code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 5. Challenge Solution - REINFORCE
#
# 
#
# In this notebook, we're going to build a REINFORCE agent that learns to play a CartPole game. OpenAI (famously funded by <NAME>) provides a package called "gym" that has a variety of environments where we can train our agents.
# ### Basic commands
#
# We are aware that it might be your first interaction with OpenAI Gym library, therefore, we are providing you with some relevant commands you might need to use throughout both parts of the challenge.
def sample_commands():
#Creating cartpole environment
env = gym.make('CartPole-v0')
#Visualize actions
env.render()
#Take action and extracting information
observation, reward, done, info = env.step(action)
#Close window
env.close()
#Extracting the number of possible actions
env.action_space
# ### Implementation
#
# Let's start building our agent by importing the required libraries and specifying parameters. You may play around with some of the values (gamma, learning rate, etc.) to see the effect it has on training.
import gym
import numpy as np
import tensorflow as tf
import keras
from keras.layers import Input, Dense
from keras.models import Sequential
# +
#####------Configuration parameters----------------------################
seed = 42
# Discount factor for past rewards
gamma = 0.99
learning_rate = 0.01
max_steps_per_episode = 10000
# -
#######------Setting up environment----------------------################
env = gym.make("CartPole-v0")
env.seed(seed)
eps = np.finfo(np.float32).eps.item() # Smallest number such that 1.0 + eps != 1.0
# ### Define a model
# Let's first build a model. It's a simple neural nets that takes in states as inputs and predicts which policy to follow. And that's it! You're soon going to see how we can train this simple model to predict the best policy in each state.
# +
num_states = env.observation_space.shape[0]
num_hiddens1 = 32
num_hiddens2 = 32
num_actions = env.action_space.n # there are two actions in this game: left and right
# define a model
inputs = Input(shape=(num_states,))
fc1 = Dense(num_hiddens1, activation='relu')(inputs)
fc2 = Dense(num_hiddens2, activation='relu')(fc1)
outputs = Dense(num_actions, activation='softmax')(fc2)
model = keras.Model(inputs=inputs, outputs=outputs)
# -
# ### Define a training process
#
# REINFORCE is one of the Policy Gradient algorithms that learns to find an optimal policy that maximises the objective function:
#
# $$
# J(\theta) = \sum_\tau P(\tau; \theta)R(\tau)
# $$
#
# where $d_{\pi}(s)$ is probability of visiting the state $s$. Since it's nearly impossible to know the exact stationary distribution of states $d_{\pi}(s)$ especially in the continuous space, we'd like to approximate the objective function. In a nutshell, the approximation represents the expected rewards with a given policy.
#
# Its derivative that we will use to update our model is
#
# $$
# \begin {split}
# \nabla J(\theta)
# & = \mathbb{E}_{\pi} [Q_{\pi}(s,a) {\nabla}_{\theta} \ln{\pi_\theta (a|s)}]
# \\ &= \mathbb{E}_{\pi} [G_t {\nabla}_{\theta} \ln{\pi_\theta (a|s)}]
# \end {split}
# $$
#
# according to the [Policy Gradient Theorem](https://lilianweng.github.io/lil-log/2018/04/08/policy-gradient-algorithms.html#policy-gradient-theorem).
#
# To train a REINFORCE agent, we have to follow the following steps:
#
# 1. Initialize the policy with random values
# 2. Sample states, actions, and rewards from one episode
# 3. Update the policy using the gradient of the objective function $J(\theta)$
# 4. Repeat 2~3 until the policy converges
#
# We just assume that the returns we get from each episode when following the current policy represents the objective function fairly well. By sampling them multiple times through thousands of episodes, we will hopefully update the model in the right direction. It's basically how most of the deep learning models work.
# +
optimizer = tf.keras.optimizers.Adam(learning_rate=learning_rate)
action_probs_history = [] # probabilities of the selected actions for each time step
rewards_history = [] # rewards for each time step
running_reward = 0
episode_count = 0
while True:
state = env.reset() # reset the environment
episode_reward = 0
# tf.GradientTape() lets us compute the gradiets of loss with respect to model weights
with tf.GradientTape() as tape:
for timestep in range(1, max_steps_per_episode):
env.render()
state = tf.convert_to_tensor(state)
state = tf.expand_dims(state, axis=0)
policy = model(state)
action = np.random.choice(num_actions, 1, p=np.squeeze(policy))[0]
action_prob = policy[0, action] # probability of the action taken
state, reward, done, _ = env.step(action)
# collect samples
rewards_history += reward, # appends a variable to the list
action_probs_history += action_prob,
episode_reward += reward
if done:
break
# Update running reward to check condition for solving
running_reward = 0.05 * episode_reward + (1 - 0.05) * running_reward
# Calculate expected value from rewards
# - At each timestep what was the total reward received after that timestep
# - Rewards in the past are discounted by multiplying them with gamma
returns = []
discounted_sum = 0
for r in rewards_history[::-1]:
discounted_sum = r + gamma * discounted_sum
returns.insert(0, discounted_sum)
# Normalize
returns = np.array(returns)
returns = (returns - np.mean(returns)) / (np.std(returns) + eps)
returns = returns.tolist()
# compute the loss
action_probs_history = tf.convert_to_tensor(action_probs_history)
cross_entropy = -tf.math.log(action_probs_history + 1e-6)
loss = tf.reduce_sum(returns * cross_entropy)
gradients = tape.gradient(loss, model.trainable_variables) # compute the gradients of loss w.r.t model parameters
optimizer.apply_gradients(zip(gradients, model.trainable_variables)) # update the parameters accordingly
action_probs_history, rewards_history = [], [] # clear the samples
# Log details
episode_count += 1
if episode_count % 10 == 0:
template = "running reward: {:.2f} at episode {}"
print(template.format(running_reward, episode_count))
if running_reward > 195: # Condition to consider the task solved
print("Solved at episode {}!".format(episode_count))
break
| Week-09/5_Challenge Solution (REINFORCE).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Download Models and Notebooks
#
# Upload this notebook to your JupyterHub's spawned server's "work" directory to setup your notebook environment with the TensorFlow Resnet model, project notebooks, and library dependencies.
# !git clone https://github.com/tensorflow/models.git
# !cd models && git checkout v1.4.0
# !git clone https://github.com/google-aai/tf-serving-k8s-tutorial.git && cd tf-serving-k8s-tutorial && git checkout strata-ny-2018
# !conda install --yes pillow
| jupyter/download_models_and_notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
### Example of many vectors connected to a single copy node ###
import numpy as np
from contractn import TN
tn = TN()
# Add central copy tensor of order 101
copy_node = tn.add_copy_node(101)
# Connect vectors to all but one edge of the copy tensor
for i in range(100):
vec = np.array([1, 0.99])
vec_node = tn.add_dense_node(vec)
# Connect i'th axis of copy_node to 0'th axis of vec_node
tn.connect_nodes(copy_node, vec_node, i, 0)
print(tn.contract()) # array([1., 0.36603234])
# %time tn.contract() # 6.85 ms
# +
### Example of Tucker vs. CP decomposition and einsum ###
import numpy as np
from contractn import TN
# Initialize TNs for 3rd-order Tucker and CP decompositions
cp = TN()
tucker = TN()
# Add central "hub" cores
cp_hub = cp.add_copy_node(3)
tucker_hub = tucker.add_dense_node(np.ones((4, 4, 4)))
# Connect each hub to three factor matrices
for i in range(3):
mat = np.eye(4, 10)
cp_mat = cp.add_dense_node(mat)
tucker_mat = tucker.add_dense_node(mat)
cp.connect_nodes(cp_hub, cp_mat, i, 0)
tucker.connect_nodes(tucker_hub, tucker_mat, i, 0)
print(cp.einsum_str) # "ac,ad,ae->cde"
print(tucker.einsum_str) # "abc,ae,bf,cg->efg"
# +
### Example of multiplying many matrices ###
import numpy as np
from contractn import TN
# Initialize and connect together vector
# to a chain of 1000 3x3 matrices
tn = TN()
prev_node = tn.add_dense_node(np.ones((3,)))
for i in range(1000):
mat_node = tn.add_dense_node(np.ones((3, 3)))
tn.connect_nodes(prev_node, mat_node, -1, 0)
prev_node = mat_node
print(tn.contract())
# [inf inf inf]
print(tn.contract(split_format=True))
# (array([1., 1., 1.]), array(1098.61228867))
# -
| contractn/notebooks/ctn_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ""
from keras.applications import InceptionV3
from keras.applications.inception_v3 import preprocess_input
from keras.preprocessing import image
from keras.models import Model
from keras.applications.imagenet_utils import decode_predictions
import numpy as np
import cv2
from cv2 import *
import matplotlib.pyplot as plt
# %matplotlib inline
import scipy as sp
from scipy.misc import toimage
import sys
sys.path.append("..")
from src.network.hiarGoogLenet import hiarGoogLeNet
from src.network.GoogleLenet import GoogLeNet
from src.network.hiarGoogLenet_high import hiarGoogLeNet_high
from src.network.hiarBayesGoogLenet import hiarBayesGoogLeNet
low_level = [27, 32, 50, 56]#, 61, 62, 63, 64
mid_level = [0, 6, 7, 8, 9, 11, 12, 13, 17, 20, 21, 22, 23, 24, 25, 26, 28, 29, 30, 33, 35, 36, 37, 38, 39, 41, 42, 43, 44, 45, 46, 47, 48, 49, 51, 52, 53, 54, 55, 57, 58, 59, 60]
high_level = [1, 2, 3, 4, 5, 10, 14, 15, 16, 18, 19, 31, 34, 40]
image_width = 75
image_height = 160
model_h = hiarGoogLeNet.build(image_height, image_width, 3, [len(low_level), len(mid_level), len(high_level)])
model_h.load_weights("/home/anhaoran/codes/pedestrian_attibutes_wpal/models/imagenet_models/hiarGoogLeNet_PETA/binary61_final_model.h5")
model_g = GoogLeNet.build(image_height, image_width, 3, 61)
model_g.load_weights("../models/imagenet_models/GoogLeNet_PETA/binary61_final_model.h5")
#model_gh = hiarGoogLeNet_high.build(image_height, image_width, 3, [len(low_level), len(mid_level), len(high_level)])
#model_gh.load_weights("../models/imagenet_models/hiarGoogLeNet_PETA/binary61_high_final_model.h5")
model_hr = hiarBayesGoogLeNet.build(image_height, image_width, 3, [len(low_level), len(mid_level), len(high_level)])
model_hr.load_weights("../models/imagenet_models/hiarBayesGoogLeNet_PETA/binary61_multi_final500_model.h5")
# -
model_g.summary()
model_hr.summary()
from src.network.hiarGoogLenetSPP import hiarGoogLeNetSPP
model_fspp = hiarGoogLeNetSPP.build(image_height, image_width, 3, [len(low_level), len(mid_level), len(high_level)])
model_fspp.summary()
# +
def load_original(img_path, img_height, img_width):
# 把原始图片压缩为 299*299大小
img = image.load_img(img_path, target_size=(image_height, image_width, 3))
plt.figure(0)
plt.subplot(211)
plt.imshow(img)
return img
"""
def load_fine_tune_googlenet_v3(img):
# 加载fine-tuning googlenet v3模型,并做预测
model = InceptionV3(include_top=True, weights='imagenet')
model.summary()
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
preds = model.predict(x)
print('Predicted:', decode_predictions(preds))
plt.subplot(212)
plt.plot(preds.ravel())
plt.show()
return model, x
"""
# +
def extract_features(model, x, layer_id, filters, layer_num_list):
'''
提取指定模型指定层指定数目的feature map并输出到一幅图上.
:param ins: 模型实例
if len(ins) != 2:
print('parameter error:(model, instance)')
return None
:param layer_id: 提取指定层特征
:param filters: 每层提取的feature map数
:param layer_num_list: 一共提取多少层feature map
:return: None
'''
#model = ins[0]
#x = ins[1]
if type(layer_id) == type(1):
model_extractfeatures = Model(inputs=model.input, outputs=model.get_layer(index=layer_id).output)
else:
model_extractfeatures = Model(inputs=model.input, outputs=model.get_layer(name=layer_id).output)
#model_extractfeatures.summary()
fc2_features = model_extractfeatures.predict(x)
if filters > len(fc2_features[0][0][0]):
print('layer number error.', len(fc2_features[0][0][0]),',',filters)
return None
for i in range(filters):
#plt.subplots_adjust(left=0, right=1, bottom=0, top=1)
plt.subplot(filters, len(layer_num_list), layer_num_list.index(layer_id) + 1 + i * len(layer_num_list) )
plt.axis("off")
if i < len(fc2_features[0][0][0]):
plt.imshow(fc2_features[0, :, :, i])
# 层数、模型、卷积核数
def extract_features_batch(model, x, filters, layer_num_list, path):
'''
批量提取特征
:param layer_num: 层数
:param model: 模型
:param filters: feature map数
:return: None
'''
#plt.figure(figsize=(filters, layer_num))
plt.subplot(filters, len(layer_num_list), 1)
for i in layer_num_list:
extract_features(model, x, i, filters, layer_num_list)
plt.savefig(path, dpi=500, quality=95)
plt.show()
def extract_features_with_layers(model, instance, layer_indexes, img_height=160, img_width=75):
'''
提取hypercolumn并可视化.
:param layers_extract: 指定层列表
:return: None
'''
hc = extract_hypercolumn(model, instance, layer_indexes, img_height, img_width)
ave = np.average(hc.transpose(1, 2, 0), axis=2)
plt.imshow(ave)
plt.show()
def extract_hypercolumn(model, instance, layer_indexes, img_height=160, img_width=75):
'''
提取指定模型指定层的hypercolumn向量
:param model: 模型
:param layer_indexes: 层id
:param instance: 输入
:return:
'''
feature_maps = []
for i in layer_indexes:
feature_maps.append(Model(inputs=model.input, outputs=model.get_layer(index=i).output).predict(instance))
hypercolumns = []
for convmap in feature_maps:
for i in range(len(convmap[0][0][0])):
upscaled = sp.misc.imresize(convmap[0, :, :, i], size=(img_height, img_width), mode="F", interp='bilinear')
hypercolumns.append(upscaled)
return np.asarray(hypercolumns)
img_path = '/home/anhaoran/data/pedestrian_attributes_PETA/PETA/3DPeS/archive/191_494_FRAME_319_RGB.bmp'
img = load_original(img_path, image_height, image_width)
img_arr = image.img_to_array(img)
img_arr = np.expand_dims(img_arr, axis=0)
print(img_arr.shape)
"""
extract_features_batch(model_h, img_arr, 10, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_h.jpg')
extract_features_batch(model_g, img_arr, 10, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_g.jpg')
extract_features_batch(model_gh, img_arr, 10, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_gh.jpg')
"""
#extract_features_with_layers(model_h, img_arr, [1, 4, 7])
#extract_features_with_layers(model_h, img_arr, [1, 4, 7, 10, 11, 14, 17])
# -
# # Low&Mid level feature map
model_h.summary()
# +
def extract_features_hiar_models(model, x, layer_id, filters, layer_num_list):
'''
提取指定模型指定层指定数目的feature map并输出到一幅图上.
:param ins: 模型实例
if len(ins) != 2:
print('parameter error:(model, instance)')
return None
:param layer_id: 提取指定层特征
:param filters: 每层提取的feature map数
:param layer_num_list: 一共提取多少层feature map
:return: None
'''
if type(layer_id) == type(1):
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(index=layer_id).output)
else:
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(name=layer_id).output)
#model_extractfeatures.summary()
fc2_features = model_extractfeatures.predict(x)
if filters > len(fc2_features[0][0][0]):
print('layer number error.', len(fc2_features[0][0][0]),',',filters)
return None
for i in range(filters):
#plt.subplots_adjust(left=0, right=1, bottom=0, top=1)
plt.subplot(filters, len(layer_num_list), layer_num_list.index(layer_id) + 1 + i * len(layer_num_list) )
plt.axis("off")
if i < len(fc2_features[0][0][0]):
plt.imshow(fc2_features[0, :, :, i])
# 层数、模型、卷积核数
def extract_features_batch_hiar_models(model, x, filters, layer_num_list, path):
'''
批量提取特征
:param layer_num: 层数
:param model: 模型
:param filters: feature map数
:return: None
'''
#plt.figure(figsize=(filters, layer_num))
plt.subplot(filters, len(layer_num_list), 1)
for i in layer_num_list:
extract_features_hiar_models(model, x, i, filters, layer_num_list)
plt.savefig(path, dpi=250, quality=95)
plt.show()
for i in range(20):
print("filter_", str(i))
extract_features_batch_hiar_models(model_h, img_arr, i, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_all_model_filter'+str(i)+'.jpg')
# -
# # base feature map
# +
def extract_features_allmodels(model, x, layer_id, filter_id, layer_num_list):
'''
提取指定模型指定层指定数目的feature map并输出到一幅图上.
:param ins: 模型实例
if len(ins) != 2:
print('parameter error:(model, instance)')
return None
:param layer_id: 提取指定层特征
:param filters: 每层提取的feature map数
:param layer_num_list: 一共提取多少层feature map
:return: None
'''
for i in range(len(model)):
if type(layer_id) == type(1):
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(index=layer_id).output)
else:
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(name=layer_id).output)
#model_extractfeatures.summary()
fc2_features = model_extractfeatures.predict(x)
if filter_id > len(fc2_features[0][0][0]):
print('layer number error.', len(fc2_features[0][0][0]),',',filter_id)
return None
#plt.subplots_adjust(left=0, right=1, bottom=0, top=1)
plt.subplot(len(model), len(layer_num_list), layer_num_list.index(layer_id) + 1 + i * len(layer_num_list) )
plt.axis("off")
if filter_id < len(fc2_features[0][0][0]):
plt.imshow(fc2_features[0, :, :, filter_id])
plt.savefig("../results/1.jpg", dpi=250, quality=95)
# 层数、模型、卷积核数
def extract_features_batch_allmodels(model, x, filter_id, layer_num_list, path):
'''
批量提取特征
:param layer_num: 层数
:param model: 模型
:param filters: feature map数
:return: None
'''
#plt.figure(figsize=(filters, layer_num))
plt.subplot(len(model), len(layer_num_list), 1)
for i in layer_num_list:
extract_features_allmodels(model, x, i, filter_id, layer_num_list)
plt.savefig(path, dpi=250, quality=95)
plt.show()
#for i in range(20):
# print("filter_", str(i))
extract_features_batch_allmodels([model_hr, model_h, model_g], img_arr, 0, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_all_model_filter'+str(0)+'.jpg')
# +
def extract_sumfeatures_allmodels(model, x, layer_id, layer_num_list):
'''
提取指定模型指定层指定数目的feature map并输出到一幅图上.
:param ins: 模型实例
if len(ins) != 2:
print('parameter error:(model, instance)')
return None
:param layer_id: 提取指定层特征
:param filters: 每层提取的feature map数
:param layer_num_list: 一共提取多少层feature map
:return: None
'''
for i in range(len(model)):
if type(layer_id) == type(1):
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(index=layer_id).output)
else:
model_extractfeatures = Model(inputs=model[i].input, outputs=model[i].get_layer(name=layer_id).output)
#model_extractfeatures.summary()
fc2_features = model_extractfeatures.predict(x)
#plt.subplots_adjust(left=0, right=1, bottom=0, top=1)
plt.subplot(len(model), len(layer_num_list), layer_num_list.index(layer_id) + 1 + i * len(layer_num_list) )
plt.axis("off")
plt.imshow(np.sum(fc2_features[0, :, :, :], axis=2))
plt.savefig("../results/1.jpg", dpi=250, quality=95)
# 层数、模型、卷积核数
def extract_sumfeatures_batch_allmodels(model, x, layer_num_list, path):
'''
批量提取特征
:param layer_num: 层数
:param model: 模型
:param filters: feature map数
:return: None
'''
#plt.figure(figsize=(filters, layer_num))
plt.subplot(len(model), len(layer_num_list), 1)
for i in layer_num_list:
extract_sumfeatures_allmodels(model, x, i, layer_num_list)
plt.savefig(path, dpi=250, quality=95)
plt.show()
#for i in range(20):
# print("filter_", str(i))
extract_sumfeatures_batch_allmodels([model_hr, model_h, model_g], img_arr, [1, 2, 3, 4, 5, 10, 15, 20, 25, 30, 40, 50, 70, 90, 110, 130], '../results/samples/sample_all_model_filter'+str(0)+'.jpg')
# -
Model(inputs=model.input, outputs=model.get_layer(index=layer_id).output)
# # Weights
from src.network.hiarBayesGoogLenet import hiarBayesGoogLeNet
model_hr = hiarBayesGoogLeNet.build(image_height, image_width, 3, [len(low_level), len(mid_level), len(high_level)])
model_hr.load_weights("/home/anhaoran/codes/pedestrian_attibutes_wpal/models/imagenet_models/hiarBayesGoogLeNet_PETA/binary61_multi_final_model.h5")
model_hr.summary()
weights = model_hr.get_layer('high_cond').get_weights()
print(weights)
print(weights[0].shape)
# +
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
plt.subplots(figsize=(9, 9))
sns.heatmap(weights[0], cmap="Blues", square=True,
xticklabels = ['personalLess15', 'personalLess30', 'personalLess45', 'personalLess60', 'personalLarger60', 'lowerBodyCapri', 'lowerBodyCasual', 'upperBodyCasual', 'personalFemale', 'lowerBodyFormal', 'upperBodyFormal', 'lowerBodyPlaid', 'personalMale', 'upperBodyPlaid'],
yticklabels = ['upperBodyLogo', 'lowerBodyThinStripes', 'upperBodyThinStripes', 'upperBodyThickStripes', 'accessoryHeadphone', 'carryingBabyBuggy', 'carryingBackpack', 'hairBald', 'footwearBoots', 'carryingOther', 'carryingShoppingTro', 'carryingUmbrella', 'carryingFolder', 'accessoryHairBand', 'accessoryHat', 'lowerBodyHotPants', 'upperBodyJacket', 'lowerBodyJeans', 'accessoryKerchief', 'footwearLeatherShoes', 'hairLong', 'lowerBodyLongSkirt', 'upperBodyLongSleeve', 'carryingLuggageCase', 'carryingMessengerBag', 'accessoryMuffler', 'accessoryNothing', 'carryingNothing', 'upperBodyNoSleeve', 'carryingPlasticBags', 'footwearSandals', 'footwearShoes', 'hairShort', 'lowerBodyShorts', 'upperBodyShortSleeve', 'lowerBodyShortSkirt', 'footwearSneakers', 'footwearStocking', 'upperBodySuit', 'carryingSuitcase', 'lowerBodySuits', 'accessorySunglasses', 'upperBodySweater', 'lowerBodyTrousers', 'upperBodyTshirt', 'upperBodyOther', 'upperBodyVNeck'])
plt.show()
# -
| notebook/plot_featuremap&weights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stock Market Analysis for Tech Stocks
# `Trying to analyse data from stock market for some technology stocks`
#
# I am using Pandas to extract and analyse the information, visualise it, and look at different ways to analyse risk of a stock, based on its performance history.
#
# `Here are the questions I am trying to answer:`
#
# 1. What was change in a stock's price over time?
# 2. What was daily return average of a stock?
# 3. What was moving average of various stocks?
# 4. What was correlation between daily returns of different stocks?
# 5. How much value do we put at risk by investing in a particular stock?
# 6. How can we attempt to predict future stock behaviour?
# +
from __future__ import division # for float division
import pandas as pd
from pandas import Series,DataFrame
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# %matplotlib inline
#To grab stock data
from pandas_datareader import DataReader
from datetime import datetime as dt
# +
#We're going to analyse stock info for Apple, Google, Microsoft, and Amazon
tech_list = ['AAPL','GOOGL','MSFT','AMZN']
# +
#Setting the end date to today
end = dt.now()
#Start date set to 1 year back
start = dt(end.year-1,end.month,end.day)
# +
#Using Yahoo Finance to grab the stock data (`YahooDailyReader`)
# Requires connection
for stock in tech_list:
globals()[stock] = DataReader(stock,'yahoo',start,end)
# -
# * Globals method sets the stock name to a global variable
# * Globals method => Apple's stock data will be stored in the AAPL global variable dataframe. Let's see if that worked
#
# `NOTE`
# * We have Date Column as index so we are geting Keyerror Date
AAPL.head()
AAPL.describe()
# * Apple last year min, max, average can be seen
AAPL.isnull().sum()
# * No null valu foud
AAPL.info()
# * Data type neads no change
# ## Change in Stock Price Over Time
# +
# stock's adjusted closing price
AAPL['Adj Close'].plot(legend=True,figsize=(12,5))
plt.show()
# -
# ## Stock's volume being traded, over time
# +
AAPL['Volume'].plot(legend=True,figsize=(12,5))
plt.show()
# -
#
# ## Moving average of various stocks in past?
# * 10days
# * 20days
# * 40days
AAPL_c = AAPL.copy()
# +
day = [20,40,60]
for movingavg in day:
col_name = 'moving_avg for %s days' %(str(movingavg))
AAPL_c[col_name] = AAPL_c['Adj Close'].rolling(window = movingavg,center=False).mean()
# -
# * Some Rolling window calculations
# * We take a window size of k at a time and perform some desired mathematical operation on it
AAPL_c.columns
AAPL_c.isnull().sum()
# +
AAPL_c[['Adj Close','moving_avg for 20 days','moving_avg for 40 days',
'moving_avg for 60 days']].plot(subplots=False,figsize=(12,5))
plt.show()
# -
# * Moving averages for more days have a smoother plot, as they're less reliable on daily fluctuations.
# * So even though, Apple's stock has a slight dip near the start of May-2021, it's generally been on an upward trend since mid-Aug-2020.
# ## Daily return average of a stock?
# * Percentage change over the adjusted closing price
AAPL['daily_return'] = AAPL['Adj Close'].pct_change()
AAPL['daily_return'].isnull().sum()
# +
AAPL['daily_return'].plot(figsize=(20,8),legend=True,linestyle='--',marker='o')
plt.show()
# +
plt.figure(figsize=(5,3),dpi=120)
sns.distplot(AAPL['daily_return'].dropna(),bins=100,color='red')
plt.show()
# -
# * Positive daily returns seem to be slightly more frequent than negative returns for Apple
# ## Correlation between daily returns of different stocks?
tech_list = ['AAPL', 'GOOGL', 'MSFT', 'AMZN']
# +
# Only 'Adj Close'
adjclose_df = DataReader(tech_list,'yahoo',start,end)['Adj Close']
# Requires connection
# -
adjclose_df.head()
adjclose_df.isnull().sum()
df = adjclose_df.pct_change()
df.tail()
df.isnull().sum()
# * Scatterplot to visualise any correlations between different stocks.
# * First we'll visualise a scatterplot for the relationship between the daily return of a stock to itself.
# +
plt.figure(figsize=(4,3),dpi=120)
sns.jointplot('GOOGL','GOOGL',df,kind='scatter',color='blue')
plt.show()
# -
# * Relationship is perfectly linear because we're trying to correlate something with itself.
# * Now check out the relationship between Google and Apple's daily returns
# +
plt.figure(figsize=(4,3),dpi=120)
sns.jointplot('GOOGL','AAPL',df,kind='reg')
plt.show()
# -
# * Minor correlation between two stocks
# +
# all other combinations
sns.pairplot(df.dropna())
plt.title('daily returns of our stocks')
plt.show()
# -
# * More clean way will be to use correalation plot
sns.heatmap(df.dropna().corr(),annot=True)
# * Amazon and Microsoft have higher correlation
# * Every company is +Vely correlated with each other
#
# ## How much value do we put at risk by investing in a particular stock?
# * Basic way to quantify risk:
# * Compare expected return (which can be the mean of the stock's daily returns) with the standard deviation of the daily returns.
rets = df.dropna()
arrowprops=dict(arrowstyle='->', connectionstyle='arc3,rad=0.5', color='red')
# +
plt.figure(figsize=(8,5))
plt.scatter(rets.mean(),rets.std(),s=20)
plt.xlabel('Expected Return')
plt.ylabel('Risk')
for label,x,y in zip(rets.columns,rets.mean(),rets.std()):
plt.annotate(label,xy=(x,y),xytext=(-110,20),textcoords = 'offset points', ha = 'right',va = 'bottom',arrowprops = dict(arrowstyle='->',connectionstyle = "arc3, rad=-0.5"))
# -
# * Focus => stock to have a high expected return and a low risk
# * Apple stocks have higher expected returns, but also have a higher risk
# ## Value At Risk
# * Amount of money we could expect to lose for a given confidence interval.
# * We'll use the `Bootstrap` method and the `Monte Carlo Method` to extract this value.
# **`Bootstrap Method`**
# * Using this method, we calculate empirical quantiles from a histogram of daily returns.
# * Quantiles help us define our confidence interval.
# +
plt.figure(figsize=(8,5))
sns.distplot(AAPL['daily_return'].dropna(),bins=100,color='red')
plt.plot()
# -
rets.head()
rets['AAPL'].quantile(0.05)
# * 0.05 empirical quantile of daily returns is at -0.033
# * This means that with 95% confidence, worst daily loss will not exceed 3.31% (of the investment).
# ## How can we attempt to predict future stock behaviour?
# **`Monte Carlo Method`**
# * In this method, we run simulations to predict future many times, and aggregate results in end for some quantifiable value.
# +
days = 365
#delta t
dt = 1/365
mu = rets.mean()['GOOGL']
sigma = rets.std()['GOOGL']
# -
#Function takes in stock price, number of days to run, mean and standard deviation values
def stock_monte_carlo(start_price,days,mu,sigma):
price = np.zeros(days)
price[0] = start_price
shock = np.zeros(days)
drift = np.zeros(days)
for x in range(1,days):
#Shock and drift formulas taken from the Monte Carlo formula
shock[x] = np.random.normal(loc=mu*dt,scale=sigma*np.sqrt(dt))
drift[x] = mu * dt
#New price = Old price + Old price*(shock+drift)
price[x] = price[x-1] + (price[x-1] * (drift[x]+shock[x]))
return price
# * We're going to run simulation of Google stocks. Let's check out the opening value of the stock.
GOOGL.head()
#
# * Let's do a simulation of 100 runs, and plot them
# +
start_price = 1447.800049 #Taken from above
plt.figure(figsize=(12,6))
for run in range(100):
plt.plot(stock_monte_carlo(start_price,days,mu,sigma))
plt.xlabel('Days')
plt.ylabel('Price')
plt.title('Monte Carlo Analysis for Google')
plt.show()
# +
runs = 10000
simulations = np.zeros(runs)
for run in range(runs):
simulations[run] = stock_monte_carlo(start_price,days,mu,sigma)[days-1]
# +
q = np.percentile(simulations,1)
plt.hist(simulations,bins=200)
plt.figtext(0.6,0.8,s="Start price: $%.2f" %start_price)
plt.figtext(0.6,0.7,"Mean final price: $%.2f" % simulations.mean())
plt.figtext(0.6,0.6,"VaR(0.99): $%.2f" % (start_price - q,))
plt.figtext(0.15,0.6, "q(0.99): $%.2f" % q)
plt.axvline(x=q, linewidth=4, color='r')
plt.title(u"Final price distribution for Google Stock after %s days" %days, weight='bold')
# -
# `OBSERVATION`
# * We can infer from this that, Google's stock is pretty stable. The starting price that we had was USD1447.800049, and average final price over 10,000 runs was USD1454.27.
#
# * Red line indicates the value of stock at risk at the desired confidence interval. For every stock, we'd be risking USD53.77, 99% of time.
| Stock-Market-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Regression Analysis:
#
# Linear regression is where we try to find best fit line between independednt and dependent variable. For example let's consider a simple linear equation of line which has only one dependent variable say x. Then the line equation is given by:
#
# $$y = b_0 + m_1x_1$$
#
# *y* is our dependent variable and *x* is our independent variable.
# *m* is the slope and *b* is the bias.
#
# ### How it works?
#
# We fit a line across our points and then calculate the error (i.e difference between predicted value and original value) and our goal is to minimize this error by adjusting our learning parameters in this case the slope(m) and bias(b).
#
# However in case of Multivariable regression, the equation looks like
#
# $$y = \theta_0 + \theta_1x_1 +\theta_2x_2 + \theta_3x_3 + ... \theta_nx_n$$
#
#
# ### Assumptions in Regression Analysis:
#
# 1. There should be linear relationship between independent and dependent variable.
# + pair plot can be used to check this.
#
#
# 2. The independent variable should not be correlated - no multicollinearity
# + scatter plot to visualize correlation effect among variables
# + VIF value <= 4 suggests no multicollinearity whereas a value of >= 10 implies serious multicollinearity.
#
#
# 3. The error terms must have constant variance - homoskedasticity
# + plot error and predicted value. It should not be funnel shaped. It should be uniform.
#
#
# 4. There should be no correlation between the residual (error) terms - no Autocorrelation.
# + residual vs fitted values plot.
#
#
# 5. The error terms must be normally distributed.
# + Q-Q plot should be linear if normally distributed
#
#
#
# ### Regression Analysis consists of 3 stages:
#
# 1. Analyzing the correlation and directionality of the data
# 2. Estimating the model, i.e., fitting the line, and
# 3. Evaluating the validity and usefulness of the model.
#
#
# ### Covariance and corelation:
#
# Covariance signifies the direction of the linear relationship between the two variables. By direction we mean if the variables are directly proportional or inversely proportional to each other.
#
# $$Cov(x,y) =\frac{\sum (x-\overline x)(y-\overline y)}{N}$$
#
# Correlation analysis is a method of statistical evaluation used to study the strength of a relationship between two, numerically measured, continuous variables. It not only shows the kind of relation (in terms of direction) but also how strong the relationship is.
#
# $$ Corr = \frac{Cov(x,y)}{\sigma_x \sigma_y}$$
#
# The correlation coefficient is a dimensionless metric and its value ranges from -1 to +1
#
# ### Normal method for finding parameters:
#
# If we consider $\theta_0 , \theta_1,..\theta_n$ as weight parameters and $x_0, x_1, x_2,.. x_n$ as independent variables. (Note: $\theta_0$ can be considered as bias and $x_0$ = 1)
#
# $$Y = \theta * X$$
#
# Then the loss function: $$J(\theta) = (Y-X\theta)^T(Y-X\theta)$$
#
# $$\theta = (X^TX)^{-1}X^TY$$
#
# Sometimes, $X^TX$ may not be invertible, or, since matrix inversion is a compute intensive operation, sometimes, it might be computationally intensive to invert $X^TX$. In such cases, we use another method called Gradient Descent
#
# ### Gradient descent for finding parameters:
#
# Gradient deacent is nothing but moving one small step down the slope till we converge to the minimum point. Since our loss is an convex function we will have only one minima.
#
# $$\theta_i = \theta_i - \alpha \frac{\partial J(\theta)}{\partial \theta_i}$$
#
#
# where i = 1,2,.. n
#
#
# ### Loss functions:
#
# $$Mean\ Squared\ Error\ or\ L1\ loss =\frac{\sum_{i=1}^n(y_i - y_i^p)^2}{n} $$
#
# $$Mean\ Absolute\ Error\ or\ L2\ Loss = \frac{\sum_{i=1}^n|y_i - y_i^p|}{n}$$
#
# $$ Huber\ Loss\ L_{\delta}(y, f(x)) =
# \begin{cases}
# \frac{1}{2} * (y - f(x))^2, \quad |y - f(x)| \leq \delta \\
# \delta * (|y - f(x)| - \frac{1}{2} * \delta), \quad otherwise
# \end{cases}$$
#
# MSE is sensitive to outliers as the error term get squared and becomes too larage.
# MAE is robust to Outliers.
#
#
# ### Goodness of fit:
# The definition of R-squared is fairly straight-forward, it is the percentage of the response variable variation that is explained by a linear model. It is also called as coefficient of determination.
#
#
# R-squared = Explained variation / Total variation
#
# $$ R-squared\ = 1 - \frac{\sum(y_i - y_i^p)^2}{\sum(y_i - \overline y)^2}$$
#
# R-squared is always between 0 and 100%:
#
# + 0% indicates that the model explains none of the variability of the response data around its mean.
# + 100% indicates that the model explains all the variability of the response data around its mean.
#
# Adding more independent variables or predictors to a regression model tends to increase the R-squared value, which tempts makers of the model to add even more variables. This is called overfitting and can return an unwarranted high R-squared value. Adjusted R-squared is used to determine how reliable the correlation is and how much it is determined by the addition of independent variables.
#
# $$Adjusted R- Squared = 1 - \frac{(1-R^2)(N-1)}{N-p-1}$$
#
# $N$ is samplesize, $p$ is number of predictors and $R^2$ is sample R squared
#
# ### Regularization Techniques:
#
# This technique discourages learning a more complex or flexible model, so as to avoid the risk of overfitting.
#
# $$L1\ or\ L_{lasso} = \frac{1}{n}\sum_{i=1}^n\bigg(y_i - \sum_{j=1}^p\theta_jx_{ij}\bigg)^2 + \lambda\sum_{i=1}^p|\theta_j|$$
#
#
# $$L2\ or\ L_{ridge} = \frac{1}{n}\sum_{i=1}^n\bigg(y_i - \sum_{j=1}^p\theta_jx_{ij}\bigg)^2 + \lambda\sum_{i=1}^p\theta_j^2$$
#
# $\lambda$ is the tuning parameter that decides how much we want to penalize the flexibility of our model.
#
# Another technique is to use validation set and train set to avoid overfitting. K- Fold cross validation.
| 08. Regression/01. Linear Regression - Theory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# # **Lab 5b - k-Means para Quantização de Atributos**
#
# #### Os algoritmos de agrupamento de dados, além de serem utilizados em análise exploratória para extrair padrões de similaridade entre os objetos, pode ser utilizado para compactar o espaço de dados.
#
# #### Neste notebook vamos utilizar nossa base de dados de Sentiment Movie Reviews para os experimentos. Primeiro iremos utilizar a técnica word2vec que aprende uma transformação dos tokens de uma base em um vetor de atributos. Em seguida, utilizaremos o algoritmo k-Means para compactar a informação desses atributos e projetar cada objeto em um espaço de atributos de tamanho fixo.
#
# #### As células-exercícios iniciam com o comentário `# EXERCICIO` e os códigos a serem completados estão marcados pelos comentários `<COMPLETAR>`.
#
# #### ** Nesse notebook: **
# #### *Parte 1:* Word2Vec
# #### *Parte 2:* k-Means para quantizar os atributos
# #### *Parte 3:* Aplicando um k-NN
# ### **Parte 0: Preliminares**
#
# #### Para este notebook utilizaremos a base de dados Movie Reviews que será utilizada para o segundo projeto.
#
# #### A base de dados tem os campos separados por '\t' e o seguinte formato:
# `"id da frase","id da sentença","Frase","Sentimento"`
#
# #### Para esse laboratório utilizaremos apenas o campo "Frase".
# +
import os
import numpy as np
def parseRDD(point):
""" Parser for the current dataset. It receives a data point and return
a sentence (third field).
Args:
point (str): input data point
Returns:
str: a string
"""
data = point.split('\t')
return (int(data[0]),data[2])
def notempty(point):
""" Returns whether the point string is not empty
Args:
point (str): input string
Returns:
bool: True if it is not empty
"""
return len(point[1])>0
filename = os.path.join("Data","MovieReviews2.tsv")
rawRDD = sc.textFile(filename,100)
header = rawRDD.take(1)[0]
dataRDD = (rawRDD
#.sample(False, 0.1, seed=42)
.filter(lambda x: x!=header)
.map(parseRDD)
.filter(notempty)
#.sample( False, 0.1, 42 )
)
print ('Read {} lines'.format(dataRDD.count()))
print ('Sample line: {}'.format(dataRDD.takeSample(False, 1)[0]))
# -
# ### **Parte 1: Word2Vec**
#
# #### A técnica [word2vec][word2vec] aprende através de uma rede neural semântica uma representação vetorial de cada token em um corpus de tal forma que palavras semanticamente similares sejam similares na representação vetorial.
#
# #### O PySpark contém uma implementação dessa técnica, para aplicá-la basta passar um RDD em que cada objeto representa um documento e cada documento é representado por uma lista de tokens na ordem em que aparecem originalmente no corpus. Após o processo de treinamento, podemos transformar um token utilizando o método [`transform`](https://spark.apache.org/docs/latest/api/python/pyspark.mllib.html#module-pyspark.mllib.feature) para transformar cada token em uma representaçã vetorial.
#
# #### Nesse ponto, cada objeto de nossa base será representada por uma matriz de tamanho variável.
#
# [word2vec]: https://code.google.com/p/word2vec/
# ### **(1a) Gerando RDD de tokens**
#
# #### Utilize a função de tokenização `tokenize` do Lab4d para gerar uma RDD `wordsRDD` contendo listas de tokens da nossa base original.
# +
# EXERCICIO
import re
split_regex = r'\W+'
stopfile = os.path.join("Data","stopwords.txt")
stopwords = set(sc.textFile(stopfile).collect())
def tokenize(string):
""" An implementation of input string tokenization that excludes stopwords
Args:
string (str): input string
Returns:
list: a list of tokens without stopwords
"""
string_list = re.split(split_regex, string)
string_list = filter(lambda w: len(w)>0, map(lambda w: w.lower(), string_list))
return [w for w in string_list if w not in stopwords]
wordsRDD = dataRDD.map(lambda x: tokenize(x[1]))
print (wordsRDD.take(1)[0])
# -
# TEST Tokenize a String (1a)
assert wordsRDD.take(1)[0]==[u'quiet', u'introspective', u'entertaining', u'independent', u'worth', u'seeking'], 'lista incorreta!'
# ### **(1b) Aplicando transformação word2vec**
#
# #### Crie um modelo word2vec aplicando o método `fit` na RDD criada no exercício anterior.
#
# #### Para aplicar esse método deve ser fazer um pipeline de métodos, primeiro executando `Word2Vec()`, em seguida aplicando o método `setVectorSize()` com o tamanho que queremos para nosso vetor (utilize tamanho 5), seguido de `setSeed()` para a semente aleatória, em caso de experimentos controlados (utilizaremos 42) e, finalmente, `fit()` com nossa `wordsRDD` como parâmetro.
# +
# EXERCICIO
from pyspark.mllib.feature import Word2Vec
model = Word2Vec() \
.setVectorSize(5) \
.setSeed(42) \
.fit(wordsRDD)
print (model.transform(u'entertaining'))
print (list(model.findSynonyms(u'entertaining', 2)))
# -
dist = np.abs(model.transform(u'entertaining')-np.array([0.0136831374839,0.00371457682922,-0.135785803199,0.047585401684,0.0414853096008])).mean()
assert dist<1e-6, 'valores incorretos'
assert list(model.findSynonyms(u'entertaining', 1))[0][0] == 'god', 'valores incorretos'
# ### **(1c) Gerando uma RDD de matrizes**
#
# #### Como primeiro passo, precisamos gerar um dicionário em que a chave são as palavras e o valor é o vetor representativo dessa palavra.
#
# #### Para isso vamos primeiro gerar uma lista `uniqueWords` contendo as palavras únicas do RDD words, removendo aquelas que aparecem menos do que 5 vezes [$^1$](#1). Em seguida, criaremos um dicionário `w2v` que a chave é um token e o valor é um `np.array` do vetor transformado daquele token[$^2$](#2).
#
# #### Finalmente, vamos criar uma RDD chamada `vectorsRDD` em que cada registro é representado por uma matriz onde cada linha representa uma palavra transformada.
#
# ##### 1
# Na versão 1.3 do PySpark o modelo Word2Vec utiliza apenas os tokens que aparecem mais do que 5 vezes no corpus, na versão 1.4 isso é parametrizado.
#
# ##### 2
# Na versão 1.4 do PySpark isso pode ser feito utilizando o método `getVectors()
# +
# EXERCICIO
uniqueWords = (wordsRDD
.<COMPLETAR>
.<COMPLETAR>
.<COMPLETAR>
.<COMPLETAR>
.collect()
)
print ('{} tokens únicos'.format(len(uniqueWords)))
w2v = {}
for w in uniqueWords:
w2v[w] = <COMPLETAR>
w2vb = sc.broadcast(w2v) # acesse como w2vb.value[w]
print ('Vetor entertaining: {}'.format( w2v[u'entertaining']))
vectorsRDD = (wordsRDD
.<COMPLETAR>
)
recs = vectorsRDD.take(2)
firstRec, secondRec = recs[0], recs[1]
print (firstRec.shape, secondRec.shape)
# -
# TEST Tokenizing the small datasets (1c)
assert len(uniqueWords) == 3388, 'valor incorreto'
assert np.mean(np.abs(w2v[u'entertaining']-[0.0136831374839,0.00371457682922,-0.135785803199,0.047585401684,0.0414853096008]))<1e-6,'valor incorreto'
assert secondRec.shape == (10,5)
# ### **Parte 2: k-Means para quantizar os atributos**
#
# #### Nesse momento é fácil perceber que não podemos aplicar nossas técnicas de aprendizado supervisionado nessa base de dados:
#
# * #### A regressão logística requer um vetor de tamanho fixo representando cada objeto
# * #### O k-NN necessita uma forma clara de comparação entre dois objetos, que métrica de similaridade devemos aplicar?
#
# #### Para resolver essa situação, vamos executar uma nova transformação em nossa RDD. Primeiro vamos aproveitar o fato de que dois tokens com significado similar são mapeados em vetores similares, para agrupá-los em um atributo único.
#
# #### Ao aplicarmos o k-Means nesse conjunto de vetores, podemos criar $k$ pontos representativos e, para cada documento, gerar um histograma de contagem de tokens nos clusters gerados.
# #### **(2a) Agrupando os vetores e criando centros representativos**
#
# #### Como primeiro passo vamos gerar um RDD com os valores do dicionário `w2v`. Em seguida, aplicaremos o algoritmo `k-Means` com $k = 200$ e $seed = 42$.
# +
# EXERCICIO
from pyspark.mllib.clustering import KMeans
vectors2RDD = sc.parallelize(np.array(list(w2v.values())),1)
print ('Sample vector: {}'.format(vectors2RDD.take(1)))
modelK = KMeans.<COMPLETAR>
clustersRDD = vectors2RDD.<COMPLETAR>
print ('10 first clusters allocation: {}'.format(clustersRDD.take(10)))
# -
# TEST Amazon record with the most tokens (1d)
assert clustersRDD.take(10)==[142, 83, 42, 0, 87, 52, 190, 17, 56, 0], 'valor incorreto'
# #### **(2b) Transformando matriz de dados em vetores quantizados**
#
# #### O próximo passo consiste em transformar nosso RDD de frases em um RDD de pares (id, vetor quantizado). Para isso vamos criar uma função quantizador que receberá como parâmetros o objeto, o modelo de k-means, o valor de k e o dicionário word2vec.
#
# #### Para cada ponto, vamos separar o id e aplicar a função `tokenize` na string. Em seguida, transformamos a lista de tokens em uma matriz word2vec. Finalmente, aplicamos cada vetor dessa matriz no modelo de k-Means, gerando um vetor de tamanho $k$ em que cada posição $i$ indica quantos tokens pertencem ao cluster $i$.
# +
# EXERCICIO
def quantizador(point, model, k, w2v):
key = <COMPLETAR>
words = <COMPLETAR>
matrix = np.array( <COMPLETAR> )
features = np.zeros(k)
for v in matrix:
c = <COMPLETAR>
features[c] += 1
return (key, features)
quantRDD = dataRDD.map(lambda x: quantizador(x, modelK, 500, w2v))
print quantRDD.take(1)
# -
# TEST Implement a TF function (2a)
assert quantRDD.take(1)[0][1].sum() == 5, 'valores incorretos'
| ATIVIDADE5/Lab04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
""" Variable exmaples
Author: <NAME>
"""
import os
os.environ['TF_CPP_MIN_LOG_LEVEL']='2'
import numpy as np
import tensorflow as tf
# -
# Example 2: assigning values to variables
with tf.device('/cpu:0'):
sess = tf.InteractiveSession()
with tf.variable_scope("foo"):
v = tf.get_variable("var1", [1])
with tf.variable_scope("foo", reuse=True):
v = tf.get_variable("var1", [1])
x = [1]
v.assign(x)
v.initializer
print v.eval()
sess.close()
| Tensorflow Tut 1/Variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from numpy.linalg import norm
import scipy.io
import utils.starplus_utils as starp
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from mpl_toolkits.axes_grid1 import ImageGrid
from utils.plotting_utils import montage_array
import time
from utils.general_utils import rescale
import tensor.tensor_product_wrapper as tp
from numpy.random import permutation
from tensor.tensor_train import ttsvd, tt_product
# +
star_plus_data = scipy.io.loadmat('data-starplus-04847-v7.mat')
roi_tensor, my_color_map, names = starp.visualize_roi(star_plus_data)
print(roi_tensor.shape)
plt.figure(1)
montage_array(roi_tensor, cmap=my_color_map, names=names)
plt.savefig('brain1.jpg')
plt.show()
tensor_PS, labels = starp.get_labels(star_plus_data)
tensor_PS = tensor_PS / norm(tensor_PS)
# tensor_PS = rescale(tensor_PS, 1, 64)
plt.figure(2)
montage_array(tensor_PS[:, :, :, 0, 0], cmap='viridis')
plt.show()
print(tensor_PS.shape)
# -
print(len(labels[0]))
shape_T = tensor_PS.shape
dim_order = permutation(np.arange(len(shape_T)))
tol = 0.5
G, ranks = ttsvd(tensor_PS, tol, dim_order=dim_order, ranks=None)
Ak = tt_product(G, shape_T, dim_order=dim_order)
print(shape_T)
print(len(shape_T))
print(dim_order)
plt.figure(3)
montage_array(Ak[:, :, :, 0, 0], cmap='viridis')
plt.show()
print(shape_T)
print(len(shape_T))
print(dim_order)
plt.figure(3)
montage_array(Ak[:, :, :, 0, 0], cmap='viridis')
plt.show()
| code_in_progress/Star_FMRI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ox)
# language: python
# name: ox
# ---
# # Use OSMnx to get street networks by place name
#
# Author: [<NAME>](https://geoffboeing.com/)
#
# - [Overview of OSMnx](http://geoffboeing.com/2016/11/osmnx-python-street-networks/)
# - [GitHub repo](https://github.com/gboeing/osmnx)
# - [Examples, demos, tutorials](https://github.com/gboeing/osmnx-examples)
# - [Documentation](https://osmnx.readthedocs.io/en/stable/)
# - [Journal article/citation](http://geoffboeing.com/publications/osmnx-complex-street-networks/)
#
# Once you've perused the features demo and overview notebooks, this notebook provides further demonstration of querying by place name with OSMnx.
# +
import osmnx as ox
# %matplotlib inline
ox.__version__
# -
# OSMnx lets you download street network data and build topologically corrected multidigraphs, project to UTM and plot the networks, and save the street network as SVGs, GraphML files, .osm files, shapefiles, or geopackages for later use. The street networks are directed and preserve one-way directionality. API responses can be cached locally so OSMnx doesn't have to request the same data from the API multiple times: saving bandwidth, increasing speed, and enabling reproducibility.
#
# You can download a street network by providing OSMnx any of the following:
# - a bounding box
# - a lat-long point plus a distance (either distance along the network, or cardinal)
# - an address plus a distance (either distance along the network, or cardinal)
# - a place name or list of place names (for OSMnx to automatically geocode and get the boundary of)
# - a polygon of the desired street network's boundaries
#
# You can also specify several different built-in network types:
# - `drive` - get drivable public streets (but not service roads)
# - `drive_service` - get drivable streets, including service roads
# - `walk` - get all streets and paths that pedestrians can use (this network type ignores one-way directionality)
# - `bike` - get all streets and paths that cyclists can use
# - `all` - download all non-private OSM streets and paths
# - `all_private` - download all OSM streets and paths, including private-access ones
#
# Or you can define your own fine-tuned network type using OSMnx's `custom_filter` parameter (to get just highways, or railways, canals, etc).
#
# ## 1. Query/download place boundaries
#
# The `graph_from_place` function uses place boundary geocoding logic to find the network within your place's boundary. Let's look briefly at this place boundary querying before we get to street networks. You can download by cities, neighborhoods, boroughs, counties, states, or countries: anything with geocodable polygon boundaries in OSM's database. Notice the polygon geometries represent political boundaries, not physical/land boundaries. OSMnx will turn your geocoded place (or multiple places) boundaries into a geopandas GeoDataFrame.
# +
# neighborhoods or boroughs
gdf = ox.geocode_to_gdf("Manhattan, New York, New York, USA")
# counties
gdf = ox.geocode_to_gdf("Cook County, Illinois, United States")
# states
gdf = ox.geocode_to_gdf("Iowa")
# -
# you can also buffer the place boundaries (0.5 km in this example)
gdf = ox.geocode_to_gdf("Piedmont, California, USA", buffer_dist=500)
# +
# you can get multiple places in a single query
gdf = ox.geocode_to_gdf(["United Kingdom", "Ireland"])
# or optionally buffer them
places = [
"Berkeley, California, USA",
"Oakland, California, USA",
"Piedmont, California, USA",
"Emeryville, California, USA",
"Alameda, Alameda County, CA, USA",
]
gdf = ox.geocode_to_gdf(places, buffer_dist=500)
# -
# The `geocode_to_gdf` function takes a `which_result` argument. Its default value `None` makes OSMnx retrieve the first result with geometry type Polygon/MultiPolygon (if one exists on OSM). Alternatively, pass an integer value as `which_result` to retrieve a specific geocoding result, regardless of its geometry type.
#
# When querying, be specific and explicit, and sanity check the results. Try passing a dict instead of a string to be more explicit.
# +
# oops, this gets the county of alameda rather than the city!
alameda1 = ox.geocode_to_gdf("Alameda, California, USA")
# this gets the city of alameda
alameda2 = ox.geocode_to_gdf(
{
"city": "Alameda",
"county": "Alameda County",
"state": "California",
"country": "USA",
}
)
# the city is a very small part of the county
alameda1 = ox.project_gdf(alameda1)
alameda2 = ox.project_gdf(alameda2)
alameda2.area.iloc[0] / alameda1.area.iloc[0]
# -
# OSM resolves 'Mexico' to Mexico City (as the first geocoding result) and returns a single point at the center of the city. If we want the boundaries of the country of Mexico, we can 1) specify which_result=None to find the first polygon and hope it's the country, or 2) pass a dict containing a structured query to specify that we want Mexico the country instead of Mexico the city.
mexico = ox.geocode_to_gdf("Mexico", which_result=1)
type(mexico["geometry"].iloc[0])
# let the geocoder find the first Polygon/MultiPolygon result
mexico = ox.geocode_to_gdf("Mexico", which_result=None)
type(mexico["geometry"].iloc[0])
# instead of a string, you can pass a dict containing a structured query for better precision
mexico = ox.geocode_to_gdf({"country": "Mexico"})
type(mexico["geometry"].iloc[0])
# you can pass multiple queries with mixed types (dicts and strings)
mx_gt_tx = ox.geocode_to_gdf([{"country": "Mexico"}, "Guatemala", {"state": "Texas"}])
mx_gt_tx = ox.project_gdf(mx_gt_tx)
ax = mx_gt_tx.plot(fc="gray", ec="w")
_ = ax.axis("off")
# If you query 'France', OSM returns the country with all its overseas territories as result 1 and European France alone as result 2. Passing `which_result=2` instead specifically retrieves the 2nd geocoding result.
france = ox.geocode_to_gdf("France", which_result=2)
france = ox.project_gdf(france)
ax = france.plot(fc="gray", ec="none")
_ = ax.axis("off")
# Finally, note that you can also query by OSM ID rather than place name by passing `by_osmid=True` to the function. See documentation for usage details.
#
# ## 2. Get street networks by place name
#
# This "by place" querying logic works the same as the place boundary querying we just saw above.
# get the walking network for piedmont
G = ox.graph_from_place("Piedmont, California, USA", network_type="walk")
# you can also get a network with a buffer distance (meters) around the place
G = ox.graph_from_place("Piedmont, California, USA", network_type="walk", buffer_dist=200)
# +
# create a network from multiple places
places = [
"Piedmont, California, USA",
{"city": "Berkeley", "state": "California"},
"Emeryville, California, USA",
]
# use retain_all to keep all disconnected subgraphs (e.g. if your places aren't contiguous)
G = ox.graph_from_place(places, network_type="drive", retain_all=True)
fig, ax = ox.plot_graph(G, node_size=0, edge_color="#FFFF5C", edge_linewidth=0.25)
# -
# or create a network from structured place queries
places = [
{"city": "Daly City", "state": "California"},
{"city": "South San Francisco", "state": "California"},
]
G = ox.graph_from_place(places, network_type="drive")
# get the network for the borough of manhattan
G = ox.graph_from_place("Manhattan, New York, New York, USA", network_type="drive")
# get the network for the financial district neighborhood in downtown LA
place = "Financial District, Los Angeles, California, USA"
G = ox.graph_from_place(place, network_type="drive")
# %%time
# get the network for all of LA
# takes a couple minutes to do all the downloading and processing
place = "Los Angeles, California, USA"
G = ox.graph_from_place(place, network_type="drive", simplify=False, retain_all=True)
# create a network constrained to the shape of hong kong island
G = ox.graph_from_place("Hong Kong Island", network_type="drive")
| notebooks/misc/osmnx_streets/official_examples/03-graph-place-queries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ! pip install ipywidgets
# ! jupyter nbextension enable --py widgetsnbextension
import ipywidgets as widgets
import IPython.display as ipd
# +
print("How we start every example.")
text = widgets.Text(description="Your name: ")
button = widgets.Button(description="Click to say Hello")
output = widgets.Output()
ipd.display(text, button, output)
def on_button_clicked(b):
with output:
print(f"Hello {text.value} !!!")
button.on_click(on_button_clicked)
# -
w = widgets.IntSlider()
print("Slide Me... ... Then run the next cell.")
ipd.display(w)
print(f"The Value of the widget is {w.value}")
# +
# this function is called when you click the button
def on_button_clicked(button):
with output:
print(f"Name {name.value}")
print(f"GPA {gpa.value}")
print(f"DOB {dob.value}")
print(f"Major {major.value}")
# this code builds the UI, and returns each of the widgets
def build_ui():
name = widgets.Text(
value='',
placeholder='',
description='Your Name:',
disabled=False
)
gpa = widgets.FloatSlider(
value=2.0,
min=0.0,
max=4.0,
step=0.1,
description='Your GPA:',
disabled=False,
continuous_update=True,
orientation='horizontal',
readout=True,
readout_format='.1f'
)
dob = widgets.DatePicker(
description='Your DOB:',
disabled=False
)
major = widgets.RadioButtons(
options=['anthropology', 'biology', 'chemistry'],
value='biology',
description='Your Major:',
disabled=False
)
button = widgets.Button(description="Click Me!")
button.on_click(on_button_clicked)
#return all the widgets
return name, gpa, dob, major, button
# main code goes here
# build the ui, return the widgets
name, gpa, dob, major, button = build_ui()
# put the widgets is a Vertical Box
box = widgets.VBox([name,gpa,dob,major,button])
#create an output widget
output = widgets.Output()
# display the box and output
ipd.display(box, output)
# -
# Want to learn more? Check out!
#
# https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20Basics.html
| Mastering-IPyWidgets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scientific Programming
# +
## Notebook settings
# multiple lines of output per cell
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# -
# >### Today
# >
# > - Scientific programming
# > - [NumPy](#NumPy)
# > - [Matplotlib](#Matplotlib)
# >
# >
# > - Regular expressions (another notebook)
# ## A Sample of Python Libraries for Scientific Programming
#
# In addition to the Standard Library, `python.org` offers an ever-growing collection of freely available components that are constantly upgraded, tested and updated. There are many modules, packages and programs available from [PyPI](https://pypi.python.org/pypi) - the Python Package Index.
# These libraries are developed either to extend the functionalities of Python, or as more specialized/efficient version of existing modules (or both). In the remaining of this tutorial we will briefly introduce two fundamental packages for scientific computing with Python: **NumPy** and **matplotlib**.
# ## NumPy
#
# The **Num**eric(al) **Py**thon package is the core library for scientific computing in Python. It enriches Python with high performance data structures, as well as with function to manipulate such data structures and for a whole range of numerical routines.
#
# **SciPy**, another core scientific package provides a wide inventory on functions that, by operating on NumPy arrays, can be used for a range of scientific applications.
# when imported, numpy is usually renamed as "np".
import numpy as np
# #### Arrays
#
# **ndarrays** (i.e. *N*-dimensional arrays) are the main data structure on which NumPy is based.
#
# ndarrays are **multidimensional** **homogeneous** arrays composed by a **predetermined number** of items:
#
#
# - the dimensionality of an ndarray is defined by its **shape** (i.e., a tuple of integers specifying the size of each dimension, a.k.a. *axes*). The number of axes of an array is called its **rank**.
#
#
# - Sometimes "dimension" is used interchangeably for the shape and the rank of an array.
#
#
# - Ndarrays are composed by items of the same type. In NumPy arrays, types are defined by the object **dtype** (data-type), and each array is associated to one and only one dtype.
#
#
# - When a ndarray is created, its size is defined and cannot be changed.
# NumPy arrays can be created in many different ways, among which by passing a regular Python list or tuple to the `array()` function:
# let's create a rank-1 array (i.e. a vector)
a = np.array([1, 2, 3, 4, 5, 6])
print(a)
# let's verify that this is a numpy array
type(a)
# this is a rank-1 array
a.ndim
# its only axis has shape [6]
a.shape
# if not explicitly specified, array() automatically infers the dtype on the basis of the type of the
# elements in the list
a.dtype
# indexing can be used to access values...
print(a[0], a[2], a[5])
# ... or to change them
a[1] = 9
a
# the rank of an array can be changed, e.g. into 2-rank array with 3 rows and 2 columns
# NB: row*col must be equal to no. of elements (6 here)
a.reshape(3, 2)
# the same syntax can be used to create rank 2 arrays
b = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
print(b)
print(b.ndim)
print(b.shape)
# but indexing is a bit more articulated
print(b[0, 0])
print(b[2, 1])
# The `dtype` option can be used to specify to the `array()` function which type of elements are in the array (see the [NumPy documentation](https://docs.scipy.org/doc/numpy-1.10.1/user/basics.types.html) for the full list of supported data-types):
c = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]], dtype = "float32")
print(c)
# Arrays with some initial content can be created by using different functions, depending on the **initial value**...
# 2-rank array of zeros
a = np.zeros((3, 4))
print (a)
# 2-rank array of ones
b = np.ones((3, 4))
print (b)
# 2-rank array with an user-selected initial value
c = np.full((3, 4), 14.05)
print(c)
# ... or on the **function** used to generate the initial values:
# 3-rank array with random values
d = np.random.random((2, 3, 4))
print(d)
# NumPy display arrays according to the following rules:
#
# - the last axis is printed left to right
#
# - the second-to-last is printed top to bottom
#
# - the rest are printed top to bottom as well, but the slices are separated by empty lines
# #### Array Indexing
#
# 1-rank array indexing, slicing and iteration works much as for the normal Python lists:
a = np.array([1, 2, 3, 4, 5, 6])
print(a[2])
print(a[2:5])
for v in a: print(v)
# When dealing with multidimensional arrays (i.e. rank > 1), it is necessary to specify an index or a slice **for each dimension**:
# +
a = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
print(a.shape)
print(a), print("--------")
print(a[1, 1]), print("--------")
print(a[0:2]), print("--------")
print(a[:, 1:3]), print("--------")
print(a[0:2, 1:3])
# -
# unspecified indices works the same as with normal lists
print(a[:2, :3])
# missing indices are considered complete slices
print(a[1])
# negative indices can be used as well
print (a[-1])
# The iteration over multidimensional arrays starts always with respect to the first axis:
for v in a: print(v)
# **boolean indexing** allows you to pick out all the array elements that satisfy a given condition
# let's create a boolean array of the same shape as "a", where each slot tells us if
# that element of "a" is an even number
even_idx = ( a % 2 == 0 )
print(even_idx)
# let's extract the even numbers in our array
print(a[even_idx])
# #### Basic Math
#
# Basic mathematical functions apply element-wise:
# e.g. for the sum
print(b + c)
# element-wise multiplication
print(b * c)
# The function `dot()` can be used to calculate the inner product:
# inner product of rank 1 matrices, or vectors (returns a number, i.e. a scalar)
print(np.dot(np.array([9, 10]), np.array([11, 12])))
# matrix-vector multiplication
print(np.dot(np.array([[1, -1, 2],[0, -3, 1]]), np.array([2, 1, 0])))
# matrix-matrix multiplication
print(np.dot(np.array([[0, 4, -2], [-4, -3, 0]]), np.array([[0, 1], [1, -1], [2, 3]])))
# Finally, Numpy functions allow to perform computation on arrays or to transform them
# compute the sum of all the elements along an axis
print(b)
print("--------")
print(np.sum(b)) # sum all element
print("--------")
print(np.sum(b, axis=0)) # think: sum over and "collapse" axis 0 (= columns)
print("--------")
print(np.sum(b, axis=1)) # think: sum over and "collapse" axis 1 (= rows)
# transpose of a matrix
f = np.arange(6).reshape(2,3)
print(f.shape)
print(f)
print("--------")
f_transp = f.T
print(f_transp.shape)
print(f_transp)
# +
# 3 different types of vectors
a_1 = np.arange(5)
print(">> 'regular' (rank 1) vector <<")
print("rank: " + str(a_1.ndim))
print("shape: " + str(a_1.shape))
print(a_1), print("--------")
a_2 = a_1.reshape(5,1)
print(">> column vector <<")
print("rank: " + str(a_2.ndim))
print("shape: " + str(a_2.shape))
print(a_2), print("--------")
a_3 = a_2.reshape(1,5)
print(">> row vector <<")
print("rank: " + str(a_3.ndim))
print("shape: " + str(a_3.shape))
print(a_3)
# -
# `np.linalg` provides access to linear algebra functions, e.g., solvers, eigenvalues/vectors, norms and decompositions.
# inverse of a matrix
a = np.array([[1., 2.], [3., 4.]])
print(a)
print(np.linalg.inv(a))
print(np.allclose(np.dot(a,np.linalg.inv(a)),np.eye(2)))
print("--------")
print(np.linalg.det(a))
# ---
# ## Matplotlib
#
# matlotlib is a 2D **plotting library**.
#
# You can have an idea of the possibilities offered by this package by exploring the [examples](http://matplotlib.org/examples/index.html) section of the documentation
#
# The easiest way to drop plots with matplotlib it is by relying on the MATLAB-like interface provided by the `pyplot` module.
# +
import matplotlib.pyplot as plt
# to make plots in IPython using Matplotlib, you must first enable IPython's matplotlib mode
# we may want to use the backend inline, that will embed plots inside the Notebook
# %matplotlib inline
# -
# ##### Line plots
#
# Line plots represents the relationship between *x*-values and *y*-values by means of a line.
#
# Graphs of this sort can be drawn by using the [`.plot()` function](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot), with its default settings
# make an array for the x-values and one array for the y-values
x = np.arange(1, 150)
y = np.log(x)
# let's create the plot
plt.plot(x, y)
# export the plot in a .pdf file
plt.savefig("data/exemplar-lineplot.pdf")
# export the plot in a .png file
plt.savefig("data/exemplar-lineplot.png")
# Multiple calls of the `.plot()` function draw **multiple line plots** in the same figure
#
# (note that matplotlib will try to automatically manage many graphical parameters among which the axis size and the line colors)
plt.plot(x, y) # think of (x, y) as an array (of rank 2) being plotted
plt.plot(x, y * 2)
# ##### Scatterplots
#
# A scatterplot represents one or more groups of observations as points in a cartesian space.
#
# The [`scatter()` function](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) implements a method to draw this kind of plots.
n = 100
x = np.random.normal(0, 1, n)
y = np.random.normal(0, 1, n)
plt.scatter(x, y)
# We can use different marker settings for `plot()` (marker type, size)
plt.plot(x, y, ".")
# Color-coding (the default behavior of both `scatter()` and `plot()` allows you to plot different groups of data points:
plt.scatter(x, y)
plt.scatter(x * 2 , y * 2) # what's happening here? (hint: array-like input to scatter...)
# ##### Manipulating the appearance of a plot
#
# Matplotlib allows you to manipulate almost every property of a plot: figure size and resolution, line appearance, axis and grid properties, textual elements ...
x = np.arange(1, 150)
y = np.log(x)
# You to change the appearance of the **line**
#
# (NOTE: See the [matplotlib documentation](https://matplotlib.org/users/colors.html) for a quick introduction on how a color can be specified by the user)
plt.plot(x, y * 2.5, color = "r") # you can change the line color
plt.plot(x, y * 2, linewidth = 0.75) # you can change the line width
plt.plot(x, y * 1.5, linestyle = '--') # you can change the line style
plt.plot(x, y, marker = '*') # you can change the marker style
# You can manually choose the **axis limits**, the **axis titles**, the **axis ticks** and **their text** (and plot a grid, if you like):
# +
plt.plot(x, y)
# set the axis limits
plt.ylim(0, 7)
plt.xlim(-5, 155) # set the x-axes limits
# let's set the y-axis text
plt.ylabel("log (x)")
# let's set the x-axis ticks and their text (in an awful way...)
plt.xticks(range(0, 151, 50), ["zero", "fifty", "one hundred", "one hundred and fifty"], size='small')
# fancy a grid?
plt.grid(True)
# -
# You can give the plot a **main title** and add a **legend**
# +
plt.plot(x, np.log10(x))
plt.plot(x, np.log(x))
plt.plot(x, np.log2(x))
# create the main title
plt.title('Logarithms Comparison')
# add a legend
plt.legend(['base 10', 'natural', 'base 2'])
# -
# You can also **annotate** some points that may be of any interest (see [this tutorial](https://matplotlib.org/users/annotations_intro.html))
plt.plot(x, y)
plt.annotate("that's log(20) !", xy = (20, np.log(20)), xytext = (35, 2),
arrowprops=dict(facecolor='r', edgecolor='None', shrink=0.1))
# ##### Bar plots
#
# Unfortunately, humans are quite bad at comparing angles, so a better way to compare the frequencies of categorical data is to use bar plots. In these graphs:
#
# - every bar is associated with a given value
#
#
# - the height of each bar is proportional to the frequency of a given value
#
#
# - all the bars have the same width
#
# These plots can be drawn by using the [`bar()`](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.bar) or the [`barh()](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.barh) functions
# +
import string
n = 12
x = np.arange(n)
y = np.random.uniform(0.5, 1.0, n)
# -
# vertical barplot
plt.bar(x, y)
plt.xticks(x, list(string.ascii_uppercase[:12]), size='small') # let's use some random letters for each value of a variable
# horizontal barplot
plt.barh(x, y)
plt.yticks(x, list(string.ascii_uppercase[:12]), size='small')
# ##### Histograms
#
# Roughly speaking, histogram are for continuous (a.k.a. numerical) variables what barplots are for categorical variables. Also in this kind of plot, indeed, values are represented by means of bars. However:
#
#
# - each bar is associated with a bin (i.e. an interval of values)
#
#
# - the area inside the bars (rather than their height) is proportional to the frequency of a value or of a set of values
#
#
# - the width of each bar is proportional to the bin size.
#
#
#
# The [`hist()`](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.hist) function implements most of the functionalities you'll need to plot histograms.
data = np.random.normal(-1, 1, 10000)
a = plt.hist(data, density = True, edgecolor='black')
# the `bins` argument allows us to specify in how many bins we want to organize our values:
plt.hist(data, bins = 50, density = True, edgecolor='black')
# we won't draw the edge of each bar
plt.hist(data, bins = 150, density = True)
# We set `density = True` in order to plot the **frequency density** of a bin (i.e. the relative frequency of a bin divided by its width), rather than its relative (or absolute) frequency (as it is in the graph bar).
#
# When the bins are equally spaced, the relative (or absolute) frequency and the frequency density are proportional. However, this proportionality is lost when the histograms plots the density of **unequally spaced** bins (i.e. when the value of the argument `bins` is a sequence).
n, bins, patches = plt.hist(data, bins = 999, density = False)
# equal bins ('density' doesn't matter for overall 'shape')
n, bins, patches = plt.hist(data, bins = 9, density = False, edgecolor='black')
patches[4].set_fc("r")
patches[5].set_fc("r")
patches[6].set_fc("r")
patches[7].set_fc("r")
patches[8].set_fc("r")
# unequal bins. density = False (relative frequency)
n, bins, patches = plt.hist(data, bins = np.array([min(data), -3, -2, -1, max(data)]), density = False, edgecolor='black')
patches[3].set_fc("r")
# unequal bins. density = True (frequency density)
n, bins, patches = plt.hist(data, bins = np.array([min(data), -3, -2, -1, max(data)]), density = True, edgecolor='black')
patches[3].set_fc("r")
# ##### Drawing several plots per canvas
#
# Multiple plots per canvas can be drawn by using the function `subplot()` to setup a grid and activate one plot per time
# +
# %matplotlib auto
x = np.arange(1, 150)
y_base10_log = np.log10(x)
y_natural_log = np.log(x)
y_base2_log = np.log2(x)
# +
# Set up a 2 x 2 grid, set the first plot as active and draw the first curve
plt.subplot(2, 2, 1)
plt.plot(x, y_base10_log)
plt.title('Base 10 Log')
# uncomment to compare subplots with the same y-axis limits
# plt.ylim(0,8)
# +
# activate the second subplot and draw the second curve
plt.subplot(2, 2, 2)
plt.plot(x, y_natural_log)
plt.title('Natural Log')
# uncomment to compare subplots with the same y-axis limits
# plt.ylim(0,8)
# +
# activate the third subplot and draw the third curve
plt.subplot(2, 2, 3)
plt.plot(x, y_base2_log)
plt.title('Base 2 Log')
# uncomment to compare subplots with the same y-axis limits
# plt.ylim(0,8)
# -
plt.tight_layout() # adjust spacing between subplots to minimize the overlaps.
plt.show()
# > **Further Reading:**
# >
# > Tutorial introductions to the Python scientific computing libraries (including other libraries that we'll briefly use in this class, such as "SciPy", "Pandas" and "scikit-learn") can be found in:
# >
# > - the [SciPy Lecture Notes](http://www.scipy-lectures.org/) ( the [pdf version](http://www.scipy-lectures.org/_downloads/ScipyLectures-simple.pdf)).
# >
# >
# > - The `python-course.eu` section dedicated to [Numerical Python](https://www.python-course.eu/numerical_programming.php).
# >
# >
# > - For a gentle introduction to the main matplotlib modules, have a look at the [beginner's guide](https://matplotlib.org/users/beginner.html) available on `matplotlib.org`.
# >
# >
# > - An excellent plotting package built on top of matplotlib is called [`Seaborn`](https://seaborn.pydata.org).
# ### Exercise 1.
#
# Plot the sine and the cosine waves in the same graph
import math
# +
x = np.arange(0, math.pi*4, 0.05)
def my_sin(d):
return [math.sin(r) for r in d]
def my_cos(d):
return [math.cos(r) for r in d]
plt.plot(x, my_sin(x))
plt.plot(x, my_cos(x))
# -
# ### Exercise 2.
#
# Read the file data/adams-hhgttg.txt and:
#
#
# - count the occurrences of each word.
#
#
# - Create a graph plotting word frequencies (on the **y-axis**) by words sorted by frequency (that is, you list the corpus words from the most frequency to the least frequent words **x-axis**). For clarity, please do not report the words in the x-axis label.
#
#
# - In a similar graph, plot the frequencies of the top-frequent 25 words.
import nltk
import re
from collections import Counter
text = open("data/adams-hhgttg.txt").read().lower()
words = nltk.regexp_tokenize(text, re.compile("[a-z]+"))
count = Counter(words).most_common()
# + [markdown] slideshow={"slide_type": "skip"}
# ---
| notebooks/4_ScientificProgramming.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// ## Basic Metrics
//
// - Counting things
// - Why is counting good?
// - Why is counting bad?
// - Word metrics, Sentence metrics, Document metrics
// - Vocabulary
// - How could these be used pedagogically?
// - What does the theory say?
//Load helper objects
import $ivy.`org.scalaj:scalaj-http_2.11:2.3.0`
import $file.imports.IO
| archive/lasi17/Notebook_03-BasicMetrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D
from tensorflow.keras.layers import MaxPooling2D
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import numpy as np
from tensorflow.keras.preprocessing import image
def load_dataset():
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
test_datagen = ImageDataGenerator(rescale = 1./255)
training_set = train_datagen.flow_from_directory('train',
target_size = (32, 32),
batch_size = 32,
class_mode = 'categorical')
test_set = test_datagen.flow_from_directory('dummy_test',
target_size = (32, 32),
batch_size = 32,
class_mode = 'categorical')
return training_set, test_set
def define_model():
classifier = Sequential()
classifier.add(Conv2D(32, (3, 3), input_shape = (32, 32, 3), activation = 'relu'))
classifier.add(MaxPooling2D(pool_size = (2, 2)))
classifier.add(Flatten())
classifier.add(Dense(units = 128, activation = 'relu'))
classifier.add(Dense(units = 10, activation = 'softmax'))
classifier.compile(optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['accuracy'])
return classifier
def evaluate_model():
dataX, dataY = load_dataset()
model = define_model()
model.fit_generator(dataX,
# steps_per_epoch = 20000,
epochs = 2,
validation_data = dataY,
# validation_steps = 3000
)
model.save('digitRecog.h5')
# + slideshow={"slide_type": "-"}
evaluate_model()
# -
from tensorflow.keras.models import load_model
def predict_digit(image_path):
model = load_model("digitRecog.h5")
individual_image=image.load_img(image_path, target_size = (32, 32))
individual_image = image.img_to_array(individual_image)
individual_image = np.expand_dims(individual_image, axis = 0)
result = model.predict(individual_image)
return result[0]
predict_digit('image.jpg')
predict_digit('zero.png')
predict_digit('two.png')
training_set.class_indices
| ImageHindiRecog.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
# implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Spracovanie textu
#
# URL https://github.com/FIIT-IAU
#
#
# ## Extrakcia čŕt z textu
#
# Aby sme mohli text klasifikovať, hľadať zhluky podobných dokumentov a pod.
#
# #### Príklad: chceme rozlíšiť, kto je autorom textu
#
# <NAME> vs. <NAME> vs. HP Lovecraft: https://www.kaggle.com/c/spooky-author-identification
#
# **Aké črty (features) by som mohol extrahovať z viet?**
# * Dĺžka vety
# * Počet slov vo vete
# * Priemerná dĺžka slov vo vete
# * Zložitosť vety (tzv. text readability metrics, napr. Flesh-Kincaid)
# * Počet spojok/predložiek/iných slovných druhov
# * **Frekvencia použitých slov - prevod vety (textu) do vektorovej reprezentácie**
#
# #### Vo všeobecnosti
#
# * Segmentácia textu
# * Prevod textu do vektorovej reprezentácie (tzv. *bag of words*)
# * Identifikácia kľúčových slov, resp. často sa spolu vyskytujúcich slov (tokenov)
# * Určovanie podobnosti dvoch textových dokumentov
#
# ## Metódy na spracovanie textu
# - Regulárne výrazy, konečné automaty, bezkontextové gramatiky
# - Pravidlové, slovníkové prístupy
# - Prístupy strojového učenia (Markovovské modely, **hlboké neurónové siete**)
#
#
# #### Väčšina metód je jazykovo-závislá
# - Veľa dostupných modelov pre angličtinu, nemčinu, španielčinu, ...
#
#
# #### Reprezentácia textu
# - Textový dokument väčšinou reprezentujeme pomocou množiny slov (angl. *bag-of-words*) = **vektorom**.
# - Zložky vektora predstavujú jednotlivé slová, resp. n-gramy zo slovníka (pre celý korpus/jazyk).
#
# Hodnotou zložiek vektora môže byť:
# * výskyt (binárne)
# * početnosť
# * frekvencia
# * váhovaná frekvencia
#
# #### Prevod textu na vektor
#
# 1. Tokenizácia (rozdelenie textu na vety, následne na slová)
# 2. Normalizácia textu
# - prevod na malé písmená
# - stemming alebo lematizácia
# - odstránenie stopslov (spojky, predložky a pod.)
# 3. Vytvorenie slovníka
# 4. Vytvorenie vektora - zložky slová zo slovníka; väčšinou riedky (angl. *sparse*; veľa núl)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tokenizácia
# + slideshow={"slide_type": "fragment"}
import nltk
# nltk.download('punkt')
text = """At eight o'clock on Thursday morning
... Arthur didn't feel very good. He closed his eyes and went to bed again."""
# + slideshow={"slide_type": "slide"}
sentences = nltk.sent_tokenize(text)
print(sentences)
# + slideshow={"slide_type": "slide"}
sent = sentences[0]
tokens = nltk.word_tokenize(sent)
print(tokens)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Normalizácia
# + slideshow={"slide_type": "fragment"}
tokens = [token.lower() for token in tokens if token not in ".,?!..."]
print(tokens)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Stemming alebo lematizácia?
#
# - Stemming vráti koreň slova. Napr. *ryba -> ryb*
# - Lematizácia prevádza slová na ich základný slovníkový tvar. Napr. *rybe -> ryba*
# - **Je to vždy jedno alebo druhé.** Prevod na koreň slova sa používa viac pri málo flexných jazykoch (napr. angličtina). Pri flexných jazykoch (napr. slovenčina) je preferovaná lematizácia.
# - **Stemming** - pre angličtinu napr. [Porterov algoritmus (1980)](https://www.cs.odu.edu/~jbollen/IR04/readings/readings5.pdf)
# - **Lematizácia** - väčšinou slovníkové metódy (morfologická databáza, resp. tvaroslovník); pre slovenčinu: https://korpus.sk/morphology_database.html
# + slideshow={"slide_type": "slide"}
porter = nltk.PorterStemmer()
stemmed = [porter.stem(token) for token in tokens]
print(stemmed)
# + slideshow={"slide_type": "slide"}
# nltk.download('wordnet')
wnl = nltk.WordNetLemmatizer()
lemmatized = [wnl.lemmatize(token) for token in tokens]
print(lemmatized)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Odstránenie stopslov
# + slideshow={"slide_type": "fragment"}
# nltk.download('stopwords')
stopwords = nltk.corpus.stopwords.words('english')
normalized_tokens = [token for token in stemmed if token not in stopwords]
print(normalized_tokens)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Prevod na vektorovú reprezentáciu
#
# Datasetom [20 newsgroups](http://qwone.com/~jason/20Newsgroups/):
#
# *"The 20 Newsgroups data set is a collection of approximately 20,000 newsgroup documents, partitioned (nearly) evenly across 20 different newsgroups. To the best of my knowledge, it was originally collected by <NAME>, probably for his Newsweeder: Learning to filter netnews paper, though he does not explicitly mention this collection. The 20 newsgroups collection has become a popular data set for experiments in text applications of machine learning techniques, such as text classification and text clustering."*
# + slideshow={"slide_type": "slide"}
from sklearn.datasets import fetch_20newsgroups
categories = ['alt.atheism', 'soc.religion.christian', 'comp.graphics', 'sci.med']
twenty_train = fetch_20newsgroups(subset='train', categories=categories, shuffle=True, random_state=42)
# + slideshow={"slide_type": "fragment"}
twenty_train.target_names
# + slideshow={"slide_type": "fragment"}
len(twenty_train.data)
# + slideshow={"slide_type": "slide"}
print("\n".join(twenty_train.data[0].split("\n")[:10]))
# + slideshow={"slide_type": "slide"}
def preprocess_text(text):
tokens = nltk.word_tokenize(text)
stopwords = nltk.corpus.stopwords.words('english')
return [token.lower() for token in tokens if token.isalpha() and token.lower() not in stopwords]
# + slideshow={"slide_type": "fragment"}
tokenized_docs = [preprocess_text(text) for text in twenty_train.data]
# + slideshow={"slide_type": "fragment"}
print(tokenized_docs[0])
# + slideshow={"slide_type": "slide"}
from gensim import corpora, models, similarities
dictionary = corpora.Dictionary(tokenized_docs)
corpus = [dictionary.doc2bow(doc) for doc in tokenized_docs]
print(corpus[10])
# + [markdown] slideshow={"slide_type": "slide"}
# ## TF-IDF = term frequency * inverse document frequency
#
# `TF` – frekvencia slova v aktuálnom dokumente
#
# `IDF` – záporný logaritmus pravdepodobnosti výskytu slova v dokumentoch korpusu (rovnaká pre všetky dokumenty)
#
# ### $ tf(t,d)=\frac{f_{t,d}}{\sum_{t' \in d}{f_{t',d}}} $
#
# ### $ idf(t,D) = -\log_2{\frac{|{d \in D: t \in d}|}{N}} = \log_2{\frac{N}{|{d \in D: t \in d}|}} $
#
# Rôzne varianty (váhovacie schémy): https://en.wikipedia.org/wiki/Tf%E2%80%93idf
# + slideshow={"slide_type": "slide"}
tfidf_model = models.TfidfModel(corpus)
tfidf_corpus = tfidf_model[corpus]
tfidf_corpus[10][:10]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Podobnosť vektorov
#
# Podobnosť pomocou Euklidovskej vzdialenosti
#
# ### $ sim(u,v) = 1- d(u,v) = 1 - \sqrt{\sum_{i=1}^{n}{(v_i-u_i)^2}} $
#
# Kosínusová podobnosť
#
# ### $sim(u,v) = cos(u,v) = \frac{u \cdot v}{||u||||v||} =\frac{\sum_{i=1}^{n}{u_iv_i}}{\sum_{i=1}^{n}{u_i}\sum_{i=1}^{n}{v_i}} $
# + slideshow={"slide_type": "slide"}
index = similarities.MatrixSimilarity(tfidf_corpus)
index[tfidf_corpus[0]]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Extrakcia čŕt pomocou scikit-learn
#
# http://scikit-learn.org/stable/modules/feature_extraction.html#text-feature-extraction
#
# https://scikit-learn.org/stable/tutorial/text_analytics/working_with_text_data.html
# + slideshow={"slide_type": "fragment"}
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
# + slideshow={"slide_type": "fragment"}
count_vect = CountVectorizer(stop_words='english')
X_train_counts = count_vect.fit_transform(twenty_train.data)
X_train_counts.shape
# + slideshow={"slide_type": "fragment"}
print(count_vect.vocabulary_.get(u'algorithm'))
# + slideshow={"slide_type": "slide"}
tfidf_transformer = TfidfTransformer()
X_train_tfidf = tfidf_transformer.fit_transform(X_train_counts)
# + [markdown] slideshow={"slide_type": "fragment"}
# Natrénujeme klasifikátor
# + slideshow={"slide_type": "fragment"}
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB().fit(X_train_tfidf, twenty_train.target)
# + slideshow={"slide_type": "slide"}
docs_new = ['God is love', 'OpenGL on the GPU is fast']
X_new_counts = count_vect.transform(docs_new)
X_new_tfidf = tfidf_transformer.transform(X_new_counts)
predicted = clf.predict(X_new_tfidf)
for doc, category in zip(docs_new, predicted):
print('%r => %s' % (doc, twenty_train.target_names[category]))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Sprehľadnenie a automatizácia predspracovania: Pipelines
#
# https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html
#
# http://zacstewart.com/2014/08/05/pipelines-of-featureunions-of-pipelines.html
# + slideshow={"slide_type": "slide"}
from sklearn.pipeline import Pipeline
text_ppl = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB())
])
# + slideshow={"slide_type": "fragment"}
text_ppl.fit(twenty_train.data, twenty_train.target)
# + slideshow={"slide_type": "fragment"}
[twenty_train.target_names[cat] for cat in text_ppl.predict(docs_new)]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Vlastný transformátor
# + slideshow={"slide_type": "fragment"}
from sklearn.base import TransformerMixin
class MyTransformer(TransformerMixin):
def __init__(self):
pass
def fit(self, X, y=None, **fit_params):
return self
def transform(self, X, **transform_params):
return X
# + [markdown] slideshow={"slide_type": "slide"}
# # Iné časté úlohy (pred)spracovania textu
# + [markdown] slideshow={"slide_type": "slide"}
# ## Rozpoznávanie slovných druhov = Part-of-Speech Tagging (POS)
#
# Slovný druh, číslo, čas, prípadne ďalšie gramatické kategórie
# + slideshow={"slide_type": "fragment"}
# nltk.download('averaged_perceptron_tagger')
tagged = nltk.pos_tag(nltk.word_tokenize(sent))
print(tagged)
# + slideshow={"slide_type": "fragment"}
# nltk.download('tagsets')
nltk.help.upenn_tagset('NNP')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Rozpoznávanie menných entít = Name Entity Recognition (NER)
#
# Osoby, organizácie, miesta a pod.
# + slideshow={"slide_type": "fragment"}
# nltk.download('maxent_ne_chunker')
# nltk.download('words')
entities = nltk.chunk.ne_chunk(tagged)
# + slideshow={"slide_type": "fragment"}
print(entities.__repr__())
# + [markdown] slideshow={"slide_type": "slide"}
# ## N-gramy
#
# Vo všobecnosti ide o postupnosť $N$ po sebe idúcich položiek. V texte väčšinou na úrovni slov.
# - bigramy
# - trigramy
# - skipgramy - $k$-skip-$n$-gramy
# - https://books.google.com/ngrams
# + slideshow={"slide_type": "slide"}
tokens = nltk.word_tokenize(sent)
bigrams = list(nltk.bigrams(tokens))
print(bigrams[:5])
# + [markdown] slideshow={"slide_type": "fragment"}
# Dá sa nastaviť aj v `CountVectorizer` transformátore.
# + slideshow={"slide_type": "fragment"}
bigram_vectorizer = CountVectorizer(ngram_range=(1, 2), token_pattern=r'\b\w+\b', min_df=1)
analyze = bigram_vectorizer.build_analyzer()
analyze('Bi-grams are cool!')
# + [markdown] slideshow={"slide_type": "slide"}
# ## WordNet
#
# * Lexikálna databáza
# * Organizovaná pomocou tzv. synsetov (množín synoným)
# * Podstatné mená, slovesá, prídavné mená, príslovky
# * Prepojenia medzi synsetmi
# * Antonymá, hyperonymá, hyponymá, holonymá, meronymá
# + slideshow={"slide_type": "slide"}
from nltk.corpus import wordnet as wn
# + slideshow={"slide_type": "fragment"}
print(wn.synsets('car'))
# + slideshow={"slide_type": "fragment"}
car = wn.synset('car.n.01')
# + slideshow={"slide_type": "fragment"}
car.lemma_names()
# + slideshow={"slide_type": "fragment"}
car.definition()
# + slideshow={"slide_type": "fragment"}
car.examples()
# + slideshow={"slide_type": "slide"}
print(car.hyponyms()[:5])
# + slideshow={"slide_type": "fragment"}
car.hypernyms()
# + slideshow={"slide_type": "fragment"}
print(car.part_meronyms()[:5])
# + slideshow={"slide_type": "fragment"}
wn.synsets('black')[0].lemmas()[0].antonyms()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Vektorová reprezentácia slov - word2vec
#
# Každé slovo má naučený vektor reálnych čísel, ktoré reprezentujú rôzne jeho vlastnosti a zachytávajú viaceré lingvistické pravidelnosti. Môžeme počítať podobnosť medzi slovami ako podobnosť dvoch vektorov.
#
# vector('Paris') - vector('France') + vector('Italy') ~= vector('Rome')
#
# vector('king') - vector('man') + vector('woman') ~= vector('queen')
#
# - https://radimrehurek.com/gensim/models/word2vec.html
# - https://medium.com/@mishra.thedeepak/word2vec-in-minutes-gensim-nlp-python-6940f4e00980
# + slideshow={"slide_type": "slide"}
from nltk.corpus import brown
nltk.download('brown')
sentences = brown.sents()
model = models.Word2Vec(sentences, min_count=1)
model.save('brown_model')
model = models.Word2Vec.load('brown_model')
# + slideshow={"slide_type": "fragment"}
# print(model.most_similar("mother"))
print(model.wv.most_similar("mother"))
# + slideshow={"slide_type": "fragment"}
# print(model.doesnt_match("breakfast cereal dinner lunch".split()))
print(model.wv.doesnt_match("breakfast cereal dinner lunch".split()))
# + [markdown] slideshow={"slide_type": "slide"}
# # Užitočné slovníky
# - ConceptNet: http://conceptnet.io/
# - Sentiment a emócie: [WordNet-Affect](http://wndomains.fbk.eu/wnaffect.html),
# - [SenticNet](https://sentic.net/),
# - [EmoSenticNet](https://www.gelbukh.com/emosenticnet/)
#
#
# # Nástroje na spracovanie textu v Pythone
#
# - [NLTK](http://www.nltk.org/)
# - [Gensim](https://radimrehurek.com/gensim/tutorial.html)
# - [sklearn.feature_extraction.text](http://scikit-learn.org/stable/modules/feature_extraction.html#text-feature-extraction)
#
# #### Nástroje (mimo Pythonu)
# - [Stanford CoreNLP](https://stanfordnlp.github.io/CoreNLP/) - rozhranie aj cez NLTK
# - [Apache OpenNLP](https://opennlp.apache.org/)
# - [WordNet](https://wordnet.princeton.edu/) - rozhranie cez NLTK
#
#
# # Extrakcia čŕt sa robí aj s inými typmi vstupov
# - Obrázky (sklearn.feature_extraction.image, [scikit-image](https://scikit-image.org/))
# - Videá ([scikit-video](http://www.scikit-video.org/stable/))
# - Signál, napr. zvuk ([scikit-signal](https://docs.scipy.org/doc/scipy/reference/signal.html), [scikit-sound](http://work.thaslwanter.at/sksound/html/))
#
#
# # Ďalšie lingvistické modely
# - [fastText](https://fasttext.cc/), [ELMo](https://allennlp.org/elmo), [BERT](https://github.com/google-research/bert), [GloVe](https://nlp.stanford.edu/projects/glove/): nejaké základné porovnanie [tu](https://www.quora.com/What-are-the-main-differences-between-the-word-embeddings-of-ELMo-BERT-Word2vec-and-GloVe)
# - [sentence embeddings](https://github.com/oborchers/Fast_Sentence_Embeddings)
# - [doc2vec](https://radimrehurek.com/gensim/models/doc2vec.html)
# - ...a ďalšie
#
# # Pre slovenčinu
#
# - [NLP4SK](http://arl6.library.sk/nlp4sk/)
# - [Slovenský národný korpus](https://korpus.sk/)
# - [word2vec](https://github.com/essential-data/word2vec-sk)
# - a [ďalšie...](https://github.com/essential-data/nlp-sk-interesting-links)
#
#
# # Zdroje
# - [<NAME>, <NAME>: Speech and Language Processing](https://web.stanford.edu/~jurafsky/slp3/)
# - http://www.nltk.org/book/
# - https://radimrehurek.com/gensim/tutorial.html
# - https://scikit-learn.org/stable/tutorial/text_analytics/working_with_text_data.html
# -
| cvicenia/tyzden-07/IAU_073_nlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# k-Nearest Neighbour Classifier using sklearn
# =================================
#
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score, confusion_matrix
# +
#Loading data and preprocessing
url='http://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data'
df=pd.read_csv(url)
df.columns=['sepal_length','sepal_width','petal_length','petal_width','flower_type']
df['flower_type'] = df['flower_type'].astype('category')
df.flower_type = df.flower_type.cat.rename_categories([0,1,2])
D=df.values
# Get the labelled set
c1=D[:20,:]; c2=D[50:70,:]; c3=D[100:120,:]
trainSet = np.concatenate((c1,c2,c3),axis=0)
# Get the testing set
c1 = D[21:50,:]; c2=D[71:100,:]; c3=D[121:,:]
testSet = np.concatenate((c1,c2,c3),axis=0)
xTrain=trainSet[:,:-1]; yTrain=trainSet[:,-1]
xTest=testSet[:,:-1]; yTest=testSet[:,-1]
# -
# create a knn classifier with K=3
clf = KNeighborsClassifier(n_neighbors=3)
clf.fit(xTrain, yTrain.astype(int))
# +
# Make predictions
#accuracy_score function nije banate hbe
yPred=clf.predict(xTest)
acc=accuracy_score(yTest.astype(int), yPred.astype(int))
print('Accuracy with 3 neighbours: ',acc)
# -
#confusion matrix function nije banate hbe
def plot_conf_mat(lTrue, lPred, title):
""" A function for plotting the confusion matrix given true and predicted labels."""
cm = confusion_matrix(lTrue.astype(int), lPred.astype(int))
print(cm)
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cm)
plt.title(title)
plt.xlabel('Predicted')
plt.ylabel('True')
plt.show()
plot_conf_mat(yTest, yPred, 'K=3')
#f = open("SRR6055422", "r")
f = open("data.csv", encoding="utf8")
text = f.readlines()
print(text)
# +
import pandas as pd
#import sys
#reload(sys)
#sys.setdefaultencoding("ISO-8859-1")
#importing the dataset
#dataset = pd.read_csv('SRR6055422.csv')
f = open("data.csv","rb")
text = f.read().decode(errors='replace')
print(text[10])
# -
#import pandas as pd
df=pd.read_csv('data.csv', encoding="ISO-8859-1")
#print(text[:1000])
df.head()
#print(text[:100000])
#c03de09c8e7a70a37fd398b725415bd6
text2 = text.split()
print(len(text2))
'''
for i in range(len(text2)):
if text2[i] == "*idx1":
print(text2[i-1])
'''
print(text[:100000])
'''
import gzip
import shutil
with gzip.open('SRR6055422.fastq.gz', 'rb') as f_in:
with open('SRR6055422.fastq', 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
'''
symbol=['@','+']
"""Stores both the sequence and the quality values for the sequence"""
f = open('SRR6055422.csv','rU')
for lines in f:
#if symbol not in lines.startwith()
data = f.readlines()
print(data)
import pandas as pd
df = pd.read_csv("data.csv")
| lab 4/lab 4/knn_using_sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TSSL Lab 3 - Nonlinear state space models and Sequential Monte Carlo
# In this lab we will make use of a non-linear state space model for analyzing the dynamics of SARS-CoV-2, the virus causing covid-19. We will use an epidemiological model referred to as a Susceptible-Exposed-Infectious-Recovered (SEIR) model. It is a stochastic adaptation of the model used by the The Public Health Agency of Sweden for predicting the spread of covid-19 in the Stockholm region early in the pandemic, see [Estimates of the peak-day and the number of infected individuals during the covid-19 outbreak in the Stockholm region, Sweden February – April 2020](https://www.folkhalsomyndigheten.se/publicerat-material/publikationsarkiv/e/estimates-of-the-peak-day-and-the-number-of-infected-individuals-during-the-covid-19-outbreak-in-the-stockholm-region-sweden-february--april-2020/).
#
# The background and details of the SEIR model that we will use are available in the document _TSSL Lab 3 Predicting Covid-19 Description of the SEIR model_ on LISAM. Please read through the model description before starting on the lab assignments to get a feeling for what type of model that we will work with.
# ---
#
# ### DISCLAIMER
# Even though we will use a type of model that is common in epidemiological studies and analyze real covid-19 data, you should _NOT_ read to much into the results of the lab. The model is intentionally simplified to fit the scope of the lab, it is not validated, and it involves several model parameters that are set somewhat arbitrarily. The lab is intended to be an illustration of how we can work with nonlinear state space models and Sequential Monte Carlo methods to solve a problem of practical interest, but the actual predictions made by the final model should be taken with a big grain of salt.
#
# ---
#
# We load a few packages that are useful for solving this lab assignment.
import pandas # Loading data / handling data frames
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = (10,6) # Increase default size of plots
# ## 3.1 A first glance at the data
# The data that we will use in this lab is a time series consisting of daily covid-19-related intensive care cases in Stockholm from March to August. As always, we start by loading and plotting the data.
data=pandas.read_csv('SIR_Stockholm.csv',header=0)
y_sthlm = data['ICU'].values
u_sthlm = data['Date'].values
ndata = len(y_sthlm)
plt.plot(u_sthlm,y_sthlm)
plt.xticks(range(0, ndata, 7), u_sthlm[::7], rotation = 90) # Show only one tick per week for clarity
plt.xlabel('Date')
plt.ylabel('New ICU cases')
plt.show()
# **Q0:** What type of values can the observations $y_t$ take? Is a Gaussian likelihood model a good choice if we want to respect the properties of the data?
# **A:** Observations $y_t$ can take only positive values (greateer than or equal to zero). Hence, a gaussian likelihood model is not an option as it would be equally likely for negative $y_t$ to be observed under that assumption.
# ## 3.2 Setting up and simulating the SEIR model
# In this section we will set up a SEIR model and use this to simulate a synthetic data set. You should keep these simulated trajectories, we will use them in the following sections.
# +
from tssltools_lab3 import Param, SEIR
"""For Stockholm the population is probably roughly 2.5 million."""
population_size = 2500000
"""" Binomial probabilities (p_se, p_ei, p_ir, and p_ic) and the transmission rate (rho)"""
pse = 0 # This controls the rate of spontaneous s->e transitions. It is set to zero for this lab.
pei = 1 / 5.1 # Based on FHM report
pir = 1 / 5 # Based on FHM report
pic = 1 / 1000 # Quite arbitrary!
rho = 0.3 # Quite arbitrary!
""" The instantaneous contact rate b[t] is modeled as
b[t] = exp(z[t])
z[t] = z[t-1] + epsilon[t], epsilon[t] ~ N(0,sigma_epsilon^2)
"""
sigma_epsilon = .1
""" For setting the initial state of the simulation"""
i0 = 1000 # Mean number of infectious individuals at initial time point
e0 = 5000 # Mean number of exposed...
r0 = 0 # Mean number of recovered
s0 = population_size - i0 - e0 - r0 # Mean number of susceptible
init_mean = np.array([s0, e0, i0, 0.], dtype=np.float64) # The last 0. is the mean of z[0]
"""All the above parameters are stored in params."""
params = Param(pse, pei, pir, pic, rho, sigma_epsilon, init_mean, population_size)
""" Create a model instance"""
model = SEIR(params)
# -
# **Q1:** Generate $10$ different trajectories of length $200$ from the model an plot them in one figure. Does the trajectories look reasonable? Could the data have been generated using this model?
#
# For reproducability, we set the seed of the random number generator to 0 before simulating the trajectories using np.random.seed(0)
#
# Save these $10$ generated trajectories for future use.
#
# *(hint: The SEIR class has a simulate method)*
np.random.seed(0)
help(model.simulate)
N=10
model_sim = model.simulate(T=200, N=N)
alpha_sim = model_sim[0]
y_sim = model_sim[1]
# Plot trajectories of simulated observations (ICU cases)
for i in range(N):
plt.plot(y_sim[0, i, :])
plt.xlabel('Time steps')
plt.ylabel('Simulated ICU cases')
# The simulated trajectories are reasonably similar to the distribution of observed data. However the range of $y_t$ is quite high in the simulated samples. But that can be tuned by adjusting the parameters of the data generating model. In conclusion the data could have been generated by this model or a similar one with different hyperparameters.
# +
# Plot the trajectories of simulated states - susceptible, exposed, infected cases and transmission rates
fig, ax = plt.subplots(nrows=2, ncols=2, sharex='all', figsize=(15,10))
fig.suptitle('State trajectories')
# Recovered state
recovered = population_size - np.sum(alpha_sim[:model.d-1, :, :], axis=0)
for i in range(N):
ax[0, 0].plot(alpha_sim[0, i, :])
ax[0, 0].set(xlabel='Time steps', ylabel='Simulated susceptible cases', title='Susceptible state')
ax[0, 1].plot(alpha_sim[1, i, :])
ax[0, 1].set(xlabel='Time steps', ylabel='Simulated exposed cases', title='Exposed state')
ax[1, 0].plot(alpha_sim[2, i, :])
ax[1, 0].set(xlabel='Time steps', ylabel='Simulated infected cases', title='Infected state')
ax[1, 1].plot(recovered[i, :])
ax[1, 1].set(xlabel='Time steps', ylabel='Simulated recovered cases', title='Recovered state')
#ax[2, 0].plot(alpha_sim[3, i, :])
#ax[2, 0].set(xlabel='Time steps', ylabel='Simulated transmission rates', title='Transmission rate')
#ax[2, 1].axis('off')
# -
# ## 3.3 Sequential Importance Sampling
#
# Next, we pick out one trajectory that we will use for filtering. We use simulated data to start with, since we then know the true underlying SEIR states and can compare the filter results with the ground truth.
# **Q2:** Implement the **Sequential Importance Sampling** algorithm by filling in the following functions.
#
# The **exp_norm** function should return the normalized weights and the log average of the unnormalized weights.
# For numerical reasons, when calculating the weights we should "normalize" the log-weights first by removing the maximal value.
#
# Let $\bar{\omega}_t = \max(\log \omega_t^i)$ and take the exponential of $\log \tilde{\omega}_t^i = \log \omega_t^i - \bar{\omega}_t$. Normalizing $\tilde{\omega}_t^i$ will yield the normalized weights!
#
# For the log average of the unnormalized weights, care has to be taken to get the correct output,
# $\log (1/N \sum_{i=1}^{N} \tilde{\omega}_t^i) = \log( 1/N \sum_{i=1}^{N} \omega_t^i) - \bar{\omega}_t$.
# We are going to need this in the future, so best to implement it right away.
#
# *(hint: look at the SEIR model class, it contains all necessary functions for propagation and weighting)*
# +
from tssltools_lab3 import smc_res
def exp_norm(logwgt):
"""
Exponentiates and normalizes the log-weights.
Parameters
----------
logwgt : ndarray
Array of size (N,) with log-weights.
Returns
-------
wgt : ndarray
Array of size (N,) with normalized weights, wgt[i] = exp(logwgt[i])/sum(exp(logwgt)),
but computed in a /numerically robust way/!
logZ : float
log of the normalizing constant, logZ = log(sum(exp(logwgt))),
but computed in a /numerically robust way/!
"""
N = len(logwgt)
C_t = max(logwgt)
logwt_tilde = logwgt - C_t
wgt = np.exp(logwt_tilde) / np.sum(np.exp(logwt_tilde))
logZ = np.log(np.sum(np.exp(logwt_tilde))) + C_t
return wgt, logZ
def ESS(wgt):
"""
Computes the effective sample size.
Parameters
----------
wgt : ndarray
Array of size (N,) with normalized importance weights.
Returns
-------
ess : float
Effective sample size.
"""
ess = np.sum(wgt)**2 / np.sum(wgt**2)
return ess
def sis_filter(model, y, N):
d = model.d
n = len(y)
# Allocate memory
particles = np.zeros((d, N, n), dtype = float) # All generated particles
logW = np.zeros((1, N, n)) # Unnormalized log-weight
W = np.zeros((1, N, n)) # Normalized weight
alpha_filt = np.zeros((d, 1, n)) # Store filter mean
N_eff = np.zeros(n) # Efficient number of particles
logZ = 0. # Log-likelihood estimate
# Filter loop
for t in range(n):
# Sample from "bootstrap proposal"
if t == 0:
particles[:, :, 0] = model.sample_state(alpha0=None, N=N) # Initialize from p(alpha_1)
logW[0, :, 0] = model.log_lik(y=y[t], alpha=particles[:, :, 0]) # Compute weights
else:
particles[:, :, t] = model.sample_state(alpha0=particles[:, :, t-1], N=N) # Propagate according to dynamics
logW[0, :, t] = model.log_lik(y=y[t], alpha=particles[:, :, t]) + logW[0, :, t-1] # Update weights
# Normalize the importance weights and compute N_eff
W[0, :, t], logZ = exp_norm(logW[0, :, t])
N_eff[t] = ESS(W[0, :, t])
# Compute filter estimates
alpha_filt[:, 0, t] = np.sum(W[0, :, t]*particles[:, :, t], axis=1)
loglik = logZ - np.log(N) # Log-likelihood
return smc_res(alpha_filt, particles, W, logW=logW, N_eff=N_eff)
# -
# **Q3:** Choose one of the simulated trajectories and run the SIS algorithm using $N = 100$ particles. Show plots comparing the filter means from the SIS algorithm with the underlying truth of the Infected, Exposed and Recovered.
#
# Also show a plot of how the ESS behaves over the run.
#
# *(hint: In the model we use the S, E, I as states, but S will be much larger than the others. To calculate R, note that S + E + I + R = Population)*
# Choose a simulated trajectory and run SIS using 100 particles
selected_trajectory = 4
sis = sis_filter(model, y=y_sim[0, selected_trajectory, :], N=100)
# +
# Plot the trajectories of filter means from SIS algorithm and the true (simulated) states
fig, ax = plt.subplots(nrows=2, ncols=2, sharex='all', figsize=(15,10))
fig.suptitle('State trajectories')
# Recovered state
recovered = population_size - np.sum(alpha_sim[:model.d-1, selected_trajectory, :], axis=0)
recovered_filter = population_size - np.sum(sis.alpha_filt[:model.d-1, 0, :], axis=0)
ax[0, 0].plot(alpha_sim[0, selected_trajectory, :], label='True state')
ax[0, 0].plot(sis.alpha_filt[0, 0, :], label='Filter mean')
ax[0, 0].set(xlabel='Time steps', ylabel='Susceptible cases', title='Susceptible state')
ax[0, 0].legend()
ax[0, 1].plot(alpha_sim[1, selected_trajectory, :], label='True state')
ax[0, 1].plot(sis.alpha_filt[1, 0, :], label='Filter mean')
ax[0, 1].set(xlabel='Time steps', ylabel='Exposed cases', title='Exposed state')
ax[0, 1].legend()
ax[1, 0].plot(alpha_sim[2, selected_trajectory, :], label='True state')
ax[1, 0].plot(sis.alpha_filt[2, 0, :], label='Filter mean')
ax[1, 0].set(xlabel='Time steps', ylabel='Infected cases', title='Infected state')
ax[1, 0].legend()
ax[1, 1].plot(recovered, label='True state')
ax[1, 1].plot(recovered_filter, label='Filter mean')
ax[1, 1].set(xlabel='Time steps', ylabel='Recovered cases', title='Recovered state')
ax[1, 1].legend()
# -
# Plot of effective sample size
plt.plot(sis.N_eff)
plt.title('Effective sample size')
plt.xlabel('Time steps')
plt.ylabel('Effective sample size')
# We observe from the plot of state trajectories that the estimated filtered distributions of the states using the SIS algorithm are good for a few initial time steps but becomes quite erratic as the time steps progresses into the future. This can be explained as soon as we look at the effective sample size (ESS) plot. The ESS drops to 1 somewhere around the 30th time step i.e. from t=30, only a single sample (particle) contributes to the estimated value of the filter distribution. This is due to the weight degeneracy problem which occurs in SIS due to repeated multiplication of weights. Hence the obtained filter estimates of the states are not reliable.
# ## 3.4 Sequential Importance Sampling with Resampling
#
# Pick the same simulated trajectory as for the previous section.
# **Q4:** Implement the **Sequential Importance Sampling with Resampling** or **Bootstrap Particle Filter** by completing the code below.
def bpf(model, y, numParticles):
d = model.d
n = len(y)
N = numParticles
# Allocate memory
particles = np.zeros((d, N, n), dtype = float) # All generated particles
logW = np.zeros((1, N, n)) # Unnormalized log-weight
W = np.zeros((1, N, n)) # Normalized weight
alpha_filt = np.zeros((d, 1, n)) # Store filter mean
N_eff = np.zeros(n) # Efficient number of particles
logZ = 0. # Log-likelihood estimate
# Filter loop
for t in range(n):
# Sample from "bootstrap proposal"
if t == 0: # Initialize from prior
particles[:, :, 0] = model.sample_state(alpha0=None, N=N)
else: # Resample and propagate according to dynamics
ind = np.random.choice(N, N, replace=True, p=W[0, :, t-1])
resampled_particles = particles[:, ind, t-1]
particles[:, :, t] = model.sample_state(alpha0=resampled_particles, N=N)
# Compute weights
logW[0, :, t] = model.log_lik(y=y[t], alpha=particles[:, :, t])
W[0, :, t], logZ_now = exp_norm(logW[0, :, t])
logZ += logZ_now - np.log(N) # Update log-likelihood estimate
N_eff[t] = ESS(W[0, :, t])
# Compute filter estimates
alpha_filt[:, 0, t] = np.sum(W[0, :, t]*particles[:, :, t], axis=1)
return smc_res(alpha_filt, particles, W, N_eff = N_eff, logZ = logZ)
# **Q5:** Use the same simulated trajectory as above and run the BPF algorithm using $N = 100$ particles. Show plots comparing the filter means from the Bootstrap Particle Filter algorithm with the underlying truth of the Infected, Exposed and Recovered. Also show a plot of how the ESS behaves over the run. Compare this with the results from the SIS algorithm.
# Choose a simulated trajectory and run BPF using 100 particles
sisr = bpf(model, y=y_sim[0, selected_trajectory, :], numParticles=100)
# +
# Plot the trajectories of filter means from SISR algorithm and the true (simulated) states
fig, ax = plt.subplots(nrows=2, ncols=2, sharex='all', figsize=(15,10))
fig.suptitle('State trajectories')
# Recovered state
recovered = population_size - np.sum(alpha_sim[:model.d-1, selected_trajectory, :], axis=0)
recovered_filter = population_size - np.sum(sisr.alpha_filt[:model.d-1, 0, :], axis=0)
ax[0, 0].plot(alpha_sim[0, selected_trajectory, :], label='True state')
ax[0, 0].plot(sisr.alpha_filt[0, 0, :], label='Filter mean')
ax[0, 0].set(xlabel='Time steps', ylabel='Susceptible cases', title='Susceptible state')
ax[0, 0].legend()
ax[0, 1].plot(alpha_sim[1, selected_trajectory, :], label='True state')
ax[0, 1].plot(sisr.alpha_filt[1, 0, :], label='Filter mean')
ax[0, 1].set(xlabel='Time steps', ylabel='Exposed cases', title='Exposed state')
ax[0, 1].legend()
ax[1, 0].plot(alpha_sim[2, selected_trajectory, :], label='True state')
ax[1, 0].plot(sisr.alpha_filt[2, 0, :], label='Filter mean')
ax[1, 0].set(xlabel='Time steps', ylabel='Infected cases', title='Infected state')
ax[1, 0].legend()
ax[1, 1].plot(recovered, label='True state')
ax[1, 1].plot(recovered_filter, label='Filter mean')
ax[1, 1].set(xlabel='Time steps', ylabel='Recovered cases', title='Recovered state')
ax[1, 1].legend()
# -
# Plot of effective sample size
plt.plot(sisr.N_eff)
plt.xlabel('Time steps')
plt.ylabel('Effective sample size')
# We observe from the state plot that the bootstrap filter estimates of the states are much more accurate than the SIS particle filter estimates. And understandably, the number of samples (particles) used to estimate the filter distribution is consistently high with only a few steps where the ESS drops below 50. Hence, the bootstrap estimates are more reliable.
# ## 3.5 Estimating the data likelihood and learning a model parameter
# In this section we consider the real data and learning the model using this data. For simplicity we will only look at the problem of estimating the $\rho$ parameter and assume that others are fixed.
#
# You are more than welcome to also study the other parameters.
# Before we begin to tweak the parameters we run the particle filter using the current parameter values to get a benchmark on the log-likelihood.
#
# **Q6:** Run the bootstrap particle filter using $N=200$ particles on the real dataset and calculate the log-likelihood. Rerun the algorithm 20 times and show a box-plot of the log-likelihood.
# +
# Run bootstrap particle filter multiple times on the real dataset
log_lik = np.zeros(20)
for i in range(20):
sisr = bpf(model, y=y_sthlm, numParticles=200)
log_lik[i] = sisr.logZ
# Box-plot of log-likelihood estimates for actual data in different runs of SISR
plt.boxplot(log_lik)
plt.title('Log-likelihood boxplot')
plt.xlabel('rho='+str(rho))
plt.ylabel('log-likelihood')
# -
# We can see that we obtain a range of log-likelihood values for different runs of the bootstrap particle filter on the same data (the real data). This variance is due to the stochastic nature of the SISR algorithm. There is randomness involved in the resampling and the propagation steps which adds variance to the log-likelihood estimates as well as the filter estimates. The median log-likelihood estimates calculated under the current parameter values is around -415 which can be used as a benchmark in parameter estimation.
# **Q7:** Make a grid of the $\rho$ parameter in the interval $[0.1, 0.9]$. Use the bootstrap particle filter to calculate the log-likelihood for each value. Run the bootstrap particle filter using $N=1000$ multiple times (20) per value and use the average as your estimate of the log-likelihood. Plot the log-likelihood function and mark the maximal value.
#
# *(hint: use np.logspace to create a grid of parameter values)*
# +
# Grid of parameter rho
rho_grid = np.linspace(start=0.1, stop=0.9, num=100)
log_lik = np.zeros((len(rho_grid), 20))
for i in range(len(rho_grid)):
# Set the rho parameter value in the model
model.set_param(rho=rho_grid[i])
for j in range(20):
# Run the bootsrtap particle filter with 1000 particles for each value of rho
sisr = bpf(model, y=y_sthlm, numParticles=1000)
log_lik[i, j] = sisr.logZ
# Average of the partile filter estimates of the log-likelihood
loglike_estimate = np.mean(log_lik, axis=1)
# +
# Maximum log-likelihood and optimal rho
max_llik = max(loglike_estimate)
max_rho = rho_grid[loglike_estimate == max_llik]
# Average log-likelihood estimates for different rho
plt.plot(rho_grid, loglike_estimate)
plt.plot(max_rho, max_llik, marker='o', color='red')
plt.axvline(x=max_rho, color='gray', ls='--')
plt.title('Maximum log-likelihood')
plt.xlabel('rho')
plt.ylabel('log-likelihood')
# -
# We get a much better log-likelihood value for $\rho$=0.72 compared to the benchmark. We can choose the optimal value for parameter $\rho$ to be approximately 0.72 where rho is the probability of exposure for each encounter with an infectious
# individual. (i.e. high risk of exposure from encounters)
# **Q8:** Run the bootstrap particle filter on the full dataset with the optimal $\rho$ value. Present a plot of the estimated Infected, Exposed and Recovered states.
# +
# Set the optimal rho parameter value in the SEIR model
rho_opt = max_rho
model.set_param(rho=rho_opt)
# Run the boostrap particle filter on full dataset with optimal rho parameter value
sisr = bpf(model, y=y_sthlm, numParticles=200)
# +
# Plot the trajectories of filter means from SISR algorithm and the true (simulated) states
fig, ax = plt.subplots(nrows=2, ncols=2, sharex='all', figsize=(15,10))
fig.suptitle('Estimated state trajectories')
# Recovered state
recovered = population_size - np.sum(sisr.alpha_filt[:model.d-1, 0, :], axis=0)
ax[0, 0].plot(sisr.alpha_filt[0, 0, :])
ax[0, 0].set(xlabel='Time steps', ylabel='Susceptible cases', title='Susceptible state')
ax[0, 1].plot(sisr.alpha_filt[1, 0, :])
ax[0, 1].set(xlabel='Time steps', ylabel='Exposed cases', title='Exposed state')
ax[1, 0].plot(sisr.alpha_filt[2, 0, :])
ax[1, 0].set(xlabel='Time steps', ylabel='Infected cases', title='Infected state')
ax[1, 1].plot(recovered)
ax[1, 1].set(xlabel='Time steps', ylabel='Recovered cases', title='Recovered state')
| Particle Filters/Code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
# 출력 옵션 변경
pd.set_option('display.max_columns', 100)
pd.set_option('display.max_rows', 1300)
# -
dlvr_raw = pd.read_csv('../data/dlvr_call.csv');
dlvr = dlvr_raw.copy()
dlvr.shape
dlvr.info()
# +
# 하나의 구 추출
dlvr = dlvr[dlvr['DLVR_STORE_SIGNGU_NM'] == '구로구']
# -
dlvr.info()
# +
# 배달일시, 지역, 업종, 매출 컬럼 선택
# 배달일시 ['PROCESS_DT'], 행정동코드 ['DLVR_DSTN_ADSTRD_CODE'], 배달상점 업종이름 (카테고리) ['DLVR_STORE_INDUTY_NM'], 배달상품금액 ['GOODS_AMOUNT']
df_dlvr = dlvr[['PROCESS_DT','DLVR_STORE_SIGNGU_NM','DLVR_STORE_INDUTY_NM','GOODS_AMOUNT']]
# +
# 업종 컬럼 값 확인
df_dlvr['DLVR_STORE_INDUTY_NM'].unique()
# +
# 업종이'배달전문업체','심부름'인 행 삭제
df_dlvr.replace(['배달전문업체','심부름'], np.nan, inplace=True)
df_dlvr = df_dlvr.dropna(axis=0)
# -
# 처리일시를 날짜 데이터 형태로 변경
df_dlvr['PROCESS_DT'] = pd.to_datetime(df_dlvr['PROCESS_DT'])
df_dlvr = df_dlvr.sort_values(by='PROCESS_DT', ascending=True)
# +
date = df_dlvr['PROCESS_DT'].unique()
df = pd.DataFrame()
df_week = pd.DataFrame(columns=['업종'])
cnt = 0
for d in date:
cnt += 1
df_date = df_dlvr[df_dlvr['PROCESS_DT'] == d]
induty = df_date['DLVR_STORE_INDUTY_NM'].unique()
for i in induty:
df_induty = df_date[df_date['DLVR_STORE_INDUTY_NM'] == i]
indSum = sum(df_induty['GOODS_AMOUNT'])
dic = { '업종' : i , '매출' : indSum}
df_week = df_week.append(dic,ignore_index=True)
if cnt % 7 == 0:
df_week_sum = df_week.groupby('업종', sort=False).sum()
df_week_sum['week'] = round(cnt/7)
df = pd.concat([df,df_week_sum],axis=0)
df_week = pd.DataFrame(columns=['업종'])
else:
pass
# -
df
df1 = df.copy()
df1
# +
# 정수형으로 변경(데이터 표기 통일)
df1.replace(np.nan, 0, inplace=True)
df1 = df1.astype('int64')
df1.info()
# -
df1.to_csv('../data/weeklySales.csv')
| project02/myanalysis/EDA4-weekly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from tools import *
from models import *
import plotly.graph_objects as go
import plotly.figure_factory as ff
from Bio.SeqUtils import GC
from Bio import SeqIO
import os
from random import sample
from plotly.subplots import make_subplots
import pickle
from scipy import stats
from collections import Counter
plt.ioff()
import warnings
warnings.filterwarnings('ignore')
# -
#for reproducibility
seed = 42
np.random.seed(seed)
torch.manual_seed(seed)
TFs = []
with open("../data/Analyzed_TFs.txt", "r") as f:
for line in f:
TFs.append(line.strip())
# +
#RECORDING THE PERFORMANCE
results = {}
new_model = {}
new_model_TL = {}
new_model_no_TL = {}
new_model_TL_freezed = {}
for i in range(1,11):
pkl_file = open("../RESULTS_50_SORTED/iterat_TL_"+str(i)+"/mccoef_old.pkl", 'rb')
mccoef_new_model = pickle.load(pkl_file)
pkl_file.close()
pkl_file = open("../RESULTS_50_SORTED/iterat_TL_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_new_model_TL = pickle.load(pkl_file)
pkl_file.close()
pkl_file = open("../RESULTS_50_SORTED/iterat_noTL_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_new_model_no_TL = pickle.load(pkl_file)
pkl_file.close()
pkl_file = open("../RESULTS_50_SORTED_BN_FR_LAYERS/iterat_TL_"+str(i)+"/mccoef.pkl",
'rb')
mccoef_new_model_TL_freezed = pickle.load(pkl_file)
pkl_file.close()
for TF in TFs:
##################################################################
if TF not in new_model.keys() and TF in mccoef_new_model.keys():
new_model[TF] = []
new_model[TF].append(mccoef_new_model[TF])
elif TF in mccoef_new_model.keys():
new_model[TF].append(mccoef_new_model[TF])
##################################################################
##################################################################
if TF not in new_model_TL.keys() and TF in mccoef_new_model_TL.keys():
new_model_TL[TF] = []
new_model_TL[TF].append(mccoef_new_model_TL[TF])
elif TF in mccoef_new_model_TL.keys():
new_model_TL[TF].append(mccoef_new_model_TL[TF])
##################################################################
##################################################################
if TF not in new_model_no_TL.keys() and TF in mccoef_new_model_no_TL.keys():
new_model_no_TL[TF] = []
new_model_no_TL[TF].append(mccoef_new_model_no_TL[TF])
elif TF in mccoef_new_model_no_TL.keys():
new_model_no_TL[TF].append(mccoef_new_model_no_TL[TF])
##################################################################
##################################################################
if TF not in new_model_TL_freezed.keys() and TF in mccoef_new_model_TL_freezed.keys():
new_model_TL_freezed[TF] = []
new_model_TL_freezed[TF].append(mccoef_new_model_TL_freezed[TF])
elif TF in new_model_TL_freezed.keys():
new_model_TL_freezed[TF].append(mccoef_new_model_TL_freezed[TF])
##################################################################
# -
new_model = pd.Series(new_model) #49 - no ARNT, because it didn't have enough data
new_model_TL = pd.Series(new_model_TL) #148
new_model_no_TL = pd.Series(new_model_no_TL) #148
new_model_TL_freezed = pd.Series(new_model_TL_freezed)
# +
new_model_TL_mean = new_model_TL.apply(lambda x: np.mean(x))
new_model_no_TL_mean = new_model_no_TL.apply(lambda x: np.mean(x))
new_model_TL_freezed_mean = new_model_TL_freezed.apply(lambda x: np.mean(x))
TL_diff_multi = new_model_TL_mean[new_model.index].subtract(new_model_no_TL_mean[new_model.index])
TL_fr_diff_multi = new_model_TL_freezed_mean[new_model.index].subtract(new_model_no_TL_mean[new_model.index])
# +
TL_diff_not_multi = new_model_TL_mean[~np.isin(new_model_TL_mean.index,
new_model.index)].subtract(new_model_no_TL_mean[~np.isin(new_model_no_TL_mean.index,
new_model.index)])
TL_fr_diff_not_multi = new_model_TL_freezed_mean[~np.isin(new_model_TL_freezed_mean.index,
new_model.index)].subtract(new_model_no_TL_mean[~np.isin(new_model_no_TL_mean.index,
new_model.index)])
# -
labels = [["in_multi"]*49, ["not_in_multi"]*99]
labels = [item for sublist in labels for item in sublist]
# +
fig = go.Figure()
fig.add_trace(go.Box(
y=list(TL_diff_multi.values)+list(TL_diff_not_multi.values),
x=labels,
name='Original_TL_vs_noTL',
marker_color='#3D9970'
))
fig.add_trace(go.Box(
y=list(TL_fr_diff_multi.values)+list(TL_fr_diff_not_multi.values),
x=labels,
name='Freezed_TL_vs_noTL',
marker_color='#FF4136'
))
#fig['layout'].update(shapes=[{'type': 'line','y0':0,
# 'y1': 0,'x0':"in_multi",
# 'x1':"not_in_multi",'xref':'x1','yref':'y1',
# 'line': {'color': 'black','width': 2.5}}])
fig.update_layout(title='TL minus noTL',
yaxis_title='Change in performance',
plot_bgcolor='rgba(0,0,0,0)', paper_bgcolor='rgba(0,0,0,0)',
boxmode='group')
fig.update_xaxes(showline=True, linewidth=2, linecolor='black')
fig.update_yaxes(showline=True, linewidth=2, linecolor='black')
fig.show()
# -
stats.ttest_ind(TL_diff_multi.values, TL_fr_diff_multi.values, equal_var = False)
stats.ttest_ind(TL_diff_not_multi.values, TL_fr_diff_not_multi.values, equal_var = False)
tfs_labels = [[tf]*10 for tf in new_model_TL.index]
tfs_labels = [item for sublist in tfs_labels for item in sublist]
# +
new_model_TL_tfs = []
for tf in new_model_TL.index:
new_model_TL_tfs = new_model_TL_tfs + new_model_TL[tf]
new_model_no_TL_tfs = []
for tf in new_model_TL.index:
new_model_no_TL_tfs = new_model_no_TL_tfs + new_model_no_TL[tf]
new_model_TL_fr_tfs = []
for tf in new_model_TL.index:
new_model_TL_fr_tfs = new_model_TL_fr_tfs + new_model_TL_freezed[tf]
# +
fig = go.Figure()
fig.add_trace(go.Box(
y=new_model_TL_tfs[120*10:],
x=tfs_labels[120*10:],
name='Original_TL',
marker_color='red',
showlegend=True
))
fig.add_trace(go.Box(
y=new_model_TL_fr_tfs[120*10:],
x=tfs_labels[120*10:],
name='Freezed_TL',
marker_color='goldenrod',
showlegend=True
))
fig.add_trace(go.Box(
y=new_model_no_TL_tfs[120*10:],
x=tfs_labels[120*10:],
name='No_TL',
marker_color='green',
showlegend=True
))
layout = go.Layout(
title = "",
xaxis = dict(
title = '',
titlefont = dict(
family = 'Courier New, monospace',
size = 18,
color = 'black'
)
),
yaxis = dict(
title = 'Mcor value',
titlefont = dict(
family = 'Courier New, monospace',
size = 18,
color = 'black'
)
)
)
#fig.update_yaxes(range=[0, 1], title= 'Mcor value', secondary_y=False)
fig.update_yaxes(range=[0, 1])
fig.update_layout(title='',
plot_bgcolor='rgba(0,0,0,0)', paper_bgcolor='rgba(0,0,0,0)',
font=dict(
family="Courier New, monospace",
size=14,
color="black"
), boxmode='group')
fig.update_layout(layout)
fig.update_layout(legend=dict(x=1.1, y=1))
fig.update_layout(autosize=False,width=1000,height=500)
fig.update_xaxes(showline=True, linewidth=2, linecolor='black')
fig.update_yaxes(showline=True, linewidth=2, linecolor='black')
fig.show()
| notebooks/TL_with_freezed_layers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Spam Classification Model (Sklearn)
# - Wrap a ML model for use as a prediction microservice in seldon-core
# - Run locally on Docker to test
# - Deploy on seldon-core running on k8s cluster
# ### Train Locally
import numpy as np
import pandas as pd
from sklearn.externals import joblib
from pathlib import Path
import string
from nltk.stem import SnowballStemmer
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import train_test_split
import pickle
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score
model_path: Path=Path('./')
# +
data = pd.read_csv("spam.csv",encoding='latin-1')
data = data.drop(["Unnamed: 2", "Unnamed: 3", "Unnamed: 4"], axis=1)
data = data.rename(columns={"v1":"class", "v2":"text"})
data.head()
def pre_process(text):
text = text.translate(str.maketrans('', '', string.punctuation))
text = [word for word in text.split() if word.lower() not in stopwords.words('english')]
words = ""
for i in text:
stemmer = SnowballStemmer("english")
words += (stemmer.stem(i))+" "
return words
features = data['text'].copy()
features = features.apply(pre_process)
vectorizer = TfidfVectorizer("english")
_features = vectorizer.fit_transform(features)
with open('skl-spam-classifier/model/vectorizer.pkl', 'wb') as vect:
pickle.dump(vectorizer, vect)
vectorizer = joblib.load(model_path.joinpath('skl-spam-classifier/model/vectorizer.pkl'))
train_x, test_x, train_y, test_y = train_test_split(_features, data['class'], test_size=0.3, random_state=0)
svc = SVC(kernel='sigmoid', gamma=1.0, probability=True)
svc.fit(train_x,train_y)
# save the model to disk
filename = 'skl-spam-classifier/model/model.pkl'
pickle.dump(svc, open(filename, 'wb'))
clf = joblib.load(model_path.joinpath(filename))
prediction = clf.predict(test_x)
accuracy_score(test_y,prediction)
# -
message = np.array(['click here to win the price'])
data = vectorizer.transform(message).todense()
probas = clf.predict_proba(data)
probas
clf.classes_
# ## Spam Classification Model (keras)
#
# - Wrap a ML model for use as a prediction microservice in seldon-core
# - Run locally on Docker to test
# - Deploy on seldon-core running on k8s cluster
# you can find data here: https://www.kaggle.com/benvozza/spam-classification/data
# ### Train Locally
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from keras.models import Model
from keras.layers import LSTM, Activation, Dense, Dropout, Input, Embedding
from keras.optimizers import RMSprop
from keras.preprocessing.text import Tokenizer
from keras.preprocessing import sequence
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
import pickle
from sklearn.externals import joblib
import tensorflow.compat.v1 as tf
tf.disable_v2_behavior()
from keras.engine.saving import model_from_json
from keras.layers import (
Bidirectional,
concatenate,
Dense,
Embedding,
LSTM,
Masking,
Reshape,
SpatialDropout1D,
TimeDistributed,
)
# +
data = pd.read_csv("spam.csv",encoding='latin-1')
data = data.drop(["Unnamed: 2", "Unnamed: 3", "Unnamed: 4"], axis=1)
data = data.rename(columns={"v1":"class", "v2":"text"})
X = data.text
Y = data['class']
le = LabelEncoder()
Y = le.fit_transform(Y)
Y = Y.reshape(-1,1)
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size=0.15)
max_words = 1000
max_len = 150
tokenizer = Tokenizer(num_words=max_words)
tokenizer.fit_on_texts(X_train)
with open('keras-spam-classifier/model/tokenizer.pkl', 'wb') as tok:
pickle.dump(tokenizer, tok)
sequences = tokenizer.texts_to_sequences(X_train)
sequences_matrix = sequence.pad_sequences(sequences,maxlen=max_len)
# +
def RNN():
inputs = Input(name='inputs',shape=[max_len])
layer = Embedding(max_words,50,input_length=max_len)(inputs)
layer = LSTM(64)(layer)
layer = Dense(256,name='FC1')(layer)
layer = Activation('relu')(layer)
layer = Dropout(0.5)(layer)
layer = Dense(1,name='out_layer')(layer)
layer = Activation('sigmoid')(layer)
model = Model(inputs=inputs,outputs=layer)
return model
model = RNN()
model.summary()
model.compile(loss='binary_crossentropy',optimizer=RMSprop(),metrics=['accuracy'])
model.fit(sequences_matrix,Y_train,batch_size=128,epochs=10,
validation_split=0.2,callbacks=[EarlyStopping(monitor='val_loss',min_delta=0.0001)])
#save model
model_json = model.to_json()
with open("keras-spam-classifier/model/architecture.json", "w") as json_file:
json_file.write(model_json)
model.save_weights("keras-spam-classifier/model/weights.h5")
# -
# ### wrap each model component using s2i
# !s2i build keras-spam-classifier/ seldonio/seldon-core-s2i-python3:1.2.2-dev spam-classifier:1.0.0.1
# !docker run --name "spam-classifier" -d --rm -p 5000:5000 spam-classifier:1.0.0.1
# !curl -g http://localhost:5000/predict --data-urlencode 'json={"data": {"names": ["message"], "ndarray": ["click here to win the price"]}}'
# !docker rm spam-classifier --force
# !s2i build keras-spam-classifier/ seldonio/seldon-core-s2i-python3:1.2.2-dev keras-spam-classifier:1.0.0.1
# !docker run --name "keras-spam-classifier" --rm -d -p 5000:5000 keras-spam-classifier:1.0.0.1
# !s2i build Translator/ seldonio/seldon-core-s2i-python3:1.2.2-dev translator:1.0.0.1
# !docker run --name "translator" -d --rm -p 5000:5000 translator:1.0.0.1
# !curl -g http://localhost:5000/transform-input --data-urlencode 'json={"data": {"names": ["message"], "ndarray": ["Wie läuft dein Tag"]}}'
# !docker rm translator --force
# !s2i build Combiner/ seldonio/seldon-core-s2i-python3:1.2.2-dev combiner:1.0.0.1
# !docker run --name "model-combiner" -d --rm -p 5000:5000 combiner:1.0.0.1
# +
# #!curl -g http://localhost:5000/aggregate --data-urlencode 'json={"data": {"names": ["message"], "ndarray": [["0.7","Spam"], ["0.80", "Spam"]]}}'
# -
# !docker rm model-combiner --force
# #### Assuming you have kubernetes cluster running and seldon-core installed, you can deploy your Machine Learning model using:
# kubectl apply -f deploy.yaml
| examples/combiners/spam_clf_combiner/spam-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### ProteinFDR 0.01 PSM FDR 0.01
# %load_ext rpy2.ipython
# +
# preprocess out_msstats.csv
import pandas as pd
def remove_decoy(Pro):
if "CONTAMINANT" in Pro or "DECOY_" in Pro:
t = []
for p in Pro.split(";"):
if "CONTAMINANT" in p or "DECOY_" in p:
pass
else:
t.append(p)
if len(t) == 0:
return None
return ";".join(t)
else:
return Pro
data = pd.read_csv("../datasets/PXD020248/out_msstats.csv", sep=",")
data['ProteinName'] = data.apply(lambda x: remove_decoy(x.ProteinName), axis=1)
data.dropna(inplace=True)
data.to_csv("../datasets/PXD020248/out_msstats_pre.csv", index=False, sep=",")
# + magic_args="-w 1600 -h 600" language="R"
# data_folder = '../datasets/PXD020248/analysis'
# setwd(data_folder)
# + magic_args="-w 1600 -h 600" language="R"
#
# library('MSstatsTMT', warn.conflicts = F, quietly = T, verbose = F)
# openms_output = './out_msstats_pre.csv'
# raw.om <- read.csv(openms_output, header = TRUE, sep = ',')
# input.om <- OpenMStoMSstatsTMTFormat(raw.om)
# head(input.om)
#
# -
# ### Protein summarization, normalization and visualization
# + magic_args="-w 1600 -h 600" language="R"
#
# protein_expression = './protein_expression.csv'
# # use MSstatsTMT for protein summarization
# quant_om.msstats <- proteinSummarization(input.om,
# method="msstats",
# global_norm=TRUE,
# reference_norm=FALSE,
# remove_norm_channel = FALSE,
# remove_empty_channel = TRUE)
#
# ## Profile plot without norm channnels and empty channels
# dataProcessPlotsTMT(data=quant_om.msstats,
# type = 'ProfilePlot',
# which.Protein = 'sp|Q9P0V3|SH3B4_HUMAN',
# width = 25, # adjust the figure width
# height = 12,
# address=FALSE)
#
# TMTProtein.Expression <- quant_om.msstats$ProteinLevelData[, c('Protein', 'Abundance', 'BioReplicate', 'Condition')]
# write.csv(TMTProtein.Expression, file=protein_expression, row.names=FALSE)
# + language="R"
#
# head(quant_om.msstats$ProteinLevelData)
# + magic_args="-w 1000 -h 600" language="R"
#
# tmp <- levels(input.om$ProteinName)
# selector <- append('allonly', tmp, 1)
# ## Quality control plot
# dataProcessPlotsTMT(data=quant_om.msstats,
# type='QCPlot',
# which.Protein=selector,
# width = 25, # adjust the figure width
# height = 12,
# address = FALSE)
# -
# ### Tests for significant changes in protein abundance across conditions
# + language="R"
#
# # Check the conditions in the protein level data
# levels(quant_om.msstats$ProteinLevelData$Condition)
# + magic_args="-w 1000 -h 600" language="R"
#
# TMT_Comp<-matrix(c(1, -1), nrow=1)
# row.names(TMT_Comp)<-c("BaP vs Control")
# colnames(TMT_Comp) = c("BaP","Control")
# TMT_Comp.pairwise <- groupComparisonTMT(quant_om.msstats, contrast.matrix = TMT_Comp, moderated = TRUE)
# head(TMT_Comp.pairwise$ComparisonResult)
# + language="R"
#
# TMT_Comp = TMT_Comp.pairwise$ComparisonResult
# write.table(TMT_Comp,"BaP_Control_Comp_result.csv",row.names=FALSE,col.names=TRUE,sep=",")
# -
# #### TMT pipeline quantifed more proteins than original result, and the most of proteins given in original result are in our result
# +
import numpy as np
from matplotlib import pyplot as plt
from venn import venn
# read the original result
OriginalResult = pd.read_excel("./1-s2.0-S0300483X20302912-mmc2.xlsx", sheet_name="sTable 2", header=3)
TMTPipelineResult = pd.read_csv("./out_msstats_pre.csv", sep=",", header=0)
TMTPipelineResult["ProteinEntry"] = TMTPipelineResult.apply(lambda x: x.ProteinName.split("|")[1], axis=1)
_, axs = plt.subplots(ncols=1,nrows=1, figsize=(6, 6))
axs.set_title("overlaping of the quantified proteins")
venn({"TMTPipeline": set(TMTPipelineResult["ProteinEntry"]), "OriginalResult": set(OriginalResult["Accession"])}, cmap=['r', 'g'], ax= axs)
# -
# ### Visualization BaP vs Control Condition and Comparing DEP
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
result = pd.read_csv('./BaP_Control_Comp_result.csv', sep=',')
result = result[['Protein', 'Label', 'log2FC', 'pvalue', 'adj.pvalue']].dropna()
result["-Log10(p-value)"] = -np.log10(result['pvalue'])
print(result.head())
result["ProteinEntry"] = result.apply(lambda x: x.Protein.split("|")[1], axis=1)
#up, normal, down
result['group'] = 'black'
result.loc[(result['log2FC'] >= 0.5)&(result['pvalue'] <= 0.05),'group'] = 'tab:red'
result.loc[(result['log2FC'] <= -0.5)&(result['pvalue'] <= 0.05),'group'] = 'tab:blue'
result.loc[result['pvalue'] > 0.05,'group'] = 'dimgrey'
print(result.head())
fig = plt.figure(figsize=(8, 32))
i = 1
for comp in np.unique(result['Label']):
data = result[result['Label'] == comp]
ax = fig.add_subplot(310 + i)
ax.scatter(data['log2FC'], data["-Log10(p-value)"], s=10, c=data['group'])
ax.set_ylabel('-Log10(pvalue)',fontweight='bold')
ax.set_xlabel('Log2FC',fontweight='bold')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.set_title(comp)
for _,row in data[(data['group'] != 'dimgrey')&(np.abs(data['log2FC']) > 0.7)].iterrows():
ax.annotate(row['ProteinEntry'], xy = (row['log2FC'], row['-Log10(p-value)']),
xytext = (row['log2FC']+0.1, row['-Log10(p-value)']+0.1))
i += 1
plt.subplots_adjust(wspace=0.5,hspace=0.5)
plt.show()
# -
# #### It seem that We found more differential expressed proteins !!!!
# +
# pvalue <= 0.05 and |log2FC| >= 0.5
_, axs = plt.subplots(ncols=1, nrows=1, figsize=(6, 6))
OrginalDEP = OriginalResult[OriginalResult["raw p-value <= 0.05 and |log2(FC)| >= 0.5"] == True]
TMTPipelineDEP = result[(result["group"] == "tab:red") | (result["group"] == "tab:blue")]
axs.set_title("overlaping of the differential expressed proteins ")
venn({'TMTPipelineDEP': set(TMTPipelineDEP["ProteinEntry"]), 'OrginalDEP': set(OrginalDEP['Accession'])},
cmap=['r', 'g'],figsize=(9, 10), ax=axs)
# -
# ### Measure reproducibility in biological replicate
# #### Overall, there is not much difference. The CV values obtained from the reanalyses were similar to the originally published datasets
# +
# How to Caculate CV !!!
from seaborn import violinplot, stripplot
data = pd.read_csv("./out_msstats_pre.csv", sep=",", header=0)
viodata = pd.DataFrame(columns=["Condition","CV[%]", "Class"])
BaPMedian = np.median(data[data["Condition"] == "BaP"]["Intensity"])
ControlMedian = np.median(data[data["Condition"] == "Control"]["Intensity"])
fillBaPMedian = {"BaP": BaPMedian, "Control": ControlMedian}
for condition in np.unique(data["Condition"]):
sdata = data[data["Condition"] == condition]
for name, group in sdata.groupby(["ProteinName"]):
# Missed Value ?
t = group["Intensity"]
# if len(set(group["BioReplicate"])) < 6:
# continue
# t = group["Intensity"].tolist()
# t.extend([fillBaPMedian[condition]] * (6 - len(set(group["BioReplicate"]))))
std = np.std(t)
mean = np.sum(group["Intensity"]) / 6
if mean != 0.0:
viodata = viodata.append({"Condition": condition, "CV[%]": round(std / mean, 3) * 100, "Class": "TMTPipeline"},
ignore_index = True)
evidata = pd.read_csv("./evidence.txt", sep="\t", header=0)
evimedian = evidata["Reporter intensity corrected 1"].tolist()
evimedian.extend(evidata["Reporter intensity corrected 2"].tolist())
evimedian.extend(evidata["Reporter intensity corrected 3"].tolist())
evimedian = np.median(evimedian)
for name, group in evidata.groupby("Proteins"):
# Missed Value ?
# if len(set(group["Experiment"])) < 2:
# continue
t = group["Reporter intensity corrected 1"].tolist()
t.extend(group["Reporter intensity corrected 2"].tolist())
t.extend(group["Reporter intensity corrected 3"].tolist())
std = np.std(t)
mean = np.sum(t) / 6
if mean != 0:
viodata = viodata.append({"Condition": "BaP", "CV[%]": round(std / mean, 3) * 100, "Class": "OriginalResult"}, ignore_index = True)
t = group["Reporter intensity corrected 4"].tolist()
t.extend(group["Reporter intensity corrected 5"].tolist())
t.extend(group["Reporter intensity corrected 6"].tolist())
std = np.std(t)
mean = np.sum(t) / 6
if mean != 0:
viodata = viodata.append({"Condition": "Control", "CV[%]": round(std / mean, 3) * 100, "Class": "OriginalResult"}, ignore_index = True)
plt.figure(figsize=(10,10))
violinplot(x="Condition", y="CV[%]", data=viodata, hue="Class", scale="count", cut=0, palette="Set2")
# stripplot(x="Condition", y="CV[%]", data=viodata[viodata["CV[%]"] > 150], jitter=0, color="c", hue="Class")
plt.ylim(0, 300)
plt.show()
print("CV of TMTPipeline in BaP Condition: %f" %(np.median(viodata[(viodata["Class"] == "TMTPipeline") & (viodata["Condition"] == "BaP")]["CV[%]"])))
print("CV of Original in BaP Condition: %f" %(np.median(viodata[(viodata["Class"] == "OriginalResult") & (viodata["Condition"] == "BaP")]["CV[%]"])))
print("CV of TMTPipeline in Control Condition: %f" %(np.median(viodata[(viodata["Class"] == "TMTPipeline") & (viodata["Condition"] == "Control")]["CV[%]"])))
print("CV of Original in Control Condition: %f" %(np.median(viodata[(viodata["Class"] == "OriginalResult") & (viodata["Condition"] == "Control")]["CV[%]"])))
# -
# ### HeatMap
# +
# Libraries
import seaborn as sns
import pandas as pd
from matplotlib import pyplot as plt
import numpy as np
# Data set
url = './protein_expression.csv'
df = pd.read_csv(url)
df2 = df.pivot_table(index='Protein', columns='Condition', values='Abundance',aggfunc=np.mean)
df2 = df2.dropna()
# plot
g = sns.clustermap(df2, method='average', metric='correlation', z_score=0, figsize=(15,15), xticklabels=True, cmap="coolwarm")
g.ax_heatmap.set_xticklabels(g.ax_heatmap.get_xmajorticklabels())#, fontsize = 5)
# tmp = g.ax_heatmap.get_xaxis()
threshold = 0.1
x_labels_ticks = g.ax_heatmap.get_xticklabels()
total_genes_above_threshold = 0
for i, xtickdata in enumerate(x_labels_ticks):
protein = xtickdata._text
if df2[protein].max() >= threshold:
# print(df2[protein])
# print("#########")
total_genes_above_threshold = total_genes_above_threshold + 1
# print(df2[protein].max())
else:
xtickdata._text = ''
#print("total_genes_above_threshold {}".format(total_genes_above_threshold))
# re set the tick labels with the modified list
g.ax_heatmap.set_xticklabels(x_labels_ticks)
plt.show()
# -
| notebook/PXD020248DownStreamAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
PATH = "../data/raw"
df = pd.read_csv(f"{PATH}/heart.csv")
# # EDA
# ## Common characteristics
print(f"The table contains {df.shape[0]} entries grouped into {df.shape[1]} columns")
# There are following columns in the given dataset:\n{
print(list(df.columns))
# For further use let's rename the columns according to the description given at the "Kaggle" source
columns = [
"age",
"sex",
"chest_pain_type",
"resting_blood_pressure",
"serum_cholestoral",
"fasting_blood_sugar",
"resting_electrocardiographic_results",
"maximum_heart_rate",
"induced_angina",
"induced_st_depression",
"st_peak_slope",
"number_of_major_vessels",
"thal",
"target"
]
df.columns = columns
# The entries in our dataframe look like
df.sample(1).T
# ## Distribution and correlation
# Let's gather some information about the dataset values distribution
df.info()
df.nunique()
df.describe().T
# Now we separate numerical and categircal features. As some threshold that makes us think of a feature as of a certain type let's take the value of 10.
threshold = 5
cat_features = df.columns[df.nunique() <= threshold][:-1].tolist()
num_features = list(df.columns[df.nunique() > threshold])
cat_features, num_features
df.corr()
g = sns.pairplot(df[num_features + ["target"]],
hue="target",
hue_order=df.target,
palette="husl")
g.savefig("pairplot.png")
| ml_project/notebooks/EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (textwiser)
# language: python
# name: textwiser
# ---
# # Document Embedding Basic Usage Example
#
# Here, we provide a basic look into how the API can be used.
import os
os.chdir('..')
# We use a sample scikit-learn dataset to test the embeddings.
# +
from sklearn.datasets import fetch_20newsgroups
categories = ['alt.atheism', 'talk.religion.misc', 'comp.graphics', 'sci.space']
newsgroups_train = fetch_20newsgroups(subset='train', categories=categories)
documents = newsgroups_train.data[:20]
# -
# In general, you can use `Embedding` models that take in raw text to convert them into real valued vectors. You can then follow them up with `Transformation` models that take in and output real valued vectors.
#
# For example, we can replicate a TfIdf to NMF featurization in the following manner:
# +
from textwiser import TextWiser, Embedding, Transformation
emb = TextWiser(Embedding.TfIdf(min_df=5), Transformation.NMF(n_components=10))
vecs = emb.fit_transform(documents)
vecs
# -
# We can just as easily try out a pooling of word embeddings:
# +
from textwiser import TextWiser, Embedding, PoolOptions, Transformation, WordOptions
emb = TextWiser(Embedding.Word(word_option=WordOptions.word2vec, pretrained='en'), Transformation.Pool(pool_option=PoolOptions.max))
vecs = emb.fit_transform(documents)
vecs
# -
# ## Schema
#
# We also provide a convenient schema for mixing and matching different word and document embeddings. This makes it easy to try out different embedding types rapidly.
#
# There are two main operations: `transform` and `concat`.
#
# The `transform` operation defines a list of operations. The first of these operations should be an `Embedding`, while the rest should be `Transformation`s. The idea is that the `Embeddings` have access to raw text and turn them into vectors, and therefore the following `Transformation`s need to operate on vectors. In PyTorch terms, this is equivalent to using `nn.Sequential`.
#
# The `concat` operation defines a concatenation of multiple embedding vectors. This can be done both at word and sentence level. In PyTorch terms, this is equivalent to using `torch.cat`.
#
# ### Sample Schema
#
# Below we outline a sample and presumably common use-case. At the root level, we have three different embeddings. The first two are different word embeddings, both are pooled (first using max pooling, second using mean pooling) to generate the sentence representations $s_1$ and $s_2$. The third is a tf-idf embedding of the document followed by a reduction of its dimensionality to 30 using NMF, generating sentence representation $s_3$. These representations are concatenated to $s_{123}$, and are fed into a final SVD transformation that brings the sentence vector back down to a manageable level ($s$).
# +
from textwiser import TextWiser, Embedding
doc_embeddings_schema = {
'transform': [
{
'concat': [
{
'transform': [
('word2vec', {'pretrained': 'en'}),
'pool'
]
},
{
'transform': [
('flair', {'pretrained': 'news-forward-fast'}),
('pool', {'pool_option': 'mean'})
]
},
{
'transform': [
'tfidf',
('nmf', { 'n_components': 10 })
]
}
]
},
'svd'
]
}
doc_embeddings = TextWiser(Embedding.Compound(schema=doc_embeddings_schema))
doc_embeddings
# -
# Once the embeddings object is initialized, we can feed in a list of text documents and get the relevant output.
doc_embeddings.fit(documents)
# +
# %%time
emb = doc_embeddings.transform(documents)
emb
# -
# You can also specify the same schema in a json file.
# +
import json
doc_embeddings = TextWiser(Embedding.Compound(schema='notebooks/schema.json'))
doc_embeddings
# -
doc_embeddings.fit_transform(documents)
| notebooks/basic_usage_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="3ad7QmOZllJr" colab_type="code" outputId="5a99bba9-8de0-43a2-d342-4795078f1b09" colab={"base_uri": "https://localhost:8080/", "height": 213}
#prepare fast.ai
# !curl https://course.fast.ai/setup/colab | bash
#mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
# <KEY>
# + id="JjsFK0eIvpT_" colab_type="code" colab={}
from fastai.tabular import *
# + [markdown] id="OeOYDo6IvpUC" colab_type="text"
# # Rossmann
# + [markdown] id="lSFHAgpkvpUD" colab_type="text"
# ## Data preparation
# + [markdown] id="hucluJ-cvpUE" colab_type="text"
# To create the feature-engineered train_clean and test_clean from the Kaggle competition data, run `rossman_data_clean.ipynb`. One important step that deals with time series is this:
#
# ```python
# add_datepart(train, "Date", drop=False)
# add_datepart(test, "Date", drop=False)
# ```
# + id="AnlhyMsdvpUF" colab_type="code" colab={}
path = Path('/content/gdrive/My Drive/DATA/rossmann')
train_df = pd.read_pickle(path/'train_clean')
# + id="rRSiQ5HUvpUI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 402} outputId="7766ebba-2991-4a1a-ba58-f05446063a74"
train_df.head().T
# + id="cWN1Bc5yvpUM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="18a3827d-8077-414c-de4e-b221de2db004"
n = len(train_df); n
# + [markdown] id="NbV6pi1LvpUQ" colab_type="text"
# ### Experimenting with a sample
# + id="b-n99BfgvpUQ" colab_type="code" colab={}
idx = np.random.permutation(range(n))[:2000]
idx.sort()
small_train_df = train_df.iloc[idx[:1000]]
small_test_df = train_df.iloc[idx[1000:]]
small_cont_vars = ['CompetitionDistance', 'Mean_Humidity']
small_cat_vars = ['Store', 'DayOfWeek', 'PromoInterval']
small_train_df = small_train_df[small_cat_vars + small_cont_vars + ['Sales']]
small_test_df = small_test_df[small_cat_vars + small_cont_vars + ['Sales']]
# + id="RHaJDDtDvpUT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="d99a91e1-0f71-414c-f427-5c402e7afcc9"
small_train_df.head()
# + id="NjcUkjYZvpUW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="4a9c3b64-bb79-4b57-805f-42117cc4cb10"
small_test_df.head()
# + id="3gi2WOtNvpUZ" colab_type="code" colab={}
categorify = Categorify(small_cat_vars, small_cont_vars)
categorify(small_train_df)
categorify(small_test_df, test=True)
# + id="yYiTT3t4vpUb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="f343702d-169f-4e3f-becd-bdd772c21e51"
small_test_df.head()
# + id="TTGvuR5_vpUg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="84ae6d90-fb64-4856-de96-4a57640fdce9"
small_train_df.PromoInterval.cat.categories
# + id="2AQOqsNhvpUi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="e470965c-c11f-43a9-894c-2c7bb0f1bcb6"
small_train_df['PromoInterval'].cat.codes[:5]
# + id="N20vJmx9vpUl" colab_type="code" colab={}
fill_missing = FillMissing(small_cat_vars, small_cont_vars)
fill_missing(small_train_df)
fill_missing(small_test_df, test=True)
# + id="s805zHq4vpUo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="49500e41-6691-4951-c182-ae6739860dd9"
small_train_df[small_train_df['CompetitionDistance_na'] == True]
# + [markdown] id="yfgGIJdWvpUr" colab_type="text"
# ### Preparing full data set
# + id="vR4RUFLNvpUt" colab_type="code" colab={}
train_df = pd.read_pickle(path/'train_clean')
test_df = pd.read_pickle(path/'test_clean')
# + id="9pGDWiZBvpUw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ada9a041-2e34-43a9-bd87-095f2466befc"
len(train_df),len(test_df)
# + id="CRcSRHdXvpUy" colab_type="code" colab={}
procs=[FillMissing, Categorify, Normalize]
# + id="2iZNrapdvpU1" colab_type="code" colab={}
cat_vars = ['Store', 'DayOfWeek', 'Year', 'Month', 'Day', 'StateHoliday', 'CompetitionMonthsOpen',
'Promo2Weeks', 'StoreType', 'Assortment', 'PromoInterval', 'CompetitionOpenSinceYear', 'Promo2SinceYear',
'State', 'Week', 'Events', 'Promo_fw', 'Promo_bw', 'StateHoliday_fw', 'StateHoliday_bw',
'SchoolHoliday_fw', 'SchoolHoliday_bw']
cont_vars = ['CompetitionDistance', 'Max_TemperatureC', 'Mean_TemperatureC', 'Min_TemperatureC',
'Max_Humidity', 'Mean_Humidity', 'Min_Humidity', 'Max_Wind_SpeedKm_h',
'Mean_Wind_SpeedKm_h', 'CloudCover', 'trend', 'trend_DE',
'AfterStateHoliday', 'BeforeStateHoliday', 'Promo', 'SchoolHoliday']
# + id="hbn_1jCYvpU4" colab_type="code" colab={}
dep_var = 'Sales'
df = train_df[cat_vars + cont_vars + [dep_var,'Date']].copy()
# + id="mAe4Cjy1vpU-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3fa864cd-f1ad-40a6-bd2c-90719c5ecb0e"
test_df['Date'].min(), test_df['Date'].max()
# + id="oW6OtnKxvpVG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06aaa56c-3234-4e04-8e4b-fc149f3e7a6b"
cut = train_df['Date'][(train_df['Date'] == train_df['Date'][len(test_df)])].index.max()
cut
# + id="tjBY2EfCvpVJ" colab_type="code" colab={}
valid_idx = range(cut)
# + id="OhVFhlnivpVL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="17cdfa86-0d43-43f8-a3e2-402a1982ced2"
df[dep_var].head()
# + id="AxCwoO4dvpVO" colab_type="code" colab={}
data = (TabularList.from_df(df, path=path, cat_names=cat_vars, cont_names=cont_vars, procs=procs,)
.split_by_idx(valid_idx)
.label_from_df(cols=dep_var, label_cls=FloatList, log=True)
.add_test(TabularList.from_df(test_df, path=path, cat_names=cat_vars, cont_names=cont_vars))
.databunch())
# + id="Xkgf5FNSvpVS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="28389bd8-94fc-4f95-9e22-6be4fff526c2"
data
# + [markdown] id="Cwsq5PJjvpVU" colab_type="text"
# ## Model
# + id="OueRGmPNvpVV" colab_type="code" colab={}
max_log_y = np.log(np.max(train_df['Sales'])*1.2)
y_range = torch.tensor([0, max_log_y], device=defaults.device)
# + id="nuaJWxb3vpVX" colab_type="code" colab={}
learn = tabular_learner(data, layers=[1000,500], ps=[0.001,0.01], emb_drop=0.04,
y_range=y_range, metrics=exp_rmspe)
# + id="E-_gboAzvpVa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 738} outputId="08b542ff-aafa-4377-fd01-f7f1f17bb205"
learn.model
# + id="uc8VtFUDvpVd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2cde4f3b-5cd8-4072-d472-a1004fa2c001"
len(data.train_ds.cont_names)
# + id="prJB5zgjvpVg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="82d73f78-f57d-488d-98cf-e5b5530e708a"
learn.lr_find()
# + id="XZU32XbKvpVk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="458e277a-19c6-420a-db24-5ff261a175b4"
learn.recorder.plot()
# + id="93MS9fzuvpVm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="d61744d6-93fe-4efc-f872-2dc4780bd7d1"
learn.fit_one_cycle(5, 1e-3, wd=0.2)
# + id="8SmcTR_rvpVo" colab_type="code" colab={}
learn.save('1')
# + id="RAgWYG1xvpVq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="49fb881b-0e40-4752-d0f5-26737c35648e"
learn.recorder.plot_losses(skip_start=10000)
# + id="0jYqfV0bvpVs" colab_type="code" colab={}
learn.load('1');
# + id="wxW9LZ1TvpVu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="bc3c26ba-3e02-49ee-aa5f-2fb27222c994"
learn.fit_one_cycle(5, 3e-4)
# + id="gPb7bN32vpVz" colab_type="code" colab={} outputId="264d29c7-7492-4b21-9439-20416a84bcfc"
learn.fit_one_cycle(5, 3e-4)
# + [markdown] id="PAIwv4gIvpV3" colab_type="text"
# (10th place in the competition was 0.108)
# + id="qKmwAKwzvpV4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="3e958f3d-5ab1-44ed-9370-6d0e25214524"
test_preds=learn.get_preds(DatasetType.Test)
test_df["Sales"]=np.exp(test_preds[0].data).numpy().T[0]
test_df[["Id","Sales"]]=test_df[["Id","Sales"]].astype("int")
test_df[["Id","Sales"]].to_csv("rossmann_submission.csv",index=False)
# + id="rXst5MkH7UB6" colab_type="code" colab={}
# !cp rossmann_submission.csv /content/gdrive/My\ Drive/DATA/rossmann
# + id="vVWcMZiI7rIi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="23f745c9-0a83-403b-95c1-27e3890d1d0e"
path
# + id="F3o1iFE-7z_c" colab_type="code" colab={}
| nbs/dl1/lesson6-rossmann.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from design_nest.eplus_components.simulation_parameters import Timestep, Verion
from design_nest.eplus_components.model import Model
v = Version()
v.name = "Version 1"
v.version_identifier = 8.9
m = Model()
m.add(v)
m.__dict__
m.save("ep.json")
m.load("ep.json")
#
m
m.__dict__
vars(m)
| tests/random_tests_with_notebooks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Part II: Exploratory Data Analysis
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv('20210315_Housesigma_just_sold_final_combined.csv')
df.head()
df.info()
df['Utility_incl'].unique()
index = np.where(df['Utility_incl']=='heat')
index
df['heat'] = 0
df['water'] = 0
df['hydro'] = 0
# +
df['heat'].iloc[index] = 1
index_2 = np.where(df['Utility_incl']=='water,heat')
df['heat'].iloc[index_2] = 1
df['water'].iloc[index_2] = 1
index_3 = np.where(df['Utility_incl']=='water')
df['water'].iloc[index_3] = 1
index_4 = np.where(df['Utility_incl']=='water,hydro,heat')
df['heat'].iloc[index_4] = 1
df['water'].iloc[index_4] = 1
df['hydro'].iloc[index_4] = 1
index_5 = np.where(df['Utility_incl']=='hydro')
df['hydro'].iloc[index_5] = 1
index_6 = np.where(df['Utility_incl']=='water,hydro')
df['water'].iloc[index_4] = 1
df['hydro'].iloc[index_4] = 1
# -
df[['water','hydro','heat']]
community_names = [i.split(' - Toronto')[0] for i in df['Community'].unique()]
df.groupby('Community')['Address'].count().plot(kind='pie', startangle = 90, )
df.groupby('House_type')['Address'].count().plot(kind='bar')
index = np.where(df['House_type']=='Condo/Apt Unit')[0]
df['House_type'].iloc[index] = 'Condo Apt'
indices = np.where((df.House_type =='Freehold Townhouse') |(df.House_type =='Co-Op Apt') | (df.House_type =='Co-Ownership Apt') |(df.House_type =='Duplex') | (df.House_type =='Other'))
df.drop(index = indices[0], inplace = True)
df.groupby('Bedroom')['Address'].count().plot(kind='bar')
df.groupby('Exposure')['Address'].count().plot(kind='bar')
index = np.where((df['Exposure']=='NS' )| (df['Exposure']=='EW'))[0]
df['Exposure'].iloc[index] = np.NaN
df['Exposure'].iloc[index]
df['Municipality'].nunique()
df.drop(columns=['Listing_num', 'Address', 'Estimated_on', 'Rental_dom', 'Rental_estimate',\
'Rental_yield', 'Property_type', 'Building_age', \
'Basement','Utility_incl', 'Listed_on', 'Updated_on', \
'Major_intersection', 'Amenities', 'Municipality'], inplace = True)
df.info()
# ### Fill NA
# Fill NA with an mean value of its bedroom size
df.groupby('Bedroom')[['Size']].mean()
# Estimating Size by the number of bedroom
df['Size'] = df['Size'].fillna(df.groupby('Bedroom')['Size'].transform('mean'))
df.groupby('Size')['Community'].count()
bins = [0, 500, 600, 700, 800, 900, 1000, 1100, 1500]
df['Size_cat'] = pd.cut(df['Size'], bins=bins, right = False, include_lowest = True)
df.groupby('Size_cat')['Community'].count()
sum(df['Size_cat'].isna())
# ### Fill NA
df.info()
df.groupby('Size_cat')[['Tax', 'Maintenance']].mean()
# Estimating Tax & Maintenance by the Size
df['Tax'] = df['Tax'].fillna(df.groupby('Size_cat')['Tax'].transform('mean'))
df['Maintenance'] = df['Maintenance'].fillna(df.groupby('Size_cat')['Maintenance'].transform('mean'))
sum(df['Exposure'].isna())
df['Exposure'].value_counts()/df['Exposure'].count()
exposure_freq = pd.Series(df['Exposure'].value_counts()/df['Exposure'].count())
exposure_freq
exposure_freq.index
import random
random_choices = random.choices(exposure_freq.index, weights = exposure_freq, k = sum(df['Exposure'].isna()))
random_choices
df[df['Exposure'].isnull()].index
null_index = np.where(df['Exposure'].isnull())[0]
for i in range(len(null_index)):
df['Exposure'].iloc[null_index[i]]=random_choices[i]
sum(df['Exposure'].isna())
df.dropna(inplace = True) # drops the Sold-price na row
df.info()
| PartI_Exploratory_Data_Analysis.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
library(tidyverse)
library(gridExtra)
str(sleep)
head(sleep)
# +
# 약제 1을 복용했을 때 수면시간의 증가 (단위는 시간이다)
y <- sleep$extra[sleep$group == 1]
y
# -
summary(y)
sd(y)
par(mfrow=c(2,2))
hist(y)
boxplot(y)
qqnorm(y); qqline(y)
hist(y, prob=TRUE)
lines(density(y), lty=2)
# +
par(mfrow=c(2,2))
hist(y)
boxplot(y)
qqnorm(y); qqline(y)
hist(y, prob=TRUE)
lines(density(y), lty=2)
png("../plots/6-1.png", 5.5, 4, units='in', pointsize=9, res=600)
y <- sleep$extra[sleep$group == 1]
opar <- par(mfrow=c(2,2))
hist(y)
boxplot(y)
qqnorm(y); qqline(y)
hist(y, prob=TRUE)
lines(density(y), lty=2)
par(opar)
dev.off()
# '일변량 t-검정(one-sample t-test)'
t.test(y)
t.test(y, alternative="greater")
# 개개인의 수면시간증가값 모형
# 평균이 0이고, 표준편차가 1.8(시간)인 종 모양의 분포(bell shaped distribution)
# N(0, 1.8^2)
curve(dnorm(x, 0, 1.8), -4, 4)
png("../plots/6-2.png", 5.5, 4, units='in', pointsize=9, res=600)
curve(dnorm(x, 0, 1.8), -4, 4)
dev.off()
# 크기가 10개인 새로운 표본
options(digits = 3)
set.seed(1606)
(y_star <- rnorm(10, 0, 1.8))
mean(y_star-0); sd(y_star)
(t_star <- mean(y_star-0) / (sd(y_star)/sqrt(length(y_star))))
(y_star <- rnorm(10, 0, 1.8))
mean(y_star-0); sd(y_star)
(t_star <- mean(y_star-0) / (sd(y_star)/sqrt(length(y_star))))
(y_star <- rnorm(10, 0, 1.8))
mean(y_star-0); sd(y_star)
(t_star <- mean(y_star-0) / (sd(y_star)/sqrt(length(y_star))))
# 10,000개의 평행우주의 표본 (각 표본은 10개의 관측치를 포함한다)
# , 그리고 각 표본의 평균값, 표본표준편차, 그리고 t-통계량 값을 계산할 수 있다:
set.seed(1606)
B <- 1e4
n <- 10
xbars_star <- rep(NA, B)
sds_star <- rep(NA, B)
ts_star <- rep(NA, B)
for(b in 1:B){
y_star <- rnorm(n, 0, 1.789)
m <- mean(y_star)
s <- sd(y_star)
xbars_star[b] <- m
sds_star[b] <- s
ts_star[b] <- m / (s/sqrt(n))
}
opar <- par(mfrow=c(2,2))
hist(xbars_star, nclass=100)
abline(v = 0.75, col='red')
hist(sds_star, nclass=100)
abline(v = 1.789, col='red')
hist(ts_star, nclass=100)
abline(v = 1.3257, col='red')
qqnorm(ts_star); qqline(ts_star)
par(opar)
png("../plots/6-3.png", 5.5*.8, 4, units='in', pointsize=9, res=600)
opar <- par(mfrow=c(2,2))
hist(xbars_star, nclass=100)
abline(v = 0.75, col='red')
hist(sds_star, nclass=100)
abline(v = 1.789, col='red')
hist(ts_star, nclass=100)
abline(v = 1.3257, col='red')
qqnorm(ts_star); qqline(ts_star)
par(opar)
dev.off()
# 우리가 관측한 t-통계량 값 1.3257은 시뮬레이션 분포에서 어디에 있는가?
length(which(ts_star > 1.3257)) / B
# 스튜던트 t 분포
# 다양한 자유도 값에 따른 t 밀도함수
# https://en.wikipedia.org/wiki/Student%27s_t-distribution
# Credit: 권용찬
nlist=c(1,2,5)
x <- seq(-5, 5, 0.05)
y <- matrix(0, nr=length(x), nc=length(nlist))
plot(x, type="n", xlab="x", ylab="P(x)",
xlim=c(-5,5), ylim=c(0, 0.45))
for( i in 1:length(nlist)){
y[,i] <- dt(x, df=nlist[i])
lines(x, y[,i], col=i, lwd=2)
}
lines(x, dnorm(x), col=4, lwd=2)
legend_text <- c(expression(paste(nu,"=1 ")),
expression(paste(nu,"=2 ")),
expression(paste(nu,"=5 ")),
expression(paste(nu,"=",infinity)))
legend("topright", legend=legend_text, lty=1, lwd=2, col=c(1:3,4),
inset=.05)
png("../plots/6-4.png", 5.5, 4, units='in', pointsize=9, res=600)
nlist=c(1,2,5)
x <- seq(-5, 5, 0.05)
y <- matrix(0, nr=length(x), nc=length(nlist))
plot(x, type="n", xlab="x", ylab="P(x)",
xlim=c(-5,5), ylim=c(0, 0.45))
for( i in 1:length(nlist)){
y[,i] <- dt(x, df=nlist[i])
lines(x, y[,i], col=i, lwd=2)
}
lines(x, dnorm(x), col=4, lwd=2)
legend_text <- c(expression(paste(nu,"=1 ")),
expression(paste(nu,"=2 ")),
expression(paste(nu,"=5 ")),
expression(paste(nu,"=",infinity)))
legend("topright", legend=legend_text, lty=1, lwd=2, col=c(1:3,4), inset=.05)
dev.off()
# 8. 신뢰구간의 의미
set.seed(1606)
(y_star <- rnorm(10, 1, 1.8))
t.test(y_star)$conf.int
(y_star <- rnorm(10, 1, 1.8))
t.test(y_star)$conf.int
(y_star <- rnorm(10, 1, 1.8))
t.test(y_star)$conf.int
library(tidyverse)
set.seed(1606)
B = 1e2
conf_intervals <-
data.frame(b=rep(NA, B),
lower=rep(NA, B),
xbar=rep(NA, B),
upper=rep(NA, B))
true_mu <- 1.0
for(b in 1:B){
(y_star <- rnorm(10, true_mu, 1.8))
conf_intervals[b, ] = c(b=b,
lower=t.test(y_star)$conf.int[1],
xbar=mean(y_star),
upper=t.test(y_star)$conf.int[2])
}
conf_intervals <- conf_intervals %>%
mutate(lucky = (lower <= true_mu & true_mu <= upper))
glimpse(conf_intervals)
table(conf_intervals$lucky)
conf_intervals %>% ggplot(aes(b, xbar, col=lucky)) +
geom_point() +
geom_errorbar(aes(ymin=lower, ymax=upper)) +
geom_hline(yintercept=true_mu, col='red')
ggsave("../plots/6-6.png", width=5.5, height=4, units='in', dpi=600)
# 6.10.2. 중심극한정리
hist(c(0, 1), nclass=100, prob=TRUE, main='Individual sleep time increase')
set.seed(1606)
B <- 1e4
n <- 10
xbars_star= rep(NA, B)
for(b in 1:B){
xbars_star[b] <- mean(sample(c(0,1), size=n, replace=TRUE))
}
hist(xbars_star, nclass=100, main='Sample mean of 10 obs')
png("../plots/6-8.png", 5.5, 4*.8, units='in', pointsize=9, res=600)
opar = par(mfrow=c(1,2))
hist(c(0, 1), nclass=100, prob=TRUE, main='Individual sleep time increase')
hist(xbars_star, nclass=100, main='Sample mean of 10 obs')
par(opar)
dev.off()
# 6.11. 모수추정의 정확도는 sqrt(n)에 비례한다.
diff(t.test(y)$conf.int)
mean(y)
diff(t.test(y)$conf.int)/2
# 자료의 incremental 가치
png("../plots/6-9.png", 5.5, 4*.8, units='in', pointsize=9, res=600)
opar = par(mfrow=c(1,2))
curve(1/sqrt(x), 1, 1000, log='x', main='s.e. vs sample size')
curve((1/sqrt(x) - 1/sqrt(x+10)) / (1/sqrt(x)), 1, 1000, log='x',
main='% decrease in s.e. \nwhen adding 10 obs')
par(opar)
dev.off()
| study/statistics/statistics-concepts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Datetime and Timedelta
#
# This chapter covers two distinct concepts, datetimes and timedeltas and how they are created and used in Pandas. A datetime represents a specific **moment** in time. A timedelta represents an **amount** of time.
#
# ## Date vs Time vs Datetime
# There is a distinction that needs to be made between the terms **date**, **time**, and **datetime**. They all three mean different things.
#
# * **date** - Only the month, day, and year. So 2016-01-01 would represent January 1, 2016 and be considered a **date**.
# * **time** - Only the hours, minutes, seconds and parts of a second (milli/micro/nano). 5 hours, 45 minutes and 6.74234 seconds for example would be considered a **time**.
# * **datetime** - A combination of the above two. Has both date (Year, Month, Day) and time (Hour, Minute, Second) components. January 1, 2016 at 5:45 p.m. would be an example of a **datetime**.
#
# The Python standard library contains the [datetime module][1]. It is a popular and important module but will not be covered here since Pandas builds its own datetime and timedelta objects that are more powerful. However, there are some notes available on datetime standard library in the **extras** directory.
#
# [1]: https://docs.python.org/3.5/library/datetime.html
# ### Datetimes in numpy
#
# numpy has its own datetime data type called [datetime64][1]. It is more powerful and flexible than core Python's datetime module. We won't be working with it either.
#
# ### pandas Timestamp
#
# pandas has its own datetime data type called a `Timestamp`, which adds more functionality to NumPy's `datetime64`.
#
# [1]: https://docs.scipy.org/doc/numpy/reference/arrays.datetime.html
# ### Creating a single Timestamp with the `to_datetime` function
#
# The `to_datetime` function converts both strings and numbers to Timestamps as well as entire arrays or Series. It is intelligent and can detect a wide variety of strings. Each of the following create a single pandas Timestamp object.
import pandas as pd
import numpy as np
pd.to_datetime('2016/1/10')
pd.to_datetime('2016-1-10')
pd.to_datetime('Jan 3, 2019 20:45.56')
pd.to_datetime('January 3, 2019 20:45.56')
pd.to_datetime('2016-01-05T05:34:43.123456789')
# ### Epoch
# The term epoch refers to the origin of a particular era. Like many other programming languages, Python uses January 1, 1970 (also known as the Unix epoch) as its epoch for keeping track of datetime. In Pandas, integers are used to represent the number of nanoseconds that have elapsed since the epoch.
#
# ### Converting numbers to Timestamps
#
# You can pass numbers to the `to_datetime` function and it will convert it to a Timestamp. It assumes you are passing in the number of nanoseconds after the epoch. The following creates the datetime that is 100 nanoseconds after Jan 1, 1970.
pd.to_datetime(100)
# ### Specify unit
#
# The default unit is nanoseconds, but you can specify a different one with the **`unit`** parameter.
# 100 seconds after the epoch
pd.to_datetime(100, unit='s')
# 20,000 days after the epoch
pd.to_datetime(20000, unit='d')
# ### Not a Series or a DataFrame
#
# When using Pandas, you are almost always working with either a Series or a DataFrame (and occasionally an Index). The Pandas Timestamp is another type unique to Pandas, but you will rarely be working with it directly.
#
# ## Why is `to_datetime` returning a `Timestamp` object?
#
# It must look a bit odd to see a Timestamp object being returned from the `to_datetime` function. The docstrings for `to_datetime` even write:
#
# > Convert argument to datetime.
#
# Technically, the object is definitely a pandas Timestamp object. We can verify this with the `type` function:
dt = pd.to_datetime(20000, unit='d')
type(dt)
# ### Datetime is common terminology in many languages
#
# The term **datetime** is common in many programming languages and this is what the Pandas documentation is referring to. The technical name of the Pandas object is indeed Timestamp, but the common name for what it represents is a datetime.
#
# ### Timestamp and datetime refer to the same thing
#
# The terms **Timestamp** and **datetime** refer to the exact same concept in pandas. Technically, each value is a Pandas `Timestamp` object but the term **datetime** is used to refer to it as well. Yes, that is extremely confusing, but hopefully now it is clear.
#
# ### Typical Timestamps in Pandas
# Typically, you will encounter Timestamps within a column of a Pandas DataFrame as we do below. Note, that the data type is `datetime64`. This is confusing, but again, Timestamp and datetime are equivalent terms.
emp = pd.read_csv('../data/employee.csv', parse_dates=['hire_date'])
emp.dtypes
# ### Each individual value in the datetime columns is a Timestamp
# If we extract the **`hire_date`** column as a Series and print out the first few rows, you will see that data type (at the bottom of the output) is still written with the word **datetime**.
hire_date = emp['hire_date']
hire_date.head()
# If we select the first value in the Series, we get a Timestamp.
hire_date.loc[0]
# ## Timestamp attributes
# These Timestamp objects have similar attributes and methods as the **`dt`** Series accessor in a previous notebook. Let's see some of these again.
ts = pd.to_datetime('Jan 3, 2019 20:45.56')
ts.day
ts.day_name()
ts.minute
# ## Timedelta - an amount of time
# A timedelta is a specific amount of time such as 20 seconds, or 13 days 5 minutes and 10 seconds. Use the **`to_timedelta`** function to create a Timedelta object. It works analogously to the **`to_datetime`** function.
#
# ### Converting strings to a Timedelta with `to_timedelta`
# A wide variety of strings are able to be converted to Timedeltas. [See the docs][1] for more info.
#
# [1]: http://pandas.pydata.org/pandas-docs/stable/timedeltas.html#to-timedelta
pd.to_timedelta('5 days 03:12:45.123')
# 10 hours and 13 microseconds
pd.to_timedelta('10h 13ms')
# ### Converting numbers to Timedeltas with `to_timedelta`
# As with **`to_datetime`**, passing a number to **`to_timedelta`** will be by default treated as the number of nanoseconds. Use the **`unit`** parameter to change the time unit.
# 123,000 nanoseconds
pd.to_timedelta(123000)
# 500 days
pd.to_timedelta(500, unit='d')
# Since years is not a standard amount, the highest unit returned is in days. You can still use 'y' to represent years with the output converted to days.
# 23 years
pd.to_timedelta(23, unit='y')
# 10 hours
pd.to_timedelta(10, 'h')
# ### No name confusion with Timedelta
# The Timedelta data type is unique to pandas just like the Timestamp object is. Pandas Timedelta is built upon NumPy's timedelta64 data type which is superior to pure Python's timedelta. Forunately, the Pandas developers used the name **Timedelta** for the data type which is the same as NumPy's.
#
# There is no name confusion here, unlike there is with **Timestamp/Datetime**.
td = pd.to_timedelta(3, 'y')
type(td)
# ## Timedelta attributes and methods
# There are many attributes and methods available to Timedelta objects. Let's see some below:
td
td.days
td.seconds
td.components
# ## Creating Timedeltas by subtracting Datetimes
# It is possible to create a Timedelta object by subtracting two Datetimes.
dt1 = pd.to_datetime('2012-12-21 5:30')
dt2 = pd.to_datetime('2016-1-1 12:45:12')
dt1
dt2
# Subtraction:
dt2 - dt1
# ### Negative Timedeltas
# A negative amount of time is possible just like any negative number is.
dt1 - dt2
# ### Math with Timedeltas
# You can do many different math operations with two Timedeltas together.
td1 = pd.to_timedelta('05:23:10')
td2 = pd.to_timedelta('00:02:20')
td1 - td2
td2 + 5 * td2
# Dividing two timedeltas will remove the units and return a number.
td1 / td2
# ### Creating Timedeltas in a DataFrame by subtracting two Datetime columns
# The bikes dataset has two datetime columns, **`starttime`** and **`stoptime`**.
bikes = pd.read_csv('../data/bikes.csv', parse_dates=['starttime', 'stoptime'])
bikes.head()
# Let's find the amount of time that elapsed between the start and stop times.
time_elapsed = bikes['stoptime'] - bikes['starttime']
time_elapsed.head()
# Since both start and stop time are datetime columns, subtracting them resulted in a timedelta column. The maximum unit of time for timedelta is days.
# ## Exercises
# ### Exercise 1
# <span style="color:green; font-size:16px">What day of the week was Jan 15, 1997?</span>
# ### Exercise 2
# <span style="color:green; font-size:16px">Was 1925 a leap year?</span>
# ### Exercise 3
# <span style="color:green; font-size:16px">What year will it be 1 million hours after the UNIX epoch?</span>
# ### Exercise 4
# <span style="color:green; font-size:16px">Create the datetime July 20, 1969 at 2:56 a.m. and 15 seconds.</span>
# ### Exercise 5
# <span style="color:green; font-size:16px"><NAME> stepped on the moon at the time in the last exercise. How many days have passed since that happened? Use the string 'today' when creating your datetime.</span>
# ### Exercise 6
# <span style="color:green; font-size:16px">Which is larger - 35 days or 700 hours?</span>
# ### Exercise 7
# <span style="color:green; font-size:16px">In a previous notebook, we were told that the employee data was retrieved on Dec 1, 2016. We used the simple calculation `2016 - emp['hire_date'].dt.year` to determine the years of experience. Can you improve upon this method to get the exact amount of years of experience and assign this as a new column named `experience`?</span>
| jupyter_notebooks/pandas/mastering_data_analysis/07. Time Series/01. Datetime and Timedelta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #### To do
# * Get Intro written with simple 3D image
# * Have as much code included in a library as possible
# * Get straight to the point
# * Then break down the function
# ### Z-slicing of triangulated surface with tool
#
# One of the fundamental algorithms in 3-axis machining is the Z-slicer
# which is the basis of the "waterline toolpath".
# It is necessary to the paths from each slice before it becomes a toolpath,
# but we will look at this later.
#
# This Z-slicing function takes in a toolshape, a Z-value, and a surface
# to be machined in the form of a set of triangles. It returns
# a set of 2D contours in the Z-plane which surround the area
# where the tool can move free of any contact with the surface.
# The points on these contours represent the positions where the
# tool is just touching the surface.
#
# +
# %matplotlib inline
from zslicing_funcs import loadplottriangles, plottoolshape, set_aspect_equal_3d
fname = "stlsamples/frameguide.stl"
tbm = loadplottriangles(fname)
ax = plottoolshape(5, 68, -2, -20, 1)
set_aspect_equal_3d();
# -
#
# which takes a toolshape (aligned with the Z-axis), a
# surface defined as set of triangles, and a horizontal plane,
# and returns a set of contours in the plane enclosing the areas
# where the tool would be in contact (or embedded) in the surface.
#
# **Tool definition**:
# Consider a ball-nosed tool of radius $r$ and tip at point
# $p = (p_x, p_y, p_z)$,
# then the set of points within this tool could be defined as:
# $$ T_p = \{q \mid \lvert q-p_{+zr} \rvert \le r \}
# \cup
# \lbrace q \mid
# q_z \ge p_z+r \text{ and }
# \lvert q_{xy} - p_{xy} \rvert \le r
# \rbrace $$
# where
# $$p_{xy} = (p_x, p_y, 0), \space p_{+zr} = (p_x, p_y, p_z+r).$$
# This is simply the union of a sphere and a cylinder extending
# upward to infinity. By convention we position the tool according to
# its tip rather than the centre of the ball (although it would be
# more mathematically natural), because it can be directly aligned --
# you can lower the tip of the tool until it is in contact
# with a flat surface, but you can't do the same with its centre.
#
# **Surface model definition**:
# Most machining algorithms require the model to be approximated
# by a (large) set of triangles. It is normally well within the
# capability of the average computer to hold such an approximation
# to a realistic tolerance within its main memory. Triangles
# are used because they are well defined, and every CAD model
# that has any sort of visualization is capable of
# outputting them owing to the fact that computer graphics
# depends on it. The memory available in the average computer
# far outstrips what is realistically demanded in terms
# of machining tolerance.
#
# A triangle $R$ is defined from three points $(p_a, p_b, p_c)$
# where:
# $$R = \lbrace p_a + \lambda (p_b - p_a) + \mu (p_c - p_a) \mid
# \lambda, \mu \geq 0 \text{ and } \lambda + \mu \leq 1 \rbrace$$
# A model $M$ is simply a set of such triangles:
# $$M = \lbrace R_i \mid i = 1, \dots, n \rbrace$$
# Often these triangles share edges and vertices to form a surface
# mesh, but it is not a requirement.
#
# **Tool area of the Z-plane**:
# Given a horizontal plane at height $z$ defined as:
# $$H_z = \lbrace (x, y, z) \mid -\infty <x<\infty, -\infty < y < \infty \rbrace$$
# then the area of impingement of the tool in this plane is
# simply defined as:
# $$ S_{T,M,z} = \lbrace p \in H_z \mid T_p \cap R \neq \emptyset
# \text{ for some } R \in M \rbrace$$
# This is a complicated way of saying that we keep all the
# points where the tool occupies the same volume as
# the surface. The boundary of this area will be the
# set of points where the tool is touching (is in contact with)
# the surface. When it goes deeper than mere touching, we call
# it "gouging".
#
# It should be obvious that:
# $$S_{T,M,z} = \bigcup_{R \in M} S_{T,R,z}$$
# where:
# $$ S_{T,R,z} = \lbrace p \in H_z \mid T_p \cap R \neq \emptyset
# \rbrace$$
# This is important, because it means we can define
# the functions to calculate $S_{T,R,z}$ based on a single triangle,
# and know how to generalize it to an model $M$ composed of a
# set of such triangles.
#
#
#
# * Get the single triangle to plot (maybe in 3D) with the toolshape
# * Get the tool shaft and ball both working
# * Plot in 2D the triangle
# * Plot the sample space around the triangles
# * Plot a series of radiating lines out from the centre point
# * Plot the combined lines between the graph paper arranged nodes
# * plot the full on subdividing contour
# * Run and plot the values at many slices
# * Plot same in 3D (if possible)
# * Commit and send to FreeCAD guy
# * This fundamental algorithm works for rest area detection, pencil milling, constant scallop.
# * I would like to have it sufficiently opened out that we can consider getting it to a form where it can run on a GPU; in which case it will out-perform the commercial systems out there.
# +
from matplotlib import pyplot as plt
from matplotlib import collections as mc
# %matplotlib inline
tbarmesh = TriangleBarMesh(flat9triangles=[[0,0,0, 1,0,1, 0.5,2,2]])
# -
[[(bar.nodeback.p.x, bar.nodefore.p.x), (bar.nodeback.p.y, bar.nodefore.p.y)] for bar in tbarmesh.bars ]
for bar in tbarmesh.bars:
plt.plot((bar.nodeback.p.x, bar.nodefore.p.x), (bar.nodeback.p.y, bar.nodefore.p.y))
ax = plt.gca()
ax.add_collection(mc.LineCollection([[(bar.nodeback.p.x, bar.nodefore.p.x), (bar.nodeback.p.y, bar.nodefore.p.y)] for bar in tbarmesh.bars ], linewidths=2))
ax.autoscale()
ax.margins(0.1)
# +
lines = [[(0, 1), (1, 1)], [(2, 3), (3, 3)], [(1, 2), (1, 3)]]
lc = mc.LineCollection(lines, linewidths=2)
ax = plt.gca()
ax.add_collection(lc)
ax.autoscale()
ax.margins(0.1)
# +
# PointZone { izone: {PZ_WITHIN_R, PZ_BEYOND_R}, r, v }
from tool_impingement import spheredistance, cylinderdistance
from tool_impingement.triangleimpingementscanning import ImpingementShape
# these upgrade the pointzone to something specific to the toolshape
def DPZsphere(p, pz, iasphere):
return spheredistance.DistPZ(p, pz)
def DLPZsphere(p, vp, r, iasphere):
return spheredistance.DistLamPZ(p, vp, r)
def DPZcylinder(p, pz, iacylinder):
return cylinderdistance.DistPZC(p, pz.r, iacylinder.zlo, iacylinder.zhi)
def DLPZcylinder(p, vp, r, iacylinder):
return cylinderdistance.DistLamPZC(p, vp, r, iacylinder.zlo, iacylinder.zhi)
class ImpingementSphere(ImpingementShape):
def __init__(self, tbarmesh, tboxing, r):
super().__init__(tbarmesh, tboxing, DPZsphere, DLPZsphere)
def Isb2dcontournormals(self):
return False
class ImpingementCylinder(ImpingementShape):
def __init__(self, tbarmesh, tboxing, r, zlo, zhi):
super().__init__(tbarmesh, tboxing, DPZcylinder, DLPZcylinder)
assert zlo <= zhi
self.zlo = zlo
self.zhi = zhi
def Isb2dcontournormals(self):
return False
# -
from tribarmes import TriangleBarMesh, SingleBoxedTriangles
tbarmesh = TriangleBarMesh(flat9triangles=[[0,0,0, 1,0,1, 0.5,2,2]])
tboxing = SingleBoxedTriangles(tbarmesh)
tboxing.CloseBoxeGenerator(0,0,0,0,0)
from barmesh import PointZone
from basicgeo import P3
sphrad = 0.5
ia = ImpingementSphere(tbarmesh, tboxing, sphrad)
pz = PointZone(0, 10, None)
p = P3(0.1, 0.1, 0.3)
ia.DistP(pz, p)
pz.r
# +
from matplotlib import pyplot as plt
# %matplotlib inline
import numpy
xs = numpy.arange(-1, 2, 0.1)
ys = numpy.arange(-1, 2, 0.1)
kxs, kys = [ ], [ ]
zslice = 0.3
kpts = [ ]
for x in xs:
kpts.append([])
for y in ys:
pz = PointZone(0, 10, None)
p = P3(x, y, 0.3)
ia.DistP(pz, p)
kpts[-1].append((p, pz))
kxs, kys = [ ], [ ]
for kx in kpts:
for ky in kx:
if ky[1].r < sphrad:
kxs.append(ky[0].x)
kys.append(ky[0].y)
plt.scatter(kxs, kys, marker=".")
from basicgeo import Along
kxs, kys = [ ], [ ]
for i in range(1, len(kpts)):
for j in range(1, len(kpts[i])):
if kpts[i][j][1].r > sphrad and kpts[i-1][j][1].r < sphrad:
pout, pin = kpts[i][j][0], kpts[i-1][j][0]
lam = ia.Cutpos(pout, pin - pout, None, sphrad)
plt.plot([pout.x, Along(lam, pout.x, pin.x)], [pout.y, Along(lam, pout.y, pin.y)], color="red")
print(lam)
# p = P3(0.1, 0.1, 0.3)
#x.DistP(pz, p)
#pz.r
# -
| zslicing_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Artificial Neural Networks
# ## 神经网络(ANN)的开山论文
# > ### 1943 A Logical Calculus of Ideas Immanent in Nervous Activity
# >### 神经活动内在性的逻辑计算(神经元M-P模型)
# 
# 
# >#### 细胞体(soma) 神经元控制中心,产生膜电位
# >#### 树突(dendron/dendrites) 神经元输入端
# >#### 轴突(axon) 末端有大量神经末梢,细胞体输出端
# >#### 突触(synapse) 神经元之间的I/O口
# # Logical Computations with Neurons
# 神经元的逻辑运算
# 神经元接受二进制输入,当达到阈值时激活输出,通过简单组合可以模拟出大部分运算
# 例如:假设神经元接受超过两个刺激则激活输出
#
# 人造神经元只能接受刺激,1957年提出的$linear\ threshold\ unit(LTU)$则可以接受数值输入,
# 其汇总所有的输入$\mathbf{z}=\mathbf{w}^T\cdot \mathbf{x}$,经过step function(阶跃函数)输出数值$step(\mathbf{z})$
# 常见的两种阶跃函数:
# 单位阶跃函数$heaviside(z)=\begin{cases}
# 0 & \text{ if } z<0 \\
# 1 & \text{ if } z\geq 0
# \end{cases}$, 符号函数$sgn(z)=\begin{cases}
# -1 & \text{ if } z<0 \\
# 0 & \text{ if } z= 0 \\
# 1 & \text{ if } z > 0
# \end{cases}$
# # The Perceptron
# 我们一般称感知机即单层$LTUs$,只有输入层和输出层构成,其中输入层中含有$bias\ neuron$偏置神经元($x_0=1$)
# **如何训练感知机(神经元)**
# 根据*Hebb's rule* ——“$\mathrm{Cells\ that\ fire\ together,\ wire\ together}$” 一起激发的神经元连接在一起
# 基于Hebb定理的学习称为Hebb Learning
# 感知机训练使用Hebb定理的变体,其着重考虑错网络误输出,更新连接上权重
#
# +
import numpy as np
from sklearn.linear_model import Perceptron
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from tensorflow_graph_in_jupyter import show_graph
import matplotlib.pylab as plt
data = load_iris()
X = data['data'][:, (2, 3)]
y = (data['target'] == 0).astype(np.int)
def plot_dataset(X, y):
plt.plot(X[:, 0][y==0], X[:, 1][y==0], 'go', label='Positive')
plt.plot(X[:, 0][y==1], X[:, 1][y==1], 'r^', label='Negative')
plt.grid(True, which='both')
plt.legend()
def plot_predictions(clf, axes):
x0s = np.linspace(axes[0], axes[1], 100)
x1s = np.linspace(axes[2], axes[3], 100)
x0, x1 = np.meshgrid(x0s, x1s)
X = np.c_[x0.ravel(), x1.ravel()]
y_pred = clf.predict(X).reshape(x0.shape)
y_decision = clf.decision_function(X).reshape(x0.shape)
plt.contourf(x0, x1, y_pred, cmap=plt.cm.brg, alpha=0.2)
plt.contourf(x0, x1, y_decision, cmap=plt.cm.brg, alpha=0.1)
plt.legend()
# x,y,z z为(x,y)点值
plot_dataset(X, y)
X_train, X_test, y_trian ,y_test = train_test_split(X, y, random_state=0)
per_clf = Perceptron(max_iter=1000) # 仅支持二分类的感知机,训练的是输入层上的权重,类似GD
per_clf.fit(X_train, y_trian)
y_pred = per_clf.predict(X_test)
print(classification_report(y_test, y_pred))
plt.title(r'navie Perceptron')
plot_predictions(per_clf, axes=[0.5,7, 0, 2.8])
plt.show()
# +
from sklearn.linear_model import SGDClassifier
plot_dataset(X, y)
X_train, X_test, y_trian ,y_test = train_test_split(X, y, random_state=0)
per_gd_clf = SGDClassifier(max_iter=1000, loss='perceptron', learning_rate='constant',
eta0=1, penalty=None, random_state=0)
per_gd_clf.fit(X_train, y_trian)
y_gd_pred = per_gd_clf.predict(X_test)
print(classification_report(y_test, y_gd_pred))
plt.title(r'navie Perceptron by SGD')
plot_predictions(per_gd_clf, axes=[0.5,7, 0, 2.8])
plt.show()
# -
# 感知机收敛理论:解不唯一
# 单层感知机无法解决非线性问题例如异或划分
# ### Mutil-Layer Perceptron($MLP$) and Backpropagation
# $\mathrm{MLP}$由多个$Hidden$层构成,故也称之为$deep\ neural\ network(DNN)$
# 反向传播算法利用梯度更新连接权重下降,所以我们用逻辑斯蒂方程代替阶跃函数,
# 此外我们引入激活函数调整逻辑斯蒂函数的输出在$[-1,1]$上
# 以使得每层的输出在开始训练的时候或多或少标准化(more or less normalized)
# 使输出在0附近和加快收敛
# 常用的激活函数($\mathrm{activation\ function}$):
# $1.\ \ hyperbolic\ tangent\ function\ \ tanh(z)=2\sigma (2z)-1$
# $2.\ \ \mathrm{The\ ReLU\ function}\ \ $ $ReLU(z)=max(z, 0)$
# [tanh is a rescaled logistic sigmoid function](https://brenocon.com/blog/2013/10/tanh-is-a-rescaled-logistic-sigmoid-function/)
# %%time
import matplotlib.pylab as plt
plt.figure(figsize=(10, 6))
x = np.linspace(-4, 4, 100000)
y1 = 1.0 / (1 + np.exp(-x))
y2 = 2.0 / (1 + np.exp(-2*x)) - 1
y3 = []
y4 = []
for i in x:
y3.append(max(0, i))
if(i < 0):
y4.append(-1.0)
elif np.allclose(i, 0):
y4.append(0)
else :
y4.append(1)
plt.title(r'$\mathrm{activation\ function}$')
plt.plot(x, y1, '--', label=r'$\sigma(z)$');
plt.plot(x, y2, '--', label=r'$tanh(z)$');
plt.plot(x, y3, label=r'ReLU');
plt.plot(x, y4, label=r'Step')
plt.ylim(-1.2, 1.2)
plt.legend(loc="lower right")
# 当class互斥时,使用Softmax代替个体激活函数
# ### FNN($feedforward\ neural\ network$)
# 前馈神经网络
# #### MLP apply in MNIST classification base on TensorFlow High-Level API
# +
# %%time
# ================================================================================================
# MLP apply in MNIST classification base on TensorFlow High-Level API
# two hidden layers; 1st: 300 neurons; 2ed: 100 neurons
# Softmax output alyer with 10 neurous
import tensorflow as tf
from sklearn.datasets import fetch_mldata
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report, accuracy_score, precision_score, recall_score
from sklearn.preprocessing import StandardScaler
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data() # 使用keras后端调取数据
X_train = X_train.astype(np.float32).reshape(-1, 28*28) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, 28*28) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=1/6)
# +
# %%time
feature_cols = [tf.feature_column.numeric_column("X", shape=[28*28])] # 数值特征列
dnn_clf = tf.estimator.DNNClassifier(hidden_units=[300,100], n_classes=10, feature_columns=feature_cols)
input_fn = tf.estimator.inputs.numpy_input_fn(x={'X':X_train}, y=y_train, num_epochs=40, batch_size=50, shuffle=True)
# -
dnn_clf.train(input_fn=input_fn)
test_input_fn = tf.estimator.inputs.numpy_input_fn(
x={'X': X_test}, y = y_test, shuffle=False # 打乱标记
)
eval_result = dnn_clf.evaluate(input_fn=test_input_fn)
print('DNNC in Test Set', eval_result)
# 查看每个class预测概率决策
y_pred_iter = dnn_clf.predict(input_fn=test_input_fn) # prediction生成器
error_cnt = 0
f = 0
for i in range(len(X_test)):
a = next(y_pred_iter)
b = [y_test[i]][0]
if not f:
print(a)
f = 1
a = a['class_ids'][0]
if a != b:
pass
# print(i, '预测:', a, '\t标记:', b)
error_cnt += 1
print(1 - error_cnt / len(X_test))
# #### DNNClassification 创建所有的含ReLU激活函数的神经元层, 输出层采取Softmax,损失函数为交叉熵
# ## Training a DNN Using Plain TensorFlow
# #### Construction Phasea
# +
import tensorflow as tf
import numpy as np
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
# -
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name='X') # 训练输入
y = tf.placeholder(tf.int32, shape=(None), name='y') # 标记
def neuron_layer(X, n_neurons, name, activation=None): # 定义神经元层
with tf.name_scope(name=name):
n_inputs = int(X.get_shape()[1]) # 该层神经元个数
stddev = 2 / np.sqrt(n_inputs)
init = tf.truncated_normal(shape=(n_inputs, n_neurons), stddev=stddev) # 截断标准偏差,加速模型收敛,见Chapter 11
W = tf.Variable(initial_value=init, name='weights') # 初始化连接层间的权重,
b = tf.Variable(tf.zeros([n_neurons]), name='biases') # 初始化为0避免了对称性
z = tf.matmul(X, W) + b
if activation is not None:
return activation(z)
else:
return z
with tf.name_scope(name='dnn'): # 定义DNNs
hidden1 = neuron_layer(X, n_hidden1, 'hidden1', activation=tf.nn.relu)
hidden2 = neuron_layer(hidden1, n_hidden2, 'hidden2', activation=tf.nn.relu)
logits = neuron_layer(hidden2, n_outputs, 'output', activation=None) # 暂不处理Softmax 输出层
with tf.name_scope('loss'): # 定义loss function
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name='loss')
lr = 0.01
with tf.name_scope('train'): # 定义优化器
optimizaer = tf.train.GradientDescentOptimizer(learning_rate=lr)
training_op = optimizaer.minimize(loss)
with tf.name_scope('eval'):
correct = tf.nn.in_top_k(logits, y, 1) # y是否是logists的第一个元素,即预测是否正确
acc = tf.reduce_mean(tf.cast(correct, tf.float32)) # 转换成float求平均
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 40
batch_size = 50
# +
# %%time
from sklearn.model_selection import train_test_split
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train = X_train.astype(np.float32).reshape(-1, n_inputs) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, n_inputs) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=1/6)
print(X_train.shape, X_val.shape)
def shufft_batch(X, y, batch_size): # 生辰batch器
rnd_idx = np.random.permutation(len(X))
n_batches = len(X) // batch_size
for batch_idx in np.array_split(rnd_idx, n_batches):
yield X[batch_idx], y[batch_idx]
# -
# %%time
with tf.Session() as sess:
init.run()
with tf.name_scope('train'):
for epoch in range(n_epochs):
for X_batch, y_batch in shufft_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X:X_batch, y:y_batch})
acc_batch = acc.eval(feed_dict={X:X_batch, y:y_batch})
acc_val = acc.eval(feed_dict={X:X_val, y:y_val})
print('Acc_batch: %.16f <====================> Acc_val: %.16f' % (acc_batch, acc_val))
saver_path = saver.save(sess, './tmp/mnist_my_final_model.ckpt')
acc_test = acc.eval(feed_dict={X:X_test, y:y_test})
print('Acc_test: %.16f' % acc_test)
import sys
from tensorflow_graph_in_jupyter import show_graph
# sys.path.append('/Users/hu-osx/Documents/Machine-Learning/handson-ml/')
show_graph(tf.get_default_graph())
# 使用TensorFlow全连接层代替手工实现的
# +
# %%time
import tensorflow as tf
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
tf.reset_default_graph()
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X_train = X_train.astype(np.float32).reshape(-1, n_inputs) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, n_inputs) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=1/6)
print(X_train.shape, X_val.shape)
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name='X') # 训练输入
y = tf.placeholder(tf.int32, shape=(None), name='y') # 标记
with tf.name_scope(name='dnn'): # 定义DNNs
he_init = tf.contrib.layers.variance_scaling_initializer()
hidden1 = tf.layers.dense(X, n_hidden1, name='hidden1', activation=tf.nn.relu)
hidden2 = tf.layers.dense(hidden1, n_hidden2, name='hidden2', activation=tf.nn.relu)
logits = tf.layers.dense(hidden2, n_outputs, name='output')
y_prob = tf.nn.softmax(logits)
with tf.name_scope('loss'): # 定义loss function
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name='loss')
lr = 0.01
with tf.name_scope('train'): # 定义优化器
optimizaer = tf.train.GradientDescentOptimizer(learning_rate=lr)
training_op = optimizaer.minimize(loss)
with tf.name_scope('eval'):
correct = tf.nn.in_top_k(logits, y, 1) # y是否是logists的第一个元素,即预测是否正确
acc = tf.reduce_mean(tf.cast(correct, tf.float32)) # 转换成float求平均
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 50
def shufft_batch(X, y, batch_size): # 生辰batch器
rnd_idx = np.random.permutation(len(X))
n_batches = len(X) // batch_size
for batch_idx in np.array_split(rnd_idx, n_batches):
yield X[batch_idx], y[batch_idx]
with tf.Session() as sess:
init.run()
with tf.name_scope('train'):
for epoch in range(n_epochs):
for X_batch, y_batch in shufft_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X:X_batch, y:y_batch})
acc_batch = acc.eval(feed_dict={X:X_batch, y:y_batch})
acc_val = acc.eval(feed_dict={X:X_val, y:y_val})
print(epoch, 'Batch acc: %.16f <====================> Valid acc: %.16f' % (acc_batch, acc_val))
saver_path = saver.save(sess, './tmp/mnist_my_final_model.ckpt')
acc_test = acc.eval(feed_dict={X:X_test, y:y_test})
print('Test acc: %.16f' % acc_test)
# Test acc: 0.9699000120162964 ('ReLU + X-init')
# Test acc: 0.9600999951362610 ('ELU' + he_init)
# show_graph(tf.get_default_graph())
# +
# 使用Softmax过的logits预测最大可能的手写体
with tf.Session() as sess:
saver.restore(sess, './tmp/mnist_my_final_model.ckpt') # 还原模型
idx = 2333
X_new_scaled = X_test[idx].reshape(1,-1)
print('label:', y_test[idx])
Z = logits.eval(feed_dict={X:X_new_scaled})
sZ = tf.nn.softmax(Z)
y_prob = np.argmax(Z, axis=1)
print('prediction:', y_prob, ', With probability', sZ.eval().flatten()[y_prob])
# -
# ### Exercises
# 5
# >$\mathbf{X}'s\ shape\ is\ None \times 10 $
# $\mathbf{W}_h's\ shape\ is\ 10\ \times 50$
| handson-ml/10_ANN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tigergraph Bindings: Demo of IT Infra Analysis
#
# Uses bindings built into PyGraphistry for Tigergraph:
#
# * Configure DB connection
# * Call dynamic endpoints for user-defined endpoints
# * Call interpreted-mode query
# * Visualize results
# ## Import and connect
# +
import graphistry
# # !pip install graphistry -q
# To specify Graphistry account & server, use:
# graphistry.register(api=3, username='...', password='...', protocol='https', server='hub.graphistry.com')
# For more options, see https://github.com/graphistry/pygraphistry#configure
# -
g = graphistry.tigergraph(
protocol='http', server='www.acme.org',
user='tigergraph', pwd='<PASSWORD>',
db='Storage', #optional
#web_port = 14240, api_port = 9000, verbose=True
)
# ## Dynamic user-defined GSQL endpoints: Call, analyze, & plot
# +
g2 = g.gsql_endpoint(
'StorageImpact', {'vertexType': 'Service', 'input': 61921, 'input.type': 'Pool'},
#{'edges': '@@edgeList', 'nodes': '@@nodeList'}
)
print('# edges:', len(g2._edges))
g2.plot()
# -
# ## On-the-fly GSQL interpreted queries: Call, analyze, & plot
# +
g3 = g.gsql("""
INTERPRET QUERY () FOR GRAPH Storage {
OrAccum<BOOL> @@stop;
ListAccum<EDGE> @@edgeList;
SetAccum<vertex> @@set;
@@set += to_vertex("61921", "Pool");
Start = @@set;
while Start.size() > 0 and @@stop == false do
Start = select t from Start:s-(:e)-:t
where e.goUpper == TRUE
accum @@edgeList += e
having t.type != "Service";
end;
print @@edgeList;
}
""",
#{'edges': '@@edgeList', 'nodes': '@@nodeList'} # can skip by default
)
print('# edges:', len(g3._edges))
g3.plot()
# -
| demos/demos_databases_apis/tigergraph/tigergraph_pygraphistry_bindings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intro Deep Learning
# %matplotlib inline
import matplotlib.pylab as plt
import numpy as np
import pandas as pd
from sklearn import linear_model
pd.set_option('display.max_rows',10)
pd.set_option('precision',3)
# Read data linear_regression_demo-master
dataframe = pd.read_fwf('linear_regression_demo-master/brain_body.txt')
datafram
x_values = dataframe[['Brain']]
x_values
y_values = dataframe[['Body']]
y_values
# # And our goal is that given a new animal's body weight will be able to predict what its brain size is.
#
# My data's got the values for the brain and the body weight and im wondering what to use to find if they relate?
#
# Lineal Regretion
#
# $y = mx + b$
dataframe.plot.scatter('Brain','Body')
# +
# body_reg.fit?
# +
# train model on data
body_reg = linear_model.LinearRegression()
body_reg.fit(dataframe[ ["Brain"] ] ,dataframe[ ["Body"]] )
# +
#ax.plot(x_values,body_reg.predict(x_values) )
ax= dataframe.plot.scatter('Brain','Body')
plt.plot(dataframe[ ["Brain"] ],body_reg.predict(dataframe[ ["Brain"]]) )
# +
# Read data linear_regression_demo-master
dataframe = pd.read_fwf('linear_regression_demo-master/brain_body.txt')
body_reg = linear_model.LinearRegression()
body_reg.fit(dataframe[ ["Brain"] ] ,dataframe[ ["Body"]] )
ax= dataframe.plot.scatter('Brain','Body')
plt.plot(dataframe[ ["Brain"] ],body_reg.predict(dataframe[ ["Brain"]]) )
# -
# # Breaking it down
#
# Linear Regression models relationship between independent and dependet variables via line of best fit
| SirajRaval_Ejemplos/IntroDeeplearning/LinealRegresion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('base')
# language: python
# name: python3
# ---
# # RESIZING AND CROPPING
# +
import cv2
img = cv2.imread('resources/titan.png')
# To check the size of image 'titan.png'
print(img.shape) # The output gives us 879 as height and 1556 as width and 3 is the chanel i.e BGR
# To resize the image
imgResize = cv2.resize(img, (300,200)) # width and height are 300,200
print(imgResize.shape)
imgCropped = img[0:400, 400:800]
# Name the window 'Output'
cv2.imshow('Output', img)
cv2.imshow('Resized Image', imgResize)
cv2.imshow('Cropped image', imgCropped)
cv2.waitKey(0)
| basics2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Guide
#
# The ArcGIS API for Python is a powerful, modern and easy to use Pythonic library to perform GIS visualization and analysis, spatial data management and GIS system administration tasks that can run both in an interactive fashion, as well as using scripts.
#
# It enables power users, system administrators and developers to leverage the rich SciPy ecosystem for automating their workflows and performing repetitive tasks using scripts. It integrates well with the IPython Notebook and enables academics, data scientists, GIS analysts and visualization enthusiasts to share geo-enriched literate programs and reproducible research with others.
#
# This guide describes how to use the ArcGIS API for Python to write Python scripts, incorporating capabilities such as mapping, query, analysis, geocoding, routing, Portal administration, and more. A great place to start developing once you've [installed the API](https://developers.arcgis.com/python/guide/Install-and-set-up/) is to browse the [sample notebooks](https://developers.arcgis.com/python/sample-notebooks/).
| guide/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os.path import join
from pyleecan.Functions.load import load
from pyleecan.definitions import DATA_DIR
from pyleecan.Classes.Skew import Skew
# Load machines
Toyota_Prius = load(join(DATA_DIR, "Machine", "Toyota_Prius.json"))
SCIM_001 = load(join(DATA_DIR, "Machine", "Railway_Traction.json"))
# Define lamination skew (optional): "linear", "vshape", "function"
SCIM_001.rotor.skew = Skew(type_skew="linear", rate=36/28, is_step=False)
Toyota_Prius.rotor.skew = Skew(type_skew="vshape", rate=2, is_step=True, Nstep=5)
# Plot skew
SCIM_001.rotor.skew.plot()
Toyota_Prius.rotor.skew.plot()
# +
### WORK IN PROGRESS
# Parametrize skew model: "uniform", "gauss" or "user-defined"
#slice_model = SliceModel(type_distribution="gauss", Nslices=7)
# Compute slices and plot
#slice_model.plot()
# -
| Tutorials/tuto_Skew.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convert to HDF5
#
# This notebook has instructions on how to prepare final HDF5 dataset. It assumes you have followed the instructions to get consolidated data in `final/` folder. Each json file has entries for that WSID indexed to a global unified index.
# !pip install h5py -q
# !pip install ftfy -q
# +
import re
import h5py # main dumping method
# import networkx as nx
import pandas as pd
import numpy as np
# import matplotlib.pyplot as plt
# # %matplotlib inline
# import orjson
# import vaex
import json
from glob import glob
from ftfy import ftfy
from io import StringIO
from dateparser import parse
from collections import Counter
from tqdm import trange
# from tqdm.notebook import trange
import math
from datetime import timedelta, date
from functools import reduce
# -
def daterange(start_date, end_date):
for n in range(int((end_date - start_date).days)):
yield start_date + timedelta(n)
# +
COLUMNS = [
("total_precipitation", "mm"),
("pressure", "mB"),
("max_pressure", "mB"),
("min_pressure", "mB"),
("radiation", "KJ/m^2"),
("temp", "C"),
("dew_point_temp", "C"),
("max_temp", "C"),
("min_temp", "C"),
("max_dew", "C"),
("min_dew", "C"),
("max_humidity", "percentage"),
("min_humidity", "percentage"),
("humidity", "percentage"),
("wind_direction", "deg"),
("wind_gust", "m/s"),
("wind_speed", "m/s")
]
COL_NAMES = [x[0] for x in COLUMNS]
UNITS = [x[1] for x in COLUMNS]
BAD_ROW_SUM = 0.0
def open_csv(f, wsid):
with open(f, "r", encoding="iso8859_1") as d:
df = pd.read_csv(
StringIO("".join(d.readlines()[8:]).replace("-9999", "0")),
sep=";",
encoding="iso8859_1"
)
df = df.drop("Unnamed: 19", axis = 1)
if wsid:
local_col_names = [f"{wsid}_{c}" for c in COL_NAMES]
else:
local_col_names = [f"{c}" for c in COL_NAMES]
df.columns = ["date", "hour"] + local_col_names
for col in local_col_names[2:]:
setattr(df, col, getattr(df, col).apply(lambda x: float(str(x).replace(",", "."))))
return df
# +
final_files = [x for x in glob("../INMET/final/*.json") if ("index" not in x and "wsids_ordered" not in x)]
print(f"final_files: {len(final_files)}")
with open("../INMET/final/index.json", "r") as f:
unified_idx = json.load(f)
print(len(unified_idx))
# -
gids_by_file = {}
for i, f in zip(trange(len(final_files)), sorted(final_files)): # sorted is very important because this way we can gaurantee ordering in array i,f in enumerate(sorted(final_files)):
with open(f, "r") as f2:
data = json.load(f2)
gids_by_file[f] = list(data.keys())
# +
# min_year = 2.5
year_wise_sample_count = {}
for min_year in np.linspace(1, 5, 9):
samples = [len(x) for x in gids_by_file.values()]
# np.mean(samples), np.median(samples)
useful_files = []
useful_files_gid = {}
for f in final_files:
# ignore all samples have < 2 years of data, ie < 24 * 365 * 2 samples => < 17520 samples
if len(gids_by_file[f]) >= 24 * 365 * min_year:
useful_files.append(f)
print(f"min: {min_year} Before: {len(gids_by_file)} After: {len(useful_files)}")
year_wise_sample_count[min_year] = len(useful_files)
# +
min_year = 4.5 # 488
samples = [len(x) for x in gids_by_file.values()]
# np.mean(samples), np.median(samples)
useful_files = []
useful_files_gid = {}
for f in final_files:
# ignore all samples have < 2 years of data, ie < 24 * 365 * 2 samples => < 17520 samples
if len(gids_by_file[f]) >= 24 * 365 * min_year:
useful_files.append(f)
useful_files = sorted(useful_files)
useful_wsids = [x[15:19] for x in useful_files]
# print(f"min: {min_year} Before: {len(gids_by_file)} After: {len(useful_files)}")
# -
# #### Positions Meta
#
# ```python
# wsmeta = pd.read_csv("../INMET/wsid_meta.csv")
# wsmeta = wsmeta.T
# headers = wsmeta.iloc[0].values.tolist()
# wsmeta = wsmeta[1:]
# wsmeta.columns = headers
# wsmeta = wsmeta[wsmeta.elev != "F"] # corrupt data
#
# wsmeta.lat = wsmeta.lat.values.astype(float)
# wsmeta.long = wsmeta.long.values.astype(float)
# wsmeta.elev = wsmeta.elev.values.astype(float)
#
# wsm = json.loads(wsmeta.to_json(orient="index"))
#
# with open("wsid_meta.json", "w") as f:
# f.write(json.dumps(wsm))
#
# ```
# +
with open("wsid_meta.json", "r") as f:
wsmeta = json.load(f)
wsmeta_ordered = []
for x in useful_wsids:
wm = wsmeta[x]
wsmeta_ordered.extend([wm["lat"], wm["long"], wm["elev"]])
wsmeta_ordered = np.array(wsmeta_ordered)
print(f"wsmeta_ordered: {wsmeta_ordered.shape}")
# +
# useful_files
# -
datetime_data = []
for k in sorted(unified_idx):
x = unified_idx[k]
mon = int(x[5:7])
day = int(x[8:10])
hrs = int(x[11:12])
datetime_data.append((mon, day, hrs))
datetime_data = np.array(datetime_data)
datetime_data
# +
# define the schema in hdf5
hdf = h5py.File("weatherGiga0.hdf5", "w")
hdf.create_dataset("wsid_meta", shape = wsmeta_ordered.shape, dtype = 'f', data = wsmeta_ordered)
hdf.create_dataset("datetime", shape = datetime_data.shape, dtype = 'i', data = datetime_data)
for _, i in zip(trange(len(unified_idx)), unified_idx):
grp = hdf.create_group(f"{i}")
data = np.zeros(shape = (len(useful_files), 17)).astype(np.float32)
grp.create_dataset("data", shape = data.shape, dtype = 'f', data = data)
grp.create_dataset("mask", shape = [data.shape[0]], dtype = 'f', data = data[:,0])
hdf.close()
# + jupyter={"outputs_hidden": true}
hdf = h5py.File("weatherGiga0.hdf5", "r+")
# sorted is very important because this way we can gaurantee ordering in array
uf = sorted(useful_files)
pb1 = trange(len(uf))
for i,f in zip(pb1, uf):
# if int(i) < 1:
# continue
w = useful_wsids[i]
pb1.set_description(f"{f}")
with open(f, "r") as f2:
data = json.loads(f2.read())
for _, gid in zip(trange(len(gids_by_file[f])), gids_by_file[f]):
this_idx_data = hdf[f"{gid}"]
if str(sum(data[gid][2:])) != "nan" and sum(data[gid][2:]) != 0:
try:
this_idx_data["data"][i] = data[gid]
this_idx_data["mask"][i] = 1
except:
continue
hdf.close()
# -
print("Complete Motherfucker")
hdf = h5py.File("weatherGiga0.hdf5", "r")
total = len(hdf)
train_idx = []
test_idx = []
for i in range(0, total, 100):
if np.random.random() < 0.9:
train_idx.append(i)
else:
test_idx.append(i)
len(train_idx), len(train_idx) / total * 100, len(test_idx), len(test_idx) / total * 100
# +
with open("train_idx.txt", "w") as f:
f.write("\n".join([str(x) for x in train_idx]))
with open("test_idx.txt", "w") as f:
f.write("\n".join([str(x) for x in test_idx]))
# -
| notebooks/prepare_hdf5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calling MOHYSE on the Raven server
#
# Here we use birdy's WPS client to launch the MOHYSE hydrological model on the server and analyze the output.
# +
from birdy import WPSClient
from example_data import TESTDATA
import datetime as dt
from urllib.request import urlretrieve
import xarray as xr
import numpy as np
from matplotlib import pyplot as plt
import os
# Set environment variable WPS_URL to "http://localhost:9099" to run on the default local server
url = os.environ.get("WPS_URL", "https://pavics.ouranos.ca/twitcher/ows/proxy/raven/wps")
wps = WPSClient(url)
# +
# The model parameters. Can either be a string of comma separated values, a list, an array or a named tuple.
# Mohyse also requires HRU parameters
params = '1.00, 0.0468, 4.2952, 2.6580, 0.4038, 0.0621, 0.0273, 0.0453'
hrus = '0.9039, 5.6179775'
# Forcing files
ts=TESTDATA['raven-mohyse-nc-ts']
# Model configuration parameters
config = dict(
start_date=dt.datetime(2000, 1, 1),
end_date=dt.datetime(2002, 1, 1),
area=4250.6,
elevation=843.0,
latitude=54.4848,
longitude=-123.3659,
)
# Let's call the model
resp = wps.raven_mohyse(ts=str(ts), params = params, hrus=hrus, **config)
# And get the response
# With `asobj` set to False, only the reference to the output is returned in the response.
# Setting `asobj` to True will retrieve the actual files and copy the locally.
[hydrograph, storage, solution, diagnostics, rv] = resp.get(asobj=True)
# -
# Since we requested output objects, we can simply access the output objects. The dianostics is just a CSV file:
print(diagnostics)
# The `hydrograph` and `storage` outputs are netCDF files storing the time series. These files are opened by default using `xarray`, which provides convenient and powerful time series analysis and plotting tools.
hydrograph.q_sim
# +
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
hydrograph.q_sim.plot()
# -
print("Max: ", hydrograph.q_sim.max())
print("Mean: ", hydrograph.q_sim.mean())
print("Monthly means: ", hydrograph.q_sim.groupby(hydrograph.time.dt.month).mean(dim='time'))
| docs/source/notebooks/running_mohyse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Load PyEpiDAG
import epidag as dag
import numpy as np
# ## Define a DAG
# Compose a script
#
# ```
# PCore Exp1 {
# # Definitions of nodes
# }
# ```
#
# Then, .. parse to a DAG
script = '''
PCore Exp1 {
n
a = 0.5
b ~ beta(1, 1)
c = min(a, b)
y ~ binom(n, c)
z = f(a,b)
}
'''
js = dag.bn_script_to_json(script)
bn = dag.BayesianNetwork(js)
bn
# ### Single value variable
#
#
# > VariableName = Number
#
# +
SingleValue = bn['a']
print('Node \'a\'')
print('Definition:', SingleValue)
print('\nFind the value')
print(SingleValue())
# -
# ### Exogenous variable
#
# > VariableName
# +
Exogenous = bn['n']
print('Node \'n\'')
print('Definition:', Exogenous)
print('\nFind the value; must append external resources')
print(Exogenous({'n': 5}))
# -
# ### Random variable
#
# > VariableName ~ p(...)
#
# ** p(...) **: a probabilidy density/mass function
# +
Random = bn['b']
print('Node \'b\'')
print('Definition:', Random)
print('\nSample a value')
print(Random())
# -
# ### Equation output
#
# > OutputName = g(...)
#
# ** g(...) **: a mathematical function
# +
Equation = bn['c']
print('Node \'c\'')
print('Definition:', Equation)
parents = {
'a': SingleValue(),
'b': Random()
}
print('\nCalculate a value')
print(Equation(parents))
# -
# ### Pseudo Node
# Pseudo nodes are the nodes which can not be implemented in simulation but in relation inference.
#
# > VariableName = f(...)
#
# ** f(...) **: a pseudo function start with ** f ** and a list of parent variables follows
# +
Random = bn['b']
print('Node \'b\'')
print('Definition:', Random)
print('\nSample a value')
print(Random())
| docs/Tutorial 1. Basic syntax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: keras_tf
# language: python
# name: keras_tf
# ---
#Source : <NAME> Blog
import numpy as np
# +
#input data
input_data = np.array([[0,0,1],[0,1,1],[1,0,1],[1,1,1]])
output_labels = np.array([[0],[1],[1],[0]])
print(input_data)
print(output_labels)
# +
#sigmoid function
def activate(x,deriv=False):
if(deriv==True):
return x*(1-x)
return 1/(1+np.exp(-x))
# +
# weight values
synaptic_weight_0 = 2 * np.random.random((3,4))-1
synaptic_weight_1 = 2 * np.random.random((4,1))-1
print(synaptic_weight_0)
print(synaptic_weight_1)
# +
for j in xrange(60000):
# Forward propagate through layers 0, 1, and 2
layer0 = input_data
layer1 = activate(np.dot(layer0,synaptic_weight_0))
layer2 = activate(np.dot(layer1,synaptic_weight_1))
#calculate error for layer 2
layer2_error = output_labels - layer2
if (j% 10000) == 0:
print "Error:" + str(np.mean(np.abs(layer2_error)))
#Use it to compute the gradient
layer2_gradient = layer2_error*activate(layer2,deriv=True)
#calculate error for layer 1
layer1_error = layer2_gradient.dot(synaptic_weight_1.T)
#Use it to compute its gradient
layer1_gradient = layer1_error * activate(layer1,deriv=True)
#update the weights using the gradients
synaptic_weight_1 += layer1.T.dot(layer2_gradient)
synaptic_weight_0 += layer0.T.dot(layer1_gradient)
# -
#testing
print(activate(np.dot(np.array([0, 1, 1]), synaptic_weight_0 )))
| Neural Net.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Demo 3: Reactive Transport
# This notebook contains a series of problems that demonstrate how to set up and run reactive transport simulations with Amanzi-ATS. It is organized in 3 subproblems that demonstrate the following processes of increasing complexity:
#
# 1. Transport of a solute dissolved in surface water.
# 2. Reactive transport of 2 solutes in the subsurface.
# 3. Integrated flow and reactive transport.
#
# The surface transport problem uses a dilution test as an example for setting solute sources and demonstrating key physical transport processes. The subsurface transport problem demonstrates how to set boundary conditions for transport and adds sorption to introduce reactions. The simulation of integrated flow and reactive transport in a hillslope .
#
# This notebook describes the problems, visualizes the results and guides the attendees to tinker with various parameters and analyze model response. The attendees should have following three directories with respective model files in them. <br>
#
# 1) surface_transport <br>
# 2) subsurface_transport <br>
# 3) hillslope <br>
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import h5py
from IPython.display import Image
# ## 1) Surface Transport
# ### 1.1) Problem Description
# The surface transport is demonstrated using a simple 1D transport in a 100 m long stream section with 3 $m^2$ cross section area. Dilution test, a common technique to measure discharge ($Q$) in the field is used as an example. In this test, solute solution of known solute concentration ($C_{in}$) is injected into the stream at a known steady rate ($Q_{in}$) and concentration time series ($C_{obs}$) also known as breakthrough curve (BTC) is measured at a downstream location. Figure below shows the schematics of the problem. <br>
#
# <img src="./pictures/surface_problem.jpg" alt="Drawing" style="width: 1000px;"/>
#
# The expected plateau concentration is calculated based on the mass conservation of the solute (see figure). The arrival time of the solute front at the measurement location is $\frac{100m}{\frac{0.03m^3/s}{3m^2}} = 10,000 s$
#
# The key features of this demo:
#
# 1) 2D mesh for surface transport <br>
# 2) Solute injection using PFLOTRAN <br>
# 3) Advection-dispersion <br>
# 4) Decay (special case) <br>
# ### 1.2) Input Specs
# Here some of the transport specific input specifications and respective parameter lists are described. The directory should have following files in the <I>surface_transport</I> directory:
#
# * surface_transport.xml (ATS input file)
# * surface_transport.in (PFLOTRAN input file)
# * hanford.dat (Database)
#
# ATS inputs for key quantities are shown below (in ATS units):
#
# <img src="./pictures/surface_problem_ATS.jpg" alt="Drawing" style="width: 800px;"/>
#
# Solute injection related inputs:
#
# * <b>Geochemical Engine</b>: To specify the geochemical engine to be used. Here we use PFLOTRAN. <br>
#
# ```xml
# <ParameterList name="surface chemistry" type="ParameterList">
# <Parameter name="PK type" type="string" value="chemistry alquimia" />
# <Parameter name="engine" type="string" value="PFloTran" />
# ```
#
# * <b>Injection rate</b>: This is provided in the form of flow rate per unit volume of injection domain, in the parameterlist <b><i>surface-water_source</i></b> <br>
#
# <img src="./pictures/surface-water_source.png" alt="Drawing" style="width: 500px;"/>
#
# * <b> Solute concentration in the injection </b>: ATS is informed that this concentration is specified through geochemical engine. <br>
#
# <img src="./pictures/geochemical_condition_surface.png" alt="Drawing" style="width: 500px;"/>
#
# And the value of concetration is provided in the PFLOTRAN input file as molarity in the <b><i>CONSTRAINT</i></b> deck <br>
#
# <img src="./pictures/inj_conc_surface.png" alt="Drawing" style="width: 100px;"/>
#
# Transport processes related inputs:
#
# * <b>Advection</b>: Flowrate in the domain (Q): This is provided in the form of flux (flowrate per unit cross-section area) in the parameterlist <b><i>surface-mass_flux</i></b> <br>
#
# <img src="./pictures/surface-mass_flux.png" alt="Drawing" style="width: 500px;"/>
#
# * <b>Dispersion</b>: Dispersion coefficient: This is provided in the parameterlist <b><i>material properties</i></b>
#
# <img src="./pictures/surface_dispersion.png" alt="Drawing" style="width: 400px;"/>
# ### 1.3) Result Visualization
# Here, we are visualizing results in the form of concentration timeseries or breakthrough curve (BTC) at a given location. This is done through <b><i>observations</i></b> parameterlist. The BTCs are written in ATS in a ".dat" file in mol-species/mol-water.
#
# <img src="./pictures/surface_observation.png" alt="Drawing" style="width: 500px;"/>
# ATS raw output unit: mol-species/mol-water in surface-outlet.dat
# Convert to mole-species/m^3-water
unit_convert_transport = 55500
plt.rcParams['figure.figsize'] = [10, 5]
path='./surface_problem/'
BTC_1 = np.loadtxt(path+'surface-outlet.dat', skiprows=11) # surface-outlet.dat is the BTC file
plt.plot(BTC_1[:,0], BTC_1[:,1]*unit_convert_transport, 'k-', label='Tracer1')
plt.title("Breakthrough Curve at 100 m from Injection", fontsize=20)
plt.xlabel("Time (s)",fontsize=16)
plt.ylabel("Concentration ($mol_C/m^3_{H2O}$)",fontsize=16)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.legend(fontsize=12, loc='lower right')
plt.show()
# ### 1.4) Exercise
# Attendees (during the session or at "home") should tinker with the injection and transport parameters and see the effects on the BTC. Some suggestions:
# 1) Double the value of <b><i>surface-mass_flux</i></b> and solute arrival should be at 5000s. However, this also increases the dilution and hence the plateau concetration should also be halved. <br>
# 2) Try higher values of dispersion coefficient <br>
# 3) Change concentration of injected solute and see the change in the plateau concentration <br>
# 4) Introduce decay of solute by using non zero reaction coefficient value in the PFLOTRAN input file <br>
# ## 2) Subsurface Transport
# ### 2.1) Problem Description
# The subsurface transport is demonstrated using a 1D reactive transport in a 100m long domain with 2m width and 1.5m depth (3D mesh: 200-1-1) with a specified concentration boundary. The flux ($q$) in the horizontal direction is specific along with dispersivities (in bear model), porosity ($\theta$), permeability ($k$) and sorption parameters ($K_d$). The reactant (Tracer1) in this example entering the domain through the specificied concentration boundary is undergoing first order decay into a product (Tracer2). Both the componenets are also undergoing equillibrium sorption in the domain.
#
# <img src="./pictures/subsurface_problem.jpg" alt="Drawing" style="width: 800px;"/>
#
# The arrival time of the solute front at the measurement location in no sorption case is $\frac{100m}{\frac{0.0025m/s}{0.25}} = 10,000 s$
#
#
# The key features of this demo:
#
# 1) 3D mesh for surface transport <br>
# 2) Specific concetration boundary <br>
# 3) Advection-dispersion-sorption <br>
# 4) First order reaction
#
# ### 2.2) Input Specs
# Here some of the transport specific input specifications and respective parameter lists are described. The directory should have following files in the <I>surface_transport</I> directory:
#
# * subsurface_transport.xml (ATS input file)
# * subsurface_transport.in (PFLOTRAN input file)
# * hanford.dat (Database)
#
# Boundary:
#
# * <b>Concentration boundary</b>: This is provided as geochemical condition in the parameterlist <b><i>boundary conditions</i></b> <br>
#
# <img src="./pictures/subsurface_boundary.png" alt="Drawing" style="width: 500px;"/>
#
# * <b> Concentration value </b>: This is provided in the PFLOTRAN input file as molarity in the <b><i>CONSTRAINT</i></b> deck <br>
#
# <img src="./pictures/subsurface_bc_conc.png" alt="Drawing" style="width: 100px;"/>
#
# Transport processes related inputs:
#
# * <b>Advection</b>: This is provided in the form of flux (flowrate per unit cross-section area, q) in the parameterlist <b><i>mass_flux</i></b> <br>
#
# <img src="./pictures/mass_flux.png" alt="Drawing" style="width: 500px;"/>
#
# * <b>Dispersion</b>: Dispersivity: Bear model is used to compute dispersion using dispersivity value in the parameterlist <b><i>material properties</i></b>
#
# <img src="./pictures/dispersivity_bear.png" alt="Drawing" style="width: 400px;"/>
#
# Reaction related inputs:
# * <b>Equillibrium Sorption</b>: Equilibrium sorption is enabled in ATS using <b><i>isotherm_kd</i></b> parameterlist
# * <b>Equillibrium Sorption Coefficient</b>: Equilibrium sorption coefficient Kd ($kg-water / m^3-bulk$) is provided in PFLOTRAN input file under <b><i>SORPTION</i></b> deck,
#
# <img src="./pictures/sorption_deck.png" alt="Drawing" style="width: 300px;"/>
#
# causing retardation by a factor as calculated below
#
# <img src="./pictures/sorption.png" alt="Drawing" style="width: 350px;"/>
#
# * <b> Reaction Coefficient </b>: The rate constant of first order conversion from Tracer1 into Tracer2 is provided in the PFLOTRAN input file under <b><i>GENERAL_REACTION</i></b> deck <br>
# ### 2.3) Result Visualization
# Here, we are visualizing results in the form of concentration timeseries or breakthrough curve (BTC) at a given location. This is done through observations parameterlist. The BTC is written in ATS in a ".dat" file. ATS yields outlet.dat file containing aqueous species and outlet_sorbed.dat. file contained sorbed species. Below, only aqeuous concentrations are showed.
plt.rcParams['figure.figsize'] = [10, 5]
path='./subsurface_problem/'
BTC_1 = np.loadtxt(path+'outlet.dat', skiprows=11) # outlet.dat is the BTC file for Aqueous species
plt.plot(BTC_1[:,0], BTC_1[:,1]*unit_convert_transport, 'k-', label='Tracer1')
plt.plot(BTC_1[:,0], BTC_1[:,2]*unit_convert_transport, 'r-', label='Tracer2')
plt.title("Breakthrough Curve at 100 m from Injection", fontsize=20)
plt.xlabel("Time (s)",fontsize=16)
plt.ylabel("Concentration ($mol_C/m^3_{H2O}$)",fontsize=16)
plt.xticks(fontsize=16)
plt.yticks(fontsize=16)
plt.legend(fontsize=12, loc='lower right')
plt.show()
# ### 2.4) Exercise
# Attendees (during the session or at "home") should tinker with transport and reaction parameters and see the effects on the BTCs. Some of the suggestions:
#
# 1) Set sorption coefficient Kd values equal to 100 for both the species in PFLOTRAN input file. This should increase the arrival time of solute fronts by the factor of retardation factor (1.4 for this case) <br>
# 2) Change reaction rate constant in PFLOTRAN input file and see the changes in the plateau concentrations of Tracer1 and Tracer2 <br>
# 3) For a non-zero sorption case, replace General Reaction deck in PFLOTRAN input file:
#
# <img src="./pictures/general_rxn.png" alt="Drawing" style="width: 250px;"/>
#
# by Radioactive Reaction deck:
#
# <img src="./pictures/radioactive_rxn.png" alt="Drawing" style="width: 250px;"/>
#
#
# The difference you see is because general reaction takes place only in the aqueous phase while radioactive decay takes place in both aqueous and sorbed phases
| 03_reactive_transport/.ipynb_checkpoints/reactive_transport_training_demo-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #!pip install keras
# #!pip python -m pip install --upgrade pip
import tensorflow as tt
from keras.datasets import fashion_mnist
(train_X,train_Y), (test_X,test_Y) = fashion_mnist.load_data()
import tensorflow as tf
# +
# #!pip install matplotlib
import numpy as np
from keras.utils import to_categorical
import matplotlib.pyplot as plt
# %matplotlib inline
print('Training data shape : ', train_X.shape, train_Y.shape)
print('Testing data shape : ', test_X.shape, test_Y.shape)
# -
#Find the unique numbers from the train labels
classes = np.unique(train_Y)
nClasses = len(classes)
print('Total number of outputs : ', nClasses)
print('Output classes : ', classes)
# +
plt.figure(figsize=[5,5])
# Display the first image in training data
plt.subplot(121)
plt.imshow(train_X[0,:,:], cmap='gray')
plt.title("Ground Truth : {}".format(train_Y[0]))
# Display the first image in testing data
plt.subplot(122)
plt.imshow(test_X[0,:,:], cmap='gray')
plt.title("Ground Truth : {}".format(test_Y[0]))
# -
train_X = train_X.reshape(-1, 28,28, 1)
test_X = test_X.reshape(-1, 28,28, 1)
train_X.shape, test_X.shape
train_X = train_X.astype('float32')
test_X = test_X.astype('float32')
train_X = train_X / 255.
test_X = test_X / 255.
# +
# Change the labels from categorical to one-hot encoding
train_Y_one_hot = to_categorical(train_Y)
test_Y_one_hot = to_categorical(test_Y)
# Display the change for category label using one-hot encoding
print('Original label:', train_Y[0])
print('After conversion to one-hot:', train_Y_one_hot[0])
# -
from sklearn.model_selection import train_test_split
train_X,valid_X,train_label,valid_label = train_test_split(train_X, train_Y_one_hot, test_size=0.2, random_state=13)
train_X.shape,valid_X.shape,train_label.shape,valid_label.shape
import keras
from keras.models import Sequential,Input,Model
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.layers.advanced_activations import LeakyReLU
# B) Activation = Sigmoid
# c) Cost Function = Cross Entropy
# D) Epochs = 20
# E) Gradient Estimation = Adam
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
batch_size = 64
epochs = 20
num_classes = 10
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
fashion_model.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_model.summary()
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout to fit model neatly
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Cross Entropy
# D) Epochs = 20
# E) Gradient Estimation = Adam
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss=keras.losses.categorical_crossentropy, optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Mean Square Error
# D) Epochs = 20
# E) Gradient Estimation = Adam
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss='mean_squared_error', optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss='mean_squared_error', optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Mean Square Error
# D) Epochs = 25
# E) Gradient Estimation = Adam
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
batch_size = 64
epochs = 25
num_classes = 10
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss='mean_squared_error', optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss='mean_squared_error', optimizer=keras.optimizers.Adam(),metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Mean Square Error
# D) Epochs = 20
# E) Gradient Estimation = SGD
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
batch_size = 64
epochs = 20
num_classes = 10
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss='mean_squared_error',y, optimizer='sgd',metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss='mean_squared_error', optimizer=keras. optimizer='sgd',metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Mean Square Error
# D) Epochs = 20
# E) Gradient Estimation = SGD
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = RandomNormal
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
#fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
#fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss='mean_squared_error',y, optimizer='sgd',metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= keras.initializers.RandomNormal(mean=0.0, stddev=0.05, seed=None)))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
#fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
#fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
#fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss='mean_squared_error', optimizer=keras. optimizer='sgd',metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
# B) Activation = linear
# c) Cost Function = Mean Square Error
# D) Epochs = 20
# E) Gradient Estimation = SGD
# F) Network Architecture = 9 Layers{3Conv2D + 3Maxpooling + 2Dense + 1Flatten}
# G) Network Initializer = Zeros
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='linear',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= kernel_initializer= keras.initializers.Zeros()))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Conv2D(64, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Conv2D(128, (3, 3), activation='linear',padding='same'))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='linear'))
fashion_model.compile(loss='mean_squared_error',y, optimizer='sgd',metrics=['accuracy'])
fashion_train = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=0)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# The model is overfitted by " " adding dropout layer to make it more fit
#adding pooling layer
fashion_model = Sequential()
fashion_model.add(Conv2D(32, kernel_size=(3, 3),activation='sigmoid',input_shape=(28,28,1),padding='same'))
fashion_model.add(Dense(128,kernel_initializer= kernel_initializer= keras.initializers.Zeros()))
fashion_model.add(MaxPooling2D((2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(64, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Conv2D(128, (3, 3), activation='sigmoid',padding='same'))
fashion_model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
fashion_model.add(Dropout(0.25))
fashion_model.add(Flatten())
#fashion_model.add(Dense(128, activation='sigmoid'))
#fashion_model.add(Dense(128, kernel_initializer='uniform', input_shape=(10,)))
#fashion_model.add(LeakyReLU(alpha=0.1))
fashion_model.add(Dense(num_classes, activation='sigmoid'))
# +
fashion_model.compile(loss='mean_squared_error', optimizer=keras. optimizer='sgd',metrics=['accuracy'])
fashion_train_dropout = fashion_model.fit(train_X, train_label, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(valid_X, valid_label))
test_eval = fashion_model.evaluate(test_X, test_Y_one_hot, verbose=1)
print('Test loss:', test_eval[0])
print('Test accuracy:', test_eval[1])
# -
accuracy = fashion_train.history['acc']
val_accuracy = fashion_train.history['val_acc']
loss = fashion_train.history['loss']
val_loss = fashion_train.history['val_loss']
epochs = range(len(accuracy))
plt.plot(epochs, accuracy, 'bo', label='Training accuracy')
plt.plot(epochs, val_accuracy, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
fashion_model.save("fashion_model_dropout.h5py")
accuracy = fashion_train_dropout.history['acc']
val_accuracy = fashion_train_dropout.history['val_acc']
loss = fashion_train_dropout.history['loss']
val_loss = fashion_train_dropout.history['val_loss']
epochs = range(len(accuracy))
plt.plot(epochs, accuracy, 'bo', label='Training accuracy')
plt.plot(epochs, val_accuracy, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
#Predict Labels
predicted_classes = fashion_model.predict(test_X)
predicted_classes = np.argmax(np.round(predicted_classes),axis=1)
predicted_classes.shape, test_Y.shape
correct = np.where(predicted_classes==test_Y)[0]
print( "Found %d correct labels"% len(correct))
for i, correct in enumerate(correct[:9]):
plt.subplot(3,3,i+1)
plt.imshow(test_X[correct].reshape(28,28), cmap='gray', interpolation='none')
plt.title("Predicted {}, Class {}".format(predicted_classes[correct], test_Y[correct]))
plt.tight_layout()
incorrect = np.where(predicted_classes!=test_Y)[0]
print ("Found %d incorrect labels" % len(incorrect))
for i, incorrect in enumerate(incorrect[:9]):
plt.subplot(3,3,i+1)
plt.imshow(test_X[incorrect].reshape(28,28), cmap='gray', interpolation='none')
plt.title("Predicted {}, Class {}".format(predicted_classes[incorrect], test_Y[incorrect]))
plt.tight_layout()
#Classification Report
from sklearn.metrics import classification_report
target_names = ["Class {}".format(i) for i in range(num_classes)]
print(classification_report(test_Y, predicted_classes, target_names=target_names))
| UntitledA4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0"
import numpy as np
import random
import tensorflow as tf
import pandas as pd
pd.set_option('display.max_columns', None)
seed_value = 0
os.environ['PYTHONHASHSEED']=str(seed_value)
random.seed(seed_value)
np.random.seed(seed_value)
tf.random.set_seed(seed_value)
# -
import ktrain
from ktrain import tabular
# # Classification and Regression on Tabular Data in `ktrain`
#
# As of v0.19.x, *ktrain* supports classification and regression on "traditional" tabular datasets. We will cover two examples in this notebook:
# - **Part I: Classification**: predicting which [Titanic passengers survived](https://www.kaggle.com/c/titanic)
# - **Part II: Regression**: predicting the age of people from [census data](http://archive.ics.uci.edu/ml/datasets/Census+Income)
#
# Let's begin with a demonstration of tabular classfication using the well-studied Titatnic dataset from Kaggle.
#
# ## Part I: Classification for Tabular Data
#
#
# ### Solving the Titanic Kaggle Challenge in `ktrain`
#
# This notebook demonstrates using *ktrain* for predicting which passengers survived the Titatnic shipwreck.
#
# The dataset can be [downloaded from Kaggle here](https://www.kaggle.com/c/titanic/overview). There is a `train.csv` with labels (i.e., `Survived`) and a `test.csv` with no labels. We will only use `train.csv` in this notebook.
#
# Let's begin by loading the data as a pandas DataFrame and inspecting it.
train_df = pd.read_csv('data/titanic/train.csv', index_col=0)
train_df.head()
# We'll drop the `Name`, `Ticket`, `Cabin` columns, as they seem like they'll be less predictive. These columns are largely unique or near-unique to passengers.
train_df = train_df.drop('Name', 1)
train_df = train_df.drop('Ticket', 1)
train_df = train_df.drop('Cabin', 1)
# *ktrain* will automatically split out a validation set if given only a training set. But, let's also manually split out a test set that we can evaluate later.
np.random.seed(42)
p = 0.1 # 10% for test set
prop = 1-p
df = train_df.copy()
msk = np.random.rand(len(df)) < prop
train_df = df[msk]
test_df = df[~msk]
train_df.shape
test_df.shape
# ### STEP 1: Load and Preprocess the Data
trn, val, preproc = tabular.tabular_from_df(train_df, label_columns=['Survived'], random_state=42)
# ##### Automated Preprocessing
# *ktrain* automatically preprocesses the dataset appropriately. Numerical columns are automatically normalized, missing values are handled, and categorical variables will be vectorized as [entity embeddings](https://arxiv.org/abs/1604.06737) for input to a neural network.
#
# ##### Auto-generated Features
# *ktrain* will auto-generate some new features. For instance, if `Age` is missing for a particular individual, an `Age_na=True` feature will be automatically added.
#
# New date features are also automatically added. This dataset does not have any **date** fields. If it did, we could populate the `date_columns` parameter to `tabular_from_df` in which case they would be used to auto-generate new features (e.g., `Day`, `Week`, `Is_month_start`, `Is_quarter_end`, etc.) using methods adapted from the **fastai** library.
#
# ##### Manually-Engineered Features
#
# In addition to these auto-generated features, one can also optionally add manually-generated, dataset-specific features to `train_df` **prior** to invoking `tabular_from_df`. For instance, the `Cabin` feature we discarded earlier might be used to extract the **deck** associated with each passenger (e.g., **B22** --> **Deck B**).
# ### STEP 2: Create a Model and Wrap in `Learner`
#
# *ktrain* uses multilayer perceptrons as the model for tabular datasets. The model can be configured with arguments to `tabular_classifier` (e.g., number and size of hidden layers, dropout values, etc.), but we will leave the defaults here.
tabular.print_tabular_classifiers()
model = tabular.tabular_classifier('mlp', trn)
learner = ktrain.get_learner(model, train_data=trn, val_data=val, batch_size=32)
# ### STEP 3: Estimate the Learning Rate
#
# Based on the plot below, we will choose a learning rate of `1e-3`.
learner.lr_find(show_plot=True, max_epochs=5)
# ### STEP 4: Train the Model
learner.fit_onecycle(5e-3, 10)
# Since we don't appear to be quite overfitting yet, we could try to train further. But, we will stop here.
#
#
# **Let's evaluate the validation set:**
learner.evaluate(val, class_names=preproc.get_classes())
# ### Make Predictions
#
# The `Predictor` for tabular datasets accepts input as a dataframe in the same format as the original training dataframe.
#
# We will use `test_df` that we created earlier.
predictor = ktrain.get_predictor(learner.model, preproc)
preds = predictor.predict(test_df, return_proba=True)
preds.shape
print('test accuracy:')
(np.argmax(preds, axis=1) == test_df['Survived'].values).sum()/test_df.shape[0]
# **Our final results as a DataFrame:**
df = test_df.copy()[[c for c in test_df.columns.values if c != 'Survived']]
df['Survived'] = test_df['Survived']
df['predicted_Survived'] = np.argmax(preds, axis=1)
df.head()
# ### Explaining Predictions
#
# We can use the `explain` method to better understand **why** a prediction was made for a particular example. Consider the passenger in the fourth row above (`PassengerID=35`) that did not survive. Although we classified this passenger correctly here, this row tends to get classified differently across different training runs. It is sometimes classified correctly (as in this run), but is also often misclassifeid.
#
# Let's better understand why.
#
# The `explain` method accepts at minimum the following three inputs:
# 1. **df**: a pandas DataFrame in the same format is the original training DataFrame
# 2. **row_index**: the DataFrame index of the example (here, we choose PassengerID=35)
# 3. **class_id**: the id of the class of interest (we choose the **Survived** class in this case)
#
# One can also replace the `row_index=35` with `row_num=3`, as both denote the fourth row.
predictor.explain(test_df, row_index=35, class_id=1)
# The plot above is generated using the [shap](https://github.com/slundberg/shap) library. You can install it with either `pip install shap` or, for *conda* users, `conda install -c conda-forge shap`. The features in red are causing our model to increase the prediction for the **Survived** class, while features in blue cause our model to *decrease* the prediction for **Survived** (or *increase* the prediction for **Not_Survived**).
#
# From the plot, we see that the predicted softmax probability for `Survived` is **50%**, which is a comparatively much less confident classification than other classifications. Why is this?
#
# We see that`Sex=male` is an influential feature that is pushing the prediction lower towards **Not_Survived**, as it was women and children given priority when allocating lifeboats on the Titanic.
#
# On the other hand, we also see that this is a First Class passenger (`Pclass=1`) with a higher-than-average `Fare` price of *82.17*. In the cell below, you'll see that the average `Fare` price is only *32*. (Moreover, this passenger embarked from Cherbourg, which has been shown to be correlated with survival.) Such features suggest that this is an upper-class, wealthier passenger and, therefore, more likely to make it onto a lifeboat and survive. We know from history that crew members were ordered to close gates that lead to the upper decks so the first and second class passengers could be evacuated first. As a result, these "upper class" features are pushing our model to increase the classification to **Survived**.
#
# **Thus, there are two opposing forces at play working against each other in this prediction,** which explains why the prediction probability is comparatively nearer to the border than other examples.
#
#
train_df['Fare'].mean()
# **NOTE**: We choose `class_id=1` in the example above because the **Survived** class of interest has an index position of 1 in the `class_names` list:
preproc.get_classes()
# Let us now look at the examples for which we were the most wrong (highest loss).
learner.view_top_losses(val_data=preproc.preprocess_test(test_df), preproc=preproc, n=3)
# The example with the highest losses are `row_num={27, 53, 19}`. Why did we get these so wrong? Let's examine `row_num=53`. Note that these IDs shown in the `view_top_losses` output are the raw row numbers, not DataFrame indices (or PassengerIDs). So, we need to use `row_num`, not `row_index` here.
#
predictor.explain(test_df, row_num=53, class_id=1)
# This is a wealthy First Class (`Pclass=1`) female passenger with a very high `Fare` price of 151.55. As mentioned above, such a passenger had a high chance for survival, which explains our model's high prediction for **Survival**. Yet, she did not survive. Upon further investigation, we can understand why. This particular passenger is **<NAME>**, a wealthy married 25-year old mother to two toddlers. When the collision occurred, her and her husband could not locate their nanny (<NAME>) and son (Trevor). So, Bess, her husband, and her 3-year-old daughter Loraine stayed behind to wait for them instead of evacuating with other First and Second Class passengers with children. They were last seen standing together smiling on the promenade deck. All three died with her daughter Loraine being the only child in 1st class and 2nd class who died on the Titanic. Their son and nanny successfully evacuated and survived.
#
# REFERENCE: [https://rt.cto.mil/stpe/](https://rt.cto.mil/stpe/)
# ### Saving and Reloading the Tabular Predictor
#
# It is easy to save and reload the predictor for deployment scenarios.
predictor.save('/tmp/titanic_predictor')
reloaded_predictor = ktrain.load_predictor('/tmp/titanic_predictor/')
reloaded_predictor.predict(test_df)[:5]
# ### Evaluating Test Sets Automatically
#
# When we evaulated the test set above, we did so manually. To evaluate a test set automatically,
# one can invoke the `learner.evaluate` method and supply a preprocessed test set as an argument:
learner.evaluate(preproc.preprocess_test(test_df), class_names=preproc.get_classes())
# The `learner.evaluate` method is simply an alias to `learner.validate`, which can also accept a dataset as an argument. If no argument is supplied, metrics will be computed for `learner.val_data`, which was supplied to `get_learner` above.
# ## Part II: Regression for Tabular Data
#
# We will briefly demonstrate tabular regression in *ktrain* by simply predicting the `age` attribute in the Census dataset available from te UCI Machine Learning repository. This is the same example used in the [AutoGluon regression example](https://autogluon.mxnet.io/tutorials/tabular_prediction/tabular-quickstart.html#regression-predicting-numeric-table-columns). Let's begin by downloading the dataset from the AutoGluon website.
import urllib.request
urllib.request.urlretrieve('https://autogluon.s3.amazonaws.com/datasets/Inc/train.csv',
'/tmp/train.csv')
# !ls /tmp/train.csv
# ### STEP 1: Load and Preprocess Data
#
# Make sure you specify `is_regression=True` here as we are predicting a numerical dependent variable (i.e., `age`).
trn, val, preproc = tabular.tabular_from_csv('/tmp/train.csv', label_columns='age',
is_regression=True, random_state=42)
# We used `tabular_from_csv` to load the dataset, but let's also quickly load as DataFrame to see it:
pd.read_csv('/tmp/train.csv').head()
# ### STEP 2: Create a Model and Wrap in `Learner`
#
# We'll use `tabular_regression_model` to create a regression model.
tabular.print_tabular_regression_models()
model = tabular.tabular_regression_model('mlp', trn)
learner = ktrain.get_learner(model, train_data=trn, val_data=val, batch_size=128)
# ### STEP 3: Estimate Learning Rate
learner.lr_find(show_plot=True)
# ### STEP 4: Train the Model
#
# According to our final validation MAE (see below), our age predictions are only off about **~7 years**.
learner.autofit(1e-3)
learner.validate()
# See the [House Price Prediction notebook](https://github.com/amaiya/ktrain/blob/master/examples/tabular/HousePricePrediction-MLP.ipynb) for another regression example.
| tutorials/tutorial-08-tabular_classification_and_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Initialization
from src.page_rank import page_rank
# ## Calculating pagerank dataframe
# Loading data and calculating the page ranks and showing the top results:
page_rank('./outputs//result.json', alpha=0.1, conv_limit=1e-12).head(20)
| pagerank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="cmEs3WWyhC4l" colab_type="text"
# # Resumo da Aula de Validação de Modelos
# + id="sAD6WK1-gdDH" colab_type="code" outputId="a3318fb7-eb82-4290-9c2a-a58b3729df9a" colab={"base_uri": "https://localhost:8080/", "height": 462}
# !pip install graphviz==0.9
# !pip install pydot
# !pip install seaborn==0.9.0
# !apt-get install grapviz
# + id="y7ik04NlDZMA" colab_type="code" outputId="42572214-d922-4378-cf93-035913147397" colab={"base_uri": "https://localhost:8080/", "height": 204}
import pandas as pd
uri = "https://gist.githubusercontent.com/guilhermesilveira/e99a526b2e7ccc6c3b70f53db43a87d2/raw/1605fc74aa778066bf2e6695e24d53cf65f2f447/machine-learning-carros-simulacao.csv"
dados = pd.read_csv(uri).drop(columns=["Unnamed: 0"], axis=1)
dados.head()
# + id="b52E4e1hjegN" colab_type="code" outputId="5f2a23fd-5f43-4a8d-85ad-a7b2d4f6f6ba" colab={"base_uri": "https://localhost:8080/", "height": 204}
# situação horrível de "azar" onde as classes estão ordenadas por padrão
dados_azar = dados.sort_values("vendido", ascending=True)
x_azar = dados_azar[["preco", "idade_do_modelo","km_por_ano"]]
y_azar = dados_azar["vendido"]
dados_azar.head()
# + id="ZNT26vFEeYmz" colab_type="code" outputId="29a8b7c8-ff63-49e6-8f49-ffe044772587" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.model_selection import cross_validate
from sklearn.dummy import DummyClassifier
import numpy as np
SEED = 301
np.random.seed(SEED)
modelo = DummyClassifier()
results = cross_validate(modelo, x_azar, y_azar, cv = 10, return_train_score=False)
media = results['test_score'].mean()
desvio_padrao = results['test_score'].std()
print("Accuracy com dummy stratified, 10 = [%.2f, %.2f]" % ((media - 2 * desvio_padrao)*100, (media + 2 * desvio_padrao) * 100))
# + id="73mCcFA_eG_K" colab_type="code" outputId="e3fd7d1e-9863-4aa2-af50-6fcd3a6c02e9" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.model_selection import cross_validate
from sklearn.tree import DecisionTreeClassifier
SEED = 301
np.random.seed(SEED)
modelo = DecisionTreeClassifier(max_depth=2)
results = cross_validate(modelo, x_azar, y_azar, cv = 10, return_train_score=False)
media = results['test_score'].mean()
desvio_padrao = results['test_score'].std()
print("Accuracy com cross validation, 10 = [%.2f, %.2f]" % ((media - 2 * desvio_padrao)*100, (media + 2 * desvio_padrao) * 100))
# + id="5C8Y6J-PGpYf" colab_type="code" outputId="b2d40790-3118-43c9-f455-dc469ba7a14e" colab={"base_uri": "https://localhost:8080/", "height": 204}
# gerando dados elatorios de modelo de carro para simulacao de agrupamento ao usar nosso estimador
np.random.seed(SEED)
dados['modelo'] = dados.idade_do_modelo + np.random.randint(-2, 3, size=10000)
dados.modelo = dados.modelo + abs(dados.modelo.min()) + 1
dados.head()
# + id="3hmjt7qPHOZY" colab_type="code" colab={}
def imprime_resultados(results):
media = results['test_score'].mean() * 100
desvio = results['test_score'].std() * 100
print("Accuracy médio %.2f" % media)
print("Intervalo [%.2f, %.2f]" % (media - 2 * desvio, media + 2 * desvio))
# + id="goijy0rSS7n-" colab_type="code" outputId="60af3f3f-7b80-4370-be22-56a24f611deb" colab={"base_uri": "https://localhost:8080/", "height": 51}
# GroupKFold em um pipeline com StandardScaler e SVC
from sklearn.model_selection import GroupKFold
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVC
from sklearn.pipeline import Pipeline
SEED = 301
np.random.seed(SEED)
scaler = StandardScaler()
modelo = SVC()
pipeline = Pipeline([('transformacao',scaler), ('estimador',modelo)])
cv = GroupKFold(n_splits = 10)
results = cross_validate(pipeline, x_azar, y_azar, cv = cv, groups = dados.modelo, return_train_score=False)
imprime_resultados(results)
# + [markdown] id="XQzKvBZdg560" colab_type="text"
# # Plotando uma Árvore de Decisão
# + id="IKD8uaWkNcUL" colab_type="code" outputId="7e1233c9-8bd3-40ed-e9e9-a6d53742818c" colab={"base_uri": "https://localhost:8080/"}
# GroupKFold para analisar como o modelo se comporta com novos grupos
from sklearn.model_selection import GroupKFold
SEED = 301
np.random.seed(SEED)
cv = GroupKFold(n_splits = 10)
modelo = DecisionTreeClassifier(max_depth=2) # A profundidade máxima de uma árvore é um Hiperparâmetro
results = cross_validate(modelo, x_azar, y_azar, cv = cv, groups = dados.modelo, return_train_score=False)
imprime_resultados(results)
# + id="7NLFnnb7aAQk" colab_type="code" outputId="8069dce9-54f1-4f55-b90a-98cba163d220" colab={"base_uri": "https://localhost:8080/"}
from sklearn.tree import export_graphviz
import graphviz
# Na validação cruzada, nós treinamos o algoritmo 10 vezes para termos uma estimativa de quão bem esse modelo funcionaria no mundo real.
# Agora queremos o modelo propriamente dito para utilizarmos na vida real.
modelo.fit(x_azar,y_azar)
features = x_azar.columns
dot_data = export_graphviz(modelo, out_file=None, filled=True, rounded=True,
class_names=["não", "sim"],
feature_names = features)
graph = graphviz.Source(dot_data)
graph
# Parâmetros que são definidos antes do treino são chamados de HIPERPARÂMETROS,
# e são diferentes de valores internos do modelo que vão sendo alterados de acordo com o que o modelo está aprendendo.
# + [markdown] id="tbXBAzodigTl" colab_type="text"
# # Testando Hiperparâmetros - 1 Dimensão
# **Profundidade Máxima da Árvore**
# + id="PycSTmcehbH0" colab_type="code" outputId="9619ae5e-db43-4617-b7d6-3b6bf54f74a3" colab={"base_uri": "https://localhost:8080/", "height": 612}
# função para rodar diversos tamanhos de arvores
def roda_arvore_decisao(max_depth):
SEED = 301
np.random.seed(SEED)
cv = GroupKFold(n_splits = 10)
modelo = DecisionTreeClassifier(max_depth = max_depth)
results = cross_validate(modelo, x_azar, y_azar, cv = cv, groups = dados.modelo, return_train_score=True) # habilitar para retornar os resultados do treino
resultado_treino = results['train_score'].mean() * 100
resultado_teste = results['test_score'].mean() * 100
print('Árvore max_depth: %d - Média Treino: %.2f - Média Teste: %.2f' % (max_depth, resultado_treino, resultado_teste))
tabela = [max_depth, resultado_treino, resultado_teste]
return tabela
resultados = [roda_arvore_decisao(i) for i in range (1, 25)] # faz um loop para chamar a função i vezes, e retornar uma lista com a tabela
resultados = pd.DataFrame(resultados, columns = ['Max_Depth','Treino','Teste']) # transforma os resultados em um DF
resultados.head()
# + id="AaXqf1SVjuXM" colab_type="code" outputId="94310131-1f16-4f07-f631-5494ff5e2cb2" colab={"base_uri": "https://localhost:8080/", "height": 301}
# importa bibliotecas para poder trabalhar com gráficos
import seaborn as sns
import matplotlib.pyplot as plt
sns.lineplot(x = 'Max_Depth', y='Treino', data = resultados)
sns.lineplot(x = 'Max_Depth', y='Teste', data = resultados)
plt.legend(['Treino','Teste'])
# Observa-se que a partir da profundidade 3 ocorre o OVERFIT, ou seja,
# os resultados de treino continuam melhorando enquanto os de teste pioram, nao possuindo a capacidade de generalização
# + id="eiuSf9WMl1Or" colab_type="code" outputId="3cae232c-ba13-44f9-d70f-06e417cb9b3f" colab={"base_uri": "https://localhost:8080/", "height": 204}
resultados.sort_values('Teste', ascending = False).head() # ordena em ordem decrescente para descobrir os melhores testes
# + [markdown] id="iQ-Ha0Wu6YSa" colab_type="text"
# # Testando Hiperparâmetros - 2 Dimensões
# **Profundidade Máxima da Árvore e Número Mínimo de Elementos em uma Folha**
# + id="Fw7R6QyprRWV" colab_type="code" colab={}
# função para retornar os resultados de treino e teste de 2 hiperparâmetros
def roda_arvore_decisao(max_depth, min_samples_leaf):
SEED = 301
np.random.seed(SEED)
cv = GroupKFold(n_splits = 10)
modelo = DecisionTreeClassifier(max_depth = max_depth, min_samples_leaf = min_samples_leaf) # acrescenta o hiperparâmetro min_samples_leaf
results = cross_validate(modelo, x_azar, y_azar, cv = cv, groups = dados.modelo, return_train_score=True)
resultado_treino = results['train_score'].mean() * 100
resultado_teste = results['test_score'].mean() * 100
print('Árvore Max_depth: %d - Min_Samples_Leaf: %.2f - Média Treino: %.2f - Média Teste: %.2f' % (max_depth, min_samples_leaf , resultado_treino, resultado_teste))
tabela = [max_depth, min_samples_leaf , resultado_treino, resultado_teste]
return tabela
# + id="FWNvRZ3S9hwC" colab_type="code" outputId="540f0130-b7a1-4062-f0a9-52d7d89494c4" colab={"base_uri": "https://localhost:8080/"}
# função que vai percorrer a arvore com diversos valores dos 2 hiperparâmetros e transformar os resultados em um DF
def busca():
resultados = []
for max_depth in range(1, 25):
for min_samples_leaf in [32, 64, 128, 256]:
tabela = roda_arvore_decisao(max_depth, min_samples_leaf)
resultados.append(tabela)
resultados = pd.DataFrame(resultados, columns = ['Max_Depth','Min_Samples_Leaf','Treino','Teste'])
return resultados
resultados = busca()
resultados.head()
# + id="N5SFa0F_-ewW" colab_type="code" outputId="d4d24108-53be-499f-ba84-2f1219100fa5" colab={"base_uri": "https://localhost:8080/"}
resultados.sort_values('Teste', ascending = False).head() # ordena em ordem decrescente para descobrir os melhores testes
# + [markdown] id="wiScijD3Hes2" colab_type="text"
# # Testando a Correlação
# + id="d5LKiFPY_7pp" colab_type="code" outputId="2786c768-8816-4037-def3-e8cfef7557c0" colab={"base_uri": "https://localhost:8080/", "height": 173}
correlacao_resultados = resultados.corr() # método do pandas que mostra a correlação dos dados, lembrando que correlação != de causualidade
correlacao_resultados.head()
# Existem inúmeras maneiras de visualisação dos resultados, escolher a que melhor se adequa a situação
# Através da algumas visualisações é possivel perceber que:
# quando max_depth cresce, o treino parece crescer também
# quando min_samples_leaf cresce, o treino cai
# quando max_depth sobe, o teste cai
# quando min_samples_leaf sobe, o teste sobe
# A partir dos resultados e da verificação da correlação altera-se os intervalos dos hiperparametros para tentar otimizar o estimador
# + id="q5HLa6F-CZna" colab_type="code" outputId="88f24cbb-f63e-413f-e579-07ea2fb3ebfe" colab={"base_uri": "https://localhost:8080/", "height": 368}
sns.heatmap(correlacao_resultados) # mapa de calor da correlação
# + id="ukY1Nn8vDWsz" colab_type="code" outputId="ed7f0195-0737-450c-c292-e01f32726a22" colab={"base_uri": "https://localhost:8080/", "height": 746}
sns.pairplot(resultados) # realiza o pareamento dos resultados, na diagonal estão os histogramas dos valores
# + id="-VCrHAwLH6an" colab_type="code" outputId="295e58b6-4657-4c55-ae56-7410eef627be" colab={"base_uri": "https://localhost:8080/", "height": 564}
# Por último, gera-se outro gráfico que consta na própria documentação do Seaborn Correlations.
# Copiei e removi apenas os trechos em que os dados são gerados e atribuídos à uma variável
sns.set(style="white")
# Generate a mask for the upper triangle
mask = np.zeros_like(correlacao_resultados, dtype=np.bool) # substituir pela variavel de correlacao_resultados
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(correlacao_resultados, mask=mask, cmap=cmap, vmax=.3, center=0, # substituir pela variavel de correlacao_resultados
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# + [markdown] id="hQmcjmvkMj5N" colab_type="text"
# # Testando Hiperparâmetros - 3 Dimensões
# **Profundidade Máxima da Árvore,
# Número Mínimo de Elementos em uma Folha e
# Número Mínimo de Elementos para Divisão**
# + id="mpBbBEILJLJZ" colab_type="code" colab={}
# função para retornar os resultados de treino e teste de 3 hiperparâmetros
def roda_arvore_decisao(max_depth, min_samples_leaf, min_samples_split):
SEED = 301
np.random.seed(SEED)
cv = GroupKFold(n_splits = 10)
modelo = DecisionTreeClassifier(max_depth = max_depth, min_samples_leaf = min_samples_leaf, min_samples_split = min_samples_split) # acrescenta o hiperparâmetro min_samples_split
results = cross_validate(modelo, x_azar, y_azar, cv = cv, groups = dados.modelo, return_train_score=True)
tempo_treino = results['fit_time'].mean() # extrair a média de tempo do treino
tempo_teste = results['score_time'].mean() # extrair a média de tempo do teste
resultado_treino = results['train_score'].mean() * 100
resultado_teste = results['test_score'].mean() * 100
# print('Árvore Max_depth: %d - Min_Samples_Leaf: %.2f - Min_Samples_Split: %d - Média Treino: %.2f - Média Teste: %.2f' % (max_depth, min_samples_leaf, min_samples_split , resultado_treino, resultado_teste))
# O print serve apenas para acompanhar enquanto roda, porém o codigo esta muito grande, não havendo necessidade para tal
tabela = [max_depth, min_samples_leaf, min_samples_split, resultado_treino, resultado_teste, tempo_treino, tempo_teste]
return tabela
# + id="40x9Al8ANQd5" colab_type="code" outputId="eafa93be-7276-48d4-dd31-620f82f6248f" colab={"base_uri": "https://localhost:8080/", "height": 204}
# função que vai percorrer a arvore com diversos valores dos 3 hiperparâmetros e transformar os resultados em um DF
def busca():
resultados = []
for max_depth in range(1, 25):
for min_samples_leaf in [32, 64, 128, 256]:
for min_samples_split in [32, 64, 128, 256]:
tabela = roda_arvore_decisao(max_depth, min_samples_leaf, min_samples_split)
resultados.append(tabela)
resultados = pd.DataFrame(resultados, columns = ['Max_Depth','Min_Samples_Leaf','Min_Samples_Split','Treino','Teste','Tempo_Treino','Tempo_Teste'])
return resultados
resultados = busca()
resultados.head()
# + id="2zpaGu69OVq5" colab_type="code" outputId="09fd3215-a48c-4e54-a1ab-8945bc8407a1" colab={"base_uri": "https://localhost:8080/", "height": 204}
correlacao_resultados = resultados.corr()
correlacao_resultados.head()
# + id="mAiiIGqPPb1-" colab_type="code" outputId="4dd0b74f-bac8-4b81-b349-5816906e6c6b" colab={"base_uri": "https://localhost:8080/", "height": 655}
sns.set(style="white")
# Generate a mask for the upper triangle
mask = np.zeros_like(correlacao_resultados, dtype=np.bool) # substituir pela variavel de correlacao_resultados
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(correlacao_resultados, mask=mask, cmap=cmap, vmax=.3, center=0, # substituir pela variavel de correlacao_resultados
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# + id="N4YYEEXIPdu3" colab_type="code" outputId="0f15a93a-f65d-4e8c-d2d0-f453f515ca70" colab={"base_uri": "https://localhost:8080/", "height": 204}
resultados.sort_values('Teste', ascending = False).head()
# Como tanto o tempo de treino quanto de teste são proximos uns dos outros, acabam não sendo tao relevantes neste exemplo
# Porem em situações na qual o tempo de processamento são mais elevedados, pode ser mais um critério de escolha de valores
# + [markdown] id="j8h13pTSUoyH" colab_type="text"
# # Busca de Hiperparâmetros com o GridSearchCV
# + id="3DlSV-tVR8DA" colab_type="code" outputId="6e73ba18-d6b0-480a-c57a-8c72d0272c1b" colab={"base_uri": "https://localhost:8080/", "height": 564}
from sklearn.model_selection import GridSearchCV # serve para buscar os melhores valores dos hiperparâmetros com validação cruzada
SEED = 301
np.random.seed(SEED)
# 4 dimensões
espaco_parametros = {
'max_depth' : [3, 5],
'min_samples_leaf' : [32,64,128],
'min_samples_split' : [32,64,128],
'criterion' : ['gini','entropy']
}
busca = GridSearchCV(DecisionTreeClassifier(),
espaco_parametros,
cv = GroupKFold(n_splits = 10))
busca.fit(x_azar, y_azar, groups = dados.modelo)
resultados = pd.DataFrame(busca.cv_results_) # o gridsearch possui um método que retorna os resultados
resultados.head()
# + id="CQP6RCURWv1K" colab_type="code" outputId="75b671bb-385d-41be-93fa-eba833aae61b" colab={"base_uri": "https://localhost:8080/", "height": 51}
# métodos do gridsearchcv que retornam respectivamente os melhores valores dos hiperparâmetros e seu resultado
print(busca.best_params_)
print(busca.best_score_ * 100)
# + id="6QhwseWTZcQJ" colab_type="code" outputId="8af36df4-7281-47b8-861d-769dc3550c46" colab={"base_uri": "https://localhost:8080/", "height": 119}
# método que retorna o melhor modelo
melhor = busca.best_estimator_
melhor
# + id="se7Qv8f6b8-X" colab_type="code" outputId="0a94b192-fc24-4fc4-a262-d20706c26d28" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import accuracy_score
# evitar esta abordagem pois estará sendo otimista, acaba incorrendo em um vício sobre os dados que já tínhamos visto
predicoes = melhor.predict(x_azar)
accuracy = accuracy_score(predicoes, y_azar)*100
print("A acurácia foi de %.2f" %accuracy)
# + [markdown] id="ky6vX5xNn5aD" colab_type="text"
# # Estimativa com Validação Cruzada Aninhada - Nested Cross Validation
# **Antes havia treinado e testado, agora vamos validar**
# + id="IDFyGrgIn4KQ" colab_type="code" colab={}
from sklearn.model_selection import cross_val_score
# scores = cross_val_score(busca, x_azar, y_azar, cv = GroupKFold(n_splits=10), groups = dados.modelo)
# ocorre um erro com o GroupKFold quando usa validação cruzada aninhada, por um bug do pandas que ainda nao foi corrigido
# Logo não conseguiremos prever o resultado para novos grupos
# Então usar o KFold normal
# + id="Fci8tl7XckLK" colab_type="code" outputId="728a5bfe-02c0-45aa-a9f2-1638fa276cd6" colab={"base_uri": "https://localhost:8080/", "height": 564}
from sklearn.model_selection import GridSearchCV, KFold
SEED = 301
np.random.seed(SEED)
# 4 dimensões
espaco_parametros = {
'max_depth' : [3, 5],
'min_samples_leaf' : [32,64,128],
'min_samples_split' : [32,64,128],
'criterion' : ['gini','entropy']
}
busca = GridSearchCV(DecisionTreeClassifier(),
espaco_parametros,
cv = KFold(n_splits = 5, shuffle = True))
busca.fit(x_azar, y_azar)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="Rg56E-rfxXS0" colab_type="code" outputId="df60b5ed-ec8a-45e7-c7cc-c72308dfb62e" colab={"base_uri": "https://localhost:8080/", "height": 357}
# validando
from sklearn.model_selection import cross_val_score
scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits = 5, shuffle = True))
scores
# + id="_gQdCuvLxvlY" colab_type="code" colab={}
def imprime_scores(scores):
media = scores.mean() * 100
desvio = scores.std() * 100
print("Accuracy médio %.2f" % media)
print("Intervalo [%.2f, %.2f]" % (media - 2 * desvio, media + 2 * desvio))
# + id="4NEmHP0VzI-S" colab_type="code" outputId="d317c676-945d-473e-d5d2-cf75df9dc70a" colab={"base_uri": "https://localhost:8080/", "height": 51}
imprime_scores(scores)
# + id="8UYDCgACzNke" colab_type="code" outputId="95070970-f4a8-46cc-8b54-ef5b94f4d110" colab={"base_uri": "https://localhost:8080/", "height": 119}
melhor = busca.best_estimator_
melhor
# + id="0QF6iva8zflf" colab_type="code" outputId="7f016fda-014d-4cef-ddbb-e2763c182c44" colab={"base_uri": "https://localhost:8080/", "height": 618}
# imprime a arvore de decisão do melhor modelo encontrado
from sklearn.tree import export_graphviz
import graphviz
features = x_azar.columns
dot_data = export_graphviz(melhor, out_file=None, filled=True, rounded=True,
class_names=["não","sim"],
feature_names=features)
graph = graphviz.Source(dot_data)
graph
# + [markdown] id="VPvpjf6X5UCS" colab_type="text"
# # Parte 2: Otimização com Exploração Aleatória
# + [markdown] id="2XLh6LIO5his" colab_type="text"
# **A diferença é que o random search (busca aleatória) procura valores aleatórios em um determinado espaço, e o grid search (busca por grade), como visto anteriormente, procura em todos os valores.**
# + id="lvR6NEFrzyWz" colab_type="code" outputId="901b9917-dba5-4af1-f659-0a166605dc4b" colab={"base_uri": "https://localhost:8080/", "height": 462}
from sklearn.model_selection import RandomizedSearchCV # algoritmo de busca aleatória
SEED = 301
np.random.seed(SEED)
# 4 dimensões e 36 espaços de hiperparâmetros
espaco_parametros = {
'max_depth' : [3, 5],
'min_samples_leaf' : [32,64,128],
'min_samples_split' : [32,64,128],
'criterion' : ['gini','entropy']
}
busca = RandomizedSearchCV(DecisionTreeClassifier(),
espaco_parametros,
n_iter = 16, # reduziu a busca de 36 para 16
cv = KFold(n_splits = 5, shuffle = True),
random_state = SEED) # passar o SEED para o random state
busca.fit(x_azar, y_azar)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + [markdown] id="-8ZKM2SZ-Roq" colab_type="text"
# **Validando o modelo treinado**
# + id="KPzgc9tt8fGy" colab_type="code" outputId="29ef8b58-b131-4d57-e348-e6df3fffd3ae" colab={"base_uri": "https://localhost:8080/", "height": 34}
scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits = 5, shuffle = True))
scores
# + id="RPmNrKu8999O" colab_type="code" outputId="83a24849-6e80-4922-ad4f-62ef41d554f1" colab={"base_uri": "https://localhost:8080/", "height": 51}
imprime_scores(scores)
# + id="m9ae6DTg-gOf" colab_type="code" outputId="09c689cb-777c-4c96-c23c-3e760812652f" colab={"base_uri": "https://localhost:8080/", "height": 119}
melhor = busca.best_estimator_
melhor
# Com isso percebe-se, que com o RandomizedSearchCV explorando menos espaços consegue resultados muito próximos do GridSearch
# + [markdown] id="GEJFNOyr7OTV" colab_type="text"
# # Customizando o Espaço de Hiperparâmetros
# **Uma busca em grid precisa de pontos específicos (discretos), portanto 1, 2, ..., 10. A busca aleatória permite espaços contínuos.**
# + id="HzeM5ygv7W4M" colab_type="code" outputId="f3a4fcf4-ac46-4885-cca0-7d61bf068558" colab={"base_uri": "https://localhost:8080/", "height": 547}
from scipy.stats import randint
SEED = 301
np.random.seed(SEED)
# 4 dimensões e 129.024 espaços de hiperparâmetros
espaco_parametros = {
'max_depth' : [3, 5, 10, 15, 20, 30, None],
'min_samples_leaf' : randint(32, 128), # chama 96 numeros aleatorios, entre 32 e 128
'min_samples_split' : randint(32, 128),
'criterion' : ['gini','entropy']
}
busca = RandomizedSearchCV(DecisionTreeClassifier(),
espaco_parametros,
n_iter = 16, # reduziu a busca de 129.024 para 16
cv = KFold(n_splits = 5, shuffle = True),
random_state = SEED)
busca.fit(x_azar, y_azar)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="Ji2tUH3Y9RM8" colab_type="code" outputId="bacd9139-7ef8-411a-d7c0-25fcf59d3fc0" colab={"base_uri": "https://localhost:8080/", "height": 153}
# validação cruzada aninhada
scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits=5, shuffle=True))
imprime_scores(scores)
melhor = busca.best_estimator_
print(melhor)
# Realizamos a busca muito mais rápida do que se fosse realizar a busca em todos os espaços de parâmetros
# + id="42iw0Yon9RyT" colab_type="code" outputId="6cbe75df-cc03-4314-cd1f-437c11a15b03" colab={"base_uri": "https://localhost:8080/", "height": 289}
resultados_ordenados_pela_media = resultados.sort_values('mean_test_score', ascending = False) # ordenar a coluna mean_test_score de forma decrescente
for indice, linha in resultados_ordenados_pela_media.iterrows(): #itrrows é um gerador de iteração que devolve dois elementos em cada uma das linhas: o índice e a linha
print("%.3f +- (%.3f) %s" % (linha.mean_test_score, linha.std_test_score*2, linha.params))
# printa as médias de cada um dos 16 resultados, o desvio padrão e os seus respectivos hiperparâmetros
# + [markdown] id="r-5MWBsOFHZB" colab_type="text"
# # Aumentando a Quantidade de Espaços Aleatórios Buscados
# + id="JFeKbLIcFCG1" colab_type="code" outputId="d3b7ae89-b3e0-4181-89fa-6a548e21f357" colab={"base_uri": "https://localhost:8080/", "height": 547}
SEED = 301
np.random.seed(SEED)
# 4 dimensões e 129.024 espaços de hiperparâmetros
espaco_parametros = {
'max_depth' : [3, 5, 10, 15, 20, 30, None],
'min_samples_leaf' : randint(32, 128),
'min_samples_split' : randint(32, 128),
'criterion' : ['gini','entropy']
}
busca = RandomizedSearchCV(DecisionTreeClassifier(),
espaco_parametros,
n_iter = 64, # Explorando 4 vezes mais o espaço, em relação a busca passada
cv = KFold(n_splits = 5, shuffle = True),
random_state = SEED)
busca.fit(x_azar, y_azar)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="4p2oVVgX-spO" colab_type="code" outputId="c31ff037-bcf3-46d3-926b-c58a598b484d" colab={"base_uri": "https://localhost:8080/", "height": 1000}
resultados_ordenados_pela_media = resultados.sort_values('mean_test_score', ascending = False)
for indice, linha in resultados_ordenados_pela_media.iterrows():
print("%.3f +- (%.3f) %s" % (linha.mean_test_score, linha.std_test_score*2, linha.params))
# Mostra as 64 buscas, sendo a primeira linha o melhor resultado
# + id="li_CEEutGouv" colab_type="code" outputId="570917fc-339e-4493-e909-594be5ca33dc" colab={"base_uri": "https://localhost:8080/", "height": 153}
# validação cruzada aninhada
scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits=5, shuffle=True))
imprime_scores(scores)
melhor = busca.best_estimator_
print(melhor)
# Percebe-se que mesmmo explorando 4 vezes mais espaços, não houve muita alteração no resultado
# + [markdown] id="l7lYV5hMP5LP" colab_type="text"
# # Comparando o GridSearchCV com o RandomizedSearchCV
# + [markdown] id="XF5apErGTLo_" colab_type="text"
# **GridSearchCV**
# + id="VA5thwhrP6Af" colab_type="code" outputId="7ce281f2-96bd-4719-ebba-383bbc487ca7" colab={"base_uri": "https://localhost:8080/", "height": 581}
from sklearn.ensemble import RandomForestClassifier # um outro algoritmo de árvore de decisão, onde ao invés de tentar uma única árvore, tenta diversas árvores
import time
SEED = 301
np.random.seed(SEED)
# 6 dimensões e 144 espaços de hiperparâmetros
espaco_parametros = {
'n_estimators' : [10,100], # a quantidade de estimadores que serão treinados
'bootstrap': [True,False], # permite definir se um mesmo elemento pode fazer parte de diferentes amostras
'max_depth' : [3, 5],
'min_samples_leaf' : [32,64,128],
'min_samples_split' : [32,64,128],
'criterion' : ['gini','entropy']
}
tic = time.time() # mede o tempo do inicio
busca = GridSearchCV(RandomForestClassifier(),
espaco_parametros,
cv = KFold(n_splits = 5, shuffle = True))
busca.fit(x_azar, y_azar)
tac = time.time() # mede o tempo do fim
tempo_decorrido = tac - tic
print("Tempo %.2f segundos" % tempo_decorrido)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="5VzUI49SR1tm" colab_type="code" outputId="2d256ba1-8b91-4b8f-c853-0ca9afe143e0" colab={"base_uri": "https://localhost:8080/", "height": 102}
resultados_ordenados_pela_media = resultados.sort_values('mean_test_score', ascending = False)
for indice, linha in resultados_ordenados_pela_media[:5].iterrows(): # apenas os 5 primeiros
print("%.3f +- (%.3f) %s" % (linha.mean_test_score, linha.std_test_score*2, linha.params))
# + [markdown] id="kiSn_3VrUG--" colab_type="text"
# **Vai demorar tanto que não consegue rodar no Colab, teria que importar pra maquina**
# + id="WfUboOSDTz-v" colab_type="code" colab={}
#tic = time.time()
#scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits=5, shuffle=True))
#tac = time.time()
#tempo_decorrido = tac - tic
#print("Tempo %.2f segundos" % tempo_passado)
#imprime_scores(scores)
#melhor = busca.best_estimator_
#print(melhor)
# + [markdown] id="zyDrwOPvU5MY" colab_type="text"
# **RandomizedSearchCV**
# + id="5_ZrvGb0U6XF" colab_type="code" outputId="89007899-73ea-4b14-f83e-4eadced439c5" colab={"base_uri": "https://localhost:8080/", "height": 445}
SEED = 301
np.random.seed(SEED)
# 6 dimensões e 144 espaços de hiperparâmetros
espaco_parametros = {
'n_estimators' : [10,100], # a quantidade de estimadores que serão treinados
'bootstrap': [True,False], # permite definir se um mesmo elemento pode fazer parte de diferentes amostras
'max_depth' : [3, 5],
'min_samples_leaf' : [32,64,128],
'min_samples_split' : [32,64,128],
'criterion' : ['gini','entropy']
}
tic = time.time() # mede o tempo do inicio
busca = RandomizedSearchCV(RandomForestClassifier(), # mudar o grid pelo random
espaco_parametros,
n_iter = 20, # realizar uma busca em 20 espaços
cv = KFold(n_splits = 5, shuffle = True))
busca.fit(x_azar, y_azar)
tac = time.time() # mede o tempo do fim
tempo_decorrido = tac - tic
print("Tempo %.2f segundos" % tempo_decorrido)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="kFckINPmVPI9" colab_type="code" outputId="bd6991f6-c004-4df4-eb88-7beb9bf4fecf" colab={"base_uri": "https://localhost:8080/", "height": 102}
resultados_ordenados_pela_media = resultados.sort_values('mean_test_score', ascending = False)
for indice, linha in resultados_ordenados_pela_media[:5].iterrows(): # apenas os 5 primeiros
print("%.3f +- (%.3f) %s" % (linha.mean_test_score, linha.std_test_score*2, linha.params))
# Em um tempo bem menor consegue-se resultados próximos ao do GridSearchCV
# + id="BW_3RpSaVe_l" colab_type="code" outputId="87f3779c-18eb-428c-a217-b9809c6175e4" colab={"base_uri": "https://localhost:8080/", "height": 187}
tic = time.time()
scores = cross_val_score(busca, x_azar, y_azar, cv = KFold(n_splits=5, shuffle=True))
tac = time.time()
tempo_passado = tac - tic
print("Tempo %.2f segundos" % tempo_passado)
imprime_scores(scores)
melhor = busca.best_estimator_
print(melhor)
# + [markdown] id="jEVSf0xqWryP" colab_type="text"
# **Aumentando o Espaço de Hiperparâmetros e a Busca Aleatória**
# + id="4nWtf50vVxFE" colab_type="code" outputId="ac023d21-2f2c-4b79-c972-18245c678866" colab={"base_uri": "https://localhost:8080/", "height": 581}
SEED = 301
np.random.seed(SEED)
# 6 dimensões e aumentou para 10.274.628 combinações
espaco_parametros = {
'n_estimators' : randint(10, 101),
'bootstrap': [True,False],
'max_depth' : randint(3, 6),
'min_samples_leaf' : randint(32, 129),
'min_samples_split' : randint(32, 129),
'criterion' : ['gini','entropy']
}
tic = time.time() # mede o tempo do inicio
busca = RandomizedSearchCV(RandomForestClassifier(),
espaco_parametros,
n_iter = 80, # aumentou a busca para 80 iterações em cima das 10 milhões de espaços possíveis
cv = KFold(n_splits = 5, shuffle = True))
busca.fit(x_azar, y_azar)
tac = time.time() # mede o tempo do fim
tempo_decorrido = tac - tic
print("Tempo %.2f segundos" % tempo_decorrido)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="UEeN2BDaXORK" colab_type="code" outputId="99c9202a-1914-47a8-b771-f02d493e9e3d" colab={"base_uri": "https://localhost:8080/", "height": 102}
resultados_ordenados_pela_media = resultados.sort_values('mean_test_score', ascending = False)
for indice, linha in resultados_ordenados_pela_media[:5].iterrows(): # apenas os 5 primeiros
print("%.3f +- (%.3f) %s" % (linha.mean_test_score, linha.std_test_score*2, linha.params))
# Mesmo nesse espaço enorme de mais de 10 milhões de combinações, não tivemos uma variabilidade muito grande de resultados.
# + [markdown] id="p_yDCBg_YlNI" colab_type="text"
# # Otimização sem validação cruzada (treino, teste e validação)
# + id="kfVLsGp9bf0P" colab_type="code" outputId="aac95d08-5643-4e37-f414-43f23cf990f2" colab={"base_uri": "https://localhost:8080/", "height": 85}
# separa antecipadamente 20% dos dados para validação
from sklearn.model_selection import train_test_split
SEED=301
np.random.seed(SEED)
x_treino_teste, x_validacao, y_treino_teste, y_validacao = train_test_split(x_azar, y_azar, test_size = 0.2, shuffle = True, stratify = y_azar)
print(x_treino_teste.shape)
print(x_validacao.shape)
print(y_treino_teste.shape)
print(y_validacao.shape)
# + id="UKNTSchyYl-7" colab_type="code" outputId="82bc011f-b39f-46dc-9eef-e491582bf390" colab={"base_uri": "https://localhost:8080/", "height": 581}
from sklearn.model_selection import StratifiedShuffleSplit
# o StratifiedShuffleSplit irá aleatorizar a ordem dos dados e quebrá-los de acordo com a estratificação dos dados que passarmos para ele
SEED=301
np.random.seed(SEED)
espaco_de_parametros = {
"n_estimators" : randint(10, 101),
"max_depth" : randint(3, 6),
"min_samples_split": randint(32, 129),
"min_samples_leaf": randint(32, 129),
"bootstrap" : [True, False],
"criterion": ["gini", "entropy"]
}
split = StratifiedShuffleSplit(n_splits = 1, test_size = 0.25) # 25 % pra teste, que equivale a 20% do total de dados (treino,teste,validação)
tic = time.time()
busca = RandomizedSearchCV(RandomForestClassifier(),
espaco_de_parametros,
n_iter = 5, # realizar apenas 5 iterações aleatórias
cv = split) # trocar o KFold pelos parâmetros do StratifiedShuffleSplit
busca.fit(x_treino_teste, y_treino_teste) # utiliza os 80% restante dos dados (pois 20% haviam sidos retirados para validação)
tac = time.time()
tempo_que_passou = tac - tic
print("Tempo %.2f segundos" % tempo_que_passou)
resultados = pd.DataFrame(busca.cv_results_)
resultados.head()
# + id="FXtMzOtqbFcX" colab_type="code" outputId="88879213-9a7a-4580-b02b-1a3e17dc611e" colab={"base_uri": "https://localhost:8080/", "height": 51}
tic = time.time()
scores = cross_val_score(busca, x_validacao, y_validacao, cv = split) # substitui pelos dados de validação, que haviam sidos separados no início
tac = time.time()
tempo_passado = tac - tic
print("Tempo %.2f segundos" % tempo_passado)
scores
# O resultado é apenas um valor - como só tivemos um teste e uma validação, remove-se a impressão da média e do intervalo
# O cross validation é um processo bastante interessante e prático.
# Porém, quando existem motivos para não utilizarmos o cross validation, devemos nos atentar a alguns detalhes importantes
# por exemplo, à perda do intervalo de resultados.
| hyperparameters_model_optimization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "skip"}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# + slideshow={"slide_type": "skip"}
df = pd.read_csv('../../pandas-videos/data/imdb_1000.csv')
# + [markdown] slideshow={"slide_type": "slide"}
# ## COMP 3122 - Artificial Intelligence with Python
# __Week 4 lab__
#
# ### [github.com/kamrik/ML1](https://github.com/kamrik/ML1)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Lab test -> home assignment
# - [exercises/numpy_assignment.py](exercises/numpy_assignment.py)
# - Submit by email
# - Starting some time this weekend the bot will respond withing several minutes with a grade
# - Up to 5 submissions, the best counts
# - I'll manually inspect some randomly selected submissions
# - If you believe there is a bug, email me or open an issue on GitHub (don't disclese your entire solution in publicly visible communication)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pandas
# - The videos (or the notebook from videos) - important
# - `df.loc[]`
# - `df.idxmax[]`
# - `df.groupby()`
# - Lab exercise - `exercises/olympic_history.ipynb`
# - Use the notebook from the videos as a reference - keep it open in another tab
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### df['column'] vs df.loc[row]
# -
df.head()
# + [markdown] slideshow={"slide_type": "slide"}
# 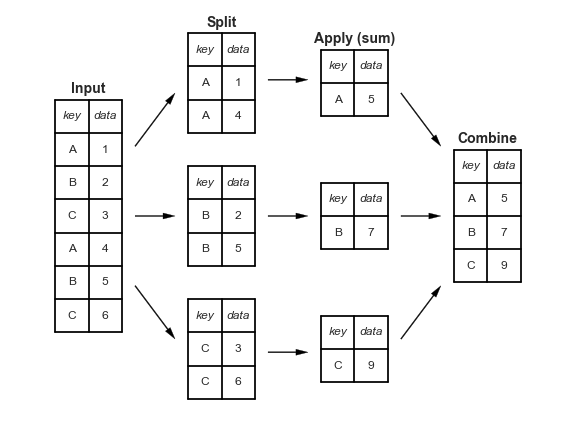
# + slideshow={"slide_type": "slide"}
df.head()
# + slideshow={"slide_type": "slide"}
df.groupby('genre').count()
# -
| ML1/lectures/04_week_lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: xpython
# language: python
# name: xpython
# ---
# # Logistic Regression: Problem solving
#
# In this session, you will predict whether or not a candy is popular based on its other properties.
# This dataset [was collected](http://fivethirtyeight.com/features/the-ultimate-halloween-candy-power-ranking/) to discover the most popular Halloween candy.
#
# | Variable | Type | Description |
# |:-----------------|:------------------|:--------------------------------------------------------------|
# | chocolate | Numeric (binary) | Does it contain chocolate? |
# | fruity | Numeric (binary) | Is it fruit flavored? |
# | caramel | Numeric (binary) | Is there caramel in the candy? |
# | peanutalmondy | Numeric (binary) | Does it contain peanuts, peanut butter or almonds? |
# | nougat | Numeric (binary) | Does it contain nougat? |
# | crispedricewafer | Numeric (binary) | Does it contain crisped rice, wafers, or a cookie component? |
# | hard | Numeric (binary) | Is it a hard candy? |
# | bar | Numeric (binary) | Is it a candy bar? |
# | pluribus | Numeric (binary) | Is it one of many candies in a bag or box? |
# | sugarpercent | Numeric (0 to 1) | The percentile of sugar it falls under within the data set. |
# | pricepercent | Numeric (0 to 1) | The unit price percentile compared to the rest of the set. |
# | winpercent | Numeric (percent) | The overall win percentage according to 269,000 matchups |
# | popular | Numeric (binary) | 1 if win percentage is over 50% and 0 otherwise |
#
# **Acknowledgements:**
#
# This dataset is Copyright (c) 2014 ESPN Internet Ventures and distributed under an MIT license.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Load the data
#
# First import `pandas`.
# -
# Load a dataframe with `"datasets/candy-data.csv"` and display it.
# Notice there is a bogus variable `competitorname` that is actually an ID, also known as an **index**.
# We saw the same thing in KNN regression with the `mpg` dataset, but that time it was the car name.
#
# Load the dataframe again, but this time use `index_col="competitorname"` to fix this.
# ## Explore the data
#
# ### Descriptive statistics
#
# Describe the data.
# Remember that for the 0/1 variables, the mean reflects the average presence of an ingredient in candy.
# For example, `chocolate` is in 43.5% of candy.
# **QUESTION:**
#
# What is the least common ingredient (there may be more than one that is the same)?
# **ANSWER: (click here to edit)**
#
# **QUESTION:**
#
# What is the most common ingredient?
# **ANSWER: (click here to edit)**
#
# **QUESTION:**
#
# Do you see any problems with the data, e.g. missing data?
# **ANSWER: (click here to edit)**
#
# ### Correlations
#
# Create and display a correlation matrix.
# **QUESTION:**
#
# What property is most positively related to being popular?
# What property is most negatively related to being popular?
# **ANSWER: (click here to edit)**
#
#
# Create a heatmap for the correlation matrix.
# Start by importing `plotly.express`.
# Create the heatmap figure
# And show it.
# **QUESTION:**
#
# What color is strongly negative, what color is zero, and what color is strongly positive?
# **ANSWER: (click here to edit)**
#
# **QUESTION:**
#
# What's going on in the lower right corner?
# **ANSWER: (click here to edit)**
#
# ### Histograms
#
# For binary variables, histograms don't tell us anything that the descriptives don't already tell us.
#
# However, there are two percent-type variables to plot, `sugarpercent` and `pricepercent`.
#
# Plot a histogram of `sugarpercent`.
# Plot a histogram of `pricepercent`.
# **QUESTION:**
#
# What can you say about the distributions of `sugarpercent` and `pricepercent`?
# Is there anything we should be concerned about?
# **ANSWER: (click here to edit)**
#
# ## Prepare train/test sets
#
# You need to split the dataframe into training data and testing data, and also separate the predictors from the class labels.
#
# Start by dropping the label, `popular`, and its counterpart, `winpercent`, to make a new dataframe called `X`.
# Save a dataframe with just `popular` in `Y`.
# Import `sklean.model_selection` to split `X` and `Y` into train and test sets.
# Now do the splits.
# ## Logistic regression model
#
# Import libraries for:
#
# - Logistic regression
# - Metrics
# - Ravel
#
# **NOTE: technically we don't need to scale anything and so don't need a pipeline.**
# **QUESTION:**
#
# Why don't we need to scale anything?
# **ANSWER: (click here to edit)**
#
# Create the logistic regression model.
# Train the logistic regression model using the splits.
# Get predictions from the model using the test data.
# ## Assessing the model
#
# Print the model accuracy.
# **QUESTION:**
#
# How does this compare to the average value of `popular`?
# Is this a good accuracy?
# **ANSWER: (click here to edit)**
#
# Print precision, recall, and F1.
# **QUESTION:**
#
# How to the precision/recall/f1 compare for unpopular (0) and popular (1)?
# **ANSWER: (click here to edit)**
#
# Make an ROC plot.
# **QUESTION:**
#
# If we decreased the recall to .66, what would the false positives be? HINT: hover your mouse over the plot line at that value.
# **ANSWER: (click here to edit)**
#
# This last part is something we didn't really get to develop in the first session, so just run the code.
#
# The odds ratio shows how much more likely a property makes the candy `popular`.
# For many of these, the property is just presence/absence.
# For example, the odds ratio of 3.06 on chocolate means that having chocolate as an ingredient makes the candy 3.06 times more popular than candy without chocolate.
pd.DataFrame( {"variable":X.columns, "odds_ratio":np.exp(np.ravel(lm.coef_)) })
# **QUESTION:**
#
# What are the top three *ingredients* that make something popular? Do any surprise you given the correlation matrix?
# **ANSWER: (click here to edit)**
#
# ## Submit your work
#
# When you have finished the notebook, please download it, log in to [OKpy](https://okpy.org/) using "Student Login", and submit it there.
#
# Then let your instructor know on Slack.
#
| Logistic-regression-PS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# ## CartPole with Q-learning
#
# Q-learning is a off-policy temporal-difference method that uses the following update:
# $$Q(S_t, A_t) \gets Q(S_t, A_t) + \alpha \left[ R_{t+1} + \gamma max_a Q(S_t+1, a) - Q(S_t, A_t) \right]$$
#
# where $Q(S_t, A_t)$ is the value of taking action $A_t$ in state $S_t$, $\alpha$ is the step-size (learning rate) and $R_{t+1}$ is the reward received. We update $Q(S_t, A_t)$ using the direct reward received plus the reward we expect to receive in future. The update can be seen as optimistic because we take the maximum expected return.
#
# The objective of the CartPole problem (from OpenAI gym: https://gym.openai.com/envs/CartPole-v0) is to teach the cart to balance a pole by applying a horizontal force to the left or right. An episode ends when the pole tips more than 15 degrees from the vertical or the cart moves more than 2.4 units from the center. In smaller problems like Grid World, a table could be used to store all possible combinations of states and actions. In more complex problems, especially those with continuous states or actions, it is impossible to completely represent all state and action pairs so an approximator (neural network) is used. The input to the neural network is a state and the output is the value of all actions (we can then choose to take the action with the highest value if we're following $\epsilon$-greedy action selection for example).
# + deletable=true editable=true
import math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import gym
import cntk
from cntk import *
from cntk.layers import *
# + [markdown] deletable=true editable=true
# The Agent class encapsulates all the necessary logic related to action action selection and model training. The network inputs and outputs depend on the environment chosen so it receives their dimensions as arguments to its constructor. I have chosen to use a network with a single hidden layer of 64 neurons and ReLu (REctified Linear Unit) activation. I calculate the loss as the mean of the square difference between the actual outputs and the expected outputs and have chosen to use a learning rate of 0.0025 and stochastic gradient descent for learning.
#
# Once again we are using $\epsilon$-greedy action selection. We generate a random number and if it is below our threshold, $\epsilon$, we pick a random action (explore), otherwise, we use the output of our model (exloit current knowledge) and the action with the highest value is selected (greedy). To see the effects of a constant versus decaying epsilon (value reduces with time), the constructor has a parameter which is a function for updating epsilon based on the number of completed episodes.
#
# The evaluate function allows us to evaluate the average performance of the agent over a given number of episodes.
# + deletable=true editable=true
class Agent:
def __init__(self, state_dim, action_dim, learning_rate, epsilon_fn):
self.state_dim = state_dim
self.action_dim = action_dim
self.learning_rate = learning_rate
self.epsilon_fn = epsilon_fn
self.epsilon = 1
# Create the neural network
self.state_var = input(state_dim, np.float32)
self.action_var = input(action_dim, np.float32)
self.model = Sequential([
Dense(64, activation=relu),
Dense(action_dim)
])(self.state_var)
loss = reduce_mean(square(self.model - self.action_var), axis=0)
lr_schedule = learning_rate_schedule(self.learning_rate, UnitType.sample)
learner = sgd(self.model.parameters, lr_schedule)
self.trainer = Trainer(self.model, loss, learner)
def predict(self, state):
"""
Feeds a state through our model to obtain the values of each action
"""
return self.model.eval(state)[0]
def act(self, state):
"""
Selects an action using the epoch-greedy approach
"""
prob = np.random.randn(1)
if prob > self.epsilon:
# exploit (greedy)
return np.argmax(self.predict(state))
else:
# explore (random action)
return np.random.randint(0, self.action_dim)
def update_epsilon(self, episode):
"""
Update the value of epsilon using the provided update function
"""
self.epsilon = self.epsilon_fn(episode)
def train(self, x, y):
"""
Performs a single gradient descent step using the provided state and target
"""
self.trainer.train_minibatch({self.state_var: x, self.action_var: y})
def evaluate(self, env, n):
"""
Computes the average performance of the trained model over n episodes
"""
episode = 0
rewards = 0
while episode < n:
s = env.reset()
done = False
while not done:
a = np.argmax(self.predict(s.astype(np.float32)))
s, r, done, info = env.step(a)
rewards += r
episode += 1
return rewards / float(n)
# + [markdown] deletable=true editable=true
# For each step of an episode, we select an action (using $\epsilon$-greedy action selection) and then execute it yielding the next state and reward as well as a boolean indicating if the next state is terminal. We then calculate the expected return as the sum of the received reward plus the discounted future reward (0 if terminal). The current state serves as our input and the expected return serves as the label (truth value) for the neural network. Finally, I store the reward for each episode so we can monitor the progress.
# + deletable=true editable=true
def train(env, agent, episodes, gamma):
"""
param env: The gym environment to train with
param agent: The agent to train
param episodes: The number of episodes to train for
param gamma: The discount factor
"""
episode = 0
rewards = 0
log_freq = 200
episode_rewards = []
s = env.reset().astype(np.float32)
while episode < episodes:
# Select an action using policy derived from Q (e-greedy)
a = agent.act(s)
# Take action and observe the next state and reward
s_, r, done, info = env.step(a)
s_ = s_.astype(np.float32)
# Compute target, y_i
y = agent.predict(s)
if done:
y[a] = r
else:
y[a] = r + gamma * np.amax(agent.predict(s_))
# Train using state and computed target
agent.train(s, y)
s = s_
rewards += r
if done:
# Episode over, reset environment
episode_rewards.append(rewards)
rewards = 0
episode += 1
agent.update_epsilon(episode)
s = env.reset().astype(np.float32)
if episode % log_freq == 0:
ave = sum(episode_rewards[(episode - log_freq):]) / float(log_freq)
print('Episode = {}, Average rewards = {}'.format(episode, ave))
return episode_rewards
# + [markdown] deletable=true editable=true
# I have decided to train for 10k episodes using a discount factor of 0.60 - for all states before the terminal state, we receive a reward of 1, so I chose to factor later rewards less. For the constant $\epsilon$ agent, I've chosen to use $\epsilon = 0.1$ and for the other, $\epsilon$ decays from a value of 1 to a minimum of 0.05 by the 8000th episode.
# + deletable=true editable=true
# Fixed value epsilon
def constant_epsilon(episode):
return 0.1
# + deletable=true editable=true
# epsilon starts with 1 and decays to a minimum of 0.05
# decay factor was chosen such that the value of epsilon is 0.05 by episode 8000
def decaying_epsilon(episode):
return max(math.exp(-3.74e-4 * episode), 0.05)
# + deletable=true editable=true
gamma = 0.60
learning_rate = 0.00025
episodes = 10000
# + deletable=true editable=true
env = gym.make('CartPole-v0')
state_dim = env.observation_space.shape
action_dim = env.action_space.n
# + deletable=true editable=true
agent1 = Agent(state_dim, action_dim, learning_rate, constant_epsilon)
rewards1 = train(env, agent1, episodes, gamma)
# + deletable=true editable=true
agent2 = Agent(state_dim, action_dim, learning_rate, decaying_epsilon)
rewards2 = train(env, agent2, episodes, gamma)
# + [markdown] deletable=true editable=true
# For visual comparison of learning with time, I have plotted the moving averages of the received rewards below:
# + deletable=true editable=true
pd.Series(rewards1).rolling(window=100).mean().plot(label='constant \u03b5')
pd.Series(rewards2).rolling(window=100).mean().plot(label='decaying \u03b5')
plt.legend()
plt.show()
# + [markdown] deletable=true editable=true
# Here, I evaluate the performance of the trained agents over 100 episodes:
# + deletable=true editable=true
eval_episodes = 100
ave_constant = agent1.evaluate(env, eval_episodes)
ave_decaying = agent2.evaluate(env, eval_episodes)
print('Average (with constant \u03b5) = {}'.format(ave_constant))
print('Average (with decaying \u03b5) = {}'.format(ave_decaying))
# + [markdown] deletable=true editable=true
# ## Conclusion
# From the plot of average reward for each model during training, we can see that learning is better with a decaying $\epsilon$ as opposed to a constant small epsilon. The results of evaluation over 100 episodes also supports this point. In practice, epsilon is usually annealed to a small value to simulate how we learn - a lot of exploration at the start but less and less as our knowledge improves.
#
# Here I run and render an episode using the better model:
# + deletable=true editable=true
s = env.reset()
done = False
while not done:
env.render()
a = np.argmax(agent2.predict(s.astype(np.float32)))
s, r, done, info = env.step(a)
env.close()
# + [markdown] deletable=true editable=true
# Obviously there is still room for improvement. You can play around with the number of layers in the network (as well as the number of neurons in each layer), a different min value for epsilon (in the exponentially decaying version), a different learner (as opposed to stochastic gradient descent), different learning rate, gamma, train for longer and so on...
#
# ## References
# 1. <NAME>, <NAME> (1998). Reinforcement Learning: An Introduction. MIT Press.
| cartpole/q_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 7
#
# # Part 1 - DT
#
# ## Capital Bikeshare data
# ## Introduction
#
# - Capital Bikeshare dataset from Kaggle: [data](https://github.com/justmarkham/DAT8/blob/master/data/bikeshare.csv), [data dictionary](https://www.kaggle.com/c/bike-sharing-demand/data)
# - Each observation represents the bikeshare rentals initiated during a given hour of a given day
# %matplotlib inline
import pandas as pd
import numpy as np
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor, export_graphviz
# read the data and set "datetime" as the index
bikes = pd.read_csv('../datasets/bikeshare.csv', index_col='datetime', parse_dates=True)
# "count" is a method, so it's best to rename that column
bikes.rename(columns={'count':'total'}, inplace=True)
# create "hour" as its own feature
bikes['hour'] = bikes.index.hour
bikes.head()
bikes.tail()
# - **hour** ranges from 0 (midnight) through 23 (11pm)
# - **workingday** is either 0 (weekend or holiday) or 1 (non-holiday weekday)
# # Exercise 7.1
#
# Run these two `groupby` statements and figure out what they tell you about the data.
# mean rentals for each value of "workingday"
bikes.groupby('workingday').total.mean()
# mean rentals for each value of "hour"
bikes.groupby('hour').total.mean()
# # Exercise 7.2
#
# Run this plotting code, and make sure you understand the output. Then, separate this plot into two separate plots conditioned on "workingday". (In other words, one plot should display the hourly trend for "workingday=0", and the other should display the hourly trend for "workingday=1".)
# mean rentals for each value of "hour"
bikes.groupby('hour').total.mean().plot()
# Plot for workingday == 0 and workingday == 1
# hourly rental trend for "workingday=0"
bikes[bikes.workingday==0].groupby('hour').total.mean().plot()
# hourly rental trend for "workingday=1"
bikes[bikes.workingday==1].groupby('hour').total.mean().plot()
# combine the two plots
bikes.groupby(['hour', 'workingday']).total.mean().unstack().plot()
# Write about your findings
# # Exercise 7.3
#
# Fit a linear regression model to the entire dataset, using "total" as the response and "hour" and "workingday" as the only features. Then, print the coefficients and interpret them. What are the limitations of linear regression in this instance?
# # Exercice 7.4
#
# Create a Decision Tree to forecast "total" by manually iterating over the features "hour" and "workingday". The algorithm must at least have 6 end nodes.
# # Exercise 7.5
#
# Train a Decision Tree using scikit-learn. Comment about the performance of the models.
# # Part 2 - Bagging
# ## Mashable news stories analysis
#
# Predicting if a news story is going to be popular
df = pd.read_csv('../datasets/mashable.csv', index_col=0)
df.head()
df.shape
X = df.drop(['url', 'Popular'], axis=1)
y = df['Popular']
y.mean()
# train/test split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1)
# # Exercise 7.6
#
# Estimate a Decision Tree Classifier and a Logistic Regression
#
# Evaluate using the following metrics:
# * Accuracy
# * F1-Score
# # Exercise 7.7
#
# Estimate 300 bagged samples
#
# Estimate the following set of classifiers:
#
# * 100 Decision Trees where max_depth=None
# * 100 Decision Trees where max_depth=2
# * 100 Logistic Regressions
# # Exercise 7.8
#
# Ensemble using majority voting
#
# Evaluate using the following metrics:
# * Accuracy
# * F1-Score
# # Exercise 7.9
#
# Estimate te probability as %models that predict positive
#
# Modify the probability threshold and select the one that maximizes the F1-Score
# # Exercise 7.10
#
# Ensemble using weighted voting using the oob_error
#
# Evaluate using the following metrics:
# * Accuracy
# * F1-Score
# # Exercise 7.11
#
# Estimate te probability of the weighted voting
#
# Modify the probability threshold and select the one that maximizes the F1-Score
# # Exercise 7.12
#
# Estimate a logistic regression using as input the estimated classifiers
#
# Modify the probability threshold such that maximizes the F1-Score
| Exercises/E7-DecisionTrees_Bagging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rucha04/example22/blob/main/Copy_of_ML_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LeIG7bjIT8Jx"
# 3. Run a logistic regression model on train data and compute confusion matrix in test set.
# + id="MluyGZc8CUMD"
# %matplotlib inline
from sklearn.datasets import load_digits
digits = load_digits()
# + colab={"base_uri": "https://localhost:8080/"} id="5m20e02JCcfB" outputId="b2b573c9-08e9-46d9-d16b-3afd39bdf9e7"
# Print to show there are 1797 images (8 by 8 images for a dimensionality of 64)
print("Image Data Shape" , digits.data.shape)
# Print to show there are 1797 labels (integers from 0-9)
print("Label Data Shape", digits.target.shape)
# + [markdown] id="hopwsP57Cj8Y"
# Showing the Images and Labels (Digits Dataset)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="MJEFmGxZCmmN" outputId="d5391bb3-868f-4809-d70d-e43d87b6af37"
import numpy as np
import matplotlib.pyplot as plt
plt.figure(figsize=(20,4))
for index, (image, label) in enumerate(zip(digits.data[0:5], digits.target[0:5])):
plt.subplot(1, 5, index + 1)
plt.imshow(np.reshape(image, (8,8)), cmap=plt.cm.gray)
plt.title('Training: %i\n' % label, fontsize = 20)
# + [markdown] id="0b6UVnXgCwi7"
# Splitting Data into Training and Test Sets (Digits Dataset)
# + id="FQkyeTPOCr5t"
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(digits.data, digits.target, test_size=0.25, random_state=0)
# + id="SLOhTA6lC4-M"
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
# + id="gop7aJpwC8BM"
logisticRegr = make_pipeline(StandardScaler(), LogisticRegression())
# + colab={"base_uri": "https://localhost:8080/"} id="SyksIzSAD5UD" outputId="d551ec90-beeb-424b-be82-79255e3c2531"
logisticRegr.fit(x_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="_TeXmaljGKMc" outputId="03d965af-64e8-423c-cf0f-13031e29d6f6"
# Returns a NumPy Array
# Predict for One Observation (image)
logisticRegr.predict(x_test[0].reshape(1,-1))
# + colab={"base_uri": "https://localhost:8080/"} id="NXw4T6eKGNvc" outputId="6711fc96-1504-4c60-8781-bd9c2f97ca94"
# Predict for Multiple Observations (images) at Once
logisticRegr.predict(x_test[0:10])
# + id="N8CPXUnOGRJG"
# Make predictions on entire test data
predictions = logisticRegr.predict(x_test)
# + colab={"base_uri": "https://localhost:8080/"} id="DsM69LRbGURX" outputId="2316d208-9af7-4acd-f024-91aeb0d3eb0f"
# Use score method to get accuracy of model
score = logisticRegr.score(x_test, y_test)
print(score)
# + [markdown] id="PTwMqHQBGbr3"
# Confusion Matrix (Digits Dataset)
# + id="-YRf12GXGcn-"
import numpy as np
import seaborn as sns
from sklearn import metrics
# + [markdown] id="w3FpYt1jGipM"
# Method 1 (Seaborn)
# + id="5RZq4EMkGj1C"
cm = metrics.confusion_matrix(y_test, predictions)
# + colab={"base_uri": "https://localhost:8080/", "height": 526} id="DLgZD6xyGnyp" outputId="5869697d-b392-4c11-9661-1bcdca5d418e"
plt.figure(figsize=(9,9))
sns.heatmap(cm, annot=True, fmt=".3f", linewidths=.5, square = True, cmap = 'Blues_r');
plt.ylabel('Actual label');
plt.xlabel('Predicted label');
all_sample_title = 'Accuracy Score: {0}'.format(score)
plt.title(all_sample_title, size = 15);
plt.savefig('toy_Digits_ConfusionSeabornCodementor.png')
#plt.show();
# + [markdown] id="qnYk3wN9GvJ8"
# Method 2 (Matplotlib)
# + colab={"base_uri": "https://localhost:8080/", "height": 657} id="vg-olkaFGrvR" outputId="503bf8e0-caf5-4103-d59c-dcb796e2f57e"
cm = metrics.confusion_matrix(y_test, predictions)
plt.figure(figsize=(9,9))
plt.imshow(cm, interpolation='nearest', cmap='Pastel1')
plt.title('Confusion matrix', size = 15)
plt.colorbar()
tick_marks = np.arange(10)
plt.xticks(tick_marks, ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"], rotation=45, size = 10)
plt.yticks(tick_marks, ["0", "1", "2", "3", "4", "5", "6", "7", "8", "9"], size = 10)
plt.tight_layout()
plt.ylabel('Actual label', size = 15)
plt.xlabel('Predicted label', size = 15)
width, height = cm.shape
for x in range(width):
for y in range(height):
plt.annotate(str(cm[x][y]), xy=(y, x),
horizontalalignment='center',
verticalalignment='center')
plt.savefig('toy_Digits_ConfusionMatplotlibCodementor.png')
#plt.show()
| Copy_of_ML_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys; sys.path.append("C:/GitWorks/pdb-profiling")
from pdb_profiling import default_config
from pdb_profiling.processors import PDB
from pdb_profiling.utils import DisplayPDB
from pdb_profiling.viewer import NGL
default_config("C:/GitWorks/pdb-profiling/test/demo")
# -
import nglview
import ipywidgets
from pandas import DataFrame
from unsync import unsync
pdb_id = '10gs'
pdb_ob = PDB(pdb_id)
pdb_ob.set_assembly().result()
DisplayPDB(dark=True).show(pdb_ob.pdb_id, pdb_ob.assembly)
# +
@unsync
async def get_interface_df(pdb_ob):
molecule_types = (
'polypeptide(L)',
'polypeptide(D)',
'polydeoxyribonucleotide',
'polyribonucleotide',
'polydeoxyribonucleotide/polyribonucleotide hybrid',
'bound',
'carbohydrate_polymer')
return DataFrame([i async for i in pdb_ob.pipe_interface_res_dict(
func='pipe_protein_else_interface',
molecule_types=molecule_types,
allow_same_class_interaction=True)])
interface_df = get_interface_df(pdb_ob).result()
interface_df.head().T
# -
rep = [{"type": "all", "params":{"assembly": "AU"}}]
views = [NGL.get_interface_view(
nglview.NGLWidget(nglview.FileStructure(f'C:/Users/Nature/Downloads/{pdb_id}.cif') , representation=rep),
interface_df.loc[i]) for i in range(4)]
# background='#272822'
ipywidgets.GridBox(views, layout=ipywidgets.Layout(grid_template_columns='1fr 1fr'))
view = nglview.show_file(f'C:/Users/Nature/Downloads/{pdb_id}.cif')
view.representation = rep
_ = [NGL.get_interface_view(view, interface_df.loc[i], background='#272822', surface_opacity_1=0.05, surface_opacity_2=0.05, surface_color_1='#F0F8FF', surface_color_2='#F0F8FF') for i in range(4)]
view
| examples/nglviewer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install atoti
class employee:
name = 'Anuj'
age = 23
bodytype = 'Athletic'
def is_he_the_man(self):
if self.name =='anuj':
print('He is my man')
else:
print('No he is not the one')
emp = employee()
emp.name
emp.is_he_the_man()
# # how to change attributes of a class
class emps:
no_working_hour = 40
# +
em1 = emps()
em2 = emps()
print(em1.no_working_hour)
print(em2.no_working_hour)
# +
emps.no_working_hour = 55
em1.no_working_hour
# -
# # Instance attribute
# +
#class attribute is common to all the instance of your class while the instance attribute is specific to each instance of your class
# -
em1.name = 'Anuj'
em2.name = 'Rudra'
em1.name
em2.name
# +
em2.color
#Here what happens is it first checks if instance attribute of color is present, if not it checks for class attribute, if not
#then it returns false / error
# -
em2.color = 'Wheatish'
em2.color
em1.no_working_hour = 66
em1.no_working_hour
em2.no_working_hour
class Employee:
def employee_details():
pass
emp = Employee()
emp.employee_details()
# +
#python by default takes
#class.instance(object)
# -
class Employee:
def employee_details(self):
self.name = 'Anuj'
print('Name = ', self.name)
emp = Employee()
emp.employee_details()
# +
#or another way by python style (less used convention)
Employee.employee_details(emp)
# employee_details() takes 0 positional arguments but 1 was given
#now you understand why this error comes up
# +
#if you don't use self object then lifespan of that particular instance is limited to that method. / function
# -
class employees:
def empl_details(self):
self.name = 'Anuj'
print('Name', self.name)
age = 23
print('age', age)
def emply_det(self):
print('printing by diff. method :', self.name)
print('Age', Age)
# +
obj = employees()
print(obj.empl_details())
# -
print(obj.emply_det())
#so you see scope of 'age' gets limited within that method while name gets printed
# # Use of static method
#
# +
#without passing self, if you just want to print a message just use decorator
#decorators are functions which takes another function and extend their functionality, it ignores the binding of object
# -
class House:
def requirements(self):
self.area = 250
print("Area of house is", self.area)
@staticmethod
def message():
print("Welcome to our house")
print("this is sum", 3+9)
house = House()
house.message()
house.requirements()
house.area
# # Use of init method
# as soon as you call the object is being created of class init method gets invoked default, use of init method is to initialize
# all the variables.
class House:
def house_det(self):
self.name = 'Bhavan'
def houses_dett(self):
print(self.name)
hous = House()
hous.houses_dett()
# +
#when you are trying to initiate the second fucntion first, remember it has no initializtion, so it fails to fetch the attribute
# +
#so __init__ is a mechanism which initialize all the attributes of our object or class before they are being used.
#special method start and end with __
# -
class House:
def __init__(self):
self.name = 'Bhavan'
def houses_dett(self):
print(self.name)
hous = House()
hous.houses_dett()
class toffee:
def __init__(self, candyname):
self.canname = candyname
def candy(self):
print("This is a", self.canname)
# +
tf1 = toffee('Sweetcandy')
tf2 = toffee('Sourcandy')
tf1.candy()
# -
tf2.candy()
# +
class Library:
def __init__(self, listofbooks):
self.available_books= listofbooks
def Displaybooks(self):
print()
print('Avaiable books:')
for i in self.available_books:
print(i)
def Lendbooks(self, requestedBook):
if requestedBook in self.available_books:
print("You have borrowed the book")
self.available_books.remove(requestedBook)
else:
print("Please enter the correct bookname")
def Update_books(self, returnedbook):
self.available_books.append(returnedbook)
print("You have returned the book")
class Customer:
def Borrow_book(self):
print('Enter the book you would like to borrow')
self.book = input()
return self.book
def Return_book(self):
print("Enter the book you would like to return")
self.book = input()
return self.book
library = Library(['Intelligent Investor', 'One Up on Wall Street', 'CPR and Pivot Points'])
customer = Customer()
while True:
print('Enter 1 to display books')
print('Enter 2 to lend books')
print('Enter 3 to return books')
print('Enter 4 to exit')
userchoice = int(input())
if userchoice is 1:
library.Displaybooks()
elif userchoice is 2:
requestedbook = customer.Borrow_book()
library.Lendbooks(requestedbook)
elif userchoice is 3:
returnedbooks = customer.Return_book()
library.Update_books(returnedbooks)
elif userchoice is 4:
exit
# -
# # Inheritance
#You can inhert other attribute of class if that class is inherited.
| oops_revision.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Overview
# 
# # Terminology
# 1. CTE: Cross Track Error
# 2.
# ## Quiz
# Which of the following is best suited to control the car?
# - [ ] Steering Constant
# - [ ] Random Steering Control
# - [x] Steer in proportion to Cross Track Error
#
# ## Quiz
# Your steering angle, alpha, equals a proportional factor of tau to the cross track error. What will happen to the car?
# # Import Libraries
import random
import numpy as np
import matplotlib.pyplot as plt
# # Robot class
class Robot(object):
def __init__(self, length=20.0):
"""
Creates robot and initializes location/orientation to 0, 0, 0.
"""
self.x = 0.0
self.y = 0.0
self.orientation = 0.0
self.length = length
self.steering_noise = 0.0
self.distance_noise = 0.0
self.steering_drift = 0.0
def set(self, x, y, orientation):
"""
Sets a robot coordinate.
"""
self.x = x
self.y = y
self.orientation = orientation % (2.0 * np.pi)
def set_noise(self, steering_noise, distance_noise):
"""
Sets the noise parameters.
"""
# makes it possible to change the noise parameters
# this is often useful in particle filters
self.steering_noise = steering_noise
self.distance_noise = distance_noise
def set_steering_drift(self, drift):
"""
Sets the systematical steering drift parameter
"""
self.steering_drift = drift
def move(self, steering, distance, tolerance=0.001, max_steering_angle=np.pi / 4.0):
"""
steering = front wheel steering angle, limited by max_steering_angle
distance = total distance driven, must be non-negative
"""
if steering > max_steering_angle:
steering = max_steering_angle
if steering < -max_steering_angle:
steering = -max_steering_angle
if distance < 0.0:
distance = 0.0
# apply noise
steering2 = random.gauss(steering, self.steering_noise)
distance2 = random.gauss(distance, self.distance_noise)
# apply steering drift
steering2 += self.steering_drift
# Execute motion
turn = np.tan(steering2) * distance2 / self.length
if abs(turn) < tolerance:
# approximate by straight line motion
self.x += distance2 * np.cos(self.orientation)
self.y += distance2 * np.sin(self.orientation)
self.orientation = (self.orientation + turn) % (2.0 * np.pi)
else:
# approximate bicycle model for motion
radius = distance2 / turn
cx = self.x - (np.sin(self.orientation) * radius)
cy = self.y + (np.cos(self.orientation) * radius)
self.orientation = (self.orientation + turn) % (2.0 * np.pi)
self.x = cx + (np.sin(self.orientation) * radius)
self.y = cy - (np.cos(self.orientation) * radius)
def __repr__(self):
return '[x=%.5f y=%.5f orient=%.5f]' % (self.x, self.y, self.orientation)
def make_robot():
"""
Resets the robot back to the initial position and drift.
You'll want to call this after you call `run`.
"""
robot = Robot()
robot.set(0, 1, 0) #set(self, x, y, orientation):
robot.set_steering_drift(10 / 180 * np.pi) #set_steering_drift(self, drift)
return robot
# # Implement P Controller
def run(robot, tau, n=100, speed=1.0):
x_trajectory = []
y_trajectory = []
for i in range(n):
cte = robot.y
steer = -tau * cte
robot.move(steer, speed)
x_trajectory.append(robot.x)
y_trajectory.append(robot.y)
return x_trajectory, y_trajectory
# +
robot = Robot()
robot.set(0, 1, 0)
x_trajectory, y_trajectory = run(robot, 0.1)
n = len(x_trajectory)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(8, 8))
ax1.plot(x_trajectory, y_trajectory, 'g', label='P controller')
ax1.plot(x_trajectory, np.zeros(n), 'r', label='reference');
ax1.legend();
# -
# # PD Controller
def run(robot, tau_p, tau_d, n=100, speed=1.0):
x_trajectory = []
y_trajectory = []
prev_cte = robot.y
for i in range(n):
cte = robot.y
diff_cte = cte - prev_cte
prev_cte = cte
steer = -tau_p * cte - tau_d * diff_cte
robot.move(steer, speed)
x_trajectory.append(robot.x)
y_trajectory.append(robot.y)
return x_trajectory, y_trajectory
# +
robot = Robot()
robot.set(0, 1, 0)
x_trajectory, y_trajectory = run(robot, 0.2, 3.0)
n = len(x_trajectory)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(8, 8))
ax1.plot(x_trajectory, y_trajectory, 'g', label='PD controller')
ax1.plot(x_trajectory, np.zeros(n), 'r', label='reference')
ax1.legend();
# -
# # PID Controller
def run(robot, tau_p, tau_d, tau_i, n=100, speed=1.0):
x_trajectory = []
y_trajectory = []
prev_cte = robot.y
int_cte = 0
for i in range(n):
cte = robot.y
diff_cte = cte - prev_cte
prev_cte = cte
int_cte += cte
steer = -tau_p * cte - tau_d * diff_cte - tau_i * int_cte
robot.move(steer, speed)
x_trajectory.append(robot.x)
y_trajectory.append(robot.y)
return x_trajectory, y_trajectory
# +
robot = Robot()
robot.set(0, 1, 0)
x_trajectory, y_trajectory = run(robot, 0.2, 3.0, 0.004)
n = len(x_trajectory)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(8,8))
ax1.plot(x_trajectory, y_trajectory, 'g', label='PID controller')
ax1.plot(x_trajectory, np.zeros(n), 'r', label='reference')
# -
# # Final PID Controller
# NOTE: We use params instead of tau_p, tau_d, tau_i
def run(robot, params, n=100, speed=1.0):
x_trajectory = []
y_trajectory = []
err = 0
prev_cte = robot.y
int_cte = 0
for i in range(2 * n):
cte = robot.y
diff_cte = cte - prev_cte
int_cte += cte
prev_cte = cte
steer = -params[0] * cte - params[1] * diff_cte - params[2] * int_cte
robot.move(steer, speed)
x_trajectory.append(robot.x)
y_trajectory.append(robot.y)
if i >= n:
err += cte ** 2
return x_trajectory, y_trajectory, err / n
# Make this tolerance bigger if you are timing out!
def twiddle(tol=0.2):
p = [0, 0, 0]
dp = [1, 1, 1]
robot = make_robot()
x_trajectory, y_trajectory, best_err = run(robot, p)
it = 0
while sum(dp) > tol:
print("Iteration {}, best error = {}".format(it, best_err))
for i in range(len(p)):
p[i] += dp[i]
robot = make_robot()
x_trajectory, y_trajectory, err = run(robot, p)
if err < best_err:
best_err = err
dp[i] *= 1.1
else:
p[i] -= 2 * dp[i]
robot = make_robot()
x_trajectory, y_trajectory, err = run(robot, p)
if err < best_err:
best_err = err
dp[i] *= 1.1
else:
p[i] += dp[i]
dp[i] *= 0.9
it += 1
return p
# +
params, err = twiddle()
print("Final twiddle error = {}".format(err))
robot = make_robot()
x_trajectory, y_trajectory, err = run(robot, params)
n = len(x_trajectory)
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(8, 8))
ax1.plot(x_trajectory, y_trajectory, 'g', label='Twiddle PID controller')
ax1.plot(x_trajectory, np.zeros(n), 'r', label='reference')
| PID Control Python/PID Control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-LOOSibGrXR_" colab_type="text"
# # Movie review classification by universal sentence encoder with 200 samples
# + [markdown] id="h_EniEMLoh65" colab_type="text"
# ## 1.Import library
# + id="ifFCFn_K93bL" colab_type="code" outputId="3857215f-c53d-4212-cb4d-8c5f2189e89a" colab={"base_uri": "https://localhost:8080/", "height": 50}
import tensorflow as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import re
import time
import seaborn as sns
import keras
import keras.layers as layers
from keras.models import Model
from keras.models import Sequential
from keras.layers import Flatten, Dense
from keras.optimizers import RMSprop,Adam
from keras import backend as K
np.random.seed(10)
keras.__version__
tf.__version__
# + [markdown] id="feCgQ0L893bb" colab_type="text"
# ## 2. Download the IMDB data as raw text
# + id="XqebCXgf-fIl" colab_type="code" colab={}
from tensorflow import keras
# Load all files from a directory in a DataFrame.
def load_directory_data(directory):
data = {}
data["sentence"] = []
data["sentiment"] = []
for file_path in os.listdir(directory):
with tf.gfile.GFile(os.path.join(directory, file_path), "r") as f:
data["sentence"].append(f.read())
data["sentiment"].append(re.match("\d+_(\d+)\.txt", file_path).group(1))
return pd.DataFrame.from_dict(data)
# Merge positive and negative examples, add a polarity column and shuffle.
def load_dataset(directory):
pos_df = load_directory_data(os.path.join(directory, "pos"))
neg_df = load_directory_data(os.path.join(directory, "neg"))
pos_df["polarity"] = 1
neg_df["polarity"] = 0
return pd.concat([pos_df, neg_df]).sample(frac=1).reset_index(drop=True)
# Download and process the dataset files.
def download_and_load_datasets(force_download=False):
dataset = tf.keras.utils.get_file(
fname="aclImdb.tar.gz",
origin="http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz",
extract=True)
train_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "train"))
test_df = load_dataset(os.path.join(os.path.dirname(dataset),
"aclImdb", "test"))
return train_df, test_df
# + id="bW_eJ4PqHS4e" colab_type="code" colab={}
train, test = download_and_load_datasets()
# + [markdown] id="M9JwrDOwpg_x" colab_type="text"
# 2000 samples are taken
# + id="NXwixSXg_AEj" colab_type="code" outputId="0cc4b07d-16f6-4375-85e7-e9eb4dba6268" colab={"base_uri": "https://localhost:8080/", "height": 195}
n=2000
x = train.sample(n)
x.head()
# + [markdown] id="y3AsFFbkpJr7" colab_type="text"
# *Column* of "sentence" is used as features and "polarity" is used as label
# + id="G3M3tVX0BK7t" colab_type="code" outputId="abcafe3f-57ae-4ecb-fd5a-c4ac83a9c1f7" colab={"base_uri": "https://localhost:8080/", "height": 121}
X=x.values[:,0:1]
print(np.shape(X))
X=X.reshape(n,)
print(np.shape(X))
X=X.tolist()
print(type(X))
print(len(X))
print(X[0:3])
# + id="2q-o_ZruB0jh" colab_type="code" outputId="0566192d-d0a0-4096-ef91-50a7c012ffe7" colab={"base_uri": "https://localhost:8080/", "height": 84}
Y=x.values[:,2:3]
print(np.shape(Y))
Y=Y.reshape(n,)
print(np.shape(Y))
Y=Y.tolist()
print(type(Y))
print(Y[0:3])
# + [markdown] id="ZQNsL9nq93bl" colab_type="text"
# ## 3. Universal Sentence Encoder is used to extract features from texts
# + id="fpDPQND093bm" colab_type="code" colab={}
module_url = "https://tfhub.dev/google/universal-sentence-encoder-large/3"
# + id="44In5u0b93bq" colab_type="code" outputId="da7f0d2e-7095-4997-80c1-5ec1758a915d" colab={"base_uri": "https://localhost:8080/", "height": 104}
# Import the Universal Sentence Encoder's TF Hub module
embed = hub.Module(module_url)
# Compute a representation for X
messages = X
# Reduce logging output.
tf.logging.set_verbosity(tf.logging.ERROR)
with tf.Session() as session:
session.run([tf.global_variables_initializer(), tf.tables_initializer()])
message_embeddings = session.run(embed(messages))
for i, message_embedding in enumerate(np.array(message_embeddings).tolist()):
message_embedding_snippet = ", ".join(
(str(x) for x in message_embedding[:3]))
# + id="CNRJVtAR93bz" colab_type="code" outputId="91619ccd-9bda-4d62-8656-31e6814be975" colab={"base_uri": "https://localhost:8080/", "height": 50}
print(type(message_embeddings))
print(np.shape(message_embeddings))
# + [markdown] id="ojr8aCwe93b9" colab_type="text"
# ## 4. Develop model for classification
#
# + id="aBp_OHWv93cA" colab_type="code" outputId="f7758843-899a-437d-a7c8-62e2332edb0e" colab={"base_uri": "https://localhost:8080/", "height": 202}
X=message_embeddings
model = Sequential()
model.add(Dense(16, input_dim=512, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.summary()
# + [markdown] id="D3BbEeOE93cK" colab_type="text"
# ## 5. Train the model with 200 samples
#
# + id="F21OeI2l93cM" colab_type="code" outputId="f5f29197-50b5-4d8d-9629-956fb0cee992" colab={"base_uri": "https://localhost:8080/", "height": 1730}
t=time.time()
model.compile(optimizer=Adam(lr=1e-3),
loss='binary_crossentropy',
metrics=['acc'])
history = model.fit(X, Y,
epochs=50,
batch_size=32,
validation_split=0.9)
#model.save_weights('pre_trained_glove_model.h5')
t2 = time.time()
print(round(t2-t, 5), 'Seconds to predict')
# + id="exJhzCEazDVd" colab_type="code" outputId="2efec150-7755-42a2-cbb3-3a83a074e9d2" colab={"base_uri": "https://localhost:8080/", "height": 349}
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.figure(figsize=(15,5))
plt.subplot(1, 2, 1)
plt.plot(epochs, acc, label='Training acc')
plt.plot(epochs, val_acc, label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('epoch')
plt.ylabel('accuracy')
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(epochs, loss, label='Training loss')
plt.plot(epochs, val_loss, label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.legend()
plt.show()
| IMDB_with_fine_tuning_by_USE_20190227.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 欢迎去[这里](https://github.com/junjiecai/jupyter_labs/tree/master/cjj_notebooks/0014_python_pitfalls)下载到本文对应的jupyter notebook文件,亲自动手实验文中的代码。
import pandas as pd
import numpy as np
from pandas import DataFrame, Series
# ## float的精度问题
# float没办法覆盖所有的数值范围,会导致数值计算的不准确。
#
# 例如,float没法表示小数点后面任意的精度
# +
def func_1():
a = 1.13445433423e-300
return a**2
func_1()
# -
# 如果的确需要如此高的精度计算,可以使用decimal.Decimal这种数据类型。
# +
def func_2():
from decimal import Decimal
d = Decimal('1.13445433423e-300')
return d**2
func_2()
# -
# 不过作为代价,Decimal数值类型会消耗更多的计算时间。下面比较两个版本的代码各执行100000次,消耗的时间
from datetime import datetime
def timing(n, func, args, kargs):
t1 = datetime.now()
for i in range(n):
func(*args,**kargs)
t2 = datetime.now()
print((t2-t1).total_seconds())
timing(100000,func_1,[],{})
timing(100000,func_2,[],{})
# 可以看到,有接近7~8倍的差距
# float同样无法准确表达过大的数值,例如下面的例子
x = 123123123123123123123123
x = float(x)
int(x)
# 可以看到, 如果将大整数转化成float,再转回成int,数值上已经发生了偏差。 不过由于python的int类型本来就可以表示任意大的整数(只要内存允许),直接使用int类型进行计算即可。
# ## pandas, numpy与大数值int
# 一些pandas(或是底层用到的numpy)的函数并不支持数值较大的int
try:
np.isinf(62226003100113821696)
except Exception as e:
print(type(e),e)
try:
Series([99898989789797878797,99898989789797878797],index = [0,1]).value_counts()
except Exception as e:
print(type(e),e)
# 如果不想损失精度的话, 部分场景可以使用decimal.Decimal解决
from decimal import Decimal
try:
print(Series([Decimal('99898989789797878797'),Decimal('99898989789797878797')],index = [0,1]).value_counts())
except Exception as e:
print(type(e),e)
# 然而,并不是所有的情况都管用
try:
np.isinf(Decimal('62226003100113821696'))
except Exception as e:
print(type(e),e)
# 想上面这种情况, 就只能转成float损失精度了。(对于例子中的np.isinf来说,精度是否降低是无关紧要的)
try:
np.isinf(float('62226003100113821696'))
except Exception as e:
print(type(e),e)
# ## 操作系统默认encoding
# ubuntu下用open新建文件的时候,默认的encoding就是utf8,因此要往文件写入中文等字符都是没有问题的。但是windows的默认encoding不是utf8(似乎是ascii?有windows的同学可以试试),则需要额外的在open中制定encoding才能顺利写入特殊的字符。
#
# 大家可以在自己的操作系统下测试一下下面代码的结果。
import sys
print(sys.getdefaultencoding())
with open('test.txt',mode = 'w') as file:
file.write('测试')
# 可以通过encoding参数强制指定encoding
with open('test.txt',mode = 'w',encoding = 'utf') as file:
file.write('测试')
# ## cx_Oracle的arraysize
conn.arrayzie = 10000
print(conn.arrayzie)
# cx_Oracle的connection有一个arraysize控制每次从数据库中一次读取的数据行数, arraysize的设置必须发生在conn.execute(sql)之前,否则是不会生效的。
# (这个例子暂时不提供实验, 以后可能要配上docker实验环境)
# ## 条件判断的括号
# 下面的例子的结果估计大部分想象的都不太一样
# +
setA = {1}
setB = {2}
True if len(setA)==1 & len(setB)==1 & len(setA|setB)==2 else False
# -
# 如果不想去费无意义的脑力去记忆运算执行顺序规则的话, 多利用括号就好。
True if (len(setA)==1) & (len(setB)==1) & (len(setA|setB)==2) else False
# ## 眼见不为实
# 在jupyter中使用pandas时, jupyter会将dataframe用比较美观的方式的展现出来。 但是特别要注意的是, 一些不同数据类型的数据, 或者带有空格的字符串, 在jupyter中是看不到区别的,例如:
df1 = DataFrame({'A':[1,2,3],'B':['a','b','c']})
df2 = DataFrame({'A':['1','2','3'],'C':['aa','ba','ca']})
df1
df2
# 看上去A列一模一样, 但是如果merge的话, 是什么也得不到的。
df1.merge(df2, on = 'A')
# 因此, 使用DataFrame的时候一定要记得检查一下相关列的数据类型是什么。
# 同理, 如果字符中含有空格, 在jupyter中也不容易被观察出来。
s1 = Series(['a','b']).to_frame()
s2 = Series(['a ','b ']).to_frame()
s1
s2
# 进行数值比较的时候实际上会被判定为False
s1==s2
# 但是如果转成list然后print出来的话,就很容易看出空格的存在。
list(s1[0])
list(s2[0])
# ## 中英文符号
# 在写代码的时候最好关闭中文输入法, 避免混入中文或全角标点, 尤其是在编写正则表达式的时候。 例如, 我们希望根据'ab'或者'ad'去分割字符串
import re
string = 'abcadfa'
re.split('ab|ad',string)
re.split('ab|ad',string)
# 第一段代码没能生效的原因是'|'是在全角状态输入的。 像这样的问题是很难观察到的。
# ## 函数默认初始值
# mutable类型的数据不要作为函数的初始值,它并不会每次使用函数的时候被初始化,而是会记忆之前的运行状态。
# mutable作为默认初始值的问题
def test(a = []):
a.append('*')
print(a)
# 运行一次的时候,正常
test()
# 运行第二次的时候
test()
# 可以看到a并不是从[]开始运行的。
#
# 如果的确需要使用[]作为初始默认值,常见的做法是接受None作为参数后,在函数内部完成初始化。
def test(a = None):
a = a or []
a.append('*')
print(a)
test()
test()
| 0003_python_pitfalls/0003_python_pitfalls_ch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Converting Models to OpenVINO IR
# +
import os
from tensorflow.python.util import deprecation
deprecation._PRINT_DEPRECATION_WARNINGS = False
import warnings
import numpy as np
import tensorflow as tf
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
from tensorflow.python.framework import graph_util
from tensorflow.python.framework import graph_io
from keras.layers import *
from keras.models import load_model
import tensorflow.keras.backend as K
import shutil, sys
import torch
from pathlib import PurePath
from keras.models import model_from_json, Sequential
# -
# ## Section 1.1: Convert Keras Models
# +
vggModel = '../vgg.json'
vggWeights = '../vgg.h5'
mlpModel = '../gsbMLP.json'
mlpWeights = '../gsbMLP.h5'
jsonModelFilePath, weightsFilePath = mlpModel, mlpWeights
# jsonModelFilePath, weightsFilePath = vggModel, vggWeights
# load model
jsonModelFile = open(jsonModelFilePath, 'r' )
base_model = jsonModelFile.read()
jsonModelFile.close()
model = model_from_json(base_model)
model.load_weights(weightsFilePath)
model.compile( loss='binary_crossentropy', optimizer='sgd', metrics=[ 'accuracy' ] )
# -
# path_to_keras = str(assets_path.joinpath('fseg-60.hdf5'))
# model = load_model(path_to_keras, custom_objects={'cross_entropy_balanced': cross_entropy_balanced})
assets_path = PurePath('.')
frozen_model_path = str(assets_path)
frozen_model_name = 'mlp_sfd.pb'
# +
tf.get_logger().setLevel('INFO')
K.set_image_data_format('channels_last')
output_node_names = [node.op.name for node in model.outputs]
# -
sess = K.get_session()
constant_graph = graph_util.convert_variables_to_constants(sess,
sess.graph.as_graph_def(),
output_node_names)
graph_io.write_graph(constant_graph, frozen_model_path,
frozen_model_name, as_text=False)
# ## Section 1.2: Convert Tensorflow Frozen Graphs
# +
# os_version = 'open_seismic:v0.0.1'
# model_optimizer_cmd = f"""
# docker run {os_version} /bin/bash executables/mo.sh -h
# """
# # ! {model_optimizer_cmd}
# ! {"mo.py -h"}
# -
# Model expects a 5-dimensional input
model.input.get_shape().as_list()
# +
# phys_mnt_vol = str(PurePath(os.getcwd()))
# docker_mnt_vol = '/mnt_vol'
# docker_frozen_model_path = f'{docker_mnt_vol}/{frozen_model_name}'
# input_shape = str([1] + model.input.get_shape().as_list()[1:]).replace(" ", "")
# data_type = 'FP32'
# docker_output_model_path = f'{docker_mnt_vol}/IR_sdf/'
# ov_model_name = 'mlp_sdf'
# configs = f"--input_model {docker_frozen_model_path} \
# --input_shape {input_shape} \
# --data_type {data_type} \
# --disable_nhwc_to_nchw \
# --output_dir {docker_output_model_path} \
# --model_name {ov_model_name}"
# model_optimizer_docker_cmd = f"docker run -v {phys_mnt_vol}:{docker_mnt_vol} {os_version} /bin/bash executables/mo.sh {configs}"
# model_optimizer_cmd # Checking the command to see if this works
# +
# # ! {model_optimizer_docker_cmd}
# +
phys_mnt_vol = str(PurePath(os.getcwd()))
phys_frozen_model_path = f'{phys_mnt_vol}/{frozen_model_name}'
input_shape = str([1] + model.input.get_shape().as_list()[1:]).replace(" ", "")
data_type = 'FP32'
phys_output_model_path = f'{phys_mnt_vol}/IR_sdf_local/'
ov_model_name = 'mlp_sdf'
configs = f"--input_model {phys_frozen_model_path} \
--input_shape {input_shape} \
--data_type {data_type} \
--disable_nhwc_to_nchw \
--output_dir {phys_output_model_path} \
--model_name {ov_model_name}"
mo_local_cmd = f"mo.py {configs}"
mo_local_cmd
# -
# ! {mo_local_cmd}
# ## Testing the Model
from openvino.inference_engine import IECore
xml_path = './IR_sdf_local/mlp_sdf.xml'
bin_path = './IR_sdf_local/mlp_sdf.bin'
ie = IECore()
pre_model = ie.read_network(model=xml_path, weights=bin_path)
model = ie.load_network(network=pre_model, device_name="CPU", num_requests=1)
input_layer = next(iter(pre_model.input_info))
n, c, h, w = pre_model.input_info[input_layer].input_data.shape
n, c, h, w
list(pre_model.outputs.values())[0].shape
| models/tensorflow_conversion/keras2tf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Support Vector Regression/Machine for Humorous Level Prediction
#
# This program is modified from https://github.com/nkartik94/Multi-Label-Text-Classification
# on 2019/11/18 by <NAME>
# ## 1. EDA: Exploratory Data Analysis
import os, sys, time
import csv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
#printmd('**bold**')
# !cd /Users/sam/GoogleDrive/指導的學生/2019_吳玟萱/2019_1104_Joke_Datasets
# !pwd
data_path = "mlabel_corpora/JokeHumorLevel.txt"
# set global variables: df
df = pd.read_csv(data_path, delimiter="\t")
#data_raw = df.loc[np.random.choice(data_raw.index, size=2000)]
print(df.shape) # same as data_raw.shape in Jupyter
# +
from sklearn.model_selection import train_test_split
# ID=L1850 為分界,之前:吳玟萱,之後:黃亭筠,均為中文系同一屆
train, test = train_test_split(df, train_size=1691, shuffle=False)
# (tempararily) set global variables: train, test
with open('mlabel_corpora/JokeHumorLevel_train.txt', 'w') as outF:
outF.write(train.to_csv(sep='\t', index=False))
with open('mlabel_corpora/JokeHumorLevel_test.txt', 'w') as outF:
outF.write(test.to_csv(sep='\t', index=False))
print(train.shape)
print(test.shape)
# -
# Do not do this, because there are many duplicate titles
# Merge Title into Content
'''
df['Content'] = df[df.columns[1:3]].apply(
lambda x: ' 。 '.join(x.dropna().astype(str)),
axis=1
)
print("Number of rows in data =",df.shape[0])
print("Number of columns in data =",df.shape[1])
print("\n")
printmd("**Sample data:**")
df.head()
'''
# ### 1.1. Checking for missing values
missing_values_check = df.isnull().sum()
print(missing_values_check)
# ### 1.2. Calculating number of jokes under each label
# Jokes with no label are considered to be clean jokes.
# Creating seperate column in dataframe to identify clean jokes.
# We use axis=1 to count row-wise and axis=0 to count column wise
def print_empty_label(df, s):
rowSums = df.iloc[:,3:].sum(axis=1)
#print(rowSums.shape)
#print(rowSums.head())
clean_comments_count = (rowSums==0).sum(axis=0)
print(f"Total number of {s} jokes = ",len(df))
print(f"Number of clean jokes in {s}= ",clean_comments_count)
print(f"Number of {s} jokes with labels =",(len(df)-clean_comments_count))
print()
print_empty_label(df, 'all')
print_empty_label(train, 'train')
print_empty_label(test, 'test')
# set global variables: categories
categories = list(df.columns.values)
print(categories)
categories = categories[3:]
print(categories)
# Calculating number of humor levels
# https://stackoverflow.com/questions/45759966/counting-unique-values-in-a-column-in-pandas-dataframe-like-in-qlik
def print_HumorLevel_count(df, categories):
for c in categories:
print(df[c].value_counts())
#print(df[c].value_counts(normalize=True))
print()
print_HumorLevel_count(df, categories)
print_HumorLevel_count(train, categories)
print_HumorLevel_count(test, categories)
# Not correct yet!!!
def plot_HumorLevel_count(df, categories):
sns.set(font_scale = 2)
plt.figure(figsize=(15,8))
ax= sns.barplot(categories, df[categories[0]].value_counts())
plt.title("Humorous Level Count", fontsize=24)
plt.ylabel('Number of jokes', fontsize=18)
plt.xlabel('Humorous Level', fontsize=18)
#adding the text labels
rects = ax.patches
#print(rects)
labels = df[categories[0]].value_counts()
#print(labels)
for rect, label in zip(rects, labels):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width()/2, height + 5, label, ha='center', va='bottom', fontsize=18)
plt.show()
# +
#plot_HumorLevel_count(df, categories)
# -
# ## 2. Data Pre-Processing
import jieba
import Stopwords
# Compute statistics of the dataset: MaxLength, MinLength, AvgChars, AvgWords
Len = df.Content.map(len)
print(f'Number of characters in all jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
Len = train.Content.map(len)
print(f'Number of characters in train jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
Len = test.Content.map(len)
print(f'Number of characters in test jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
# set global variables: data
data = df
#data = df.loc[np.random.choice(df.index, size=3365)]
data.shape
# +
import nltk
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
import re
import sys
import warnings
if not sys.warnoptions:
warnings.simplefilter("ignore")
# -
# ### 2.1. Cleaning Data
# +
def cleanHtml(sentence):
cleanr = re.compile('<.*?>')
cleantext = re.sub(cleanr, ' ', str(sentence))
return cleantext
def cleanPunc(sentence): #function to clean the word of any punctuation or special characters
cleaned = re.sub(r'[?|!|\'|"|#]',r'',sentence)
cleaned = re.sub(r'[.|,|)|(|\|/]',r' ',cleaned)
cleaned = cleaned.strip()
cleaned = cleaned.replace("\n"," ")
return cleaned
def keepAlpha(sentence):
alpha_sent = ""
for word in sentence.split():
alpha_word = re.sub('[^a-z A-Z]+', ' ', word)
alpha_sent += alpha_word
alpha_sent += " "
alpha_sent = alpha_sent.strip()
return alpha_sent
# -
import Stopwords # import my own module with STOP_WORDS
from nltk.stem import PorterStemmer, WordNetLemmatizer
ps = PorterStemmer()
wnl = WordNetLemmatizer()
# +
def clean_text(text):
'''
Given a raw text string, return a clean text string.
Example:
input: "Years passed. 多少 年过 去 了 。 "
output: "years passed.多少年过去了。"
'''
text = str(text)
text = text.lower() # 'years passed. 多少 年过 去 了 。'
# Next line will remove redundant white space for jeiba to cut
text = re.sub(r'\s+([^a-zA-Z0-9.])', r'\1', text) # years passed.多少年过去了。
# see: https://stackoverflow.com/questions/16720541/python-string-replace-regular-expression
text = text.strip(' ')
return text
def clean_words(text, RmvStopWord=True, RmvMark=True):
words = jieba.lcut(text)
# print("After jieba.lcut():", words)
# WL = [ w
WL = [ ps.stem(w)
# WL = [ wnl.lemmatize(w)
for w in words
if (not re.match(r'\s', w)) # remove white spaces
and (RmvMark==False or not re.match(r'\W', w)) # remove punctuations
# and (RmvMark==False or not re.match('^[a-z_]$', w)) # remove punctuations
# and (RmvMark==False or w not in PUNCTUATIONS)
and (RmvStopWord==False or w not in Stopwords.STOP_WORDS)
and (not re.match(r'^\d+$', w)) # remove digit
]
WL = " ".join(WL)
return WL
# -
print(data.head())
data['Content'] = data['Content'].str.lower()
#data['Content'] = data['Content'].apply(cleanHtml)
#data['Content'] = data['Content'].apply(cleanPunc)
#data['Content'] = data['Content'].apply(keepAlpha)
data['Content'] = data['Content'].apply(clean_text)
data['Content'] = data['Content'].apply(clean_words)
#data.head()
# ### 2.2. Removing Stop Words
# +
stop_words = set(stopwords.words('english'))
stop_words.update(['zero','one','two','three','four','five','six','seven','eight','nine','ten','may','also','across','among','beside','however','yet','within'])
re_stop_words = re.compile(r"\b(" + "|".join(stop_words) + ")\\W", re.I)
def removeStopWords(sentence):
global re_stop_words
return re_stop_words.sub(" ", sentence)
data['Content'] = data['Content'].apply(removeStopWords)
#data.head()
# -
# ### 2.3. Stemming
# +
stemmer = SnowballStemmer("english")
def stemming(sentence):
stemSentence = ""
for word in sentence.split():
stem = stemmer.stem(word)
stemSentence += stem
stemSentence += " "
stemSentence = stemSentence.strip()
return stemSentence
data['Content'] = data['Content'].apply(stemming)
data.head()
# -
# ### 2.4. Train-Test Split
# +
from sklearn.model_selection import train_test_split
# set global variables: train, test
#train, test = train_test_split(data, random_state=42, test_size=0.10, shuffle=True)
train, test = train_test_split(data, random_state=42, train_size=1691, shuffle=False)
print(train.shape)
print(test.shape)
# -
# set global variables: train_text, test_text
train_text = train['Content']
test_text = test['Content']
# Compute statistics of the dataset: MaxLength, MinLength, AvgChars, AvgWords
Len = data.Content.map(lambda x: len(x.split()))
print(f'Number of words in all jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
Len = train.Content.map(lambda x: len(x.split()))
print(f'Number of words in train jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
Len = test.Content.map(lambda x: len(x.split()))
print(f'Number of words in test jokes: Max={max(Len)}, Min={min(Len)}, Avg={sum(Len)/len(Len)}')
# ### 2.6 TF-IDF
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm
#vectorizer = TfidfVectorizer(strip_accents='unicode', analyzer='word', ngram_range=(1,3), norm='l2')
vectorizer = TfidfVectorizer(strip_accents='unicode', analyzer='word',
ngram_range=(1,2), norm='l2')
vectorizer.fit(train_text)
vectorizer.fit(test_text)
# +
# Set global variables:
x_train = vectorizer.transform(train_text)
y_train = train.drop(labels = ['ID', 'Title', 'Content'], axis=1)
#print(y_train.head())
train_yL = y_train['HumorLevel']
#print(type(train_yL), "\n", train_yL.head())
x_test = vectorizer.transform(test_text)
y_test = test.drop(labels = ['ID', 'Title', 'Content'], axis=1)
test_yL = y_test['HumorLevel']
# label encode the target variable
# http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html
LabEncoder = preprocessing.LabelEncoder() # convert label name to label int
train_y = LabEncoder.fit_transform(train_yL)
test_y = LabEncoder.fit_transform(test_yL)
Num_Classes = len(LabEncoder.classes_)
# +
time_TfidfVector = time.time()
def Create_TFxIDF(data_text, train_text, test_text):
# word level tf-idf
#tfidf_vect = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}', max_features=10000)
tfidf_vect = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}',
stop_words=Stopwords.STOP_WORDS, max_df=0.95, min_df=1, max_features=10000)
tfidf_vect.fit(data_text)
xtrain_tfidf = tfidf_vect.transform(train_text)
xtest_tfidf = tfidf_vect.transform(test_text)
print(f"xtrain_tfidf.shape:{xtrain_tfidf.shape}, xtest_tfidf.shape: {xtest_tfidf.shape}")
# word level ngram tf-idf
tfidf_vect_ngram = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}',
stop_words=Stopwords.STOP_WORDS, max_df=0.95, min_df=1,
ngram_range=(1,3), max_features=10000)
tfidf_vect_ngram.fit(data_text)
xtrain_tfidf_ngram = tfidf_vect_ngram.transform(train_text)
xtest_tfidf_ngram = tfidf_vect_ngram.transform(test_text)
print(f"xtrain_tfidf_ngram.shape:{xtrain_tfidf.shape}, xtest_tfidf_ngram.shape: {xtest_tfidf.shape}")
# character level ngram tf-idf
tfidf_vect_ngram_chars = TfidfVectorizer(analyzer='char', token_pattern=r'\w{1,}',
stop_words=Stopwords.STOP_WORDS, max_df=0.95, min_df=1,
ngram_range=(1,3), max_features=10000)
tfidf_vect_ngram_chars.fit(data_text)
xtrain_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(train_text)
xtest_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(test_text)
print(f"xtrain_tfidf_ngram_chars.shape:{xtrain_tfidf.shape}, xtest_tfidf_ngram_chars.shape: {xtest_tfidf.shape}")
print("It takes %4.2f seconds to convert 3 TFxIDF vectors."%(time.time()-time_TfidfVector))
return (xtrain_tfidf, xtest_tfidf,
xtrain_tfidf_ngram, xtest_tfidf_ngram,
xtrain_tfidf_ngram_chars, xtest_tfidf_ngram_chars,
tfidf_vect, tfidf_vect_ngram, tfidf_vect_ngram_chars)
# Set global variables:
(xtrain_tfidf, xtest_tfidf,
xtrain_tfidf_ngram, xtest_tfidf_ngram,
xtrain_tfidf_ngram_chars, xtest_tfidf_ngram_chars,
tfidf_vect, tfidf_vect_ngram, tfidf_vect_ngram_chars) = Create_TFxIDF(data.Content, train_text, test_text)
# -
# re-assign x_train and x_test to what we want
#x_train, x_test, vectorizer = xtrain_tfidf, xtest_tfidf, tfidf_vect
#x_train, x_test, vectorizer = xtrain_tfidf_ngram, xtest_tfidf_ngram, tfidf_vect_ngram
#x_train, x_test, vectorizer = xtrain_tfidf_ngram_chars, xtest_tfidf_ngram_chars, tfidf_vect_ngram_chars
print(x_train.shape, x_test.shape)
#print(x_train)
# ### 2.6 Count Vector
# +
time_CountVector = time.time()
def Create_CountVector(data_text, train_text, test_text):
# Create a count vectorizer object.
# It takes the steps of prepocessing, tokenizer, stopwording, ...
# http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html
count_vect = CountVectorizer(analyzer='word', token_pattern=r'\w{1,}',
stop_words=Stopwords.STOP_WORDS, max_df=0.95, min_df=1)
count_vect.fit(data_text)
# transform the training and validation data using count vectorizer object
xtrain_count = count_vect.transform(train_text)
xtest_count = count_vect.transform(test_text)
print("It takes %4.2f seconds to convert count vectors."%(time.time()-time_CountVector))
return(xtrain_count, xtest_count, count_vect)
# Set global variables:
(xtrain_count, xtest_count, count_vect) = Create_CountVector(data.Content, train_text, test_text)
# +
def Print_count_vect(xtrain_count, xtest_count, count_vect):
print(type(count_vect), count_vect)
print(type(xtrain_count), type(xtest_count))
print("xtrain_count.shape:", xtrain_count.shape)
print("xtest_count.shape :", xtest_count.shape)
# https://stackoverflow.com/questions/36967666/transform-scipy-sparse-csr-to-pandas
# from scipy.sparse.csr import csr_matrix
# A = csr_matrix([[1, 0, 2], [0, 3, 0]]); print(A)
# df = pd.DataFrame(A.toarray()); print(df)
#print(xtrain_count)
#print(xtest_count[0, 0:10])
print("\nUsed stop words: ", count_vect.get_stop_words())
Print_count_vect(xtrain_count, xtest_count, count_vect)
# -
# ## 3. Support Vector Regression
# +
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import Pipeline
from sklearn.metrics import mean_squared_error, mean_absolute_error, median_absolute_error
from sklearn.metrics import explained_variance_score, r2_score
# -
# define the evaluation metrics
def print_svr_report(y_true, prediction):
#print('max_error: %1.4f'%(max_error(y_true, prediction)))
print('mean_squared_error: %1.4f'%(mean_squared_error(y_true, prediction)))
print('mean_absolute_error: %1.4f'%(mean_absolute_error(y_true, prediction)))
print('r2_score: %1.4f'%(r2_score(y_true, prediction)))
print('explained_variance_score: %1.4f'%(explained_variance_score(y_true, prediction)))
print(f'y_true.shape={y_true.shape}, prediction.shape={prediction.shape}')
#type(y_true<class 'pandas.core.frame.DataFrame'>, type(prediction)=<class 'numpy.ndarray'>
print(f'type(y_true{type(y_true)}, type(prediction)={type(prediction)}')
#pred = pd.DataFrame(data=prediction, columns=[y_true.columns]) # cannot have the same column name with y_true
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.reset_index.html
pred = pd.DataFrame(data=y_true.reset_index(drop=True), columns=['HumorLevel'])
pred['Funiness'] = prediction
pred['FuninessLevel'] = pred['Funiness'].map(round)
#print('y_true.head() :\n', y_true.head())
print('pred.head() :\n', pred.head())
print(y_true[y_true.columns[0]].value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
print(pred.FuninessLevel.value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
# https://stackoverflow.com/questions/52777668/python-pandas-compare-two-columns-for-equality-and-result-in-third-dataframe
pred['result'] = np.where(pred['HumorLevel'] == pred['FuninessLevel'], 1, 0)
print(pred['result'].value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
print(pred['result'].value_counts(normalize=True, sort=False, ascending=False, bins=None, dropna=True))
# Next line refers to: http://scikit.ml/tutorial.html
from sklearn.svm import SVR, LinearSVR
# Next line refers to https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(x_train, y_train)
print("reg.score:", reg.score(x_train, y_train))
predictions = reg.predict(x_test)
print_svr_report(y_test, predictions)
# +
# %%time
# https://scikit-learn.org/stable/modules/svm.html#regression
classifier = LinearSVR()
classifier.fit(x_train, y_train)
predictions = classifier.predict(x_test)
print_svr_report(y_test, predictions)
# +
# %%time
# https://scikit-learn.org/stable/modules/svm.html#regression
classifier = SVR()
classifier.fit(x_train, y_train)
predictions = classifier.predict(x_test)
print_svr_report(y_test, predictions)
# +
with open('out/HumorLevel_True.txt', 'w') as outF:
outF.write(y_test.to_csv(sep='\t', index=False))
# https://stackoverflow.com/questions/36967666/transform-scipy-sparse-csr-to-pandas
with open('out/HumorLevel_Pred.txt', 'w') as outF:
outF.write(pd.DataFrame(predictions, columns=list(y_test.columns)).to_csv(sep='\t', index=False))
# -
# ## 4. use classification for prediction Humorous Level
# ### 4.1 Define metrics
def train_predict(classifier, feature_vector_train, label, feature_vector_test):
# fit the training dataset on the classifier
classifier.fit(feature_vector_train, label)
# predict the labels on test dataset
return classifier.predict(feature_vector_test), classifier
def tcfunc(x, n=4): # trancate a number to have n decimal digits
d = '0' * n
d = int('1' + d)
# https://stackoverflow.com/questions/4541155/check-if-a-number-is-int-or-float
if isinstance(x, (int, float)): return int(x * d) / d
return x
# http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
#import itertools # replace this line by next line on 2019/01/03, because cannot find itertools for Python 3.6.7
import more_itertools
import matplotlib.pyplot as plt
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, numpy.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
#print(cm) # print out consufion matrix
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = numpy.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
# for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
# Replace the above line by the next line on 2019/01/03, because cannot find itertools for Python 3.6.7
for i, j in more_itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# use global variables:
# test_y
# LabEncoder.classes_
def show_confusion_matrix(predictions):
# Compute confusion matrix
cnf_matrix = confusion_matrix(test_y, predictions)
numpy.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=LabEncoder.classes_ ,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=LabEncoder.classes_ , normalize=True,
title='Normalized confusion matrix')
plt.show()
# +
# http://scikit-learn.org/stable/modules/model_evaluation.html
from sklearn.metrics import precision_recall_fscore_support
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
# use a global variable: test_y
def show_Result(predictions):
print(predictions[:10])
# http://scikit-learn.org/stable/modules/generated/sklearn.metrics.f1_score.html
# print("MicroF1 = %0.4f, MacroF1=%0.4f" %
# (metrics.f1_score(test_y, predictions, average='micro'),
# metrics.f1_score(test_y, predictions, average='macro')))
# https://stackoverflow.com/questions/455612/limiting-floats-to-two-decimal-points
# http://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_fscore_support.html
print("\tPrecision\tRecall\tF1\tSupport")
(Precision, Recall, F1, Support) = list(map(tcfunc,
precision_recall_fscore_support(test_y, predictions, average='micro')))
print("Micro\t{}\t{}\t{}\t{}".format(Precision, Recall, F1, Support))
(Precision, Recall, F1, Support) = list(map(tcfunc,
precision_recall_fscore_support(test_y, predictions, average='macro')))
print("Macro\t{}\t{}\t{}\t{}".format(Precision, Recall, F1, Support))
if True:
#if False:
print(confusion_matrix(test_y, predictions))
try:
print(classification_report(test_y, predictions, digits=4))
except ValueError:
print('May be some category has no predicted samples')
show_confusion_matrix(predictions)
# https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.reset_index.html
y_true = pd.DataFrame(test_y, columns=['HumorLevel'])
pred = pd.DataFrame(data=y_true.reset_index(drop=True), columns=['HumorLevel'])
pred['FuninessLevel'] = predictions
print(y_true[y_true.columns[0]].value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
print(pred.FuninessLevel.value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
# https://stackoverflow.com/questions/52777668/python-pandas-compare-two-columns-for-equality-and-result-in-third-dataframe
pred['result'] = np.where(pred['HumorLevel'] == pred['FuninessLevel'], 1, 0)
print(pred['result'].value_counts(normalize=False, sort=False, ascending=False, bins=None, dropna=True))
print(pred['result'].value_counts(normalize=True, sort=False, ascending=False, bins=None, dropna=True))
# +
# This function is modified from: https://gist.github.com/bbengfort/044682e76def583a12e6c09209c664a1
# and from: https://stackoverflow.com/questions/26976362/how-to-get-most-informative-features-for-scikit-learn-classifier-for-different-c
# This function only works for binary classes
def most_informative_feature_for_class(vectorizer, classifier, labels, n=10):
coefs = sorted( # Zip the feature names with the coefs and sort
zip(classifier.coef_[0], vectorizer.get_feature_names()))
topn = zip(coefs[:n], coefs[:-(n+1):-1])
# Create two columns with most negative and most positive features.
for (cp, fnp), (cn, fnn) in topn:
print("\t%.4f\t%-15s\t\t%.4f\t%-15s" % (cp, fnp, cn, fnn))
# nltk.classify.NaiveBayesClassifier has a show_most_informative_features()
# You may compare the result here with those at: https://www.twilio.com/blog/2017/09/sentiment-analysis-python-messy-data-nltk.html
# -
# ### 4.2 Run classifiers
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm
from sklearn import decomposition, ensemble
# +
def Run_NaiveBayes():
time_NaiveBayes = time.time()
# Naive Bayes on Count Vectors
predict, clf = train_predict(naive_bayes.MultinomialNB(), xtrain_count, train_y, xtest_count)
print("\nNB, Count Vectors: ")
show_Result(predict)
most_informative_feature_for_class(count_vect, clf, train_yL, n=10)
# Naive Bayes on Word Level TF IDF Vectors
predict, clf = train_predict(naive_bayes.MultinomialNB(), xtrain_tfidf, train_y, xtest_tfidf)
print("\nNB, WordLevel TF-IDF: ")
show_Result(predict)
#most_informative_feature_for_class(vectorizer, classifier, classlabel, n=10)
most_informative_feature_for_class(tfidf_vect, clf, train_y, n=10)
# Naive Bayes on Ngram Level TF IDF Vectors
predict, clf = train_predict(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram, train_y, xtest_tfidf_ngram)
print("\nNB, N-Gram Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram, clf, train_y, n=10)
# Naive Bayes on Character Level TF IDF Vectors
predict, clf = train_predict(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram_chars, train_y, xtest_tfidf_ngram_chars)
print("NB, CharLevel Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram_chars, clf, train_y, n=10)
print("\nIt takes %4.2f seconds for Naive Bayes."%(time.time()-time_NaiveBayes))
Run_NaiveBayes()
# +
def Run_LogisticRegret():
time_LogisticRegret = time.time()
# Linear Classifier on Count Vectors
predict, clf = train_predict(linear_model.LogisticRegression(), xtrain_count, train_y, xtest_count)
print("\nLR, Count Vectors: ")
show_Result(predict)
most_informative_feature_for_class(count_vect, clf, train_y, n=10)
# Linear Classifier on Word Level TF IDF Vectors
predict, clf = train_predict(linear_model.LogisticRegression(), xtrain_tfidf, train_y, xtest_tfidf)
print("\nLR, WordLevel TF-IDF: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect, clf, train_y, n=10)
# Linear Classifier on Ngram Level TF IDF Vectors
predict, clf = train_predict(linear_model.LogisticRegression(), xtrain_tfidf_ngram, train_y, xtest_tfidf_ngram)
print("\nLR, N-Gram Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram, clf, train_y, n=10)
# Linear Classifier on Character Level TF IDF Vectors
predict, clf = train_predict(linear_model.LogisticRegression(), xtrain_tfidf_ngram_chars, train_y, xtest_tfidf_ngram_chars)
print("\nLR, CharLevel Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram_chars, clf, train_y, n=10)
print("\nIt takes %4.2f seconds for Logistic Regression."%(time.time()-time_LogisticRegret))
Run_LogisticRegret()
# +
def Run_SVM():
time_LinearSVM = time.time()
# Use of class_weight='balanced' decrease accuracy, although PCWeb is unbalanced
#accuracy = train_model(svm.SVC(class_weight='balanced'), xtrain_count, train_y, xtest_count)
# LinearSVC() is much much better than SVC()
predict, clf = train_predict(svm.LinearSVC(), xtrain_count, train_y, xtest_count)
print("\nSVM, Count Vectors: ")
show_Result(predict)
most_informative_feature_for_class(count_vect, clf, train_y, n=10)
predict, clf = train_predict(svm.LinearSVC(), xtrain_tfidf, train_y, xtest_tfidf)
print("\nSVM, WordLevel TF-IDF: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect, clf, train_y, n=10)
predict, clf = train_predict(svm.LinearSVC(), xtrain_tfidf_ngram, train_y, xtest_tfidf_ngram)
print("\nSVM, N-Gram Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram, clf, train_y, n=10)
predict, clf = train_predict(svm.LinearSVC(), xtrain_tfidf_ngram_chars, train_y, xtest_tfidf_ngram_chars)
print("\nSVM, CharLevel Vectors: ")
show_Result(predict)
most_informative_feature_for_class(tfidf_vect_ngram_chars, clf, train_y, n=10)
print("\nIt takes %4.2f seconds for Linear SVM."%(time.time()-time_LinearSVM))
Run_SVM()
# +
def Run_RdnForest():
time_RdnForest = time.time()
# RF on Count Vectors
predict, clf = train_predict(ensemble.RandomForestClassifier(), xtrain_count, train_y, xtest_count)
print("\nRF, Count Vectors: ")
show_Result(predict)
#most_informative_feature_for_class(count_vect, clf, train_y, n=10)
#'RandomForestClassifier' object has no attribute 'coef_'
# RF on Word Level TF IDF Vectors
predict, clf = train_predict(ensemble.RandomForestClassifier(), xtrain_tfidf, train_y, xtest_tfidf)
print("\nRF, WordLevel TF-IDF: ")
show_Result(predict)
predict, clf = train_predict(ensemble.RandomForestClassifier(), xtrain_tfidf_ngram, train_y, xtest_tfidf_ngram)
print("\nRF, N-Gram Vectors: ")
show_Result(predict)
predict, clf = train_predict(ensemble.RandomForestClassifier(), xtrain_tfidf_ngram_chars, train_y, xtest_tfidf_ngram_chars)
print("\nRF, CharLevel Vectors: ")
show_Result(predict)
print("\nIt takes %4.2f seconds for Random Forest."%(time.time()-time_RdnForest))
Run_RdnForest()
# -
| tools/JokeHumorLevel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
from fastai import *
from fastai.vision import *
# -
# # Dogs and cats
# ## Resnet 34
path = untar_data(URLs.DOGS)
path
data = ImageDataBunch.from_folder(path, ds_tfms=get_transforms(), size=224).normalize(imagenet_stats)
img,label = data.valid_ds[-1]
img.show(title=data.classes[label])
learn = create_cnn(data, models.resnet34, metrics=accuracy)
learn.fit_one_cycle(1)
learn.unfreeze()
learn.fit_one_cycle(6, slice(1e-5,3e-4), pct_start=0.05)
accuracy(*learn.TTA())
# ## rn50
learn = create_cnn(data, models.resnet50, metrics=accuracy)
learn.fit_one_cycle(6)
learn.unfreeze()
learn.fit_one_cycle(6, slice(1e-5,3e-4), pct_start=0.05)
accuracy(*learn.TTA())
| examples/dogs_cats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import math
print(math.tanh(-2.2)) #Garbage truck
print(math.tanh(.1)) #Bear
print(math.tanh(2.5)) #Good doggo
import torch.nn as nn
import torch.optim as optim
# Using __call__ rather than forward
# +
#. __call__
# y = model(x) # correct
# y = model.forward(x) # Silent error. Don’t do it!
# -
linear_model = nn.Linear(1, 2)
input = torch.randn(2, 1)
output = linear_model(input)
print(output.size())
# +
# 6.2.2 linear model
# nn.Linear accepts three arguments: the number of input features, the number of output features, and whether the linear
# model includes a bias or not (defaulting to True, here):
linear_model = nn.Linear(1, 1) #y = xA^T + b
t_un_val = torch.randn(2,1)
linear_model(t_un_val)
# -
linear_model.weight
linear_model.bias
x = torch.ones(1)
linear_model(x)
# +
# BATCHING INPUTS
# create an input tensor of size B × Nin,
# where B is the size of the batch and Nin is the number of input features
x = torch.ones(10,1)
linear_model(x)
# +
t_c = [0.5, 14.0, 15.0, 28.0, 11.0, 8.0, 3.0, -4.0, 6.0, 13.0, 21.0]
t_u = [35.7, 55.9, 58.2, 81.9, 56.3, 48.9, 33.9, 21.8, 48.4, 60.4, 68.4]
t_c = torch.tensor(t_c).unsqueeze(1) # Adds the extra dimension at axis 1,在索引1添加额外的维度将输入变换
t_u = torch.tensor(t_u).unsqueeze(1)
print(t_u.shape)
t_u
# +
n_samples = t_u.shape[0]
n_val = int(0.2 * n_samples)
shuffled_indices = torch.randperm(n_samples)
train_indices = shuffled_indices[:-n_val]
val_indices = shuffled_indices[-n_val:]
train_indices, val_indices
# -
# 训练集
train_t_u = t_u[train_indices]
train_t_c = t_c[train_indices]
# 验证集
val_t_u = t_u[val_indices]
val_t_c = t_c[val_indices]
#输入标准化处理
train_t_un = 0.1 * train_t_u
val_t_un = 0.1 * val_t_u
linear_model = nn.Linear(1,1)
optimizer = optim.SGD(linear_model.parameters(),
lr = 1e-2)
linear_model.parameters()
list(linear_model.parameters())
def training_loop(n_epochs, optimizer, model, loss_fn, t_u_train, t_u_val,t_c_train, t_c_val):
for epoch in range(1, n_epochs + 1):
t_p_train = model(t_u_train)
loss_train = loss_fn(t_p_train, t_c_train)
t_p_val = model(t_u_val)
loss_val = loss_fn(t_p_val, t_c_val)
optimizer.zero_grad()
loss_train.backward()
optimizer.step()
if epoch == 1 or epoch % 1000 == 0:
print(f"Epoch {epoch}, Training loss {loss_train.item():.4f},"
f" Validation loss {loss_val.item():.4f}")
training_loop(
n_epochs = 3000,
optimizer = optimizer,
model = linear_model,
loss_fn = nn.MSELoss(),
t_u_train = train_t_un,
t_u_val = val_t_un,
t_c_train = train_t_c,
t_c_val = val_t_c)
print(linear_model.weight)
print(linear_model.bias)
# +
# seq model
seq_model = nn.Sequential(
nn.Linear(1, 13),
nn.Tanh(),
nn.Linear(13, 1))
seq_model
# -
[param.shape for param in seq_model.parameters()]# 检查参数
# model.parameters() will collect weight and bias from both the first and second linear modules
# identify parameters by name,通过名字区分参数,调用named_parameters
for name, param in seq_model.named_parameters():
print(name, param.shape)
# +
from collections import OrderedDict
# 有序字典显示model
seq_model = nn.Sequential(OrderedDict([
('hidden_linear', nn.Linear(1, 8)),#隐藏层linear
('hidden_activation', nn.Tanh()),#隐藏层激活函数tanh
('output_linear', nn.Linear(8, 1))#输出层linear
]))
seq_model
# -
# 查看更多子模型的解释信息
for name, param in seq_model.named_parameters():
print(name, param.shape)
# 查看具体一层的参数值
seq_model.output_linear.bias
# 训练集
train_t_u2 = t_u[train_indices]
train_t_c2 = t_c[train_indices]
# 验证集
val_t_u2 = t_u[val_indices]
val_t_c2 = t_c[val_indices]
#输入标准化处理
train_t_un2 = 0.1 * train_t_u
val_t_un2 = 0.1 * val_t_u
# +
# 在最终轮之后查看的梯度结果
optimizer = optim.SGD(seq_model.parameters(), lr=1e-3)
training_loop(
n_epochs = 5000,
optimizer = optimizer,
model = seq_model,
loss_fn = nn.MSELoss(),
t_u_train = train_t_un2,
t_u_val = val_t_un2,
t_c_train = train_t_c2,
t_c_val = val_t_c2)
print('output', seq_model(val_t_un2))
print('answer', val_t_c2)
print('hidden', seq_model.hidden_linear.weight.grad)
# +
# 6.3.3 和线性模型比较
from matplotlib import pyplot as plt
t_range = torch.arange(20., 90.).unsqueeze(1)
fig = plt.figure(dpi=600)
plt.xlabel("Fahrenheit")
plt.ylabel("Celsius")
plt.plot(t_u.numpy(), t_c.numpy(), 'o')
plt.plot(t_range.numpy(), seq_model(0.1 * t_range).detach().numpy(), 'c-')
plt.plot(t_u.numpy(), seq_model(0.1 * t_u).detach().numpy(), 'kx')
# -
| chap6/chap6_v0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pre-process input data for coastal variable extraction
#
# Author: <NAME>; <EMAIL>
#
# ***
#
# Pre-process files to be used in extractor.ipynb (Extract barrier island metrics along transects). See the project [README](https://github.com/esturdivant-usgs/BI-geomorph-extraction/blob/master/README.md) and the Methods Report (Zeigler et al., in review).
#
#
# ## Pre-processing steps
#
# 1. Pre-created geomorphic features: dunes, shoreline points, armoring.
# 2. Inlets
# 3. Shoreline
# 4. Transects - extend and sort
# 5. Transects - tidy
#
#
# ## Notes:
# This process requires some manipulation of the spatial layers by the user. When applicable, instructions are described in this file.
#
# ***
#
# ## Import modules
import os
import sys
import pandas as pd
import numpy as np
import arcpy
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
import core.functions_warcpy as fwa
import core.functions as fun
from core.setvars import *
# ### Initialize variables
#
# Change the filename variables to match your local files. They should be in an Esri file geodatabase named site+year.gdb in your project directory, which will be the value of the variable `home`.
#
# Input the site, year, and project directory path. `setvars.py` retrieves the pre-determined values for that site in that year from `configmap.py`. The project directory will be used to set up your workspace. It's hidden for security – sorry! I recommend that you type the path somewhere and paste it in.
# +
# Inputs - vector
orig_trans = os.path.join(home, 'trans_orig')
# Extended transects: NASC transects extended and sorted, ready to be the base geometry for processing
extendedTrans = os.path.join(home, 'Monomoy2014_extTrans_null')
# Tidied transects: Extended transects without overlapping transects
extTrans_tidy = os.path.join(home, 'Monomoy_tidyTrans')
# Geomorphology points: positions of indicated geomorphic features
ShorelinePts = os.path.join(home, 'Monomoy2014_SLpts') # shoreline
dlPts = os.path.join(home, 'Monomoy2014_DLpts') # dune toe
dhPts = os.path.join(home, 'Monomoy2014_DHpts') # dune crest
# Inlet lines: polyline feature classes delimiting inlet position. Must intersect the full island shoreline
inletLines = os.path.join(home, 'Monomoy2014_inletLines')
# Full island shoreline: polygon that outlines the island shoreline, MHW on oceanside and MTL on bayside
barrierBoundary = os.path.join(home, 'bndpoly_manual_2sl')
# Elevation grid: DEM of island elevation at either 5 m or 1 m resolution
elevGrid = os.path.join(home, 'Monomoy2014_DEM_5m')
# ---
# OPTIONAL - comment out each one that is not available
# ---
# Study area boundary; manually digitize if the barrier island study area does not end at an inlet.
# SA_bounds = os.path.join(home, 'SA_bounds')
# Armoring lines: digitize lines of shorefront armoring to be used if dune toe points are not available.
# armorLines = os.path.join(home, 'armorLines')
# -
# ## Prepare input layers
# +
datapre = '14CNT01'
csvpath = os.path.join(proj_dir, 'Input_Data', '{}_morphology'.format(datapre), '{}_morphology.csv'.format(datapre))
# state = sitevals['state']
dt_fc, dc_fc, sl_fc = fwa.MorphologyCSV_to_FCsByFeature(csvpath, state, proj_code, csv_fill = 999, fc_fill = -99999, csv_epsg=4326)
# -
# ### Dunes
# Display the points and the DEM in a GIS to check for irregularities. For example, if shoreline points representing a distance less than X m are visually offset from the general shoreline, they should likely be removed. Another red flag is when the positions of dlows and dhighs in relation to the shore are illogical, i.e. dune crests are seaward of dune toes. Address these irregularities by manually deleting points. Delete conservatively.
# ### Armoring
# If the dlows do not capture the entire top-of-beach due to atypical formations caused by anthropogenic modification, you may need to digitize the beachfront armoring. The next code block will generate an empty feature class. Refer to the DEM and orthoimagery. If there is no armoring in the study area, continue. If there is armoring, use the Editing toolbar to add lines to the feature class that trace instances of armoring. Common manifestations of what we call armoring are sandfencing and sandbagging and concrete seawalls.
#
# If there is no armoring file in the project geodatabase, the extractor script will notify you that it is proceeding without armoring.
#
# *__Requires manipulation in GIS__*
arcpy.CreateFeatureclass_management(home, os.path.basename(armorLines), 'POLYLINE', spatial_reference=utmSR)
print("{} created. Now manually digitize the shorefront armoring.".format(armorLines))
# ### Inlets
# We also need to manually digitize inlets if an inlet delineation does not already exist. To do, the code below will produce the feature class. After which, use the Editing toolbar to create a line where the oceanside shore meets a tidal inlet. If the study area includes both sides of an inlet, that inlet will be represented by two lines. The inlet lines are used to define the bounds of the oceanside shore, which is also considered the point where the oceanside shore meets the bayside. Inlet lines must intersect the MHW contour.
#
# What do we do when the study area and not an inlet is the end?
#
# *__Requires manipulation in GIS__*
# manually create lines that correspond to end of land and cross the MHW line (use bndpoly/DEM)
arcpy.CreateFeatureclass_management(home, os.path.basename(inletLines), 'POLYLINE', spatial_reference=utmSR)
print("{} created. Now we'll stop for you to manually create lines at each inlet.".format(inletLines))
# ### Shoreline
# The shoreline is produced through a combination of the DEM and the shoreline points. The first step converts the DEM to both MTL and MHW contour polygons. Those polygons are combined to produce the full shoreline, which is considered to fall at MHW on the oceanside and MTL on the bayside (to include partially submerged wetland).
#
# If the study area does not end cleanly at an inlet, create a separate polyline feature class (default name is 'SA_bounds') and add lines that bisect the shoreline; they should look and function like inlet lines. Specify this in the arguments for DEMtoFullShorelinePoly() and CreateShoreBetweenInlets().
#
# At some small inlets, channel depth may be above MTL. In this case, the script left to its own devices will leave the MTL contour between the two inlet lines. This can be rectified after processing by deleting the mid-inlet features from the temp file 'shore_2split.'
# manually create lines that correspond to end of land and cross the MHW line (use bndpoly/DEM)
if len(SA_bounds) and not arcpy.Exists(SA_bounds):
arcpy.CreateFeatureclass_management(home, 'SA_bounds', 'POLYLINE', spatial_reference=utmSR)
print("{} created. \nNow we'll stop for you to manually create lines at each inlet.".format(SA_bounds))
bndpoly = fwa.DEMtoFullShorelinePoly(elevGrid, sitevals['MTL'], sitevals['MHW'], inletLines, ShorelinePts)
print('Select features from {} that should not be included in the final shoreline polygon. '.format(bndpoly))
# *__Requires display in GIS__*
#
# User input is required to identify only the areas within the study area and eliminate isolated landmasses that are not. Once the features to delete are selected, either delete in the GIS or run the code below. Make sure the bndpoly variable matches the layer name in the GIS.
#
# __Do not...__ select the features in ArcGIS and then run DeleteFeatures in this Notebook Python kernel. That will delete the entire feature class.
#
# ```
# arcpy.DeleteFeatures_management(bndpoly)
# ```
#
# The next step snaps the boundary polygon to the shoreline points anywhere they don't already match and as long as as they are within 25 m of each other.
bndpoly = os.path.join(home, 'bndpoly_manual')
barrierBoundary = fwa.NewBNDpoly(bndpoly, ShorelinePts, barrierBoundary, '25 METERS', '50 METERS')
# #### ShoreBetweenInlets
# This step could be moved out of pre-processing and into the extractor because it doesn't require user input.
shoreline = fwa.CreateShoreBetweenInlets(barrierBoundary, inletLines, shoreline,
ShorelinePts, proj_code, SA_bounds)
# ### Transects - extend, sort, and tidy
#
# Create extendedTrans, which are NASC transects for the study area extended to cover the island, with gaps filled, and sorted in the field sort_ID.
#
# #### 1. Extend the transects and use a copy of the lines to fill alongshore gaps
# +
# Initialize temp file names
trans_extended = os.path.join(arcpy.env.scratchGDB, 'trans_extended')
trans_presort = trans_extended+'_presort'
# Delete transects over 200 m outside of the study area.
if input("Need to remove extra transects? 'y' if barrierBoundary should be used to select. ") == 'y':
fwa.RemoveTransectsOutsideBounds(orig_trans, barrierBoundary)
# Extend transects and create blank duplicate to use to fill gaps
fwa.ExtendLine(orig_trans, trans_extended, extendlength, proj_code)
fwa.CopyAndWipeFC(trans_extended, trans_presort, ['sort_ID'])
print("MANUALLY: use groups of existing transects in new FC '{}' to fill gaps.".format(trans_presort))
# -
# *__Requires manipulation in GIS__*
#
# 1. Edit the trans_presort_temp feature class. __Move and rotate__ groups of transects to fill in gaps that are greater than 50 m alongshore. There is no need to preserve the original transects, but avoid overlapping the transects with each other and with the originals. Do not move any transects slightly. If they are moved, they will not be deleted in the next stage. If you slightly move any, you can either undo or delete that line entirely.
fwa.RemoveDuplicates(trans_presort, trans_extended, barrierBoundary)
# #### 2. Sort the transects along the shore
# Usually if the shoreline curves, we need to identify different groups of transects for sorting. This is because the GIS will not correctly establish the alongshore order by simple ordering from the identified sort_corner. If this is the case, answer __yes__ to the next prompt.
sort_lines = fwa.SortTransectPrep(spatialref=utmSR)
sort_lines = os.path.join(arcpy.env.scratchGDB, 'sort_lines')
# *__Requires manipulation in GIS__*
#
# The last step generated an empty sort lines feature class if you indicated that transects need to be sorted in batches to preserve the order. Now, the user creates lines that will be used to spatially sort transects in groups.
#
# For each group of transects:
#
# 1. __Create a new line__ in 'sort_lines' that intersects all transects in the group. The transects intersected by the line will be sorted independently before being appended to the preceding groups. (*__add example figure__*)
# 2. __Assign values__ for the fields 'sort,' 'sort_corner,' and 'reverse.' 'sort' indicates the order in which the line should be used and 'sort_corn' indicates the corner from which to perform the spatial sort ('LL', 'UL', etc.). 'reverse' indicates whether the order should be reversed (roughly equivalent to 'DESCENDING').
# 3. Run the following code to create a new sorted transect file.
fwa.SortTransectsFromSortLines(trans_presort, extendedTrans, sort_lines, tID_fld)
arcpy.Describe(orig_trans).spatialReference.name
orig_trans = fwa.ReProject(orig_trans, orig_trans+'_utm', proj_code=arcpy.Describe(extendedTrans).spatialReference.factoryCode)
arcpy.Describe(orig_trans).spatialReference.name
# +
extendedTrans = os.path.join(home, 'mon_trans')
# project originals to UTM if they're not already
# orig_trans = fwa.ReProject(orig_trans, orig_trans+'_utm', proj_code=arcpy.Describe(extendedTrans).spatialReference.factoryCode)
# If there's no overlap between transect and original transect, set TRANSECTID, TRANSORDER, LRR to fill
fldsToWipe = ['TRANSECTID', 'TRANSORDER', 'LRR']
# 2. Remove orig transects from manually created transects
# If any of the original extended transects (with null values) are still present in trans_presort, delete them.
for row in arcpy.da.SearchCursor(orig_trans, ['SHAPE@']):
nasc_line = row[0]
buff = nasc_line.buffer(10)
with arcpy.da.UpdateCursor(extendedTrans, ['SHAPE@', 'TRANSECTID', 'TRANSORDER']) as cursor:
for trow in cursor:
tran = trow[0]
# if not buff.overlaps(tran): # none of the geometries overlap
# trow[1] = -99
# trow[2] = -99
# # trow[3] = -99
# cursor.updateRow(trow)
if buff.disjoint(tran): # all of them are disjoint even after reprojecting the originals
trow[1] = -99
trow[2] = -99
# trow[3] = -99
cursor.updateRow(trow)
# -
# ## 3. Tidy the extended (and sorted) transects to remove overlap
#
# *__Requires manipulation in GIS__*
#
# Overlapping transects cause problems during conversion to 5-m points and to rasters. We create a separate feature class with the 'tidied' transects, in which the lines don't overlap. This is largely a manually process with the following steps:
#
# 1. Select transects to be used to split other transects. Prioritize transects that a) were originally from NASC, b) have dune points within 25 m, and c) are oriented perpendicular to shore. (add example figure)
# 2. Use the Copy Features geoprocessing tool to copy only the selected transects into a new feature class. If desired, here is the code that could be used to copy the selected features and clear the selection:
# ```python
# arcpy.CopyFeatures_management(extendedTrans, overlapTrans_lines)
# arcpy.SelectLayerByAttribute_management(extendedTrans, "CLEAR_SELECTION")
# ```
# 3. Run the code below to split the transects at the selected lines of overlap.
overlapTrans_lines = os.path.join(arcpy.env.scratchGDB, 'overlapTrans_lines')
if not arcpy.Exists(overlapTrans_lines):
overlapTrans_lines = input("Filename of the feature class of only 'boundary' transects: ")
arcpy.Intersect_analysis([extendedTrans, overlapTrans_lines], trans_x,
'ALL', output_type="POINT")
arcpy.SplitLineAtPoint_management(extendedTrans, trans_x, extTrans_tidy)
# Delete the extraneous segments manually. Recommended:
#
# 1. Using Select with Line draw a line to the appropriate side of the boundary transects. This will select the line segments that need to be deleted.
# 1. Delete the selected lines.
# 1. Remove any remaining overlaps entirely by hand. Use the Split Line tool in the Editing toolbar to split lines to be shortened at the points of overlap. Then delete the remnant sections.
# ## Join anthro data to transects
#
# 1. Convert xls spreadsheet to points
# 2. Select the first points along each transects and create new FC
# 3. Spatial Join the new FC to the updated transects
# - one to one
# - keep all target features
# - keep only the ID fields and the three anthro fields (and the transect fields [LRR, etc.]?)
# - intersect
#
# 4. Join the transect values to the pts based on sort_ID
#
# Input shapefiles
shlpts_shp = os.path.join(proj_dir, 'rock14_shlpts.shp')
dlpts_shp = os.path.join(proj_dir, 'rock14_dlowpts.shp')
dhpts_shp = os.path.join(proj_dir, 'rock14_dhighpts.shp')
trans_shp = os.path.join(proj_dir, 'rock_trans.shp')
shoreline_shp = os.path.join(proj_dir, 'rock14_shoreline.shp')
| vol2/prepper_mon14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Crash course in Jupyter and Python
#
# - Introduction to Jupyter
# - Using Markdown
# - Magic functions
# - REPL
# - Saving and exporting Jupyter notebooks
# - Python
# - Data types
# - Operators
# - Collections
# - Functions and methods
# - Control flow
# - Loops, comprehension
# - Packages and namespace
# - Coding style
# - Understanding error messages
# - Getting help
# ## Class Repository
#
# Course material will be posted here. Please make any personal modifications to a **copy** of the notebook to avoid merge conflicts.
#
# https://github.com/cliburn/sta-663-2019.git
# ## Introduction to Jupyter
#
# - [Official Jupyter docs](https://jupyter.readthedocs.io/en/latest/)
# - User interface and kernels
# - Notebook, editor, terminal
# - Literate programming
# - Code and markdown cells
# - Menu and toolbar
# - Key bindings
# - Polyglot programming
# ### Using Markdown
#
# - What is markdown?
# - Headers
# - Formatting text
# - Syntax-highlighted code
# - Lists
# - Hyperlinks and images
# - LaTeX
# ### Magic functions
#
# - [List of magic functions](https://ipython.readthedocs.io/en/stable/interactive/magics.html)
# - `%magic`
# - Shell access
# - Convenience functions
# - Quick and dirty text files
# ### REPL
#
# - Read, Eval, Print, Loop
# - Learn by experimentation
# ### Saving and exporting Jupyter notebooks
#
# - The File menu item
# - Save and Checkpoint
# - Exporting
# - Close and Halt
# - Cleaning up with the Running tab
# ## Introduction to Python
#
# - [Official Python docs](https://docs.python.org/3/)
# - [Why Python?](https://insights.stackoverflow.com/trends?tags=python%2Cjavascript%2Cjava%2Cc%2B%2B%2Cr%2Cjulia-lang%2Cscala&utm_source=so-owned&utm_medium=blog&utm_campaign=gen-blog&utm_content=blog-link&utm_term=incredible-growth-python)
# - General purpose language (web, databases, introductory programming classes)
# - Language for scientific computation (physics, engineering, statistics, ML, AI)
# - Human readable
# - Interpreted
# - Dynamic typing
# - Strong typing
# - Multi-paradigm
# - Implementations (CPython, PyPy, Jython, IronPython)
# ### Data types
#
# - boolean
# - int, double, complex
# - strings
# - None
# ### Operators
#
# - mathematical
# - logical
# - bitwise
# - membership
# - identity
# - assignment and in-place operators
# - operator precedence
# ### Collections
#
# - Sequence containers - list, tuple
# - Mapping containers - set, dict
# - The [`collections`](https://docs.python.org/2/library/collections.html) module
# ### Functions and methods
#
# - Anatomy of a function
# - Docstrings
# - Class methods
# ### Control flow
#
# - if and the ternary operator
# - Checking conditions - what evaluates as true/false?
# - if-elif-else
# - while
# - break, continue
# - pass
# ### Loops and comprehensions
#
# - for, range, enumerate
# - lazy and eager evaluation
# - list, set, dict comprehensions
# - generator expression
# ### Packages and namespace
#
# - Modules (file)
# - Package (hierarchical modules)
# - Namespace and naming conflicts
# - Using `import`
# - [Batteries included](https://docs.python.org/3/library/index.html)
# ### Coding style
#
# - [PEP 8 — the Style Guide for Python Code](https://pep8.org/)
#
# ### Understanding error messages
#
# - [Built-in exceptions](https://docs.python.org/3/library/exceptions.html)
# ### Getting help
#
# - `?foo`, `foo?`, `help(foo)`
# - Use a search engine
# - Use `StackOverflow`
# - Ask your TA
| notebook/S01_Jupyter_and_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 8 Analyzing Sentence Structure
# ## 8.1 Some Grammatical Dilemmas
# ### Ubiquitous Ambiguity
import nltk
groucho_grammar = nltk.CFG.fromstring("""
S -> NP VP
PP -> P NP
NP -> Det N | Det N PP | 'I'
VP -> V NP | VP PP
Det -> 'an' | 'my'
N -> 'elephant' | 'pajamas'
V -> 'shot'
P -> 'in'
""")
sent = ['I', 'shot', 'an', 'elephant', 'in', 'my', 'pajamas']
parser = nltk.ChartParser(groucho_grammar)
for tree in parser.parse(sent):
print(tree)
# ## 8.3 Context-Free Grammar
# ### A Simple Grammar
grammar1 = nltk.CFG.fromstring("""
S -> NP VP
VP -> V NP | V NP PP
PP -> P NP
V -> "saw" | "ate" | "walked"
NP -> "John" | "Mary" | "Bob" | Det N | Det N PP
Det -> "a" | "an" | "the" | "my"
N -> "man" | "dog" | "cat" | "telescope" | "park"
P -> "in" | "on" | "by" | "with"
""")
sent = "Mary saw Bob".split()
rd_parser = nltk.RecursiveDescentParser(grammar1)
for tree in rd_parser.parse(sent):
print(tree)
# ### Recursion in Syntactic Structure
grammar2 = nltk.CFG.fromstring("""
S -> NP VP
NP -> Det Nom | PropN
Nom -> Adj Nom | N
VP -> V Adj | V NP | V S | V NP PP
PP -> P NP
PropN -> 'Buster' | 'Chatterer' | 'Joe'
Det -> 'the' | 'a'
N -> 'bear' | 'squirrel' | 'tree' | 'fish' | 'log'
Adj -> 'angry' | 'frightened' | 'little' | 'tall'
V -> 'chased' | 'saw' | 'said' | 'thought' | 'was' | 'put'
P -> 'on'
""")
# ## 8.4 Parsing with Context-Free Grammar
# ### Recursive Descent Parsing
rd_parser = nltk.RecursiveDescentParser(grammar1)
sent = 'Mary saw a dog'.split()
for t in rd_parser.parse(sent):
print(t)
# ### Shift-Reduce Parsing
sr_parser = nltk.ShiftReduceParser(grammar1)
sent = 'Mary saw a dog'.split()
for tree in sr_parser.parse(sent):
print(tree)
# ### Well-Formed Substring Tables
def init_wfst(tokens, grammar):
numtokens = len(tokens)
wfst = [[None for i in range(numtokens+1)] for j in range(numtokens+1)]
for i in range(numtokens):
productions = grammar.productions(rhs=tokens[i])
wfst[i][i+1] = productions[0].lhs()
return wfst
def complete_wfst(wfst, tokens, grammar, trace=False):
index = dict((p.rhs(), p.lhs()) for p in grammar.productions())
numtokens = len(tokens)
for span in range(2, numtokens+1):
for start in range(numtokens+1-span):
end = start + span
for mid in range(start+1, end):
nt1, nt2 = wfst[start][mid], wfst[mid][end]
if nt1 and nt2 and (nt1,nt2) in index:
wfst[start][end] = index[(nt1,nt2)]
if trace:
print("[%s] %3s [%s] %3s [%s] ==> [%s] %3s [%s]" % \
(start, nt1, mid, nt2, end, start, index[(nt1,nt2)], end))
return wfst
def display(wfst, tokens):
print('\nWFST ' + ' '.join(("%-4d" % i) for i in range(1, len(wfst))))
for i in range(len(wfst)-1):
print("%d " % i, end=" ")
for j in range(1, len(wfst)):
print("%-4s" % (wfst[i][j] or '.'), end=" ")
print()
tokens = "I shot an elephant in my pajamas".split()
wfst0 = init_wfst(tokens, groucho_grammar)
display(wfst0, tokens)
wfst1 = complete_wfst(wfst0, tokens, groucho_grammar)
display(wfst1, tokens)
wfst1 = complete_wfst(wfst0, tokens, groucho_grammar, trace=True)
# ## 8.5 Dependencies and Dependency Grammar
groucho_dep_grammar = nltk.DependencyGrammar.fromstring("""
'shot' -> 'I' | 'elephant' | 'in'
'elephant' -> 'an' | 'in'
'in' -> 'pajamas'
'pajamas' -> 'my'
""")
print(groucho_dep_grammar)
pdp = nltk.ProjectiveDependencyParser(groucho_dep_grammar)
sent = 'I shot an elephant in my pajamas'.split()
trees = pdp.parse(sent)
for tree in trees:
print(tree)
# ## 8.6 Grammar Development
# ### Treebanks and Grammars
from nltk.corpus import treebank
t = treebank.parsed_sents('wsj_0001.mrg')[0]
print(t)
def filter(tree):
child_nodes = [child.label() for child in tree
if isinstance(child, nltk.Tree)]
return (tree.label() == 'VP') and ('S' in child_nodes)
[subtree for tree in treebank.parsed_sents()
for subtree in tree.subtrees(filter)]
from collections import defaultdict
entries = nltk.corpus.ppattach.attachments('training')
table = defaultdict(lambda: defaultdict(set))
for entry in entries:
key = entry.noun1 + '-' + entry.prep + '-' + entry.noun2
table[key][entry.attachment].add(entry.verb)
for key in sorted(table):
if len(table[key]) > 1:
print(key, 'N:', sorted(table[key]['N']), 'V:', sorted(table[key]['V']))
nltk.corpus.sinica_treebank.parsed_sents()[3450].draw()
# ### Pernicious Ambiguity
grammar = nltk.CFG.fromstring("""
S -> NP V NP
NP -> NP Sbar
Sbar -> NP V
NP -> 'fish'
V -> 'fish'
""")
tokens = ["fish"] * 5
cp = nltk.ChartParser(grammar)
for tree in cp.parse(tokens):
print(tree)
# ### Weighted Grammar
def give(t):
return t.label() == 'VP' and len(t) > 2 and t[1].label() == 'NP'\
and (t[2].label() == 'PP-DTV' or t[2].label() == 'NP')\
and ('give' in t[0].leaves() or 'gave' in t[0].leaves())
def sent(t):
return ' '.join(token for token in t.leaves() if token[0] not in '*-0')
def print_node(t, width):
output = "%s %s: %s / %s: %s" %\
(sent(t[0]), t[1].label(), sent(t[1]), t[2].label(), sent(t[2]))
if len(output) > width:
output = output[:width] + "..."
print(output)
for tree in nltk.corpus.treebank.parsed_sents():
for t in tree.subtrees(give):
print_node(t, 72)
grammar = nltk.PCFG.fromstring("""
S -> NP VP [1.0]
VP -> TV NP [0.4]
VP -> IV [0.3]
VP -> DatV NP NP [0.3]
TV -> 'saw' [1.0]
IV -> 'ate' [1.0]
DatV -> 'gave' [1.0]
NP -> 'telescopes' [0.8]
NP -> 'Jack' [0.2]
""")
print(grammar)
viterbi_parser = nltk.ViterbiParser(grammar)
for tree in viterbi_parser.parse(['Jack', 'saw', 'telescopes']):
print(tree)
| Demo Codes/ch08.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784')
# -
mnist
# cd ..
# + pycharm={"name": "#%%\n"}
type(mnist)
X, y = mnist['data'], mnist['target']
X_train = X[:60000]
y_train = y[:60000]
X_test = X[60000:]
y_test = y[60000:]
from knn import KNNClassifier
knn_csf = KNNClassifier(5)
knn_csf.fit(X_train, y_train)
# -
knn_csf.predict(X_test[:20])
y_test[:20]
# 图像展示
some_digit_image = X[2395].reshape(28, 28)
import matplotlib
import matplotlib.pyplot as plt
plt.imshow(some_digit_image, cmap = matplotlib.cm.binary)
plt.show()
knn_csf.score(X_test[910:920], y_test[910:920])
| notebook/knn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.2 64-bit (''private_playground-5JPsKkiQ'': pipenv)'
# name: python3
# ---
import numpy as np
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from scipy import stats
from matplotlib import pyplot as plt
plt.rcParams['figure.figsize'] = [14, 9]
number_of_obs = 1000
# my_seed = 42
# np.random.seed(my_seed)
mvnorm = stats.multivariate_normal(mean=[0, 0], cov=[[1., 0.9],
[0.9, 1.]])
X_norm = mvnorm.rvs(number_of_obs)
X = stats.norm().cdf(X_norm)
X1 = X[:,0]
X2 = X[:,1]
np.corrcoef(X1,X2)
plt.plot(list(range(len(X1))), X1, 'o')
plt.hist(X1, bins=100)
plt.show()
plt.plot(X1, X2, 'o')
residuals = stats.norm(loc = 0, scale = 1).rvs(number_of_obs)
y = 30 * X1 - 10 * X2 + residuals
lr = LinearRegression()
lr.fit(X,y)
lr.coef_
plt.plot(X2, y, 'o')
# +
my_list = [],
my_dict = {},
my_comment = # # #. #
my_string = "string"
my_label = 'label'
'laksdfj'
my_label = 'label_2'
my_multiplication = a * b
my_func(param=12)
my_escape = "asd\@sd"
my_dict = {'key_1': "value 1"}
| data_analytics/collinearity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
#theano imports
import lasagne
import theano
import theano.tensor as T
import sys
sys.setrecursionlimit(100000)
floatX = theano.config.floatX
# -
# %load_ext autoreload
# %autoreload 2
# # Stack-augmented RNN
# 
#
#
# This notebook reproduces experiment from article http://arxiv.org/abs/1503.01007
# ## Task and architecture
#
# __The task__ is to train an RNN to generate sequences from expression `|`$ a^n b^m c^{n+m} $
# * n and m are integers
# * Performance metrics:
# * Correct format - `|`, some a, some b, some c
# * ||aaacbbcba would be an example of incorrect one
# * Number of C letters to be as close to sum of A band B letters as possible
# * Ideally, #c = #a + #b
#
#
# Model we use:
# * Stack-augmented RNN
#
# Trained as a language model (predicting next symbols given preceding history)
# ### Sequence generator
# +
def generate_sequence(batch_size = 10,crop_length = 100 ):
"""
Generates sequence from pattern [0 1*n 2*m 3*(n+m)]
"""
sequences=[]
for i in range(batch_size):
seq = [0]
while len(seq) < crop_length:
n,m = np.random.randint(1,10,2)
seq += [0]+ [1]*n+[2]*m+[3]*(n+m)
seq = seq[:crop_length]
sequences.append(seq)
return np.array(sequences,dtype='int32')
alphabet = np.array(list('|abc'))
# -
#quality evaluation
from operator import add
import re
def get_metrics(gen_sequences,alphabet):
strings = map(lambda v:reduce(add,v), map(alphabet.__getitem__,gen_sequences))
#at least one complete string
strings = filter(lambda v: '|' in v,strings)
#cut off last unfinished string, if any
strings = map(lambda v: v[:-1-v[::-1].index('|')],strings)
all_strings = '|'.join(strings)
seqs = filter(len,all_strings.split('|'))
matches = map(lambda seq:re.match(r"^a+b+c+",seq),seqs)
is_correct = map( lambda seq,m: m is not None and m.pos==0 and m.endpos == len(seq),
seqs, matches)
correct_seqs = np.array(seqs)[np.array(is_correct,dtype='bool')]
seqs_error = map(lambda s: s.count('a')+ s.count('b') - s.count('c'),correct_seqs)
return np.mean(is_correct), np.mean(np.abs(seqs_error))
# ### samples
# %%time
map(''.join,map(alphabet.__getitem__,generate_sequence(5,100)))
# # Constants
# +
# length of training/generation sequence
SEQ_LENGTH = 50
# Sequences in a minibatch
BATCH_SIZE = 100
# Total epochs
N_EPOCHS = 5000
#how often to print metrics
REPORT_RATE = 100
# +
#Reference sequences for training
sequences_batch = T.matrix(dtype="int32",name="reference_sequences")
#Batch size (theano expression)
batch_size = sequences_batch.shape[0]
# -
# # Define RNN
#
# A "simple" scheme of what was there in the paper, followed exactly by the code below
#
# * Time goes from left to right
# * Layers start at the bottom and go up
# * memory states are green, memory updates are purple
# * inputs and outputs are blue traingles
# * transformations are in pale yellow
#
# 
# +
import lasagne
from lasagne.layers import DenseLayer, ConcatLayer, InputLayer, EmbeddingLayer
import agentnet
from agentnet.resolver import ProbablisticResolver
from agentnet.memory import GRUMemoryLayer, StackAugmentation
from agentnet.agent import Recurrence
# +
#input letter goes here
output_shape = (None,)
observation_layer = InputLayer(output_shape,name="obs_input")
##Outputs
# next letter probabilities
probability_layer = lasagne.layers.DenseLayer(gru,
num_units = n_tokens,
nonlinearity= lasagne.nonlinearities.softmax,
name="policy_original")
#resolver picks a particular letter in generation mode
resolver = ProbablisticResolver(probability_layer,
assume_normalized=True,
name="resolver")
#verify that letter shape matches
assert tuple(lasagne.layers.get_output_shape(resolver)) == tuple(output_shape)
#define a dictionary that maps agent's "next memory" output to its previous memory input
from collections import OrderedDict
memory_dict = OrderedDict([
(gru,prev_gru_layer),
(next_stack, prev_stack_layer),
])
# -
# # Training mode
#
# * Here we define an in-training version of out generator.
# * To speed up training, we will feed agent with a reference sequences.
# * So, agent pretends to generate letters, but actually is fed with the reference ones.
#
#define an input layer that stores sequences
sequences_input_layer = InputLayer((None,SEQ_LENGTH),
input_var= sequences_batch,
name = "reference sequences"
)
<Main stuff goes here,see lecture instructions>
# # Unroll the recurrence
#
# * returns sessions of shape [batch_size, sequence_length, variable_size] for each variable
#
# * this is still happening inside a lasagne networks
# * which allows you to, for example, create a recurrence inside another recurrence
# +
#get lasagne layers for sequences
# dict of RNN/stack sequences, (actions and probabilities output)
recurrent_states, wouldbe_letters_layer, probas_seq_layer= <get the sequence layers>
# gru state sequence (can extract stack likewise)
rnn_seq_layer = recurrent_states[gru]
# for this task we only need probabilities :)
# probas_seq_layer
# -
probas_seq = lasagne.layers.get_output(probas_seq_layer)
# ### The rest is a regular lasagne
# ### Weights
#Obtain them
weights = lasagne.layers.get_all_params(probas_seq_layer,
trainable=True,
)
weights
total_weights = int(T.sum([T.prod(w.shape) for w in weights]).eval())
print "Total weights:", total_weights
# # Loss function
#
# simple crossentropy
# +
# take all predicitons but for last one (we don't have it's 'next')
predicted_probas = probas_seq[:,:-1]
# crop probabilities to avoid -Inf logarithms
predicted_probas = T.maximum(predicted_probas,1e-10)
# correct answers
references = sequences_batch[:,1:]
# familiar lasagne crossentropy
model_loss = lasagne.objectives.categorical_crossentropy(
predicted_probas.reshape([-1,n_tokens]),
references.ravel()
).mean()
# -
#Regularizer for kicks
from lasagne.regularization import regularize_network_params, l2
reg_l2 = regularize_network_params(resolver,l2)*10**-5
loss = model_loss + reg_l2
updates = lasagne.updates.adadelta(loss,weights,learning_rate=0.1)
# # Compile the training and evaluation steps
#
# * Since we unroll scans, the first compile may take approx. 3.5 cups of tea
# * If you want it to just compile something, consider either of the two:
# * turning off optimizer
# * reducing sequence length to, say, 20
train_fun = theano.function([sequences_batch],[loss],updates=updates)
evaluation_fun = theano.function([sequences_batch],[loss,model_loss,reg_l2])
# # Generating letters recurrently
# * In this mode, generator's i+1'st input is it's i-th output
# * the only difference in terms of code is that instead of reading letters from sequence,
# * we feed bach the previously generated ones
# +
# Recurrent state dictionary: maps state outputs to prev_state inputs
feedback_dict = OrderedDict([
<your code>
])
# generation batch size
gen_batch_size = T.scalar('generated batch size','int32')
#define a lasagne recurrence that actually generates letters
# it will be used to generate sample sequences.
generator_active = Recurrence(
<your code>
)
# +
#get sequences as before
generator_memory, generator_outputs = generator_active.get_sequence_layers()
action_seq_layer, gen_probabilities_layer = generator_outputs
generated_action_seq = lasagne.layers.get_output(action_seq_layer)
# -
# ~0.5 tea cups
get_sequences = theano.function([gen_batch_size],generated_action_seq)
#sample output
map(alphabet.__getitem__,get_sequences(3))
# # Training loop:
#
# * Generating sequences
# * Training on them
# * Printing metrics every once in a while
# +
from collections import defaultdict
metrics = defaultdict(dict)
# +
for i in range(N_EPOCHS):
#generate reference
new_batch = generate_sequence(BATCH_SIZE,SEQ_LENGTH)
#feed the monster
train_fun(new_batch)
#Display metrics once in a while
if i % REPORT_RATE==0:
loss_components = evaluation_fun(new_batch)
print "iter:%i\tfull:%.5f\tllh:%.5f\treg:%.5f"%tuple([i]+map(float,loss_components))
metrics['crossentropy'][i]=float(loss_components[1])
examples = get_sequences(1000)
correctness_ratio,c_count_mae = get_metrics(examples,alphabet)
metrics["correctness_rates"][i] = correctness_ratio
metrics["c_count_errors"][i]=c_count_mae
print "Correct sequences ratio: %.5f"%(correctness_ratio)
print "C-s count mean abs error: %.5f"%(c_count_mae)
for tid_line in examples[:3]:
print ' '.join(map(alphabet.__getitem__,tid_line))
# -
for metric in metrics:
plt.figure(figsize=[14,8])
plt.plot(*zip(*sorted(metrics[metric].items(),key=lambda (k,v):k)),label=metric)
plt.legend(loc='best')
plt.ylabel("popugai")
plt.xlabel("epoch")
plt.grid()
| week5/week5_STACK_RNN.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// ## Java in a Jupyter Notebook, basically runs jshell.
System.out.println("hello jupyter " + " java world");
// #### No need to have a class, can run code segments.
String twice(String s) {
return s + s;
}
System.out.println(twice("happy"));
int[] numbers = {2,4,5,7};
for(int i: numbers) {
System.out.println(i);
}
// #### Create a bubblesort class.
// Using a class to encapsulate the process.
// The utility swap method simply swaps two elements of an array.
// The main work is carried in the sort method.
// <img src="images/bubble_sort.png" alt="Quick Union Observation" style="height: 500px; width:700px;"/>
// +
class BubbleSort {
private int[] arr;
private int nElements;
public void swap(int index1, int index2) {
int temp = arr[index1];
arr[index1] = arr[index2];
arr[index2] = temp;
}
public BubbleSort(int max) {
arr = new int[max]; // creates array, elements zeroed
nElements = 0;
}
public void insert(int item) {
arr[nElements] = item;
nElements++;
}
public void display() {
System.out.println("Array has the following elements: ");
for (int i : arr) {
System.out.println(i);
}
}
public void sort() {
int in;
int out;
for(out = nElements-1; out > 1; out--){
for(in=0;in < out; in++) {
if (arr[in] > arr[in+1])
swap(in,in+1);
}
}
}
} // end class
// -
BubbleSort numbers = new BubbleSort(6);
numbers.insert(8);
numbers.insert(3);
numbers.insert(1);
numbers.insert(30);
numbers.insert(14);
numbers.insert(28);
numbers.display();
numbers.sort();
numbers.display();
// #### We can improve upon the algorithm slightly by adding a flag to the swap section.
// i.e. set the flag to false and each time a swap is made set the flag to true. if the flag is flag after the inner loop has completed and before the outer loop starts again then the list is in order. So in effect your steps would be
// - Start outer loop
// - initialise flag to false
// - complete inner loop setting flag to true if a swap is made
// - check flag , if it false then you are done.
// - continue with outer loop
| bubble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/macsakini/AI-Modelling-/blob/main/rlmodel.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="SKL-Xuun0kwd"
import gym
import keras
import tensorflow
#import keras_rl2
import random
import numpy as np
# + id="BAb-6fe71Bfq"
class BalanceEnv(gym.Env):
#metadata = {‘render.modes’: [‘human’]}
def __init__(self):
#0 is going down, 1 is going up
self.action_space = gym.spaces.Discrete(2,)
#ranges from o to 7 metres
self.observation_space = gym.spaces.Box(low = np.array([0]),high = np.array([7]))
self.game_length = 20
self.state = 3.5 + random.randint(-1,1)
pass
def step(self, action):
if action == 0:
self.state -= 1.25
self.game_length -= 1
else:
self.state += 1.25
self.game_length -= 1
if self.state >= 0 and self.state<= 7:
reward = 1
else:
reward = -1
if self.game_length <= 0:
done = True
elif self.state < 0:
done = True
elif self.state > 7:
done =True
else:
done = False
# Apply temperature noise
#self.state += ra
info = {}
return self.state, reward, done, info
def render(self):
pass
def reset(self):
self.game_length = 50
self.state = 3.5 + random.randint(-1,1)
return self.state
# + colab={"base_uri": "https://localhost:8080/"} id="Ab7fj1rN4dh2" outputId="fe49b59b-0dae-4eae-974d-7fc84473da44"
env = BalanceEnv()
# + [markdown] id="gNQDvHQL6hRJ"
#
# + colab={"base_uri": "https://localhost:8080/"} id="1Uu1e4zK6iGq" outputId="c8a1ce10-d156-4c65-b9ed-23e0d61491d9"
env.observation_space.sample()
# + colab={"base_uri": "https://localhost:8080/"} id="ccjvOgto6pOZ" outputId="a6d1525f-54f8-4b5a-e707-3c0a667c8959"
episodes = 10
observation = env.reset()
for episode in range(1, episodes + 1):
state = env.reset()
done = False
score = 0
while not done:
#env.render()
action = env.action_space.sample()
n_state, reward, done, info = env.step(action)
#print(n_state)
score += reward
print('Episode:{} Score:{} Steps:{}'.format(episode, score, n_state))
# + id="-Yxt79eOHpgs"
import numpy as np
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Flatten
from tensorflow.keras.optimizers import Adam
# + id="Z81n3PFE7MjG"
states = env.observation_space.shape
actions = env.action_space.n
# + id="oQ2uAV2JBbAA"
def build_model(states, actions):
model = tensorflow.keras.Sequential()
model.add(Dense(24, activation='relu', input_shape=states))
model.add(Dense(24, activation='relu'))
model.add(Dense(actions, activation='linear'))
return model
# + colab={"base_uri": "https://localhost:8080/"} id="ksIbQUxxIDIE" outputId="d80154ea-776b-4ef1-c7b7-62b1178347f3"
model = build_model(states, actions)
model.summary()
# + id="Hnl50Hy9IPUz"
from rl.agents import DQNAgent
from rl.policy import BoltzmannQPolicy
from rl.memory import SequentialMemory
# + id="9AYUvQNaIcRG"
def build_agent(model, actions):
policy = BoltzmannQPolicy()
memory = SequentialMemory(limit=50000, window_length=1)
dqn = DQNAgent(model=model, memory=memory, policy=policy,
nb_actions=actions, nb_steps_warmup=10, target_model_update=1e-2)
return dqn
# + colab={"base_uri": "https://localhost:8080/"} id="fbn81Yd2IiX7" outputId="d22177ae-20e7-49de-cebf-ef034aa3c1e2"
dqn = build_agent(model, actions)
dqn.compile(Adam(learning_rate = 1e-3), metrics=['mae'])
dqn.fit(env, nb_steps=4000, visualize=False, verbose=1)
# + colab={"base_uri": "https://localhost:8080/"} id="xQl69FoXKn-v" outputId="8b4b2e93-4a16-435c-fa83-b8aab1f0b45f"
scores = dqn.test(env, nb_episodes=10, visualize=False)
print(np.mean(scores.history['episode_reward']))
print(scores)
# + id="dFoDaaG-QqOj"
dqn.save_weights('dqn_weights.h5f', overwrite=True)
# + [markdown] id="94vrVhgVKxZy"
#
| rlmodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ### %run
# %run hello.py
hello("123")
# # ### %timeit
# 时间测试
# 自动选择测试次数 看执行效率
# %timeit L = [i**2 for i in range(1000)]
# %timeit L = [i**2 for i in range(1000000)]
# %%timeit
# 单元格内所有的
# %%单元测试符
L = []
for n in range(1000):
L.append(n**2)
# # ### %time
# %time L = [i**2 for i in range(1000)]
# %%time
L = []
for n in range(1000):
L.append(n**2)
| Python399MoocCourse/class3/.ipynb_checkpoints/3.2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from sklearn.neighbors import NearestNeighbors
from sklearn import preprocessing
# +
#Load the data
train = pd.read_pickle('X_train_features.pkl')
test = pd.read_pickle('X_test_features.pkl')
#Create list of columns that have >25 unique values and are dtype=float
def find_float_columns(df):
float_columns = []
for col in df:
if len(df[col].unique()) > 25 and df[col].dtype in ['float16','float32','float64']:# and train[col].dtype == 'object':
float_columns.append(col)
return float_columns
float_cols = find_float_columns(train)
# +
# Replace N/As
def fillna_and_inf_df(float_columns,df):
for col in float_columns:
df[col] = df[col].fillna(value=-99)
if np.isfinite(df[col]).all()== False:
df[col] = df[col].replace([np.inf], 10000)
return df
train = fillna_and_inf_df(float_cols,train)
test = fillna_and_inf_df(float_cols,test)
# Scale to mean = 0 and SD = 1
train_numerics = preprocessing.scale(train[float_columns])
test_numerics = preprocessing.scale(test[float_columns])
# -
neigh = NearestNeighbors(n_neighbors = 3)
neigh.fit(scaled_numeric_df)
neigh.kneighbors()
| ieee_fraud/Basic_KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#for Manupulation
import numpy as np
import pandas as pd
#for visulaization
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# +
#For reading the csv file
df = pd.read_csv('TRAIN (2).csv')
# -
df.head()
df.shape
# +
#Creating copy of Train Data Set
data = df.copy()
# -
df.isnull().sum()
df.describe().transpose()
# +
#Sales
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
sns.displot(df['Sales'],kde=False,bins=30)
# +
#Holidays
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
sns.countplot(x='Holiday',data=df)
# +
#Various stores and the specific amount of stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Store_Type'].unique())
sns.countplot(df['Store_Type'],data=df,order=order)
# +
#Various locations and the locations which have maximum stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Location_Type'].unique())
sns.countplot(df['Location_Type'],data=df,order=order)
# +
#The Number of stores in each locations
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Store_Type'].unique())
sns.countplot(df['Store_Type'],data=df,order=order,hue='Location_Type')
# +
#The discount given by each stores.
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Store_Type'].unique())
sns.countplot(df['Store_Type'],data=df,order=order,hue='Discount')
# +
#Various Regions and the Regions which have maximum stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Region_Code'].unique())
sns.countplot(df['Region_Code'],data=df,order=order)
# +
#The Number of stores in each Regions
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df['Store_Type'].unique())
sns.countplot(df['Store_Type'],data=df,order=order,hue='Region_Code')
# -
data['Date'] = pd.to_datetime(data['Date'])
type(data['Date'])
# +
latest = data['Date'].max()
oldest = data['Date'].min()
data['Days'] = (data['Date'] - oldest).dt.days
data.tail(10)
# -
data = data.drop(['ID','Date','#Order'],axis=1)
data
# +
storetype_dummies = pd.get_dummies(data['Store_Type'])
data = pd.concat([data.drop('Store_Type',axis=1),storetype_dummies],axis=1)
# +
locationtype_dummies = pd.get_dummies(data['Location_Type'])
data = pd.concat([data.drop('Location_Type',axis=1),locationtype_dummies],axis=1)
regioncode_dummies = pd.get_dummies(data['Region_Code'])
data = pd.concat([data.drop('Region_Code',axis=1),regioncode_dummies],axis=1)
data
# +
from sklearn import preprocessing
le = preprocessing.LabelEncoder()
le_discount = le.fit(data['Discount'])
data['Discount'] = le_discount.transform(data['Discount'])
data
# -
data.dtypes
# +
#testing and training the data
X = data.drop(['Sales'],axis=1)
y = data['Sales']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=101)
# -
from sklearn.ensemble import RandomForestRegressor
model_fr = RandomForestRegressor(n_estimators=800)
model_fr.fit(X_train,y_train)
predictions_fr = model_fr.predict(X_test)
predictions_fr
from sklearn.metrics import mean_squared_log_error
mean_squared_log_error(y_test,predictions_fr)
model_fr.score(X_train,y_train)
# +
#For reading the csv file
df_test = pd.read_csv('TEST_FINAL.csv')
# -
df_test.head()
df_test['Store_Type'].unique()
# +
#creating copy of test dataset
data_test = df_test.copy()
# -
df_test.isnull().sum()
df_test.describe().transpose()
# +
#Holidays
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
sns.countplot(x='Holiday',data=df_test)
# +
#Various stores and the specific amount of stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Store_Type'].unique())
sns.countplot(df_test['Store_Type'],data=df_test,order=order)
# +
#Various locations and the locations which have maximum stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Location_Type'].unique())
sns.countplot(df_test['Location_Type'],data=df_test,order=order)
# +
#The Number of stores in each locations
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Store_Type'].unique())
sns.countplot(df_test['Store_Type'],data=df_test,order=order,hue='Location_Type')
# +
#The discount given by each stores.
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Store_Type'].unique())
sns.countplot(df_test['Store_Type'],data=df_test,order=order,hue='Discount')
# +
#Various Regions and the Regions which have maximum stores
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Region_Code'].unique())
sns.countplot(df_test['Region_Code'],data=df_test,order=order)
# +
#The Number of stores in each Regions
plt.figure(figsize=(10,6))
sns.set_style('darkgrid')
order = sorted(df_test['Store_Type'].unique())
sns.countplot(df_test['Store_Type'],data=df_test,order=order,hue='Region_Code')
# -
data_test['Date'] = pd.to_datetime(data_test['Date'])
type(data_test['Date'])
# +
latest_test = data_test['Date'].max()
oldest_test = data_test['Date'].min()
data_test['Days'] = (data_test['Date'] - oldest_test).dt.days
data_test.tail(10)
# +
storetype_dummies = pd.get_dummies(data_test['Store_Type'])
data_test = pd.concat([data_test.drop('Store_Type',axis=1),storetype_dummies],axis=1)
locationtype_dummies = pd.get_dummies(data_test['Location_Type'])
data_test = pd.concat([data_test.drop('Location_Type',axis=1),locationtype_dummies],axis=1)
regioncode_dummies = pd.get_dummies(data_test['Region_Code'])
data_test = pd.concat([data_test.drop('Region_Code',axis=1),regioncode_dummies],axis=1)
data_test
# -
le_discount = le.fit(data_test['Discount'])
data_test['Discount'] = le_discount.transform(data_test['Discount'])
data_test = data_test.drop(['ID','Date'],axis=1)
data_test
predictions_test = model_fr.predict(data_test)
predictions_test
# +
dfid = df_test['ID']
df_submission = pd.concat([dfid, pd.DataFrame(predictions_test).rename(columns = {0 : 'Sales'})], axis=1)
df_submission
# -
df_submission.to_csv('submission_final_fr_1.csv',index=False)
| Project 1 Using Random Forest using Store Id 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import accuracy_score
from tqdm import tqdm
# OLS
from statsmodels.formula.api import ols
import statsmodels.api as sm
# VIF 확인
from statsmodels.stats.outliers_influence import variance_inflation_factor
from sklearn.metrics import mean_absolute_error
import warnings
# +
import warnings
import platform
from matplotlib import font_manager, rc
path = "c:/Windows/Fonts/malgun.ttf"
if platform.system() == 'Darwin':
font_name = 'AppleGothic'
rc('font', family='AppleGothic')
elif platform.system() == 'Windows':
font_name = font_manager.FontProperties(fname=path).get_name()
rc('font', family=font_name)
else:
font_name = font_manager.FontProperties(fname="/usr/share/fonts/nanumfont/NanumGothic.ttf")
rc('font', family="NanumGothic")
warnings.simplefilter(action='ignore')
# -
# !pip install lightgbm
# +
# 라이브러리 import
from sklearn.datasets import load_breast_cancer
from sklearn.datasets import load_wine
from sklearn.model_selection import train_test_split
from lightgbm import LGBMClassifier, plot_importance
from matplotlib import pyplot as plt
from lightgbm import plot_importance
from lightgbm import plot_tree
from sklearn.metrics import classification_report
# -
import lightgbm as lgb
total = pd.read_csv("total_data(2016포함).csv")
total = total[(total["year"] != "2020(상반기)") & (total["year"] != "2020(하반기)")]
col_name_x = total.columns[2:-5]
col_name_y = total.columns[-5:]
total[["year", "q1", "q2", "q3", "q4", "q5"]]
X = total[col_name_x]
y = total[col_name_y]
x_train = X.loc[:163]
x_test = X.loc[164:]
y_train = y.loc[:163]
y_test = y.loc[164:]
[["year", "q1", "q2", "q3", "q4", "q5"]]
y_train[["year", "q1"]]
# +
y1_train = y_train[["q1"]]
y1_test = y_test[["q1"]]
y2_train = y_train["q2"]
y2_test = y_test["q2"]
y3_train = y_train["q3"]
y3_test = y_test["q3"]
y4_train = y_train["q4"]
y4_test = y_test["q4"]
y5_train = y_train["q5"]
y5_test = y_test["q5"]
# +
std_scaler = StandardScaler()
std_scaler.fit(x_train)
x_train_s = pd.DataFrame(std_scaler.transform(x_train), columns = x_train.columns)
# x_train_s = sm.add_constant(x_train, has_constant = "add")
x_test_s = std_scaler.transform(x_test)
x_test_s = pd.DataFrame(std_scaler.transform(x_test), columns = x_test.columns)
# x_test_s = sm.add_constant(x_test, has_constant = "add")
# +
b1 = ['report_sx_여성', 'vio_cnt', 'theft_violence_x', 'robber_murder_x',
'theft_violence_y', 'robber_murder_y', '가해_10대이하', '피해_80대이상', 'for_합계',
'popu_2030', 'popu_o60', 'for_u20', 'for_2030', 'for_4050', 'for_o60',
'single_합계', 'single_2030', 'single_4050', 'single_o60', 'single_f_합계',
'single_f_4050', 'single_f_o60']
b2 = ['robber_murder_y', 'single_o60', '가해_10대이하', 'for_u20',
'robber_murder_x', 'for_합계', '피해_80대이상', 'vio_cnt', 'single_합계',
'single_f_합계', 'theft_violence_y', 'theft_violence_x', 'for_o60',
'single_4050', 'single_f_4050', 'popu_o60', 'single_f_o60',
'report_sx_여성', 'for_2030', 'for_4050']
b3 = ['robber_murder_y', 'single_f_2030', '가해_80대이상', 'single_2030',
'single_4050', '피해_70대', 'traffic_x', 'adult', 'robber_murder_x',
'mur_rob_cnt', '가해_70대', 'theft_violence_y', 'for_4050', 'single_합계',
'for_o60', 'theft_violence_x', '5m_crm_yn', 'for_합계', 'for_2030',
'bell']
b4 = ['mur_rob_cnt', 'for_4050', 'for_합계', 'single_4050', 'single_합계', 'cctv',
'for_2030', 'theft_violence_x', 'for_o60', '가해_여', 'car_cnt']
b5 = ['single_o60', 'single_f_o60', 'single_f_합계', 'for_2030', 'single_2030',
'for_합계', 'for_o60', 'popu_o60', 'vio_cnt', 'single_4050', 'popu_2030',
'for_u20', 'single_f_4050', 'robber_murder_x', 'single_합계', '가해_여',
'for_4050', '피해_80대이상', 'robber_murder_y', 'theft_violence_x']
# -
# # LGB
# 장점)
#
# - 학습하는데 걸리는 시간이 적다, 빠른 속도
#
# - 메모리 사용량이 상대적으로 적은편이다
#
# - categorical feature들의 자동 변환과 최적 분할
#
# - GPU 학습 지원
#
# 단점)
#
# - 작은 dataset을 사용할 경우 과적합 가능성이 크다 (일반적으로 10,000개 이하의 데이터를 적다고 한다)
import lightgbm as lgbm
reg = lgbm.LGBMRegressor()
lgb_model = lgb.LGBMClassifier(max_depth = 10, n_estimators = 100, learning_rate = 0.01).fit(x_train, y5_train.astype("int"))
a = lgb_model.predict(x_test)
mean_absolute_error(a, y5_test)
fig, ax = plt.subplots(figsize = (20, 20))
plot_importance(lgb_model, ax=ax)
plt.show(30)
| 3.Model_code/LGB_MODEL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #%matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from flask import Flask, jsonify
import numpy as np
import pandas as pd
import datetime as dt
from scipy import stats
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine,inspect, func
engine = create_engine("sqlite:///Resources/hawaii.sqlite")
inspector = inspect(engine)
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect=True)
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Climate Analysis
# ## Precipitation Analysis
# +
# Design a query to retrieve the last 12 months of precipitation data and plot the results
inspector.get_table_names()
columns = inspector.get_columns('measurement')
# Select only the date and prcp values.
session.query(Measurement.date, Measurement.prcp).all()
# +
# Calculate the date 1 year ago from the last data point in the database
# Query to find the last data point in the database
last_date = session.query(func.max(Measurement.date)).scalar()
# The date 1 year ago from the last data point in the database
date_one_yr_ago_dt = dt.datetime.strptime(last_date, '%Y-%m-%d') - dt.timedelta(days=365)
query_date = date_one_yr_ago_dt.strftime('%Y-%m-%d')
# -
# Perform a query to retrieve the date and precipitation scores
last_year = session.query(Measurement.date, Measurement.prcp).\
filter(Measurement.date >= '2016-08-23').all()
last_year
# Save the query results as a Pandas DataFrame and set the index to the date column
last_year_df = pd.DataFrame(last_year)
last_year_df = last_year_df.set_index("date")
last_year_df.head()
# Sort the dataframe by date
sorted_df = last_year_df.sort_values(by = "date")
sorted_df.head()
# +
# Use Pandas Plotting with Matplotlib to plot the data
# Set plot and plot the chart
plt.figure(figsize=(15, 10))
plt.plot(sorted_df.index, sorted_df['prcp'])
# Add legned
plt.legend(['precipitation'], loc="upper right",fontsize=15)
# set x location
xloc = ['2016-08-23', '2016-10-01', '2016-11-09', '2016-12-19', '2017-01-28', '2017-03-09', '2017-04-18','2017-05-31','2017-07-10']
# Assign xticks
plt.xticks(xloc, rotation='vertical',fontsize=15)
# Set Labels & Title
plt.xlabel('Date', fontsize=15)
plt.ylabel("Inches",fontsize=15)
plt.title(f"Precipitation (inches)in Honolulu, Hawaii from \n {query_date} to {last_date}",fontsize=20, fontweight = 'bold')
plt.yticks(size=15)
# Asign xlim and ylim
plt.xlim(0,370)
plt.ylim(-0.4,7)
# Save Figure
plt.savefig("Images/Precipitation_Plot.png")
# Show plot
plt.show()
plt.tight_layout()
# -
# Use Pandas to calcualte the summary statistics for the precipitation data
sorted_df.describe()
# ## Stations Analysis
# Design a query to show how many stations are available in this dataset?
#Inspect the station table
columns = inspector.get_columns('station')
session.query(Station.id).count()
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
ste = [Station.station, func.count(Measurement.station)]
session.query(*ste).filter(Station.station == Measurement.station).group_by(Station.station).\
order_by(func.count(Measurement.station).desc()).all()
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature of the most active station?
sel = [func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs)]
session.query(*sel).filter(Measurement.station == ste[0]).all()
# +
# Choose the station with the highest number of temperature observations.
session.query(Measurement.station, func.count(Measurement.tobs)).\
group_by(Measurement.station).\
order_by(func.count(Measurement.tobs).desc()).first() # 'USC00519281'
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
results = session.query(Measurement.tobs).\
filter(Measurement.date.between(query_date,last_date),\
Measurement.station == 'USC00519281').all()
# Plot the results as a histogram
plt.figure(figsize=(15,10))
plt.hist(np.ravel(results), bins=12)
plt.title('Temperatures Observed At Station USC00519281 (Last 12 Months)',fontsize=20, fontweight = 'bold')
plt.xlabel('Temperature',fontsize=15)
plt.ylabel('Frequency',fontsize=15)
plt.legend(['tobs'], loc = 'upper right')
plt.yticks(size=15)
plt.xticks(size=15)
plt.show()
# -
# ## Step 2 - Climate App
# For this part the solution is provided in python "app.py" file in the main folder.
# ## Bonus Challenge Assignment
# ### Temperature Analysis I
# +
# Identify the average temperature in June at all stations across all available years in the dataset.
avg_june_temp_tuple_list = session.query(func.avg(Measurement.tobs)).\
filter(func.strftime("%m", Measurement.date) == "06").all()
average_june_temp = list(np.ravel(avg_june_temp_tuple_list))[0]
# Identify the average temperature in December at all stations across all available years in the dataset.
avg_dec_temp_tuple_list = session.query(func.avg(Measurement.tobs)).\
filter(func.strftime("%m", Measurement.date) == "12").all()
average_dec_temp = list(np.ravel(avg_dec_temp_tuple_list))[0]
print(f"The average temperature in June at all stations across all available years in the dataset is {average_june_temp} F.")
print(f"The average temperature in December at all stations across all available years in the dataset is {average_dec_temp} F.")
# +
june_temp_tuple_list = session.query(Measurement.tobs).\
filter(func.strftime("%m", Measurement.date) == "06").all()
june_temp_list = list(np.ravel(june_temp_tuple_list))
# Remove nulls (if any)
clean_june_temp_list = []
for temp in june_temp_list:
if temp != None :
clean_june_temp_list.append(temp)
dec_temp_tuple_list = session.query(Measurement.tobs).\
filter(func.strftime("%m", Measurement.date) == "12").all()
dec_temp_list = list(np.ravel(dec_temp_tuple_list))
# Remove nulls (if any)
clean_dec_temp_list = []
for temp in dec_temp_list:
if temp != None :
clean_dec_temp_list.append(temp)
# Scatter Plot of Data
plt.subplot(2, 1, 1)
plt.scatter(range(len(clean_june_temp_list)), clean_june_temp_list, label="June Temperatures")
plt.scatter(range(len(clean_dec_temp_list)), clean_dec_temp_list, label="December Temperatures")
plt.legend()
# Histogram Plot of Data
plt.subplot(2, 1, 2)
plt.hist(clean_june_temp_list, 10, density=True, alpha=0.7, label="June Temperatures")
plt.hist(clean_dec_temp_list, 10, density=True, alpha=0.7, label="December Temperatures")
plt.axvline(np.mean(clean_june_temp_list), color='k', linestyle='dashed', linewidth=1)
plt.axvline(np.mean(clean_dec_temp_list), color='k', linestyle='dashed', linewidth=1)
plt.legend()
plt.savefig('./Images/june_dec_scatterplot_histogram.png')
# -
# ## Unpaired (Independent t-test)¶
# - Independent t-tests compare the means of 2 independent populations (June temperatures in Hawaii and December temperatures in Hawaii).
# - We want to use an independent t-test because we want to compare the means of two independent populations. A paired t-test (one sample t-test) looks at comparing the sample to the population, which we don't want in this case.
# - Assumptions - Data are normally distributed, data are independent, data are homogenous (The standard deviations are roughly equal).
stats.ttest_ind(june_temp_list, dec_temp_list, equal_var=False)
# ### Temperature Analysis II
# +
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# -
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
yearly_temp = calc_temps('2016-08-23', '2017-08-23')
yearly_temp
# +
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
tmax = yearly_temp[0][2]
tmin = yearly_temp[0][0]
peak_to_peak = tmax - tmin # This will be our error line
tavg = yearly_temp[0][1] # This will be the height of our graph
# Plot
fig, ax = plt.subplots(figsize = (5, 10)) # Create figure & axis objects
ax.bar(x = 1, height = tavg, yerr = peak_to_peak/2, width = 0.4,color = 'coral', alpha = 0.5) # Plotting
ax.set_xticks([0])
plt.yticks(size=14)
# "Labels"
plt.title("Trip Avg Temp")
plt.ylabel("Temp (F)")
plt.savefig("Images/Trip Avg Temp.png")
plt.tight_layout()
# -
# ### Daily Rainfall Average
# +
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
results = session.query(Measurement.station, Station.name, Station.latitude,\
Station.longitude, Station.elevation, func.avg(Measurement.prcp)).\
filter(Measurement.station == Station.station,\
Measurement.date.between(query_date, last_date)).\
group_by(Measurement.station).\
order_by(Measurement.prcp.desc()).\
all()
print(results)
# +
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# +
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
start_date = dt.datetime(2018, 1, 1)
end_date = dt.datetime(2018, 1, 7)
# Use the start and end date to create a range of dates
datelist = pd.date_range(start_date, periods=7).tolist()
# Stip off the year and save a list of %m-%d strings
dates = []
for date in datelist:
dates.append(dt.datetime.strftime(date, '%m-%d'))
# Loop through the list of %m-%d strings and calculate the normals for each date
normals = []
for date in dates:
normals.append(daily_normals(date))
print("Chosen Dates: Aug 23 to Aug 29 (7 day trip)")
for normal in normals:
print(normal)
# -
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
normal_list = []
for normal in normals:
normal_list.append(np.ravel(normal))
trip_normals_df = pd.DataFrame(normal_list, columns = ['tmin', 'tavg', 'tmax'])
trip_normals_df.index = [str(date.strftime('%Y-%m-%d')) for date in datelist]
trip_normals_df
# +
# Plot the daily normals as an area plot with `stacked=False`
fig, ax = plt.subplots(figsize=(10,7))
trip_normals_df.plot(kind='area', stacked=False, alpha=0.25, ax=ax)
plt.title("Daily Normals",fontsize=15)
plt.ylabel("Date",fontsize=15)
plt.xlabel("Temperature",fontsize=15)
plt.ylim(0,80)
plt.xlim(0,4)
plt.yticks(size=14)
plt.xticks(np.arange(7), trip_normals_df.index,fontsize=15)
plt.setp(ax.xaxis.get_majorticklabels(), rotation=30, ha="right" )
plt.show()
# -
| .ipynb_checkpoints/climate-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from bs4 import BeautifulSoup #to beautify xml output
from lxml import etree as ET #read and write xml files
pd.options.display.max_columns=50
import os
from os import listdir
from os.path import isfile, join
import time
# -
# Read the trips file from SF-CHAMP output
trip=pd.read_table('SF_CHAMP_Outputs/_trip_2.dat',sep='\t',header='infer')
trip.head()
mzone = pd.read_csv('SF_CHAMP_Outputs/mzone_short.csv') #MZONE coordinates in EPSG 26910 projection
mzone.drop(columns=['Unnamed: 0'],inplace=True)
mzone.head()
trip=trip[(trip['otaz']<1000)|(trip['dtaz']<1000)]
len(trip)
# +
#decoding the columns
def trip_purpose(x):
#(0=home, 1=work, 2=school, 3=escort, 4=personal business (& medical),
#5=shopping, 6=meal, 7=social (& recreation), 8=recreation (H version only)
#9=medical (H version only), 10=change mode at a park and ride lot
#according to the sf-light scenario population.xml all the following purposes are valid
#maybe needs activity-intercepts or activity-parameters
if x==0:
return 'home'
elif x==1:
return 'work'
elif x==2:
return 'school'
elif x==3:
return 'escort'
elif x==4:
return 'business'
elif x==5:
return 'shopping'
elif x==6:
return "meal"
else:
return 'other'
def convert_time(x):
#ARRTM, DEPTM, ENDTM etc is in minutes after 3 AM
return time.strftime("%H:%M:%S", time.gmtime(x*60+180))
def mode_type(df):
#Trip main mode type (1=walk, 2=bike, 3=sov, 4=hov 2, 5=hov 3+,
#6=walk to transit, 7=park and ride, 8=school bus, 9=TNC, 10=other – survey only)
if df['mode']==1:
return 'walk'
elif df['mode']==2:
return 'bike'
elif df['mode']==3:
return 'car'
elif df['mode']==4:
#return 'car'
if df['dorp']==1:
return 'car'
else:
return 'passenger'
elif df['mode']==5:
#return 'car'
if df['dorp']==1:
return 'car'
else:
return 'passenger'
elif df['mode']==6:
return 'walk_transit'
elif df['mode']==7:
return 'drive_transit'
elif df['mode']==9:
return 'ride_hail'
else:
return 'other'
def trip_type_location(df):
if df['otaz']<1000 and df['dtaz']<1000:
return 'int-int'
elif df['otaz']<1000 and df['dtaz']>1000:
return 'int-ext'
elif df['otaz']>1000 and df['dtaz']>1000:
return 'ext-ext'
else:
return 'ext-int'
def data_cleanup(trip_test,mzone):
trip_test['hhno']=trip_test['hhno'].astype(str)
trip_test['pno']=trip_test['pno'].astype(str)
trip_test['person_id']=trip_test['hhno']+'-'+trip_test['pno']
trip_test['trip_type']=trip_test.apply(trip_type_location,axis=1)
plans=trip_test[[
'person_id', #format is hh number-person number
'trip_type',#i-x type
'opurp',#The purpose at the trip origin (0=home, 1=work, 2=school, 3=escort, 4=personal business (& medical), 5=shopping, 6=meal, 7=social (& recreation), 8=recreation (H version only) 9=medical (H version only), 10=change mode at a park and ride lot
'dpurp',#the purpose at trip destination
#'oadtyp',#Trip origin address type (1=home, 2=usual work location, 3=usual school location, 4=other location in region, 5=out of region/missing (survey data only), 6=inserted change mode location for park and ride
#'dadtyp',#trip dest. address type
'opcl',#trip origin parcel
'dpcl',#trip dest parcel
#'otaz',#origin TAZ
#'dtaz',#dest TAZ
'mode',#Trip mode (1=walk, 2=bike, 3=sov, 4=hov 2, 5=hov 3+, 6=walk to transit, 7=park and ride, 8=school bus, 9=TNC, 10=other – survey only)
'dorp',#For auto trips, 1=driver, 2=passenger; for transit trips, is set to the total walk access+egress time, in integer minutes
'deptm',#The trip departure time, in minutes after midnight (or hours*100+minute for estimation mode)
'arrtm',#trip arrival time
'endacttm'#activity end time
]]
plans['trip_origin_purpose']=plans['opurp'].apply(trip_purpose)
plans['trip_dest_purpose']=plans['dpurp'].apply(trip_purpose)
plans['departure_time']=plans['deptm'].apply(convert_time)
plans['arrival_time']=plans['arrtm'].apply(convert_time)
plans['end_activity_time']=plans['endacttm'].apply(convert_time)
plans['mode']=plans.apply(mode_type,axis=1)
plans=plans[['person_id', 'trip_type', 'opcl', 'dpcl',
'mode', 'dorp', 'trip_origin_purpose','trip_dest_purpose',
'departure_time', 'arrival_time', 'end_activity_time']]
plans.sort_values(['person_id','departure_time'],inplace=True)
# if trip_type=='all internal':
# plans_df=plans[plans['trip_type']=='int-int']
# elif trip_type=='all within':
# plans_df=plans[plans['trip_type']!='ext-ext']
# else:
# plans_df=plans
plans_maz_orig=pd.merge(plans,mzone,left_on=['opcl'],right_on=['MAZID'],how='left')
plans_maz_all=pd.merge(plans_maz_orig,mzone,left_on=['dpcl'],right_on=['MAZID'],how='left')
plans_maz_all=plans_maz_all.rename(columns={'X_COORD_x':'X_ORIG',
'Y_COORD_x':'Y_ORIG',
'X_COORD_y':'X_DEST',
'Y_COORD_y':'Y_DEST'})
plans_maz_all.drop(columns=['MAZID_x','MAZID_y'],inplace=True)
plans_maz_all=plans_maz_all.drop_duplicates()
trip_test=None
mzone=None
return (plans_maz_all)
# -
# %%time
plans_df=data_cleanup(trip,mzone)
plans_df.trip_type.value_counts()
plans_df[plans_df['person_id']=='5000-1']
plans_df['mode'].value_counts()
plans_df[plans_df['person_id']=='1-1']
# +
from lxml import etree as ET
parser = ET.XMLParser(remove_blank_text=True)
template = ET.parse('SF_CHAMP_Converted/sf-within-int-trips-temp.xml',parser)
population_tag = template.getroot()
# -
def create_xml(plans_dataframe,size):
for i in range(0,len(plans_df.person_id.unique())+1,size):
j=i+size
test_df=plans_dataframe[plans_dataframe['person_id'].isin(plans_dataframe.person_id.unique()[i:j])]
parser = ET.XMLParser(remove_blank_text=True)
template = ET.parse('SF_CHAMP_Converted/sf-within-ix-trips.xml',parser)
population_tag = template.getroot()
for i in test_df['person_id'].unique():
person_id=str(i)
person_tag = ET.SubElement(population_tag, 'person')
person_tag.set('id',person_id)
plan_tag = ET.SubElement(person_tag, 'plan')
plan_tag.set('selected', 'yes')
df=test_df[test_df['person_id']==i]
act_tag = ET.SubElement(plan_tag, 'activity')
act_tag.set('type',df.iloc[0,df.columns.get_loc('trip_origin_purpose')])
act_tag.set('end_time',df.iloc[0,df.columns.get_loc('departure_time')])#
act_tag.set('y',str(df.iloc[0,df.columns.get_loc('Y_ORIG')]))
act_tag.set('x',str(df.iloc[0,df.columns.get_loc('X_ORIG')]))
for x in range(0,len(df.values)):
leg_tag = ET.SubElement(plan_tag, 'leg')
leg_tag.set('mode',df.values[x,df.columns.get_loc('mode')])
act_tag = ET.SubElement(plan_tag, 'activity')
act_tag.set('type',df.values[x,df.columns.get_loc('trip_dest_purpose')])
act_tag.set('end_time',df.values[x,df.columns.get_loc('end_activity_time')])
act_tag.set('y',str(df.values[x,df.columns.get_loc('Y_DEST')]))
act_tag.set('x',str(df.values[x,df.columns.get_loc('X_DEST')]))
population_tag.append(person_tag)
tree=ET.ElementTree(population_tag)
tree.write('SF_CHAMP_Converted/sf-within-ix-trips.xml', pretty_print=True, xml_declaration=True, encoding="utf-8")
# %%time
create_xml(plans_df,10000)
| convert/Script/01 SF-CHAMP Trips to MATSim format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import matplotlib.pyplot as plt
from pymedphys.level1.mudensity import *
DATA_DIRECTORY = "../../tests/data"
DELIVERY_DATA_FILEPATH = os.path.join(DATA_DIRECTORY, 'mu_density_example_arrays.npz')
regress_test_arrays = np.load(DELIVERY_DATA_FILEPATH)
# +
mu = regress_test_arrays['mu']
mlc = regress_test_arrays['mlc']
jaw = regress_test_arrays['jaw']
leaf_pair_widths = np.array(AGILITY_LEAF_PAIR_WIDTHS)
max_leaf_gap=400
grid_resolution=2.5
time_steps=50
mu, mlc, jaw = remove_irrelevant_control_points(mu, mlc, jaw)
# -
def determine_full_grid(max_leaf_gap, grid_resolution, leaf_pair_widths):
leaf_x = np.arange(
-max_leaf_gap/2,
max_leaf_gap/2 + grid_resolution,
grid_resolution).astype('float')
_, initial_leaf_grid_y_pos = determine_leaf_y(leaf_pair_widths)
total_leaf_widths = np.sum(leaf_pair_widths)
top_grid_pos = (
np.ceil((total_leaf_widths/2 - initial_leaf_grid_y_pos) / grid_resolution) *
grid_resolution + initial_leaf_grid_y_pos)
bot_grid_pos = (
initial_leaf_grid_y_pos -
np.ceil((total_leaf_widths/2 + initial_leaf_grid_y_pos) / grid_resolution) *
grid_resolution)
grid_y = np.arange(
bot_grid_pos, top_grid_pos + grid_resolution, grid_resolution)
grid_xx, grid_yy = np.meshgrid(leaf_x, grid_y)
return grid_xx, grid_yy
def determine_calc_grid_and_adjustments(mlc, jaw, leaf_pair_widths,
grid_resolution):
min_y = np.min(-jaw[:, 0])
max_y = np.max(jaw[:, 1])
leaf_y, initial_leaf_grid_y_pos = determine_leaf_y(leaf_pair_widths)
top_grid_pos = (
np.ceil((max_y - initial_leaf_grid_y_pos) / grid_resolution) *
grid_resolution + initial_leaf_grid_y_pos)
bot_grid_pos = (
initial_leaf_grid_y_pos -
np.ceil((-min_y + initial_leaf_grid_y_pos) / grid_resolution) *
grid_resolution)
grid = dict()
grid['jaw'] = np.arange(
bot_grid_pos, top_grid_pos + grid_resolution, grid_resolution
).astype('float')
grid_leaf_map = np.argmin(
np.abs(grid['jaw'][:, None] - leaf_y[None, :]), axis=1)
adjusted_grid_leaf_map = grid_leaf_map - np.min(grid_leaf_map)
leaves_to_be_calced = np.unique(grid_leaf_map)
adjusted_mlc = mlc[:, leaves_to_be_calced, :]
min_x = np.floor(
np.min(-adjusted_mlc[:, :, 0]) / grid_resolution) * grid_resolution
max_x = np.ceil(
np.max(adjusted_mlc[:, :, 1]) / grid_resolution) * grid_resolution
grid['mlc'] = np.arange(
min_x, max_x + grid_resolution, grid_resolution
).astype('float')
return grid, adjusted_grid_leaf_map, adjusted_mlc
# +
i = 3
control_point_slice = slice(i, i + 2, 1)
current_mlc = mlc[control_point_slice, :, :]
# current_jaw = jaw[control_point_slice, :]
current_jaw = np.array([
[2, 1],
[16, -4]
])
delivered_mu = np.diff(mu[control_point_slice])
(
grid, grid_leaf_map, current_mlc
) = determine_calc_grid_and_adjustments(
current_mlc, current_jaw,
leaf_pair_widths, grid_resolution)
# -
current_mlc
current_jaw
delivered_mu
grid
grid_leaf_map
# +
positions = {
'mlc': {
1: (-current_mlc[0, :, 0], -current_mlc[1, :, 0]), # left
-1: (current_mlc[0, :, 1], current_mlc[1, :, 1]) # right
},
'jaw': {
1: (-current_jaw[0::-1, 0], -current_jaw[1::, 0]), # bot
-1: (current_jaw[0::-1, 1], current_jaw[1::, 1]) # top
}
}
positions
# -
def calc_blocked_t(travel_diff, grid_resolution):
blocked_t = np.ones_like(travel_diff) * np.nan
fully_blocked = travel_diff <= -grid_resolution/2
fully_open = travel_diff >= grid_resolution/2
blocked_t[fully_blocked] = 1
blocked_t[fully_open] = 0
transient = ~fully_blocked & ~fully_open
blocked_t[transient] = (
(-travel_diff[transient] + grid_resolution/2) /
grid_resolution)
assert np.all(~np.isnan(blocked_t))
return blocked_t
# +
multiplier = -1
device = 'mlc'
start, end = positions[device][multiplier]
dt = (end - start) / (time_steps - 1)
travel = start[None, :] + np.arange(0,time_steps)[:, None] * dt[None, :]
np.shape(travel)
np.shape(multiplier * (grid[device][None,None, :] - travel[:, :, None]))
# +
blocked_by_device = {}
for device, value in positions.items():
blocked_by_device[device] = dict()
for multiplier, (start, end) in value.items():
dt = (end - start) / 49
travel = start[None, :] + np.arange(0,time_steps)[:, None] * dt[None, :]
travel_diff = multiplier * (grid[device][None, None, :] - travel[:, :, None])
blocked_by_device[device][multiplier] = calc_blocked_t(
travel_diff, grid_resolution)
# +
device_open = {}
for device, value in blocked_by_device.items():
device_sum = np.sum(np.concatenate([
np.expand_dims(blocked, axis=0)
for _, blocked in value.items()
], axis=0), axis=0)
device_open[device] = 1 - device_sum
# -
np.shape(device_open['mlc'])
np.shape(device_open['jaw'])
mlc_open = device_open['mlc'][:, grid_leaf_map, :]
jaw_open = device_open['jaw'][:, 0, :]
np.shape(mlc_open)
np.shape(jaw_open)
open_t = mlc_open * jaw_open[:, :, None]
open_fraction = np.mean(open_t, axis=0)
np.shape(open_fraction)
grid['mlc']
grid['jaw']
plt.pcolormesh(grid['mlc'], grid['jaw'], open_fraction)
plt.colorbar()
plt.title('MU density')
plt.xlabel('MLC direction (mm)')
plt.ylabel('Jaw direction (mm)')
plt.axis('equal')
plt.gca().invert_yaxis()
current_mlc
current_jaw
| examples/archive/mudensity/03_fractional_mudensity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating Shakespeare Sonnets with Deep Learning
# ---
# Creating a character-level language model trained on <NAME>'s sonnets.
#
# ## Data Loading & Visualization
# +
import numpy as np
import sys
import os
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(21)
print("Numpy version:", np.__version__)
# -
# Our corpus is composed of all of <NAME>'s sonnets concatenated into a single text file. The sonnets were retrieved from [shakespeares-sonnets.com](http://www.shakespeares-sonnets.com/all.php). There are a total of 154 sonnets in the corpus.
# +
with open('sonnets.txt', 'r') as f:
data = f.read().lower()
print("Corpus length: %d characters" % len(data))
# +
# Visualize sonnet character length
sonnets = data.split('\n\n')
sonnet_lens = [len(sonnet) for sonnet in sonnets]
print('Average sonnet length: %.2f characters' % np.mean(sonnet_lens))
plt.figure(figsize=(15,10))
plt.bar([i for i in range(1, len(sonnets)+1)], sonnet_lens)
plt.title('Number of Characters per sonnet')
plt.ylabel('# Characters')
plt.xlabel('Sonnets')
plt.show()
# -
# Sonnet 145 is Shakespeare's shortest sonnet at 506 characters long while sonnet 11 is the longest with 673 characters. Since the average sonnet length is ~608 characters long when we generate characters later we will generate 600 characters to create 1 sonnet.
# ## Vectorization
# ---
# Now we will set the maximum length a sequence can have, how many characters to skip when we grab a new sample, and create a vector of targets (the next character in the sequence). The `sentences` variable will hold all our training samples where each sample will be 40 characters from the dataset offset by 3 characters from the previous sample. Here is an example where `maxlen` is 40 and `step` is 3:
#
# **Sentence 1:**
#
# 'from fairest creatures we desire increas'
#
# **Target 1:**
#
# 'e'
#
# **Sentence 2:**
#
# 'm fairest creatures we desire increase,\n'
#
# **Target 2:**
#
# 't'
#
# **Sentence 3:**
#
# 'airest creatures we desire increase,\ntha'
#
# **Target 3:**
#
# 't'
#
# For context here are the first 100 characters in the corpus:
#
# 'from fairest creatures we desire increase,\nthat thereby beauty's rose might never die,\nbut as the ri'
# +
# Max length of each sequence
maxlen = 40
# Sample new sequence every step characters
step = 3
sentences = []
targets = []
# Loop through sonnets and create sequences and associated targets
for i in range(0, len(data) - maxlen, step):
sentences.append(data[i:i + maxlen])
targets.append(data[maxlen + i])
print("Number of sequences:", len(sentences))
# Grab all unique characters in corpus
chars = sorted(list(set(data)))
print("Number of unique characters:", len(chars))
# Dictionary mapping unique character to integer indices
char_indices = dict((char, chars.index(char)) for char in chars)
# -
# Now we will vectorize our data into a one-hot encoded tensor of shape **(num_sequences, max_sequence_length, num_characters)**.
# +
# Vectorize sequences and targets
x = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for j, char in enumerate(sentence):
x[i, j, char_indices[char]] = 1
y[i, char_indices[targets[i]]] = 1
print("Size of training sequences:", x.shape)
print("Size of training targets:", y.shape)
# -
# ## Build Model
# ---
# Here we will build and compile the deep learning model using Keras. The model will consist of 1 LSTM layer with a fully connected layer on top. The output has a softmax activation function applied so the output can be represented as a probability for each character in our list of possible characters.
# +
from keras.models import Sequential
from keras.layers import Dense, LSTM
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(chars))))
model.add(Dense(len(chars), activation='softmax'))
model.summary()
# +
from keras.optimizers import SGD
optimizer = SGD(lr=0.01, momentum=0.9, nesterov=True)
model.compile(optimizer=optimizer, loss='categorical_crossentropy')
# -
# The model will output a probability value for each character possible. Instead of choosing the character with the highest probability, we will reweight the probabilities and sample from them based on a "temperature" value. The higher the temperature the more likely a random character will be chosen, the lower the temperature the more deterministic the model will behave.
def sample(preds, temperature=1.0):
''' Reweight the predicted probabilities and draw sample from newly created probability distribution. '''
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
# Now the model will be trained on the text and targets. After training for 1 epoch, a random sequence will be chosen from the training corpus and fed into the model. Using this "seed text" we will predict the next 600 characters at different temperatures and store them in different text files locally.
# +
epochs = 60
loss = [] # Custom history list to save model's loss
# Create directory to store generated text
base_dir = 'generated_text'
if not os.path.isdir(base_dir):
os.mkdir(base_dir)
for epoch in range(1, epochs+1):
print("Epoch", epoch)
# Fit model for 1 epoch then generate text given a seed.
history = model.fit(x, y, batch_size=128, epochs=1)
loss.append(history.history['loss'][0])
# Create directory to store text for each epoch
epoch_dir = os.path.join(base_dir, 'epoch_' + str(epoch))
if not os.path.isdir(epoch_dir):
os.mkdir(epoch_dir)
# Select a random seed text to feed into model and generate text
start_idx = np.random.randint(0, len(data) - maxlen - 1)
seed_text = data[start_idx:start_idx + maxlen]
for temp in [0.2, 0.5, 1.0, 1.3]:
generated_text = seed_text
temp_file = 'epoch' + str(epoch) + '_temp' + str(temp) + '.txt'
file = open(os.path.join(epoch_dir, temp_file), 'w')
file.write(generated_text)
# Predict and generate 600 characters (approx. 1 sonnet length)
for i in range(600):
# Vectorize generated text
sampled = np.zeros((1, maxlen, len(chars)))
for j, char in enumerate(generated_text):
sampled[0, j, char_indices[char]] = 1.
# Predict next character
preds = model.predict(sampled, verbose=0)[0]
pred_idx = sample(preds, temperature=temp)
next_char = chars[pred_idx]
# Append predicted character to seed text
generated_text += next_char
generated_text = generated_text[1:]
# Write to text file
file.write(next_char)
print('Temp ' + str(temp) + " done.")
file.close()
# -
# ## Generating New Sonnets
# ---
# Here we will pick a random seed text from the training data and predict 600 (average sonnet length) new characters using our newly trained model. We will also use a temperature of 0.5 because that gives a good balance between randomness and deterministic behaviour.
def generate_sonnet(temp):
''' Given a temperature, generate a new sonnet '''
start_idx = np.random.randint(0, len(data) - maxlen - 1)
new_sonnet = data[start_idx:start_idx + maxlen]
sys.stdout.write(new_sonnet)
for i in range(600):
# Vectorize generated text
sampled = np.zeros((1, maxlen, len(chars)))
for j, char in enumerate(new_sonnet):
sampled[0, j, char_indices[char]] = 1.
# Predict next character
preds = model.predict(sampled, verbose=0)[0]
pred_idx = sample(preds, temperature=temp)
next_char = chars[pred_idx]
# Append predicted character to seed text
new_sonnet += next_char
new_sonnet = new_sonnet[1:]
# Print to console
sys.stdout.write(next_char)
sys.stdout.flush()
# Generate new sonnets at 0.5 temperature
generate_sonnet(0.5)
# Generate new sonnets at 0.2 temperature
generate_sonnet(0.2)
# Based on the model's output above it learns to format the sonnet correctly most of the time and seems to have learned to separate words with spaces which is impressive since this is a character-level language model. Something interesting it does is it adds punctuation to the end of most lines which is exactly what Shakespeare does in his sonnets. The output where the temperature is 0.2 has a lot of repetition in it. It is probably entering a loop where it sees some sequence of characters before it and predicts the same characters over and over again. This is evident in the last few lines where words like "see the seed" and "the sor" and "and the" appear frequently. Although this behaviour can be expected with such a low temperature. There is not much change to the probability distribution after reweighting the model's outputs.
#
# The first example at temperature 0.5 has more word variety but also more spelling mistakes due to the fact that there is more randomness involved in choosing the next character.
#
# Below the temperature is higher meaning that there will be more random guessing for the next character which explains why there are a lot of spelling mistakes and bizarre words. I think 0.5 is a good temperature for resampling probabilities because too high (1.0 and up) will cause the model to output lot of random letters and will end up being a jumbled mess.
# Generate new sonnet with 1.0 temp
generate_sonnet(1.0)
# Plot model loss over epochs
plt.plot([i for i in range(1, epochs+1)], loss)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Model Loss')
plt.show()
# Now we will save the final model for future use.
# Save model
model.save('shakespeare_sonnet_model.h5')
# ## Things To Try
# ---
# 1. Stack recurrent layers
# 2. Add bidirectional lstm layers
# 3. Change maximum length of sequences (`maxlen` variable)
# 4. Change step size (`step` variable)
# 5. Train over longer epochs
# 6. Change number of lstm units
# 7. Experiment with different optimizers (rmsprop, adam)
# 8. Try word-level language model
# +
from keras.models import load_model
# Load model
model = load_model('shakespeare_sonnet_model.h5')
| Generate-Sonnets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# pypreclin preprocessing
# =======================
#
# Credit: <NAME> & <NAME>
#
# WORK IN PROGRESS
#
# The preprocessing and analysis of nonhuman primate (NHP) magnetic resonance
# imaging (MRI) data presents some unique challenges.
#
# 1. non standard orientation, sphinx position.
# 2. strong intensity bias.
# 3. non brain tissues.
#
# Over the years, we have created our own custom solution that solves these
# problems.
#
# This example shows how to preprocess a functional timeserie without
# fielmap or reverse phase encoded images. One can inpsect the called function to
# apply only a single step.
#
# First checks
# ------------
#
# In order to test if the 'pypreclin' package is installed on your machine, you
# can try to import it.
#
#
from pypreclin.workflow.preproc_fmri import prepoc
from pprint import pprint
# Load a test dataset
# -------------------
#
# Now load a toy single run dataset with 10 volumes.
#
#
rootdir = "/tmp"
funcfile = os.path.join(rootdir, "")
anatfile = ""
sid = ""
tr = ""
outdir = ""
template = ""
jipdir = ""
# Run
# ---
#
# Finally run all the prepocessing steps. In order to avoid the installation
# of all dependencies, please use the provided container.
#
#
outputs = preproc(
funcfile=funcfile,
anatfile=anatfile,
sid=sid,
outdir=outdir,
repetitiontime=tr,
template=template,
jipdir="/jip/bin",
erase=False,
resample=False,
interleaved=False,
sliceorder="ascending",
realign_dof=6,
realign_to_vol=True,
warp=False,
warp_njobs=1,
warp_index=8,
warp_file=None,
warp_restrict=[0, 1, 0],
anatorient="RAS",
funcorient="RAS",
kernel_size=3,
fslconfig="/etc/fsl/5.0.11/fsl.sh",
normalization_trf=None,
coregistration_trf=None,
recon1=False,
recon2=False,
auto=True,
verbose=2)
pprint(outputs)
| examples/.ipynb_checkpoints/preproc-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Latihan Materi Fungsi Lanjutan
#
# **Penulis**: <NAME> | **Lisensi**: CC-BY-SA 4.0
# ## Soal 1
#
# Silakan unduh berkas [ini](https://files.ui.ac.id/f/b8be8be421/) sebagai data yang akan digunakan pada soal ini.
#
# Diberikan sebuah List berisi 100 ribu angka yang dibuat secara acak. Angka tersebut merepresentasikan suatu panjang dari kubus yang akan dihitung volumenya. Tugasmu adalah mengimplementasikan program yang dapat menghitung seluruh volume dari kubus dengan panjang yang disediakan oleh List. Ada beberapa versi yang dapat kamu buat dengan program ini:
#
# 1. Penggunaan iterasi konvensional dengan menghitung volume kemudian menimpa panjang di posisi yang sama (*in-place modification*).
# 2. Penggunaan fungsi <code>map</code> dengan membuat sebuah objek Map baru (tidak perlu dikonversi ke List).
# 3. Penggunaan iterasi dengan pendefinisian **Iterator** sendiri dengan mencetak seluruh kemungkinan volume secara bertahap.
#
# Untuk versi pertama dan kedua, silakan gunakan *template* yang sudah tersedia di bawah ini untuk mengukur waktu eksekusi dari blok program tersebut. Idealnya, versi kedua akan dijalankan lebih cepat dibandingkan versi pertama.
#
# > **Catatan**
# > Kamu bisa langsung mengolah programmu di sini :)
# +
import datetime
"""
Fungsi menggunakan iterasi biasa dilakukan di sini
"""
time1start = datetime.datetime.now()
# Isi kode kamu di sini
print("Elapsed time: {}".format(datetime.datetime.now()-time1start))
"""
Fungsi menggunakan map dilakukan di sini
"""
time2start = datetime.datetime.now()
# Isi kode kamu di sini
print("Elapsed time: {}".format(datetime.datetime.now()-time2start))
| Latihan Fungsi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from addit.ncf import conti, nc2D, nc3D
import jax.numpy as jnp
import numpy as np
def inc1D(x,xv):
"""integrated neighbouring contribution array on a line.
Args:
x: x values (N)
xv: x grid (Ng)
Returns:
integrated neighbouring contribution for i
"""
indarr=jnp.arange(len(xv))
vcl=vmap(conti,(0,None,None),0)
return jnp.sum(vcl(indarr,x,xv),axis=1)
# +
from jax.lax import scan
from jax import jit,vmap
@jit
def inc2D(x,y,xv,yv):
"""integrated 2D neighbouring contribution (memory reduced sum).
Args:
x: x values
y: x values
xv: x grid
yv: x grid
Returns:
neighbouring contribution
Note:
This function computes \sum_n fx_n \otimes fy_n,
where fx_n and fy_n are the n-th NCFs for 1D.
A direct sum uses huge RAM.
In this function, we use jax.lax.scan to compute the sum
Example:
>>>N=10000
>>> xv=jnp.linspace(0,1,11) #grid
>>> yv=jnp.linspace(0,1,11) #grid
>>> x=np.random.rand(N)
>>> y=np.random.rand(N)
>>> val=inc2D(x,y,xv,yv)
The direct sum is computed as
>>> valdirect=jnp.sum(nc2D(x,y,xv,yv),axis=2)
>>> jnp.mean(jnp.sqrt((val/valdirect-1.0)**2))
>>> DeviceArray(2.0836995e-07, dtype=float32)
"""
Ngx=len(xv)
Ngy=len(yv)
indarrx=jnp.arange(Ngx)
indarry=jnp.arange(Ngy)
vcl=vmap(conti,(0,None,None),0)
fx=vcl(indarrx,x,xv) # Ngx x N memory
fy=vcl(indarry,y,yv) # Ngy x N memory
#jnp.sum(fx[:,None]*fy[None,:],axis=2) Ngx x Ngy x N -> huge memory
fxy=jnp.vstack([fx,fy]).T
def fsum(x,arr):
null=0.0
fx=arr[0:Ngx]
fy=arr[Ngx:Ngx+Ngy]
val=x+fx[:,None]*fy[None,:]
return val, null
init0=jnp.zeros((Ngx,Ngy))
val,null=scan(fsum,init0,fxy)
return val
# -
N=10000
xv=jnp.linspace(0,1,11) #grid
yv=jnp.linspace(0,1,15) #grid
x=np.random.rand(N)
y=np.random.rand(N)
val=inc2D(x,y,xv,yv)
valdirect=jnp.sum(nc2D(x,y,xv,yv),axis=2) # direct sum (lots memory)
jnp.mean(jnp.sqrt((val/valdirect-1.0)**2))
import matplotlib.pyplot as plt
fig=plt.figure()
ax=fig.add_subplot(121)
a=plt.imshow(val)
plt.colorbar(a,shrink=0.3)
plt.title("INCF")
ax=fig.add_subplot(122)
a=plt.imshow((val/valdirect-1.0)*100)
plt.colorbar(a,shrink=0.3)
plt.title("difference (%)")
plt.show()
@jit
def inc3D(x,y,z,xv,yv,zv):
"""integrated neighbouring contribution for 3D (memory reduced sum).
Args:
x: x values
y: y values
z: z values
xv: x grid
yv: y grid
zv: z grid
Returns:
integrated neighbouring contribution
Note:
This function computes \sum_n fx_n \otimes fy_n \otimes fz_n,
where fx_n, fy_n, and fz_n are the n-th NCFs for 1D.
A direct sum uses huge RAM.
In this function, we use jax.lax.scan to compute the sum
Example:
>>>N=10000
>>> xv=jnp.linspace(0,1,11) #grid
>>> yv=jnp.linspace(0,1,11) #grid
>>> zv=jnp.linspace(0,1,11) #grid
>>> x=np.random.rand(N)
>>> y=np.random.rand(N)
>>> z=np.random.rand(N)
>>> val=inc3D(x,y,z,xv,yv,zv)
The direct sum is computed as
>>> valdirect=jnp.sum(nc3D(x,y,z,xv,yv,zv),axis=3) # direct sum (lots memory)
>>> jnp.mean(jnp.sqrt((val/valdirect-1.0)**2))
>>> DeviceArray(9.686315e-08, dtype=float32)
"""
Ngx=len(xv)
Ngy=len(yv)
Ngz=len(zv)
indarrx=jnp.arange(Ngx)
indarry=jnp.arange(Ngy)
indarrz=jnp.arange(Ngz)
vcl=vmap(conti,(0,None,None),0)
fx=vcl(indarrx,x,xv) # Ngx x N memory
fy=vcl(indarry,y,yv) # Ngy x N memory
fz=vcl(indarrz,z,zv) # Ngz x N memory
fxyz=jnp.vstack([fx,fy,fz]).T
def fsum(x,arr):
null=0.0
fx=arr[0:Ngx]
fy=arr[Ngx:Ngx+Ngy]
fz=arr[Ngx+Ngy:Ngx+Ngy+Ngz]
val=x+fx[:,None,None]*fy[None,:,None]*fz[None,None,:]
return val, null
init0=jnp.zeros((Ngx,Ngy,Ngz))
val,null=scan(fsum,init0,fxyz)
return val
N=10000
xv=jnp.linspace(0,1,11) #grid
yv=jnp.linspace(0,1,13) #grid
zv=jnp.linspace(0,1,15) #grid
x=np.random.rand(N)
y=np.random.rand(N)
z=np.random.rand(N)
val=inc3D(x,y,z,xv,yv,zv)
valdirect=jnp.sum(nc3D(x,y,z,xv,yv,zv),axis=3) # direct sum (lots memory)
jnp.mean(jnp.sqrt((val/valdirect-1.0)**2))
| snippets/dev/Integrated neighbouring contribution function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cub_data_science
# language: python
# name: cub_data_science
# ---
# # Data Analysis
# ## Link to data: https://www.kaggle.com/fedesoriano/company-bankruptcy-prediction
# +
# Import packages
import numpy as np
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from pprint import pprint
from pickle import dump
from random import sample as r_sample
# Pandas df print formating
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
pd.set_option('display.width', 75)
pd.set_option('display.colheader_justify', 'center')
pd.set_option('display.precision', 3)
# Plotting packages
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (10, 5)
plt.style.use('fivethirtyeight')
# -
# ## Load data
raw_data = pd.read_csv('raw_data.csv')
# ## View and explore data
pprint(raw_data.columns.values.tolist())
# ### Check for nan values
raw_data.isnull().values.any()
# ### Check max values
raw_data.max()
# ### Check min values
raw_data.min()
# ### Plot hist of target values of bankrupcty
dist_y_plot = raw_data["Bankrupt?"].value_counts().plot(kind='bar',rot=0)
plt.title("Spread of target values")
plt.ylabel("Count")
plt.xlabel("0=no and 1=yes")
plt.show(dist_y_plot)
# ## Split into Train and Val
col_names = raw_data.columns
X, y = pd.DataFrame(raw_data.iloc[:,1:]), pd.DataFrame(raw_data.iloc[:,0])
print("X shape: {}".format(X.shape))
print("y shape: {}".format(y.shape))
X.head()
y.head()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=2)
print("Number of train samples: {}".format(len(y_train)))
print("Number of test samples: {}".format(len(y_test)))
print("X_train shape: {}".format(X_train.shape))
print("y_train shape: {}".format(y_train.shape))
print("X_test shape: {}".format(X_test.shape))
print("X_test shape: {}".format(y_test.shape))
# ## Scale input data
# +
# Declare scaler object
scaler = MinMaxScaler()
# Scale w.r.t train data
scaler.fit(X_train)
# Scale train
X_train = scaler.transform(X_train)
# Scale test
X_test = scaler.transform(X_test)
# save the scaler
dump(scaler, open('scaler.pkl', 'wb'))
# load the scaler
# scaler = load(open('scaler.pkl', 'rb'))
# -
# ## Balance train data
# ### Intial test for sanity
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, classification_report
# logistic regression object
lr = LogisticRegression()
# train the model on train set
lr.fit(X_train, y_train.values[:,0])
predictions = lr.predict(X_test)
# print classification report
print(classification_report(y_test.values[:,0], predictions))
# -
# ## Apply SMOTE to balance data
# +
print("Before OverSampling, counts of label '1': {}".format(sum(y_train.values == 1)))
print("Before OverSampling, counts of label '0': {} \n".format(sum(y_train.values == 0)))
from imblearn.over_sampling import SMOTE
oversample = SMOTE()
X_train, y_train = oversample.fit_resample(X_train, y_train.values)
print('After OverSampling, the shape of train_X: {}'.format(X_train.shape))
print('After OverSampling, the shape of train_y: {} \n'.format(y_train.shape))
print("After OverSampling, counts of label '1': {}".format(sum(y_train == 1)))
print("After OverSampling, counts of label '0': {}".format(sum(y_train == 0)))
# -
# ### Post test for sanity
# +
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import confusion_matrix, classification_report
# logistic regression object
lr = LogisticRegression()
# train the model on train set
lr.fit(X_train, y_train)
predictions = lr.predict(X_test)
# print classification report
print(classification_report(y_test.values[:,0], predictions))
# -
# ## Save new data
pd.DataFrame(X_train,columns=col_names[1:]).to_csv("X_train.csv",index=False)
pd.DataFrame(X_test,columns=col_names[1:]).to_csv("X_test.csv",index=False)
pd.DataFrame(y_train,columns=[col_names[0]]).to_csv("y_train.csv",index=False)
pd.DataFrame(y_test,columns=[col_names[0]]).to_csv("y_test.csv",index=False)
| ml_dev_tutorial/Data/dataPrep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="x275tSx2asW0"
# # Anomaly Detection - KDD'99 Dataset
# -
# In week 4 of the inzva Applied AI program we will spend most of our time on Anomaly Detection techniques.
#
# **This notebook includes**
#
# *1- Isolation Forest*
#
# *2- Local Outlier Factor*
#
# *3- Autoencoder*
#
# *4 - XGBoosting*
#
# We first start with obtaining the data and creating training and test sets. In this notebook, we create our models only for Probing attack detection.
#
#
# + [markdown] colab_type="text" id="sDK-7yCKbWL8"
# ## 1. Data
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="lOPme80Z1TKg" outputId="91802941-a964-488b-fcca-d97dd7b9132a"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={} colab_type="code" id="ZCSpFnNe1UlI"
import pandas as pd
kdd = pd.read_csv('/content/gdrive/My Drive/kddcup.data/kdd.csv')
# + colab={} colab_type="code" id="EI2zzt-K_2mL"
kdd = kdd.iloc[:,1:43]
# + colab={} colab_type="code" id="m-tUOxq4_4xU"
kdd = kdd.drop(['Protocol Type', 'Service', 'Flag'], axis = 1)
# + colab={} colab_type="code" id="qZTBqq6xxzdS"
kdd_train = kdd.iloc[0:102563, :]
# + colab={} colab_type="code" id="JQ2KBBexxzdU"
kdd_test = kdd.iloc[102563:183737, :]
# + colab={} colab_type="code" id="gemL8-7axzdW"
kdd_train_probe = kdd_train[(kdd_train.Type_Groups == 'Normal') | (kdd_train.Type_Groups == 'Probe')]
# + colab={} colab_type="code" id="BCklITVMxzdZ"
kdd_test_probe = kdd_test[(kdd_test.Type_Groups == 'Normal') | (kdd_test.Type_Groups == 'Probe')]
# + [markdown] colab_type="text" id="VGnCZb2kghyf"
# Normals are encoded as 1, anomalies are encoded as -1. Use this version if you want to apply Isolation Forest or Local Outlier Factor. Otherwise, please run the code in the next chunk.
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="iNwybydCxzda" outputId="8b6f9d40-9d39-4115-e436-f2efbdd7dbee"
import numpy as np
kdd_train_probe['Type_Groups'] = np.where(kdd_train_probe['Type_Groups'] == 'Normal', 1, -1)
kdd_test_probe['Type_Groups'] = np.where(kdd_test_probe['Type_Groups'] == 'Normal', 1, -1)
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="fq9aMv8iACrc" outputId="ae9b1dcf-f56f-4e0b-9dd1-e7c85eff3c54"
kdd_train_probe['Type_Groups'] = np.where(kdd_train_probe['Type_Groups'] == 'Normal', 0, 1)
kdd_test_probe['Type_Groups'] = np.where(kdd_test_probe['Type_Groups'] == 'Normal', 0, 1)
# + [markdown] colab_type="text" id="h0VhI_Yohn5v"
# ## 1.1 Isolation Forest
# + colab={} colab_type="code" id="hPRl1mULdBK8"
x_train = kdd_train_probe.drop(['Type_Groups'], axis = 1)
y_train = kdd_train_probe['Type_Groups']
# + colab={} colab_type="code" id="xTk1whWyiSr9"
x_test = kdd_test_probe.drop(['Type_Groups'], axis = 1)
y_test = kdd_test_probe['Type_Groups']
# + colab={} colab_type="code" id="tkh6vw8tdh8A"
clfIF = IsolationForest(max_samples=0.25, random_state=11, contamination=0.15, n_estimators=100, n_jobs=-1)
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="kHrohm2_dmap" outputId="42c25894-53a3-465e-e590-81103d213d85"
clfIF.fit(x_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="euBiTJ5LeGl8" outputId="9a79f048-6631-40ad-b03c-515ff94d8fc2"
y_pred_train = clfIF.predict(x_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="goT-4ZQBfL1H" outputId="8eca80fa-ce2e-4530-cce3-71a7dfc55dcc"
np.unique(y_pred_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="TxmfACrjeNl8" outputId="09683a4e-07b0-4603-cb5c-0356db77192f"
conf_matrix = confusion_matrix(y_train, y_pred_train)
conf_matrix
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="TJJDsmz7iNAR" outputId="eb2bba32-abe6-47c1-cc6a-26e06b0ad345"
y_pred_test = clfIF.predict(x_test)
conf_matrix_if_test = confusion_matrix(y_test, y_pred_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="8BM4PcCElPf6" outputId="222283f5-0117-43dd-e625-e59e0a460879"
conf_matrix_if_test
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="5gNlUF0DmtoO" outputId="3cc30c30-a5af-4745-fb7c-0f14b6312959"
print(classification_report(y_test, y_pred_test))
# + colab={} colab_type="code" id="Tp_WKBzfcKFb"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import itertools
from sklearn.ensemble import IsolationForest
from sklearn.neighbors import LocalOutlierFactor
from sklearn import svm
from sklearn.neighbors import NearestNeighbors
import seaborn as sns
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import recall_score
from sklearn.metrics import roc_auc_score
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
def plot_confusion_matrix(cm, title, classes=['abnormal', 'normal'],
cmap=plt.cm.Blues, save=False, saveas="MyFigure.png"):
# print Confusion matrix with blue gradient colours
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=90)
plt.yticks(tick_marks, classes)
fmt = '.1%'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
if save:
plt.savefig(saveas, dpi=100)
# + [markdown] colab_type="text" id="oDNzbAGJhd4I"
# ## 1.3 Local Outlier Factor
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="KxpmhajEhGyL" outputId="64cadcaf-00fc-4bf2-d84e-fba164b427ad"
clfLOF = LocalOutlierFactor(n_neighbors=15, metric='euclidean', algorithm='auto', contamination=0.15, n_jobs=-1)
clfLOF.fit(x_train, y_train)
# + colab={} colab_type="code" id="2fO9j-HPh3Fi"
y_pred_train_lof = clfLOF.fit_predict(x_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="brYaF_LgjX72" outputId="1fb9f3e8-06d7-4b78-9523-f34b8b5dbb7e"
conf_matrix_lof = confusion_matrix(y_train, y_pred_train_lof)
conf_matrix_lof
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="fMymZ_MNlG1O" outputId="c91baf3f-41ce-43db-94e7-7aa137b4b2eb"
y_pred_test_lof = clfLOF.fit_predict(x_test, y_test)
conf_matrix_lof_test = confusion_matrix(y_test, y_pred_test_lof)
conf_matrix_lof_test
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="oQHzcfiRlmP0" outputId="d1722fd8-3782-4aaf-d336-a23bb4296acf"
print(classification_report(y_test, y_pred_test_lof))
# + [markdown] colab_type="text" id="FrY00ZRxn9lS"
# ## 1.4 Autoencoder
# + colab={"base_uri": "https://localhost:8080/", "height": 80} colab_type="code" id="QzQ5iH-OpgQP" outputId="50344490-d847-4e47-ddef-dca5e56cc890"
import pickle
from scipy import stats
import tensorflow as tf
from pylab import rcParams
from sklearn.model_selection import train_test_split
from keras.models import Model, load_model
from keras.layers import Input, Dense
from keras.callbacks import ModelCheckpoint, TensorBoard
from keras import regularizers
# + [markdown] colab_type="text" id="6qZu26TppGkF"
# ## 1.4.1 Probing Attack Detection
# + [markdown] colab_type="text" id="PEijPpzhXh5t"
# ## We need to create Validation Test
# + colab={"base_uri": "https://localhost:8080/", "height": 162} colab_type="code" id="RXYZxz5DXmat" outputId="3688ae8c-4f8d-4d5a-e4a6-fc7b521fd728"
kdd_test_probe.groupby('Type_Groups').count()
# + colab={"base_uri": "https://localhost:8080/", "height": 162} colab_type="code" id="K_uA_W-XX67U" outputId="ba511863-f343-480e-a445-f55bf99b1fba"
kdd_test_probe.iloc[14000:34000,:].groupby('Type_Groups').count()
# + colab={} colab_type="code" id="0NnV4TglYmJD"
kdd_valid_probe = kdd_test_probe.iloc[14000:34000,:]
# + colab={} colab_type="code" id="E_AkIPVsZeBL"
kdd_test_v2_probe = pd.concat([kdd_test_probe.iloc[0:14000,:], kdd_test_probe.iloc[34001:64759,:]])
# + colab={"base_uri": "https://localhost:8080/", "height": 162} colab_type="code" id="firflwnBY1yR" outputId="83c45372-703e-4c62-f449-d4bf3be4c2fa"
kdd_test_v2_probe.groupby('Type_Groups').count()
# + [markdown] colab_type="text" id="iKgW7zMsecSp"
# Now that we obtained Train, Test and Validation sets, we can train our data and optimize the reconstruction error threshold using test set. Then, we will perform our actual prediction tast by using Validation set.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="zZF3-B2GzG4H" outputId="0b06807c-0555-4e05-faa3-3505036b18dd"
X_train, X_test = kdd_train_probe, kdd_test_v2_probe
X_train = X_train[X_train.Type_Groups == 0]
X_train = X_train.drop(['Type_Groups'], axis=1)
y_test = X_test['Type_Groups']
X_test = X_test.drop(['Type_Groups'], axis=1)
X_train = X_train.values
X_test = X_test.values
X_train.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 139} colab_type="code" id="ddmJ-5CSpPrw" outputId="c2002f28-cdcb-4fd0-a861-9304aa064da0"
input_dim = X_train.shape[1]
encoding_dim = 14
input_layer = Input(shape=(input_dim, ))
encoder = Dense(encoding_dim, activation="tanh",
activity_regularizer=regularizers.l1(10e-5))(input_layer)
encoder = Dense(int(encoding_dim / 2), activation="relu")(encoder)
decoder = Dense(int(encoding_dim), activation='tanh')(encoder)
decoder = Dense(input_dim, activation='relu')(decoder)
autoencoder_corr = Model(inputs=input_layer, outputs=decoder)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="5CWGL4-b2umu" outputId="be1b73ba-3a41-475c-86eb-a84fbd5ac7c7"
nb_epoch = 34
batch_size = 100
batch_index=0
autoencoder_corr.compile(optimizer='adam',
loss='mean_squared_error',
metrics=['accuracy'])
checkpointer = ModelCheckpoint(filepath="model.h5", #TO SAVE THE MODEL
verbose=0,
save_best_only=True)
tensorboard = TensorBoard(log_dir='./logs',
histogram_freq=0,
write_graph=True,
write_images=True)
history = autoencoder_corr.fit(X_train, X_train, #INPUT AND EXPECTED OUTPUT ARE THE SAME.
epochs=nb_epoch,
batch_size=batch_size,
shuffle=True,
validation_data=(X_test, X_test),
verbose=1,
callbacks=[checkpointer, tensorboard]).history
# + [markdown] colab_type="text" id="p1wb7_MsBO05"
# # Tune the threshold value using training set
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="v_WijHwGBOjs" outputId="8f51e62e-756c-4b69-9172-a7d00f9f284b"
predictions = autoencoder_corr.predict(X_test)
mse = np.mean(np.power(X_test - predictions, 2), axis=1)
error_df = pd.DataFrame({'reconstruction_error': mse,
'true_class': y_test})
error_df.describe()
# + colab={} colab_type="code" id="NWhNe_P7BOVz"
threshold = np.arange(0, 1, 0.01)
results = np.zeros(100)
probe = np.zeros(100)
i = 0
for t in threshold:
y_pred = [1 if e > t else 0 for e in error_df.reconstruction_error.values]
conf_matrix = confusion_matrix(error_df.true_class, y_pred)
results[i] = (conf_matrix[0,0] + conf_matrix[1,1]) / 64759
probe[i] = conf_matrix[1,1] / 4166
i = i + 1
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="vEPiSbh8YNDc" outputId="661da4c8-6a92-41fb-c14d-8ba14cbf0a61"
predictions = autoencoder_corr.predict(X_test)
mse = np.mean(np.power(X_test - predictions, 2), axis=1)
error_df = pd.DataFrame({'reconstruction_error': mse,
'true_class': y_test})
error_df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="UlWYF4Cf3XLL" outputId="5b17ed7c-0d35-4b11-c4cd-e8b5de79af53"
threshold = 0.13 # 0.20 de iyi
LABELS = ["Normal", "Probing"]
y_pred = [1 if e > threshold else 0 for e in error_df.reconstruction_error.values]
conf_matrix = confusion_matrix(error_df.true_class, y_pred)
#plt.figure(figsize=(12, 12))
#sns.heatmap(conf_matrix, xticklabels=LABELS, yticklabels=LABELS, annot=True, fmt="d");
#plt.title("Confusion matrix")
#plt.ylabel('True class')
#plt.xlabel('Predicted class')
#plt.show()
conf_matrix
# + colab={} colab_type="code" id="wJEkAmeC3v1v"
from sklearn.metrics import (confusion_matrix, precision_recall_curve, auc,
roc_curve, recall_score, classification_report, f1_score,
precision_recall_fscore_support)
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="oyHacenx3VJu" outputId="506301f5-0c2c-4f9a-9098-0b7c3fced5ce"
fpr, tpr, thresholds = roc_curve(error_df.true_class, error_df.reconstruction_error)
roc_auc = auc(fpr, tpr)
plt.title('Receiver Operating Characteristic')
plt.plot(fpr, tpr, label='AUC = %0.4f'% roc_auc)
plt.legend(loc='lower right')
plt.plot([0,1],[0,1],'r--')
plt.xlim([-0.001, 1])
plt.ylim([0, 1.001])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show();
# + colab={} colab_type="code" id="4HhIPx8t3XP7"
threshold = np.arange(0, 1, 0.01)
results = np.zeros(100)
probe = np.zeros(100)
i = 0
for t in threshold:
y_pred = [1 if e > t else 0 for e in error_df.reconstruction_error.values]
conf_matrix = confusion_matrix(error_df.true_class, y_pred)
results[i] = (conf_matrix[0,0] + conf_matrix[1,1]) / 44758
probe[i] = conf_matrix[1,1] / 2922
i = i + 1
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="ki6gdyl6jGo4" outputId="f08c288f-3eb8-4be5-d11a-3f8aba14d29e"
probe
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="Y4pippgbjLHG" outputId="c95b3652-782d-4f72-fa6b-f713c4209714"
threshold
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="6ZxbaJfHG24M" outputId="e2134a91-9504-4207-e88c-bf2a7bc7fac9"
import matplotlib.pyplot as plt
plt.plot(threshold, probe)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="pS90TZJcHGGD" outputId="285fccd4-dafb-42a2-ab38-2333d3178ad1"
plt.plot(threshold, results)
# + [markdown] colab_type="text" id="rXN_FzmBjUfP"
# Let's try if we can find a better threshold value between 0.14 and 0.18.
# + colab={} colab_type="code" id="8kLPqf61BAf6"
threshold = np.arange(0.3, 0.5, 0.001)
results = np.zeros(200)
probe = np.zeros(200)
i = 0
for t in threshold:
y_pred = [1 if e > t else 0 for e in error_df.reconstruction_error.values]
conf_matrix = confusion_matrix(error_df.true_class, y_pred)
results[i] = (conf_matrix[0,0] + conf_matrix[1,1]) / 44758
probe[i] = conf_matrix[1,1] / 2922
i = i + 1
# + colab={"base_uri": "https://localhost:8080/", "height": 408} colab_type="code" id="zI558plHkG08" outputId="389a9a29-84c5-4bd3-b279-8d6a9e274afa"
threshold
# + colab={"base_uri": "https://localhost:8080/", "height": 697} colab_type="code" id="Cj9GcA5gkUrH" outputId="4c457248-c370-4232-db63-af70868dc5e5"
probe
# + colab={"base_uri": "https://localhost:8080/", "height": 697} colab_type="code" id="cmJ2X6z7klyj" outputId="226f9166-9438-49bc-d375-74dec3fe34e9"
results
# + [markdown] colab_type="text" id="vydMN6zBmjDq"
# Let's fix our threshold value to 0.499 and apply it to Validation Set.
# + colab={} colab_type="code" id="JvofCF2WmnyC"
X_valid = kdd_valid_probe
y_valid = X_valid['Type_Groups']
X_valid = X_valid.drop(['Type_Groups'], axis=1)
X_valid = X_valid.values
# + colab={"base_uri": "https://localhost:8080/", "height": 297} colab_type="code" id="zc_9M1VNm38D" outputId="eca32605-5914-4e72-d1e7-7bcad0cdf0f9"
predictions = autoencoder_corr.predict(X_valid)
mse = np.mean(np.power(X_valid - predictions, 2), axis=1)
error_df = pd.DataFrame({'reconstruction_error': mse,
'true_class': y_valid})
error_df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="Tb9UyZXmnA03" outputId="e0e6f281-8b93-43f1-e1f7-3e4c57f4d3aa"
threshold = 0.499 # 0.20 de iyi
LABELS = ["Normal", "Probing"]
y_pred = [1 if e > threshold else 0 for e in error_df.reconstruction_error.values]
conf_matrix = confusion_matrix(error_df.true_class, y_pred)
conf_matrix
# + [markdown] colab_type="text" id="2a7efGoZLk1O"
# ## XGBoosting
# + colab={"base_uri": "https://localhost:8080/", "height": 139} colab_type="code" id="tzb8DuA_7GDw" outputId="9779a015-2ac0-4148-9d0b-261f8e780760"
from numpy import loadtxt
from sklearn.metrics import accuracy_score
import os
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import f1_score, roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.neural_network import MLPClassifier
from imblearn.pipeline import make_pipeline as make_pipeline_imb
from imblearn.over_sampling import SMOTE
from imblearn.metrics import classification_report_imbalanced
import xgboost as xgb
from xgboost import XGBClassifier
import pandas as pd
import matplotlib.pyplot as plt
from collections import Counter
import time
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="7wFt0NEX6gWA" outputId="cf12015b-e77f-484f-f645-87ea629f86a2"
kdd_train_probe.head()
# + colab={} colab_type="code" id="Dj2ak4hnLmbq"
def xgb_classifier(X_train, X_test, y_train, y_test, useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
"""
# {'learning_rate': 0.1, 'max_depth': 5, 'min_child_weight': 3} 0.862920874517388
# {'colsample_bytree': 1.0, 'gamma': 0.2} 0.871
# {'gamma': 0.2, 'scale_pos_weight': 1} 0.8702009952422571
# {'subsample': 0.6} 0.864310306628855
"""
alg = XGBClassifier(learning_rate=0.1,
n_estimators=140,
max_depth=5, # maximum number of features used in each tree
min_child_weight=3,
gamma=0.2,
subsample=0.6, #hich is for each tree the % of rows taken to build the tree.
#I recommend not taking out too many rows, as performance will drop a lot. Take values from 0.8 to 1.
colsample_bytree=1.0, # number of columns used by each tree.
#In order to avoid some columns to take too much credit for the prediction
#(think of it like in recommender systems when you recommend the most purchased products
#and forget about the long tail), take out a good proportion of columns.
#Values from 0.3 to 0.8 if you have many columns (especially if you did one-hot encoding),
#or 0.8 to 1 if you only have a few columns.
objective='binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
if useTrainCV:
print("Start Feeding Data")
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(X_train.values, label=y_train.values)
# xgtest = xgb.DMatrix(X_test.values, label=y_test.values)
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,
early_stopping_rounds=early_stopping_rounds)
alg.set_params(n_estimators=cvresult.shape[0])
#
print('Start Training')
alg.fit(X_train, y_train, eval_metric='auc')
# param_test1 = {}
# gsearch1 = GridSearchCV(estimator=XGBClassifier(learning_rate=0.1, n_estimators=140, max_depth=5,
# min_child_weight=3, gamma=0.2, subsample=0.8,
# colsample_bytree=1.0,
# objective='binary:logistic', nthread=4, scale_pos_weight=1,
# seed=27),
# param_grid=param_test1,
# scoring='f1',
# n_jobs=4, iid=False, cv=5)
# gsearch1.fit(X_train, y_train)
# print(gsearch1.cv_results_, gsearch1.best_params_, gsearch1.best_score_)
#
print("Start Predicting")
predictions = alg.predict(X_test)
pred_proba = alg.predict_proba(X_test)[:, 1]
#
print("\nResulting Metrics")
print("Accuracy : %.4g" % metrics.accuracy_score(y_test, predictions))
print("AUC : %f" % metrics.roc_auc_score(y_test, pred_proba))
print("F1 Score : %f" % metrics.f1_score(y_test, predictions))
feat_imp = alg.feature_importances_
feat = X_train.columns.tolist()
# clf.best_estimator_.booster().get_fscore()
res_df = pd.DataFrame({'Features': feat, 'Importance': feat_imp}).sort_values(by='Importance', ascending=False)
res_df.plot('Features', 'Importance', kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
plt.show()
print(res_df)
print(res_df["Features"].tolist())
# + colab={} colab_type="code" id="B4gndvi-6FsL"
y_train = kdd_train_probe['Type_Groups']
# + colab={} colab_type="code" id="F_gWM69b6vdw"
X_train = kdd_train_probe.drop(['Type_Groups'], axis = 1)
# + colab={} colab_type="code" id="VX90qotY6z-L"
y_test = kdd_test_probe['Type_Groups']
X_test = kdd_test_probe.drop(['Type_Groups'], axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="1sikZut16_gj" outputId="a4701674-657a-4d01-af3a-c7227b815400"
xgb_classifier(X_train, X_test, y_train, y_test)
# + colab={} colab_type="code" id="RGFn4DS27Ela"
| Applied AI Study Group #3 - June 2020/week4/1 - Anomaly Detection Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
pd.set_option('display.max_columns', None)
airbnb=pd.read_csv('Data-Project.csv',index_col=False)
airbnb.head(2)
yelp=pd.read_csv('yelp_cleaned.csv',index_col=False)
yelp.drop(['Unnamed: 0'],axis=1,inplace=True)
yelp=yelp[yelp['borough']!='New Jersey']
# +
categories = []
for iter in yelp['category title']:
categories.extend(str.split(str(iter), sep=','))
categories = pd.Series(categories)
categories = categories.str.lstrip()
categories.value_counts()
# -
def unpack_categories(df, column):
for index, value in df[[column]].iterrows():
df.loc[index, 'is_restaurant'] = ('Bars' not in str(value))
df.loc[index, 'is_bar'] = ('Bars' in str(value)) | ('Nightlife' in str(value))
return df
yelp_df_copy = yelp.copy()
yelp_df_copy = unpack_categories(yelp_df_copy, 'category title')
yelp_df_copy.is_restaurant.value_counts()
airbnb_nhood=pd.DataFrame(airbnb['neighborhood'].value_counts().reset_index())
airbnb_nhood.isnull().sum()
yelp_nhood = pd.DataFrame(yelp_df_copy['neighborhood'].value_counts().reset_index())
match_hoods = pd.merge(airbnb_nhood, yelp_nhood,on=['index'], how='outer',suffixes=('_airbnb', '_yelp'))
# +
#Count records where there is no match on each side.
airbnb_nomatch = match_hoods.neighborhood_airbnb[match_hoods.neighborhood_yelp.isnull()]
print('There are ' + str(airbnb_nomatch.sum()) + ' AirBnB listings with no match in the Yelp data.')
yelp_nomatch = match_hoods.neighborhood_yelp[match_hoods.neighborhood_airbnb.isnull()]
print('There are ' + str(yelp_nomatch.sum()) + ' Yelp listings with no match in the AirBnB data.')
# -
match_hoods=match_hoods.dropna()
yelp_df_copy=yelp_df_copy[~(yelp_df_copy.neighborhood.isin(["Yonkers", "New Rochelle"]))]
# +
#hAVERSINE: https://stackoverflow.com/questions/19412462/getting-distance-between-two-points-based-on-latitude-longitude
from math import sin, cos, sqrt, atan2, radians
def compute_distance(lat1, lon1, lat2, lon2):
# approximate radius of earth in km
R = 6373.0
lat1 = radians(lat1)
lon1 = radians(lon1)
lat2 = radians(lat2)
lon2 = radians(lon2)
dlon = lon2 - lon1
dlat = lat2 - lat1
a = sin(dlat / 2)**2 + cos(lat1) * cos(lat2) * sin(dlon / 2)**2
c = 2 * atan2(sqrt(a), sqrt(1 - a))
distance = R * c
return distance
# -
yelp_restaurants=yelp_df_copy.loc[yelp_df_copy['is_restaurant'] == True]
yelp_restaurants.drop(['is_bar'],axis=1,inplace=True)
yelp_bar=yelp_df_copy.loc[yelp_df_copy['is_bar'] == True]
yelp_bar.drop(['is_restaurant'],axis=1,inplace=True)
a_hoods = airbnb[['id_airbnb', 'neighborhood', 'latitude', 'longitude']]
a_hoods.head()
y_hoods_r = yelp_restaurants[['id', 'neighborhood', 'latitude', 'longitude','review_count','rating']]
y_hoods_r.head()
y_hoods_b = yelp_bar[['id', 'neighborhood', 'latitude', 'longitude','review_count','rating']]
y_hoods_b.head()
restaurants = pd.merge(a_hoods, y_hoods_r, how='inner', on='neighborhood', suffixes = ('_airbnb', '_yelp'))
restaurants.shape
restaurants['distance'] = list(map(compute_distance, restaurants['latitude_airbnb'],
restaurants['longitude_airbnb'], restaurants['latitude_yelp'],
restaurants['longitude_yelp']))
restaurants_1m = restaurants[restaurants['distance']<=1]
#filter for distance < .5
restaurants_2m =restaurants[restaurants['distance']<=2]
print(restaurants_1m.shape)
print(restaurants_2m.shape)
restaurants_1m
#For everything within .1 mile:
restaurants_1mile = restaurants_1m.groupby(['id_airbnb']).agg({'id':'count',
'review_count':'sum',
'rating':'mean'})
restaurants_1mile = restaurants_1mile.rename(columns={'id':'total_restaurants_1mile',
'review_count':'total_restaurants_reviews_1mile',
'rating':'restaurants_avg_rating_1mile'})
restaurants_1mile.head()
#For everything within .1 mile:
restaurants_2mile = restaurants_2m.groupby(['id_airbnb']).agg({'id':'count',
'review_count':'sum',
'rating':'mean'})
restaurants_2mile = restaurants_2mile.rename(columns={'id':'restaurants_yelp_2mile',
'review_count':'total_restaurants_reviews_2mile',
'rating':'restaurants_avg_rating_2mile'})
all_restaurants = pd.merge(restaurants_1mile , restaurants_2mile , how='right', on='id_airbnb')
all_restaurants.head()
bars = pd.merge(a_hoods, y_hoods_b, how='inner', on='neighborhood', suffixes = ('_airbnb', '_yelp'))
#print(bars.shape)
bars['distance'] = list(map(compute_distance, bars['latitude_airbnb'],
bars['longitude_airbnb'], bars['latitude_yelp'],
bars['longitude_yelp']))
#print(bars.head())
#filter for distance < 1
bars_1m = bars[bars['distance']<=1]
#filter for distance < 2
bars_2m =bars[bars['distance']<=2]
#print(bars_1.shape)
#print(bars_5.shape)
#For everything within .1 mile:
bars_1mile = bars_1m.groupby(['id_airbnb']).agg({'id':'count',
'review_count':'sum',
'rating':'mean'})
bars_1mile.head()
bars_1mile = bars_1mile.rename(columns={'id':'total_bars_1mile',
'review_count':'total_bars_reviews_1mile',
'rating':'bars_avg_rating_1mile'})
#print(bars_1mile.head())
#For everything within .1 mile:
bars_2mile = bars_2m.groupby(['id_airbnb']).agg({'id':'count',
'review_count':'sum',
'rating':'mean'})
bars_2mile = bars_2mile.rename(columns={'id':'bars_yelp_2mile',
'review_count':'total_bars_reviews_2mile',
'rating':'bars_avg_rating_2mile'})
all_bars = pd.merge(bars_1mile , bars_2mile , how='right', on='id_airbnb')
print(all_bars.head())
rest_df=pd.merge(all_restaurants, all_bars, on='id_airbnb', how='left')
airbnb_df=pd.merge(airbnb, rest_df, on='id_airbnb', how='left')
airbnb_df.head(2)
airbnb_df.isnull().sum()
airbnb_df.to_csv('Data-Project.csv')
| Jupyter Notebook/Merge/Restaurants-1-2mile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Input and Output
#
# ## Print Formatting
#
# We've already seen the print command.
#
# In Python 3, print is actually a function not a statement
print 'This is a string'
# +
x = 'String'
print 'Place my variable here: ' + x
# +
x = 'String'
print 'Place my variable here: %s' %(x)
# +
x = 123.45
print 'Place my variable here: %s' %(x)
# -
print 'Floating point number: %1.2f' %(13.145)
print 'Floating point number: %1.3f' %(13.145)
print 'Floating point number: %1.1f' %(13.145)
print 'Floating point number: %1.10f' %(13.145)
print 'Floating point number: %25.10f' %(13.145)
# ## Conversion Format Methods
#
# * Convert to string
#
print 'Convert to string %r' %(123)
print 'First; %s, Second: %s, Third: %s' %('hi!', 'two', 3)
print 'First; %s, Second: %s' %(2,2)
print 'First: {x} Second: {x}'.format(x='inserted')
print 'First: {x} New: {y} Second: {x}'.format(x='inserted', y = 'two')
# ## References
# 1. [Python Formatting](https://pyformat.info)
| notebooks/02b_input_and_output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estrutura de Repetição: for
#
# ### Funcionamento:
# + active=""
# for i in range(n):
# repetir código n vezes
# -
for i in range(5):
print('diego')
#i é o indice - é a contagem da execução
for i in range(5):
print(i)
# - Imagine que você está construindo uma automação para enviar todo dia por e-mail um resumo da produção de uma fábrica. Construa um código que exiba a quantidade produzida de cada os produto nesse "e-mail"
# +
produtos = ['coca', 'pepsi', 'guarana', 'sprite', 'fanta']
producao = [15000, 12000, 13000, 5000, 250]
for i in range(5):
print('{} unidade produzidas de {}'.format(producao[i], produtos[i]))
# -
produtos = ['coca', 'pepsi', 'guarana', 'sprite', 'fanta']
producao = [15000, 12000, 13000, 5000, 250]
tamanho_lista = len(produtos)
for i in range(tamanho_lista):
print('{} unidade produzidas de {}'.format(producao[i], produtos[i]))
| Arquivo aulas/Modulo 6/MOD6-Aula1(for).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## Exploring the City of Saint Louis public safety data with Apache Spark 2.3
# The Saint Louis OpenData project contains hundreds of datasets for the city of Saint Louis. Open government data has the potential to increase the quality of life for residents, create more efficient government services, better public decisions, and even create new local businesses and services.
# 
# We start by mounting the Amazon S3 storage to the notebook.
# +
ACCESS_KEY = "<KEY>"
SECRET_KEY = "<KEY>"
ENCODED_SECRET_KEY = SECRET_KEY.replace("/", "%2F")
AWS_BUCKET_NAME = "xxxxx-xxxx"
MOUNT_NAME = "input"
dbutils.fs.mount("s3a://%s:%s@%s" % (ACCESS_KEY, ENCODED_SECRET_KEY, AWS_BUCKET_NAME), "/mnt/%s" % MOUNT_NAME)
display(dbutils.fs.ls("/mnt/%s" % MOUNT_NAME))
# -
# List all mounted points
display(dbutils.fs.mounts())
# The 2020 safety data of Saint Louis city is uploaded to S3 in csv format and pulled down to Databricks for analysis. You can list the file with the `%fs ls` command
# %fs ls /mnt/input/input
# Note, I combined 12 csv files into 1 file to get a full year of data. I downloaded directly from this link: http://www.slmpd.org/Crimereports.shtml
# I'm using "spark" as an entry point into all functionality in Spark 2.3.
spark
# Using the SparkSession, create a DataFrame from the CSV file by inferring the schema.
crimeDataDF = spark.read.csv('/mnt/input/input/2020stlcrimedata.csv', header=True, inferSchema=True)
crimeDataDF.count()
# Display the data using a display function by Databricks.
display(crimeDataDF)
# Notice that the above cell takes ~2 seconds to run b/c it is inferring the schema by sampling the file and reading through it.
#
# Inferring the schema works for ad-hoc analysis against smaller datasets. But when working on terabyte of data, it's better to provide an **explicit pre-defined schema manually**, so there's no inferring cost:
from pyspark.sql.types import StructType, StructField, IntegerType, StringType, BooleanType, DecimalType
# Note that we are removing all space characters from the col names to prevent errors when writing to Parquet later
crimeSchema = StructType(
[
StructField('Complaint', StringType(), True),
StructField('CodedMonth', StringType(), True),
StructField('DateOccur', StringType(), True),
StructField('FlagCrime', StringType(), True),
StructField('FlagUnfounded', StringType(), True),
StructField('FlagAdministrative', StringType(), True),
StructField('Count', IntegerType(), True),
StructField('FlagCleanup', StringType(), True),
StructField('Crime', IntegerType(), True),
StructField('District', IntegerType(), True),
StructField('Description', StringType(), True),
StructField('ILEADSAddress', IntegerType(), True),
StructField('ILEADSStreet', StringType(), True),
StructField('Neighborhood', IntegerType(), True),
StructField('LocationName', StringType(), True),
StructField('LocationComment', StringType(), True),
StructField('CADAddress', IntegerType(), True),
StructField('CADStreet', StringType(), True),
StructField('XCoord', StringType(), True),
StructField('YCoord', StringType(), True)
]
)
crimeDataSDF = spark.read.csv('/mnt/input/input/2020stlcrimedata.csv', header=True, schema=crimeSchema)
display(crimeDataSDF)
# The csv file contains null records so we drop all null records from the table.
crimeDataSDF.na.drop(subset="Complaint")
crimeDataSDF.count()
# Look at the first 5 records in the DataFrame:
crimeDataSDF.show(5)
# Print just the column names in the DataFrame:
crimeDataSDF.columns
# Count how many rows total there are in DataFrame (and see how long it takes to do a full scan from remote disk/S3):
crimeDataSDF.count()
# There are over ~46 thousand rows in the DataFrame and it takes ~2 seconds to do a full read of it.
# ### **Analysis with PySpark DataFrames and Spark SQL API**
# Create a temp view to use spark.sql
crimeDataSDF.createOrReplaceTempView("crimesql")
# **Q-1) How many different types of calls were made to the Police Department?**
crimeDataSDF.select('Description').distinct().show(35, False)
display(sqlContext.sql("SELECT Description FROM crimesql GROUP BY description"), limit=35)
# The queries above show the different type of calls to the police department.
# **Q-2) How many incidents of each call type were there?**
display(crimeDataSDF.select('Description').groupBy('Description').count().orderBy('Count', ascending=False))
display(sqlContext.sql("SELECT Description, count(*) as count FROM crimesql GROUP BY description ORDER BY count desc"))
# Seems like the Saint Louis City Police department is called for leaving crime scene far more than any other type. Note that the above command took about 3 seconds to execute. In an upcoming section, we'll cache the data into memory for up to 100x speed increases.
# **Q-4) What is the most dangerous month in Saint Louis city?**
display(crimeDataSDF.select('CodedMonth').groupBy('CodedMonth').count().orderBy('Count',ascending=False))
display(sqlContext.sql("select CodedMonth, count(*) as count from crimesql group by CodedMonth ORDER BY count DESC"))
# Seems like August 2020 is the most dangerous month in Saint Louis city and June comes in second.
# ###  ** Doing Date/Time Analysis**
# **Q-4) How many service calls were logged on July 4th?**
# Notice that the date or time columns (DateOccur) is currently being interpreted as strings, rather than date or time objects:
crimeDataSDF.printSchema()
# Let's use the unix_timestamp() function to convert the string into a timestamp:
#
# https://people.apache.org/~pwendell/spark-nightly/spark-master-docs/latest/api/python/pyspark.sql.html?highlight=spark#pyspark.sql.functions.from_unixtime
from pyspark.sql.functions import *
# +
# Note that PySpark uses the Java Simple Date Format patterns
from_pattern1 = 'MM/dd/yyyy'
to_pattern1 = 'MM/dd/yyyy'
from_pattern2 = 'MM/dd/yyyy HH:mm'
to_pattern2 = 'MM/dd/yyyy HH:mm'
crimeDataSTsDF = crimeDataSDF \
.withColumn('DateOccurTS', unix_timestamp(crimeDataSDF['DateOccur'], from_pattern2).cast("timestamp")) \
.drop('DateOccur')
# -
crimeDataSTsDF.printSchema()
crimeDataSTsDF.createOrReplaceTempView("crimesql_ts")
# Notice that the formatting of the timestamps is now different:
# Note that July 4th, is the 185th day of the year in 2020.
#
# Filter the DF down to just 2020 and days of year equal 185:
crimeDataSTsDF.filter(year('DateOccurTs') == '2020').filter(dayofyear('DateOccurTs') == 185).groupBy(dayofyear('DateOccurTs')).count().orderBy(dayofyear('DateOccurTs')).show()
display(sqlContext.sql("select count(*) as count FROM crimesql_ts WHERE cast(DateOccurTs as date)='2020-07-04'"))
They were 133 calls made on July 4, 2020.
# **Q-5) How many service calls were logged in the first week of January 2020?**
# Note that we can narrow down to the 2020 year and look at the first 7 days.
crimeDataSTsDF.filter(year('DateOccurTs') == '2020').filter(dayofyear('DateOccurTs') >= 1).filter(dayofyear('DateOccurTs') <= 7).groupBy(dayofyear('DateOccurTs')).count().orderBy(dayofyear('DateOccurTs')).show()
display(sqlContext.sql("SELECT cast(DateOccurTs as date) as Date, count(*) as count FROM crimesql_ts Where cast(DateOccurTs as date) between '2020-01-01' and '2020-01-07' Group BY cast(DateOccurTs as date) ORDER BY Date "))
# Visualize the results in a bar graph:
display(crimeDataSTsDF.filter(year('DateOccurTs') == '2020').filter(dayofyear('DateOccurTs') >= 1).filter(dayofyear('DateOccurTs') <= 7).groupBy(dayofyear('DateOccurTs')).count().orderBy(dayofyear('DateOccurTs')))
# ###  ** Memory, Caching and write to Parquet**
# The DataFrame is currently comprised of 2 partitions:
crimeDataSTsDF.rdd.getNumPartitions()
# the repartition to 3 so that the data is divided evenly among 3 slots on Databrick Community Edition.
crimeDataSTsDF.repartition(3).createOrReplaceTempView("crimeDataVIEW");
spark.catalog.cacheTable("crimeDataVIEW")
# Call .count() to materialize the cache
spark.table("crimeDataVIEW").count()
crimeDataDF = spark.table("crimeDataVIEW")
# Once the data is cached, the full table scan from Amazon S3 took 1/10 of a second verus 2 seconds as before.
crimeDataDF.count()
spark.catalog.isCached("crimeDataVIEW")
# The 3 partitions are now cached in memory to match the Databrick 3 slots.
# 
# We can check the Spark UI to see the 3 partitions in memory:
# Now that our data has the correct date/time types for each column and it is correctly partitioned, let's write it down as a parquet file for future loading:
# %fs ls /mnt/input/input
crimeDataDF.write.format('parquet').save('dbfs:/mnt/input/input/data')
# Now the directory should contain 3 .gz compressed Parquet files (one for each partition):
# %fs ls dbfs:/mnt/input/input/data
# Here's how you can easily read the parquet file from S3 in the future:
tempDF = spark.read.parquet('dbfs:/mnt/input/input/data/')
display(tempDF.limit(2))
# The possibilities are endless with these datasets.
This notebook was inspired by Sameer Farooqui at Databricks.
| stl-crime-data-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:gsoc_ensembl]
# language: python
# name: conda-env-gsoc_ensembl-py
# ---
# +
####################################################################################################
# Copyright 2019 <NAME> and EMBL-European Bioinformatics Institute
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
####################################################################################################
# -
# # Approach :
# 1. Filter 'all_transcript_data.csv' having gbkey as ncRNA. Save the RefSeq Gene IDs and RefSeq Transcript ID columns.
# 2. Filter 'all_gene_data.csv' whose gene IDs lie in the 'RefSeq Gene ID' collected in point 1 above.
# 3. Filter 'all_exon_data.csv' having gbkey as ncRNA.
# +
import pandas as pd
df = pd.read_csv('/all_gene_data/all_transcript_data.csv', index_col=0)
# -
df.head(5)
df.drop('SNO', axis=1,inplace=True)
len(df[df.columns[1]])
# df[df.columns[5]]
from tqdm import tqdm
# ## Below code for getting all transcript data having gbkey as ncRNA
# %%time
refseq_gene_ids = []
ncrna_transcript_index_to_drop = []
for i in tqdm(range(len(df[df.columns[1]]))):
if df[df.columns[5]][i] != 'ncRNA':
ncrna_transcript_index_to_drop.append(i)
elif df[df.columns[5]][i] == 'ncRNA':
refseq_gene_ids.append(df[df.columns[1]][i])
df.drop(ncrna_transcript_index_to_drop, axis=0,inplace=True)
s_no = []
for i in range(len(df[df.columns[1]])):
s_no.append(i+1)
df['SNO'] = s_no
df = df[['SNO','Index', 'RefSeq Gene ID','RefSeq Transcript ID', 'Dbxref', 'Parent', 'gbkey', 'Type', 'Seq_id', 'Strand','Start', 'End','No. of Exons','Transcript Length']]
df.reset_index(drop=True, inplace=True)
df.head(5)
df.to_csv('/all_gbkey=ncRNA_data/all_ncrna_transcript_data.csv')
# # gene data ncrna
df = pd.read_csv('/refseq_numeric/all_gene_data/all_gene_data.csv', index_col=0)
df.drop('SNO', axis=1,inplace=True)
# +
# df.head(5)
# -
# %%time
s = set(refseq_gene_ids)
drop_values = []
for i in tqdm(range(len(df[df.columns[1]]))):
if df[df.columns[1]][i] in s:
continue
else:
drop_values.append(i)
df.drop(drop_values, axis=0,inplace=True)
# +
s_no = []
for i in range(len(df[df.columns[1]])):
s_no.append(i+1)
df['SNO'] = s_no
# +
# df.head(2)
# -
df = df[['SNO','Index', 'RefSeq Gene ID', 'Dbxref','Name', 'gbkey', 'Biotype', 'Type', 'Seq_id', 'Strand','Start', 'End', 'No. of Transcripts']]
df.reset_index(drop=True, inplace=True)
# +
# df.head(5)
# +
# len(df[df.columns[1]])
# -
df.to_csv('/all_gbkey=ncRNA_data/all_ncrna_gene_data.csv')
# ## Exon data ncrna below
import pandas as pd
df = pd.read_csv('/all_gene_data/all_exon_data.csv', index_col=0)
df.drop('SNO', axis=1,inplace=True)
# +
# df.head(5)
# +
ncrna_exon_index_to_drop = []
for i in tqdm(range(len(df[df.columns[1]]))):
if df[df.columns[6]][i] != 'ncRNA':
ncrna_exon_index_to_drop.append(i)
# -
df.drop(ncrna_exon_index_to_drop, axis=0,inplace=True)
# +
# len(df[df.columns[1]])
# -
s_no = []
for i in range(len(df[df.columns[1]])):
s_no.append(i+1)
df['SNO'] = s_no
df = df[['SNO','Index', 'RefSeq Gene ID','RefSeq Transcript ID','RefSeq Exon ID','Dbxref', 'Parent', 'gbkey', 'Type', 'Seq_id', 'Strand','Start', 'End','Exon Length']]
df.reset_index(drop=True, inplace=True)
df.head(5)
df.to_csv('/all_gbkey=ncRNA_data/all_ncrna_exon_data.csv')
# # Once you load the csv files, do not forget to reset the index values ! [ df.reset_index(drop=True, inplace=True) ]. Default index values are not sequential (Since they got dropped)
| RefSeq-analysis/data_acquisition/refseq_gff_data/all_gene&lncRNA_data/python_scripts/get_all_ncrna_data_from_refseq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook is a copy of David's notebook of the same name, but using the Metapack datapackages.
# + deletable=true editable=true
import pandas as pd
# For development and debugging
# #%load_ext autoreload
# #%autoreload 2
# + deletable=true editable=true
import metatab
p = metatab.open_package('/Volumes/Storage/proj/virt-proj/data-projects/healthy-food-access/packages/fns.usda.gov-f2s_census/fns.usda.gov-f2s_census-2015-1')
p
# + deletable=true editable=true
r = p.resource('census')
r
# + [markdown] deletable=true editable=true
# ### Below is the data as of the latest F2S Census (6/27/16)
# #### Original data can be found at https://farmtoschoolcensus.fns.usda.gov
# I filtered the data below for schools in CA, in San Diego County, and that have either already been participating in F2S in some capacity or have just started in the 2014-2015 school year.
# + deletable=true editable=true
# Read in the excel file and display
census_df = r.dataframe()
# + deletable=true editable=true
census_df.loc[:4].T
# + deletable=true editable=true
# Filter the dataframe, fill na values, and display
census_df_CA = census_df[census_df['state'] == 'CA']
census_df_SD = census_df_CA[census_df_CA['coname'] == 'SAN DIEGO COUNTY']
census_df_SD = census_df_SD[census_df_SD['f2s'] <= 2]
census_df_SD.fillna(0, inplace=True)
census_df_SD.head()
# + deletable=true editable=true
# Perform calculations and display
print('Number of school districts currently participating in F2S or started in 2014-2015 school year:',
len(list(census_df_SD['sfaname'].unique())))
print('Number of schools in those school districts:',
census_df_SD['schoolnum'].astype(int).sum())
print('Number of school gardens in those schools/districts:',
census_df_SD['gardennum'].astype(int).sum())
# -
| users/eric/F2S Census Data Wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="oDVJp_KOV2YL"
# # Data
# + [markdown] id="s7TgOjQLxJkj"
# https://scikit-learn.org/stable/modules/classes.html#module-sklearn.datasets
#
# + id="sPs2xux8--o5" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="d136490f-c925-47ae-9d34-620129945428"
# Mount the drive to load datasets
from google.colab import drive
drive.mount('/drive')
# + id="bhmhsIkxFSVf"
# Import libraries
import math
import numpy as np
import pandas as pd
import seaborn as sns
from collections import Counter
import random
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from sklearn.mixture import GaussianMixture
from sklearn.neighbors import LocalOutlierFactor
from sklearn.datasets import make_moons, make_blobs
from sklearn.covariance import EllipticEnvelope
from sklearn.metrics import classification_report,accuracy_score, f1_score, confusion_matrix, roc_auc_score, average_precision_score
from sklearn.svm import OneClassSVM
from sklearn.ensemble import IsolationForest
from sklearn.cluster import DBSCAN
from sklearn.preprocessing import MinMaxScaler
# + [markdown] id="twsDBvxDtWqB"
# ### Synthetic data creation
# + id="vWRyFz675EFB"
# All the below steps create synthetic dataset for testing different anamoly detection models
syntetic_dataset = {}
syntetic_dataset['num'] = [[-1.1], [0.2], [101.1], [0.3], [1], [0.5]]
n_samples = 300
outliers_fraction = 0.15
n_outliers = int(outliers_fraction * n_samples)
n_inliers = n_samples - n_outliers
rng = np.random.RandomState(42)
blobs_params = dict(random_state=0, n_samples=n_inliers, n_features=2)
syntetic_dataset['norm'] = make_blobs(centers=[[0, 0], [0, 0]],
cluster_std=0.5,
**blobs_params)[0]
syntetic_dataset['2cluster'] = make_blobs(centers=[[2, 2], [-2, -2]],
cluster_std=[0.5, 0.5],
**blobs_params)[0]
syntetic_dataset['2moon'] = 4. * (make_moons(n_samples=n_samples,
noise=.05,
random_state=0)[0]
- np.array([0.5, 0.25]))
for dataset in syntetic_dataset.keys():
# Add outliers
if dataset == 'num':
continue
syntetic_dataset[dataset] = np.concatenate([syntetic_dataset[dataset],
rng.uniform(low=-6, high=6,
size=(n_outliers, 2))],
axis=0)
# + [markdown] id="ClnZ6m6AZC3j"
# #### Plotting the Synthetic Data
# + id="6_3LcNIU9kP_" colab={"base_uri": "https://localhost:8080/", "height": 324} outputId="bfec102a-1fdc-4733-bd9f-cc8aaf81f4b6"
plt.figure(figsize=(12, 4))
plt.subplots_adjust(left=.02,
right=.98,
bottom=.001,
top=.96,
wspace=.05,
hspace=.01)
plt.subplot(1, 3,1)
plt.scatter(syntetic_dataset['norm'][:,0],syntetic_dataset['norm'][:,1])
plt.subplot(1, 3,2)
plt.scatter(syntetic_dataset['2cluster'][:,0],syntetic_dataset['2cluster'][:,1])
plt.subplot(1, 3,3)
plt.scatter(syntetic_dataset['2moon'][:,0],syntetic_dataset['2moon'][:,1])
plt.show()
# + [markdown] id="gd7G2F8kB7TG"
# ## KDD Cup 1999 Data
# This is the data set used for The Third International Knowledge Discovery and Data Mining Tools Competition. The competition task was to build a network intrusion detector, a predictive model capable of distinguishing between ''bad'' connections, called intrusions or attacks, and ''good'' normal connections. This database contains a standard set of data to be audited, which includes a wide variety of intrusions simulated in a military network environment.
#
# The dataset is about 708MB and contains about 4.9M connections. For each connection, the data set contains information like the number of bytes sent, login attempts, TCP errors, and so on.
# + id="0DjoRObDB6QH" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="88fb2740-ca23-4a40-d484-68588c29fa65"
from sklearn.datasets import fetch_kddcup99
dataset = fetch_kddcup99(shuffle=True,percent10=True) # all the features or subset='http' for http features
kdd99_x = dataset.data
kdd99_y = dataset.target
print('data shape:',kdd99_x.shape)
print('target shape:',kdd99_y.shape)
# + [markdown] id="Udx1vR1GRmvs"
# ## Credit Card Fraud Detection
# The datasets contains transactions made by credit cards in September 2013 by european cardholders. This dataset presents transactions that occurred in two days, where we have 492 frauds out of 284,807 transactions. The dataset is highly unbalanced, the positive class (frauds) account for 0.172% of all transactions.
#
# It contains only numerical input variables which are the result of a PCA transformation.
#
# Features V1, V2, ... V28 are the principal components obtained with PCA, the only features which have not been transformed with PCA are 'Time' and 'Amount'.
# + id="EsEgZNuvRouR" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="c2a9a835-fb15-4c26-cf27-c6c92545270c"
# Load the credit card fraud dataset
creditcard_df = pd.read_csv('/drive/My Drive/anomaly_data/creditcard.csv')
print(creditcard_df.columns)
creditcard_df.fillna(0, inplace=True)
# + id="idL5wL9uAjmz" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="b90efbef-e581-4607-d93e-1f59129b3282"
# Distribution of fraud (anamoly) in the dataset
creditcard_Fraud = creditcard_df[creditcard_df['Class']==1]
creditcard_Normal = creditcard_df[creditcard_df['Class']==0]
outlier_fraction = len(creditcard_Fraud)/float(len(creditcard_Normal))
count_classes = pd.value_counts(creditcard_df['Class'], sort = True)
count_classes.plot(kind = 'bar', rot=0)
plt.title("Transaction Class Distribution")
plt.xticks(range(2))
plt.xlabel("Class")
plt.ylabel("Frequency");
# + id="hK_r8luKAx7T" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="8fd50f50-c222-4ce2-8619-3b7fb3d0e38f"
creditcard_df.Amount.describe()
# + [markdown] id="b_HojonkZVkU"
# #### EDA - Fraud vs. Normal transactions
# + id="jhNmYsvLIgL-" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b5920deb-5331-4624-9cba-ba7a29f0ffd0"
features=['V17','V14', 'V11', 'V4', 'V15', 'V13','Amount']
nplots=np.size(features)
plt.figure(figsize=(15,4*nplots))
gs = gridspec.GridSpec(nplots,1)
for i, feat in enumerate(features):
ax = plt.subplot(gs[i])
sns.distplot(creditcard_df[feat][creditcard_df.Class==1], bins=30)
sns.distplot(creditcard_df[feat][creditcard_df.Class==0], bins=30)
ax.legend(['fraudulent', 'non-fraudulent'],loc='best')
ax.set_xlabel('')
ax.set_title('Distribution of feature: ' + feat)
# + id="WSfdDWCaPOFd" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="3ca945b3-bd13-4eae-a8c3-93f8c1e2d42c"
# Filter the columns to remove data we do not want
columns = [c for c in creditcard_df.columns if c not in ["Class"]]
# Store the variable we are predicting
target = "Class"
# Define a random state
state = np.random.RandomState(42)
creditcard_X = creditcard_df[columns]
creditcard_Y = creditcard_df[target]
creditcard_X_outliers = state.uniform(low=0, high=1, size=(creditcard_X.shape[0], creditcard_X.shape[1]))
# Print the shapes of X & Y
print(creditcard_X.shape)
print(creditcard_Y.shape)
np.unique(np.array(creditcard_Y), return_counts=True)
# Scaling the data
scaler = MinMaxScaler()
creditcard_X_scaled = scaler.fit_transform(creditcard_X)
# + [markdown] id="gOCRNWasA7TV"
# ## MNIST
# + id="dXvOIwIqjtZq"
# Load the MNIST dataset
mnist_df = pd.read_csv('/drive/My Drive/anomaly_data/mnist_train.csv', header=None)
anomalous_classes =[7]
contamination= 0.0005
mnist_df.describe()
mnist_data = mnist_df.values[:, 1:]
mnist_label = mnist_df.values[:, :1]
mnist_label = list(map(int, mnist_label))
classes, counts = np.unique(mnist_label, return_counts=True)
mnist_x = []
mnist_y = []
dropout_prob = contamination * (counts.sum() - counts[anomalous_classes].sum())
dropout_prob /= counts[anomalous_classes].sum() * (1 - contamination)
for i, (image, label) in enumerate(zip(mnist_data, mnist_label)):
if label in anomalous_classes and random.random() >= dropout_prob:
continue
mnist_x.append(image),
mnist_y.append(int(label in anomalous_classes))
mnist_x = np.array(mnist_x)
mnist_y = np.array(mnist_y)
scaler = MinMaxScaler()
mnist_x_scaled = scaler.fit_transform(mnist_x)
# + id="KL3wsI3q0PkD" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6b6686a9-3c52-4524-c3da-8cb6003433a0"
np.unique(np.array(mnist_y), return_counts=True)
# + [markdown] id="1JenpU2SMJk8"
# #Some functions
# + [markdown] id="DMRr8etmMUHp"
# ###Performance evaluation
# + id="HQKoQIAlOESt"
def find_best_threshold(y_true, y_pred_prob):
best_f1_score = 0
best_threshold = 0
for threshold_val in np.arange(0.1, 0.95, 0.05):
y_pred = y_pred_prob >= threshold_val
f1_score = f1_score(y_true, y_pred)
if best_f1_score < f1_score:
best_threshold = threshold_val
best_f1_score = f1_score
return float(best_threshold)
# + id="Nm-DKUg8MOMo"
def eval(lab, pred):
c = Counter()
c.update(pred.round())
print('prediction frequency: ')
print(c)
print(f"Accuracy Score: {accuracy_score(lab, pred.round())}")
print(f"F1 Score: {f1_score(lab, pred.round())}")
print(f"Average Precision Score: {average_precision_score(lab, pred.round())}")
print(f"AUROC Score: {roc_auc_score(lab, pred)}")
print(f"Confusion Matrix:\n {confusion_matrix(lab, pred.round())}")
print("Classification Report :")
print(classification_report(lab,pred.round()))
def eval_prob(lab, pred, threshold=0.5):
c = Counter()
c.update(pred>threshold)
print('prediction frequency: ')
print(c)
print(f"Accuracy Score: {accuracy_score(lab, pred>threshold)}")
print(f"F1 Score: {f1_score(lab, pred>threshold)}")
print(f"Average Precision Score: {average_precision_score(lab, pred>threshold)}")
print(f"AUROC Score: {roc_auc_score(lab, pred)}")
print(f"Confusion Matrix:\n {confusion_matrix(lab, pred>threshold)}")
print("Classification Report :")
print(classification_report(lab,pred>threshold))
# + [markdown] id="UwdnQagJGrJ_"
# ##### Visualization
# + id="lBRVhZcjGvzE"
def visualization(X,p,threshold=0.5):
plt.figure(figsize=(10,10))
plt.scatter(X[:,0],X[:,1],c=p,cmap='viridis',marker='x')
outliers = np.nonzero(p<threshold)[0]
plt.scatter(X[outliers,0],X[outliers,1],marker="o",facecolor="none",edgecolor="r",s=70);
plt.colorbar();
# + [markdown] id="pnqLUsxdV2qv"
# # Isolation forest model - Unsupervised model (No labels required)
#
#
# Isolation Forest is based on the Decision Tree algorithm. It isolates the outliers by randomly selecting a feature from the given set of features and then randomly selecting a split value between the max and min values of that feature. This random partitioning of features will produce shorter paths in trees for the anomalous data points, thus distinguishing them from the rest of the data.
# + id="Lul6rynxVTGm" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="447b6f2e-f961-46da-892c-41a89cb7ca98"
# Creating a small sample data to understand the isolation ofrest model
X = np.array([[-1, -1], [-2, -1], [-3, -2], [0, 0], [-20, 50], [3, 5]])
clf = IsolationForest(n_estimators=2, warm_start=True)
clf.fit(X) # fit 2 trees
print(clf.predict(X))
print(clf.decision_function(X))
clf.set_params(n_estimators=20) # adding more trees
clf.fit(X) # fit the added trees
print(clf.predict(X))
print(clf.decision_function(X))
# + [markdown] id="l6jAOERAZ5b0"
# Above example clearly shows that isolation forest is able to detect the outlier points from the dataset. In the output predictions all the outliers are marked as -1. Also, decision_function() helps to find out the values of anamoly scores
# + [markdown] id="ApV4puK6R2X7"
# ##Syntetic dataset
# + id="tDI4gXzzYIPT" colab={"base_uri": "https://localhost:8080/", "height": 596} outputId="4ab7e5f1-1e23-48d2-c77f-486588e576cc"
syn_x = syntetic_dataset['2moon']
np.random.seed(1)
clf = IsolationForest(contamination=outliers_fraction)
preds = clf.fit_predict(syn_x)
des = clf.decision_function(syn_x)
visualization(syn_x,preds)
# visualization(syn_x,des>.01)
# + [markdown] id="kM1SGVw1Q91m"
# ##Credit Card Fraud Dataset
# + id="9fXNPEyxW_zB" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="77ce387d-1c71-4325-f1b5-253fc66225cf"
X = creditcard_X
Y = creditcard_Y
# + [markdown] id="eUs6dNFyLaa6"
# ###Exercise 1
# Train a model using isolation forest and test different hyper-parameter to get the best result. you can find the list of all hyper parameters here: [*IsolationForest*](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.IsolationForest.html#sklearn.ensemble.IsolationForest)
#
# Challenge : Report your best F1-score on slack! Let’s see who got the best score!!
#
# y_pred: output prediction
#
# + id="oc1wCEfsLMN4"
print(np.unique(np.array(y_pred), return_counts=True))
y_pred[y_pred == 1] = 0
y_pred[y_pred == -1] = 1
# + id="Cg050S5tXbtb" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="8bbfca0b-a2a6-4072-f053-314f75bd86ff"
eval(Y,y_pred)
# + [markdown] id="kMvJRwQ-V284"
# #One-SVM
#
# When modeling one class, the algorithm captures the density of the majority class and classifies examples on the extremes of the density function as outliers. This modification of SVM is referred to as One-Class SVM.
# + [markdown] id="mj556NbDSc37"
# ## Synthetic datasets
# + id="hByDEGiXSfgc" colab={"base_uri": "https://localhost:8080/", "height": 596} outputId="dbff4ca9-f784-487b-e3c0-9cde0cac237e"
syn_x = syntetic_dataset['2cluster']
m = OneClassSVM(nu=outliers_fraction, kernel="rbf", gamma=0.1)
preds = m.fit_predict(syn_x)
score = m.score_samples(syn_x)
# visualization(syn_x,score,8.6)
visualization(syn_x,preds)
# + [markdown] id="Lbt8ykriWxbj"
# ## Credit Card Fraud Dataset
# + id="XsVz65SsWuH8"
X=creditcard_X_scaled[:20000,:]
Y=creditcard_Y[:20000]
# + [markdown] id="SaQpR9CYN4uS"
#
#
# ###Exercise 2
# Train a model using OneClassSVM and test different hyper-parameter to get the best result. you can find the list of all hyper parameters here: [OneClassSVM](https://scikit-learn.org/stable/modules/generated/sklearn.svm.OneClassSVM.html#sklearn.svm.OneClassSVM)
#
# Challenge : Report your best F1-score on slack! Let’s see who got the best score!!
#
# y_pred: output prediction
#
# + id="b9EvLmluNkde"
print(np.unique(np.array(y_pred), return_counts=True))
y_pred[y_pred == 1] = 0
y_pred[y_pred == -1] = 1
# + id="0SxOIPzhThge"
eval(Y,y_pred)
# + [markdown] id="3edeHGA4V3Md"
# #Clustering
# + [markdown] id="COAPozduV3Xz"
# ##K-means
# + id="BrRAXQeEiAie" colab={"base_uri": "https://localhost:8080/", "height": 596} outputId="b31d9895-1a8f-48b5-9a9b-08f9e39ffbfc"
syn_x = syntetic_dataset['2cluster']
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=3)
kmeans.fit(syn_x)
centers = kmeans.cluster_centers_
preds = kmeans.predict(syn_x)
preds
visualization(syn_x,preds,0)
# + id="NQYqsDMSnatf" colab={"base_uri": "https://localhost:8080/", "height": 647} outputId="1a0911ab-d9d5-4e10-ae09-5a175dd2144d"
# All the points that are farther than the formed clusters are treated as anamolies
np.unique(preds)
clusters = kmeans.cluster_centers_[preds]
print(kmeans.cluster_centers_)
dist = np.average([np.sum(c1 - c2)**2 for c1, c2 in zip(syn_x, clusters)])
dist_all = np.array([float(np.sum(c1 - c2)**2<dist) for c1, c2 in zip(syn_x, clusters)])
visualization(syn_x,dist_all)
# + id="3lG6l7HexfZ_" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="f7e33719-b0fa-4029-c6e7-d7f45fe464db"
print(dist)
print(np.unique(np.array(dist_all), return_counts=True))
# + [markdown] id="JZ8Vr6JiWY3w"
# ##DBSCAN
# + [markdown] id="bSOZuM9ywQ4g"
# DBSCAN will output an array of -1’s and 0’s, where -1 indicates an outlier. Below, I visualize outputted outliers in red by plotting two variables.
#
# More info: https://medium.com/@elutins/dbscan-what-is-it-when-to-use-it-how-to-use-it-8bd506293818
# + [markdown] id="BhWqMq5KDpdt"
# ##synthetic dataset
# + id="0pX1YaMiWcEw" colab={"base_uri": "https://localhost:8080/", "height": 596} outputId="b104b2ee-3712-4d4c-ea53-be6aa7b842f4"
m = DBSCAN(min_samples = 3, eps = 1)
clusters = m.fit_predict(syntetic_dataset['2cluster'])
list(clusters).count(-1)
visualization(syntetic_dataset['2cluster'],clusters,0)
# + id="SMP_8QgiYxsS" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="660a7b57-39ce-49a0-b070-6235d01594d8"
clusters
# + [markdown] id="KEfjnUFl4jGo"
# ##Credit Card Fraud Dataset
# + id="7VgRqUe5wInl"
X=creditcard_X_scaled[:20000,:]
Y=creditcard_Y[:20000]
# + [markdown] id="GirJQi4hOuMz"
#
# ###Exercise 3
# Train a model using DBSCAN and test different hyper-parameter to get the best result. you can find the list of all hyper parameters here: [DBSCAN](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.DBSCAN.html#sklearn.cluster.DBSCAN)
#
# Challenge : Report your best F1-score on slack! Let’s see who got the best score!!
#
# y_pred: output prediction
#
# + id="l4et_tdKOtTD"
print(np.unique(np.array(clusters), return_counts=True))
clusters[clusters != -1] = 0
clusters[clusters == -1] = 1
# + id="8cnOgMnBwUwG" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="4b3f2934-a39f-43fe-83fa-7a97226cf39e"
eval(Y,clusters)
# + [markdown] id="R3II81-HQKNx"
# #Shuffling
# + [markdown] id="Wuw8uNLxP1LF"
# ##Credit Card Fraud Dataset
# + id="lnzNDa6AQObk" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dcb1cf66-80ee-4871-cccf-76ed0cd7eb1d"
X = creditcard_X_scaled
Y = creditcard_Y
X_pure=[X[i] for i in range(len(X)) if Y[i]==0]
Y_pure=[Y[i] for i in range(len(Y)) if Y[i]==0]
X1 = np.array(X_pure)
X2 = np.array(X_pure)
X2 = X2.T
for i in range(len(X2)):
np.random.shuffle(X2[i])
X2 = X2.T
print(X1.shape, X2.shape)
# + id="ea7AKK5wQRxP" colab={"base_uri": "https://localhost:8080/", "height": 717} outputId="fa4dd2f0-3562-487f-ac79-71e39925b75a"
from sklearn.neural_network import MLPClassifier
clf = MLPClassifier(hidden_layer_sizes=(64,),max_iter=20)
X_all = np.concatenate((X1, X2))
Y_all = Y_pure + [1]*len(X2)
clf.fit(X_all, Y_all)
pred_all = clf.predict(X_all)
eval(Y_all, pred_all)
pred_val = clf.predict(X)
eval(Y, pred_val)
# + id="8Hpisc-iLPc3" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="9a8fb002-886f-44a3-f2c1-f561e35c4d59"
pred_val = clf.predict_proba(X)[:,1]
eval_prob(Y, pred_val,0.95)
# + [markdown] id="G_2SujJPPZzj"
# ###Exercise 4
# Shuffle each coloumn individually and train a model based on it and then combine then(ensemble) to do the prediction!
# + [markdown] id="DtVfUNPRZvES"
# #Exercise 5
# Test these models on MNIST, KDD cup dataset and report you best score! :-)
| anomaly and novelty detection/Session 2/Anomaly_session2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploratory Data Analysis in Action - Loading and exploring the data set
# **Import statements**
#
#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# **Global settings**
pd.options.display.max_rows = 999
pd.options.display.max_columns = 100
plt.rcParams["figure.figsize"] = [15,6]
# ## Inspect the raw data set
#
# It is good practice to inspect the data set before loading into Python. If we are dealing with text-like data we may inspect the data set using our favorite text editor or simply using the shell.
#
# When working with Jupyter notebooks we may use the notebook magic command `!` to execute shell commands form within a code cell.
#
# The data set is found in the `data` folder:
#
# ./data/operations.csv
#
#
# In order to look at the first 20 lines of the text file we use the bash command `head -20 PATH_TO_FILE/FILENAME`.
# +
### your code here ...
# -
# ## Load data set
PATH = "./data/"
df_raw = pd.read_csv(PATH + "operations.csv", low_memory=False)
# ## Review proporties of the pandas object
# **Number of rows and columns**
df_raw.shape
# **Memory usage** (Check out the [blog entry of <NAME>](https://www.dataquest.io/blog/pandas-big-data/) for a deep dive into memory management with pandas)
df_raw.info(verbose=False, memory_usage='deep')
# **Inspect columns and row**
df_raw.head()
# 
#
# Source: Wikipedia [B24](https://de.wikipedia.org/wiki/Consolidated_B-24), [A36](https://de.wikipedia.org/wiki/North_American_A-36)
df_raw.columns
# ### Data types
df_raw.dtypes
# ### Explore variables of interest
# > **Challange:** Explore the numeric variable `Aircraft Lost`
# +
### your code here ...
# -
# > **Challange:** Explore the categorical variable `Target Country`
# +
### your code here ...
| Workshop_Data_Science/Exploratory Data Analysis WW2 - Loading and exploring the data set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: epa1
# language: python
# name: epa1
# ---
import yaml
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# +
# input data from queries executed in DBeaver
# define cohort (label) years
cohort_years = [2007, 2008, 2009, 2010, 2011, 2012, 2013, 2014]
# define number of handlers in each cohort
# (cohort_year, label_year): (2006, 2007), (2007, 2008), (2008, 2009), (2009, 2010),
# (2010, 2011), (2011, 2012), (2012, 2013)
val_with_history = [572, 583, 594, 599, 575, 619, 691, 689]
val_without_history = [769, 732, 755, 1563, 772, 1037, 1619, 2597]
# input number of handlers with a label in each cohort
# (cohort_year, feature_start_year, feature_end_year): (2006, 2002, 2006), (2007, 2003, 2007),
# (2008, 2004, 2008), (2009, 2005, 2009), (2010, 2006, 2010), (2011, 2007, 2011), (2012, 2008, 2012),
# (2013, 2009, 2013)
val_with_history_with_label = [158, 192, 161, 195, 180, 201, 201, 209]
val_without_history_with_label = [74, 94, 110, 85, 64, 71, 45, 60]
# +
# calculate exclusive sets of handlers with no label
val_with_history_no_label = []
for i in range(len(val_with_history)):
with_history_no_label = val_with_history[i] - val_with_history_with_label[i]
val_with_history_no_label.append(with_history_no_label)
val_without_history_no_label = []
for i in range(len(val_without_history)):
without_history_no_label = val_without_history[i] - val_without_history_with_label[i]
val_without_history_no_label.append(without_history_no_label)
# -
# create empty dataframe for cohort data
cohorts = pd.DataFrame(index=cohort_years)
# load data into dataframe
cohorts['val_with_history'] = val_with_history
cohorts['val_without_history'] = val_without_history
cohorts['val_with_history_with_label'] = val_with_history_with_label
cohorts['val_without_history_with_label'] = val_without_history_with_label
cohorts['val_with_history_no_label'] = val_with_history_no_label
cohorts['val_without_history_no_label'] = val_without_history_no_label
# plot cohort data as stacked bar chart
fig, ax = plt.subplots(figsize=(11,8))
ax.set_title('Validation cohort sizes by year (with history)')
ax.set_ylabel('Number of handlers')
ax.set_xlabel('Validation (label) year')
ax.set_ylim(0, 2650)
cohorts.plot(y=['val_with_history_with_label', 'val_with_history_no_label'], kind='bar', stacked=True, ax=ax)
for p in ax.patches:
width, height = p.get_width(), p.get_height()
x, y = p.get_xy()
ax.text(x+width/2,
y+height/2,
'{:.0f}'.format(height),
horizontalalignment='center',
verticalalignment='center',
color='white',
fontweight='bold')
plt.savefig('cohort_wHist.png')
# plot cohort data as stacked bar chart
fig, ax = plt.subplots(figsize=(11,8))
ax.set_title('Validation cohort sizes by year (without history)')
ax.set_ylabel('Number of handlers')
ax.set_xlabel('Validation (label) year')
ax.set_ylim(0, 2650)
cohorts.plot(y=['val_without_history_with_label', 'val_without_history_no_label'], kind='bar', stacked=True, ax=ax)
for p in ax.patches:
width, height = p.get_width(), p.get_height()
x, y = p.get_xy()
ax.text(x+width/2,
y+height/2,
'{:.0f}'.format(height),
horizontalalignment='center',
verticalalignment='center',
color='white',
fontweight='bold')
plt.savefig('cohort_woHist.png')
| cohort_sizes_20201116.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook 1: Why is Machine Learning difficult?
# ## Overview
#
# In this notebook, we will get our hands dirty trying to gain intuition about why machine learning is difficult.
#
# Our task is going to be a simple one, fitting data with polynomials of different order. Formally, this goes under the name of polynomial regression. Here we will do a series of exercises that are intended to give the reader intuition about the major challenges that any machine learning algorithm faces.
#
# ## Learning Goal
#
# We will explore how our ability to predict depends on the number of data points we have, the "noise" in the data, and our knowledge about relevant features. The goal is to build intuition about why prediction is difficult and discuss general strategies for overcoming these difficulties.
#
#
# ## The Prediction Problem
#
# Consider a probabilistic process that gives rise to labeled data $(x,y)$. The data is generated by drawing samples from the equation
# $$
# y_i= f(x_i) + \eta_i,
# $$
# where $f(x_i)$ is some fixed, but (possibly unknown) function, and $\eta_i$ is a Gaussian, uncorrelate noise variable such that
# $$
# \langle \eta_i \rangle=0 \\
# \langle \eta_i \eta_j \rangle = \delta_{ij} \sigma
# $$
# We will refer to the $f(x_i)$ as the **true features** used to generate the data.
#
# To make prediction, we will consider a family of functions $g_\alpha(x;\theta_\alpha)$ that depend on some parameters $\theta_\alpha$. These functions respresent the **model class** that we are using to try to model the data and make predictions. The $g_\alpha(x;\theta_\alpha)$ encode the class of **features** we are using to represent the data.
#
# To learn the parameters $\boldsymbol{\theta}$, we will train our models on a **training data set** and then test the effectiveness of the model on a <i>different</i> dataset, the **test data set**. The reason we must divide our data into a training and test dataset is that the point of machine learning is to make accurate predictions about new data we have not seen. As we will see below, models that give the best fit to the training data do not necessarily make the best predictions on the test data. This will be a running theme that we will encounter repeatedly in machine learning.
#
#
# For the remainder of the notebook, we will focus on polynomial regression. Our task is to model the data with polynomials and make predictions about the new data that we have not seen.
# We will consider two qualitatively distinct situations:
# <ul>
# <li> In the first case, the process that generates the underlying data is in the model class we are using to make predictions. For polynomial regression, this means that the functions $f(x_i)$ are themselves polynomials.
# <li>In the second case, our data lies outside our model class. In the case of polynomial regression, this could correspond to the case where the $f(x_i)$ is a 10-th order polynomial but $g_\alpha(x;\theta_\alpha)$ are polynomials of order 1 or 3.
# </ul>
#
# In the exercises and discussion we consider 3 model classes:
# <ul>
# <li> the case where the $g_\alpha(x;\theta_\alpha)$ are all polynomials up to order 1 (linear models),
# <li> the case where the $g_\alpha(x;\theta_\alpha)$ are all polynomials up to order 3,
# <li> the case where the $g_\alpha(x;\theta_\alpha)$ are all polynomials up to order 10.
# </ul>
#
# To measure our ability to predict, we will learn our parameters by fitting our training dataset and then making predictions on our test data set. One common measure of predictive performance of our algorithm is to compare the predictions,$\{y_j^\mathrm{pred}\}$, to the true values $\{y_j\}$. A commonly employed measure for this is the sum of the mean square-error (MSE) on the test set:
# $$
# MSE= \frac{1}{N_\mathrm{test}}\sum_{j=1}^{N_\mathrm{test}} (y_j^\mathrm{pred}-y_j)^2
# $$
# We will return to this in later notebooks. For now, we will try to get a qualitative picture by examining plots on test and training data.
#
# ## Fitting vs. predicting when the data is in the model class
#
#
# We start by considering the case:
# $$
# f(x)=2x.
# $$
# Then the data is clearly generated by a model that is contained within all three model classes we are using to make predictions (linear models, third order polynomials, and tenth order polynomials).
#
#
# Run the code for the following cases:
# <ul>
# <li> For $f(x)=2x$, $N_{\mathrm{train}}=10$ and $\sigma=0$ (noiseless case), train the three classes of models (linear, third-order polynomial, and tenth order polynomial) for a training set when $x_i \in [0,1]$. Make graphs comparing fits for different order of polynomials. Which model fits the data the best?
# <li> Do you think that the data that has the least error on the training set will also make the best predictions? Why or why not? Can you try to discuss and formalize your intuition? What can go right and what can go wrong?
# <li>Check your answer by seeing how well your fits predict newly generated test data (including on data outside the range you fit on, for example $x \in [0,1.2]$) using the code below. How well do you do on points in the range of $x$ where you trained the model? How about points outside the original training data set?
# <li>Repeat the exercises above for $f(x)=2x$, $N_{\mathrm{train}}=10$, and $\sigma=1$. What changes?
# <li>Repeat the exercises above for $f(x)=2x$, $N_{\mathrm{train}}=100$, and $\sigma=1$. What changes?
# <li> Summarize what you have learned about the relationship between model complexity (number of parameters), goodness of fit on training data, and the ability to predict well.
# </ul>
#
#
# ## Fitting vs. predicting when the data is not in the model class
# Thus far, we have considered the case where the data is generated using a model contained in the model class. Now consider $f(x)=2x-10x^5+15x^{10}$. *Notice that the for linear and third-order polynomial the true model $f(x)$ is not contained in model class $g_\alpha(x)$* .
#
# <ul>
# <li> Repeat the exercises above fitting and predicting for $f(x)=2x-10x^5+15x^{10}$ for $N_{\mathrm{train}}=10,100$ and $\sigma=0,1$. Record your observations.
# <li> Do better fits lead to better predictions?
# <li> What is the relationship between the true model for generating the data and the model class that has the most predictive power? How is this related to the model complexity? How does this depend on the number of data points $N_{\mathrm{train}}$ and $\sigma$?
# <li> Summarize what you think you learned about the relationship of knowing the true model class and predictive power.
#
# # Training the models:
#This is Python Notebook to walk through polynomial regression examples
#We will use this to think about regression
import numpy as np
# %matplotlib inline
from sklearn import datasets, linear_model
from sklearn.preprocessing import PolynomialFeatures
from sklearn.metrics import mean_squared_error
from matplotlib import pyplot as plt
# ## Training over different true functions, models, and noise
# +
sigma_train_list = [0.001, 1]
fig, ax_list = plt.subplots(ncols=2, nrows=len(sigma_train_list), sharey=True, figsize=(14, 6))
N_train = int(1e1)
x = np.linspace(0.05, 0.95, N_train)
xplot = np.linspace(0.02, 0.98, 200) # grid of points, some are in the training set, some are not
for row, sigma_train in enumerate(sigma_train_list):
# Train on integers
# Draw Gaussian random noise
s = sigma_train * np.random.randn(N_train)
y_data_dict = {"linear": 2 * x + s, "tenth": 2 * x - 10 * x**5 + 15 * x**10 +s}
fit_dict = dict()
poly_degrees = [3, 10]
for fig_count, (model, y) in enumerate(y_data_dict.items()):
ax = ax_list[row][fig_count]
# Linear Regression : create linear regression object
clf = linear_model.LinearRegression()
clf.fit(x[:, np.newaxis], y)
fit_dict[1] = clf
ax.plot(x, y, "x")
# Use fitted linear model to predict the y value:
y_pred = clf.predict(xplot[:, np.newaxis])
linear_plot = ax.plot(xplot, y_pred, label='deg 1')
for deg in poly_degrees:
poly_features = PolynomialFeatures(deg)
X = poly_features.fit_transform(x[:, np.newaxis])
clf = linear_model.LinearRegression()
clf.fit(X, y)
fit_dict[deg] = clf
Xplot = poly_features.fit_transform(xplot[:,np.newaxis])
poly_plot = ax.plot(xplot, clf.predict(Xplot), label='deg {}'.format(deg))
Title="%s, N=%i, $\sigma=%.2f$"%(model, N_test, sigma_train)
ax.set_title(Title+" (train)")
ax.legend()
fig.subplots_adjust(wspace=0.05, hspace=0.4)
# +
# np.linalg.pinv(PolynomialFeatures(1).fit_transform(x[:, np.newaxis])) @ y
# +
N_test = 20
max_x = 1.2
x_test = max_x * np.random.random(N_test)
x_plot = np.linspace(0, max_x, 200)
fig, ax_list = plt.subplots(ncols=2, nrows=len(sigma_train_list), sharey=True, figsize=(14, 6))
for row, sigma_test in enumerate(sigma_train_list):
s = sigma_test * np.random.randn(N_test)
y_data_dict = {"linear": 2 * x_test + s, "tenth": 2 * x_test - 10 * x_test**5 + 15 * x_test**10 +s}
for fig_count, (model, y) in enumerate(y_data_dict.items()):
ax = ax_list[row][fig_count]
ax.plot(x_test, y, "x")
for deg, clf in fit_dict.items():
if deg > 1:
X_test = PolynomialFeatures(deg).fit_transform(x_plot[:, np.newaxis])
ax.plot(x_plot, clf.predict(X_test), label="deg {}".format(deg))
else:
ax.plot(x_plot, clf.predict(x_plot[:, np.newaxis]), label="deg {}".format(deg))
Title="%s, N=%i, $\sigma=%.2f$"%(model, N_test, sigma_test)
ax.set_title(Title+" (test)")
ax.legend()
#ax.set_ylim(-50, 200)
fig.subplots_adjust(wspace=0.05, hspace=0.4)
# -
# ## Observations
#
# - Nice tool called PolynomialFeatures to get feature matrix for a given polynomial degree. Model parameters can be obtained from matrix inversion
# - Complex models "overfit" when training size is small. Also do poorly outside fit range. Therefore simple models although far from reality may do better when limited data is present
#
| updated_notebooks/001-ML_is_difficult.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Some example outputs and instructions to the N-body simulation code.
# Hi, this section will show you some of the outputs from my N-body simulation code. At the end of this section, you may even be able to create your own universe. If you want to download the code and run it yourself, make sure you read the README file so you will know how the code work and the requirements for the code to run. If you are just here for fun and want to see some fun simulations of the universe, then buckle up, you are in for a ride.
# ## Unit, unit and unit
from IPython.display import Image
Image(filename='unit joke.jpg')
# Run the cell above to see the image. The image is obtained from https://www.pinterest.com.au/pin/90986854941568312/.
# The most common system of units are the metric or imperial system. However, cosmologists work on very large scales. Therefore, it is difficult to express the mass or distance in the astronmy context with the usual system of units. Thus, astronomers and cosmologists have their own system of units. Here are some units that are commonly used in astronomy:
#
# Solar mass is the mass of our sun.
#
# Parsec is a distance unit. 1 parsec is approximately three light years.
#
# In my simulation, the mass is in terms of 10^5 solar masses and the distance in the unit of kiloparsec.
# ## Warning
# The simulations from the code are very very rough approximation of motion of stars or galaxies in the universe. Want to see more accurate simulation? Check out the videos on this website https://wwwmpa.mpa-garching.mpg.de/galform/virgo/millennium/.
# ## Two body simulation
# Why don't we start from something easy and make sure the code is producing the correct output. The simplest way to do that is to put two particles in the simulation and see how they interact with each other. The only force in my code is the gravitational force. The gravitational force is an attractive force, so we expect the two objects will come closer and closer to each other. Try to run the following code to see whether this is happening.
# %matplotlib inline
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import time
import sys
from IPython.display import HTML
def find_particle(position, x, y, z):
"""
Parameters
----------
position : array_like
This is the position of all particles in the simulation.
x : array_like
The array contains the grid point in the x direction.
y : array_like
The array contains the grid point in the y direction.
z : array_like
The array contains the grid point in the z direction.
Returns
-------
num : List
Number of particles in each grid.
particle_in_cell : List
The index of particles in each grid cell.
"""
particle_in_cell =[]
num = []
#Initialise the output list
limit_x = len(x)-1
limit_y = len(y)-1
limit_z = len(z)-1
#Number of computation requires for each dimension.
counter_x = 0
counter_y = 0
counter_z = 0
#This counts the number of cells that have been computed in each direction.
for i in range(limit_x*limit_y*limit_z):
position_x = position[0,:]
position_y = position[1,:]
position_z = position[2,:]
#The poistion of the particle in each direction.
xx = np.where(position_x < x[counter_x+1], position_x, 0)
yy = np.where(position_y < y[counter_y+1], position_y, 0)
zz = np.where(position_z < z[counter_z+1], position_z, 0)
#Find the particles in the position array which are to the right of the desired grid. For such particle
#replace the position with zero.
xxx = np.where(xx > x[counter_x], xx, 0)
yyy = np.where(yy > y[counter_y], yy, 0)
zzz = np.where(zz > z[counter_z], zz, 0)
#Find the particles in the position array which are to the left of the desired grid. For such particle
#replace the position with zero.
index_x = np.nonzero(xxx)[0]
index_y = np.nonzero(yyy)[0]
index_z = np.nonzero(zzz)[0]
#Find the index of the particle which are nonzero. These particles are located in the desired grid.
#print(index_x, index_y, index_z)
xy = np.intersect1d(index_x, index_y, assume_unique=True)
xyz = np.intersect1d(xy, index_z, assume_unique=True)
#The codes above finds the index of particle in the desired grid. The index in each array is unique.
if (len(xyz != 0)):
num.append(len(xyz))
particle_in_cell.append(xyz)
#Append the particle in the grid and the number of particle in the grid if the number of particles in
#the grid is nonzero.
counter_x += 1
#Move to the grid at the right
if (counter_x == limit_x):
#This means it completes calculate the particles in the grid in a row. Advance to the next row.
counter_x = 0
counter_y += 1
if (counter_y == limit_y):
#This moves to the next layer of the xy-plane.
counter_y = 0
counter_z += 1
return num, particle_in_cell
def apply_boundary(p, Nd, Np):
"""
Parameters
----------
p : array_like
Position of all particle in the array.
Nd : int
Number of dimensions of the simulation.
Np : int
Number of particles in the simulation.
Returns
-------
p : array_like
The position of particles after applying the boundary condition.
"""
# Modify to apply your chosen boundary conditions
position_x = p[0,:]
position_y = p[1,:]
position_z = p[2,:]
#The position of particles in the x, y and z position.
#The following lines will find the particle outside the simulation and move it back to the simulation
#based on the periodic boundary condition.
xx = np.where(position_x > x_bound, (position_x-x_bound)-x_bound, position_x)
xxx = np.where(xx < -x_bound, x_bound - (-x_bound - xx), xx)
yy = np.where(position_y > y_bound, (position_y-y_bound)-y_bound, position_y)
yyy = np.where(yy < -y_bound, y_bound - (-y_bound - yy),yy)
zz = np.where(position_z > z_bound, (position_z-z_bound)-z_bound, position_z)
zzz = np.where(zz < -z_bound, z_bound - (-z_bound - zz), zz)
p = np.concatenate((xxx, yyy, zzz), axis = 0)
p = np.reshape(p, (Nd, Np))
#Reconstruct the array for position of particles.
return p
def center_of_mass(particle_in_cell, num, mass, position):
"""
Parameters
----------
particle_in_cell : List
The list that contain the index of particles in each grid cell.
num : List
The list contains the number of particles in each grid cell.
mass : array_like
The mass of all the particles.
position : array_like
The position of all the partiles.
Returns
-------
result : List
The center of mass position in each grid cell.
total_mass : List
The total mass of all particles in each grid cell.
"""
result = []
total_mass = []
#Initialise the output lists
position_x = position[0,:]
position_y = position[1,:]
position_z = position[2,:]
for i in range(len(particle_in_cell)):
COM_x = 0.0
COM_y = 0.0
COM_z = 0.0
M_total = 0.0
#Initialise the center of mass position and the total mass of particles in the grid
for j in range(num[i]):
COM_x += mass[particle_in_cell[i][j]]*position_x[particle_in_cell[i][j]]
COM_y += mass[particle_in_cell[i][j]]*position_y[particle_in_cell[i][j]]
COM_z += mass[particle_in_cell[i][j]]*position_z[particle_in_cell[i][j]]
M_total += mass[particle_in_cell[i][j]]
#Calculate the center off mass
result.append(np.array([COM_x/M_total, COM_y/M_total, COM_z/M_total]))
total_mass.append(M_total)
return result, total_mass
def position2grid(particle_index, particle_in_cell):
"""
Parameters
----------
particle_index : array_like
Index of all particles in the simulation.
particle_in_cell : List
The index of particle in each grid cell.
Returns
-------
result : List
It matches the index of particle to the index of the grid it is in.
"""
result = []
for i in range(len(particle_index)):
for j in range(len(particle_in_cell)):
size = (np.intersect1d(np.array([i]), particle_in_cell[j]))
#Determine whether the particle is in the grid[j]
if (size.size > 0):#If the particle is in grid[j], the size of the array will be nonzero.
#Since the index of particle is also unique, we are certain that when the size of array is not zero.
#we find the cell of which the particle is in
break
result.append(np.array([i,j]))
return result
def accel_grid(COM, total_mass, mass, index, p, particle, particle_in_cell, num, grid_length, smooth):
"""
Parameters
----------
COM : List
Center of mass of all grid cells.
total_mass : List
Mass of all particles in a single grid cell.
mass : array_like
The mass of each individual particle.
index : int
The index of the grid cell.
p : array_like
Position of all the particle.
particle : int
Index of the particle.
particle_in_cell : List
The list contains the index of particles in each grid cell.
num : List
Number of particles in each grid cell.
grid_length : The length of the gridcell
A reference length. If the distance between particle and center of mass of any grid is below the
reference length. We will calulate the interaction in particle-particle basis (similar to P3M).
Returns
-------
float
The acceleration in the x-direction.
float
The acceleration in the y-direction.
float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
smooth_grid = grid_length #The smoothen scale which is set to length of the grid.
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
for i in range(len(COM)):
r_dash = np.sqrt((COM[i][0]-p[0, particle])**2 + (COM[i][1]-p[1, particle])**2 + (COM[i][2]-p[2,particle])**2)
#The distance between the particle and the center of mass of particles.
if (r_dash <= grid_length):
#If less than the grid size, calculate the force using individual particle.
accel_x += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[0]
accel_y += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[1]
accel_z += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[2]
else:
#Larger than the gridsize, calculate the force with center of mass.
r = np.sqrt((COM[i][0]-p[0, particle])**2 + (COM[i][1]-p[1, particle])**2 + (COM[i][2]-p[2,particle])**2 + smooth_grid**2)
accel_x += G*total_mass[i]*(COM[i][0]-p[0, particle])/r**3
accel_y += G*total_mass[i]*(COM[i][1]-p[1, particle])/r**3
accel_z += G*total_mass[i]*(COM[i][2]-p[2, particle])/r**3
return accel_x, accel_y, accel_z
#Acceleration of the particles
def accel_particle(p, particle, mass, index, particle_in_cell, smooth):
"""
Parameters
----------
p : array_like
The psition of all particles.
particle : int
The index of the particle.
mass : array_like
The mass of all particles.
index : int
The index of the grid of which the particle is in.
particle_in_cell : List
The index of particles in each grid cell.
Returns
-------
float
The acceleration in the x-direction.
float
The acceleration in the y-direction.
float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
#smooth = 1.0 #The smoothen scale is 100 pc which is bigger than the size of globular cluster (around 0.01 kpc, smallest possible
#mass) and the size of a dwarf galaxy (around 1 kpc, largest possible mass)
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
total = particle_in_cell[index]
#This is the collection of all particles in a specific grid.
for i in range(len(total)):
if (total[i] != particle):
#Calculate the force on the particle individually.
r = np.sqrt((p[0,total[i]]-p[0, particle]+smooth)**2 + (p[1,total[i]]-p[1, particle]+smooth)**2 + (p[2,total[i]]-p[2,particle]+smooth)**2)
accel_x += G*mass[total[i]]*(p[0,total[i]]-p[0, particle])/r**3
accel_y += G*mass[total[i]]*(p[1,total[i]]-p[1, particle])/r**3
accel_z += G*mass[total[i]]*(p[2,total[i]]-p[2, particle])/r**3
return accel_x, accel_y, accel_z
def acceleration(p, num, Np, mass, smooth):
"""
Parameters
----------
p : array_like
Position of all particles.
num : List
The number of particle in each grid.
Np : int
Total number of particles.
mass : array_like
The mass of each particle.
Returns
-------
float
The acceleration in the x-direction.
float
The acceleration in the y-direction.
float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
smooth = 1.0 #The smoothen scale is 100 pc which is bigger than the size of globular cluster (around 0.01 kpc, smallest possible
#mass) and the size of a dwarf galaxy (around 1 kpc, largest possible mass)
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
for i in range(Np):
if (i != num):
r = np.sqrt((p[0,i]-p[0, num])**2 + (p[1,i]-p[1, num])**2 + (p[2,i]-p[2,num])**2 + smooth**2)
accel_x += G*mass[i]*(p[0,i]-p[0, num])/r**3
accel_y += G*mass[i]*(p[1,i]-p[1, num])/r**3
accel_z += G*mass[i]*(p[2,i]-p[2, num])/r**3
return accel_x, accel_y, accel_z
def recession_vel(position, H_0):
v_rec = position*Hubble_convert(H_0) #return the recession velocity in kpc/Myr
return v_rec
def Hubble_convert(H_0):
result = H_0*1000.0*3.1536*10**13/(3.09*10**16)/10**6 #This formula convert the Hubble parameter from
#km/s/Mpc to Myr^-1 in order to match the unit convention in this program
return result
mass_min = 10**4
mass_max = 10**5
bound_xy = 10
bound_z = 10
method = 1
grid_xy = 5
grid_z = 5
Np = 2
Nt = 100
dt = 0.1
v_max = 0.2/np.sqrt(3.0)
H_0 = 70
smooth = 1
if (method == 0):
text = 'Grid'
elif (method == 1):
text = 'Exact'
t0 = time.time()
# For reproducibility, set a seed for randomly generated inputs. Change to your favourite integer.
np.random.seed(4080)
# Set the number of spatial dimensions (at least 2)
Nd = 3
#The array that contains the index of the particles.
particle_index = np.arange(Np)
# Set how long the animation should dispay each timestep (in milliseconds).
frame_duration = 100
#boundary for x, y and z
x_bound = bound_xy
y_bound = bound_xy
z_bound = bound_z
# Set initial positions at random within box
# position_xy = (np.random.normal(loc = 0.0, scale = 4, size = (2, Np)))
# #position_xy = (1.0-2.0*np.random.random(size = (2, Np)))*bound_xy/2
# #Gaussian initial condition in the xy plane.
# position_z = (1.0-2.0*np.random.random(size = (1, Np)))*z_bound
# #Random distribution in the z direction.
#This gives the initial condition
#position = np.concatenate((position_xy, position_z), axis = 0)
position = np.random.normal(loc = 0.0, scale = 2, size = (Nd, Np))
#position = (1.0-2.0*np.random.random(size=(Nd,Np)))*x_bound/2
#position = apply_boundary(position, Nd, Np)
#position_1 = np.random.normal(loc = -15.0, scale = 5, size = (3, 100))
#position_2 = np.random.normal(loc = 15.0, scale = 5, size = (3, 100))
#position = np.concatenate((position_1, position_2), axis = 1)
mass = 10**(np.random.random(size=(Np))*(np.log10(mass_max)-np.log10(mass_min)) + np.log10(mass_min))
#The mass range of dark matter halos. The minimum and maximum mass is determined by the user.
#velocity = np.random.normal(loc=0.0, scale = 0.03, size = (Nd, Np))
velocity = (1.0-2.0*np.random.random(size = (Nd, Np)))*v_max
#velocity_1 = np.full((Nd, 100), v_max)
#velocity_2 = np.full((Nd, 100), -v_max)
#velocity = np.concatenate((velocity_1, velocity_2), axis = 1)
#velocity = np.zeros((Nd, Np))
#print(velocity)
position += velocity/2.0*dt #first step of Leapfrog method.
position = apply_boundary(position, Nd, Np)#Apply the periodic boundary condition
position_new = np.reshape(np.concatenate((position[0,:], position[1,:])), (2, Np)).T
#Position_new is a 2xNp matrix. The first column is the x position of particles and the second column
#is the y position.
#print(position_new[:,0])
gridsize = np.array([grid_xy,grid_xy,grid_z])
#This array contains the number of grids in each direction.
x = np.linspace(-x_bound, x_bound, gridsize[0]+1)
y = np.linspace(-y_bound, y_bound, gridsize[1]+1)
z = np.linspace(-z_bound, z_bound, gridsize[2]+1)
#x, y and z are the grid in each direction.
grid_length = np.sqrt(((x[1]-x[0])/2)**2 + ((y[1]-y[0])/2)**2 + ((z[1]-z[0])/2)**2)
#The length of the grid.
# Set the axes on which the points will be shown
plt.ion() # Set interactive mode on
fig = plt.figure(figsize=(8,8)) # Create frame and set size
ax = plt.axes() # Set the axes as the only set (you may wish to change this later)
ax.set_xlabel('kpc')
ax.set_ylabel('kpc')
ax.set_title('Collision of two galaxies')
# Create command which will plot the positions of the particles
scat = plt.scatter(position_new[:,0], position_new[:,1], s= (np.log10(mass))**2)
#This is a scatter plot. It takes in column vector of x and y position. s is the size of the particle which depends
#on the mass of the particle.
# Define procedure to update positions at each timestep
def update(i):
global position,velocity, mass, Nd, Np, particle_index, method, H_0 # Get positions and velocities, mass,
#number of particles, index of particles and the method of evaluation.
year = i*dt #Shows how many year has passed since the initial condition.
#points.set_label('%lf Myrs'%year)
scat.set_label('%lf Myrs'%year)
num, particle_in_cell = find_particle(position, x, y, z)#This returns the index of particles in each grid
#and the number of particles in each grid.
COM, total_mass = center_of_mass(particle_in_cell, num, mass, position)
#This returns the center of mass, and the total mass in each grid.
index = position2grid(particle_index, particle_in_cell)#This maps the particle index to the grid it is in.
accel = np.zeros(shape = position.shape)#The acceleration of the particle.
if (method == 0):
for i in range (Np):
particle = i #The index of the particle.
grid_index = index[i][1] #The index of the grid.
accel[0,i], accel[1,i], accel[2,i] = accel_grid(COM, total_mass, mass, grid_index, position, particle, particle_in_cell, num, grid_length, smooth)
#This gives the acceleration of the particle with the grid method
elif (method == 1):
for i in range(Np):
accel[0, i], accel[1, i], accel[2, i] = acceleration(position, i, Np, mass, smooth)
#This returns the acceleration of the particle with the exact method.
else:
sys.exit("Invalid input for method of simulation.")
velocity += accel*dt #Calculate the new velocity with Euler's method.
#print(position[:,35])
position += (velocity+recession_vel(position,H_0))*dt # Increment positions according to their velocites
#print(accel[:,35])
position = apply_boundary(position, Nd, Np) # Apply boundary conditions
ax.set_xlim(-x_bound-x_bound*Hubble_convert(H_0)*year,x_bound+x_bound*Hubble_convert(H_0)*year) # Set x-axis limitslpitns,
ax.set_ylim(-y_bound-y_bound*Hubble_convert(H_0)*year,y_bound+y_bound*Hubble_convert(H_0)*year) # Set y-axis limits
scat.set_offsets(np.reshape(np.concatenate((position[0,:], position[1,:])), (2, Np)).T)#This line of code basically
#combine the array of x and p position and then transpose it. This is because the scatter plot only accepts
#column vector of x and y position.
plt.legend()#Display the time in the lower right corner.
return scat,
# Create animation
# https://matplotlib.org/api/_as_gen/matplotlib.animation.FuncAnimation.html
plt.rcParams['animation.ffmpeg_path'] = "./ffmpeg/bin/ffmpeg.exe"
ani = animation.FuncAnimation(fig, update, frames=Nt,interval = frame_duration, blit = True)
FFwriter = animation.FFMpegWriter()
ani.save("twobody.mp4", writer = FFwriter)
t1 = time.time()
print(t1-t0) #Prints out the time require to complete the simulation.
plt.show()
# -
# Run the cell below to see the video.
# +
from IPython.display import Video
Video("twobody.mp4")
# -
# Looks like the gravity is working properly, you can see the two particles are getting closer closer together overtime. Let's move on to our next demonstration.
# ## How are the galaxies formed???
# You may have heard of the cosmological principle that states the universe is satistically homogeneous (same property in all position) and isotropic (same property in all direction). This means on large scale, particles are just randomly distributed in space. However, how galaxies which has a much higher density than the average density of the universe form? It turns out since the gravitational force are attarctive, if the density of a region is much higher than the average density of the universe. The particles will begain cluster together and form structures. The next video will show how this happens.
# ### Remark
# This section is meant to be interactive. However, this is my first time using Jupyter notebook and I don't want to add a 600 line code cell each time to generate the simulation. Therefore, if you want to generate the video yourself, just go the the python code above and change some initial conditions. The inital condition for the next video is (just change these parameters and leave other parameters the same):
#
# position = (1.0-2.0*np.random.random(size=(Nd,Np)))*x_bound/2
#
# velocity = np.random.normal(loc=0.0, scale = 0.1, size = (Nd, Np))
#
# Np = 100
#
# bound_xy = 20
#
# bound_z = 20
#
# dt = 0.5
# +
from IPython.display import Video
Video("gravitational collapse.mp4")
# -
# Initially, the particles are distributed randomly on from -10 kpc to 10 kpc in each direction. Then we can see due to the graviational effect, all particles start to fall inwards into the center. This is because particles that are close together will first start to cluster together which increases the density in that region. From Newton's law of gravity, larger mass lead to a stronger gravitational force. Therefore, other particles that are initially far away starts to feel the force and pull to the cluster of mass. This feedback loop continues until all partciles are pulled towards the center of mass of the system. When some particle with smaller mass travel close to the center of the galaxy, it will get a huge boost in velocity accouting to Newton's law of gravity. Consequently, it is being flung out the galaxy.
# ### Initial conditions
# Some of you may try to input different initial conditions and find out the video you get is totally different from the one I have shown here and maybe wondering how do I know which initial condition to use to produce this kind of simulation. There are some general tricks to choose an initial condition. Let's take the simulation above as an example.
#
# A typical size of galaxies are around tens of kiloparsecs. For example, the diameter of the Milky way is around 32 kiloparsecs. The initial distribution of particles has a diameter around 20 kiloparsecs which is similar to the size of the normal galaxy.
#
# Another component in Newton's law of gravity is the mass of the particle. The total mass of the particle in the simulation above is around 500 billion solar masses, which is also a typical mass for a galaxy. The mass of particle is different. If the size of the particle is bigger, it means the particle has a bigger mass.
#
# The soften length is a parameter which determine the scale of which the simulation is no longer able to resolve. I determine the soften length from the typical size of the maximum particle possible in the simulation. For example, the maximum mass possible in the above simulation is 10 billion solar masses which is a typical mass of a dwarf galaxy. The typical length of the dwarf galaxy is around 1 kiloparsec, so the soften length is chosen to be 1 kiloparsec.
#
# A general rule of thumb for the time step is the ratio between the soften length and the typical velocity of particles.
# ## Particles???
# Some of you may wonder what is the physical representation of the particle in the simulation. In reality, they are dark matter halos. The dark matter halos are cosmological structures in the universe that are decoupled from the expansion of the universe and the matter inside are bound by gravity.
# ## How do we know dark matter exist ???
# As you may have heard, dark matter does not interact with light so they are not visible to us, so how do we know dark matter exist. Well, we know the amount of luminous (baryonic) matter density is about 10 % of the dark matter density. Why don't we reduce the mass of the dark matter halo to 10% of its original mass and see what happen.
#
# Initial conditions:
# The same initial condition as the galaxy formation simulation except set mass_min and mass_max to 10 % of its original value.
# +
from IPython.display import Video
Video("gravitational collpase baryon only.mp4")
# -
# As you can see from the video above, the particles do not cluster together as in the previous video. Consequently, the galaxy is not formed in this simulation. This shows baryon alone is not able to form galaxies. There are exotic matters that have mass but not interact with light in order for galaxies to form. This is one of the strongest evidence that support the dark matter theory.
# Well, what if the density of baryon in some regions is so high so it may be able to form a galaxy in the first place? The next video will show even that's the case, particles will soon disperse.
#
# Inital conidtions:
#
# position = np.random.normal(loc = 0.0, scale = 4, size = (Nd, Np))
#
# velocity = (1.0-2.0*np.random.random(size = (Nd, Np)))*v_max
#
# The inital position changes from a random distribution to a Gaussian distribution since most stars are concentrated in the buldge area and the outskirts of galaxy contain fewer stars. I choose the initial velocity profile to be random which is not a very accurate representation of velocities of stars in the galaxy. This is due to the velocity of stars in the galaxy is hard to model. The maximum velocity of particle in the simulation is 0.2 kpc/Myr which is around 200 km/s similar to the velocity of stars around the disk of the Milky way.
# +
from IPython.display import Video
Video("galaxy baryon only.mp4")
# -
# # Expanding universe
# The universe is undergoing a period of accelerating expandsion due to the mysterious dark energy. In order to incorperate the expansion of the universe into the simulation, the velocities the particles have two components. The peculiar velocity due to the gravitational interaction with other particles in the simulation and the recession velocity due to the expansion of the universe. We can determine the recession velocity by multiplying the Hubble constant with the distance of the particle to the observer. In the simulation, we assume the observer is located at the origin. In the kiloparsec scale, the effect of recession velocity on the motion of the stars is neglectable. However, the recession velocity becomes important when we do a simulation on the scale of megaparsec. The following video shows the motion of galaxies in a galaxy cluster which size is usually on the order of megaparsec.
#
# Initial conditions:
#
# mass_min = 10**8
#
# mass_max = 10**9
#
# velocity = np.zeros((Nd, Np))
#
# bound_xy = bound_z = 10**4
#
# position = (1.0-2.0*np.random.random(size=(Nd,Np)))*x_bound/2
#
# smooth = 10**3
#
# dt = 50
#
# The typical mass of clusters are around 10**15 solar masses, so the min_mass and max_max are adjusted such that the total mass of particles in the simulation is around the same mass as the cluster.
# +
from IPython.display import Video
Video("galaxy cluster.mp4")
# -
# Initially the galaxies are randomly distributed, but after some time, the galaxies are pulled together due to the gravitational force and start to form galaxy clusters. The x and y axis are expanding due to the expansion of the universe. In this case, the gravitational force betweeen particles are so strong such that it defeats the expansion of the universe. Therefore, a galaxy cluster is able to form at the end. Galaxy clusters are the largest gravitational bound object in the universe.
#
# # Grid approximation
# All simulations above are produced using the exact solution with Newton's law of gravity. However, this process is time consuming since we have to calculate the force on one particle from all other particles and then repeat this for all particles. This means the total number of computation requires is on the order of $N_p^{2}$ where $N_p$ is the number of particles in the simulation. This is a bad news because it will take a significantly longer time to complete the simulation if we double the amount of particles in the simulation. My code tackles this problem with the grid approximation techniue which is a simplified version of the tree code.
# Firstly, like the tree code, the grid approximation method will first divide the simulation into a number of grid cells. However, in tree code, we stop dividing when their is only one particle in each grid. However, in the grid approximation method, the number of grids is given by the user. Then we calculate the center of mass of all grid cells and filter out the grid cell that has zero particles. Lastly, we compute the force on a particle by computing the force between the particle and the center of mass of other grid cells of the distance between the particle and the center of mass is larger than the size of the grid cell. Otherwise, we calculate the force between the particle and the particles in other grid cells.
from IPython.display import Image
Image(filename='table.png')
# THe table above demonstrates the amount of time require to run one frame of the simulation for the grid approxomation and the exact method. When the number of particles are less than 100, the grid approximation only takes slightly shorter time than the exact method to finish the simulation. However, this situtation changes drastically when the number of particles are larger than 100. At around 300 particles, the grid approximation only takes one second less than the exact method to finish the simulation. At around 1000 particles, the grid approximation now only takes around half of the time of the exact method to finish the simulation.
from IPython.display import Image
Image(filename='method comparison.png')
# The figure above shows the logarithmic plot of the running time aginast the number of particles. the straight line is the linear regression model for each method. The linear regression model demonstrates the running time for the exact method is proportional to $N^{1.6 \pm 0.1}$ while the grid approximation method is proportional to $N^{1.2 \pm 0.1}$. Therefore, if you have a lot of particles in the simulation. The grid approximation method will run much quicker than the exact solution. Run the next two cell will show you the approximation method gives a very similar simulation to the exact method. The first video is the exact method and the second video is the approximation method.
# +
from IPython.display import Video
Video("gravitational collapse.mp4")
# +
from IPython.display import Video
Video("gravitational collapse grid.mp4")
# -
# ## Some tips
# The running time for the grid approximation method depends on the number of cells in each direction and possibly the initial conditions. To achieve optimal efficiency, you can run the grid approximation for one frame and see which grid size gives the fastest running time. Typically 4 to 6 grids in each direction is enough if the number of particles is between one hundred to few thoudsands.
# # Galaxy statistics
# We can obtain a lot of information about the universe from analysing the clustering of galaxies. In cosmology, there are two main two statistical quantities used to measure the degree of clustering of galaxies. One of them is the two-point correlation function and the other one is the power spectrum. The two-point correlation function is related to the power spectrum by Fourier transform. In our case, the two-point correlation function is measure in unit of distance while the power spectrum depends on the wavenumber which is proportional to the inverse of distance. The Fourier transform is just a mathematical transformation to link this two quantities together.
# The two-point correlation function measure the excess (compare to random distribution) probability of finding distance of galaxy with distance r. In my simulation, the two point correlation is calculated through the Peebles-Hauser estimator: $$ \xi = \frac{DD(r)-RR(r)}{RR(r)} $$.
# The number of pairs separated by distance r in the dataset is denoted by DD(r) and RR(r) represents the number of pair separated by distance r in a random distributed data set. RR(r) can be determined analytically by (http://articles.adsabs.harvard.edu/pdf/1986ApJ...301...70R): $$ RR(r) = \frac{N^2}{V} V_{shell}$$
# The total number of particles in denoted by N, the volume of the simulation is denoted by V and the volume of the shell with size dr is denoted by $V_{shell}$. In my code, the volume of the shell is the difference between the volume of sphere with radius r amd r+dr. Although the simulation is a cube, the periodic boundary condition allows us to approximate the volume of the simulation as a sphere. The power spectrum is related to the two-point correlation function by $$ P(k) = 2 \pi \int_{0}^{\infty} dr r^2 \frac{\sin{(k r)}}{k r} \xi(r) $$
# The power spectrum is just the Fourier transform of the two-point correlation function. Hence, both the power spectrum and two-point correlation function gives the same information about the clustering of galaxies. In our case, the correlation function is discrete. There are two different ways to evaluate the power spectrum. The first method is to use the trapezoidal rule to calculate the power spectrum and the second way is to use an interpolation spline to calculate the power spectrum and then evaluate the integral numerically.
# ## Compare trapezoidal rule to numerical integration.
# +
from IPython.display import Video
Video("integration.mp4")
# -
# The video above shows there is no difference when we use spline integration or the trapezoidal rule. Both methods give exactly the same power spectrum. However, it normally takes about 10 seconds for the numerical simulation to complete while the trapezoidal rule is usually done in one second. Therefore, the trapzoidal rule is more time efficient.
# ### Remark
# To calculate the spline of the power spectrum in the above figure. We first need to numerically integrate the correlation function to get the power spectrum. Then we use another cubic spline to smooth this power spectrum and then plot the new cubic spline. The code for this is included in the test3.py. For the rest of the presentation, I will use the trapezoidal rule to calculate the power spectrum since it is more time efficient. Then I will use the cubic spline again to smooth the power spectrum. Additionally, the power spectrum in the above figure is normalized. However, if we normalized the power spectrum, it is harder to see how the power spectrum will change over time. This is because the normalization constant of the power spectrum is usually much larger than the chnages in the power spectrum over time. Therefore, I will also use the original version of the power spectrum for the rest of my presentation.
# ## An oscillatory power spectrum???
# The power spectrum above is very different to the power spectrum we see in real life. Usually the matter power spectrum will follow some sort of power law of the wavenumber k. Then why does the power spectrum above shows an oscillatory behaviour.
# ### Warning: math incoming (not interested in math, go to the next section)
# The sinc function in the integrand is highly oscillatory. This could cause the power spectrum to show oscillatory behaviour as well. Here is the (not so rigorous) mathematical proof:
#
# First consider the trapezoidal rule. We are just truncating the correlation function. When we only have one pair of galaxies, the integral becomes: $$ \int_x^{x+dx} x^2 \frac{\sin{k x}}{k x} \xi(x) + \int_0^x x^2 \frac{\sin{k x}}{k x} \times -1 + \int_{x+dx}^{x_{max}} x^2 \frac{\sin{k x}}{k x} \times -1. $$ Here, the maximum separation inside the simulation is denoted by $ x_{max} $ and since we are binning the correlation function, the value of correlation function between x and x + dx is constant. Let's call that constant c. Then the correlation function simplifies to $$ c (\frac{\sin{k (x+dx)} - k (x+dx) \cos{k (x+dx)}} {k^3} - \frac{\sin{k x} - k x \cos{k x}} {k^3}) - \frac{\sin{k x} - k x \cos{k x}} {k^3} - (\frac{\sin{k x_{max}} - k x_{max} \cos{k x_{max}}} {k^3} - \frac{\sin{k (x+dx)} - k (x+dx) \cos{k (x+dx)}} {k^3})$$ You can see the power spectrum is now just the sum of cosine and sine functions. The same situation happens when you have multiple pairs. For bins that have pairs of galaxies, the integrand will become the value of correlation function in that bin times the sinc function and $x^2$. When their is no pair, the integrand is then minus of the sinc function times $x^2$. Both integral will gives back sine and cosine function which are oscillatory. Therefore, the power spectrum is also oscillatory (I just realized the proof here is for the rectangular rule. However, the proof for trapezoidal rule is pretty similar except $\xi(x)$ between x and x+dx is a linear function connects $\xi(x)$ and $\xi(x+dx)$. The highest order of the linear function is x, integration by parts will show the final function is still the sum of sine and cosine function).
#
# Then consider the correlation function is smooth, then the power spectrum of one pair of galaxies at x = a becomes: $$ \lim_{dx \to 0} \int_x^{x+dx} x^2 \frac{\sin{k x}}{k x} \xi(x) + \int_0^x x^2 \frac{\sin{k x}}{k x} \times -1 + \int_{x+dx}^{x_{max}} x^2 \frac{\sin{k x}}{k x} \times -1 $$ Then the correlation function behaves like a kronecker delta function and the integral is simplifed to $$ a^2 \frac{\sin{a k}}{a k} \xi(a) - (\frac{\sin{k x_{max}} - k x_{max} \cos{k x_{max}}} {k^3}) $$ In our simulation the correlation function is smooth by a cubic spline. Therefore, to the leading order, the correlation function is proportional to $x^3$. From integration by parts, even we have multiple pairs of galaxies, the integral will still gives a sum of cosine and sine functtion.Therefore, in this case, the power spectrum is still a sum of cosine and sine function. Consequently, as shown in the figure above, smoothing the correlation function does not reduce the oscillatory behaviour of the power spectrum. This shows unless we have a smooth continous correlation function to begin with, smoothing it with a polynomial spline will not reduce the oscillatory behaviour.
# The proper way to calculate the power spectrum is computing the density perturbation at different position and Fourier transform the density perturbation into Fourier space. The power spectrum is defined as the variance of the density perturbation in Fourier space with different wavenumber (there maybe a factor of $(2 \pi)^3$ depends on the form of Fourier transform). Therefore, the power spectrum in the code is just a rough approximation of the actual power spectrum.
# # Some examples of the power spectrum and the correlation function.
# +
from IPython.display import Video
Video("collapse.mp4")
# -
# The propose of the spline of both correlation function and power spectrum is to make the graph look smoother. The initial condition here is:
#
# position = np.random.normal(loc = 0.0, scale = 4, size = (Nd, Np))
#
# velocity = v_max*(1-2*np.random.random((Nd,Np)))
#
# v_max= 0.2
#
# smooth = 1.0
#
# dt = 0.5
#
# H_0 = 70.0
#
# mass_min = 10**4
#
# mass_max = 10**5
# The video above shows how different particle will cluster together if the initial position of the particles follow a Gaussian distribution. Since initially the particles follow a Gaussian distribution, we can see the value of the correlation function is high at low separation and at high separation, the correlation function approaches -1. There is a large overdensity around the center of the box, this attarct all particles inside the simulation toward the center. We can see the correlation function at low sparation jumps up as expected. After some time, some of the particles are being flung out of the center due to the large gravitational force at the center. The correlation function at the center decreases. The limit of the x-axis is set by the largest separation where the correlation function is greater than zero. As you can see in the simulation, the largest separation with positive correlation function first decreases because particles are falling into the center and then increases since some particles are being flung out of the center.
#
# For the power spectrum, the part with high wavenumber changes significantly during the simulation. Higer wavenumber corresponds to smaller scale. The simulation shows the particles are cluster together into smaller and smaller scale, so the power spectrum on small sclae changes. However, the power spectrum on large scale (small k) stays the same since their scale is similar to the size of the simulation. On these scales, the there is not much changes such as no particle leave or enter the system. Therefore, the power spectrum on large scale stays the same. In this video, we did not normalize the power spectrum, the unit of the power spectrum should be $(kpc)^3$.
# +
from IPython.display import Video
Video("collision.mp4")
# -
# The video above shows how two galaxies will interact with each other when they collide. The initial condition are
#
# Np = 200
#
# Nt = 300
#
# mass_min = 10**3
#
# mass_max = 10**4
#
# position_1 = np.random.normal(loc = -15.0, scale = 5, size = (3, 100))
#
# position_2 = np.random.normal(loc = 15.0, scale = 5, size = (3, 100))
#
# position = np.concatenate((position_1, position_2), axis = 1)
#
# velocity_1 = np.full((Nd, 100), v_max)
#
# velocity_2 = np.full((Nd, 100), -v_max)
#
# velocity = np.concatenate((velocity_1, velocity_2), axis = 1)
#
# smooth = 1.0
#
# dt = 0.5
#
# H_0 = 70.0
# As two galaxies approach each other, we can see the correlation function at low separation increases significantly. It reaches the maximum approximately when the two galaxies are merge together. After the collision, both galaxies becomes more disperse but with a much denser core. Some of the particles are being flung out during the collision due to the extremely high gravitaitonal potential at the center. After the collision, the correlation function at low separation decreases as two galaxies moves further away.
#
# For the powerspectrum, similar to the last video, the small scale power spectrum changes significantly and the power spectrum at large scale stay the same.
# There are two apparent inaccurate description of galaxy collision in the video above. Firstly, both galaxies collapse onto itself even before the collision occurs. This is could be because the mass of the single particle is still a little too high. The simulation still requires some extra fine-tuning in order to match the real condition. Secondly, we may expact to have two peaks in the correlation function, one at low separation and the other at around the mean separation of the two galaxies. However, this is due to the analytical formula for the separation of two galaxies in a random distribution (RR(r)) is proportional to $r^2$ to the leading order (For a sphere, V_shell is proportional to $(r+dr)^3-r^3$ which is proportional to $r^2$ to the leading order). Therefore, for the random distribution, when the size of the bin is fixed, we are expected to find more pairs at a larger separation. Consequently, this reduce the correlation function at large separation and we don't see two peaks in the correlation function.
# # Try it yourself
# The following cell will include the python code that calculate the motion of particles, correlation function and the power spectrum together. You can play with it and see how particles behave based on your initial condition.
# +
import numpy as np
from pylab import *
import matplotlib.pyplot as plt
import matplotlib.animation as animation
from mpl_toolkits.mplot3d import Axes3D
from scipy.interpolate import CubicSpline
from scipy import integrate
import time
from scipy.interpolate import InterpolatedUnivariateSpline
# This is a simple skeleton code just to give some ideas.
# It plots collisionles particles moving at random in a cubic box in the main panel
# and shows the distribution of their separations in one of two other panels.
def find_particle(position, x, y, z):
"""
This function finds the index of particles in each grid cell.
Parameters
----------
position : array_like
This is the position of all particles in the simulation.
x : array_like
The array contains the grid point in the x direction.
y : array_like
The array contains the grid point in the y direction.
z : array_like
The array contains the grid point in the z direction.
Returns
-------
num : List
Number of particles in each grid.
particle_in_cell : List
The index of particles in each grid cell.
"""
particle_in_cell =[]
num = []
#Initialise the output list
limit_x = len(x)-1
limit_y = len(y)-1
limit_z = len(z)-1
#Number of computation requires for each dimension.
counter_x = 0
counter_y = 0
counter_z = 0
#This counts the number of cells that have been computed in each direction.
for i in range(limit_x*limit_y*limit_z):
position_x = position[0,:]
position_y = position[1,:]
position_z = position[2,:]
#The poistion of the particle in each direction.
xx = np.where(position_x < x[counter_x+1], position_x, 0)
yy = np.where(position_y < y[counter_y+1], position_y, 0)
zz = np.where(position_z < z[counter_z+1], position_z, 0)
#Find the particles in the position array which are to the right of the desired grid. For such particle
#replace the position with zero.
xxx = np.where(xx > x[counter_x], xx, 0)
yyy = np.where(yy > y[counter_y], yy, 0)
zzz = np.where(zz > z[counter_z], zz, 0)
#Find the particles in the position array which are to the left of the desired grid. For such particle
#replace the position with zero.
index_x = np.nonzero(xxx)[0]
index_y = np.nonzero(yyy)[0]
index_z = np.nonzero(zzz)[0]
#Find the index of the particle which are nonzero. These particles are located in the desired grid.
#print(index_x, index_y, index_z)
xy = np.intersect1d(index_x, index_y, assume_unique=True)
xyz = np.intersect1d(xy, index_z, assume_unique=True)
#The codes above finds the index of particle in the desired grid. The index in each array is unique.
if (len(xyz != 0)):
num.append(len(xyz))
particle_in_cell.append(xyz)
#Append the particle in the grid and the number of particle in the grid if the number of particles in
#the grid is nonzero.
counter_x += 1
#Move to the grid at the right
if (counter_x == limit_x):
#This means it completes calculate the particles in the grid in a row. Advance to the next row.
counter_x = 0
counter_y += 1
if (counter_y == limit_y):
#This moves to the next layer of the xy-plane.
counter_y = 0
counter_z += 1
return num, particle_in_cell
def apply_boundary(p, Nd, Np):
"""
This function applies the periodic boundary condition to the position of particles.
Parameters
----------
p : array_like
Position of all particle in the array.
Nd : int
Number of dimensions of the simulation.
Np : int
Number of particles in the simulation.
Returns
-------
p : array_like
The position of particles after applying the boundary condition.
"""
# Modify to apply your chosen boundary conditions
position_x = p[0,:]
position_y = p[1,:]
position_z = p[2,:]
#The position of particles in the x, y and z position.
#The following lines will find the particle outside the simulation and move it back to the simulation
#based on the periodic boundary condition.
xx = np.where(position_x > x_bound, (position_x-x_bound)-x_bound, position_x)
xxx = np.where(xx < -x_bound, x_bound - (-x_bound - xx), xx)
yy = np.where(position_y > y_bound, (position_y-y_bound)-y_bound, position_y)
yyy = np.where(yy < -y_bound, y_bound - (-y_bound - yy),yy)
zz = np.where(position_z > z_bound, (position_z-z_bound)-z_bound, position_z)
zzz = np.where(zz < -z_bound, z_bound - (-z_bound - zz), zz)
p = np.concatenate((xxx, yyy, zzz), axis = 0)
p = np.reshape(p, (Nd, Np))
#Reconstruct the array for position of particles.
return p
def center_of_mass(particle_in_cell, num, mass, position):
"""
This function calculates the center of mass of all particles in the same grid cell.
Parameters
----------
particle_in_cell : List
The list that contain the index of particles in each grid cell.
num : List
The list contains the number of particles in each grid cell.
mass : array_like
The mass of all the particles.
position : array_like
The position of all the partiles.
Returns
-------
result : List
The center of mass position in each grid cell.
total_mass : List
The total mass of all particles in each grid cell.
"""
result = []
total_mass = []
#Initialise the output lists
position_x = position[0,:]
position_y = position[1,:]
position_z = position[2,:]
for i in range(len(particle_in_cell)):
COM_x = 0.0
COM_y = 0.0
COM_z = 0.0
M_total = 0.0
#Initialise the center of mass position and the total mass of particles in the grid
for j in range(num[i]):
COM_x += mass[particle_in_cell[i][j]]*position_x[particle_in_cell[i][j]]
COM_y += mass[particle_in_cell[i][j]]*position_y[particle_in_cell[i][j]]
COM_z += mass[particle_in_cell[i][j]]*position_z[particle_in_cell[i][j]]
M_total += mass[particle_in_cell[i][j]]
#Calculate the center off mass
result.append(np.array([COM_x/M_total, COM_y/M_total, COM_z/M_total]))
total_mass.append(M_total)
return result, total_mass
def position2grid(particle_index, particle_in_cell):
"""
This function matches the index of the particle to the index of grid cell it is in.
Parameters
----------
particle_index : array_like
Index of all particles in the simulation.
particle_in_cell : List
The index of particle in each grid cell.
Returns
-------
result : List
It matches the index of particle to the index of the grid it is in.
"""
result = []
for i in range(len(particle_index)):
for j in range(len(particle_in_cell)):
size = (np.intersect1d(np.array([i]), particle_in_cell[j]))
#Determine whether the particle is in the grid[j]
if (size.size > 0):#If the particle is in grid[j], the size of the array will be nonzero.
#Since the index of particle is also unique, we are certain that when the size of array is not zero.
#we find the cell of which the particle is in
break
result.append(np.array([i,j]))
return result
def accel_grid(COM, total_mass, mass, index, p, particle, particle_in_cell, num, grid_length, smooth):
"""
This uses the center of mass to calculate the acceleration of a particle.
Parameters
----------
COM : List
Center of mass of all grid cells.
total_mass : List
Mass of all particles in a single grid cell.
mass : array_like
The mass of each individual particle.
index : int
The index of the grid cell.
p : array_like
Position of all the particle.
particle : int
Index of the particle.
particle_in_cell : List
The list contains the index of particles in each grid cell.
num : List
Number of particles in each grid cell.
grid_length : The length of the gridcell
A reference length. If the distance between particle and center of mass of any grid is below the
reference length. We will calulate the interaction in particle-particle basis (similar to P3M).
Returns
-------
float
The acceleration in the x-direction.
float
The acceleration in the y-direction.
float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
smooth_grid = grid_length #The smoothen scale which is set to length of the grid.
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
for i in range(len(COM)):
r_dash = np.sqrt((COM[i][0]-p[0, particle])**2 + (COM[i][1]-p[1, particle])**2 + (COM[i][2]-p[2,particle])**2)
#The distance between the particle and the center of mass of particles.
if (r_dash <= grid_length):
#If less than the grid size, calculate the force using individual particle.
accel_x += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[0]
accel_y += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[1]
accel_z += accel_particle(p, particle, mass, i, particle_in_cell, smooth)[2]
else:
#Larger than the gridsize, calculate the force with center of mass.
r = np.sqrt((COM[i][0]-p[0, particle])**2 + (COM[i][1]-p[1, particle])**2 + (COM[i][2]-p[2,particle])**2 + smooth_grid**2)
accel_x += G*total_mass[i]*(COM[i][0]-p[0, particle])/r**3
accel_y += G*total_mass[i]*(COM[i][1]-p[1, particle])/r**3
accel_z += G*total_mass[i]*(COM[i][2]-p[2, particle])/r**3
return accel_x, accel_y, accel_z
#Acceleration of the particles
def accel_particle(p, particle, mass, index, particle_in_cell, smooth):
"""
This calculates the acceleration of particle on a particle-particle basis.
Parameters
----------
p : array_like
The psition of all particles.
particle : int
The index of the particle.
mass : array_like
The mass of all particles.
index : int
The index of the grid of which the particle is in.
particle_in_cell : List
The index of particles in each grid cell.
Returns
-------
accel_x: float
The acceleration in the x-direction.
accel_y: float
The acceleration in the y-direction.
accel_z: float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
#smooth = 1.0 #The smoothen scale is 100 pc which is bigger than the size of globular cluster (around 0.01 kpc, smallest possible
#mass) and the size of a dwarf galaxy (around 1 kpc, largest possible mass)
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
total = particle_in_cell[index]
#This is the collection of all particles in a specific grid.
for i in range(len(total)):
if (total[i] != particle):
#Calculate the force on the particle individually.
r = np.sqrt((p[0,total[i]]-p[0, particle])**2 + (p[1,total[i]]-p[1, particle])**2 + (p[2,total[i]]-p[2,particle])**2 + smooth**2)
accel_x += G*mass[total[i]]*(p[0,total[i]]-p[0, particle])/r**3
accel_y += G*mass[total[i]]*(p[1,total[i]]-p[1, particle])/r**3
accel_z += G*mass[total[i]]*(p[2,total[i]]-p[2, particle])/r**3
return accel_x, accel_y, accel_z
def recession_vel(position, H_0):
"""
This calculates the recession velocity of the particle at a given position
Parameters
----------
position : array_like
The position of every particle in the simulation
H_0 : float
The value of Hubble constant in km/s/Mpc
Returns
-------
v_rec : array_like
The recession velocity of the particle at each position.
"""
v_rec = position*Hubble_convert(H_0) #return the recession velocity in kpc/Myr
return v_rec
def Hubble_convert(H_0):
"""
Converts the Hubble parameter from km/s/Mpc to Myr^-1
Parameters
----------
H_0 : float
The Hubble parameter in km/s/Mpc.
Returns
-------
result : float
The Hubble parameter in Myr^-1.
"""
result = H_0*1000.0*3.1536*10**13/(3.09*10**16)/10**6 #This formula convert the Hubble parameter from
#km/s/Mpc to Myr^-1 in order to match the unit convention in this program
return result
#Acceleration of the particles
def acceleration(p, num, Np, mass, smooth):
"""
This uses the exact method to calculate the force on a particle.
Parameters
----------
p : array_like
Position of all particles.
num : List
The number of particle in each grid.
Np : int
Total number of particles.
mass : array_like
The mass of each particle.
Returns
-------
accel_x: float
The acceleration in the x-direction.
accel_y: float
The acceleration in the y-direction.
accel_z: float
The acceleration in the z-direction.
"""
G = 4.452*10**(-7) #in unit of kpc^3/10^5 solar masses/Myr^2
accel_x = 0.0
accel_y = 0.0
accel_z = 0.0
for i in range(Np):
if (i != num):
r = np.sqrt((p[0,i]-p[0, num])**2 + (p[1,i]-p[1, num])**2 + (p[2,i]-p[2,num])**2 + smooth**2)
accel_x += G*mass[i]*(p[0,i]-p[0, num])/r**3
accel_y += G*mass[i]*(p[1,i]-p[1, num])/r**3
accel_z += G*mass[i]*(p[2,i]-p[2, num])/r**3
return accel_x, accel_y, accel_z
def separation(p):
"""
This code clulates the seperation between two particles
Parameters
----------
p : array_like
The position of all particles in the simulation.
Returns
-------
float
The separation between all particles in the simulation.
"""
# Function to find separations from position vectors
s = (p[:,None,:] - p[:,:,None]) # find N x N x Nd matrix of particle separations
return np.sum(s**2,axis=0)**0.5 # return N x N matrix of scalar separations
#Basic user interface. Ask the user to input the following parameter
print('To modify the initial condition, please modify the code directly.')
mass_min = eval(input("The minimum nonzero resolution mass (in 10^5 solar masses): "))
mass_max = eval(input("The maximum mass of the dark matter halo (in 10^5 solar masses): "))
bound_xyz = eval(input("The length of simulation in the xy plane (in kpc): "))
method = eval(input("Evaluation method, enter 0 for grid approximation and 1 for exact solution: "))
grid_xyz = eval(input("Total number of grids in the x or y position: "))
Np = eval(input("Total number of particles: "))
Nt = eval(input("Total number of time steps: "))
dt = eval(input("Time step (in Myr): "))
v_max = eval(input("The maximum drift velocity (in kpc/Myr): "))
H_0 = eval(input("The Hubble parameter (in km/s/Mpc). For static universe, enter 0.0: "))
smooth = eval(input("The soften length of the simulation (in kpc): "))
t_0 = time.time()
# For reproducibility, set a seed for randomly generated inputs. Change to your favourite integer.
np.random.seed(4080)
# Set the number of spatial dimensions (at least 2)
Nd = 3
# Set how long the animation should dispay each timestep (in milliseconds).
frame_duration = 100
#boundary for x, y and z
x_bound = bound_xyz
y_bound = bound_xyz
z_bound = bound_xyz
# Set initial positions at random within box
# position_xy = (np.random.normal(loc = 0.0, scale = 4, size = (2, Np)))
# position_z = np.random.random(size = (1, Np))*z_bound
# position = np.concatenate((position_xy, position_z), axis = 0)
#position = np.random.normal(loc = 0.0, scale = 4, size = (Nd, Np))
position_1 = np.random.normal(loc = -15.0, scale = 5, size = (3, 100))
position_2 = np.random.normal(loc = 15.0, scale = 5, size = (3, 100))
position = np.concatenate((position_1, position_2), axis = 1)
# Set initial velocities to be random fractions of the maximum
#velocity = v_max*(1-2*np.random.random((Nd,Np)))
velocity_1 = np.full((Nd, 100), v_max)
velocity_2 = np.full((Nd, 100), -v_max)
velocity = np.concatenate((velocity_1, velocity_2), axis = 1)
mass = 10**(np.random.random(size=(Np))*(np.log10(mass_max)-np.log10(mass_min)) + np.log10(mass_min))
position += velocity/2.0*dt #first step of Leapfrog method.
position = apply_boundary(position, Nd, Np)#Apply the periodic boundary condition
position_new = np.reshape(np.concatenate((position[0,:], position[1,:])), (2, Np)).T
#Position_new is a 2xNp matrix. The first column is the x position of particles and the second column
#is the y position.
separation_max = np.sqrt(3.0)*x_bound*2.0
#The maximum separation possible in the simulation.
# Set the axes on which the points will be shown
plt.ion() # Set interactive mode on
fig = figure(figsize=(12,6)) # Create frame and set size
subplots_adjust(left=0.1, bottom=0.1, right=0.95, top=0.95,wspace=0.15,hspace=0.2)
ax1 = subplot(121) # For normal 2D projection
ax1.set_xlabel('kpc')
ax1.set_ylabel('kpc')
ax1.set_title('Collision of two galaxies')
# # Create command which will plot the positions of the particles
scat = plt.scatter(position_new[:,0], position_new[:,1], s= (np.log10(mass))**2)
ax2 = subplot(222) # Create second set of axes as the top right panel in a 2x2 grid
xlabel('Separation (kpc)')
ylabel('Correlation function')
dx= 0.5 # Set width of x-axis bins
xb = np.arange(0, separation_max+dx,dx)
xb[0] = 1e-6 # Shift first bin edge by a fraction to avoid showing all the zeros (a cheat, but saves so much time!)
line, = ax2.plot([],[],drawstyle='steps-post', label='data') # Define a command that plots a line in this panel
smooth_line, = ax2.plot([],[],drawstyle='steps-post',label='Spline') #This plots the spline interpolation of the
#correlation function
ax2.legend(loc='upper right')
ax4 = plt.subplot(224) # Create last set of axes as the bottom right panel in a 2x2 grid
ax4.set_xscale('log')
ax4.set_yscale('log')
#Set both the x and y axis on a log scale.
plane, = ax4.plot([], [], drawstyle = 'steps-post', label='data') #This plots the power spectrum
smooth_plane, = ax4.plot([], [], drawstyle = 'steps-post', label='spline') #This plots the spline interpolation
#of the power spectrum.
xlabel('Wavenumber (kpc^-1)')
ylabel('Power spectrum (kpc^3)')
ax4.legend(loc='best')
# Define procedure to update positions at each timestep
def update(i):
global position,velocity, dx, mass, smooth # Get positions and velocities and bin width
N = position.shape[1]
year = i*dt #Shows how many year has passed since the initial condition.
scat.set_label('%lf Myrs'%year)
ax1.legend(loc='upper right')#Display the time in the lower right corner.
accel = np.empty(shape = position.shape)
for i in range(N):
accel[0, i], accel[1, i], accel[2, i] = acceleration(position, i, Np, mass, smooth)
velocity += accel
position += (velocity+recession_vel(position,H_0))*dt # Increment positions according to their velocites
#The total velocity is the sum of the peculiar velocity and the recession velocity.
position = apply_boundary(position, Nd, Np) # Apply boundary conditions
ax1.set_xlim(-x_bound-x_bound*Hubble_convert(H_0)*year,x_bound+x_bound*Hubble_convert(H_0)*year) # Set x-axis limits
ax1.set_ylim(-y_bound-y_bound*Hubble_convert(H_0)*year,y_bound+y_bound*Hubble_convert(H_0)*year) # Set y-axis limits
#points.set_data(position[0,:], position[1,:]) # Show 2D projection of first 2 position coordinates
scat.set_offsets(np.reshape(np.concatenate((position[0,:], position[1,:])), (2, Np)).T)#This line of code basically
DD = np.ravel(tril(separation(position)))#The separation of all particles in the dataset.
factor = Np**2/((2*x_bound)**3) #This is the number density of pair of particles in the simulation. Since we use
#periodic boundary condition, we can also consider our simulation in a sphere.
h_DD, x_DD = histogram(DD,bins=xb) #The number of pairs of galaxies in each bin.
h = np.zeros(len(h_DD)) #Correlation function
x_max = 0.0#The mximum value in the x-axis
for i in range(len(h_DD)):
h[i] = h_DD[i]/((4.0/3.0*np.pi*(xb[i+1]**3-xb[i]**3)*factor))-1.0 # calculate the correlation function
#using the estimator
if (h[i] > 0):
x_max = x_DD[i] #Find the largest separation where the correlation function is greater than 0.
line.set_data(x_DD[:-1],h) # Set the new data for the line in the 2nd panel
ax2.set_xlim(0, x_max)
ax2.set_ylim(-1, np.amax(h)+5)
variable_x = x_DD[:-1]
cs = CubicSpline(variable_x, h) #Find a spline interpolation between the bin and the correlation function
x = np.linspace(xb[0], np.sqrt(3.0)*2.0*x_bound, num=10000)
smooth_plot = cs(x) #Plot the correlation function with the spline interpolation.
smooth_line.set_data(x, smooth_plot)
k = 2.0*np.pi/(x_DD[:-1]) #The wavenumber.
k_min = np.amin(k) #The minimum wavenumber
k_max = 2.0*np.pi/smooth #the maximum wavenumber is fixed to the wavenumber of the soften length. This is
#because any scale below the soften length is in accurate.
k_order_max = int(np.floor(np.log10(k_max))) + 1
k_order_min = int(np.ceil(np.log10(np.amin(k)))) - 1
#all possible k are between 10^k_order_min and 10^k_order_max
N_op = k_order_max - k_order_min
#Number of steps to go from k_order_min to k_order_max
x_axis = []
order = k_order_min
#The following for loop will put each interval from 10^n to 10^(n+1) into 100 smaller intervals. This will
#help to smooth the power spectrum and make the spline interpolation easier later.
for i in range(N_op):
segment = np.linspace(10**order, 10**(order+1), 100, endpoint=False)
order += 1
x_axis.append(segment)
x_axis = np.array(x_axis)
size_x, size_y = x_axis.shape
x_axis = np.reshape(x_axis, (size_x*size_y))
PS = [] #The power spectrum
k_eff = [] #Wavenumbers between k_min and k_max
for i in range(len(x_axis)):
if ((x_axis[i] < k_min) or (x_axis[i] > k_max)):
continue
PS.append(np.trapz(variable_x**2*h*np.sin(x_axis[i]*variable_x)/(x_axis[i]*variable_x)*2.0*np.pi))
k_eff.append(x_axis[i])
PS = np.array(PS)
k_eff = np.array(k_eff)
ax4.set_xlim(k_min, k_max)
ax4.set_ylim(1, np.amax(PS)) #Since the y axis is set to log scale. The minimum value of y cannot be less than zero.
cs_ps = CubicSpline(k_eff, PS) #Spline interpolate the power spectrum.
k_final = np.linspace(k_min, k_max, 10000)
PS_spline = cs_ps(k_final) #The spline interpolation of the power spectrum.
smooth_plane.set_data(k_final, PS_spline)
plane.set_data(k_eff,PS)
return scat, plane, smooth_line, line, smooth_plane # Plot the points and the line
# Create animation
# https://matplotlib.org/api/_as_gen/matplotlib.animation.FuncAnimation.html
ani = animation.FuncAnimation(fig, update, frames=Nt,interval = frame_duration)
#plt.show()
ani.save("panels.mp4")
t_1 = time.time()
print(t_1-t_0)
# -
# ## You have reached the end of the presentation.
| presentation/presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bayes Decision Rule
# *Complete and hand in this completed worksheet (including its outputs and any supporting code outside of the worksheet) with your assignment submission. Please check the pdf file for more details.*
#
# In this exercise you will:
#
# - implement the calculation of **likelihood** of each features given particular class
# - implement the calculation of **posterior** of each class given particular feature
# - implement the calculation of **minimal total risk** of bayes decision rule
# +
# some basic imports
import scipy.io as sio
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
data = sio.loadmat('data.mat')
x1_train, x1_test, x2_train, x2_test = data['x1_train'], data['x1_test'], data['x2_train'], data['x2_test']
all_x = np.concatenate([x1_train, x1_test, x2_train, x2_test], 1)
data_range = [np.min(all_x), np.max(all_x)]
# +
from get_x_distribution import get_x_distribution
train_x = get_x_distribution(x1_train, x2_train, data_range)
test_x = get_x_distribution(x1_test, x2_test, data_range)
# -
# ## Part 1 likelihood
# Your code for this section will be written inside **bayes_decision_rule/likehood.py**.
# +
from likelihood import likelihood
l = likelihood(train_x)
width = 0.35
p1 = plt.bar(np.arange(data_range[0], data_range[1] + 1), l.T[:,0], width)
p2 = plt.bar(np.arange(data_range[0], data_range[1] + 1) + width, l.T[:,1], width)
plt.xlabel('x')
plt.ylabel('$P(x|\omega)$')
plt.legend((p1[0], p2[0]), ('$\omega_1$', '$\omega_2$'))
plt.axis([data_range[0] - 1, data_range[1] + 1, 0, 0.5])
plt.show()
# +
#TODO
#compute the number of all the misclassified x using maximum likelihood decision rule
# begin answer
misclassfied = 0
C, N = test_x.shape
for c in range(C):
# in each class of test data
for n in range(N):
# for each n s.t. x = n
if(l[c][n] < l[1-c][n]):
# misclassified according to ML dicision rule
misclassfied += test_x[c][n]
print(misclassfied/np.sum(test_x))
# end answer
# -
# ## Part 2 posterior
# Your code for this section will be written inside **bayes_decision_rule/posterior.py**.
# +
from posterior import posterior
p = posterior(train_x)
width = 0.35
p1 = plt.bar(np.arange(data_range[0], data_range[1] + 1), p.T[:,0], width)
p2 = plt.bar(np.arange(data_range[0], data_range[1] + 1) + width, p.T[:,1], width)
plt.xlabel('x')
plt.ylabel('$P(\omega|x)$')
plt.legend((p1[0], p2[0]), ('$\omega_1$', '$\omega_2$'))
plt.axis([data_range[0] - 1, data_range[1] + 1, 0, 1.2])
plt.show()
# +
#TODO
#compute the number of all the misclassified x using optimal bayes decision rule
# begin answer
misclassfied = 0
C, N = test_x.shape
for c in range(C):
# in each class of test data
for n in range(N):
# for each n s.t. x = n
if(p[c][n] < p[1-c][n]):
# misclassified according to ML dicision rule
misclassfied += test_x[c][n]
print(misclassfied/np.sum(test_x))
# end answer
# -
# ## Part 3 risk
#
# +
risk = np.array([[0, 1], [2, 0]])
#TODO
#get the minimal risk using optimal bayes decision rule and risk weights
# begin answer
# recalculate prior, likelihood, posterior and p(x) from test data and train data
x = test_x + train_x
C, N = x.shape
prior = np.array([np.sum(x[0,:]), np.sum(x[1,:])])/np.sum(x)
l = likelihood(x)
p = posterior(x)
px = np.zeros(N)
for xi in range(N):
for c in range(C):
px[xi] += l[c][xi]*prior[c]
minRisk = 0
for xi in range(N):
ni, nj = risk.shape
riskList = np.zeros(ni)
for i in range(ni):
for j in range(nj):
riskList[i] += risk[i][j]*p[j][xi]
minRisk += np.min(riskList)*px[xi]
print(minRisk)
# end answer
# -
| hw1/hw1_code/bayes_decision_rule/run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Before your start:
# - Read the README.md file
# - Comment as much as you can and use the resources in the README.md file
# - Happy learning!
# # Challenge 1 - Working with JSON files
#
# Import the pandas library
# +
# Your import here:
# -
# #### After importing pandas, let's find a dataset. In this lesson we will be working with a NASA dataset.
#
# Run the code in the cell below to load the dataset containing information about asteroids that have landed on earth. This piece of code helps us open the URL for the dataset and deocde the data using UTF-8.
# +
# Run this code
from urllib.request import urlopen
import json
response = urlopen("https://data.nasa.gov/resource/y77d-th95.json")
json_data = response.read().decode('utf-8', 'replace')
# -
# In the next cell, load the data in `json_data` and load it into a pandas dataframe. Name the dataframe `nasa`.
# +
# Your code here:
# -
# Now that we have loaded the data, let's examine it using the `head()` function.
# +
# Your code here:
# -
# #### The `value_counts()` function is commonly used in pandas to find the frequency of every value in a column.
#
# In the cell below, use the `value_counts()` function to determine the frequency of all types of asteroid landings by applying the function to the `fall` column.
# +
# Your code here:
# -
# Finally, let's save the dataframe as a json file again. Since we downloaded the file from an online source, the goal of saving the dataframe is to have a local copy. Save the dataframe using the `orient=records` argument and name the file `nasa.json`.
# +
# Your code here:
# -
# # Challenge 2 - Working with CSV and Other Separated Files
#
# csv files are more commonly used as dataframes. In the cell below, load the file from the URL provided using the `read_csv()` function in pandas. Starting version 0.19 of pandas, you can load a csv file into a dataframe directly from a URL without having to load the file first like we did with the JSON URL. The dataset we will be using contains informtaions about NASA shuttles.
#
# In the cell below, we define the column names and the URL of the data. Following this cell, read the tst file to a variable called `shuttle`. Since the file does not contain the column names, you must add them yourself using the column names declared in `cols` using the `names` argument. Additionally, a tst file is space separated, make sure you pass ` sep=' '` to the function.
# +
# Run this code:
cols = ['time', 'rad_flow', 'fpv_close', 'fpv_open', 'high', 'bypass', 'bpv_close', 'bpv_open', 'class']
tst_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/statlog/shuttle/shuttle.tst'
# +
# Your code here:
# -
# Let's verify that this worked by looking at the `head()` function.
# +
# Your code here:
# -
# To make life easier for us, let's turn this dataframe into a comma separated file by saving it using the `to_csv()` function. Save `shuttle` into the file `shuttle.csv` and ensure the file is comma separated and that we are not saving the index column.
# +
# Your code here:
# -
# # Challenge 3 - Working with Excel Files
#
# We can also use pandas to convert excel spreadsheets to dataframes. Let's use the `read_excel()` function. In this case, `astronauts.xls` is in the same folder that contains this notebook. Read this file into a variable called `astronaut`.
#
# Note: Make sure to install the `xlrd` library if it is not yet installed.
# +
# Your code here:
# -
# Use the `head()` function to inspect the dataframe.
# +
# Your code here:
# -
# Use the `value_counts()` function to find the most popular undergraduate major among all astronauts.
# +
# Your code here:
# -
# Due to all the commas present in the cells of this file, let's save it as a tab separated csv file. In the cell below, save `astronaut` as a tab separated file using the `to_csv` function. Call the file `astronaut.csv` and remember to remove the index column.
# +
# Your code here:
# -
# # Bonus Challenge - Fertility Dataset
#
# Visit the following [URL](https://archive.ics.uci.edu/ml/datasets/Fertility) and retrieve the dataset as well as the column headers. Determine the correct separator and read the file into a variable called `fertility`. Examine the dataframe using the `head()` function.
# +
# Your code here:
# -
| module-1/lab-import-export/your-code/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'''No.53
Find the contiguous subarray within an array (containing at least one number) which has the largest sum.
For example, given the array [-2,1,-3,4,-1,2,1,-5,4],
the contiguous subarray [4,-1,2,1] has the largest sum = 6.
'''
def maxSubArray(self, nums):
"""
:type nums: List[int]
:rtype: int
"""
rsum=maxsum=nums[0]
for i in nums[1:]:
rsum=max(i,rsum+i)
maxsum=max(maxsum,rsum)
return maxsum
# +
'''No.121
Say you have an array for which the ith element is the price of a given stock on day i.
If you were only permitted to complete at most one transaction (ie, buy one and sell one share of the stock), design an algorithm to find the maximum profit.
Example 1:
Input: [7, 1, 5, 3, 6, 4]
Output: 5
max. difference = 6-1 = 5 (not 7-1 = 6, as selling price needs to be larger than buying price)
Example 2:
Input: [7, 6, 4, 3, 1]
Output: 0
In this case, no transaction is done, i.e. max profit = 0.
'''
def maxProfit(self, prices):
"""
:type prices: List[int]
:rtype: int
"""
'''
time : O(n)
space: O(1)
'''
if prices==None or len(prices)<2: return 0
mini , money = prices[0] , 0
for moneytoday in prices:
mini=min(mini,moneytoday) #find the minimum in the list
money=max( money , moneytoday - mini) #find the maximum
return money
| 53, 121.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## SLU07 - Advanced Statistics: Learning notebook
# In this notebook we will cover the following:
#
# - Probability Distributions
#
# - Normal
# - Binomial
# - Geometric
# - Exponential
# - Poisson
# - Student's T
#
# - Point estimates
# - Confidence intervals
#
# - Significance tests
# - One-sample T-test
# - Chi-squared goodness of fit test
#
# ## Imports
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# this allows us to see our plots in the notebook:
% matplotlib inline
# -
# A probability distribution is a mathematical function that describes the values (and the respective probabilities) that a random variable can assume.
#
# Now that sounds scary, but this is not your average statistics class! We're not gonna ask you to memorize these formulas. You'll just need to know:
#
# > 1. What distribution you're looking at
# > 2. What functions you can use
#
# To do all of this programatically, we'll use a library called [SciPy](https://www.scipy.org/), which has a really cool statistics module.
import scipy.stats as stats
# ___________
# # Normal Distribution
#
# The normal distribution is a continous probability distribution, and its the most common distribution you'll find.
#
# It describes well a lot of random phenomenons, such as human height.
# The notation is:
#
#
# # $ {\mathcal {N}}(\mu ,\sigma ^{2}) $
#
#
# The parameters are just:
#
#
# ### $ mean = {\mathcal \mu \in \mathbb {R} } $
# ### $ variance = { \sigma ^{2}>0} $
# ### Example
# Let's model the height of men in Portugal.
#
# We know that human height is normally distributed, so our choice of using a normal distribution is solid.
#
# We just need the parameters, which are:
#
# > $\mathcal \mu$ = 174 cm
#
# > $\sigma ^{2}$ = $ 8.2 ^ {2}$ cm
#
#
# So our normal distribution is:
# ## $ {\mathcal {N}}(174, 67.24) $
# Let's generate some 10000 datapoints from this distribution.
# Since we are using a normal distribution, we'll use `stats.norm`.
#
# We'll use its `.rsv` method to generated some data from this dristribution. You can see that here, `loc` is the mean, and `scale` is the standard deviation (which gets squared by the function)
normal_data = stats.norm.rvs(size=10000, loc=174, scale=8.2, random_state=0)
# Let's plot it!
pd.Series(normal_data).plot(kind="hist", bins=50)
# You can see the distictive bell-shape. The height of most men sits around in the center, with the extreme values being rarer.
#
# Now that we have the distribution, we can use some functions to gain insights.
# ## Cumulative distribution function: `.cdf()`
#
# This function tells us the probability that the random variable will assume a value less than the one you provide.
# Hugo is 193 cm tall. What percentage of men are shorter than him?
stats.norm.cdf(x=193, # Cutoff value (quantile) to check
loc=174, # Mean
scale=8.2) # Standard Deviation
# He's taller than almost 99% of men!
# ## Percent point function: `.ppf()`
#
# This function is the inverse of `.cdf()`; Instead of inputing a quantile and reciveing a probablity, we input a probability and recieve a quantile.
# Below which height are 90% of men?
stats.norm.ppf(q=0.9, # Cutoff value (quantile) to check
loc=174, # Mean
scale=8.2) # Standard Deviation
# This means that 90% of men are under 184.5 cm of height.
# ## Probability density function: `.pdf()`
#
# This function gives us the relatively likelihood of the random variable assuming a certain value.
#
# For example, the relative likelihood of a randomly chosen man from this population being 170 cm tall:
stats.norm.pdf(x=170, # Value to check
loc=174, # Distribution start
scale=8.2) # Distribution end
# And now the relative likelihood of a randomdly chosen man from this population being 150 cm tall:
#
stats.norm.pdf(x=150, # Value to check
loc=174, # Distribution start
scale=8.2) # Distribution end
# You can think of these values as the probability density at value: it's more likely for a man to be 170 cm tall as opposed to 150 cm tall.
# ___________
# # Binomial Distribution
#
# The binomial distribution is a discrete probability distribution that models the number of sucesses in a set of independent trials.
#
# It describes random phenomenenon such as the number of heads you'll get, when you flip a coin a number of times.
# The notation is:
#
# # $ B(n, p) $
#
# The parameters are just:
#
# ### $ n ∈ N0 $ - number of trials
#
# ### $ p ∈ [0,1] $ - success probability in each trial
# ### Example
# Let's model the number of heads we get when we flip a coin 10 times. This is a fair coin, so the chance of getting heads at each trial is 50%.
#
# So our parameters are:
#
# > n = 10
#
# > p = 0.5
#
#
# So our binomial distribution is:
# ## $ B(10, 0.5) $
# Let's generate some 10000 datapoints from this distribution.
# This means we'll performing 10000 experiments, in which we flip a coin 10 times.
#
# Since we are using a binomial distribution, we'll use `stats.binom`.
#
# We'll use its `.rsv` method to generated the data. You can see that here, `n` is the number of trials, and `p` is the probability of success in each trial.
binomial_data = stats.binom.rvs(size=10000, n=10, p=0.5, random_state=0)
# Let's plot it!
pd.Series(binomial_data).plot(kind="hist", bins = 50)
# Now, unlike the normal distribution, this is a discrete distribution, meaning that the random variable can only assume discrete integer values. It does however sort of look like a normal distribution, in the sense that it is symmetric, but that changes when you use a `p` different from 0.5.
#
# Let's now toss a biased coin. This a coin that is more likely to land on heads than tails.
biased_coin_data = stats.binom.rvs(size=10000, n=10, p=0.8, random_state=0)
# And let's plot it.
pd.Series(biased_coin_data).plot(kind="hist", bins=50)
# You can see that this biased coin is more likely to get more heads in 10 trials than the fair coin. The distribution "shifted" to the right, so to speak.
#
# Let's now use some functions to gain insights.
# ## Cumulative distribution function: `.cdf()`
#
# This function tells us the probability that the random variable will assume a value less (or equal) to the one you provide.
#
# Let's find out the probability of getting 7 heads in 10 trials, with this biased coin.
#
stats.binom.cdf(k=7, # Probability of k = 7 heads or less
n=10, # In 10 trials
p=0.8) # And success probability 0.8
# If you want to ask the question "what is the probablity of getting at least 7 heads in 10 trails?", you are actually asking "What is the probability of NOT getting 6 or less heads in 10 trials".
#
# We express the "NOT" by subtracting the probability from 1, like this:
1 - stats.binom.cdf(k=6, # Probability of k = 6 heads or less
n=10, # In 10 trials
p=0.8) # And success probability 0.8
# ## Probability mass function: `.pmf()`
# Before, we used `.pdf()` to check the probability density on a certain point of a continuous probability density function. However, the binomial distribution is a discrete probability distribution, so instead we use `.pmf()` to check the proportion of observations at a certain point.
#
# Let's find out the probability of getting __exactly__ 5 heads in 10 trails, on our biased coin.
stats.binom.pmf(k=5, # Probability of k = 5 heads
n=10, # With 10 flips
p=0.8) # And success probability 0.5
# _______
# # Geometric Distribution and Exponential distribution
#
# These distributions are useful when you want to model the time it takes for an event to occur.
#
# The geometric distribution is a discrete distribution, and it's useful for modelling things like number of times you need to flip a coin before you see heads.
#
# The exponential distribution is its continuous analogue, and it's useful for modelling things like the time you need to wait before your bus arrives, knowing that there is a bus every 15 minutes.
#
# While these distributions are useful, we have a lot of ground to cover so we can't explain them here.
#
# Refer to these resources if you later want to learn more about them:
# ________
# # Poisson Distribution
#
# This distribution is useful when you want to model the probability of the number of times an event is likely to occur, within a certain timeframe.
#
# It's useful to model things such as the number of pacients an hospital will recieve within an hour.
#
# While also a very useful distribution, we have a lot to cover, so I leave it up to you to learn more about it if you want.
# __________
# # Point estimates
#
# We need to know 2 important concepts:
#
# > 1. Population
# > 2. Sample
#
# Imagine you're selling a product, and your product is in fact so popular, that you have tens of thousands of customers. Now, you'd like to know more about your customers, like what their average age is.
#
# Now, if we could reach out to every single one of your customers and ask them their age, we could calculate the __Population mean__.
#
# But since we can't do that, we have to settle for asking a sample of costumers their age, and calculating the __Sample mean__.
#
# What this means, is that we can use the __Sample mean__ to estimate the __Population mean__.
#
# Let's give it a try:
np.random.seed(3)
population_ages1 = stats.poisson.rvs(loc=18, mu=35, size=150000)
population_ages2 = stats.poisson.rvs(loc=18, mu=10, size=100000)
population_ages = pd.Series(np.concatenate((population_ages1, population_ages2)))
population_ages.mean()
age_sample = population_ages.sample(500, random_state=0)
age_sample.mean()
# Not too bad, right? With just 500 people, we got a pretty good estimate of the mean population age.
#
# If we wanted to be even more sure, we could take several samples, to check if that value doesn't change widly a lot from sample to sample. You see where I'm getting at? We plot out sampling distribution!
#
# Let's do that then. We'll take a sample of 500, take the mean of that sample, and record that mean. We do that process and bunch of times.
#
# Let's try doing that 20 times.
# +
seed_number = 0
point_estimates = [] # Make empty list to hold point estimates
for x in range(20): # Generate 200 samples
np.random.seed(seed_number)
sample = np.random.choice(a= population_ages, size=500)
point_estimates.append( sample.mean() )
seed_number += 1
pd.DataFrame(point_estimates).hist(bins=30)
# -
# The distribution doesn't seem very evident here. Let's take more 50 samples, and see if it helps.
# +
seed_number = 0
point_estimates = [] # Make empty list to hold point estimates
for x in range(50): # Generate 200 samples
np.random.seed(seed_number)
sample = np.random.choice(a= population_ages, size=500)
point_estimates.append( sample.mean() )
seed_number += 1
pd.DataFrame(point_estimates).hist(bins=30)
# -
# It's starting to look like something even seen before, no? Let's take 1000 samples!
# +
seed_number = 0
point_estimates = [] # Make empty list to hold point estimates
for x in range(1000): # Generate 200 samples
np.random.seed(seed_number)
sample = np.random.choice(a= population_ages, size=500)
point_estimates.append( sample.mean() )
seed_number += 1
pd.DataFrame(point_estimates).hist(bins=30)
# -
# Cool, isn't it? If we take the mean of a lot of samples, the resulting distribution will be a normal distribution! This is one of the most concepts in probability theory, and it's called the Central Limit Theorem. It tells us that distribution of many sample means will be normally distributed, no matter the underlying distribution.
#
# Wait, no matter the underlying distribution? Let's look at the distribution of ages for a sample of our population.
pd.DataFrame(population_ages.sample(500)).hist(bins=58,
range=(17.5,75.5))
# Well, the distribution of ages on our sample doesn't look normally distributed at all. Maybe this is just an anomally?
# Let's now take a look at the distribution of ages for our entire population.
pd.DataFrame(population_ages).hist(bins=58,
range=(17.5,75.5))
# Yup, the distribution of the population is the same; the distribution of the sample mirrors it.
#
# This is what we mean by "distribution of many sample means will be normally distributed, no matter the underlying distribution". And this is an extremely useful concept, because it means we can apply a bunch of statistical techniques that assume that our distribution is normal!
#
# Now, remember that we were trying to estimate the mean age of our population using the mean age of a sample. Let's assume we can only take a single sample of 500.
#
# We're going to get a value for the mean age, sure, but as we've seen before, that value is subject to randomness, and that randomness is described by a normal distribution.
#
# Maybe what we want is not to present a single value for our estimated population mean, but a confidence interval; An interval of values, for which we can say something like: "We are 95% confident that the population mean is between 42 and 44."
#
# The cool thing about our sampling distribution for the mean being a normal distribution, is that it makes it easy to calculate those confidence intervals. Let me show you.
# # Confidence Intervals
# The way we calculate a confidence interval is by taking point estimate (the mean) and then adding and subtracting a margin of error.
#
# The formula for the margin of error is:
#
# # $ Z * \frac{\sigma}{\sqrt{n}} $
#
# The parameters are:
#
# __Z__ - it stands for Z-score, which is the number of standard deviations from the mean that you need to capture the desired confidence level.
#
# >For example, if we wanted a confidence interval of 95%, we could use a Z-score of 2, because roughly 95% of the data is within 2 standard deviations from the mean. But to be more exact, we can use `stats.norm.ppf()` to get the Z-score, by inputing the quantile.
#
# $ {\sigma} $ - standard deviation of the population.
#
# > Uh oh. You might be asking yourself, how are we supposed to know the standard deviation of the population, if all we have access is a sample? We'll see ahead a stategy to deal with this.
#
# __n__ - the number of samples.
#
# Since we don't have access to the standard deviation of the population, we can use the standard deviation of the sample instead. But by doing this, we are introducing a source of error; To compensate for it, instead of using the Z-score, we'll use something called the T-score.
#
# The T-score comes from a special distribution, called the Student's T-distribution. It resembles the normal distribution, except it gets wider if the sample size is low, and as the sample size increases, it becomes equal to the Normal distribution.
# The T-distribution needs a parameter called the `degrees of freedom`, which is just the sample size minus 1.
#
# Let's see:
# 
# As you can see, as we increase our sample size, the T-distribution gets closer to the Normal distribution.
#
# So we end up with:
#
# # $ T * \frac{\sigma}{\sqrt{n}} $
#
# In which T is the T-score, $\sigma$ is the standard deviation of the sample, and n is the number of samples.
#
# Let's put it all together, and calculate the 95% confidence interval for the mean age of the population!
# First we get the mean age in the sample:
mean_sample_age = age_sample.mean()
# Now we get the T-score. Since we want a 95% confidence interval, that means our significance level ($\alpha$) is 0.05.
#
# And since the distribution has two tails, we need to do:
#
# 1 - 0.95 = 0.05
#
# 0.05 / 2 = 0.025
#
# 1 - 0.025 = 0.975
#
# 0.975 is then the quantile we want.
#
# `df`, aka degrees of freedom, is 499 because it's the sample size minus 1.
t_critical = stats.t.ppf(q = 0.975, df=499)
# Now we need $\sigma$, which is the standard deviation for the sample:
std_sample_age = age_sample.std()
# And $\sqrt{n}$, which is just the square root for the number of samples.
sqrt_of_n_samples = np.sqrt(500)
# Putting it all together:
# +
error_margin = t_critical * std_sample_age / sqrt_of_n_samples
confidence_interval = (mean_sample_age - error_margin,
mean_sample_age + error_margin)
print(confidence_interval)
# -
# Tada! Now, while it was important to understand how a confidence interval is calculated, for convinience, we can do it all using `t.interval()`!
stats.t.interval(alpha = 0.95, # Confidence level
df= 499, # Degrees of freedom
loc = mean_sample_age, # Sample mean
scale = std_sample_age / sqrt_of_n_samples) # Standard deviation of the sample divided by the square root of the number of samples
# ___
# # Hypotesis testing
# Hypthesis testing is based on making a statement, the "null hypothesis", that nothing interesting is going on between the variables you're studying. Then, you at the evidence, and ask yourself "Does the evidence support this idea that nothing interesting is going on?".
#
# For example, imagine you take the data you have on your clients, `age_sample`. For the purposes of this, we're going to consider this as our population.
#
# Now, you're selling your product on a given day, and you got 15 new clients. You ask yourself: "Is the mean age of the costumers who bought my product today different from the mean age of my typical costumers?"
#
# In other words, you want to know if the possible difference in mean age is so great, that it would be very unlikely for it to happen by pure chance. A difference so significant, that makes you think some other factor may be at hand.
#
# When the difference is so great that it is very unlikely for it to happen by pure chance, you reject the "null hypothesis" (that nothing interesting is going on) in favor of the "alternative hypothesis" (that something interesting is going on). In our case, it could be that on this particular day there was a school holiday, causing a bunch of kids to come to your store.
#
# Now, to define what we mean by "very unlikely for it to happen by pure chance", we have to define what very unlikely is. I won't go very deep into the pitfalls associated with defining a confidence level, but a very common confidence level is $\alpha$ = 0.05.
#
# When doing a significance test, there two possible types of error:
#
# __Type I error__
# > You reject the null hypothesis in favor of the alternative hypothesis, but the null hypothesis was actually true. Also know as a false positive. With a $\alpha$ = 0.05 significance level, you expect this type of error to happen 5% of the time. The higher the $\alpha$, the less likely you are to make this error.
#
#
# __Type II error__
# >You do not reject the null hypothesis in favor of the alternative hypothesis, but the alternative hypothesis was actually true. Also know as a false negative. The lower the $\alpha$, the less likely you are to make this error.
#
# Now, what we want to do is to check if the mean age from our new sample differs from the mean age of our population. for that, we're going to use a __One Sample T-Test__. For that, we'll use `stats.ttest_1samp()`.
# # One Sample T-Test
#
# Let's first check the mean age of the costumers we have on record:
# +
mean_population_age = age_sample.mean() # Remember, right now we're considering the data we have on 500 costumers as our population
mean_population_age
# -
# Let's now check the mean age of the clients we had today:
# +
new_client_ages = np.array([15, 13, 14, 54, 16, 12, 10, 16, 14, 12, 10, 13, 60, 42, 11])
new_client_ages.mean()
# -
# Alright, the mean age seems pretty different; But how do we know if this difference is significant? In other words, how do we know if a result as extreme as this is unlikely to happen due to chance?
#
# For that, we'll use the One Sample T-Test, which compares the mean of a new sample with the mean of the population, and tests if the difference is significant.
stats.ttest_1samp(a= new_client_ages, # New client ages
popmean= mean_population_age) # Population mean
# The test returned a p-value of 0.00016374485160662263;
#
# Since this value is lower than the $\alpha$ = 0.05 confidence level we defined, we reject the null hypothesis; We say that the evidence supports the alternative hypothesis; That some other factor is at play, because it's unlikely to get a result as extreme as this by pure chance.
#
# There are types of T-tests, adequate for other situations, which we can't cover here. You can find them [here](https://hamelg.blogspot.pt/2015/11/python-for-data-analysis-part-24.html).
#
# Now what if you want to investigate a categorical variable? You're going to need another type of test, called a Chi-Squared Test.
# # Chi-Squared Goodness-Of-Fit Test
# The Chi-Squared goodness-of-fit test allows us to test if a sample of categorical data matches an expected distribution.
#
# For example, let's say we have 4 different products: A, B, C.
#
# We expect A to represent on average 0.2 of our sales, B on average 0.5, and C on average 0.3.
#
# Let's say today our sales were:
# +
sales_df = pd.DataFrame({
'A': 37,
'B': 110,
'C': 50
}, index=['sales'])
sales_df
# -
# A total of:
sales_df.loc['sales'].sum()
# The expected counts are:
print('Expected count for 197 sales: ')
print('Product A: ' + str(197 * 0.2))
print('Product B: ' + str(197 * 0.5))
print('Product C: ' + str(197 * 0.3))
# We want to know if the observed counts are in line with the expected counts (the null hypothesis), or if the observed count differ from the expected counts (alternative hypothesis).
#
# We'll define $\alpha$ = 0.05 as our significance level.
# To do the test itself, we use `stats.chisquare()`.
stats.chisquare(f_obs= [37, 110, 50], # Array of observed counts
f_exp= [39.4, 98.5, 59.1]) # Array of expected counts
# As you can see, the p-value is much higher than our significance level, so we do not reject the null hypothesis; the observed counts are in line with the expected counts.
# # Wrap up
#
# That was a lot to cover! There are all sorts of distributions and statistical tests, but we can use them easily using python libraries such as [Scipy.stats](https://docs.scipy.org/doc/scipy/reference/stats.html).
#
# To learn more about using statistics in practice, I recommend you look at this brilliant series of blog posts on [data analysis in python](https://hamelg.blogspot.pt/2015/12/python-for-data-analysis-index.html) (you'll see that a lot of the content in this learning unit was inspired / borrowed from these blogposts!)
#
# To learn more about statistics on a theoretical standpoint, I recommend [this free book](https://www.openintro.org/stat/textbook.php?stat_book=os) which will teach you all the foundations you need in a motivating way!
| units/SLU07_Advanced_Statistics/Learning notebook - SLU07 (Advanced Statistics).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mounesi/pa/blob/master/notebooks/random_forest_Irvington_validation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="SAf7JfNWeQ7l"
# Read Original data from this
# [paper](https://www.irvingtonny.gov/DocumentCenter/View/8036/Public-Works-Roadway-Pavement-Report?bidId=)
# + colab_type="code" id="rmVo9_BIdMLu" colab={}
# cp drive/My\ Drive/data/pa/dataIrvingtonVillage/ListofRoadwaysIrvington_plot.csv ./data.csv
# + colab_type="code" id="L4kOsAv5eQdd" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="a0ea3efe-889c-41a3-fe85-d5c6bf98c203"
import pandas as pd
df = pd.read_csv('data.csv', index_col=0)
df.head()
# + [markdown] colab_type="text" id="9fQkfnlp8_L6"
# Clean Data: read nulls, drop extra rows, fix formating
# + colab_type="code" id="8sba3ujk8unN" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="5d88d455-240e-4cea-e5b9-16ec1d64126e"
# Count the NaN under an entire DataFrame
df.isnull().sum()
# + colab_type="code" id="7LPjvm438upa" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="a6f007f8-d4f1-4ff0-84cf-562892fafd01"
#Drop the Date RESURFACED and NOTES and num
df.drop(columns=['DATE RESURFACED', 'NOTES'], inplace=True)
# droping rows with Nan in the geo_loc
df.dropna(subset=['geo_loc'], inplace=True)
# Convert Year from string to numeric
df["Year RESURFACED"].replace(',','', regex=True, inplace=True)
df[["Year RESURFACED"]] = df[["Year RESURFACED"]].apply(pd.to_numeric)
# replace the missing columns with the average
column_means = df.mean()
column_means['Year RESURFACED'] = round(column_means['Year RESURFACED'],0)
column_means['Pavement Rate Overall'] = round(column_means['Pavement Rate Overall'],0)
df = df.fillna(column_means)
# Check for NaN under a single DataFrame column:
df.isnull().any()
# + [markdown] colab_type="text" id="MlHeIIRuVbBc"
# visualize of the data
# + colab_type="code" id="USAjenrPVfFr" colab={"base_uri": "https://localhost:8080/", "height": 914} outputId="f7269ad7-8932-4dfd-991c-caac6b2df5cf"
# plot Year_RESURFACED_vs_Pavement_Rate_Total
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
sns.set(color_codes=True)
g = sns.jointplot(x="Year RESURFACED", y="Pavement Rate Total", data=df, kind="kde", color="magenta")
g.plot_joint(plt.scatter, c="w", s=30, linewidth=2, marker="+")
g.ax_joint.collections[0].set_alpha(0)
g.set_axis_labels("$Year\ Resrufaced$", "$Pavement\ Score$");
g.savefig("Year_RESURFACED_vs_Pavement_Rate_Total.png")
h = sns.jointplot(x="AVERAGE WIDTH (FT)", y="Pavement Rate Total", data=df, kind="kde", color="c")
h.plot_joint(plt.scatter, c="b", s=30, linewidth=2, marker="+")
h.ax_joint.collections[0].set_alpha(0)
h.set_axis_labels("$AVERAGE\ WIDTH\ (FT)$", "$Pavement\ Score$");
h.savefig("AVERAGE_WIDTH_(FT)_vs_Pavement_Rate_Total.png")
# + [markdown] colab_type="text" id="nYkzNvtUDhdN"
# Geo 2
# + colab_type="code" id="9DmQiiMFDgxG" colab={"base_uri": "https://localhost:8080/", "height": 547} outputId="8161f9a4-eb8f-4aac-aba2-edb95cb02586"
import folium
m = folium.Map(location=[41.039262, -73.866576], zoom_start=10)
m
# + [markdown] colab_type="text" id="JoU8Y9ulpCMR"
# Get long lat data
# + colab_type="code" id="vA7wuuN1uw-t" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c51d48cc-0388-4942-aaef-c31c3d3be22b"
# !pip install geopandas
# !pip install contextily
# + colab_type="code" id="CnAPtW4NrPDx" colab={}
df = pd.concat([df, df['geo_loc'].str.split(', ', expand=True).astype(float)], axis=1)
# + colab_type="code" id="CBiB9Y_ku_eh" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="5bc5720b-603e-41d4-c497-ee95f0a2cd61"
df.rename(columns={0:'Latitude' , 1:'Longitude'},inplace=True)
df.head()
# + colab_type="code" id="7l81LOa77EKc" colab={"base_uri": "https://localhost:8080/", "height": 631} outputId="abdf79a4-22fd-40ac-e441-947d31b81f78"
import matplotlib.pyplot as plt
from shapely.geometry import Point
import geopandas as gpd
import pandas as pd
import contextily as ctx
df['coords'] = list(zip(df.Longitude, df.Latitude))
geo_df = gpd.GeoDataFrame(
df, crs ={'init': 'epsg:4326'},
geometry = df['coords'].apply(Point)
).to_crs(epsg=3857)
# ... and make the plot
ax = geo_df.plot(
figsize= (10, 10),
alpha = 1
)
ctx.add_basemap(ax, zoom=15)
ax.set_axis_off()
plt.title('Irvington village')
plt.savefig("Irvington_Village")
plt.show()
# + colab_type="code" id="zoEwViqRLUUw" colab={}
#https://residentmario.github.io/geoplot/quickstart/quickstart.html
#gplt.pointplot(df, projection=gcrs.AlbersEqualArea(), hue='DIP', legend=True)
# + colab_type="code" id="I2RHwQqlMqxt" colab={}
# + [markdown] colab_type="text" id="UGPbkL2LO0ri"
# # **Inferecing**
# + colab_type="code" id="z74PpNEsOy-7" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="172e7318-0b84-4789-80b1-29762d7f5697"
# Changin tensorflow version to 1.15
# %tensorflow_version 1.x
# pycocotools must be installed
# !pip install pycocotools
#clone github
# !git clone --depth 1 https://github.com/tensorflow/models
# Define the research folder
import os
RESEARCH_DIR = os.path.join('/','content', 'models', 'research')
# + colab_type="code" id="9ZE_XvWYOzC8" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="88d1002d-99ac-4c5d-e7a4-df342e9f0c05"
# cd {RESEARCH_DIR}
# + colab_type="code" id="z0ypnvzyOzFe" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="7f3a5039-c15c-4039-b3c7-85d6a1786794"
# !pip install tf_slim
# + colab_type="code" id="o8_F5LMMOzBP" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="35357650-1d1d-47b9-981c-d3a9dcce8842"
#compiling the proto buffers (not important to understand for this project but you can learn more about them here: https://developers.google.com/protocol-buffers/)
# !protoc object_detection/protos/*.proto --python_out=.
# exports the PYTHONPATH environment variable with the reasearch and slim folders' paths
os.environ['PYTHONPATH'] += f':/content/models/research/:/content/models/research/slim/'
# testing the model builder
# !python object_detection/builders/model_builder_test.py
# + [markdown] colab_type="text" id="w6DlKzaqRCQG"
#
# + colab_type="code" id="r4kro6bePxn0" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7dcaf38f-a599-467b-b108-cbc97eb218ec"
# cd /content
# + colab_type="code" id="471xLY86Qe03" colab={}
#DATA
# Read it from Goolge street view
# + colab_type="code" id="u9MSi_s9Qe3e" colab={}
# TRAINED_MODEL
import os
MODEL_FOLDER_NAME_GDRIVE = 'distress_ssd_mobilenet_v2_batch_Size_16_epochs_200000_valid_8' # will change for different models
MODEL_WEIGHT_GDIR = os.path.join('/', 'content', 'drive', 'My\ Drive', 'data', 'pa', 'MODEL_ARCHIVE', MODEL_FOLDER_NAME_GDRIVE)
MODEL_WEIGHT_DIR = os.path.join('/', 'content', MODEL_FOLDER_NAME_GDRIVE )
# ! cp -rR {MODEL_WEIGHT_GDIR} {MODEL_WEIGHT_DIR}
# + colab_type="code" id="BJziSr0yQe74" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="623127a8-b8b9-4f14-c8fc-be4b568d2497"
MODEL_WEIGHT_DIR
# + colab_type="code" id="WZGnZuFmQe6y" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="d7c68be5-42b4-45b8-a673-c9b40cf2e43c"
FINE_TUNED_DIR = os.path.join(MODEL_WEIGHT_DIR, 'fine_tuned_model')
CONFIG_PATH = os.path.join(FINE_TUNED_DIR,'pipeline.config')
# #!cat {CONFIG_PATH}
#check the modified config which will be used
# !cat {CONFIG_PATH} | grep num_step
# !cat {CONFIG_PATH} | grep num_classes
# !cat {CONFIG_PATH} | grep input_path
# !cat {CONFIG_PATH} | grep abel_map_path
# !cat {CONFIG_PATH} | grep weight:
# !cat {CONFIG_PATH} | grep num_examples
# !cat {CONFIG_PATH} | grep batch_size
# !cat {CONFIG_PATH} | grep " type:"
# + colab_type="code" id="RtCaPUDBSpFU" colab={}
# OUTPUT DIR FOR MULITPLE FILE
RESULT_OUTPUT_DIR_MULTIPLE = os.path.join('/', 'content', 'result_output_dir_multiple' )
# ! mkdir {RESULT_OUTPUT_DIR_MULTIPLE}
# + colab_type="code" id="c3y76_AWSpJ1" colab={}
# path to the frozen graph:
PATH_TO_FROZEN_GRAPH = os.path.join(FINE_TUNED_DIR, 'frozen_inference_graph.pb')
# path to the label map
PATH_TO_LABEL_MAP = os.path.join(FINE_TUNED_DIR, 'label_map.pbtxt')
# + colab_type="code" id="ivEDzIAWTVAk" colab={}
# path to a single image
frame_index = 11
#IMAGE_NAME = f'image ({frame_index}).png'
#PATH_TO_SINGLE_IMAGE = os.path.join(INPUT_TEST_DIR, IMAGE_NAME)
# To save
OUTPUT_NAME = f'single_output ({frame_index}).png'
#PATH_SAVE_SINGLE_IMAGE = os.path.join(RESULT_OUTPUT_DIR , OUTPUT_NAME)
# + colab_type="code" id="vTd426x3TVFJ" colab={}
# + colab_type="code" id="1NypGNyEZ5Om" colab={}
# rotatory angle download
import requests
GOOGLE_KEY = "<KEY>"
IMG_WIDTH, IMG_HEIGHT = (600, 600)
ANGLE_SHIFT = 4 # 360 / 90
PITCH_ANG = -60 # should stay constant
"""
geo1 = {
"addr" : "W Post Rd, White Plains, NY",
"geoloc": (41.0176417,-73.7772902)
}
geo2 = {
"addr" : "W Post Rd, White Plains, NY",
"geoloc": (41.020012, -73.776141)
}
geo3 = {
"addr" : "W Post Rd, White Plains, NY",
"geoloc": (41.022670, -73.774865)
}
geo4 = {
"addr" : "W Post Rd, White Plains, NY",
"geoloc": (41.024367, -73.772952)
}
geo5 = {
"addr" : "E Post Rd, White Plains, NY",
"geoloc": (41.026623, -73.769262)
}
geo6 = {
"addr" : "E Post Rd, White Plains, NY",
"geoloc": (41.029930, -73.763814)
}
geo7 = {
"addr" : "E Post Rd, White Plains, NY",
"geoloc": (41.029816, -73.764083)
}
geo7 = {
"addr" : "E Post Rd, White Plains, NY",
"geoloc": (41.031143, -73.762136)
}
geo8 = {
"addr" : "N Broadway, White Plains, NY",
"geoloc": (41.034627, -73.763780)
}
geo9 = {
"addr" : "N Broadway, White Plains, NY",
"geoloc": (41.040754, -73.767644)
}
geo10 = {
"addr" : "N Broadway, White Plains, NY",
"geoloc": (41.043902, -73.768910)
}
geo_list = [geo1, geo2, geo3, geo4, geo5,
geo6, geo7, geo8, geo9, geo10]
"""
def get_image(loc_lat, loc_long , heading):
image_url = f"https://maps.googleapis.com/maps/api/streetview?size={IMG_WIDTH}x{IMG_HEIGHT}&location={loc_lat},{loc_long}&heading={heading}&pitch={PITCH_ANG}&key={GOOGLE_KEY}"
img_data = requests.get(image_url).content
return img_data
# + colab_type="code" id="kxmImTX5eKrV" colab={}
# !mkdir street_dataset
# !mkdir street_dataset/mobilenet_test
# + colab_type="code" id="tOq6cV78tJr6" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="1617a2a5-080e-4231-ef7d-f102bd58f22d"
df.head()
# + colab_type="code" id="N_ACKlBDum-W" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ec21d5f-8a6c-44d3-87a1-e45a2ffca91a"
loc_long = len(df['Longitude'].iloc[:])
loc_long
# + colab_type="code" id="4qhjeYIda4cm" colab={}
# for single image
c = 0 # geo list number
i_ang = 0 # angle shift
#loc_lat, loc_long = geo_list[c]['geoloc']
loc_lat = df['Latitude'].iloc[c]
loc_long = df['Longitude'].iloc[c]
heading = 360 / ANGLE_SHIFT * i_ang # to cover 360 degree
img_data = get_image(loc_lat, loc_long , heading)
image_name = f'image{4*c +i_ang}.jpg'
#image = Image.frombytes('RGBA', (128,128), img_data, 'raw')
with open(f'./street_dataset/{image_name}', 'wb') as handler:
handler.write(img_data)
# + id="V0yt9vsnb1uo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fbd44e64-ee90-413d-9d41-5afbd49add85"
# cd ../../
# + id="p0OC6mSWadmU" colab_type="code" colab={}
# for all images
c = 0 # geo list number
i_ang = 0 # angle shift
#loc_lat, loc_long = geo_list[c]['geoloc']
for c in range(len(df['Longitude'].iloc[:])):
for i_ang in range(4):
loc_lat = df['Latitude'].iloc[c]
loc_long = df['Longitude'].iloc[c]
heading = 360 / ANGLE_SHIFT * i_ang # to cover 360 degree
img_data = get_image(loc_lat, loc_long , heading)
image_name = f'image{4*c +i_ang}.jpg'
#image = Image.frombytes('RGBA', (128,128), img_data, 'raw')
with open(f'./street_dataset/{image_name}', 'wb') as handler:
handler.write(img_data)
# + id="jVStT0u6YzG2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0f08e949-79a2-4e0b-8749-684c2bbcd4bd"
image_name
# + id="oVpt1MEHT_Ez" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4371213f-4b30-47ef-bd64-908709f6d9fd"
# cd /content/models/research/
# + colab_type="code" id="fHYnPbj-a5Kr" colab={}
# for multiple images
import time
import tensorflow as tf
from object_detection.utils import label_map_util
import numpy as np
import cv2
import tensorflow as tf
from object_detection.utils import visualization_utils as vis_util
NUM_CLASSES = 1
THRESH= 0.1
# Image Frame Size (480x480 default)
IMAGE_WIDTH = 600
IMAGE_HEIGHT = 600
TIMER = 1
ITIMER = 0
PATH_TO_LABELS = PATH_TO_LABEL_MAP
# Read Labels
LABEL_MAP = label_map_util.load_labelmap(PATH_TO_LABELS)
CATEGORIES = label_map_util.convert_label_map_to_categories(LABEL_MAP,
max_num_classes=
NUM_CLASSES,
use_display_name
=True)
CATEGORY_INDEX = label_map_util.create_category_index(CATEGORIES)
# + id="TEOMmgdmdEm5" colab_type="code" colab={}
def read_single_image(i, test_dir_path):
'''
read the image data
'''
path = test_dir_path+"/image{}.jpg".format(i)
i+=1
print(path)
img = cv2.imread(path) # reading the img
#get image shape
#width, height, ch = img.shape
#select suqare part of image if needed
#if width != height:
# img = img[0:959 , 0:959]
#resize the image if needed
#if width>IMAGE_WIDTH or height>IMAGE_HEIGHT:
# img = cv2.resize(img, (IMAGE_WIDTH, IMAGE_HEIGHT)) # Resize image to see all
return img
# + id="gUpkhk3adWBS" colab_type="code" colab={}
TEST_DIR_PATH = '/content/street_dataset'
# + id="K2QyGUredsOa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="4a9d3fd2-012b-4afb-ac93-5270af5f17f0"
img_sample = read_single_image(1, TEST_DIR_PATH)
cv2.imshow(img_sample)
# + id="GlLK7hzFdXIO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e6eb4799-f00a-4f72-fb03-da63f8aa6935"
pwd
# + colab_type="code" id="HC3u4P-djdk5" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c3d1d7d0-8257-4344-d619-c46a0fab6053"
# setup
def setupgraph(path_to_model):
'''
import unserialized graph file
'''
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(path_to_model, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
return detection_graph
########################################################################################################################################################
# The main code
def run_inference(image, sess):
'''
run interence
'''
# Get handles to input and output tensors
ops = tf.get_default_graph().get_operations()
all_tensor_names = {output.name for op in ops for output in op.outputs}
# make dictonary of tensor names
tensor_dict = {}
for key in ['num_detections', 'detection_boxes', 'detection_scores',
'detection_classes', 'detection_masks']:
tensor_name = key + ':0'
if tensor_name in all_tensor_names:
tensor_dict[key] = tf.get_default_graph().get_tensor_by_name(tensor_name)
image_tensor = tf.get_default_graph().get_tensor_by_name('image_tensor:0')
# Run inference
output_dict = sess.run(tensor_dict, feed_dict={image_tensor: np.expand_dims(image, 0)})
# all outputs are float32 numpy arrays, so convert types as appropriate
output_dict['num_detections'] = int(output_dict['num_detections'][0]) #num detections not used in mobile model
output_dict['detection_classes'] = output_dict['detection_classes'][0].astype(np.uint8)
output_dict['detection_boxes'] = output_dict['detection_boxes'][0]
output_dict['detection_scores'] = output_dict['detection_scores'][0]
if 'detection_masks' in output_dict:
output_dict['detection_masks'] = output_dict['detection_masks'][0]
return output_dict
########################################################################################################################################################
def read_video(desired_frame, cap):
'''
Read the video
'''
img_resized = 0
if cap.isOpened():
ret, frame = cap.read()
img1024 = frame[896:1920 , 26:1050]
img_resized = cv2.resize(img1024, (IMAGE_WIDTH, IMAGE_HEIGHT)) # Resize image to see all
if False:
#convert to gray and then present it as RGB (to test if gray hurts performance)
img_gray = cv2.cvtColor(img_resized, cv2.COLOR_BGR2GRAY)
img_resized = cv2.cvtColor(img_gray,cv2.COLOR_GRAY2RGB)
else:
print('video file not open')
return img_resized
########################################################################################################################################################
def read_single_image(i, test_dir_path):
'''
read the image data
'''
path = test_dir_path+"/image{}.jpg".format(i)
i+=1
print(path)
img = cv2.imread(path) # reading the img
#get image shape
#width, height, ch = img.shape
#select suqare part of image if needed
#if width != height:
# img = img[0:959 , 0:959]
#resize the image if needed
#if width>IMAGE_WIDTH or height>IMAGE_HEIGHT:
# img = cv2.resize(img, (IMAGE_WIDTH, IMAGE_HEIGHT)) # Resize image to see all
return img
########################################################################################################################################################
def visulization(image_np, output_dict, category_index, out, i, test_dir_path, thresh, save):
'''
Code To Generate Images and Videos With Results Drawn On
'''
print(f'test dir path is: {test_dir_path}')
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
output_dict['detection_boxes'],
output_dict['detection_classes'],
output_dict['detection_scores'],
category_index,
max_boxes_to_draw=20,
min_score_thresh=thresh,
instance_masks=output_dict.get('detection_masks'),
use_normalized_coordinates=True,
line_thickness=2)
#For DEBUG SAVE EACH FRAME with top score in name
if True:
test_score = str(int(100*output_dict['detection_scores'][0]))
# save image as jpg
save_image_paths = test_dir_path+'/mobilenet_test/testCrackOut{}'.format(i)+'_Score_'+test_score+'.jpg'
print(f'frame is saved at save_image_paths: {save_image_paths}')
cv2.imwrite(save_image_paths, image_np)
return image_np
if save == 1:
#for presentation uses, save frames to video
print('saving video')
out.write(image_np)
########################################################################################################################################################
def get_videos():
'''
read video and creat output video holder
'''
# get video
cap = cv2.VideoCapture('./Trailing3.MKV')
# setup output
out = cv2.VideoWriter('crack_video.avi',cv2.VideoWriter_fourcc('M','J','P','G'), 15, (IMAGE_WIDTH, IMAGE_HEIGHT))
return cap, out
########################################################################################################################################################
'''
Main Code To Run
'''
PATH_TO_MODEL = PATH_TO_FROZEN_GRAPH
detection_graph = setupgraph(PATH_TO_MODEL)
#TEST_DIR_PATH = './street_dataset'
TEST_DIR_PATH = '/content/street_dataset'
pa_rate = []
cap, out = get_videos()
# get graph and start session
with detection_graph.as_default():
with tf.Session() as sess:
# use session to run loop over all images
startoverall = time.time()
frames = ANGLE_SHIFT * len(df['Longitude'])
for i, image_frame in enumerate(range(frames)):
# Load Input Data (video or image)... THIS IS A SLOW STEP
#image_np = read_video(image_frame, cap)
image_np = read_single_image(i, TEST_DIR_PATH)
# inference and check time
start = time.time()
output_dict = run_inference(image_np, sess)
ttime = time.time()-start
za = (output_dict['detection_scores'] > THRESH).sum()
pa_rate.append(za)
if ITIMER:
print('Finiahed Image: '+str(i)+'... Total time (sec): '+str(round(ttime,3))+'... FPS: '+str(round(1/ttime,3)))
if True:
visulization(image_np, output_dict, CATEGORY_INDEX, out, i, TEST_DIR_PATH, THRESH, save=0)
if TIMER:
#measure time completed
endoverall = time.time()-startoverall
print('Main Loop Average FPS: '+str(round(frames/endoverall,3)))
# clean up
cap.release()
out.release()
# + colab_type="code" id="xzMBtv0UzYvq" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="57d598d1-057d-406f-b249-4d9b4644437f"
len(pa_rate)
# + colab_type="code" id="5rA1jdc3Yu2v" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e360ffea-728a-4838-9a24-01238e5a5f5a"
# cd {RESEARCH_DIR}
# + colab_type="code" id="npuTbLl1yiS3" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6494f4b5-8edb-42b9-af82-f9dcc9d0c123"
len(pa_rate)
# + colab_type="code" id="JMHZ2w-EyiYj" colab={}
pa_local= []
pa_count = []
c_num = 15
coef = 0.1
for i in range(len(df['Longitude'])):
pa_sum = 0
for j in range(ANGLE_SHIFT):
pa_sum += pa_rate[4*i + j]
pa_count.append(pa_sum)
pa_temp = 9 * np.exp(-pa_sum*coef)
pa_local.append(pa_temp)
# + id="V1fuHGvqfRtM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b0fb8bb8-8405-434f-bbc5-67ebc9804493"
len(pa_count)
# + colab_type="code" id="5XWDOoSW-8Br" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e46d927a-77d2-4630-ecd9-84bf3f1dc561"
pa_local
# + id="L40FLlSXf0WG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="4eab1cfe-70bf-4b11-96fb-f4e647381a9d"
df.head()
# + colab_type="code" id="SxBnNAC_yiV5" colab={"base_uri": "https://localhost:8080/", "height": 844} outputId="668c0277-a415-4a06-d836-3018894576b6"
# Using DataFrame.insert() to add a column
df.insert(2, "machine_vision_prediction", pa_local, True)
df.insert(2, "crack_count", pa_count, True)
# Observe the result
df
# + colab_type="code" id="g_cdpkJuPcC4" colab={"base_uri": "https://localhost:8080/", "height": 441} outputId="4a1822a9-7330-48dc-bf6f-8a7003d0b9f4"
# plot Year_RESURFACED_vs_Pavement_Rate_Total
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
sns.set(color_codes=True)
g = sns.jointplot(x="machine_vision_prediction", y="Pavement Rate Total", data=df, kind="kde", color="c")
g.plot_joint(plt.scatter, c="w", s=30, linewidth=2, marker="+")
g.ax_joint.collections[0].set_alpha(0)
g.set_axis_labels("$machine vision prediction$", "$Pavement\ Score$");
g.savefig("Machine Vision vs_Pavement_Rate_Total.png")
# + colab_type="code" id="xnuk7V92Dx9K" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="68ed2f6b-0229-466a-ef68-4bdd994ea1a4"
# plot Year_RESURFACED_vs_Pavement_Rate_Total
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from scipy import stats
sns.set(color_codes=True)
g = sns.jointplot(x="Year RESURFACED", y="machine_vision_prediction", data=df, kind="kde", color="c")
g.plot_joint(plt.scatter, c="w", s=30, linewidth=2, marker="+")
g.ax_joint.collections[0].set_alpha(0)
g.set_axis_labels("$Year RESURFACED$", "$machine_vision_prediction$");
g.savefig("Year\ RESURFACED vs machine vision prediction.png")
# + colab_type="code" id="CUERJ7n-PTJc" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3a21b154-cf04-4c5c-b335-8107c0d9d30e"
((df['machine_vision_prediction'] - df['Pavement Rate Total']) ** 2).mean() ** .5
# + id="CL_2zWGhgfx_" colab_type="code" colab={}
# + id="r5v9R41MggIY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="d9344bac-32f9-448e-feab-2fcfc59895c6"
df.head()
# + [markdown] id="v2DbO4M8ghWB" colab_type="text"
# # random farest
# + id="OlERS4_LhiDT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="94eed88c-0baf-4700-fec7-3138025de441"
# Use numpy to convert to arrays
import numpy as np
# Labels are the values we want to predict
labels = np.array(df['Pavement Rate Total'])
# Remove the labels from the features
# axis 1 refers to the columns
#df2= df.drop('Pavement Rate Total','ROAD NAME', 'geo_loc', 'AREA (SF)', 'AREA (SY)', 'Pavement Rate Overall', 'Latitude', 'Longitude', 'coords', 'geometry' , axis = 1)
#df2= df[['crack_count','LENGTH\n(FT)', 'AVERAGE WIDTH (FT)', 'Year RESURFACED']]
df2= df[['crack_count','Year RESURFACED']]
df2.head()
# + id="QkTBPfhljpyS" colab_type="code" colab={}
# Saving feature names for later use
feature_list = list(df2.columns)
# Convert to numpy array
df2 = np.array(df2)
# + id="iQhrE3ZWhiVt" colab_type="code" colab={}
# Using Skicit-learn to split data into training and testing sets
from sklearn.model_selection import train_test_split
# Split the data into training and testing sets
train_features, test_features, train_labels, test_labels = train_test_split(df2, labels, test_size = 0.25, random_state = 42)
# + id="-cHbeeZEhh_0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="be658fa2-22f8-461d-8501-29498e909525"
print('Training Features Shape:', train_features.shape)
print('Training Labels Shape:', train_labels.shape)
print('Testing Features Shape:', test_features.shape)
print('Testing Labels Shape:', test_labels.shape)
# + id="uwJAsK00lkM0" colab_type="code" colab={}
# The baseline predictions are the historical averages
#baseline_preds = test_features[:, feature_list.index('average')]
# Baseline errors, and display average baseline error
#baseline_errors = abs(baseline_preds - test_labels)
#print('Average baseline error: ', round(np.mean(baseline_errors), 2))
# + id="EvyupMvdlySZ" colab_type="code" colab={}
# Import the model we are using
from sklearn.ensemble import RandomForestRegressor
# Instantiate model with 1000 decision trees
rf = RandomForestRegressor(n_estimators = 1000, random_state = 42)
# Train the model on training data
rf.fit(train_features, train_labels);
# + id="SeonDtWNl4bu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a2bcdf95-b3af-4ae3-949d-b404edad6c5e"
# Use the forest's predict method on the test data
predictions = rf.predict(test_features)
# Calculate the absolute errors
errors = abs(predictions - test_labels)
# Print out the mean absolute error (mae)
print('Mean Absolute Error:', round(np.mean(errors), 2), 'rate.')
# + id="SGSaObVdmT2d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="796db821-bb5c-4c2d-a6b0-ed704ce9f73c"
# Calculate mean absolute percentage error (MAPE)
mape = 100 * (errors / test_labels)
# Calculate and display accuracy
accuracy = 100 - np.mean(mape)
print('Accuracy:', round(accuracy, 2), '%.')
# + id="w8kmtChqmbBb" colab_type="code" colab={}
# Import tools needed for visualization
from sklearn.tree import export_graphviz
import pydot
# Pull out one tree from the forest
tree = rf.estimators_[5]
# Import tools needed for visualization
from sklearn.tree import export_graphviz
import pydot
# Pull out one tree from the forest
tree = rf.estimators_[5]
# Export the image to a dot file
export_graphviz(tree, out_file = 'tree.dot', feature_names = feature_list, rounded = True, precision = 1)
# Use dot file to create a graph
(graph, ) = pydot.graph_from_dot_file('tree.dot')
# Write graph to a png file
graph.write_png('tree.png')
# + id="esqqwJcumpsY" colab_type="code" colab={}
# Limit depth of tree to 3 levels
rf_small = RandomForestRegressor(n_estimators=10, max_depth = 3)
rf_small.fit(train_features, train_labels)
# Extract the small tree
tree_small = rf_small.estimators_[5]
# Save the tree as a png image
export_graphviz(tree_small, out_file = 'small_tree.dot', feature_names = feature_list, rounded = True, precision = 1)
(graph, ) = pydot.graph_from_dot_file('small_tree.dot')
graph.write_png('/content/small_tree.png');
# + id="tJ0SZ3SNmq-1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="41df0bc8-e6a1-479b-f444-5e0e4b9e35dd"
pwd
# + id="zVvq5Mkqn__7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e9856d95-32e2-4bb1-82cd-eee60f80e933"
# Get numerical feature importances
importances = list(rf.feature_importances_)
# List of tuples with variable and importance
feature_importances = [(feature, round(importance, 2)) for feature, importance in zip(feature_list, importances)]
# Sort the feature importances by most important first
feature_importances = sorted(feature_importances, key = lambda x: x[1], reverse = True)
# Print out the feature and importances
[print('Variable: {:20} Importance: {}'.format(*pair)) for pair in feature_importances];
# + id="k0kop6I4p29o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 411} outputId="4cf7e561-2a82-4a9b-93fe-705d090675ce"
# Import matplotlib for plotting and use magic command for Jupyter Notebooks
import matplotlib.pyplot as plt
# %matplotlib inline
# Set the style
plt.style.use('fivethirtyeight')
# list of x locations for plotting
x_values = list(range(len(importances)))
# Make a bar chart
plt.bar(x_values, importances, orientation = 'vertical')
# Tick labels for x axis
plt.xticks(x_values, feature_list, rotation='vertical')
# Axis labels and title
plt.ylabel('Importance'); plt.xlabel('Variable'); plt.title('Variable Importances');
# + [markdown] colab_type="text" id="M3LDIV3vxiQi"
# #Practice
# + colab_type="code" id="3sB2SEvddTmW" colab={}
index = df.index
index
# + colab_type="code" id="TjfxTV65gw9Y" colab={}
columns = df.columns
columns
# + colab_type="code" id="a1ShlDvSinaq" colab={}
values = df.values
values
# + colab_type="code" id="1dbtrqCbssia" colab={}
# Check for NaN under a single DataFrame column:
df['geo_loc'].isnull().any()
# Count the NaN under a single DataFrame column:
df['geo_loc'].isnull().sum()
# Count the NaN under an entire DataFrame
df.isnull().sum()
# + colab_type="code" id="IgLNeAYDu4N9" colab={}
# droping rows with Nan in the geo_loc
df.dropna(subset=['geo_loc'], inplace=True)
# Count the NaN under a single DataFrame column:
df['geo_loc'].isnull().sum()
# + colab_type="code" id="pc-c6_WZxkyG" colab={}
import numpy as np
df1 = pd.DataFrame({"name": [12, np.nan, 20],
"num": [20, 14, np.nan],
"age": ['20', '14.43', '13,6'],
"toy": [np.nan, 'Batmobile,', 'Bul,lwhip'],
"born": [pd.NaT, pd.Timestamp("1940-04-25"),
pd.NaT]})
df1
# + colab_type="code" id="xbjgWCbXxkz5" colab={}
df1.dropna(subset=['name'], inplace=True)
# + colab_type="code" id="dlmTwQ2dxk10" colab={}
df1
# + colab_type="code" id="ZtSDLK53-EBY" colab={}
column_means = df1.mean()
# + colab_type="code" id="6dWJ3k6T-nRq" colab={}
column_means
# + colab_type="code" id="aFaZPcwA-EDi" colab={}
df1 = df1.fillna(column_means)
# + colab_type="code" id="JBEjy4c5FjPV" colab={}
df1['age'].replace(',','', regex=True, inplace=True)
# + colab_type="code" id="bJFWja86-EFk" colab={}
df1
# + colab_type="code" id="U8nj_JAJBvMb" colab={}
df6 = pd.to_numeric(df1['age'], downcast='integer')
# + colab_type="code" id="8TJs8v8gHm4w" colab={}
df6
| notebooks/random_forest_Irvington_validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import jsonlines
from tqdm import tqdm
import os
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID" # see issue #152
os.environ["CUDA_VISIBLE_DEVICES"] = ""
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (20.0, 10.0)
# -
#cond_ops = ['=', '>', '<', 'OP']
cond_ops = [[1,0,0], [0,1,0], [0,0,1], 'OP']
# +
#Paths
train_db_path='WikiSQL/data/train.db'
train_data_json_path='WikiSQL/data/train.jsonl'
train_table_json_path='WikiSQL/data/train.tables.jsonl'
dev_db_path='WikiSQL/data/dev.db'
dev_data_json_path='WikiSQL/data/dev.jsonl'
dev_table_json_path='WikiSQL/data/dev.tables.jsonl'
test_db_path='WikiSQL/data/test.db'
test_data_json_path='WikiSQL/data/test.jsonl'
test_table_json_path='WikiSQL/data/test.tables.jsonl'
# -
def get_col_tokens(table_json_path):
tokens=[]
with jsonlines.open(table_json_path) as reader:
for obj in tqdm(reader):
for header in obj['header']:
tokens.append(header)
return tokens
def get_question_tokens(data_json_path):
tokens=[]
with jsonlines.open(data_json_path) as reader:
for obj in tqdm(reader):
tokens.append(obj['question'])
return tokens
texts=[]
texts.extend(get_col_tokens(train_table_json_path))
texts.extend(get_col_tokens(dev_table_json_path))
texts.extend(get_col_tokens(test_table_json_path))
texts.extend(get_question_tokens(train_data_json_path))
texts.extend(get_question_tokens(dev_data_json_path))
texts.extend(get_question_tokens(test_data_json_path))
tokenizer=Tokenizer(char_level=True,lower=True)
tokenizer.fit_on_texts(texts)
max_token_index=len(tokenizer.index_word.keys())+1
max_token_index
def text2seq(text):
return np.ravel([tokenizer.texts_to_sequences([text])])
max_len=0
for text in texts:
if len(text2seq(text))>max_len:
max_len=len(text2seq(text))
max_len
def text2seq(text):
return np.ravel(pad_sequences(tokenizer.texts_to_sequences([text]),maxlen=max_len,padding='post'))
df = pd.Series(texts)
df.str.len().hist(bins=200)
plt.show()
del texts
#make a dict to keep track of all the columns of all the tables
def get_table(table_json_path):
tables={}
with jsonlines.open(table_json_path) as reader:
for obj in tqdm(reader):
tables[obj['id']]={'headers':[],'types':[]}
tables[obj['id']]['headers']=obj['header']
tables[obj['id']]['types']=obj['types']
return tables
train_tables=get_table(train_table_json_path)
dev_tables=get_table(dev_table_json_path)
test_tables=get_table(test_table_json_path)
def get_where_cols_value(dict_obj,tables):
temp=tables[dict_obj['table_id']]['headers']
where_cols = []
values=[]
ops=[]
for cond in dict_obj['sql']['conds']:
where_cols.append(temp[cond[0]])
value=str(cond[2]).lower()
start=dict_obj['question'].lower().find(value)
end=start+len(value)
values.append((start,end))
ops.append(cond_ops[cond[1]])
return where_cols,values,ops
def get_dataset(data_json_path,table):
#read the jsonl file
X=[]
y=[]
global tokenizer
with jsonlines.open(data_json_path) as reader:
for obj in tqdm(reader):
where_cols,values,ops=get_where_cols_value(obj,table)
question=obj['question']
#tokenized representation
question=text2seq(question)
for i in range(len(where_cols)):
X.append([question,text2seq(where_cols[i]),ops[i]])
y.append(values[i])
return X,y
X_train,y_train=get_dataset(train_data_json_path,train_tables)
X_dev,y_dev=get_dataset(dev_data_json_path,dev_tables)
#dimension
d=64
from keras.models import Model
from keras.layers import Input, GRU, Dense ,Flatten , Concatenate,Bidirectional,Average,RepeatVector,Flatten
from keras.layers.embeddings import Embedding
from keras.constraints import max_norm
from keras import regularizers
# +
# Define an input sequence and process it.
question_input = Input(shape=(max_len,),name='Q_input')
column_input = Input(shape=(max_len,),name='C_input')
ops_input = Input(shape=(3,),name='Ops_input')
embedding= Embedding(max_token_index, d, input_length=max_len,name='embedding')
# embedding_C= Embedding(max_token_index, d, input_length=max_len,name='embedding_C')
# embeddings_constraint=max_norm(2.),
# embeddings_regularizer=regularizers.l1_l2(l1=0.001, l2=0.001))
Q_embedding= embedding(question_input)
C_embedding= embedding(column_input)
encoder_question = Bidirectional(GRU(d, return_state=True))
Q_act , Q_state_h1, Q_state_h2 = encoder_question(Q_embedding)
encoder_column = Bidirectional(GRU(d, return_state=True))
C_act , C_state_h1, C_state_h2 = encoder_column(C_embedding)
con=Concatenate()([Q_act,Q_state_h1,Q_state_h2,C_act,C_state_h1,C_state_h2,ops_input])
start=Dense(max_len,activation='softmax',name='start')(con)
con_2=Concatenate()([con,start])
end=Dense(max_len,activation='softmax',name='end')(con_2)
model = Model([question_input, column_input,ops_input], [start,end])
model.summary()
# +
#from sklearn.externals import joblib
#wb=joblib.load('embedding select col.joblib')
#model.layers[2].set_weights(wb)
#model.layers[2].trainable = False
# +
import keras_contrib
from keras import optimizers
opt = optimizers.Adam(lr=0.0001)
# -
opt=keras_contrib.optimizers.padam.Padam()
# Run training
model.compile(optimizer=opt, loss='categorical_crossentropy',metrics=['accuracy'])
def generator(X,y,batch_size=32):
indices=range(len(X))
while True:
# Select files (paths/indices) for the batch
batch_indices= np.random.choice(a = indices,size = batch_size)
X_Q_batch=[]
X_C_batch=[]
X_O_batch=[]
y_start_batch=[]
y_end_batch=[]
for index in batch_indices:
X_Q_batch.append(X[index][0])
X_C_batch.append(X[index][1])
X_O_batch.append(X[index][2])
y_start_batch.append(np.eye(max_len)[y[index][0]])
y_end_batch.append(np.eye(max_len)[y[index][1]])
yield( [np.array(X_Q_batch),np.array(X_C_batch),np.array(X_O_batch)], [np.array(y_start_batch),np.array(y_end_batch)])
batch_size = 128 # Batch size for training.
epochs = 200 # Number of epochs to train for.
train_generator=generator(X_train,y_train,batch_size=batch_size)
dev_generator=generator(X_dev,y_dev,batch_size=batch_size)
from keras.callbacks import EarlyStopping, ModelCheckpoint
# Set callback functions to early stop training and save the best model so far
callbacks = [EarlyStopping(monitor='val_loss', patience=15),
ModelCheckpoint(filepath='where_value_best_model.h5', monitor='val_loss', save_best_only=True)]
train_steps=len(X_train)/batch_size
dev_steps=len(X_dev)/batch_size
model.load_weights('where_value_best_model.h5')
history=model.fit_generator(train_generator, steps_per_epoch=train_steps, epochs=epochs, verbose=1,
callbacks=callbacks, validation_data=dev_generator,
validation_steps=dev_steps,max_queue_size=10)
# +
# Get training and test loss histories
training_loss = history.history['loss']
test_loss = history.history['val_loss']
# Create count of the number of epochs
epoch_count = range(1, len(training_loss) + 1)
plt.figure(figsize=(20,10))
# Visualize loss history
plt.plot(epoch_count, training_loss, 'r--')
plt.plot(epoch_count, test_loss, 'b-')
plt.legend(['Training Loss', 'Validation Loss'])
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.show();
# -
X_test,y_test=get_dataset(test_data_json_path,test_tables)
test_generator=generator(X_test,y_test,batch_size=batch_size)
test_steps=len(X_test)/batch_size
model.evaluate_generator(dev_generator, steps=dev_steps, verbose=1)
model.evaluate_generator(test_generator, steps=test_steps, verbose=1)
| Train/SQL Where Value.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import nltk
# +
# nltk.download()
# -
import pandas as pd
# +
#TEST DATA
test = [
('The beer was good.', 'pos'),
('I do not enjoy my job', 'neg'),
("I ain't feeling dandy today.", 'neg'),
("I feel amazing!", 'pos'),
('Gary is a friend of mine.', 'pos'),
("I can't believe I'm doing this.", 'neg')
]
#TRAIN DATA
train = [('I love this sandwich.', 'pos'),
('This is an amazing place!', 'pos'),
('I feel very good about these beers.', 'pos'),
('This is my best work.', 'pos'),
("What an awesome view", 'pos'),
('I do not like this restaurant', 'neg'),
('I am tired of this stuff.', 'neg'),
("I can't deal with this", 'neg'),
('He is my sworn enemy!', 'neg'),
('My boss is horrible.', 'neg')]
# -
train = pd.DataFrame(train, columns=['sentences','sentiments'])
test = pd.DataFrame(test, columns=['sentences','sentiments'])
train
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
train['sentiments'] = encoder.fit_transform(train['sentiments'])
test['sentiments'] = encoder.fit_transform(test['sentiments'])
from nltk.corpus import stopwords
from string import punctuation
stuff_to_be_removed = list(stopwords.words("english"))+list(punctuation)
from nltk.tokenize import word_tokenize
from nltk.stem.lancaster import LancasterStemmer
from nltk.stem import WordNetLemmatizer
lancaster_stemmer = LancasterStemmer()
stemmer = LancasterStemmer()
lemmatizer = WordNetLemmatizer()
# +
# nltk.download('stopwords')
# -
train_corpus = train['sentences'].tolist()
test_corpus = test['sentences'].tolist()
train_label = train['sentiments'].tolist()
test_label = test['sentiments'].tolist()
corpus = train_corpus + test_corpus
labels = train_label + test_label
corpus
final_corpus = []
for i in range(len(corpus)):
text = word_tokenize(corpus[i].lower())
text = [lancaster_stemmer.stem(y) for y in text if y not in stuff_to_be_removed]
sent = " ".join(text)
final_corpus.append(sent)
# +
# nltk.download('punkt')
# -
labels
df = pd.DataFrame(final_corpus, columns = ['sentences'])
df['sentiments'] = labels
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
cv = CountVectorizer()
tfidf = TfidfVectorizer()
vector = tfidf.fit_transform(df['sentences'])
X = pd.DataFrame(vector.toarray(), columns=list(tfidf.get_feature_names()))
y = df['sentiments']
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split as tts
from sklearn.metrics import classification_report, confusion_matrix, accuracy_score
rfc = RandomForestClassifier(random_state=42)
X_train, X_test, y_train, y_test = tts(X,y,test_size = 0.25, random_state= 42)
rfc.fit(X_train, y_train)
rfc.score(X_test, y_test)
y_pred = rfc.predict(X_test)
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
# Logistic regression
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression(random_state=42)
log_reg.fit(X_train, y_train)
y_pred = log_reg.predict(X_test)
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
# Naive Bayes
from sklearn.naive_bayes import MultinomialNB
nb = MultinomialNB()
nb.fit(X_train,y_train)
y_pred = nb.predict(X_test)
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
gnb.fit(X_train,y_train)
y_pred = gnb.predict(X_test)
print(classification_report(y_test,y_pred))
print(accuracy_score(y_test,y_pred))
| C22_Natural Language Processing/NLP - Text Sentiment Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: bigdata
# language: python
# name: bigdata
# ---
# <a href="https://colab.research.google.com/github/SLCFLAB/Data-Science-Python/blob/main/Day%204/4_1.data_scraping.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="4Zy8dj0HFe1Y"
import requests
from bs4 import BeautifulSoup
import time
import pandas as pd
import numpy as np
import os
# + [markdown] id="I-mhkHr5YtuD"
# # References
# https://docs.python-requests.org/en/master/
#
#
# + [markdown] id="qMilnEXgACbV"
# # Request
# + colab={"base_uri": "https://localhost:8080/"} id="UzALRtxiYr3r" outputId="63d1d5fb-6c3f-49cd-a214-5c8aaad2cf30"
r = requests.get("https://www.wooribank.com/",
timeout = 15, # 1초 동안 무응답 시 에러처리
headers = {
"User-agent" : "Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_7) \
AppleWebKit/537.36 (KHTML, like Gecko) Chrome/98.0.4758.109"
}
)
r.encoding = "utf-8" # encoding 방식 지정해주기
print(r.status_code)
print(r.encoding)
# print(r.text)
# + colab={"base_uri": "https://localhost:8080/"} id="dyob5-ljZv02" outputId="da58ec3e-454f-441f-e615-8c52abc04608"
# 다른 페이지들도 가져와보기
naver_webtoon_html = requests.get("https://comic.naver.com/webtoon/weekday",
timeout = 15, # 1초 동안 무응답 시 에러처리
headers = {
"User-agent" : "Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_7) \
AppleWebKit/537.36 (KHTML, like Gecko) Chrome/98.0.4758.109"
}
)
print(naver_webtoon_html.status_code)
# print(naver_webtoon_html.text)
# + colab={"base_uri": "https://localhost:8080/"} id="CbJ6bCmQagle" outputId="caf5a687-a8fb-4149-a912-7e58e53c742b"
# 다른 페이지들도 가져와보기
melon_html = requests.get("https://www.melon.com/chart/index.htm",
timeout = 15, # 1초 동안 무응답 시 에러처리,
headers = {
"User-agent" : "Mozilla/5.0 (Macintosh; Intel Mac OS X 10_15_7) \
AppleWebKit/537.36 (KHTML, like Gecko) Chrome/98.0.4758.109"
}
)
print(melon_html.status_code)
# print(melon_html.text)
# + [markdown] id="lbCCR3rrYr3t"
# # BeautifulSoup
# + colab={"base_uri": "https://localhost:8080/"} id="2wxHzQtvYr3u" outputId="2ad50c4a-d76b-4b79-8885-859562b4f13a"
soup = BeautifulSoup(r.content, 'html.parser')
# print(soup.prettify())
print(soup.title)
print(soup.title.string)
# + colab={"base_uri": "https://localhost:8080/"} id="WtOR4Y3LiGL2" outputId="77e88e85-656b-45fc-9216-0faa0ecf0f4a"
soup.select("div[class='tit-5 absolute js-display-hover']")
# + colab={"base_uri": "https://localhost:8080/"} id="jU6XSg_uiJhr" outputId="4a7efb8d-e224-4cec-df69-781dd711d9ca"
soup.select("div[class='tit-5 absolute js-display-hover'] li")
# + colab={"base_uri": "https://localhost:8080/"} id="eUxrC5lIYr3v" outputId="966cbf24-20a3-42a3-93d8-b1b62fe855c8"
for item in soup.select("div[class='tit-5 absolute js-display-hover'] li"):
print(item.text)
# + colab={"base_uri": "https://localhost:8080/"} id="RM2FPiF6jhUk" outputId="001293ac-ebc2-4a39-ceb2-8a76f7d55fcb"
soup.select_one("div[class='tit-5 absolute js-display-hover'] li")
# + colab={"base_uri": "https://localhost:8080/"} id="E-e3NaBkj9HK" outputId="2228eef3-5ca3-4da5-f0d5-2926cb1b53b0"
soup.find("div",{"class":"tit-5 absolute js-display-hover"})
# + colab={"base_uri": "https://localhost:8080/"} id="Yd6ymY_OlD92" outputId="bf609473-8523-404c-ba83-06c1b8d8128d"
soup.find("div",{"class":"tit-5 absolute js-display-hover"}).find_all("li")
# + [markdown] id="cMP-EWjfYr3v"
# # Request, BS4 실습
# + id="FpuJnmYmnq45"
# 연습용 페이지 크롤링 (To-do)
# https://dataquestio.github.io/web-scraping-pages/ids_and_classes.html
# "First-outer paragraph" 출력해보기
# + id="uKxCKAmwYr3w"
# 오늘의 코스피 받아오기 (To-do)
# + id="cNsFPmWStwS9"
# 네이버웹툰 https://comic.naver.com/webtoon/weekday 실시간 인기 급상승 웹툰 1위-10위 제목 뽑아보기
# + id="Wa53XQe2qj7O"
# National Weather Service 페이지 크롤링
# https://forecast.weather.gov/MapClick.php?lat=37.7772&lon=-122.4168#.Yk05PBNBxFw
# + id="FU3bY1AaYr3x"
# 시간별 날씨 (To-do)
# https://weather.naver.com/today/
# 시간대별로 날씨 뽑아와서 dictionary에 저장하기
# + id="AAoNHMQauDH0"
# Challenging
# Melon playlist
# https://www.melon.com/chart/index.htm
# 1위-100위 노래 제목 뽑기
# + id="vT8wPlyryb_-"
| Day 4/4_1.data_scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID-19 Reopening / Recovery Indicators
#
# * [Federal Gating Criteria](https://www.whitehouse.gov/wp-content/uploads/2020/04/Guidelines-for-Opening-Up-America-Again.pdf)
# * [State Gating Criteria](https://covid19.ca.gov/roadmap-counties/)
# * [WHO Testing and Positivity Rate Guidelines](https://coronavirus.jhu.edu/testing/testing-positivity)
# * More [background articles](https://github.com/CityOfLosAngeles/covid19-indicators/blob/master/reopening-sources.md)
# * NY, Chicago, LA, and CA indicators [provided in our GitHub](https://github.com/CityOfLosAngeles/covid19-indicators/blob/master/Reopening_Indicators_Comparison.xlsx)
#
# This notebook gives helpful hints for working with our COVID-19 data in Jupyter Notebooks. It covers:
# * How to import CSVs hosted on ESRI
# * How to use functions `utils.py` to get the cases and deaths charts for any US county, state, or MSA.
# * How to use the functions in `utils.py` to get the hospitalizations charts for any CA county.
# +
import pandas as pd
from processing_utils import utils
from datetime import date
# -
# ## Import data from ESRI CSV
# The [README](https://github.com/CityOfLosAngeles/aqueduct/blob/master/dags/public-health/covid19/README.md#helpful-hints) gave some helpful hints for how to read in CSVs that are hosted by ESRI.
US_COUNTY_URL = "http://lahub.maps.arcgis.com/sharing/rest/content/items/782ca660304a4bdda1cc9757a2504647/data"
CROSSWALK_URL = (
"https://raw.githubusercontent.com/CityOfLosAngeles/covid19-indicators/master/data/"
"msa_county_pop_crosswalk.csv"
)
df = pd.read_csv(US_COUNTY_URL, dtype={"fips": "str"})
df.head(2)
crosswalk = pd.read_csv(CROSSWALK_URL)
crosswalk.head(2)
# ## County indicators for cases and deaths
# * Uses the `county_case_charts` function within `utils`
# * 7-day rolling average shown for new cases
# * 7-day rolling average shown for new deaths
#
# The function takes `COUNTY INFORMATION` as a string and `START_DATE` as a datetime variable, and returns the charts and the dataframe used to create the chart.
#
# `COUNTY_INFORMATION` takes form of {COUNTY_NAME, STATE_NAME} or {5-digit county FIPS code}:
# * {COUNTY_NAME}: with our without " County" e.g. "Alameda County" or "Alameda"
# * {STATE_NAME}: full state name or state abbreviation e.g. "California" or "CA"
# * {5-DIGIT COUNTY FIPS CODE}: a [list of county FIPS found here.](https://www.nrcs.usda.gov/wps/portal/nrcs/detail/national/home/?cid=nrcs143_013697) e.g. "06075" for San Francisco
#
# `START_DATE` takes form of a datetime variable
# * Use the `datetime` package
# * Put in form date(YYYY, M, D), such as (2020, 5, 1) or (2020, 4, 10)
start_date = date(2020, 5, 1)
cook = utils.county_case_charts("Cook, Illinois", start_date)
cook.head()
# Other acceptable examples are:
# ```
# start_date = date(2020, 5, 15)
# dallas = utils.county_case_charts("Dallas, TX", start_date)
# atlanta = utils.county_case_charts("Fulton, Georgia", start_date)
# ```
# ## State indicators for cases and deaths
# * Uses the `state_case_charts` function within `utils`
# * 7-day rolling average shown for new cases
# * 7-day rolling average shown for new deaths
#
# The function takes `STATE_INFORMATION` as a string and `START_DATE` as a datetime variable, and returns the charts and the dataframe used to create the chart.
#
# `STATE_INFORMATION` takes form of {STATE_NAME}:
# * {STATE_NAME}: full state name or state abbreviation e.g. "California" or "CA"
start_date = date(2020, 3, 1)
ny = utils.state_case_charts("New York", start_date)
ny.head()
# Other acceptable examples are:
# ```
# start_date = date(2020, 2, 15)
# tn = utils.state_case_charts("Tennessee", start_date)
# fl = utils.state_case_charts("FL", start_date)
# ```
# ## MSA indicators for cases and deaths
# * Uses the `msa_case_charts` function within `utils`
# * 7-day rolling average shown for new cases
# * 7-day rolling average shown for new deaths
#
# The function takes `METROPOLITAN_INFORMATION` as a string and `START_DATE` as a datetime variable, and returns the charts and the dataframe used to create the chart. [Our crosswalk makes this information available.](https://github.com/CityOfLosAngeles/covid19-indicators/blob/master/data/msa_county_pop_crosswalk.csv)
#
# `METROPOLITAN_INFORMATION` takes form of {CBSA NAME} or {CBSA CODE}:
# * {CBSA NAME}: full metropolitan area name or any subset of it e.g. "Boston-Cambridge-Newton, MA-NH" or "Boston-Cambridge"
# * {CBSA CODE}: e.g. "14460" for Boston-Cambridge-Newton, MA-NH or "31080" for Los Angeles-Long Beach-Anaheim, CA
start_date = date(2020, 5, 1)
boston_msa = utils.msa_case_charts("Boston-Cambridge", start_date)
boston_msa.head()
# Other acceptable examples are:
# ```
# start_date = date(2020, 4, 1)
# dfw = utils.msa_case_charts("19100", start_date)
# durham = utils.msa_case_charts("Durham-Chapel Hill", start_date)
# ```
# ## CA COVID-hospitalizations
# * Uses the `county_covid_hospital_charts` function within `utils`
# * 7-day rolling average shown for all COVID-related hospitalizations and COVID-ICU hospitalizations (subset of all hospitalizations)
start_date = date(2020, 4, 30)
orange = utils.county_covid_hospital_charts("Orange, CA", start_date)
orange.head()
# Other acceptable examples are:
# ```
# start_date = date(2020, 5, 15)
# sf = utils.county_covid_hospital_charts("06075", start_date)
# santa_clara = utils.county_covid_hospital_charts("Santa Clara County", start_date)
# sd = utils.county_covid_hospital_charts("San Diego County, CA", start_date)
# ```
| notebooks/sample-indicators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R [conda env:larval_gonad]
# language: R
# name: conda-env-larval_gonad-r
# ---
# # Testis Replicate 1 scRNA-Seq UMI Cutoff 5,000
# This is the seurat analysis for testis replicate 1 after setting the UMI cuotff of 5,000
options(repr.plot.width=10, repr.plot.height=10)
# +
DATA <- '../output/2018-03-21_explore_umi_testis1_5000.tsv'
OUTDIR <- '../output/testis1_scRNAseq_umi_5k'
REFERENCES_DIR <- Sys.getenv('REFERENCES_DIR')
NAME <- 'TestisForce3k'
# Get list of mitochondiral genes
fbgn2chrom <- read.table('../output/fbgn2chrom.tsv', header=T)
fbgn2symbol <- read.csv(file.path(REFERENCES_DIR, 'dmel/r6-16/fb_annotation/dmel_r6-16.fb_annotation'), header=T, sep = '\t')[, c('gene_symbol', 'primary_FBgn')]
mito <- fbgn2chrom[fbgn2chrom$chrom == 'chrM', 'FBgn']
# -
source('../lib/seurat.R')
library(Seurat)
library(dplyr)
library(Matrix)
dat <- read.table(DATA, header=TRUE)
# +
# Initialize the Seurat object with the raw (non-normalized data).
# Keep all genes expressed in >= 3 cells (~0.1% of the data). Keep all cells with at least 200 detected genes
sobj <- CreateSeuratObject(raw.data = dat, min.cells = 3, min.genes = 200, project = NAME)
nCells <- dim(sobj@meta.data)[1]
# calculate the percent genes on chrom M
mask <- row.names(sobj@raw.data) %in% mito
percent.mito <- Matrix::colSums(sobj@raw.data[mask, ]/Matrix::colSums(sobj@raw.data)) * 100
sobj <- AddMetaData(object = sobj, metadata = percent.mito, col.name = "percent_mito")
# -
nCells
VlnPlot(object = sobj, features.plot = c('nGene', 'nUMI', 'percent_mito'), nCol = 2)
par(mfrow = c(2, 1))
GenePlot(object = sobj, gene1 = 'nUMI', gene2 = 'percent_mito')
GenePlot(object = sobj, gene1 = 'nUMI', gene2 = 'nGene')
# Filter Genes based on low and high gene counts
sobj <- FilterCells(object=sobj, subset.names=c("nGene"), low.thresholds=c(200), high.thresholds=c(6000))
dim(sobj@meta.data)[1]
# + code_folding=[]
sobj <- NormalizeData(object = sobj, normalization.method = "LogNormalize", scale.factor = 1e4)
sobj <- FindVariableGenes(object = sobj, mean.function = ExpMean, dispersion.function = LogVMR,
x.low.cutoff = 0.01,
x.high.cutoff = 2.8,
y.cutoff = 0.5,
y.high.cutoff = Inf
)
# -
length(x = sobj@var.genes)
sobj <- ScaleData(object = sobj, vars.to.regress = c("nUMI"), display.progress = F)
# + code_folding=[]
### Perform linear dimensional reduction
sobj <- RunPCA(object = sobj, pc.genes = sobj@var.genes, do.print = FALSE, pcs.print = 1:5, genes.print = 5, pcs.compute = 100)
# -
PrintPCA(object = sobj, pcs.print = 1:5, genes.print = 5, use.full = FALSE)
VizPCA(object = sobj, pcs.use = 1:3)
PCAPlot(object = sobj, dim.1 = 1, dim.2 = 2)
# + code_folding=[]
# ProjectPCA scores each gene in the dataset (including genes not included in the PCA) based on their correlation
# with the calculated components. Though we don't use this further here, it can be used to identify markers that
# are strongly correlated with cellular heterogeneity, but may not have passed through variable gene selection.
# The results of the projected PCA can be explored by setting use.full=T in the functions above
sobj <- ProjectPCA(object = sobj, do.print = F)
# -
PCElbowPlot(object = sobj, num.pc=30)
sobj <- JackStraw(object = sobj, num.replicate = 100, do.print = FALSE, num.pc = 30)
JackStrawPlot(object = sobj, nCol = 6, PCs = 1:30)
# + code_folding=[]
sobj <- FindClusters(
object = sobj,
reduction.type = "pca",
dims.use = 1:30,
resolution = c(0.4, 0.6, 1.0, 1.2, 1.4),
print.output = 0,
save.SNN = TRUE,
)
PrintFindClustersParams(object = sobj)
# + code_folding=[]
### Run Non-linear dimensional reduction (tSNE)
sobj <- RunTSNE(object = sobj, dims.use = 1:30, do.fast = TRUE)
# -
TSNEPlot(object = sobj, group.by='res.1.2')
dump_seurat(object = sobj, dir = OUTDIR)
# Save cluster info
params <- c(0.4, 0.6, 1.0, 1.2, 1.4)
for (i in params) {
name <- paste0('res.', i)
fname <- paste0('biomarkers_', i, '.tsv')
sobj <- SetAllIdent(sobj, id = name)
markers <- FindAllMarkers(object = sobj, only.pos = TRUE, min.pct = 0.25, thresh.use = 0.25, print.bar = FALSE)
markers = merge(fbgn2symbol, markers, by.x='primary_FBgn', by.y='gene', all.y=T)
save_biomarkers(markers = markers, dir = OUTDIR, fname = fname)
}
| notebook/2018-03-21_testis1_5k_umi.ipynb |