=em_weight_posterior_kwargs,
posterior_update_epoch_start=posterior_update_epoch_start,
posterior_eval_epoch_frequency=posterior_eval_epoch_frequency,
batch_size=batch_size,
pre_train_lr=pre_train_lr,
pre_train_n_epochs=pre_train_n_epochs,
swa_lr=swa_lr,
swa_weight_decay=swa_weight_decay,
swa_n_epochs=swa_n_epochs,
n_trials=10
)
# +
relative_distances_from_mean_mean = np.mean(relative_distances_from_mean)
relative_distances_from_mean_std_error = sem(relative_distances_from_mean)
relative_distances_from_cov_mean = np.mean(relative_distances_from_cov)
relative_distances_from_cov_std_error = sem(relative_distances_from_cov)
wasserstein_mean = np.mean(wasserstein_distances)
wasserstein_std_error = sem(wasserstein_distances)
print('Mean Relative Distance from Mean = {:.4f} +- {:.4f}'.format(
relative_distances_from_mean_mean, relative_distances_from_mean_std_error,
))
print('Mean Relative Distance from Covar = {:.4f} +- {:.4f}'.format(
relative_distances_from_cov_mean, relative_distances_from_cov_std_error,
))
print('Mean Wasserstein Distance = {:.4f} +- {:.4f}'.format(
wasserstein_mean, wasserstein_std_error,
))
# -
# ### Plot the SGD iterates for a single trial
# +
results = run_trial(
n_samples=n_samples,
n_features=n_features,
feature_covar=feature_covar,
alpha=alpha,
beta=beta,
latent_dim=latent_dim,
gradient_weight_posterior_kwargs=gradient_weight_posterior_kwargs,
em_weight_posterior_kwargs=em_weight_posterior_kwargs,
posterior_update_epoch_start=posterior_update_epoch_start,
posterior_eval_epoch_frequency=posterior_eval_epoch_frequency,
batch_size=batch_size,
pre_train_lr=pre_train_lr,
pre_train_n_epochs=pre_train_n_epochs,
swa_lr=swa_lr,
swa_weight_decay=swa_weight_decay,
swa_n_epochs=swa_n_epochs,
random_seed=0,
)
W = results['W']
w_pretrained = results['w_pretrained']
true_posterior_mean = results['true_posterior_mean'].numpy()
true_posterior_covar = results['true_posterior_covar'] .numpy()
empirical_mean = results['empirical_mean'].numpy()
empirical_covar = results['empirical_covar'].numpy()
# +
def plot_pdf(W: np.ndarray, mean: np.ndarray, covar: np.ndarray, w_pretrained: np.ndarray, w_swa: np.ndarray, ax: Axes):
plt.rcParams.update({'font.size': 12})
w1, w2 = W[:, 0], W[:, 1]
border = 0.05
x_max = max(w1.max(), w_pretrained[0]) + border
x_min = min(w1.min(), w_pretrained[0]) - border
y_max = max(w2.max(), w_pretrained[1]) + border
y_min = min(w2.min(), w_pretrained[1]) - border
X, Y = np.mgrid[x_min:x_max:.01, y_min:y_max:.01]
pos = np.dstack((X, Y))
rv = multivariate_normal(mean, covar)
c = ax.contourf(X, Y, rv.pdf(pos))
plt.colorbar(c, ax=ax)
ax.scatter(w1, w2, color='red', s=10, alpha=0.7, label='Sampled weights')
ax.scatter(w_pretrained[0], w_pretrained[1], marker='o', color='white', s=200, label='Pre-trained weights')
ax.scatter(w_swa[0], w_swa[1], marker='*', color='black', s=200, label='SWA solution')
fig, axes = plt.subplots(1, 1, figsize=(8, 6))
plot_pdf(W, true_posterior_mean, true_posterior_covar, w_pretrained, empirical_mean, axes)
plt.legend()
png_path = f'../../thesis/plots/linear_model_weight_iterates__lr={swa_lr}__lambda={swa_weight_decay}.png'
plt.savefig(png_path, format='png')
plt.show()
# -
# ### Plot the window averages of the SGD iterates for the same data
# +
window_size = 50 #int(n_samples / batch_size)
n_windows = int(len(W) / window_size)
W_averages = np.array([x.mean(axis=0) for x in np.split(W, n_windows)])
fig, axes = plt.subplots(1, 1, figsize=(8, 6))
plot_pdf(W_averages, true_posterior_mean, true_posterior_covar, w_pretrained, W_averages.mean(axis=0), axes)
plt.legend()
png_path = f'../../thesis/plots/linear_model_average_weight_iterates__lr={swa_lr}__lambda={swa_weight_decay}__batch_size={batch_size}__window_size={window_size}.png'
plt.savefig(png_path, format='png')
plt.show()
# -
| experiments/notebooks/LinearRegressionSGDIterates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn.functional as F
# replace following class code with an easy sequential network
class Net(torch.nn.Module):
def __init__(self, n_feature, n_hidden, n_output):
super(Net, self).__init__()
self.hidden = torch.nn.Linear(n_feature, n_hidden) # hidden layer
self.predict = torch.nn.Linear(n_hidden, n_output) # output layer
def forward(self, x):
x = F.relu(self.hidden(x)) # activation function for hidden layer
x = self.predict(x) # linear output
return x
net1 = Net(1, 10, 1)
# easy and fast way to build your network
net2 = torch.nn.Sequential(
torch.nn.Linear(1, 10),
torch.nn.ReLU(),
torch.nn.Linear(10, 1)
)
print(net1) # net1 architecture
"""
Net (
(hidden): Linear (1 -> 10)
(predict): Linear (10 -> 1)
)
"""
print(net2) # net2 architecture
"""
Sequential (
(0): Linear (1 -> 10)
(1): ReLU ()
(2): Linear (10 -> 1)
)
"""
# -
| chapter6/code/.ipynb_checkpoints/torch_neuralnetwork-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
GRAMMAR="""
@@grammar::CALC
start = expression $ ;
expression
=
| expression '+' term
| expression '-' term
| expression cmpop expression
| term
;
term
=
| term '*' factor
| term '/' factor
| factor
;
factor
=
| '(' expression ')'
| latex
| symbol
;
cmpop
=
| Equal
| NotEq
| Lt
| LtE
| Gt
| GtE ;
Equal = '=' ;
NotEq = '!=' ;
Lt = '<' ;
LtE = '<=';
Gt = '>';
GtE = '>=' ;
latex
=
|func_lim
|func_int
|func_sum
|func_prod
|func_log
|func_ln
|func_sin
|func_cos
|func_tan
|func_csc
|func_sec
|func_cot
|func_arcsin
|func_arccos
|func_arctan
|func_arccsc
|func_arcsec
|func_arccot
|func_sinh
|func_cosh
|func_tanh
|func_arsinh
|func_arcosh
|func_artanh
|func_sqrt
| greek_alpha
| greek_beta
| greek_gamma
| greek_mu
| greek_phi
| greek_pi
| greek_varphi
| accent_hat
| accent_bar
| accent_prime
| cmd_times
| cmd_cdot
| cmd_div
| cmd_frac
| cmd_mathig
;
cmd_times = '\\times';
cmd_cdot = '\\cdot';
cmd_div = '\\div';
cmd_frac = '\\frac';
cmd_mathig = '\\mathit';
accent_hat = "\\hat"
;
accent_bar = "\\bar"
;
accent_prime = "\\prime"
;
greek_alpha =
| "\\alpha"
| "\\Alpha"
;
greek_beta =
| "\\beta"
| "\\Beta"
;
greek_gamma =
| "\\gamma"
| "\\Gamma"
;
greek_mu =
| "\\mu"
| "\\Mu"
;
greek_phi =
| "\\phi"
| "\\Phi"
;
greek_pi =
| "\\pi"
| "\\Pi"
;
greek_varphi =
| "\\varphi"
| "\\VarPhi"
;
l_paren = '(';
r_paren = ')';
l_brace = '{';
r_brace = '}';
l_bracket = '[';
r_bracket = ']';
symbol
=
| variable
| constant
;
variable
=
| 'x'
| 'y'
| 'z'
;
constant = real ;
func_lim = "\\lim" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_int = "\\int" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_sum = "\\sum" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_prod = "\\prod" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_log = "\\log" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_ln = "\\ln" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_sin = "\\sin" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_cos = "\\cos" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_tan = "\\tan" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_csc = "\\csc" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_sec = "\\sec" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_cot = "\\cot" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arcsin = "\\arcsin" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arccos = "\\arccos" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arctan = "\\arctan" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arccsc = "\\arccsc" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arcsec = "\\arcsec" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arccot = "\\arccot" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_sinh = "\\sinh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_cosh = "\\cosh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_tanh = "\\tanh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arsinh = "\\arsinh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_arcosh = "\\arcosh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_artanh = "\\artanh" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
func_sqrt = "\\sqrt" (l_paren|l_brace) (expression|variable|constant) (r_paren|r_brace) ;
LIM_APPROACH_SYM
=
|'\\to'
| '\\rightarrow'
| '\\Rightarrow'
| '\\longrightarrow'
| '\\Longrightarrow'
;
real
=
|scientific
|float
|integer;
scientific = /[0-9]*\.*[0-9]+[eEdD][+-]?[0-9]+/ ;
float = /-?[0-9]*.[0-9]+/
;
integer = /\d+/
;
"""
import tatsu
from tatsu.util import asjson
def valid_expression(expression):
try:
print(expression)
#grammar = open("grammars/calc.ebnf").read()
parser = tatsu.compile(GRAMMAR)
ast = parser.parse(expression)
return ast
except Exception as e:
print(e)
return False
def parse_expression(expression):
# if expression is valid, return ast
ast = valid_expression(expression)
if ast:
import pprint
# pprint.pprint(ast, indent=2, width=20)
#data = json.dumps(asjson(pprint.pprint(ast,indent=2, width=20)))
data = json.dumps(asjson(ast), indent=2)
#store_results(data)
return data
return None
# -
valid_expression('1 + 2 = \\log(x)')
| example/AMS_LaTex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reset
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
# These are some parameters to make figures nice (and big)
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
plt.rcParams['figure.figsize'] = 16,8
params = {'legend.fontsize': 'x-large',
'figure.figsize': (15, 5),
'axes.labelsize': 'x-large',
'axes.titlesize':'x-large',
'xtick.labelsize':'x-large',
'ytick.labelsize':'x-large'}
plt.rcParams.update(params)
# -
# # Exercise 1: Unfair dice
# Consider a pair of unfair dice. The probabilities for the two dice are as follows:
#
# |Roll|Probability Dice 1|Probability Dice 2
# |---|---|---|
# |1|1/8|1/10|
# |2|1/8|1/10|
# |3|1/8|1/10|
# |4|1/8|1/10|
# |5|1/8|3/10|
# |6|3/8|3/10|
#
# ## Question:
# Use the law of total probability. to compute the probability of rolling a total of 11.
#
# ### Answer
# We denote by $S$ the sum of the dice and by $D_1$ the value of the roll of dice 1
# $$P(S=11)=\sum_{n=1}^{6}P(S=11|D_{1}=n)$$
# $$P(S=11)=P(S=11|D_{1}=5)\cdot P(D_{1}=5)+P(S=11|D_{1}=6)\cdot P(D_{1}=6)$$
# $$P(S=11)=P(D_{2}=6)\cdot P(D_{1}=5)+P(D_{2}=6)\cdot P(D_{1}=5)$$
# $$P(S=11)=3/10\cdot1/8+3/10\cdot3/8=10/80=1/8$$
#
# <hr style="border:2px solid black"> </hr>
# # Exercise 2: Covariance vs independence
# Consider two random variables, $X$ and $Y$. $X$ is uniformly distributed over the interval $\left[-1,1\right]$:
#
# $$X\sim U[-1,1],$$
#
# while $Y$ is normally distributed (Gaussian), with a variance equal to $X^{2}$. We would denote this as:
# $$Y|X\sim\mathcal{N}\left(0,X^{2}\right),$$
# to imply that
# $$P(Y=y|X=x)=p(y|x)=\left(2\pi x^2\right)^{-1/2}\exp\left[-\frac{1}{2}\left(\frac{y}{x}\right)^2\right]$$
# The two random variables are obviously not independent. Indepencene requires $p(y|x)=p(y)$, which in turn would imply $p(y)=p(y|x_1)p(y|x_2)$ for $x_1\neq x_2$.
# ## Question 1 (Theory):
# Prove analyitically that $Cov(X,Y)=0$.<br>
# *Hint:* Use the relation $p(x,y)=p(y|x)p(x)$ to compute $E(XY)$. Alternatively, you can use the same relation to first prove $E(E(Y|X))$.
#
# ### Answer:
# $$Cov(X,Y)=E(XY)-E(X)E(Y)=E(XY)$$
# $$=\int_{-1}^{1}\int_{-\infty}^{\infty}x\cdot y\cdot p(x,y)\cdot dx\cdot dy=\int_{-1}^{1}\int_{-\infty}^{\infty}y\cdot x\cdot p(y|x)p(x)\cdot dx\cdot dy$$
# $$=\int_{-1}^{1}\left[\int_{-\infty}^{\infty}y\cdot p(y|x)\cdot dy\right]x\cdot dx$$
# $$=\int_{-1}^{1}\left[\int_{-\infty}^{\infty}y\cdot\frac{1}{\sqrt{2\pi x^{2}}}e^{-\frac{1}{2}\left(\frac{y}{x}\right)^{2}}\right]x\cdot dx$$
# The inner integral is just the expected value of $y$ for a constant $x$, $E(Y|X)$ and it is zero, since $Y|X\sim\mathcal{N}\left(0,X^{2}\right)$. Thus, since the integrand is zero, the whole intergral is zero.
# ## Question 2 (Numerical):
# Show, numerically, that expected covariance is zero.
# 1. Draw $n$ samples $(x_j,y_j)$ of $(X,Y)$ and plot $y_j$ vs $x_j$ for $n=100$:
# 2. Compute the sample covariance $s_{n-1}=\frac{1}{n-1}\sum_{j=1}^{n}(y_j-\overline y)$ of $X,Y$ for $n=100$. Repeat the experiment a large number of times (e.g. $M=10,000$) and plot the sampling distribution of $s_{100-1}$. What is the mean of the sampling distribution.
# 3. Now increase the sample size up to $n=100,000$ and plot the value of the sample covariance as a function of $n$. By the Law of Large Numbers you should see it asymptote to zero
#
# ### Answer
# +
#2.1
Ndraws=100
X=stats.uniform.rvs(loc=-1,scale=2,size=Ndraws);
Y=np.zeros([Ndraws])
for i in range(Ndraws):
Y[i]=stats.norm.rvs(loc=0,scale=np.abs(X[i]),size=1)
plt.plot(X,Y,'.')
scov=1/(Ndraws-1)*np.sum((X-np.mean(X))*(Y-np.mean(Y)))
print(scov)
# +
#2.2
M=1000
Ndraws=100
scov=np.zeros(M);
for j in range(M):
X=stats.uniform.rvs(loc=-1,scale=2,size=Ndraws);
Y=np.zeros([Ndraws]);
for i in range(Ndraws):
Y[i]=stats.norm.rvs(loc=0,scale=np.abs(X[i]),size=1);
scov[j]=1/(Ndraws-1)*np.sum((X-np.mean(X))*(Y-np.mean(Y)));
plt.hist(scov,rwidth=0.98);
print(np.mean(scov))
# +
#2.3
Ndraws=100000
scov=np.zeros(Ndraws)
X=stats.uniform.rvs(loc=-1,scale=2,size=Ndraws)
Y=np.zeros([Ndraws])
for i in range(Ndraws):
Y[i]=stats.norm.rvs(loc=0,scale=np.abs(X[i]),size=1)
if i>1:
scov[i]=1/(i-1)*np.sum((X[0:i]-np.mean(X[0:i]))*(Y[0:i]-np.mean(Y[0:i])))
plt.plot(scov)
plt.grid()
# -
# <hr style="border:2px solid black"> </hr>
# # Exercise 3: Central Limit Theorem
# The central limit theorem says that the distribution of the sample mean of **any** random variable approaches a normal distribution.
#
# **Theorem** Let $ X_1, \cdots , X_n $ be $n$ independent and identically distributed (i.i.d) random variables with expectation $\mu$ and variance $\sigma^2$. The distribution of the sample mean $\overline X_n=\frac{1}{n}\sum_{i=1}^n X_i$ approaches the distribution of a gaussian
#
# $$\overline X_n \sim \mathcal N (\mu,\sigma^2/n),$$
# for large $n$.
#
# In this exercise, you will convince yourself of this theorem numerically. Here is a recipe for how to do it:
# - Pick your probability distribution. The CLT even works for discrete random variables!
# - Generate a random $n \times m$ matrix ($n$ rows, $m$ columns) of realizations from that distribution.
# - For each column, find the sample mean $\overline X_n$ of the $n$ samples, by taking the mean along the first (0-th) dimension. You now have $m$ independent realizations of the sample mean $\overline X_n$.
# - You can think of each column as an experiment where you take $n$ samples and average over them. We want to know the distribution of the sample-mean. The $m$ columns represent $m$ experiments, and thus provide us with $m$ realizations of the sample mean random variable. From these we can approximate a distribution of the sample mean (via, e.g. a histogram).
# - On top of the histogram of the sample mean distribution, plot the pdf of a normal distribution with the same process mean and process variance as the sample mean of the distribution of $\overline X_n$.
#
#
# ## Question 1: Continuous random variables:
# Demonstrate, numerically, that the sample mean of a number of Gamma-distributed random variables is approximately normal. https://en.wikipedia.org/wiki/Gamma_distribution
#
# Plot the distribution of the sample mean for $n=[1,5,25,100]$,using $m=10,000$, and overlay it with a normal pdf. For best visualization,use values of $\alpha=1$ loc$=0$, scale=$1$ for the gamma distribution; 30 bins for the histogram; and set the x-limits of [3,6] for all four values of $n$.
#
# ### Answer:
# +
m=10000
n=[1,5,20,100]
Nbins=30
fig,ax=plt.subplots(4,1,figsize=[8,8])
alpha=1;
loc=0;
scale=1;
for j in range(4):
x=stats.gamma.rvs(alpha,loc=loc,scale=scale,size=[n[j],m])
sample_mean=np.mean(x,axis=0);
z=np.linspace(0,5,100);
norm_pdf=stats.norm.pdf(z,loc=np.mean(sample_mean),scale=np.std(sample_mean));
ax[j].hist(sample_mean,Nbins,rwidth=1,density=True)
ax[j].plot(z,norm_pdf);
ax[j].set_xlim(left=0,right=4)
# -
# ## Question 2: Discrete random variables:
# Demonstrate, numerically, that the sample mean of a large number of random dice throws is approximately normal.
#
# Simulate the dice using a discrete uniform random variables <code>stats.randint.rvs</code>, taking values from 1 to 6 (remember Python is right exclusive). The sample mean $\overline X_n$ is thus equivalnt to the average value of the dice throw $n$ throws.
#
# Plot the normalized (density=True) histogram for $n=[1,2,25,200]$, using $m=100,000$, and overlay it with a normal pdf. For best visualization use 50 bins for the histogram, and set the x-limits of [1,6] for all four values of $n$.
# ### Answer
# +
m=100000
n=[1,2,25,200]
Nbins=50
fig,ax=plt.subplots(4,1,figsize=[16,8])
alpha=1;
loc=0;
scale=1;
for j in range(4):
x=stats.randint.rvs(1,7,size=[n[j],m])
sample_mean=np.mean(x,axis=0);
z=np.linspace(0,7,1000);
norm_pdf=stats.norm.pdf(z,loc=np.mean(sample_mean),scale=np.std(sample_mean));
ax[j].hist(sample_mean,Nbins,rwidth=1,density=True)
ax[j].plot(z,norm_pdf);
ax[j].set_xlim(left=1,right=6)
# -
# ## Question 3: Precip in Urbana
# Plot the histograms of precipitation in urbana on hourly, daily, monthly, and annual time scales. What do you observe?
#
# For convenience, I've downloaded 4-times daily hourly data from ERA5 for the gridcell representing Urbana. We'll use xarray since it makes it very easy to compute daily-, monthly-, and annual-total precipitation.
#
# The cell below computes hourly, daily, monthly, and annual values of precipitation. All you have to do is plot their histograms
# +
import xarray as xr
#convert from m/hr to inches/hr, taking into account we only sample 4hrs of the day
ds=xr.open_dataset('/data/keeling/a/cristi/SIMLES/data/ERA5precip_urbana_1950-2021.nc');
unit_conv=1000/24.5*6
pr_hr =ds.tp*unit_conv;
pr_day =pr_hr.resample(time='1D').sum('time')
pr_mon=pr_hr.resample(time='1M').sum('time')
pr_yr =pr_hr.resample(time='1Y').sum('time')
Nbins=15;
# -
# ### Answer
Nbins=15
fig,ax=plt.subplots(2,2,figsize=[12,12]);
ax[0,0].hist(pr_hr,Nbins,rwidth=0.9);
ax[0,1].hist(pr_day,Nbins,rwidth=0.9);
ax[1,0].hist(pr_mon,Nbins,rwidth=0.9);4
ax[1,1].hist(pr_yr,Nbins,rwidth=0.9);
# <hr style="border:2px solid black"> </hr>
# # Exercise 4: Houston precipitation return times via MLE
# In the wake of <NAME>, many have described the assocaited flooding as a "500-year event". How can this be, given that in most places there are only a few decades of data available? In this exercise we apply a simple (and most likely wrong) methodology to estimate _return periods_, and comment on the wisdom of that concept.
#
# Let's load and get to know the data. We are looking at daily precip data (in cm) at Beaumont Research Center and Port Arthur, two of the weather stations in the Houston area that reported very high daily precip totals.
#
# The data comes from NOAA GHCN:<br>
# https://www.ncdc.noaa.gov/cdo-web/datasets/GHCND/stations/GHCND:USC00410613/detail<br>
# https://www.ncdc.noaa.gov/cdo-web/datasets/GHCND/stations/GHCND:USW00012917/detail
#
# read data and take a cursory look
#df=pd.read_csv('/data/keeling/a/cristi/SIMLES/data/Beaumont_precip.csv')
df=pd.read_csv('/data/keeling/a/cristi/SIMLES/data/PortArthur_precip.csv')
df.head()
# +
# plot raw precipitation
precip_raw=df['PRCP'].values
precip_raw=precip_raw[np.isnan(precip_raw)==False] # take out nans
fig,ax=plt.subplots(1,1)
ax.plot(precip_raw)
ax.set_xlabel('day since beginning of record')
ax.set_ylabel('Daily Precip (cm)')
# +
# Plot the histogram of the data.
# For distributions such as a gamma distribution it makes sense to use a logarithmic axis.
#define bin edges and bin widths.
# we'll use the maximum value in the data to define the upper limit
bin_edge_low=0
bin_edge_high=np.round(np.max(precip_raw)+1);
bin_width=0.25
bin_edges=np.arange(bin_edge_low,bin_edge_high,bin_width)
fig,ax=plt.subplots(1,2)
ax[0].hist(precip_raw,bin_edges,rwidth=0.9);
ax[0].set_xlabel('daily precip (cm)')
ax[0].set_ylabel('count (number of days)')
ax[0].grid()
ax[1].hist(precip_raw,bin_edges,rwidth=0.9)
ax[1].set_yscale('log')
ax[1].grid()
ax[1].set_xlabel('daily precip (cm)')
ax[1].set_ylabel('count (number of days)')
# +
# the jump in the first bin indicates a probability mass at 0 ( a large number of days do not see any precipitation).
# Let's only look at days when it rains. While we're at it, let's clean NaNs as well.
precip=precip_raw[precip_raw>0.01]
# Plot the histogram of the data
fig,ax=plt.subplots(1,2)
ax[0].hist(precip,bin_edges,rwidth=0.9);
ax[0].set_xlabel('daily precip (cm)')
ax[0].set_ylabel('count (number of days)')
ax[0].grid()
ax[0].set_xlabel('daily precip (cm)')
ax[0].set_ylabel('count (number of days)')
ax[1].hist(precip,bin_edges,rwidth=0.9)
ax[1].set_yscale('log')
ax[1].grid()
ax[1].set_xlabel('daily precip (cm)')
ax[1].set_ylabel('count (number of days)')
# -
# ## Question 1:
# Fit an gamma distribution to the data, using the <code>stats.gamma.fit</code> method to obtain maximum likelihood estimates for the parameters.
# Show the fit by overlaying the pdf of the gamma distribution with mle parameters on top of the histogram of daily precipitation at Beaumont Research Center.
#
# Hints:
# - you'll need to show a *density* estimate of the histogram, unlike the count i.e. ensure <code>density=True</code>.
# - The method will output the thre parameters of the gamma random variable: <code>a,loc,scale</code> (see documentation <a href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.gamma.html"> here</a>). So you'll need to call it as <code>alpha_mle,loc_mle,scale_mle=stats.gama.fit( .... )</code>
#
# ### Answer:
# +
alpha_mle,loc_mle,scale_mle=stats.gamma.fit(precip)
x_plot=np.linspace(0,np.max(precip),200)
gamma_pdf=stats.gamma.pdf(x_plot,alpha_mle,loc_mle,scale_mle)
# Plot the histogram of the data
fig,ax=plt.subplots(1,2)
ax[0].hist(precip,bin_edges,rwidth=0.9,density=True);
ax[0].set_xlabel('daily precip (cm)')
ax[0].set_ylabel('count (number of days)')
ax[1].hist(precip,bin_edges,rwidth=0.9,density=True)
ax[1].set_yscale('log')
ax[0].plot(x_plot,gamma_pdf)
ax[1].plot(x_plot,gamma_pdf)
# -
np.max(precip)
# ## Question 2:
# Compute the return time of the rainiest day recorded at Beaumont Research Center (in years).
#
# What does this mean? The rainiest day at Beaumont brought $x$ cm. The return time represents how often we would expect to get $x$ cm or more of rain at Beaumont.
#
# To compute the return time we need to compute the probability of daily rain >$x$ cm. The inverse of this probability is the frequency of daily rain >$x$ cm.
#
# For example, if the probability of daily rain > 3 cm =1/30, it means we would expect that it rains 3 cm or more once about every 30 day, and we would say 3 cm is a 10 day event.
#
# For the largest precip event the probability will be significantly smaller, and thus the return time significantly larger
#
# *Hint*: Remember that the probability of daily rain being *less* than $x$ cm is given by the CDF: $$F(x)=P(\text{daily rain}<x\text{ cm})$$.
# *Hint*: The answer should only take a very small number of lines of code
# ### Answer
gamma_F=stats.gamma.cdf(x_plot,alpha_mle,loc_mle,scale_mle)
prob=1-stats.gamma.cdf(np.max(precip),alpha_mle,loc_mle,scale_mle)
1/prob/365
# ## Question 3:
# Repeat the analysis for the Port Arthur data. If you fit a Gamma ditribution and compute the return time of the largest daily rain event, what is the return time?
#
# Does that seem reasonable? Why do you think the statistical model fails here? Think of the type of precipitation events that make up the precipitation data at Port Arthur
#
# {
# "tags": [
# "margin",
# ]
# }
# ### Answer
# +
# read data and take a cursory look
df=pd.read_csv('/data/keeling/a/cristi/SIMLES/data/PortArthur_precip.csv')
df.head()
# plot raw precipitation
precip_raw=df['PRCP'].values
precip_raw=precip_raw[np.isnan(precip_raw)==False] # take out nans
precip=precip_raw[precip_raw>0.01]
alpha_mle,loc_mle,scale_mle=stats.gamma.fit(precip)
x_plot=np.linspace(0,np.max(precip),200)
gamma_pdf=stats.gamma.pdf(x_plot,alpha_mle,loc_mle,scale_mle)
# Plot the histogram of the data
fig,ax=plt.subplots(1,2)
ax[0].hist(precip,bin_edges,rwidth=0.9,density=True);
ax[0].set_xlabel('daily precip (cm)')
ax[0].set_ylabel('count (number of days)')
ax[1].hist(precip,bin_edges,rwidth=0.9,density=True)
ax[1].set_yscale('log')
ax[0].plot(x_plot,gamma_pdf)
ax[1].plot(x_plot,gamma_pdf)
# -
gamma_F=stats.gamma.cdf(x_plot,alpha_mle,loc_mle,scale_mle)
prob=1-stats.gamma.cdf(np.max(precip),alpha_mle,loc_mle,scale_mle)
1/prob/365
| content/Module01/M01_Lab_Answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import math
df = pd.read_csv('titanic.csv')
list(df)
df = df[['Pclass', 'Name', 'Sex', 'Age', 'SibSp', 'Parch', 'Embarked', 'Survived']]
mapping_dict = df.groupby('Pclass')['Age'].mean().to_dict()
mapping_dict
mapping_dict.get(1)
def fill_age(row):
if (math.isnan(row['Age'])):
return mapping_dict.get(row['Pclass'])
return row['Age']
df['Age'] = df.apply(fill_age, axis=1)
df.info()
df.drop('Name', axis=1, inplace=True)
df
pd.get_dummies(df['Sex'])
gender_series = pd.get_dummies(df['Sex'], drop_first=True)
embarked_series = pd.get_dummies(df['Embarked'], drop_first=True)
df = pd.concat([df, gender_series, embarked_series], axis = 1)
df.drop(['Sex', 'Embarked'], axis=1, inplace=True)
final = df[['Pclass', 'Age', 'SibSp', 'Parch', 'male', 'Q', 'S', 'Survived']]
final
final.to_csv('cleaned_data.csv', index=False)
| data_wrangling_and_eda/data_wrangling_continued.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''base'': conda)'
# name: python3
# ---
# # Analysis of WASP-189 (Phase Curve) using `pycheops`
#
# In the following notebook, we will analyse the data from `CHEOPS` visit 1 data of HD106315b using `pycheops`.
#
# The method is similar to that used in the analysis of KELT-11b data ([https://github.com/Jayshil/pycheops-tutorials/blob/main/KELT-11/p2_kelt11_extra_decorr_parm.ipynb](https://github.com/Jayshil/pycheops-tutorials/blob/main/KELT-11/p2_kelt11_extra_decorr_parm.ipynb)).
import numpy as np
import matplotlib.pyplot as plt
from astropy.io import fits
from dace.cheops import Cheops
import pycheops
import re
from pycheops.utils import phaser
from kelp import Model, Planet, Filter
from scipy.optimize import minimize
from emcee import EnsembleSampler
from multiprocessing import Pool
from corner import corner
# ## Downloading the data
# +
# Downloading data
dd = pycheops.Dataset('CH_PR100036_TG000701_V0200')
tim, fl, fle = dd.get_lightcurve(aperture='DEFAULT', decontaminate=True)#, reject_highpoints=True)
# To clip outliers (I would, in general, not prefer using this)
tim, fl, fle = dd.clip_outliers(verbose=True);
# Plotting the data
plt.figure(figsize=(16,9))
plt.errorbar(tim, fl, yerr=fle, fmt='.', c='orangered')
plt.xlabel('Time (BJD)')
plt.ylabel('Normalised Flux')
plt.title('Transit lightcurve for WASP-189b (Aperture: DEFAULT)')
plt.grid()
# -
# ### Planetary check
#
# We may want to check the position of various planetary bodies with respect to the target star, because if some of the bodies is very near to the target star then we may want to correct for their effects.
dd.planet_check()
# ## Renormalising the lightcurve
#
# It already seems pretty flatten -- and applying the function would only reduce the quality of the data.
# +
# This code is taken from the pycheops example notebook
dep, dep_err = 0.00499886*1e6, 1.8378e-05*1e6
P = 2.7240338
BJD_0 = 2456706.4558
cycle = round((dd.bjd_ref-BJD_0)/P)
T_0 = BJD_0 - dd.bjd_ref + (cycle+0)*P
T_1 = BJD_0 - dd.bjd_ref + (cycle+1)*P
T_ec = T_0 + (P/2)
D = dep/1e6 # Depth stored in ppm
W = 0.1819/P # Width stored in days
try:
f_c = 0.#np.sqrt(0.093)*np.sin(67.0*np.pi/180)
f_s = 0.#np.sqrt(0.093)*np.cos(67.0*np.pi/180)
except:
# From Pepper et al., 2017
ecosw = 0.#ufloat(-0.004,0.05099)
esinw = 0.#ufloat(0.031,0.055)
ecc = usqrt(ecosw**2+esinw**2)
f_s = 0.#esinw/usqrt(ecc) # f_s = sqrt(e)sin(omega) = e.sin(omega)/sqrt(e)
f_c = 0.#ecosw/usqrt(ecc) # f_c = sqrt(e)cos(omega) = e.cos(omega)/sqrt(e)
# Let's take data from first transit only
"""
mask1 = tim > T_0-(W*P/2)
mask2 = tim < T_0+(W*P/2)
mask3 = tim > T_ec - (W*P/2)
mask4 = tim < T_ec + (W*P/2)
mask5 = tim > T_1-(W*P/2)
mask6 = tim < T_1+(W*P/2)
"""
mask = np.zeros(len(tim), dtype=bool)
for i in range(len(tim)):
if (tim[i] > T_0-(W*P/2)) and (tim[i] < T_0+(W*P/2)):
mask[i] = True
elif (tim[i]>T_ec-(W*P/2)) and (tim[i]<T_ec+(W*P/2)):
mask[i] = True
elif (tim[i]>T_1-(W*P/2)) and (tim[i]<T_1+(W*P/2)):
mask[i] = True
print(mask)
tim, fl, fle = dd.mask_data(mask)
# Plotting the data
plt.figure(figsize=(16,9))
plt.errorbar(tim, fl, yerr=fle, fmt='.', c='orangered')
plt.axvline(T_ec, c='k', lw=3)
plt.axvline(x=T_0, c='k', lw=3)
plt.axvline(x=T_1, c='k', lw=3)
plt.xlabel('Time (BJD)')
plt.ylabel('Normalised Flux')
plt.title('Phase curve for WASP-189b (Aperture: DEFAULT)')
plt.grid()
# -
tim, fl, fle = dd.flatten(T_0, P*W)
# Plotting the data
plt.figure(figsize=(16,9))
plt.errorbar(tim, fl, yerr=fle, fmt='.', c='orangered')
plt.axvline(T_ec, c='k', lw=3)
plt.axvline(x=T_0, c='k', lw=3)
plt.axvline(x=T_1, c='k', lw=3)
plt.xlabel('Time (BJD)')
plt.ylabel('Normalised Flux')
plt.title('Phase curve for HD106315b (Aperture: DEFAULT)')
plt.grid()
# ## Detrending (or, decorrelating) the dataset
#
# To perform this operation of detrending we may want to look at diagnostic report for this observations. The diagnostic report consist of various plots of flux as a function of several instrumental properties like roll angle of the spacecraft, centroid position etc. This would enable us to see if there is any trend going on with instruments so that we can take care of it. So, first let's see the diagnostic report...
dd.diagnostic_plot()
dd.should_I_decorr()
dd.decorr(dfdx=True, dfdy=True, dfdt=True, dfdbg=True, dfdsinphi=True, dfdcosphi=True)
# # Modelling the phasecurve
phs = phaser(tim,2.72403380,0.43646240,-0.5)
planet = Planet(per=2.72403380, t0=0.43646240, inc=84.7812, rp=0.07070261, ecc=0., w=90., a=4.71779818, t_secondary=1.7984793, rp_a=0.01498636,\
fp=1., T_s=8000., limb_dark='quadratic', u=[0.4,0.2])
filt = Filter.from_name("CHEOPS")
# +
def pc_model(p, x):
"""
Phase curve model with two free parameters
"""
offset, c_11, a_b= p
C = [[0],
[0, c_11, 0]]
model = Model(hotspot_offset=offset, alpha=0.6,
omega_drag=4.5, A_B=a_b, C_ml=C, lmax=1,
planet=planet, filt=filt)
fl1 = model.thermal_phase_curve(x, f=2**-0.5, check_sorted=False).flux
fl_norm = fl1/np.max(fl1)
return fl_norm
def lnprior(p):
"""
Log-prior: sets reasonable bounds on the fitting parameters
"""
offset, c_11, a_b = p
if (offset > np.max(tim) or offset < np.min(tim) or c_11 > 1 or c_11 < 0 or a_b < 0 or a_b > 1):
return -np.inf
return 0
def lnlike(p, x, y, yerr):
"""
Log-likelihood: via the chi^2
"""
return -0.5 * np.sum((pc_model(p, x) - y)**2 / yerr**2)
def lnprob(p, x, y, yerr):
"""
Log probability: sum of lnlike and lnprior
"""
lp = lnprior(p)
if np.isfinite(lp):
return lp + lnlike(p, x, y, yerr)
return -np.inf
# +
initp = np.array([-0.7, 0.1, 0.5])
bounds = [[0, 2], [0.1, 1]]
soln = minimize(lambda *args: -lnprob(*args),
initp, args=(tim, fl, fle),
method='powell')
soln
# +
flux_model = pc_model(soln.x, tim)
residuals = fl-flux_model
figure, ax = plt.subplots(2, 1, sharex = True, figsize=(16,9))
ax[0].scatter(tim, fl, marker='.')
ax[0].plot(tim, flux_model, "r")
ax[1].scatter(tim, residuals, marker='.')
ax[0].set_ylabel("Flux")
ax[1].set_ylabel("Residuals")
ax[1].set_xlabel("Time")
ax[1].grid()
ax[0].grid()
# -
def lsq(p):
chi = ((pc_model(p,tim)-f)/fle)**2
ab = np.sum(chi)
return ab
| WASP-189/Phase_curves/p1_wasp189_phase_curve_kelp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import json
def main():
base_library_url = "http://127.0.0.1:5000/api/v1/resources/books/"
_id = 0
library_url = base_library_url + f"?id={_id}"
req = requests.get(library_url)
while req.json() != []:
print(req.json())
print()
_id += 1
library_url = base_library_url + f"?id={_id}"
req = requests.get(library_url)
print(req, req.text)
main()
| Remote Library Client.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Minibatch Stochastic Gradient Descent
// :label:`sec_minibatch_sgd`
//
// So far we encountered two extremes in the approach to gradient based learning: :numref:`sec_gd` uses the full dataset to compute gradients and to update parameters, one pass at a time. Conversely :numref:`sec_sgd` processes one observation at a time to make progress. Each of them has its own drawbacks. Gradient Descent is not particularly *data efficient* whenever data is very similar. Stochastic Gradient Descent is not particularly *computationally efficient* since CPUs and GPUs cannot exploit the full power of vectorization. This suggests that there might be a happy medium, and in fact, that's what we have been using so far in the examples we discussed.
//
// ## Vectorization and Caches
//
// At the heart of the decision to use minibatches is computational efficiency. This is most easily understood when considering parallelization to multiple GPUs and multiple servers. In this case we need to send at least one image to each GPU. With 8 GPUs per server and 16 servers we already arrive at a minibatch size of 128.
//
// Things are a bit more subtle when it comes to single GPUs or even CPUs. These devices have multiple types of memory, often multiple type of compute units and different bandwidth constraints between them. For instance, a CPU has a small number of registers and then L1, L2 and in some cases even L3 cache (which is shared between the different processor cores). These caches are of increasing size and latency (and at the same time they are of decreasing bandwidth). Suffice it to say, the processor is capable of performing many more operations than what the main memory interface is able to provide.
//
// * A 2GHz CPU with 16 cores and AVX-512 vectorization can process up to $2 \cdot 10^9 \cdot 16 \cdot 32 = 10^{12}$ bytes per second. The capability of GPUs easily exceeds this number by a factor of 100. On the other hand, a midrange server processor might not have much more than 100 GB/s bandwidth, i.e., less than one tenth of what would be required to keep the processor fed. To make matters worse, not all memory access is created equal: first, memory interfaces are typically 64 bit wide or wider (e.g., on GPUs up to 384 bit), hence reading a single byte incurs the cost of a much wider access.
// * There is significant overhead for the first access whereas sequential access is relatively cheap (this is often called a burst read). There are many more things to keep in mind, such as caching when we have multiple sockets, chiplets and other structures. A detailed discussion of this is beyond the scope of this section. See e.g., this [Wikipedia article](https://en.wikipedia.org/wiki/Cache_hierarchy) for a more in-depth discussion.
//
// The way to alleviate these constraints is to use a hierarchy of CPU caches which are actually fast enough to supply the processor with data. This is *the* driving force behind batching in deep learning. To keep matters simple, consider matrix-matrix multiplication, say $\mathbf{A} = \mathbf{B}\mathbf{C}$. We have a number of options for calculating $\mathbf{A}$. For instance we could try the following:
//
// 1. We could compute $\mathbf{A}_{ij} = \mathbf{B}_{i,:} \mathbf{C}_{:,j}^\top$, i.e., we could compute it element-wise by means of dot products.
// 1. We could compute $\mathbf{A}_{:,j} = \mathbf{B} \mathbf{C}_{:,j}^\top$, i.e., we could compute it one column at a time. Likewise we could compute $\mathbf{A}$ one row $\mathbf{A}_{i,:}$ at a time.
// 1. We could simply compute $\mathbf{A} = \mathbf{B} \mathbf{C}$.
// 1. We could break $\mathbf{B}$ and $\mathbf{C}$ into smaller block matrices and compute $\mathbf{A}$ one block at a time.
//
// If we follow the first option, we will need to copy one row and one column vector into the CPU each time we want to compute an element $\mathbf{A}_{ij}$. Even worse, due to the fact that matrix elements are aligned sequentially we are thus required to access many disjoint locations for one of the two vectors as we read them from memory. The second option is much more favorable. In it, we are able to keep the column vector $\mathbf{C}_{:,j}$ in the CPU cache while we keep on traversing through $B$. This halves the memory bandwidth requirement with correspondingly faster access. Of course, option 3 is most desirable. Unfortunately, most matrices might not entirely fit into cache (this is what we are discussing after all). However, option 4 offers a practically useful alternative: we can move blocks of the matrix into cache and multiply them locally. Optimized libraries take care of this for us. Let us have a look at how efficient these operations are in practice.
// +
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.7.0-SNAPSHOT
// %maven ai.djl:basicdataset:0.7.0-SNAPSHOT
// %maven org.slf4j:slf4j-api:1.7.26
// %maven org.slf4j:slf4j-simple:1.7.26
// %maven ai.djl.mxnet:mxnet-engine:0.7.0-SNAPSHOT
// %maven ai.djl.mxnet:mxnet-native-auto:1.7.0-a
// -
// %load ../utils/plot-utils
// %load ../utils/Functions.java
// %load ../utils/StopWatch.java
// %load ../utils/Training.java
// %load ../utils/Accumulator.java
// +
import ai.djl.ndarray.index.NDIndex;
import ai.djl.Model;
import ai.djl.metric.Metrics;
import ai.djl.ndarray.NDArray;
import ai.djl.ndarray.NDList;
import ai.djl.ndarray.NDManager;
import ai.djl.ndarray.types.DataType;
import ai.djl.ndarray.types.Shape;
import ai.djl.nn.Blocks;
import ai.djl.nn.SequentialBlock;
import ai.djl.nn.core.Linear;
import ai.djl.repository.Repository;
import ai.djl.training.DefaultTrainingConfig;
import ai.djl.training.EasyTrain;
import ai.djl.training.Trainer;
import ai.djl.training.dataset.Batch;
import ai.djl.training.dataset.Dataset;
import ai.djl.training.evaluator.Accuracy;
import ai.djl.training.initializer.NormalInitializer;
import ai.djl.training.listener.TrainingListener;
import ai.djl.training.loss.Loss;
import ai.djl.training.optimizer.Optimizer;
import ai.djl.training.optimizer.learningrate.LearningRateTracker;
import org.apache.commons.lang3.ArrayUtils;
import ai.djl.translate.TranslateException;
import ai.djl.basicdataset.AirfoilRandomAccess;
// -
NDManager manager = NDManager.newBaseManager();
StopWatch stopWatch = new StopWatch();
NDArray A = manager.zeros(new Shape(256, 256));
NDArray B = manager.randomNormal(new Shape(256, 256));
NDArray C = manager.randomNormal(new Shape(256, 256));
// Element-wise assignment simply iterates over all rows and columns of $\mathbf{B}$ and $\mathbf{C}$ respectively to assign the value to $\mathbf{A}$.
// Compute A = B C one element at a time
stopWatch.start();
for (int i = 0; i < 256; i++) {
for (int j = 0; j < 256; j++) {
A.set(new NDIndex(i, j),
B.get(new NDIndex(String.format("%d, :", i)))
.dot(C.get(new NDIndex(String.format(":, %d", j)))));
}
}
stopWatch.stop();
// A faster strategy is to perform column-wise assignment.
// Compute A = B C one column at a time
stopWatch.start();
for (int j = 0; j < 256; j++) {
A.set(new NDIndex(String.format(":, %d", j)), B.dot(C.get(new NDIndex(String.format(":, %d", j)))));
}
stopWatch.stop();
// Last, the most effective manner is to perform the entire operation in one block. Let us see what the respective speed of the operations is.
// +
// Compute A = B C in one go
stopWatch.start();
A = B.dot(C);
stopWatch.stop();
// Multiply and add count as separate operations (fused in practice)
float[] gigaflops = new float[stopWatch.getTimes().size()];
for (int i = 0; i < stopWatch.getTimes().size(); i++) {
gigaflops[i] = (float)(2 / stopWatch.getTimes().get(i));
}
System.out.printf("Performance in Gigaflops: element %.3f, column %.3f, full %.3f", gigaflops[0], gigaflops[1], gigaflops[2]);
// -
// ## Minibatches
//
// :label:`sec_minibatches`
//
// In the past we took it for granted that we would read *minibatches* of data rather than single observations to update parameters. We now give a brief justification for it. Processing single observations requires us to perform many single matrix-vector (or even vector-vector) multiplications, which is quite expensive and which incurs a significant overhead on behalf of the underlying deep learning framework. This applies both to evaluating a network when applied to data (often referred to as inference) and when computing gradients to update parameters. That is, this applies whenever we perform $\mathbf{w} \leftarrow \mathbf{w} - \eta_t \mathbf{g}_t$ where
//
// $$\mathbf{g}_t = \partial_{\mathbf{w}} f(\mathbf{x}_{t}, \mathbf{w})$$
//
// We can increase the *computational* efficiency of this operation by applying it to a minibatch of observations at a time. That is, we replace the gradient $\mathbf{g}_t$ over a single observation by one over a small batch
//
// $$\mathbf{g}_t = \partial_{\mathbf{w}} \frac{1}{|\mathcal{B}_t|} \sum_{i \in \mathcal{B}_t} f(\mathbf{x}_{i}, \mathbf{w})$$
//
// Let us see what this does to the statistical properties of $\mathbf{g}_t$: since both $\mathbf{x}_t$ and also all elements of the minibatch $\mathcal{B}_t$ are drawn uniformly at random from the training set, the expectation of the gradient remains unchanged. The variance, on the other hand, is reduced significantly. Since the minibatch gradient is composed of $b := |\mathcal{B}_t|$ independent gradients which are being averaged, its standard deviation is reduced by a factor of $b^{-\frac{1}{2}}$. This, by itself, is a good thing, since it means that the updates are more reliably aligned with the full gradient.
//
// Naively this would indicate that choosing a large minibatch $\mathcal{B}_t$ would be universally desirable. Alas, after some point, the additional reduction in standard deviation is minimal when compared to the linear increase in computational cost. In practice we pick a minibatch that is large enough to offer good computational efficiency while still fitting into the memory of a GPU. To illustrate the savings let us have a look at some code. In it we perform the same matrix-matrix multiplication, but this time broken up into "minibatches" of 64 columns at a time.
stopWatch.start();
for (int j = 0; j < 256; j+=64) {
A.set(new NDIndex(String.format(":, %d:%d", j, j + 64)),
B.dot(C.get(new NDIndex(String.format(":, %d:%d", j, j + 64)))));
}
stopWatch.stop();
System.out.printf("Performance in Gigaflops: block %.3f\n", 2 / stopWatch.getTimes().get(3));
// As we can see, the computation on the minibatch is essentially as efficient as on the full matrix. A word of caution is in order. In :numref:`sec_batch_norm` we used a type of regularization that was heavily dependent on the amount of variance in a minibatch. As we increase the latter, the variance decreases and with it the benefit of the noise-injection due to batch normalization. See e.g., :cite:`Ioffe.2017` for details on how to rescale and compute the appropriate terms.
//
// ## Reading the Dataset
//
// Let us have a look at how minibatches are efficiently generated from data. In the following we use a dataset developed by NASA to test the wing [noise from different aircraft](https://archive.ics.uci.edu/ml/datasets/Airfoil+Self-Noise) to compare these optimization algorithms. For convenience we only use the first $1,500$ examples. The data is whitened for preprocessing, i.e., we remove the mean and rescale the variance to $1$ per coordinate.
// +
NDManager manager = NDManager.newBaseManager();
public AirfoilRandomAccess getDataCh11(int batchSize, int n) throws IOException, TranslateException {
// Load data
AirfoilRandomAccess airfoil =
AirfoilRandomAccess.builder()
.optUsage(Dataset.Usage.TRAIN)
.setSampling(batchSize, true)
.build();
// Select Features
airfoil.addAllFeatures();
// Prepare Data
airfoil.prepare();
// Select first n cases
airfoil.selectFirstN(n);
// Remove the mean and rescale variance to 1 for all features
airfoil.whitenAll();
return airfoil;
}
// -
// ## Implementation from Scratch
//
// Recall the minibatch SGD implementation from :numref:`sec_linear_scratch`. In the following we provide a slightly more general implementation. For convenience it has the same call signature as the other optimization algorithms introduced later in this chapter. Specifically, we add the status
// input `states` and place the hyperparameter in dictionary `hyperparams`. In
// addition, we will average the loss of each minibatch example in the training
// function, so the gradient in the optimization algorithm does not need to be
// divided by the batch size.
public class Optimization {
public static void sgd(NDList params, NDList states, Map<String, Float> hyperparams) {
for (int i = 0; i < params.size(); i++) {
NDArray param = params.get(i);
// Update param
// param = param - param.gradient * lr
param.subi(param.getGradient().mul(hyperparams.get("lr")));
}
}
}
// Next, we implement a generic training function to facilitate the use of the other optimization algorithms introduced later in this chapter. It initializes a linear regression model and can be used to train the model with minibatch SGD and other algorithms introduced subsequently.
public static float evaluateLoss(Iterable<Batch> dataIterator, NDArray w, NDArray b) {
Accumulator metric = new Accumulator(2); // sumLoss, numExamples
for (Batch batch : dataIterator) {
NDArray X = batch.getData().head();
NDArray y = batch.getLabels().head();
NDArray yHat = Training.linreg(X, w, b);
float lossSum = Training.squaredLoss(yHat, y).sum().getFloat();
metric.add(new float[]{lossSum, (float) y.size()});
batch.close();
}
return metric.get(0) / metric.get(1);
}
public static class LossTime {
public float[] loss;
public float[] time;
public LossTime(float[] loss, float[] time) {
this.loss = loss;
this.time = time;
}
}
public void plotLossEpoch(float[] loss, float[] epoch) {
Table data = Table.create("data")
.addColumns(
DoubleColumn.create("epoch", Functions.floatToDoubleArray(epoch)),
DoubleColumn.create("loss", Functions.floatToDoubleArray(loss))
);
display(LinePlot.create("loss vs. epoch", data, "epoch", "loss"));
}
// +
import ai.djl.training.GradientCollector;
import ai.djl.engine.Engine;
public float[] arrayListToFloat (ArrayList<Double> arrayList) {
float[] ret = new float[arrayList.size()];
for (int i = 0; i < arrayList.size(); i++) {
ret[i] = arrayList.get(i).floatValue();
}
return ret;
}
@FunctionalInterface
public static interface TrainerConsumer {
void train(NDList params, NDList states, Map<String, Float> hyperparams);
}
public static LossTime trainCh11(TrainerConsumer trainer, NDList states, Map<String, Float> hyperparams,
AirfoilRandomAccess dataset,
int featureDim, int numEpochs) {
NDManager manager = NDManager.newBaseManager();
NDArray w = manager.randomNormal(0, 0.01f, new Shape(featureDim, 1), DataType.FLOAT32);
NDArray b = manager.zeros(new Shape(1));
w.attachGradient();
b.attachGradient();
NDList params = new NDList(w, b);
int n = 0;
StopWatch stopWatch = new StopWatch();
stopWatch.start();
float lastLoss = -1;
ArrayList<Double> loss = new ArrayList<>();
ArrayList<Double> epoch = new ArrayList<>();
for (int i = 0; i < numEpochs; i++) {
for (Batch batch : dataset.getData(manager)) {
int len = (int) dataset.size() / batch.getSize(); // number of batches
NDArray X = batch.getData().head();
NDArray y = batch.getLabels().head();
NDArray l;
try (GradientCollector gc = Engine.getInstance().newGradientCollector()) {
NDArray yHat = Training.linreg(X, params.get(0), params.get(1));
l = Training.squaredLoss(yHat, y).mean();
gc.backward(l);
}
trainer.train(params, states, hyperparams);
n += X.getShape().get(0);
if (n % 200 == 0) {
stopWatch.stop();
lastLoss = evaluateLoss(dataset.getData(manager), params.get(0), params.get(1));
loss.add((double) lastLoss);
double lastEpoch = 1.0 * n / X.getShape().get(0) / len;
epoch.add(lastEpoch);
stopWatch.start();
}
batch.close();
}
}
plotLossEpoch(arrayListToFloat(loss), arrayListToFloat(epoch));
System.out.printf("loss: %.3f, %.3f sec/epoch\n", lastLoss, stopWatch.avg());
return new LossTime(arrayListToFloat(loss), arrayListToFloat(stopWatch.cumsum()));
}
// -
// Let us see how optimization proceeds for batch gradient descent. This can be achieved by setting the minibatch size to 1500 (i.e., to the total number of examples). As a result the model parameters are updated only once per epoch. There is little progress. In fact, after 6 steps progress stalls.
// +
public static LossTime trainSgd(float lr, int batchSize, int numEpochs) throws IOException, TranslateException {
AirfoilRandomAccess dataset = getDataCh11(batchSize, 1500);
int featureDim = dataset.getFeatureArraySize();
Map<String, Float> hyperparams = new HashMap<>();
hyperparams.put("lr", lr);
return trainCh11(Optimization::sgd, new NDList(), hyperparams, dataset, featureDim, numEpochs);
}
LossTime gdRes = trainSgd(1f, 1500, 10);
// -
// When the batch size equals 1, we use SGD for optimization. For simplicity of implementation we picked a constant (albeit small) learning rate. In SGD, the model parameters are updated whenever an example is processed. In our case this amounts to 1500 updates per epoch. As we can see, the decline in the value of the objective function slows down after one epoch. Although both the procedures processed 1500 examples within one epoch, SGD consumes more time than gradient descent in our experiment. This is because SGD updated the parameters more frequently and since it is less efficient to process single observations one at a time.
LossTime sgdRes = trainSgd(0.005f, 1, 2);
// Last, when the batch size equals 100, we use minibatch SGD for optimization. The time required per epoch is longer than the time needed for SGD and the time for batch gradient descent.
LossTime mini1Res = trainSgd(0.4f, 100, 2);
// Reducing the batch size to 10, the time for each epoch increases because the workload for each batch is less efficient to execute.
LossTime mini2Res = trainSgd(0.05f, 10, 2);
// Finally, we compare the time versus loss for the preview four experiments. As can be seen, despite SGD converges faster than GD in terms of number of examples processed, it uses more time to reach the same loss than GD because that computing gradient example by example is not efficient. Minibatch SGD is able to trade-off the convergence speed and computation efficiency. A minibatch size 10 is more efficient than SGD; a minibatch size 100 even outperforms GD in terms of runtime.
public String[] getTypeArray(LossTime lossTime, String name) {
String[] type = new String[lossTime.time.length];
for (int i = 0; i < type.length; i++) {
type[i] = name;
}
return type;
}
// Converts a float array to a log scale
float[] convertLogScale(float[] array) {
float[] newArray = new float[array.length];
for (int i = 0; i < array.length; i++) {
newArray[i] = (float) Math.log10(array[i]);
}
return newArray;
}
float[] time = ArrayUtils.addAll(ArrayUtils.addAll(gdRes.time, sgdRes.time),
ArrayUtils.addAll(mini1Res.time, mini2Res.time));
float[] loss = ArrayUtils.addAll(ArrayUtils.addAll(gdRes.loss, sgdRes.loss),
ArrayUtils.addAll(mini1Res.loss, mini2Res.loss));
String[] type = ArrayUtils.addAll(ArrayUtils.addAll(getTypeArray(gdRes, "gd"),
getTypeArray(sgdRes, "sgd")),
ArrayUtils.addAll(getTypeArray(mini1Res, "batch size = 100"),
getTypeArray(mini1Res, "batch size = 10")));
Table data = Table.create("data")
.addColumns(
DoubleColumn.create("log time (sec)", Functions.floatToDoubleArray(convertLogScale(time))),
DoubleColumn.create("loss", Functions.floatToDoubleArray(loss)),
StringColumn.create("type", type)
);
LinePlot.create("loss vs. time", data, "log time (sec)", "loss", "type");
// ## Concise Implementation
//
// In DJL, we can use the `Optimizer` package to access different optimization algorithms. This is used to implement a generic training function. We will use this throughout the current chapter.
public static void trainConciseCh11(Optimizer sgd, AirfoilRandomAccess dataset,
int numEpochs) {
// Initialization
NDManager manager = NDManager.newBaseManager();
SequentialBlock net = new SequentialBlock();
Linear linear = Linear.builder().setUnits(1).build();
net.add(linear);
net.setInitializer(new NormalInitializer());
Model model = Model.newInstance("concise implementation");
model.setBlock(net);
Loss loss = Loss.l2Loss();
DefaultTrainingConfig config = new DefaultTrainingConfig(loss)
.optOptimizer(sgd)
.addEvaluator(new Accuracy()) // Model Accuracy
.addTrainingListeners(TrainingListener.Defaults.logging()); // Logging
Trainer trainer = model.newTrainer(config);
int n = 0;
StopWatch stopWatch = new StopWatch();
stopWatch.start();
trainer.initialize(new Shape(10, 5));
Metrics metrics = new Metrics();
trainer.setMetrics(metrics);
float lastLoss = -1;
ArrayList<Double> lossArray = new ArrayList<>();
ArrayList<Double> epochArray = new ArrayList<>();
for (Batch batch : trainer.iterateDataset(dataset)) {
int len = (int) dataset.size() / batch.getSize(); // number of batches
NDArray X = batch.getData().head();
EasyTrain.trainBatch(trainer, batch);
trainer.step();
n += X.getShape().get(0);
if (n % 200 == 0) {
stopWatch.stop();
stopWatch.stop();
lastLoss = evaluateLoss(dataset.getData(manager), linear.getParameters().get(0).getValue().getArray()
.reshape(new Shape(dataset.getFeatureArraySize(), 1)),
linear.getParameters().get(1).getValue().getArray());
lossArray.add((double) lastLoss);
double lastEpoch = 1.0 * n / X.getShape().get(0) / len;
epochArray.add(lastEpoch);
stopWatch.start();
}
batch.close();
}
plotLossEpoch(arrayListToFloat(lossArray), arrayListToFloat(epochArray));
System.out.printf("loss: %.3f, %.3f sec/epoch\n", lastLoss, stopWatch.avg());
}
// Using DJL to repeat the last experiment shows identical behavior.
// +
AirfoilRandomAccess airfoilDataset = getDataCh11(10, 1500);
LearningRateTracker lrt = LearningRateTracker.fixedLearningRate(0.05f);
Optimizer sgd = Optimizer.sgd().setLearningRateTracker(lrt).build();
trainConciseCh11(sgd, airfoilDataset, 2);
// -
// ## Summary
//
// * Vectorization makes code more efficient due to reduced overhead arising from the deep learning framework and due to better memory locality and caching on CPUs and GPUs.
// * There is a trade-off between statistical efficiency arising from SGD and computational efficiency arising from processing large batches of data at a time.
// * Minibatch stochastic gradient descent offers the best of both worlds: computational and statistical efficiency.
// * In minibatch SGD we process batches of data obtained by a random permutation of the training data (i.e., each observation is processed only once per epoch, albeit in random order).
// * It is advisable to decay the learning rates during training.
// * In general, minibatch SGD is faster than SGD and gradient descent for convergence to a smaller risk, when measured in terms of clock time.
//
// ## Exercises
//
// 1. Modify the batch size and learning rate and observe the rate of decline for the value of the objective function and the time consumed in each epoch.
// 1. Read the DJL documentation and explore the different learning rate trackers in `ai.djl.training.optimizer.learningrate` to see how they affect training. Try using a `FactorTracker` to reduce the learning rate to 1/10 of its previous value after each epoch.
// 1. Compare minibatch SGD with a variant that actually *samples with replacement* from the training set. What happens?
// 1. An evil genie replicates your dataset without telling you (i.e., each observation occurs twice and your dataset grows to twice its original size, but nobody told you). How does the behavior of SGD, minibatch SGD and that of gradient descent change?
| jupyter/d2l-java/chapter_optimization/minibatch-sgd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <NAME> - UPV/EHU 2016
# +
##################################################################################################################
### WISE utilities:
##################################################################################################################
def download_wise_fits(img_id, img_ra, img_dec, img_arcsec, wise_band):
"""
fits file info: ( hdu_list.info() )
No. Name Type Cards Dimensions Format
0 PRIMARY PrimaryHDU 67 (XXX, XXX) float32
"""
fits_file = '%s/FITS/IMG_%06d_WISE,_%04d_arcsec,_band_%i.fits' % (raw_dir, img_id, img_arcsec, wise_band)
temp_dir = '%s/Temp_files' % data_dir
count = 0
while (not os.path.isfile(fits_file)) and (count < 50):
count += 1
try:
base_url = 'http://irsa.ipac.caltech.edu/ibe/search/wise/allwise/p3am_cdd?columns=coadd_id,'
query_band = 'band&where=band%20IN%20' + '(%i)' % (wise_band)
query_pos = '&POS=%.4f,%.4f&INTERSECT=CENTER&mcen' % (img_ra, img_dec)
final_url = base_url + query_band + query_pos
print 'Query URL:\n', final_url
urllib.urlretrieve(final_url, '%s/query_url.dat' % temp_dir)
print '\nData received:'
# !cat '$temp_dir/query_url.dat'
# !tail -n 1 '$temp_dir/query_url.dat' > '$temp_dir/query_data.dat'
meta_data = np.genfromtxt('%s/query_data.dat' % temp_dir, dtype='str')
params = {'coadd_id':meta_data[-2] , 'band':wise_band}
params['coaddgrp'] = params['coadd_id'][:2]
params['coadd_ra'] = params['coadd_id'][:4]
path = str.format('{coaddgrp:s}/{coadd_ra:s}/{coadd_id:s}/{coadd_id:s}-w{band:1d}-int-3.fits?',**params)
cutoff_url = 'center=%.4f,%.4fdeg&size=%i,%iarcsec&gzip=false' % (img_ra, img_dec, img_arcsec, img_arcsec)
img_url = 'http://irsa.ipac.caltech.edu/ibe/data/wise/allwise/p3am_cdd/' + path + cutoff_url
print '\nDownloading fits image from:\n%s' % img_url
urllib.urlretrieve(img_url, fits_file)
print '\nFits image saved in:\n%s\n\n******************************\n' % fits_file
except:
pass
def get_wise_image_data(img_id, img_arcsec, wise_band):
fits_file = '%s/WISE/FITS/IMG_%06d_WISE,_%04d_arcsec,_band_%i.fits' % (raw_dir, img_id, img_arcsec, wise_band)
img_data = fits.getdata(fits_file)
img_data = np.rot90(np.rot90(np.fliplr(img_data)))
img_data = np.array(img_data, dtype=np.float)
return img_data
def wise_fits_to_png(img_id, img_arcsec, img_pix, wise_band, circles=False, **circles_kwargs):
png_file = '%s/WISE/PNG/IMG_%06d_WISE,_%04d_arcsec,_band_%i.png' % (raw_dir, img_id, img_arcsec, wise_band)
img_data = get_wise_image_data(img_id, img_arcsec, wise_band)
if not circles:
fig = plot_image_data(img_data)
elif circles:
fig = plot_image_data_circles(img_data, img_arcsec, circles_kwargs['r_bins'])
fig.savefig(png_file)
##################################################################################################################
### Planck utilities:
##################################################################################################################
def read_planck_map(map_file, display=False):
fits_file = '%s/Planck/FITS_maps/%s' % (raw_dir, map_file)
planck_map = hp.read_map(fits_file)
Nside = np.int(np.sqrt(planck_map.size/12.))
if display:
hp.mollview(planck_map, nest=False, title=map_file)
return planck_map, Nside
def get_planck_patch(planck_map, clu_id, img_pix, zoom_factor):
Nside = np.int(np.sqrt(planck_map.size/12.))
clu = Cluster(redm, clu_id)
clu_long = clu.l.radian # l: gal longitude, in radians
clu_lat = clu.b.radian # b: gal latitude, in radians
map2D = np.zeros((img_pix, img_pix))
ix, iy = np.ogrid[0:img_pix, 0:img_pix]
X = 2.2/zoom_factor*(ix - img_pix/2.)/img_pix
Y = 2.2/zoom_factor*(iy - img_pix/2.)/img_pix
rho = np.sqrt(X**2 + Y**2) + 10.**(-9)
C = np.arcsin(rho)
Arg1 = np.cos(C)*np.sin(clu_lat)
Arg2 = (Y*np.sin(C)*np.cos(clu_lat))/rho
Arg3 = rho*np.cos(clu_lat)*np.cos(C) - Y*np.sin(clu_lat)*np.sin(C)
Arg4 = X*np.sin(C)
Arg5 = Arg4/Arg3
pix_lat = np.arcsin(Arg1 + Arg2)
pix_long = clu_long + np.arctan(Arg5)
fix_mask = Arg3 <= 0
pix_long[fix_mask] = np.pi + pix_long[fix_mask]
theta_ring = np.pi/2.0 - pix_lat
phi_ring = pix_long
iring = ang2pix(Nside, theta_ring, phi_ring, nest=False)
map2D = planck_map[iring]
StartPhi = pix_lat[img_pix/2,0]*(180.0/np.pi)
EndPhi = pix_lat[img_pix/2,-1]*(180.0/np.pi)
FOV_deg = np.abs(EndPhi - StartPhi)
map2D = np.array(map2D, dtype=np.float16)
return map2D, FOV_deg
def cluster_patch(clu_id, band, zoom_factor):
patch_file = '%s/Planck_patchs/%s/Patch_%06d_%02d.npy' % (data_dir, band, clu_id, zoom_factor)
if os.path.isfile(patch_file):
map2D = np.load(patch_file)
map2D = np.array(map2D, dtype=np.float)
return map2D
elif not os.path.isfile(patch_file):
print '***ERROR: patch not found...***'
##################################################################################################################
### SDSS, Planck & ROSAT utilities:
##################################################################################################################
#def download_sdss(img_dir, img_redm_id, img_ra, img_dec, img_scale, img_arcsec, img_pix_size):
#
# base_url = 'http://skyservice.pha.jhu.edu/DR12/ImgCutout/getjpeg.aspx?'
# query_url = 'ra=%.10f&dec=%.10f&scale=%.4f&width=%i&height=%i' % (img_ra, img_dec, img_scale, img_pix_size, img_pix_size)
# img_url = base_url + query_url
# urllib.urlretrieve(img_url, '%s/SDSS/IMG_%06d_SDSS,_%i_arcsec.png' % (img_dir, img_redm_id, img_arcsec))
#def process_planck(img_dir, img_id, img_pix_size):
#
# raw_file = '%s/Planck/RAW/SZ_%06d' % (img_dir, img_id)
# png_file = '%s/Planck/PNG/IMG_%06d.png' % (img_dir, img_id)
#
# if os.path.isfile(raw_file):
# img = Image.open(raw_file)
# img = img.resize((img_pix_size, img_pix_size), Image.ANTIALIAS)
# img.save(png_file)
#def process_rosat(img_dir, img_id, img_pix_size):
#
# raw_file = '%s/ROSAT/RAW/ROSAT_R6_%06d.gif' % (img_dir, img_id)
# png_file = '%s/ROSAT/PNG/IMG_ROSAT_R6_%06d.png' % (img_dir, img_id)
#
# if os.path.isfile(raw_file):
# img = Image.open(raw_file)
# img = img.resize((img_pix_size, img_pix_size), Image.ANTIALIAS)
# img.save(png_file)
##################################################################################################################
### WISE flux & light analysis:
##################################################################################################################
def radial_binning(min_arcsec, max_arcsec, nbins):
r_bins = np.linspace(min_arcsec, max_arcsec, nbins+1)
return r_bins
def compute_flux_distri(img_data, img_arcsec, r_bins):
arcsecs_in_pix = img_arcsec/img_data.shape[0]
lx, ly = img_data.shape
X, Y = np.ogrid[0:lx, 0:ly]
dist_matrix = distances_image(img_pix)*arcsecs_in_pix ## distances from center in arcsecs
fluxes = []
distances = (r_bins[:-1] + r_bins[1:])/2.
for n in range(len(r_bins)-1):
r_min = r_bins[n]
r_max = r_bins[n+1]
radial_mask = (dist_matrix >= r_min) & (dist_matrix < r_max)
radial_area = np.pi*(r_max**2-r_min**2)
flux = img_data[radial_mask].sum()/radial_area
fluxes.append(flux)
return np.array(fluxes)
def compute_light_distri(img_data, img_arcsec, r_bins):
arcsecs_in_pix = img_arcsec/image_data.shape[0]
lx, ly = img_data.shape
X, Y = np.ogrid[0:lx, 0:ly]
dist_matrix = np.sqrt((X - lx/2)**2 + (Y - ly/2)**2)*arcsecs_in_pix ## distances from center in arcsecs
lights = []
distances = (r_bins[:-1] + r_bins[1:])/2.
for n in range(len(r_bins)-1):
r_max = r_bins[n+1]
radial_mask = (dist_matrix < r_max)
radial_area = np.pi*(r_max**2)
light = img_data[radial_mask].sum()/radial_area
lights.append(light)
return np.array(lights)
def plot_flux_distribution(out_dir, img_redm_id, wise_bands, fluxes, cent_fit, back_fit, r_bins):
"""Flux of cluster"""
out_file = '%s/FLUX/%06d_flux.png' % (out_dir, img_redm_id)
distances = (r_bins[:-1] + r_bins[1:])/2.
dist_fit = np.arange(distances[0], distances[-1], 1.)
colors = ['red', 'black', 'blue']
freqs = ['3.4', '4.6', '12', '22']
figsizex, figsizey = 6.5, 4
fig = plt.figure(figsize=(figsizex, figsizey))
ax = fig.add_subplot(1, 1, 1)
for band in range(len(wise_bands)):
plt.plot(distances, fluxes[band], color=colors[band], label='W%i: %s $\mu$m' % (wise_bands[band], freqs[band]))
plt.plot(dist_fit, back_fit + func_fit(dist_fit, *cent_fit), 'k', label='fit flux')
plt.plot(dist_fit, np.full(len(dist_fit), back_fit), 'g:', label='fit background', linewidth=3)
plt.plot(dist_fit, func_fit(dist_fit, *cent_fit), 'b:', label='fit exp', linewidth=3)
plt.grid(alpha=0.5)
plt.xlabel(r'r [arcsecs]')
plt.ylabel(r'flux [DI arsec$^{-2}$]')
ax.set_xlim([r_bins[0], r_bins[-1]])
#ax.set_xscale('log')
#ax.set_yscale('log')
ax.set_ylim([0.,6.])
ax.set_xlim([0.,120.])
plt.legend(loc='upper right', frameon=False, labelspacing=0.5)
fig.savefig(out_file)
plt.close()
def plot_light_distribution(out_dir, img_redm_id, wise_bands, lights, cent_fit, back_fit, fit_light, half_light, r_bins):
"""Flux of cluster"""
out_file = '%s/LIGHT/%06d_light.png' % (out_dir, img_redm_id)
distances = (r_bins[:-1] + r_bins[1:])/2.
dist_fit = np.arange(distances[0], distances[-1], 0.1)
colors = ['red', 'black', 'blue']
freqs = ['3.4', '4.6', '12', '22']
figsizex, figsizey = 6.5, 4
fig = plt.figure(figsize=(figsizex, figsizey))
ax = fig.add_subplot(1, 1, 1)
for band in range(len(wise_bands)):
plt.plot(distances, lights[band], color=colors[band], label='W%i: %s $\mu$m' % (wise_bands[band], freqs[band]))
plt.plot(dist_fit, fit_light, 'k', label='fit flux')
plt.plot(dist_fit, np.full(len(dist_fit), back_fit), 'g:', label='fit background', linewidth=3)
plt.plot([half_light, half_light], [0., 10000.], 'b', label='half-light', linewidth=1)
plt.grid(alpha=0.5)
plt.xlabel(r'r [arcsecs]')
plt.ylabel(r'Surface brightness [DI arcecs$^{-2}$]')
ax.set_xlim([r_bins[0], r_bins[-1]])
#ax.set_xscale('log')
#ax.set_yscale('log')
ax.set_ylim([0.,6.])
ax.set_xlim([0.,120.])
plt.legend(loc='upper right', frameon=False, labelspacing=0.5)
fig.savefig(out_file)
plt.close()
def func_fit(x, *p):
a, b = p
return a*np.exp(-x/b)
def fit_flux_distri(fluxes, r_bins):
distances = (r_bins[:-1] + r_bins[1:])/2.
back_mask = distances >= 80.
back_fit = np.median(fluxes[back_mask])
err = lambda p: np.mean((func_fit(distances,*p)-(fluxes-back_fit))**2)
p_init = [1., 25]
cent_fit = minimize(err, p_init, bounds=[(0., None),(0., None)], method="L-BFGS-B").x
chi2 = err(cent_fit)
#p_opt = curve_fit(func, distances, fluxes, maxfev=100000)[0]
return cent_fit, back_fit, chi2
def check_light_distri(lights, cent_fit, back_fit, r_bins):
distances = (r_bins[:-1] + r_bins[1:])/2.
dist_fit = np.arange(distances[0], distances[-1], 0.1)
total_light = []
total_back_light = []
for d in range(len(dist_fit)):
bin_light = integrate.quad(lambda x: (back_fit + func_fit(x,*cent_fit))*2*np.pi*x, 0., dist_fit[d])[0]
bin_back_light = integrate.quad(lambda x: (back_fit)*2*np.pi*x, 0., dist_fit[d])[0]
total_light.append(bin_light/(np.pi*dist_fit[d]**2))
total_back_light.append(bin_back_light/(np.pi*dist_fit[d]**2))
fit_light = np.array(total_light)
sum_light = 0.
for d in range(len(dist_fit)):
sum_light += fit_light[d]
if (sum_light > (fit_light.sum() - np.array(total_back_light).sum())/2.):
half_light = dist_fit[d]
break
return fit_light, half_light
def process_half_light(z, angle_arcsecs):
arcsecs_in_rad = 360.*3600./(2.*np.pi)
ad_dist = Planck13.angular_diameter_distance(z).value
physical_transverse_dist = ad_dist*(angle_arcsecs/arcsecs_in_rad)
return physical_transverse_dist
| Python_scripts/Utilities_satellites.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Novelty
#
# ### CARLE uses two novelty-based reward proxies: Random Network Distillation and Autoencoder Loss
#
#
# <img src="../../assets/oscillator_policy_life_ae_rnd.gif" width=60%>
#
# <NAME> et al. “Exploration by Random Network Distillation.” ArXiv abs/1810.12894 (2019): n. pag.
# ### In CARLE's implementation, the autoencoder loss reward uses a fully convolutional network while the random network distillation random and prediction networks each use a fully connected head. This gives the AE reward translation invariance while the RND reward depends partially on where a pattern appears and jumps for edge crossings.
| notebooks/ieee_cog_2021/slide_6.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Literal Includes
#
# Testing literal Includes
# + hide-output=false
import numpy as np
a = np.random.rand(1000)
print(a)
# -
# should list a python highlighted program
# + [markdown] hide-output=false
# ```julia
# # function to calculate the volume of a sphere
# function sphere_vol(r)
# # julia allows Unicode names (in UTF-8 encoding)
# # so either "pi" or the symbol π can be used
# return 4/3*pi*r^3
# end
# ```
#
# -
# this is a julia highlighted program but is in a markdown cell as the default language for the notebook is **python**
#
# Literal includes can also have hidden output
# + hide-output=true
import numpy as np
a = np.random.rand(1000)
print(a)
| tests/base/ipynb/literal_include.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import, division, print_function
import tensorflow as tf
import numpy as np
# +
# MNIST dataset parameters
num_classes = 10 # total classes (0-9 digits).
num_features = 784 # data features (img shape: 28 * 28)
# Training parameters
learning_rate = 0.001
training_steps = 3000
batch_size = 256
display_step = 100
# Network parameters.
n_hidden_1 = 128 # 1st layer number of neurons.
n_hidden_2 = 256 # 2nd layer number of neurons.
# -
# Prepare MNIST data.
from tensorflow.keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# Convert to float32.
x_train, x_test = np.array(x_train, np.float32), np.array(x_test, np.float32)
# Flatten images to 1-D vector of 784 features (28*28)
x_train, x_test = x_train.reshape([-1, num_features]), x_test.reshape([-1, num_features])
# Normalize images value from [0, 255] to [0, 1].
x_train, x_test = x_train / 255., x_test / 255.
# Use tf.data API to shuffle and batch data.
train_data = tf.data.Dataset.from_tensor_slices((x_train, y_train))
train_data = train_data.repeat().shuffle(5000).batch(batch_size).prefetch(1)
# +
# Store layers weight & bias
# A random value generator to initialize weights.
random_normal = tf.initializers.RandomNormal()
weights = {
'h1' : tf.Variable(random_normal([num_features, n_hidden_1])),
'h2' : tf.Variable(random_normal([n_hidden_1, n_hidden_2])),
'out' : tf.Variable(random_normal([n_hidden_2, num_classes]))
}
biases = {
'b1' : tf.Variable(tf.zeros([n_hidden_1])),
'b2' : tf.Variable(tf.zeros([n_hidden_2])),
'out' : tf.Variable(tf.zeros([num_classes]))
}
# -
# Create model.
def neural_net(x):
# Hidden fully connected layer with 128 neurons.
layer_1 = tf.add(tf.matmul(x, weights['h1']), biases['b1'])
# Apply sigmoid to layer_1 output for non-linearity.
layer_1 = tf.nn.sigmoid(layer_1)
# Hidden fully connected layer with 256 neurons.
layer_2 = tf.add(tf.matmul(layer_1, weights['h2']), biases['b2'])
# Apply sigmoid to layer_2 output for non-linearity.
layer_2 = tf.nn.sigmoid(layer_2)
# Output fully connected layer with a neuron for each class.
out_layer = tf.matmul(layer_2, weights['out']) + biases['out']
# Apply softmax to normalize the logits to a probability distribution.
return tf.nn.softmax(out_layer)
# +
# Cross-Entropy loss function.
def cross_entropy(y_pred, y_true):
# Encode label to a one hot vector.
y_true = tf.one_hot(y_true, depth=num_classes)
# Clip prediction values to avoid log(0) error.
y_pred = tf.clip_by_value(y_pred, 1e-9, 1.)
# Compute cross-entropy.
return tf.reduce_mean(-tf.reduce_sum(y_true * tf.math.log(y_pred)))
# Accuracy metric.
def accuracy(y_pred, y_true):
# Predicted class is the index of highest score in prediction vector (i.e. argmax).
correct_prediction = tf.equal(tf.argmax(y_pred, 1), tf.cast(y_true, tf.int64))
return tf.reduce_mean(tf.cast(correct_prediction, tf.float32), axis=-1)
# Stochastic gradient descent optimizer.
optimizer = tf.optimizers.SGD(learning_rate)
# -
# Optimization process.
def run_optimization(x, y):
# Wrap computation inside a GradientTape for automatic differentiation.
with tf.GradientTape() as g:
pred = neural_net(x)
loss = cross_entropy(pred, y)
# Variables to update, i.e. trainable variables.
trainable_variables = list(weights.values()) + list(biases.values())
# Compute gradients.
gradients = g.gradient(loss, trainable_variables)
# Update W and b following gradients.
optimizer.apply_gradients(zip(gradients, trainable_variables))
# + jupyter={"source_hidden": true}
# Run training for the given number of steps.
for step, (batch_x, batch_y) in enumerate(train_data.take(training_steps), 1):
# Run the optimization to update W and b values.
run_optimization(batch_x, batch_y)
if step % display_step == 0:
pred = neural_net(batch_x)
loss = cross_entropy(pred, batch_y)
acc = accuracy(pred, batch_y)
print("step: %i, loss: %f, accuracy: %f" % (step, loss, acc))
# -
# Test model on validation set.
pred = neural_net(x_test)
print("Test Accuracy: %f" % accuracy(pred, y_test))
# VIsualize predictions.
import matplotlib.pyplot as plt
# +
# Predict 5 images from validation set.
n_images = 5
test_images = x_test[:n_images]
predictions = neural_net(test_images)
# Display image and model prediction.
for i in range(n_images):
plt.imshow(np.reshape(test_images[i], [28, 28]), cmap='gray')
plt.show()
print("Model prediction: %i" % np.argmax(predictions.numpy()[i]))
# -
| neural_network_raw_typing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_style("whitegrid")
# -
# # Model: xgboost
# +
from sklearn.metrics import classification_report, confusion_matrix, plot_confusion_matrix, roc_auc_score, roc_curve, precision_recall_curve
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.compose import ColumnTransformer
from sklearn.model_selection import train_test_split
from xgboost.sklearn import XGBClassifier
# -
# ## Data
#
# Load the dataset, applying no major transformations to it.
data = pd.read_csv('../dataset/creditcard.csv')
data.head()
X = data.drop(columns=['Class'])
y = data['Class']
# Since the data is largely unbalanced we must use a stratified sampling to make sure we get both negative and positive samples to train with.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=0, stratify=y)
# ## Pipeline (build)
# +
numeric_feature_indexes = slice(0, 30)
preprocessor = ColumnTransformer(
transformers=[
('num', StandardScaler(), numeric_feature_indexes),
])
pipeline = Pipeline([
('preprocessor', preprocessor),
('classifier', XGBClassifier())
])
# +
num_features_type_map = {feature: 'float64' for feature in X_train.columns[numeric_feature_indexes]}
X_train = X_train.astype(num_features_type_map)
X_test = X_test.astype(num_features_type_map)
# -
# ## Pipeline (train)
model = pipeline.fit(X_train, y_train, classifier__eval_metric='auc')
model
# ## Pipeline (evaluate)
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
disp = plot_confusion_matrix(model, X_test, y_test, display_labels=['normal', 'fraudulent'], cmap=plt.cm.Blues)
disp.ax_.grid(False)
# Some great material is available here: https://machinelearningmastery.com/roc-curves-and-precision-recall-curves-for-classification-in-python/
# +
y_pred_proba = pipeline.predict_proba(X_test)[::,1]
fpr, tpr, _ = roc_curve(y_test, y_pred_proba)
auc = roc_auc_score(y_test, y_pred_proba)
fig, ax = plt.subplots(figsize=(5,5))
ax.plot(fpr,tpr,label=f"auc {auc:2.2f}")
ax.legend(loc=4)
ax.set_xlabel('False Positive Rate')
ax.set_ylabel('True Positive Rate');
# +
precision, recall, _ = precision_recall_curve(y_test, y_pred_proba)
fig, ax = plt.subplots(figsize=(5,5))
no_skill = len(y_test[y_test==1]) / len(y_test)
ax.plot([0, 1], [no_skill, no_skill], linestyle='--', label='No Skill')
ax.plot(recall, precision)
ax.set_xlabel('Precision')
ax.set_ylabel('Recall');
| classification-fraud-detection/solution-2-service-composition/2.0 model [sklearn-xgboost].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py3)
# language: python
# name: py3
# ---
# # DL-Tabular
# For this purpose, we will use the heart disease UCI data
# +
# https://towardsdatascience.com/pytorch-tabular-binary-classification-a0368da5bb89
# https://towardsdatascience.com/pytorch-basics-intro-to-dataloaders-and-loss-functions-868e86450047
# +
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
import seaborn as sns
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
import shap
# -
data = pd.read_csv("C:/Users/joaof/Downloads/processed.cleveland.data",header=None)
headers=["age","sex","cp","trestbps","chol","fbs","restecg","thalach","exang","oldpeak","slope","ca","thal","num"]
data.columns=headers
data.head()
# Since we want to create a simple classification problem, we will transform the variable num into a dual category, presence (1,2,3,4) or absence disease (0)
data['target'] = data['num'].apply(lambda x: 1 if x>=1 else 0)
data.drop(columns=['num'],inplace=True)
data.describe(include="all")
data.info()
sns.countplot(x = 'target', data=data)
sns.countplot(x = 'ca', data=data)
sns.countplot(x = 'thal', data=data)
# since they are so low, lets remove the "?"
data.drop(data[data["thal"]=="?"].index,inplace=True)
data.drop(data[data["ca"]=="?"].index,inplace=True)
data.reset_index(inplace=True,drop=True)
y=data.iloc[:,-1]
X=data.iloc[:,0:-1]
X_train,X_test,y_train,y_test = train_test_split(X,y, test_size=0.2,random_state=42)
sns.countplot(x = y_train)
sns.countplot(x = y_train)
scaler = StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
len(X_train[0])
EPOCHS = 50
BATCH_SIZE = 64
LEARNING_RATE = 0.001
# +
#dataloader
## train data
class trainData(Dataset):
def __init__(self, X_data, y_data):
self.X_data = X_data
self.y_data = y_data
def __getitem__(self, index):
return self.X_data[index], self.y_data[index]
def __len__ (self):
return len(self.X_data)
train_data = trainData(torch.FloatTensor(X_train),
torch.FloatTensor(y_train))
## test data
class testData(Dataset):
def __init__(self, X_data):
self.X_data = X_data
def __getitem__(self, index):
return self.X_data[index]
def __len__ (self):
return len(self.X_data)
test_data = testData(torch.FloatTensor(X_test))
# -
train_loader = DataLoader(dataset=train_data, batch_size=BATCH_SIZE, shuffle=True)
test_loader = DataLoader(dataset=test_data, batch_size=1)
class binaryClassification(nn.Module):
def __init__(self):
super(binaryClassification, self).__init__()
# Number of input features is 12.
self.layer_1 = nn.Linear(13, 64)
self.layer_2 = nn.Linear(64, 64)
self.layer_out = nn.Linear(64, 1)
self.relu = nn.ReLU()
self.dropout = nn.Dropout(p=0.1)
self.batchnorm1 = nn.BatchNorm1d(64)
self.batchnorm2 = nn.BatchNorm1d(64)
def forward(self, inputs):
x = self.relu(self.layer_1(inputs))
x = self.batchnorm1(x)
x = self.relu(self.layer_2(x))
x = self.batchnorm2(x)
x = self.dropout(x)
x = self.layer_out(x)
return x
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
model = binaryClassification()
model.to(device)
print(model)
criterion = nn.BCEWithLogitsLoss()
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
def binary_acc(y_pred, y_test):
y_pred_tag = torch.round(torch.sigmoid(y_pred))
correct_results_sum = (y_pred_tag == y_test).sum().float()
acc = correct_results_sum/y_test.shape[0]
acc = torch.round(acc * 100)
return acc
model.train()
for e in range(1, EPOCHS+1):
epoch_loss = 0
epoch_acc = 0
for X_batch, y_batch in train_loader:
X_batch, y_batch = X_batch.to(device), y_batch.to(device)
optimizer.zero_grad()
y_pred = model(X_batch)
loss = criterion(y_pred, y_batch.unsqueeze(1))
acc = binary_acc(y_pred, y_batch.unsqueeze(1))
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
print(f'Epoch {e+0:03}: | Loss: {epoch_loss/len(train_loader):.5f} | Acc: {epoch_acc/len(train_loader):.3f}')
| classification-dl/classification-dl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#DATA generator
import random
def makeTerrainData(n_points=1000):
###############################################################################
### make the toy dataset
random.seed(42)
grade = [random.random() for ii in range(0,n_points)]
bumpy = [random.random() for ii in range(0,n_points)]
error = [random.random() for ii in range(0,n_points)]
y = [round(grade[ii]*bumpy[ii]+0.3+0.1*error[ii]) for ii in range(0,n_points)]
for ii in range(0, len(y)):
if grade[ii]>0.8 or bumpy[ii]>0.8:
y[ii] = 1.0
### split into train/test sets
X = [[gg, ss] for gg, ss in zip(grade, bumpy)]
split = int(0.75*n_points)
X_train = X[0:split]
X_test = X[split:]
y_train = y[0:split]
y_test = y[split:]
grade_sig = [X_train[ii][0] for ii in range(0, len(X_train)) if y_train[ii]==0]
bumpy_sig = [X_train[ii][1] for ii in range(0, len(X_train)) if y_train[ii]==0]
grade_bkg = [X_train[ii][0] for ii in range(0, len(X_train)) if y_train[ii]==1]
bumpy_bkg = [X_train[ii][1] for ii in range(0, len(X_train)) if y_train[ii]==1]
# training_data = {"fast":{"grade":grade_sig, "bumpiness":bumpy_sig}
# , "slow":{"grade":grade_bkg, "bumpiness":bumpy_bkg}}
grade_sig = [X_test[ii][0] for ii in range(0, len(X_test)) if y_test[ii]==0]
bumpy_sig = [X_test[ii][1] for ii in range(0, len(X_test)) if y_test[ii]==0]
grade_bkg = [X_test[ii][0] for ii in range(0, len(X_test)) if y_test[ii]==1]
bumpy_bkg = [X_test[ii][1] for ii in range(0, len(X_test)) if y_test[ii]==1]
test_data = {"fast":{"grade":grade_sig, "bumpiness":bumpy_sig}
, "slow":{"grade":grade_bkg, "bumpiness":bumpy_bkg}}
return X_train, y_train, X_test, y_test
# return training_data, test_data
# +
import matplotlib.pyplot as plt
import pylab as pl
import numpy as np
% matplotlib inline
def prettyPicture(clf, X_test, y_test):
x_min = 0.0; x_max = 1.0
y_min = 0.0; y_max = 1.0
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, m_max]x[y_min, y_max].
h = .01 # step size in the mesh
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.pcolormesh(xx, yy, Z, cmap=pl.cm.seismic)
# Plot also the test points
grade_sig = [X_test[ii][0] for ii in range(0, len(X_test)) if y_test[ii]==0]
bumpy_sig = [X_test[ii][1] for ii in range(0, len(X_test)) if y_test[ii]==0]
grade_bkg = [X_test[ii][0] for ii in range(0, len(X_test)) if y_test[ii]==1]
bumpy_bkg = [X_test[ii][1] for ii in range(0, len(X_test)) if y_test[ii]==1]
plt.scatter(grade_sig, bumpy_sig, color = "b", label="fast")
plt.scatter(grade_bkg, bumpy_bkg, color = "r", label="slow")
plt.legend()
plt.xlabel("bumpiness")
plt.ylabel("grade")
plt.savefig("test.png")
# -
# # Naive Bayes
# +
import numpy as np
import pylab as pl
features_train, labels_train, features_test, labels_test = makeTerrainData()
grade_fast = [features_train[ii][0] for ii in range(0, len(features_train)) if labels_train[ii]==0]
bumpy_fast = [features_train[ii][1] for ii in range(0, len(features_train)) if labels_train[ii]==0]
grade_slow = [features_train[ii][0] for ii in range(0, len(features_train)) if labels_train[ii]==1]
bumpy_slow = [features_train[ii][1] for ii in range(0, len(features_train)) if labels_train[ii]==1]
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
clf.fit(features_train, labels_train)
pred = clf.predict(features_test)
from sklearn.metrics import accuracy_score
print('ACCURACY:',accuracy_score(pred, labels_test)*100)
prettyPicture(clf, features_test, labels_test)
# output_image("test.png", "png", open("test.png", "rb").read())
# -
# # Support Vector Machine
# +
import matplotlib.pyplot as plt
import copy
import numpy as np
import pylab as pl
features_train, labels_train, features_test, labels_test = makeTerrainData()
from sklearn.svm import SVC
clf = SVC(C=1.0,kernel="rbf",gamma=700)
clf.fit(features_train, labels_train)
pred = clf.predict(features_test)
from sklearn.metrics import accuracy_score
acc = accuracy_score(pred, labels_test)
print('ACCURACY:',acc*100)
prettyPicture(clf, features_test, labels_test)
# -
| ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preparing the data
# +
# Importing the libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Loading the data
# https://archive.ics.uci.edu/ml/datasets/statlog+(australian+credit+approval)
df = pd.read_csv("Credit_Card_Applications.csv")
df.head()
# +
# Split the data
X = df.iloc[:, :-1].values
y = df.iloc[:, -1].values # Class if credit application is approved or not
# Feature Scaling
from sklearn.preprocessing import MinMaxScaler
mms = MinMaxScaler(feature_range = (0, 1))
X = mms.fit_transform(X)
# -
# # Self Organizing Map model
# ## Training the model
from minisom import MiniSom
# Initiliaze the SOM model
som = MiniSom(x = 10, y = 10, input_len = X.shape[1],
sigma = 1.0, learning_rate = 0.5,
random_seed = 123) # (x, y) = SOM output grid dimension
# Initiliaze random weights for the model
som.random_weights_init(data = X)
# Train the SOM model
som.train_random(data = X, num_iteration = 500)
# ## Visualising the results
# Depending on the winning node with a high Mean Interneuron Distance (MID), we will identify potential fraud cases. After visualising the SOM below, the outliers with white color and high MID and also got the approval of the bank represent the potential fraud cases.
plt.figure(figsize = (12, 10))
plt.bone() # Initiate the frame
plt.pcolor(som.distance_map().T) # Color the distance matrix
plt.colorbar() # Legend for colors
markers = ['o', 's'] # o : circle, s: square
colors = ['r', 'g'] # Red = rejected, Green = approved
for i, x in enumerate(X):
w = som.winner(x) # Take the customer's winning node coordinates
plt.plot(w[0] + 0.5, # + 0.5 for centrering the marker
w[1] + 0.5,
markers[y[i]],
markeredgecolor = colors[y[i]],
markerfacecolor = 'None',
markeredgewidth = 1.5,
markersize = 20)
plt.show()
# ## Finding the potential frauds
# According the mapping above, the outliers with high MIDs at (5, 6), (4, 7) and (5, 8) coordinates include potential fraud cases.
mappings = som.win_map(X)
frauds = np.concatenate((mappings[(5, 6)], mappings[(4, 7)], mappings[(5, 8)]))
frauds = mms.inverse_transform(frauds)
results = pd.DataFrame([int(i) for i in frauds[:, 0]],
columns = ['Fraud Customer ID'])
results.style
# # Building Artificial Neural Network
# +
# Creating the matrix of features
customers = df.iloc[:, 1:].values
# Creating the dependent variable
is_fraud = np.zeros(df.shape[0], dtype = np.int64)
for i in range(df.shape[0]):
if df.iloc[i, 0] in frauds:
is_fraud[i] = 1
# -
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
customers = sc.fit_transform(customers)
# +
import tensorflow as tf
import tensorflow.keras.layers as F
def build_ann():
model = tf.keras.models.Sequential()
model.add(F.Dense(units = 2, kernel_initializer = 'uniform', activation = 'relu', input_dim = customers.shape[1]))
model.add(F.Dense(units = 1, kernel_initializer = 'uniform', activation = 'sigmoid'))
model.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
return model
# -
# Training the ANN model
ann = build_ann()
ann.fit(customers, is_fraud, batch_size = 1, epochs = 3)
# Get the predictions and rank their probabilities
y_pred = ann.predict(customers)
y_pred = np.concatenate((df.iloc[:, 0:1].values, y_pred), axis = 1)
y_pred = y_pred[y_pred[:, 1].argsort()[::-1]]
probs_per_customer = pd.DataFrame(y_pred, columns = ['CustomerID', 'Fraud Probability'])
probs_per_customer['CustomerID'] = probs_per_customer['CustomerID'].astype(np.int64)
probs_per_customer.head(10).style
| Deep Learning/Self Organizing Maps/Credit Fraud Detection with SOM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
import tensorflow as tf
tf.__version__
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/home/victor/MNIST_data/", one_hot=True)
len(mnist.train.images)
# ### 2d tensor that will have the imae values
x = tf.placeholder(tf.float32, [None, 784])
# ### variables to be optimized, model to obtain
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# ### Implementing the model
# +
# y = tf.nn.softmax(tf.matmul(x, W) + b) #commented when using tf.nn.softmax_cross_entropy_with_logits()
# -
# ### a variable to use in loss function
y_ = tf.placeholder(tf.float32, [None, 10])
# ### Cross entropy as cost function
# +
#cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(y), reduction_indices=[1]))
# -
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y_, logits=y))
# ### Applying minimization to the loss function
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
# ### Launch the model
sess = tf.InteractiveSession()
# ### Initialize the variables
tf.global_variables_initializer().run()
# ### Train the model
for _ in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(100)
sess.run(train_step, feed_dict={x: batch_xs, y_: batch_ys})
# ## "Using small batches of random data is called stochastic training -- in this case, stochastic gradient descent"
batch_xs.shape #images
batch_ys.shape #label
# ### Evaluate the model
correct_prediction = tf.equal(tf.argmax(y,1), tf.argmax(y_,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print(sess.run(accuracy, feed_dict={x: mnist.test.images, y_: mnist.test.labels}))
# acurácia para a croos-entropy "manual" 0.9148
| MNIST_For_ML_Beginners.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
import numpy as np
import random
import string # to process standard python strings
# +
f=open('chatbot1.txt','r',errors = 'ignore')
raw=f.read()
raw=raw.lower()# converts to lowercase
nltk.download('punkt') # first-time use only
nltk.download('wordnet') # first-time use only
sent_tokens = nltk.sent_tokenize(raw)# converts to list of sentences
word_tokens = nltk.word_tokenize(raw)# converts to list of words
# +
sent_tokens[:2]
['a chatbot (also known as a talkbot, chatterbot, bot, im bot, interactive agent, or artificial conversational entity) is a computer program or an artificial intelligence which conducts a conversation via auditory or textual methods.',
'such programs are often designed to convincingly simulate how a human would behave as a conversational partner, thereby passing the turing test.']
word_tokens[:2]
['a', 'chatbot', '(', 'also', 'known']
# +
lemmer = nltk.stem.WordNetLemmatizer()
#WordNet is a semantically-oriented dictionary of English included in NLTK.
def LemTokens(tokens):
return [lemmer.lemmatize(token) for token in tokens]
remove_punct_dict = dict((ord(punct), None) for punct in string.punctuation)
def LemNormalize(text):
return LemTokens(nltk.word_tokenize(text.lower().translate(remove_punct_dict)))
# +
GREETING_INPUTS = ("hello", "hi", "greetings", "sup", "what's up","hey",)
GREETING_RESPONSES = ["hi", "hey", "*nods*", "hi there", "hello", "I am glad! You are talking to me"]
def greeting(sentence):
for word in sentence.split():
if word.lower() in GREETING_INPUTS:
return random.choice(GREETING_RESPONSES)
# -
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
# +
def response(user_response):
robo_response=''
sent_tokens.append(user_response)
TfidfVec = TfidfVectorizer(tokenizer=LemNormalize, stop_words='english')
tfidf = TfidfVec.fit_transform(sent_tokens)
vals = cosine_similarity(tfidf[-1], tfidf)
idx=vals.argsort()[0][-2]
flat = vals.flatten()
flat.sort()
req_tfidf = flat[-2]
if(req_tfidf==0):
robo_response=robo_response+"I am sorry! I don't understand you"
return robo_response
else:
robo_response = robo_response+sent_tokens[idx]
return robo_response
# -
'''flag=True
print("ROBO: My name is Robo. I will answer your queries about Chatbots. If you want to exit, type Bye!")
while(flag==True):
user_response = input()
user_response=user_response.lower()
if(user_response!='bye'):
if(user_response=='thanks' or user_response=='thank you' ):
flag=False
print("ROBO: You are welcome..")
else:
if(greeting(user_response)!=None):
print("ROBO: "+greeting(user_response))
else:
print("ROBO: ",end="")
print(response(user_response))
sent_tokens.remove(user_response)
else:
flag=False
print("ROBO: Bye! take care..")'''
response('i am worried about my future')
from GAD_FUTURE_EVENTS import future
future('future')
| ChatBot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# author: <NAME>
# -
# ## NumPy
#
# NumPy is the fundamental package for scientific computing with Python. It contains among other things:
#
# - a powerful N-dimensional array object
# - sophisticated (broadcasting) functions
# - tools for integrating C/C++ and Fortran code
# - useful linear algebra, Fourier transform, and random number capabilities
#
# Besides its obvious scientific uses, NumPy can also be used as an efficient multi-dimensional container of generic data. Arbitrary data-types can be defined.
# This allows NumPy to seamlessly and speedily integrate with a wide variety of databases.
#
# **Note:** NumPy stands for Numeric Python. It's easy, fast and allow calculations over entire arrays. Find more [here](http://www.numpy.org/)
# ## Basic usage
# let's import library first
import numpy as np
# from Python list to Numpy array
python_list = [1.2, 3.4, 6, 99]
np_array = np.array(python_list)
print(np_array)
# add/substract/divide/multiply by scalar
np_array+4.5, np_array-4.5, np_array/4.5, np_array*4.5
# subsetting
np_array[np_array<3]
# shape of array
print(np_array.shape) # column vector
# +
np.logical_and(np_array > 2, np_array < 20) # logical_or, logical_not is also available
# select result
print(np_array[np.logical_and(np_array > 2, np_array < 20)])
# -
# for loop over 2D numpy array (https://docs.scipy.org/doc/numpy/reference/arrays.nditer.html#arrays-nditer)
for val in np.nditer(np.array([[1,2,3], [3,2,1]])):
print(val)
# ## Basic Statistics
# let's generate some data
data_points = np.random.randint(200, size=100)
data_points
np.mean(data_points), np.median(data_points), np.std(data_points), np.max(data_points), np.min(data_points)
# ## Matrices (2D Arrays)
# +
# 2D array
np_array2D = np.array([[1,2,3], [3,2,1], [5,6,2]])
print(f'Our 2D array: \n {np_array2D}')
print(f'\nShape of 2D array is {np_array2D.shape}') # matrix
# slice 2D array
np_array2D_sliced = np_array2D[1:, :-1]
print(f'\nOur 2D array (first row and last column sliced): \n {np_array2D_sliced}')
# +
# matrix multiplication
A = np.random.randint(25, size=(3,3))
B = np.random.randint(45, size=(3,3))
print(f'A matrix : \n {A}')
print(f'\nB matrix : \n {B}')
print(f'\nA*B element-wise: \n {A*B}')
print(f'\nA*B matrix multiplication : \n {A.dot(B)}')
| Modules/1-Preparation/notebooks/intro_to_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# +
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, BatchNormalization, LocallyConnected2D, Permute
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
import keras.losses
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import pandas as pd
import os
import pickle
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
import isolearn.io as isoio
import isolearn.keras as isol
from genesis.visualization import *
from genesis.generator import *
from genesis.predictor import *
from genesis.optimizer import *
from definitions.generator.aparent_deconv_conv_generator_concat import load_generator_network, get_shallow_copy_function
from definitions.predictor.aparent_w_dense_functional import load_saved_predictor
import sklearn
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from scipy.stats import pearsonr
import seaborn as sns
from matplotlib import colors
class IdentityEncoder(iso.SequenceEncoder) :
def __init__(self, seq_len, channel_map) :
super(IdentityEncoder, self).__init__('identity', (seq_len, len(channel_map)))
self.seq_len = seq_len
self.n_channels = len(channel_map)
self.encode_map = channel_map
self.decode_map = {
nt: ix for ix, nt in self.encode_map.items()
}
def encode(self, seq) :
encoding = np.zeros((self.seq_len, self.n_channels))
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
return encoding
def encode_inplace(self, seq, encoding) :
for i in range(len(seq)) :
if seq[i] in self.encode_map :
channel_ix = self.encode_map[seq[i]]
encoding[i, channel_ix] = 1.
def encode_inplace_sparse(self, seq, encoding_mat, row_index) :
raise NotImplementError()
def decode(self, encoding) :
seq = ''
for pos in range(0, encoding.shape[0]) :
argmax_nt = np.argmax(encoding[pos, :])
max_nt = np.max(encoding[pos, :])
seq += self.decode_map[argmax_nt]
return seq
def decode_sparse(self, encoding_mat, row_index) :
raise NotImplementError()
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
# +
class GenesisMonitor(Callback):
def __init__(self, generator_model, sequence_encoder, run_dir="", run_prefix="", n_sequences=32, batch_size=32, input_tensor_funcs=None) :
self.generator_model = generator_model
self.batch_size = batch_size
self.n_sequences = n_sequences
self.input_tensor_funcs = input_tensor_funcs
self.sequence_encoder = sequence_encoder
self.run_prefix = run_prefix
self.run_dir = run_dir
if not os.path.exists(self.run_dir): os.makedirs(self.run_dir)
seqs = self._sample_sequences()
self._store_sequences(seqs, 0)
def _sample_sequences(self) :
n_batches = self.n_sequences // self.batch_size
self.input_tensors = [self.input_tensor_funcs[i](i) for i in range(len(self.input_tensor_funcs))]
gen_bundle = self.generator_model.predict(x=self.input_tensors, batch_size=self.batch_size)
_, _, _, _, _, sampled_pwm, _, _, _ = gen_bundle
seqs = [
self.sequence_encoder.decode(sampled_pwm[i, 0, :, :, 0]) for i in range(sampled_pwm.shape[0])
]
return seqs
def _store_sequences(self, seqs, epoch) :
#Save sequences to file
with open(self.run_dir + self.run_prefix + "_epoch_" + str(epoch) + "_" + str(self.n_sequences) + "_sequences.txt", "wt") as f:
for i in range(len(seqs)) :
f.write(seqs[i] + "\n")
def on_epoch_end(self, epoch, logs={}) :
seqs = self._sample_sequences()
self._store_sequences(seqs, epoch)
# +
def get_latent_margin_sample_cosine(margin=0.5) :
def latent_margin_sample_cosine(seed1, seed2) :
rescaled_seed1 = seed1#(seed1 + 1.) / 2.
rescaled_seed2 = seed2#(seed2 + 1.) / 2.
mean_sample_ent = K.sum(rescaled_seed1 * rescaled_seed2, axis=-1) / (K.sqrt(K.sum(rescaled_seed1**2, axis=-1)) * K.sqrt(K.sum(rescaled_seed2**2, axis=-1)))
margin_sample_ent = K.switch(mean_sample_ent > K.constant(margin, shape=(1,)), mean_sample_ent - margin, K.zeros_like(mean_sample_ent))
return margin_sample_ent
return latent_margin_sample_cosine
#Define target isoform loss function
def get_isoform_loss(target_isos, fitness_target=1.0, fitness_weight=2.0, isoform_start=80, isoform_end=115, use_start=0, use_end=70, use_target_bits=1.8, cse_start=70, cse_end=76, cse_target_bits=1.8, dse_start=76, dse_end=125, dse_target_bits=1.8, entropy_weight=0.0, entropy_loss_mode='margin', sequence_similarity_weight=0.0, sequence_similarity_margin=0.0, dense_1_similarity_weight=0.0, dense_1_similarity_margin=0.0, punish_dn_cse=0.0, punish_up_c=0.0, punish_dn_c=0.0, punish_up_g=0.0, punish_dn_g=0.0, punish_up_aa=0.0, punish_dn_aa=0.0) :
target_iso = np.zeros((len(target_isos), 1))
for i, t_iso in enumerate(target_isos) :
target_iso[i, 0] = t_iso
masked_use_entropy_mse = get_target_entropy_sme_masked(pwm_start=use_start, pwm_end=use_end, target_bits=use_target_bits)
cse_entropy_mse = get_target_entropy_sme(pwm_start=cse_start, pwm_end=cse_end, target_bits=cse_target_bits)
masked_dse_entropy_mse = get_target_entropy_sme_masked(pwm_start=dse_start, pwm_end=dse_end, target_bits=dse_target_bits)
if entropy_loss_mode == 'margin' :
masked_use_entropy_mse = get_margin_entropy_ame_masked(pwm_start=use_start, pwm_end=use_end, min_bits=use_target_bits)
cse_entropy_mse = get_margin_entropy_ame(pwm_start=cse_start, pwm_end=cse_end, min_bits=cse_target_bits)
masked_dse_entropy_mse = get_margin_entropy_ame_masked(pwm_start=dse_start, pwm_end=dse_end, min_bits=dse_target_bits)
punish_dn_cse_func = get_punish_cse(pwm_start=74, pwm_end=dse_end)
punish_up_c_func = get_punish_c(pwm_start=use_start, pwm_end=use_end)
punish_dn_c_func = get_punish_c(pwm_start=dse_start, pwm_end=dse_end)
punish_up_g_func = get_punish_g(pwm_start=use_start, pwm_end=use_end)
punish_dn_g_func = get_punish_g(pwm_start=use_start, pwm_end=use_end)
punish_up_aa_func = get_punish_aa(pwm_start=use_start, pwm_end=use_end)
punish_dn_aa_func = get_punish_aa(pwm_start=dse_start, pwm_end=dse_end)
pwm_sample_entropy_func = get_pwm_margin_sample_entropy_masked(pwm_start=70-60, pwm_end=76+60, margin=sequence_similarity_margin, shift_1_nt=True)
dense_1_sample_entropy_func = get_latent_margin_sample_cosine(margin=dense_1_similarity_margin)
def loss_func(loss_tensors) :
_, _, _, sequence_class, pwm_logits_1, pwm_logits_2, pwm_1, pwm_2, sampled_pwm_1, sampled_pwm_2, mask, sampled_mask, iso_pred_1, _, iso_score_pred_1, _, dense_1_pred_1, iso_pred_2, _, _, _, dense_1_pred_2 = loss_tensors
#Create target isoform with sample axis
iso_targets = K.constant(target_iso)
iso_true = K.gather(iso_targets, sequence_class[:, 0])
iso_true = K.tile(K.expand_dims(iso_true, axis=-1), (1, K.shape(sampled_pwm_1)[1], 1))
#Specify costs
iso_loss = fitness_weight * K.mean(K.maximum(-K.print_tensor(iso_score_pred_1[..., 0], message="iso_score_pred_1=") + fitness_target, K.zeros_like(iso_score_pred_1[..., 0])), axis=1)
seq_loss = 0.0
seq_loss += punish_dn_cse * K.mean(punish_dn_cse_func(sampled_pwm_1), axis=1)
seq_loss += punish_up_c * K.mean(punish_up_c_func(sampled_pwm_1), axis=1)
seq_loss += punish_dn_c * K.mean(punish_dn_c_func(sampled_pwm_1), axis=1)
seq_loss += punish_up_g * K.mean(punish_up_g_func(sampled_pwm_1), axis=1)
seq_loss += punish_dn_g * K.mean(punish_dn_g_func(sampled_pwm_1), axis=1)
seq_loss += punish_up_aa * K.mean(punish_up_aa_func(sampled_pwm_1), axis=1)
seq_loss += punish_dn_aa * K.mean(punish_dn_aa_func(sampled_pwm_1), axis=1)
entropy_loss = entropy_weight * ((masked_use_entropy_mse(pwm_1, mask) if use_target_bits is not None else 0.0) + (cse_entropy_mse(pwm_1) if cse_target_bits is not None else 0.0) + (masked_dse_entropy_mse(pwm_1, mask) if dse_target_bits is not None else 0.0))
sequence_similarity_loss = sequence_similarity_weight * K.mean(pwm_sample_entropy_func(sampled_pwm_1, sampled_pwm_2, sampled_mask), axis=1)
dense_1_similarity_loss = dense_1_similarity_weight * K.mean(dense_1_sample_entropy_func(dense_1_pred_1, dense_1_pred_2), axis=1)
#Compute total loss
total_loss = iso_loss + seq_loss + entropy_loss + sequence_similarity_loss + dense_1_similarity_loss
return total_loss
return loss_func
class EpochVariableCallback(Callback):
def __init__(self, my_variable, my_func):
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_end(self, epoch, logs={}):
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#Function for running GENESIS
def run_genesis(run_prefix, sequence_templates, loss_func, library_contexts, model_path, batch_size=32, n_samples=1, n_epochs=10, steps_per_epoch=100, n_intermediate_sequences=960) :
#Build Generator Network
_, generator = build_generator(batch_size, len(sequence_templates[0]), load_generator_network, n_classes=len(sequence_templates), n_samples=n_samples, sequence_templates=sequence_templates, batch_normalize_pwm=False)
#Build Validation Generator Network
_, val_generator = get_generator_copier(generator)(batch_size, len(sequence_templates[0]), get_shallow_copy_function(generator), n_classes=len(sequence_templates), n_samples=n_samples, sequence_templates=sequence_templates, batch_normalize_pwm=False, validation_sample_mode='sample', supply_inputs=True)
#Build Predictor Network and hook it on the generator PWM output tensor
_, predictor = build_predictor_w_adversary(generator, load_saved_predictor(model_path, library_contexts=library_contexts), batch_size, n_samples=n_samples, eval_mode='sample')
#Build Loss Model (In: Generator seed, Out: Loss function)
_, loss_model = build_loss_model(predictor, loss_func)
#Specify Optimizer to use
opt = keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
#Compile Loss Model (Minimize self)
loss_model.compile(loss=lambda true, pred: pred, optimizer=opt)
#Randomized validation tensors
val_random_tensor_funcs = [
lambda i: np.array(np.zeros(n_intermediate_sequences)).reshape(-1, 1),
lambda i: np.random.uniform(-1, 1, (n_intermediate_sequences, 100)),
lambda i: np.random.uniform(-1, 1, (n_intermediate_sequences, 100))
]
#Standard sequence decoder
acgt_encoder = IdentityEncoder(205, {'A':0, 'C':1, 'G':2, 'T':3})
#Build callback for printing intermediate sequences
random_genesis_monitor = GenesisMonitor(val_generator, acgt_encoder, run_dir="./samples/" + run_prefix + "/", run_prefix="intermediate", n_sequences=n_intermediate_sequences, batch_size=batch_size, input_tensor_funcs=val_random_tensor_funcs)
#Fit Loss Model
train_history = loss_model.fit(
[], np.ones((1, 1)),
epochs=n_epochs,
steps_per_epoch=steps_per_epoch,
callbacks=[random_genesis_monitor]
)
train_history = None
return generator, predictor, train_history
# +
#Specfiy file path to pre-trained predictor network
save_dir = os.path.join(os.getcwd(), '../../../aparent/saved_models')
saved_predictor_model_name = 'aparent_plasmid_iso_cut_distalpas_all_libs_no_sampleweights_sgd.h5'
saved_predictor_model_path = os.path.join(save_dir, saved_predictor_model_name)
# +
#Maximize isoform proportions for all native minigene libraries
sequence_templates = [
'TCCCTACACGACGCTCTTCCGATCTNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNAATAAANNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNNAATAAATTGTTCGTTGGTCGGCTTGAGTGCGTGTGTCTCGTTTAGATGCTGCGCCTAACCCTAAGCAGATTCTTCATGCAATTG'
]
library_contexts = [
'simple'
]
# +
#Train APA Cleavage GENESIS Network
print("Training GENESIS")
model_prefix = "genesis_apa_max_isoform_simple_25000_updates_similarity_margin_seq_05_lat_07_latent_penalty_earthmover_weight_01_target_14"
#Number of PWMs to generate per objective
batch_size = 64
#Number of One-hot sequences to sample from the PWM at each grad step
n_samples = 10
#Number of epochs per objective to optimize
n_epochs = 250
#Number of steps (grad updates) per epoch
steps_per_epoch = 100
#Number of sequences to sample and store for each epoch
n_intermediate_sequences = 960
K.clear_session()
loss = get_isoform_loss(
[1.0],
fitness_target=14.0,
fitness_weight=0.1,
use_start=25,
use_end=70,
use_target_bits=1.8,
cse_start=70,
cse_end=76,
cse_target_bits=None,#1.8,
dse_start=76,
dse_end=121,
dse_target_bits=1.8,
entropy_weight=1.0,
sequence_similarity_weight=5.0,
sequence_similarity_margin=0.5,
dense_1_similarity_weight=5.0,
dense_1_similarity_margin=0.7,
punish_dn_cse=0.0,
punish_up_c=0.0,
punish_dn_c=0.0,
punish_up_g=0.0,
punish_dn_g=0.0,
punish_up_aa=0.0,
punish_dn_aa=0.0,
)
generator_model, predictor_model, train_history = run_genesis(model_prefix, [sequence_templates[0]], loss, [library_contexts[0]], saved_predictor_model_path, batch_size, n_samples, n_epochs, steps_per_epoch, n_intermediate_sequences)
generator_model.get_layer('lambda_rand_sequence_class').function = lambda inp: inp
generator_model.get_layer('lambda_rand_input_1').function = lambda inp: inp
generator_model.get_layer('lambda_rand_input_2').function = lambda inp: inp
predictor_model.get_layer('lambda_rand_sequence_class').function = lambda inp: inp
predictor_model.get_layer('lambda_rand_input_1').function = lambda inp: inp
predictor_model.get_layer('lambda_rand_input_2').function = lambda inp: inp
# Save model and weights
save_dir = 'saved_models'
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_name = model_prefix + '_generator.h5'
model_path = os.path.join(save_dir, model_name)
generator_model.save(model_path)
print('Saved trained model at %s ' % model_path)
model_name = model_prefix + '_predictor.h5'
model_path = os.path.join(save_dir, model_name)
predictor_model.save(model_path)
print('Saved trained model at %s ' % model_path)
# +
#Specfiy file path to pre-trained predictor network
save_dir = os.path.join(os.getcwd(), '../../../aparent/saved_models')
saved_predictor_model_name = 'aparent_plasmid_iso_cut_distalpas_all_libs_no_sampleweights_sgd.h5'
saved_predictor_model_path = os.path.join(save_dir, saved_predictor_model_name)
saved_predictor = load_model(saved_predictor_model_path)
acgt_encoder = IdentityEncoder(205, {'A':0, 'C':1, 'G':2, 'T':3})
# +
#Load GENESIS models and predict sample sequences
model_prefix = "genesis_apa_max_isoform_simple_25000_updates_similarity_margin_seq_03_lat_095_latent_penalty_earthmover_weight_01_target_14"
batch_size = 64
sequence_template = sequence_templates[0]
save_dir = os.path.join(os.getcwd(), 'saved_models')
model_name = model_prefix + '_generator.h5'
model_path = os.path.join(save_dir, model_name)
generator = load_model(model_path, custom_objects={'st_sampled_softmax': st_sampled_softmax, 'st_hardmax_softmax': st_hardmax_softmax})
n = batch_size
sequence_class = np.array([0] * n).reshape(-1, 1) #np.random.uniform(-6, 6, (n, 1)) #
noise_1 = np.random.uniform(-1, 1, (n, 100))
noise_2 = np.random.uniform(-1, 1, (n, 100))
pred_outputs = generator.predict([sequence_class, noise_1, noise_2], batch_size=batch_size)
_, _, _, optimized_pwm, _, sampled_pwm, _, _, _ = pred_outputs
#Make predictions using black box model
fake_lib = np.zeros((optimized_pwm.shape[0], 13))
fake_lib[:, 5] = 1.
fake_d = np.ones((optimized_pwm.shape[0], 1))
iso_pred, cut_pred = saved_predictor.predict(x=[sampled_pwm[:, 0, ...], fake_lib, fake_d], batch_size=batch_size)
for pwm_index in range(16) :
print("iso_pred = " + str(iso_pred[pwm_index, 0]))
print("score_pred = " + str(np.log(iso_pred[pwm_index, 0] / (1. - iso_pred[pwm_index, 0]))))
pwm = np.expand_dims(optimized_pwm[pwm_index, :, :, 0], axis=0)
cut = np.expand_dims(cut_pred[pwm_index, :], axis=0)
iso = np.expand_dims(iso_pred[pwm_index], axis=0)
plot_seqprop_logo(pwm, iso, cut, annotate_peaks='max', sequence_template=sequence_templates[0], figsize=(12, 1.5), width_ratios=[1, 8], logo_height=0.8, usage_unit='fraction', plot_start=70-49, plot_end=76+49, save_figs=False, fig_name=model_prefix + "_pwm_index_" + str(pwm_index), fig_dpi=150)
# +
#Load GENESIS models and predict sample sequences
n = 1000
n_slack = 0.05 * n
n_ceil = int((n + n_slack) / batch_size) * batch_size + batch_size
sequence_class = np.array([0] * n_ceil).reshape(-1, 1) #np.random.uniform(-6, 6, (n, 1)) #
noise_1 = np.random.uniform(-1, 1, (n_ceil, 100))
noise_2 = np.random.uniform(-1, 1, (n_ceil, 100))
pred_outputs = generator.predict([sequence_class, noise_1, noise_2], batch_size=batch_size)
_, _, _, optimized_pwm, _, sampled_pwm, _, _, _ = pred_outputs
pwms = optimized_pwm[:, :, :, 0]
onehots = sampled_pwm[:, 0, :, :, 0]
#Make predictions using black box model
fake_lib = np.zeros((optimized_pwm.shape[0], 13))
fake_lib[:, 5] = 1.
fake_d = np.ones((optimized_pwm.shape[0], 1))
iso_pred, _ = saved_predictor.predict(x=[sampled_pwm[:, 0, ...], fake_lib, fake_d], batch_size=batch_size)
prob_pred = np.ravel(iso_pred)
score_pred = np.log(prob_pred / (1. - prob_pred))
sort_index = np.argsort(score_pred)[::-1]
pwms = pwms[sort_index][:n]
onehots = onehots[sort_index][:n]
score_pred = score_pred[sort_index][:n]
prob_pred = prob_pred[sort_index][:n]
# +
import seaborn as sns
#Target vs. Engineered Isoform Log Odds
save_figs = False
print("mean proportion = " + str(round(np.mean(prob_pred), 4)))
print("std proportion = " + str(round(np.std(prob_pred), 4)))
print("mean score = " + str(round(np.mean(score_pred), 4)))
print("std score = " + str(round(np.std(score_pred), 4)))
print("-------------------------")
f = plt.figure(figsize=(6, 4))
sns.violinplot(data=[score_pred])
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Fitness Score (log)', fontsize=18)
plt.tight_layout()
if save_figs :
plt.savefig(model_prefix + "_fitness_score_violin.png", transparent=True, dpi=150)
plt.savefig(model_prefix + "_fitness_score_violin.eps")
plt.savefig(model_prefix + "_fitness_score_violin.svg")
plt.show()
f = plt.figure(figsize=(6, 4))
sns.stripplot(data=[score_pred], jitter=1.)
plt.xlim(-0.25, 0.25)
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Fitness Score (log)', fontsize=18)
plt.tight_layout()
if save_figs :
plt.savefig(model_prefix + "_fitness_score_stripplot.png", transparent=True, dpi=150)
plt.savefig(model_prefix + "_fitness_score_stripplot.eps")
plt.savefig(model_prefix + "_fitness_score_stripplot.svg")
plt.show()
# +
#Calculate average/std nucleotide entropy
nt_entropies = []
for j in range(onehots.shape[1]) :
if sequence_templates[0][j] == 'N' :
p_A = np.sum(onehots[:, j, 0]) / n
p_C = np.sum(onehots[:, j, 1]) / n
p_G = np.sum(onehots[:, j, 2]) / n
p_T = np.sum(onehots[:, j, 3]) / n
nt_entropy = 0
if p_A * p_C * p_G * p_T > 0. :
nt_entropy = - (p_A * np.log2(p_A) + p_C * np.log2(p_C) + p_G * np.log2(p_G) + p_T * np.log2(p_T))
nt_entropies.append(nt_entropy)
nt_entropies = np.array(nt_entropies)
print("Mean NT Entropy = " + str(round(np.mean(nt_entropies), 4)))
print("Std NT Entropy = " + str(round(np.std(nt_entropies), 4)))
#Calculate hexamer entropies
hexamer_encoder = isol.NMerEncoder(n_mer_len=6, count_n_mers=True)
hexamers = isol.SparseBatchEncoder(encoder=hexamer_encoder)([
acgt_encoder.decode(onehots[i, :, :]) for i in range(onehots.shape[0])
])
hexamer_sum = np.ravel(hexamers.sum(axis=0))
hexamers_probs = hexamer_sum / np.sum(hexamer_sum)
n_nonzero_hexamers = len(np.nonzero(hexamer_sum > 0)[0])
print("Number of unique hexamers = " + str(n_nonzero_hexamers))
hexamer_entropy = -1. * np.sum(hexamers_probs[hexamer_sum > 0] * np.log2(hexamers_probs[hexamer_sum > 0]))
print("Hexamer Entropy = " + str(hexamer_entropy))
#Calculate average/std hexamer entropy
nonzero_index = np.nonzero(hexamer_sum > 0)[0]
hexamer_entropies = []
for j in range(n_nonzero_hexamers) :
p_on = len(np.nonzero(hexamers[:, nonzero_index[j]] > 0)[0]) / hexamers.shape[0]
p_off = 1. - p_on
hexamer_entropy = 0
if p_on * p_off > 0. :
hexamer_entropy = -(p_on * np.log2(p_on) + p_off * np.log2(p_off))
hexamer_entropies.append(hexamer_entropy)
hexamer_entropies = np.array(hexamer_entropies)
print("Mean Binary Hexamer Entropy = " + str(round(np.mean(hexamer_entropies), 4)))
print("Std Binary Hexamer Entropy = " + str(round(np.std(hexamer_entropies), 4)))
# +
import editdistance
#Calculate random pair-wise edit distances
save_figs = False
seqs = [
acgt_encoder.decode(onehots[i, :, :]) for i in range(onehots.shape[0])
]
shuffle_index = np.arange(len(seqs))
np.random.shuffle(shuffle_index)
distances = []
for i in range(len(seqs)) :
if i == shuffle_index[i] :
continue
seq_1 = seqs[i]
seq_2 = seqs[shuffle_index[i]]
dist = editdistance.eval(seq_1, seq_2)
distances.append(dist)
import seaborn as sns
distances = np.array(distances) / np.sum([1 if sequence_templates[0][j] == 'N' else 0 for j in range(len(sequence_templates[0]))])
print("mean distance/nt = " + str(round(np.mean(distances), 4)))
print("std distance/nt = " + str(round(np.std(distances), 4)))
print("-------------------------")
f = plt.figure(figsize=(6, 4))
sns.violinplot(data=[distances])
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Edit distance / nucleotide', fontsize=18)
plt.tight_layout()
if save_figs :
plt.savefig(model_prefix + "_edit_distance_violin.png", transparent=True, dpi=150)
plt.savefig(model_prefix + "_edit_distance_violin.eps")
plt.savefig(model_prefix + "_edit_distance_violin.svg")
plt.show()
f = plt.figure(figsize=(6, 4))
sns.stripplot(data=[distances], jitter=1.)
plt.xlim(-0.25, 0.25)
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Edit distance / nucleotide', fontsize=18)
plt.tight_layout()
if save_figs :
plt.savefig(model_prefix + "_edit_distance_stripplot.png", transparent=True, dpi=150)
plt.savefig(model_prefix + "_edit_distance_stripplot.eps")
plt.savefig(model_prefix + "_edit_distance_stripplot.svg")
plt.show()
# +
plot_n_seqs = 100
plot_start = 70-49
plot_end = 76+49
save_figs = False
flat_pwms = np.zeros((pwms.shape[0], pwms.shape[1]))
for i in range(pwms.shape[0]) :
for j in range(pwms.shape[1]) :
max_nt_ix = np.argmax(pwms[i, j, :])
flat_pwms[i, j] = max_nt_ix + 1
flat_pwms = flat_pwms[:plot_n_seqs, plot_start:plot_end]
cmap = colors.ListedColormap(['red', 'blue', 'orange', 'darkgreen'])
bounds=[0, 1, 2, 3, 4, 5]
norm = colors.BoundaryNorm(bounds, cmap.N)
f = plt.figure(figsize=(4, 12))
plt.imshow(flat_pwms, aspect='equal', interpolation='nearest', origin='lower', cmap=cmap, norm=norm)
plt.xticks([], [])
plt.yticks([], [])
plt.tight_layout()
if save_figs :
plt.savefig(model_prefix + "_diversity_seqs.png", transparent=True, dpi=150)
plt.savefig(model_prefix + "_diversity_seqs.svg")
plt.savefig(model_prefix + "_diversity_seqs.eps")
plt.show()
# -
#Get latent space predictor
saved_predictor_w_dense = Model(
inputs = saved_predictor.inputs,
outputs = saved_predictor.outputs + [saved_predictor.get_layer('dropout_1').output]
)
saved_predictor_w_dense.compile(loss='mse', optimizer=keras.optimizers.SGD(lr=0.1))
# +
#Load GENESIS models and predict sample sequences
batch_size = 64
n = 4096#10000
n_slack = 0#0.05 * n
n_ceil = int((n + n_slack) / batch_size) * batch_size
if n_ceil < n :
n_ceil += batch_size
sequence_class = np.array([0] * n_ceil).reshape(-1, 1) #np.random.uniform(-6, 6, (n, 1)) #
noise_1 = np.random.uniform(-1, 1, (n_ceil, 100))
noise_2 = np.random.uniform(-1, 1, (n_ceil, 100))
pred_outputs = generator.predict([sequence_class, noise_1, noise_2], batch_size=batch_size)
_, _, _, optimized_pwm, _, sampled_pwm, _, _, _ = pred_outputs
pwms = optimized_pwm[:, :, :, 0]
onehots = sampled_pwm[:, 0, :, :, 0]
#Make predictions using black box model
fake_lib = np.zeros((optimized_pwm.shape[0], 13))
fake_lib[:, 5] = 1.
fake_d = np.ones((optimized_pwm.shape[0], 1))
iso_pred, _, dense_pred = saved_predictor_w_dense.predict(x=[sampled_pwm[:, 0, ...], fake_lib, fake_d], batch_size=batch_size)
prob_pred = np.ravel(iso_pred)
score_pred = np.log(prob_pred / (1. - prob_pred))
sort_index = np.argsort(score_pred)[::-1]
pwms = pwms[sort_index][:n]
onehots = onehots[sort_index][:n]
score_pred = score_pred[sort_index][:n]
prob_pred = prob_pred[sort_index][:n]
dense_pred = dense_pred[sort_index][:n]
# -
#Save sequences to file
with open(model_prefix + "_4096_sequences.txt", "wt") as f:
for i in range(onehots.shape[0]) :
seq = acgt_encoder.decode(onehots[i])
f.write(seq + "\n")
# +
#Load GENESIS models and predict sample sequences
n = 4096
upsamples = [1, 10, 100]
for upsample in upsamples :
print("Upsampling = " + str(int(upsample)) + "X.")
n_ceil = int((n * upsample) / batch_size) * batch_size + batch_size
sequence_class = np.array([0] * n_ceil).reshape(-1, 1) #np.random.uniform(-6, 6, (n, 1)) #
noise_1 = np.random.uniform(-1, 1, (n_ceil, 100))
noise_2 = np.random.uniform(-1, 1, (n_ceil, 100))
pred_outputs = generator.predict([sequence_class, noise_1, noise_2], batch_size=batch_size)
_, _, _, optimized_pwm, _, sampled_pwm, _, _, _ = pred_outputs
pwms = optimized_pwm[:, :, :, 0]
onehots = sampled_pwm[:, 0, :, :, 0]
#Make predictions using black box model
fake_lib = np.zeros((optimized_pwm.shape[0], 13))
fake_lib[:, 5] = 1.
fake_d = np.ones((optimized_pwm.shape[0], 1))
iso_pred, _ = saved_predictor.predict(x=[sampled_pwm[:, 0, ...], fake_lib, fake_d], batch_size=batch_size)
prob_pred = np.ravel(iso_pred)
score_pred = np.log(prob_pred / (1. - prob_pred))
sort_index = np.argsort(score_pred)[::-1]
pwms = pwms[sort_index][:n]
onehots = onehots[sort_index][:n]
score_pred = score_pred[sort_index][:n]
prob_pred = prob_pred[sort_index][:n]
#Save sequences to file
with open(model_prefix + "_4096_sequences_upsampling_" + str(int(upsample)) + ".txt", "wt") as f:
for i in range(onehots.shape[0]) :
seq = acgt_encoder.decode(onehots[i])
f.write(seq + "\n")
print("mean proportion = " + str(round(np.mean(prob_pred), 4)))
print("std proportion = " + str(round(np.std(prob_pred), 4)))
print("mean score = " + str(round(np.mean(score_pred), 4)))
print("std score = " + str(round(np.std(score_pred), 4)))
print("-------------------------")
f = plt.figure(figsize=(6, 4))
sns.violinplot(data=[score_pred])
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Fitness Score (log)', fontsize=18)
plt.tight_layout()
plt.show()
seqs = [
acgt_encoder.decode(onehots[i, :, :]) for i in range(onehots.shape[0])
]
shuffle_index = np.arange(len(seqs))
np.random.shuffle(shuffle_index)
distances = []
for i in range(len(seqs)) :
if i == shuffle_index[i] :
continue
seq_1 = seqs[i]
seq_2 = seqs[shuffle_index[i]]
dist = editdistance.eval(seq_1, seq_2)
distances.append(dist)
distances = np.array(distances) / np.sum([1 if sequence_templates[0][j] == 'N' else 0 for j in range(len(sequence_templates[0]))])
print("mean distance/nt = " + str(round(np.mean(distances), 4)))
print("std distance/nt = " + str(round(np.std(distances), 4)))
print("-------------------------")
f = plt.figure(figsize=(6, 4))
sns.violinplot(data=[distances])
plt.xticks([], [])
plt.yticks(fontsize=14)
plt.ylabel('Edit distance / nucleotide', fontsize=18)
plt.tight_layout()
plt.show()
# -
| analysis/apa/apa_max_isoform_genesis_simple_predictor_latent_similarity_cosine_earthmover.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # INFO 3402 – Week 05: Histograms and Perception
#
# [<NAME>, Ph.D.](http://brianckeegan.com/)
# [Assistant Professor, Department of Information Science](https://www.colorado.edu/cmci/people/information-science/brian-c-keegan)
# University of Colorado Boulder
#
# Copyright and distributed under an [MIT License](https://opensource.org/licenses/MIT)
# ## Setup
# +
import numpy as np
np.set_printoptions(suppress=True)
import pandas as pd
idx = pd.IndexSlice
pd.options.display.max_columns = 100
# -
# Load the Boulder and Broomfield weather data from Week 1.
boulder_df = pd.read_csv('../Week 01 - Loading and Documentation/boulder_weather.csv',
header=1,
sep='|'
)
boulder_df.head()
# ## Background
#
# We've discussed the *shape* of DataFrames in terms of rows, columns, MultiIndicies, and tidy data. But for our first encounter with data visualization, we want to explore the *shape* of data values themselves. A classic and still influential practice is to visualize the distribution of data values as histograms.
#
# We are going to spend a lot of time for the rest of class working with the [matplotlib](https://matplotlib.org/) data visualization library. Many other parts of Python's data analysis ecosystem, including pandas and seaborn, interface with matplotlib for visualization. In particular, we are going to use matplotlib's "[object-oriented](https://matplotlib.org/stable/tutorials/introductory/usage.html?highlight=object%20oriented#coding-styles)" coding style for constucting and customizing visualizations. Here is the "anatomy" of a matplotlib figure with the corresponding methods for changing that part of a figure. We'll be referencing this a lot!
#
# 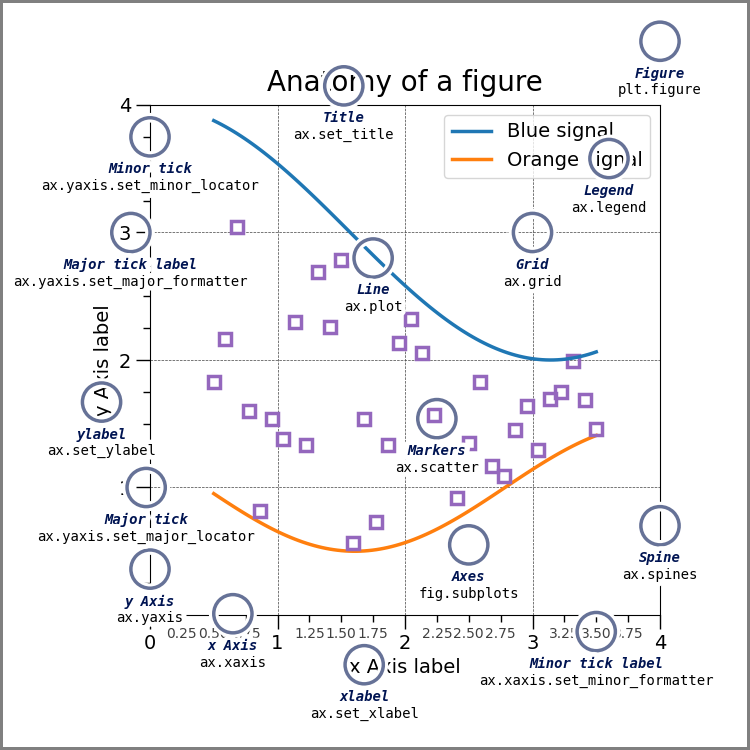
# ## Setting up matplotlib
#
# When we use matplotlib within a Jupyter Notebook, we have to run a special command called a "[cell magic](https://ipython.readthedocs.io/en/stable/interactive/magics.html)" so that the images it generates are included in the notebook. If you didn't have the `%matplotlib inline` cell magic, you would need to run a command like `plt.show()` or `plt.savefig()` to see the output of your visualization.
#
# We also construct and manage plots through matplotlib's `pyplot` ([docs](https://matplotlib.org/stable/api/_as_gen/matplotlib.pyplot.html)) interface. The `pyplot` module contains many functions for improving plots.
#
# **Take-away 01**: If you're using matplotlib in a Jupyter Notebook, include the `%matplotlib inline` command.
# **Take-away 02**: Always import and alias the pyplot interface at a minimum: `import matplotlib.pyplot as plt`
# +
# %matplotlib inline
import matplotlib.pyplot as plt
# -
# ### Exercise 01: Load Broomfield weather data
broomfield_df = pd.read_csv('../Week 01 - Loading and Documentation/broomfield_weather.csv',
header=1,
sep='|'
)
broomfield_df.head()
# ## Creating and customizing figures and subplots
#
# This is our first stop on the "object-oriented" coding style in matplotlib. Before we do anything, we just want to create a figure called `f` containing a single subplot called `ax`. These are empty until we fill them.
f, ax = plt.subplots()
# We can call the `f` Figure object and it will return the (now-empty) visualization.
f
# The `f` object contains an enormous number of methods and attributes, many of which we will explore in the weeks to come. The `.get_axes()` method retrieves the subplot objects (called axes) within the figure object. There's only one subplot in this figure, so it's a list of one ax. Later, we will see how to make figures with multiple subplots (or see [this tutorial](https://matplotlib.org/stable/tutorials/intermediate/arranging_axes.html) if you can't wait).
f.get_axes()
# The `ax` object (or the `AxesSubplot:` object within the figure's list of axes) have their own methods and attributes. The `.get_children()` shows all the components of this "empty" plot:
# * `spines` for the four borders (top, bottom, right, left)
# * the `XAxis` and `YAxis` with the tick labels 0.0 to 1.0
# * empty `Text` objects for the xlabel, ylabel, and title
# * a `Rectangle` patch object that overlaps with the spines.
ax.get_children()
# The rectangle object is a box that has a height and width of 1 and starts at the origin (0,0).
rect = ax.get_children()[-1]
rect.get_height(), rect.get_width(), rect.get_xy()
# We could mutate this Rectangle: color it red and give it a new width and height.
rect.set_facecolor('red')
rect.set_height(.5)
rect.set_width(.5)
# If we check the `f` object again, we can see this rectangle within the axes object within the figure object is now red and has half the height and width as before.
f
# ### Exercise 02: Mutate a patch edgecolor and linewidth
#
# Mutate the `rect` object ([docs](https://matplotlib.org/stable/api/_as_gen/matplotlib.patches.Rectangle.html)) so that it has an "edgecolor" of green and "linewidth" of 10. Show the result in `f`.
rect.set_edgecolor('green')
rect.set_linewidth(10)
rect.set_linestyle('--')
f
# ## One-dimensional histogram
#
# We're going to make a simple histogram using the weather data we loaded above.
#
# First, retrieve the values of the Boulder "TEMP" Series as a numpy `array` using the `.values` attribute and store as `boulder_temps`.
boulder_temps = boulder_df['TEMP'].values
boulder_temps
# Now make an empty figure and subplot.
f,ax = plt.subplots()
# Now plot a histogram of `boulder_temps` in the `ax` object. The `.hist` method returns three values:
#
# * The count of values in each bin
# * The bounadies of each bin
# * The `BarContainer` object that populates/replaces the contents of the empty `ax`
ax.hist(boulder_temps)
# But where's the plot? Check `f`.
f
# If you do the create the figure and subplot and make the histogram in the same cell (which we'll do from now on) the figure appears automatically.
#
# The other output from the `hist` method is still included: if you want to silence this, add a semicolon to the end of the line.
f,ax = plt.subplots()
ax.hist(boulder_temps); # Silence the bin counts and boundaries
# ### Number of bins
# We can also change the number of bins in the histogram with the "bins" parameter. Start with few bins.
f,ax = plt.subplots()
ax.hist(boulder_temps,bins=5);
# Try a large number of bins. We start to see there might be two-ish peaks to the data, which would make sense for different temperatures in the winter and summer.
f,ax = plt.subplots()
ax.hist(boulder_temps,bins=50);
# Try a really big number of bins. The bimodal distribution is perhaps clearer here, but this probably more bins than intended for a histogram.
# +
f,ax = plt.subplots()
ax.hist(boulder_temps,bins=500)
ax.set_xlabel('Temperature (F)')
ax.set_ylabel('Count (Days)')
ax.set_title('Temperatures in Boulder, Colorado (2010-2021)')
f.savefig('too_many_bins.png',facecolor='w')
# -
# ### Label axes
#
# One of the most essential visualization practices is to label our axes. Add an x-label and y-label.
# +
f,ax = plt.subplots()
ax.hist(boulder_temps,bins=25)
ax.set_xlabel('Temperature (F)')
ax.set_ylabel('Count (Days)')
ax.set_title('Temperatures in Boulder, Colorado (2010-2021)')
# -
# ### Exercise 03: Plot a histogram of Broomfield's temperatures
#
# Create an empty figure and subplot, extract the "TEMP" values from `broomfield_df`, make and display a histogram, and label your axes.
# +
f,ax = plt.subplots()
ax.hist(broomfield_df['TEMP'],bins=25);
# +
f,a_subplot = plt.subplots()
broomfield_df['TEMP'].plot.hist(ax=a_subplot,bins=50)
# -
# ## Skewed data
#
# The weather data has nice statistical and visual properties of being (approximately) normally distributed. Many kinds of data do not have this property and instead show a strong skew with low counts being significantly more represented than high counts.
#
# Load in the county COVID data from Week 3 as `co_covid_df`.
co_covid_df = pd.read_csv('../Week 03 - Combining and Validation/co_county_covid.csv')
co_covid_df.head()
# Extract the values for the "Cases" column and make a histogram. There's an enormous number of county-months where there are 0 or few cases and a handful of county-months where there are thousands of cases. This is not an informative data visualization.
# +
covid_cases = co_covid_df['Cases'].values
f,ax = plt.subplots()
ax.hist(covid_cases)
ax.set_xlabel('Cases')
ax.set_ylabel('Count (Monthly)')
ax.set_title('COVID-19 Cases in Colorado Counties')
f.savefig('county_cases.png',facecolor='w')
# -
# Even adding more bins does not improve this visualization.
# +
f,ax = plt.subplots()
ax.hist(covid_cases,bins=50)
ax.set_xlabel('Cases')
ax.set_ylabel('Count (Monthly)')
ax.set_title('COVID-19 Cases in Colorado Counties')
# -
# A common approach to handling skewed data like this is to use a log-transformation. Rather than the bin sizes being linear (1 to 10, 11 to 20, *etc*.) the bin sizes are exponential (1 to 10, 11 to 100, 101 to 1000, *etc*.).
#
# A numpy function called `linspace` ([docs](https://numpy.org/doc/stable/reference/generated/numpy.linspace.html)) can illustrate creating 10 linearly-spaced bins from 1 to 1000 and a similar function called `geomspace` ([docs](https://numpy.org/doc/stable/reference/generated/numpy.geomspace.html)) to create 10 exponentially-spaced bins from 1 to 1000.
np.linspace(1,10000,11)
np.geomspace(1,10000,11) # The start and stop values are in base units 10^0 = 1, 10^4 = 10000
# We can pass these bin boundaries into the "bins" parameter for `hist`.
# +
f,ax = plt.subplots()
ax.hist(covid_cases,bins=np.linspace(1,10000,11)) # Linear bin sizes
ax.set_xlabel('Cases')
ax.set_ylabel('Count (Monthly)')
ax.set_title('COVID-19 Cases in Colorado Counties')
# -
# Use the geometrically-spaced bins instead. There's definitely something different going on here!
# +
f,ax = plt.subplots()
ax.hist(covid_cases,bins=np.geomspace(1,10000,11)) # Geometric bin sizes
ax.set_xlabel('Cases')
ax.set_ylabel('Count (Monthly)')
ax.set_title('COVID-19 Cases in Colorado Counties')
# -
# Because we're using geometric instead of linear bins, we should also use a geometric scale for the x and y-axes instead of a linear scale. Use the `.set_xscale()` ([docs](https://matplotlib.org/stable/api/_as_gen/matplotlib.axes.Axes.set_xscale.html)) method to turn the x-axis ticks into a geometric rather than linear progression. This is called a "semi-log" plot.
#
# You could also make the y-axis have a geometric/logarithmic progression (uncomment the last line with `.set_yscale('log')`. This would be a "log-log" plot.
# +
f,ax = plt.subplots()
ax.hist(covid_cases,bins=np.geomspace(1,10000,11)) # Geometric bin sizes
ax.set_xlabel('Cases')
ax.set_ylabel('Count (Monthly)')
ax.set_title('Monthly COVID-19 Cases in Colorado Counties')
ax.set_xscale('log')
# ax.set_yscale('log')
# -
# ### Advanced: Why should I care about log-binned data?
# (This won't be on the quiz, but if you're curious, read on)
#
# Make some data following different distributions: [exponential](https://numpy.org/doc/stable/reference/random/generated/numpy.random.exponential.html), [lognormal](https://numpy.org/doc/stable/reference/random/generated/numpy.random.lognormal.html), [pareto](https://numpy.org/doc/stable/reference/random/generated/numpy.random.pareto.html), and [power](https://numpy.org/doc/stable/reference/random/generated/numpy.random.power.html).
# +
# https://stackoverflow.com/a/58331860/1574687
# Rescale data to be (0,1)
from sklearn.preprocessing import minmax_scale
_size = 1000
# Make random distributions of different shapes
random_exponential = minmax_scale(np.random.exponential(size=_size),feature_range=(0,1))
random_lognormal = minmax_scale(np.random.lognormal(size=_size),feature_range=(0,1))
random_pareto = minmax_scale(np.random.pareto(.9,size=_size),feature_range=(0,1))
random_power = minmax_scale(np.random.power(.1,size=_size),feature_range=(0,1))
# -
# On linear axes and bins these distributions look pretty similar.
# +
# Make four subplots
f,(ax0,ax1,ax2,ax3) = plt.subplots(
nrows = 1,
ncols = 4,
figsize=(16,4)
)
# Histograms for each subplot
ax0.hist(random_exponential,bins=25)
ax1.hist(random_lognormal,bins=25)
ax2.hist(random_pareto,bins=25)
ax3.hist(random_power,bins=25)
# Set titles
ax0.set_title('Exponential')
ax1.set_title('Log-normal')
ax2.set_title('Pareto')
ax3.set_title('Power')
# Super-title and cleanup
f.suptitle('Long-tailed distributions')
f.set_facecolor('w')
f.tight_layout()
# -
# On log axes and bins they look very different!
# +
# Make four subplots
f,(ax0,ax1,ax2,ax3) = plt.subplots(
nrows = 1,
ncols = 4,
figsize = (16,4),
subplot_kw = {'xscale':'log'}
)
# Histograms for each subplot with geomspace bins
ax0.hist(random_exponential,bins=np.geomspace(1e-3,1e0,25))
ax1.hist(random_lognormal,bins=np.geomspace(1e-3,1e0,25))
ax2.hist(random_pareto,bins=np.geomspace(1e-4,1e0,25))
ax3.hist(random_power,bins=np.geomspace(1e-3,1e0,25))
# Set titles
ax0.set_title('Exponential')
ax1.set_title('Log-normal')
ax2.set_title('Pareto')
ax3.set_title('Power')
# Super-title and cleanup
f.suptitle('Long-tailed distributions')
f.set_facecolor('w')
f.tight_layout()
# -
# ### Exercise 04: Make a log-log histogram of deaths
#
# Create an empty figure and subplot, extract the "Deaths" values from `co_covid_df`, make a histogram with an appropriate number and size of bins, label your axes, and rescale the axes.
co_covid_df.head()
# +
f,covid_deaths_ax = plt.subplots()
co_covid_df['Deaths'].plot.hist(ax=covid_deaths_ax)
# -
no_neg_deaths = co_covid_df[co_covid_df['Deaths'] > 0]
f,ax = plt.subplots()
no_neg_deaths['Deaths'].plot.hist(ax=ax,bins=np.geomspace(1,40,10))
ax.set_xscale('log')
# ## Visualizing multiple histograms
#
# ([Adapted from this gallery](https://matplotlib.org/stable/gallery/statistics/histogram_multihist.html))
#
# We may want to visualize more than one histogram at once. Let's work with the "TEMP" data from the Boulder and Broomfield data.
#
# These data are different sizes reflecting different dates, start by inner joining them together so they are the same size. I'm using the "suffixes" parameter because the column names are otherwise identical (tidy data violation!).
boulder_df.shape, broomfield_df.shape
# +
both_df = pd.merge(
left = boulder_df,
right = broomfield_df,
left_on = 'DATE',
right_on = 'DATE',
how = 'inner',
suffixes = ('_BOULDER','_BROOMFIELD')
)
both_df.head()
# -
boulder_temps = both_df['TEMP_BOULDER'].values
broomfield_temps = both_df['TEMP_BROOMFIELD'].values
# Option 1 is to make side-by-side plots. The "ncols" parameter is changed from default 1 to 2. Now the `subplots` function produces two subplot axes that I capture as `ax0` and `ax1`.
f,(ax0,ax1) = plt.subplots(1,2)
# +
f,(ax0,ax1) = plt.subplots(
nrows=1,
ncols=2,
figsize=(8,4),
sharey=True
)
ax0.hist(boulder_temps,bins=25)
ax1.hist(broomfield_temps,bins=25)
ax0.set_xlabel('Temperature (F)')
ax0.set_ylabel('Count (Days)')
ax0.set_title('Boulder, Colorado')
ax1.set_xlabel('Temperature (F)')
ax1.set_ylabel('Count (Days)')
ax1.set_title('Broomfield, Colorado')
f.suptitle('Temperatures 2010-2021',fontweight='heavy')
f.tight_layout()
f.savefig('hist_temp_boulder_broomfield.png',facecolor='w')
# -
# Option 2 is to make the histogram on a single plot but with different bars for each city. This is hard with default matplotlib, but is easier with pandas and seaborn.
#
# stack ([docs](https://numpy.org/doc/stable/reference/generated/numpy.stack.html)) the two arrays together (similar to a concat).
# Stack the numpy arrays together
stacked_arrays = np.stack([boulder_temps,broomfield_temps],axis=1)
boulder_temps.shape, broomfield_temps.shape, stacked_arrays.shape
# +
f,ax = plt.subplots()
ax.hist(
stacked_arrays,
bins = 15,
histtype = 'bar',
color = ['tab:red','tab:blue'],
label = ['Boulder','Broomfield']
)
ax.set_xlabel('Temperature (F)')
ax.set_ylabel('Count (Days)')
ax.set_title('Temperatures (2010-2021)',fontweight='heavy')
ax.legend(prop={'size': 10})
# -
# Option 3 is to stack the bars on top of each other. This is perceptually hard to compare to identify differences in many cases.
# +
f,ax = plt.subplots()
ax.hist(
stacked_arrays,
bins = 15,
histtype = 'bar',
stacked = True,
color = ['tab:red','tab:blue'],
label = ['Boulder','Broomfield']
)
ax.set_xlabel('Temperature (F)')
ax.set_ylabel('Count (Days)')
ax.set_title('Temperatures (2010-2021)',fontweight='heavy')
ax.legend(prop={'size': 10})
# -
# Plotting a cumulative histogram.
# +
f,ax = plt.subplots()
ax.hist(
stacked_arrays,
bins = 15,
histtype = 'bar',
cumulative = True,
color = ['tab:red','tab:blue'],
label = ['Boulder','Broomfield']
)
ax.set_xlabel('Temperature (F)')
ax.set_ylabel('Count (Days, cuml.)')
ax.set_title('Temperatures (2010-2021)',fontweight='heavy')
ax.legend(prop={'size': 10})
# -
# Making multiple plots with pandas.
co_covid_df.head()
co_covid_df['County'].unique()
13*5
axs.shape
co_covid_df.head()
# +
f,axs = plt.subplots(nrows=13,ncols=5,figsize=(10,20))
co_covid_df.plot.hist(
column='Cases',
by=['County'],
ax=axs,
legend=False,
# bins=np.geomspace(1,5000,10)
);
f.tight_layout()
# -
# ## pandas methods
#
# Everything we were doing above involved an extra step to extract data in pandas as numpy arrays and some new functions to manipulate those numpy arrays. Let's stay in pandas-land!
#
# You can call the `.hist()` method ([docs](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.hist.html)) on a pandas Series and it will return a figure and subplot with the histogram. There are some stylistic differences, but this is much easier!
both_df['TEMP_BOULDER'].hist()
# You could also call `.hist()` on a DataFrame and get a histogram for each.
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].hist()
# The `.plot.hist()` method is distinct and will draw stacked, cumulative, horizontal and others.
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].plot.hist()
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].plot.hist(stacked=True)
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].plot.hist(orientation='horizontal')
# You can also pass pandas's subplots back to a figure you've already setup with `subplots`. Just pass the axes created by subplots to the "ax" parameter inside the pandas `.hist()` method.
# +
f,(ax0,ax1) = plt.subplots(ncols=2,figsize=(8,4))
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].plot.hist(stacked=True,ax=ax0,legend=False)
both_df[['TEMP_BOULDER','TEMP_BROOMFIELD']].plot.hist(orientation='horizontal',ax=ax1)
# -
# ### Exercise 05: Make a histogram of COVID cases and deaths
no_neg_deaths.head(2)
# +
f,(ax0,ax1) = plt.subplots(1,2)
no_neg_deaths['Cases'].plot.hist(ax=ax0)
no_neg_deaths['Deaths'].plot.hist(ax=ax1)
# ax0.set_xscale('log')
# ax1.set_xscale('log')
# -
no_neg_deaths['Death'].cumsum()
# +
f,(ax0,ax1) = plt.subplots(1,2)
no_neg_deaths['Cases'].plot.hist(ax=ax0,bins=np.geomspace(1,10000,10))
no_neg_deaths['Deaths'].plot.hist(ax=ax1,bins=np.geomspace(1,100,10))
ax0.set_xscale('log')
ax1.set_xscale('log')
# -
# ## Advanced: Two-dimensional histogram
#
# You can visualize the counts of two series of data. This is a bit like a terrain map, darker colors are lower and brighter colors are higher. Basically, how many days did Boulder and Broomfield have these temperature values? Unsurprisingly, two nearby cities have similar temperatures: all the values lie along the diagonal, there are no days when it's 80 in Boulder and 20 in Broomfield.
# +
f,ax = plt.subplots(figsize=(7,6))
counts, xedges, yedges, im = ax.hist2d(
x = boulder_temps,
y = broomfield_temps,
bins = 25
)
ax.set_xlabel('Temperatures in Boulder (F)')
ax.set_ylabel('Temperatures in Broomfield (F)')
f.colorbar(im,ax=ax)
# -
| Week 05 - Histograms/Week 05 - Lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import numpy as np
import pandas as pd
import os
# +
#%%
# Get the Trading Calendar, run from a zip35 code environment
with open('trading_calendar.csv') as f:
reader = csv.reader(f)
data = list(reader)
arr = np.array(data)
trading_days = arr.ravel()
#%%
# Function to format the csv files for the csvdir.py Zipline Bundle Ingest process
def format_bundle(indir, outdir):
count = 0
for f in os.listdir(indir): # For Production
df = pd.read_csv('{}/{}'.format(indir, f), index_col='date')
# I need to add some logic here to force the number of rows to equal the number of expected trading days
df = df.reindex(trading_days)
# Export it in the csvdir format needed for the ziplien bundle ingestion process
df.reset_index(inplace=True)
# Check if there is there is any divident, if not make it zero
if not 'dividend' in df.columns:
df['dividend'] = 0.0
df = df[['date', 'open_adj', 'high_adj', 'low_adj', 'close_adj', 'volume_adj', 'dividend']]
df['dividend'].fillna(0.00, inplace=True)
df['ratio'] = 1 # Since I alread did all the adjusting
df.rename(columns={'open_adj': 'open',
'high_adj': 'high',
'low_adj': 'low',
'close_adj': 'close',
'volume_adj': 'volume'
}, inplace=True)
# Round the numbers in the dataframe
df = df.round({'open':2,
'high':2,
'low':2,
'close':2,
'volume':1,
'dividend':2})
df.to_csv('{}/{}'.format(outdir, f), index=False)
count += 1
return ('{} files was adjusted'.format(count))
#%% Execute the function to format the files
format_bundle('../../data/rawdata/historical', '../../data/celandata/historical')
# %%
# -
| notebooks/jsirab/Analysis/Cleaning_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Filter
#
# You can filter the rows of a table with [Table.filter](https://hail.is/docs/devel/hail.Table.html#hail.Table.filter). This returns a table of those rows for which the expression evaluates to `True`.
# + slideshow={"slide_type": "fragment"}
import hail as hl
import matplotlib.pyplot as plt
import seaborn
# %matplotlib inline
hl.utils.get_movie_lens('data/')
users = hl.read_table('data/users.ht')
# + slideshow={"slide_type": "fragment"}
users.filter(users.occupation == 'programmer').count()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Annotate
#
# You can add new fields to a table with [annotate](https://hail.is/docs/devel/hail.Table.html#hail.Table.annotate). Let's mean-center and variance-normalize the `age` field.
# + slideshow={"slide_type": "slide"}
stats = users.aggregate(hl.agg.stats(users.age))
missing_occupations = hl.set(['other', 'none'])
t = users.annotate(
cleaned_occupation = hl.cond(missing_occupations.contains(users.occupation),
hl.null('str'),
users.occupation))
t.show()
# + [markdown] slideshow={"slide_type": "fragment"}
# Note: `annotate` is functional: it doesn't mutate `users`, but returns a new table. This is also true of `filter`. In fact, all operations in Hail are functional.
# -
users.describe()
# + [markdown] slideshow={"slide_type": "slide"}
# There are two other annotate methods: [select](https://hail.is/docs/devel/hail.Table.html#hail.Table.select) and [transmute](https://hail.is/docs/devel/hail.Table.html#hail.Table.transmute). `select` returns a table with an entirely new set of fields. `transmute` replaces any fields mentioned on the right-hand side with the new fields, but leaves unmentioned fields unchanged. `transmute` is useful for transforming data into a new form. How about some examples?
# + slideshow={"slide_type": "slide"}
(users.select(len_occupation = hl.len(users.occupation))
.describe())
# + slideshow={"slide_type": "slide"}
(users.transmute(
cleaned_occupation = hl.cond(missing_occupations.contains(users.occupation),
hl.null(hl.tstr),
users.occupation))
.describe())
# + [markdown] slideshow={"slide_type": "slide"}
# Finally, you can add global fields with [annotate_globals](https://hail.is/docs/devel/hail.Table.html#hail.Table.annotate_globals). Globals are useful for storing metadata about a dataset or storing small data structures like sets and maps.
# + slideshow={"slide_type": "fragment"}
t = users.annotate_globals(cohort = 5, cloudable = hl.set(['sample1', 'sample10', 'sample15']))
t.describe()
# -
t.cloudable
t.cloudable.value
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercises
#
#
# - [Z-score normalize](https://en.wikipedia.org/wiki/Standard_score) the age field of `users`.
# - Convert `zip` to an integer. Hint: Not all zipcodes are US zipcodes! Use [hl.int32](https://hail.is/docs/devel/functions/constructors.html#hail.expr.functions.int32) to convert a string to an integer. Use [StringExpression.matches](https://hail.is/docs/devel/expressions.html#hail.expr.expression.StringExpression.matches) to see if a string matches a regular expression.
# -
| python/hail/docs/tutorials/07-filter-annotate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="7whD0jTtpK9A"
# # Installation & Versioning
# + [markdown] id="9VC5cxNDpZFf"
# Use the function %matplotlib inline to enable the inline plotting in this notebook.
#
#
# + id="c9GiiO1zId5d"
# %matplotlib inline
# + [markdown] id="_fIvdsyMqaE5"
# Install Triton via pip:
# + id="2BBwoJK-qWey" colab={"base_uri": "https://localhost:8080/"} outputId="741be77b-f9fd-4177-bdec-59a73f5d7c22"
# !pip install triton==2.0.0.dev20220206
# + [markdown] id="CL477-IeId5g"
#
# # Layer Normalization
#
# + id="HXOCNfDMId5h"
import pkg_resources
# Import from latest release tag==2.0.0.dev20220206
pkg_resources.require("triton==2.0.0.dev20220206")
import torch
import triton
import triton.language as tl
try:
# This is https://github.com/NVIDIA/apex, NOT the apex on PyPi, so it
# should not be added to extras_require in setup.py.
import apex
HAS_APEX = True
except ModuleNotFoundError:
HAS_APEX = False
# Forward Pass
@triton.jit
def _layer_norm_fwd_fused(X, Y, W, B, M, V, stride, N, eps,
BLOCK_SIZE: tl.constexpr):
# position of elements processed by this program
row = tl.program_id(0)
cols = tl.arange(0, BLOCK_SIZE)
mask = cols < N
# offset data pointers to start at the row of interest
X += row * stride
Y += row * stride
# load data and cast to float32
x = tl.load(X + cols, mask=mask, other=0).to(tl.float32)
# compute mean
mean = tl.sum(x, axis=0) / N
# compute std
xmean = tl.where(mask, x - mean, 0.)
var = tl.sum(xmean * xmean, axis=0) / N
rstd = 1 / tl.sqrt(var + eps)
xhat = xmean * rstd
# write-back mean/rstd
tl.store(M + row, mean)
tl.store(V + row, rstd)
# multiply by weight and add bias
w = tl.load(W + cols, mask=mask)
b = tl.load(B + cols, mask=mask)
y = xhat * w + b
# write-back
tl.store(Y + cols, y, mask=mask)
# Backward pass (DX + partial DW + partial DB)
@triton.jit
def _layer_norm_bwd_dx_fused(DX, DY, DW, DB, X, W, B, M, V, Lock, stride, N, eps,
GROUP_SIZE_M: tl.constexpr, BLOCK_SIZE_N: tl.constexpr):
# position of elements processed by this program
row = tl.program_id(0)
cols = tl.arange(0, BLOCK_SIZE_N)
mask = cols < N
# offset data pointers to start at the row of interest
X += row * stride
DY += row * stride
DX += row * stride
# offset locks and weight/bias gradient pointer
# each kernel instance accumulates partial sums for
# DW and DB into one of GROUP_SIZE_M independent buffers
# these buffers stay in the L2, which allow this kernel
# to be fast
lock_id = row % GROUP_SIZE_M
Lock += lock_id
Count = Lock + GROUP_SIZE_M
DW = DW + lock_id * N + cols
DB = DB + lock_id * N + cols
# load data to SRAM
x = tl.load(X + cols, mask=mask, other=0).to(tl.float32)
dy = tl.load(DY + cols, mask=mask, other=0).to(tl.float32)
w = tl.load(W + cols, mask=mask).to(tl.float32)
mean = tl.load(M + row)
rstd = tl.load(V + row)
# compute dx
xhat = (x - mean) * rstd
wdy = w * dy
xhat = tl.where(mask, xhat, 0.)
wdy = tl.where(mask, wdy, 0.)
mean1 = tl.sum(xhat * wdy, axis=0) / N
mean2 = tl.sum(wdy, axis=0) / N
dx = (wdy - (xhat * mean1 + mean2)) * rstd
# write-back dx
tl.store(DX + cols, dx, mask=mask)
# accumulate partial sums for dw/db
partial_dw = (dy * xhat).to(w.dtype)
partial_db = (dy).to(w.dtype)
while tl.atomic_cas(Lock, 0, 1) == 1:
pass
count = tl.load(Count)
# first store doesn't accumulate
if count == 0:
tl.atomic_xchg(Count, 1)
else:
partial_dw += tl.load(DW, mask=mask)
partial_db += tl.load(DB, mask=mask)
tl.store(DW, partial_dw, mask=mask)
tl.store(DB, partial_db, mask=mask)
# release lock
tl.atomic_xchg(Lock, 0)
# Backward pass (total DW + total DB)
@triton.jit
def _layer_norm_bwd_dwdb(DW, DB, FINAL_DW, FINAL_DB, M, N,
BLOCK_SIZE_M: tl.constexpr, BLOCK_SIZE_N: tl.constexpr):
pid = tl.program_id(0)
cols = pid * BLOCK_SIZE_N + tl.arange(0, BLOCK_SIZE_N)
dw = tl.zeros((BLOCK_SIZE_M, BLOCK_SIZE_N), dtype=tl.float32)
db = tl.zeros((BLOCK_SIZE_M, BLOCK_SIZE_N), dtype=tl.float32)
for i in range(0, M, BLOCK_SIZE_M):
rows = i + tl.arange(0, BLOCK_SIZE_M)
mask = (rows[:, None] < M) & (cols[None, :] < N)
offs = rows[:, None] * N + cols[None, :]
dw += tl.load(DW + offs, mask=mask, other=0.)
db += tl.load(DB + offs, mask=mask, other=0.)
sum_dw = tl.sum(dw, axis=0)
sum_db = tl.sum(db, axis=0)
tl.store(FINAL_DW + cols, sum_dw, mask=cols < N)
tl.store(FINAL_DB + cols, sum_db, mask=cols < N)
class LayerNorm(torch.autograd.Function):
@staticmethod
def forward(ctx, x, normalized_shape, weight, bias, eps):
# allocate output
y = torch.empty_like(x)
# reshape input data into 2D tensor
x_arg = x.reshape(-1, x.shape[-1])
M, N = x_arg.shape
mean = torch.empty((M, ), dtype=torch.float32, device='cuda')
rstd = torch.empty((M, ), dtype=torch.float32, device='cuda')
# Less than 64KB per feature: enqueue fused kernel
MAX_FUSED_SIZE = 65536 // x.element_size()
BLOCK_SIZE = min(MAX_FUSED_SIZE, triton.next_power_of_2(N))
if N > BLOCK_SIZE:
raise RuntimeError("This layer norm doesn't support feature dim >= 64KB.")
# heuristics for number of warps
num_warps = min(max(BLOCK_SIZE // 256, 1), 8)
# enqueue kernel
_layer_norm_fwd_fused[(M,)](x_arg, y, weight, bias, mean, rstd,
x_arg.stride(0), N, eps,
BLOCK_SIZE=BLOCK_SIZE, num_warps=num_warps)
ctx.save_for_backward(x, weight, bias, mean, rstd)
ctx.BLOCK_SIZE = BLOCK_SIZE
ctx.num_warps = num_warps
ctx.eps = eps
return y
@staticmethod
def backward(ctx, dy):
x, w, b, m, v = ctx.saved_tensors
# heuristics for amount of parallel reduction stream for DG/DB
N = w.shape[0]
GROUP_SIZE_M = 64
if N <= 8192: GROUP_SIZE_M = 96
if N <= 4096: GROUP_SIZE_M = 128
if N <= 1024: GROUP_SIZE_M = 256
# allocate output
locks = torch.zeros(2 * GROUP_SIZE_M, dtype=torch.int32, device='cuda')
_dw = torch.empty((GROUP_SIZE_M, w.shape[0]), dtype=x.dtype, device=w.device)
_db = torch.empty((GROUP_SIZE_M, w.shape[0]), dtype=x.dtype, device=w.device)
dw = torch.empty((w.shape[0],), dtype=w.dtype, device=w.device)
db = torch.empty((w.shape[0],), dtype=w.dtype, device=w.device)
dx = torch.empty_like(dy)
# enqueue kernel using forward pass heuristics
# also compute partial sums for DW and DB
x_arg = x.reshape(-1, x.shape[-1])
M, N = x_arg.shape
_layer_norm_bwd_dx_fused[(M,)](dx, dy, _dw, _db, x, w, b, m, v, locks,
x_arg.stride(0), N, ctx.eps,
BLOCK_SIZE_N=ctx.BLOCK_SIZE,
GROUP_SIZE_M=GROUP_SIZE_M,
num_warps=ctx.num_warps)
grid = lambda meta: [triton.cdiv(N, meta['BLOCK_SIZE_N'])]
# accumulate partial sums in separate kernel
_layer_norm_bwd_dwdb[grid](_dw, _db, dw, db, GROUP_SIZE_M, N,
BLOCK_SIZE_M=32,
BLOCK_SIZE_N=128)
return dx, None, dw, db, None
layer_norm = LayerNorm.apply
def test_layer_norm(M, N, dtype, eps=1e-5, device='cuda'):
# create data
x_shape = (M, N)
w_shape = (x_shape[-1], )
weight = torch.rand(w_shape, dtype=dtype, device='cuda', requires_grad=True)
bias = torch.rand(w_shape, dtype=dtype, device='cuda', requires_grad=True)
x = -2.3 + 0.5 * torch.randn(x_shape, dtype=dtype, device='cuda')
dy = .1 * torch.randn_like(x)
x.requires_grad_(True)
# forward pass
y_tri = layer_norm(x, w_shape, weight, bias, eps)
y_ref = torch.nn.functional.layer_norm(x, w_shape, weight, bias, eps).to(dtype)
# backward pass (triton)
y_tri.backward(dy, retain_graph=True)
dx_tri, dw_tri, db_tri = [_.grad.clone() for _ in [x, weight, bias]]
x.grad, weight.grad, bias.grad = None, None, None
# backward pass (torch)
y_ref.backward(dy, retain_graph=True)
dx_ref, dw_ref, db_ref = [_.grad.clone() for _ in [x, weight, bias]]
# compare
triton.testing.assert_almost_equal(y_tri, y_ref)
triton.testing.assert_almost_equal(dx_tri, dx_ref)
triton.testing.assert_almost_equal(db_tri, db_ref, decimal=1)
triton.testing.assert_almost_equal(dw_tri, dw_ref, decimal=1)
@triton.testing.perf_report(
triton.testing.Benchmark(
x_names=['N'],
x_vals=[512 * i for i in range(2, 32)],
line_arg='provider',
line_vals=['triton', 'torch'] + (['apex'] if HAS_APEX else []),
line_names=['Triton', 'Torch'] + (['Apex'] if HAS_APEX else []),
styles=[('blue', '-'), ('green', '-'), ('orange', '-')],
ylabel='GB/s',
plot_name='layer-norm-backward',
args={'M': 4096, 'dtype': torch.float16, 'mode': 'backward'}
)
)
def bench_layer_norm(M, N, dtype, provider, mode='backward', eps=1e-5, device='cuda'):
# create data
x_shape = (M, N)
w_shape = (x_shape[-1], )
weight = torch.rand(w_shape, dtype=dtype, device='cuda', requires_grad=True)
bias = torch.rand(w_shape, dtype=dtype, device='cuda', requires_grad=True)
x = -2.3 + 0.5 * torch.randn(x_shape, dtype=dtype, device='cuda')
dy = .1 * torch.randn_like(x)
x.requires_grad_(True)
# utility functions
if provider == 'triton':
y_fwd = lambda: layer_norm(x, w_shape, weight, bias, eps)
if provider == 'torch':
y_fwd = lambda: torch.nn.functional.layer_norm(x, w_shape, weight, bias, eps)
if provider == 'apex':
apex_layer_norm = apex.normalization.FusedLayerNorm(w_shape).to(x.device).to(x.dtype)
y_fwd = lambda: apex_layer_norm(x)
# forward pass
if mode == 'forward':
gbps = lambda ms: 2 * x.numel() * x.element_size() / ms * 1e-6
ms, min_ms, max_ms = triton.testing.do_bench(y_fwd, rep=500)
# backward pass
if mode == 'backward':
gbps = lambda ms: 3 * x.numel() * x.element_size() / ms * 1e-6
y = y_fwd()
ms, min_ms, max_ms = triton.testing.do_bench(lambda: y.backward(dy, retain_graph=True),
grad_to_none=[x], rep=500)
return gbps(ms), gbps(max_ms), gbps(min_ms)
bench_layer_norm.run(save_path='.', print_data=True)
| python/tutorials/jupyter_notebooks/05_layer_norm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rSE6Xk6x71gI" colab_type="text"
# # Hands-On Session No. 1
# ## (Get to know PyTorch, Gradient Descent on Linear Regression)
# + id="lYnvYDMrAaIi" colab_type="code" colab={}
import torch
import matplotlib.pyplot as plt
# + [markdown] id="CtRg7T4x8aE9" colab_type="text"
# ## Create Data
#
# 1. Create an an array of x-values (range between 0 and 1).
# 2. Choose some coefficient theta
# 3. Create the y values: x-values multiplied by theta, plus some gaussian noise
# 4. Plot x,y data
# + id="7zyteipk8S0O" colab_type="code" colab={}
# complete your code here
# real_theta = ??
# + [markdown] id="0KsDxXu2_JZU" colab_type="text"
# ## Goal
#
# Our loss function is:
# $$ L = \frac{1}{2} \Vert y - \theta X \Vert^2_2 $$
#
# Our goal is to solve for best $\theta$ that describes the linear relation between the data (x, y)
#
# + [markdown] id="YM4s6KJZ86zn" colab_type="text"
# ## Optional: Solve for theta using normal equations
# + id="6fmtxGvg898J" colab_type="code" colab={}
# complete your code here
# + [markdown] id="UQxRviVj9SGV" colab_type="text"
# ## Solving using gradient descent
#
# 1. compute the analytic gradient $\frac{\partial L}{\partial \theta}$
# 2. solve for theta using gradient descent method
# 3. change the gradient computation to that give by pytorch (instead of using the analytic one)
# + id="tB8ApQT79ARB" colab_type="code" colab={}
# %matplotlib inline
import time
import pylab as pl
from IPython import display
# theta = ? # <-------- fill initial theta in ??
plt.ion()
plt.plot(X.squeeze().numpy(), y.squeeze().numpy(), '.b')
graph = plt.plot(X.squeeze().numpy(), torch.zeros_like(X).squeeze().numpy(), 'r')[0]
losss = []
for i in range(T):
# compute next theta using gradient descent
# Next lines are for visualization (uncomment when you're ready)
##########################################################################
# # plot intermediate
# loss = ?? ; losss.append(loss.item()) # <------ fill loss in ??
# if i % 10 == 0:
# print('{}: loss: {:.3g} theta: {:.3g}'.format(i, loss.item(), theta))
# display.clear_output(wait=True)
# graph.set_ydata((theta * X).squeeze().numpy())
# display.display(plt.gcf())
# time.sleep(0.1)
##########################################################################
print('real theta: ', real_theta)
print('found theta:', theta)
# plt.figure(); plt.plot(range(len(losss)), losss, 'k')
# + [markdown] id="jOH27XJU9_Ha" colab_type="text"
# ## Bonus Questions:
#
#
# (1) Add bias to data and solve for both theta and data using gradient descent (optional: using normal equations)
#
# (2a) Create data for some polynom of degree d. Solve for coefficients using gradient descent (optional: using normal equations).
#
# (2b) Create data with few samples (~5-10). What do you see for larger d?
# + id="dKp9lUnh-8g_" colab_type="code" colab={}
| 01-hands_on (niv).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="1czVdIlqnImH"
# # Evaluating GANs
# + [markdown] colab_type="text" id="1KD3ZgLs80vY"
# ### Goals
# In this notebook, you're going to gain a better understanding of some of the challenges that come with evaluating GANs and a response you can take to alleviate some of them called Fréchet Inception Distance (FID).
#
# ### Learning Objectives
# 1. Understand the challenges associated with evaluating GANs.
# 2. Write code to evaluate the Fréchet Inception Distance.
#
#
# + [markdown] colab_type="text" id="YcjvPOSMSCzL"
# ## Challenges With Evaluating GANs
#
# #### Loss is Uninformative of Performance
# One aspect that makes evaluating GANs challenging is that the loss tells us little about their performance. Unlike with classifiers, where a low loss on a test set indicates superior performance, a low loss for the generator or discriminator suggests that learning has stopped.
#
#
# #### No Clear Non-human Metric
# If you define the goal of a GAN as "generating images which look real to people" then it's technically possible to measure this directly: [you can ask people to act as a discriminator](https://arxiv.org/abs/1904.01121). However, this takes significant time and money so ideally you can use a proxy for this. There is also no "perfect" discriminator that can differentiate reals from fakes - if there were, a lot of machine learning tasks would be solved ;)
#
# In this notebook, you will implement Fréchet Inception Distance, one method which aims to solve these issues.
# + [markdown] colab_type="text" id="wU8DDM6l9rZb"
# ## Getting Started
# For this notebook, you will again be using [CelebA](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html). You will start by loading a pre-trained generator which has been trained on CelebA.
# + [markdown] colab_type="text" id="gsTReUckdFQh"
# Here, you will import some useful libraries and packages. You will also be provided with the generator and noise code from earlier assignments.
# + colab={} colab_type="code" id="JfkorNJrnmNO"
import torch
import numpy as np
from torch import nn
from tqdm.auto import tqdm
from torchvision import transforms
from torchvision.datasets import CelebA
from torchvision.utils import make_grid
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
torch.manual_seed(0) # Set for our testing purposes, please do not change!
class Generator(nn.Module):
'''
Generator Class
Values:
z_dim: the dimension of the noise vector, a scalar
im_chan: the number of channels in the images, fitted for the dataset used, a scalar
(CelebA is rgb, so 3 is your default)
hidden_dim: the inner dimension, a scalar
'''
def __init__(self, z_dim=10, im_chan=3, hidden_dim=64):
super(Generator, self).__init__()
self.z_dim = z_dim
# Build the neural network
self.gen = nn.Sequential(
self.make_gen_block(z_dim, hidden_dim * 8),
self.make_gen_block(hidden_dim * 8, hidden_dim * 4),
self.make_gen_block(hidden_dim * 4, hidden_dim * 2),
self.make_gen_block(hidden_dim * 2, hidden_dim),
self.make_gen_block(hidden_dim, im_chan, kernel_size=4, final_layer=True),
)
def make_gen_block(self, input_channels, output_channels, kernel_size=3, stride=2, final_layer=False):
'''
Function to return a sequence of operations corresponding to a generator block of DCGAN;
a transposed convolution, a batchnorm (except in the final layer), and an activation.
Parameters:
input_channels: how many channels the input feature representation has
output_channels: how many channels the output feature representation should have
kernel_size: the size of each convolutional filter, equivalent to (kernel_size, kernel_size)
stride: the stride of the convolution
final_layer: a boolean, true if it is the final layer and false otherwise
(affects activation and batchnorm)
'''
if not final_layer:
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.BatchNorm2d(output_channels),
nn.ReLU(inplace=True),
)
else:
return nn.Sequential(
nn.ConvTranspose2d(input_channels, output_channels, kernel_size, stride),
nn.Tanh(),
)
def forward(self, noise):
'''
Function for completing a forward pass of the generator: Given a noise tensor,
returns generated images.
Parameters:
noise: a noise tensor with dimensions (n_samples, z_dim)
'''
x = noise.view(len(noise), self.z_dim, 1, 1)
return self.gen(x)
def get_noise(n_samples, z_dim, device='cpu'):
'''
Function for creating noise vectors: Given the dimensions (n_samples, z_dim)
creates a tensor of that shape filled with random numbers from the normal distribution.
Parameters:
n_samples: the number of samples to generate, a scalar
z_dim: the dimension of the noise vector, a scalar
device: the device type
'''
return torch.randn(n_samples, z_dim, device=device)
# + [markdown] colab_type="text" id="qRk_8azSq3tF"
# ## Loading the Pre-trained Model
#
# Now, you can set the arguments for the model and load the dataset:
# * z_dim: the dimension of the noise vector
# * image_size: the image size of the input to Inception (more details in the following section)
# * device: the device type
# + colab={"base_uri": "https://localhost:8080/", "height": 426} colab_type="code" executionInfo={"elapsed": 5223, "status": "error", "timestamp": 1599108736989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjA3YFIDW6U361yE3hLUnt-lzGRhMwlG-B2_mQLtQ=s64", "userId": "16768126786232632622"}, "user_tz": 240} id="UXptQZcwrBrq" outputId="b7ef6127-ba5d-4f7b-c437-c8d75d67a6b0"
z_dim = 64
image_size = 299
device = 'cuda'
transform = transforms.Compose([
transforms.Resize(image_size),
transforms.CenterCrop(image_size),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5)),
])
in_coursera = True # Set this to false if you're running this outside Coursera
if in_coursera:
import numpy as np
data = torch.Tensor(np.load('fid_images_tensor.npz', allow_pickle=True)['arr_0'])
dataset = torch.utils.data.TensorDataset(data, data)
else:
dataset = CelebA(".", download=True, transform=transform)
# + [markdown] colab_type="text" id="VJvvJ7GDVcyi"
# Then, you can load and initialize the model with weights from a pre-trained model. This allows you to use the pre-trained model as if you trained it yourself.
# + colab={} colab_type="code" id="m5-yxnpDVSzv"
gen = Generator(z_dim).to(device)
gen.load_state_dict(torch.load(f"pretrained_celeba.pth", map_location=torch.device(device))["gen"])
gen = gen.eval()
# + [markdown] colab_type="text" id="Zvis3TCd8tmI"
# ## Inception-v3 Network
# Inception-V3 is a neural network trained on [ImageNet](http://www.image-net.org/) to classify objects. You may recall from the lectures that ImageNet has over 1 million images to train on. As a result, Inception-V3 does a good job detecting features and classifying images. Here, you will load Inception-V3 as `inception_model`.
#
# <!--
# In the past, people would use a pretrained Inception network to identify the classes of the objects generated by a GAN and measure how similar the distribution of classes generated was to the true image (using KL divergence). This is known as inception score.
#
# However, there are many problems with this metric. Barratt and Sharma's 2018 "[A Note on the Inception Score](https://arxiv.org/pdf/1801.01973.pdf)" highlights many issues with this approach. Among them, they highlight its instability, its exploitability, and the widespread use of Inception Score on models not trained on ImageNet. -->
#
#
#
# + colab={} colab_type="code" id="1tPRtB993v68"
from torchvision.models import inception_v3
inception_model = inception_v3(pretrained=False)
inception_model.load_state_dict(torch.load("inception_v3_google-1a9a5a14.pth"))
inception_model.to(device)
inception_model = inception_model.eval() # Evaluation mode
# + [markdown] colab_type="text" id="9iYUtcRU-woT"
# ## Fréchet Inception Distance
#
# Fréchet Inception Distance (FID) was proposed as an improvement over Inception Score and still uses the Inception-v3 network as part of its calculation. However, instead of using the classification labels of the Inception-v3 network, it uses the output from an earlier layer—the layer right before the labels. This is often called the feature layer. Research has shown that deep convolutional neural networks trained on difficult tasks, like classifying many classes, build increasingly sophisticated representations of features going deeper into the network. For example, the first few layers may learn to detect different kinds of edges and curves, while the later layers may have neurons that fire in response to human faces.
#
# To get the feature layer of a convolutional neural network, you can replace the final fully connected layer with an identity layer that simply returns whatever input it received, unchanged. This essentially removes the final classification layer and leaves you with the intermediate outputs from the layer before.
#
# <details>
#
# <summary>
# <font size="3" color="green">
# <b>Optional hint for <code><font size="4">inception_model.fc</font></code></b>
# </font>
# </summary>
#
# 1. You may find [torch.nn.Identity()](https://pytorch.org/docs/master/generated/torch.nn.Identity.html) helpful.
#
# </details>
# + colab={} colab_type="code" id="y2KBoaJEXH29"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED CELL: inception_model.fc
# You want to replace the final fully-connected (fc) layer
# with an identity function layer to cut off the classification
# layer and get a feature extractor
#### START CODE HERE ####
inception_model.fc = torch.nn.Identity()
#### END CODE HERE ####
# + colab={} colab_type="code" id="_txoiugobfck"
# UNIT TEST
test_identity_noise = torch.randn(100, 100)
assert torch.equal(test_identity_noise, inception_model.fc(test_identity_noise))
print("Success!")
# + [markdown] colab_type="text" id="BKzLLQI_XLdw"
# ### Fréchet Distance
# Fréchet distance uses the values from the feature layer for two sets of images, say reals and fakes, and compares different statistical properties between them to see how different they are. Specifically, Fréchet distance finds the shortest distance needed to walk along two lines, or two curves, simultaneously. The most intuitive explanation of Fréchet distance is as the "minimum leash distance" between two points. Imagine yourself and your dog, both moving along two curves. If you walked on one curve and your dog, attached to a leash, walked on the other at the same pace, what is the least amount of leash that you can give your dog so that you never need to give them more slack during your walk? Using this, the Fréchet distance measures the similarity between these two curves.
#
# The basic idea is similar for calculating the Fréchet distance between two probability distributions. You'll start by seeing what this looks like in one-dimensional, also called univariate, space.
# + [markdown] colab_type="text" id="fb5z23wVFE3m"
#
# #### Univariate Fréchet Distance
# You can calculate the distance between two normal distributions $X$ and $Y$ with means $\mu_X$ and $\mu_Y$ and standard deviations $\sigma_X$ and $\sigma_Y$, as:
#
# $$d(X,Y) = (\mu_X-\mu_Y)^2 + (\sigma_X-\sigma_Y)^2 $$
#
# Pretty simple, right? Now you can see how it can be converted to be used in multi-dimensional, which is also called multivariate, space.
# + [markdown] colab_type="text" id="f1syunux1Jon"
# #### Multivariate Fréchet Distance
# **Covariance**
#
# To find the Fréchet distance between two multivariate normal distributions, you first need to find the covariance instead of the standard deviation. The covariance, which is the multivariate version of variance (the square of standard deviation), is represented using a square matrix where the side length is equal to the number of dimensions. Since the feature vectors you will be using have 2048 values/weights, the covariance matrix will be 2048 x 2048. But for the sake of an example, this is a covariance matrix in a two-dimensional space:
#
# $\Sigma = \left(\begin{array}{cc}
# 1 & 0\\
# 0 & 1
# \end{array}\right)
# $
#
# The value at location $(i, j)$ corresponds to the covariance of vector $i$ with vector $j$. Since the covariance of $i$ with $j$ and $j$ with $i$ are equivalent, the matrix will always be symmetric with respect to the diagonal. The diagonal is the covariance of that element with itself. In this example, there are zeros everywhere except the diagonal. That means that the two dimensions are independent of one another, they are completely unrelated.
#
# The following code cell will visualize this matrix.
# + colab={} colab_type="code" id="qDLxAxYo-v0y"
#import os
#os.environ['KMP_DUPLICATE_LIB_OK']='True'
from torch.distributions import MultivariateNormal
import seaborn as sns # This is for visualization
mean = torch.Tensor([0, 0]) # Center the mean at the origin
covariance = torch.Tensor( # This matrix shows independence - there are only non-zero values on the diagonal
[[1, 0],
[0, 1]]
)
independent_dist = MultivariateNormal(mean, covariance)
samples = independent_dist.sample((10000,))
res = sns.jointplot(samples[:, 0], samples[:, 1], kind="kde")
plt.show()
# + [markdown] colab_type="text" id="HkEtuo_k2ED5"
# Now, here's an example of a multivariate normal distribution that has covariance:
#
# $\Sigma = \left(\begin{array}{cc}
# 2 & -1\\
# -1 & 2
# \end{array}\right)
# $
#
# And see how it looks:
#
# + colab={} colab_type="code" id="SQvjyAmw2SGl"
mean = torch.Tensor([0, 0])
covariance = torch.Tensor(
[[2, -1],
[-1, 2]]
)
covariant_dist = MultivariateNormal(mean, covariance)
samples = covariant_dist.sample((10000,))
res = sns.jointplot(samples[:, 0], samples[:, 1], kind="kde")
plt.show()
# + [markdown] colab_type="text" id="uf3kci9d2ROC"
# **Formula**
#
# Based on the paper, "[The Fréchet distance between multivariate normal distributions](https://core.ac.uk/reader/82269844)" by <NAME> (1982), the Fréchet distance between two multivariate normal distributions $X$ and $Y$ is:
#
# $d(X, Y) = \Vert\mu_X-\mu_Y\Vert^2 + \mathrm{Tr}\left(\Sigma_X+\Sigma_Y - 2 \sqrt{\Sigma_X \Sigma_Y}\right)$
#
# Similar to the formula for univariate Fréchet distance, you can calculate the distance between the means and the distance between the standard deviations. However, calculating the distance between the standard deviations changes slightly here, as it includes the matrix product and matrix square root. $\mathrm{Tr}$ refers to the trace, the sum of the diagonal elements of a matrix.
#
# Now you can implement this!
#
# <details>
#
# <summary>
# <font size="3" color="green">
# <b>Optional hints for <code><font size="4">frechet_distance</font></code></b>
# </font>
# </summary>
#
# 1. You want to implement the above equation in code.
# 2. You might find the functions `torch.norm` and `torch.trace` helpful here.
# 3. A matrix_sqrt function is defined for you above -- you need to use it instead of `torch.sqrt()` which only gets the elementwise square root instead of the matrix square root.
# 4. You can also use the `@` symbol for matrix multiplication.
# </details>
# + colab={} colab_type="code" id="iOlCmNPiuuhK"
import scipy
# This is the matrix square root function you will be using
def matrix_sqrt(x):
'''
Function that takes in a matrix and returns the square root of that matrix.
For an input matrix A, the output matrix B would be such that B @ B is the matrix A.
Parameters:
x: a matrix
'''
y = x.cpu().detach().numpy()
y = scipy.linalg.sqrtm(y)
return torch.Tensor(y.real, device=x.device)
# + colab={} colab_type="code" id="_hLWk57s91it"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: frechet_distance
def frechet_distance(mu_x, mu_y, sigma_x, sigma_y):
'''
Function for returning the Fréchet distance between multivariate Gaussians,
parameterized by their means and covariance matrices.
Parameters:
mu_x: the mean of the first Gaussian, (n_features)
mu_y: the mean of the second Gaussian, (n_features)
sigma_x: the covariance matrix of the first Gaussian, (n_features, n_features)
sigma_y: the covariance matrix of the second Gaussian, (n_features, n_features)
'''
#### START CODE HERE ####
return (mu_x - mu_y).dot(mu_x - mu_y) + torch.trace(sigma_x) + torch.trace(sigma_y) - 2*torch.trace(matrix_sqrt(sigma_x @ sigma_y))
#### END CODE HERE ####
# + colab={} colab_type="code" id="Pphv97XEgPDh"
# UNIT TEST
mean1 = torch.Tensor([0, 0]) # Center the mean at the origin
covariance1 = torch.Tensor( # This matrix shows independence - there are only non-zero values on the diagonal
[[1, 0],
[0, 1]]
)
dist1 = MultivariateNormal(mean1, covariance1)
mean2 = torch.Tensor([0, 0]) # Center the mean at the origin
covariance2 = torch.Tensor( # This matrix shows dependence
[[2, -1],
[-1, 2]]
)
dist2 = MultivariateNormal(mean2, covariance2)
assert torch.isclose(
frechet_distance(
dist1.mean, dist2.mean,
dist1.covariance_matrix, dist2.covariance_matrix
),
4 - 2 * torch.sqrt(torch.tensor(3.))
)
assert (frechet_distance(
dist1.mean, dist1.mean,
dist1.covariance_matrix, dist1.covariance_matrix
).item() == 0)
print("Success!")
# + [markdown] colab_type="text" id="r9dMgbxGCTno"
# ## Putting it all together!
# Now, you can apply FID to your generator from earlier.
#
# You will start by defining a bit of helper code to preprocess the image for the Inception-v3 network:
# <!-- This isn't exactly what FID is meant for, since inception scores expect a natural image, but it should give a rough idea of the diversity and quality of your images. [TODO: move to bottom since image net is trained on nature (cat, dog) images, fidelity (quality)] -->
# + colab={} colab_type="code" id="s2ZMTdzMCZtn"
def preprocess(img):
img = torch.nn.functional.interpolate(img, size=(299, 299), mode='bilinear', align_corners=False)
return img
# + [markdown] colab_type="text" id="hzXQTFv_UNjm"
# Then, you'll define a function to calculate the covariance of the features that returns a covariance matrix given a list of values:
# + colab={} colab_type="code" id="4BEbwlGLiPWJ"
import numpy as np
def get_covariance(features):
return torch.Tensor(np.cov(features.detach().numpy(), rowvar=False))
# + [markdown] colab_type="text" id="Sw5obaXjVv_v"
# Finally, you can use the pre-trained Inception-v3 model to compute features of the real and fake images. With these features, you can then get the covariance and means of these features across many samples.
#
# First, you get the features of the real and fake images using the Inception-v3 model:
# + colab={} colab_type="code" id="qQwHSAhrf0hX"
fake_features_list = []
real_features_list = []
gen.eval()
n_samples = 512 # The total number of samples
batch_size = 4 # Samples per iteration
dataloader = DataLoader(
dataset,
batch_size=batch_size,
shuffle=True)
cur_samples = 0
with torch.no_grad(): # You don't need to calculate gradients here, so you do this to save memory
try:
for real_example, _ in tqdm(dataloader, total=n_samples // batch_size): # Go by batch
real_samples = real_example
real_features = inception_model(real_samples.to(device)).detach().to('cpu') # Move features to CPU
real_features_list.append(real_features)
fake_samples = get_noise(len(real_example), z_dim).to(device)
fake_samples = preprocess(gen(fake_samples))
fake_features = inception_model(fake_samples.to(device)).detach().to('cpu')
fake_features_list.append(fake_features)
cur_samples += len(real_samples)
if cur_samples >= n_samples:
break
except:
print("Error in loop")
# + [markdown] colab_type="text" id="LUrJ_ZEZXkvu"
# Then, you can combine all of the values that you collected for the reals and fakes into large tensors:
# + colab={} colab_type="code" id="UmiOuDulqDTC"
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# UNIT TEST COMMENT: Needed as is for autograding
fake_features_all = torch.cat(fake_features_list)
real_features_all = torch.cat(real_features_list)
# + [markdown] colab_type="text" id="41XEEZRoXxN1"
# And calculate the covariance and means of these real and fake features:
# + colab={} colab_type="code" id="quNhtmwaNhpT"
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED CELL
# Calculate the covariance matrix for the fake and real features
# and also calculate the means of the feature over the batch (for each feature dimension mean)
#### START CODE HERE ####
mu_fake = fake_features_all.mean(0)
mu_real = real_features_all.mean(0)
sigma_fake = get_covariance(fake_features_all)
sigma_real = get_covariance(real_features_all)
#### END CODE HERE ####
# + colab={} colab_type="code" id="zC1spPHPxkOQ"
assert tuple(sigma_fake.shape) == (fake_features_all.shape[1], fake_features_all.shape[1])
assert torch.abs(sigma_fake[0, 0] - 2.5e-2) < 1e-2 and torch.abs(sigma_fake[-1, -1] - 5e-2) < 1e-2
assert tuple(sigma_real.shape) == (real_features_all.shape[1], real_features_all.shape[1])
assert torch.abs(sigma_real[0, 0] - 3.5768e-2) < 1e-4 and torch.abs(sigma_real[0, 1] + 5.3236e-4) < 1e-4
assert tuple(mu_fake.shape) == (fake_features_all.shape[1],)
assert tuple(mu_real.shape) == (real_features_all.shape[1],)
assert torch.abs(mu_real[0] - 0.3099) < 0.01 and torch.abs(mu_real[1] - 0.2721) < 0.01
assert torch.abs(mu_fake[0] - 0.37) < 0.05 and torch.abs(mu_real[1] - 0.27) < 0.05
print("Success!")
# + [markdown] colab_type="text" id="gyHx-bBqo5MU"
# At this point, you can also visualize what the pairwise multivariate distributions of the inception features look like!
# + colab={} colab_type="code" id="fBOp8wfNkce1"
indices = [2, 4, 5]
fake_dist = MultivariateNormal(mu_fake[indices], sigma_fake[indices][:, indices])
fake_samples = fake_dist.sample((5000,))
real_dist = MultivariateNormal(mu_real[indices], sigma_real[indices][:, indices])
real_samples = real_dist.sample((5000,))
import pandas as pd
df_fake = pd.DataFrame(fake_samples.numpy(), columns=indices)
df_real = pd.DataFrame(real_samples.numpy(), columns=indices)
df_fake["is_real"] = "no"
df_real["is_real"] = "yes"
df = pd.concat([df_fake, df_real])
sns.pairplot(df, plot_kws={'alpha': 0.1}, hue='is_real')
# + [markdown] colab_type="text" id="Ja7DQ9SSX0u2"
# Lastly, you can use your earlier `frechet_distance` function to calculate the FID and evaluate your GAN. You can see how similar/different the features of the generated images are to the features of the real images. The next cell might take five minutes or so to run in Coursera.
# + colab={} colab_type="code" id="qS8kklILkmyg"
with torch.no_grad():
print(frechet_distance(mu_real, mu_fake, sigma_real, sigma_fake).item())
# + [markdown] colab_type="text" id="Rnt8jEP6Fh70"
# You'll notice this model gets a pretty high FID, likely over 30. Since lower is better, and the best models on CelebA get scores in the single-digits, there's clearly a long way to go with this model. You can use FID to compare different models, as well as different stages of training of the same model.
| Evaluating_GANs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
a = np.asarray([[1,2,3,4], [5,6,7,8]])
def mod_arr(a):
a[0,0] = 100
return a
print(a)
print(mod_arr(a))
print(a)
# +
a = np.asarray([[1,2,3,4], [5,6,7,8]])
def mod_arr(a):
x = a
x[0,0] = 100
return x
print(a)
print(mod_arr(a))
print(a)
# -
import pandas as pd
df=pd.DataFrame({'banana':[1,2,3,4], 'bean':[5,6,7,8]})
x = df['banana'] # this is not assigned by value, but by reference!!!!
df
x
x[3] = 10
x
df
# +
# so it changes the dataframe!!!
# +
# This doesn't happen to immuntable types, like strings:
# +
a = 'babbelbubbel'
def mod_arr(x):
#x = a
x = 'Z'+x
return x
print(a)
print(mod_arr(a))
print(a)
# -
| notebooks/.ipynb_checkpoints/pass_by_ref_or_value_python-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Consul Intro
#
# Please see the [README.md](https://github.com/micahhausler/consul-demo/blob/master/README.md) and get your consul cluster up and running
# ## Setting a key/value with `requests`
# For full documentation, see the [Consul http-api](https://www.consul.io/docs/agent/http.html)
# +
import requests
base_url = 'http://192.168.59.103:8500/v1/kv/'
response = requests.put(base_url + 'key1', data="value1")
print(response.text)
# -
# ## Getting a key/value with `python-consul`
# For full documentation, see python-consul [documentation](http://python-consul.readthedocs.org/en/latest/#consul-kv)
# +
from consul import Consul
c = Consul('192.168.59.103')
index, data = c.kv.get('key1')
print(data['Value'].decode('utf8'))
# -
# ## Other features:
#
# * Locks
# * DNS for service discovery
# * Limit access to key/value subsets with ACL's and tokens
| Consul.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python で気軽に化学・化学工学
# # 第 8 章 モデル y = f(x) を構築して、新たなサンプルの y を推定する
# ## 8.7.4 ランダムフォレスト (Random Forests, RF)
# ### 回帰分析
# ## Jupyter Notebook の有用なショートカットのまとめ
# - <kbd>Esc</kbd>: コマンドモードに移行(セルの枠が青)
# - <kbd>Enter</kbd>: 編集モードに移行(セルの枠が緑)
# - コマンドモードで <kbd>M</kbd>: Markdown セル (説明・メモを書く用) に変更
# - コマンドモードで <kbd>Y</kbd>: Code セル (Python コードを書く用) に変更
# - コマンドモードで <kbd>H</kbd>: ヘルプを表示
# - コマンドモードで <kbd>A</kbd>: ひとつ**上**に空のセルを挿入
# - コマンドモードで <kbd>B</kbd>: ひとつ**下**に空のセルを挿入
# - コマンドモードで <kbd>D</kbd><kbd>D</kbd>: セルを削除
# - <kbd>Ctrl</kbd>+<kbd>Enter</kbd>: セルの内容を実行
# - <kbd>Shift</kbd>+<kbd>Enter</kbd>: セルの内容を実行して下へ
# わからないことがありましたら、関係する単語やエラーの文章などでウェブ検索してご自身で調べてみましょう。
# ### 沸点のデータセット (descriptors_8_with_boiling_point.csv)
# Hall and Story が収集した[沸点のデータセット](https://pubs.acs.org/doi/abs/10.1021/ci960375x)。294 個の化合物について、沸点 (Boiling Point) が測定されており、8 つの分子記述子 (特徴量) で化学構造が数値化されています。特徴量は、分子量 (MolWt)、水素原子以外の原子で計算された分子量 (HeavyAtomMolWt)、価電子の数 (NumValenceElectrons)、水素原子以外の原子の数 (HeavyAtomCount)、窒素原子と酸素原子の数 (NOCount)、水素原子と炭素原子以外の原子の数 (NumHeteroatoms)、回転可能な結合の数 (NumRotatableBonds)、環の数 (RingCount) です。
import pandas as pd # pandas のインポート
dataset = pd.read_csv('descriptors_8_with_boiling_point.csv', index_col=0, header=0) # 沸点のデータセットの読み込み
x = dataset.iloc[:, 1:] # 記述子を 説明変数 x とします
y = dataset.iloc[:, 0] # 沸点を目的変数 y とします
# トレーニングデータとテストデータの分割
from sklearn.model_selection import train_test_split
# ランダムにトレーニングデータとテストデータとに分割。random_state に数字を与えることで、別のときに同じ数字を使えば、ランダムとはいえ同じ結果にすることができます
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=94, shuffle=True, random_state=99)
# DT モデルと同様にして RF モデルにおいても、一般的に x や y の標準化 (オートスケーリング) は行いません。
# RF の実行
from sklearn.ensemble import RandomForestRegressor # 回帰分析用の RF の実行に使用
model = RandomForestRegressor(n_estimators=500, max_features=0.5, oob_score=True) # RF モデルの宣言
model.fit(x_train, y_train) # RF モデル構築
# 構築された RF モデルにおける説明変数 x の重要度
model.feature_importances_ # 特徴量の重要度。array 型で出力されます
importances = pd.DataFrame(model.feature_importances_) # pandas の DataFrame 型に変換
importances.index = x_train.columns # 説明変数に対応する名前を、元のデータの説明変数名に
importances.columns = ['importances'] # 列名を変更
importances # 念のため確認
importances.to_csv('importances.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# Out Of Bag (OOB) における r<sup>2</sup>
model.oob_score_
# トレーニングデータの y の値の推定
estimated_y_train = pd.DataFrame(model.predict(x_train)) # トレーニングデータの y の値を推定し、pandas の DataFrame 型に変換
estimated_y_train.index = x_train.index # サンプル名を、元のデータのサンプル名に
estimated_y_train.columns = ['estimated_y'] # 列名を変更
estimated_y_train # 念のため確認
estimated_y_train.to_csv('estimated_y_train.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# トレーニングデータの y の実測値 vs. 推定値プロット
import matplotlib.pyplot as plt
import matplotlib.figure as figure # 図の調整に使用
plt.rcParams['font.size'] = 18 # 横軸や縦軸の名前の文字などのフォントのサイズ
plt.figure(figsize=figure.figaspect(1)) # 図の形を正方形に
plt.scatter(y_train, estimated_y_train.iloc[:, 0]) # 散布図。estimated_y_train は 200×1 の行列のため、0 列目を選択する必要があります
y_max = max(y_train.max(), estimated_y_train.iloc[:, 0].max()) # 実測値の最大値と、推定値の最大値の中で、より大きい値を取得
y_min = min(y_train.min(), estimated_y_train.iloc[:, 0].min()) # 実測値の最小値と、推定値の最小値の中で、より小さい値を取得
plt.plot([y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], [y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], 'k-') # 取得した最小値-5%から最大値+5%まで、対角線を作成
plt.ylim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # y 軸の範囲の設定
plt.xlim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # x 軸の範囲の設定
plt.xlabel("actual y") # x 軸の名前
plt.ylabel("estimated y") # y 軸の名前
plt.show() # 以上の設定で描画
# トレーニングデータの r<sup>2</sup>, MAE
from sklearn import metrics
metrics.r2_score(y_train, estimated_y_train) # r2
metrics.mean_absolute_error(y_train, estimated_y_train) # MAE
# テストデータの y の値の推定。トレーニングデータをテストデータに変えるだけで、実行する内容はトレーニングデータのときと同じです
estimated_y_test = pd.DataFrame(model.predict(x_test)) # テストデータの y の値を推定し、pandas の DataFrame 型に変換
estimated_y_test.index = x_test.index # サンプル名を、元のデータのサンプル名に
estimated_y_test.columns = ['estimated_y'] # 列名を変更
estimated_y_test # 念のため確認
estimated_y_test.to_csv('estimated_y_test.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# テストデータの y の実測値 vs. 推定値プロット
plt.rcParams['font.size'] = 18 # 横軸や縦軸の名前の文字などのフォントのサイズ
plt.figure(figsize=figure.figaspect(1)) # 図の形を正方形に
plt.scatter(y_test, estimated_y_test.iloc[:, 0]) # 散布図。estimated_y_train は 200×1 の行列のため、0 列目を選択する必要があります
y_max = max(y_test.max(), estimated_y_test.iloc[:, 0].max()) # 実測値の最大値と、推定値の最大値の中で、より大きい値を取得
y_min = min(y_test.min(), estimated_y_test.iloc[:, 0].min()) # 実測値の最小値と、推定値の最小値の中で、より小さい値を取得
plt.plot([y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], [y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], 'k-') # 取得した最小値-5%から最大値+5%まで、対角線を作成
plt.ylim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # y 軸の範囲の設定
plt.xlim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # x 軸の範囲の設定
plt.xlabel("actual y") # x 軸の名前
plt.ylabel("estimated y") # y 軸の名前
plt.show() # 以上の設定で描画
# テストデータの r<sup>2</sup>, MAE
metrics.r2_score(y_test, estimated_y_test) # r2
metrics.mean_absolute_error(y_test, estimated_y_test) # MAE
# OOB を用いた説明変数 x の割合の最適化
import numpy as np # NumPy のインポート
ratios_of_x = np.arange(0.1, 1.1, 0.1) # 用いる説明変数の割合の候補
ratios_of_x # 念のため確認
r2_oob = [] # 空の list。説明変数の数の割合ごとに、OOB における r2 を入れていきます
for ratio_of_x in ratios_of_x:
model = RandomForestRegressor(n_estimators=500, max_features=ratio_of_x, oob_score=True, random_state=1)
model.fit(x_train, y_train)
r2_oob.append(model.oob_score_)
import matplotlib.pyplot as plt # 図の描画に使用
# 結果の確認
plt.rcParams['font.size'] = 18
plt.scatter(ratios_of_x, r2_oob)
plt.xlabel('ratio of x')
plt.ylabel('r2 for OOB')
plt.show()
optimal_ratio_of_x = ratios_of_x[r2_oob.index(max(r2_oob))] # OOB における r2 が最大となる選択する x の割合
optimal_ratio_of_x # 念のため確認
# RF モデルの構築および予測
model = RandomForestRegressor(n_estimators=500, max_features=optimal_ratio_of_x, oob_score=True)
model.fit(x_train, y_train) # RF モデル構築
# 構築された RF モデルにおける説明変数 x の重要度
model.feature_importances_ # 特徴量の重要度。array 型で出力されます
importances = pd.DataFrame(model.feature_importances_) # pandas の DataFrame 型に変換
importances.index = x_train.columns # 説明変数に対応する名前を、元のデータの説明変数名に
importances.columns = ['importances'] # 列名を変更
importances # 念のため確認
importances.to_csv('importances.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# トレーニングデータの y の値の推定
estimated_y_train = pd.DataFrame(model.predict(x_train)) # トレーニングデータの y の値を推定し、pandas の DataFrame 型に変換
estimated_y_train.index = x_train.index # サンプル名を、元のデータのサンプル名に
estimated_y_train.columns = ['estimated_y'] # 列名を変更
estimated_y_train # 念のため確認
estimated_y_train.to_csv('estimated_y_train.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# トレーニングデータの y の実測値 vs. 推定値プロット
import matplotlib.pyplot as plt
import matplotlib.figure as figure # 図の調整に使用
plt.rcParams['font.size'] = 18 # 横軸や縦軸の名前の文字などのフォントのサイズ
plt.figure(figsize=figure.figaspect(1)) # 図の形を正方形に
plt.scatter(y_train, estimated_y_train.iloc[:, 0]) # 散布図。estimated_y_train は 200×1 の行列のため、0 列目を選択する必要があります
y_max = max(y_train.max(), estimated_y_train.iloc[:, 0].max()) # 実測値の最大値と、推定値の最大値の中で、より大きい値を取得
y_min = min(y_train.min(), estimated_y_train.iloc[:, 0].min()) # 実測値の最小値と、推定値の最小値の中で、より小さい値を取得
plt.plot([y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], [y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], 'k-') # 取得した最小値-5%から最大値+5%まで、対角線を作成
plt.ylim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # y 軸の範囲の設定
plt.xlim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # x 軸の範囲の設定
plt.xlabel("actual y") # x 軸の名前
plt.ylabel("estimated y") # y 軸の名前
plt.show() # 以上の設定で描画
# トレーニングデータの r<sup>2</sup>, MAE
from sklearn import metrics
metrics.r2_score(y_train, estimated_y_train) # r2
metrics.mean_absolute_error(y_train, estimated_y_train) # MAE
# テストデータの y の値の推定。トレーニングデータをテストデータに変えるだけで、実行する内容はトレーニングデータのときと同じです
estimated_y_test = pd.DataFrame(model.predict(x_test)) # テストデータの y の値を推定し、pandas の DataFrame 型に変換
estimated_y_test.index = x_test.index # サンプル名を、元のデータのサンプル名に
estimated_y_test.columns = ['estimated_y'] # 列名を変更
estimated_y_test # 念のため確認
estimated_y_test.to_csv('estimated_y_test.csv') # csv ファイルに保存。同じ名前のファイルがあるときは上書きされますので注意してください
# テストデータの y の実測値 vs. 推定値プロット
plt.rcParams['font.size'] = 18 # 横軸や縦軸の名前の文字などのフォントのサイズ
plt.figure(figsize=figure.figaspect(1)) # 図の形を正方形に
plt.scatter(y_test, estimated_y_test.iloc[:, 0]) # 散布図。estimated_y_train は 200×1 の行列のため、0 列目を選択する必要があります
y_max = max(y_test.max(), estimated_y_test.iloc[:, 0].max()) # 実測値の最大値と、推定値の最大値の中で、より大きい値を取得
y_min = min(y_test.min(), estimated_y_test.iloc[:, 0].min()) # 実測値の最小値と、推定値の最小値の中で、より小さい値を取得
plt.plot([y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], [y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)], 'k-') # 取得した最小値-5%から最大値+5%まで、対角線を作成
plt.ylim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # y 軸の範囲の設定
plt.xlim(y_min - 0.05 * (y_max - y_min), y_max + 0.05 * (y_max - y_min)) # x 軸の範囲の設定
plt.xlabel("actual y") # x 軸の名前
plt.ylabel("estimated y") # y 軸の名前
plt.show() # 以上の設定で描画
# テストデータの r<sup>2</sup>, MAE
metrics.r2_score(y_test, estimated_y_test) # r2
metrics.mean_absolute_error(y_test, estimated_y_test) # MAE
# 自分のデータセットをお持ちの方は、そのデータセットでも今回の内容を確認してみましょう。
| sample_program_8_7_4_rf_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PhilReinholdPygrape tutorial 5: Measurement and register operations
#
# This notebook is based on the original examples written by Phil,
# they can be found in [this directory](https://github.com/tesla-cat/yarn/tree/master/examplesFromForkedLibraries/PhilReinholdPygrape).
# Specifically, it contains the following:
#
# - `cavity_register.py`
#
# Ruiqi, 28 Jun 2020
import numpy as np
from qutip import *
from qutip.qip.algorithms import qft
from yarn.PhilReinholdPygrape import *
from yarn.qutipHelpers import (
plotWignersIntermediateStates,
plotOccupationsIntermediateStates,
plotExpectation, cat
)
# ## parameters
# ### 1. operation
# +
operateOnQubitN = 1
operationIsCoherent = True
operationTypes = ['measurement','register']
operationType = operationTypes[0]
measurementShouldInvert = True
measurementNames = ['projectionX','projectionY','projectionZ']
measurementName = measurementNames[0]
operationName = 'qft'
# -
# ### 2. system
dimC = 20
dimQ = 2
numQ = 3
# ### 3. frequency (GHz)
driveC = driveQ = 1e-3
chi = -2.95e-3
chiPrime = 1e-6
anharmonicityQ = -215e-3
kerr = -5.4e-6
# ### 4. time (ns)
relaxationC = 2e7
relaxationQ = 70e3
dephasingQ = 22e3
# ### 5. optimization
useLoss = True
maxIter = 120 * 2**numQ
maxAmp = 16
# ## operators
# ### 1. projection operators for measurement
g, e = fock(2,0), fock(2,1)
projectionZ = ket2dm( e )
projectionX = ket2dm( (g + e).unit() )
projectionY = ket2dm( (g + 1j*e).unit() )
# ### 2. operators for register
# +
pauli = {'x':sigmax(), 'y':sigmay(), 'z':sigmaz()}
def rotation(axis,angle):
return ( 1j*angle/2*pauli[axis] ).expm()
pauliX = sigmax()
pauliZ = sigmaz()
rotationXpiOver2 = rotation('x',np.pi/2)
rotationYpiOver2 = rotation('y',np.pi/2)
# -
# ## define the grape setup
# +
inits = np.concatenate([
np.array([tensor(basis(dimQ, 0), basis(dimC, i)).full()[:,0] for i in range(n)]),
np.array([tensor(basis(dimQ, 1), basis(dimC, i)).full()[:,0] for i in range(n)]),
])
U = np.identity(dimC*dimQ, dtype=np.complex)
U[:n,:n] = final_op[:n,:n]
U[dimC:dimC+n,dimC:dimC+n] = final_op[n:,n:]
U[:n,dimC:dimC+n] = final_op[:n,n:]
U[dimC:dimC+n,:n] = final_op[n:,:n]
finals = []
for i in range(inits.shape[0]):
finals.append(np.dot(U,inits[i,:]))
finals = np.array(finals)
print(inits)
print(finals)
# -
def makeSetup(dimC, dimQ):
Hdrift, HcontrolList = make_hmt(
dimC, dimQ, chi, chiPrime,
kerr, anharmonicityQ, driveQ, driveC
)
# Incoherent, don't need cavity drive
if not operationIsCoherent:
HcontrolList = HcontrolList[:2,:,:]
if operationType == 'measurement':
identities = projections = flips = [qeye(2) for _ in range(numQ+1)]
identity = tensor(identities)
# project
projections[operateOnQubitN-1] = eval(measurementName)
projections.reverse() # why?
project = tensor(projections)
# flip
flips[-1] = sigmax()
flips.reverse()
flip = tensor(flips)
# operation = flip * <project+> + identity * <project->
if not measurementShouldInvert:
operation = flip * project + (identity - project)
else:
operation = project + flip * (identity - project)
inits = np.concatenate([
np.array([qutip.tensor(qutip.basis(nq, 0), qutip.basis(nc, i)).full()[:,0] for i in range(n)]),
np.array([qutip.tensor(qutip.basis(nq, 1), qutip.basis(nc, i)).full()[:,0] for i in range(n)]),
])
U = np.identity(nc*nq, dtype=np.complex)
U[:n,:n] = final_op[:n,:n]
U[nc:nc+n,nc:nc+n] = final_op[n:,n:]
U[:n,nc:nc+n] = final_op[:n,n:]
U[nc:nc+n,:n] = final_op[n:,:n]
finals = []
for i in range(inits.shape[0]):
finals.append(np.dot(U,inits[i,:]))
finals = np.array(finals)
# Register operations
else:
ops = [qutip.identity(2) for _ in range(N_BITS)]
if opname == 'x':
ops[BIT_N-1] = qutip.sigmax()
elif opname == 'x2':
ops[BIT_N-1] = (1j*np.pi/4*qutip.sigmax()).expm()
elif opname == 'y2':
ops[BIT_N-1] = (1j*np.pi/4*qutip.sigmay()).expm()
elif opname == 'z':
ops[BIT_N-1] = qutip.sigmaz()
elif opname == 'qft':
xs = np.arange(n) / float(n)
ops = [qutip.Qobj(np.array(
[1/np.sqrt(n)*np.exp(1j*2*np.pi*i*xs)
for i in range(n)]))]
ops.reverse()
op = qutip.tensor(ops).full()
inits = np.array([qutip.tensor(qutip.basis(nq, 0), qutip.basis(nc, i)).full()[:,0] for i in range(n)])
finals = np.zeros((n,nq*nc), dtype=np.complex)
finals[:n,:n] = op
a, ad, b, bd = make_ops(nc, nq)
if USE_LOSS:
loss_vec = np.ones(nc*nq) - ((a.dag()*a) * 0.5/T1cav + (b.dag()*b) * 0.5/T1q).diag()
else:
loss_vec = None
return StateTransferSetup(H0, Hcs, inits, finals, coherent=coherent, loss_vec=loss_vec)
| trash/PhilReinholdPygrape-tutorial-5-Measurement-and-register-operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="FA0fl6F8bWgl"
# ## CCNSS 2018 Module 1: Neurons, synapses and networks
# # Tutorial 1: Wilson-Cowan equations
# [source](https://colab.research.google.com/drive/16strzPZxTEqR2owgSh6NNLlj2j7MNOQb)
#
# Please execute the cell below to initalise the notebook environment.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="I3mWe1tibWgn"
import matplotlib.pyplot as plt # import matplotlib
import numpy as np # import numpy
import scipy as sp # import scipy
import math # import basic math functions
import random # import basic random number generator functions
fig_w, fig_h = (6, 4)
plt.rcParams.update({'figure.figsize': (fig_w, fig_h)})
# + [markdown] colab_type="text" id="3ZsFEVzWbWgp"
# ## Objectives
# In this notebook we will introduce the *Wilson-Cowan* rate model, and use it to learn more about phase planes, nullclines, and attractors.
#
# ** Background paper:**
# * <NAME> and <NAME> (1972) Excitatory and inhibitory interactions in localized populations of model neurons. Biophysical Journal 12.
#
# + [markdown] colab_type="text" id="vrj-kuMQbWgq"
# ## Background
#
# The Wilson-Cowan equations model the mean-field (i.e., average across the population) dynamics of two coupled populations of excitatory (E) and inhibitory (I) neurons:
#
# \begin{align}
# &\tau_E \frac{dE}{dt} = -E + (1 - r E) F(w_{EE}E -w_{EI}I + I_{ext};a,\theta)\\
# &\tau_I \frac{dI}{dt} = -I + (1 - r I) F(w_{IE}E -w_{II}I;a,\theta)
# \end{align}
#
# $E(t)$ represents the average activation of the excitatory population, and $I(t)$ the activation of the inhibitory population. The parameters $\tau_E$ and $\tau_I$ control the timescales of each population. The connection strengths are given by: $w_{EE}$ (E to E), $w_{EI}$ (I to E), $w_{IE}$ (E to I), and $w_{II}$ (I to I). Refractory effects are modelled through the parameter $r$, and $I_{ext}$ represents external input to the excitatory population.
#
#
#
# The function F describes the population activation function. We assume F to be a sigmoidal function, which is parameterized by its gain $a$ and threshold $\theta$.
#
# $$ F(x;a,\theta) = \frac{1}{1+\exp\{-a(x-\theta)\}} - \frac{1}{1+\exp\{a\theta\}}$$
#
# The argument $x$ represents the input to the population. Note that the the second term is chosen so that $F(0;a,\theta)=0$.
#
# To start, execute the cell below to initialise the simulation parameters.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="IjzkBDOubWgr"
dt = 0.1
# Connection weights
wEE = 12
wEI = 4
wIE = 13
wII = 11
# Refractory parameter
r = 1
# External input
I_ext = 0
# Excitatory parameters
tau_E = 1 # Timescale of excitatory population
a_E = 1.2 # Gain of excitatory population
theta_E = 2.8 # Threshold of excitatory population
# Inhibitory parameters
tau_I = 1 # Timescale of inhibitory population
a_I = 1 # Gain of inhibitory population
theta_I = 4 # Threshold of inhibitory population
# + [markdown] colab_type="text" id="-bwsyFppbWgt"
# **EXERCISE 1**
#
# Fill in the function below to define the activation function F as a function of its input x, and arguments a, and $\theta$. Verify your function by evaluating the excitatory activation function for $x = 0,3,6$. Then plot F for both E and I population parameters over $0 \leq x \leq 10$.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="vV95DJ7HSux3"
def F(x,a,theta):
"""Population activation function.
Arguments:
x -- the population input
a -- the gain of the function
theta -- the threshold of the function
Returns:
y -- the population activation response
"""
# insert your code here
return y
# insert your code here
# + [markdown] colab_type="text" id="M79NVeesQlaI"
# **EXPECTED OUTPUT**
#
# ```
# 0.0
# 0.5261444259857104
# 0.9453894296980492
# ```
# 
# + [markdown] colab_type="text" id="2yrNXJIybWgy"
# **Exercise 2:** Fill in the function below to simulate the dynamics of the Wilson-Cowan equation for up to $t_{max}=15$ with steps of $dt$. Remember from the LIF tutorial that we can numerically integrate the ODEs by replacing the derivatives with their discretized approximations:
#
# \begin{align}
# &\frac{dE}{dt} \to \frac{E[k+\Delta t]-E[k]}{\Delta t} \hspace{5 mm}\text{ and }\hspace{5mm}\frac{dI}{dt} \to \frac{I[k+\Delta t]-I[k]}{\Delta t}\\
# \end{align}
#
# Then simulate the dynamics of the population starting from initial condition $E_0=I_0=0.2$ and plot the results. What is the steady state solution? Then, also plot the dynamics starting from $E_0=I_0=0.25$ and plot the solution (in dashed lines). Now what is the steady state solution?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="lSWgVDrlUzOn"
def simulate_wc(t,E0,I0):
"""Simulate the Wilson-Cowan equations.
Arguments:
t -- time (vector)
E0 -- initial condition weeof the excitatory population
I0 -- initial condition of the inhibitory population
Returns:
E -- Activity of excitatory population (vector)
I -- Activity of inhibitory population (vector)
"""
# insert your code here
return E,I
# insert your code here
# + [markdown] colab_type="text" id="UPcE4xdqVElr"
# **EXPECTED OUTPUT**
#
# 
# + [markdown] colab_type="text" id="T4-eiSSCbWg1"
# **Exercise 3:** Now use the same function to simulate the Wilson Cowan equations for different initial conditions from $0.01 \leq E_0 \leq 1$ and $0.01 \leq I_0 \leq 1$ with stepsize 0.1. For each initial condition, find the steady state value to which $E$ and $I$ converge. There are several ways to do this. A simple way to do this is to check, for each initial condition, that the last two points in the simulation are within 1% of each other:
#
# $$ \frac{E(t_{max})-E(t_{max}-dt)}{E(t_{max})} \leq 0.01 $$
#
# Use the following code within your for loops to throw an error in case the trajectories have not converged:
# ``raise ValueError('Has not converged.')``
#
# Then you can just keep increasing $t_{max}$ until every initial condition converges. Plot the steady state values ($E$ vs. $I$) What fixed points do you see?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="WT8ZhrW5eDYU"
# insert your code here
# + [markdown] colab_type="text" id="7xeKqybCatOT"
# **EXPECTED OUTPUT**
#
# 
# + [markdown] colab_type="text" id="cgiP1SJKbWg6"
# **Exercise 4**: To make the phase plane plot, we first need to determine the inverse of F. To calculate the inverse, set y = F(x), and then solve for x. Then, fill out the function below to define the inverse activation function $F^{-1}$. Check that this is the correct inverse function by testing $F^{-1}$ for $x=0,0.5,0.9$, and then plotting x against $F^{-1}(F(x))$ for $0\leq x\leq1$ (use the excitatory population parameters).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="GAmeU929fQ_d"
def F_inv(x,a,theta):
"""Define the inverse of the population activation function.
Arguments:
x -- the population input
a -- the gain of the function
theta -- the threshold of the function
Returns:
y -- value of the inverse function
"""
# insert your code here
return y
# insert your code here
# + [markdown] colab_type="text" id="oZ5sSDgAdu87"
# **EXPECTED OUTPUT**
#
# ```
# 0.0
# 2.9120659956266
# 5.002378884081663
# ```
# 
#
#
# + [markdown] colab_type="text" id="WFtmR-6ZbWg-"
# **Exercise 5:** Now, derive the E and I nullclines, in terms of the inverse function $F^{-1}$. To do this, set $\frac{dE}{dt}=0$ and solve for $I$, then set $\frac{dI}{dt}=0$ and solve for $E$. Then, fill out the two functions below to calculate the I nullcline (over $-0.01 \leq I \leq 0.3$) and E nullcline (over $-0.01 \leq E \leq 0.48$). First test the value of the I nullcline for $I=0.1$, then test the E nullcline for $E=0.1$. Then use these functions to plot the nullclines in phase space (E vs. I). What fixed points do you see? Compare the intersections of the nullclines with the steady state values you observed numerically in Exercise 3.
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="RIq3rCcMiQZ2"
def get_E_nullcline(E):
"""Solve for I along the E nullcline (dE/dt = 0).
Arguments:
E -- values of E over which the nullcline is computed
Returns:
I -- values of I along the nullcline for each E
"""
# insert your code here
return I
def get_I_nullcline(I):
"""Solve for E along the I nullcline (dI/dt = 0).
Arguments:
I -- values of I over which the nullcline is computed
Returns:
E -- values of E along the nullcline for each I
"""
# insert your code here
return E
# insert your code here
# + [markdown] colab_type="text" id="op1XBT2nd2-o"
# **EXPECTED OUTPUT**
# ```
# 0.24546433162390224
# -0.029802383619274175
# ```
#
# 
# + [markdown] colab_type="text" id="I5kMzrGKbWhC"
# **Exercise 6:** Now, on top of the nullclines, plot some sample trajectories starting with different initial conditions, for $0 \leq E_0 \leq 1$ and $0 \leq I_0 \leq 1$. How many attractors do you see?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hF-AGDtojEOC"
# insert your code here
# + [markdown] colab_type="text" id="6YuW8b_yd43s"
# **EXPECTED OUTPUT**
#
# 
# + [markdown] colab_type="text" id="kJjtMK8TbWhG"
# **Exercise 7:** Repeat the previous exercise while varying the recurrent excitatory connectivity over the following values: $w_{EE}=5,10,12,15$. What is happening? Can you find a value of wEE where a qualitative transformation occurs? What does this tell you about how increasing recurrent connectivity affects the dynamics?
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="wE63frPAjig6"
# insert your code here
# + [markdown] colab_type="text" id="YjCtkDVXd7OS"
# **EXPECTED OUTPUT**
#
# 
# 
# 
# 
| module1/1_wilson-cowan_equations/1_wilson-cowan_equations_old.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Module 1: Introduction to SageMaker Feature Store
# **Note:** Please set kernel to `Python 3 (Data Science)` and select instance to `ml.t3.medium`
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Setup](#Setup)
# 1. [Load and explore datasets](#Load-and-explore-datasets)
# 1. [Create feature definitions and groups](#Create-feature-definitions-and-groups)
# 1. [Ingest data into feature groups](#Ingest-data-into-feature-groups)
# 1. [Get feature record from the Online feature store](#Get-feature-record-from-the-Online-feature-store)
# 1. [List feature groups](#List-feature-groups)
# # Background
# In this notebook, you will learn how to create **3** feature groups for `customers`, `products` and `orders` datasets
# in the SageMaker Feature Store. You will then learn how to ingest the feature
# columns into the created feature groups (both the Online and the Offline store) using SageMaker Python SDK. You will also see how to get an ingested feature record from the Online store. In the end, you will know how to list all the feature groups created within the Feature Store and delete them.
#
# **Note:** The feature groups created in this notebook will be used in the upcoming modules.
#
# # Setup
# #### Imports
from sagemaker.feature_store.feature_group import FeatureGroup
from time import gmtime, strftime, sleep
from random import randint
import pandas as pd
import numpy as np
import subprocess
import sagemaker
import importlib
import logging
import time
import sys
if sagemaker.__version__ < '2.48.1':
subprocess.check_call([sys.executable, '-m', 'pip', 'install', 'sagemaker==2.48.1'])
importlib.reload(sagemaker)
logger = logging.getLogger('__name__')
logger.setLevel(logging.DEBUG)
logger.addHandler(logging.StreamHandler())
logger.info(f'Using SageMaker version: {sagemaker.__version__}')
logger.info(f'Using Pandas version: {pd.__version__}')
# #### Essentials
sagemaker_session = sagemaker.Session()
role = sagemaker.get_execution_role()
default_bucket = sagemaker_session.default_bucket()
logger.info(f'Default S3 bucket = {default_bucket}')
prefix = 'sagemaker-feature-store'
region = sagemaker_session.boto_region_name
# # Load and explore datasets
customers_df = pd.read_csv('.././data/transformed/customers.csv')
customers_df.head(5)
customers_df.dtypes
customers_df['customer_id'] = customers_df['customer_id'].astype('string')
customers_df['event_time'] = customers_df['event_time'].astype('string')
customers_df.dtypes
products_df = pd.read_csv('.././data/transformed/products.csv')
products_df.head(5)
products_df['product_id'] = products_df['product_id'].astype('string')
products_df['event_time'] = products_df['event_time'].astype('string')
products_df.dtypes
orders_df = pd.read_csv('.././data/transformed/orders.csv')
orders_df
orders_df['order_id'] = orders_df['order_id'].astype('string')
orders_df['customer_id'] = orders_df['customer_id'].astype('string')
orders_df['product_id'] = orders_df['product_id'].astype('string')
orders_df['event_time'] = orders_df['event_time'].astype('string')
orders_df.dtypes
customers_count = customers_df.shape[0]
# %store customers_count
products_count = products_df.shape[0]
# %store products_count
orders_count = orders_df.shape[0]
# %store orders_count
# # Create feature definitions and groups
current_timestamp = strftime('%m-%d-%H-%M', gmtime())
# prefix to track all the feature groups created as part of feature store champions workshop (fscw)
fs_prefix = 'fscw-'
customers_feature_group_name = f'{fs_prefix}customers-{current_timestamp}'
# %store customers_feature_group_name
products_feature_group_name = f'{fs_prefix}products-{current_timestamp}'
# %store products_feature_group_name
orders_feature_group_name = f'{fs_prefix}orders-{current_timestamp}'
# %store orders_feature_group_name
logger.info(f'Customers feature group name = {customers_feature_group_name}')
logger.info(f'Products feature group name = {products_feature_group_name}')
logger.info(f'Orders feature group name = {orders_feature_group_name}')
customers_feature_group = FeatureGroup(name=customers_feature_group_name, sagemaker_session=sagemaker_session)
products_feature_group = FeatureGroup(name=products_feature_group_name, sagemaker_session=sagemaker_session)
orders_feature_group = FeatureGroup(name=orders_feature_group_name, sagemaker_session=sagemaker_session)
customers_feature_group.load_feature_definitions(data_frame=customers_df)
products_feature_group.load_feature_definitions(data_frame=products_df)
orders_feature_group.load_feature_definitions(data_frame=orders_df)
# Let's create the feature groups now
def wait_for_feature_group_creation_complete(feature_group):
status = feature_group.describe().get('FeatureGroupStatus')
print(f'Initial status: {status}')
while status == 'Creating':
logger.info(f'Waiting for feature group: {feature_group.name} to be created ...')
time.sleep(5)
status = feature_group.describe().get('FeatureGroupStatus')
if status != 'Created':
raise SystemExit(f'Failed to create feature group {feature_group.name}: {status}')
logger.info(f'FeatureGroup {feature_group.name} was successfully created.')
customers_feature_group.create(s3_uri=f's3://{default_bucket}/{prefix}',
record_identifier_name='customer_id',
event_time_feature_name='event_time',
role_arn=role,
enable_online_store=True)
wait_for_feature_group_creation_complete(customers_feature_group)
products_feature_group.create(s3_uri=f's3://{default_bucket}/{prefix}',
record_identifier_name='product_id',
event_time_feature_name='event_time',
role_arn=role,
enable_online_store=True)
wait_for_feature_group_creation_complete(products_feature_group)
orders_feature_group.create(s3_uri=f's3://{default_bucket}/{prefix}',
record_identifier_name='order_id',
event_time_feature_name='event_time',
role_arn=role,
enable_online_store=True)
wait_for_feature_group_creation_complete(orders_feature_group)
# # Ingest data into feature groups
# +
# %%time
logger.info(f'Ingesting data into feature group: {customers_feature_group.name} ...')
customers_feature_group.ingest(data_frame=customers_df, max_processes=16, wait=True)
logger.info(f'{len(customers_df)} customer records ingested into feature group: {customers_feature_group.name}')
# +
# %%time
logger.info(f'Ingesting data into feature group: {products_feature_group.name} ...')
products_feature_group.ingest(data_frame=products_df, max_processes=16, wait=True)
logger.info(f'{len(products_df)} product records ingested into feature group: {products_feature_group.name}')
# +
# %%time
logger.info(f'Ingesting data into feature group: {orders_feature_group.name} ...')
orders_feature_group.ingest(data_frame=orders_df, max_processes=16, wait=True)
logger.info(f'{len(orders_df)} order records ingested into feature group: {orders_feature_group.name}')
# -
# # Get feature record from the Online feature store
featurestore_runtime_client = sagemaker_session.boto_session.client('sagemaker-featurestore-runtime', region_name=region)
# Retrieve a record from customers feature group
customer_id = f'C{randint(1, 10000)}'
logger.info(f'customer_id={customer_id}')
feature_record = featurestore_runtime_client.get_record(FeatureGroupName=customers_feature_group_name,
RecordIdentifierValueAsString=customer_id)
feature_record
# # List feature groups
sagemaker_client = sagemaker_session.boto_session.client('sagemaker', region_name=region)
response = sagemaker_client.list_feature_groups()
for fg in response['FeatureGroupSummaries']:
fg_name = fg['FeatureGroupName']
print(f'Found feature group: {fg_name}')
| 01-module-feature-store-foundations/m1_nb1_introduction_to_feature_store.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import pandas as pd
import json
cdf = pd.read_csv('census_data.csv',index_col='state', encoding='utf-8') #census data
hdf = pd.read_csv('health.csv', encoding='utf-8') #health data from the cdc
#print(df.corr()['depression'])
# -
cdf.info()
hdf.info()
# +
correlation_results = []
results_dict = {}
for i in hdf.Question.unique():
#break down into specific questions
q_df = hdf[hdf['Question'] == i]
for a in q_df.Response.unique():
#break down into specific responses for each question
r_df = q_df[q_df['Response'] == a].set_index('Locationdesc')
# rename data_values to the question and response
question = str(r_df['Question'].unique()[0])
response = str(a)
name = question + "__" + response
#array of values connected to the above question and response
cor_query = r_df["Data_value"].rename(name)
#print(cor_query)
# append the data_value of this particular questions and response by state to the census data
try:
#print("Mining correlations for....")
#print(name)
correlations = pd.concat([cdf, cor_query], axis=1).corr()[name]
#print("correlations \n")
indexer = 0
for index, value in correlations.iteritems():
if value > .65 and value < 1:
print(correlations.name)
print(value)
print(index)
results_dict[correlations.name] = {
'correlation':value,
"values":[cor_query.values],
index:[cdf[index].values],
'states': cdf.index.values,
}
correlation_results.append(
{correlations.name:[{
'correlation':value,
"values":[cor_query.values],
index:[cdf[index].values],
'states': cdf.index.values,
}
]})
indexer += 1
print("---" * 20)
elif value < -.65:
print(correlations.name)
print(value)
print(index)
print("---" * 20)
results_dict[correlations.name] = {
'correlation':value,
"values":[cor_query.values],
index:[cdf[index].values],
'states': cdf.index.values,
}
correlation_results.append(
{correlations.name:[{
'correlation':value,
correlations.name + "_values":[cor_query.values],
index:[cdf[index].values],
'states': cdf.index.values,
}
]})
except Exception as e:
print(e)
print(f"cannot process {name}")
# -
#
results_dict["Was there a time in the past 12 months when you needed to see a doctor but could not because of cost?__Yes"]
# +
# columns from the Census data we want to pull for graphing
blindness = pd.Series(results_dict["Are you blind or do you have serious difficulty seeing, even when wearing glasses?__Yes"]['values'][0]).rename('blindness')
depression = pd.Series(cdf['depression'].values).rename('percentDepressed')
arthritis = pd.Series(results_dict["Adults who have been told they have arthritis (variable calculated from one or more BRFSS questions)__Yes"]['values'][0]).rename('Arthritis')
poverty_line = pd.Series(cdf['Total Civilian Noninstitutionalized Population; Estimate; 100 to 149 percent of the poverty level'].values).rename("povertyPop")
state_abbr = pd.Series(cdf["locationAbbr"].values).rename('stateAbbr')
states = pd.Series(results_dict["Are you blind or do you have serious difficulty seeing, even when wearing glasses?__Yes"]['states']).rename('states')
medianIncome = pd.Series(cdf['medianIncomeAll'].values).rename('medianIncome')
doc_too_expensive = pd.Series(results_dict["Was there a time in the past 12 months when you needed to see a doctor but could not because of cost?__Yes"]['values'][0]).rename('healthcare_unaffordable')
long_term_unemployment = pd.Series(results_dict["What is your employment status?__No work >yr"]["values"][0]).rename('unemployment')
# -
df = pd.concat(
[
state_abbr,
states,
poverty_line,
blindness,
depression,
arthritis,
medianIncome,
doc_too_expensive,
long_term_unemployment],
axis=1).set_index('stateAbbr').dropna()
df
df.to_csv('data.csv', encoding='utf-8')
| data/correlation_extractor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Dmitri9149/Transformer_From_Scratch/blob/main/Final_Transformer_MXNet_12800_11_10_20.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="se3XU8H6kLYd" outputId="55883163-c99b-4f56-e43f-713a3e07e890" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !pip install -U mxnet-cu101==1.7.0
# !pip install d2l==0.14.4
# + id="nIZmHf96M8DY"
# ### !pip install ipython-autotime
# ### %load_ext autotime
# + id="VlxeCFu4pTB6"
import math
from d2l import mxnet as d2l
from mxnet import np, npx
from mxnet.gluon import nn
from mxnet import np, npx, init, gluon, autograd
import collections
import os
import time
npx.set_np()
from mxnet import autograd, np, npx
# + [markdown] id="I9SiUMN4xLyq"
# The code for Transformer from scratch is collected here. The code is mostly from http://d2l.ai/chapter_attention-mechanisms/transformer.html . I did many comments to the code at most difficult points. I hope my additional code and comments will help in better understanding of the Transformer.
# This is the original article for the Transformer :
# <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … <NAME>. (2017). Attention is all you need. Advances in neural information processing systems (pp. 5998–6008).
#
# The future work:
# 1. To learn the Transformer on big data set.
# 2. Transation from (to) English to Finnish language.
# 3. Modify the architecture of the Transformer.
# 4. Better tokenization and preprocessing.
#
#
#
# + [markdown] id="VsTaZurJGlAN"
# ### Attention Mechanism
#
# + [markdown] id="5xCH_Y6CG3RN"
# #### Masked softmax
#
# This is importand auxiliary function.
# """ The masked softmax takes a 3-dimensional input and enables us to filter out some elements by specifying a valid length for the last dimension.... As a result, any value outside the valid length will be masked as 0.""" (citation from d2l.ai).
# The notion of valid length come from the need to add special <pad> token if our sentence is shorter than length we use for all sentencies in batches. The <pad> tokens will not participate in prediction.
#
# My comments are started with ### ,
# the comments with one # are from the original d2l.ai code.
#
# Some functions for plotting and downloading of specific files from specific places are still taken from the d2l.ai library on GitHub : https://github.com/d2l-ai/d2l-en/blob/master/d2l/mxnet.py But the biggest par of the code is collected here (and commented).
# + id="q5eCmYbQkVZ3"
### from d2l.ai
def masked_softmax(X, valid_len):
"""Perform softmax by filtering out some elements."""
# X: 3-D tensor, valid_len: 1-D or 2-D tensor
### why 3-D tensor ?
### first dimention; we will quantify samples within batch,
### so, the first dimention determines the number of samples in the batch
###
### second dimention; we will quantify queries,
### we may have several queries,
### the second dimention determines the number of queries
###
### we may set up the valid lengths same for every sample in the
### batch, i.e 1-D valid-lengh with size (batch_size, )
### the same means : independent of the queries
### On the contarry: we may set up valid lengths individually for every
### sample in a batch and for every query,
### in this case it will be 2-D valid length
### with size (batch size, number of queries)
###
### Third parameter will correspond to the number of key/value pairs
###
### We may need the valid_length when: 1. we <pad> the end of a sentence: it is too
### short, shorter than num_steps ; 2. when we use the valid_lenght in decoder
### via training, and every word in target sentence is used as query: the query
### can (or may ?) see the all words to the left, but not to the right (see the
### encoder decoder code below). To handle the case we use valid_length too.
###
if valid_len is None:
return npx.softmax(X)
else:
shape = X.shape
if valid_len.ndim == 1:
valid_len = valid_len.repeat(shape[1], axis=0)
else:
valid_len = valid_len.reshape(-1)
# Fill masked elements with a large negative, whose exp is 0
X = npx.sequence_mask(X.reshape(-1, shape[-1]), valid_len, True,
axis=1, value=-1e6)
return npx.softmax(X).reshape(shape)
# + id="846OVWClkhCa" outputId="2fe99c5d-8f46-4739-8a83-f7984dabb042" colab={"base_uri": "https://localhost:8080/", "height": 116}
### from d2l.ai
masked_softmax(np.random.uniform(size=(2, 2, 4)), np.array([2, 3]))
### 2 - number of samples in the batch
### 2 - we make deal with 2 queries
### 4 - four key/value pairs
### for the first sample in our batch , from 4 pairs we will take
### into account only results from first 2 pairs, the rest will be multiplied by 0,
### because that pairs correspond to <pad> tokens
### for the second sample (4 key/value pairs) we will take into account
### only results for first 3 key/value pairs (the rest will masked with 0,
### because the rest pairs correspond to <pad> tokens)
### this is the meaning of np.array([2,3]) as valid length
### the velid length is not dependent from queries in this case
# + id="Ty6KkQs0kiYF" outputId="b78f1b02-19c1-4343-d57f-0d394789dcbd" colab={"base_uri": "https://localhost:8080/", "height": 77}
### from d2l.ai
npx.batch_dot(np.ones((2, 1, 3)), np.ones((2, 3, 2)))
# + id="WWRQYp1sPO_F" outputId="b1aede4f-79cb-4f6a-8b68-e7c20141bf02" colab={"base_uri": "https://localhost:8080/", "height": 176}
### one more example with 1-D valid length
valid_length = np.array([2,3])
### the shape is (2,) : one dimentional length
print('valid_length shape= ', valid_length.shape)
masked_softmax(np.random.uniform (size =(2, 3, 5)), valid_length )
# + id="n_Vwfo60JtQa"
# + id="277f-M9HPTAl" outputId="4f425407-d8cc-4c4d-8668-6860fcd048ea" colab={"base_uri": "https://localhost:8080/", "height": 334}
### if we declare 2-D valid_length
valid_length = np.array([[3, 5, 4], [2,4, 1], [1,4, 3],[1,2,3]])
print('valid_length shape= ', valid_length.shape)
masked_softmax(np.random.uniform(size = (4, 3, 5)), valid_length)
### Let us consider the first sample in our batch
### [[0.21225105, 0.31475353, 0.4729953 , 0. , 0. ,
### 0. ],
### [0.19417836, 0.20596693, 0.16711308, 0.15453914, 0.27820238,
### 0. ],
### [0.2753876 , 0.21671425, 0.30811197, 0.19978616, 0. ,
### 0. ]],
### from third dimention in np.random.uniform(size = (4, 3, 5)) we may see it correspond to
### 5 key/value pairs (that is why the length of the lines is 5)
### second dimention in np.random.uniform(size = (4, 3, 5)) means the results are obtained from
### 3 queries, that is why there are 3 lines corresponding to every batch
###
### Below we may see there are 4 groups, because the first dimention, the
### number of samples, is 4 (batch size)
###
### np.array([[3, 5, 4], [2,4, 1], [1,4, 3],[1,2,3]])
### is 2-D array (of size 4 * 3 in our case))
### 4 is batch size, 3 is number of queries : we have 4 groups with 3 lines in each;
### the [3,5,4] subarray correspond to the first sample in the batch,
### in the first group : the first line has first 3 non zero elements,
### the second line 5 first non zero and third line 4 first non zero elements.
# + [markdown] id="qxiF2bdujQHW"
# ### Dot product attention
#
# #### Why we need it, how it is calculated
#
#
# + [markdown] id="GabuXpxG54vE"
# We have query with dimension `d`.
# We have #kv_pairs: key/value pairs. Every key and value are vectors of dimension `d`. We pass the query trought the 'grid' with the leng of #kv_pairs and get #kv_pairs of scores. How it works within the pass: we make dot product of query with every of #kv_pairs keys in the 'grid' and get a #kv_pairs scores. We also normilize the scores by dividing on $\sqrt{d}$.
# If we have batch with size batch_size and number of queries = #queries, we will get tensor of scores of size (batch_size, #queries, #kv_pairs).
# In this way we receive the attention_weights tensor.
# We also have tensor 'value' of values of size (batch_size, #kv_pairs, dim_v).
# Finally, using npx.batch_dot(attention_weights, value) we will get tensor of size (batch_size, #queries, dim_v) which corresponf of the 'passing' our queries throught the 'grid' of key/value pairs: for every query, for every sample in the batch we will get the transformed vector of size dim_v.
#
# + id="bCGtMEgIkmRK"
### from d2l.ai book
class DotProductAttention(nn.Block):
def __init__(self, dropout, **kwargs):
super(DotProductAttention, self).__init__(**kwargs)
self.dropout = nn.Dropout(dropout)
# `query`: (`batch_size`, #queries, `d`)
# `key`: (`batch_size`, #kv_pairs, `d`)
# `value`: (`batch_size`, #kv_pairs, `dim_v`)
# `valid_len`: either (`batch_size`, ) or (`batch_size`, xx)
def forward(self, query, key, value, valid_len=None):
d = query.shape[-1]
# Set transpose_b=True to swap the last two dimensions of key
scores = npx.batch_dot(query, key, transpose_b=True) / math.sqrt(d)
attention_weights = self.dropout(masked_softmax(scores, valid_len))
return npx.batch_dot(attention_weights, value)
# + id="3JF5Sxa6kqDq"
if False:
### the code from d2l.ai
atten = DotProductAttention(dropout=0.5)
atten.initialize()
### batch size of 2, #kv_pairs = 10, every key is vector of size 2 with
### ones : (1.,1.)
keys = np.ones((2, 10, 2))
### we start with vector which keep float numbers from 0 to 39;
### reshape it to tensor which model one sample batch with 10 key/value pairs and
### dimension of values dim_v = 4; finally we repeat the construction to get 2
### similar samples (batch with 2 samples).
values = np.arange(40).reshape(1, 10, 4).repeat(2, axis=0)
atten(np.ones((2, 1, 2)), keys, values, np.array([2, 6]))
# + id="WklkVqgAq8lC"
if False:
atten = DotProductAttention(dropout=0.5)
atten.initialize()
keys = np.ones((3,10,5)) # keys in batch of size 3; for every line in batch we have
### 10 keys/values pairs ; where every key is 5 dimentional vector (and value will be 7 dimentional vector);
### each key is forming pair with value, there are 10 such pairs
values = np.arange(70).reshape(1,10,7).repeat(3, axis =0) # values in batch of
### size 3 ; 10 values with 7 dimentional vector each;
### in our batch the 3 samples are identical by construction
queries = np.ones((3,4,5)) # quiries in batch of size 3, there are 4 queries,
### where every query is vector of size 5 (same size as for key)
atten(queries, keys, values, np.array([3, 8, 6])) # values in batch of size 3 ;
### 4 quiry per every sample in batch where every query is vector of size 5
### the valid_len is 1-D
### for the 3 samples the valid_length have size 3 , 8 , 6 ;
### size 3 for first sample , ....., ..... size 6 for the last sample
### the outputs are:
### for every entry in the batch (for every of the 3 samples)
### for every of 4 queries
### total : 3*4 = 12 final values: vectors of size 7
### the values are different for different samples in the batch ,
### because we used different valid length,
### but for every sample group in the batch (same sample, different queries),
### all 4 final values are the same:
### even we use 4 queries, all the quiries are equal in our case
# + [markdown] id="1qHhjcNvy3xJ"
# ### Multihead Attention
#
# """ The *multi-head attention* layer consists of $h$ parallel self-attention layers, each one is called a *head*. For each head, before feeding into the attention layer, we project the queries, keys, and values with three dense layers with hidden sizes $p_q$, $p_k$, and $p_v$, respectively. The outputs of these $h$ attention heads are concatenated and then processed by a final dense layer.
#
#
# 
#
#
# Assume that the dimension for a query, a key, and a value are $d_q$, $d_k$, and $d_v$, respectively. Then, for each head $i=1,\ldots, h$, we can train learnable parameters
# $\mathbf W_q^{(i)}\in\mathbb R^{p_q\times d_q}$,
# $\mathbf W_k^{(i)}\in\mathbb R^{p_k\times d_k}$,
# and $\mathbf W_v^{(i)}\in\mathbb R^{p_v\times d_v}$. Therefore, the output for each head is
#
# $$\mathbf o^{(i)} = \mathrm{attention}(\mathbf W_q^{(i)}\mathbf q, \mathbf W_k^{(i)}\mathbf k,\mathbf W_v^{(i)}\mathbf v),$$
#
# where $\textrm{attention}$ can be any attention layer, such as the `DotProductAttention` and `MLPAttention` as we introduced in :numref:`sec_attention`.
#
#
#
# After that, the output with length $p_v$ from each of the $h$ attention heads are concatenated to be an output of length $h p_v$, which is then passed the final dense layer with $d_o$ hidden units. The weights of this dense layer can be denoted by $\mathbf W_o\in\mathbb R^{d_o\times h p_v}$. As a result, the multi-head attention output will be
#
# $$\mathbf o = \mathbf W_o \begin{bmatrix}\mathbf o^{(1)}\\\vdots\\\mathbf o^{(h)}\end{bmatrix}.$$
#
#
# Now we can implement the multi-head attention. Assume that the multi-head attention contain the number heads `num_heads` $=h$, the hidden size `num_hiddens` $=p_q=p_k=p_v$ are the same for the query, key, and value dense layers. In addition, since the multi-head attention keeps the same dimensionality between its input and its output, we have the output feature size $d_o =$ `num_hiddens` as well. """ (citation from d2l.ai book).
#
# + [markdown] id="J618Qc3qMu1j"
# There are some problems in the d2l.ai text, there is stated :
# $p_q$ = $p_k$ = $p_v$ = num_hiddens,
# and
# $d_o =$ `num_hiddens` as well.
# So, we have $W_o$ transformation from input of size (num_heads * num_hiddens) to output of size (num_hiddens). If h > 1, the input size and output size can not be equal. But in the PyTorch code in the d2l.ai we have:
# self.W_o = nn.Linear(num_hiddens, num_hiddens, bias=bias)
# with equal input and output. It is hidden in the d2l.ai
# MXNet code: self.W_o = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False), because in the
# case of Gluon Dense layer we state only output dimension (num_hiddens in the case). The input dimension is not stated.
#
# There is also assumed in the code below (from d2l.ai book), the num_hiddens is multiple of num_heads. No assumptions about it in the main text of the book. But in the d2l.ai code the assumption is used.
#
# The ony interpretation to the code below I may give now:
# $p_v$ * num_heads=num_hiddens (same for $p_q$ = $p_k$ = $p_v$),
# but not $p_v$=num_hiddens.
# I will interpret the code with the assumption.
#
#
# + id="2WvAyhjFk4qS"
### from d2l.ai
class MultiHeadAttention(nn.Block):
def __init__(self, num_hiddens, num_heads, dropout, use_bias=False, **kwargs):
super(MultiHeadAttention, self).__init__(**kwargs)
self.num_heads = num_heads
self.attention = d2l.DotProductAttention(dropout)
### here, as I understand, the num_hiddens = num_heads * p_v
### where p_v (see the text above) is the dimension of the vector
### to which a query is transformed by single head,
### the size of p_v is to be (num_hidden/num_heads)
### it explains what the code below do
self.W_q = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
self.W_k = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
self.W_v = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
### if every head transform query of size `dim` = num_hiddens to
### p_v = p_q = p_k = (num_hidden/num_heads), when we
### concatenate num_heads of such queries, we will get
### vector of size num_hidden again;
### it explains the input / output dimensions for W_o :
### input and output have same dimension = num_hiddens
self.W_o = nn.Dense(num_hiddens, use_bias=use_bias, flatten=False)
### every query generate num_heads outputs , which we cancatenate in
### one vector of dimention num_hiddens : so the output of every query is
### of size num_heads / num_hiddens;
### to apply self-attention we de-cancatenate the combined output
### to hum_heads of separate outputs from every query
### with size (num_hiddens / num_heads), and
### simultaneously recombine them in single batch (with size num_heads),
### which increase the total batch size to (batch_size * num_heads)
### We have to correct the valid_length to take into account
### the num_heads query transformtions are combined now in single batch.
### After application of self_attention, we make the reverse operation:
### locate the batch samples which correspond to the outputs of the same query
### in different heads, and concatenate them again in one combined output.
### The number of batches decrease and the length of output increase by the
### same factor num_heads.
### These are the roles of transpose_qkv , transpose_output functions below:
def forward(self, query, key, value, valid_len):
# For self-attention, `query`, `key`, and `value` shape:
# (`batch_size`, `seq_len`, `dim`), where `seq_len` is the length of
# input sequence. `valid_len` shape is either (`batch_size`, ) or
# (`batch_size`, `seq_len`).
# Project and transpose `query`, `key`, and `value` from
# (`batch_size`, `seq_len`, `num_hiddens`) to
# (`batch_size` * `num_heads`, `seq_len`, `num_hiddens` / `num_heads`)
query = transpose_qkv(self.W_q(query), self.num_heads)
key = transpose_qkv(self.W_k(key), self.num_heads)
value = transpose_qkv(self.W_v(value), self.num_heads)
if valid_len is not None:
# Copy `valid_len` by `num_heads` times
if valid_len.ndim == 1:
valid_len = np.tile(valid_len, self.num_heads)
else:
valid_len = np.tile(valid_len, (self.num_heads, 1))
# For self-attention, `output` shape:
# (`batch_size` * `num_heads`, `seq_len`, `num_hiddens` / `num_heads`)
output = self.attention(query, key, value, valid_len)
# `output_concat` shape: (`batch_size`, `seq_len`, `num_hiddens`)
output_concat = transpose_output(output, self.num_heads)
return self.W_o(output_concat)
# + id="7yIscKTuk6nL"
### from d2l.ai
def transpose_qkv(X, num_heads):
# Input `X` shape: (`batch_size`, `seq_len`, `num_hiddens`).
# Output `X` shape:
# (`batch_size`, `seq_len`, `num_heads`, `num_hiddens` / `num_heads`)
X = X.reshape(X.shape[0], X.shape[1], num_heads, -1)
# `X` shape:
# (`batch_size`, `num_heads`, `seq_len`, `num_hiddens` / `num_heads`)
X = X.transpose(0, 2, 1, 3)
# `output` shape:
# (`batch_size` * `num_heads`, `seq_len`, `num_hiddens` / `num_heads`)
output = X.reshape(-1, X.shape[2], X.shape[3])
return output
### from d2l.ai
def transpose_output(X, num_heads):
# A reversed version of `transpose_qkv`
X = X.reshape(-1, num_heads, X.shape[1], X.shape[2])
X = X.transpose(0, 2, 1, 3)
return X.reshape(X.shape[0], X.shape[1], -1)
# + id="n5rGL8xek-US"
if False:
### from d2l.ai
### num_hiddens = 100, num_heads=10
cell = MultiHeadAttention(100, 10, 0.5)
cell.initialize()
X = np.ones((2, 4, 5))
valid_len = np.array([2, 3])
cell(X, X, X, valid_len).shape
# + id="R83Fn2Sn-CWL"
if False:
### it correspond to scenario size of embedding is 512 ; num_heads = 8 ;
### num_hiddens = 512
cell = MultiHeadAttention(512, 8, 0.5)
cell.initialize()
# num of batches is 3 ; seq_len is 20 ; size of embedding is 512
X = np.ones((3, 20, 512))
valid_len = np.array([15,17,12])
cell(X, X, X, valid_len).shape
# + [markdown] id="GTS9pWEQ_MZw"
# ### Position-wise encoding
#
# Two 1 * 1 convolutional layers are applied. Extract
# position independent features of word representations (in the same way the convolution layers are applied in image recognition networks).
# """ Similar to the multi-head attention, the position-wise feed-forward network will only change the last dimension size of the input—the feature dimension. In addition, if two items in the input sequence are identical, the according outputs will be identical as well. """ (citation from d2l.ai)
# + id="zJFb4Mq3lD6Y"
### from d2l.ai
class PositionWiseFFN(nn.Block):
def __init__(self, ffn_num_hiddens, pw_num_outputs, **kwargs):
super(PositionWiseFFN, self).__init__(**kwargs)
self.dense1 = nn.Dense(ffn_num_hiddens, flatten=False,
activation='relu')
self.dense2 = nn.Dense(pw_num_outputs, flatten=False)
def forward(self, X):
return self.dense2(self.dense1(X))
# + id="lEFrwk6klIq8"
if False:
ffn = PositionWiseFFN(4, 8)
ffn.initialize()
ffn(np.ones((2, 3, 4)))[0]
# + [markdown] id="xJz6_7utLNvG"
# ### Add and Norm
# """ we add a layer that contains a residual structure and a layer normalization after both the multi-head attention layer and the position-wise FFN network. Layer normalization is similar to batch normalization ........ One difference is that the mean and variances for the layer normalization are calculated along the last dimension, e.g X.mean(axis=-1) instead of the first batch dimension, e.g., X.mean(axis=0). Layer normalization prevents the range of values in the layers from changing too much, which allows faster training and better generalization ability. """ (citation from d2l.ai)
# + id="q31gjuTWLyUg"
if False:
### from d2l.ai
layer = nn.LayerNorm()
layer.initialize()
batch = nn.BatchNorm()
batch.initialize()
X = np.array([[1, 2], [2, 3]])
# Compute mean and variance from `X` in the training mode
with autograd.record():
print('layer norm:', layer(X), '\nbatch norm:', batch(X))
# + [markdown] id="WaYlKHljMB6l"
#
# """AddNorm accepts two inputs X and Y. We can deem X as the original input in the residual network, and Y as the outputs from either the multi-head attention layer or the position-wise FFN network. In addition, we apply dropout on Y for regularization.""" citation from d2l.ai
# + id="KTwTY_wtlNP6"
### from d2l.ai
class AddNorm(nn.Block):
def __init__(self, dropout, **kwargs):
super(AddNorm, self).__init__(**kwargs)
self.dropout = nn.Dropout(dropout)
self.ln = nn.LayerNorm()
def forward(self, X, Y):
return self.ln(self.dropout(Y) + X)
# + id="w61BnqdalQbB"
if False:
### d2l.ai
add_norm = AddNorm(0.5)
add_norm.initialize()
add_norm(np.ones((2, 3, 4)), np.ones((2, 3, 4))).shape
# + [markdown] id="iKDZY55-A59e"
# ### Positional Encoding
# + id="hKa5KWQglWH_"
### I used the code as alternative to the original positional encoding;
### just encode position of words (tokens) in sentence ,
### it changes the results , but the results are quite well.
if False:
### from d2l.ai
class PositionalEncoding(nn.Block):
def __init__(self, num_hiddens, dropout, max_len=100):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(dropout)
# Create a long enough `P`
### max_len correspond to sequence length ;
### num_hiddens correspond to embedding size
###
self.P = np.zeros((1, max_len, num_hiddens))
### X = np.arange(0, max_len).reshape(-1, 1) / np.power(
### 10000, np.arange(0, num_hiddens, 2) / num_hiddens)
### self.P[:, :, 0::2] = np.sin(X)
### self.P[:, :, 1::2] = np.cos(X)
###################### my code be carefull !!!!!
X = np.arange(0, max_len).reshape(-1, 1) / max_len
### 10000, np.arange(0, num_hiddens, 2) / num_hiddens)
self.P[:, :, 0::1] = np.sin(X)
### self.P[:, :, 1::2] = np.cos(X)
################################
def forward(self, X):
X = X + self.P[:, :X.shape[1], :].as_in_ctx(X.ctx)
return self.dropout(X)
# + id="SvVLBV6jMqyz"
### from d2l.ai
class PositionalEncoding(nn.Block):
def __init__(self, num_hiddens, dropout, max_len=1000):
super(PositionalEncoding, self).__init__()
self.dropout = nn.Dropout(dropout)
# Create a long enough `P`
### max_len correspond to sequence length ;
### num_hiddens correspond to embedding size
self.P = np.zeros((1, max_len, num_hiddens))
X = np.arange(0, max_len).reshape(-1, 1) / np.power(
10000, np.arange(0, num_hiddens, 2) / num_hiddens)
self.P[:, :, 0::2] = np.sin(X)
self.P[:, :, 1::2] = np.cos(X)
def forward(self, X):
X = X + self.P[:, :X.shape[1], :].as_in_ctx(X.ctx)
return self.dropout(X)
# + id="I_2Ew2GWlZUh"
if False:
### from d2l.ai
### num_hiddens = 20 , dropout = 0
pe = PositionalEncoding(20, 0)
pe.initialize()
### we assume batch_size = 1; max_length = 100 correspond to tokens (here words) in our line;
### num_hiddens = 20 (embedding size)
###
Y = pe(np.zeros((1, 100, 20)))
### dim correspond to coordinate in embedding vector of out tokens (words)
d2l.plot(np.arange(100), Y[0, :, 4:8].T, figsize=(6, 2.5),
legend=["dim %d" % p for p in [4, 5, 6, 7]])
# + [markdown] id="k2GyVY8oey9I"
# ### Encoder
#
# """Armed with all the essential components of Transformer, let us first build a Transformer encoder block. This encoder contains a multi-head attention layer, a position-wise feed-forward network, and two “add and norm” connection blocks. As shown in the code, for both of the attention model and the positional FFN model in the EncoderBlock, their outputs’ dimension are equal to the num_hiddens. This is due to the nature of the residual block, as we need to add these outputs back to the original value during “add and norm”. """ (citation from d2l.ai)
# + id="rm2rOdPXlckg"
### from d2l.ai
### this block will not change the input shape
class EncoderBlock(nn.Block):
def __init__(self, num_hiddens, ffn_num_hiddens, num_heads, dropout,
use_bias=False, **kwargs):
super(EncoderBlock, self).__init__(**kwargs)
self.attention = MultiHeadAttention(num_hiddens, num_heads, dropout,
use_bias)
self.addnorm1 = AddNorm(dropout)
self.ffn = PositionWiseFFN(ffn_num_hiddens, num_hiddens)
self.addnorm2 = AddNorm(dropout)
def forward(self, X, valid_len):
### we sum the original input to the attention block and the output from the
### block + we normilize the result using AddNorm
Y = self.addnorm1(X, self.attention(X, X, X, valid_len))
return self.addnorm2(Y, self.ffn(Y))
# + [markdown] id="CAAQoTC30nzJ"
# """ Now it comes to the implementation of the entire Transformer encoder. With the Transformer encoder, $n$ blocks of `EncoderBlock` stack up one after another. Because of the residual connection, the embedding layer size $d$ is same as the Transformer block output size. Also note that we multiply the embedding output by $\sqrt{d}$ to prevent its values from being too small. """ (citation from d2l.ai)
# + id="izJGhrKRauzo"
### from d2l.ai
class Encoder(nn.Block):
"""The base encoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Encoder, self).__init__(**kwargs)
def forward(self, X, *args):
raise NotImplementedError
# + id="c2wamZIhllpO"
### from d2l.ai
class TransformerEncoder(Encoder):
def __init__(self, vocab_size, num_hiddens, ffn_num_hiddens,
num_heads, num_layers, dropout, use_bias=False, **kwargs):
super(TransformerEncoder, self).__init__(**kwargs)
self.num_hiddens = num_hiddens
self.embedding = nn.Embedding(vocab_size, num_hiddens)
self.pos_encoding = PositionalEncoding(num_hiddens, dropout)
self.blks = nn.Sequential()
for _ in range(num_layers):
self.blks.add(
EncoderBlock(num_hiddens, ffn_num_hiddens, num_heads, dropout,
use_bias))
def forward(self, X, valid_len, *args):
X = self.pos_encoding(self.embedding(X) * math.sqrt(self.num_hiddens))
for blk in self.blks:
X = blk(X, valid_len)
return X
# + [markdown] id="tyZgVJaW5IpA"
# ### Decoder
#
# """ During training, the output for the $t$-query could observe all the previous key-value pairs. It results in an different behavior from prediction. Thus, during prediction we can eliminate the unnecessary information by specifying the valid length to be $t$ for the $t^\textrm{th}$ query. """
# (citation from d2l.ai)
# + id="S_YlpVF7lsMR"
### from d2l.ai
class DecoderBlock(nn.Block):
# `i` means it is the i-th block in the decoder
### the i will be initialized from the TransformerDecoder block
### the block will be used in TransformerDecoder in stack ,
### several blocks will be aranged in sequence, output from
### one block will be input to the next blosk
def __init__(self, num_hiddens, ffn_num_hiddens, num_heads,
dropout, i, **kwargs):
super(DecoderBlock, self).__init__(**kwargs)
self.i = i
### in the block we will aplly (MultiHeadAttention + AddNorm)
### and again (MultiHeadAttention + AddNorm) ;
### then we will apply PositionWiseFFN
self.attention1 = MultiHeadAttention(num_hiddens, num_heads, dropout)
self.addnorm1 = AddNorm(dropout)
self.attention2 = MultiHeadAttention(num_hiddens, num_heads, dropout)
self.addnorm2 = AddNorm(dropout)
self.ffn = PositionWiseFFN(ffn_num_hiddens, num_hiddens)
self.addnorm3 = AddNorm(dropout)
def forward(self, X, state):
### we use state [0] and state[1] to keep output from TransformerEncoder :
### enc_outputs and enc_valid_length;
### which correspond to sentences we are translating (sentences in language FROM
### which we translate);
### the state [0] and state [1] are received from TransformerDecoder
### enclosing block as shared parameter;
enc_outputs, enc_valid_len = state[0], state[1]
# `state[2][i]` contains the past queries for this block
### in the first block (i = 1) , at this place in code,
### the queries = None, see the code in TransformetEncoder :
###
### def init_state(self, enc_outputs, enc_valid_len, *args):
### return [enc_outputs, enc_valid_len, [None]*self.num_layers]
###
### TransformerEncoder is initialized from EncoderDecoder
### using the 'init_state' function (see above) , as
### we can see, the fird element in array is None for self.layers;
### the 'init_state' determines the 'state' in TransformerEncoder,
### in the code above we use state[0] and state[1] to determine
### 'enc_outputs', 'enc_valid_len' in this block
if state[2][self.i] is None:
key_values = X
else:
### queries from previous block are added to the current queries and
### used together as key value pairs
key_values = np.concatenate((state[2][self.i], X), axis=1)
state[2][self.i] = key_values
if autograd.is_training():
### here are are in training mode
### below in 'attention' function we will use X as queries,
### X correspond to all words in the target sentence within training;
### seq_len correspond to the length of the whole target sentence;
### we will use seq_len queries, for every sample in the batch;
### for us important the following:
### first query from the sentence has to be constrained
### to first key_value pair; second: to the first two key_value pairs,
### etc...
### that is why the valid_len is generated in the way:
batch_size, seq_len, _ = X.shape
# Shape: (batch_size, seq_len), the values in the j-th column
# are j+1
### while training we take into account the result of passing a query
### in target sentence throught the 'grid' of key/value pairs to the
### left of the query ;
### every query in the target sequence has its own valid_len and
### the valid_len correspond to the position of a query in the
### sentence
valid_len = np.tile(np.arange(1, seq_len + 1, ctx=X.ctx),
(batch_size, 1))
else:
valid_len = None
### the attention mechanism is used on key_values corresponding
### to the target sentence key_values (then AddNorm is applied)
X2 = self.attention1(X, key_values, key_values, valid_len)
Y = self.addnorm1(X, X2)
### the attention mechanism is used on TransformerEncoder outputs
### key_values as the 'grid' (then AddNorm is applied);
### the key/values are the learned pairs
### which are originated from the source sentence
Y2 = self.attention2(Y, enc_outputs, enc_outputs, enc_valid_len)
Z = self.addnorm2(Y, Y2)
return self.addnorm3(Z, self.ffn(Z)), state
# + id="iDNchRW5aPfd"
### from d2l.ai
class Decoder(nn.Block):
"""The base decoder interface for the encoder-decoder architecture."""
def __init__(self, **kwargs):
super(Decoder, self).__init__(**kwargs)
def init_state(self, enc_outputs, *args):
raise NotImplementedError
def forward(self, X, state):
raise NotImplementedError
# + id="MOF871y_lxOS"
### from d2l.ai
class TransformerDecoder(Decoder):
def __init__(self, vocab_size, num_hiddens, ffn_num_hiddens,
num_heads, num_layers, dropout, **kwargs):
super(TransformerDecoder, self).__init__(**kwargs)
self.num_hiddens = num_hiddens
self.num_layers = num_layers
self.embedding = nn.Embedding(vocab_size, num_hiddens)
self.pos_encoding = PositionalEncoding(num_hiddens, dropout)
### sequential application of several DecoderBlock's
self.blks = nn.Sequential()
for i in range(num_layers):
self.blks.add(
DecoderBlock(num_hiddens, ffn_num_hiddens, num_heads,
dropout, i))
self.dense = nn.Dense(vocab_size, flatten=False)
def init_state(self, enc_outputs, env_valid_len, *args):
return [enc_outputs, env_valid_len, [None]*self.num_layers]
def forward(self, X, state):
X = self.pos_encoding(self.embedding(X) * math.sqrt(self.num_hiddens))
for blk in self.blks:
X, state = blk(X, state)
return self.dense(X), state
# + id="RwO0LYrSS6i1"
### from d2l.ai
### this block couples together TransformerEncoder and TransformerDecoder
###
class EncoderDecoder(nn.Block):
"""The base class for the encoder-decoder architecture."""
def __init__(self, encoder, decoder, **kwargs):
super(EncoderDecoder, self).__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
def forward(self, enc_X, dec_X, *args):
### the enc_outputs are moved to decoder from encoder;
### the coupling happens in this point of code
enc_outputs = self.encoder(enc_X, *args)
### initial decoder state: dec_state is calculated using the dec_outputs
### and used as 'state' in TransformerDecoder
dec_state = self.decoder.init_state(enc_outputs, *args)
### use initial state + input dec_X to the decoder to calculate
### the decoder output
return self.decoder(dec_X, dec_state)
# + [markdown] id="XrqvERfDfILO"
# ### Training
# + id="NKwxMS4ZqAdM"
### from d2l.ai
### because of the padding (and valid_length) we have to filter out some entries
class MaskedSoftmaxCELoss(gluon.loss.SoftmaxCELoss):
# `pred` shape: (`batch_size`, `seq_len`, `vocab_size`)
# `label` shape: (`batch_size`, `seq_len`)
# `valid_len` shape: (`batch_size`, )
def forward(self, pred, label, valid_len):
# weights shape: (batch_size, seq_len, 1)
weights = np.expand_dims(np.ones_like(label), axis=-1)
weights = npx.sequence_mask(weights, valid_len, True, axis=1)
return super(MaskedSoftmaxCELoss, self).forward(pred, label, weights)
# + id="_9vQqtmNqItq"
if False:
### from d2l.ai
loss = MaskedSoftmaxCELoss()
loss(np.ones((3, 4, 10)), np.ones((3, 4)), np.array([4, 2, 0]))
# + id="o70lOEd4ryf8"
### from d2l.ai
### prevents too high gradients
def grad_clipping(model, theta):
"""Clip the gradient."""
if isinstance(model, gluon.Block):
params = [p.data() for p in model.collect_params().values()]
else:
params = model.params
norm = math.sqrt(sum((p.grad ** 2).sum() for p in params))
if norm > theta:
for param in params:
param.grad[:] *= theta / norm
# + id="c4IEuqKWH0Oa"
### from d2l.ai
### accumulate results in one array, auxiliary function
class Accumulator:
"""For accumulating sums over `n` variables."""
def __init__(self, n):
self.data = [0.0] * n
def add(self, *args):
self.data = [a + float(b) for a, b in zip(self.data, args)]
def reset(self):
self.data = [0.0] * len(self.data)
def __getitem__(self, idx):
return self.data[idx]
# + id="2nalImXie535"
### from d2l.ai
def train_s2s_ch9(model, data_iter, lr, num_epochs, device):
model.initialize(init.Xavier(), force_reinit=True, ctx=device)
trainer = gluon.Trainer(model.collect_params(),
'adam', {'learning_rate': lr})
loss = MaskedSoftmaxCELoss()
animator = d2l.Animator(xlabel='epoch', ylabel='loss',
xlim=[1, num_epochs], ylim=[0, 0.75])
for epoch in range(1, num_epochs + 1):
timer = d2l.Timer()
metric = d2l.Accumulator(2) # loss_sum, num_tokens
### use data_iter from load_data_nmt to get X and Y which include:
### the source and target
### sentence representations + X_vlen and Y_vlen : the valid lenghts of
### the sentencies
for batch in data_iter:
X, X_vlen, Y, Y_vlen = [x.as_in_ctx(device) for x in batch]
Y_input, Y_label, Y_vlen = Y[:, :-1], Y[:, 1:], Y_vlen-1
with autograd.record():
Y_hat, _ = model(X, Y_input, X_vlen, Y_vlen)
l = loss(Y_hat, Y_label, Y_vlen)
l.backward()
grad_clipping(model, 1)
num_tokens = Y_vlen.sum()
trainer.step(num_tokens)
metric.add(l.sum(), num_tokens)
if epoch % 10 == 0:
animator.add(epoch, (metric[0]/metric[1],))
print(f'loss {metric[0] / metric[1]:.3f}, {metric[1] / timer.stop():.1f} '
f'tokens/sec on {str(device)}')
# + [markdown] id="u_sxg49EICVl"
# ### Reading and Processing the Text
# + id="rgUlZVmd1IHE"
### from d2l.ai
def download_extract(name, folder=None):
"""Download and extract a zip/tar file."""
fname = download(name)
base_dir = os.path.dirname(fname)
data_dir, ext = os.path.splitext(fname)
if ext == '.zip':
fp = zipfile.ZipFile(fname, 'r')
elif ext in ('.tar', '.gz'):
fp = tarfile.open(fname, 'r')
else:
assert False, 'Only zip/tar files can be extracted.'
fp.extractall(base_dir)
return os.path.join(base_dir, folder) if folder else data_dir
# + [markdown] id="cvPS1MbLGVab"
# """ ... a dataset that contains a set of English sentences with the corresponding French translations. As can be seen that each line contains an English sentence with its French translation, which are separated by a TAB.""" (citation from d2l.ai)
# + id="N6qY1LlJhALj"
### d2l.ai
### the data for the translation are prepared by the d2l.ai project (book)
d2l.DATA_HUB['fra-eng'] = (d2l.DATA_URL + 'fra-eng.zip',
'94646ad1522d915e7b0f9296181140edcf86a4f5')
def read_data_nmt():
data_dir = d2l.download_extract('fra-eng')
with open(os.path.join(data_dir, 'fra.txt'), 'r') as f:
return f.read()
# + id="oZ1WHzBLjIID" outputId="6875b3fb-ce10-499d-f212-acf9bd71e6a2" colab={"base_uri": "https://localhost:8080/", "height": 196}
raw_text = read_data_nmt()
print(raw_text[0:106])
# + id="Gue5_rT_GL65"
### from d2l.ai
def preprocess_nmt(text):
def no_space(char, prev_char):
return char in set(',.!') and prev_char != ' '
text = text.replace('\u202f', ' ').replace('\xa0', ' ').lower()
out = [' ' + char if i > 0 and no_space(char, text[i-1]) else char
for i, char in enumerate(text)]
return ''.join(out)
# + id="X9iThu9ymSAj" outputId="6061971a-4e27-462f-da83-48d883601e36" colab={"base_uri": "https://localhost:8080/", "height": 156}
### from d2l.ai
text = preprocess_nmt(raw_text)
print(text[0:95])
# + id="_Dm_Tx5vGCSl"
### from d2l.ai
def tokenize_nmt(text, num_examples=None):
source, target = [], []
for i, line in enumerate(text.split('\n')):
if num_examples and i > num_examples:
break
parts = line.split('\t')
if len(parts) == 2:
source.append(parts[0].split(' '))
target.append(parts[1].split(' '))
return source, target
# + id="CEQ6O9wRmdn_" outputId="bb89429f-9fa0-4793-f590-46446f1463b7" colab={"base_uri": "https://localhost:8080/", "height": 57}
### from d2l.ai
source, target = tokenize_nmt(text)
source[0:3], target[0:3]
# + [markdown] id="kQbNVIJjm5Km"
# #### Histogram of the number of tokens per sentence
#
# There are mostly 5 token sentencies, num of tokens is
# usually less than 10..15.
# + id="BtOp7UHxm2GI" outputId="dff7b7c0-b797-470e-d947-3f3cde18dd23" colab={"base_uri": "https://localhost:8080/", "height": 244}
### from d2l.ai
d2l.set_figsize()
d2l.plt.hist([[len(l) for l in source], [len(l) for l in target]],
label=['source', 'target'])
d2l.plt.legend(loc='upper right');
# + [markdown] id="QAJ31deVz58_"
# ### Vocabulary
# + id="u-ZnTJKdxw2s"
### from d2l.ai
def count_corpus(tokens):
"""Count token frequencies."""
# Here `tokens` is a 1D list or 2D list
if len(tokens) == 0 or isinstance(tokens[0], list):
# Flatten a list of token lists into a list of tokens
tokens = [token for line in tokens for token in line]
return collections.Counter(tokens)
# + id="yvc84yOHxVWo"
### from d2l.ai
class Vocab:
"""Vocabulary for text."""
def __init__(self, tokens=None, min_freq=0, reserved_tokens=None):
if tokens is None:
tokens = []
if reserved_tokens is None:
reserved_tokens = []
# Sort according to frequencies
counter = count_corpus(tokens)
self.token_freqs = sorted(counter.items(), key=lambda x: x[0])
self.token_freqs.sort(key=lambda x: x[1], reverse=True)
# The index for the unknown token is 0
self.unk, uniq_tokens = 0, ['<unk>'] + reserved_tokens
uniq_tokens += [token for token, freq in self.token_freqs
if freq >= min_freq and token not in uniq_tokens]
self.idx_to_token, self.token_to_idx = [], dict()
for token in uniq_tokens:
self.idx_to_token.append(token)
self.token_to_idx[token] = len(self.idx_to_token) - 1
def __len__(self):
return len(self.idx_to_token)
def __getitem__(self, tokens):
if not isinstance(tokens, (list, tuple)):
return self.token_to_idx.get(tokens, self.unk)
return [self.__getitem__(token) for token in tokens]
def to_tokens(self, indices):
if not isinstance(indices, (list, tuple)):
return self.idx_to_token[indices]
return [self.idx_to_token[index] for index in indices]
# + id="8XIGBHAEz83o" outputId="6e71de82-effb-4b14-bdd7-d03ceea13861" colab={"base_uri": "https://localhost:8080/", "height": 37}
### from d2l.ai
src_vocab = Vocab(source, min_freq=3,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
len(src_vocab)
# + [markdown] id="WlymolB-zvf-"
# ### Loading the dataset
# + id="hT96fUzTF5LG"
### from d2l.ai
def truncate_pad(line, num_steps, padding_token):
if len(line) > num_steps:
return line[:num_steps] # Trim
return line + [padding_token] * (num_steps - len(line)) # Pad
### the <pad> is represented by number 1 in Vocabuary
# + id="xcCKS0imn8Ll" outputId="b0a17a35-ec1a-4e31-cf33-2909da189a59" colab={"base_uri": "https://localhost:8080/", "height": 37}
### from d2l.ai
truncate_pad(src_vocab[source[0]], 10, src_vocab['<pad>'])
# + id="30na3aM9FuuO"
### from d2l.ai
def build_array(lines, vocab, num_steps, is_source):
lines = [vocab[l] for l in lines]
if not is_source:
lines = [[vocab['<bos>']] + l + [vocab['<eos>']] for l in lines]
array = np.array([truncate_pad(
l, num_steps, vocab['<pad>']) for l in lines])
valid_len = (array != vocab['<pad>']).sum(axis=1)
return array, valid_len
# + id="_C9-MV_DtO3b"
### from d2l.ai
def load_array(data_arrays, batch_size, is_train=True):
"""Construct a Gluon data iterator."""
dataset = gluon.data.ArrayDataset(*data_arrays)
return gluon.data.DataLoader(dataset, batch_size, shuffle=is_train)
# + id="DgDvaAHrFS29"
### from d2l.ai
### quite importand function to construct dataset for training (data_iter)
### from original data
def load_data_nmt(batch_size, num_steps, num_examples=12800):
text = preprocess_nmt(read_data_nmt())
source, target = tokenize_nmt(text, num_examples)
src_vocab = Vocab(source, min_freq=3,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
tgt_vocab = Vocab(target, min_freq=3,
reserved_tokens=['<pad>', '<bos>', '<eos>'])
src_array, src_valid_len = build_array(
source, src_vocab, num_steps, True)
tgt_array, tgt_valid_len = build_array(
target, tgt_vocab, num_steps, False)
data_arrays = (src_array, src_valid_len, tgt_array, tgt_valid_len)
data_iter = load_array(data_arrays, batch_size)
return src_vocab, tgt_vocab, data_iter
# + id="E2Vbr344rbhI"
### from d2l.ai
def try_gpu(i=0):
"""Return gpu(i) if exists, otherwise return cpu()."""
return npx.gpu(i) if npx.num_gpus() >= i + 1 else npx.cpu()
# + [markdown] id="uM6PC5noDevE"
# ### Model: training and prediction
# + id="zlHL925Pl4VU" outputId="841323c4-2bc0-469a-87bc-3a83347166c9" colab={"base_uri": "https://localhost:8080/", "height": 302}
### the code from d2l.ai
### estimate the execution time for the cell in seconds
start = time.time()
num_hiddens, num_layers, dropout, batch_size, num_steps = 32, 2, 0.0, 128, 10
lr, num_epochs, device = 0.001, 600, try_gpu()
ffn_num_hiddens, num_heads = 64, 4 ### num_hiddens is to be a multiple of num_heads !!
src_vocab, tgt_vocab, train_iter = load_data_nmt(batch_size, num_steps)
encoder = TransformerEncoder(
len(src_vocab), num_hiddens, ffn_num_hiddens, num_heads, num_layers,
dropout)
decoder = TransformerDecoder(
len(src_vocab), num_hiddens, ffn_num_hiddens, num_heads, num_layers,
dropout)
model = EncoderDecoder(encoder, decoder)
train_s2s_ch9(model, train_iter, lr, num_epochs, device)
### estimate the execution time for the cell
end = time.time()
print(end - start)
# + id="cmPoZHgvBhw8"
### from d2l.ai
def predict_s2s_ch9(model, src_sentence, src_vocab, tgt_vocab, num_steps,
device):
src_tokens = src_vocab[src_sentence.lower().split(' ')]
enc_valid_len = np.array([len(src_tokens)], ctx=device)
src_tokens = truncate_pad(src_tokens, num_steps, src_vocab['<pad>'])
enc_X = np.array(src_tokens, ctx=device)
# Add the batch size dimension
enc_outputs = model.encoder(np.expand_dims(enc_X, axis=0),
enc_valid_len)
dec_state = model.decoder.init_state(enc_outputs, enc_valid_len)
dec_X = np.expand_dims(np.array([tgt_vocab['<bos>']], ctx=device), axis=0)
predict_tokens = []
for _ in range(num_steps):
Y, dec_state = model.decoder(dec_X, dec_state)
# The token with highest score is used as the next time step input
dec_X = Y.argmax(axis=2)
py = dec_X.squeeze(axis=0).astype('int32').item()
if py == tgt_vocab['<eos>']:
break
predict_tokens.append(py)
return ' '.join(tgt_vocab.to_tokens(predict_tokens))
# + id="hxmimesrTg9T" outputId="254fc611-6be4-47ed-9bd6-ea8d91e90a67" colab={"base_uri": "https://localhost:8080/", "height": 453}
for sentence in ['Go .', 'Wow !', "I'm OK .", 'I won !',
'Let it be !', 'How are you ?', 'How old are you ?',
'Cats are cats, dogs are dogs .', 'My friend lives in US .',
'He is fifty nine years old .', 'I like music and science .',
'I love you .', 'The dog is chasing the cat .',
'Somewhere on the earth .', 'Do not worry !',
'Sit down, please !', 'Not at all !', 'It is very very strange .',
'Take it into account .', 'The dark side of the moon .',
'Come on !', 'We are the champions, my friends .']:
print(sentence + ' => ' + predict_s2s_ch9(
model, sentence, src_vocab, tgt_vocab, num_steps, device))
| experiments/Final_Transformer_MXNet_12800_11_10_20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from source.conversation_helper import conversation_filter
from source.visualization_helper import init_plot_style
init_plot_style()
ds = pd.read_csv('/Users/john/data/twitter/tweets_ec_nd_conv.csv', usecols=['id', 'screen_name', 'text', 'hashtags','num_replies', 'conversation_id'])
ds.shape
conv = conversation_filter(ds)
conv.shape
# # text
conv['textlen'] = conv.text.str.len();
#textlen=ds.groupby('textlen').size()
#textlen.reset_index(inplace=True)
fig = plt.figure()
conv.textlen.plot.hist( bins=100)
#plt.xscale('log')
plt.title('number of chars in tweets')
plt.xlabel('chars')
plt.xlim([0, 180])
#plt.xscale('log')
fig.autofmt_xdate()
# # hashtags
s = conv["hashtags"].str.split(';', expand=True).stack()
i = s.index.get_level_values(0)
s.index = s.index.droplevel(-1)
ds_hashtags = ds[['id']].loc[i].copy()
ds_hashtags['hashtags'] = s.values
ds_hashtags.head()
perc = ds_hashtags.id.nunique() / ds.shape[0]
print(perc)
ht=ds_hashtags.groupby('hashtags').size()
top=ht.sort_values(ascending=False).head(50)
x=range(1, len(top)+1,1)
y=top.values
plt.scatter(x, y, marker='x')
plt.yscale('log')
plt.xlabel('top 50 hashtags')
plt.ylabel('tweets')
# +
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
from nltk.tokenize import word_tokenize
import re
use_stemmer = False
stemmer = SnowballStemmer("spanish")
stopset = set(stopwords.words('english') + stopwords.words('spanish'))
def tokenize(text):
text = re.sub(r"(?:\#|@|https?\://)\S+", "", text)
text = re.sub("[^a-zA-Z] ","",text)
text = re.sub(r'\b\w{1,2}\b',' ',text)
text = re.sub(r'http[\bs]*',' ',text)
tokens = word_tokenize(text)
if use_stemmer:
tokens = [stemmer.stem(t) for t in tokens]
return tokens
words = ' '.join(conv['text'])
use_stemmer=False
word_list = tokenize(words)
word_list = [word for word in word_list if word not in stopset]
print(f'number of words with stop words removal: {len(word_list)}')
wds = pd.DataFrame(word_list)
wds.columns = ['word']
wds=wds.word.value_counts().reset_index()
wds.columns=['word', 'count']
wds=wds[wds['count']>3]
print(f'number of words (TF min:3): {len(wds)}')
# -
wds2=wds.sort_values('count', ascending=False).head(1000)
from wordcloud import WordCloud
filtered_words = ' '.join(wds2.word)
wordcloud = WordCloud(background_color='white', width=500, height=300).generate(filtered_words)
plt.imshow(wordcloud)
plt.axis("off")
# # text analysis
questions=conv[conv.text.str.endswith('?')]
print(questions.shape)
questions.head()
admiration=conv[conv.text.str.endswith('!')]
print(admiration.shape)
admiration.head()
suspense=conv[conv.text.str.endswith('..')]
print(suspense.shape)
suspense.head()
| exploratory_text.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Welcome Home
#
# Usually when Steve comes home, there is no one at home. Steve can get lonely at times especially after long hard battle with creepers and zombies.
#
#
# In this programming adventure we'll make Minecraft display a warm and friendly welcome message when Steve comes home. We'll test your program by exploring the world and then come back home to a friendly welcome. Along the way we will learn about coordinate systems which help us locate objects in a game. We will also learn about variables and conditions.
# ## Coordinate Systems
#
# From your math classes, you will remember coordinate systems to locate points in a plane. The points (2,3), (-3,1) and (-1,-2.5) are shown in the grid below.
#
# 
#
# The Minecraft coordinate grid is shown below:
#
# 
#
# In Minecraft, when you move East, your X-coordinate increases and when you move South, your Z-coordinate increases. Let's confirm this through a few Minecraft exercises.
#
#
# ## Task 1: Moving in Minecraft coordinate systems
#
# In Minecraft look at Steve's coordinates. Now move Steve to any other position. See how his coordinates change as you move?
#
# - [ ] Change your direction so that only the Xcoordinate moves when you move forward or back.
#
# - [ ] Change your direction so only the Z-coordinate moves when you move forward or back.
# ## Task 2: Write a program to show Steve's position on the screen
# Remember functions from the first Adventure? A function lets us do things in a computer program or in the minecraft game. The function **getTilePos()** get the players position as (x,y,z) coordinates in Minecraft. Let's using this function to print Steve's position as he moves around. We need to store Steve's position when we call the function **getTilePos()** so that we can print the position later. We can use a program **variable** to store the position. A variable has a name and can be used to store values. We'll call our variable **pos** for position and it will contain the Steve's position. When we want to print the position, we print the values of the position x,y and z coordinates using another function **print()** which prints any strings you give it.
#
# Start up minecraft and type the following in a new cell.
#
# ```python
# from mcpi.minecraft import *
# mc = Minecraft.create()
# pos = mc.player.getTilePos()
# print(pos.x)
# print(pos.y)
# print(pos.z)
# ```
#
# When you run your program by pressing Ctrl+Enter in the program cell, you should now see Steve's position printed.
#
# ** Great Job!**
#
from mcpi.minecraft import *
import time
mc = Minecraft.create()
# Type Task 2 program here
# ## Task 3: Prettying up messages
# The messages we printed are somewhat basic and can be confusing since we don't know which number is x,y or z. Why not print a message that is more useful. Often messages are built by attaching strings and data. Try typing
#
# ```python
# "my name is " + "Steve"
# ```
#
# in a code cell. What message gets printed? Now try
#
# ```python
# "my age is " + 10
# ```
#
# Hmmm... That did not work :( Strings can only be attached or _concatenated_ with other strings. In order to attach a number to string, we need to convert the number into a printable string. We will use another function **str()** which returns a printable string of its arguments. Since x,y,z coordinates are numbers, we need to convert them to strings in order to print them with other strings. Too how the str() function works type the following in a code cell and run.
#
# ```python
# "my age is " + str(10)
# ```
#
# What gets printed by the line below?
#
# ```python
# "x = " + str(10) + ",y = " + str(20) + ",z = " + str(30)
# ```
#
# You now have all the information you need to print a pretty message.
#
# - [ ] Modify your program to print a pretty message shown below to correctly print Steve's position
# Steve's position is: x = 10,y = 20,z = 30
# - [ ] Modify your program to use a variable names _message_ to store the pretty message and then print the message
# Hint:
#
# ```python
# message = ...
# print(message)
# ```
#
## Type Task 3 program here
# ## Task 4: Display Steve's coordinates in Minecraft
# For this task instead of printing Steve's coordinates, lets display them in Minecraft using the **postToChat()** function from Adventure1
#
# You should see a message like the one below once you run your program.
#
while True:
time.sleep(1)
## Type Task 4 program here
# ## Home
# In Minecraft move to a location that you want to call home and place a Gold block there. Move Steve on top of the Gold block and write down his coordinates. Lets save these coordinates in the variables **home_x** and **home_z**. We will use these variables to detect when Steve returns home.
#
## Change these values for your home
home_x = 0
home_z = 0
# ## Is Steve home?
# Now the magic of figuring out if Steve is home. As Steve moves in Minecraft, his x and z coordinates change. We can detect that Steve is home when his coordinates are equal to the cordinates of his home! To put it in math terms, Steve is home when
#
# $$
# (pos_{x},pos_{z}) = (home_{x},home_{z})
# $$
#
# In the program we can write the math expression as
#
# ```python
# pos.x == home_x and pos.z == home_z
# ```
#
# We can an **if** program block to check if Steve's coordinates equal his home coordinates. An **if** block is written as shown below
#
# ```python
# if (condition):
# do something 1
# do something 2
# ```
#
# Lets put this all together in the program below
# ```python
# while True:
# time.sleep(1)
# pos = mc.player.getTilePos()
# if (pos.x == home_x and pos.z == home_z):
# mc.postToChat("Welcome home Steve.")
# # the rest of your program from task 4
# ```
#
# What happens when you run around in Minecraft and return to the gold block that is your home? That warm message makes Steve happy. He can now be well rested for battling creepers the next day.
## Type Task 5 program here
# ## Recap
#
# In this adventure you learned about **coordinates**, **variables** and **if conditions**. You used your knowledge to greet Steve with a warm message when he returns home.
#
# ** Great Job! **
#
#
| notebooks/Adventure2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + hideCode=false hideOutput=false
# -*- utf-8 -*-
from setup import *
import pandas as pd
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
import seaborn as sns
# + hideCode=false hideOutput=true
# notebook setup when plotting
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
# -
# # Fonts
# check avalable fonts
plot.check_fonts() # function from setup
# set specific font
plt.rcParams["font.family"] = 'Noto Sans CJK JP'
plt.rcParams["font.weight"] = "bold"
# # Figure setup
# ## Matplotlib
# set the size and the dpi of the figure
plt.figure(figsize=(25,3), dpi= 80)
# subplot position for one specific figure
plt.subplot(rows,cols,position)
# turn off the axises
plt.axis('off')
# +
# # ???
plt.rcParams['axes.unicode_minus'] = False
# -
# update plotting params
large = 22; med = 16; small = 12
params = {'axes.titlesize': large, 'legend.fontsize': med, 'figure.figsize': (16, 10), 'axes.labelsize': med,
'xtick.labelsize': med, 'ytick.labelsize': med, 'figure.titlesize': large}
plt.rcParams.update(params)
# update plotting style
plt.style.use('seaborn-whitegrid')
# set label names
plt.ylabel('name')
plt.xlabel('name')
# use specific font in one specific place
font2 = {'weight' : 'normal','size' : 20, }
plt.ylabel('{}评分'.format(scale),font=font2)
# show a pre-existing picture
im = plt.imread('image.png')
plt.imshow(im)
# +
# # ???
plt.tick_params(labelsize=20)
# -
# set the range of y axis
plt.ylim(0, y_max*1.3)
# use this function before you save a figure incase of there was too much white space
plt.tight_layout()
# ### axes
ax = plt.gca() # get current axes
ax.xaxis.set_ticks_position('top')
ax.invert_yaxis()
# 将右边和上边的边框(脊)的颜色去掉
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
# set ticks positions
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
# ## Seaborn
# Font scale
sns.set(font_scale=1)
sns.set_style("white")
# # Plot
# ## Matplotlib
# direct plot from pandas dataframe
pd.DataFrame().value_counts(normalize=True).plot(kind='pie', shadow=True, autopct='%.2f%%', title="abc", colormap='gray_r',fontsize=20)
# plot annotation
plt.annotate(r'%s'%sig, xy=(x0, y0), xycoords='data', xytext=(-15, +1),
textcoords='offset points', fontsize=25,color="black")
# plot a dot
plt.plot(1.1,y1,'o',color='black')
# plot a line
plt.plot([1,2], [3,4],linestyle="dashed", color = 'black',linewidth=0.8) # [x1,x2], [y1,y2]
# bar plot
plt.bar(x, scale_mean, width=width, yerr = scale_std, tick_label=labels, linewidth=1.5,label='Control',color='gray')
# ## Seaborn
import numpy as np
mask = np.zeros_like(df1.corr())
print(mask.shape)
for i in range(len(mask)):
for j in range(i+1, len(mask[0])):
mask[i][j] = True
plt.figure(figsize=(15,15))
sns.heatmap(df1.corr(),cmap="RdBu_r",annot=True, mask=mask)
sns.heatmap(heatmap_data, cmap='Reds_r',annot=True)
sns.histplot(hist_data, color='gray')
# linear model plot
sns.lmplot(y, x, hue, data, palette='gray',markers=['o','^'])
# regressiong plot
sns.regplot(x_label, y_label, dataframe)
# + hideCode=true hideOutput=false hidePrompt=false
# get the data of all scales
sql = 'SELECT * FROM Scale '
dbcr.execute(sql)
Scale = pd.DataFrame(dbcr.fetchall(), columns=get_columns('Scale'))
Scale_Filtered = Scale[Scale['ID'].isin(ID)]
Scale_Filtered = Scale_Filtered[~Scale_Filtered['timepoint'].isin([4])].set_index(['ID']) # 有一个两次后测2的 通过~取反,选取不包含数字4的行
# + hideCode=true hideOutput=false
Scale_corr_1= Scale_Filtered[Scale_Filtered['timepoint'].isin([1])].drop(index='P_010')[columns_for_anal] # Reorder
Scale_corr_1[['Age','ASIA','Injured_seg','Seg_count']] = get_patinet_info(Scale_corr_1, ['Age','ASIA_Pretest','Injured_Segment','Count_of_Injured_Segment'])
missing_pt_corr = pd.DataFrame(Scale_corr_1.isnull().sum(axis=0), columns=['关联分析']).drop(index=['timepoint']).T
# + hideCode=true
# 缺失数据统计
missing = pd.concat([missing_pt_1,missing_pt_2, missing_paired, missing_pt_corr])
#save_table(missing,"缺失值")
| analysis_behaviour_data/plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Activity 7.02: Visualizing Stock Prices with Bokeh
# This activity will combine most of what you have already learned about Bokeh. You will also need the skills you have acquired while working with Pandas.
#
# We will create an interactive visualization that displays a candle stick plot, which is often used when handling stock price data.
# We will be able to compare two stocks with each other by selecting them from dropdowns.
# A RangeSlider will allow us to restrict the displayed date range in the requested year 2016.
# Depending on what graph we choose, we will either see the candle stick visualization or a simple line plot displaying the volume of the selected stock.
# The dataset of this exercise contains temporal stock price data.
# This means we'll be looking at data over a range of time.
# #### Loading our dataset
# +
# importing the necessary dependencies
import pandas as pd
from bokeh.io import output_notebook
output_notebook()
# -
# loading the Dataset with geoplotlib
dataset = pd.read_csv('../../Datasets/stock_prices.csv')
# looking at the dataset
dataset.head()
# Just as in the previous exercise, we want to map the date column to another column with the shortened date that only contains the year, month, and day.
# +
# mapping the date of each row to only the year-month-day format
from datetime import datetime
def shorten_time_stamp(timestamp):
shortened = timestamp[0]
if len(shortened) > 10:
parsed_date=datetime.strptime(shortened, '%Y-%m-%d %H:%M:%S')
shortened=datetime.strftime(parsed_date, '%Y-%m-%d')
return shortened
dataset['short_date'] = dataset.apply(lambda x: shorten_time_stamp(x), axis=1)
# -
# **Note:**
# The exectuion of the cell will take a moment since it's a fairly large dataset.
# Please be patient.
# looking at the dataset with shortened dat
dataset.head()
# **Note:**
# The last, newly added, column now holds the timestamp without the hour, minute, and second information.
# ---
# #### Building an interactive visualization
# There are many options when it comes to choosing which interactivity to use.
# Since the goal of this activity is to be able to compare two stocks with each other in terms of traded volume and the high/low and open/close prices over a time range, we will need widgets to select elements and a slider to select a given range.
# Considering that we have to options of display, we also need a way to select either one or the other.
#
# At the end of this task, you will have something comparable to image below implemented and ready to compare data from the dataset.
#
# <img src="./assets/candle_plot.png" width=500 align="left"/>
# importing the necessary dependencies
from bokeh.plotting import figure, show
from ipywidgets import interact, widgets
# Before we go in and implement the plotting methods, we want to set up the interactivity widgets.
# **Please scroll down** to the cell that says `# extracing the necessary data` before implementing the plotting.
#
# Still make sure to execute the cells below that will simply `pass` and do nothing for now.
# At the moment, our `show()` in the last cell will not render any elements into our visualization.
# We will start with the, so called, candle stick visualization which is often used with stock price data.
#
# The already defined method below gets our `plot` object, a `stock_name`, a `stock_range` containing the data for the, with the widgets, selected range, and a color for the line.
# We will use those arguments to create the candle sticks. They basically contain a `segment` that creates the vertical line, and either a green or red `vbar` to color code whether the close price is lower than the open price.
#
#
# Once you have created the candles, we also want to have a continuous line running through the mean (high, low) point of each candle. So you have to calculate the mean for every high, low pair and then plot those data points with a line with the given `color`.
#
# Make sure to reference the example provided in the Bokeh library here, you can adapt the code in there to our arguments.
# https://bokeh.pydata.org/en/latest/docs/gallery/candlestick.html
#
# After you are done implementing the `add_candle_plot` method, scroll down and run the `@interact` cell again.
# You will now see the candles being displayed for the two selected stocks.
# **Note:**
# Since we are providing the `plot` as a reference to the method, we don't need to return anything since we are mutating our passed in plot.
def add_candle_plot(plot, stock_name, stock_range, color):
inc_1 = stock_range.close > stock_range.open
dec_1 = stock_range.open > stock_range.close
w = 0.5
plot.segment(stock_range['short_date'], stock_range['high'],
stock_range['short_date'], stock_range['low'],
color="grey")
plot.vbar(stock_range['short_date'][inc_1], w,
stock_range['high'][inc_1], stock_range['close'][inc_1],
fill_color="green", line_color="black",
legend_label=('Mean price of ' + stock_name), muted_alpha=0.2)
plot.vbar(stock_range['short_date'][dec_1], w,
stock_range['high'][dec_1], stock_range['close'][dec_1],
fill_color="red", line_color="black",
legend_label=('Mean price of ' + stock_name), muted_alpha=0.2)
stock_mean_val=stock_range[['high', 'low']].mean(axis=1)
plot.line(stock_range['short_date'], stock_mean_val,
legend_label=('Mean price of ' + stock_name), muted_alpha=0.2,
line_color=color, alpha=0.5)
# The last missing step is implementing the plotting of the lines if the `volume` value is selected.
# We've created simple lines in the previous exercise, so this should not be a problem.
#
# One additional interaction feature is to have an interactive legend that allows us to "mute", meaning grey out, each stock in the visualization.
# To make our legend interactive please take a look at the documentation for the legend feature.
# https://bokeh.pydata.org/en/latest/docs/user_guide/interaction/legends.html
# **Note:**
# Don't forget to update your `add_canlde_plot` `vbar`s and `segment` to also include the `muted_alpha` parameter. Otherwise you won't be able to mute the stocks in the "open-close" visualization.
# method to build the plot
def get_plot(stock_1, stock_2, date, value):
stock_1 = dataset[dataset['symbol'] == stock_1]
stock_2 = dataset[dataset['symbol'] == stock_2]
stock_1_name=stock_1['symbol'].unique()[0]
stock_1_range=stock_1[(stock_1['short_date'] >= date[0]) & (stock_1['short_date'] <= date[1])]
stock_2_name=stock_2['symbol'].unique()[0]
stock_2_range=stock_2[(stock_2['short_date'] >= date[0]) & (stock_2['short_date'] <= date[1])]
plot=figure(title='Stock prices',
x_axis_label='Date',
x_range=stock_1_range['short_date'],
y_axis_label='Price in $USD',
plot_width=800,
plot_height=500)
plot.xaxis.major_label_orientation = 1
plot.grid.grid_line_alpha=0.3
if value == 'open-close':
add_candle_plot(plot, stock_1_name, stock_1_range, 'blue')
add_candle_plot(plot, stock_2_name, stock_2_range, 'orange')
if value == 'volume':
plot.line(stock_1_range['short_date'], stock_1_range['volume'],
legend_label=stock_1_name, muted_alpha=0.2)
plot.line(stock_2_range['short_date'], stock_2_range['volume'],
legend_label=stock_2_name, muted_alpha=0.2,
line_color='orange')
plot.legend.click_policy="mute"
return plot
# We want to **start implementing our visualization here**.
#
# In the following cells, we will extract the necessary data which will be provided to the widget elements.
# In the first cell we want to extract the following information:
# - a list of unique stock names that are present in the dataset
# - a list of all short_dates that are in 2016
# - a sorted list of unique dates generated from the previous list of dates from 2016
# - a list with the values `open-close` and `volume`
#
# Once we have this information in place, we can start building our widgets.
# extracing the necessary data
stock_names=dataset['symbol'].unique()
dates_2016=dataset[dataset['short_date'] >= '2016-01-01']['short_date']
unique_dates_2016=sorted(dates_2016.unique())
value_options=['open-close', 'volume']
# Given the extracted information from the cell above, we can now define the widgets and provide the available options to it.
#
# As mentioned in the introduction, we want to have several interactive features including:
# - two `Dropdown`s with which we can select two stocks that should be compared to each other
# - the first dropdown by default should have the `AAPL` stock selected, named "Compare: "
# - the second dropdown by default should have the `AON` stock selected, named "to: "
#
#
# - a `SelectionRange` which will allow us to select a range of dates from the extracted list of unique 2016 dates
# - by default, the first 25 dates should be selected, named "From-To"
# - make sure to disable the `continuous_update` parameter here
# - adjust the layout width to 500px to make sure the dates are displayed correctly
#
#
# - a `RadioButton` group that provides the options "open-close" and "volume"
# - by default, "open-close" should be selected, named "Metric"
# +
# setting up the interaction elements
drp_1=widgets.Dropdown(options=stock_names,
value='AAPL',
description='Compare:')
drp_2=widgets.Dropdown(options=stock_names,
value='AON',
description='to:')
range_slider=widgets.SelectionRangeSlider(options=unique_dates_2016,
index=(0,25),
continuous_update=False,
description='From-To',
layout={'width': '500px'})
value_radio=widgets.RadioButtons(options=value_options,
value='open-close',
description='Metric')
# -
# **Note:**
# As mentioned in the previous exercise, we can also make use of the widgets described here: https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html
# After setting up the widgets, we can the method that will be called with each update of the interaction widgets.
# As seen in the previous exercise, we will use the `@interact` decorator for this.
#
# Instead of value ranges or lists, we will provide the variable names of our already created widgets in the decorator.
# The method will get 4 arguments, `stock_1`, `stock_2`, `date`, and `value`.
# Since we have already set up the empty method that will return a plot above, we can call `show()` with the method call inside to show the result once it is returned from the `get_stock_for_2016` method.
#
# Once you've build the widgets, upon execution, you will see them being displayed below the cell.
# We are now ready to to **scroll up and implement the plotting** with Bokeh.
# creating the interact method
@interact(stock_1=drp_1, stock_2=drp_2, date=range_slider, value=value_radio)
def get_stock_for_2016(stock_1, stock_2, date, value):
show(get_plot(stock_1, stock_2, date, value))
# This is a nice example that shows us how much interaction we can add to a visualization with very simple techniques such as using the interact functionality.
#
# **Note:**
# Think about what else you could add/change for this visualization. Maybe we don't only want to display 2016 but be able to select the year we want to display. Maybe we want to compare different years with each other.
# There are endless options.
#
| Chapter07/Activity7.02/Activity7.02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Processing Results from Solving Dynamic Programming Problem using Grids
#
# There are several state variables. We solved for the optimal choices and values along grids of the state variables.
#
# There are several matrixes/tensors etc that store the value function policy function results where each dimension corresponds to a discretized state space grid.
#
# There are also several (unique-valued) grid that store the grid values at which we solved the problems.
# ## Program
# +
ff_dyna_sup_expand_grids <- function(ar.st.vars, list.ar.fl,
list.ts.valpolmat) {
# val <- array(c(1, 1.5, 0, 2, 0, 4, 0, 3), dim=c(2, 2, 2))
# pol <- array(runif(8), dim=c(2, 2, 2))
# list.ts.valpolmat <- list(val=val, pol=pol)
# ar.fl.x <- c(1.1,2.3)
# ar.fl.y <- c(2.1,3.3)
# ar.fl.z <- c(3.1,4.3)
# ar.st.vars <- c('vx', 'vy', 'vz')
# list.ar.fl <- list(ar.fl.x, ar.fl.y, ar.fl.z)
mt.fl.expanded <- do.call(expand.grid, list.ar.fl)
colnames(mt.fl.expanded) <- ar.st.vars
df.out <- as_tibble(bind_cols(mt.fl.expanded, list.ts.valpolmat))
return(df.out)
}
# -
# ## Load Data
# +
# Library
library(tidyverse)
library(AER)
library(R.matlab)
# Load Sample Data
setwd('C:/Users/fan/R4Econ/_data/')
matfile <- 'vf_az_p_ga_sa0.mat'
mat.out <- readMat(matfile)
# -
# ## Test Program
# +
# 1 is r1c1t1, 1.5 in r2c1t1, 0 in r1c2t1, etc.
# Three dimensions, row first, column second, and tensor third
val <- array(c(1, 1.5, 0, 2, 0, 4, 0, 3), dim=c(2, 2, 2))
pol <- array(runif(8), dim=c(2, 2, 2))
# dim(x)
# print(x)
# Suppose the three dimensions above come from some value function problem with three states each solved along a grid
ar.fl.x <- c(1.1,2.3)
ar.fl.y <- c(2.1,3.3)
ar.fl.z <- c(3.1,4.3)
ar.st.vars <- c('vx', 'vy', 'vz')
# Inputs
list.ar.fl <- list(ar.fl.x, ar.fl.y, ar.fl.z)
list.ts.valpolmat <- tibble(val=as.numeric(val), pol=as.numeric(pol))
# -
# Invoke Program
ff_dyna_sup_expand_grids(ar.st.vars, list.ar.fl, list.ts.valpolmat)
# ## Line by Line
# +
# 1 is r1c1t1, 1.5 in r2c1t1, 0 in r1c2t1, etc.
# Three dimensions, row first, column second, and tensor third
val <- array(c(1, 1.5, 0, 2, 0, 4, 0, 3), dim=c(2, 2, 2))
pol <- array(runif(8), dim=c(2, 2, 2))
# dim(x)
# print(x)
# Suppose the three dimensions above come from some value function problem with three states each solved along a grid
ar.fl.x <- c(1.1,2.3)
ar.fl.y <- c(2.1,3.3)
ar.fl.z <- c(3.1,4.3)
ar.st.vars <- c('vx', 'vy', 'vz')
# -
# Input
list.ar.fl <- list(ar.fl.x, ar.fl.y, ar.fl.z)
list.ts.valpolmat <- tibble(val=as.numeric(val), pol=as.numeric(pol))
# +
# Generating Combined Dataframe, Model Solution as Dataframe
mt.fl.expanded <- do.call(expand.grid, list.ar.fl)
colnames(mt.fl.expanded) <- ar.st.vars
as_tibble(bind_cols(mt.fl.expanded, list.ts.valpolmat))
# as_tibble(bind_cols(mt.fl.expanded, list.ts.valpolmat))
# for (i in 1:length(list.input)) {
# df.fl.expanded <- df.fl.expanded %>% mutate(val = val)
# }
| dynamic/support/expandgrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/data-datum/AnalisisyVisualizacion/blob/master/diplo_vis_punto1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="n9PuYzpFJa8W"
# # Visualización de datos sysarmy y análisis exploratorio
#
#
#
#
#
# + id="gxSTkgXlJWfR"
import io
import matplotlib
import matplotlib.pyplot as plt
import numpy
import pandas as pd
import seaborn
seaborn.set_context('talk')
# + id="BGrRzTuQJ31o"
url = 'https://cs.famaf.unc.edu.ar/~mteruel/datasets/diplodatos/sysarmy_survey_2020_processed.csv'
df = pd.read_csv(url)
# + colab={"base_uri": "https://localhost:8080/", "height": 302} id="sMXqawSsKFgi" outputId="0c8177ce-df74-4bb7-89b0-793d2f8f7456"
df[:3]
# + [markdown] id="7O_7awpkJ-1j"
# # Análisis Descriptivo
# + id="hpNwUZjmLUpq"
relevant_columns = ['tools_programming_languages', 'salary_monthly_NETO']
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="sFasu6NNJ7y3" outputId="593ad880-8e1a-4c3c-dbfb-74f670929257"
# Convert the comma-separated string of languages to a list of string.
# Remove 'ninguno de los anteriores' option, spaces and training commas.
def split_languages(languages_str):
if not isinstance(languages_str, str):
return []
# Remove 'other' option
languages_str = languages_str.lower()\
.replace('ninguno de los anteriores', '')
# Split string into list of items
# Remove spaces and commas for each item
return [lang.strip().replace(',', '')
for lang in languages_str.split()]
# Create a new column with the list of languages
df.loc[:, 'cured_programming_languages'] = df.tools_programming_languages\
.apply(split_languages)
if 'cured_programming_languages' not in relevant_columns:
relevant_columns.append('cured_programming_languages')
# Duplicate each row of df for each programming language
# mentioned in the response.
# We only include in df_lang the columns we are going to analyze later, so we
# don't duplicate innecesary information.
df_lang = df.cured_programming_languages\
.apply(pd.Series).stack()\
.reset_index(level=-1, drop=True).to_frame()\
.join(df[relevant_columns])\
.rename(columns={0: 'programming_language'})
# Horrible programming style! But a lot of data science code can be written with
# as concatenations of functions (pipelines), and there's no elegant way of
# doing that on Python.
df_lang[:5]
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="mz70t-z_Lbkh" outputId="7971069f-dc58-4dae-90a8-64fd154e7444"
language_count = df_lang.programming_language.value_counts()\
.reset_index()\
.rename(columns={'index': 'language', 'programming_language': 'frequency'})
language_count[:10]
# + id="VCv8nzkeLq1u"
#hasta aca nos dieron ellos
# + colab={"base_uri": "https://localhost:8080/", "height": 319} id="NGP12ZVALsHq" outputId="d4133377-0b56-4474-83a4-779eb0841fe9"
#barplot vamos a plotear los primeros 20
languages_20 = language_count[:19]
import seaborn as sns
sns.barplot(x="frequency", y="language", data=languages_20)
# + colab={"base_uri": "https://localhost:8080/", "height": 687} id="wCcfAELuL2IV" outputId="990e5018-3e9e-4608-c537-0052cffdb6f5"
#cuales son los lenguajes asociados a los mejores sueldos
plt.figure(figsize=(40,20))
sns.boxplot(x="salary_monthly_NETO", y="programming_language", data=df_lang)
#definitivamente hay q limpiar estos datos
# + id="s0uGTfEUQQx4"
#vamos a filtrar los lenguajes que estan orientados a front-end programming
front_end = ['basic','vb','vb5','vb6', 'vba','vbscript','visual','.net', 'net', 'asp', 'c#']
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="0uhVtZN-NKns" outputId="549e7d9e-955a-48f6-dec9-2a1d295497ba"
df_lang.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="5rFPQDZ6OSrn" outputId="0403403e-2f9d-4c78-ff58-4e2cdcc37426"
web=df_lang[df_lang['programming_language'].isin(['basic','vb','vb5','vb6', 'vba','vbscript','visual','.net', 'net', 'asp', 'c#'])]
web.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="MsZsBLssYWu0" outputId="1d1e5947-ebb3-4393-b813-76487d3771e0"
sns.boxplot(x="salary_monthly_NETO", y="programming_language", data=web)
# + id="XVhGl2_CabXm"
web_design=['react','elm','dart','ruby','kotlin','typescript','javascript', 'html', 'css', 'php']
# + id="x4EdN-sAdnsI"
web_d=df_lang[df_lang['programming_language'].isin(['react','elm','dart','ruby','kotlin','typescript','javascript', 'html', 'css', 'php'])]
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="EP8nTRZihsgf" outputId="700fc8e9-2b46-4451-a90d-3602089d1457"
web_d.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="8DJbxqL6jX8P" outputId="5a604773-d5f1-4bc3-c6cc-79d0bb2aef70"
sns.boxenplot(x="salary_monthly_NETO", data=web_d)
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="DhrNA4P1eCGY" outputId="65f6fa50-368f-4ca5-fbf4-854bcaa9e747"
sns.boxplot(x="salary_monthly_NETO", y="programming_language", data=web_d)
#existen muchos valores extremos
# + colab={"base_uri": "https://localhost:8080/", "height": 536} id="FMOVOxYTefN0" outputId="2b53e308-810f-43e7-8b69-80ed61ad97bd"
#veamos si mejora la visualización utilizando boxeplot en vez de boxplot
plt.figure(figsize=(12, 8))
sns.boxenplot(x="salary_monthly_NETO", y="programming_language", data=web_d)
# + [markdown] id="Xc-FRCSow22U"
# ### 1er paso: eliminar react y elm
# + colab={"base_uri": "https://localhost:8080/", "height": 536} id="cU6Y1doZlSnF" outputId="accc6404-a7f7-4263-84a5-8d16134a5659"
#1er paso: eliminar elm y react
web_d1=df_lang[df_lang['programming_language'].isin(['dart','ruby','kotlin','typescript','javascript', 'html', 'css', 'php'])]
plt.figure(figsize=(12, 8))
sns.boxenplot(x="salary_monthly_NETO", y="programming_language", data=web_d1)
# + [markdown] id="YPajkY4Ow7J9"
# ### 2do paso: eliminar valores extremos con PERCENTILES
# + colab={"base_uri": "https://localhost:8080/"} id="oxDaX3XqmCX_" outputId="e87e4b4c-659a-40db-95a9-5b9abbac55ef"
#2do paso: eliminar valores extremos
web_d1['salary_monthly_NETO'].quantile([.95, .98, .99, .995, .998])
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="dPYyYoJ7nk6F" outputId="f6dbb99a-24ab-46c0-9cf3-51c2c84ab078"
fig, axes = plt.subplots(figsize=(12, 12), nrows=3)
max_limit = web_d1['salary_monthly_NETO'].quantile(.98)
data = web_d1[web_d1['salary_monthly_NETO'] < max_limit]['salary_monthly_NETO']
sns.histplot(x=data, ax=axes[0])
sns.boxplot(x=data, ax=axes[1])
sns.boxenplot(x=data, ax=axes[2])
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="ky-NNKsIC-Q5" outputId="627e3efa-b138-42f4-80b6-be3311221fdf"
web_d1.describe()
# + [markdown] id="MS3Q1m8C61Vz"
# ### 2.1 Eliminar valores extremos usando RANGOS INTERCUARTILICOS
# + id="f-h9jwdYDPjQ"
def clean_outliers(dataset, column_name):
"""Returns dataset removing the outlier rows from column @column_name."""
interesting_col = dataset[column_name]
# Here we can remove the outliers from both ends, or even add more restrictions.
mask_outlier = (
numpy.abs(interesting_col - interesting_col.mean()) <= (2.5 * interesting_col.std()))
return dataset[mask_outlier]
# + id="Bc1rvCtLFSG9"
salary_col='salary_monthly_NETO'
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="usf059hZDcOa" outputId="73425629-5799-4a4b-ba46-3045b864d508"
fig, axes = plt.subplots(figsize=(12, 12), nrows=3)
max_limit = web_d1[salary_col].quantile(.98)
data = clean_outliers(web_d1, salary_col)[salary_col]
sns.histplot(x=data, ax=axes[0])
sns.boxplot(x=data, ax=axes[1])
sns.boxenplot(x=data, ax=axes[2])
# + [markdown] id="a2v812Vp7zO7"
# # Exploramos subconjunto de ciencia de datos
# + id="7fKrCTAz735-"
'python',
'r',
'dax', DATA ANALYSIS EXPRESSIONS
'regex'
'db',
'esql',
'mongodb',
'nosql',
'oracle',
'pl-sql','pl/sql', 'plsql',
'sql',
'bi',
'adabas'
# + id="wHVgTeH88RrT"
ds_data=df_lang[df_lang['programming_language'].isin(['python','r','mongodb','sql','nosql','bi','pl-sql','pl/sql', 'plsql', 'db', 'esql', 'oracle', 'adabas'])]
# + colab={"base_uri": "https://localhost:8080/", "height": 536} id="53jx24vA-ZG4" outputId="497b5a60-e496-42da-8e81-8568d263cf9a"
plt.figure(figsize=(12, 8))
sns.boxplot(x="salary_monthly_NETO", y="programming_language", data=ds_data)
# + colab={"base_uri": "https://localhost:8080/", "height": 536} id="3HA_yz7X-_nR" outputId="e5cb3dec-f639-4f1c-8e4b-8291232096db"
#eliminar los lenguajes q tienen muy pocos casos
#pasar a boxenplot
ds_data1=df_lang[df_lang['programming_language'].isin(['python','r','sql','pl-sql','pl/sql', 'plsql', 'esql', 'oracle'])]
plt.figure(figsize=(12, 8))
sns.boxenplot(x="salary_monthly_NETO", y="programming_language", data=ds_data1)
# + [markdown] id="OIlEfwNoF3ML"
# # Eliminar outliers con PERCENTILES
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="oFz9trAO_gBm" outputId="c0f88375-152e-488a-9f74-db2a3ec6d705"
fig, axes = plt.subplots(figsize=(12, 12), nrows=3)
max_limit = ds_data1['salary_monthly_NETO'].quantile(.98)
data = ds_data1[ds_data1['salary_monthly_NETO'] < max_limit]['salary_monthly_NETO']
sns.histplot(x=data, ax=axes[0])
sns.boxplot(x=data, ax=axes[1])
sns.boxenplot(x=data, ax=axes[2])
# + [markdown] id="JXXICPbWF7r1"
# # Eliminar outliers con RANGO INTERCUARTILICO
# + colab={"base_uri": "https://localhost:8080/", "height": 753} id="YUPenYR4BCqT" outputId="1de36688-1acf-4a52-f218-04536176fd7a"
fig, axes = plt.subplots(figsize=(12, 12), nrows=3)
max_limit = ds_data1[salary_col].quantile(.98)
data = clean_outliers(ds_data1, salary_col)[salary_col]
sns.histplot(x=data, ax=axes[0])
sns.boxplot(x=data, ax=axes[1])
sns.boxenplot(x=data, ax=axes[2])
| diplo_vis_punto1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Standard errors for calibrated parameters: Neoclassical Growth Model example
#
# *We are grateful to <NAME> for suggesting this example. Any errors are our own.*
#
#
# In this notebook we will work through the basic logic of [Cocci & Plagborg-Møller (2021)](https://scholar.princeton.edu/mikkelpm/calibration) in the context of calibrating a simple version of the Neoclassical Growth Model (NGM). Though the model is highly stylized, it helps provide intuition for our procedures. Please see our paper for other, arguably more realistic, empirical applications.
#
#
# ## Model
#
# We consider the simplest version of the NGM without population growth or technological growth. As explained in section 3.4 of [Dirk Krueger's lecture notes](https://perhuaman.files.wordpress.com/2014/06/macrotheory-dirk-krueger.pdf) (note the difference in notation), this model implies three key steady-state equations:
# 1. **Euler equation:** $r = \rho$, where $\rho$ is the household discount rate and $r$ is the real interest rate.
# 2. **Capital accumulation:** $\frac{I}{K} = \delta$, where $\delta$ is the capital depreciation rate and $I/K$ is the ratio of investment to capital stock.
# 3. **Rental rate of capital:** $\frac{K}{Y} = \frac{\alpha}{\rho+\delta}$, where $\alpha$ is the capital elasticity in the production function and $\frac{K}{Y}$ is the capital-output ratio.
#
# We want to use these equations to calibrate (i.e., estimate) the parameters $\rho$, $\delta$, and $\alpha$.
#
#
# ## Estimation
#
# We can measure the steady-state values of the variables on the left-hand side of the above equations by computing sample averages of the relevant time series over a long time span. Denote these sample averages by $\hat{r}$, $\widehat{\frac{I}{K}}$, and $\widehat{\frac{K}{Y}}$, respectively. We can then obtain natural method-of-moment estimates of the three parameters as follows:
# $$\hat{\rho} = \hat{r},\quad \hat{\delta}=\widehat{\frac{I}{K}},\quad \hat{\alpha}=\widehat{\frac{K}{Y}}\left(\hat{\rho}+\hat{\delta} \right).$$
#
#
# ## Standard errors
#
# The sample averages above are subject to statistical noise due to the finite time sample. This statistical noise obviously carries over to the estimated parameters. To gauge the extent of the noise, we seek to compute standard errors for the estimated parameters.
#
# The key ingredients into calculating standard errors for the parameters are the standard errors for the three sample averages. Denote these by $\hat{\sigma}\left(\hat{r}\right)$, $\hat{\sigma}\left(\widehat{\frac{I}{K}}\right)$, and $\hat{\sigma}\left(\widehat{\frac{K}{Y}}\right)$. To compute these standard errors, one would have to apply a formula that accounts for serial correlation in the data, such as the [Newey-West long-run variance estimator](https://www.stata.com/manuals16/tsnewey.pdf). Let's assume that we have already done this.
#
# It's immediate what the standard errors for $\hat{\rho}$ and $\hat{\delta}$ are: They simply equal $\hat{\sigma}\left(\hat{r}\right)$ and $\hat{\sigma}\left(\widehat{\frac{I}{K}}\right)$, respectively. However, the standard error for $\hat{\alpha}$ is not so obvious, as this estimator depends implicitly on all three sample averages:
# $$\hat{\alpha}=\widehat{\frac{K}{Y}}\left(\hat{r}+\widehat{\frac{I}{K}}\right) \approx \alpha + x_1\left(\hat{r}-r\right) + x_2\left(\widehat{\frac{I}{K}}-\frac{I}{K}\right) + x_3\left(\widehat{\frac{K}{Y}}-\frac{K}{Y}\right),$$
# where the last approximation is a [delta method](https://en.wikipedia.org/wiki/Delta_method) linearization with
# $$x_1=x_2=\frac{K}{Y},\quad x_3=r+\frac{I}{K}.$$
# Since $\hat{\alpha}$ is approximately a linear combination of several sample averages, computing its standard error requires not just the standard errors for the individual sample averages, but also their correlations.
#
#
# ## Limited-information inference
#
# If we observed annual data on the real interest rate, capital, investment, and output, it would not be too difficult to estimate the cross-correlations of the sample averages. This could again be done using the Newey-West estimator of the $3 \times 3$ long-run variance-covariance matrix. Yet, in practice we may face several potential complicating factors:
# 1. **Non-overlapping samples:** Perhaps we do not observe all time series over the same time span. This makes it difficult to apply the usual Newey-West formulas.
# 2. **Different data frequencies:** Perhaps the real interest rate series is obtained from daily yields on inflation-protected bonds, while the other time series are annual. This again complicates the econometric analysis.
# 3. **Finite-sample accuracy:** The Newey-West procedure is known to suffer from small-sample biases when the data exhibits strong time series dependence. Trying to exploit estimates of the correlations of the sample averages could therefore lead to distorted inference in realistic sample sizes.
# 4. **Non-public data:** Perhaps some of the sample averages were not computed by ourselves, but only obtained from other papers (say, a paper that imputes real interest rates by feeding bond yields through a structural model). Those other papers may report the standard errors for their respective individual moments, but not the correlations with other moments that our calibration relies on.
#
# A pragmatic limited-information approach would therefore be to give up on computing the precise standard error of $\hat{\alpha}$ and instead compute an *upper bound* on it. We seek an upper bound that depends only on the standard errors of the individual moments, not their correlations.
#
# The key to obtaining such a bound is the following inequality for random variables $X$ and $Y$:
# $$\begin{align*}
# \text{Std}(X+Y) &= \sqrt{\text{Var}(X+Y)} \\
# &= \sqrt{\text{Var}(X) + \text{Var}(Y)+2\text{Corr}(X,Y)\text{Std}(X)\text{Std}(Y)} \\
# &\leq \sqrt{\text{Var}(X) + \text{Var}(Y)+2\text{Std}(X)\text{Std}(Y)} \\
# &= \sqrt{(\text{Std}(X)+\text{Std}(Y))^2} \\
# &= \text{Std}(X)+\text{Std}(Y).
# \end{align*}$$
# Applying this logic to the earlier approximation for $\hat{\alpha}$, we get the bound (up to a small approximation error)
# $$\text{Std}(\hat{\alpha}) \leq |x_1|\text{Std}\left(\hat{r}\right) + |x_2|\text{Std}\left(\widehat{\frac{I}{K}}\right) + |x_3|\text{Std}\left(\widehat{\frac{K}{Y}}\right).$$
# We can therefore compute an upper bound for the standard error of $\hat{\alpha}$ as follows:
# $$\hat{\sigma}(\hat{\alpha}) \leq |\hat{x}_1|\hat{\sigma}\left(\hat{r}\right) + |\hat{x}_2|\hat{\sigma}\left(\widehat{\frac{I}{K}}\right) + |\hat{x}_3|\hat{\sigma}\left(\widehat{\frac{K}{Y}}\right),$$
# where
# $$\hat{x}_1=\hat{x}_2=\widehat{\frac{K}{Y}},\quad \hat{x}_3=\left(\hat{r}+\widehat{\frac{I}{K}} \right).$$
# Notice that this upper bound only depends on things that we know: the sample averages themselves and their individual standard errors (but not the correlations across moments).
#
# It's impossible to improve the bound without further knowledge of the correlation structure: The bound turns out to equal the actual standard error when the three sample averages are perfectly correlated with each other. This is proved in Lemma 1 in [our paper](https://scholar.princeton.edu/mikkelpm/calibration). For this reason, we refer to the standard error bound as the *worst-case standard error*.
#
#
# ## Numerical example
#
# Our software package makes it easy to calculate worst-case standard errors. As an illustration, suppose the sample averages (with standard errors in parentheses) equal
# $$\hat{r}=0.02\;(0.002), \quad \widehat{\frac{I}{K}}=0.08\;(0.01), \quad \widehat{\frac{K}{Y}} = 3\;(0.1).$$
# We define the model equations and data as follows. Let $\theta=(\rho,\delta,\alpha)$ and $\mu=(r,\frac{I}{K},\frac{K}{Y})$ denote the vectors of parameters and moments, respectively.
# +
import numpy as np
from stderr_calibration import MinDist # Minimum distance routines
# Define mapping from parameters to moments
h = lambda theta: np.array([theta[0],theta[1],theta[2]/(theta[0]+theta[1])])
# Define empirical moments and their s.e.
mu = np.array([0.02,0.08,3])
sigma = np.array([0.002,0.01,0.1])
# Define MinDist object used in later analysis
obj = MinDist(h,mu,moment_se=sigma)
# -
# We can now estimate the parameters and compute their worst-case standard errors:
param_estim = np.array([mu[0],mu[1],mu[2]*(mu[0]+mu[1])]) # Closed-form formula for estimates
res = obj.fit(param_estim=param_estim,eff=False)
print('Parameter estimates')
print(res['estim'])
print('Worst-case standard errors')
print(res['estim_se'])
# (Note: The derivatives required to compute $\hat{x}_1,\hat{x}_2,\hat{x}_3$ are produced under the hood by the software using finite differences. We could have also computed the parameter estimates $\hat{\rho},\hat{\delta},\hat{\alpha}$ numerically if we didn't have a closed-form formula. See our [other example](https://mikkelpm.github.io/stderr_calibration_python/example.html) for details.)
#
#
# ## Over-identification test
#
# The textbook NGM also implies that the steady-state labor share of income should equal $1-\alpha$. Suppose we measure the sample average of the labor share to be 0.6 with a standard error of 0.01. We wish to test the over-identifying restriction that the earlier estimate of $\hat{\alpha}$ is consistent with this moment. We can do this as follows.
# +
# Define expanded mapping that also includes over-identifying moment
h_expand = lambda theta: np.append(h(theta),1-theta[2])
# Define expanded empirical moments and standard errors
mu_expand = np.append(mu,0.6)
sigma_expand = np.append(sigma,0.01)
# Define new MinDist object and fit
obj_expand = MinDist(h_expand,mu_expand,moment_se=sigma_expand)
res_expand = obj_expand.fit(param_estim=param_estim,weight_mat=np.diag(np.array([1,1,1,0])),eff=False)
# Same parameter estimates as before (weight matrix indicates that these estimates do not use the fourth moment)
# Over-identification test
res_overid = obj_expand.overid(res_expand)
print('Error in matching non-targeted moment')
print(res_overid['moment_error'][3]) # The non-targeted moment is the fourth one
print('Standard error')
print(res_overid['moment_error_se'][3])
print('t-statistic')
print(res_overid['tstat'][3])
# -
# Since the absolute value of the t-statistic lies between 1.64 and 1.96, we can reject the validity of the model at the 10% significance level, but not at the 5% level.
#
# The over-identification test checks how different the estimate $\hat{\alpha}$ would have been if we had instead computed it as the sample average capital share of income $1-0.6=0.4$. Is the difference in parameter estimates between the two calibration strategies too large to be explained by statistical noise?
#
#
# ## Other features in the paper
#
# The above NGM example is very simple and stylized. In [our paper](https://scholar.princeton.edu/mikkelpm/calibration) we extend the basic ideas along various dimensions that are relevant for applied research. For example:
# - The matched moments need not be simple sample averages, but could be regression coefficients, quantiles, etc. The moments need not be related to steady-state quantities, but could involve essentially any feature of the available data.
# - The calibration (method-of-moments) estimator need not be available in closed form (usually one would obtain it by numerical optimization).
# - If some, but not all, of the correlations between the empirical moments are known, this can be exploited to sharpen inference.
# - If we have more moments to match than parameters to estimate, we can compute the optimal weighting of the moments that minimizes the worst-case standard errors of the parameters.
# - If we are interested in a function of the model parameters (such as a counterfactual quantity) rather than the parameters *per se*, we can compute worst-case standard errors for that function, too.
# - If we are interested in testing several parameter restrictions at once, a joint test is available that has valid size asymptotically.
# - All computational routines can handle models with relatively large numbers of parameters and moments.
| example_ngm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/a-forty-two/COG_GN22CDBDS001_MARCH_22/blob/main/MORE_PANDAS_Auto_MPG.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zeX0eDXRxdg6"
# # MPG Cars
# + [markdown] id="gZk-UJbYxdg-"
# ### Introduction:
#
# The following exercise utilizes data from [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Auto+MPG)
#
# ### Step 1. Import the necessary libraries
# + id="cz2NoCX3xdg_"
# + [markdown] id="6o7DIoiTxdhA"
# ### Step 2. Import the first dataset [cars1](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/05_Merge/Auto_MPG/cars1.csv) and [cars2](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/05_Merge/Auto_MPG/cars2.csv).
# + id="6eThsl46xjNY"
# https://raw.githubusercontent.com/a-forty-two/COG_GN22CDBDS001_MARCH_22/main/cars1.csv
# https://raw.githubusercontent.com/a-forty-two/COG_GN22CDBDS001_MARCH_22/main/cars2.csv
# + [markdown] id="9XdlJPXzxdhB"
# ### Step 3. Assign each to a variable called cars1 and cars2
# + id="GPz7DuOLxdhB"
# + [markdown] id="OXlLuT7KxdhC"
# ### Step 4. Oops, it seems our first dataset has some unnamed blank columns, fix cars1
# + id="txC6GcQ3xdhC"
# + [markdown] id="Cu0zwlMqxdhD"
# ### Step 5. What is the number of observations in each dataset?
# + id="RYQvsY8YxdhD"
# + [markdown] id="54ohJazRxdhE"
# ### Step 6. Join cars1 and cars2 into a single DataFrame called cars
# + id="DArHdd61xdhF"
# + [markdown] id="PVZT4IabxdhG"
# ### Step 7. Oops, there is a column missing, called owners. Create a random number Series from 15,000 to 73,000.
# + id="CTT_4ix-xdhG"
# + [markdown] id="bqbBmlxPxdhG"
# ### Step 8. Add the column owners to cars
# + id="dJxRs7tixdhH"
| notebooks/MORE_PANDAS_Auto_MPG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Quiz -Week 6A
# ##Q1.
#
# * The figure below shows two positive points (purple squares) and two negative points (green circles):
#
# 
#
# * That is, the training data set consists of:
#
# * (x1,y1) = ((5,4),+1)
# * (x2,y2) = ((8,3),+1)
# * (x3,y3) = ((7,2),-1)
# * (x4,y4) = ((3,3),-1)
#
# * Our goal is to find the __maximum-margin linear classifier__ for this data. In easy cases, the shortest line between a positive and negative point has a __perpendicular bisector that separates the points__. If so, the perpendicular bisector is surely the maximum-margin separator. Alas, in this case, the closest pair of positive and negative points, x2 and x3, have a perpendicular bisector that misclassifies x1 as negative, so that won't work.
#
# * The __next-best possibility__ is that we can find a pair of points on one side (i.e., either two positive or two negative points) such that __a line parallel to the line through these points__ is the maximum-margin separator. In these cases, the limit to how far from the two points the parallel line can get is determined by the closest (to the line between the two points) of the points on the other side. For our simple data set, this situation holds.
#
# * Consider all possibilities for boundaries of this type, and express the boundary as w.x+b=0, such that w.x+b≥1 for positive points x and w.x+b≤-1 for negative points x. Assuming that w = (w1,w2), identify in the list below the true statement about one of w1, w2, and b.
# +
import numpy as np
p1 = (5, 4)
p2 = (8, 3)
p3 = (7, 2)
p4 = (3, 3)
def calc_wb(p1, p2):
dx = ( p1[0] - p2[0] )
dy = ( p1[1] - p2[1] )
return ( ( float(dy) *2 / float(dy - dx), float(-dx)*2 / float(dy - dx) ),\
(dx*p2[1] - dy * p2[0])*2 / float(dy - dx) + 1) # b = dx*y1 - dy*x1
def cal_margin(w, b, pt):
return w[0] * pt[0] + w[1] * pt[1] + b
w, b = calc_wb(p1, p2)
print "w for p1, p2: " + str(w)
print "b for p1, p2: " + str(b)
print "==========================="
print cal_margin(w, b, p1)
print cal_margin(w, b, p2)
print cal_margin(w, b, p3)
print cal_margin(w, b, p4)
print
w, b = calc_wb(p4, p3)
print "w for p1, p2: " + str(w)
print "b for p1, p2: " + str(b)
print "==========================="
print cal_margin(w, b, p1)
print cal_margin(w, b, p2)
print cal_margin(w, b, p3)
print cal_margin(w, b, p4)
# -
# ##Q2.
#
#
# * Consider the following training set of 16 points. The eight purple squares are positive examples, and the eight green circles are negative examples.
#
# 
#
# * We propose to use the diagonal line with slope +1 and intercept +2 as a decision boundary, with positive examples above and negative examples below. However, like any linear boundary for this training set, some examples are misclassified. We can measure the goodness of the boundary by computing all the slack variables that exceed 0, and then using them in one of several objective functions. In this problem, we shall only concern ourselves with computing the slack variables, not an objective function.
#
# * To be specific, suppose the boundary is written in the form w.x+b=0, where w = (-1,1) and b = -2. Note that we can scale the three numbers involved as we wish, and so doing changes the margin around the boundary. However, we want to consider this specific boundary and margin.
#
# * Determine the slack for each of the 16 points. Then, identify the correct statement in the list below.
# +
w = (-1, 1)
b = -2
def cal_margin(w, b, pt):
return w[0] * pt[0] + w[1] * pt[1] + b
print cal_margin(w, b, (7, 10) )
print cal_margin(w, b, (7, 8) )
print cal_margin(w, b, (3, 4) )
print cal_margin(w, b, (3, 4) )
# -
# ##Q3.
#
# * Below we see a set of 20 points and a decision tree for classifying the points.
#
# 
# 
#
# * To be precise, the 20 points represent (Age,Salary) pairs of people who do or do not buy gold jewelry. Age (appreviated A in the decision tree) is the x-axis, and Salary (S in the tree) is the y-axis. Those that do are represented by gold points, and those that do not by green points. The 10 points of gold-jewelry buyers are:
#
# * (28,145), (38,115), (43,83), (50,130), (50,90), (50,60), (50,30), (55,118), (63,88), and (65,140).
#
# * The 10 points of those that do not buy gold jewelry are:
#
# * (23,40), (25,125), (29,97), (33,22), (35,63), (42,57), (44, 105), (55,63), (55,20), and (64,37).
#
# * Some of these points are correctly classified by the decision tree and some are not. Determine the classification of each point, and then indicate in the list below the point that is misclassified.
# +
def predict_by_tree(pt):
if pt[0] < 45:
if pt[1] < 110:
print "Doesn't buy"
else:
print "Buy"
else:
if pt[1] < 75:
print "Doesn't buy"
else:
print "Buy"
predict_by_tree((43, 83))
predict_by_tree((55, 118))
predict_by_tree((65, 140))
predict_by_tree((28, 145))
print "=============="
predict_by_tree((65, 140))
predict_by_tree((25, 125))
predict_by_tree((44, 105))
predict_by_tree((35, 63))
# -
# # Quiz Week 6A.
# ## Q1.
#
# * Using the matrix-vector multiplication described in Section 2.3.1, applied to the matrix and vector:
#
# <pre>
#
# | 1 2 3 4 | | 1 |
# | 5 6 7 8 | * | 2 |
# | 9 10 11 12 | | 3 |
# | 13 14 15 16 | | 4 |
#
# </pre>
#
# * Apply the Map function to this matrix and vector. Then, identify in the list below, one of the key-value pairs that are output of Map.
#
#
# ## Solution 1.
#
# The matrix-vector product is the vector x of length n, whose ith element xi is given by
#
# $$
# \begin{equation}
# x_i = \sum_{ j = 1}^n m_{ij} \cdot v_j
# \end{equation}
# $$
#
# * From each matrix element mij it produces the key-value pair ( i, $m_{ij} \cdot v_j$ ).
# * Thus, all terms of the sum that make up the component $x_i$ of the matrix-vector product will get the same key, i.
# +
import numpy as np
mat = np.array([ [1, 2, 3, 4],
[5, 6, 7, 8],
[9, 10,11,12],
[13,14,15,16] ])
vec = np.array([1, 2, 3, 4])
def key_val(mat, vec):
pair = dict()
for idx, row in enumerate(mat):
# pair[idx + 1] = np.dot(row, vec)
pair[idx + 1] = row * vec
return pair
print key_val(mat, vec)
# -
# ## Q2.
# * Suppose we use the algorithm of __Section 2.3.10__ to compute the __product of matrices M and N__. Let M have x rows and y columns, while N has y rows and z columns. As a function of x, y, and z, express the answers to the following questions:
# * The output of __the Map function__ has __how many different keys__? How many __key-value pairs__ are there with each key? How many __key-value pairs are there in all__?
# * The input to the Reduce function has how many keys? What is the length of the value (a list) associated with each key?
#
# ## Solution 2.
#
# 1. Diffrent keys output of Map function => x * z
# 2. Key-value pairs with each key => 2 * y
# 3. Key value pairs in all => 2 * x * y * z
# 4. Key Input to Reduce function =>
# 5. Length of value list = 2 * y
# ##Q3.
# * Suppose we use the __two-stage algorithm of Section 2.3.9__ to compute the product of matrices M and N. Let M have x rows and y columns, while N has y rows and z columns. As a function of x, y, and z, express the answers to the following questions:
# * The output of the first Map function has __how many different keys__? How many key-value pairs are there with each key? How many key-value pairs are there in all?
# * The output of the __first Reduce function has how many keys__? What is __the length of the value (a list) associated with each key__?
# * The output of the second Map function has how many different keys? How many key-value pairs are there with each key? How many key-value pairs are there in all?
# * Then, identify the true statement in the list below.
#
#
# ## Solution 3.
#
# 1. Different keys of first map => y
# 2. Different key-value pairs of each key => y * x + y * z
# 2. Key-value pairs in all => y * ( y * x + y * z)
# ## Q4.
#
# * Suppose we have the following relations:
# <pre>
#
# R S
#
# __A__ __B__ __B__ __C__
# 0 1 0 1
# 1 2 1 2
# 2 3 2 3
#
# </pre>
#
# * and we take their __natural join__ by the algorithm of Section 2.3.7. Apply the Map function to the tuples of these relations. Then, construct the elements that are input to the Reduce function. Identify one of these elements in the list below.
# * Map Results:
# <pre>
# (1, (R, 0))
# (2, (R, 1))
# (3, (R, 2))
# (0, (S, 1))
# (1, (S, 2))
# (2, (S, 3))
# </pre>
| week6/.ipynb_checkpoints/Quiz-Week6-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm, datasets
iris = datasets.load_iris()
X = iris.data[:, :2]
y = iris.target
C = 1.0
svc = svm.SVC(kernel='linear', C=1).fit(X,y)
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
h = (x_max / x_min)/100
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
plt.subplot(1, 1, 1)
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.8)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.Paired)
plt.xlabel('Sepal length')
plt.ylabel('Sepal width')
plt.xlim(xx.min(), xx.max())
plt.title('SVC with linear kernel')
plt.show()
| svm/svmimplementation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating adversarial images with Keras
# Presenting an adversarial mask/patch generator
# ## Random image transformations for use with the adversarial patch
from patch.transformations import test_random_transform
for i in range(2):
print("Test image with random transform: %s" % (i+1))
test_random_transform(min_scale=0.25, max_scale=2.0, max_rotation=22.5)
# ## Create adversarial patch for target label
# +
import json
from patch.model_state import ModelContainer
with open('config.json') as json_file:
config = json.load(json_file)
STEPS = config["patch_epochs"]
# custom_weights_path points to a pre-trained model on 10 classes of images
mc = ModelContainer("SimpleVGG16", custom_weights_path="cool_net.h5")
print("Training SimpleVGG16")
mc.reset_patch()
for i in range(STEPS):
loss = mc.train_step(scale=(0.1, 1.0))
if i % int(STEPS/10) == 0:
print("[%s] loss: %s" % (i, loss))
print("Done training!")
patch = mc.patch()
# +
from patch.constants import PATCH_SHAPE
from patch.image import show
from patch.model_state import circle_mask
def show_patch(patch):
circle = circle_mask(PATCH_SHAPE)
show(circle * patch + (1-circle))
show_patch(patch)
# -
# ## Test result by applying the patch to images in the validation set
# +
import matplotlib.pyplot as plt
import time
import math
import numpy as np
from patch.constants import BATCH_SIZE, TARGET_LABEL, LABELS
def label_to_name(label):
return LABELS[label]
def report(model, step=None, show_images=False, n=400, verbose=True, scale=(0.1, 1.0)):
"""Prints a report on how well the model is doing.
If you want to see multiple samples, pass a positive int to show_images
Model can be a ModelContainer instance, or a string. If it's a string, we
lookup that model name in the MultiModel
"""
start = time.time()
# n examples where target was in top 5
top_5 = 0
# n examples where target was top 1
wins = 0
# n examples in total
n_batches = int(math.ceil(float(n) / BATCH_SIZE))
total = BATCH_SIZE * n_batches
loss = 0
for b in range(n_batches):
if isinstance(model, str):
loss_per_example, probs, patched_imgs = M.inference_batch(model, scale=scale)
else:
loss_per_example, probs, patched_imgs = model.inference_batch(scale=scale)
loss += np.mean(loss_per_example)
for i in range(BATCH_SIZE):
top_labels = np.argsort(-probs[i])[:5]
if TARGET_LABEL in top_labels:
top_5 += 1
if top_labels[0] == TARGET_LABEL:
wins += 1
loss = loss / n_batches
top_5p = int(100 * float(top_5) / total)
winp = int(100 * float(wins) / total)
if step is not None:
r = 'Step: {} \t'.format(step)
else:
r = ''
r += 'LogLoss: {:.1f} \tWin Rate: {}%\t Top5: {}%\tn: {}'.format(math.log(loss), winp, top_5p, total)
if verbose:
print(r)
if show_images:
if show_images is True:
show_images = 1
_visualize_example(patched_imgs, probs, loss_per_example, show_images)
elapsed = time.time() - start
return {'logloss': math.log(loss), 'win': winp, 'top5': top_5p, 'time': elapsed, 'loss': loss}
def _visualize_example(patched_imgs, probs, loss_per_example, n_reports=1):
for i in range(n_reports):
show(patched_imgs[i])
predictions_str = ''
top_label_ids = np.argsort(-probs[i])[:5]
for label in top_label_ids:
p = probs[i][label]
name = label_to_name(label)
if len(name) > 30:
name = name[:27] + "..."
if name == "toaster":
predictions_str += "\033[1m"
name = name.ljust(30, " ")
predictions_str += "{} {:.2f}".format(name, p)
if name.startswith("toaster"):
predictions_str += "\033[0m"
predictions_str += "\n"
#predictions_str += "\033[1mLogLoss: {:.1f}\033[0m\n".format(math.log(loss_per_example[i]))
print(predictions_str)
report(mc, n=32, show_images=3, scale=0.4)
| adversarial_patch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Purpose
#
# To reduce the size of the model while maintaining maximal information content, I will cluster all of the channels of each contact map and take a respresentative channel from each cluster.
# A little magic
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# Imports
import h5py
from matplotlib import pyplot as plt
import numpy as np
from scipy.cluster.hierarchy import dendrogram, linkage
from scipy.sparse import coo_matrix
import sklearn.cluster
import torch
import itertools
from tqdm import *
# ## Handle reading the HDF5 file containing all of the data
# +
# Make a counts matrix to store co-clustering of
counts = np.zeros((12, 12))
with h5py.File('../../data/contacts.hdf5', 'r') as h5file:
keys = list(h5file.keys())
for entry in tqdm(keys):
atomtypes = h5file[entry]['memberships'][:]
memberships = h5file[entry]['memberships'][:]
target = torch.from_numpy(h5file[entry]['target'][:].astype(np.int64))
target[:, 2] = target[:, 2] - 3
n_res = np.max(memberships[:, 0]) + 1
val = torch.ones(len(target))
size = torch.Size([n_res, n_res, 12])
contact_map = torch.sparse.FloatTensor(target.t(), val, size)
contact_map = contact_map.to_dense().numpy().transpose((2, 0, 1)).reshape(12, -1)
n_clust = 4
clustering = sklearn.cluster.AgglomerativeClustering(n_clust).fit(contact_map)
labels = clustering.labels_
for i in range(n_clust):
channels = np.where(labels == i)[0]
for j in itertools.product(channels, repeat=2):
counts[int(j[0]), int(j[1])] += 1
# -
plt.imshow(counts / len(keys), cmap='gray')
| notebooks/exploratory_data_analysis/contact_map_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jeu du serpent - *Snake*
# Dans ce chapitre, nous allons mettre à profit la plupart des techniques que nous avons apprises jusqu'ici afin de construire une implémentation réaliste du jeu bien connu du [serpent](https://fr.wikipedia.org/wiki/Snake_(genre_de_jeu_vid%C3%A9o)) (*Snake game* [en](https://en.wikipedia.org/wiki/Snake_(video_game_genre))).
# <img src="attachment:snake.jpeg"/>
# Comme nous allons développer une application plus substantielle cette fois, de manière incrémentale, vous trouverez certainement plus commode d'utiliser un éditeur de texte (aussi simple que Notepad ou aussi sophistiqué qu'un IDE). À chaque étape, écrivez votre code dans l'éditeur et sauvegarder la nouvelle version dans un fichier texte - utiliser alors le bouton Load de l'ARMlite pour charger la dernière version du code.
# ## Un serpent qui bouge
# Nous allons commencer par écrire un simple boucle qui va déplacer la tête du serpent d'un pixel vers la droite à chaque itération. Dans cette version, le serpent va grandir continuellement, ce qui n'est pas ce que nous voulons pour le jeu final, mais nous améliorerons ce point un peu plus tard:
# ```
# // Définir les registres
# // R0-R2: réservés pour des usages temporaires
# // Variables globales
# // R3: position de la queue
# // R4: position de la tête
# // Constantes:
# MOV R10, #.PixelScreen
# MOV R11, #.green // couleur du serpent
#
# // Initialiser le jeu
# MOV R3, #1084 // pour la queue et ...
# MOV R4, #1088 // pour la tête à côté (4 octets = 1 mot = 1 pixel)
# STR R11, [R10+R3]
# STR R11, [R10+R4]
#
# // rafraîchissement ou mise à jour (update):
# maj:
# ADD R4, R4, #4 // incrémenter la position de la tête
# STR R11, [R10+R4] //dessiner la nouvelle tête
# B maj
# ```
# #### Exercice 39
#
# Charger le code et faites le fonctionner en mode pas à pas pendant quelques itérations de façon à bien comprendre ce qu'il fait.
#
# Ensuite faite-le tourner normalement. Quel problèmes observe-t-on immédiatement?
#
# **Sauvegarder votre programme**
# ___
# ## Controler la fréquence de rafraîchissement
# Parmi les problèmes trouvés à la première étape, le plus immédiat est la vitesse. La meilleur façon de la controler est d'utiliser les interruptions de l'horloge. Effectuer les modifications suivantes au code:
# ```
# // Définir les registres
# ...
#
# // Configurer les interruptions mais sans les activer pour l'instant
# MOV R0, #maj
# STR R0, .ClockISR
# MOV R0, #50
# STR R0, .ClockInterruptFrequency
#
# // Initialiser le jeu
# ...
# // ici, nous sommes prêt à gérer les interruptions
# MOV R0, #1
# STR R0, .InterruptRegister
#
# mainLoop: b mainLoop // pour occuper le processeur en attente d'interruption
#
# // rafraîchissement ou mise à jour (update)
# // piloté par l'horloge
# maj:
# ADD R4, R4, #4 // incrémenter la position de la tête
# STR R11, [R10+R4] // dessiner la nouvelle tête
# RFE
# ```
# *Notes*:
# - Nous configurons les interruptions d'horloge au début (par convention), mais les interruptions ne sont pas activées tant que le jeu n'a pas été initialisé - autrement la routine maj pourrait être appelée avant même qu'il n'y ait un serpent à déplacer.
# - **ClockInterruptFrequency** est défini en millisecondes. Ici, nous l'avons configuré pour qu'une interruption horloge ait lieu toutes les 50 millisecondes.
# - Comme **maj** est déclenchée par la survenue d'une interruption, elle se termine par `RFE` à la place de `B maj`.
# - **maj** ne sauvegarde aucun registre sur la pile car le seul registre qu'elle modifie est R4 qui est une variable globale (partagée par tout le code).
# #### Exercice 40
#
# Effectuer les changements indiqués et faites tourner le programme.
#
# Modifier **ClockInterruptFrequency** en augmentant ou en diminuant sa valeur (par rapport à 50 millisecondes)
#
# Comment pourrions-nous faire pour permettre à l'utilisateur de modifier la vitesse ce qui correspondrait à différents niveaux de difficulté? Seule une description est attendue, pas la peine de la coder pour l'instant.
#
# **Sauvegarder votre programme**
# ___
# ## Changer la direction avec les touches H, J, K, L
# Ensuite, nous allons permettre à l'utilisateur de modifier la direction du serpent en utilisant les touches H (gauche), J (bas), K (haut), L (droite). Nous pourrions vérifier si des touches ont été utilisées dans la boucle principale, mais une solution plus élégante est d'utiliser les interruptions clavier. Voici cette routine d'interruption:
# ```
# // routine d'interruption clavier
# // Si la touche est valide (H,J,K,L), la transférer dans R7
# clavier: PUSH {R0}
# LDR R0, .LastKey //lire la dernière touche enfoncée (mais sans attendre ...)
# CMP R0, #74 // touche J (haut)
# BEQ majTouche
# CMP R0, #75 // touche K (bas)
# BEQ majTouche
# CMP R0, #72 // touche H (gauche)
# BEQ majTouche
# CMP R0, #76 // touche L (droite)
# BEQ majTouche
# B .+2 // branchement inconditionnel deux instructions plus bas (POP...)
# majTouche: MOV R7, R0
# POP {R0}
# RFE
# ```
# *Notes*:
# - Cette nouvelle routine peut être placée après la routine **maj**.
# - `B .+2` signifie «effectuer un branchement deux instructions plus loin» (i.e. sauter par dessus la prochaîne instruction). Lorsqu'on effectue des petits sauts localement, cette syntaxe évite l'encombrement d'avoir à définir une nouvelle étiquette.
# - Ici, nous sauvegardons le registre R0 sur la pile car il peut être utilisé à d'autres fins lorsque la routine est appelée. En revanche, R7 a une portée globale.
# Et voici les modifications nécessaires à apporter en début de programme - l'une pour la définition d'une nouvelle variable globale, l'autre pour la configuration d'une interruption supplémentaire:
# ```
# // Définir les registres
# ...
# // R7: valeur ASCII de la dernière touche enfoncée
# ...
# //initialiser le jeu
# ...
# MOV R7, #76 //Au départ le serpent se déplace vers la droite
# // ici, nous sommes prêt à gérer les interruptions
# ...
#
# // Configurer les interruptions mais sans les activer pour l'instant
# ...
# MOV R0, #clavier
# STR R0, .KeyboardISR
# MOV R0, #1
# STR R0, .KeyboardMask
#
# ...
# ```
# À présent, nous devons exploiter la dernière touche utilisée (dans R7) afin de contrôler la direction du mouvement dans la routine **maj**:
# ```
# // rafraîchissement ou mise à jour (update):
# maj:
# CMP R7, #74
# BEQ bas
# CMP R7, #75
# BEQ haut
# CMP R7, #72
# BEQ gauche
# CMP R7, #76
# BEQ droite
# gauche: SUB R4, R4, #4
# B repeindre
# droite: ADD R4, R4, #4
# B repeindre
# haut: SUB R4, R4, #256 // -64*4=-256 -> monter d'une ligne
# B repeindre
# bas: ADD R4, R4, #256 // +64*4=256 -> descendre d'une ligne
# repeindre:
# STR R11, [R10+R4]
# RFE
# ```
# #### Exercice 41
#
# Ajouter ces portions de code à l'ancien (à la bonne place) puis faites tourner le programme. Vérifiez que tout fonctionne bien en déplaçant le serpent dans les quatre directions (si c'est difficile allonger la durée entre deux interruptions horloge) puis faites une capture d'écran montrant la zone d'affichage.
#
# Pourquoi le fait de sortir de l'écran par le haut ou par le bas produit une erreur alors que ce n'est pas le cas lorsqu'on sort par la gauche ou par la droite?
#
# **Sauvegarder votre programme**
# ____
# ## Heurter un côté fait perdre
# Dans le vrai jeu, si le serpent heurte l'un des quatre bords de la scène, on perd. Les bords haut et bas de la scène sont simples à traiter: nous pouvons tester si la valeur de R4 est inférieure à 0 ou supérieure à l'adresse du dernier pixel: 12284 (=64\*48\*4 - le 4 car un pixel fait 4 octets). Pour les bords gauche et droit, il va nous falloir regarder les 8 bits de poids faibles de l'index du pixel en utilisant un ET logique avec #255 (0x000000ff).
# - Si le serpent bouge vers la gauche, nous devons agir au moment où ces 8 bits de poids faibles prennent la valeur 0,
# - Si le serpent bouge vers la droite, nous devons agir au moment où ces 8 bits de poids faibles prennent la valeur #252 (car nous enlevons 4 à chaque fois, non 1).
#
# Voici les modifications à appliquer au coeur de la routine de mise à jour:
#
# ```
# ...
# gauche: AND R0, R4, #255
# CMP R0, #0
# BEQ perdu
# SUB R4, R4, #4
# B repeindre
# droite: AND R0, R4, #255
# CMP R0, #252
# BEQ perdu
# ADD R4, R4, #4
# B repeindre
# haut: SUB R4, R4, #256 // -64*4=-256 -> monter d'une ligne
# CMP R4, #0
# BLT perdu
# B repeindre
# bas: ADD R4, R4, #256 // +64*4=256 -> descendre d'une ligne
# MOV R0, #12284
# CMP R4, R0
# BGT perdu
# ...
# ```
# Nous devons aussi définir l'étiquette **perdu** qu'on peut mettre à la toute fin du code:
#
# ```
# ...
# perdu:
# MOV R0, #msgPerdu
# STR R0, .WriteString
# HALT
#
# msgPerdu: .ASCIZ " Game Over!\n"
# ```
# #### Exercice 42
#
# Modifier le code en incorporant ces changements et faites tourner le programmes quatre fois de façon à vous assurer que le jeu s'arrête correctement lorsqu'on touche un des quatres bords, mais que vous pouvez tout de même tourner juste avant.
#
# (En programmation on parle de (test des) *conditions aux limites* \[ *edge* ou *boundary conditions* \] - dans ce cas, les conditions correspondent à une limite «physique»!)
#
# Faites une copie d'écran montrant un virage juste avant la collision.
#
# **Sauvegarder votre programme**
# _____
# ## Le serpent ne peut pas se traverser lui même!
# Nous pouvons aussi ajouter la règle que le jeu est perdu si la tête du serpent repasse sur son corps. La manière la plus simple de faire cela est de vérifier si le tête s'apprête à bouger sur un pixel qui est déjà vert.
# ```
# ...
# repeindre:
# // commencer par vérifier que le serpent ne repasse pas sur lui-même
# LDR R0, [R10+R4] // lire sur l'écran le contenu du pixel où l'on s'apprête à mettre la tête
# CMP R0, R11 // ce pixel est-il vert ..
# BEQ perdu // si oui, c'est perdu
# STR R11, [R10+R4]
# RFE
# ```
# #### Exercice 43
#
# Effectuer ce changement au code et faites tourner le programme. Prendre une capture d'écran qui montre que le jeu se termine si le serpent se traverse.
#
# Que se passe-t-il si l'on inverse la direction du mouvement? (on va vers la gauche puis vers la droite par exemple)
#
# **Sauvegarder votre programme**
# ___________
# ## Créer une pomme à une position aléatoire
# À présent ajoutons une pomme, dans un position aléatoire à l'écran. Alors, à chaque fois que le serpent «mange» (passe sur) la pomme, nous ajouterons 1 au score du joueur et générerons une pomme à une nouvelle position aléatoire. Nous avons besoin de produire un nombre aléatoire dans l'intervalle 0-12280, et il doit être divisible par 4. Par défaut, la directive `.Random` de l'ARMlite produit un nombre aléatoire de 32-bits (un mot). Nous pouvons le combiner avec `AND` et le *masque de bits* `0b00000000'00000011'11111111'11111100` (soit 0x3ffc) pour le ramener à un nombre divisible par 4 (car les 2 bits de poids faibles sont à zéro) de l'intervalle 0-16380 ( car $3\cdot 16^3+15\cdot 16^2+15\cdot 16+12=16380$ ). Ensuite, il nous suffira de tester si le résultat est dans l'intervalle 0-12280 sinon nous relancerons «le dé» à nouveau.
#
# Premièrement, nous définirons une nouvelle constante pour la couleur de la pomme, ainsi qu'une nouvelle variable globale pour tenir le compte du nombre de pommes mangés:
#
# ```
# // Définir les registres
# ...
# // R8: Nombre de pommes mangées
# // Constantes:
# ...
# MOV R12, #.red // couleur d'une pomme
# ...
# //initialiser le jeu
# MOV R8, #0
# ...
# ```
# La sous-routine qui suit génére l'index d'un pixel au hasard dans l'intervalle requis et peint la pomme sur ce pixel. Elle vérifie aussi que nous ne plaçons pas la pomme sur le corps du serpent, c'est-à-dire sur un pixel déjà peint en vert:
#
# ```
# // Produit une pomme dans une position au hasard mais valide
# creerPomme: PUSH {R0, R1, LR}
# auHasard:
# LDR R1, .Random // obtenir un mot de 32-bits au hasard
# MOV R0, #0x3ffc
# AND R1, R1, R0 // conserver les bits de positions 2-14
# MOV R0, #12284
# CMP R1, R0
# BGT auHasard // relancer si on est pas dans l'intervalle correct
# LDR R0, [R10+R1]
# CMP R0, R11
# BEQ auHasard // relancer si on tombe sur le corps du serpent
# STR R12, [R10+R1] // dessiner la pomme
# POP {R0, R1, LR}
# RET
# ```
# *Notes*:
# - Cette nouvelle routine peut être placée à n'importe quel endroit pourvu qu'elle ne se chevauche pas avec une routine existante. L'auteur la place juste avant **perdu** et vous suggère d'en faire autant pour rester consistent avec ce livre.
# - Du fait qu'il s'agit d'une sous-routine qui sera donc appelée en utilisant l'instruction `BL` (ce n'est pas une routine d'interruption), elle se termine avec `RET`.
# - En fait, les instructions `PUSH` et `POP` ne sont pas strictement nécessaires pour ce code (car R0 et R1 ne sont utilisés que de façon locale et qu'il n'y pas d'appel à une autre sous-routine à l'intérieur de celle-ci qui risquerait de modifier le point de retour `LR`), mais nous les avons inclus pour suivre les «bonnes pratiques».
# Nous pouvons à présent appeler la sous-routine **creerPomme** dans la zone d'initialisation du jeu (mais toujours avant d'activer les interruptions):
#
# ```
# // Initialiser le jeu
# ...
# BL creerPomme
# // ici, nous sommes prêt à gérer les interruptions
# ...
# ```
#
# Ensuite, à la fin de la routine de mise à jour de l'écran - **maj** et à l'intérieur de la zone **repeindre**, nous pouvons tester si la pomme a été mangée et, dans ce cas, mettre à jour le score (nombre de pommes mangées) et créer une nouvelle pomme:
# ```
# repeindre:
# ...
# BEQ perdu // pour repère d'insertion!
# CMP R0, R12 // Vérifier si le pixel a la couleur d'une pomme ...
# BNE .+3 // ... et sauter directement à l'affichage de la tête dans ce cas
# ADD R8, R8, #1 // Incrémenter le «score»
# BL creerPomme
# STR R11, [R10+R4]
# RFE
# ```
# Enfin, nous pouvons aussi écrire le score dans la console à la fin du jeu en modifiant la séquence **perdu** (à la fin mais avant `HALT`):
#
# ```
# perdu:
# ...
# MOV R0, #score
# STR R0, .WriteString
# STR R8, .WriteUnsignedNum
# HALT
#
# msgPerdu: .ASCIZ " Game Over!\n"
# score: .ASCIZ "Votre score: "
# ```
# #### Exercice 44
#
# Effectuer tous les changements indiqués. Prendre une capture d'écran montrant le serpent et une pomme.
#
# Jouez à nouveau, en mangeant au moins deux pommes puis prendre à nouveau une capture d'écran de la scène finale et de votre score.
#
# **Sauvegarder votre programme**
# _____
# ## Le serpent ne grandit que s'il mange une pomme
# Notre jeu dans sa version courante n'est pas comme le vrai jeu du serpent: le serpent démarre avec deux segments de long, mais il ne grandit que lorsqu'il mange une pomme. Implémenter cela est un bon challenge, car il nous allons avoir besoin de suivre non seulement la tête et la queue du serpent mais aussi chaque partie du corps - de façon que la queue suive la tête selon le même chemin.
# Si nous sauvegardons chaque adresse mémoire qui correspond à une partie du corps du serpent dans une structure de données *file* \[ *queue data structure* \], alors à chaque fois qu'on déplace la tête nous pouvons «enfiler» sa nouvelle adresse (celle du pixel qui la représente), c'est-à-dire la placer en fin de file et, tout en déplaçant la queue du serpent en avant, nous pouvons «défiler» (sortir de la file) sa position située au début de la file (Il est probablement peu intuitif qu'on «enfile» en fin de file et qu'on «défile» au début de la file - techniquement la queue est au début de la file et la tête à la fin).
#
# Une **file** est une collection de mots successifs en mémoire. Nous pouvons définir une telle collection avec une étiquette à la toute fin de notre code:
#
# ```
# msgPerdu: .ASCIZ " Game Over!\n"
# score: .ASCIZ "Votre score: "
# .ALIGN 256
# corps: // la file des parties du corps du serpent; commence ici et se poursuit aux adresses suivantes
# ```
#
# *Note*: Nous avons besoin de `.ALIGN` afin de nous assurer que la file démarre sur le début d'un mot (contrainte d'alignement) - cela pourrait ne pas être le cas car les données qui précèdent sont rangées en mémoire comme des suites d'octets et non de mots. `.ALIGN 4` produirait l'effet voulu (démarrage à la prochaine adresse multiple de 4 libre), mais `.ALIGN 256` positionne la file au tout début de la prochaîne page mémoire, ce qui est plus commode pour la visualiser.
# Nous avons aussi besoin de définir deux registres supplémentaires, qui nous servirons de pointeurs sur le début et la fin de la file:
#
# ```
# // Définir les registres
# ...
# // R5: pointeur sur le début de la file (pour la queue)
# // R6: pointeur sur la fin de la file (pour la tête)
# ...
# ```
# Puis d'initialiser ces deux pointeurs:
#
# ```
# // Initialiser le jeu
# ...
# STR R11, [R10+R4] // REPERE
# MOV R5, #corps // adresse du début de la file (pour la queue)
# ADD R6, R5, #4 // adresse de fin de la file (pour la tête)
# STR R3, [R5] // R3 pointe sur l'adresse de la queue
# STR R4, [R6] // R4 pointe sur l'adresse de la tête
# MOV R7, #76 // Au départ le serpent se déplace vers la droite
# BL creerPomme // REPERE
# ...
# ```
# *Note*: Ici nous faisons un usage effectif de l'adressage indirect. Dans R5, par exemple, il n'y a pas la valeur (#.green) de la queue du serpent, mais l'adresse en mémoire du pixel qui représente la queue du serpent.
# À présent, à l'intérieur du code de **repeindre**, tout en peignant la tête en vert à sa nouvelle position, nous devons ajouter sa nouvelle position à la fin de la file «corps». Ensuite, sauf dans le cas où le serpent vient de manger une pomme, nous devons repeindre le pixel de queue en blanc à nouveau et défiler (sortir de la file) la référence à ce pixel. (si le serpent a mangé une pomme, nous sautons par dessus le code qui bouge la queue ce qui a pour effet de faire grandir le serpent d'un pixel.)
#
# ```
# repeindre:
# // commencer par vérifier que le serpent ne repasse pas sur lui-même
# LDR R0, [R10+R4] // lire sur l'écran le contenu du pixel où l'on s'apprête à mettre la tête
# CMP R0, R11 // ce pixel est-il vert ..
# BEQ perdu // si oui, c'est perdu
# ADD R6, R6, #4 // Incrémenter le pointeur de fin de file d'un mot
# STR R4, [R6] // enfiler la nouvelle position de la tête (en fin de file)
# CMP R0, R12 // Vérifier si le pixel a la couleur d'une pomme ...
# BEQ manger
# MOV R0, #.white
# STR R0, [R10+R3] // blanchir l'ancienne position de la queue
# ADD R5, R5, #4 // Incrémenter le pointeur de début de file (défiler)
# LDR R3, [R5] // mettre à jour le pointeur de la queue
# // BNE .+3 à supprimer
# B .+3 // ... et sauter directement à l'affichage de la tête dans ce cas
# manger: ADD R8, R8, #1 // Incrémenter le «score»
# BL creerPomme
# STR R11, [R10+R4]
# RFE
# ```
# #### Exercice 45
#
# Incorporer ces changements dans votre code. Vous devriez voir que votre serpent ne change pas de longueur tant qu'il n'a pas manger de pomme.
#
# Faites grandir votre serpent d'au moins deux pixels (donc deux pommes à manger) puis faites une copie d'écran montrant la scène finale ainsi que votre score.
#
# Lorsque le programme est à l'arrêt, aller à la page mémoire 002 et faites une capture d'écran de cette page mémoire.
#
# Décrivez avec vos propres mots ce que détiennent ces mots mémoire.
# ____
# Bien que notre programme fonctionne correctement du point de vue de l'utilisateur, ce n'est pas une bonne implémentation.
#
# Le problème est qu'à chaque mouvement du serpent, la partie active de la file (les positions mémoire entre le début et la fin de la file) ne cesse d'avancer dans la mémoire, en laissant des données «mortes» derrière elle. Il se peut même que la file finisse par atteindre la fin de la mémoire quand bien même la longueur effective du serpent resterait assez petite.
#
# L'ARMlite dispose d'1 Mo de mémoire, ainsi, même si le serpent avance de 10 pixels par seconde et en supposant que vous fassiez exprès de louper la pomme afin de faire durer le jeu le plus longtemps possible, il vous faudrait plusieurs heures avant d'avoir saturé complètement la mémoire:
#
# (1s consomme $10\times 4=40$ octets soit $60\times 60 \times 40 < 2\times 10^5$ octets par heure donc plus de 5 heures pour 1Mo$\approx 10^6$o)
#
# Mais, d'un programme qui fonctionne de cette manière, on dit qu'il a des «fuites mémoires» \[ *memory leak* \] et aucun programmeur qui se respecte ne laisserait son code dans cet état.
# ## Implémentation d'une file circulaire \[ *circular queue* \]
# La solution, comme vous le savez peut-être si vous avez déjà étudié les structures de données, est de fixer la longueur maximale de la file et de laisser les pointeurs revenir en début de file lorsqu'ils atteignent son extrémité droite.
#
# Quelle taille choisir pour notre file? Étant donné que, dans cette résolution (64x32), l'écran ne dispose que de 3072 pixels, même un excellent joueur ne pourra rendre le serpent plus long que cela. Nous réserverons donc 3072 mots en mémoire pour la file avec ce changement:
#
# ```
# .ALIGN 256
# corps: .BLOCK 3072 //on réserve un bloc de 3072 mots pour notre file
# limite: //adresse du mot juste après la fin de la file
# ```
# Nous modifions à présent **repeindre**:
#
# ```
# repeindre:
# ...
# BEQ perdu // si oui, c'est perdu
# ADD R6, R6, #4 // Incrémenter le pointeur de fin de file d'un mot
# CMP R6, #limit // le pointeur a-t-il atteint la fin de file? ...
# BLT .+2
# MOV R6, #corps // ...si oui, le remettre au début
# STR R4, [R6] // enfiler la nouvelle position de la tête (en fin de file)
# ...
# ADD R5, R5, #4 // Incrémenter le pointeur de début de file (défiler)
# CMP R5, #limit // similaire au pointeur de tête pour la queue
# BLT .+2
# MOV R5, #corps
# LDR R3, [R5] // mettre à jour le pointeur de la queue
# ...
# ```
# #### Exercice 46
#
# Mettre en oeuvre ces modifications et, après avoir vérifier qu'il se charge correctement, **sauvegarder le programme**.
#
# Ensuite, passer en mode «player» en ajoutant la query string
#
# https://peterhigginson.co.uk/ARMlite/?profile=player
#
# et amusez-vous bien!
# ____
# ## Améliorations possibles
# Si vous avez un peu de temps pour améliorer le jeu, voici quelques suggestions, mais vous pouvez en trouver d'autres par vous-même:
# - À la fin du jeu, donner à l'utilisateur la possibilité de rejouer sans avoir à arrêter puis redémarrer le programme.
# - Faire en sorte que la position de démarrage et le mouvement initial du serpent soit aléatoire.
# - Donner au joueur la possibilité, au démarrage du programme, de modifier la vitese du jeu en saisissant un nombre lequel sera «décodé» pour régler de façon approprié l'intervalle de rafraichissement **ClockInterruptFrequency**.
# - Augmenter la fréquence d'horlorge au fur et à mesure que des pommes sont mangées.
# - Créer un ou plusieurs obstacles, aléatoirement, que le serpent ne doit pas toucher.
# - Ignorer les mouvements contraires plutôt que de faire mourir le joueur instantanément.
| 06_machine_physique/Initiation_au_langage_d_assemblage/Partie2/08_jeu_du_serpent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Predicting the Success of Cyber-Related Terrorist Attacks:
# + [markdown] slideshow={"slide_type": "subslide"}
# This dataset is from Global Terrorism Database curated by the University of Maryland:
#
# National Consortium for the Study of Terrorism and Responses to Terrorism (START). (2018). Global Terrorism Database [Data file]. Retrieved from https://www.start.umd.edu/gtd.
#
# It is a rich dataset with numerous variables and plenty of opportunities for analysis. In this project, we will focus on predicting the 'success' of attacks which are related to cyber-events or have consequences for cyber-infrastructure (we will describe these characteristics in further detail below.
#
# To begin, we will take a look at how this study classifies a successful terrorist attack and distinguishes it from an unsuccessful attack. Below, is their description from pages 11 and 26 of their code-book (__[GTD Global Terrorism Database. Codebook: Inclusion Criteria and Variables](http://www.start.umd.edu/gtd/downloads/Codebook.pdf)__):
#
# > "The GTD does not include plots or conspiracies that are not enacted, or at least attempted. For
# an event to be included in the GTD, the attackers must be “out the door,” en route to execute
# the attack. Planning, reconnaissance, and acquiring supplies do not meet this threshold.
# The GTD does include attacks that were attempted but ultimately unsuccessful. The
# circumstances vary depending on tactics (for details see the success variable, below). However,
# in general if a bomb is planted but fails to detonate; if an arsonist is intercepted by authorities
# before igniting a fire; or, if an assassin attempts and fails to kill his or her intended target, the
# attack is considered for inclusion in the GTD, and marked success=0." P. 11
#
# > "Success of a terrorist strike is defined according to the tangible effects of the attack.
# Success is not judged in terms of the larger goals of the perpetrators. For example, a
# bomb that exploded in a building would be counted as a success even if it did not
# succeed in bringing the building down or inducing government repression.
# The definition of a successful attack depends on the type of attack. Essentially, the
# key question is whether or not the attack type took place. If a case has multiple
# attack types, it is successful if any of the attack types are successful, with the
# exception of assassinations, which are only successful if the intended target is killed.
# 1 = "Yes" The incident was successful.
# 0 = "No" The incident was not successful." P. 26
#
# Thus, our focus below will be on using the data collected to build a model which will successfully predict the success of a terror attack. Below, we will begin importing and working with our data, and explanations and analysis will follow when pertinent.
# + slideshow={"slide_type": "skip"}
import pandas as pd
import numpy as np
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn as skl
import gc
import sys
import re
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# + [markdown] slideshow={"slide_type": "slide"}
# ### 1. Importing, Cleaning and General Overview:
#
# #### A. Importing Data.
# + slideshow={"slide_type": "fragment"}
df = pd.read_excel('globalterrorismdb_0718dist.xlsx',
usecols = 'A, I, K, M, S:W, AA:AB, AD, AJ, AL, AM, AN, AP, BG, BM:BN, BQ, CE, CG, DA',
dtype = {'summary':str, 'motive':str})
# + slideshow={"slide_type": "skip"}
# Renaming our columns for usability:
cyber_data = pd.DataFrame(df.rename({'eventid':'event_id', 'doubtterr':'doubt', 'attacktype1_txt':'attack_1txt',
'targtype1_txt':'target_1txt', 'targsubtype1':'sub_target',
'targsubtype1_txt':'sub_targettxt', 'target1':'specific_target',
'natlty1_txt':'victim_nationalitytxt', 'gname':'group_name',
'guncertain1':'group_attrib_crtainty', 'individual':'unaffil_individ',
'weaptype1':'weapon', 'weaptype1_txt':'weapontxt',
'weapsubtype1':'sub_weapon', 'weapsubtype1_txt':'sub_weapontxt'}, axis = 1))
# + slideshow={"slide_type": "skip"}
# Memory Mitigation:
del df
gc.collect()
# + slideshow={"slide_type": "skip"}
cyber_shape_1 = cyber_data.shape
cyber_shape_1
# + [markdown] slideshow={"slide_type": "slide"}
# #### B. Selecting Rows Specific to Cyber-Related Terrorism:
#
# In order to filter the dataset and focus our inquiry on cyber-related events, we will use the following regex statements. This statement attempts to focus on communication platforms (cellular, internet, radio) and its infrastructure (to a certain extent).
# + slideshow={"slide_type": "subslide"}
# Regex Filter which fills na in this column with the value: 'Unknown'
specific = cyber_data.specific_target.str.contains(r'(internet|cell+|radio|communic+|emai+|cyb+|web|hac+)',
na = 'Unknown', flags = re.IGNORECASE)
specific_true = specific.loc[specific == True].keys()
specific_unknown = specific.loc[specific == 'Unknown'].keys()
# same for motive column.
motive = cyber_data.motive.str.contains(r'(internet|cell+|radio|comm+|infor+|emai+|cyb+|web|hac+)',
na = 'Unknown', flags = re.IGNORECASE)
motive_true = motive.loc[motive == True].keys()
# same for summary column.
summary = cyber_data.summary.str.contains(r'(internet|cell+|radio|comm+|infor+|emai+|cyb+|web|hac+)',
na = 'Unknown', flags = re.IGNORECASE)
summary_true = summary.loc[summary == True].keys()
# + slideshow={"slide_type": "subslide"}
# Combining the above results into a dataframe and looking at the shape:
cyber_data = cyber_data.loc[(cyber_data.index.isin(specific_true)) | (cyber_data.index.isin(motive_true)) |
(cyber_data.index.isin(summary_true)) | (cyber_data.index.isin(specific_unknown))]
cyber_data.shape
# + [markdown] slideshow={"slide_type": "slide"}
# #### C. Splitting Our Train/Test Data:
#
# Below, we dropped some specific columns in dataframe in order to make future processing more efficient. The 'event_id' column wasn't numerically significant and created problems when getting dummies was necessary. The same occured with 'summary' and 'motive.' These last two columns, however, will come in handy when considering avenues for further research, which we will discuss below.
# + slideshow={"slide_type": "fragment"}
from sklearn.model_selection import cross_val_score, train_test_split
# Defining our Input and Output data:
# (Cleaning afterwards to prevent leakage)
X = cyber_data.drop(['event_id', 'success', 'summary', 'motive'], axis = 1)
Y = pd.DataFrame(cyber_data['success'])
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = .25)
X_train_start_index = X_train.index
X_test_start_index = X_test.index
# + slideshow={"slide_type": "skip"}
print(X_train.shape)
print(X_test.shape)
# + slideshow={"slide_type": "skip"}
# Memory Mitigation (Resource cited below):
ipython_vars = ['In', 'Out', 'exit', 'quit', 'get_ipython', 'ipython_vars']
cleaner = sorted([(x, sys.getsizeof(globals().get(x))) for x in dir() if not x.startswith('_') and x not in sys.modules and x not in ipython_vars],
key=lambda x: x[1], reverse=True)
# + slideshow={"slide_type": "skip"}
del X, Y, cleaner, specific, motive, summary, summary_true, motive_true, specific_true, specific_unknown
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# #### D. Checking Nulls:
# + slideshow={"slide_type": "skip"}
train_nulls = pd.DataFrame(X_train.isna().sum())
train_nulls = train_nulls.loc[train_nulls[0] != 0]
test_nulls = pd.DataFrame(X_test.isna().sum())
test_nulls = test_nulls.loc[test_nulls[0] != 0]
# + [markdown] slideshow={"slide_type": "fragment"} variables={"test_nulls": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>city</th>\n <td>7</td>\n </tr>\n <tr>\n <th>sub_targettxt</th>\n <td>184</td>\n </tr>\n <tr>\n <th>corp1</th>\n <td>553</td>\n </tr>\n <tr>\n <th>specific_target</th>\n <td>154</td>\n </tr>\n <tr>\n <th>victim_nationalitytxt</th>\n <td>27</td>\n </tr>\n <tr>\n <th>group_attrib_crtainty</th>\n <td>15</td>\n </tr>\n <tr>\n <th>sub_weapontxt</th>\n <td>359</td>\n </tr>\n </tbody>\n</table>\n</div>", "train_nulls": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>city</th>\n <td>25</td>\n </tr>\n <tr>\n <th>sub_targettxt</th>\n <td>603</td>\n </tr>\n <tr>\n <th>corp1</th>\n <td>1696</td>\n </tr>\n <tr>\n <th>specific_target</th>\n <td>482</td>\n </tr>\n <tr>\n <th>victim_nationalitytxt</th>\n <td>100</td>\n </tr>\n <tr>\n <th>group_attrib_crtainty</th>\n <td>34</td>\n </tr>\n <tr>\n <th>sub_weapontxt</th>\n <td>1036</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Our nulls here are concentrated in a few specific columns - and only one is in a numeric column.
# <head>
# <table>
# <tr>
# <td> {{train_nulls}} </td>
# <td> {{test_nulls}} </td>
# </tr>
# </table>
# </head>
#
# As such, we will do the following:
#
# 1. Fill na's in text columns with 'Unknown'
# 2. Drop the na's in group_attrib_crtainty just prior to modeling since it is a low-enough total that we can drop it without significant consequences for our analysis.
# + [markdown] slideshow={"slide_type": "slide"}
# #### E. Isolating Columns:
#
# We want to be sure we are only using the numeric columns that are significant (i.e. binary or numerically related to the values they contain) and not arbitrary categoricals (such as using numbers to classify one instance over another). Initially, we imported more of the dataset which included different types of data-types. After running the notebook as it was closer to its final form, it took considerably longer to load, at which point we dropped most of these columns from our initial read-in of the data. We account for a few lingering columns below:
# + slideshow={"slide_type": "fragment"}
cyber_train_X = X_train[['country_txt', 'region_txt', 'city', 'crit1', 'crit2', 'crit3', 'doubt',
'suicide', 'attack_1txt', 'target_1txt', 'sub_targettxt', 'corp1',
'specific_target', 'victim_nationalitytxt', 'group_name', 'unaffil_individ', 'weapontxt',
'sub_weapontxt']]
cyber_train_X = cyber_train_X.fillna('Unknown')
cyber_train_X['group_attrib_crtainty'] = X_train[['group_attrib_crtainty']]
# Making sure input and output dataframes still have the same amount of rows:
cyber_train_Y = Y_train.iloc[Y_train.index == cyber_train_X.index]
# + slideshow={"slide_type": "skip"}
# Applying the same process above to our test data:
cyber_test_X = X_test[['country_txt', 'region_txt', 'city', 'crit1', 'crit2', 'crit3', 'doubt',
'suicide', 'attack_1txt', 'target_1txt', 'sub_targettxt', 'corp1',
'specific_target', 'victim_nationalitytxt', 'group_name', 'unaffil_individ', 'weapontxt',
'sub_weapontxt']]
cyber_test_X.fillna('Unknown')
cyber_test_X['group_attrib_crtainty'] = X_test[['group_attrib_crtainty']]
# Making sure input and output dataframes still have the same amount of rows:
cyber_test_Y = Y_test.iloc[Y_test.index == cyber_test_X.index]
# + slideshow={"slide_type": "skip"}
del X_train, X_test, Y_train, Y_test
# + slideshow={"slide_type": "skip"}
print(cyber_train_X.shape)
print(cyber_test_X.shape)
# + [markdown] slideshow={"slide_type": "slide"}
# #### F. Applying Filters to Text Columns.
#
# The cells below are an attempt to consolidate (or group) some of the values together with a few certain columns we will be focusing on. Given that these columns have a considerable number of unique values, when getting dummies later, it will greatly increase the size of our feature set. While an increased feature-set is not necessarily a bad thing, preventing the size from becoming too large will aid in our explanatory power later on. In other words, our feature set will have a comprehensible size, allowing us to explain the characteristics around a successful attack. Otherwise, we run the risk of a feature set which is too large for a human to understand and too many features - making it difficult to see which columns/characteristics are significant for our analysis.
#
# Below, we focus mainly on the types of weapons, the named terrorist groups and the cities attacked. We also consolidated some of the sub_targets into larger groups. After running our preliminary models, we found that targets and sub_targets were significant to our models, so we conducted some further feature engineering afterwards, which we will discuss later.
# + [markdown] slideshow={"slide_type": "subslide"}
# There were a number of resources which were helpful in this grouping process, which we will lay out here:
#
# Regarding the 'sub_target' column (which provides further details regarding the primary target) we mainly referred to the codebook accompanying the study (as mentioned above: (__[GTD Global Terrorism Database. Codebook: Inclusion Criteria and Variables](http://www.start.umd.edu/gtd/downloads/Codebook.pdf)__)). The grouping below was rather simple - primarily placing a few types of sub-targets together when it would not negatively impact our explanatory capabilites down the road.
# + [markdown] slideshow={"slide_type": "subslide"}
# The 'city' and 'group_name' columns were more technical and involved some outside research. The groupings below are aimed at linking cities together if they fall within an ideological, religious, environmental or political umbrella in which certain terrorist groups are interested. For example, some middle-eastern cities are considered to lean more 'Sunni' as opposed to 'Shia', while others are 'split.' Some South American cities lie within an area experiencing heavy gang activity (such as the Northern Triangle between Mexico and Panama). Our goal with these groupings was to combine cities whenever they had a common interest factor for terrorist events, in the hopes that it would consolidate their correlation and aid in our predictive models.
#
# These groupings, however, can be improved upon with further in-depth research. Our time with this project was somewhat limited and there are a handful of regions we were unable to group together or research. Additionally, our expertise in global terrorism is slight in comparison to those working in the field, which would benefit from an expert team member when creating these filters. That said, it would be highly interesting to continue improving these classifications, especially given the wealth of information and databases made available by respected international research organizations, which we will list here:
# + [markdown] slideshow={"slide_type": "subslide"}
# Some cities and groups were classified according to their religious leanings:
#
# - Some Middle-Eastern cities were grouped under a Sunni/Shia or Sunni/Shia Split category. These resources were helpful in ascertaining where a city fell with respect to these religious tendencies:
# - A New York Times article by <NAME>, <NAME> and <NAME> on January 5th, 2016, entitled: __[Behind Stark Political Divisions, a More
# Complex Map of Sunnis and Shiites.](https://www.nytimes.com/interactive/2016/01/04/world/middleeast/sunni-shiite-map-middle-east-iran-saudi-arabia.html)__
# - A blog post by <NAME> on August 14th, 2014, entitled: __[Carte religieuse de l’Iran et ses voisins](http://zakhor-online.com/?attachment_id=7932)__.
# - <NAME> wrote a great article for Vox on March 26th, 2015 which has a lot of great maps for reference: __[40 maps that explain the Middle East](https://www.vox.com/a/maps-explain-the-middle-east)__
# The Gulf 2000 Project has a plethora of maps and resources regarding issues in the Middle-East, and specifically this page by Dr. <NAME> entitled __[Atlas of the Islamic World and Vicinity
# (Infographs, Maps and Statistics Collection).](http://gulf2000.columbia.edu/maps.shtml)__ Some other maps we used from this site were: - __[A map of West Africa](http://gulf2000.columbia.edu/images/maps/West_Africa_Religion_lg.png)__
# - __[A map of Libya](http://gulf2000.columbia.edu/images/maps/Libya_Religion_Western_Sector_lg.png)__
# - __[A Shia territories map](http://gulf2000.columbia.edu/images/maps/ShiasReligionCore_lg.png)__
#
# The Crisis Group also has a number of useful resources and articles regarding these topics. Specifically regarding Colombia and South America, we referenced and article entitled __[Colombia’s Armed Groups Battle for the Spoils of Peace](https://www.crisisgroup.org/latin-america-caribbean/andes/colombia/63-colombias-armed-groups-battle-spoils-peace)__ from Report 63 / Latin America & Caribbean 19 OCTOBER 2017. There was an interesting map by <NAME> (International Crisis Group 2017), entitled __[Map of Armed Groups and Coca Crops in Colombia, 2017.](https://www.crisisgroup.org/latin-america-caribbean/andes/colombia/63-colombias-armed-groups-battle-spoils-peace#map-5700-8)__.
# + [markdown] slideshow={"slide_type": "subslide"}
# We'll also take this opportunity to list out further databases and resources we used for this project:
#
# For 'group_names' we referenced START's resources again (as they have a wealth of resources surrounding the database we used and the elements it describes. Here, they list the names of __['Big, Allied and Dangerous' terrorist groups,](http://www.start.umd.edu/baad/database)___ which we used as the basis for a majority of our classification.
#
# The __[SATP website](http://www.satp.org/conflict-maps/bangladesh)__ was instrumental in gaining insight into Asian conflicts. Within this website, we referenced a specific article regarding the
# __[Nepalise Maoist conflicts](http://www.satp.org/terrorist-profile/nepal/communist-party-of-nepal-maoist)__, which described their objectives, operating areas, leaders, etc. Towards the bottom of the page, it includes a ranking by the Nepalese Home Ministry, of areas in Nepal according to their sensitivity to these issues. We then used this __[map](https://reliefweb.int/map/nepal/nepal-regions-zones-and-districts-cities-april-2015)__ from ReliefWeb to help us locate smaller cities in pertinent areas.
# + [markdown] slideshow={"slide_type": "subslide"}
# South America and the Northern Triangle:
#
# Again, there are a number of databases and materials available from top-level international organizations. Regarding the Northern Triangle (namely Guatemala, Honduras, El Salvador) we referenced the following articles:
# - __[Central America’s Violent Northern Triangle](https://www.cfr.org/backgrounder/central-americas-violent-northern-triangle)__By <NAME> and <NAME> (updated June 26th, 2018)
# - The Insight Crime Organiztion has a main page for each country involved in frequent conflicts__[such as this one regarding El Salvador](https://www.insightcrime.org/el-salvador-organized-crime-news/)__
# - Relief Web also had a variety of resources we used here, specifically a __[main search page](https://reliefweb.int/country/slv)__ for each country that leads to reports, infographics and the like.
# - The __[Humanitarian Reponse Group](https://www.humanitarianresponse.info/en/infographics)__ also has great resources and infographics.
# - The __[United Nations Regional Information Centre](https://www.unric.org/en/databases/26912-refugees-humanitarian-affairs-and-migration)__ points to a number of outside resources depending on what one is looking for.
# - In refrencing hotspots of violence in Guatemala, we used Relief Web's summary on __[Humanitarian Needs Overview for Guatemala in 2017](https://reliefweb.int/sites/reliefweb.int/files/resources/20180315_SUMMARY_HNO%20GT_ENG.pdf)__
# - Regarding the same of El Salvador, we used Relief Web's __[Map of El Salvador.](https://reliefweb.int/sites/reliefweb.int/files/resources/20180405%20monthly%20humanitarian%20snapshot%20-%20ENG.pdf)__
# - For Honduras we referred to an article entitled __['Northern Triangle is World's Extortion Hotspot'](https://www.insightcrime.org/news/brief/northern-triangle-world-extortion-hotspot/)__ by <NAME> and <NAME> on JULY 1st, 2015. This article has a map that was helpful in locating our cities.
# - Regarding Columbia, we frequently referred to the __[Crisis Group's resources.](https://www.crisisgroup.org/)__
# There were two articles in particular we referred to here. The first was entitled 'Colombia’s Armed Groups Battle for the Spoils of Peace' (which we already referenced above). The second was a map entitled __['Map of ELN Presence in Colombia in 2012 and 2018, and Expansion between 2012 and 2018'](https://www.crisisgroup.org/latin-america-caribbean/andes/colombia/68-missing-peace-colombias-new-government-and-last-guerrillas#map-6189-1)__ within an article named __['The Missing Peace: Colombia’s New Government and Last Guerrillas'](https://www.crisisgroup.org/latin-america-caribbean/andes/colombia/68-missing-peace-colombias-new-government-and-last-guerrillas)__ from their Report 68 / Latin America & Caribbean on July 12th, 2018.
# + [markdown] slideshow={"slide_type": "subslide"}
# Other General References include:
#
# - __[Homeland Security Digital Library](https://www.hsdl.org/?collection&id=2167)__
# - __[U Mass Lowell](https://www.uml.edu/Research/CTSS/Online-Resources.aspx)__ Has a great page with links to further resources.
# - The Council on Foreign Relations has a page entitled __[Invisible Armies Insurgency Tracker: A Visual History of Guerrilla Warfare From 1775 to 2012 (from April 18th, 2013](https://www.cfr.org/wars-and-warfare/invisible-armies-insurgency-tracker/p29917)__. As they describe it at the top of the page: "The interactive Invisible Armies Insurgency Tracker presents a database of insurgencies from 1775 to 2012. It supplements the comprehensive historical narrative in Invisible Armies: An Epic History of Guerrilla Warfare from Ancient Times to the Present, by CFR Senior Fellow Max Boot."
# - West Point has __[The Combating Terrorism Center](https://ctc.usma.edu/regions/middle-east/)__ which is incredibly helpful and insightful with their background information.
# - __[The Terrorism Research and Analysis Consortium](https://www.trackingterrorism.org/region/afghanistan)__ has a lot of information - mostly private and paid, but one can at least browse the surface of what they offer (we also used some of their 'vulnerable cities' classifications in our cells below).
# + [markdown] slideshow={"slide_type": "skip"}
# ##### 1. Defining Categories for Values within 'Sub_targettxt', 'Group_Name' and 'City':
# + slideshow={"slide_type": "slide"}
# Here is the function we will use to take our list of values and replace it with specific group/category names:
# We will do this for each of the test and training sets:
def magic_value_replacer(df, column, variable, string):
df[column] = df[column].replace(variable, string)
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Sub_targettxt:
# + slideshow={"slide_type": "skip"}
industrial = ['Gas/Oil/Electric', 'Industrial/Textiles/Factory', 'Farm/Ranch', 'Mining', 'Construction']
white_collar = ['Restaurant/Bar/Café', 'Bank/Commerce', 'Multinational Corporation',
'Medical/Pharmaceutical', 'Retail/Grocery/Bakery (including cell phone shops and generic shops)',
'Hotel/Resort', 'Entertainment/Cultural/Stadium/Casino', 'Private Security Company/Firm',
'Legal Services', 'Retail/Grocery/Bakery']
gov_figure1 = ['Judges/Attorneys/Courts', 'Judge/Attorney/Court',
'Government Personnel (excluding police, military)']
gov_figure2 = ['Politician or Political Party Movement/Meeting/Rally', 'Royalty', 'Head of State',
'Election Related', 'Election-related']
pol_facilities = ['Police Buildings (Headquarters/Stations/School)',
'Police Patrol (including vehicles and convoys)', 'Police Checkpoint', 'Prison/Jail',
'Police Building (headquarters, station, school)']
mil_facilities = ['Military Barracks/Base/Headquarters/Checkpost', 'Military Recruiting Station/Academy',
'Military Weaponry', 'Military Aircraft', 'Military Maritime', 'Paramilitary',
'Military Transportation/Vehicle (excluding convoys)', 'Military Checkpoint']
mil_personnel = ['Military Unit/Patrol/Convoy', 'Non-combatant Personnel',
'Military Personnel (soldiers, troops, officers, forces)']
gov_diplomatic = ['Diplomatic Personnel (outside of embassy, consulate)', 'Embassy/Consulate', 'NATO',
'International Organization (peacekeeper, aid agency, compound)']
educational = ['Teacher/Professor/Instructor', 'School/University/Educational Building', 'Other Personnel']
food_water = ['Food Supply', 'Water Supply']
internet_comm_information = ['Newspaper Journalist/Staff/Facility', 'Radio Journalist/Staff/Facility',
'Television Journalist/Staff/Facility', 'Other (including online news agencies)',
'Radio', 'Internet Infrastructure', 'Television', 'Electricity',
'Telephone/Telegraph']
religious = ['Religion Identified', 'Religious Figure', 'Place of Worship', 'Affiliated Institution']
political = ['Protrainer', 'Political Party Member/Rally', 'Party Official/Candidate/Other Personnel',
'Party Office/Facility', 'Rally']
mass_socio = ['Refugee (including Camps/IDP/Asylum Seekers)', 'Named Civilian', 'Student',
'Race/Ethnicity Identified', 'Farmer', 'Vehicles/Transportation', 'Marketplace/Plaza/Square',
'Village/City/Town/Suburb', 'House/Apartment/Residence', 'Laborer (General)/Occupation Identified',
'Procession/Gathering (funeral, wedding, birthday, religious)', 'Civilian Maritime',
'Public Areas (e.g., Public garden, parking lot, garage, beach, public buildings, camps)',
'Public Area (garden, parking lot, garage, beach, public building, camp)', 'Port',
'Memorial/Cemetery/Monument', 'Museum/Cultural Center/Cultural House', 'Labor Union Related',
'Tourism Travel Agency', 'Tour Bus/Van/Vehicle', 'Tourist', 'Other Facility', 'Airport',
'train/train Tracks/ Trolley', 'Bus Station/Stop', 'Subway', 'Bridge/Car Tunnel',
'Highway/Road/Toll/Traffic Signal', 'Taxi/Rickshaw', 'Bus (excluding tourists)',
'Commercial Maritime', 'Train/Train Tracks/Trolley', 'Aircraft (not at an airport)',
'Airline Officer/Personnel', 'Aircraft (not at an airport)',
'Demilitarized Zone (including Green Zone)']
first_responders = ['Clinics', 'Fire Fighter/Truck', 'Ambulance']
other_utilities = [ 'Gas', 'Oil', 'Oil Tanker']
# + [markdown] slideshow={"slide_type": "fragment"}
# ###### specific_target:
#
# There were some duplicates here that were related to what we are looking for, so I went ahead and combined them in the hopes that they might lead to some more insights.
# + slideshow={"slide_type": "fragment"}
comm_related = ['Cell tower', 'Cell phone tower', 'Cell Tower', 'Cell Phone Tower', 'Cell Phone Shop',
'Telecommunication Tower', 'Telecommunications Tower', 'Radio Stations', 'Radio Station',
'Radio station', 'radio station', 'Radio station antenna', 'A mobile phone tower',
'A mobile tower was targeted in the attack.', 'A Globe Telecom cell site', 'Internet Cafe',
'Telecommunications office', 'Telecommunication Institute', 'Communications Tower',
'Telecommunications Mast', 'An internet cafe']
polling_areas = ['Polling Station', 'Polling Center', 'Polling Stations', 'Polling Booth']
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Group_name: (Grouping by Ideology, Political Tendencies, Etc.)
# + slideshow={"slide_type": "skip"}
palestinian_separatists = ['Hamas (Islamic Resistance Movement)', 'Palestinian Islamic Jihad (PIJ)',
'Popular Front for the Liberation of Palestine (PFLP)', 'Popular Resistance Committees',
'Al-Fatah']
militants = ['Militants', 'Gunmen']
asian_separatists = ['Abu Sayyaf Group (ASG)', 'Colonel Karuna Faction', 'Eastern Turkistan Islamic Movement (ETIM)',
'Free Aceh Movement (GAM)', '<NAME> (Jtmm)',
'<NAME>- Goit (Jtmm-G)',
'<NAME>a- Jwala Singh (Jtmm-J)',
'<NAME>orcha- Rajan Mukti (Jtmm-R)',
'Liberation Tigers of Tamil Eelam (LTTE)', 'Moro Islamic Liberation Front (MILF)',
'Runda Kumpulan Kecil (Rkk)', 'Terai Army']
middle_eastern_separatists = ['Haqqani Network', 'Harkatul Jihad-E-Islami', 'Lashkar-E-Taiba (Let)',
'Kurdistan Workers\' Party (PKK)', 'Lashkar-E-Balochistan', 'Chechen Rebels',
'Free Syrian Army', 'Caucasus Emirate', 'Baloch Republican Army (BRA)',
'Ansar Al-Islam', 'Kurdistan Free Life Party', 'Baloch Liberation Front (Blf)',
'Baloch Liberation Army (BLA)', 'Ansar Al-Sharia (Libya)', 'Jaish-E-Mohammad (Jem)',
'Riyadus-Salikhin Reconnaissance And Sabotage Battalion Of Chechen Martyrs',
'Hizbul Mujahideen (Hm)', 'Southern Mobility Movement (Yemen)',
'Supreme Council For Islamic Revolution In Iraq (Sciri)']
indian_separatists = ['Dima Halao Daoga (Dhd)', 'Black Widows', 'Garo National Liberation Army',
'Kangleipak Communist Party (KCP)', 'National Democratic Front of Bodoland (NDFB)',
'National Liberation Front of Tripura (NLFT)', 'People\'s Liberation Army (PLA)',
'United Liberation Front of Assam (ULFA)', 'United National Liberation Front (UNLF)',
'Karbi Longri North Cachar Liberation Front (Klnlf)',
'National Socialist Council of Nagaland-Isak-Muivah (NSCN-IM)',
'People\'s Revolutionary Party of Kangleipak (PREPAK)']
NW_indian_groups = ['Lashkar-e-Jhangvi', 'Sipah-e-Sahaba/Pakistan (SSP)', 'Hizbul Mujahideen (HM)',
'Baloch Liberation Front (BLF)', 'Baloch Young Tigers (BYT)', 'Baloch Young Tigers (BYT)',
'Baloch Liberation Army (BLA)', 'Baloch Republican Army (BRA)', 'United Baloch Army (UBA)',
'Free Balochistan Army (FBA)', 'Baloch Nationalists']
SE_indian_groups = ['Communist Party of India - Maoist (CPI-Maoist)', 'Indian Mujahideen',
'Jama\'atul Mujahideen Bangladesh (JMB)', 'Bangladesh Sarbahara Party',
'Purbo Banglar Communist Party', 'Harkatul Jihad-e-Islami',
'National Socialist Council of Nagaland-Unification (NSCN-U)',
'Kanglei Yawol Kanna Lup (KYKL)', 'Kuki Tribal Militants', 'Kuki National Front (KNF)',
'United Kuki Liberation Front (UKLF) - India', 'Hill Tiger Force (HTF)',
'National Socialist Council of Nagaland-Khaplang (NSCN-K)',
'National Socialist Council of Nagaland-Isak-Muivah (NSCN-IM)',
'Hynniewtrep National Liberation Council (HNLC)']
african_political = ['National Union for the Total Independence of Angola (UNITA)']
irish_separatists = ['Real Irish Republican Army (RIRA)', 'Oglaigh Na Heireann', 'Irish Republican Army (IRA)']
FARC_left_right = ['National Liberation Army of Colombia (ELN)', 'Popular Liberation Army (EPL)',
'Revolutionary Armed Forces of Colombia (FARC)', 'United Self Defense Units of Colombia (AUC)']
middle_eastern_religious = ['Al-Gama\'at Al-Islamiyya (IG)', 'Al-Nusrah Front', 'Al-Qa\'ida',
'Al-Qa\'ida in the Arabian Peninsula (AQAP)', 'Al-Shabaab', 'Ansar Al-Islam',
'Ansar Al-Sharia (Libya)', 'Al-Qa\'ida in the Lands of the Islamic Maghreb (AQLIM)',
'Asa\'Ib Ahl Al-Haqq', 'Caucasus Emirate', 'Eritrean Islamic Jihad Movement (EIJM)',
'Great Eastern Islamic Raiders Front (Ibda-C)', 'Hizbul Al Islam (Somalia)',
'Islamic Courts Union (ICU)', 'Islamic State of Iraq and al Sham (ISIS)',
'Islamic Movement of Uzbekistan (IMU)', 'Jamiat Ul-Mujahedin (Jum)',
'Jundallah', 'Mahdi Army', 'Taliban', 'Tehrik-i-Taliban Pakistan (TTP)',
'Muslim extremists', 'Armed Islamic Group (GIA)', 'Sunni Muslim extremists',
'Al-Qaida in the Islamic Maghreb (AQIM)', 'Al-Qaida', 'Al-Qaida in Iraq',
'Islamic State of Iraq and the Levant (ISIL)',
'Al-Qaida in the Arabian Peninsula (AQAP)']
israel_palestine_lebanon = ['Anti-Semitic extremists', 'Hezbollah']
asian_african_religious = ['Students Islamic Movement of India (Simi)', 'Ranbir Sena', 'Jemaah Islamiya (JI)',
'Movement for Oneness and Jihad in West Africa (MUJAO)', 'Lord\'s Resistance Army (LRA)',
'Boko Haram']
# + [markdown] slideshow={"slide_type": "skip"}
# ###### City: (Grouping by Predominate Religion, Political Party or Conflict Issue)
# + slideshow={"slide_type": "skip"}
sunni_cities = ['Mosul', 'Kirkuk', 'Sanandaj', 'Ramadi', 'Trabzone', 'Diarbekir',
'Damascus', 'Gwadar', 'Zahedan', 'Kandahar', 'Khiva', 'Fallujah',
'Dakhla', 'Tajura', 'Sabrata', 'Azizia', 'Kasabat', 'Misrata', 'Tripoli',
'Takrit', 'Tikrit']
shia_cities = ['Mecca', 'Najaf', 'Karbala', 'Samarra', 'Ahwaz', 'Basra',
'Medina', 'Tabriz', 'Tunceli', 'Zahran', 'Tehran', 'Rasht', 'Bojnurd',
'Hillah', 'Diwania', 'Khalis', 'Dujali', 'Balad', 'Khanaqin',
'Sargodha', 'Dadu', 'Moro']
split_cities = ['Kirmanshah', 'Baghdad', 'Kadhimia', 'Kuwait', 'Kars', 'Maras',
'Ankara', 'Sivas', 'Aleppo', 'Beirut', 'Abha', 'Jizan', 'Qazvin',
'Gunbad', 'Ashgabat', 'Mashhad', 'Herat', 'Merv', 'Charju', 'Bukhara',
'Samarkand', 'Mazari Sharif', 'Kandahar', 'Lar', 'Bandar Abbas', 'Dubai',
'Abu Dhabi', 'Tashkent', 'Erzurum', 'Konya', 'Izmir', 'Bursa', 'Istanbul',
'Tarhuna', ]
ibadi_cities_libya = ['Nalut', 'Zentan', 'Gharian', 'Dafnia', 'Abu Kammash', 'Zuwara']
columbia_eln_cities = ['Riosucio', 'Buenaventura', 'Cali', 'Popayán', 'Bucaramanga',
'Barrancabermeja', 'Cucuta', 'Santa Rita']
maoist_insurgency = ['Bhagalpur', 'Arwal', 'Khagaria', 'Rohtas', 'Kaimur',
'Bhabua', 'Munger', 'Monghyr', 'Vaishali',
'Dhanbad', 'Pakur', 'Koderma', 'Palamu', 'Balaghat',
'Katni', 'Khandwa', 'Rajgarh', 'Shajapur']
w_africa_muslim = ['Touba', 'N\'Djamena', 'Maiduguri', 'Zaria', 'Sokoto', 'Kenema',
'Cetoua', 'Mopte', 'Bobo-Dioulasso', 'Kayes', 'Monrovia']
w_africa_mixed = ['Dogondutchi', 'Niamey', 'Parakou', 'Abuja', 'Kaduna', 'Kankan',
'Lagos', 'Port Harcourt', 'Couala', 'Yaounde', 'Kumasi', 'Bamako',
'Bertoua', 'Liberville', 'Port-Gentil', 'Zinder', 'Ouagadougou',
'Freetown', 'Conakry', 'Bissau', 'Banjul', 'Dakar']
w_africa_christian = ['Benin City', 'Onitsha', 'Abidjan', 'Takoradi', 'Accra', 'Lome']
# Nepal Maoist Conflict - Class A According to SATP
Nepal_maoist_A = ['Musikot', 'Rukumkot', 'Jajarkot District', 'Salyan', 'Pyuthan', 'Gajul', 'Rank', 'Budagaun',
'Kalikot District', 'Rolpa', 'Rolpa District', 'Rukum District', 'Khalanga']
# Nepal Maoist Conflict - Class B According to SATP
Nepal_maoist_B = ['Charikot', 'Dolakha', 'Jiri', 'Ramechhap', 'Sindhuli Garhi', 'Sindhuli District', 'Dhungrebas',
'Panaoti', 'Gorkha', 'Tulsipur', 'Ghorahi', 'Surkhet', 'Birendranagar',
'Accham', 'Kamal Bajar', 'Dang', 'Dang District']
# Nepal Maoist Conflict - Class C According to SATP
Nepal_maoist_C = ['Khotang Bajar', 'Khotang District', 'Khandanda', 'Okhaldhunga', 'Rumjatar', 'Udayapur Garhi',
'Rasuwa District', 'Gaighat', 'Hitura', 'Makwanpur Garhi', 'Patan', 'Baglung', 'Dhorpatan',
'Bardiya', 'Gulariya', 'Dailekh', 'Dailekh District', 'Jumla', 'Dhading District',
'Udayapur District', 'Lalitpur', 'Hetauda', 'Gulariya']
# N_Triangle_S.America:
northern_triangle = ['Tegucigalpa', 'San Pedro Sula', 'Guatemala City', 'Villa Nueva', 'Villa Canales',
'Mixco', 'San Jan Sacatepequez', 'Chinautla', 'Escuintla', 'Jalapa', 'Puerto Barrios',
'Morales', 'La Libertad', 'Nueva Concepcion', 'Metapan', 'Acajutla', 'Sonsonate',
'Izalco', 'San Salvador', 'Apopa', 'Zaragoza', 'Colon', 'Santa Tecla', 'Usulutan',
'San Miguel', 'La Union']
# + [markdown] slideshow={"slide_type": "skip"}
# ##### 2. Implementing the Filtering Function:
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Sub_targettxt:
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_train_X, 'sub_targettxt', industrial, 'Industrial')
magic_value_replacer(cyber_train_X, 'sub_targettxt', white_collar, 'White_collar')
magic_value_replacer(cyber_train_X, 'sub_targettxt', gov_figure1, 'Gov_Figure1')
magic_value_replacer(cyber_train_X, 'sub_targettxt', gov_figure2, 'Gov_Figure2')
magic_value_replacer(cyber_train_X, 'sub_targettxt', pol_facilities, 'Police_Facilities')
magic_value_replacer(cyber_train_X, 'sub_targettxt', mil_facilities, 'Military_Facilities')
magic_value_replacer(cyber_train_X, 'sub_targettxt', mil_personnel, 'Military_Personnel')
magic_value_replacer(cyber_train_X, 'sub_targettxt', gov_diplomatic, 'Gov_Diplomatic')
magic_value_replacer(cyber_train_X, 'sub_targettxt', educational, 'Educational')
magic_value_replacer(cyber_train_X, 'sub_targettxt', food_water, 'Food_Water')
# magic_value_replacer(cyber_train_X, 'sub_targettxt', internet_comm_information, 'Info/Comm/Internet')
magic_value_replacer(cyber_train_X, 'sub_targettxt', religious, 'Religious')
magic_value_replacer(cyber_train_X, 'sub_targettxt', political, 'Political')
magic_value_replacer(cyber_train_X, 'sub_targettxt', mass_socio, 'Mass_Socio')
magic_value_replacer(cyber_train_X, 'sub_targettxt', first_responders, 'First_Responders')
magic_value_replacer(cyber_train_X, 'sub_targettxt', other_utilities, 'Other_Utilities')
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_test_X, 'sub_targettxt', industrial, 'Industrial')
magic_value_replacer(cyber_test_X, 'sub_targettxt', white_collar, 'White_collar')
magic_value_replacer(cyber_test_X, 'sub_targettxt', gov_figure1, 'Gov_Figure1')
magic_value_replacer(cyber_test_X, 'sub_targettxt', gov_figure2, 'Gov_Figure2')
magic_value_replacer(cyber_test_X, 'sub_targettxt', pol_facilities, 'Police_Facilities')
magic_value_replacer(cyber_test_X, 'sub_targettxt', mil_facilities, 'Military_Facilities')
magic_value_replacer(cyber_test_X, 'sub_targettxt', mil_personnel, 'Military_Personnel')
magic_value_replacer(cyber_test_X, 'sub_targettxt', gov_diplomatic, 'Gov_Diplomatic')
magic_value_replacer(cyber_test_X, 'sub_targettxt', educational, 'Educational')
magic_value_replacer(cyber_test_X, 'sub_targettxt', food_water, 'Food_Water')
# magic_value_replacer(cyber_test_X, 'sub_targettxt', internet_comm_information, 'Info/Comm/Internet')
magic_value_replacer(cyber_test_X, 'sub_targettxt', religious, 'Religious')
magic_value_replacer(cyber_test_X, 'sub_targettxt', political, 'Political')
magic_value_replacer(cyber_test_X, 'sub_targettxt', mass_socio, 'Mass_Socio')
magic_value_replacer(cyber_test_X, 'sub_targettxt', first_responders, 'First_Responders')
magic_value_replacer(cyber_test_X, 'sub_targettxt', other_utilities, 'Other_Utilities')
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Specific_target:
# + slideshow={"slide_type": "subslide"}
magic_value_replacer(cyber_train_X, 'specific_target', comm_related, 'Comm-Related')
magic_value_replacer(cyber_train_X, 'specific_target', polling_areas, 'Polling_Areas')
magic_value_replacer(cyber_test_X, 'specific_target', comm_related, 'Comm-Related')
magic_value_replacer(cyber_test_X, 'specific_target', polling_areas, 'Polling_Areas')
# Also applying this to cyber_data for some visualizations:
magic_value_replacer(cyber_data, 'specific_target', comm_related, 'Comm-Related')
magic_value_replacer(cyber_data, 'specific_target', polling_areas, 'Polling_Areas')
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Group_name: (Grouping by Ideology, Political Tendencies, Etc.)
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_train_X, 'group_name', palestinian_separatists, 'Palestinian_Separatists')
magic_value_replacer(cyber_train_X, 'group_name', militants, 'Militants')
magic_value_replacer(cyber_train_X, 'group_name', asian_separatists, 'Asian_Separatists')
magic_value_replacer(cyber_train_X, 'group_name', middle_eastern_separatists, 'Middle_Eastern_Separatists')
magic_value_replacer(cyber_train_X, 'group_name', indian_separatists, 'Indian_Separatists')
magic_value_replacer(cyber_train_X, 'group_name', NW_indian_groups, 'NW_Indian_Groups')
magic_value_replacer(cyber_train_X, 'group_name', SE_indian_groups, 'NW_Indian_Groups')
magic_value_replacer(cyber_train_X, 'group_name', african_political, 'African_Political')
magic_value_replacer(cyber_train_X, 'group_name', irish_separatists, 'Irish_Separatists')
magic_value_replacer(cyber_train_X, 'group_name', FARC_left_right, 'FARC_left_right')
magic_value_replacer(cyber_train_X, 'group_name', middle_eastern_religious, 'Middle_Eastern_Religious')
magic_value_replacer(cyber_train_X, 'group_name', israel_palestine_lebanon, 'Israel_Palestinian_Lebanon')
magic_value_replacer(cyber_train_X, 'group_name', asian_african_religious, 'Asian_African_Religious')
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_test_X, 'group_name', palestinian_separatists, 'Palestinian_Separatists')
magic_value_replacer(cyber_test_X, 'group_name', militants, 'Militants')
magic_value_replacer(cyber_test_X, 'group_name', asian_separatists, 'Asian_Separatists')
magic_value_replacer(cyber_test_X, 'group_name', middle_eastern_separatists, 'Middle_Eastern_Separatists')
magic_value_replacer(cyber_test_X, 'group_name', indian_separatists, 'Indian_Separatists')
magic_value_replacer(cyber_test_X, 'group_name', NW_indian_groups, 'NW_Indian_Groups')
magic_value_replacer(cyber_test_X, 'group_name', SE_indian_groups, 'NW_Indian_Groups')
magic_value_replacer(cyber_test_X, 'group_name', african_political, 'African_Political')
magic_value_replacer(cyber_test_X, 'group_name', irish_separatists, 'Irish_Separatists')
magic_value_replacer(cyber_test_X, 'group_name', FARC_left_right, 'FARC_left_right')
magic_value_replacer(cyber_test_X, 'group_name', middle_eastern_religious, 'Middle_Eastern_Religious')
magic_value_replacer(cyber_test_X, 'group_name', israel_palestine_lebanon, 'Israel_Palestinian_Lebanon')
magic_value_replacer(cyber_test_X, 'group_name', asian_african_religious, 'Asian_African_Religious')
# + [markdown] slideshow={"slide_type": "skip"}
# ###### City: (Grouping by Predominate Religion, Political Party or Conflict Issue)
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_train_X, 'city', sunni_cities, 'Sunni_Cities')
magic_value_replacer(cyber_train_X, 'city', shia_cities, 'Shia_Cities')
magic_value_replacer(cyber_train_X, 'city', split_cities, 'Split_Cities')
magic_value_replacer(cyber_train_X, 'city', ibadi_cities_libya, 'Ibadi_Cities_Libya')
magic_value_replacer(cyber_train_X, 'city', columbia_eln_cities, 'Columbia_ELN_Cities')
magic_value_replacer(cyber_train_X, 'city', maoist_insurgency, 'Maoist_Insurgency')
magic_value_replacer(cyber_train_X, 'city', w_africa_muslim, 'W_Africa_muslim')
magic_value_replacer(cyber_train_X, 'city', w_africa_mixed, 'W_Africa_mixed')
magic_value_replacer(cyber_train_X, 'city', w_africa_christian, 'W_Africa_christian')
magic_value_replacer(cyber_train_X, 'city', Nepal_maoist_A, 'Nepal_Maoist_A')
magic_value_replacer(cyber_train_X, 'city', Nepal_maoist_B, 'Nepal_Maoist_B')
magic_value_replacer(cyber_train_X, 'city', Nepal_maoist_C, 'Nepal_Maoist_C')
magic_value_replacer(cyber_train_X, 'city', northern_triangle, 'Northern_Triangle')
# + slideshow={"slide_type": "skip"}
magic_value_replacer(cyber_test_X, 'city', sunni_cities, 'Sunni_Cities')
magic_value_replacer(cyber_test_X, 'city', shia_cities, 'Shia_Cities')
magic_value_replacer(cyber_test_X, 'city', split_cities, 'Split_Cities')
magic_value_replacer(cyber_test_X, 'city', ibadi_cities_libya, 'Ibadi_Cities_Libya')
magic_value_replacer(cyber_test_X, 'city', columbia_eln_cities, 'Columbia_ELN_Cities')
magic_value_replacer(cyber_test_X, 'city', maoist_insurgency, 'Maoist_Insurgency')
magic_value_replacer(cyber_test_X, 'city', w_africa_muslim, 'W_Africa_muslim')
magic_value_replacer(cyber_test_X, 'city', w_africa_mixed, 'W_Africa_mixed')
magic_value_replacer(cyber_test_X, 'city', w_africa_christian, 'W_Africa_christian')
magic_value_replacer(cyber_test_X, 'city', Nepal_maoist_A, 'Nepal_Maoist_A')
magic_value_replacer(cyber_test_X, 'city', Nepal_maoist_B, 'Nepal_Maoist_B')
magic_value_replacer(cyber_test_X, 'city', Nepal_maoist_C, 'Nepal_Maoist_C')
magic_value_replacer(cyber_test_X, 'city', northern_triangle, 'Northern_Triangle')
# + slideshow={"slide_type": "skip"}
print(cyber_train_X.shape)
print(cyber_test_X.shape)
# + slideshow={"slide_type": "skip"}
# Mitigating some memory issues:
del industrial, white_collar, gov_figure1, gov_figure2, pol_facilities, mil_facilities, mil_personnel
del gov_diplomatic, educational, food_water, internet_comm_information, religious, political, mass_socio,
del sunni_cities, shia_cities, split_cities, ibadi_cities_libya, columbia_eln_cities, maoist_insurgency
del w_africa_muslim, w_africa_mixed, w_africa_christian, Nepal_maoist_A, Nepal_maoist_B, Nepal_maoist_C
del northern_triangle, african_political, asian_separatists, middle_eastern_separatists, first_responders
del FARC_left_right, middle_eastern_religious, israel_palestine_lebanon, asian_african_religious, militants
del indian_separatists, NW_indian_groups, SE_indian_groups, irish_separatists, palestinian_separatists,
del other_utilities, comm_related, polling_areas
gc.collect()
# + slideshow={"slide_type": "skip"}
ipython_vars = ['In', 'Out', 'exit', 'quit', 'get_ipython', 'ipython_vars']
cleaner = sorted([(x, sys.getsizeof(globals().get(x))) for x in dir() if not x.startswith('_') and x not in sys.modules and x not in ipython_vars],
key=lambda x: x[1], reverse=True)
# + [markdown] slideshow={"slide_type": "slide"}
# ### 2. Preliminary Visualizations and Exploration:
#
# Here we take a look at some of the correlations and relationships between our features/columns and see what we might want to focus on with our models.
# + slideshow={"slide_type": "skip"}
success_class_balance = pd.DataFrame(cyber_data['success'].value_counts())
cyber_data_description = pd.DataFrame(cyber_data.describe())
cyber_data_objects = pd.DataFrame(cyber_data.dtypes.loc[cyber_data.dtypes == 'O'])
cyber_data_ints = pd.DataFrame(cyber_data.dtypes.loc[cyber_data.dtypes == 'int64'])
cyber_data_floats = pd.DataFrame(cyber_data.dtypes.loc[cyber_data.dtypes == 'float'])
# + [markdown] slideshow={"slide_type": "fragment"} variables={"success_class_balance": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>success</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>1</th>\n <td>11994</td>\n </tr>\n <tr>\n <th>0</th>\n <td>1261</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Based on our look at value counts, it looks like we will need to keep our class imbalance in mind as we continue. Once we get to running our models, we will use a balanced accuracy score in order to evaluate our model with a more accurate perspective.
#
# {{success_class_balance}}
#
# The descriptive statistics below give us a nice layout of our more common values for each feature, number of unique values and their distributions as well. After this, it looks as if our feature object types are relatively workable. We should be fine in that regard.
# + [markdown] slideshow={"slide_type": "subslide"} variables={"cyber_data_description": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>event_id</th>\n <th>crit1</th>\n <th>crit2</th>\n <th>crit3</th>\n <th>doubt</th>\n <th>success</th>\n <th>suicide</th>\n <th>group_attrib_crtainty</th>\n <th>unaffil_individ</th>\n <th>property</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>count</th>\n <td>1.325500e+04</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n <td>13206.000000</td>\n <td>13255.000000</td>\n <td>13255.000000</td>\n </tr>\n <tr>\n <th>mean</th>\n <td>2.008238e+11</td>\n <td>0.988759</td>\n <td>0.987401</td>\n <td>0.948397</td>\n <td>-0.047680</td>\n <td>0.904866</td>\n <td>0.040890</td>\n <td>0.190292</td>\n <td>0.005734</td>\n <td>-0.937156</td>\n </tr>\n <tr>\n <th>std</th>\n <td>9.704446e+08</td>\n <td>0.105430</td>\n <td>0.111540</td>\n <td>0.221233</td>\n <td>1.204459</td>\n <td>0.293411</td>\n <td>0.198043</td>\n <td>0.392547</td>\n <td>0.075507</td>\n <td>3.455691</td>\n </tr>\n <tr>\n <th>min</th>\n <td>1.970011e+11</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>-9.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>-9.000000</td>\n </tr>\n <tr>\n <th>25%</th>\n <td>2.008010e+11</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n </tr>\n <tr>\n <th>50%</th>\n <td>2.011100e+11</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n </tr>\n <tr>\n <th>75%</th>\n <td>2.014062e+11</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>1.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>0.000000</td>\n <td>1.000000</td>\n </tr>\n <tr>\n <th>max</th>\n <td>2.017123e+11</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n <td>1.000000</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# > Descriptive Statistics:
# {{cyber_data_description}}
# + slideshow={"slide_type": "subslide"}
cyber_data.describe(include = 'O')
# + [markdown] slideshow={"slide_type": "subslide"} variables={"cyber_data_floats": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>doubt</th>\n <td>float64</td>\n </tr>\n <tr>\n <th>group_attrib_crtainty</th>\n <td>float64</td>\n </tr>\n </tbody>\n</table>\n</div>", "cyber_data_ints": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>event_id</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>crit1</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>crit2</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>crit3</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>success</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>suicide</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>unaffil_individ</th>\n <td>int64</td>\n </tr>\n <tr>\n <th>property</th>\n <td>int64</td>\n </tr>\n </tbody>\n</table>\n</div>", "cyber_data_objects": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>country_txt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>region_txt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>city</th>\n <td>object</td>\n </tr>\n <tr>\n <th>summary</th>\n <td>object</td>\n </tr>\n <tr>\n <th>attack_1txt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>target_1txt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>sub_targettxt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>corp1</th>\n <td>object</td>\n </tr>\n <tr>\n <th>specific_target</th>\n <td>object</td>\n </tr>\n <tr>\n <th>victim_nationalitytxt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>group_name</th>\n <td>object</td>\n </tr>\n <tr>\n <th>motive</th>\n <td>object</td>\n </tr>\n <tr>\n <th>weapontxt</th>\n <td>object</td>\n </tr>\n <tr>\n <th>sub_weapontxt</th>\n <td>object</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# <head>
# <table>
# <tr>
# <td> {{cyber_data_objects}} </td>
# <td> {{cyber_data_ints}} </td>
# <td> {{cyber_data_floats}} </td>
# </tr>
# </table>
# </head>
#
#
#
# + slideshow={"slide_type": "subslide"}
cyber_data.specific_target.value_counts().head(10)
# + [markdown] slideshow={"slide_type": "skip"}
# #### B. Correlation:
# + slideshow={"slide_type": "slide"}
f, ax = plt.subplots(figsize = (11, 9))
ax1 = sns.heatmap(cyber_data.corr(), annot = True)
plt.show()
# + [markdown] slideshow={"slide_type": "skip"}
# #### C. Most Active Groups:
# + slideshow={"slide_type": "skip"}
# Most prolific groups:
actives = cyber_train_X['group_name'].value_counts().head(11).drop('Unknown')
mask3 = cyber_train_X['group_name'].map(lambda x: x in actives)
actives_df = cyber_train_X[mask3]
temp_output = cyber_train_Y.loc[actives_df.index]
beta_1 = actives.keys()
gamma_1 = actives.values
# Most affected countries:
hot_countries = cyber_train_X['country_txt'].value_counts().head(10)
mask3 = cyber_train_X['country_txt'].map(lambda x: x in hot_countries)
hot_countries_df = cyber_train_X[mask3]
phi = hot_countries_df['country_txt'].value_counts().head(10)
temp_output2 = cyber_train_Y.loc[hot_countries_df.index]
beta_2 = phi.keys()
gamma_2 = phi.values
# + slideshow={"slide_type": "slide"}
f, (ax1, ax2) = plt.subplots(1, 2, figsize = (17, 7))
plt.subplot(1, 2, 1)
ax1 = sns.pointplot(x = actives_df['group_name'], y = temp_output['success'])
ax1.set_xticklabels(labels = beta_1, rotation = '80', fontdict = {'fontsize':10})
ax1.set_xlabel('Name of Terrorist Group', fontdict = {'fontsize':12})
ax1.set_ylabel('Success Rate', fontdict = {'fontsize':12})
ax1.set_title('Most Active Terror Groups and Their Success Rates')
plt.subplots_adjust(wspace = .4)
plt.subplot(1, 2, 2)
ax2 = sns.pointplot(x = beta_2, y = gamma_2)
ax2.set_xticklabels(labels = beta_2, rotation = '80', fontdict = {'fontsize':12})
ax2.set_xlabel('Name of Country', fontdict = {'fontsize':12})
ax2.set_ylabel('Number of Successful Attacks', fontdict = {'fontsize':12})
ax2.set_title('Countries with Most Activity and Number of Successful Attacks')
plt.subplots_adjust(wspace = .3, hspace = .3)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# #### E. Highly Targeted Areas:
#
# The next few graphs take a look at the target feature (i.e. the feature that describes what sector is targeted within the attacks) and how they are distributed according to other features. After having run some preliminary feature selections, our algorithims highlighted the importance of the target features, so we wanted to take a look at it in more-depth.
# + slideshow={"slide_type": "skip"}
# Gathering the top-10 targets:
big_targets = cyber_data['target_1txt'].value_counts().head(5)
big_targets_mask = cyber_data['target_1txt'].apply(lambda x: x in big_targets)
targeted_df = cyber_data[big_targets_mask]
# + slideshow={"slide_type": "subslide"}
# Visualization Parameters:
countplot_kwargs = {'edgecolor':'black',
'linewidth':.85,
'alpha':.85}
countplot_rc = {'figure.dpi': 90,
'font.size': 20}
# Plot set-up:
plt.figure(figsize = (20, 10))
plt.rc(countplot_rc)
ax1 = sns.countplot(x = 'target_1txt', hue = 'region_txt', data = targeted_df,
orient = 'h', palette = 'Paired', **countplot_kwargs)
ax1.legend(loc = "upper right")
plt.ylabel("Count")
plt.xlabel("Targeted Sectors")
plt.title("Geographical Location for Targeted Areas")
plt.xticks()
plt.show()
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = (20, 10))
plt.rc(countplot_rc)
ax2 = sns.countplot(x = 'target_1txt', hue = 'attack_1txt', data = targeted_df, orient = 'h',
palette = 'Paired', **countplot_kwargs)
ax2.legend(loc = 'upper right')
plt.ylabel("Count")
plt.xlabel("Targeted Sectors")
plt.title("Targeted Areas and Attack Method")
plt.show()
# + slideshow={"slide_type": "subslide"}
plt.figure(figsize = (20, 10))
plt.rc(countplot_rc)
ax3 = sns.countplot(x = 'target_1txt', hue = 'weapontxt', data = targeted_df, orient = 'h',
palette = 'Paired', **countplot_kwargs)
ax3.legend(loc = 'upper right')
plt.ylabel("Count")
plt.xlabel("Targeted Sectors")
plt.title("Targeted Areas and Weapon Type")
plt.show()
# + slideshow={"slide_type": "skip"}
del actives, actives_df, temp_output, beta_1, beta_2, gamma_1, gamma_2,
del hot_countries, hot_countries_df, phi, temp_output2, mask3
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ### 3. Feature Selection:
#
# Getting closer to modeling and using feature selection algorithms to see what will help us best minimize our feature set while maintaining the most amount of variation in our data.
#
# We mainly focused on Select KBest within sklearn and PCA analysis to give us two perspectives on the data (one that we can parse out as humans, namely Select KBest, and one that is mostly computationally described, namely PCA).
# + [markdown] slideshow={"slide_type": "subslide"}
# #### A. Select K-Best:
#
# ##### Assessing the Overall DataFrame and its Features:
# + slideshow={"slide_type": "skip"}
from sklearn import feature_selection
# + slideshow={"slide_type": "fragment"}
# Getting dummies on our training and test sets (slight wrangling involved):
# First dropping any na's:
cyber_train_X.dropna(axis = 0, inplace = True)
cyber_test_X.dropna(axis = 0, inplace = True)
# Then grabbing an index to make sure we maintain our train/test split:
train_index = cyber_train_X.index
test_index = cyber_test_X.index
# Dummy-time (we combined the dataframes here to make sure we didn't get duplicated dummies in both
# training and test sets):
dummy_prep = pd.concat([cyber_train_X, cyber_test_X]).drop(['group_name', 'city', 'weapontxt'], axis = 1)
dummy_1 = pd.get_dummies(dummy_prep)
# Re-filtering our Training/Test Inputs:
cyber_train_dummy_X = dummy_1.loc[train_index]
cyber_test_dummy_X = dummy_1.loc[test_index]
# Re-filtering our Training/Test Outputs:
cyber_train_dummy_Y = cyber_train_Y.loc[train_index]
cyber_test_dummy_Y = cyber_test_Y.loc[test_index]
# + slideshow={"slide_type": "subslide"}
# Making sure we have the same sizes still:
print(cyber_train_dummy_X.shape)
print(cyber_test_dummy_X.shape)
# + slideshow={"slide_type": "skip"}
# Fitting SelectKBest to our Features and output:
# Here we tried a number of sizes: 20, 30, 500 and 700. We found that 25-30 got us the best results.
KBest_1 = feature_selection.SelectKBest(k = 25)
cyber_train_KBest1 = KBest_1.fit_transform(cyber_train_dummy_X, cyber_train_dummy_Y)
# Transforming the test-set
cyber_test_KBest1 = KBest_1.transform(cyber_test_dummy_X)
# + slideshow={"slide_type": "skip"}
# Creating a DF with the top 25 features:
feature_mask = KBest_1.get_support(indices = True)
KBest_1_features = pd.DataFrame(cyber_train_dummy_X.columns[feature_mask])
# + [markdown] slideshow={"slide_type": "subslide"} variables={"KBest_1_features": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>country_txt_Nepal</td>\n </tr>\n <tr>\n <th>1</th>\n <td>country_txt_Uruguay</td>\n </tr>\n <tr>\n <th>2</th>\n <td>attack_1txt_Armed Assault</td>\n </tr>\n <tr>\n <th>3</th>\n <td>attack_1txt_Assassination</td>\n </tr>\n <tr>\n <th>4</th>\n <td>attack_1txt_Bombing/Explosion</td>\n </tr>\n <tr>\n <th>5</th>\n <td>attack_1txt_Facility/Infrastructure Attack</td>\n </tr>\n <tr>\n <th>6</th>\n <td>attack_1txt_Hostage Taking (Kidnapping)</td>\n </tr>\n <tr>\n <th>7</th>\n <td>target_1txt_Government (General)</td>\n </tr>\n <tr>\n <th>8</th>\n <td>target_1txt_Private Citizens & Property</td>\n </tr>\n <tr>\n <th>9</th>\n <td>target_1txt_Unknown</td>\n </tr>\n <tr>\n <th>10</th>\n <td>sub_targettxt_Gov_Figure1</td>\n </tr>\n <tr>\n <th>11</th>\n <td>sub_targettxt_Unknown</td>\n </tr>\n <tr>\n <th>12</th>\n <td>corp1_Indian Armed Forces</td>\n </tr>\n <tr>\n <th>13</th>\n <td>corp1_Not Applicable</td>\n </tr>\n <tr>\n <th>14</th>\n <td>corp1_Unified Communist Party of Nepal: Maoist...</td>\n </tr>\n <tr>\n <th>15</th>\n <td>specific_target_Election Rally</td>\n </tr>\n <tr>\n <th>16</th>\n <td>specific_target_Internet Providers</td>\n </tr>\n <tr>\n <th>17</th>\n <td>specific_target_Local Jewish Community Member</td>\n </tr>\n <tr>\n <th>18</th>\n <td>specific_target_Unknown</td>\n </tr>\n <tr>\n <th>19</th>\n <td>victim_nationalitytxt_Nepal</td>\n </tr>\n <tr>\n <th>20</th>\n <td>victim_nationalitytxt_Unknown</td>\n </tr>\n <tr>\n <th>21</th>\n <td>sub_weapontxt_Arson/Fire</td>\n </tr>\n <tr>\n <th>22</th>\n <td>sub_weapontxt_Letter Bomb</td>\n </tr>\n <tr>\n <th>23</th>\n <td>sub_weapontxt_Other Explosive Type</td>\n </tr>\n <tr>\n <th>24</th>\n <td>sub_weapontxt_Unknown Gun Type</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Here, we are getting a lot of return from the attack, target, sub-target and a few country features. The doubt column indicates whether doubt exists regarding the classification of this incident as a terrorist incident, as opposed to some sort of other crime. High-correlation here would make sense, but for now, we will focus on the columns below as the doubt feature is potentially over-correlated.
#
# {{KBest_1_features}}
# + [markdown] slideshow={"slide_type": "slide"}
# ##### Assessing Individual Features:
#
# Running SelectKBest on a few features individually to see which of their values is selected as most important. This will help us consolidate some of those values and make a more specific dataframe.
# + [markdown] slideshow={"slide_type": "fragment"}
# ###### Group_name:
# + slideshow={"slide_type": "skip"}
# Getting dummies on our training and test sets (slight wrangling involved):
names_train_X = cyber_train_X['group_name']
names_test_X = cyber_test_X['group_name']
names_train_index = cyber_train_X['group_name'].index
names_test_index = cyber_test_X['group_name'].index
names_dummy_prep = pd.concat([names_train_X, names_test_X])
names_dummy_1 = pd.get_dummies(names_dummy_prep)
# Filtering Training/Test Inputs:
names_train_dummy_X = names_dummy_1.loc[names_train_index]
names_test_dummy_X = names_dummy_1.loc[names_test_index]
# Filtering Training Outputs:
names_train_dummy_Y = cyber_train_Y.loc[names_train_index]
names_test_dummy_Y = cyber_test_Y.loc[names_test_index]
# -
names_train_X
# + slideshow={"slide_type": "skip"}
# Fitting model to our features and output.
# Again, we tried 30, 50 and 150, here. It seemed like 25-30 features gave us the best results.
KBest_names = feature_selection.SelectKBest(k = 25)
names_train_KBest = KBest_names.fit_transform(names_train_dummy_X, names_train_dummy_Y)
# Transforming our test set.
names_test_KBest = KBest_names.transform(names_test_dummy_X)
# + slideshow={"slide_type": "skip"}
# Summarizing the scores for those top 25 features in a df:
names_mask = KBest_names.get_support(indices = True)
KBest_names_features = pd.DataFrame(names_train_dummy_X.columns[names_mask])
# + [markdown] slideshow={"slide_type": "subslide"} variables={"KBest_names_features": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>Armed Commandos of Liberation</td>\n </tr>\n <tr>\n <th>1</th>\n <td>Black September</td>\n </tr>\n <tr>\n <th>2</th>\n <td>Communist Party of Nepal (People's War Group)</td>\n </tr>\n <tr>\n <th>3</th>\n <td>Communist Party of Nepal - Maoist (CPN-Maoist-...</td>\n </tr>\n <tr>\n <th>4</th>\n <td>Communist Party of Nepal-Maoist (Baidya)</td>\n </tr>\n <tr>\n <th>5</th>\n <td>Hekla Reception Committee-Initiative for More ...</td>\n </tr>\n <tr>\n <th>6</th>\n <td>Informal Anarchist Federation</td>\n </tr>\n <tr>\n <th>7</th>\n <td>Jihadi-inspired extremists</td>\n </tr>\n <tr>\n <th>8</th>\n <td>Left-Wing Militants</td>\n </tr>\n <tr>\n <th>9</th>\n <td>Maoist Communist Party of Manipur</td>\n </tr>\n <tr>\n <th>10</th>\n <td>Maoists</td>\n </tr>\n <tr>\n <th>11</th>\n <td>Mexican Revolutionary Movement</td>\n </tr>\n <tr>\n <th>12</th>\n <td>NW_Indian_Groups</td>\n </tr>\n <tr>\n <th>13</th>\n <td>Orly Organization</td>\n </tr>\n <tr>\n <th>14</th>\n <td>Pattani United Liberation Organization (PULO)</td>\n </tr>\n <tr>\n <th>15</th>\n <td>Riyadus-Salikhin Reconnaissance and Sabotage B...</td>\n </tr>\n <tr>\n <th>16</th>\n <td>Sabaot Land Defense Force (SLDF)</td>\n </tr>\n <tr>\n <th>17</th>\n <td>Scottish Socialist Republican League</td>\n </tr>\n <tr>\n <th>18</th>\n <td>September 11</td>\n </tr>\n <tr>\n <th>19</th>\n <td>Supporters of <NAME> dia Wamba</td>\n </tr>\n <tr>\n <th>20</th>\n <td>The 78 Unemployed</td>\n </tr>\n <tr>\n <th>21</th>\n <td>The Extraditables</td>\n </tr>\n <tr>\n <th>22</th>\n <td>The Justice Department</td>\n </tr>\n <tr>\n <th>23</th>\n <td>The World United Formosans for Independence (W...</td>\n </tr>\n <tr>\n <th>24</th>\n <td>Unknown</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# It looks like some of our groupings from above made it into the top-25 features, which is reassuring that our efforts above produced a result. This also gives us a slightly broader view of the terrorist groups that are highly active, as opposed to overly-specific groups that might wash out the activity in other countries.
#
# {{KBest_names_features}}
#
# This also points to another issue for consideration when performing future work on the dataset. When making these groups and filters, one will want to be sure to create groups that consider as much of the globe as possible. Otherwise, one would run the risk of coagulating a few groups together from one area, thereby increasing their significance, and over-powering the significance of other groups. The groups we created above were an attempt at creating well-represented portions of the globe. It would benefit, however, from more time and research so as to further tweak these groupings towards a higher accuracy.
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### City:
# + slideshow={"slide_type": "skip"}
# Getting dummies on our training and test sets (slight wrangling involved):
city_train_X = cyber_train_X['city']
city_test_X = cyber_test_X['city']
city_train_index = cyber_train_X['city'].index
city_test_index = cyber_test_X['city'].index
city_dummy_prep = pd.concat([city_train_X, city_test_X])
city_dummy_1 = pd.get_dummies(city_dummy_prep)
# Training/Test Inputs:
city_train_dummy_X = city_dummy_1.loc[city_train_index]
city_test_dummy_X = city_dummy_1.loc[city_test_index]
# Training Output:
city_train_dummy_Y = cyber_train_Y.loc[city_train_index]
city_test_dummy_Y = cyber_test_Y.loc[city_test_index]
# + slideshow={"slide_type": "skip"}
# Fitting model to our features and output.
KBest_city = feature_selection.SelectKBest(k = 25) # Tried 30 and 150. 25-30 was our best range.
city_train_KBest = KBest_city.fit_transform(city_train_dummy_X, city_train_dummy_Y)
# Transforming our test set:
city_test_KBest = KBest_city.transform(city_test_dummy_X)
# + slideshow={"slide_type": "skip"}
# Summarizing the scores for those top 25 features in a df:
pd.set_option('max_rows', 101)
city_mask = KBest_city.get_support(indices = True)
KBest_city_features = pd.DataFrame(city_train_dummy_X.columns[city_mask])
# + [markdown] slideshow={"slide_type": "subslide"} variables={"KBest_city_features": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>Aguada</td>\n </tr>\n <tr>\n <th>1</th>\n <td>Ajdari</td>\n </tr>\n <tr>\n <th>2</th>\n <td>Bihsud district</td>\n </tr>\n <tr>\n <th>3</th>\n <td>Brussels</td>\n </tr>\n <tr>\n <th>4</th>\n <td>Daraa</td>\n </tr>\n <tr>\n <th>5</th>\n <td>Dhangadhi</td>\n </tr>\n <tr>\n <th>6</th>\n <td>Dublin</td>\n </tr>\n <tr>\n <th>7</th>\n <td>Hind Khel</td>\n </tr>\n <tr>\n <th>8</th>\n <td>Imphal</td>\n </tr>\n <tr>\n <th>9</th>\n <td>Isulan</td>\n </tr>\n <tr>\n <th>10</th>\n <td>Jos</td>\n </tr>\n <tr>\n <th>11</th>\n <td>Kharan district</td>\n </tr>\n <tr>\n <th>12</th>\n <td>Lawdar district</td>\n </tr>\n <tr>\n <th>13</th>\n <td>London</td>\n </tr>\n <tr>\n <th>14</th>\n <td>Los Angeles</td>\n </tr>\n <tr>\n <th>15</th>\n <td>Mogadishu</td>\n </tr>\n <tr>\n <th>16</th>\n <td>Nabalawag</td>\n </tr>\n <tr>\n <th>17</th>\n <td>Ordzhonikidzevskaya</td>\n </tr>\n <tr>\n <th>18</th>\n <td>Paris</td>\n </tr>\n <tr>\n <th>19</th>\n <td>Siraha district</td>\n </tr>\n <tr>\n <th>20</th>\n <td>Sirnoo</td>\n </tr>\n <tr>\n <th>21</th>\n <td>Split_Cities</td>\n </tr>\n <tr>\n <th>22</th>\n <td>Sunni_Cities</td>\n </tr>\n <tr>\n <th>23</th>\n <td>The Hague</td>\n </tr>\n <tr>\n <th>24</th>\n <td>Zinjibar</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# {{KBest_city_features}}
# + [markdown] slideshow={"slide_type": "subslide"}
# ###### Specific_target:
# + slideshow={"slide_type": "skip"}
# Getting dummies on our training and test sets (slight wrangling involved):
spec_targ_train_X = cyber_train_X['specific_target']
spec_targ_test_X = cyber_test_X['specific_target']
spec_targ_train_index = cyber_train_X['specific_target'].index
spec_targ_test_index = cyber_test_X['specific_target'].index
spec_targ_dummy_prep = pd.concat([spec_targ_train_X, spec_targ_test_X])
spec_targ_dummy_1 = pd.get_dummies(spec_targ_dummy_prep)
# Training/Test Inputs:
spec_targ_train_dummy_X = spec_targ_dummy_1.loc[spec_targ_train_index]
spec_targ_test_dummy_X = spec_targ_dummy_1.loc[spec_targ_test_index]
# Training Output:
spec_targ_train_dummy_Y = cyber_train_Y.loc[spec_targ_train_index]
spec_targ_test_dummy_Y = cyber_test_Y.loc[spec_targ_test_index]
# + slideshow={"slide_type": "skip"}
# Fitting the model to our features and training output:
KBest_spec_targ = feature_selection.SelectKBest(k = 25) # Tried 5, but 3 was best.
spec_targ_train_KBest = KBest_spec_targ.fit_transform(spec_targ_train_dummy_X, spec_targ_train_dummy_Y)
# Transforming our test set:
spec_targ_test_KBest = KBest_spec_targ.transform(spec_targ_test_dummy_X)
# + slideshow={"slide_type": "skip"}
# Summarizing the scores for those top 3 features in a df:
spec_targ_mask = KBest_spec_targ.get_support(indices = True)
KBest_spec_targ_features = pd.DataFrame(spec_targ_train_dummy_X.columns[spec_targ_mask])
# + [markdown] slideshow={"slide_type": "subslide"} variables={"KBest_spec_targ_features": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>0</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>A bus terminal</td>\n </tr>\n <tr>\n <th>1</th>\n <td>Bus Park</td>\n </tr>\n <tr>\n <th>2</th>\n <td>Chair: <NAME></td>\n </tr>\n <tr>\n <th>3</th>\n <td>Civilians</td>\n </tr>\n <tr>\n <th>4</th>\n <td>Comm-Related</td>\n </tr>\n <tr>\n <th>5</th>\n <td>Commander</td>\n </tr>\n <tr>\n <th>6</th>\n <td>Election Rally</td>\n </tr>\n <tr>\n <th>7</th>\n <td>Internet Providers</td>\n </tr>\n <tr>\n <th>8</th>\n <td>Judge</td>\n </tr>\n <tr>\n <th>9</th>\n <td>Local Jewish Community Member</td>\n </tr>\n <tr>\n <th>10</th>\n <td>Railway Tracks</td>\n </tr>\n <tr>\n <th>11</th>\n <td>Residence of Leader: Tajmir Khan</td>\n </tr>\n <tr>\n <th>12</th>\n <td>The target was a polling station.</td>\n </tr>\n <tr>\n <th>13</th>\n <td>Town of El Doncello</td>\n </tr>\n <tr>\n <th>14</th>\n <td>Unknown</td>\n </tr>\n <tr>\n <th>15</th>\n <td>four researchers at the Tulane Regional Primat...</td>\n </tr>\n <tr>\n <th>16</th>\n <td><NAME>illos, subdirector radio supe...</td>\n </tr>\n <tr>\n <th>17</th>\n <td>home of <NAME>. transport and...</td>\n </tr>\n <tr>\n <th>18</th>\n <td>researcher, <NAME>, at the So...</td>\n </tr>\n <tr>\n <th>19</th>\n <td>six researchers at Yerkes Regional Primate Res...</td>\n </tr>\n <tr>\n <th>20</th>\n <td>six researchers at the University of Washingto...</td>\n </tr>\n <tr>\n <th>21</th>\n <td>some researchers at the University of Minnesot...</td>\n </tr>\n <tr>\n <th>22</th>\n <td>three researchers at the University of Califor...</td>\n </tr>\n <tr>\n <th>23</th>\n <td>trucking company in Arcata, California</td>\n </tr>\n <tr>\n <th>24</th>\n <td>two primate researchers, <NAME> and Ji...</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# From here, we can infer that while the data-set we are looking at somehow has consequences for cyber infrastructre or networks (including email and social networks as well as the physical networks), there seems to be a correlation with bombings and incendiary methods as well. As such, we cannot rule out the possibility that cyber-related attacks are often related to physical or other forms of terrorist attacks (at least in the current analysis we are doing here).
#
# {{KBest_spec_targ_features}}
# + [markdown] slideshow={"slide_type": "slide"}
# #### B. Principle Component Analysis:
# ##### Assessing the Overall DataFrame:
# + slideshow={"slide_type": "fragment"}
from sklearn.decomposition import PCA
# Using the whole dataset (cyber_train_data):
PCA_1 = PCA(n_components = 25) # Tried 30 and 500
cyber_train_PCA = PCA_1.fit_transform(cyber_train_dummy_X)
cyber_test_PCA = PCA_1.transform(cyber_test_dummy_X)
# + [markdown] slideshow={"slide_type": "skip"}
# ##### Assessing Individual Features
#
# ###### Group_name:
# + slideshow={"slide_type": "skip"}
# Now looking at group_name values:
PCA_name = PCA(n_components = 25) # Tried 30 and 150
name_train_PCA = PCA_name.fit_transform(names_train_dummy_X)
name_test_PCA = PCA_name.transform(names_test_dummy_X)
# + [markdown] slideshow={"slide_type": "skip"}
# ###### City:
# + slideshow={"slide_type": "skip"}
# Then some city stuff:
PCA_cities = PCA(n_components = 25) # Tried 30 and 150
cities_train_PCA = PCA_cities.fit_transform(city_train_dummy_X)
cities_test_PCA = PCA_cities.transform(city_test_dummy_X)
# + [markdown] slideshow={"slide_type": "skip"}
# ###### Specific_target:
# + slideshow={"slide_type": "skip"}
# Specific_target:
PCA_spec_targ = PCA(n_components = 3) # Tried 5
spec_targ_train_PCA = PCA_spec_targ.fit_transform(spec_targ_train_dummy_X)
spec_targ_test_PCA = PCA_spec_targ.transform(spec_targ_test_dummy_X)
# + [markdown] slideshow={"slide_type": "slide"}
# #### C. Creating Specified DF with Selected Features:
#
# ##### DataFrame from KBest Algorithms:
# + slideshow={"slide_type": "fragment"}
# Training:
alpha = pd.DataFrame(cyber_train_KBest1, columns = KBest_1_features[0])
beta = pd.DataFrame(names_train_KBest, columns = KBest_names_features[0])
gamma = pd.DataFrame(city_train_KBest, columns = KBest_city_features[0])
delta = pd.DataFrame(spec_targ_train_KBest, columns = KBest_spec_targ_features[0])
KBest_train_X = pd.concat([alpha, beta, gamma, delta], axis = 1)
# + slideshow={"slide_type": "skip"}
# Test:
alpha2 = pd.DataFrame(cyber_test_KBest1, columns = KBest_1_features[0])
beta2 = pd.DataFrame(names_test_KBest, columns = KBest_names_features[0])
gamma2 = pd.DataFrame(city_test_KBest, columns = KBest_city_features[0])
delta2 = pd.DataFrame(spec_targ_test_KBest, columns = KBest_spec_targ_features[0])
KBest_test_X = pd.concat([alpha2, beta2, gamma2, delta2], axis = 1)
# + slideshow={"slide_type": "skip"}
del alpha, alpha2, city_dummy_1, city_dummy_prep, big_targets, beta,
# + slideshow={"slide_type": "skip"}
del beta2, delta, delta2,
del dummy_1
# + slideshow={"slide_type": "skip"}
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### DataFrame from PCA:
# + slideshow={"slide_type": "fragment"}
# Training:
cyber_train_PCA = pd.DataFrame(cyber_train_PCA)
name_train_PCA = pd.DataFrame(name_train_PCA)
cities_train_PCA = pd.DataFrame(cities_train_PCA)
spec_targ_train_PCA = pd.DataFrame(spec_targ_train_PCA)
PCA_train_X = pd.DataFrame()
PCA_train_X = pd.concat([cyber_train_PCA, name_train_PCA, cities_train_PCA, spec_targ_train_PCA], axis = 1)
# + slideshow={"slide_type": "skip"}
# Test:
cyber_test_PCA = pd.DataFrame(cyber_test_PCA)
name_test_PCA = pd.DataFrame(name_test_PCA)
cities_test_PCA = pd.DataFrame(cities_test_PCA)
spec_targ_test_PCA = pd.DataFrame(spec_targ_test_PCA)
PCA_test_X = pd.DataFrame()
PCA_test_X = pd.concat([cyber_test_PCA, name_test_PCA, cities_test_PCA, spec_targ_test_PCA], axis = 1)
# + slideshow={"slide_type": "skip"}
del cyber_train_PCA, name_train_PCA, cities_train_PCA, spec_targ_train_PCA, cyber_test_PCA
del name_test_PCA, cities_test_PCA, spec_targ_test_PCA
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ### 4. Preliminary Models:
#
# Starting with a simple Logistic Regression since our output feature is binary. Afterwards, we will move on to our Random Forest, Support Vector Classifier and Gradient Booster. We chose these models since they often work well with binary output features. These models will also be discussed below in more detail.
# + slideshow={"slide_type": "skip"}
prelim_results = pd.DataFrame(columns=['Test B.A. Score', 'FP', 'FN', 'Mean CV Score', 'CV Std'])
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Logistic Regression:
#
# ##### KBest Features:
# + slideshow={"slide_type": "skip"}
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, balanced_accuracy_score
from sklearn.model_selection import cross_val_score
KBest_logistic = LogisticRegression()
KBest_logistic = KBest_logistic.fit(KBest_train_X, cyber_train_dummy_Y)
KBest_logistic_train_pred_ = KBest_logistic.predict(KBest_train_X)
KBest_logistic_test_pred_ = KBest_logistic.predict(KBest_test_X)
# Evaluation:
# Confustion Matrices:
KBest_logistic_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_logistic_train_pred_, labels = [0, 1])
KBest_logistic_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_logistic_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
KBest_logistic_cv = cross_val_score(KBest_logistic, KBest_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at balanced accuracy/f1 scores:
KBest_logistic_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_logistic_train_pred_)
KBest_logistic_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_logistic_test_pred_)
# + slideshow={"slide_type": "slide"}
plusminus = u"\u00B1"
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_logistic_cv.mean(),
plusminus, KBest_logistic_cv.std()))
print("The cv scores are: {}".format(KBest_logistic_cv))
conf_df = pd.DataFrame(KBest_logistic_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_logistic_train))
conf_df2 = pd.DataFrame(KBest_logistic_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_logistic_test))
prelim_results = prelim_results.append({'Test B.A. Score': KBest_logistic_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_logistic_cv.mean(),
'CV Std': KBest_logistic_cv.std()}, ignore_index = True)
# + slideshow={"slide_type": "subslide"}
fig, (ax1, ax2) = plt.subplots(nrows = 1, ncols = 2, figsize = (10, 4), sharey = True)
ax1 = sns.countplot(KBest_logistic_test_pred_, ax = ax1)
ax1.set_title("Prediction Results")
ax1.set_xticklabels(labels = ['Unsuccessful', 'Successful'])
ax2 = sns.countplot(cyber_test_Y.success, ax = ax2)
ax2.set_title("Actual Results")
ax2.set_xticklabels(labels = ['Unsuccessful', 'Successful'])
plt.show()
# + slideshow={"slide_type": "skip"}
del KBest_logistic, KBest_logistic_train_pred_, KBest_logistic_test_pred_ , KBest_logistic_confusion_train
del KBest_logistic_confusion_test, KBest_logistic_cv, KBest_logistic_train, KBest_logistic_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### PCA Features:
# + slideshow={"slide_type": "skip"}
PCA_logistic = LogisticRegression()
PCA_logistic = PCA_logistic.fit(PCA_train_X, cyber_train_dummy_Y)
PCA_logistic_train_pred_ = PCA_logistic.predict(PCA_train_X)
PCA_logistic_test_pred_ = PCA_logistic.predict(PCA_test_X)
# Evaluation:
# Confustion Matrices:
PCA_logistic_confusion_train = confusion_matrix(cyber_train_dummy_Y, PCA_logistic_train_pred_, labels = [0, 1])
PCA_logistic_confusion_test = confusion_matrix(cyber_test_dummy_Y, PCA_logistic_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
PCA_logistic_cv = cross_val_score(PCA_logistic, PCA_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at balanced accuracy/f1 scores:
PCA_logistic_train = balanced_accuracy_score(cyber_train_dummy_Y, PCA_logistic_train_pred_)
PCA_logistic_test = balanced_accuracy_score(cyber_test_dummy_Y, PCA_logistic_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(PCA_logistic_cv.mean(),
plusminus, PCA_logistic_cv.std()))
print("The cv scores are: {}".format(PCA_logistic_cv))
conf_df = pd.DataFrame(PCA_logistic_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(PCA_logistic_train))
conf_df2 = pd.DataFrame(PCA_logistic_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(PCA_logistic_test))
prelim_results = prelim_results.append({'Test B.A. Score': PCA_logistic_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': PCA_logistic_cv.mean(),
'CV Std': PCA_logistic_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "fragment"}
# Now, while we do have some decent scores here, especially regarding the true positive predictions, we do have a class imbalance issue we will need to take into consideration (as demonstrated below). One way of doing that will work on increasing the accuracy of our true negatives and focusing less on our true positives. It will also help to take into consideration our False Negatives and Positives. Lowering these will help strengthen our model and give us more predictive integrity. In other words, we want to make sure we avoid as many False Negatives (i.e. instances where our model does not predict a terrorist attack, when instead one does, in fact, occur) as possible. We also want to be sure we are keeping our False Positive count as low as possible since responding to predicted terrorist incidents when there are none will exhaust resources and employees - in turn taking away their energies from realistic threats.
#
# In order to do so, we have implemented the Balanced Accuracy score, which gives us an average of our False Negatives and Positives. It allows us to consider another aspect of the model results outside of the training and test-set scores (which only allow us to see on aspect of a model's predictive results). The Balanced Accuracy is a nice addition to the confusion matrix, which gives us the hard-numbers which are factored into the Balanced Accuracy score. In assessing the models above along with those below, we will be looking at all of these evaluation methods in order to determine which model is the best and make our reasoning as well-rounded as possible.
# + slideshow={"slide_type": "skip"}
print(cyber_train_dummy_Y.success.value_counts())
print(cyber_test_dummy_Y.success.value_counts())
# + slideshow={"slide_type": "skip"}
del PCA_logistic, PCA_logistic_train_pred_, PCA_logistic_test_pred_ , PCA_logistic_confusion_train
del PCA_logistic_confusion_test, PCA_logistic_cv, PCA_logistic_train, PCA_logistic_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# #### Random Forest:
#
# ##### KBest Features:
# + slideshow={"slide_type": "skip"}
from sklearn.ensemble import RandomForestClassifier
KBest_forest = RandomForestClassifier(n_estimators = 30, max_depth = 12)
KBest_forest = KBest_forest.fit(KBest_train_X, cyber_train_dummy_Y)
KBest_forest_train_pred_ = KBest_forest.predict(KBest_train_X)
KBest_forest_test_pred_ = KBest_forest.predict(KBest_test_X)
# Evaluation:
# Confustion Matrices:
KBest_forest_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_forest_train_pred_, labels = [0, 1])
KBest_forest_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_forest_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
KBest_forest_cv = cross_val_score(KBest_forest, KBest_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at balanced accuracy/f1 scores:
KBest_forest_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_forest_train_pred_)
KBest_forest_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_forest_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_forest_cv.mean(),
plusminus, KBest_forest_cv.std()))
print("The cv scores are: {}".format(KBest_forest_cv))
conf_df = pd.DataFrame(KBest_forest_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_forest_train))
conf_df2 = pd.DataFrame(KBest_forest_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_forest_test))
print('\n\nFeature Importances:\n')
feat_imp = pd.DataFrame(KBest_forest.feature_importances_.round(2), index = KBest_train_X.columns,
columns = ["Importances"])
display(feat_imp.sort_values('Importances', ascending = False))
prelim_results = prelim_results.append({'Test B.A. Score': KBest_forest_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_forest_cv.mean(),
'CV Std': KBest_forest_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "slide"}
# ##### PCA Features
# + slideshow={"slide_type": "skip"}
PCA_forest = RandomForestClassifier()
PCA_forest = PCA_forest.fit(PCA_train_X, cyber_train_dummy_Y)
PCA_forest_train_pred_ = PCA_forest.predict(PCA_train_X)
PCA_forest_test_pred_ = PCA_forest.predict(PCA_test_X)
# Evaluation:
# Confusion Matrices:
PCA_forest_confusion_train = confusion_matrix(cyber_train_dummy_Y, PCA_forest_train_pred_, labels = [0, 1])
PCA_forest_confusion_test = confusion_matrix(cyber_test_dummy_Y, PCA_forest_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
PCA_forest_cv = cross_val_score(PCA_forest, PCA_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking and balanced accuracy/f1 scores:
PCA_forest_train = balanced_accuracy_score(cyber_train_dummy_Y, PCA_forest_train_pred_)
PCA_forest_test = balanced_accuracy_score(cyber_test_dummy_Y, PCA_forest_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(PCA_forest_cv.mean(),
plusminus, PCA_forest_cv.std()))
print("The cv scores are: {}".format(PCA_forest_cv))
conf_df = pd.DataFrame(PCA_forest_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(PCA_forest_train))
conf_df2 = pd.DataFrame(PCA_forest_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(PCA_forest_test))
prelim_results = prelim_results.append({'Test B.A. Score': PCA_forest_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': PCA_forest_cv.mean(),
'CV Std': PCA_forest_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# Here, we can see quite a bit of over-fitting given that the training score and balanced accuracy scores are nearly perfect, while the test score is about 5 percentage points lower than the training score and over 20 points below the balanced accuracy (a drastic difference compared to what we have seen thus far). The nice aspect about the Random Forest, however, is that it lets us look into the feature importances which were used in creating the model, which we saw in the KBest model above.
# + slideshow={"slide_type": "skip"}
del PCA_forest, PCA_forest_train_pred_, PCA_forest_test_pred_ , PCA_forest_confusion_train,
del PCA_forest_confusion_test, PCA_forest_cv, PCA_forest_train, PCA_forest_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# #### Support Vector Classifier:
#
# ##### KBest Features:
# + slideshow={"slide_type": "skip"}
from sklearn.svm import SVC
KBest_SVC = SVC()
KBest_SVC = KBest_SVC.fit(KBest_train_X, cyber_train_dummy_Y)
KBest_SVC_train_pred_ = KBest_SVC.predict(KBest_train_X)
KBest_SVC_test_pred_ = KBest_SVC.predict(KBest_test_X)
# Evaluation:
# Confusion matrices:
KBest_SVC_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_SVC_train_pred_, labels = [0, 1])
KBest_SVC_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_SVC_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
KBest_SVC_cv = cross_val_score(KBest_SVC, KBest_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at balanced accuracy/f1 scores:
KBest_SVC_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_SVC_train_pred_)
KBest_SVC_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_SVC_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_SVC_cv.mean(),
plusminus, KBest_SVC_cv.std()))
print("The cv scores are: {}".format(KBest_SVC_cv))
conf_df = pd.DataFrame(KBest_SVC_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_SVC_train))
conf_df2 = pd.DataFrame(KBest_SVC_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_SVC_test))
prelim_results = prelim_results.append({'Test B.A. Score': KBest_SVC_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_SVC_cv.mean(),
'CV Std': KBest_SVC_cv.std()}, ignore_index = True)
# + slideshow={"slide_type": "skip"}
del KBest_SVC, KBest_SVC_train_pred_, KBest_SVC_test_pred_ , KBest_SVC_confusion_train
del KBest_SVC_confusion_test, KBest_SVC_cv, KBest_SVC_train, KBest_SVC_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### PCA Features
# + slideshow={"slide_type": "skip"}
PCA_SVC = SVC()
PCA_SVC = PCA_SVC.fit(PCA_train_X, cyber_train_dummy_Y)
PCA_SVC_train_pred_ = PCA_SVC.predict(PCA_train_X)
PCA_SVC_test_pred_ = PCA_SVC.predict(PCA_test_X)
# Evaluation:
# Confusion Matrices:
PCA_SVC_confusion_train = confusion_matrix(cyber_train_dummy_Y, PCA_SVC_train_pred_, labels = [0, 1])
PCA_SVC_confusion_test = confusion_matrix(cyber_test_dummy_Y, PCA_SVC_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
PCA_SVC_cv = cross_val_score(PCA_SVC, PCA_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at the F1/balanced accuracy scores:
PCA_SVC_train = balanced_accuracy_score(cyber_train_dummy_Y, PCA_SVC_train_pred_)
PCA_SVC_test = balanced_accuracy_score(cyber_test_dummy_Y, PCA_SVC_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(PCA_SVC_cv.mean(),
plusminus, PCA_SVC_cv.std()))
print("The cv scores are: {}".format(PCA_SVC_cv))
conf_df = pd.DataFrame(PCA_SVC_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(PCA_SVC_train))
conf_df2 = pd.DataFrame(PCA_SVC_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(PCA_SVC_test))
prelim_results = prelim_results.append({'Test B.A. Score': PCA_SVC_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': PCA_SVC_cv.mean(),
'CV Std': PCA_SVC_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "subslide"}
# These support vector models are considerably stable in the sense that they are able to achieve similar test scores throughout various rounds, which is good. The balanced accuracy scores are decent and the amount of error is considerably less than the models above. Let's take a look at one more round of model-type before we make this our official model.
# + slideshow={"slide_type": "skip"}
del PCA_SVC, PCA_SVC_train_pred_, PCA_SVC_test_pred_ , PCA_SVC_confusion_train
del PCA_SVC_confusion_test, PCA_SVC_cv, PCA_SVC_train, PCA_SVC_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# #### Gradient Boosting:
#
# ##### KBest Features:
# + slideshow={"slide_type": "skip"}
from sklearn.ensemble import GradientBoostingClassifier
KBest_GBC = GradientBoostingClassifier()
KBest_GBC = KBest_GBC.fit(KBest_train_X, cyber_train_dummy_Y)
KBest_GBC_train_pred_ = KBest_GBC.predict(KBest_train_X)
KBest_GBC_test_pred_ = KBest_GBC.predict(KBest_test_X)
# Evaluation:
# Confusion matrices:
KBest_GBC_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_GBC_train_pred_, labels = [0, 1])
KBest_GBC_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_GBC_test_pred_, labels = [0, 1])
# Cross-validation and train/test scores:
KBest_GBC_cv = cross_val_score(KBest_GBC, KBest_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at the F1/balanced-accuracy scores:
KBest_GBC_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_GBC_train_pred_)
KBest_GBC_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_GBC_test_pred_)
# + slideshow={"slide_type": "slide"}
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_GBC_cv.mean(),
plusminus, KBest_GBC_cv.std()))
print("The cv scores are: {}".format(KBest_GBC_cv))
conf_df = pd.DataFrame(KBest_GBC_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_GBC_train))
conf_df2 = pd.DataFrame(KBest_GBC_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_GBC_test))
print('\n\nFeature Importances:\n')
feat_imp = pd.DataFrame(KBest_GBC.feature_importances_.round(2), index = KBest_train_X.columns,
columns = ["Importances"])
display(feat_imp.sort_values('Importances', ascending = False))
prelim_results = prelim_results.append({'Test B.A. Score': KBest_GBC_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_GBC_cv.mean(),
'CV Std': KBest_GBC_cv.std()}, ignore_index = True)
# + slideshow={"slide_type": "skip"}
del KBest_GBC, KBest_GBC_train_pred_, KBest_GBC_test_pred_ , KBest_GBC_confusion_train
del KBest_GBC_confusion_test, KBest_GBC_cv, KBest_GBC_train, KBest_GBC_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"}
# ##### PCA Features
# + slideshow={"slide_type": "skip"}
PCA_GBC = GradientBoostingClassifier()
PCA_GBC = PCA_GBC.fit(PCA_train_X, cyber_train_dummy_Y)
PCA_GBC_train_pred_ = PCA_GBC.predict(PCA_train_X)
PCA_GBC_test_pred_ = PCA_GBC.predict(PCA_test_X)
# Evaluation:
# Confusion matrices:
PCA_GBC_confusion_train = confusion_matrix(cyber_train_dummy_Y, PCA_GBC_train_pred_, labels = [0, 1])
PCA_GBC_confusion_test = confusion_matrix(cyber_test_dummy_Y, PCA_GBC_test_pred_, labels = [0, 1])
# Cross_validation and train/test score:
PCA_GBC_cv = cross_val_score(PCA_GBC, PCA_train_X, cyber_train_dummy_Y,
scoring = 'balanced_accuracy', cv = 5)
# Looking at the F1/balanced accuracy scores:
PCA_GBC_train = balanced_accuracy_score(cyber_train_dummy_Y, PCA_GBC_train_pred_)
PCA_GBC_test = balanced_accuracy_score(cyber_test_dummy_Y, PCA_GBC_test_pred_)
# + slideshow={"slide_type": "slide"}
plusminus = u"\u00B1"
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(PCA_GBC_cv.mean(),
plusminus, PCA_GBC_cv.std()))
print("The cv scores are: {}".format(PCA_GBC_cv))
conf_df = pd.DataFrame(PCA_GBC_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(PCA_GBC_train))
conf_df2 = pd.DataFrame(PCA_GBC_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(PCA_GBC_test))
prelim_results = prelim_results.append({'Test B.A. Score': PCA_GBC_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': PCA_GBC_cv.mean(),
'CV Std': PCA_GBC_cv.std()}, ignore_index = True)
# + slideshow={"slide_type": "skip"}
del PCA_GBC, PCA_GBC_train_pred_, PCA_GBC_test_pred_ , PCA_GBC_confusion_train
del PCA_GBC_confusion_test, PCA_GBC_cv, PCA_GBC_train, PCA_GBC_test
gc.collect()
# + [markdown] slideshow={"slide_type": "slide"} variables={"prelim_results": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>Test B.A. Score</th>\n <th>FP</th>\n <th>FN</th>\n <th>Mean CV Score</th>\n <th>CV Std</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>0.564348</td>\n <td>184.0</td>\n <td>7.0</td>\n <td>0.573343</td>\n <td>0.007393</td>\n </tr>\n <tr>\n <th>1</th>\n <td>0.551107</td>\n <td>189.0</td>\n <td>13.0</td>\n <td>0.556829</td>\n <td>0.005586</td>\n </tr>\n <tr>\n <th>2</th>\n <td>0.528060</td>\n <td>200.0</td>\n <td>1.0</td>\n <td>0.560620</td>\n <td>0.005684</td>\n </tr>\n <tr>\n <th>3</th>\n <td>0.633375</td>\n <td>149.0</td>\n <td>63.0</td>\n <td>0.641248</td>\n <td>0.017357</td>\n </tr>\n <tr>\n <th>4</th>\n <td>0.500000</td>\n <td>212.0</td>\n <td>0.0</td>\n <td>0.500000</td>\n <td>0.000000</td>\n </tr>\n <tr>\n <th>5</th>\n <td>0.500000</td>\n <td>212.0</td>\n <td>0.0</td>\n <td>0.500000</td>\n <td>0.000000</td>\n </tr>\n <tr>\n <th>6</th>\n <td>0.571182</td>\n <td>181.0</td>\n <td>8.0</td>\n <td>0.578078</td>\n <td>0.017042</td>\n </tr>\n <tr>\n <th>7</th>\n <td>0.598945</td>\n <td>168.0</td>\n <td>20.0</td>\n <td>0.602553</td>\n <td>0.007509</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# #### Summary of First Models:
#
# > Based on these preliminary results, we were able to get relatively similar results from nearly all models, but it looks like our Random Forest Classifier did best (with the Support Vector Classifier coming in at a close second). In addition, the Random Forest model did best with the KBest feature-set. We will try and focus on the Random Forest model with our next phase and tune the parameters to see if we can optimize those results.
#
# > Results Table:
#
# {{prelim_results}}
#
# > Now onto tuning our model!
# + slideshow={"slide_type": "skip"}
del PCA_spec_targ, PCA_name, PCA_cities, PCA_1, KBest_spec_targ, KBest_names, conf_df, conf_df2
del KBest_city, KBest_1, city_mask, feature_mask, names_mask, spec_targ_mask, feature_selection
del KBest_spec_targ_features, KBest_city_features, KBest_names_features, KBest_1_features
del spec_targ_test_index, city_test_dummy_Y, names_test_dummy_Y, spec_targ_test_dummy_Y, spec_targ_test_dummy_X
del X_test_start_index, city_test_index, names_test_index, spec_targ_train_index, city_train_dummy_Y
del names_train_dummy_Y, spec_targ_train_dummy_Y, spec_targ_train_dummy_X, city_train_index, names_train_index
del names_test_dummy_X, names_train_X, city_train_X, spec_targ_train_X, names_test_X, city_test_X, spec_targ_test_X
del PosT2, PosT, names_train_dummy_X, city_test_dummy_X, city_train_dummy_X
gc.collect()
# + slideshow={"slide_type": "skip"}
# These are the usual ipython objects, including this one you are creating
ipython_vars = ['In', 'Out', 'exit', 'quit', 'get_ipython', 'ipython_vars']
# Get a sorted list of the objects and their sizes
cleaner = sorted([(x, sys.getsizeof(globals().get(x))) for x in dir() if not x.startswith('_') and x not in sys.modules and x not in ipython_vars], key=lambda x: x[1], reverse=True)
# + [markdown] slideshow={"slide_type": "slide"}
# ### 5. Improving our Scores:
#
# #### A. Investigating the Data Itself:
#
#
# + slideshow={"slide_type": "skip"}
def RF_KBest_Eval():
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_forest_cv.mean(),
plusminus, KBest_forest_cv.std()))
print("The cv scores are: {}".format(KBest_forest_cv))
conf_df = pd.DataFrame(KBest_forest_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("\nTraining-Set Metrics:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_forest_train))
conf_df2 = pd.DataFrame(KBest_forest_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test-Set Metrics:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_forest_test))
print('\n\nFeature Importances:\n')
feat_imp = pd.DataFrame(KBest_forest.feature_importances_.round(2), index = KBest_train_X.columns,
columns = ["Importances"])
display(feat_imp.sort_values('Importances', ascending = False))
# + slideshow={"slide_type": "fragment"}
RF_KBest_Eval()
# + slideshow={"slide_type": "skip"}
# our comm-related df:
comms = cyber_data.loc[cyber_data['specific_target'] == 'Comm-Related']
# Isolating Assassination instances (in our visualization dataframe so we can look at how our outcome variable
# correlates with them):
attacks_as = cyber_train_X.loc[cyber_train_X['attack_1txt'] == 'Assassination']
# Retrieving our 'success' columns for comparison:
success_mask = cyber_train_Y.index.map(lambda x: x in attacks_as.index)
s_col = cyber_train_Y[success_mask]
attacks_as['success'] = s_col['success']
attacks_success = pd.DataFrame(attacks_as.success.value_counts())
## Locating the successful instances:
success_as = attacks_as.loc[attacks_as['success'] == 1]
fails_as = attacks_as.loc[attacks_as['success'] == 0]
# + [markdown] slideshow={"slide_type": "subslide"} variables={"attacks_success": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>success</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>1</th>\n <td>601</td>\n </tr>\n <tr>\n <th>0</th>\n <td>307</td>\n </tr>\n </tbody>\n</table>\n</div>", "pd.DataFrame(cyber_data.success.value_counts())": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>success</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>1</th>\n <td>11994</td>\n </tr>\n <tr>\n <th>0</th>\n <td>1261</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Now that we've seen what our preliminary model gave as our most important feature (i.e. where the attack type is an assassination), we have a difficult challenge on our hands since most of our frequent values in a handful of features are 'Unknown.' How does one predict given that our most-common evidence falls into an 'Unknown' category? Let's look at the data for within the umbrella 'assassination attacks' and check out some class-balances.
#
# <head>
# <table>
# <th style = 'text-align:center'> Entire Data-set </th>
# <th style = 'text-align:center'> Data Grouped by Assassination </th>
# <tr>
# <td> {{pd.DataFrame(cyber_data.success.value_counts())}} </td>
# <td> {{attacks_success}} </td>
# </tr>
# </table>
# </head>
#
# Here, we can see that we went from a 90% imbalance to a 50% imbalance and a distribution where our unsuccessful attacks represent one-sixth of their entire class.
# + [markdown] slideshow={"slide_type": "subslide"} variables={"fails_as.describe(include = 'O')": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>country_txt</th>\n <th>region_txt</th>\n <th>city</th>\n <th>attack_1txt</th>\n <th>target_1txt</th>\n <th>sub_targettxt</th>\n <th>corp1</th>\n <th>specific_target</th>\n <th>victim_nationalitytxt</th>\n <th>group_name</th>\n <th>weapontxt</th>\n <th>sub_weapontxt</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>count</th>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n <td>307</td>\n </tr>\n <tr>\n <th>unique</th>\n <td>34</td>\n <td>9</td>\n <td>198</td>\n <td>1</td>\n <td>14</td>\n <td>24</td>\n <td>189</td>\n <td>293</td>\n <td>38</td>\n <td>35</td>\n <td>5</td>\n <td>17</td>\n </tr>\n <tr>\n <th>top</th>\n <td>Iraq</td>\n <td>South Asia</td>\n <td>Sunni_Cities</td>\n <td>Assassination</td>\n <td>Government (General)</td>\n <td>Gov_Figure1</td>\n <td>Iraqi Police Service (IPS)</td>\n <td>Commander</td>\n <td>Iraq</td>\n <td>Unknown</td>\n <td>Explosives</td>\n <td>Unknown Gun Type</td>\n </tr>\n <tr>\n <th>freq</th>\n <td>68</td>\n <td>120</td>\n <td>33</td>\n <td>307</td>\n <td>100</td>\n <td>57</td>\n <td>20</td>\n <td>6</td>\n <td>68</td>\n <td>180</td>\n <td>190</td>\n <td>74</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Here is a closer look at Assassination attacks in our training set that failed:
#
# {{fails_as.describe(include = 'O')}}
# + [markdown] slideshow={"slide_type": "subslide"} variables={"fails_as.loc[fails_as['group_name'] == 'Unknown'].describe(include = 'O')": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>country_txt</th>\n <th>region_txt</th>\n <th>city</th>\n <th>attack_1txt</th>\n <th>target_1txt</th>\n <th>sub_targettxt</th>\n <th>corp1</th>\n <th>specific_target</th>\n <th>victim_nationalitytxt</th>\n <th>group_name</th>\n <th>weapontxt</th>\n <th>sub_weapontxt</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>count</th>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n <td>180</td>\n </tr>\n <tr>\n <th>unique</th>\n <td>24</td>\n <td>9</td>\n <td>113</td>\n <td>1</td>\n <td>10</td>\n <td>19</td>\n <td>115</td>\n <td>173</td>\n <td>25</td>\n <td>1</td>\n <td>4</td>\n <td>15</td>\n </tr>\n <tr>\n <th>top</th>\n <td>Iraq</td>\n <td>Middle East & North Africa</td>\n <td>Sunni_Cities</td>\n <td>Assassination</td>\n <td>Government (General)</td>\n <td>Police Security Forces/Officers</td>\n <td>Iraqi Police Service (IPS)</td>\n <td>Commander</td>\n <td>Iraq</td>\n <td>Unknown</td>\n <td>Explosives</td>\n <td>Unknown Gun Type</td>\n </tr>\n <tr>\n <th>freq</th>\n <td>57</td>\n <td>79</td>\n <td>27</td>\n <td>180</td>\n <td>56</td>\n <td>35</td>\n <td>20</td>\n <td>5</td>\n <td>57</td>\n <td>180</td>\n <td>110</td>\n <td>49</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# ...and a closer look at the same set above, with an additional filter: 'Unknown group-names'. Here, it is important to note the very slight difference in the frequency of the feature 'Sunni_cities':
# {{fails_as.loc[fails_as['group_name'] == 'Unknown'].describe(include = 'O')}}
# + [markdown] slideshow={"slide_type": "subslide"} variables={"pd.DataFrame(attacks_as.loc[(attacks_as['city'] == 'Sunni_Cities')].success.value_counts())": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>success</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>0</th>\n <td>33</td>\n </tr>\n <tr>\n <th>1</th>\n <td>18</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Looking at failed assassination attempts within Sunni_cities, we see our class imbalance has switched:
#
# {{pd.DataFrame(attacks_as.loc[(attacks_as['city'] == 'Sunni_Cities')].success.value_counts())}}
#
# + [markdown] slideshow={"slide_type": "fragment"} variables={"pd.DataFrame(fails_as.city.value_counts().head(5))": "<div>\n<style scoped>\n .dataframe tbody tr th:only-of-type {\n vertical-align: middle;\n }\n\n .dataframe tbody tr th {\n vertical-align: top;\n }\n\n .dataframe thead th {\n text-align: right;\n }\n</style>\n<table border=\"1\" class=\"dataframe\">\n <thead>\n <tr style=\"text-align: right;\">\n <th></th>\n <th>city</th>\n </tr>\n </thead>\n <tbody>\n <tr>\n <th>Sunni_Cities</th>\n <td>33</td>\n </tr>\n <tr>\n <th>Unknown</th>\n <td>14</td>\n </tr>\n <tr>\n <th>Mogadishu</th>\n <td>14</td>\n </tr>\n <tr>\n <th>Split_Cities</th>\n <td>13</td>\n </tr>\n <tr>\n <th>Benghazi</th>\n <td>5</td>\n </tr>\n </tbody>\n</table>\n</div>"}
# Here are our top 5 cities in this scenario:
#
# {{pd.DataFrame(fails_as.city.value_counts().head(5))}}
# + slideshow={"slide_type": "subslide"}
cyber_train_X.loc[(cyber_train_X['city'] == 'Sunni_Cities') |
(cyber_train_X['city'] == 'Mogadishu') &
(cyber_train_X['attack_1txt'] == 'Assassination'), 'Barometer'] = 1
Barometer_train = cyber_train_X['Barometer'].replace(np.nan, 0)
Barometer_train = Barometer_train.reset_index().drop('index', axis = 1)
# + slideshow={"slide_type": "skip"}
cyber_test_X.loc[(cyber_test_X['city'] == 'Sunni_Cities') |
(cyber_test_X['city'] == 'Mogadishu') &
(cyber_test_X['attack_1txt'] == 'Assassination'), 'Barometer'] = 1
Barometer_test = cyber_test_X['Barometer'].replace(np.nan, 0)
Barometer_test = Barometer_test.reset_index().drop('index', axis = 1)
# + slideshow={"slide_type": "fragment"}
KBest_train_X3 = KBest_train_X
KBest_test_X3 = KBest_test_X
KBest_train_X3 = pd.concat([KBest_train_X3, Barometer_train], axis = 1)
KBest_test_X3 = pd.concat([KBest_test_X3, Barometer_test], axis = 1)
# + [markdown] slideshow={"slide_type": "slide"}
# ### 6. Tuning our Final Model:
#
# Here, we will try and use a for loop to run through a set of parameters, which we will then use to visualize the parameters which will optimize our model's predictive power. For the moment, the parameters we will focus on are the 'n_estimators' and 'max_depth.'
# + slideshow={"slide_type": "fragment"}
# Double checking the shape of our engineered feature-set:
print(KBest_train_X3.shape)
print(KBest_test_X3.shape)
# + [markdown] slideshow={"slide_type": "slide"}
# #### A. Using GridSearchCV
# + slideshow={"slide_type": "fragment"}
from sklearn.model_selection import GridSearchCV
KBest_Grid = RandomForestClassifier(criterion = 'entropy')
params = [{'n_estimators': np.arange(40, 300, 20),
'max_depth': np.arange(3, 16, 1)}]
params_2 = [{'n_estimators': np.arange(300, 20, -20),
'max_depth': np.arange(15, 2, -1)}]
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### 1. Random Forest with Original Data-Set:
# + slideshow={"slide_type": "fragment"}
forest_grid = GridSearchCV(KBest_Grid, params, cv = 5, scoring = 'balanced_accuracy')
forest_grid.fit(KBest_train_X, cyber_train_dummy_Y)
# + slideshow={"slide_type": "skip"}
# These are the usual ipython objects, including this one you are creating
ipython_vars = ['In', 'Out', 'exit', 'quit', 'get_ipython', 'ipython_vars']
# Get a sorted list of the objects and their sizes
cleaner = sorted([(x, sys.getsizeof(globals().get(x))) for x in dir() if not x.startswith('_') and x not in sys.modules and x not in ipython_vars], key=lambda x: x[1], reverse=True)
# + slideshow={"slide_type": "skip"}
del dummy_prep, names_dummy_1, names_dummy_prep, spec_targ_dummy_prep, PCA_test_X, PCA_train_X
del attacks_as, comms, success_mask, big_targets_mask
# + slideshow={"slide_type": "skip"}
gc.collect()
# + slideshow={"slide_type": "subslide"}
forest_grid_2 = GridSearchCV(KBest_Grid, params_2, cv = 5, scoring = 'balanced_accuracy')
forest_grid_2.fit(KBest_train_X, cyber_train_dummy_Y)
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### 2. Random Forest with New Feature:
# + slideshow={"slide_type": "fragment"}
forest_grid_3 = GridSearchCV(KBest_Grid, params, cv = 5, scoring = 'balanced_accuracy')
forest_grid_3.fit(KBest_train_X3, cyber_train_dummy_Y)
# + slideshow={"slide_type": "subslide"}
forest_grid_4 = GridSearchCV(KBest_Grid, params_2, cv = 5, scoring = 'balanced_accuracy')
forest_grid_4.fit(KBest_train_X3, cyber_train_dummy_Y)
# + slideshow={"slide_type": "subslide"}
print(forest_grid.best_params_)
print(forest_grid_2.best_params_)
print(forest_grid_3.best_params_)
print(forest_grid_4.best_params_)
# + slideshow={"slide_type": "fragment"}
print(forest_grid.best_score_)
print(forest_grid_2.best_score_)
print(forest_grid_3.best_score_)
print(forest_grid_4.best_score_)
# + [markdown] slideshow={"slide_type": "slide"}
# #### B. Custom GridSearch:
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### 1. RandomForest with KBest (original features):
# + slideshow={"slide_type": "fragment"}
param_dict = {'n_estimators': [80, 100, 120, 80, 100, 120],
'max_depth' : [15, 16, 17, 13, 14, 12]}
param_df = pd.DataFrame(param_dict)
# + slideshow={"slide_type": "subslide"}
results_list = pd.DataFrame(columns=['B.A. Score', 'FP', 'FN', 'Mean CV Score', 'CV Std'])
plusminus = u"\u00B1"
for row in param_df.itertuples(index = False):
KBest_RF = RandomForestClassifier(n_estimators = row[0], max_depth = row[1])
KBest_RF = KBest_RF.fit(KBest_train_X, cyber_train_dummy_Y)
KBest_RF_train_pred_ = KBest_RF.predict(KBest_train_X)
KBest_RF_test_pred_ = KBest_RF.predict(KBest_test_X)
# Evaluation:
# Confusion matrices:
KBest_RF_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_RF_train_pred_, labels = [0, 1])
KBest_RF_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_RF_test_pred_, labels = [0, 1])
# Cross_validation and train/test score:
KBest_RF_cv = cross_val_score(KBest_RF, KBest_train_X, cyber_train_dummy_Y, cv = 5,
scoring = 'balanced_accuracy')
# Looking at the F1/balanced accuracy scores:
KBest_RF_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_RF_train_pred_)
KBest_RF_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_RF_test_pred_)
# Getting some scores on cross-validation, False Negatives and Positives and Balanced Accuracy:
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_RF_cv.mean(),
plusminus, KBest_RF_cv.std()))
conf_df = pd.DataFrame(KBest_RF_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("Training set results:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_RF_train))
conf_df2 = pd.DataFrame(KBest_RF_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test set results:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_RF_test))
print('-----------------')
results_list = results_list.append({'B.A. Score': KBest_RF_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_RF_cv.mean(),
'CV Std': KBest_RF_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "subslide"} variables={"results_list": {}}
# Here are the results we received for Random Forest Model (with tweaked parameters and our engineered feature):
#
# {{results_list}}
# + [markdown] slideshow={"slide_type": "slide"}
# ##### 2. RandomForest with KBest (engineered feature added):
# + slideshow={"slide_type": "subslide"}
results_list_2 = pd.DataFrame(columns=['B.A. Score', 'FP', 'FN', 'Mean CV Score', 'CV Std'])
plusminus = u"\u00B1"
for row in param_df.itertuples(index = False):
KBest_RF = RandomForestClassifier(n_estimators = row[0], max_depth = row[1])
KBest_RF = KBest_RF.fit(KBest_train_X3, cyber_train_dummy_Y)
KBest_RF_train_pred_ = KBest_RF.predict(KBest_train_X3)
KBest_RF_test_pred_ = KBest_RF.predict(KBest_test_X3)
# Evaluation:
# Confusion matrices:
KBest_RF_confusion_train = confusion_matrix(cyber_train_dummy_Y, KBest_RF_train_pred_, labels = [0, 1])
KBest_RF_confusion_test = confusion_matrix(cyber_test_dummy_Y, KBest_RF_test_pred_, labels = [0, 1])
# Cross_validation and train/test score:
KBest_RF_cv = cross_val_score(KBest_RF, KBest_train_X3, cyber_train_dummy_Y, cv = 5,
scoring = 'balanced_accuracy')
# Looking at the F1/balanced accuracy scores:
KBest_RF_train = balanced_accuracy_score(cyber_train_dummy_Y, KBest_RF_train_pred_)
KBest_RF_test = balanced_accuracy_score(cyber_test_dummy_Y, KBest_RF_test_pred_)
# Getting some scores on cross-validation, False Negatives and Positives and Balanced Accuracy:
print("\nThe mean cross-validation score is: {:.2%} {}{:.2%}".format(KBest_RF_cv.mean(),
plusminus, KBest_RF_cv.std()))
conf_df = pd.DataFrame(KBest_RF_confusion_train)
FP = conf_df.loc[0, 1]
FN = conf_df.loc[1, 0]
NegT = conf_df.iloc[0].sum()
PosT = conf_df.iloc[1].sum()
print("Training set results:")
print(conf_df)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP, (FP / NegT)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN, (FN / PosT)))
print('Balanced Accuracy: {:.2%}\n'.format(KBest_RF_train))
conf_df2 = pd.DataFrame(KBest_RF_confusion_test)
FP2 = conf_df2.loc[0, 1]
FN2 = conf_df2.loc[1, 0]
NegT2 = conf_df2.iloc[0].sum()
PosT2 = conf_df2.iloc[1].sum()
print("Test set results:")
print(conf_df2)
print('\nFalse Positive/Type I Error: {} ({:.2%})'.format(FP2, (FP2 / NegT2)))
print('False Negative/Type II Error: {} ({:.2%})'.format(FN2, (FN2 / PosT2)))
print('Balanced Accuracy: {:.2%}'.format(KBest_RF_test))
print('-----------------')
results_list_2 = results_list_2.append({'B.A. Score': KBest_RF_test,
'FP': FP2,
'FN': FN2,
'Mean CV Score': KBest_RF_cv.mean(),
'CV Std': KBest_RF_cv.std()}, ignore_index = True)
# + [markdown] slideshow={"slide_type": "subslide"} variables={"results_list_2": {}}
# Here are the results we received for Random Forest Model (with tweaked parameters and our engineered feature):
#
# {{results_list_2}}
# + [markdown] slideshow={"slide_type": "slide"}
# ### 7. Final Analysis, Considerations and Avenues for Further Research:
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Potential Weaknesses:
#
# While we were able to get some decent scores, there are a few things to not regarding this model and its dataset. First of all, the filters would greatly benefit from further research and detailed attention. The above filters are merely a start and represent a base amount of research. It could certainly serve as a starting point for someone with expertise in the field.
#
# In addition, the filters we applied could be susceptible to a certain amount of bias: the filter itself will not be able to catch all instances of cyber-related terrorism conclusively; more likely than not it will miss one certain aspect or another. Increasing the number of eyes on the model and data will help with this.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Further research:
#
# Given the richness of the variables within this dataset, there are plenty of avenues for further research. One could use Regex to search the summary and motive columns for further detail and insight into the nature of these cyber-related tasks. The filter above is relatively broad and potentially encapsulates instances that many might not consider related to 'cyber' events. It would be interesting to create a more intricate regex filter which could give us a more detailed understanding of the 'cyber' aspect of these events: How specifically are they related and in what manners? What geographical locations tend to be hotbeds for such activity. What targets are specified and why? The 'motive' feature, in particular, could have extensive benefits with prediction, depending on the vocabulary used by those conducting the study and entering the data.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Explanatory v. Predictive Power:
#
# With the current analysis, our goal lies mainly in the predictive power of our model - in other words, its ability to fit with the current data and produce an output with completely new data - preferably an output with as little variation in this new accuracy score as possible. We are trying to look at the trends in our current data and use it to identify the classifications and probabilities in which new or future observations will fall. An explanatory model, with regards to a dataset describing Terrorism, might have interest to a scholar or an academic institution in the sense that it reveals behaviors and patterns of observations that have already occurred, but it has no bearing on future observations. It would offer no suggestions on what these patterns might say regarding incoming data (and in this case, future terror attacks).
#
# The caveat, however, is making sure that our model adapts well to new input (or test) data so that the variation between training and test results can be as minimal as possible (i.e. so the difference between the model's prediction and reality is minimal). Predictive analysis, especially in a terror-related context, involves considerably more risk than an explanatory model, and should be handled with an intense attention to detail and accuracy.
#
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# General References:
#
# Referenced the following sites for honing my knowledge of the models, python, etc:
#
# 1. https://www.analyticsvidhya.com/blog/2017/06/a-comprehensive-guide-for-linear-ridge-and-lasso-regression/
# 2. https://machinelearningmastery.com/feature-selection-machine-learning-python/
# 3. https://medium.com/@pushkarmandot/what-is-the-significance-of-c-value-in-support-vector-machine-28224e852c5a
# 4. A big shout out to <NAME> for pointing out this link to me (submitted by the user 'Abdou' on Stack Overflow: https://stackoverflow.com/questions/40993626/list-memory-usage-in-ipython-and-jupyter)
| Unit_3_Capstone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="AzA7D_5U4ZCb"
# 論文
# https://arxiv.org/abs/2112.05131
#
# GitHub
# https://github.com/sxyu/svox2
#
# <a href="https://colab.research.google.com/github/kaz12tech/ai_demos/blob/master/Plenoxels_demo.ipynb" target="_blank"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="y_L3U0xl4ZCe"
# # ランタイムの設定
# 「ランタイム」→「ランタイムのタイプを変更」→「ハードウェアアクセラレータ」をGPUに変更
# + [markdown] id="ICKFXCTV4ZCg"
# # 実行方法
# 「ランタイム」→「すべてのセルを実行」を選択
# + [markdown] id="P-jy3f_w4ZCg"
# # GPU確認
# + colab={"base_uri": "https://localhost:8080/"} id="kNjB8L3X4ZCh" outputId="50e6b09c-2dda-49a3-dd66-78ff7bbc3103"
# !nvidia-smi
# + [markdown] id="DZ03KqwrGof0"
# # GoogleDriveのマウント
# + colab={"base_uri": "https://localhost:8080/"} id="DQvO6TunGn05" outputId="46377b2a-6c79-4c90-f5b4-00f56934b728"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="UKRsZxQs4ZCi"
# # GitHubからPlenoxelsのソースコードを取得
# + colab={"base_uri": "https://localhost:8080/"} id="8lDsOunm4ZCj" outputId="ab45525d-62f0-4162-e450-be0a31d401ac"
# %cd /content/
# !git clone https://github.com/sxyu/svox2.git
# + id="m96BO0z1_JX6"
# !sed -E -i "s/\{minv=:/minv=\{minv:/g" /content/svox2/opt/opt.py
# !sed -E -i "s/\{meanv=:/meanv=\{meanv:/g" /content/svox2/opt/opt.py
# !sed -E -i "s/\{maxv=:/maxv=\{maxv:/g" /content/svox2/opt/opt.py
# !sed -E -i "s/\{minv=:/minv=\{minv:/g" /content/svox2/opt/render_imgs.py
# !sed -E -i "s/\{meanv=:/meanv=\{meanv:/g" /content/svox2/opt/render_imgs.py
# !sed -E -i "s/\{maxv=:/maxv=\{maxv:/g" /content/svox2/opt/render_imgs.py
# + [markdown] id="qbF9na1649Cg"
# # ライブラリのインストール
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="7NNxBjDN475O" outputId="b090b381-650b-47f9-be4a-2f638bc8ce0b"
# %cd /content/svox2
# !apt install ninja-build -y
# !pip install imageio-ffmpeg
# !pip install ipdb
# !pip install lpips
# !pip install pymcubes
# !pip install .
# + [markdown] id="A1-I1eWCMESQ"
# # ライブラリのインポート
# + colab={"base_uri": "https://localhost:8080/"} id="S61FL0CZMGcj" outputId="ccf15c77-1347-45c3-ccc5-0273e140e1cd"
import os
from moviepy.editor import *
from moviepy.video.fx.resize import resize
# + [markdown] id="a0TcvrZo6Brm"
# # 学習済みモデルのダウンロード
# gdownを使用してダウンロードできないため
# https://drive.google.com/drive/folders/1SOEJDw8mot7kf5viUK9XryOAmZGe_vvE のckpt_syn.tar.gzを右クリックし「ドライブにショートカットを追加」を選択し、マイドライブを選択後、「ショートカットを追加」
# 自身のGoogle Driveにショートカットを作成
# + colab={"base_uri": "https://localhost:8080/"} id="ukIsH0C96D52" outputId="4ff5eaaa-263c-4b11-f7fe-71c5ebd6e16f"
# %cd /content/svox2
# !mkdir ckpt
# %cd ckpt
if not os.path.exists("/content/svox2/ckpt/ckpt_syn.tar.gz"):
# !cp /content/drive/MyDrive/ckpt_syn.tar.gz /content/svox2/ckpt
# !tar -xvf ./ckpt_syn.tar.gz > /dev/null
# + [markdown] id="dRAI3ZBqKg_I"
# # データセットのダウンロード
# gdownを使用してダウンロードできないため
# https://drive.google.com/drive/folders/128yBriW1IG_3NJ5Rp7APSTZsJqdJdfc1 のnerf_synthetic.zipを右クリックし「ドライブにショートカットを追加」を選択し、マイドライブを選択後、「ショートカットを追加」
# 自身のGoogle Driveにショートカットを作成
# + colab={"base_uri": "https://localhost:8080/"} id="XnEAYnLEKeya" outputId="a2268186-3118-4f2a-925e-f5123cf28aff"
# %cd /content/svox2/
# !mkdir datasets
# %cd datasets
if not os.path.exists("/content/svox2/datasets/nerf_synthetic.zip"):
# !cp /content/drive/MyDrive/nerf_synthetic.zip /content/svox2/datasets/
# !unzip ./nerf_synthetic.zip > /dev/null
# + [markdown] id="LXh32gxmI2Ev"
# # 自由視点画像の生成
# + colab={"base_uri": "https://localhost:8080/"} id="p3rpWSs0I6mI" outputId="8b1b5872-0854-46d5-aea7-23fc8966a6cf"
# %cd /content/svox2/
# !mkdir -p /content/svox2/results/hotdog
# !cp /content/svox2/ckpt/256_to_512_fasttv/hotdog/ckpt.npz /content/svox2/results/hotdog
# + colab={"base_uri": "https://localhost:8080/"} id="6B5_P2DPcQ45" outputId="9bdfff01-5d83-475b-8c15-5f6667083722"
# %%time
# %cd /content/svox2/opt
# !python render_imgs.py /content/svox2/results/hotdog/ckpt.npz /content/svox2/datasets/nerf_synthetic/hotdog/
# + [markdown] id="ANrHoGs5DOqo"
# ## 自由視点画像の表示
# + colab={"base_uri": "https://localhost:8080/", "height": 476} id="1bdKvRmqDTzZ" outputId="8dacb23a-bfaf-482b-ead7-4ee8aef18ba6"
# %cd /content/svox2/
results_video = "/content/svox2/results/hotdog/test_renders.mp4"
clip = VideoFileClip(results_video)
clip = resize(clip, height=420)
clip.ipython_display()
# + colab={"base_uri": "https://localhost:8080/"} id="P7R2PSiJe6e0" outputId="11d9feb8-0a41-4cb6-c04e-87ee4c6cb36c"
# %cd /content/svox2/
# !mkdir -p /content/svox2/results/materials
# !cp /content/svox2/ckpt/256_to_512_fasttv/materials/ckpt.npz /content/svox2/results/materials
# + colab={"base_uri": "https://localhost:8080/"} id="AFejSEtTfNN5" outputId="ce771380-c5a0-40da-d067-47b8680ea3d3"
# %%time
# %cd /content/svox2/opt
# !python render_imgs_circle.py /content/svox2/results/materials/ckpt.npz /content/svox2/datasets/nerf_synthetic/materials/
# + colab={"base_uri": "https://localhost:8080/", "height": 476} id="bU9iUUMmfXjT" outputId="9ca0ee6e-8063-4e0d-917f-8d77fc2f04bf"
# %cd /content/svox2/
results_video = "/content/svox2/results/materials/circle_renders.mp4"
clip = VideoFileClip(results_video)
clip = resize(clip, height=420)
clip.ipython_display()
| Plenoxels_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Strategy-proof monetary anchoring
# ============
# *Please allow Binder to be loaded (<1 min), then you can look at the code, change parameters and run cell by cell as in any Jupyter Notebook (Maj+Enter to run each cell once, or use menu shortcut to run all cells under Cell > "Run All").*
#
# --
# ### Motivation
#
# Mundell’s redundancy (or "n-1") problem illustrates how international finance can be interpreted by some as a zero-sum game, with many prisoners dilemma problems. Such a view logically leads some countries to try to "game" the international monetary system, first and foremost through their managed floating currencies (hence the vocabulary around "manipulation" on what concerns exchange rate levels).
#
#
# Here we provide a model and simulations in an interactive Notebook format to load and play directly with the Matlab code **to examine how international monetary systems relying on specified nominal anchors (silver and gold, sterling then USD, bancor sometimes maybe)** - though the most "natural" form of anchoring, as it provides a solution to commitment and reneging issues - **create incentives that might be correlated with hegemonic wars** *(political scientists - Wallerstein, Modelski, Organski or more recently Goldstein - have long explored these cycles, but economists also advanced some causal hypothesis. <NAME> in his "Forme di fenomeni economici e previsioni" for instance postulated that ascending and descending phases in international trade are linked to more or less available saving levels, and that period of social unrest and cross-country wars occur in such descending phases)*.
#
#
# --
# ### Safe asset shortage or financial cycle buildup - the two sides of the same (anchor-) coin ?
#
# Since the dollar unpegged from gold and some currencies started to float, exchange rate manipulation has been tolerated up to some point - principally because of the increased difficulty in pinpointing exactly what the "fair" parity between the two legs of a currency pair would be, which itself would influence in return all other currency pairs these two legs are involved in.
#
# The term "fair" itself changed meaning, between a Gold Standard world and one in which the global loosening of the monetary anchoring (no rules determines how much USD can or should be printed to optimize the global economy) and in which cooperation (or the lack of) between central banks determine world interest rates.
#
# **The exchange rate indeterminancy in today's world might be quite exarcebated by such a loosening of the anchoring**, as the seminal result by <NAME> Wallace (1981) that the nominal exchange rate is indeterminate in a world with pure interest rate targets has been extended (Caballero, Farhi, Gourinchas, 2015) to the case when the global economy is in a liquidity trap. However, while there are now several competing theories aiming at explaining the secular decline in global real interest rates over much of the past 30 years, none of them explicitly examines how the anchoring of the international monetary system might (or might not) have played a role in causing this trend.
#
# **That is why we will try to provide here a framework based on modelling the anchoring as a mechanism design problem**, to explicit parameters that changed between now and the Gold Standard, before/after 1971, and which might provide a framework linking the safe asset shortage view of today's monetary anchoring, to Borio's financial cycle buildup warning (a possible consequence of the more flexible anchoring). **Such a framework would also allow to explore the different conditions under which yield curve control by the Fed would work without raising inflation, conditions which would lead to parallel financial systems to emerge, and conditions which might lead to Triffin events (and how they would unfold).**
#
#
# --
# ### A role for mechanism design
#
# Central banks' reserve management operations and the decision process revolving around them naturally call for game theory models, such as those underpinning the contract theory, borrowing and lending and to some extent medieval village economy literature.
#
# We will start first with a most basic multiagent asset allocation scheme, that of Bogolmonaia and Moulin (2001)'s cake-eating game, which we will progressively complexify and merge with standard open macroeconomy models. Indeed we believe that this model provides a close analogy to real life operations by central bank officers, and can complement existing models such as Mundell-Fleming how safety considerations are being carried out dynamically in time, along with risk vs returns logics. Furthermore, this highly abstract model can merge well with existing macroeconomy models, **by enabling calibration of values in stylized model such as the Safe Asset Scarcity and Aggregate Demand version of IS/LM developed by Caballero, Farhi and Gourinchas (AER, 2016)**. Indeed, we believe that the superposition of games with incentives on top of existing models are not competing but complementary views, just as Roth's suspension bridge building analogy illustrates how in addition of "simple, beautiful and indispensable physics", *"bridge design also concerns metal fatigue, soil mechanics, and the sideways forces of waves and wind"*. **International macroeconomics also unfolds within a game of economic competition and collaboration, whose rules - even if they "might be impossible to be answered analytically"** - must still be explored (and maybe computed), to allow "bridges designed on the same basic model to be built longer and stronger over time, as the complexities and how to deal with them become better understood" (Roth, 2002).
#
# --
# ### Our model' basic building block
#
# We chose this framework first because of its proximity with real life reserve related decisions taken by central banks. Second because of its good axiomatic properties (**”equal treatment of equals”**) on which to later build a close to optimal scheme, and third because of its simplicity and versatility suited for calibraton and simulation.
#
# Here are the features of our version of the model :
#
# - Time is discrete, in rounds.
# - At the start of each round each country generates one "cake" (debt)
# - Each country simultaneously eats each others' cakes (debt) at different speed and following different rankings (so when a piece of cake is finished the countries that were eating eat move to the item below in their rankings)
# - Each country stores what he ate in his reserves - the endowments
# - If at the end of a round a country's cake has not been completed absorbed (by itself and/or by other countries) it suffers a devaluation - i.e. every the value of pieces of cake from this country store in others' endowments is decreased, and its interest rate rises.
# - Countries' ranking preferences are function of their trading preferences, their trading partners' debts' "safety", interest rate, size, and some randomness. **See in more details below how rankings and cakes are generated by each country at each round, and on how endowments change according to countries' behaviours.**
#
# +
# Exogeneous parameters of the simulation to be played with in the rest of this notebook
number_of_countries = 6 %total number of countries in the simulation
number_of_rounds = 7 %total number of "cake eating rounds"
interest_rate = (0.03).*ones(number_of_countries,1); %initial interest rates - more below on how interest rates change according to how successfully debt is absorbeb at each round.
interest_rate_history = zeros(number_of_countries,number_of_rounds);
uni=0.7; %uni is used in the random generation of the trade preference matrix - a number_of_countries x number_of_countries matrix in which the number line i, column j indicates how interconnected the economies of countries i and j are. This random generation is done with the function R = sprandsym(n,density) - in our case n=number_of_countries and density = uni, which returns a symmetric random, n-by-n, sparse matrix with approximately density*n*n nonzeros; each entry is the sum of one or more normally distributed random samples, and (0 <= density <= 1) .For calibration uni=0.3 creates with good probability if there are 6 countries in total 3 trade coalitions, one with 3 countries, one with 2, and 1 in autarky.
# %If we were in Matlab we would have added the parameter rc=0.02, still for this sprandsym function, which denotes how inequal trade between countries can be (the smaller the more). Indeed, R = sprandsym(n,density,rc) returns a matrix with a reciprocal condition number equal to rc. The distribution of entries is nonuniform; it is roughly symmetric about 0; all are in [−1,1]. If rc is a vector of length n, then R has eigenvalues rc. Thus, if rc is a positive (nonnegative) vector then R is a positive (nonnegative) definite matrix. In either case, R is generated by random Jacobi rotations applied to a diagonal matrix with the given eigenvalues or condition number.
# -
#
# --
# ### The link with the Safe Asset Shortage hypothesis, and Borio's financial cycle build up hypothesis
#
# From the parameters of the cake eating model below, and they interact with each other, we can attempt to draw a few parallels with the competing hypothesis of Caballero, Farhi and Gourinchas, and of Borio.
#
# In fact, let's first note how the cake-eating model itself **is based** off scarcity - indeed if central banks *could* absorb all imbalances and debt at each round then in our model there would no default, no devaluation, and all assets would be considered safe. However given the limited *duration* parameter *(see the parameter duration_T_updated_history introduced below, noting the records of how long was the timer for each round. A round ends when this timer ends, or when countries ate all the cakes if faster than the timer, with the timer changing at each round depending on the total cake sizes and total eating speed ratio, so that it increases rather steadily)*, only the countries that first manages to "offload" their debts avoid devaluation and maintain a "safe" reputation.
#
# Second, let's note how the two parameters of how much cakes (ie debts) each country is generating, and how much cakes each country can absorb, at each round, are proxies of how developed/stretched a country's financial system is. One can note how past history of safety in a country can lead to an expanding emission/absorption cycle, which might eventually burst à la Minsky, related to Borio's hypothesis.
#
# Hence two scenarii : one in which a few safe countries' debts are pursued in higher demand than what they emit (safe asset shortage), which lowers the interest rate. But this scarcity only incentivizes them to stretch more their financial systems so that they both emit and absorb more pieces of cake. Up until a situation in which they emitted too much pieces of cake compared to the demand, in which case they suffer a devaluation requiring them to either emit less pieces of cake, absborb more of their own (through decreased interest rate as described by Borio), or convince more countries to absorb theirs first.
#
# *NB : explore how this could be linked back to Caballero, Farhi and Gourinchas' IS-LM equations*
# +
# We initialize here the recording vectors that will collect the history of the simulation, to plot the graphs at the end of this notebook
endowments_reserves_history = zeros(number_of_countries,number_of_rounds); %The records of how much cakes in total each country has accumulated at each round (see cell just below for the definition of a cake)
endowments_debts_history = zeros(number_of_countries,number_of_rounds); %The records of how much cakes (ie debts) in total each country has generated up to round t (see cell just below for the definition of a cake)
sum_cakes_history = zeros(1,number_of_rounds); %The records of each country's cake size for each round (see cell just below for the definition of a cake)
sum_of_generated_cakes = 0; %for cumulated
abso_speed_history = zeros(number_of_countries,number_of_rounds); %The records of the eating speed of each country for each round (see cell just below for the definition of the eating speed)
abso_history = zeros(1,number_of_rounds);
sum_of_abso_speed = 0; %for cumulated
devaluation_history = zeros(number_of_countries,number_of_rounds); %The records of the cumulated number of times which each country devalued up to round i
ranking_history = zeros(number_of_countries,number_of_countries); %The records of the order in which each country ranked other countries' cake to be eaten at each round(see cell just below for the definition of a cake)
history_counter = ones(number_of_countries,1); %used in which round to determine sizes of generated cakes and cake eating speeds (see within round cell below)
deval_history = zeros(number_of_countries,1); %idem
duration_T_updated_history = zeros(1,number_of_rounds); %The records of how long was the timer for each round (see cell just below for the definition of a timer)
# -
# ## Examples of figures from paper that can be obtained with this notebook :
#
# **First a 6 countries in 3 trade groups, 7 rounds simulation (just 7 rounds are enough to see trends !)**
#
# From this first short simulation note that :
#
# - the size of cakes, eating speed etc... tend to powers law i.e. one country (here country 1) gains much more than any more in that trading group (mathematical proof in paper). Intuition is that as it is perceived "safer" by other countries (because it devalues less) the "cakes" (debts) it generates are always eaten first by other countries, and thus incentivizes (through lower interest rate) country 1 to generate bigger cake, invest more in its cake eating speed, invest more in other countries' debts as well, etc.
#
# - the country that would gain the most is determined by initial conditions (in this very easy example everything is initialized at random, but looking at the initial ranking - the higher the score the most sought after a ranking is - and the initial interest rate we can tell us that country 1 will be winning !)
#
# - the country that would gain the most in a short run trade group is not necessarily the one determined by fundamentals - i.e. not the most connected one or the most prudent one *(in the trade group between country 1, 4 and 6 country 1 is neither the most connected - since country 6 has more connections than it does - nor the most self preserving - since country 4 ranks its own debts to others' higher first)*
#
# <img src="Screen%20Shot%202020-04-02%20at%2010.14.45%20AM.png" />
#
#
# **A second 6 countries in 3 trade groups, 7 rounds simulation**
#
# - Here the winner is country 3 - which is less connected (only to country 5) than the cluster with countries 1, 2 and 4
#
# - The worse off country in the group (1, 2 and 4) is doing even worse than country 6 in autarky (extreme case of the model, because country 4 started (randomly) at the least preferred country in country rankings, and failed to get its cakes eaten at the end of round 1, thus devalues round 2, its interest rate surge, and it gets stuck in a devaluation sprial from round 1 to 4.
#
# - Note that here country 1 had better initial conditions to be the be winner of the simulations (best ranked country at the start, lowest interest rate at the start). But its trading partners - country 2 and 4, are the ones devaluing the most
#
# <img src="Screen%20Shot%202020-04-02%20at%2010.44.32%20AM.png" />
#
#
# **Finally a 9 countries, 1000 rounds simulation**
#
# The figures below illustrate the long rung trends in this model :
#
# - Reduction of average global rates can happen (see rounds 120 to 220 on both the cumulated. This is when the total eating speed is faster than the timer, hence the reduced number of devaluations rounds 120 to 220
#
# - Triffin events exist : the yellow country (country 3) started as the winner of this simulation, until around round 220 at which point it is overtaken by country 1 in deep blue and don't stop devaluing until the end of the 1000 rounds. This corresponds to fundamental analysis (country 3 is only trading with 2 countries whereas country 1 with 4. But countries 6 and 9 for instance are both connected to 4 trading partners, with stronger ties among some of them and with themselves. These two countries are respectly the 2nd and 3rd most sought after countries, both in rankings, in upward trends in rankings, and in how much of their cakes are absorbed by others as reserve).
#
# *Note that these long term analysis will be nuanced by a more calibrated simulation in the notebooks on 1945-now and after Covid-19, available here and here, in which the cake eating speed and cake generation speed are governed by more complicated but more realistic
#
# <img src="Screen%20Shot%202020-04-02%20at%2011.07.19%20AM.png" />
#
# <img src="Screen%20Shot%202020-04-02%20at%2010.55.01%20AM.png" />
# +
# We initialize here the intermediary parameters that will be used within each round
endowments = zeros(number_of_countries,number_of_countries); % Where countries store what "cakes" (debts) they ate during each round. At the end of each round a country's cake that hasn't been totally eaten will decrease the parts of all reserves that contained pieces of cakes (from all previous rounds, not just this round) from that country (it's a devaluation - and we record it in deval_history = deval_history + 1). Else if the cake has been totally it counts as a success and we record it in history_counter = history_counter + 1
abso_speed = 100.*rand([number_of_countries,1]); % The eating speed of the countries - here initialized at random. During the rest of the game these speed will evolve depending on the history_counters and devaluation_counters of each country
abso_speed_init = abso_speed;
initial_pos = 100.*rand([number_of_countries,1]); % Cakes generated by each country, initialized at random. Then at the start of each round the size of cakes being generated by each country are function of how safe this country is (history_counter - deval_history) plus a random number different for each country (see further in the code)
initial_pos_init =initial_pos;
initial_pos_round = initial_pos;
cakes_from_previous_round = initial_pos;
duration_T = 1; % There's a timer for each rouund. A round ends when this timer ends, or when countries ate all the cakes if faster than the timer (the timer changes at each round depending on the total cake sizes and total eating speed ratio, so that it increases rather steadily)
fraction_ranking = (1/3); % Used for calibration
duration_T_updated = duration_T;
successfully_loaded_cake_last_round = zeros(number_of_countries,1);
# +
# random initialization of ranking between countries, before it is impacted by trade preference and readjusted within each rounds
ranked_list = zeros(number_of_countries,number_of_countries);
for j=1:number_of_countries
ranked_list(j,:) = randperm(number_of_countries);
end
# +
# Initialization of trade preference, and impact on ranking among countries just defined above.
% NOTE THAT THESE CAN BE PLAYED AROUND, AND CALIBRATED AS IN THE 1945-NOW Notebook !!!
trading_preferences = ceil(abs(full(sprandsym(number_of_countries,uni)))); %NOTE THAT IN MATLAB FULL CODE IS
# %trading_preferences = ceil(abs(full(sprandsym(number_of_countries,uni,rc,2)))); with rc discussed in the intro, for unequality in initial distribution
for k=1:number_of_countries
% we first find a country's preferred trading partners and rank them among themselves
[~,trading_order]=sort(trading_preferences(k,:).*(ones(1,number_of_countries)+endowments(k,:)),'descend'); %we rank by first trade preference, then by the size of trading partners. Note that this is modelled more finely in the 1945-now simulation notebook
% the 1945-now version :
%[~,trading_order]=sort(trading_preferences(k,:).*(ones(1,number_of_countries)+max(0,successfully_loaded_cake_last_round(k,1).*(history_counter(k,1)-deval_history(k,1)).*endowments(k,:)*(interest_rate(k,1)))),'descend');
% then we adjust the country's rankings by moving these trading partners up the list
res=sum(trading_preferences(k,:)~=0);
if res>0
for l=1:res
ressort_count=find(ranked_list(k,:)==trading_order(l));
if ressort_count>l
for m=1:(ressort_count-l)
ranked_list(k,ressort_count+1-m)=ranked_list(k,ressort_count-m);
end
ranked_list(k,l)=trading_order(l);
else
end
end
end
end
ranked_list_true_init = ranked_list %print this ranking matrix so that you can see; each line i is the ranking list of a country i, from top priority left to last choice right.
# +
# Now we go into rounds of cake eating, that are looped over. Everything important happens in this cell !
for i=1:number_of_rounds
cakes = max(0,successfully_loaded_cake_last_round.*(history_counter-deval_history)+rand([number_of_countries,1]));
abso_speed = max(0.1,(history_counter-deval_history)+rand([number_of_countries,1]));
% At the start of the round cakes (ie debt) are generated by each country, and each country is assigned a new eating speed, both depending only on how successful it history is in getting its debt absorbed.
% Plus we just add a bit of random perturbations to make it more realistic. But very simple model here, that is made more sophisticated in the other simulations (see the two other notebooks
# %we record all of this for graphs later
cakes_from_previous_round = cakes;
sum_of_generated_cakes = sum_of_generated_cakes + cakes;
sum_of_abso_speed = sum_of_abso_speed + abso_speed;
abso_history(1,i)=sum(abso_speed);
sum_cakes_history(1,i)=sum(cakes);
initial_pos_round = cakes;
% The counter of a round is set up as the max cake size generated at this round
duration_T_updated = max(1,max(cakes));
% We initialize the counting
time_counter=0;
cakes_counter=0;
rank_counter = ones(number_of_countries,1);
removed_cake_row = zeros(number_of_countries,1);
k=1;
latest_finished_cake = 0;
will_be_totally_consumed = zeros(number_of_countries,1);
% And the countries start eating ! Until all the cakes are eaten or the counter is down to zero.
while (duration_T_updated>0 && cakes_counter<number_of_countries)
initial_pos_updated = cakes;
diff_between_wished_and_offered = zeros(number_of_countries,1);
% We go through the preference list of all the countries - if the cake item at the top of each country's remaining list is still there they'll eat it, else we move down an item in that country's list
% For that we first figure out which cake pieces are sought after by the most countries x their eating speed
for l=1:number_of_countries
if removed_cake_row(ranked_list(l,1))>0
o=1;
u=1;
if removed_cake_row(ranked_list(l,o+1),1)>0
u=o+1;
while ((u<(number_of_countries-1)) && (removed_cake_row(ranked_list(l,u+1),1)>0))
u=u+1;
end
ranked_list(l,o)=ranked_list(l,u+1);
else
ranked_list(l,o)=ranked_list(l,o+1);
end
rank_counter(j,1)=rank_counter(j,1)+1;
end
end
for j=1:number_of_countries
if removed_cake_row(j,1)>0
else
if sum(abso_speed(ranked_list(:,1)==j))>cakes(j,1)
will_be_totally_consumed(j,1)=will_be_totally_consumed(j,1)+1;
diff_between_wished_and_offered(j,1) = sum(abso_speed(ranked_list(:,1)==j))/initial_pos_updated(j,1);
end
end
end
if sum(diff_between_wished_and_offered)==0
for j=1:number_of_countries
if removed_cake_row(j,1)>0
else
diff_between_wished_and_offered(j,1) = sum(abso_speed(ranked_list(:,1)==j))/initial_pos_updated(j,1);
will_be_totally_consumed(j,1)=will_be_totally_consumed(j,1)+1;
end
end
else
end
% Now that we know which cake pieces are sought after by which countries x their eating speed we can sort them
[~,consumption_order]=sort(diff_between_wished_and_offered,'descend');
% Just some safety checks
if sum(diff_between_wished_and_offered)==0
[ii,jj]=find(~removed_cake_row);
if size(ii)==1
consumption_order(1)=ii;
else
for j=1:size(ii)
consumption_order(j)=ii(j);
end
end
else
end
% Now we know which cake will be eaten !
% Let's share it across countries according to their eating speed - we move these to the relevant slots in the endowments matrix.
% With just a special attention if the cake piece being eaten won't be finished in time (the "else" to this "if" condition)
if (duration_T_updated-cakes(consumption_order(1),1))>0
duration_T_updated = duration_T_updated - cakes(consumption_order(1),1);
for j=1:number_of_countries
if ranked_list(j,1)==consumption_order(1)
endowments(j,consumption_order(1)) = endowments(j,consumption_order(1))+(abso_speed(j,1)./(sum(abso_speed(ranked_list(:,1)==consumption_order(1)))))*initial_pos_updated(consumption_order(1),1);
elseif cakes(ranked_list(j,1),1)==0
removed_cake_row(ranked_list(j,1),1)=removed_cake_row(ranked_list(j,1),1)+1;
else
endowments(j,ranked_list(j,1)) = endowments(j,ranked_list(j,1))+(abso_speed(j,1)*cakes(consumption_order(1),1))/(sum(abso_speed(ranked_list(:,1)==consumption_order(1))));
cakes(ranked_list(j,1),1)=initial_pos_updated(ranked_list(j,1),1)-(sum(abso_speed(ranked_list(:,1)==ranked_list(j,1)))*cakes(consumption_order(1),1))/(sum(abso_speed(ranked_list(:,1)==consumption_order(1))));
end
end
time_counter = time_counter + cakes(consumption_order(1),1);
cakes(consumption_order(1),1) = 0;
removed_cake_row(consumption_order(1),1)=removed_cake_row(consumption_order(1),1)+1;
% With just a special attention if the cake piece being eaten won't be finished in time (the "else" to this "if" condition)
else
cakes(consumption_order(1),1) = cakes(consumption_order(1),1) - duration_T_updated;
time_counter = time_counter + duration_T_updated;
for j=1:number_of_countries
if ranked_list(j,1)==consumption_order(1)
endowments(j,consumption_order(1)) = endowments(j,consumption_order(1))+(abso_speed(j,1)./(sum(abso_speed(ranked_list(:,1)==consumption_order(1)))))*duration_T_updated;
elseif cakes(ranked_list(j,1),1)==0
removed_cake_row(ranked_list(j,1),1)=removed_cake_row(ranked_list(j,1),1)+1;
else
endowments(j,ranked_list(j,1)) = endowments(j,ranked_list(j,1))+(abso_speed(j,1)*duration_T_updated)/(sum(abso_speed(ranked_list(:,1)==consumption_order(1))));
cakes(ranked_list(j,1),1)=cakes(ranked_list(j,1),1)-(abso_speed(j,1)*duration_T_updated)/(sum(abso_speed(ranked_list(:,1)==consumption_order(1))));
end
end
duration_T_updated = 0;
end
latest_finished_cake = consumption_order(1);
cakes_counter = cakes_counter + 1;
k=k+1;
duration_T_updated_history(1,i)=duration_T_updated;
end
% At the end of a round we devalue all countries that didn't manage to get all its debt absorbed during this round, proportionally to how much of it is left at the end of the round
for j=1:number_of_countries
if (initial_pos_round(j,1)-cakes(j,1))>0
endowments(:,j)=endowments(:,j)*((initial_pos_round(j,1)-cakes(j,1))/initial_pos_round(j,1));
else
end
if cakes(j,1)==0
history_counter(j,1) = history_counter(j,1) + 1;
successfully_loaded_cake_last_round(j,1) = 1;
interest_rate(j,1)=interest_rate(j,1)./(1+rand(1,1));
else
interest_rate(j,1)=min(1,max(interest_rate_history(j,1),interest_rate(j,1).*(1+rand(1,1))));
deval_history(j,1) = deval_history(j,1)+1 ;
end
end
% And we update the ranking list for each country after the events of this round, in a similar fashion as what happened initially on the cell just above on this notebook.
[~,success_order]=sort(history_counter,'descend');
ranked_list = repmat(success_order',number_of_countries,1);
for k=1:number_of_countries
r1 = randperm(number_of_countries,2*round(number_of_countries*fraction_ranking));
for j=1:round(number_of_countries*fraction_ranking)
inter=ranked_list(k,r1(1,j));
ranked_list(k,r1(1,j))=ranked_list(k,r1(1,2*j));
ranked_list(k,r1(1,2*j))=inter;
end
end
for k=1:number_of_countries
[~,trading_order]=sort(trading_preferences(k,:).*(ones(1,number_of_countries)+max(0,successfully_loaded_cake_last_round(k,1).*(history_counter(k,1)-deval_history(k,1)).*endowments(k,:)*(interest_rate(k,1)))),'descend');
res=sum(trading_preferences(k,:)~=0);
if res>0
for l=1:res
ressort_count=find(ranked_list(k,:)==trading_order(l));
if ressort_count>l
for m=1:(ressort_count-l)
ranked_list(k,ressort_count+1-m)=ranked_list(k,ressort_count-m);
end
ranked_list(k,l)=trading_order(l);
else
end
end
end
interest_rate_history(k,i) = interest_rate(k,1);
end
ranked_list_init = ranked_list;
endowments_reserves_history(:,i) = sum(endowments,2);
endowments_debts_history(:,i) = sum(endowments,1)';
abso_speed_history(:,i) = abso_speed;
devaluation_history(:,i) = deval_history;
ranking_history(:,:,i) = ranked_list_init;
end
# -
# ### That was the core of the simulation code ! Now one can just plot everything
#
# So the graphs can either be all in one cell, or be plot individually one per cell. I'll let the reader copy paste the relevant pieces of code accordingly, that can be put in a following single cell as shown for example below.
#
# *Also note that the digraph function used to plot the graph of the trade preferences among countries is not implemented yet in Octave. So the code is provided here but won't produce this specific graph.
# +
sum_ranking_history = zeros(number_of_countries,number_of_rounds);
i=1;
for j=1:number_of_countries
tag{j}=strcat('country ',num2str(j));
for k=1:number_of_countries
[~,indexes]=sort(ranking_history(k,:,i));
score_for_one_country = ((number_of_countries+1)*ones(1,1)-indexes);
sum_ranking_history(j,1) = sum_ranking_history(j,1)+score_for_one_country(1,j);
end
end
sum_ranking_history_b = sum_ranking_history;
for i=2:number_of_rounds
for j=1:number_of_countries
for k=1:number_of_countries
[~,indexes]=sort(ranking_history(k,:,i));
score_for_one_country = ((number_of_countries+1)*ones(1,1)-indexes);
sum_ranking_history(j,i) = sum_ranking_history(j,i)+score_for_one_country(1,j);
end
end
sum_ranking_history(:,i)=(sum_ranking_history(:,i)+sum_ranking_history(:,i-1));%./(mean(sum_ranking_history(:,i-1)));
sum_ranking_history_b(:,i)=sum_ranking_history(:,i)-sum_ranking_history(:,i-1);%./(mean(sum_ranking_history(:,i-1)));
end
figure;
A=trading_preferences;
G = digraph(A);
LWidths = 5*G.Edges.Weight/max(G.Edges.Weight);
subplot(2,3,1);
plot(G,'LineWidth',LWidths)
title('Trade preferences among 6 (numbered) countries graph')
subplot(2,3,2);
plot(interest_rate_history')
title('Interest rate history')
# %legend(tag)
subplot(2,3,3);
plot(abso_speed_history')
title('Cake eating speed history')
# %legend(tag)
subplot(2,3,4);
plot(sum_ranking_history_b')
title('Ranking history by country')
legend(tag)
subplot(2,3,5);
plot(endowments_debts_history')
title('Cumulated debts generated by country history')
# %legend(tag)
subplot(2,3,6);
plot(endowments_reserves_history')
title('Cumulated reserves by country history')
# %legend(tag)
# -
# ### **Calibration on 1945 - now and post Covid debt simulations**
#
# We present here the detailed code used for the simulations. For that we first calibrate the model to match the main trends and events observed from 1945 to now (2020). Being able to capture these features of past history will provide confidence and intuition in the model's inner workings. That lead us to provide several scenarii in how the models' parameters will change following Covid-19's increase in debts *(these parameters from the model being for instance the change in debt levels, while debt absorption speed are maintained roughly constant. Different degrees in shifts in trade and supply chain preferences are also considered)*. We thus generate 50 more rounds after a common departing point, and report here the different outcomes, and the different probabilities of each outcome (since they are intervention of random perturbations in the model the probabilities of each outcome is calculated based on how many of different random sequence generate this specific outcome).
#
# ### Results
#
# The simulations of extra Covid debts show that :
#
# - **in the case of everything constant (pre Trump US and China trade relations)** debts are overall sustainable, with adjustements in reserve levels and contraction of countries' economies depending on how affected by the crisis they were.
#
# <img src="Screen%20Shot%202020-04-06%20at%2010.05.35%20AM.png" />
#
# - **similar results in the case of everything constant, with just additional reliance on trade with China** (for instance for masks, medical supplies etc). The economic contraction is smaller, and the global reserve contraction as well. This could be interpreted by the fact that the additional trade with China is flushed in its reserve (the increase can be seen in the graph below), used to acquire other countries' debt (inferred from the lowering of global interest rates) and thus sustaining economic activity
#
# <img src="Screen%20Shot%202020-04-06%20at%2011.13.15%20AM.png" />
#
# - **in the case of everything constant among all countries except China and US, who in that scenario stop trading with each other**, they are small probabilities of a global contraction (less reserves, less debts, less GDP growth) that preserves the overall shares of each country in the global economy - except for China in the global economy which suffers more. However in these small probability event the US go through an initial bubble fueled by the decrease of China, that then collapses back to the initial level (and note that after this collapse China's share is also back to the initial level - but with the cost of a global contraction, especially for all EM markets). **And with the caveat provided just above that this debt model doesn't take into account other incentives and consequences created by such dynamics - small probabilities in economic contraction in one country might leading to escalating tensions on other dimensions as well*
#
# <img src="Screen%20Shot%202020-04-04%20at%209.50.20%20PM.png" />
#
# - **in the case of stopping trade between China and the US AND additional reliance by other countries on China**, there is high probability for a sudden takeover of the role of the US as reserve currency China, accompanied by strong contraction for every other country. **Again, with the previous caveat of unaccounted destabilizing geopolitical consequences**
#
# <img src="Screen%20Shot%202020-04-04%20at%209.57.55%20PM.png" />
#
#
# - **finally, in the case of our proposed stabilization scheme** *(details in the paper - in the model it is just reflected as more willingness from participating countries to accept others' devaluation - see the commented code in the last cells of this notebook. Un-implemented here is also the extra information each country will have on how much latitude they will have at each round on their debt, which should also help in addition of the effects produced here to stabilize debts, interest rates and balance global reserve currencies)* the effects of all above scenarii are smoothen, with much more balanced reserve currencies (at least 3 currencies accounting each more than 20% of the global share of reserves, with the exact currencies - most often Germany ie the Euro, or the JPY - switching depending on the random shocks, but without adverse effects on all countries - including on the US even though the dollar loses its monopoly as reserve currency)
#
# Examples from one representative random initialization
# <img src="Screen%20Shot%202020-04-04%20at%2010.12.50%20PM.png" />
#
# Examples from another representative random initilization
# <img src="Screen%20Shot%202020-04-04%20at%2010.13.52%20PM.png" />
#
# ### A quick review of our strategy proof scheme :
#
# At the core of our proposal resides the *multi-currency* nature of the market making activity of international banks, which we aim at replicating at a central bank level. A first underlying reason would be that of *collection of risk* - in the line of bigger risks for higher returns. Additionally (and that is a second reason for emphasizing the multi-currency aspect) in our case the additional risk is that of adding on more exotic currencies such as the South East Asian ones in a reserve basket previously not containing them - that could thus act as an additional cushion to mitigate external shock on them.
#
# It is then possible to see that for such a **multi-currency reserve balance sheet** the profit objective of traditional private market maker is here also aligned with the volatility stabilization objective of the central banks involved, and that the mechanisms underlying these forms of interventions would be conceptually the same as the ones through which financiers' alterations to their balance sheets affect both the level and volatility of exchange rates.
#
# The design question then, critical to the feasibility of such a multi-currencies account for instance, is on **how to create and foster this form of collaboration among different central banks**.
#
# ### Computing these multilateral balance sheets, through privacy-preserving methods
#
# Privacy-preserving methods like multiparty computation *(referring to the financial adaptation of Abbe, Khandani, Lo, 2012, which build on Yao 1982; Goldreich, Micali, and Wigderson 1987; <NAME>, and Wigderson 1988; <NAME>, and Damgard 1988; Beaver, Micali, and Rogaway
# 1990; Cramer et al. 1999)* provide a clear and useful framework on which to **reach agreement on the different exchange and interest rate objectives pursued by respective central banks**. This is why this section will build on the language and framework of these privacy-preserving methods - which again don't have to be *actually* implemented for the interventions to be conducted.
#
# 1. Thus central banks would first each allocate some funds denominated in their national currencies, plus portions of their reserves in each of the currencies they would like to stabilize their exchange rates with.
# 2. They would then set-up the daily volatility bands they want their national currencies to stay in, which proportion of reserves they are ready to commit to the operation, and what amount of risk or leverage they are ready to bear on any derivative instrument used in the process (for convenience let's imagine the updates are only feasible at a regular interval - for instance once a day or once a week. We are then in a discreet time setting). These three points are where privacy-preserving methods might be applied, since participant central banks wouldn't want for instance *the entirety* of the reserves they engaged in that fund to be potentially used to defend the currency of another participating country)
# 3. Thus a daily total amount of reserves committed to each currency could be computed using the aforementioned methods, without individual inputs from each central bank being revealed). Note that the viewing of the results of these computations can also be restrained so that each central bank only sees the pooling of reserves allocated to him - and not that allocated to others. In this fashion, both inputs and outputs of these forms of computations have modular privacy settings, that can be deployed according to participants preferences.
#
# The previous paragraph described mostly a pooling of reserves, quite similar to what already exists in the Chiang Mai Initiative, but **applied on a continuous and systematic basis rather than just during times of extreme stress** (in fact one of the goals of this automatic and systematic market smoothing is to potentially provide in-commensurable psychological relief and precious time for governments to *avoid* times of extreme stress).
#
# There are also **other benefits to having a common multicurrency account: first, the mechanisms derived to stabilize a currency vs the international currencies part of the reserves portfolio can now also be used to also stabilize more exotic currency pairs** (but among natural trading partners) such as the MYR/IDR.
#
# Furthermore, **our scheme would constitute an implementation of the "optimal capital controls**, that should lean against the wind by requiring a temporary tax on inflows and a subsidy on outflows (if dealing with sudden inflows, the opposite if dealing with a sudden stop)" from Farhi and Werning (2013).
#
# Finally, it would **help by design including more currencies in the global reserve distribution and alleviating the "safe asset shortage"** (Caballero, Farhi, Gourinchas, 2017).
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example 4: Flexural parameters at single grid cell
# All the previous examples have led us to the calculation of the wavelet admittance and coherence functions at every grid cell. However, the main purpose ot `PlateFlex` is to estimate (and map) the flexural parameters of the lithosphere from the inversion of the admittance and coherence functions. We will explore this step-by-step again, starting with the estimation of flexural parameters at a single grid cell.
#
# Let's start again by loading the topography data into a `TopoGrid` object and the Bouguer gravity anomaly data into a `BougGrid` object, which are then both inserted into a `Project` object.
# +
import numpy as np
import pandas as pd
from plateflex import TopoGrid, BougGrid, Project
# Read header (first line) of data set using pandas to get grid parameters
xmin, xmax, ymin, ymax, zmin, zmax, dx, dy, nx, ny = \
pd.read_csv('../data/Topo_NA.xyz', sep='\t', nrows=0).columns[1:].values.astype(float)
# Change type of nx and ny from float to integers
nx = int(nx); ny = int(ny)
# Read topography and Bouguer anomaly data
topodata = pd.read_csv('../data/Topo_NA.xyz', sep='\t', \
skiprows=1, names=['x', 'y', 'z'])['z'].values.reshape(ny,nx)[::-1]
bougdata = pd.read_csv('../data/Bouguer_NA.xyz', sep='\t', \
skiprows=1, names=['x', 'y', 'z'])['z'].values.reshape(ny,nx)[::-1]
# Load the data as `TopoGrid` and `BougGrid` objects
topo = TopoGrid(topodata, dx, dy)
boug = BougGrid(bougdata, dx, dy)
# Create contours
contours = topo.make_contours(0.)
# Make mask
mask = (topo.data < -500.)
# Define new project
project = Project(grids=[topo, boug])
# Initialize project
project.init()
# Calculate wavelet admittance and coherence
project.wlet_admit_coh()
# Take random cell value within grid and set as attribute
cell = (250, 100)
project.cell = cell
# -
# We are ready to estimate the flexural parameters for the specified grid cell. Before going further, let's remind ourselves of the following:
project.__dict__.keys()
# You notice that the project contains new attributes `rhoc` and `zc`, which can be used in the estimation of the flexural parameters. To check how these are defined, simply print them:
print(project.rhoc)
print(project.zc)
# By default these attributes are equal to `None`. This implies that the softwares will use default vaules for those fields during estimation of the flexural parameters.
#
# Let's explore the default parameters of the model.
# +
# First we need to import `plateflex`
import plateflex
# Then print out the default parameters
plateflex.get_conf_flex()
# -
# In this default model, the crust is 35 km thick and has a density of 2700 kg/m^3, the mantle density is 3200 kg/m^3 and water depth has been set to 0 m. Furthermore, the fluid density (fluid above topography) has been set to that of air (0 kg/m^3). Finally, during initialization of the project, the software recognized that the `GravGrid` object was of type `BougGrid`, which updated the `boug` variable to 1 (`True`).
#
# <div class="alert alert-block alert-info">
# <b>Note:</b> The variables `rhof` and `wd` do not need to be changed. The code will automatically extract the water depth at the specified grid cell and determined whether to use `rhow` or `rhoa` for the fluid density above the topography.
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>Warning 1:</b> It is highly recommmended not to change the variable `boug`, since the calculation of the predicted admittance and coherence depends on that flag and will give erroneous results if mis-used.
# </div>
#
# <div class="alert alert-block alert-warning">
# <b>Warning 2:</b> It is the responsibility of the user to specify the correct type of `GravGrid` to use when loading the data set of gravity anomalies.
# </div>
#
# Although some of these values appear to be reasonable, it is always possible to change some of them - in particular `rhoc`, `rhom`, `zc`, `rhow` or `rhoa` (although the default values for `rhom`, `rhow` and `rhoa` are fairly good estimates for this particular problem and should not affect the solution).
# +
# Change crustal thickness and density
from plateflex.flex import conf_flex as cf
cf.zc = 40.e3 # in meters
cf.rhoc = 2670. # in kg/m^3
plateflex.get_conf_flex()
# -
# Next, you probably remember that during initialization, the `inverse` attribute was set to `L2` (check this with `print(project.inverse)`). This implies that by default the software will use a non-linear least-squares method to estimate the flexural parameters. The two available options are `'L2'` and `'bayes'`, the latter based on a probabilistic estimation method based on bayesian inference. A couple of things to note:
#
# <div class="alert alert-block alert-info">
# <b>Note 1:</b> The two methods should give very similar results. Differences arise mainly because the 'bayes' approach draws a finite number of samples from the posterior distribution, and equality is achieved only for infinite sampling of the posterior and for the maximum a posteriori (MAP) estimate (because the mean estimate of the posterior can be biased if there is more than one mode).
# </div>
#
# <div class="alert alert-block alert-info">
# <b>Note 2:</b> The 'L2' approach is <b>MUCH</b> faster than the 'bayes' approach. However, the 'bayes' approach provides useful statistics for the inference step.
# </div>
#
# Let's examine the estimation of the flexural parameters using the default `'L2'` approach, by only specifying the cell location (tuple):
# +
cell = (250, 200)
# Perform estimation
project.estimate_cell(cell)
# Print summary
print(project.summary)
# Plot observed and predicted (best-fit) functins:
project.plot_functions()
# -
# Now do the same using the `'bayes'` approach
# +
# Switch inverse attribute
project.inverse = 'bayes'
# Perform estimation
project.estimate_cell(cell)
# Print summary
print(project.summary)
# Plot observed and predicted (best-fit) functions using the MAP estimate:
project.plot_functions(est='MAP')
# Plot stats for estimate
project.plot_bayes_stats()
# -
# In the background, the method has done a joint inversion of the admittance and coherence functions to estimate the flexural parameters. This is the default setting for the estimation, but can be changed by specifying a different `atype` values (default: `atype='joint'`; other available options are `atype='admit'` and `atype='coh'`):
# +
# Switch back to `L2`
project.inverse = 'L2'
# Perform estimation, print summary and plot
project.estimate_cell(cell, atype='admit')
print(project.summary)
project.plot_functions()
# -
# Perform estimation, print summary and plot
project.estimate_cell(cell, atype='coh')
print(project.summary)
project.plot_functions()
# And the same with the `'bayes'` approach
# +
# Switch again to `bayes`
project.inverse = 'bayes'
# Perform estimation, print summary and plot
project.estimate_cell(cell, atype='admit')
print(project.summary)
project.plot_functions(est='MAP')
project.plot_bayes_stats()
# -
# Perform estimation, print summary and plot
project.estimate_cell(cell, atype='coh')
print(project.summary)
project.plot_functions(est='MAP')
project.plot_bayes_stats()
# You can sometimtes see in the last example (all runs of the `'bayes'` approach are different) that the sampling has flirted with high `F` values, thus giving a biased (lower) "mean" estimate. This effect could also potentially bias the distribution of `Te` values. The `'bayes'` approach is therefore showing that the inversion of the coherence function alone can give non-unique results.
#
# We are now ready to map out these parameters over the whole grid! Check out Example 5.
| plateflex/examples/Notebooks/Ex4_estimate_flex_parameters_cell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# +
import matplotlib as mpl
mpl.rc('font', family='NanumBarunGothic')
# -
df = pd.read_csv("../data/BC_Card_edit.csv")
print(df.info())
df.head()
df['기준년월'].value_counts()
df = df[df['고객소재지_광역시도'] == '서울특별시']
df_2103 = df[df['기준년월'] == 202103]
df_2103.head(10)
sigoongu = list(set(df_2103['고객소재지_시군구']))
print(len(sigoongu))
# +
big_category = list(set(df_2103['품목대분류명']))
print(len(big_category))
middle_category = list(set(df_2103['품목중분류명']))
print(len(middle_category))
# -
lifestyle = list(set(df_2103['가구생애주기']))
print(len(lifestyle))
lifestyle
# +
print('지역별 매출')
print(df_2103['고객소재지_시군구'].value_counts())
df_2103_total = df_2103['고객소재지_시군구'].value_counts().tolist()
plt.bar(sigoongu, df_2103_total)
plt.xticks(rotation = 90)
plt.show()
# +
# 시군구 - 매출건수(품목대분류명)
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
sigoongu_moneycount_large = pd.DataFrame(pack, index=sigoongu)
sigoongu_moneycount_large = sigoongu_moneycount_large.fillna(0)
plt.figure(figsize=(20,10))
plt.pcolor(sigoongu_moneycount_large)
plt.xticks(np.arange(0.5, len(sigoongu_moneycount_large.columns), 1), sigoongu_moneycount_large.columns, rotation=90)
plt.yticks(np.arange(0.5, len(sigoongu_moneycount_large.index), 1), sigoongu_moneycount_large.index)
plt.xlabel('__', fontsize=14)
plt.ylabel('__', fontsize=14)
plt.colorbar()
plt.show()
# +
# 시군구 - 매출건수(품목대분류명)
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False)
# +
# 각 시군구 내에서 매출건수(품목대분류명) 기준으로 순위
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False, axis = 1)
# +
# 클러스터링
# 시군구 - 매출건수(품목대분류명)
import numpy as np
from sklearn.cluster import KMeans
import seaborn as sns
datapoints = sigoongu_moneycount_large.values
kmeans = KMeans(n_clusters=5).fit(datapoints)
sigoongu_moneycount_large['cluster_id'] = kmeans.labels_
for i in range(5):
temp = sigoongu_moneycount_large[sigoongu_moneycount_large['cluster_id'] == i]
print(temp['cluster_id'])
# +
# 시군구 - 매출건수(품목중분류명)
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
sigoongu_moneycount_middle = pd.DataFrame(pack, index=sigoongu)
sigoongu_moneycount_middle = sigoongu_moneycount_middle.fillna(0)
plt.figure(figsize=(40,10))
plt.pcolor(sigoongu_moneycount_middle)
plt.xticks(np.arange(0.5, len(sigoongu_moneycount_middle.columns), 1), sigoongu_moneycount_middle.columns, rotation=90)
plt.yticks(np.arange(0.5, len(sigoongu_moneycount_middle.index), 1), sigoongu_moneycount_middle.index)
plt.xlabel('__', fontsize=14)
plt.ylabel('__', fontsize=14)
plt.colorbar()
plt.show()
# +
# 시군구 - 매출건수(품목중분류명) - 순위
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False)
# +
# 각 시군구 내에서 매출건수(품목중분류명) 기준으로 순위
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출건수'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False, axis = 1)
# +
# 클러스터링
# 시군구 - 매출건수(품목중분류명)
import numpy as np
from sklearn.cluster import KMeans
import seaborn as sns
datapoints = sigoongu_moneycount_middle.values
kmeans = KMeans(n_clusters=5).fit(datapoints)
sigoongu_moneycount_middle['cluster_id'] = kmeans.labels_
for i in range(5):
temp = sigoongu_moneycount_middle[sigoongu_moneycount_middle['cluster_id'] == i]
print(temp['cluster_id'])
# +
# 시군구 - 매출금액(품목대분류명)
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
sigoongu_money_large = pd.DataFrame(pack, index=sigoongu)
sigoongu_money_large = sigoongu_money_large.fillna(0)
plt.figure(figsize=(20,10))
plt.pcolor(sigoongu_money_large)
plt.xticks(np.arange(0.5, len(sigoongu_money_large.columns), 1), sigoongu_money_large.columns, rotation=90)
plt.yticks(np.arange(0.5, len(sigoongu_money_large.index), 1), sigoongu_money_large.index)
plt.xlabel('__', fontsize=14)
plt.ylabel('__', fontsize=14)
plt.colorbar()
plt.show()
# +
# 시군구 - 매출금액(품목대분류명) - 순위 매기기
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending= False)
# +
# 각 시군구 내에서 매출금액(품목대분류명) 기준 순위
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목대분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목대분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending= False, axis = 1)
# +
# 클러스터링
# 시군구 - 매출금액(품목대분류명)
import numpy as np
from sklearn.cluster import KMeans
import seaborn as sns
datapoints = sigoongu_money_large.values
kmeans = KMeans(n_clusters=5).fit(datapoints)
sigoongu_money_large['cluster_id'] = kmeans.labels_
for i in range(5):
temp = sigoongu_money_large[sigoongu_money_large['cluster_id'] == i]
print(temp['cluster_id'])
# +
# 시군구 - 매출금액(품목중분류명)
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
sigoongu_money_middle = pd.DataFrame(pack, index=sigoongu)
sigoongu_money_middle = sigoongu_money_middle.fillna(0)
plt.figure(figsize=(40,10))
plt.pcolor(sigoongu_money_middle)
plt.xticks(np.arange(0.5, len(sigoongu_money_middle.columns), 1), sigoongu_money_middle, rotation=90)
plt.yticks(np.arange(0.5, len(sigoongu_money_middle.index), 1), sigoongu_money_middle.index)
plt.xlabel('__', fontsize=14)
plt.ylabel('__', fontsize=14)
plt.colorbar()
plt.show()
# +
# 시군구 - 매출금액(품목중분류명) - 순위 매기기
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False)
# +
# 각 시군구 내에서 매출금액(품목중분류명) 기준 순위
pack = []
for gu in sigoongu:
temp = df_2103[df_2103['고객소재지_시군구'] == gu]
title = list(set(temp['품목중분류명']))
title_count = {}
for t in title:
temp_b = temp[temp['품목중분류명']==t]
count = temp_b['매출금액'].sum()
title_count[t] = count
pack.append(title_count)
big_pack = pd.DataFrame(pack, index=sigoongu)
big_pack = big_pack.fillna(0)
big_pack.rank(method = 'min', ascending = False, axis = 1)
# +
# 클러스터링
# 시군구 - 매출금액(품목중분류명)
import numpy as np
from sklearn.cluster import KMeans
import seaborn as sns
datapoints = sigoongu_money_middle.values
kmeans = KMeans(n_clusters=5).fit(datapoints)
sigoongu_money_middle['cluster_id'] = kmeans.labels_
for i in range(5):
temp = sigoongu_money_middle[sigoongu_money_middle['cluster_id'] == i]
print(temp['cluster_id'])
| BCCard/2103_visualize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
all_states2 = pd.read_csv('./data/table2_all.csv', low_memory=False)
type(all_states2)
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# 3/23
import pandas as pd
df = pd.read_csv('data/df_extra_clean.csv', low_memory=False)
df
| data-collection-master/APIs/census_master/notebooks/.ipynb_checkpoints/monday_3_2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#CAPITALIZE
s=input().split()
for i in s:
print(i[0].upper()+i[1:],end="")
print(' ',end="")
# +
#Check Strict Superset
s=set(map(int,input().split()))
x=int(input())
f1=1
def check(a):
f=0
if a.issubset(s):
f=1
return f
while(x>0):
s1=set(map(int,input().split()))
t=check(s1)
f1*=t
x-=1
if f1==1:
print("True")
else:
print("False")
# -
#Cheeck Subset
x=int(input())
while(x>0):
l1=int(input())
s1=set(map(int,input().split()))
l2=int(input())
s2=set(map(int,input().split()))
if(s1.issubset(s2)):
print("True")
else:
print("False")
x-=1
# +
#Find a string
s=input()
l=len(s)
s1=input()
c=0
for i in s1:
co1=0
for j in s:
if i==j:
ch=s[co1:co1+l]
if ch==s1:
c=c+1
co1=co1+1
print(c);
# +
#merge the tools
s=input()
x=int(input())
y=x
def no_repeat(s):
l=len(s)
new=""
for i in range(l-1,-1,-1):
c=0
for j in range(i,-1,-1):
if(s[i]==s[j]):
c+=1
if c==1:
new=new+s[i]
print(new[::-1])
l=len(s)
c=l//x+1
start=0
end=l
while(c>0):
sub=s[start:x]
no_repeat(sub)
start+=y
x+=y
c-=1
# -
#Minion game
s=input();
s1=0
s2=0
vow='AEIOUaeiou'
for i in range(len(s)):
if s[i] not in vow:
s1=s1+len(s[i:])
else:
s2=s2+len(s[i:])
if s1>s2:
print("Stuart ",end="")
print(s1)
elif s2>s1:
print("Kevin ",end="")
print(s1)
else:
print("Draw")
# +
#mutation
s=input();
k,ch= input().split()
k1=int(k);
new=s[:k1]+ch+s[k1+1:];
print(new)
# -
#String Formatting
x=int(input())
for i in range(1,x+1):
octal=oct(i)
hexa=hex(i)
binary=bin(i)
print(i,octal,hexa,binary,sep=' ')
# +
#String Validators
from ast import Expr
s=input();
f1=f2=f3=f4=f5=0;
for i in s:
if i.isalnum():
f1=1;
break;
for i in s:
if i.isalpha():
f2=1
break;
for i in s:
if i.isdigit():
f3=1
break
for i in s:
if i.islower():
f4=1
break
for i in s:
if i.isupper():
f5=1
break
if f1==1:
print("True")
else:
print("False")
if f2==1:
print("True")
else:
print("False")
if f3==1:
print("True")
else:
print("False")
if f4==1:
print("True")
else:
print("False")
if f5==1:
print("True")
else:
print("False")
# -
#String-split-join
list=a.split()
l=len(list)
c=0
for i in list:
if c==l-1:
print(i)
else:
print(i+'-',end="")
c=c+1
#swap
def swap_case(s):
x=s.swapcase()
return x
s = input()
result = swap_case(s)
print(result)
# +
tempCodeRunnerFile
x=(int)(input())
s=list(map(int,input().split()))
for i in s:
c=0
for j in s:
if i==j:
c=c+1
if c>1:
break
if c==1:
print(i)
# -
#Text Wrap
s=input()
x=(int)(input())
y=x
ini=0
while (ini<=len(s)):
new=s[ini:x]
print(new)
ini=ini+y
x=x+4
# +
#The Captain's Room
x=(int)(input())
s=list(map(int,input().split()))
for i in s:
c=0
for j in s:
if i==j:
c=c+1
if c>1:
break
if c==1:
print(i)
# -
#What's your name
s=input()
s1=input()
print('Hello'+' '+s+' '+s1+': '+'You just delved into python.')
# +
#No Idea!
c=list(map(int,input().split()))
s=list(map(int,input().split()))
s1=list(map(int,input().split()))
s2=list(map(int,input().split()))
happy=0
for i in s1:
for j in s:
if i==j:
happy+=1
for i in s2:
for j in s:
if i==j:
happy-=1
print(happy)
# -
#Set Mutations
x=int(input())
a=set(map(int,input().split()))
c=int(input())
while(c>0):
s=input().split()
a1=set(map(int,input().split()))
if s[0]=='update':
a.update(a1)
if s[0]=='intersection_update':
a.intersection_update(a1)
if s[0]=='symmetric_difference_update':
a.symmetric_difference_update(a1)
if s[0]=='difference_update':
a.difference_update(a1)
if s[0]=='intersection_update':
a.intersection_update(a1)
c-=1
sum=0
for i in a:
sum+=i
print(sum)
| 'Python_string.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Regression is basically a process which predicts the relationship between x and y based on features.
# This time we are going to practice Linear Regression with Boston House Price Data that are already embedded in scikit-learn datasets
#
# **Useful functions**
# - sklearn.metrics.mean_squared_error: famous evaluation method (MSE)
#
# - np.sqrt(x): square root of tensor x
#
# - linear_model.coef_ : get `Regression coefficient` of the fitted linear model
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn.datasets as datasets
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
BOSTON_DATA = datasets.load_boston()
# TODO: get rid of the semicolon and see how the data look like.
BOSTON_DATA;
# ## Simple EDA
# Load both boston data and target, and convert it as dataframe.
def add_target_to_data(dataset):
# TODO: make the raw dataset cleaner and easier to process -> use dataframe
df = pd.DataFrame(dataset.data, columns=dataset.feature_names)
# TODO: put the target data (price) to the dataframe we just made.
print("Before adding target: ", df.shape)
df['PRICE'] = dataset.target
print("After adding target: {} \n {}\n".format(df.shape, df.head(2)))
return df
"""
10 features as default.
Why didn't I put all the 13 features? Because n_row=2 and n_col=5 as default.
It will create 10 graphs for each features.
"""
def plotting_graph(df, features, n_row=2, n_col=5):
fig, axes = plt.subplots(n_row, n_col, figsize=(16, 8))
assert len(features) == n_row * n_col
# TODO: Draw a regression graph using seaborn's regplot
for i, feature in enumerate(features):
row = int(i / n_col)
col = i % n_col
sns.regplot(x=feature, y='PRICE', data=df, ax=axes[row][col])
plt.show()
def split_dataframe(df):
label_data = df['PRICE']
# others without PRICE
# axis!! --> Whether to drop labels from the index (0 or ‘index’) or columns (1 or ‘columns’).
input_data = df.drop(['PRICE'], axis=1)
# TODO: split! Set random_state if you want consistently same result
input_train, input_eval, label_train, label_eval = train_test_split(input_data, label_data, test_size=0.3,
random_state=42)
return input_train, input_eval, label_train, label_eval
boston_df = add_target_to_data(BOSTON_DATA)
features = ['RM', 'ZN', 'INDUS', 'NOX', 'AGE', 'PTRATIO', 'LSTAT', 'RAD', 'CRIM', 'B']
plotting_graph(boston_df, features, n_row=2, n_col=5)
'''
The correlation coefficient ranges from -1 to 1.
If the value is close to 1, it means that there is a strong positive correlation between the two variables.
When it is close to -1, the variables have a strong negative correlation.
'''
correlation_matrix = boston_df.corr().round(2)
correlation_matrix
sns.heatmap(correlation_matrix, cmap="YlGnBu")
plt.show()
# ## Prediction with Linear Regression
# +
X_train, X_test, Y_train, Y_test = split_dataframe(boston_df)
# TODO: Load your machine learning model
model = LinearRegression()
# TODO: Train!
model.fit(X_train, Y_train)
# TODO: make prediction with unseen data!
pred = model.predict(X_test)
expectation = Y_test
# TODO: what is mse between the answer and your prediction?
lr_mse = mean_squared_error(expectation, pred)
# TODO: RMSE
lr_rmse = np.sqrt(lr_mse)
print('LR_MSE: {0:.3f}, LR_RMSE: {1:.3F}'.format(lr_mse, lr_rmse))
# Regression Coefficient
print('Regression Coefficients:', np.round(model.coef_, 1))
# sort from the biggest
coeff = pd.Series(data=model.coef_, index=X_train.columns).sort_values(ascending=False)
print(coeff)
# -
plt.scatter(expectation, pred)
plt.plot([0, 50], [0, 50], '--k')
plt.xlabel('Expected price')
plt.ylabel('Predicted price')
plt.tight_layout()
# ## Prediction with other Regression methods
#
# - **Ridge, Lasso and ElasticNet**
# - **Gradient Boosting Regressor**
# - **XG Boost**
# - **SGD Regressor**
# According to sklearn's official documentation,
#
# "SGDRegressor is well suited for regression problems with a large number of training samples (> 10,000), for other problems we recommend Ridge, Lasso, or ElasticNet."
# !pip install xgboost
from sklearn.linear_model import Ridge, Lasso, ElasticNet, SGDRegressor
from sklearn.ensemble import GradientBoostingRegressor
from xgboost import XGBRegressor
# +
# Try tuning the hyper-parameters
models = {
"Ridge" : Ridge(),
"Lasso" : Lasso(),
"ElasticNet" : ElasticNet(),
"Gradient Boosting" : GradientBoostingRegressor(),
"SGD" : SGDRegressor(max_iter=1000, tol=1e-3),
"XGB" : XGBRegressor(objective ='reg:linear', colsample_bytree = 0.3, learning_rate = 0.1,
max_depth = 5, alpha = 10, n_estimators = 10)
}
pred_record = {}
# -
for name, model in models.items():
# TODO: Load your machine learning model
curr_model = model
# TODO: Train!
curr_model.fit(X_train, Y_train)
# TODO: make prediction with unseen data!
pred = curr_model.predict(X_test)
expectation = Y_test
# TODO: what is mse between the answer and your prediction?
mse = mean_squared_error(expectation, pred)
# TODO: RMSE
rmse = np.sqrt(mse)
print('{} MSE: {}, {} RMSE: {}'.format(name, mse, name, rmse))
pred_record.update({name : pred})
# +
prediction = pred_record["SGD"]
plt.scatter(expectation, prediction)
plt.plot([0, 50], [0, 50], '--k')
plt.xlabel('Expected price')
plt.ylabel('Predicted price')
plt.tight_layout()
# +
prediction = pred_record["XGB"]
plt.scatter(expectation, prediction)
plt.plot([0, 50], [0, 50], '--k')
plt.xlabel('Expected price')
plt.ylabel('Predicted price')
plt.tight_layout()
# +
prediction = pred_record["Gradient Boosting"]
plt.scatter(expectation, prediction)
plt.plot([0, 50], [0, 50], '--k')
plt.xlabel('Expected price')
plt.ylabel('Predicted price')
plt.tight_layout()
# -
# ## A Little Taster Session for Neural Network
# +
from tensorflow import keras
from tensorflow.keras.layers import add, Dense, Activation
def neural_net():
model = keras.Sequential()
model.add(Dense(512, input_dim=BOSTON_DATA.data.shape[1]))
model.add(Activation('relu'))
model.add(Dense(256))
model.add(Activation('relu'))
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(64))
model.add(Activation('relu'))
model.add(Dense(1))
return model
# -
model = neural_net()
model.summary()
model.compile(loss='mse', optimizer='adam', metrics=['accuracy'])
# +
history = model.fit(X_train, Y_train, epochs=100)
loss, test_acc = model.evaluate(X_test, Y_test)
print('Test Loss : {:.4f} | Test Accuracy : {}'.format(loss, test_acc))
# +
prediction = model.predict(X_test)
plt.scatter(expectation, prediction)
plt.plot([0, 50], [0, 50], '--k')
plt.xlabel('Expected price')
plt.ylabel('Predicted price')
plt.tight_layout()
# -
| notebooks/Session02-BostonHousePrice-Solution.ipynb |