code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook Intentions
#
# The data types used by MEPS are a bit archaic. Additionally since 2017 several files have been formatted in a way such that they can only be accessed using SAS.
#
# In this notebook we will attempt to unpack data contained in the ascii files using the data parameters intended for R developers. If we can successfully do this we won't require the use of pyr2 and the maintainence of a second language in the codebase (R).
#
# +
import os
from os.path import expanduser
import sys
sys.path.append(os.path.join(expanduser("~"), "meps"))
from zipfile import ZipFile
from meps_db.components.populators import BaseComponentsPopulator as bcp
from meps_db.components.reference import FYCDF_PUF_SSP_LOOKUP
from meps_db.utilities.universal_utilities import UniversalUtilityFunctions as util
# -
# get path
zip_path = bcp.get_zip_path(zip_type="consolidated", year=2018, year_lookup=FYCDF_PUF_SSP_LOOKUP)
print(f"Path to .dat file: {zip_path}")
# +
# unzip
filename = zip_path.split("/")[-1]
unzip_path = zip_path.replace(filename, "")
unzipped_filename = filename.split("dat.zip")[0] + ".dat"
with ZipFile(zip_path,"r") as zip_ref:
zip_ref.extractall(unzip_path)
print(f"Unzipped: {filename} to {unzip_path} as {unzipped_filename}")
# -
# store ascii
with open(os.path.join(unzip_path, unzipped_filename), 'rb') as f:
ascii_text = f.read()
# load R parameters
puf_params = util.load_data_from_file(
file_path=os.path.join(
expanduser("~"),
"meps",
"meps_dev",
"meps_db",
"components",
"populator_support",
unzipped_filename.strip('.dat'),
),
file_format="json"
)
# +
# test first respondent
resp = 0
row = {}
for start, end, name, dtype in zip(
puf_params["position_start"],
puf_params["postion_end"],
puf_params["var_names"],
puf_params["var_types"]
):
val = ascii_text[start-1:end].decode("utf-8").strip()
typed_val = float(val) if dtype == "n" else str(val)
row.update({name: typed_val})
for key in list(row.keys())[:10]:
print(f"{key}: {row[key]}")
# +
# test full ascii text
data = []
resp = 0
row_data = ascii_text.decode("utf-8").split("\r\n")
for row in row_data[:-1]:
resp_dict = {}
for start, end, name, dtype in zip(
puf_params["position_start"],
puf_params["postion_end"],
puf_params["var_names"],
puf_params["var_types"]
):
val = row[start-1:end].strip()
typed_val = float(val) if dtype == "n" else str(val)
resp_dict.update({name: typed_val})
data.append(resp_dict)
|
sandbox/20201_03_upack_ascii/unpack_ascii.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Selección óptima de portafolios I
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/thumb/6/6e/Separation_theorem_of_MPT.svg/2000px-Separation_theorem_of_MPT.svg.png" width="400px" height="400px" />
#
# En la clase pasada vimos que:
# - La LAC describe las posibles selecciones de riesgo-rendimiento entre un activo libre de riesgo y un activo riesgoso.
# - Su pendiente es igual al radio de Sharpe del activo riesgoso.
# - La asignación óptima de capital para cualquier inversionista es el punto tangente de la curva de indiferencia del inversionista con la LAC (depende de las preferencias particulares - aversión al riesgo).
#
# Para todo lo anterior, supusimos que ya teníamos el portafolio óptimo (activo riesgoso).
#
# En el siguiente análisis:
#
#
# **Objetivos:**
# - ¿Cuál es el portafolio óptimo de activos riesgosos?
# - ¿Cuál es el mejor portafolio de activos riesgosos?
# - Es un portafolio eficiente en media-varianza.
# - Problema: dado un conjunto de activos riesgosos, ¿cómo construimos la mejor combinación?
#
# *Referencia:*
# - Notas del curso "Portfolio Selection and Risk Management", Rice University, disponible en Coursera.
# ___
# ## 1. Maximizando el radio de Sharpe
#
# ### ¿Qué pasa si tenemos dos activos riesgosos?
#
# Cuando tenemos dos o más activos riesgosos, tenemos disponibles diferentes LAC. ¿Qué significan sus pendientes?
#
# <font color=blue> Ver en el tablero.</font>
#
# Pregunta:
# - ¿Qué es lo que se quiere?
# **Conclusión:**
# - El mejor portafolio de activos no depende de las preferencias individuales, y por tanto va a ser igual para todos.
# - Dicho mejor portafolio maximiza el radio de Sharpe.
# - A este portafolio lo llamaremos el portafolio eficiente en media-varianza (EMV)
# **Idea principal: el portafolio óptimo de activos riesgosos es independiente de las preferencias del inversionista.**
# - El portafolio EMV determina el portafolio óptimo de activos riesgosos.
# - Todos tendremos el mismo portafolio de activos riesgosos (EMV), y lo combinaremos con el activo libre de reisgo de acuerdo con las preferencias de cada uno de nosotros (aversión al riesgo).
# - La LAC combinando el activo libre de riesgo y el portafolio EMV, se vuelve el conjunto de portafolios eficientes.
# Entonces, se deben seguir los siguientes pasos:
# 1. Crear la frontera media-varianza.
# 2. Encontrar el portafolio que maximize el radio de Sharpe (portafolio EMV).
# 3. Construir la frontera eficiente (LAC) del punto $(0,r_f)$ al punto $(\sigma_s,E[r_s])$ del portafolio EMV.
# 4. Combinar de acuerdo a sus preferencias.
# ___
# ## 2. Solución analítica del portafolio EMV: caso con dos activos.
#
# Queremos solucionar el siguiente problema:
#
# \begin{align}
# \max_{w_1,w_2} &\quad \frac{E[r_p]-r_f}{\sigma_p}\\
# \text{s.a.} &\quad E[r_p]=w_1E[r_1]+w_2E[r_2]\\
# &\quad \sigma_p=\sqrt{w_1^2\sigma_1^2+w_2^2\sigma_2^2+2w_1w_2\rho_{12}\sigma_1\sigma_2}\\
# &\quad w_1+w_2=1, \quad w_1,w_2\geq0
# \end{align}
# el cual es equivalente a
#
# \begin{align}
# \max_{w_1} &\quad \frac{w_1E[r_1]+(1-w_1)E[r_2]-r_f}{\sqrt{w_1^2\sigma_1^2+(1-w_1)^2\sigma_2^2+2w_1(1-w_1)\rho_{12}\sigma_1\sigma_2}}\\
# \text{s.a.} &\quad 0\leq w_1\leq1
# \end{align}
# **Actividad.**
# El anterior es un problema de maximizar una función de una variable en un dominio cerrado. No debaría representar dificultad.
#
# Encontrar la solución analítica a este problema.
#
# Quien primero lo haga, y salga a explicarlo al tablero, le subo alguna tarea, quiz o el primer examen a 100.
#
# Deben llegar a:
#
# $$w_{1,EMV}=\frac{(E[r_1]-r_f)\sigma_2^2-(E[r_2]-r_f)\sigma_{12}}{(E[r_2]-r_f)\sigma_1^2+(E[r_1]-r_f)\sigma_2^2-((E[r_1]-r_f)+(E[r_2]-r_f))\sigma_{12}}.$$
# Si nadie lo ha hecho en 30 min., procederé a hacerlo yo.
#
# **Nota:**
# - así como obtuvimos una expresión para el peso del portafolio de mínima varianza con dos activos, obtenemos una expresión para el peso del portafolio Eficiente en Media-Varianza.
# - Estas actividades son sin duda un buen ejercicio, y se pueden replicar usando técnicas de varias variables (multiplicadores de Lagrange) cuando se tengan más de dos activos.
# - Sin embargo, la complejidad del problema crece considerablemente con el número de variables, y la solución analítica deja de ser viable cuando mencionamos que un portafolio bien diversificado consta aproximadamente de 50-60 activos.
# - En esos casos, este problema se soluciona con rutinas numéricas que hagan la optimización por nosotros.
# - Por eso, les enseño cómo resolver este problema con optimizadores numéricos, porque son una solución viable y escalable a más variables.
# ## 3. Ejemplo ilustrativo.
#
# Retomamos el ejemplo de mercados de acciones en los países integrantes del $G5$: EU, RU, Francia, Alemania y Japón.
# Importamos pandas y numpy
import pandas as pd
import numpy as np
# +
# Resumen en base anual de rendimientos esperados y volatilidades
annual_ret_summ = pd.DataFrame(columns=['EU', 'RU', 'Francia', 'Alemania', 'Japon'], index=['Media', 'Volatilidad'])
annual_ret_summ.loc['Media'] = np.array([0.1355, 0.1589, 0.1519, 0.1435, 0.1497])
annual_ret_summ.loc['Volatilidad'] = np.array([0.1535, 0.2430, 0.2324, 0.2038, 0.2298])
annual_ret_summ.round(4)
# -
# Matriz de correlación
corr = pd.DataFrame(data= np.array([[1.0000, 0.5003, 0.4398, 0.3681, 0.2663],
[0.5003, 1.0000, 0.5420, 0.4265, 0.3581],
[0.4398, 0.5420, 1.0000, 0.6032, 0.3923],
[0.3681, 0.4265, 0.6032, 1.0000, 0.3663],
[0.2663, 0.3581, 0.3923, 0.3663, 1.0000]]),
columns=annual_ret_summ.columns, index=annual_ret_summ.columns)
corr.round(4)
# Supondremos, además, que la tasa libre de riesgo es $r_f=5\%$.
# Tasa libre de riesgo
# Entonces, supondremos que tenemos disponibles los activos correspondientes a los mercados de acciones de EU y Japón, y en adición el activo libre de riesgo.
# #### 1. Construir la frontera de mínima varianza
# +
# Vector de w variando entre 0 y 1 con n pasos
# Rendimientos esperados individuales
# Activo1: EU, Activo2:Japon
# Volatilidades individuales
# Correlacion
# -
# DataFrame de portafolios:
# 1. Índice: i
# 2. Columnas 1-2: w, 1-w
# 3. Columnas 3-4: E[r], sigma
# 4. Columna 5: Sharpe ratio
# Importar librerías de gráficos
# Gráfica de dispersión de puntos coloreando
# de acuerdo a SR
# #### 2. Encontrar el portafolio que maximiza el radio de Sharpe (EMV)
# Primero, encontramos este portafolio con la fórmula que obtuvimos:
# Fórmula que obtuvimos
# Ahora sí, con la función scipy.optimize.minimize
# Importar el módulo optimize
# +
## Construcción de parámetros
## Activo 1: EU, Activo 2: Japon
# 1. Sigma: matriz de varianza-covarianza
# 2. Eind: rendimientos esperados activos individuales
# -
# Función objetivo (-SR)
# +
# Dato inicial
# Cotas de las variables
#bnds = ((0,1),(0,1))#
# Restricciones
# +
# Optimización numérica
# Resultado
# -
# Con lo anterior, podemos obtener datos de rendimiento esperado y volatilidad del portafolio EMV
# Rendimiento esperado y volatilidad del portafolio EMV
# Gráfica de dispersión de puntos coloreando
# de acuerdo a SR, y portafolio EMV
# #### 3. Construir LAC
# Ahora, dibujamos la LAC, combinando el portafolio EMV con el activo libre de riesgo:
# Vector de wp variando entre 0 y 1.5 con n pasos
# DataFrame de CAL:
# 1. Índice: i
# 2. Columnas 1-2: wp, wrf
# 3. Columnas 3-4: E[r], sigma
# 4. Columna 5: Sharpe ratio
# Gráfica de dispersión de puntos coloreando
# de acuerdo a SR, portafolio EMV y LAC
# #### 4. Combinación óptima de acuerdo a preferencias
#
# Con los datos anteriores, y la caracterización de aversión al riesgo, se escoge la combinación óptima entre el portafolio EMV y el activo libre de riesgo de acuerdo a:
#
# $$w^\ast=\frac{E[r_s-r_f]}{\gamma\sigma_s^2}.$$
# Para gamma=7
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
|
Modulo3/Clase14_SeleccionOptimaPortI.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:MachineLearning]
# language: python
# name: conda-env-MachineLearning-py
# ---
# ## Image classifier (least squares method)
#
# This is simple classifier based on [least squares method](https://en.wikipedia.org/wiki/Least_squares).
# First of all, let's import numpy and time libraries.
import numpy as np
import time
# Now, let's read the data. It is written in two files (<i>mnist_train.csv</i> and <i>mnist_test.csv</i>) picked from [here](https://pjreddie.com/projects/mnist-in-csv/).
#
# <b>Attention!</b> Reading and parsing big csv files needs some memory (up to a couple of gigabytes) and some calculation power, so it's better to run the following block of code only once.
train_data = np.loadtxt("mnist_train.csv", delimiter=',')
test_data = np.loadtxt("mnist_test.csv", delimiter=',')
# As you can see, we have 60k vectors representing images in `train_data` and 10k vectors in `test_data`. Every vector has 785 values - first one is the tag (a digit from 0 to 9) and next 784 numbers represent the image of this digit (actually it is reshaped 28x28 matrix of pixels, modified for easier processing).
print(train_data.shape)
print(test_data.shape)
# So, let's start. Our algorithm has complexity of $O(N \times M \times L)$, where $N$ is the size of train selection, $M$ is the size of test selection and $L$ represents the length of vectors in selections. Therefore, we need to choose subsets of our selections and work with it, no with the full data.
#
# We still can take full data, but then computations will require a lot of time.
train_size = 10000
test_size = 1000
IMAGE_LENGTH = test_data.shape[1] - 1
# +
def get_random_subset(data, new_size):
old_size = data.shape[0]
indexes = np.random.choice(old_size, new_size, replace = False)
labels = data[indexes, 0].astype(int)
images = data[indexes, 1:]
return labels, images
train_labels, train_img = get_random_subset(train_data, train_size)
test_labels, test_img = get_random_subset(test_data, test_size)
# -
# Now, let's write our classifier itself. As said above, we will use the least squares method.
#
# How does it work?
# For every sample from test selection (which is a vector), we need to find the closest (by Euclidean distance) vector from train selection. The label of the closest vector will be the "predicted" label of vector from test selection.
#
#
# <b>Comment:</b> for small sizes of test and train subsets we could vectorize our algorithm by creating matrix of Euclidean distance between each pair of vectors. However, it will rise memory usage which is critical for huge subsets of images.
def classify_image(test_img, train_img, train_labels):
DM = np.square(test_img - train_img).sum(axis = 1)
index = DM.argmin(axis = 0)
return train_labels[index]
# Now let's test our algorithm. We will also measure time of execution using `time` library.
# +
# %%time
predicted_results = [classify_image(test, train_img, train_labels)
for test in test_img]
success_count = (predicted_results == test_labels).sum()
accuracy = success_count / test_size
print("SAMPLES COUNT :", train_size)
print("TESTS COUNT :", test_size)
print("ACCURACY :", np.round(accuracy * 100, 2), '%')
# -
# Well done!
#
# Now let's see how accuracy depends on size of train subset. Since small subsets has more contribution of random in result accuracy, we need to choose different subsets and run our classifier on each. As a result, we will take the average value of accuracy as our accuracy for a specific size of subset.
def get_average_accuracy(test_img, train_data, train_subset_size, iterations_count):
total_accuracy = 0
for i in range(iterations_count):
train_labels, train_img = get_random_subset(train_data, train_subset_size)
predicted_results = [classify_image(test, train_img, train_labels)
for test in test_img]
success_count = (predicted_results == test_labels).sum()
total_accuracy += success_count / test_labels.shape[0]
return total_accuracy / iterations_count
# +
# %%time
size_of_subset_v = [50, 200, 500, 1000, 2000, 5000, 10000]
count_of_iters_v = [100, 40, 20, 15, 10, 3, 1]
accuracy_info = [get_average_accuracy(test_img, train_data, size, iters)
for size, iters in zip(size_of_subset_v, count_of_iters_v)]
# -
# Let's visualize our results using $matplotlib$. Note that x-axis is logarithmically scaled.
# +
import matplotlib.pyplot as plt
plt.semilogx(size_of_subset_v, accuracy_info, color = "g")
plt.scatter(size_of_subset_v, accuracy_info, color = "b")
plt.xlabel("Size of train subset")
plt.ylabel("Accuracy")
plt.grid(axis="y", which="both", linestyle='--')
plt.show()
# -
# As it was expected, the accuracy depends on size of train subset.
#
# #### Summary:
# Least squares method is quite good for such type of dataset. It is easy-to-understand and easy-to-code, but it is very sensitive for bad data. Plus this method has to work with the full train dataset every time we want to get prediction.
#
# So, you could choose this method if:
# * Your dataset is well-separated and clear (has no wrong data)
# * It's easy to separate test data using the small subset of train data
# * You have no time for understanding and coding better and more complex soutions
#
|
practice_1/ImageClassifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: blog
# language: python
# name: blog
# ---
# +
# In this post I am going to try to explain how to do the CSS scrolling colour effect.
# If it comes out nicely I will put it in the blog but for now I like the original more.
# Pinetools, CSS @keysomethings filter animation, feColorMatrix, contrast, brightness, greyscale, whatever.
# also to remember is that I had to move some of the css from the body background to the side panels.
# -
|
posts/wip/css_animation_filters.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import os
from collections import namedtuple
from pyspark.sql import SparkSession, Row
sc = spark.sparkContext
sc.setLogLevel('WARN')
# +
# clean old table
answer_df = spark.sql("select * from hive_table limit 5")
assert answer_df.first().X_1 == 1.624
assert answer_df.first().X_8 == -0.761
answer_df.show()
# -
spark.sql("select count(*) as row_count from hive_table").show()
spark.sql("show tables").show()
spark.sql("show databases").show()
x = spark.sql("show create table hive_table").toPandas()
print(x.createtab_stmt[0])
|
project/spark_hive_read_table.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import fire_simulation as fs
# This is a simple simulation for fire on homogeneous field with no wind: fire spread rate is ($\sqrt{n+1}$ - $\sqrt{n}$)r .
# Black shows burned area,red shows burning area, and white shows unburned area.
fs.homofire_nowind()
# This is a simple simulation for fire on homogeneous field with constant wind. Black shows burned area, red shows burning area,
# and white shows unburned area.
fs.homofire_cwind()
# This is a simple simulation for fire on homogeneous field with constant wind which has a direction change with time.
fs.homofire_dwind()
# this is the 2-D wind-driven fire simulation. Input the wind direction, 'N', 'S', 'E', 'W', 'NW', 'NE', 'SE', 'SW'
fs.wind_fire_simulation('SE')
|
fire_behavior/Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ssCOanHc8JH_"
# # Training Adversarial Inverse RL and State Marginal Matching Agents in Brax
#
# In [Brax Training](https://colab.research.google.com/github/google/brax/blob/main/notebooks/training.ipynb) we tried out [gym](https://gym.openai.com/)-like environments and PPO, SAC, evolutionary search, and trajectory optimization algorithms. We can build various RL algorithms on top of these ultra-fast implementations. This colab runs a family of [adversarial inverse RL](https://arxiv.org/abs/1911.02256) algorithms, which includes [GAIL](https://papers.nips.cc/paper/2016/hash/cc7e2b878868cbae992d1fb743995d8f-Abstract.html) and [AIRL](https://arxiv.org/abs/1710.11248) as special cases. These algorithms minimize D(p(s,a), p\*(s,a)) or D(p(s), p\*(s)), the divergence D between the policy's state(-action) marginal distribution p(s,a) or p(s), and a given target distribution p\*(s,a) or p\*(s). As discussed in [f-MAX](https://arxiv.org/abs/1911.02256), these algorithms could also be used for [state-marginal matching](https://arxiv.org/abs/1906.05274) RL besides imitation learning. Let's try them out!
#
# This provides a bare bone implementation based on minimal modifications to the
# baseline [PPO](https://github.com/google/brax/blob/main/brax/training/ppo.py),
# enabling training in a few minutes. More features, tunings, and benchmarked results will be added soon, including:
# * Support for training a mixture of policies
# * Examples for imitation learning
# + [markdown] id="VYe1kc3a4Oxc"
#
#
# ```
# # This is formatted as code
# ```
#
# [](https://colab.research.google.com/github/google/brax/blob/main/notebooks/braxlines/irl_smm.ipynb)
# + id="_sOmCoOrF0F8"
#@title Install Brax and some helper modules
#@markdown ## ⚠️ PLEASE NOTE:
#@markdown This colab runs best using a TPU runtime. From the Colab menu, choose Runtime > Change runtime type, then select 'TPU' in the dropdown.
from datetime import datetime
import functools
import os
import jax
import jax.numpy as jnp
from IPython.display import HTML, clear_output
import matplotlib.pyplot as plt
try:
import brax
except ImportError:
# !pip install git+https://github.com/google/brax.git@main
clear_output()
import brax
import tensorflow_probability as tfp
from brax.io import html
from brax.experimental.composer import composer
from brax.experimental.composer import observers
from brax.experimental.braxlines.training import ppo
from brax.experimental.braxlines.irl_smm import utils as irl_utils
tfp = tfp.substrates.jax
tfd = tfp.distributions
def visualize(sys, qps, save_path: str = None):
"""Renders a 3D visualization of the environment."""
if save_path:
html.save_html(save_path, sys, qps, make_dir=True)
return HTML(html.render(sys, qps))
if 'COLAB_TPU_ADDR' in os.environ:
from jax.tools import colab_tpu
colab_tpu.setup_tpu()
# + id="NaJDZqhCLovU"
#@title Visualizing pre-included Brax environments
env_name = "ant" # @param ['ant', 'halfcheetah', 'ant_cheetah', 'uni_ant', 'bi_ant']
env_space = "vel" # @param ['vel', 'pos', 'ang']
exp_name = "smm_multimode" # @param ['smm', 'smm_multimode', 'smm_multimode3', 'smm_maxent']
algo_name = "gail2" # @param ['gail', 'airl', 'gail2', 'fairl']
disc_arch = "mlp" # @param ['linear', 'mlp']
logits_clip_range = 10.0# @param {'type': 'number'}
normalize_obs_for_disc = False # @param {'type': 'boolean'}
balance_data_for_disc = True # @param {'type': 'boolean'}
env_indices = {
'vel': { # x-y velocity
'ant': (13, 14),
'humanoid': (22, 23),
'halfcheetah': (11,),
'uni_ant': (('vel:torso_ant1', 0),('vel:torso_ant1', 1)),
'bi_ant': (('vel:torso_ant1', 0),('vel:torso_ant2', 0)),
},
'ang': { # angular velocity
'ant': (17,),
'uni_ant': (('ang:torso_ant1', 2),),
},
}[env_space][env_name]
base_env_fn = composer.create_fn(env_name=env_name)
base_env = base_env_fn()
disc_arch = {
'linear': (),
'mlp': (32, 32),
}[disc_arch]
if exp_name in ['smm', 'smm_multimode', 'smm_multimode3', 'smm_maxent']:
disc_fn = functools.partial(
irl_utils.IRLDiscriminator,
input_size=len(env_indices),
obs_indices=env_indices,
include_action=False,
arch=disc_arch,
logits_clip_range=logits_clip_range,
)
else:
raise NotImplementedError(exp_name)
disc = disc_fn(reward_type=algo_name,
normalize_obs=normalize_obs_for_disc,
balance_data=balance_data_for_disc,
env=base_env)
extra_params = disc.init_model(rng=jax.random.PRNGKey(seed=0))
env_fn = irl_utils.create_fn(env_name=env_name, disc=disc)
env = env_fn()
# Visualize in 3D
env = env_fn()
jit_env_reset = jax.jit(env.reset)
state = jit_env_reset(rng=jax.random.PRNGKey(seed=0))
clear_output() # clear out jax.lax warning before rendering
visualize(env.sys, [state.qp])
# + id="rM7nNiXJU-4s"
#@title Generate and visualize target data distribution p\*(s, a) or p\*(s)
N = 250# @param{type: 'integer'}
def draw_2d_uniform(rng, N, low, high):
rng, key = jax.random.split(rng)
dist = tfd.Uniform(low=jnp.array(low), high=jnp.array(high))
data_2d = dist.sample(sample_shape=N, seed=key)
return rng, data_2d
rng = jax.random.PRNGKey(seed=0)
if exp_name == 'smm':
rng, target_data_2d = draw_2d_uniform(rng, N=N, low=[-6.,-0.5], high=[-4.,0.5])
target_data = target_data_2d[..., :len(env_indices)]
elif exp_name == 'smm_multimode':
rng, sample1 = draw_2d_uniform(rng, N=N, low=[-6.,-0.5], high=[-4.,0.5])
rng, sample2 = draw_2d_uniform(rng, N=N, low=[4.,-0.5], high=[6.,0.5])
target_data_2d = jnp.concatenate([sample1, sample2], axis=0)
target_data = target_data_2d[..., :len(env_indices)]
elif exp_name == 'smm_multimode3':
rng, sample1 = draw_2d_uniform(rng, N=N, low=[-2.,-2.], high=[-1.,-1.])
rng, sample2 = draw_2d_uniform(rng, N=N, low=[1.,-2.], high=[2.,-1.])
rng, sample3 = draw_2d_uniform(rng, N=N, low=[-0.5,1.], high=[0.5,2.])
target_data_2d = jnp.concatenate([sample1, sample2, sample3], axis=0)
target_data = target_data_2d[..., :len(env_indices)]
elif exp_name == 'smm_maxent':
rng, target_data_2d = draw_2d_uniform(rng, N=N, low=[-6.,-6.], high=[6.,6.])
target_data = target_data_2d[..., :len(env_indices)]
else:
raise NotImplementedError(exp_name)
disc.set_target_data(target_data)
print(f'target_data={target_data.shape}')
lim = jnp.max(jnp.abs(target_data_2d)) + 0.5
plt.scatter(x=target_data_2d[:, 0],
y=target_data_2d[:, 1],
c=jnp.array([0,0,1]))
plt.xlim((-lim, lim))
plt.ylim((-lim, lim))
plt.title('target (e.g. x-y velocities)')
plt.show()
# + id="4vgMSWODfyMC"
#@title Training some pre-included Brax environments
num_timesteps_multiplier = 2# @param {type: 'integer'}
# We determined some reasonable hyperparameters offline and share them here.
n = num_timesteps_multiplier
train_fn = functools.partial(
ppo.train, num_timesteps = 50_000_000*n, log_frequency = 20,
reward_scaling = 10, episode_length = 1000, normalize_observations = True,
action_repeat = 1, unroll_length = 5, num_minibatches = 32,
num_update_epochs = 4, discounting = 0.95, learning_rate = 3e-4,
entropy_cost = 1e-2, num_envs = 2048, batch_size = 1024
)
times = [datetime.now()]
plotdata = {}
plotkeys = ['eval/episode_reward', 'losses/disc_loss', 'losses/total_loss',
'losses/policy_loss', 'losses/value_loss', 'losses/entropy_loss']
grid = jnp.linspace(-6.5, 6.5, 25)
xgrid, ygrid = jnp.meshgrid(grid, grid)
datagrid = jnp.concatenate([xgrid.reshape(-1, 1), ygrid.reshape(-1, 1)], axis=-1)
lim = jnp.max(jnp.abs(datagrid)) + 0.5
def progress(num_steps, metrics, optimizer_params):
times.append(datetime.now())
for key, v in metrics.items():
plotdata[key] = plotdata.get(key, dict(x=[], y=[]))
plotdata[key]['x'] += [num_steps]
plotdata[key]['y'] += [v]
clear_output(wait=True)
num_figs = len(plotkeys) + 1
fig, axs = plt.subplots(ncols=num_figs, figsize=(3.5*num_figs, 3))
# plot learning curves
for i, key in enumerate(plotkeys):
if key in plotdata:
axs[i].plot(plotdata[key]['x'], plotdata[key]['y'])
axs[i].set(xlabel='# environment steps', ylabel=key)
axs[i].set_xlim([0, train_fn.keywords['num_timesteps']])
# plot discriminator visualization
distgrid = disc.dist(datagrid[..., :len(env_indices)], params=optimizer_params['extra'])
probsgrid = jax.nn.sigmoid(distgrid.logits)
print(f'disc probs: max={probsgrid.max()}, min={probsgrid.min()}')
colors = jnp.clip(jnp.array([[-2, 0, 2]]) * (probsgrid-0.5), a_min=0)
axs[-1].scatter(x=datagrid[:, 0],
y=datagrid[:, 1],
c=colors)
axs[-1].set_xlim((-lim, lim))
axs[-1].set_ylim((-lim, lim))
axs[-1].set(title='discriminator output (red=0, black=0.5, blue=1)')
fig.tight_layout()
plt.show()
extra_loss_fns = dict(disc_loss=disc.disc_loss_fn)
inference_fn, params, _ = train_fn(environment_fn=env_fn,
progress_fn=progress, extra_params=extra_params,
extra_loss_fns=extra_loss_fns)
print(f'time to jit: {times[1] - times[0]}')
print(f'time to train: {times[-1] - times[1]}')
# + id="RNMLEyaTspEM"
#@title Visualizing a trajectory of the learned inference function
seed = 0 # @param {'type': 'integer'}
save_path = None # @param {'type': 'raw'}
save_path = save_path.format(
date=datetime.now().strftime('%Y%m%d'),
env_space=env_space,
env_name=env_name,
exp_name=exp_name,
algo_name=algo_name) if save_path else save_path
jit_env_step = jax.jit(env.step)
jit_inference_fn = jax.jit(inference_fn)
qps, states = [], []
state = env.reset(rng=jax.random.PRNGKey(seed=seed))
while not state.done:
qps.append(state.qp)
states.append(state)
act = jit_inference_fn(params, state.obs, state.rng)
state = jit_env_step(state, act, params[0], params[-1])
visualize(env.sys, qps, save_path=save_path)
# + id="p5eWOxg7RmQQ"
#@title Visualizing skills of the learned inference function in 2D plot
import numpy as np
from itertools import product
num_samples = 5 # @param {type: 'integer'}
time_subsampling = 10 # @param {type: 'integer'}
time_last_n = 500 # @param {type: 'integer'}
seed = 0 # @param {type: 'integer'}
# Reset and run environment
batch_env = env_fn(batch_size=num_samples)
state = batch_env.reset(
jnp.array([jax.random.PRNGKey(seed+i) for i in range(num_samples)]))
states = [state]
jit_step = jax.jit(batch_env.step)
jit_inference_fn = jax.jit(inference_fn)
while not state.done.all():
act = jit_inference_fn(params, state.obs, state.rng[0])
state = jit_step(state, act, params[0], params[-1])
states.append(state)
# Get indices of interest
obses_full = jnp.stack([state.obs for state in states],
axis=0)
obses = obses_full[-time_last_n:][::time_subsampling]
env_vars = batch_env.disc.index_obs(obses) # [T, num_samples, #env_indices]
target_vars = target_data
if env_vars.shape[-1] == 1:
env_vars = jnp.concatenate([env_vars, jnp.zeros(env_vars.shape)], axis=-1)
target_vars = jnp.concatenate([target_vars, jnp.zeros(target_vars.shape)], axis=-1)
print(f'env_vars.shape={env_vars.shape}')
print(f'target_vars.shape={target_vars.shape}')
env_vars_flat = env_vars.reshape(-1, 2)
target_vars_flat = target_vars.reshape(-1, 2)
# Plot
lim = jnp.max(jnp.abs(
jnp.concatenate([env_vars_flat, target_vars_flat], axis=0))) + 0.5
fig, axs = plt.subplots(ncols=2, figsize=(3.5*3, 3))
fig.tight_layout()
axs[0].set(title='agent policy')
axs[0].set_xlim([-lim, lim])
axs[0].set_ylim([-lim, lim])
axs[0].scatter(x=env_vars_flat[:, 0], y=env_vars_flat[:, 1], c=[1,0,0], alpha=0.3)
axs[1].set(title='target')
axs[1].set_xlim([-lim, lim])
axs[1].set_ylim([-lim, lim])
axs[1].scatter(x=target_vars_flat[:, 0], y=target_vars_flat[:, 1], c=[0,0,1], alpha=0.3)
plt.show()
|
notebooks/braxlines/irl_smm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob
import pandas as pd
import numpy as np
from datetime import datetime, timedelta
def create_Summary(project,row):
project['sentDate'] = pd.to_datetime(project['sentDate'],format='%Y-%m-%d %H:%M:%S.%f')
project = project[project['sentDate'].notnull()]
project = project.sort_values('sentDate')
df = pd.DataFrame()
df.loc[row,'Project'] = project['projectId'][project.index[0]]
project.groupby('fromOrganizationId')
df.loc[row,'Number_of_Organizations'] = len(project.groupby('fromOrganizationId').count())
df.loc[row,'Project_Duration'] = int((project['sentDate'][project.index[-1]] - project['sentDate'][project.index[0]]).days)
project.groupby('fromUserId')
df.loc[row,'Number_of_Users'] = len(project.groupby('fromUserId').count())
project.groupby('correspondenceTypeId')
df.loc[row,'Kinds_Of_Correspondence'] = len(project.groupby('correspondenceTypeId').count())
return df
# +
# Reading all the .csv files into one
path =r'C:\Users\212342133\Documents\Python Scripts\1_Exploration\Aconex\Data+science+-+test+(June+2017)\Data science - test (June 2017)\correspondence_data'
allFiles = glob.glob(path + "/*.csv")
frame = pd.DataFrame()
list_ = []
for index,file_ in enumerate(allFiles):
df = pd.read_csv(file_, index_col=None, header=0)
df = create_Summary(df,index)
list_.append(df)
frame_summary = pd.concat(list_)
# -
|
Aconex.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Climate Change and Deaths Caused by Lung Cancer Analysis
# In this analysis, we would like to see the correlation between climate change and deaths caused by lung cancer, specifically at the top 5 countries with the highest population in the world.
# 1. China
# 2. India
# 3. United States
# 4. Indonesia
# 5. Brazil
#
#
# %matplotlib inline
# Dependencies and Set Up
import pandas as pd
import numpy as np
import requests
import json
import matplotlib.pyplot as plt
from scipy import stats
# Read csv for temperature by countries from 1991 to 2016
temp_china = pd.read_csv("./Resources/temperature_1991_2016_China.csv")
temp_india = pd.read_csv("./Resources/temperature_1991_2016_India.csv")
temp_usa = pd.read_csv("./Resources/temperature_1991_2016_USA.csv")
temp_indonesia = pd.read_csv("./Resources/temperature_1991_2016_Indonesia.csv")
temp_brazil = pd.read_csv("./Resources/temperature_1991_2016_Brazil.csv")
# Check and print the temperature data (China)
temp_china.head()
# +
# Grouping the DataFrame by year
temp_china_by_year = temp_china.groupby(["Year"])
# Calculate the average temperature by year and print in DataFrame
temp_china_by_year_mean = pd.DataFrame(temp_china_by_year["Temperature - (Celsius)"].mean())
temp_china_by_year_mean.head()
# +
# # Plot the graph based on mean temperature in China by year (1991 to 2016)
# plt.plot(temp_china_by_year_mean.index, temp_china_by_year_mean["Temperature - (Celsius)"],
# color="green")
# plt.show()
# +
# Perform a linear regression on the temperature year by year
year = temp_china_by_year_mean.index
temp = temp_china_by_year_mean["Temperature - (Celsius)"]
(slope, intercept, r_value, p_value, std_err) = stats.linregress(year, temp)
# Get regression values
regress_values = year * slope + intercept
# print(regress_values)
# +
# Create plot for temperature in China from 1991 to 2016 with the line regression
plt.plot(temp_china_by_year_mean.index, temp_china_by_year_mean["Temperature - (Celsius)"],
color="green")
plt.plot(year, regress_values, color="red")
plt.title("Temperature (C) in China from 1991 to 2016")
plt.xlabel("Year")
plt.ylabel("Temperature (C)")
# Save the image of the plot in "Images" folder
plt.savefig("./Images/temp_china.png")
plt.show()
# -
# Check and print the temperature data (India)
temp_india.head()
# +
# Grouping the DataFrame by year
temp_india_by_year = temp_india.groupby(["Year"])
# Calculate the average temperature by year and print in DataFrame
temp_india_by_year_mean = pd.DataFrame(temp_india_by_year["Temperature - (Celsius)"].mean())
temp_india_by_year_mean.head()
# +
# Perform a linear regression on the temperature year by year
year = temp_india_by_year_mean.index
temp = temp_india_by_year_mean["Temperature - (Celsius)"]
(slope, intercept, r_value, p_value, std_err) = stats.linregress(year, temp)
# Get regression values
regress_values = year * slope + intercept
# print(regress_values)
# +
# Create plot for temperature in China from 1991 to 2016 with the line regression
plt.plot(temp_india_by_year_mean.index, temp_india_by_year_mean["Temperature - (Celsius)"],
color="orange")
plt.plot(year, regress_values, color="blue")
plt.title("Temperature (C) in India from 1991 to 2016")
plt.xlabel("Year")
plt.ylabel("Temperature (C)")
# Save the image of the plot in "Images" folder
plt.savefig("./Images/temp_india.png")
plt.show()
# -
# Check and print the temperature data (USA)
temp_usa.head()
# +
# Grouping the DataFrame by year
temp_usa_by_year = temp_usa.groupby(["Year"])
# Calculate the average temperature by year and print in DataFrame
temp_usa_by_year_mean = pd.DataFrame(temp_usa_by_year["Temperature - (Celsius)"].mean())
temp_usa_by_year_mean.head()
# +
# Perform a linear regression on the temperature year by year
year = temp_usa_by_year_mean.index
temp = temp_usa_by_year_mean["Temperature - (Celsius)"]
(slope, intercept, r_value, p_value, std_err) = stats.linregress(year, temp)
# Get regression values
regress_values = year * slope + intercept
# print(regress_values)
# +
# Create plot for temperature in China from 1991 to 2016 with the line regression
plt.plot(temp_usa_by_year_mean.index, temp_usa_by_year_mean["Temperature - (Celsius)"],
color="orange")
plt.plot(year, regress_values, color="blue")
plt.title("Temperature (C) in United States from 1991 to 2016")
plt.xlabel("Year")
plt.ylabel("Temperature (C)")
# Save the image of the plot in "Images" folder
plt.savefig("./Images/temp_usa.png")
plt.show()
# -
# Check and print the temperature data (Indonesia)
temp_indonesia.head()
# +
# Grouping the DataFrame by year
temp_indonesia_by_year = temp_indonesia.groupby(["Year"])
# Calculate the average temperature by year and print in DataFrame
temp_indonesia_by_year_mean = pd.DataFrame(temp_indonesia_by_year["Temperature - (Celsius)"].mean())
temp_indonesia_by_year_mean.head()
# +
# Perform a linear regression on the temperature year by year
year = temp_indonesia_by_year_mean.index
temp = temp_indonesia_by_year_mean["Temperature - (Celsius)"]
(slope, intercept, r_value, p_value, std_err) = stats.linregress(year, temp)
# Get regression values
regress_values = year * slope + intercept
# print(regress_values)
# +
# Create plot for temperature in China from 1991 to 2016 with the line regression
plt.plot(temp_indonesia_by_year_mean.index, temp_indonesia_by_year_mean["Temperature - (Celsius)"],
color="orange")
plt.plot(year, regress_values, color="blue")
plt.title("Temperature (C) in Indonesia from 1991 to 2016")
plt.xlabel("Year")
plt.ylabel("Temperature (C)")
# Save the image of the plot in "Images" folder
plt.savefig("./Images/temp_indonesia.png")
plt.show()
# -
# Check and print the temperature data (Brazil)
temp_brazil.head()
# +
# Grouping the DataFrame by year
temp_brazil_by_year = temp_brazil.groupby(["Year"])
# Calculate the average temperature by year and print in DataFrame
temp_brazil_by_year_mean = pd.DataFrame(temp_brazil_by_year["Temperature - (Celsius)"].mean())
temp_brazil_by_year_mean.head()
# +
# Perform a linear regression on the temperature year by year
year = temp_brazil_by_year_mean.index
temp = temp_brazil_by_year_mean["Temperature - (Celsius)"]
(slope, intercept, r_value, p_value, std_err) = stats.linregress(year, temp)
# Get regression values
regress_values = year * slope + intercept
# print(regress_values)
# +
# Create plot for temperature in China from 1991 to 2016 with the line regression
plt.plot(temp_brazil_by_year_mean.index, temp_brazil_by_year_mean["Temperature - (Celsius)"],
color="orange")
plt.plot(year, regress_values, color="blue")
plt.title("Temperature (C) in Brazil from 1991 to 2016")
plt.xlabel("Year")
plt.ylabel("Temperature (C)")
# Save the image of the plot in "Images" folder
plt.savefig("./Images/temp_brazil.png")
plt.show()
# -
# Read the csv for the annual CO2 emission by country
CO2_emission = pd.read_csv("./Resources/annual_co2_emissions_by_region.csv")
CO2_emission.head()
# Rename the column name
CO2_emission = CO2_emission.rename(
columns = {"Entity": "Country", "Annual CO2 emissions (tonnes )": "CO2 emissions (tonnes)"})
CO2_emission.head()
# +
# Extract only China data
columns = ["Country", "Year", "CO2 emissions (tonnes)"]
CO2_emission_china = CO2_emission.loc[(CO2_emission["Country"] == "China"), columns]
CO2_emission_china.head()
# +
# Extract China data for 1991 to 2016 only
CO2_emission_china = CO2_emission_china.set_index("Year")
years = np.arange(1991, 2017, 1)
years_91_16 = []
for year in years:
years_91_16.append(year)
# years_91_16
CO2_emission_china = CO2_emission_china.loc[years_91_16]
CO2_emission_china.head(10)
# +
# Extract only India data
columns = ["Country", "Year", "CO2 emissions (tonnes)"]
CO2_emission_india = CO2_emission.loc[(CO2_emission["Country"] == "India"), columns]
CO2_emission_india.head()
# +
# Extract India data for 1991 to 2016 only
CO2_emission_india = CO2_emission_india.set_index("Year")
CO2_emission_india = CO2_emission_india.loc[years_91_16]
CO2_emission_india.head(10)
# +
# Extract only United States data
columns = ["Country", "Year", "CO2 emissions (tonnes)"]
CO2_emission_usa = CO2_emission.loc[(CO2_emission["Country"] == "United States"), columns]
CO2_emission_usa.head()
# +
# Extract United States data for 1991 to 2016 only
CO2_emission_usa = CO2_emission_usa.set_index("Year")
CO2_emission_usa = CO2_emission_usa.loc[years_91_16]
CO2_emission_usa.head(10)
# +
# Extract only Indonesia data
columns = ["Country", "Year", "CO2 emissions (tonnes)"]
CO2_emission_indonesia = CO2_emission.loc[(CO2_emission["Country"] == "Indonesia"), columns]
CO2_emission_indonesia.head()
# +
# Extract Indonesia data for 1991 to 2016 only
CO2_emission_indonesia = CO2_emission_indonesia.set_index("Year")
CO2_emission_indonesia = CO2_emission_indonesia.loc[years_91_16]
CO2_emission_indonesia.head(10)
# +
# Extract only Brazil data
columns = ["Country", "Year", "CO2 emissions (tonnes)"]
CO2_emission_brazil = CO2_emission.loc[(CO2_emission["Country"] == "Brazil"), columns]
CO2_emission_brazil.head()
# +
# Extract Brazil data for 1991 to 2016 only
CO2_emission_brazil = CO2_emission_brazil.set_index("Year")
CO2_emission_brazil = CO2_emission_brazil.loc[years_91_16]
CO2_emission_brazil.head(10)
# -
# Read the csv for total cancer deaths by cancer types
cancer_deaths = pd.read_csv("./Resources/total_cancer_deaths_by_type.csv")
cancer_deaths.head()
# Seeing the list of column names
list(cancer_deaths.columns)
# Extracting the columns for Country/Entity, Year, and deaths because of lung cancer
lung_cancer_deaths = cancer_deaths.loc[:, ["Entity", "Year", "Tracheal, bronchus, and lung cancer (deaths)"]]
lung_cancer_deaths.head()
# Rename the column name
lung_cancer_deaths = lung_cancer_deaths.rename(columns = {"Entity": "Country"})
lung_cancer_deaths.head()
# +
# Extract the deaths caused by lung cancer for China only
lung_cancer_deaths_china = lung_cancer_deaths.loc[lung_cancer_deaths["Country"] == "China"]
# Set index as year and extract the deaths caused by lung cancer in China for year 1991 to 2016 only
lung_cancer_deaths_china = lung_cancer_deaths_china.set_index("Year")
lung_cancer_deaths_china = lung_cancer_deaths_china.loc[years_91_16]
lung_cancer_deaths_china.head(10)
# +
# Extract the deaths caused by lung cancer for India only
lung_cancer_deaths_india = lung_cancer_deaths.loc[lung_cancer_deaths["Country"] == "India"]
# Set index as year and extract the deaths caused by lung cancer in India for year 1991 to 2016 only
lung_cancer_deaths_india = lung_cancer_deaths_india.set_index("Year")
lung_cancer_deaths_india = lung_cancer_deaths_india.loc[years_91_16]
lung_cancer_deaths_india.head(10)
# +
# Extract the deaths caused by lung cancer for United States only
lung_cancer_deaths_usa = lung_cancer_deaths.loc[lung_cancer_deaths["Country"] == "United States"]
# Set index as year and extract the deaths caused by lung cancer in United States for year 1991 to 2016 only
lung_cancer_deaths_usa = lung_cancer_deaths_usa.set_index("Year")
lung_cancer_deaths_usa = lung_cancer_deaths_usa.loc[years_91_16]
lung_cancer_deaths_usa.head(10)
# +
# Extract the deaths caused by lung cancer for Indonesia only
lung_cancer_deaths_indonesia = lung_cancer_deaths.loc[lung_cancer_deaths["Country"] == "Indonesia"]
# Set index as year and extract the deaths caused by lung cancer in Indonesia for year 1991 to 2016 only
lung_cancer_deaths_indonesia = lung_cancer_deaths_indonesia.set_index("Year")
lung_cancer_deaths_indonesia = lung_cancer_deaths_indonesia.loc[years_91_16]
lung_cancer_deaths_indonesia.head(10)
# +
# Extract the deaths caused by lung cancer for Brazil only
lung_cancer_deaths_brazil = lung_cancer_deaths.loc[lung_cancer_deaths["Country"] == "Brazil"]
# Set index as year and extract the deaths caused by lung cancer in Brazil for year 1991 to 2016 only
lung_cancer_deaths_brazil = lung_cancer_deaths_brazil.set_index("Year")
lung_cancer_deaths_brazil = lung_cancer_deaths_brazil.loc[years_91_16]
lung_cancer_deaths_brazil.head(10)
# -
# Read the csv for total population by region
pop = pd.read_csv("./Resources/total_population_by_region.csv")
pop.head()
# +
# Extract the data for China and year from 1991 to 2016 only
pop_91_16 = pop.loc[:,["Country Name",
"1991", "1992", "1993", "1994", "1995",
"1996", "1997", "1998", "1999", "2000",
"2001", "2002", "2003", "2004", "2005",
"2006", "2007", "2008", "2009", "2010",
"2011", "2012", "2013", "2014", "2015", "2016"]]
# Set index as Country
pop_91_16 = pop_91_16.set_index("Country Name")
# Transpose the columns and rows
pop_91_16 = pd.DataFrame.transpose(pop_91_16)
pop_91_16.head()
# -
pop_91_16 = pop_91_16.rename_axis("Year", axis=1)
pop_91_16.head()
# Extract the population data for China only and rename the column name to "Population"
pop_china = pop_91_16.loc[:, ["China"]]
pop_china = pop_china.rename(columns = {"China": "Population"})
pop_china.index = pop_china.index.astype("int64")
pop_china.head()
# Extract the population data for India only and rename the column name to "Population"
pop_india = pop_91_16.loc[:, ["India"]]
pop_india = pop_india.rename(columns = {"India": "Population"})
pop_india.index = pop_india.index.astype("int64")
pop_india.head()
# Extract the population data for United States only and rename the column name to "Population"
pop_usa = pop_91_16.loc[:, ["United States"]]
pop_usa = pop_usa.rename(columns = {"United States": "Population"})
pop_usa.index = pop_usa.index.astype("int64")
pop_usa.head()
# Extract the population data for Indonesia only and rename the column name to "Population"
pop_indonesia = pop_91_16.loc[:, ["Indonesia"]]
pop_indonesia = pop_indonesia.rename(columns = {"Indonesia": "Population"})
pop_indonesia.index = pop_indonesia.index.astype("int64")
pop_indonesia.head()
# Extract the population data for Brazil only and rename the column name to "Population"
pop_brazil = pop_91_16.loc[:, ["Brazil"]]
pop_brazil = pop_brazil.rename(columns = {"Brazil": "Population"})
pop_brazil.index = pop_brazil.index.astype("int64")
pop_brazil.head()
lung_cancer_deaths_china.head()
lung_cancer_deaths_china = lung_cancer_deaths_china.rename_axis(index=None, columns="Year")
lung_cancer_deaths_china.head()
# +
# Merge population data with the total deaths caused by lung cancer deaths, to get the percentage of people
# that were died because of lung cancer in each country
lung_cancer_deaths_total_pop_china = pop_china.merge(lung_cancer_deaths_china, how="outer",
left_index=True, right_index=True)
lung_cancer_deaths_pct_china = \
lung_cancer_deaths_total_pop_china["Tracheal, bronchus, and lung cancer (deaths)"] / \
lung_cancer_deaths_total_pop_china["Population"] * 100
lung_cancer_deaths_total_pop_china["Tracheal, bronchus, and lung cancer deaths (%)"] = lung_cancer_deaths_pct_china
# The following output is the percentage table.
lung_cancer_deaths_total_pop_china
# +
# Plot both CO2 emission and lung cancer deaths data for China in 2 graphs but side by side and share the x-axis
years = np.arange(1991, 2017, 1)
years_label = []
for year in years:
years_label.append(year)
fig, [ax1,ax2] = plt.subplots(1,2, figsize=(16,6), sharex=True)
ax1.plot(years, CO2_emission_china["CO2 emissions (tonnes)"],
color="red", linewidth=1)
ax1.set_xlabel("Year")
ax1.set_ylabel("CO2 Emissions in China (Tonnes)", color="red")
ax1.set_xticks(years_label)
ax1.set_xticklabels(years_label, rotation=45)
ax1.set_title("CO2 Emission in China from 1991 to 2016")
# ax1.grid()
ax2.plot(years, lung_cancer_deaths_total_pop_china["Tracheal, bronchus, and lung cancer deaths (%)"],
color="blue", linewidth=1)
ax2.set_xlabel("Year")
ax2.set_ylabel("Percentage of Tracheal, bronchus, and lung cancer deaths", color="blue")
ax2.set_xticks(years_label)
ax2.set_xticklabels(years_label, rotation=45)
ax2.set_title("Deaths Caused by Tracheal, Bronchus, Lung Cancer in China from 1991 to 2016 (%)")
# ax2.grid()
fig.tight_layout(pad=3.0)
plt.show()
# -
lung_cancer_deaths_india = lung_cancer_deaths_india.rename_axis(index=None, columns="Year")
lung_cancer_deaths_india.head()
# +
# Merge population data with the total deaths caused by lung cancer deaths, to get the percentage of people
# that died because of lung cancer in each country
lung_cancer_deaths_total_pop_india = pop_india.merge(lung_cancer_deaths_india, how="outer",
left_index=True, right_index=True)
lung_cancer_deaths_pct_india = \
lung_cancer_deaths_total_pop_india["Tracheal, bronchus, and lung cancer (deaths)"] / \
lung_cancer_deaths_total_pop_india["Population"] * 100
lung_cancer_deaths_total_pop_india["Tracheal, bronchus, and lung cancer deaths (%)"] = lung_cancer_deaths_pct_india
# The following output is the percentage table.
lung_cancer_deaths_total_pop_india
# +
# Plot both CO2 emission and lung cancer deaths data for India in 2 graphs but side by side and share the x-axis
years = np.arange(1991, 2017, 1)
years_label = []
for year in years:
years_label.append(year)
fig, [ax1,ax2] = plt.subplots(1,2, figsize=(16,6), sharex=True)
ax1.plot(years, CO2_emission_india["CO2 emissions (tonnes)"],
color="red", linewidth=1)
ax1.set_xlabel("Year")
ax1.set_ylabel("CO2 Emissions in India (Tonnes)", color="red")
ax1.set_xticks(years_label)
ax1.set_xticklabels(years_label, rotation=45)
ax1.set_title("CO2 Emission in India from 1991 to 2016")
# ax1.grid()
ax2.plot(years, lung_cancer_deaths_total_pop_india["Tracheal, bronchus, and lung cancer deaths (%)"],
color="blue", linewidth=1)
ax2.set_xlabel("Year")
ax2.set_ylabel("Percentage of Tracheal, bronchus, and lung cancer deaths", color="blue")
ax2.set_xticks(years_label)
ax2.set_xticklabels(years_label, rotation=45)
ax2.set_title("Deaths Caused by Tracheal, Bronchus, Lung Cancer in India from 1991 to 2016 (%)")
# ax2.grid()
fig.tight_layout(pad=3.0)
plt.show()
# -
lung_cancer_deaths_usa = lung_cancer_deaths_usa.rename_axis(index=None, columns="Year")
lung_cancer_deaths_usa.head()
# +
# Merge population data with the total deaths caused by lung cancer deaths, to get the percentage of people
# that died because of lung cancer in each country
lung_cancer_deaths_total_pop_usa = pop_usa.merge(lung_cancer_deaths_usa, how="outer",
left_index=True, right_index=True)
lung_cancer_deaths_pct_usa = \
lung_cancer_deaths_total_pop_usa["Tracheal, bronchus, and lung cancer (deaths)"] / \
lung_cancer_deaths_total_pop_usa["Population"] * 100
lung_cancer_deaths_total_pop_usa["Tracheal, bronchus, and lung cancer deaths (%)"] = lung_cancer_deaths_pct_usa
# The following output is the percentage table.
lung_cancer_deaths_total_pop_usa
# +
# Plot both CO2 emission and lung cancer deaths data for United States in 2 graphs but side by side and share the x-axis
years = np.arange(1991, 2017, 1)
years_label = []
for year in years:
years_label.append(year)
fig, [ax1,ax2] = plt.subplots(1,2, figsize=(16,6), sharex=True)
ax1.plot(years, CO2_emission_usa["CO2 emissions (tonnes)"],
color="red", linewidth=1)
ax1.set_xlabel("Year")
ax1.set_ylabel("CO2 Emissions in USA (Tonnes)", color="red")
ax1.set_xticks(years_label)
ax1.set_xticklabels(years_label, rotation=45)
ax1.set_title("CO2 Emission in USA from 1991 to 2016")
# ax1.grid(axis="y")
ax2.plot(years, lung_cancer_deaths_total_pop_usa["Tracheal, bronchus, and lung cancer deaths (%)"],
color="blue", linewidth=1)
ax2.set_xlabel("Year")
ax2.set_ylabel("Percentage of Tracheal, bronchus, and lung cancer deaths", color="blue")
ax2.set_xticks(years_label)
ax2.set_xticklabels(years_label, rotation=45)
ax2.set_title("Deaths Caused by Tracheal, Bronchus, Lung Cancer in USA from 1991 to 2016 (%)")
# ax2.grid()
fig.tight_layout(pad=3.0)
plt.show()
# -
lung_cancer_deaths_indonesia = lung_cancer_deaths_indonesia.rename_axis(index=None, columns="Year")
lung_cancer_deaths_indonesia.head()
# +
# Merge population data with the total deaths caused by lung cancer deaths, to get the percentage of people
# that died because of lung cancer in each country
lung_cancer_deaths_total_pop_indonesia = pop_indonesia.merge(lung_cancer_deaths_indonesia, how="outer",
left_index=True, right_index=True)
lung_cancer_deaths_pct_indonesia = \
lung_cancer_deaths_total_pop_indonesia["Tracheal, bronchus, and lung cancer (deaths)"] / \
lung_cancer_deaths_total_pop_indonesia["Population"] * 100
lung_cancer_deaths_total_pop_indonesia["Tracheal, bronchus, and lung cancer deaths (%)"] = lung_cancer_deaths_pct_indonesia
# The following output is the percentage table.
lung_cancer_deaths_total_pop_indonesia
# +
# Plot both CO2 emission and lung cancer deaths data for Indonesia in 2 graphs but side by side and share the x-axis
years = np.arange(1991, 2017, 1)
years_label = []
for year in years:
years_label.append(year)
fig, [ax1,ax2] = plt.subplots(1,2, figsize=(16,6), sharex=True)
ax1.plot(years, CO2_emission_indonesia["CO2 emissions (tonnes)"],
color="red", linewidth=1)
ax1.set_xlabel("Year")
ax1.set_ylabel("CO2 Emissions in Indonesia (Tonnes)", color="red")
ax1.set_xticks(years_label)
ax1.set_xticklabels(years_label, rotation=45)
ax1.set_title("CO2 Emission in Indonesia from 1991 to 2016")
# ax1.grid()
ax2.plot(years, lung_cancer_deaths_total_pop_indonesia["Tracheal, bronchus, and lung cancer deaths (%)"],
color="blue", linewidth=1)
ax2.set_xlabel("Year")
ax2.set_ylabel("Percentage of Tracheal, bronchus, and lung cancer deaths", color="blue")
ax2.set_xticks(years_label)
ax2.set_xticklabels(years_label, rotation=45)
ax2.set_title("Deaths Caused by Tracheal, Bronchus, Lung Cancer in Indonesia from 1991 to 2016 (%)")
# ax2.grid()
fig.tight_layout(pad=3.0)
plt.show()
# -
lung_cancer_deaths_brazil = lung_cancer_deaths_brazil.rename_axis(index=None, columns="Year")
lung_cancer_deaths_brazil.head()
# +
# Merge population data with the total deaths caused by lung cancer deaths, to get the percentage of people
# that died because of lung cancer in each country
lung_cancer_deaths_total_pop_brazil = pop_brazil.merge(lung_cancer_deaths_brazil, how="outer",
left_index=True, right_index=True)
lung_cancer_deaths_pct_brazil = \
lung_cancer_deaths_total_pop_brazil["Tracheal, bronchus, and lung cancer (deaths)"] / \
lung_cancer_deaths_total_pop_brazil["Population"] * 100
lung_cancer_deaths_total_pop_brazil["Tracheal, bronchus, and lung cancer deaths (%)"] = lung_cancer_deaths_pct_brazil
# The following output is the percentage table.
lung_cancer_deaths_total_pop_brazil
# +
# Plot both CO2 emission and lung cancer deaths data for Brazil in 2 graphs but side by side and share the x-axis
years = np.arange(1991, 2017, 1)
years_label = []
for year in years:
years_label.append(year)
fig, [ax1,ax2] = plt.subplots(1,2, figsize=(16,6), sharex=True)
ax1.plot(years, CO2_emission_brazil["CO2 emissions (tonnes)"],
color="red", linewidth=1)
ax1.set_xlabel("Year")
ax1.set_ylabel("CO2 Emissions in Brazil (Tonnes)", color="red")
ax1.set_xticks(years_label)
ax1.set_xticklabels(years_label, rotation=45)
ax1.set_title("CO2 Emission in Brazil from 1991 to 2016")
# ax1.grid()
ax2.plot(years, lung_cancer_deaths_total_pop_brazil["Tracheal, bronchus, and lung cancer deaths (%)"],
color="blue", linewidth=1)
ax2.set_xlabel("Year")
ax2.set_ylabel("Percentage of Tracheal, bronchus, and lung cancer deaths", color="blue")
ax2.set_xticks(years_label)
ax2.set_xticklabels(years_label, rotation=45)
ax2.set_title("Deaths Caused by Tracheal, Bronchus, Lung Cancer in Brazil from 1991 to 2016 (%)")
# ax2.grid()
fig.tight_layout(pad=3.0)
plt.show()
# -
# Export the CO2 emission data to csv file
CO2_emission_china.to_csv("./Results/CO2_emission_china.csv")
CO2_emission_india.to_csv("./Results/CO2_emission_india.csv")
CO2_emission_usa.to_csv("./Results/CO2_emission_usa.csv")
CO2_emission_indonesia.to_csv("./Results/CO2_emission_indonesia.csv")
CO2_emission_brazil.to_csv("./Results/CO2_emission_brazil.csv")
# Export the lung cancer deaths data to csv file
lung_cancer_deaths_total_pop_china.to_csv("./Results/lung_cancer_deaths_china.csv")
lung_cancer_deaths_total_pop_india.to_csv("./Results/lung_cancer_deaths_india.csv")
lung_cancer_deaths_total_pop_usa.to_csv("./Results/lung_cancer_deaths_usa.csv")
lung_cancer_deaths_total_pop_indonesia.to_csv("./Results/lung_cancer_deaths_indonesia.csv")
lung_cancer_deaths_total_pop_brazil.to_csv("./Results/lung_cancer_deaths_brazil.csv")
|
Climate_Change_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Implementing a Recommender System with SageMaker, Tensfolow, and Keras
#
# ## Model: dense_5_Multiply_50_embeddings_10_epochs_dropout
#
# ## _**Making Product - Shoes Recommendations Using Neural Networks and Embeddings**_
#
#
# ## Background
#
# #### In many ways, recommender systems were a catalyst for the current popularity of machine learning. One of Amazon's earliest successes was the "Customers who bought this, also bought..." feature, while the million dollar Netflix Prize spurred research, raised public awareness, and inspired numerous other data science competitions.
#
# #### Recommender systems can utilize a multitude of data sources and ML algorithms, and most combine various unsupervised, supervised, and reinforcement learning techniques into a holistic framework. However, the core component is almost always a model which which predicts a user's rating (or purchase) for a certain item based on that user's historical ratings of similar items as well as the behavior of other similar users. The minimal required dataset for this is a history of user item ratings. In our case, we'll use 1 to 5 star ratings from over 2M Amazon customers. More details on this dataset can be found at its [AWS Public Datasets page](https://s3.amazonaws.com/amazon-reviews-pds/readme.html).
#
# #### Matrix factorization has been the cornerstone of most user-item prediction models. This method starts with the large, sparse, user-item ratings in a single matrix, where users index the rows, and items index the columns. It then seeks to find two lower-dimensional, dense matrices which, when multiplied together, preserve the information and relationships in the larger matrix.
#
# 
#
# ### ** Matrix factorization has been extended and genarlized with deep learning and embeddings. These techniques allows us to introduce non-linearities for enhanced performance and flexibility. This notebook will fit a neural network-based model to generate recommendations for the Amazon dataset. It will start by exploring our data in the notebook and even training a model on a sample of the data. Later we'll expand to the full dataset and fit our model using a SageMaker managed training cluster. We'll then deploy to an endpoint and check our method.
#
# ---
#
# ## Setup
#
# #### _This notebook was created and tested on an ml.p2.xlarge notebook instance._
#
# #### Let's start by specifying:
#
# #### - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# #### - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the `get_execution_role()` call with the appropriate full IAM role arn string(s).
#
# ---
#
# Frame the recommendation system as a rating prediction machine learning problem and create a hybrid architecture that mixes the collaborative and content based filtering approaches:
# - Collaborative part: Predict items ratings in order to recommend to the user items that he is likely to rate high.
# - Content based: use metadata inputs (such as price and title) about items to find similar items to recommend.
#
# ### - Create 2 explicit recommendation engine models based on 2 machine learning architecture using Keras:
# 1. a matrix factorization model
# 2. a deep neural network model.
#
#
# ### Compare the results of the different models and configurations to find the "best" predicting model
#
# ### Used the best model for recommending items to users
# +
### name of model
modname = 'dense_5_Multiply_50_embeddings_10_epochs_dropout'
### size of embedding
embedding_size = 50
### number of epochs
num_epochs = 10
# +
import sys
# !{sys.executable} -m pip install --upgrade pip
# !{sys.executable} -m pip install sagemaker-experiments
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install numpy
# !{sys.executable} -m pip install matplotlib
# !{sys.executable} -m pip install boto3
# !{sys.executable} -m pip install sagemaker
# !{sys.executable} -m pip install pyspark
# !{sys.executable} -m pip install ipython-autotime
# !{sys.executable} -m pip install surprise
# !{sys.executable} -m pip install smart_open
# !{sys.executable} -m pip install pyarrow
# !{sys.executable} -m pip install fastparquet
# +
# Check Jave version
# # !sudo yum -y update
# -
# Need to use Java 1.8.0
# !sudo yum remove jre-1.7.0-openjdk -y
# !java -version
# +
# # !sudo update-alternatives --config java
# -
# !pip install pyarrow fastparquet
# !pip install ipython-autotime
# !pip install tqdm pydot pydotplus pydot_ng
# +
#### To measure all running time
# https://github.com/cpcloud/ipython-autotime
# %load_ext autotime
# +
# %pylab inline
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
import re
import seaborn as sbn
import nltk
import tqdm as tqdm
import sqlite3
import pandas as pd
import numpy as np
from pandas import DataFrame
import string
import pydot
import pydotplus
import pydot_ng
import pickle
import time
import gzip
import os
os.getcwd()
import matplotlib.pyplot as plt
from math import floor,ceil
#from nltk.corpus import stopwords
#stop = stopwords.words("english")
from nltk.stem.porter import PorterStemmer
english_stemmer=nltk.stem.SnowballStemmer('english')
from nltk.tokenize import word_tokenize
from sklearn.metrics import accuracy_score, confusion_matrix,roc_curve, auc,classification_report, mean_squared_error, mean_absolute_error
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.svm import LinearSVC
from sklearn.neighbors import NearestNeighbors
from sklearn.linear_model import LogisticRegression
from sklearn import neighbors
from scipy.spatial.distance import cosine
from sklearn.feature_selection import SelectKBest
from IPython.display import SVG
# Tensorflow
import tensorflow as tf
#Keras
from keras.models import Sequential, Model, load_model, save_model
from keras.callbacks import ModelCheckpoint
from keras.layers import Dense, Activation, Dropout, Input, Masking, TimeDistributed, LSTM, Conv1D, Embedding
from keras.layers import GRU, Bidirectional, BatchNormalization, Reshape
from keras.optimizers import Adam
from keras.layers.core import Reshape, Dropout, Dense
from keras.layers.merge import Multiply, Dot, Concatenate
from keras.layers.embeddings import Embedding
from keras import optimizers
from keras.callbacks import ModelCheckpoint
from keras.utils.vis_utils import model_to_dot
# +
import pandas as pd
import boto3
import sagemaker
from sagemaker import get_execution_role
from sagemaker.session import Session
from sagemaker.analytics import ExperimentAnalytics
import gzip
import json
from pyspark.ml import Pipeline
from pyspark.sql.types import StructField, StructType, StringType, DoubleType
from pyspark.ml.feature import StringIndexer, VectorIndexer, OneHotEncoder, VectorAssembler
from pyspark.sql.functions import *
# spark imports
from pyspark.sql import SparkSession
from pyspark.sql.functions import UserDefinedFunction, explode, desc
from pyspark.sql.types import StringType, ArrayType
from pyspark.ml.evaluation import RegressionEvaluator
import os
import pandas as pd
import pyarrow
import fastparquet
# from pandas_profiling import ProfileReport
# -
# ### Set and Check GPUs
# +
#Session
from keras import backend as K
def set_check_gpu():
cfg = K.tf.ConfigProto()
cfg.gpu_options.per_process_gpu_memory_fraction =1 # allow all of the GPU memory to be allocated
# for 8 GPUs
# cfg.gpu_options.visible_device_list = "0,1,2,3,4,5,6,7" # "0,1"
# for 1 GPU
cfg.gpu_options.visible_device_list = "0"
#cfg.gpu_options.allow_growth = True # # Don't pre-allocate memory; dynamically allocate the memory used on the GPU as-needed
#cfg.log_device_placement = True # to log device placement (on which device the operation ran)
sess = K.tf.Session(config=cfg)
K.set_session(sess) # set this TensorFlow session as the default session for Keras
print("* TF version: ", [tf.__version__, tf.test.is_gpu_available()])
print("* List of GPU(s): ", tf.config.experimental.list_physical_devices() )
print("* Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID";
# set for 8 GPUs
# os.environ["CUDA_VISIBLE_DEVICES"] = "0,1,2,3,4,5,6,7";
# set for 1 GPU
os.environ["CUDA_VISIBLE_DEVICES"] = "0";
# Tf debugging option
tf.debugging.set_log_device_placement(True)
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
# print(tf.config.list_logical_devices('GPU'))
print(tf.config.experimental.list_physical_devices('GPU'))
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
# -
set_check_gpu()
# reset GPU memory& Keras Session
def reset_keras():
try:
del classifier
del model
except:
pass
K.clear_session()
K.get_session().close()
# sess = K.get_session()
cfg = K.tf.ConfigProto()
cfg.gpu_options.per_process_gpu_memory_fraction
# cfg.gpu_options.visible_device_list = "0,1,2,3,4,5,6,7" # "0,1"
cfg.gpu_options.visible_device_list = "0" # "0,1"
cfg.gpu_options.allow_growth = True # dynamically grow the memory used on the GPU
sess = K.tf.Session(config=cfg)
K.set_session(sess) # set this TensorFlow session as the default session for Keras
# ---
# ## Data - https://s3.amazonaws.com/amazon-reviews-pds/tsv/index.txt
#
# ### Explore
#
# Let's start by bringing in our dataset from an S3 public bucket.
# More details on this dataset can be found at its [AWS Public Datasets page](https://s3.amazonaws.com/amazon-reviews-pds/readme.html).
#
# _Note, because this dataset is over a half gigabyte, the load from S3 may take ~10 minutes. Also, since Amazon SageMaker Notebooks start with a 5GB persistent volume by default, and we don't need to keep this data on our instance for long, we'll bring it to the temporary volume (which has up to 20GB of storage)._
# !aws s3 ls s3://amazon-reviews-pds/tsv/
# !mkdir -p ../data
# !aws s3 cp s3://amazon-reviews-pds/tsv/amazon_reviews_us_Shoes_v1_00.tsv.gz ../data
# Let's read the data into a [Pandas DataFrame](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.html) so that we can begin to understand it.
#
# *Note, we'll set `error_bad_lines=False` when reading the file in as there appear to be a very small number of records which would create a problem otherwise.*
# !ls -alh ../data
df = pd.read_csv('../data/amazon_reviews_us_Shoes_v1_00.tsv.gz', delimiter='\t', error_bad_lines=False)
# # ** Amazon product dataset data analysis
# We can see this dataset includes information like:
#
# - `marketplace`: 2-letter country code (in this case all "US").
# - `customer_id`: Random identifier that can be used to aggregate reviews written by a single author.
# - `review_id`: A unique ID for the review.
# - `product_id`: The Amazon Standard Identification Number (ASIN). `http://www.amazon.com/dp/<ASIN>` links to the product's detail page.
# - `product_parent`: The parent of that ASIN. Multiple ASINs (color or format variations of the same product) can roll up into a single parent parent.
# - `product_title`: Title description of the product.
# - `product_category`: Broad product category that can be used to group reviews (in this case this products).
# - `star_rating`: The review's rating (1 to 5 stars).
# - `helpful_votes`: Number of helpful votes for the review.
# - `total_votes`: Number of total votes the review received.
# - `vine`: Was the review written as part of the [Vine](https://www.amazon.com/gp/vine/help) program?
# - `verified_purchase`: Was the review from a verified purchase?
# - `review_headline`: The title of the review itself.
# - `review_body`: The text of the review.
# - `review_date`: The date the review was written.
#
# For this example, let's limit ourselves to `customer_id`, `product_id`, and `star_rating`. Including additional features in our recommendation system could be beneficial, but would require substantial processing (particularly the text data) which would take us beyond the scope of this notebook.
#
# *Note: we'll keep `product_title` on the dataset to help verify our recommendations later in the notebook, but it will not be used in algorithm training.*
# ### Because most people haven't use most products, and people rate fewer products than we actually watch, we'd expect our data to be sparse. Our algorithm should work well with this sparse problem in general, but we may still want to clean out some of the long tail. Let's look at some basic percentiles to confirm.
df.head()
# shape of data
df.shape
# Describing the data set
df.describe()
# checking if there is any null data or not
df.isnull().sum()
# remove numm data
df = df.dropna()
df.head(n=3)
# checking if there is any null data or not
df.isnull().sum()
# Describing the data according to the ratings
df.groupby('star_rating').describe()
df.columns
df = df[['customer_id', 'product_id', 'star_rating', 'product_title', 'helpful_votes']]
# ## Select voted review only
df.shape
df = df[df['helpful_votes'] > 0]
df.shape
4358333-1106199
# +
customers = df['customer_id'].value_counts()
products = df['product_id'].value_counts()
quantiles = [0, 0.01, 0.02, 0.03, 0.04, 0.05, 0.1, 0.25, 0.5, 0.75, 0.9, 0.95, 0.96, 0.97, 0.98, 0.99, 1]
print('customers\n', customers.quantile(quantiles))
print('products\n', products.quantile(quantiles))
# -
#
#
# ### Clean
#
# #### As we can see, only about 5% of customers have rated 2 or more products, and only 10% of products have been rated by 2+ customers.
# ## Let's filter out this long tail.
# +
customers = customers[customers >= 2]
products = products[products >= 2]
reduced_df = df.merge(pd.DataFrame({'customer_id': customers.index})).merge(pd.DataFrame({'product_id': products.index}))
# -
reduced_df.shape
reduced_df.to_csv('../data/amazon_reviews_us_Shoes_v1_00_help_voted_And_cut_lognTail.csv', index_label=False)
# !aws s3 cp ../data/amazon_reviews_us_Shoes_v1_00_help_voted_And_cut_lognTail.csv s3://dse-cohort5-group1/Keras-DeepRecommender-Shoes/data/amazon_reviews_us_Shoes_v1_00_help_voted_And_cut_lognTail.csv
# Now, we'll recreate our customer and product lists since there are customers with more than 5 reviews, but all of their reviews are on products with less than 5 reviews (and vice versa).
customers = reduced_df['customer_id'].value_counts()
products = reduced_df['product_id'].value_counts()
# Next, we'll number each user and item, giving them their own sequential index. This will allow us to hold the information in a sparse format where the sequential indices indicate the row and column in our ratings matrix.
# +
customer_index = pd.DataFrame({'customer_id': customers.index, 'user': np.arange(customers.shape[0])})
product_index = pd.DataFrame({'product_id': products.index,
'item': np.arange(products.shape[0])})
reduced_df = reduced_df.merge(customer_index).merge(product_index)
print(reduced_df.shape)
reduced_df.head()
# -
# Thus I have 1069568 rows and 8 columns dataset.
#
# ## 2. Arrange and clean the data
# Rearrange the columns by relevance and rename column names
review_data = reduced_df
review_data.columns
# +
review_data = review_data[['customer_id', 'product_id', 'star_rating', 'product_title', 'helpful_votes', 'user', 'item']]
review_data.rename(columns={ 'star_rating': 'score','customer_id': 'user_id', 'user': 'user_name'}, inplace=True)
#the variables names after rename in the modified data frame
list(review_data)
# -
review_data.head(n=3)
# +
# review_data["score"]
# -
# Ratings distribution using pandas:
review_data["score"] = review_data["score"].fillna(review_data["score"].median())
review_data["score"].describe()
# Plot the distribution
review_data["score"].hist(bins=10)
# ## The median in both datasets is 5. This means that the data is skewed towards high ratings.
# This is a common bias in internet ratings, where people tend to rate items that they liked, and rarely spend time to comment something they dislike or are indiferent to. This will have a huge impact on the way I model the recommendation problem.
#
# ### Key conclusions from above:
#
# - Reviews are skewed towards positive
# - Many people agree with score 5 reviews
#
# ### Arrange and clean the data
# - Cleaning, handling missing data, normalization, etc:
# - For the algorithm in keras to work, remap all item_ids and user_ids to an interger between 0 and the total number of users or the total number of items
review_data.columns
review_data.head(n=2)
items = review_data.product_id.unique()
item_map = {i:val for i,val in enumerate(items)}
inverse_item_map = {val:i for i,val in enumerate(items)}
review_data["old_item_id"] = review_data["product_id"] # copying for join with metadata
review_data["item_id"] = review_data["product_id"].map(inverse_item_map)
items = review_data.item_id.unique()
print ("We have %d unique items in metadata "%items.shape[0])
# +
users = review_data.user_id.unique()
user_map = {i:val for i,val in enumerate(users)}
inverse_user_map = {val:i for i,val in enumerate(users)}
review_data["old_user_id"] = review_data["user_id"]
review_data["user_id"] = review_data["user_id"].map(inverse_user_map)
items_reviewed = review_data.product_id.unique()
review_data["old_item_id"] = review_data["product_id"] # copying for join with metadata
review_data["item_id"] = review_data["product_id"].map(inverse_item_map)
items_reviewed = review_data.item_id.unique()
users = review_data.user_id.unique()
helpful_votes = review_data.helpful_votes.unique()
# -
print ("We have %d unique users"%users.shape[0])
print ("We have %d unique items reviewed"%items_reviewed.shape[0])
# We have 192403 unique users in the "small" dataset
# We have 63001 unique items reviewed in the "small" dataset
# #### We have 94852 unique users
# #### We have 97758 unique items reviewed
review_data.head(3)
# ### Check the Distribution of number of ratings per user:
users_ratings = review_data['old_user_id'].value_counts().reset_index()
users_ratings.columns= ['old_user_id','num_ratings']
users_ratings['num_ratings'].describe()
# ### The distribution of number of ratings per user is very skewed in both datasets, with 50% of people having done a small number of reviews, and few made many ratings.
# ### I will check if it gives us enough information for generating good recommendations.
#
# ### * Check the Distribution of the number of ratings per item:
review_data.columns
review_data.head(n=10)
# ### To evaluate the model, I randomly separate the data into a training and test set.
ratings_train, ratings_test = train_test_split( review_data, test_size=0.1, random_state=0)
ratings_train.shape
ratings_test.shape
ratings_train.head(n=2)
# # **Define embeddings
# ### The $\underline{embeddings}$ are low-dimensional hidden representations of users and items,
# ### i.e. for each item I can find its properties and for each user I can encode how much they like those properties so I can determine attitudes or preferences of users by a small number of hidden factors
#
# ### Throughout the training, I learn two new low-dimensional dense representations: one embedding for the users and another one for the items.
#
# +
# declare input embeddings to the model
#User input
user_id_input = Input(shape=[1], name='user')
#Item Input
item_id_input = Input(shape=[1], name='item')
#helpful_votes
helpful_votes_id_input = Input(shape=[1], name='helpful_votes')
# define the size of embeddings as a parameter
user_embedding_size = embedding_size # Check 5, 10 , 15, 20, 50
item_embedding_size = embedding_size # Check 5, 10 , 15, 20, 50
helpful_votes_embedding_size = embedding_size # Check 5, 10 , 15, 20, 50
# apply an embedding layer to all inputs
user_embedding = Embedding(output_dim=user_embedding_size, input_dim=users.shape[0],
input_length=1, name='user_embedding')(user_id_input)
item_embedding = Embedding(output_dim=item_embedding_size, input_dim=items_reviewed.shape[0],
input_length=1, name='item_embedding')(item_id_input)
helpful_votes_embedding = Embedding(output_dim=helpful_votes_embedding_size, input_dim=helpful_votes.shape[0],
input_length=1, name='price_embedding')(helpful_votes_id_input)
# reshape from shape (batch_size, input_length,embedding_size) to (batch_size, embedding_size).
user_vecs = Reshape([user_embedding_size])(user_embedding)
user_vecs = Dropout(0.8)(user_vecs)
item_vecs = Reshape([item_embedding_size])(item_embedding)
item_vecs = Dropout(0.8)(item_vecs)
helpful_votes_vecs = Reshape([helpful_votes_embedding_size])(helpful_votes_embedding)
helpful_votes_vecs = Dropout(0.8)(helpful_votes_vecs)
# -
# # 2. Deep Recommender
#
# ### Instead of taking a dot product of the user and the item embedding, concatenate or multiply them and use them as features for a neural network.
# ### Thus, we are not constrained to the dot product way of combining the embeddings, and can learn complex non-linear relationships.
#
# 
#
#
#
#
#
# !mkdir -p ../models
# Try add dense layers on top of the embeddings before merging (Comment to drop this idea.)
user_vecs = Dense(64, activation='relu')(user_vecs)
user_vecs = Dropout(0.4)(user_vecs)
item_vecs = Dense(64, activation='relu')(item_vecs)
item_vecs = Dropout(0.4)(item_vecs)
helpful_votes_vecs = Dense(64, activation='relu')(helpful_votes_vecs)
item_vecs = Dropout(0.4)(item_vecs)
# +
# Concatenate the item embeddings :
# item_vecs_complete = Concatenate()([item_vecs, helpful_votes_vecs])
# Concatenate user and item embeddings and use them as features for the neural network:
# input_vecs = Concatenate()([user_vecs, item_vecs_complete]) # can be changed by Multiply
#input_vecs = Concatenate()([user_vecs, item_vecs]) # can be changed by Multiply
# Multiply user and item embeddings and use them as features for the neural network:
input_vecs = Multiply()([user_vecs, item_vecs]) # can be changed by concat
# Dropout is a technique where randomly selected neurons are ignored during training to prevent overfitting
input_vecs = Dropout(0.4)(input_vecs)
# Check one dense 128 or two dense layers (128,128) or (128,64) or three denses layers (128,64,32))
# First layer
# Dense(128) is a fully-connected layer with 128 hidden units.
# Use rectified linear units (ReLU) f(x)=max(0,x) as an activation function.
x = Dense(128, activation='relu')(input_vecs)
x = Dropout(0.4)(x) # Add droupout or not # To improve the performance
# Next Layers
x = Dense(128, activation='relu')(x) # Add dense again or not
x = Dropout(0.4)(x) # Add droupout or not # To improve the performance
x = Dense(64, activation='relu')(x) # Add dense again or not
x = Dropout(0.4)(x) # Add droupout or not # To improve the performance
x = Dense(32, activation='relu')(x) # Add dense again or not #
x = Dropout(0.4)(x) # Add droupout or not # To improve the performance
# The output
y = Dense(1)(x)
# +
# create model
model = Model(inputs=
[
user_id_input,
item_id_input
],
outputs=y)
# compile model
model.compile(loss='mse',
optimizer="adam" )
# set save location for model
save_path = "../models"
thename = save_path + '/' + modname + '.h5'
mcheck = ModelCheckpoint(thename, monitor='val_loss', save_best_only=True)
# fit model - increate batch_size to 64
history = model.fit([ratings_train["user_id"]
, ratings_train["item_id"]
]
, ratings_train["score"]
, batch_size=64
, epochs=num_epochs
, validation_split=0.1
, callbacks=[mcheck]
, shuffle=True)
# -
# !mkdir -p ../histories
# +
# Save the fitted model history to a file
with open('../histories/' + modname + '.pkl' , 'wb') as file_pi: pickle.dump(history.history, file_pi)
print("Save history in ", '../histories/' + modname + '.pkl')
# +
def disp_model(path,file,suffix):
model = load_model(path+file+suffix)
## Summarise the model
model.summary()
# Extract the learnt user and item embeddings, i.e., a table with number of items and users rows and columns, with number of columns is the dimension of the trained embedding.
# In our case, the embeddings correspond exactly to the weights of the model:
weights = model.get_weights()
print ("embeddings \ weights shapes",[w.shape for w in weights])
return model
model_path = "../models/"
# +
def plt_pickle(path,file,suffix):
with open(path+file+suffix , 'rb') as file_pi:
thepickle= pickle.load(file_pi)
plot(thepickle["loss"],label ='Train Error ' + file,linestyle="--")
plot(thepickle["val_loss"],label='Validation Error ' + file)
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Error")
##plt.ylim(0, 0.1)
return pd.DataFrame(thepickle,columns =['loss','val_loss'])
hist_path = "../histories/"
# -
print(model_path)
print(modname)
model=disp_model(model_path, modname, '.h5')
# Display the model using keras
SVG(model_to_dot(model).create(prog='dot', format='svg'))
x=plt_pickle(hist_path , modname , '.pkl')
x.head(20).transpose()
|
Keras-DeepRecommender-Shoes/2_Modeling/dense_5_Multiply_50_embeddings_10_epochs_dropout.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Face Detection Using CNN Algorithms
# >**This project is for building CNN architecture for facial features and face detection by plotting points on the facial features and box around the face**
# +
# import packages
import PIL
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import keras
import tensorflow as tf
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPooling2D
from keras.preprocessing import image
# %matplotlib inline
# -
# ## 1. Data Wrangling
# ### 1.1. Data Loading
# >
# **Data Source:** https://www.kaggle.com/jessicali9530/celeba-dataset
# But we are going to **use only the data of the first 35000 images** as a start then we are going to update the model step by step beacuse the data is very very big **(1 GB)**
# +
# This cell for the values that will be used across the entire notebook
# pathes
key_points_data_path = "./data/list_landmarks_align_celeba.csv"
images_data_path = "./data/images"
# data size
images_data_size = 35000
# originsl image dimensions
x_org = 178 # original x value
y_org = 218 # original y value
# new image dimensions
x_ = 45 # new value of x
image_size_ratio = x_org / y_org # dimensions ratio
y_ = int(image_size_ratio * x_) # new value of y
# image sizes
original_image_size = (x_org, y_org)
new_image_size = (x_,y_)
# the image size that will be used in the training process
image_size_training = new_image_size
# check the new size value
new_image_size
# -
# #### 1.1.1. Key points data
# +
# load the dataset (key points)
df_org = pd.read_csv(key_points_data_path)
df_org = df_org[:images_data_size]
# check
df_org.head(3)
# -
df_org.info()
# #### 1.1.2. Images Data
# +
# convert images to arrays to use it in training
images_data = list()
for idx in range(df_org.shape[0]):
# to get the path based on index
path = "{}/{}".format(str(images_data_path),str(df_org.iloc[idx].image_id))
# to read the image
image = PIL.Image.open(path).resize(image_size_training)
image_array = np.asarray(image) / 255
# append the image array to images_data
images_data.append(image_array)
# convert images_data to be array not list
images_data = np.array(images_data)
# check
images_data.shape
# -
# test
plt.imshow(images_data[18]);
# Final data sets which we will work with
print("Images Data Arrays Shape:", images_data.shape)
print("Key Points Data Shape:", df_org.shape)
# ### 1.2. Data Cleaning (Preprocessing)
# check if there is any null values in the data
df_org.isnull().sum()
# check the numerical values properties in the data
df_org.describe()
# > **Data is clean and ready for analysis**
# ### 1.3. Images reading and plotting
# function to read images based on index
def image_array(index, size=image_size_training, path=images_data_path):
"""
This functions is for converting images to arrays to deal with it in the model.
Input: index of the image that we want to convert to array
size of the image that we want for the array of the image
path of the images data to get the image
Output: the image array as numpy array
"""
# to get the path based on index
path = "{}/{}".format(str(path),str(df_org.iloc[index].image_id))
# to read the image
image = PIL.Image.open(path).resize(size)
image_array = np.asarray(image)
return image_array
# function to get a list of all key points of the face
def image_key_points_list(index, df = df_org):
"""
This function for getting the key points on the face as list to deal with it in plotting sections
"""
# box dictionary
points_list = [df.iloc[index].lefteye_x,
df.iloc[index].lefteye_y,
df.iloc[index].righteye_x,
df.iloc[index].righteye_y,
df.iloc[index].nose_x,
df.iloc[index].nose_y,
df.iloc[index].leftmouth_x,
df.iloc[index].leftmouth_y,
df.iloc[index].rightmouth_x,
df.iloc[index].rightmouth_y]
return points_list
# function to plot the image with green box around the faces
def plotting_image_with_box(index, df = df_org, size=original_image_size):
"""
This function for plotting the image with points on facial features and box around the face
"""
test_image = image_array(index, size)
points_list = image_key_points_list(index, df)
# face points
le_x, le_y, re_x, re_y = points_list[0], points_list[1], points_list[2], points_list[3]
n_x, n_y = points_list[4], points_list[5]
lm_x, lm_y, rm_x, rm_y = points_list[6], points_list[7], points_list[8], points_list[9]
# Create figure and axes
fig, ax = plt.subplots()
# plot the image
ax.imshow(test_image)
# plot the points on the face
ax.plot([le_x,re_x,n_x,lm_x,rm_x], [le_y,re_y,n_y,lm_y,rm_y], 'bo-')
# plot the box around the face
width = abs(le_x-rm_x-60)
height = abs(le_y-rm_y-75)
rect = patches.Rectangle((le_x-30, le_y-40), width, height, linewidth=4, edgecolor='g', facecolor='none')
ax.add_patch(rect);
# test
index = 18
plotting_image_with_box(index)
# ## 2. Preparing the data for ML model
# ### 2.1. Rescaling key points to be consistent with the new image size for training
# +
# # copy a version from the data to prepare it for analysis
df = df_org.copy()
# check
df.head(3)
# +
# function for updating key points for a new size
def rescale_key_points(oldsize=original_image_size, newsize=image_size_training):
"""
This function is for rescaling the key points from the original scale to a nwe scale from our chossen and we reduce
the image size to make the analysis faster and using lower memory
"""
# old and nwe sizes (x,y) values
x_axis_old = oldsize[0]
y_axis_old = oldsize[1]
x_axis_new = newsize[0]
y_axis_new = newsize[1]
x_ratio = x_axis_new / x_axis_old
y_ratio = y_axis_new / y_axis_old
# converting the keypoints values to be trained with the new size of the images
keypoints_x = ['lefteye_x', 'righteye_x', 'nose_x', 'leftmouth_x', 'rightmouth_x']
keypoints_y = ['lefteye_y', 'righteye_y', 'nose_y', 'leftmouth_y', 'rightmouth_y']
df[keypoints_x] = (df[keypoints_x] * x_ratio).astype('int')
df[keypoints_y] = (df[keypoints_y] * y_ratio).astype('int')
return 0
# call the function
rescale_key_points()
# check
df.head()
# -
# ### 2.2. Split the data into training and test datasets
# +
# training data
train_labels = df[:images_data_size - 1000]
train_images = images_data[:images_data_size - 1000]
# test data (1000 sample)
test_labels = df[images_data_size - 1000 + 1:]
test_images = images_data[images_data_size - 1000 + 1:]
# -
# ## 2.3 Split the data into train and validation datasets
# +
# droping image_id column as we will not use it in the model
y = train_labels.drop(['image_id'], axis = 1) # labels
X = train_images # images
# check
y.head(2)
# +
# to split the data into two sets (one for training and one for testing)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.17,random_state = 42)
# check the ratio
X_val.shape[0]/X_train.shape[0]
# -
# ## 3. Building and training the model
# +
# diminsions of the image in the traing process
x_ = image_size_training[0]
y_ = image_size_training[1]
# build the model
model = Sequential()
model.add(Conv2D(filters=8, kernel_size=(3, 3), padding='same', activation="relu", input_shape=(y_,x_,3)))
model.add(Conv2D(filters=8, kernel_size=(3, 3), padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.2))
model.add(Conv2D(filters=16, kernel_size=(3, 3), padding='same', activation='relu'))
model.add(Conv2D(filters=16, kernel_size=(3, 3), padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.2))
model.add(Conv2D(filters=32, kernel_size=(3, 3), padding='same', activation="relu"))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.2))
model.add(Conv2D(filters=64, kernel_size=(3, 3), padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(32, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(16, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='relu'))
# -
# get the summary of the model layers
model.summary()
# compile the model
model.compile(optimizer='adam', loss='mean_squared_error', metrics=['accuracy'])
# fitting the model with our data
training_process = model.fit(X_train, y_train, epochs=200, validation_data=(X_val, y_val), batch_size=300, shuffle=True)
# ## 4. Test the model
def predictions_test_model(index):
img = tf.keras.preprocessing.image.load_img("{}/0{}.jpg".format(images_data_path, index),target_size=(y_,x_,3))
img = tf.keras.preprocessing.image.img_to_array(img)
img = img/255
points_list = model.predict(img.reshape(1,y_,x_,3)).astype('int')[0]
# converting key points values to the original size
x_ratio = 1.05 * (original_image_size[0] / image_size_training[0])
y_ratio = 1.085 * (original_image_size[1] / image_size_training[1])
"""
In the previous ratios we multiply them by contant to reduce the noise that happened when we rescaled the points in
the previous training, there is no meaning for these numbers (i just pick them with trails)
"""
points_list[0] = int(points_list[0] * x_ratio)
points_list[2] = int(points_list[2] * x_ratio)
points_list[4] = int(points_list[4] * x_ratio)
points_list[6] = int(points_list[6] * x_ratio)
points_list[8] = int(points_list[8] * x_ratio)
points_list[1] = int(points_list[1] * y_ratio)
points_list[3] = int(points_list[3] * y_ratio)
points_list[5] = int(points_list[5] * y_ratio)
points_list[7] = int(points_list[7] * y_ratio)
points_list[9] = int(points_list[9] * y_ratio)
return points_list
# function to plot the image with green box around the faces
def test_image_with_box_plot(index, pred_or_actual = 'pred', pointsColor='bo-' ,boxcolor='g'):
img = tf.keras.preprocessing.image.load_img("{}/0{}.jpg".format(images_data_path, index),target_size=(y_org,x_org,3))
img = tf.keras.preprocessing.image.img_to_array(img)
test_image = img/255
# predictions of key points on the face
if pred_or_actual == 'pred':
points_list = predictions_test_model(index) # this for predections of the model
elif pred_or_actual == 'actual':
points_list = image_key_points_list(index) # this for the actual labels of the test data
# face points
le_x, le_y, re_x, re_y = points_list[0], points_list[1], points_list[2], points_list[3]
n_x, n_y = points_list[4], points_list[5]
lm_x, lm_y, rm_x, rm_y = points_list[6], points_list[7], points_list[8], points_list[9]
# Create figure and axes
fig, ax = plt.subplots()
# plot the image
ax.imshow(test_image)
# plot the points on the face
ax.plot([le_x,re_x,n_x,lm_x,rm_x], [le_y,re_y,n_y,lm_y,rm_y], pointsColor)
# plot the box around the face
width = abs(le_x-rm_x-60)
height = abs(le_y-rm_y-75)
rect = patches.Rectangle((le_x-30, le_y-40), width, height, linewidth=4, edgecolor=boxcolor, facecolor='none')
ax.add_patch(rect);
return points_list
# test the model performace
index = 34100
print('RED box for predections\n')
print('GREEN box for actual labels\n')
test_image_with_box_plot(index, pred_or_actual = 'pred', pointsColor='mo-' ,boxcolor='r')
test_image_with_box_plot(index, pred_or_actual = 'actual')
# +
# losses of both training and validation sets
loss = training_process.history['loss']
val_loss = training_process.history['val_loss']
# plot both losses
plt.plot(loss)
plt.plot(val_loss)
plt.legend(['loss', 'val_loss']);
# -
# accuracy of the model
accuracy = model.evaluate(X_val,y_val)[1] * 100
print("Accuracy of the model = ", round(accuracy,2))
# ## 5. Save the model
# saving the model as h5 file
model.save('model.h5')
|
Face-Detection/Facial-Features-and-Face-Detection-CNN/Facial-Features-and-Face-Detection-CNN.ipynb
|
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from utils import *
from IPython.display import display,HTML
# # NLP Deep Dive: RNNs
# ## Text Preprocessing
# ### Tokenization
# ### Word Tokenization with fastai
from fastai2.text.all import *
path = untar_data(URLs.IMDB)
files = get_text_files(path, folders = ['train', 'test', 'unsup'])
txt = files[0].open().read(); txt[:75]
spacy = WordTokenizer()
toks = first(spacy([txt]))
print(coll_repr(toks, 30))
first(spacy(['The U.S. dollar $1 is $1.00.']))
tkn = Tokenizer(spacy)
print(coll_repr(tkn(txt), 31))
defaults.text_proc_rules
coll_repr(tkn('© Fast.ai www.fast.ai/INDEX'), 31)
# ### Subword Tokenization
txts = L(o.open().read() for o in files[:2000])
def subword(sz):
sp = SubwordTokenizer(vocab_sz=sz)
sp.setup(txts)
return ' '.join(first(sp([txt]))[:40])
subword(1000)
subword(200)
subword(10000)
# ### Numericalization with fastai
toks = tkn(txt)
print(coll_repr(tkn(txt), 31))
toks200 = txts[:200].map(tkn)
toks200[0]
num = Numericalize()
num.setup(toks200)
coll_repr(num.vocab,20)
nums = num(toks)[:20]; nums
' '.join(num.vocab[o] for o in nums)
# ### Putting Our Texts into Batches for a Language Model
# + hide_input=false
stream = "In this chapter, we will go back over the example of classifying movie reviews we studied in chapter 1 and dig deeper under the surface. First we will look at the processing steps necessary to convert text into numbers and how to customize it. By doing this, we'll have another example of the PreProcessor used in the data block API.\nThen we will study how we build a language model and train it for a while."
tokens = tkn(stream)
bs,seq_len = 6,15
d_tokens = np.array([tokens[i*seq_len:(i+1)*seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# + hide_input=true
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15:i*15+seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# + hide_input=true
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15+seq_len:i*15+2*seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# + hide_input=true
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15+10:i*15+15] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# -
nums200 = toks200.map(num)
dl = LMDataLoader(nums200)
x,y = first(dl)
x.shape,y.shape
' '.join(num.vocab[o] for o in x[0][:20])
' '.join(num.vocab[o] for o in y[0][:20])
# ## Training a Text Classifier
# ### Language Model Using DataBlock
# +
get_imdb = partial(get_text_files, folders=['train', 'test', 'unsup'])
dls_lm = DataBlock(
blocks=TextBlock.from_folder(path, is_lm=True),
get_items=get_imdb, splitter=RandomSplitter(0.1)
).dataloaders(path, path=path, bs=128, seq_len=80)
# -
dls_lm.show_batch(max_n=2)
# ### Fine-Tuning the Language Model
learn = language_model_learner(
dls_lm, AWD_LSTM, drop_mult=0.3,
metrics=[accuracy, Perplexity()]).to_fp16()
learn.fit_one_cycle(1, 2e-2)
# ### Saving and Loading Models
learn.save('1epoch')
learn = learn.load('1epoch')
learn.unfreeze()
learn.fit_one_cycle(10, 2e-3)
learn.save_encoder('finetuned')
# ### Text Generation
TEXT = "I liked this movie because"
N_WORDS = 40
N_SENTENCES = 2
preds = [learn.predict(TEXT, N_WORDS, temperature=0.75)
for _ in range(N_SENTENCES)]
print("\n".join(preds))
# ### Creating the Classifier DataLoaders
dls_clas = DataBlock(
blocks=(TextBlock.from_folder(path, vocab=dls_lm.vocab),CategoryBlock),
get_y = parent_label,
get_items=partial(get_text_files, folders=['train', 'test']),
splitter=GrandparentSplitter(valid_name='test')
).dataloaders(path, path=path, bs=128, seq_len=72)
dls_clas.show_batch(max_n=3)
nums_samp = toks200[:10].map(num)
nums_samp.map(len)
learn = text_classifier_learner(dls_clas, AWD_LSTM, drop_mult=0.5,
metrics=accuracy).to_fp16()
learn = learn.load_encoder('finetuned')
# ### Fine-Tuning the Classifier
learn.fit_one_cycle(1, 2e-2)
learn.freeze_to(-2)
learn.fit_one_cycle(1, slice(1e-2/(2.6**4),1e-2))
learn.freeze_to(-3)
learn.fit_one_cycle(1, slice(5e-3/(2.6**4),5e-3))
learn.unfreeze()
learn.fit_one_cycle(2, slice(1e-3/(2.6**4),1e-3))
# ## Disinformation and Language Models
# ## Conclusion
# ## Questionnaire
# 1. What is "self-supervised learning"?
# 1. What is a "language model"?
# 1. Why is a language model considered self-supervised?
# 1. What are self-supervised models usually used for?
# 1. Why do we fine-tune language models?
# 1. What are the three steps to create a state-of-the-art text classifier?
# 1. How do the 50,000 unlabeled movie reviews help us create a better text classifier for the IMDb dataset?
# 1. What are the three steps to prepare your data for a language model?
# 1. What is "tokenization"? Why do we need it?
# 1. Name three different approaches to tokenization.
# 1. What is `xxbos`?
# 1. List four rules that fastai applies to text during tokenization.
# 1. Why are repeated characters replaced with a token showing the number of repetitions and the character that's repeated?
# 1. What is "numericalization"?
# 1. Why might there be words that are replaced with the "unknown word" token?
# 1. With a batch size of 64, the first row of the tensor representing the first batch contains the first 64 tokens for the dataset. What does the second row of that tensor contain? What does the first row of the second batch contain? (Careful—students often get this one wrong! Be sure to check your answer on the book's website.)
# 1. Why do we need padding for text classification? Why don't we need it for language modeling?
# 1. What does an embedding matrix for NLP contain? What is its shape?
# 1. What is "perplexity"?
# 1. Why do we have to pass the vocabulary of the language model to the classifier data block?
# 1. What is "gradual unfreezing"?
# 1. Why is text generation always likely to be ahead of automatic identification of machine-generated texts?
# ### Further Research
# 1. See what you can learn about language models and disinformation. What are the best language models today? Take a look at some of their outputs. Do you find them convincing? How could a bad actor best use such a model to create conflict and uncertainty?
# 1. Given the limitation that models are unlikely to be able to consistently recognize machine-generated texts, what other approaches may be needed to handle large-scale disinformation campaigns that leverage deep learning?
|
_notebooks/clean/10_nlp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="hX4n9TsbGw-f"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="0nbI5DtDGw-i"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="AOpGoE2T-YXS"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/text/word2vec">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/word2vec.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/word2vec.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/word2vec.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="haJUNjSB60Kh"
# # Word2Vec
# + [markdown] id="99d4ky2lWFvn"
# Word2Vec is not a singular algorithm, rather, it is a family of model architectures and optimizations that can be used to learn word embeddings from large datasets. Embeddings learned through Word2Vec have proven to be successful on a variety of downstream natural language processing tasks.
#
# Note: This tutorial is based on [Efficient Estimation of Word Representations in Vector Space](https://arxiv.org/pdf/1301.3781.pdf) and
# [Distributed
# Representations of Words and Phrases and their Compositionality](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf). It is not an exact implementation of the papers. Rather, it is intended to illustrate the key ideas.
#
# These papers proposed two methods for learning representations of words:
#
# * **Continuous Bag-of-Words Model** which predicts the middle word based on surrounding context words. The context consists of a few words before and after the current (middle) word. This architecture is called a bag-of-words model as the order of words in the context is not important.
# * **Continuous Skip-gram Model** which predict words within a certain range before and after the current word in the same sentence. A worked example of this is given below.
#
#
# You'll use the skip-gram approach in this tutorial. First, you'll explore skip-grams and other concepts using a single sentence for illustration. Next, you'll train your own Word2Vec model on a small dataset. This tutorial also contains code to export the trained embeddings and visualize them in the [TensorFlow Embedding Projector](http://projector.tensorflow.org/).
#
# + [markdown] id="xP00WlaMWBZC"
# ## Skip-gram and Negative Sampling
# + [markdown] id="Zr2wjv0bW236"
# While a bag-of-words model predicts a word given the neighboring context, a skip-gram model predicts the context (or neighbors) of a word, given the word itself. The model is trained on skip-grams, which are n-grams that allow tokens to be skipped (see the diagram below for an example). The context of a word can be represented through a set of skip-gram pairs of `(target_word, context_word)` where `context_word` appears in the neighboring context of `target_word`.
# + [markdown] id="ICjc-McbaVTd"
# Consider the following sentence of 8 words.
# > The wide road shimmered in the hot sun.
#
# The context words for each of the 8 words of this sentence are defined by a window size. The window size determines the span of words on either side of a `target_word` that can be considered `context word`. Take a look at this table of skip-grams for target words based on different window sizes.
# + [markdown] id="YKE87IKT_YT8"
# Note: For this tutorial, a window size of *n* implies n words on each side with a total window span of 2*n+1 words across a word.
# + [markdown] id="RsCwQ07E8mqU"
# 
# + [markdown] id="gK1gN1jwkMpU"
# The training objective of the skip-gram model is to maximize the probability of predicting context words given the target word. For a sequence of words *w<sub>1</sub>, w<sub>2</sub>, ... w<sub>T</sub>*, the objective can be written as the average log probability
# + [markdown] id="pILO_iAc84e-"
# 
# + [markdown] id="Gsy6TUbtnz_K"
# where `c` is the size of the training context. The basic skip-gram formulation defines this probability using the softmax function.
# + [markdown] id="P81Qavbb9APd"
# 
# + [markdown] id="axZvd-hhotVB"
# where *v* and *v<sup>'<sup>* are target and context vector representations of words and *W* is vocabulary size.
# + [markdown] id="SoLzxbqSpT6_"
# Computing the denominator of this formulation involves performing a full softmax over the entire vocabulary words which is often large (10<sup>5</sup>-10<sup>7</sup>) terms.
# + [markdown] id="Y5VWYtmFzHkU"
# The [Noise Contrastive Estimation](https://www.tensorflow.org/api_docs/python/tf/nn/nce_loss) loss function is an efficient approximation for a full softmax. With an objective to learn word embeddings instead of modelling the word distribution, NCE loss can be [simplified](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) to use negative sampling.
# + [markdown] id="WTZBPf1RsOsg"
# The simplified negative sampling objective for a target word is to distinguish the context word from *num_ns* negative samples drawn from noise distribution *P<sub>n</sub>(w)* of words. More precisely, an efficient approximation of full softmax over the vocabulary is, for a skip-gram pair, to pose the loss for a target word as a classification problem between the context word and *num_ns* negative samples.
# + [markdown] id="Cl0rSfHjt6Mf"
# A negative sample is defined as a (target_word, context_word) pair such that the context_word does not appear in the `window_size` neighborhood of the target_word. For the example sentence, these are few potential negative samples (when `window_size` is 2).
#
# ```
# (hot, shimmered)
# (wide, hot)
# (wide, sun)
# ```
# + [markdown] id="kq0q2uqbucFg"
# In the next section, you'll generate skip-grams and negative samples for a single sentence. You'll also learn about subsampling techniques and train a classification model for positive and negative training examples later in the tutorial.
# + [markdown] id="mk4-Hpe1CH16"
# ## Setup
# + id="RutaI-Tpev3T"
import io
import re
import string
import tensorflow as tf
import tqdm
from tensorflow.keras import Model
from tensorflow.keras.layers import Dot, Embedding, Flatten
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
# + id="10pyUMFkGKVQ"
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# + id="XkJ5299Tek6B"
SEED = 42
AUTOTUNE = tf.data.AUTOTUNE
# + [markdown] id="RW-g5buCHwh3"
# ### Vectorize an example sentence
# + [markdown] id="y8TfZIgoQrcP"
# Consider the following sentence:
# `The wide road shimmered in the hot sun.`
#
# Tokenize the sentence:
# + id="bsl7jBzV6_KK"
sentence = "The wide road shimmered in the hot sun"
tokens = list(sentence.lower().split())
print(len(tokens))
# + [markdown] id="PU-bs1XtThEw"
# Create a vocabulary to save mappings from tokens to integer indices.
# + id="UdYv1HJUQ8XA"
vocab, index = {}, 1 # start indexing from 1
vocab['<pad>'] = 0 # add a padding token
for token in tokens:
if token not in vocab:
vocab[token] = index
index += 1
vocab_size = len(vocab)
print(vocab)
# + [markdown] id="ZpuP43Dddasr"
# Create an inverse vocabulary to save mappings from integer indices to tokens.
# + id="o9ULAJYtEvKl"
inverse_vocab = {index: token for token, index in vocab.items()}
print(inverse_vocab)
# + [markdown] id="n3qtuyxIRyii"
# Vectorize your sentence.
#
# + id="CsB3-9uQQYyl"
example_sequence = [vocab[word] for word in tokens]
print(example_sequence)
# + [markdown] id="ox1I28JRIOdM"
# ### Generate skip-grams from one sentence
# + [markdown] id="t7NNKAmSiHvy"
# The `tf.keras.preprocessing.sequence` module provides useful functions that simplify data preparation for Word2Vec. You can use the `tf.keras.preprocessing.sequence.skipgrams` to generate skip-gram pairs from the `example_sequence` with a given `window_size` from tokens in the range `[0, vocab_size)`.
#
# Note: `negative_samples` is set to `0` here as batching negative samples generated by this function requires a bit of code. You will use another function to perform negative sampling in the next section.
#
# + id="USAJxW4RD7pn"
window_size = 2
positive_skip_grams, _ = tf.keras.preprocessing.sequence.skipgrams(
example_sequence,
vocabulary_size=vocab_size,
window_size=window_size,
negative_samples=0)
print(len(positive_skip_grams))
# + [markdown] id="uc9uhiMwY-AQ"
# Take a look at few positive skip-grams.
# + id="SCnqEukIE9pt"
for target, context in positive_skip_grams[:5]:
print(f"({target}, {context}): ({inverse_vocab[target]}, {inverse_vocab[context]})")
# + [markdown] id="_ua9PkMTISF0"
# ### Negative sampling for one skip-gram
# + [markdown] id="Esqn8WBfZnEK"
# The `skipgrams` function returns all positive skip-gram pairs by sliding over a given window span. To produce additional skip-gram pairs that would serve as negative samples for training, you need to sample random words from the vocabulary. Use the `tf.random.log_uniform_candidate_sampler` function to sample `num_ns` number of negative samples for a given target word in a window. You can call the function on one skip-grams's target word and pass the context word as true class to exclude it from being sampled.
#
# + [markdown] id="AgH3aSvw3xTD"
# Key point: *num_ns* (number of negative samples per positive context word) between [5, 20] is [shown to work](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) best for smaller datasets, while *num_ns* between [2,5] suffices for larger datasets.
# + id="m_LmdzqIGr5L"
# Get target and context words for one positive skip-gram.
target_word, context_word = positive_skip_grams[0]
# Set the number of negative samples per positive context.
num_ns = 4
context_class = tf.reshape(tf.constant(context_word, dtype="int64"), (1, 1))
negative_sampling_candidates, _, _ = tf.random.log_uniform_candidate_sampler(
true_classes=context_class, # class that should be sampled as 'positive'
num_true=1, # each positive skip-gram has 1 positive context class
num_sampled=num_ns, # number of negative context words to sample
unique=True, # all the negative samples should be unique
range_max=vocab_size, # pick index of the samples from [0, vocab_size]
seed=SEED, # seed for reproducibility
name="negative_sampling" # name of this operation
)
print(negative_sampling_candidates)
print([inverse_vocab[index.numpy()] for index in negative_sampling_candidates])
# + [markdown] id="8MSxWCrLIalp"
# ### Construct one training example
# + [markdown] id="Q6uEWdj8vKKv"
# For a given positive `(target_word, context_word)` skip-gram, you now also have `num_ns` negative sampled context words that do not appear in the window size neighborhood of `target_word`. Batch the `1` positive `context_word` and `num_ns` negative context words into one tensor. This produces a set of positive skip-grams (labelled as `1`) and negative samples (labelled as `0`) for each target word.
# + id="zSiZwifuLvHf"
# Add a dimension so you can use concatenation (on the next step).
negative_sampling_candidates = tf.expand_dims(negative_sampling_candidates, 1)
# Concat positive context word with negative sampled words.
context = tf.concat([context_class, negative_sampling_candidates], 0)
# Label first context word as 1 (positive) followed by num_ns 0s (negative).
label = tf.constant([1] + [0]*num_ns, dtype="int64")
# Reshape target to shape (1,) and context and label to (num_ns+1,).
target = tf.squeeze(target_word)
context = tf.squeeze(context)
label = tf.squeeze(label)
# + [markdown] id="OIJeoFCAwtXJ"
# Take a look at the context and the corresponding labels for the target word from the skip-gram example above.
# + id="tzyCPCuZwmdL"
print(f"target_index : {target}")
print(f"target_word : {inverse_vocab[target_word]}")
print(f"context_indices : {context}")
print(f"context_words : {[inverse_vocab[c.numpy()] for c in context]}")
print(f"label : {label}")
# + [markdown] id="gBtTcUVQr8EO"
# A tuple of `(target, context, label)` tensors constitutes one training example for training your skip-gram negative sampling Word2Vec model. Notice that the target is of shape `(1,)` while the context and label are of shape `(1+num_ns,)`
# + id="x-FwkR8jx9-Z"
print("target :", target)
print("context :", context)
print("label :", label)
# + [markdown] id="4bRJIlow4Dlv"
# ### Summary
# + [markdown] id="pWkuha0oykG5"
# This picture summarizes the procedure of generating training example from a sentence.
#
# + [markdown] id="_KlwdiAa9crJ"
# 
# + [markdown] id="9wmdO_MEIpaM"
# ## Compile all steps into one function
#
# + [markdown] id="iLKwNAczHsKg"
# ### Skip-gram Sampling table
# + [markdown] id="TUUK3uDtFNFE"
# A large dataset means larger vocabulary with higher number of more frequent words such as stopwords. Training examples obtained from sampling commonly occurring words (such as `the`, `is`, `on`) don't add much useful information for the model to learn from. [Mikolov et al.](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) suggest subsampling of frequent words as a helpful practice to improve embedding quality.
# + [markdown] id="bPtbv7zNP7Dx"
# The `tf.keras.preprocessing.sequence.skipgrams` function accepts a sampling table argument to encode probabilities of sampling any token. You can use the `tf.keras.preprocessing.sequence.make_sampling_table` to generate a word-frequency rank based probabilistic sampling table and pass it to `skipgrams` function. Take a look at the sampling probabilities for a `vocab_size` of 10.
# + id="Rn9zAnDccyRg"
sampling_table = tf.keras.preprocessing.sequence.make_sampling_table(size=10)
print(sampling_table)
# + [markdown] id="EHvSptcPk5fp"
# `sampling_table[i]` denotes the probability of sampling the i-th most common word in a dataset. The function assumes a [Zipf's distribution](https://en.wikipedia.org/wiki/Zipf%27s_law) of the word frequencies for sampling.
# + [markdown] id="mRHMssMmHgH-"
# Key point: The `tf.random.log_uniform_candidate_sampler` already assumes that the vocabulary frequency follows a log-uniform (Zipf's) distribution. Using these distribution weighted sampling also helps approximate the Noise Contrastive Estimation (NCE) loss with simpler loss functions for training a negative sampling objective.
# + [markdown] id="aj--8RFK6fgW"
# ### Generate training data
# + [markdown] id="dy5hl4lQ0B2M"
# Compile all the steps described above into a function that can be called on a list of vectorized sentences obtained from any text dataset. Notice that the sampling table is built before sampling skip-gram word pairs. You will use this function in the later sections.
# + id="63INISDEX1Hu"
# Generates skip-gram pairs with negative sampling for a list of sequences
# (int-encoded sentences) based on window size, number of negative samples
# and vocabulary size.
def generate_training_data(sequences, window_size, num_ns, vocab_size, seed):
# Elements of each training example are appended to these lists.
targets, contexts, labels = [], [], []
# Build the sampling table for vocab_size tokens.
sampling_table = tf.keras.preprocessing.sequence.make_sampling_table(vocab_size)
# Iterate over all sequences (sentences) in dataset.
for sequence in tqdm.tqdm(sequences):
# Generate positive skip-gram pairs for a sequence (sentence).
positive_skip_grams, _ = tf.keras.preprocessing.sequence.skipgrams(
sequence,
vocabulary_size=vocab_size,
sampling_table=sampling_table,
window_size=window_size,
negative_samples=0)
# Iterate over each positive skip-gram pair to produce training examples
# with positive context word and negative samples.
for target_word, context_word in positive_skip_grams:
context_class = tf.expand_dims(
tf.constant([context_word], dtype="int64"), 1)
negative_sampling_candidates, _, _ = tf.random.log_uniform_candidate_sampler(
true_classes=context_class,
num_true=1,
num_sampled=num_ns,
unique=True,
range_max=vocab_size,
seed=SEED,
name="negative_sampling")
# Build context and label vectors (for one target word)
negative_sampling_candidates = tf.expand_dims(
negative_sampling_candidates, 1)
context = tf.concat([context_class, negative_sampling_candidates], 0)
label = tf.constant([1] + [0]*num_ns, dtype="int64")
# Append each element from the training example to global lists.
targets.append(target_word)
contexts.append(context)
labels.append(label)
return targets, contexts, labels
# + [markdown] id="shvPC8Ji2cMK"
# ## Prepare training data for Word2Vec
# + [markdown] id="j5mbZsZu6uKg"
# With an understanding of how to work with one sentence for a skip-gram negative sampling based Word2Vec model, you can proceed to generate training examples from a larger list of sentences!
# + [markdown] id="OFlikI6L26nh"
# ### Download text corpus
#
# + [markdown] id="rEFavOgN98al"
# You will use a text file of Shakespeare's writing for this tutorial. Change the following line to run this code on your own data.
# + id="QFkitxzVVaAi"
path_to_file = tf.keras.utils.get_file('shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
# + [markdown] id="sOsbLq8a37dr"
# Read text from the file and take a look at the first few lines.
# + id="lfgnsUw3ofMD"
with open(path_to_file) as f:
lines = f.read().splitlines()
for line in lines[:20]:
print(line)
# + [markdown] id="gTNZYqUs5C2V"
# Use the non empty lines to construct a `tf.data.TextLineDataset` object for next steps.
# + id="ViDrwy-HjAs9"
text_ds = tf.data.TextLineDataset(path_to_file).filter(lambda x: tf.cast(tf.strings.length(x), bool))
# + [markdown] id="vfsc88zE9upk"
# ### Vectorize sentences from the corpus
# + [markdown] id="XfgZo8zR94KK"
# You can use the `TextVectorization` layer to vectorize sentences from the corpus. Learn more about using this layer in this [Text Classification](https://www.tensorflow.org/tutorials/keras/text_classification) tutorial. Notice from the first few sentences above that the text needs to be in one case and punctuation needs to be removed. To do this, define a `custom_standardization function` that can be used in the TextVectorization layer.
# + id="2MlsXzo-ZlfK"
# Now, create a custom standardization function to lowercase the text and
# remove punctuation.
def custom_standardization(input_data):
lowercase = tf.strings.lower(input_data)
return tf.strings.regex_replace(lowercase,
'[%s]' % re.escape(string.punctuation), '')
# Define the vocabulary size and number of words in a sequence.
vocab_size = 4096
sequence_length = 10
# Use the text vectorization layer to normalize, split, and map strings to
# integers. Set output_sequence_length length to pad all samples to same length.
vectorize_layer = TextVectorization(
standardize=custom_standardization,
max_tokens=vocab_size,
output_mode='int',
output_sequence_length=sequence_length)
# + [markdown] id="g92LuvnyBmz1"
# Call `adapt` on the text dataset to create vocabulary.
#
# + id="seZau_iYMPFT"
vectorize_layer.adapt(text_ds.batch(1024))
# + [markdown] id="jg2z7eeHMnH-"
# Once the state of the layer has been adapted to represent the text corpus, the vocabulary can be accessed with `get_vocabulary()`. This function returns a list of all vocabulary tokens sorted (descending) by their frequency.
# + id="jgw9pTA7MRaU"
# Save the created vocabulary for reference.
inverse_vocab = vectorize_layer.get_vocabulary()
print(inverse_vocab[:20])
# + [markdown] id="DOQ30Tx6KA2G"
# The vectorize_layer can now be used to generate vectors for each element in the `text_ds`.
# + id="yUVYrDp0araQ"
# Vectorize the data in text_ds.
text_vector_ds = text_ds.batch(1024).prefetch(AUTOTUNE).map(vectorize_layer).unbatch()
# + [markdown] id="7YyH_SYzB72p"
# ### Obtain sequences from the dataset
# + [markdown] id="NFUQLX0_KaRC"
# You now have a `tf.data.Dataset` of integer encoded sentences. To prepare the dataset for training a Word2Vec model, flatten the dataset into a list of sentence vector sequences. This step is required as you would iterate over each sentence in the dataset to produce positive and negative examples.
#
# Note: Since the `generate_training_data()` defined earlier uses non-TF python/numpy functions, you could also use a `tf.py_function` or `tf.numpy_function` with `tf.data.Dataset.map()`.
# + id="sGXoOh9y11pM"
sequences = list(text_vector_ds.as_numpy_iterator())
print(len(sequences))
# + [markdown] id="tDc4riukLTqg"
# Take a look at few examples from `sequences`.
#
# + id="WZf1RIbB2Dfb"
for seq in sequences[:5]:
print(f"{seq} => {[inverse_vocab[i] for i in seq]}")
# + [markdown] id="yDzSOjNwCWNh"
# ### Generate training examples from sequences
# + [markdown] id="BehvYr-nEKyY"
# `sequences` is now a list of int encoded sentences. Just call the `generate_training_data()` function defined earlier to generate training examples for the Word2Vec model. To recap, the function iterates over each word from each sequence to collect positive and negative context words. Length of target, contexts and labels should be same, representing the total number of training examples.
# + id="44DJ22M6nX5o"
targets, contexts, labels = generate_training_data(
sequences=sequences,
window_size=2,
num_ns=4,
vocab_size=vocab_size,
seed=SEED)
print(len(targets), len(contexts), len(labels))
# + [markdown] id="97PqsusOFEpc"
# ### Configure the dataset for performance
# + [markdown] id="7jnFVySViQTj"
# To perform efficient batching for the potentially large number of training examples, use the `tf.data.Dataset` API. After this step, you would have a `tf.data.Dataset` object of `(target_word, context_word), (label)` elements to train your Word2Vec model!
# + id="nbu8PxPSnVY2"
BATCH_SIZE = 1024
BUFFER_SIZE = 10000
dataset = tf.data.Dataset.from_tensor_slices(((targets, contexts), labels))
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
print(dataset)
# + [markdown] id="tyrNX6Fs6K3F"
# Add `cache()` and `prefetch()` to improve performance.
# + id="Y5Ueg6bcFPVL"
dataset = dataset.cache().prefetch(buffer_size=AUTOTUNE)
print(dataset)
# + [markdown] id="1S-CmUMszyEf"
# ## Model and Training
# + [markdown] id="sQFqaBMPwBqC"
# The Word2Vec model can be implemented as a classifier to distinguish between true context words from skip-grams and false context words obtained through negative sampling. You can perform a dot product between the embeddings of target and context words to obtain predictions for labels and compute loss against true labels in the dataset.
# + [markdown] id="oc7kTbiwD9sy"
# ### Subclassed Word2Vec Model
# + [markdown] id="Jvr9pM1G1sQN"
# Use the [Keras Subclassing API](https://www.tensorflow.org/guide/keras/custom_layers_and_models) to define your Word2Vec model with the following layers:
#
#
# * `target_embedding`: A `tf.keras.layers.Embedding` layer which looks up the embedding of a word when it appears as a target word. The number of parameters in this layer are `(vocab_size * embedding_dim)`.
# * `context_embedding`: Another `tf.keras.layers.Embedding` layer which looks up the embedding of a word when it appears as a context word. The number of parameters in this layer are the same as those in `target_embedding`, i.e. `(vocab_size * embedding_dim)`.
# * `dots`: A `tf.keras.layers.Dot` layer that computes the dot product of target and context embeddings from a training pair.
# * `flatten`: A `tf.keras.layers.Flatten` layer to flatten the results of `dots` layer into logits.
#
# With the subclassed model, you can define the `call()` function that accepts `(target, context)` pairs which can then be passed into their corresponding embedding layer. Reshape the `context_embedding` to perform a dot product with `target_embedding` and return the flattened result.
# + [markdown] id="KiAwuIqqw7-7"
# Key point: The `target_embedding` and `context_embedding` layers can be shared as well. You could also use a concatenation of both embeddings as the final Word2Vec embedding.
# + id="i9ec-sS6xd8Z"
class Word2Vec(Model):
def __init__(self, vocab_size, embedding_dim):
super(Word2Vec, self).__init__()
self.target_embedding = Embedding(vocab_size,
embedding_dim,
input_length=1,
name="w2v_embedding")
self.context_embedding = Embedding(vocab_size,
embedding_dim,
input_length=num_ns+1)
self.dots = Dot(axes=(3, 2))
self.flatten = Flatten()
def call(self, pair):
target, context = pair
word_emb = self.target_embedding(target)
context_emb = self.context_embedding(context)
dots = self.dots([context_emb, word_emb])
return self.flatten(dots)
# + [markdown] id="-RLKz9LFECXu"
# ### Define loss function and compile model
#
# + [markdown] id="I3Md-9QanqBM"
# For simplicity, you can use `tf.keras.losses.CategoricalCrossEntropy` as an alternative to the negative sampling loss. If you would like to write your own custom loss function, you can also do so as follows:
#
# ``` python
# def custom_loss(x_logit, y_true):
# return tf.nn.sigmoid_cross_entropy_with_logits(logits=x_logit, labels=y_true)
# ```
#
# It's time to build your model! Instantiate your Word2Vec class with an embedding dimension of 128 (you could experiment with different values). Compile the model with the `tf.keras.optimizers.Adam` optimizer.
# + id="ekQg_KbWnnmQ"
embedding_dim = 128
word2vec = Word2Vec(vocab_size, embedding_dim)
word2vec.compile(optimizer='adam',
loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] id="P3MUMrluqNX2"
# Also define a callback to log training statistics for tensorboard.
# + id="9d-ftBCeEZIR"
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir="logs")
# + [markdown] id="h5wEBotlGZ7B"
# Train the model with `dataset` prepared above for some number of epochs.
# + id="gmC1BJalEZIY"
word2vec.fit(dataset, epochs=20, callbacks=[tensorboard_callback])
# + [markdown] id="wze38jG57XvZ"
# Tensorboard now shows the Word2Vec model's accuracy and loss.
# + id="22E9eqS55rgz"
#docs_infra: no_execute
# %tensorboard --logdir logs
# + [markdown] id="awF3iRQCZOLj"
# <!-- <img class="tfo-display-only-on-site" src="images/word2vec_tensorboard.png"/> -->
# + [markdown] id="TaDW2tIIz8fL"
# ## Embedding lookup and analysis
# + [markdown] id="Zp5rv01WG2YA"
# Obtain the weights from the model using `get_layer()` and `get_weights()`. The `get_vocabulary()` function provides the vocabulary to build a metadata file with one token per line.
# + id="_Uamp1YH8RzU"
weights = word2vec.get_layer('w2v_embedding').get_weights()[0]
vocab = vectorize_layer.get_vocabulary()
# + [markdown] id="gWzdmUzS8Sl4"
# Create and save the vectors and metadata file.
# + id="VLIahl9s53XT"
out_v = io.open('vectors.tsv', 'w', encoding='utf-8')
out_m = io.open('metadata.tsv', 'w', encoding='utf-8')
for index, word in enumerate(vocab):
if index == 0:
continue # skip 0, it's padding.
vec = weights[index]
out_v.write('\t'.join([str(x) for x in vec]) + "\n")
out_m.write(word + "\n")
out_v.close()
out_m.close()
# + [markdown] id="1T8KcThhIU8-"
# Download the `vectors.tsv` and `metadata.tsv` to analyze the obtained embeddings in the [Embedding Projector](https://projector.tensorflow.org/).
# + id="lUsjQOKMIV2z"
try:
from google.colab import files
files.download('vectors.tsv')
files.download('metadata.tsv')
except Exception:
pass
# + [markdown] id="iS_uMeMw3Xpj"
# ## Next steps
#
# + [markdown] id="BSgAZpwF5xF_"
# This tutorial has shown you how to implement a skip-gram Word2Vec model with negative sampling from scratch and visualize the obtained word embeddings.
#
# * To learn more about word vectors and their mathematical representations, refer to these [notes](https://web.stanford.edu/class/cs224n/readings/cs224n-2019-notes01-wordvecs1.pdf).
#
# * To learn more about advanced text processing, read the [Transformer model for language understanding](https://www.tensorflow.org/tutorials/text/transformer) tutorial.
#
# * If you’re interested in pre-trained embedding models, you may also be interested in [Exploring the TF-Hub CORD-19 Swivel Embeddings](https://www.tensorflow.org/hub/tutorials/cord_19_embeddings_keras), or the [Multilingual Universal Sentence Encoder](https://www.tensorflow.org/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder)
#
# * You may also like to train the model on a new dataset (there are many available in [TensorFlow Datasets](https://www.tensorflow.org/datasets)).
#
|
src/text/word2vec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="Rc3gPsMdLgmK"
# # T019 · Molecular dynamics simulation
#
# Authors:
#
# - <NAME>, CADD seminar 2020, Charité/FU Berlin
# - <NAME>, 2020/21, Internship at [Volkamer Lab, Charité](https://volkamerlab.org/)
# - <NAME>, 2020, [Open Force Field Consortium](https://openforcefield.org/)
# - <NAME>, 2020/21, [Volkamer Lab, Charité](https://volkamerlab.org/)
# - <NAME>, 2020/21, [Volkamer Lab, Charité](https://volkamerlab.org/)
# + [markdown] id="ehs-GJfkwUap"
# __Note__
#
# This talktorial was designed to be used with [Google Colab](https://colab.research.google.com/github/volkamerlab/teachopencadd/blob/1bd7cb0c9f6379aebc0c1a0b1c7413685910cffa/teachopencadd/talktorials/019_md_simulation/talktorial.ipynb). It is also possible to use it on a local computer. However, performing the molecular dynamics simulation may take a considerably long time if no GPU is available.
#
# Also, note that this talktorial **will not run on Windows** for the time being (check progress in [this issue](https://github.com/volkamerlab/teachopencadd/issues/136)).
# + [markdown] id="v0WOINBLExEO"
# ## Aim of this talktorial
# + [markdown] id="ETO2kR5CE6-E"
# In this talktorial, we will learn why molecular dynamics (MD) simulations are important for drug design and which steps are necessary to perform an MD simulation of a protein in complex with a ligand. The kinase EGFR will serve as sample system for simulation.
# + [markdown] id="1KVizU3UVCxF"
# ### Contents in *Theory*
#
# - Molecular dynamics
# - Force fields
# - Boundary conditions
# - MD simulations and drug design
# - EGFR kinase
# + [markdown] id="kdamNfk7VCxF"
# ### Contents in *Practical*
#
# - Installation on Google Colab
# - Adjust environment for local installations running on Linux or MacOS
# - Import dependencies
# - Download PDB file
# - Prepare the protein ligand complex
# - Protein preparation
# - Ligand preparation
# - Merge protein and ligand
# - MD simulation set up
# - Force field
# - System
# - Perform the MD simulation
# - Download results
# + [markdown] id="mNvsYi3OLgmU"
# ### References
#
# - Review on the impact of MD simulations in drug discovery ([_J Med Chem_ (2016), **59**(9), 4035‐4061](https://doi.org/10.1021/acs.jmedchem.5b01684))
# - Review on the physics behind MD simulations and best practices ([_Living J Comp Mol Sci_ (2019), **1**(1), 5957](https://doi.org/10.33011/livecoms.1.1.5957))
# - Review on force fields ([_J Chem Inf Model_ (2018), **58**(3), 565-578](https://doi.org/10.1021/acs.jcim.8b00042))
# - Review on EGFR in cancer ([_Cancers (Basel)_ (2017), **9**(5), 52](https://dx.doi.org/10.3390%2Fcancers9050052))
# - Summarized statistical knowledge from Pierre-<NAME> ([Théorie Analytique des Probabilités _Gauthier-Villars_ (1820), **3**)](https://archive.org/details/uvrescompltesde31fragoog/page/n15/mode/2up)
# - Inspired by a notebook form <NAME> ([github](https://github.com/jaimergp/uab-msc-bioinf/blob/master/MD%20Simulation%20and%20Analysis%20in%20a%20Notebook.ipynb))
# - Repositories of [OpenMM](https://github.com/openmm/openmm) and [OpenMM Forcefields](https://github.com/openmm/openmmforcefields), [RDKit](https://github.com/rdkit/rdkit), [PyPDB](https://github.com/williamgilpin/pypdb), [MDTraj](https://github.com/mdtraj/mdtraj), [PDBFixer](https://github.com/openmm/pdbfixer)
# - Wikipedia articles about [MD simulations](https://en.wikipedia.org/wiki/Molecular_dynamics), [AMBER](https://en.wikipedia.org/wiki/AMBER) and [force fields](https://en.wikipedia.org/wiki/Force_field_(chemistry)) in general
# + [markdown] id="WpSLhe8ZLgmd"
# ## Theory
# + [markdown] id="lxhBZyifLgmm"
# ### Molecular dynamics
# + [markdown] id="svQEDsBCwUa5"
# Molecular dynamics is a computational method analyzing the movements and interactions of atoms and molecules of a defined system. The method stems from theoretical physics, where it was developed in the 1950s (Alder and Wainwright in [_J Chem Phys_ (1959), **31**(2), 459](https://doi.org/10.1063/1.1730376)), although the ideas behind it can be dated much earlier:
#
# > An intelligence which could, at any moment, comprehend all the forces by which nature is animated and the respective positions of the beings of which it is composed, and moreover, if this intelligence were far-reaching enough to subject these data to analysis, it would encompass in that formula both the movements of the largest bodies in the universe and those of the lightest atom: to it nothing would be uncertain, and the future, as well as the past, would be present to its eyes. The human mind offers us, in the perfection which it has given to astronomy, a faint sketch of this intelligence. (<NAME>, 1820)
#
# + [markdown] id="kLgNBvtIm7S3"
# Let us just take this statement by Laplace as the ideological substrate underneath molecular dynamics simulations. In other terms, we can approximate the behavior of a physical system by knowing the characteristics of its components and applying Newton's laws of motion. By solving the equations of motion, we can obtain a molecular trajectory of the system, which is a series of snapshots with the positions and velocities of all its particles, as well as its potential energy. To do so, we define functions, called force fields, which provide an approximate description of all the forces applied to each particle in the system. We then use numerical integrators to solve the initial value problem for the system and obtain the trajectory. As it sounds, the process requires quite a bit of processing power and it was only few years ago that MD started seeing a more widespread use, especially in the field of computational chemistry and biology, as well as in drug discovery ([_J Med Chem_ (2016), **59**(9), 4035‐4061](https://doi.org/10.1021/acs.jmedchem.5b01684)).
# + [markdown] id="yXlInBPrwUa6"
# 
#
# **Figure 1**: Molecular dynamics simulation of the rotation of a supramolecule composed of three molecules in a confined nanoscopic pore (Palma et al. via [Wikimedia](https://commons.wikimedia.org/w/index.php?curid=34866205)).
# + [markdown] id="8-uCG3KPwUa8"
# ### Force fields
#
#
# + [markdown] id="NC8t-i-cwUbB"
# Force fields describe the forces between atoms within and between molecules. They are parametric equations with components for different forces (bond stretching, van-der-Waals and more). The parameter values are usually derived experimentally and change for each MD scenario, depending on the molecules involved and the simulation settings. The result is a mathematical description of the energy landscape of the system, in which the forces acting on each particle result from the gradient of the potential energy with respect to the coordinates of the atoms.
#
# Several force fields are available, each with its own characteristics ([_J Chem Inf Model_ (2018), **58**(3), 565-578](https://doi.org/10.1021/acs.jcim.8b00042)). In this notebook, we will use a member of the AMBER force field family, which are widely used for MD simulations of proteins. Their functional form is:
#
# $$V(r^N) = \sum_{i \in bonds}k_{bi} (l_i-l^0_i)^2 + \sum_{i \in angles}k_{ai}(\theta_i - \theta^0_i)^2 + \sum_{i\in torsions} \sum_n \frac{1}{2} V_i^n[1+cos(nw_i-\gamma_i)]$$
# $$+ \sum_{j=1}^{N-1}\sum_{I=j+1}^{N} f_{ij}\in ij [(\frac{r^0_{ij}}{r_{ij}})^{12}-2(\frac{r^0_{ij}}{r_{ij}})^{6}]+\frac{q_iq_j}{4\pi \in_0 r_{ij}}$$
# + [markdown] id="pjSsc4l-wUbB"
# The formula consists of a sum of different components. The first three components contain information about bond lengths, angles and torsions (intramolecular forces). The last component describes intermolecular, non-bonded forces like van-der-Waals forces and electrostatic interactions. The various parameters, denoted by a superscript 0, depend on the force field used and vary between all members of the AMBER force field family. Note that these force fields assume fixed-charge particles and do not allow polarization, nor do they consider how a local charge influences its surroundings.
#
# The following visual representation of force fields components shows the same concepts in a more intuitive way.
# + [markdown] id="6aDDjIqYwUbC"
# 
#
# **Figure 2**: Components of a molecular mechanics force field (Edboas via [Wikimedia](https://commons.wikimedia.org/w/index.php?curid=4194424)).
# + [markdown] id="3J7LdqzKB0PM"
# ### Boundary conditions
#
# Often, molecular systems are simulated in a box filled with solvent such as water. These boxes are of finite size, which results in problems for molecules at or near the box boundaries. With which molecules should those interact? Periodic boundary conditions can avoid such boundary artifacts by simulating a theoretically infinite system. Molecules at one boundary of the box thereby interact with molecules at the boundary on the other side of the box. This mimics a situation, in which the simulation box is surrounded by replicas of itself. When visualizing such MD simulations, one can often observe that particles leave the box at one side (Fig. 3). However, they re-appear at the same time on the other side of the box with the same velocity. For simulations under periodic boundary conditions, it is recommended to use a simulation box large enough, so that the simulated macromolecule does not come into contact with neighboring images of itself.
# + [markdown] id="fdnT_He0VCxF"
# 
#
# **Figure 3**: Molecular dynamics simulation of water molecules with periodic boundary conditions (Kmckiern via [Wikimedia](https://commons.wikimedia.org/wiki/File:MD_water.gif)).
# + [markdown] id="zjcLLVbKNt-F"
# ### MD simulations and drug design
#
# MD simulations give valuable insights into the highly dynamic process of ligand binding to their target. When a ligand (or a drug) approaches a macromolecule (protein) in solution, it encounters a structure in constant motion. Also, ligands may induce conformational changes in the macromolecule that can best accommodate the small molecule. Such conformations may not be discovered with static methods. Accordingly, binding sites that are not observed in static ligand-free structures, but can be discovered with MD simulations, are sometimes called *cryptic binding sites* ([_J Med Chem_ (2016), **59**(9), 4035‐4061](https://doi.org/10.1021/acs.jmedchem.5b01684)). The identification of such binding sites with MD simulation can kickstart new drug discovery campaigns. Later in the drug discovery process, MD simulations can also be used to estimate the quality of computationally identified small molecules before performing more costly and time-intensive *in vitro* tests. Altogether, MD simulations pose a valuable asset in computational drug design.
# + [markdown] id="z-ZEaeB2n4hg"
# ### EGFR kinase
#
# The **E**pidermal **G**rowth **F**actor **R**eceptor (EGFR) is an important drug target with implications in cancer and inflammation ([Wikipedia](https://en.wikipedia.org/wiki/Epidermal_growth_factor_receptor)). It is a transmembrane protein with an extracellular receptor domain and an intracellular kinase domain. The binding of the endogenous ligand epidermal growth factor results in activation of the kinase domain via dimerization and autophosphorylation. The activated kinase domain can then phosphorylate downstream signaling proteins triggering DNA synthesis and cell proliferation ([_Cancers (Basel)_ (2017), **9**(5), 52](https://dx.doi.org/10.3390%2Fcancers9050052)). Inhibition of this kinase is the underlying mechanism of action of several already approved cancer drugs ([DrugBank](https://go.drugbank.com/bio_entities/BE0000767)). In this talktorial, we use the PDB structure **3POZ** of this kinase, which is in complex with the small molecule inhibitor **03P**, to perform an MD simulation ([PDB: 3POZ](https://www.rcsb.org/structure/3poz)).
# + [markdown] id="fARXaAnzwUau"
# 
#
# **Figure 4**: Structure 3POZ of the EGFR kinase domain bound to inhibitor 03P ([PDB: 3POZ](https://www.rcsb.org/structure/3poz)).
# + [markdown] id="dkhvXl0HLgnb"
# ## Practical
#
# We will now proceed to perform an MD simulation using the molecular dynamics engine [OpenMM](https://github.com/openmm/openmm), a high performance toolkit for molecular simulation. It is open source and can be used as application or library. We will use it as Python library.
# + [markdown] id="7I_BbH58n4h2"
# ### Installation on Google Colab
#
# The following code cells will install all required packages, if you are working on [Google Colab](https://colab.research.google.com/notebooks/intro.ipynb). Installing the [condacolab](https://github.com/jaimergp/condacolab) package will restart the kernel, which is intended. This notebook can also be used on a local computer but requires considerable computing power.
# + colab={"base_uri": "https://localhost:8080/"} id="K78GGT60Nu8p" outputId="40251762-3bfd-474a-8dda-c030b2a22b1d"
try:
import google.colab
# !pip install condacolab
import condacolab
condacolab.install()
except ModuleNotFoundError:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="VbqoZFShp00I" outputId="0f44a9dd-0cfd-49bc-dd46-969b50c2a9e8"
try:
import condacolab
from google.colab import files
from IPython.display import clear_output
condacolab.check()
# !conda install -q -y -c conda-forge mdtraj openmm openmmforcefields openff-toolkit pdbfixer pypdb rdkit
except ModuleNotFoundError:
on_colab = False
else:
#check if installation was succesful
try:
import rdkit
on_colab = True
clear_output() # clear the excessive installation outputs
print("Dependencies successfully installed!")
except ModuleNotFoundError:
print("Error while installing dependencies!")
# -
# ### Adjust environment for local installations running on Linux or MacOS
import sys
if not on_colab and sys.platform.startswith(("linux", "darwin")):
# !conda install -q -y -c conda-forge openmmforcefields
# ### Import dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="3g6vsWucLgng" outputId="daef1613-b0bf-4929-d5ed-8c3137151645"
import copy
from pathlib import Path
import requests
from IPython.display import display
import numpy as np
from rdkit import Chem
from rdkit.Chem import Draw
from rdkit.Chem import AllChem
import mdtraj as md
import pdbfixer
import simtk.openmm as mm
import simtk.openmm.app as app
from simtk.openmm import unit
from openff.toolkit.topology import Molecule, Topology
from openmmforcefields.generators import GAFFTemplateGenerator
# + id="w5g1k527SexH"
# create data directory if not exists
HERE = Path(_dh[-1])
DATA = HERE / "data"
DATA.mkdir(exist_ok=True)
# + [markdown] id="ltkfdrHvn4iI"
# ### Download PDB file
# The Protein Data Bank ([PDB](https://www.rcsb.org/)) allows for easy downloading of files via url.
# + id="Ecabaj2Y9kte"
pdbid = "4EIY"
ligand_name = "ZMA"
pdb_path = DATA / f"{pdbid}.pdb"
pdb_url = f"https://files.rcsb.org/download/{pdbid}.pdb"
# -
r = requests.get(pdb_url)
r.raise_for_status()
with open(pdb_path, "wb") as f:
f.write(r.content)
# + [markdown] id="krsfBvoenaYu"
# ### Prepare the protein ligand complex
# + [markdown] id="t3mGSg8LLgop"
# #### Protein preparation
#
# A crucial part for successful simulation is a correct and complete system. Crystallographic structures retrieved from the Protein Data Bank often miss atoms, mainly hydrogens, and may contain non-standard residues. In this talktorial, we will use the Python package [PDBFixer](https://github.com/openmm/pdbfixer) to prepare the protein structure. However, co-crystallized ligands are not handled well by [PDBFixer](https://github.com/openmm/pdbfixer) and will thus be prepared separately.
# + id="pA9Vct0iNRhl"
def prepare_protein(
pdb_file, ignore_missing_residues=True, ignore_terminal_missing_residues=True, ph=7.0
):
"""
Use pdbfixer to prepare the protein from a PDB file. Hetero atoms such as ligands are
removed and non-standard residues replaced. Missing atoms to existing residues are added.
Missing residues are ignored by default, but can be included.
Parameters
----------
pdb_file: pathlib.Path or str
PDB file containing the system to simulate.
ignore_missing_residues: bool, optional
If missing residues should be ignored or built.
ignore_terminal_missing_residues: bool, optional
If missing residues at the beginning and the end of a chain should be ignored or built.
ph: float, optional
pH value used to determine protonation state of residues
Returns
-------
fixer: pdbfixer.pdbfixer.PDBFixer
Prepared protein system.
"""
fixer = pdbfixer.PDBFixer(str(pdb_file))
fixer.removeHeterogens() # co-crystallized ligands are unknown to PDBFixer
fixer.findMissingResidues() # identify missing residues, needed for identification of missing atoms
# if missing terminal residues shall be ignored, remove them from the dictionary
if ignore_terminal_missing_residues:
chains = list(fixer.topology.chains())
keys = fixer.missingResidues.keys()
for key in list(keys):
chain = chains[key[0]]
if key[1] == 0 or key[1] == len(list(chain.residues())):
del fixer.missingResidues[key]
# if all missing residues shall be ignored ignored, clear the dictionary
if ignore_missing_residues:
fixer.missingResidues = {}
fixer.findNonstandardResidues() # find non-standard residue
fixer.replaceNonstandardResidues() # replace non-standard residues with standard one
fixer.findMissingAtoms() # find missing heavy atoms
fixer.addMissingAtoms() # add missing atoms and residues
fixer.addMissingHydrogens(ph) # add missing hydrogens
return fixer
# + id="bxuyg-oxOtvn"
# prepare protein and build only missing non-terminal residues
prepared_protein = prepare_protein(pdb_path, ignore_missing_residues=False)
# + [markdown] id="_wR47KGjnP2O"
# #### Prepare ligand
# + [markdown] id="8VoOjhASZ4Jh"
# After preparing the protein, we turn our attention to the ligand. Again, we need to add hydrogens, but also need to make sure the bond orders are correctly assigned, since some PDB entries may contain errors. We use the Python package [RDKit](https://github.com/rdkit/rdkit), an open source cheminformatics library.
# We will provide the correct protonation state and bond orders to [RDKit](https://github.com/rdkit/rdkit) via a SMILES string. Uncharged isomeric SMILES strings for each co-crystallized ligand can be found in their respective [PDB](https://www.rcsb.org) entry. The ligand of PDB entry [3POZ](https://www.rcsb.org/structure/3poz) has the name [03P](https://www.rcsb.org/ligand/03P). If a ligand is likely to bind in its charged form or as a specific tautomer, such characteristics need to be incorporated into the SMILES string.
# + id="si7ARKHoabal"
def prepare_ligand(pdb_file, resname, smiles, depict=True):
"""
Prepare a ligand from a PDB file via adding hydrogens and assigning bond orders. A depiction
of the ligand before and after preparation is rendered in 2D to allow an inspection of the
results. Huge thanks to @j-wags for the suggestion.
Parameters
----------
pdb_file: pathlib.PosixPath
PDB file containing the ligand of interest.
resname: str
Three character residue name of the ligand.
smiles : str
SMILES string of the ligand informing about correct protonation and bond orders.
depict: bool, optional
show a 2D representation of the ligand
Returns
-------
prepared_ligand: rdkit.Chem.rdchem.Mol
Prepared ligand.
"""
# split molecule
rdkit_mol = Chem.MolFromPDBFile(str(pdb_file))
rdkit_mol_split = Chem.rdmolops.SplitMolByPDBResidues(rdkit_mol)
# extract the ligand and remove any already present hydrogens
ligand = rdkit_mol_split[resname]
ligand = Chem.RemoveHs(ligand)
# assign bond orders from template
reference_mol = Chem.MolFromSmiles(smiles)
prepared_ligand = AllChem.AssignBondOrdersFromTemplate(reference_mol, ligand)
prepared_ligand.AddConformer(ligand.GetConformer(0))
# protonate ligand
prepared_ligand = Chem.rdmolops.AddHs(prepared_ligand, addCoords=True)
# 2D depiction
if depict:
ligand_2d = copy.deepcopy(ligand)
prepared_ligand_2d = copy.deepcopy(prepared_ligand)
AllChem.Compute2DCoords(ligand_2d)
AllChem.Compute2DCoords(prepared_ligand_2d)
display(
Draw.MolsToGridImage(
[ligand_2d, prepared_ligand_2d], molsPerRow=2, legends=["original", "prepared"]
)
)
# return ligand
return prepared_ligand
# + [markdown] id="GkqslZRknFbq"
# Calling this function with the isomeric SMILES string taken from the PDB entry for [03P](https://www.rcsb.org/ligand/03P) returns a correctly prepared ligand. Printed is a 2D-representation of the original and the prepared ligand for inspection.
# + colab={"base_uri": "https://localhost:8080/", "height": 217} id="04v75eTlcEtx" outputId="800d782f-8e3c-40c7-bd2d-3339fc8e8b54"
smiles = "c1cc(oc1)c2nc3nc(nc(n3n2)N)NCCc4ccc(cc4)O"
rdkit_ligand = prepare_ligand(pdb_path, ligand_name, smiles)
# + [markdown] id="NP9WekSTO5N3"
# #### Merge protein and ligand
#
# In the next step, we want to merge the prepared protein and ligand structures using the Python package [MDTraj](https://github.com/mdtraj/mdtraj). [MDTraj](https://github.com/mdtraj/mdtraj) can handle the prepared protein, which is currently a [PDBFixer](https://github.com/openmm/pdbfixer) molecule, a format that has a topology and atom positions similar to and usually interchangeable with [OpenMM Modeller](http://docs.openmm.org/latest/userguide/application.html#model-building-and-editing) topologies and positions. For the ligand however, we need to do several conversions, since it is currently an [RDKit](https://github.com/rdkit/rdkit) molecule.
# + id="UwehzZrhpDnD"
def rdkit_to_openmm(rdkit_mol, name="LIG"):
"""
Convert an RDKit molecule to an OpenMM molecule.
Inspired by @hannahbrucemcdonald and @glass-w.
Parameters
----------
rdkit_mol: rdkit.Chem.rdchem.Mol
RDKit molecule to convert.
name: str
Molecule name.
Returns
-------
omm_molecule: simtk.openmm.app.Modeller
OpenMM modeller object holding the molecule of interest.
"""
# convert RDKit to OpenFF
off_mol = Molecule.from_rdkit(rdkit_mol)
# add name for molecule
off_mol.name = name
# add names for atoms
element_counter_dict = {}
for off_atom, rdkit_atom in zip(off_mol.atoms, rdkit_mol.GetAtoms()):
element = rdkit_atom.GetSymbol()
if element in element_counter_dict.keys():
element_counter_dict[element] += 1
else:
element_counter_dict[element] = 1
off_atom.name = element + str(element_counter_dict[element])
# convert from OpenFF to OpenMM
off_mol_topology = off_mol.to_topology()
mol_topology = off_mol_topology.to_openmm()
mol_positions = off_mol.conformers[0]
# convert units from Ångström to Nanometers
for atom in mol_positions:
coords = atom / atom.unit
atom = (coords / 10.0) * unit.nanometers # since openmm works in nm
# combine topology and positions in modeller object
omm_mol = app.Modeller(mol_topology, mol_positions)
return omm_mol
# + id="I77OHmv47vEN"
omm_ligand = rdkit_to_openmm(rdkit_ligand, ligand_name)
# + [markdown] id="dqY5AE0JGIgL"
# Now protein and ligand are both in [OpenMM](https://github.com/openmm/openmm) like formats and can be merged with [MDTraj](https://github.com/mdtraj/mdtraj).
# + id="8gJMRq8fZYOU"
def merge_protein_and_ligand(protein, ligand):
"""
Merge two OpenMM objects.
Parameters
----------
protein: pdbfixer.pdbfixer.PDBFixer
Protein to merge.
ligand: simtk.openmm.app.Modeller
Ligand to merge.
Returns
-------
complex_topology: simtk.openmm.app.topology.Topology
The merged topology.
complex_positions: simtk.unit.quantity.Quantity
The merged positions.
"""
# combine topologies
md_protein_topology = md.Topology.from_openmm(protein.topology) # using mdtraj for protein top
md_ligand_topology = md.Topology.from_openmm(ligand.topology) # using mdtraj for ligand top
md_complex_topology = md_protein_topology.join(md_ligand_topology) # add them together
complex_topology = md_complex_topology.to_openmm()
# combine positions
total_atoms = len(protein.positions) + len(ligand.positions)
# create an array for storing all atom positions as tupels containing a value and a unit
# called OpenMM Quantities
complex_positions = unit.Quantity(np.zeros([total_atoms, 3]), unit=unit.nanometers)
complex_positions[: len(protein.positions)] = protein.positions # add protein positions
complex_positions[len(protein.positions) :] = ligand.positions # add ligand positions
return complex_topology, complex_positions
# + id="ZXmeeV0ZPOho"
complex_topology, complex_positions = merge_protein_and_ligand(prepared_protein, omm_ligand)
print("Complex topology has", complex_topology.getNumAtoms(), "atoms.")
# NBVAL_CHECK_OUTPUT
# + [markdown] id="2HAmjMG5Lgpd"
# ### MD simulation set up
#
# We can now use the prepared complex to set up the MD simulation.
# + [markdown] id="1B9DRdICny_f"
# #### Force field
#
# Common force fields like AMBER have parameters for amino acids, nucleic acids, water and ions and usually offer several options to choose from depending on your aim. We use the `amber14-all.xml` force field file, which is shipped with OpenMM and includes parameters for proteins, DNA, RNA and lipids. For solvation we use the standard three-site [water model](https://en.wikipedia.org/wiki/Water_model) [**TIP3P**](https://aip.scitation.org/doi/10.1063/1.445869).
#
# Parameters for ligands however are not included. To generate these parameters, we can use the **G**eneral **A**MBER **F**orce**F**ield ([GAFF](http://ambermd.org/antechamber/gaff.html)), which is implemented in the Python package [OpenMM Forcefields](https://github.com/openmm/openmmforcefields). The following function generates a force field object holding standard AMBER parameters and additionally includes parameters for a small molecule if required.
# + id="EFHKhU6v1k-h"
def generate_forcefield(
rdkit_mol=None, protein_ff="amber14-all.xml", solvent_ff="amber14/tip3pfb.xml"
):
"""
Generate an OpenMM Forcefield object and register a small molecule.
Parameters
----------
rdkit_mol: rdkit.Chem.rdchem.Mol
Small molecule to register in the force field.
protein_ff: string
Name of the force field.
solvent_ff: string
Name of the solvent force field.
Returns
-------
forcefield: simtk.openmm.app.Forcefield
Forcefield with registered small molecule.
"""
forcefield = app.ForceField(protein_ff, solvent_ff)
if rdkit_mol is not None:
gaff = GAFFTemplateGenerator(
molecules=Molecule.from_rdkit(rdkit_mol, allow_undefined_stereo=True)
)
forcefield.registerTemplateGenerator(gaff.generator)
return forcefield
# + id="5is-oVrO2Fjy"
forcefield = generate_forcefield(rdkit_ligand)
# + [markdown] id="cGrreylan6XS"
# #### System
# + [markdown] id="DzO12MFp2P48"
# With our configured force field we can now use the [OpenMM Modeller](http://docs.openmm.org/latest/userguide/application.html#model-building-and-editing) class to create the MD environment, a simulation box which contains the complex and is filled with a solvent. The standard solvent is water with a specified amount of ions. The size of the box can be determined in various ways. We define it with a padding, which results in a cubic box with dimensions dependent on the largest dimension of the complex.
#
# > Note this step can take a long time, in the order of minutes, depending on your hardware.
# + id="c4LEvWMI8ash"
modeller = app.Modeller(complex_topology, complex_positions)
modeller.addSolvent(forcefield, padding=1.0 * unit.nanometers, ionicStrength=0.15 * unit.molar)
# + [markdown] id="TwAEe5d8n4jf"
# With our solvated system and force field, we can finally create an [OpenMM System](http://docs.openmm.org/development/api-python/generated/openmm.openmm.System.html#openmm.openmm.System) and set up the simulation.
# Additionally to the system the simulation needs an integrator. An [OpenMM Integrator](http://docs.openmm.org/development/api-python/library.html#integrators) defines a method for simulating a system by integrating the equations of motion. The chosen **Langevin Integrator** uses Langevin equations. A list of all different kinds of integrators can be found in the [OpenMM Docs](http://docs.openmm.org/development/api-python/library.html#integrators). For further insight into the **Langevin Integrator**, we recommend reading about Langevin equations, e.g. on [Wikipedia](https://en.wikipedia.org/wiki/Langevin_equation).
# + id="n52Cmkr4dpI-"
system = forcefield.createSystem(modeller.topology, nonbondedMethod=app.PME)
integrator = mm.LangevinIntegrator(
300 * unit.kelvin, 1.0 / unit.picoseconds, 2.0 * unit.femtoseconds
)
simulation = app.Simulation(modeller.topology, system, integrator)
simulation.context.setPositions(modeller.positions)
# + [markdown] id="JwEgrFkILgqL"
# ### Perform the MD simulation
# Now that everything is set up, we can perform the simulation. We need to set starting positions and minimize the energy of the system to get a low energy starting configuration, which is important to decrease the chance of simulation failures due to severe atom clashes. The energy minimized system is saved.
# + id="EgkE1EUGLgqP"
simulation.minimizeEnergy()
with open(DATA / "topology.pdb", "w") as pdb_file:
app.PDBFile.writeFile(
simulation.topology,
simulation.context.getState(getPositions=True, enforcePeriodicBox=True).getPositions(),
file=pdb_file,
keepIds=True,
)
# + [markdown] id="KAwpxWM7ozSo"
# Once the minimization has finished, we can perform the MD simulation. In this talktorial, we will do a short simulation for illustration. Simulations for research purposes span several nanoseconds, even up to microseconds. We will simulate only 100 ps of molecular dynamics corresponding to 50k steps of 2 fs each. We save molecular "snapshots" every 10 ps (5000 steps), for a total of 10 frames. The results are saved in an .xtc file, which contains the coordinates of all the atoms at a given time point. Together with the PDB file of the energy minimized system written before, it gives us all the information needed for later analysis.
#
# **Note**: This talktorial will only generate a 20 fs trajectory, if not on Google Colab. However, if you have a good GPU available, you can also increase the simulation time.
# + id="E8t-0ZxFLgqh"
# output settings
if on_colab:
steps = 50000 # corresponds to 100 ps
write_interval = 5000 # write every 10 ps
log_interval = 2500 # log progress to stdout every 5 ps
else:
steps = 10 # corresponds to 20 fs
write_interval = 1 # write every 2 fs
log_interval = 1 # log progress to stdout every 2 fs
simulation.reporters.append(
md.reporters.XTCReporter(file=str(DATA / "trajectory.xtc"), reportInterval=write_interval)
)
simulation.reporters.append(
app.StateDataReporter(
sys.stdout,
log_interval,
step=True,
potentialEnergy=True,
temperature=True,
progress=True,
remainingTime=True,
speed=True,
totalSteps=steps,
separator="\t",
)
)
# + [markdown] id="J3gGRFBFozSp"
# The velocities for all particles in the system are randomly chosen from a distribution at the given temperature. We chose 300 Kelvin, which is some degrees above room temperature.
# A random seed is generated, but could be explicitly given to reproduce results.
#
# Then the simulation is performed by taking the steps defined before.
# + colab={"base_uri": "https://localhost:8080/"} id="7B7AiVO3fr03" outputId="1efb21e9-74de-47a4-dce9-961a514175f0"
simulation.context.setVelocitiesToTemperature(300 * unit.kelvin)
simulation.step(steps) # perform the simulation
# -
# Check the trajectory exists and is not empty
(DATA / "trajectory.xtc").stat().st_size > 0
# NBVAL_CHECK_OUTPUT
# + [markdown] id="M_Wf2prXqoif"
# ### Download results
#
# You can execute the following cell if you are working on Google Colab to download the MD simulation results.
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="FfcMX8G1i0RY" outputId="84b171f2-7ae1-42fd-cce6-9fa34431a88a"
if on_colab:
files.download(DATA / "topology.pdb")
files.download(DATA / "trajectory.xtc")
# + [markdown] id="aAleBNfKyhCd"
# ## Discussion
#
# We have successfully performed an MD simulation of a protein ligand complex. However, we simulated only a considerably short time to keep the execution time of the talktorial short. To address critical questions in drug design, longer simulations are often required.
# MD simulations are still too computationally costly to be useful for this purpose. Thus, so-called enhanced sampling methods were developed, that aim to accelerate the conformational sampling. Some of the most common methods are discussed in the **Further reading** section below.
# Furthermore, we did not include an equilibration step, which is commonly used to slowly heat up the system from 0 to 300 K before starting the simulation and might be important when simulating more sensitive systems including lipid bilayers. The protonation of ligand and protein was done separately, which is suboptimal, since protonation states of protein residues and ligand affect each other. However, we did not find a free and open-source solution meeting all requirements. Suggestions are very welcome!
# If you want to learn how to visualize and analyze the trajectory, you can refer to **Talktorial T020** in this repository.
# + [markdown] id="qFsGM5zEozSq"
# ## Quiz
#
# * Which inter- and intramolecular forces are being considered in the AMBER force field? Can you think of any forces not taken into account?
# * Would you expect to see the exact same simulation results when running the notebook twice with the same parameters?
# * Try doing a short (10ps, snapshot every 1ps) simulation of a protein without a ligand. You can find a broad variety of structures on [PDB](https://www.rcsb.org/) or you can use the EGFR kinase and remove the ligand.
# + [markdown] id="DYy_qmUJF1W6"
# ## Further reading
# + [markdown] id="IWYS2Q40u_dl"
# ### Enhanced sampling methods
#
# In theory, unbiased MD simulations should be capable of simulating binding and unbinding events of a drug molecule and its macromolecular target. However, the timescale of binding and unbinding events lies in the millisecond to second range. Enhanced sampling methods aim to accelerate the conformational sampling ([_J Med Chem._ 2016, **59(9)**, 4035-61](https://doi.org/10.1021/acs.jmedchem.5b01684)).
#
# One of these is **Free energy perturbation (FEP)** (also called alchemical free energy calculation), which computes the free energy difference when going from a state A to another state B. It is often employed in lead optimization to evaluate small modification at the ligand, that may boost the binding affinity for the desired target. The ligand from state A is thereby gradually transformed into the ligand of state B by simulating several intermediate ("alchemical") states ([alchemistry](http://www.alchemistry.org/wiki/Main_Page)).
#
# Another technique for free-energy calculations is **Umbrella sampling (US)**. US enforces sampling along a collective variable (CV) by performing staged simulations with an energetic bias. The bias usually takes the form of a harmonic potential, hence the term "umbrella". Its goal is to sample high-energy regions along the CV. However, the use in drug design is limited by the high computational cost.
#
# In contrast, **Steered MD (SMD)** follows a different approach: it applies external forces to the system. Those forces are time-dependent and facilitate the unbinding of the ligand from the target. The SMD calculates the final force exerted on the system. The unbinding force profile can then be used to filter hits from docking calculations and to discriminate active from inactive molecules.
|
teachopencadd/talktorials/T019_md_simulation/talktorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center><font color='Red' text align='center'>The Mnist Dataset</font></center>
# The dataset should be stored in the root foler of the project directory, in a folder called "data".
#
# The sets can be downloaded from http://yann.lecun.com/exdb/mnist/
#
# They are stored in gzip format so we must deal with decompressing the files.
# +
import gzip
with gzip.open('data/t10k-images-idx3-ubyte.gz', 'rb') as f:
file_content = f.read()
#Print first 4 elements of the file to see did it open. Expected result: \x00\x00\x08\x03
file_content[0:4]
# -
# ref : { https://docs.python.org/2/library/gzip.html }
# Our test set looks like this:
# 
type(file_content)
#Check to see if the values match up. Expected result: 2051
int.from_bytes(file_content[0:4], byteorder='big')
#Expected result: 10000
int.from_bytes(file_content[4:8], byteorder='big')
#Expected result: 28
int.from_bytes(file_content[8:12], byteorder='big')
#Expected result: 28
int.from_bytes(file_content[12:16], byteorder='big')
#Expected result: unasigned so should be 0
int.from_bytes(file_content[16:20], byteorder='big')
# ## Reading a single image
# Here we will try to read and display the image
# +
#Need numpy todeal with the array
import numpy as np
image = ~np.array(list(file_content[16:800])).reshape(28,28).astype(np.uint8)
# +
#Display the image
# %matplotlib inline
import matplotlib.pyplot as plt
plt.imshow(image, cmap='gray')
# -
# Reading a label, this should match up with the above image so we can see that the dataset is correct.
# +
# Adapted from: https://docs.python.org/3/library/gzip.html
import gzip
with gzip.open('data/t10k-labels-idx1-ubyte.gz', 'rb') as f:
labels = f.read()
# display the image label
int.from_bytes(labels[8:9], byteorder="big")
# -
#to get the further images we need to go every 784 bytes after the first 16 bytes in the file
image2 = ~np.array(list(file_content[800:1584])).reshape(28,28).astype(np.uint8)
plt.imshow(image2, cmap='gray')
# display the image label
int.from_bytes(labels[9:10], byteorder="big")
image3 = ~np.array(list(file_content[1584:2368])).reshape(28,28).astype(np.uint8)
plt.imshow(image3, cmap='gray')
# display the image label
int.from_bytes(labels[10:11], byteorder="big")
# looks like we are reading the images in fine.
# # End
|
mnist-dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas - DataFrames 3
import pandas as pd
import numpy as np
np.random.seed(101)
# Index Levels
outside = 'G1 G1 G1 G2 G2 G2'.split()
inside = [1,2,3,1,2,3]
hier_index = list(zip(outside, inside))
hier_index = pd.MultiIndex.from_tuples(hier_index)
outside
inside
hier_index
list(zip(outside, inside))
# Let's create a multi level index
df = pd.DataFrame(np.random.randn(6,2), hier_index, ['A', 'B'])
df
# Select a group
df.loc['G1']
# Call from outside index, then call inner index
df.loc['G1'].loc[1]
# Check index names
df.index.names
# Rename the index names
df.index.names = ['Groups', 'Num']
df
# +
# Let's grab G1-B
# -
df.loc['G2'].loc[2]['B']
# Grab all B items in G2
df.loc['G2']['B']
# ### Cross section
# Used only when we have a multi level index
df.xs('G1')
# Let's get all values where 'Num' in 1. Since Num is an inner level
# this allows deeper cross sections easily
df.xs(1, level='Num')
df
|
python-datasci-bootcamp/03-Python-for-Data-Analysis-Pandas/t_dataframes_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Confusion plots
# Copyright (c) 2020-2021 <NAME>, MIT License.
# ## Preliminaries
# +
# %matplotlib inline
from pathlib import Path
from sklearn.cluster import SpectralCoclustering
from tqdm import tqdm
import numpy as np
import seaborn as sns
import pandas as pd
import pylab as plt
import sys
sys.path.append("..")
from eval_segmentation import boundaries_to_intervals, intervals_to_boundaries, get_intervals_from_dir
from phoneseg_algorithms import get_segment_intervals, l2_segmentation, l2_n_segments, benji_l2_n_segments
# +
# Dataset
dataset = "buckeye"
split = "val"
model = "vqvae"
phoneseg_tag = "phoneseg_l2"
# phoneseg_tag = "phoneseg_l2.4"
# Directories
indices_dir = Path("../exp/")/model/dataset/split/"indices"
z_dir = Path("../exp/")/model/dataset/split/"auxiliary_embedding2"
phoneseg_dir = Path("../exp/")/model/dataset/split/phoneseg_tag/"intervals"
phoneref_dir = Path("../data/")/dataset/"phone_intervals"
audio_dir = Path("../../VectorQuantizedCPC/datasets/")/dataset/split
# -
# ## Read segmentation and reference
# +
# Read segmentation
segmentation_interval_dict = {}
segmentation_interval_dict = get_intervals_from_dir(phoneseg_dir)
utterances = segmentation_interval_dict.keys()
# Read phone reference
phone_ref_interval_dict = get_intervals_from_dir(phoneref_dir, utterances)
# -
# ## Overlap
# Get overlap
overlap_dict = {} # overlap["307"]["dh"] is frames of code 307 overlapping with "dh"
phones = set()
for utt_key in tqdm(phone_ref_interval_dict):
for interval_left, interval_right, code_index in segmentation_interval_dict[utt_key]:
code_index = int(code_index)
if code_index not in overlap_dict:
overlap_dict[code_index] = {}
for gt_left, gt_right, phone in phone_ref_interval_dict[utt_key]:
phones.add(phone)
if gt_right <= interval_left or gt_left >= interval_right:
continue
overlap = interval_right - interval_left
if gt_left > interval_left:
overlap -= gt_left - interval_left
if gt_right < interval_right:
overlap -= interval_right - gt_right
if phone not in overlap_dict[code_index]:
overlap_dict[code_index][phone] = 0
overlap_dict[code_index][phone] += overlap
# +
# Counts
code_indices = sorted([code_index for code_index in overlap_dict])
phones = sorted(phones)
counts = np.zeros((len(code_indices), len(phones)))
for i_code, code_index in enumerate(code_indices):
for i_phone, phone in enumerate(phones):
if phone in overlap_dict[code_index]:
counts[i_code, i_phone] = overlap_dict[code_index][phone]
# Normalize counts to get P(phone|code)
confusion_matrix = counts.T
confusion_matrix = confusion_matrix / np.sum(confusion_matrix, axis=0, keepdims=True)
# +
# Cluster
df = pd.DataFrame(confusion_matrix, index=phones, columns=range(confusion_matrix.shape[-1]))
model = SpectralCoclustering(n_clusters=20, random_state=0, svd_method="arpack", n_init=50)
model.fit(df.values)
# Reindex dataframe so that clustered phones are adjacent to eachother
df2 = df.reindex(index=[df.index[i] for i in np.argsort(model.row_labels_)], columns=[df.columns[i] for i in np.argsort(model.column_labels_)])
# -
# Plot
plt.figure(figsize=(22, 10))
sns.heatmap(df2, vmin=0, vmax=0.5, cbar=False, cmap="Blues", xticklabels=[])
# +
# Total number of frames
print("Total no. frames: {:d}".format(int(np.sum(counts))))
# Remove low count phones and codes
drop_phones = []
drop_codes = []
for i_phone, phone in enumerate(phones):
n_frames = np.sum(counts[:, i_phone])
if n_frames < 500:
drop_phones.append(phone)
# print("Dropping:", phone)
for i_code, code_index in enumerate(code_indices):
n_frames = np.sum(counts[i_code, :])
if n_frames < 100:
drop_codes.append(i_code)
# print("Dropping:", code_index)
df3 = df.drop(drop_phones)
df3 = df3.drop(columns=drop_codes)
# Cluster
model3 = SpectralCoclustering(n_clusters=20, random_state=0, svd_method="arpack", n_init=50)
model3.fit(df3.values)
# Reindex dataframe so that clustered phones are adjacent to eachother
df4 = df3.reindex(index=[df3.index[i] for i in np.argsort(model3.row_labels_)], columns=[df3.columns[i] for i in np.argsort(model3.column_labels_)])
# -
# Plot
plt.figure(figsize=(22, 10))
sns.heatmap(df4, vmin=0.1, vmax=0.4, cbar=False, cmap="Blues", xticklabels=[], yticklabels=[])
|
vqseg/notebooks/confusion_plots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import kurtosis
from scipy.stats import skew
time=pd.read_csv('delivery_time.csv')
time
time=time.rename(columns={'Delivery Time': 'DelT', 'Sorting Time': 'SortT'})
time.shape
time.dtypes
time.tail
time.info()
time[time.duplicated()].shape
time
time_cleaned1=time.drop_duplicates()
time_cleaned1.shape
time_cleaned1
time.duplicated()
time[time.duplicated()]
time_cleaned1['DelT'].hist()
time_cleaned1['SortT'].hist()
time_box=time_cleaned1.dropna()
time_box
plt.boxplot(time_box)
time.describe()
print(kurtosis(time.DelT))
# +
print(kurtosis(time.SortT))
# -
print(skew(time.DelT))
print(skew(time.SortT))
time.corr
import seaborn as sns
sns.pairplot(time)
corrMatrix=time.corr()
sns.heatmap(corrMatrix, annot=True)
plt.show()
cols=time.columns
colours=['#FF0000','#00FFFF']
sns.heatmap(time[cols].isnull(), cmap=sns.color_palette(colours))
time.boxplot(column='SortT')
time.boxplot(column='DelT')
time['DelT'].value_counts().plot.bar()
time['SortT'].value_counts().plot.bar()
sns.distplot(time['DelT'])
sns.distplot(time['SortT'])
import statsmodels.formula.api as smf
model = smf.ols("DelT~SortT",data = time).fit()
sns.regplot(x="DelT", y="SortT", data=time);
model.params
print(model.tvalues, '\n', model.pvalues)
(model.rsquared,model.rsquared_adj)
model.summary()
time_1=time
time_1['DelT'] = np.log(time_1['DelT'])
time_1['SortT'] = np.log(time_1['SortT'])
sns.distplot(time_1['DelT'])
fig = plt.figure()
sns.distplot(time_1['SortT'])
fig = plt.figure()
model_2 = smf.ols("SortT~DelT",data = time_1).fit()
model_2.summary()
time_2=time
time_1['DelT'] = np.log(time_1['DelT'])
sns.distplot(time_1['DelT'])
fig = plt.figure()
sns.distplot(time_1['SortT'])
fig = plt.figure()
model_3 = smf.ols("SortT~DelT",data = time_1).fit()
model_3.summary()
time_3=time
time_1['DelT'] = np.log(time_1['DelT'])
sns.distplot(time_1['DelT'])
fig = plt.figure()
sns.distplot(time_1['SortT'])
fig = plt.figure()
model_4 = smf.ols("SortT~DelT",data = time_1).fit()
model_4.summary()
pred = model.predict(time)
pred1 = model.predict(time)
import matplotlib.pyplot as plt
plt.scatter(x= time.SortT, y= time.DelT, color= 'red')
plt.plot(time.SortT, pred,color= 'blue')
plt.xlabel("Sorting time")
plt.ylabel("Delivery time")
plt.scatter(x= time.SortT, y= time.DelT, color= 'red')
plt.plot(time.SortT, pred1,color= 'blue')
plt.xlabel("Sorting time")
plt.ylabel("Delivery time")
|
Assignment 4 ( Q no-1) Simple Linear Regression (1).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import jax
import jax.numpy as np
import numpy as onp
from scipy.stats import chi2
from jax.scipy.stats import bernoulli, norm
from jax import random, grad, jit, value_and_grad, vmap
from jax import tree_multimap, tree_map
from jax.flatten_util import ravel_pytree
from collections import namedtuple
from copy import deepcopy
from cycler import cycler
from functools import partial
from matplotlib import cm, rc
import matplotlib.pyplot as plt
from synthetic_data import toy_logistic_data
# %matplotlib inline
# -
# Make sure to create this path
PATH = './figures'
# matplotlib Global Settings
rc('lines', lw=2)
rc('axes', lw=1.2, titlesize='large', labelsize='x-large')
rc('legend', fontsize='x-large')
rc('font', family='serif')
w, b, X, Y = toy_logistic_data(100, 82)
data = (X, Y[:, 0])
def get_line_coords(w, b):
w1, w2 = w
x1s = np.linspace(-1, 1, 100)
x2s = -(w1 * x1s + b) / (w2 + 1e-7)
return x1s, x2s
plt.scatter(X[:, 0], X[:, 1], c=Y)
x1s, x2s = get_line_coords(w, b)
plt.plot(x1s, x2s)
LocScaleParameters = namedtuple('LocScaleParameters', ['loc', 'log_scale'])
def gaussian_sample(params, key, num_samples=1):
mean = params.loc
std_dev = np.exp(params.log_scale)
samples = mean[np.newaxis, ...] + random.normal(key, shape=(num_samples, *mean.shape)) * std_dev
return samples
# +
def gaussian_log_pdf(params, x):
assert x.shape == params.loc.shape, "Input must have the same shape as the event. Use vmap for batching."
return np.sum(norm.logpdf(x, loc=params.loc, scale=np.exp(params.log_scale)))
def batch_log_pdf(params, x):
log_pdf = vmap(gaussian_log_pdf, in_axes=(None, 0))(params, x)
return log_pdf
# -
LinearModel = namedtuple('LinearModel', ('w', 'b'))
Hyperparameters = namedtuple('Hyperparameters', ('likelihood_parameters', 'prior_parameters'))
# +
def create_linear_model(features):
w = LocScaleParameters(loc=np.zeros((features, )), log_scale=np.zeros((features, )))
b = LocScaleParameters(loc=np.zeros(()), log_scale=np.zeros(()))
return LinearModel(w=w, b=b)
def logistic_regression_prior(features):
w = LocScaleParameters(loc=np.zeros((features, )), log_scale=np.zeros((features, )))
b = LocScaleParameters(loc=np.zeros(()), log_scale=np.zeros(()))
return LinearModel(w=w, b=b)
# -
def model_map(fn, model):
model_type = type(model)
new_model = model_type(*map(fn, model))
return new_model
def model_multimap(fn, model, extra_args):
model_type = type(model)
new_model = model_type(*map(fn, model, *extra_args))
return new_model
logistic_regression_prior_params = logistic_regression_prior(2)
logistic_regression_posterior = create_linear_model(2)
hyperparameters = Hyperparameters(likelihood_parameters=None, prior_parameters=logistic_regression_prior_params)
def predict(samples, x):
w = samples.w
b = samples.b
logits = np.dot(w, x) + b
return jax.nn.sigmoid(logits)
def bernoulli_logpmf(k, p):
tol = 1e-7
p = np.clip(p, tol, 1 - tol)
return k * np.log(p) + (1 - k) * np.log(1 - p)
def neg_likelihood(samples, data):
x, y = data
y_pred = vmap(predict, in_axes=(None, 0))(samples, x).T # SxB
logprob = vmap(bernoulli_logpmf, in_axes=(None, 0))(y, y_pred)
return -np.sum(logprob, axis=1)
def reparam_log_likelihood(samples, data):
return - neg_likelihood(samples, data)
def vi_objective(variational_parameters, hyperparameters, data, key, num_samples=1):
sampling_keys = random.split(key, num=2)
samples = model_multimap(lambda x, y: gaussian_sample(x, y, num_samples=num_samples),
variational_parameters, (sampling_keys, ))
exp_log_likelihood = reparam_log_likelihood(samples, data)
prior_parameters = hyperparameters.prior_parameters
exp_log_prior = model_multimap(batch_log_pdf, prior_parameters, (samples, ))
exp_log_posterior = model_multimap(batch_log_pdf, variational_parameters, (samples, ))
elbo_samples = (exp_log_likelihood - sum(exp_log_posterior) + sum(exp_log_prior))
return - np.mean(elbo_samples)
vi_loss_closure = jit(partial(vi_objective, hyperparameters=hyperparameters, data=data, num_samples=5))
vi_loss_value_and_grad = jit(value_and_grad(vi_objective), static_argnums=(1, 2, 4))
def varKL_objective(variational_parameters, hyperparameters, data, key, num_samples=1):
sampling_keys = random.split(key, num=2)
samples = model_multimap(lambda x, y: jax.lax.stop_gradient(gaussian_sample(x, y, num_samples=num_samples)),
variational_parameters, (sampling_keys, ))
exp_log_likelihood = reparam_log_likelihood(samples, data)
prior_parameters = hyperparameters.prior_parameters
exp_log_prior = model_multimap(batch_log_pdf, prior_parameters, (samples, ))
exp_log_posterior = model_multimap(batch_log_pdf, variational_parameters, (samples, ))
elbo_samples = (exp_log_likelihood - sum(exp_log_posterior) + sum(exp_log_prior))
return 0.5 * np.var(elbo_samples, ddof=1)
varKL_loss_value_and_grad = jit(value_and_grad(varKL_objective), static_argnums=(1, 2, 4))
def bbvi_objective(variational_parameters, hyperparameters, data, key, num_samples=1):
sampling_keys = random.split(key, num=2)
samples = model_multimap(lambda x, y: gaussian_sample(x, y, num_samples=num_samples),
variational_parameters, (sampling_keys, ))
samples = jax.lax.stop_gradient(samples)
exp_log_likelihood = reparam_log_likelihood(samples, data)
prior_parameters = hyperparameters.prior_parameters
exp_log_prior = model_multimap(batch_log_pdf, prior_parameters, (samples, ))
exp_log_posterior = model_multimap(batch_log_pdf, variational_parameters, (samples, ))
elbo_samples = (exp_log_likelihood - sum(exp_log_posterior) + sum(exp_log_prior))
loss_samples = jax.lax.stop_gradient(elbo_samples) * sum(exp_log_posterior)
return - np.mean(loss_samples)
bbvi_loss_value_and_grad = jit(value_and_grad(bbvi_objective), static_argnums=(1, 2, 4))
def log_posterior(variational_parameters, data, key, num_samples=1):
sampling_keys = random.split(key, num=2)
samples = model_multimap(lambda x, y: gaussian_sample(x, y, num_samples=num_samples),
variational_parameters, (sampling_keys, ))
samples = jax.lax.stop_gradient(samples)
exp_log_posterior = model_multimap(batch_log_pdf, variational_parameters, (samples, ))
return - np.mean(sum(exp_log_posterior))
score_function = jit(grad(log_posterior), static_argnums=(1, 3))
def gd_update(param, grad, learning_rate):
return param - learning_rate * grad
learning_rate = 0.001
key = random.PRNGKey(42)
param_periods = [deepcopy(logistic_regression_posterior)]
for i in range(100):
_, key = random.split(key)
# loss, gradients = vi_loss_value_and_grad(logistic_regression_posterior, hyperparameters, data, key, 5)
loss, gradients = varKL_loss_value_and_grad(logistic_regression_posterior, hyperparameters, data, key, 5)
update_fn = partial(gd_update, learning_rate=learning_rate)
updates = tree_multimap(update_fn, logistic_regression_posterior, (gradients))
logistic_regression_posterior = LinearModel(*updates)
param_periods.append(deepcopy(logistic_regression_posterior))
print("Loss =", loss)
# +
key = random.PRNGKey(42)
def sample_grads(params, key, num_samples):
varKL_loss_grads = []
bbvi_loss_grads = []
cv_bbvi = []
single_sample_grads = []
single_sample_cv = []
single_sample_value = []
for i in range(1000):
key, _ = random.split(key)
# VARIANCE LOSS
_, g = varKL_loss_value_and_grad(params, hyperparameters, data, key, num_samples)
g, _ = ravel_pytree(g)
varKL_loss_grads.append(g)
## BBVI LOSS
_, g = bbvi_loss_value_and_grad(params, hyperparameters, data, key, num_samples)
g, _ = ravel_pytree(g)
bbvi_loss_grads.append(g)
## CV BBVI LOSS
cv = score_function(params, data, key, num_samples)
cv, _ = ravel_pytree(cv)
cv_bbvi.append(cv)
key, _ = random.split(key)
## Single sample grad
_, g= bbvi_loss_value_and_grad(params, hyperparameters, data, key, 1)
g, _ = ravel_pytree(g)
single_sample_grads.append(g)
## Single samples CV
cv = score_function(params, data, key, 1)
cv, _ = ravel_pytree(cv)
single_sample_cv.append(cv)
## Single sample value
v, _ = vi_loss_value_and_grad(params, hyperparameters, data, key, 1)
single_sample_value.append(v)
varKL_loss_grads = np.stack(varKL_loss_grads)
bbvi_loss_grads = np.stack(bbvi_loss_grads)
cv_bbvi = np.stack(cv_bbvi)
single_sample_grads = np.stack(single_sample_grads)
single_sample_cv = np.stack(single_sample_cv)
single_sample_value = np.stack(single_sample_value)
return varKL_loss_grads, bbvi_loss_grads, cv_bbvi, single_sample_grads, single_sample_cv, single_sample_value
# -
def compute_cv_coeff(control_variates, gradients):
coeff = []
for i in range(gradients.shape[1]):
cov = onp.cov(control_variates[:, i], gradients[:, i], rowvar=False)
coeff.append(cov[0, 1] / cov[0, 0])
return np.stack(coeff)
def compute_delta(control_variates, function_values):
coeff = []
for i in range(control_variates.shape[1]):
cov = onp.cov(control_variates[:, i] ** 2, function_values, rowvar=False)
var = np.var(control_variates[:, i])
coeff.append(cov[0, 1] / var)
return np.stack(coeff)
def compute_chi2_confidence(x, df):
chi_2_interval = chi2.ppf([0.01, 0.99], df=df)
return df * x[None, :] / chi_2_interval[:, None]
CV_NUM_SAMPLES = [2, 1000]
def run(params, key):
results_dict = {
'bbvi': [],
'optimal_cv_bbvi': [],
'variance_loss': [],
}
for params in param_periods:
samples = sample_grads(params, key, 4)
varKL_loss_grads, bbvi_loss_grads, cv_bbvi, single_sample_grads, single_samples_cv, single_sample_value = samples
perm = onp.random.permutation(len(cv_bbvi))
cv_variance = []
for cv_num_samples in CV_NUM_SAMPLES:
optimal_cv_bbvi_coeff = compute_cv_coeff(single_samples_cv[perm[:cv_num_samples]], single_sample_grads[perm[:cv_num_samples]])
optimal_cv_bbvi_loss_grads = bbvi_loss_grads - optimal_cv_bbvi_coeff[None, :] * cv_bbvi
cv_variance.append(np.var(optimal_cv_bbvi_loss_grads, axis=0, ddof=1))
results_dict['variance_loss'].append(np.var(varKL_loss_grads, axis=0, ddof=1))
results_dict['bbvi'].append(np.var(bbvi_loss_grads, axis=0, ddof=1))
results_dict['optimal_cv_bbvi'].append(cv_variance)
return results_dict
results_dict = run(logistic_regression_posterior, key)
# +
import pickle
with open('./results/logistic_regression_variance_trace.pk', 'wb') as f:
pickle.dump(results_dict, f)
# with open('./results/logistic_regression_variance_trace.pk', 'rb') as f:
# results_dict = pickle.load(f)
# -
def get_title(idx):
if idx % 2 == 0:
return f'mean'
else:
return 'log standard deviation'
def plot_trace(results_dict, param_idx, log_scale=True):
bbvi_colours = ['C7', 'C0', 'C5', 'C6', 'C7', 'C8', 'C9']
def plot_single_trace(results, name, c, ls='-'):
stacked = results[:, param_idx]
xx = np.arange(0, stacked.shape[0], step=1)
plt.plot(xx, stacked, label=name, c=c, ls=ls, alpha=0.8)
bounds = compute_chi2_confidence(stacked, df=999)
plt.fill_between(xx, bounds[0, :], bounds[1, :], alpha=0.2, color=c)
plt.figure(figsize=(5, 3))
plot_single_trace(np.stack(results_dict['bbvi']), name='Reinforce', c='C1')
names = ['Sampled estimator', 'Oracle estimator']
for i, s in enumerate(CV_NUM_SAMPLES):
plot_single_trace(np.array(results_dict['optimal_cv_bbvi'])[:, i, :],
name=names[i], c=bbvi_colours[i])
plot_single_trace(np.stack(results_dict['variance_loss']), name='VarGrad', c='C3')
# plt.xticks(np.arange(len(NUM_SAMPLES)), NUM_SAMPLES)
plt.xlabel('epoch')
plt.ylabel('Variance')
if log_scale:
plt.yscale('log')
# plt.title(f'Logistic regression gradient variance w.r.t. variational {get_title(param_idx)}', fontsize='x-large')
plt.grid(axis='y', alpha=0.2)
# plt.legend(loc='lower center', bbox_to_anchor=(0.5, -0.8), ncol=2, fontsize='large', frameon=False)
plt.legend(loc='upper left', ncol=1, fontsize='medium', frameon=False)
plt.savefig(f'{PATH}/variance_wrt_iterations_{param_idx}.pdf', bbox_inches='tight')
for i in range(6):
plot_trace(results_dict, i)
|
notebooks/logistic_regression_gradient_variance_trace_plots_paper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
from tf.app import use
A = use("bhsa", hoist=globals())
query = """
clause
phrase
=: word sp#verb
"""
results = A.search(query)
# ```1.15s 188532 results```
#
# Above was done with cythonized, but unmodified text-fabric.
query = """
clause
phrase
=: word sp#verb
"""
results = A.search(query)
# ```1.78s 188532 results```
#
# This was done with pure python text-fabric.
|
data9/tfcython.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seml
import pandas as pd
from matplotlib import pyplot as plt
results = seml.get_results('gr_benchmark', to_data_frame=True)
results.groupby(['config.arguments.train_scheme', 'config.arguments.prune_criterion', 'config.arguments.ood_data_set'])['result.filename'].agg('max')
results['config.arguments.epsilons'] = results['config.arguments.epsilons'].astype(str)
|
notebooks/SEML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.__version__
import numpy as ban
ban.__version__
ar = np.array(10)
ar
ar.shape #to know how many rows and col are there
ar.ndim # to find dimesions
ar.size ## to find size
ar.dtype
ar.dtype.name # to find date type
#list
li = [10,20,30,40]
ar = np.array(li)
ar
ar.shape
np.append(li,50)
ar.size
ar.ndim
ar.dtype
ar.dtype.name
#tuple
tu = 10,2,0,30,3,4,12,65
ar= np.array(tu)
ar
ar.shape # to find rows and columns
ar.size # to find elements in given
ar.dtype # to find data type
ar.dtype.name
#set
se = {10,2,3,6,3}
b = np.array(se)
b
b.dtype
b.dtype.name
b.size
#dictionary
di = {'key1':'20', 'key2':'30'}
c = np.array(di)
c
c.size
c.ndim
c.dtype
# N-Dimentional List
nli = [[[[1,2,3],[2,3,4],[3,4,5]]]]
d= np.array(nli)
d
d.shape
d.size
d.ndim
#matrix
m = [[1,2,],[2,3]]
e = np.matrix(m)
e
e.shape
import numpy as np
d10=[[[[[[[[[[1,2],[2,3],[4,5]]]]]]]]]]
f = np.array(d10)
f
# +
A = [[1, 4, 5, 12],
[-5, 8, 9, 0],
[-6, 7, 11, 19]]
print("A =", A)
print("A[1] =", A[1]) # 2nd row
print("A[1][2] =", A[1][2]) # 3rd element of 2nd row
print("A[0][-1] =", A[0][-1]) # Last element of 1st Row
A = [[1, 4, 5, 12], [-5, 8, 9, 0], [-6, 7, 11, 19]]
A[1] = [-5, 8, 9, 0]
A[1][2] = 9
A[0][-1] = 0
# +
#pratice
# -
import numpy as np
a = np.array([1, 2, 3]) # Create a rank 1 array
#np.rank(a)
print(type(a)) # Prints "<class 'numpy.ndarray'>"
print(a.shape) # Prints "(3,)"
print(a[0], a[1], a[2]) # Prints "1 2 3"
a[0] = 5 # Change an element of the array
print(a)
b = np.array([[1,2,3],[4,5,6]]) # Create a rank 2 array
print(b.shape) # Prints "(2, 3)"
print(b[0, 0], b[0, 1], b[1, 0]) # Prints "1 2 4"
# +
import numpy as np
a = np.zeros((2,2)) # Create an array of all zeros
print(a) # Prints "[[ 0. 0.]
# [ 0. 0.]]"
b = np.ones((1,2)) # Create an array of all ones
print(b) # Prints "[[ 1. 1.]]"
c = np.full((2,2), 7) # Create a constant array
print(c) # Prints "[[ 7. 7.]
# [ 7. 7.]]"
d = np.eye(2) # Create a 2x2 identity matrix
print(d) # Prints "[[ 1. 0.]
# [ 0. 1.]]"
e = np.random.random((2,2)) # Create an array filled with random values
print(e) # Might print "[[ 0.91940167 0.08143941]
# [ 0.68744134 0.87236687]]"
# +
import numpy as np
# Create the following rank 2 array with shape (3, 4)
# [[ 1 2 3 4]
# [ 5 6 7 8]
# [ 9 10 11 12]]
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
# Use slicing to pull out the subarray consisting of the first 2 rows
# and columns 1 and 2; b is the following array of shape (2, 2):
# [[2 3]
# [6 7]]
b = a[:2, 1:3]
# A slice of an array is a view into the same data, so modifying it
# will modify the original array.
print(a[0, 1]) # Prints "2"
b[0, 0] = 77 # b[0, 0] is the same piece of data as a[0, 1]
print(a[0, 1])
# +
import numpy as np
# Create the following rank 2 array with shape (3, 4)
# [[ 1 2 3 4]
# [ 5 6 7 8]
# [ 9 10 11 12]]
a = np.array([[1,2,3,4], [5,6,7,8], [9,10,11,12]])
# Two ways of accessing the data in the middle row of the array.
# Mixing integer indexing with slices yields an array of lower rank,
# while using only slices yields an array of the same rank as the
# original array:
row_r1 = a[1, :] # Rank 1 view of the second row of a
row_r2 = a[1:2, :] # Rank 2 view of the second row of a
print(row_r1, row_r1.shape) # Prints "[5 6 7 8] (4,)"
print(row_r2, row_r2.shape) # Prints "[[5 6 7 8]] (1, 4)"
# We can make the same distinction when accessing columns of an array:
col_r1 = a[:, 1]
col_r2 = a[:, 1:2]
print(col_r1, col_r1.shape) # Prints "[ 2 6 10] (3,)"
print(col_r2, col_r2.shape) # Prints "[[ 2]
# [ 6]
# [10]] (3, 1)"
print(row_r1)
print(row_r2)
# -
|
20-01-2020-numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext lab_black
import os
import glob
import pathlib
import pandas as pd
from selenium import webdriver
from selenium.webdriver.common.by import By
from selenium.webdriver.chrome.options import Options
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.chrome.webdriver import WebDriver
from selenium.webdriver.support import expected_conditions as EC
this_dir = pathlib.Path(os.path.abspath(""))
data_dir = this_dir / "data"
# Configure the browser
options = Options()
options.add_argument("--headless")
options.add_argument("--no-sandbox")
options.add_argument("--disable-gpu")
options.add_argument("--disable-dev-shm-usage")
options.add_argument("--disable-extensions")
driver = webdriver.Chrome(options=options)
# Get the page
driver.get(
"https://datavisualization.cdph.ca.gov/t/LNC/views/COVIDSNFDASHV3/COVIDSNFDASH?:embed=y&:showVizHome=no&:host_url=https%3A%2F%2Fdatavisualization.cdph.ca.gov%2F&:embed_code_version=3&:tabs=yes&:toolbar=yes&:showAppBanner=false&:display_spinner=no&:loadOrderID=0"
)
# Pull the date
def get_date():
wait = WebDriverWait(driver, 300)
wait.until(
EC.visibility_of_element_located(
(By.ID, "tableau_base_widget_QuickFilterPanel_0")
)
)
date_filter = driver.find_element_by_id("tableau_base_widget_QuickFilterPanel_0")
date_string = date_filter.find_elements_by_xpath(
"//div[contains(@class, 'tabComboBoxName')]"
)[-1].text
return pd.to_datetime(date_string).date()
date = get_date()
# Routine to pull the download button
def get_download_button():
wait = WebDriverWait(driver, 300)
wait.until(EC.visibility_of_element_located((By.ID, "download-ToolbarButton")))
btn = driver.find_element_by_id("download-ToolbarButton")
assert btn.text == "Download"
return btn
# Routine to pull data for values
def get_data(element_id):
# WAit for the box
wait = WebDriverWait(driver, 300)
wait.until(EC.visibility_of_element_located((By.ID, element_id)))
# Get it and click it twice
box = driver.find_element_by_id(element_id)
box.click()
box.click()
# Click the download button
download_button = get_download_button()
download_button.click()
# Wait for the popup
wait.until(EC.visibility_of_element_located((By.XPATH, "//fieldset")))
fieldset = driver.find_element_by_xpath("//fieldset")
# Get the Data button and click it
buttons = fieldset.find_elements_by_xpath("//button")
data_button = next(e for e in buttons if e.text == "Data")
data_button.click()
# Move to the new tab
driver.switch_to.window(driver.window_handles[1])
# Get the data
wait.until(EC.visibility_of_element_located((By.XPATH, "//td")))
value = int(driver.find_element_by_xpath("//td").text)
# Close out
driver.close()
driver.switch_to.window(driver.window_handles[0])
# Return the data
return value
# Pull the four we're after
staff_active_cases = get_data("tabZoneId74")
patients_active_cases = get_data("tabZoneId124")
patients_deaths = get_data("tabZoneId78")
staff_deaths = get_data("tabZoneId86")
patients_confirmed_cases = get_data("tabZoneId76")
staff_confirmed_cases = get_data("tabZoneId84")
data = dict(
date=date,
staff_active_cases=staff_active_cases,
patients_active_cases=patients_active_cases,
patients_confirmed_cases=patients_confirmed_cases,
patients_deaths=patients_deaths,
staff_confirmed_cases=staff_confirmed_cases,
staff_deaths=staff_deaths,
)
data
# Kill the browser
driver.quit()
# Export
df = pd.DataFrame([data])
df.to_csv(data_dir / f"{date}.csv", index=False)
csv_list = [
i
for i in glob.glob(str(data_dir / "*.csv"))
if not str(i).endswith("timeseries.csv")
]
df_list = []
for csv in csv_list:
file_date = csv.split("/")[-1].replace(".csv", "")
df = pd.read_csv(csv, parse_dates=["date"])
df_list.append(df)
df = pd.concat(df_list).sort_values(["date"])
df.to_csv(data_dir / "timeseries.csv", index=False)
|
skilled-nursing-totals/scrape.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b1f363f4-31f7-4462-899e-de8e638ed2d7", "showTitle": false, "title": ""}
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('nlp').getOrCreate()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "adcfba8f-42f1-4fcb-aaf5-501b73b60113", "showTitle": false, "title": ""}
from pyspark.ml.feature import Tokenizer,RegexTokenizer
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2e8c0e73-fba1-4367-9844-a7dd83e6215a", "showTitle": false, "title": ""}
from pyspark.sql.functions import col,udf
from pyspark.sql.types import IntegerType
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6e0a038a-64ca-44cf-ac34-0ae94b4b9975", "showTitle": false, "title": ""}
sentence_df = spark.createDataFrame([
(0, 'Hello everyone and welcome to the tools for natural language processing lecture part 1.')
,(1, 'Before we jump straight into the code along project I want to take a brief moment to explore a few of the tools that SPARC has for dealing with text data understanding these tools.')
,(2, 'Were going to be able to use them easily in our custom coatl project.')
,(3, 'So were going to learn a lot of these basic features that youll find yourself using all the time.')
,(4, 'If,you,end,up,dealing,with,texts,they,show,a,spark,and,Python.')
],['id','sentence'])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e6c3b710-4508-4c06-b31f-daa0591de12b", "showTitle": false, "title": ""}
sentence_df.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b47cd53c-69fe-4534-89db-1eafc117acff", "showTitle": false, "title": ""}
tokenizer = Tokenizer(inputCol='sentence',outputCol='words')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "536d6ac3-afbe-4bf4-a71a-47c5da02c9be", "showTitle": false, "title": ""}
regex_tokenizer = RegexTokenizer(inputCol='sentence',outputCol='words',pattern='\\W')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "83ed6ec2-d180-4c19-aa0b-25fadad166b5", "showTitle": false, "title": ""}
count_tokens=udf(lambda words:len(words),IntegerType())
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "68f90785-8f1c-4f44-989c-552d82cdcf79", "showTitle": false, "title": ""}
df_tokenized = tokenizer.transform(sentence_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4af468b1-f15d-40de-9f81-cb936b3bbed5", "showTitle": false, "title": ""}
df_tokenized.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "61c63ef4-7c30-4daa-ab0a-f37c0d4ab355", "showTitle": false, "title": ""}
df_tokenized.withColumn('tokens',count_tokens(col('words'))).show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "43727bf6-084a-4b79-94fa-e5239eccb4fa", "showTitle": false, "title": ""}
regex_tokenized = regex_tokenizer.transform(sentence_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "b06105bf-cb2e-49e3-b087-78191e0adb6f", "showTitle": false, "title": ""}
regex_tokenized.withColumn('tokens',count_tokens(col('words'))).show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4368dc52-c4b3-433f-af02-2872b5af2460", "showTitle": false, "title": ""}
from pyspark.ml.feature import StopWordsRemover
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "aaeb21fa-6594-404e-b222-e2b477a9b1b8", "showTitle": false, "title": ""}
sentence_df2 = spark.createDataFrame([
(0, ['Hello','everyone','and','welcome to the tools for natural language processing lecture part 1.'])
,(1, ['Before','we','jump','straight into the code along project I want to take a brief moment to explore a few of the tools that SPARC has for dealing with text data understanding these tools.'])
,(2, ['Were','going','to','be able to use them easily in our custom coatl project.'])
,(3, ['So','were','going','to learn a lot of these basic features that youll find yourself using all the time.'])
,(4, ['If','you','end','up','dealing','with','texts','they','show','a','spark','and','Python.'])
],['id','tokens'])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "3adc15d6-d15c-4395-a479-de1001ce7c58", "showTitle": false, "title": ""}
sentence_df2.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "31c44d74-8ee4-48b5-907a-93c19a143495", "showTitle": false, "title": ""}
remover = StopWordsRemover(inputCol='tokens',outputCol='filtered')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4fdb297c-5287-4698-9e18-7b296fd03516", "showTitle": false, "title": ""}
remover.transform(sentence_df2).show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2cd620f8-aed9-4984-904b-22ed2015c707", "showTitle": false, "title": ""}
from pyspark.ml.feature import NGram
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e918ff1c-042b-44dd-9c0e-8d626e2ac385", "showTitle": false, "title": ""}
word_df = spark.createDataFrame([
(0, ['Hello','everyone','and','welcome to the tools for natural language processing lecture part 1.'])
,(1, ['Before','we','jump','straight into the code along project I want to take a brief moment to explore a few of the tools that SPARC has for dealing with text data understanding these tools.'])
,(2, ['Were','going','to','be able to use them easily in our custom coatl project.'])
,(3, ['So','were','going','to learn a lot of these basic features that youll find yourself using all the time.'])
,(4, ['If','you','end','up','dealing','with','texts','they','show','a','spark','and','Python.'])
],['id','words'])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0fd5f195-4602-43c0-8748-3fc6325fb165", "showTitle": false, "title": ""}
ngram = NGram(n=2,inputCol='words',outputCol='grams')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a4a728f6-95d4-4d1f-973e-3d1681a874e1", "showTitle": false, "title": ""}
ngram.transform(word_df).show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "099b1549-3841-47ca-aec7-ccb4c73536e4", "showTitle": false, "title": ""}
ngram.transform(word_df).select('grams').show(truncate=False)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c255a2a5-7c29-4a87-88c3-6e83b6e3d7f5", "showTitle": false, "title": ""}
from pyspark.ml.feature import HashingTF,IDF,Tokenizer
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c6ed583a-fda4-4e5d-a533-d36d2b3492e0", "showTitle": false, "title": ""}
df_tokenized.show(truncate=False)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "99d76c60-85df-47bc-9c6d-e842652259f6", "showTitle": false, "title": ""}
hashing_tf = HashingTF(inputCol='words',outputCol='rawFeatures')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ea0b3043-4730-400d-9362-15647bbbd6a4", "showTitle": false, "title": ""}
df_featurized = hashing_tf.transform(df_tokenized)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "460cd30c-ab63-4ab4-b559-dbfb05e288b1", "showTitle": false, "title": ""}
idf = IDF(inputCol='rawFeatures',outputCol='features')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "600bc283-0561-45e0-a3db-11cd626a89d1", "showTitle": false, "title": ""}
idf_model = idf.fit(df_featurized)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0fba12b6-15a7-4ac2-a39a-cfb91c3d53d3", "showTitle": false, "title": ""}
df_rescaled = idf_model.transform(df_featurized)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d1b7f3a2-7b02-4a6d-bac2-791a44d958dc", "showTitle": false, "title": ""}
df_rescaled.select('id','features').show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "501151d3-6030-442e-96ab-3bab59a52ae2", "showTitle": false, "title": ""}
from pyspark.ml.feature import CountVectorizer
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e12c16a6-8a95-42f0-be2a-b9e5040c20bd", "showTitle": false, "title": ""}
df = spark.createDataFrame([
(0,'a b c'.split(' '))
,(1, 'a b b c a'.split(' '))
], ['id', 'words'])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c96fc0e6-1ccd-4325-92d5-68e02d9914d1", "showTitle": false, "title": ""}
cv = CountVectorizer(inputCol='words',outputCol='features',vocabSize=3,minDF=2.0)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e3fbc13d-b8b5-442f-bccd-80f0243e5c6c", "showTitle": false, "title": ""}
cv_model = cv.fit(df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "97f1333f-a0d3-4bcd-a3ee-5168bc1aa2ec", "showTitle": false, "title": ""}
results = cv_model.transform(df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e17798e5-4535-4bc8-9c3b-3507bcc1b13c", "showTitle": false, "title": ""}
results.show(truncate=False)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e96d10ff-1e69-413c-aa80-aeb7deb48c02", "showTitle": false, "title": ""}
df_spam = spark.read.csv('/FileStore/tables/SMSSpamCollection',inferSchema=True,sep='\t')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "046e8b75-540a-4507-8a95-e057973fa985", "showTitle": false, "title": ""}
df_spam.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d39a485b-5eac-4add-bd5e-24d56e630a1f", "showTitle": false, "title": ""}
df_spam = df_spam.withColumnRenamed('_c0','class').withColumnRenamed('_c1','text')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f8181e85-b980-4da4-9738-a56490127e1c", "showTitle": false, "title": ""}
from pyspark.sql.functions import length
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a55b122b-268d-48ae-8b02-3e978c8e9aad", "showTitle": false, "title": ""}
df_spam = df_spam.withColumn('length',length(df_spam['text']))
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "2776bf82-1676-4160-b9af-b71e5f1dea38", "showTitle": false, "title": ""}
df_spam.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "621563aa-470c-4dc1-b850-5b8347ed05cf", "showTitle": false, "title": ""}
df_spam.groupBy('class').mean().show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6ca2f656-e4d4-4f86-8d6d-76b76b38ec17", "showTitle": false, "title": ""}
from pyspark.ml.feature import StringIndexer
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "9f04ca1a-b376-48fe-9286-c4b2251aaf40", "showTitle": false, "title": ""}
spam_tokenizer = Tokenizer(inputCol='text',outputCol='token_text')
spam_stop_remover = StopWordsRemover(inputCol='token_text',outputCol='stop_token')
spam_count_vectorizer = CountVectorizer(inputCol='stop_token',outputCol='count_vec')
idf = IDF(inputCol='count_vec',outputCol='tf_idf')
ham_spam_to_numeric = StringIndexer(inputCol='class',outputCol='label')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ad868b12-4f9e-40f5-bd81-5c1b85543de6", "showTitle": false, "title": ""}
from pyspark.ml.feature import VectorAssembler
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d7823b8c-1a65-4546-9578-1a4644899fa6", "showTitle": false, "title": ""}
data_cleanser = VectorAssembler(inputCols=['tf_idf','length'],outputCol='features')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e585f6b9-d74a-408f-bd5d-5d3db7acc687", "showTitle": false, "title": ""}
from pyspark.ml.classification import NaiveBayes
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "99b26797-f0d8-40ae-8f5a-57bb62d8b8fd", "showTitle": false, "title": ""}
nb = NaiveBayes()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "35fba2be-943f-432b-94af-bfb8a2106740", "showTitle": false, "title": ""}
from pyspark.ml import Pipeline
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f1356e24-e3d1-48f2-8a1d-ecabce71560b", "showTitle": false, "title": ""}
data_preprocessing_pipeline = Pipeline(stages=[ham_spam_to_numeric
,spam_tokenizer
,spam_stop_remover
,spam_count_vectorizer
,idf
,data_cleanser])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "73637388-478d-4a73-8d43-58a123db3af2", "showTitle": false, "title": ""}
job_clean = data_preprocessing_pipeline.fit(df_spam)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5c78bcd8-d426-46e3-808f-acde0598a005", "showTitle": false, "title": ""}
df_clean = job_clean.transform(df_spam)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7f19dadc-fe08-46dd-950d-8ef28f451519", "showTitle": false, "title": ""}
df_clean.columns
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "eefac25e-8b47-4196-9828-6d4b8d4c0e79", "showTitle": false, "title": ""}
df_clean = df_clean.select('label','features')
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8119569f-3b77-4de3-84d5-11f7d5c7f6a1", "showTitle": false, "title": ""}
df_clean.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "30646127-98aa-4370-a85f-ff49b2076166", "showTitle": false, "title": ""}
train_df,test_df = df_clean.randomSplit([0.7,0.3])
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "fa60cee4-521a-41a3-9c9e-86adc8e3d223", "showTitle": false, "title": ""}
spam_detector = nb.fit(train_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4528affa-8e7d-4835-b338-63c78a401fb8", "showTitle": false, "title": ""}
df_spam.printSchema()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "24520f5e-e40a-4ce3-8122-959714293368", "showTitle": false, "title": ""}
test_results = spam_detector.transform(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "55aad8c6-2cc0-4263-9837-0fed5355536e", "showTitle": false, "title": ""}
test_results.show()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f748f4ee-d6ae-4eb1-9746-08ba3e2c3887", "showTitle": false, "title": ""}
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "61fe9ac0-654b-43c2-99d8-6723af2136e3", "showTitle": false, "title": ""}
acc_eval = MulticlassClassificationEvaluator()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "30671af3-0b3c-4b3a-a29f-9a219cf5e022", "showTitle": false, "title": ""}
acc_nb = acc_eval.evaluate(test_results)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "d36e3f08-b3a3-4b08-8036-dbe442ce73a0", "showTitle": false, "title": ""}
print('Accuracy of Naive Bayes Model')
print(acc_nb)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8b3bc340-285d-44e5-99a8-d0eb85418ebc", "showTitle": false, "title": ""}
from pyspark.ml.classification import LogisticRegression,DecisionTreeClassifier,RandomForestClassifier,GBTClassifier
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f13addf4-e904-4f20-a4e6-d594c19ec92e", "showTitle": false, "title": ""}
logreg = LogisticRegression()
dtc = DecisionTreeClassifier()
rfc = RandomForestClassifier()
gbt = GBTClassifier()
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8f132421-8ada-4898-84d2-e86a11181493", "showTitle": false, "title": ""}
spam_detector_logreg = logreg.fit(train_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "17c554c2-bf3f-4e1a-afdb-a731911eaf7b", "showTitle": false, "title": ""}
logreg_test_results = spam_detector_logreg.transform(test_df)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5d8d8f88-8524-4b4c-b130-e5e089f0e459", "showTitle": false, "title": ""}
acc_logreg = acc_eval.evaluate(logreg_test_results)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "da15af23-6641-44e5-97eb-31cefaa87258", "showTitle": false, "title": ""}
print('Accuracy of Logistic Regression Model')
print(acc_logreg)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "0cbf38ad-9c7f-4765-a93f-0751d8651bc4", "showTitle": false, "title": ""}
spam_detector_dtc = dtc.fit(train_df)
dtc_test_results = spam_detector_dtc.transform(test_df)
acc_dtc = acc_eval.evaluate(dtc_test_results)
print('Accuracy of Decision Tree Model')
print(acc_dtc)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "a67c399d-6f62-4be0-9dc8-3a1edcd359cc", "showTitle": false, "title": ""}
spam_detector_rfc = rfc.fit(train_df)
rfc_test_results = spam_detector_rfc.transform(test_df)
acc_rfc = acc_eval.evaluate(rfc_test_results)
print('Accuracy of Random Forest Model')
print(acc_rfc)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "5cb695ed-fb75-45c1-9b00-c09bd262de89", "showTitle": false, "title": ""}
spam_detector_gbt = gbt.fit(train_df)
gbt_test_results = spam_detector_gbt.transform(test_df)
acc_gbt = acc_eval.evaluate(gbt_test_results)
print('Accuracy of Gradient Boosted Trees Model')
print(acc_gbt)
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "e69892a9-3390-42c6-8037-e70e7a17d59b", "showTitle": false, "title": ""}
|
projects/Spark and Python for Big Data with PySpark/Udemy_Section 16 Natural Language Processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P2i_HYQtRLUr" colab_type="text"
# ### Below is the code for 'Rotation-equivariant convolutional neural network ensembles in image processing', published in UbiComp/ISWC '19 Adjunct 2019.
#
# Abstract of Essay:
# For the present engineering of neural networks, rotation invariant is hard to be obtained. Rotation symmetry is an important characteristic in our physical world. In image recognition, using rotated images would largely decrease the performance of neural networks. This situation seriously hindered the application of neural networks in the real-world, such as human tracking, self-driving cars, and intelligent surveillance. In this paper, we would like to present a rotation-equivariant design of convolutional neural network ensembles to counteract the problem of rotated image processing task. This convolutional neural network ensembles combine multiple convolutional neural networks trained by different ranges of rotation angles respectively. In our proposed theory, the model lowers the training difficulty by learning with smaller separations of random rotation angles instead of a huge one. Experiments are reported in this paper. The convolutional neural network ensembles could reach 96.35% on rotated MNIST datasets, 84.9% on rotated Fashion-MNIST datasets, and 91.35% on rotated KMNIST datasets. These results are comparable to current state-of-the-art performance.
#
# + [markdown] id="3Wa_knoiCf_g" colab_type="text"
# ## Step 1: Import useful packages
# + id="OQq2SdEqCWJk" colab_type="code" colab={}
import torch
from torch import nn, optim
from torch.utils.data import DataLoader
import torchvision
import torchvision.transforms as transforms
from torchvision.transforms import Compose, ToTensor, Normalize, Resize
import torchvision.transforms.functional as TF
from torchvision.models.resnet import ResNet, BasicBlock
from torchvision.datasets import MNIST
from tqdm.autonotebook import tqdm
from sklearn.metrics import precision_score, recall_score, f1_score, accuracy_score
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
import inspect
import time
import numpy as np
# + [markdown] id="M-BsCZFFHAKE" colab_type="text"
# ## Step 2: Load and normalizing the MNIST training datasets
#
# + id="WzCJEVmsHAfB" colab_type="code" colab={}
means = deviations = [0.5]
train_transform=[]
start_angle=-270
end_angle=0
rotate_angle=45
ensemble_num=(360//rotate_angle)+1
for i in range(ensemble_num):
train_transform.append(transforms.Compose([transforms.RandomRotation([start_angle,end_angle]),
transforms.ToTensor(),
transforms.Normalize(means, deviations)]))
start_angle=-270+rotate_angle*(i+1)
end_angle=rotate_angle*(i+1)
# add trainset
trainset=[]
for i in range(ensemble_num):
trainset.append(torchvision.datasets.MNIST(root='./data', train=True,
download=True, transform=train_transform[i]))
# add trainloader
trainloader=[]
for i in range(ensemble_num):
trainloader.append(torch.utils.data.DataLoader(trainset[i], batch_size=128,
shuffle=True, num_workers=2))
# + [markdown] id="4A1_RXaV3vLc" colab_type="text"
# ## Step 3: Define a Convolutional Neural Network
# + id="4in2zxJ23vrT" colab_type="code" colab={}
class MnistResNet(nn.Module):
def __init__(self):
super(MnistResNet, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 32, kernel_size=5, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.layer2 = nn.Sequential(
nn.Conv2d(32, 64, kernel_size=5, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.drop_out = nn.Dropout()
self.fc1 = nn.Linear(7 * 7 * 64, 1000)
self.fc2 = nn.Linear(1000, 10)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = out.reshape(out.size(0), -1)
out = self.drop_out(out)
out = self.fc1(out)
out = self.fc2(out)
return out
# + [markdown] id="xU-HThkx4ubE" colab_type="text"
# ## Step 4: Define Loss functions and optimizers
# + id="k-JJT8oj4utW" colab_type="code" colab={}
#define loss function
criterion = nn.CrossEntropyLoss()
# use GPU
device = torch.device("cuda:0")
# define models
model=[]
for i in range(ensemble_num):
model.append(MnistResNet().to(device))
# define optimizers
optimizer=[]
for i in range(ensemble_num):
optimizer.append(optim.SGD(model[i].parameters(), lr=0.01, momentum=0.9, weight_decay=5e-4))
# + [markdown] id="Wq7IC55F8c91" colab_type="text"
# ## Step 5:Train each ensemble network
# + id="zymEtbj78dUx" colab_type="code" outputId="62bfca14-98d7-456c-e0f1-7a9ef8aab612" colab={"base_uri": "https://localhost:8080/", "height": 1000}
epoch_range = 6
# training loop + eval loop
for ensemble_id in range(ensemble_num):
running_loss = 0.0
print("Loss of ensemble model",ensemble_id)
for epoch in range(epoch_range):
for i, data in enumerate(trainloader[ensemble_id], 0):
# get the inputs
inputs, labels = data
# print(labels.numpy().shape)
inputs, labels = inputs.to(device), labels.to(device)
# zero the parameter gradients
optimizer[ensemble_id].zero_grad()
# forward + backward + optimize
outputs = model[ensemble_id](inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer[ensemble_id].step()
# print statistics
running_loss += loss.item()
if i % 20 == 19: # print every 2000 mini-batches
print('[%d, %5d] loss: %.6f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
# + [markdown] id="2Sxfc4GQAhWK" colab_type="text"
# ## Step 6: Form the encoded sets
# + id="DHMwONS6Ahoy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a34c5d6d-a788-423a-a084-9b1995290d0e"
correct = 0
train_data = []
train_label = []
for i in range(ensemble_num):
with torch.no_grad():
for data in trainloader[i]:
images, labels = data
images, labels = images.to(device), labels.to(device)
outputs=[]
for i in range(ensemble_num):
outputs.append(model[i](images))
outputs[i]=outputs[i].cpu().numpy()
labels = labels.cpu().numpy()
for i in range(len(outputs[0])):
d = np.concatenate((outputs[0][i], outputs[1][i]), axis=None)
d = np.concatenate((d, outputs[2][i]), axis=None)
d = np.concatenate((d, outputs[3][i]), axis=None)
d = np.concatenate((d, outputs[4][i]), axis=None)
d = np.concatenate((d, outputs[5][i]), axis=None)
d = np.concatenate((d, outputs[6][i]), axis=None)
train_data.append(d)
train_label.append(labels[i])
print(len(train_label))
# + [markdown] id="7BgrQnHzH1Sp" colab_type="text"
# ## Step 7: Test correction rate under several Classifers
# + id="usoSmNmKH1mo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="18c59974-bca8-4c06-929a-181fe16b1a43"
#Correction rate under KNeighborsClassifier
X_train, X_test, y_train, y_test = train_test_split(train_data, train_label, test_size=0.05, random_state=0)
neigh = KNeighborsClassifier(n_neighbors=10)
neigh.fit(X_train, y_train)
pred = neigh.predict(X_test)
total = 0
correct = 1
for i in range(len(y_test)):
total += 1
if y_test[i] == pred[i]:
correct += 1
print("The correction rate under KNN is:")
print(correct * 1.0 / total * 1.0)
# + id="CMy-cF-sKgVr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="8b6cf5dd-3479-44cb-8f8a-aca9e0894006"
#Correction rate under DecisionTreeClassifier
X_train, X_test, y_train, y_test = train_test_split(train_data, train_label, test_size=0.05, random_state=0)
tree = DecisionTreeClassifier(random_state=0)
tree.fit(X_train, y_train)
pred = tree.predict(X_test)
total = 0
correct = 1
for i in range(len(y_test)):
total += 1
if y_test[i] == pred[i]:
correct += 1
print("The correction rate under Decision Tree is:")
print(correct * 1.0 / total * 1.0)
# + id="TkQeQJczK3qq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="829adead-f65b-4842-f7e4-f4f86f5ab5a8"
#Correction rate under RandomForestClassifier
X_train, X_test, y_train, y_test = train_test_split(train_data, train_label, test_size=0.05, random_state=0)
forest = RandomForestClassifier(n_estimators=100, max_depth=6, random_state=0)
forest.fit(X_train, y_train)
pred = forest.predict(X_test)
total = 0
correct = 1
for i in range(len(y_test)):
total += 1
if y_test[i] == pred[i]:
correct += 1
print("The correction rate under Random Forest is:")
print(correct * 1.0 / total * 1.0)
# + id="4mm4OVFeLiUq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="11916ad5-c892-4120-e116-3a62c37a357f"
#Correction rate under MLPClassifier
X_train, X_test, y_train, y_test = train_test_split(train_data, train_label, test_size=0.05, random_state=0)
mlp = MLPClassifier(alpha=1, max_iter=1000)
mlp.fit(X_train, y_train)
pred = mlp.predict(X_test)
total = 0
correct = 1
for i in range(len(y_test)):
total += 1
if y_test[i] == pred[i]:
correct += 1
print("The correction rate under MLPClassifier is:")
print(correct * 1.0 / total * 1.0)
|
Rotation_Equivariant_CNN_Ensembles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Name: <NAME>
# Nov 23, 2021
# -
# ### Introduction to Numpy
#
# 1. NumPy is a linear algebra library for python
# 2. Numpy is also incredibly fast, as it has its binding to C libraries
# 3. It is highle recommended to install python using anaconda distribution to make sure all underlying dependencies (such as Linear Algebra libraries) all sync up with the use of conda install
# 4. if you have Anaconda, install Numpy by going to your terminal or command prompt and typing:
# ```conda install numpy
# pip install numpy```
import numpy as np
np.version
# ### Numpy Arrays - np.array()
#
# 1. Numpy arrays essentially come in two flavours: vectors and matrcies
# 2. Vectors are either 1-d Arrays or 2-d Matrices (but you should note a matrix can still have only one row or one column).
#casting normal python list to a numpy array
my_list = [1,2,3,4,5,6,7,8,9,10]
arr = np.array(my_list)
arr
# casting list of list to 2-d numpy array - matrix
my_mat = [[1,2,3],[4,5,6],[7,8,9]]
np.array(my_mat)
# ### Usually, We use Numpy built-in generation methods to create arrays a lot faster and a lot simpler.
# ### Most Commonly used are:
# +
# similar to python's range method
np.arange(start=0,stop=10)
# python range returns range object. it needs to be typecasted to list
# -
np.arange(0,10,step=2) #step =2 , to get even numbers
np.arange(10) # start and step are optional
# +
# we cannot create 2d array with np.arange()
# -
# generate arrays with all zeros
np.zeros(3) # 1-d array
np.zeros((2,3)) # pass tuple for matrix with rows and cols in a tuple
# generate arrays with all ones
np.ones(4)
np.ones((5,5))
# +
# evenly spaced numbers over a specified interval
np.linspace(0,5,10) # third argument: number of evenly spaced points you need
# range also provides evenly spaced values but third argumnet is step size
# (5-0)/9 should be the step size while using range
# -
np.linspace(2,3,5) # (3-2)/4 -> each step size
# +
# CREATING IDENTITY MATRIX
np.eye(4) # it is two dimensional square matrix with diagonal elements 1
# -
# ### Creating Array with Random Samples from Uniform Distribution:
# Discrete Unifrom Distribtion: https://www.youtube.com/watch?v=cyIEhL92wiw
# <br>Continuous Uniform Distribution: https://www.youtube.com/watch?v=j8XLYFzTJzE
# +
# Creating matrix with random numbers
# Creates an array of the given shape and populate it with random samples from a uniform probability distribution over [0, 1).
np.random.rand(5) # 1-d array
# -
np.random.rand(5,5) # 2-d array; we dont pass tuple here
# Random Samples from normal distribution (bell-shaped) centered around 0 or gaussian distribution
np.random.randn(5)
np.random.randn(4,4) # 2-d array; we dont pass tuple here
# random integer between low to high value
np.random.randint(1,100) # 1- inclusive, 100-exclusive
np.random.randint(1,100, 10) # third arg - number of random integers we want
# ### Attributes and Methods of an Array
arr = np.arange(16)
arr # 1-dimensional array of 0 upto 24
ran_arr = np.random.randint(0,50, 10)
ran_arr
# +
# RESHAPE METHOD - returns the same data but in new shape
# it useful for converting 1d array produced from methods like np.arange(), np.linspace() which does not have functionality to
# produce 2d arrays
arr.reshape(4,4) # pass the new dimensions we want
# -
arr
arr.reshape(8,2)
arr.reshape(16,1)
arr.reshape(1,16) # notice that it is 2d array
# +
# we might get an error if we cannot fill up that matrix completely
arr.reshape(5,10)
# -
arr.reshape(2,3)
# +
# rows * columns = number of elements in the array
# -
# MAX VALUE
ran_arr.max()
ran_arr.min()
arr.max() # works even for 2-dimensional arrays
# GETTING MIN INDEX & MAX INDEX
ran_arr.argmax()
arr = arr.reshape(5,5)
arr.argmax() # By default, the index is into the flattened array, otherwise along the specified axis.
# +
# SHAPE
arr.shape
# -
ran_arr.shape # indicates its 1-dimensional array
arr.dtype # data type in the array
# ### QUICK REVIEW
#
# 1. Create numpy arrays:
# - Casting a list - `np.array()`
# - Built-in functions - `np.arange()`, `np.zeros()`, `np.ones()`, `np.linspace()`, `np.eye()`
# - Random generation functions - `np.random.rand()`, `np.random.randn()`, `np.random.randint()`
# 2. Methods: `arr.reshape()`, `arr.min()`, `arr.max()`, `arr.argmin()`, `arr.argmax()`, `arr.shape`, `arr.dtype`
|
S05NumPy/L01ArrayCreation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### This jupyter notebooks provides the code for classifying signals using the Discrete Wavelet Transform.
# ### To get some more background information, please have a look at the accompanying blog-post:
# ### http://ataspinar.com/2018/12/21/a-guide-for-using-the-wavelet-transform-in-machine-learning/
# +
import os
from time import perf_counter
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import pywt
from collections import defaultdict, Counter
import keras
from keras.layers import Conv1D, BatchNormalization, Dense, Flatten, Activation
from tensorflow.keras.layers.experimental import preprocessing
from keras.models import Sequential
from keras.callbacks import History, EarlyStopping
history = History()
# -
# # 1. Loading the UCI HAR dataset
# download dataset from https://archive.ics.uci.edu/ml/datasets/Human+Activity+Recognition+Using+Smartphones
# +
activities_description = {
1: 'walking',
2: 'walking upstairs',
3: 'walking downstairs',
4: 'sitting',
5: 'standing',
6: 'laying'
}
def read_signals(filename):
with open(filename, 'r') as fp:
data = fp.read().splitlines()
data = map(lambda x: x.rstrip().lstrip().split(), data)
data = [list(map(float, line)) for line in data]
return data
def read_labels(filename):
with open(filename, 'r') as fp:
activities = fp.read().splitlines()
activities = list(map(lambda x: int(x)-1, activities))
return activities
def randomize(dataset, labels):
permutation = np.random.permutation(labels.shape[0])
shuffled_dataset = dataset[permutation, :, :]
shuffled_labels = labels[permutation]
return shuffled_dataset, shuffled_labels
DATA_FOLDER = '../datasets/UCI HAR Dataset/'
INPUT_FOLDER_TRAIN = DATA_FOLDER+'train/Inertial Signals/'
INPUT_FOLDER_TEST = DATA_FOLDER+'test/Inertial Signals/'
INPUT_FILES_TRAIN = ['body_acc_x_train.txt', 'body_acc_y_train.txt', 'body_acc_z_train.txt',
'body_gyro_x_train.txt', 'body_gyro_y_train.txt', 'body_gyro_z_train.txt',
'total_acc_x_train.txt', 'total_acc_y_train.txt', 'total_acc_z_train.txt']
INPUT_FILES_TEST = ['body_acc_x_test.txt', 'body_acc_y_test.txt', 'body_acc_z_test.txt',
'body_gyro_x_test.txt', 'body_gyro_y_test.txt', 'body_gyro_z_test.txt',
'total_acc_x_test.txt', 'total_acc_y_test.txt', 'total_acc_z_test.txt']
LABELFILE_TRAIN = DATA_FOLDER+'train/y_train.txt'
LABELFILE_TEST = DATA_FOLDER+'test/y_test.txt'
train_signals, test_signals = [], []
for input_file in INPUT_FILES_TRAIN:
signal = read_signals(INPUT_FOLDER_TRAIN + input_file)
train_signals.append(signal)
train_signals = np.transpose(np.array(train_signals), (1, 2, 0))
for input_file in INPUT_FILES_TEST:
signal = read_signals(INPUT_FOLDER_TEST + input_file)
test_signals.append(signal)
test_signals = np.transpose(np.array(test_signals), (1, 2, 0))
train_labels = read_labels(LABELFILE_TRAIN)
test_labels = read_labels(LABELFILE_TEST)
[no_signals_train, no_steps_train, no_components_train] = np.shape(train_signals)
[no_signals_test, no_steps_test, no_components_test] = np.shape(train_signals)
no_labels = len(np.unique(train_labels[:]))
print("The train dataset contains {} signals, each one of length {} and {} components ".format(no_signals_train, no_steps_train, no_components_train))
print("The test dataset contains {} signals, each one of length {} and {} components ".format(no_signals_test, no_steps_test, no_components_test))
print("The train dataset contains {} labels, with the following distribution:\n {}".format(np.shape(train_labels)[0], Counter(train_labels[:])))
print("The test dataset contains {} labels, with the following distribution:\n {}".format(np.shape(test_labels)[0], Counter(test_labels[:])))
uci_har_signals_train, uci_har_labels_train = randomize(train_signals, np.array(train_labels))
uci_har_signals_test, uci_har_labels_test = randomize(test_signals, np.array(test_labels))
# -
# # 2. Generating features for the UCI-HAR features
def get_uci_har_features(dataset, labels, waveletname, truncated_len=None):
uci_har_features = []
# Take the DWT of each component and concat them end-to-end
for signal_no in range(0, len(dataset)):
features = []
for signal_comp in range(0,dataset.shape[2]):
signal = dataset[signal_no, :, signal_comp]
list_coeff = pywt.wavedec(signal, waveletname, mode='per')
# convert the wavelet decomposition to array
comp_dwt=[]
for coeff in list_coeff:
comp_dwt.extend(coeff)
features.append(comp_dwt[:truncated_len])
uci_har_features.append(features)
X = np.array(uci_har_features).transpose(0,2,1)
Y = labels
return X, Y
# +
waveletname = 'rbio3.1'
truncate_len = 8
t_start = perf_counter()
x_train, y_train = get_uci_har_features(uci_har_signals_train, uci_har_labels_train, \
waveletname, truncated_len=truncate_len)
x_test, y_test = get_uci_har_features(uci_har_signals_test, uci_har_labels_test, \
waveletname, truncated_len=truncate_len)
t_stop = perf_counter()
t_diff = t_stop-t_start
print ('Time for DWT preprocessing {} seconds'.format(t_diff))
print ('Training data shape: {}'.format(x_train.shape))
print ('Test data shape: {}'.format(x_test.shape))
# -
# # 3. Classifying the train and test sets
# +
num_classes = 6
batch_size = 16
epochs = 128
input_shape = np.shape(x_train)[1:]
print('input_shape: {}'.format(input_shape))
# convert the data to the right type
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
# convert class vectors to binary class matrices - this is for use in the
# categorical_crossentropy loss below
y_train = list(y_train)
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = list(y_test)
y_test = keras.utils.to_categorical(y_test, num_classes)
# +
model = Sequential()
normalizer = preprocessing.Normalization()
normalizer.adapt(x_train)
model.add(keras.Input(shape=input_shape))
model.add(normalizer)
model.add(Dense(16, kernel_regularizer=keras.regularizers.l1_l2(l1=1e-4,l2=1e-4)))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dense(32, kernel_regularizer=keras.regularizers.l1_l2(l1=5e-4,l2=5e-4)))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dense(32, kernel_regularizer=keras.regularizers.l1_l2(l1=5e-4,l2=5e-4)))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Dense(16, kernel_regularizer=keras.regularizers.l1_l2(l1=5e-4,l2=5e-4)))
model.add(BatchNormalization())
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(num_classes, kernel_regularizer=keras.regularizers.l1_l2(l1=1e-4,l2=1e-4),\
activation='softmax'))
model.summary()
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
es = EarlyStopping(monitor='val_loss', verbose=0, patience=8)
t_start = perf_counter()
model.fit(x_train, y_train, batch_size=batch_size,
epochs=epochs, verbose=0,
validation_data=(x_test, y_test),
callbacks=[history,es])
t_stop = perf_counter()
t_diff = t_stop-t_start
print ('Time to train the network {} seconds'.format(t_diff))
train_score = model.evaluate(x_train, y_train, verbose=0)
print('Train loss: {}, Train accuracy: {}'.format(train_score[0], train_score[1]))
test_score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss: {}, Test accuracy: {}'.format(test_score[0], test_score[1]))
# +
fig, axarr = plt.subplots(figsize=(14,7), ncols=2)
axarr[0].plot(history.history['accuracy'], label='train accuracy')
axarr[0].plot(history.history['val_accuracy'], label='test accuracy')
axarr[0].set_xlabel('Number of Epochs', fontsize=18)
axarr[0].set_ylabel('Accuracy', fontsize=18)
axarr[0].set_ylim([0.5,1])
axarr[0].legend()
axarr[1].plot(history.history['loss'], label='train loss')
axarr[1].plot(history.history['val_loss'], label='test loss')
axarr[1].set_xlabel('Number of Epochs', fontsize=18)
axarr[1].set_ylabel('Loss', fontsize=18)
axarr[1].legend()
plt.show()
# -
|
notebooks/WV11 - Evaluating truncated DWT preprocessing for classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 데모 - 우리는 AI를 사용하여, 수송에 영향을 주고 수익 증가 기회가 되기도 하는, 날씨 상태의 변경에 대응하는 공급망을 만들어 가상의 글로벌 소매 기업을 도와 줄 것입니다
#
# 
#
# ## 공급망이란 무엇입니까?
#
# 
#
# - 공급망이란 기업과 기업의 공급자 사이에서 특정 상품을 생산하여 분배하는 네트워크입니다. 공급망은 제품이나 서비스가 소비자에게 도달하는 일련의 과정을 말합니다.
# - 최적화된 공급망은 비용을 낮춰주고 더 빠른 생산 사이클을 형성할 수 있도록 해줍니다. 따라서 공급망 관리는 아주 중요한 과정입니다.
# - 공급망은 물리적인 상품이 많은 세상에서 아주 중요한 요소입니다. 기업은 공급망을 최적화하고 개선하는 데 지속적으로 많은 비용과 노력을 투자하며, 이는 멈추지 않고 계속될 것입니다.
# - 5%의 수송 비용을 줄일 수 있다면 큰 변화가 일어납니다.
# - 수송망 관리 시스템은 특히 많은 부분으로 나누어지는 제조 과정의 복잡도와 비용을 줄일 수 있습니다.
# - 1 의류 제조업자는 원자재를 생산 라인으로 옮길 것입니다.
# - 2 그러면 제조업자는 기계를 가동하고 원자재를 사용하는 다른 일을 수행하는 데 노동 비용을 소모할 것입니다.
# - 3 제품이 생산되고 나면 소비자에게 팔릴 때까지 포장 및 저장될 것입니다.
#
# 
#
# ## 공급망 문제
#
# 
#
# - 공급망을 계획하는 아주 비싼 인력을 많이 고용해야 합니다.
# - 공급망의 각 단계별로
# - 보통 다른 기능이나 파트너와 충돌을 일으킵니다.
# - 각각 국지적으로만 최적화되어 더 큰 기회를 놓치고 있습니다.
# - 이미 오래되버린 데이터로 작업하기 때문에 나쁜 결정으로 이어지고 있습니다.
# - 지나치게 단순화된 모델을 사용하여 실제 세상과 관련이 별로 없습니다.
#
# 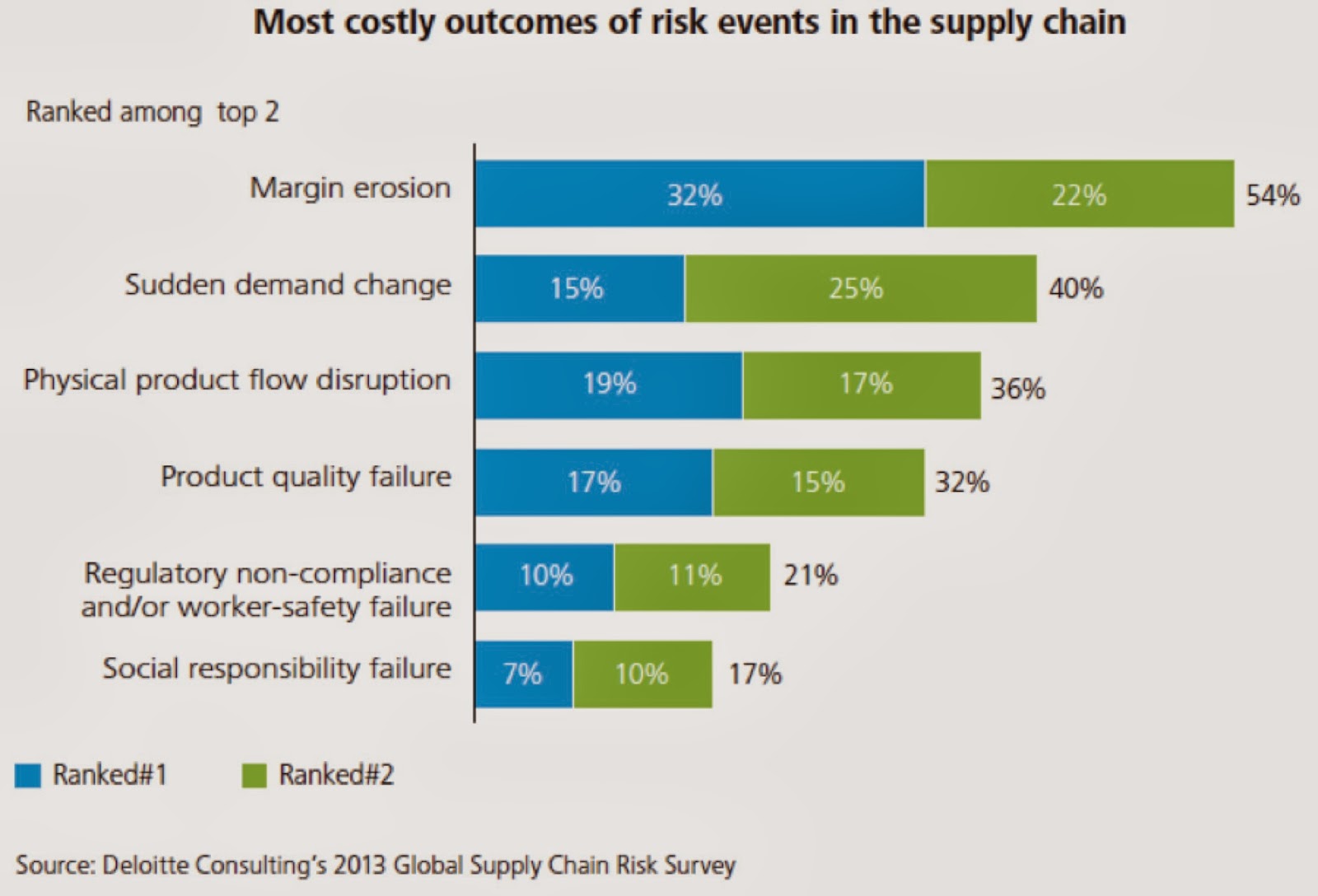
#
# ## 스타트업은 AI로 공급망을 어떻게 향상시킬 수 있을까요?
#
# 
#
# - 1 실시간 데이터에 접근합니다.
# - 오늘날의 많은 공급망은 오래 전 데이터를 바탕으로 계획을 세웠으며, 이는 최선의 선택이 아니기에 의사 결정이 다소 부실할 수 있습니다.
# - 2 커뮤니티 (다중 집단의) 데이터에 접근합니다 (기업 외부에 있는 데이타는 매우 유용합니다)
# - 3 광범위한 네트워크의 목적성을 지원합니다.
# - AI 솔루션은 공급망 내의 제약 상황을 직면했을 때에도, 글로벌 고객들의 목적을 달성을 지원해야 합니다.
# - 4 의사결정 과정은 수익이 증대되는 방향으로 되어야 하며, 변경 시의 비용도 고려해야 합니다.
# - AI툴은 의사결정할 시, 수익 증대와 변경 비용 간의 트레이드오프도 고려해야 합니다.
# - 5 의사결정 과정은 지속적이고 자기 학습적이어야 하며 스스로를 모니터링 해야합니다.
#
# 
#
# - 변동성과 지연은 상호 순환적인 문제이며, 실행 효율은 끊임없이 바뀝니다. AI 시스템은 이 문제를 단기적으로 보아서는 안되고, 끊임없이 주시해야 합니다.
# - 6 AI 엔진은 자동으로 의사를 결정해야 합니다.
# - 알고리즘이 똑똑한 결정만을 하는 게 아니라, 그것을 자동으로 수행할 수 있어야 유의미한 값을 얻을 수 있습니다.
# - 7 AI 엔진은 확장성이 있어야 합니다.
# - 큰 규모의 공급망은 수억개는 아니더라도 수백만개의 비축 장소가 있습니다. AI 솔루션은 반드시 큰 규모의 데이터에 대해서도 빠르고 현명한 판단을 내려야 합니다.
# - 8 사용자가 시스템과 맞물릴 방법이 있어야 합니다.
# - AI는 “블랙 박스” 안에서 동작하고 있어서는 안됩니다. 사용자 인터페이스는 사용자가 결정의 기준이나 파급 효과를 볼 수 있게 해야 하며, AI가 해결할 수 없는 이슈를 사용자가 이해할 수 있도록 해야 합니다.
#
# 
#
# ## AI가 이 문제를 해결할 수 있는 다양한 방법에는 무엇이 있습니까?
#
# ##### Forrester의 최근 설문 조사에 따르면, 오직 13퍼센트의 기업만이 물류에 AI 시스템 도입을 위해 선도 및 투자하는 조직이 있다고 답했습니다.
# ##### 공급망에서 생성되지만 충분히 이용되지 않은 데이터와 공급망이 결합된다면 어마어마한 파괴력이 있습니다. AI + IOT + 블록 체인 = 자동화된 공급망 (스스로 지각하고, 스스로 계획하고, 스스로 결정합니다)
#
# 
#
# ### 조달 운영을 위한 챗봇
#
# 
#
# - 조달 관련 작업을 효과적으로 하기 위해서는 챗봇이 자동화되고 개선되어야 하며, 이를 위해 견고하고 똑똑한 데이터셋이 필요합니다.
# - ‘procuebot’은
# - 1 간단한 대화중 공급자에게 말을 걸 수 있습니다.
# - 2 관리 및 준수 자료에 대한 행동을 설정하고, 공급자에게 행동을 취할 수 있습니다.
# - 3 구매 요청을 자동으로 배치합니다.
# - 4 물품 조달 기능 또는 공급자(들) 집합과 관련한 내부 질문을 조사하고 대답합니다.
# - 5 송장과 지불/구매 요청의 수신/파일링/문서화 (Smith 2016).
# - http://chymebot.com/ 이 하나의 예시입니다.
#
# ### 공급망 계획을 위한 머신러닝
#
# - 재고량, 수요, 공급의 예측을 도울 수 있습니다.
# - 공급망 관리 전문가들은 똑똑한 알고리즘과 머신-머신 간의 빅데이터 분석을 토대로 최선의 시나리오를 받을 것입니다.
# - 균형잡힌 수요와 공급으로 물품 배송을 최적화할 수 있고, 사람의 분석이 필요하지 않으며, 대신 성공적인 파라메터를 설정하기 위한 행동만이 필요합니다.
# - https://www.clearmetal.com/ 이 하나의 예시입니다.
#
# ### 창고 관리를 위한 머신 러닝
#
# 
#
# - 공급망에는 적절한 창고 및 재고 관리가 필수적입니다.
# - 수요 예측을 고려하지 않는 공급 결함(공급 과잉 또는 결핍)이 소비자 대상의 기업/소매상에게는 큰 재앙이 될 수 있습니다.
# - 머신러닝은 무한히 예측을 제공해주며, 그 예측들은 다시 스스로를 개선시킵니다.
# - 이러한 기능들로 창고 관리를 개선할 수 있습니다.
# - https://rubikloud.com/ 이 하나의 예시입니다.
#
# ### 물류와 배송을 위한 자율주행차
#
# 
#
# - 더 빠르고 정확한 배송은 시간과 비용을 줄여줍니다.
# - 또한 환경 친화적인 운영 요소를 추가해주고, 인건비를 감소시켜줍니다. 무엇보다도 경쟁사와의 격차를 벌여줍니다.
# - 법적으로 운전자는 8시간의 휴식 없이는, 하루에 11시간을 운전할 수 없지만, 운전자가 없는 트럭은 24시간 운행이 가능합니다.
# - 즉 이 기술은 25퍼센트의 비용으로 미국의 운송 네트워크 효율을 두배로 늘여줍니다.
# - wwww.comma.ai 가 하나의 예시입니다.
#
# ### 데이터 정리와 데이터 신뢰도 향상을 위한 자연어 처리
#
# 
#
# - 자연어 처리 기술은 공급자와 관련된 데이터셋을 만들 수 있으며, 언어 장벽 때문에 손대지 못한 정보를 해석할 수 있습니다.
# - 기업의 사회적 책임이나 지속가능성 및 관리의 관점에서, 자연어 처리 기술은 이전에는 구매자-공급자 간 언어 장벽 때문에 하지 못했던 감사 및 준수 과정을 간소화할 수 있습니다.
# - https://www.datalogue.io/
#
# ### 공급자 선택 및 관리를 위한 머신러닝과 예측분석
#
# 
#
# - 공급자 리스크는 글로벌 브랜드에게는 일종의 족쇄입니다.
# - 공급자의 운영이나 평판 문제로, 당신의 회사가 책임을 지게될 수도 있습니다.
# - 하지만 공급자 선택 및 리스크 관리를 위한 최선의 시나리오가 있다면 어떨까요?
# - 공급자 평가, 회계 감사, 신용 평점과 같이 공급자 관리에서 발생하는 데이터셋은 향후의 결정에서 중요한 기초 자료가 됩니다.
# - 머신러닝과 알고리즘의 도움으로, 이러한 수작업은 자동화가 될 수 있습니다.
# - AI가 최선의 공급자를 얄려줄 수 있습니다.
# +
|
AI for Supply Chains.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (TensorFlow 1.15 Python 3.7 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:ap-northeast-1:102112518831:image/tensorflow-1.15-cpu-py37-ubuntu18.04-v7
# ---
# # Keras を 使った mnist 分類器の Neo コンパイルとコンパイルモデルの推論
# * まずは Neo を前提条件である (keras の場合) Keras 2.2.4 と neo モデルのランタイム(dlr) を インストール
# !pip install -U keras==2.2.4
# !pip install matplotlib
# !pip install dlr
import tensorflow as tf
from sagemaker.tensorflow import TensorFlow
from tensorflow.keras.datasets import mnist
import numpy as np
from matplotlib import pyplot as plt
import keras
import sagemaker, os
from time import sleep
from uuid import uuid4
import dlr
print(keras.__version__)
print(tf.__version__)
# +
work_dir = './work'
# !mkdir -p {work_dir}
TRAIN_X_PATH = os.path.join(work_dir,'train_X.npy')
TEST_X_PATH = os.path.join(work_dir,'test_X.npy')
TRAIN_Y_PATH = os.path.join(work_dir,'train_y.npy')
TEST_Y_PATH = os.path.join(work_dir,'test_y.npy')
(train_X, train_y), (test_X, test_y) = mnist.load_data()
train_X = (train_X-127.5)/127.5
test_X = (test_X-127.5)/127.5
# channel last
train_X = train_X.reshape((train_X.shape[0],train_X.shape[1],train_X.shape[2],1))
test_X = test_X.reshape((test_X.shape[0],test_X.shape[1],test_X.shape[2],1))
# one-hot
train_y = np.eye(10)[train_y]
test_y = np.eye(10)[test_y]
np.save(TRAIN_X_PATH,train_X)
np.save(TEST_X_PATH,test_X)
np.save(TRAIN_Y_PATH,train_y)
np.save(TEST_Y_PATH,test_y)
# -
# check shapes
print(train_X.shape,train_y.shape,test_X.shape,test_y.shape)
role = sagemaker.get_execution_role()
sess = sagemaker.session.Session()
train_X_uri = sess.upload_data(path=TRAIN_X_PATH, bucket=sess.default_bucket(), key_prefix='sagemaker/mnist')
train_y_uri = sess.upload_data(path=TRAIN_Y_PATH, bucket=sess.default_bucket(), key_prefix='sagemaker/mnist')
test_X_uri = sess.upload_data(path=TEST_X_PATH, bucket=sess.default_bucket(), key_prefix='sagemaker/mnist')
test_y_uri = sess.upload_data(path=TEST_Y_PATH, bucket=sess.default_bucket(), key_prefix='sagemaker/mnist')
print(train_X_uri)
print(train_y_uri)
print(test_X_uri)
print(test_y_uri)
# ## Classifier Train
estimator = TensorFlow(
entry_point='./src/classifier_train.py',
role=role,
instance_count=1,
instance_type='ml.g4dn.xlarge',
framework_version='1.13', # SageMaker Neo を使うために keras 2.2.4 に合わせる
py_version='py3',
hyperparameters={
'epochs':30,
'increment':'False'
}
)
# +
# %%time
print(train_X_uri[:-11]) # dir以下全て
estimator.fit({
'train': train_X_uri[:-11],
})
# -
classifier_model_uri = estimator.latest_training_job.describe()['ModelArtifacts']['S3ModelArtifacts']
print(classifier_model_uri)
# ## Compile
import boto3
sm_client = boto3.client('sagemaker')
classifier_output_s3_location = f's3://{sagemaker.session.Session().default_bucket()}/sagemaker/mnist/classifier/compilied_model'
print(classifier_output_s3_location)
# ### Classifier のコンパイル
classifier_compile_jobname = f'classifier-{str(uuid4())}'
print(classifier_compile_jobname)
response = sm_client.create_compilation_job(
CompilationJobName=classifier_compile_jobname,
RoleArn=role,
InputConfig={
'S3Uri': classifier_model_uri,
'DataInputConfig': '{"input_1":[1,1,28,28]}',
'Framework': 'KERAS',
},
OutputConfig={
'S3OutputLocation': classifier_output_s3_location,
'TargetDevice': 'ml_c5',
},
StoppingCondition={
'MaxRuntimeInSeconds': 900,
'MaxWaitTimeInSeconds': 900
},
)
# ### コンパイルの完了待ち
while True:
response = sm_client.describe_compilation_job(CompilationJobName=classifier_compile_jobname)
status = response['CompilationJobStatus']
if status in ['COMPLETED','FAILED','STOPPED'] :
print('!')
print(status)
classifier_neo_model_uri = response['ModelArtifacts']['S3ModelArtifacts']
break
else:
print('.',end='')
sleep(5)
print(classifier_neo_model_uri)
# ### ファイルの配置
# +
neo_dir = './neo_model'
# classifier download
# !aws s3 cp {classifier_model_uri} {work_dir}/
# !tar zxvf {work_dir}/model.tar.gz -C ./
# !aws s3 cp {classifier_neo_model_uri} {work_dir}/
# !mkdir -p {neo_dir}
# !tar zxvf {work_dir}/model-ml_c5.tar.gz -C {neo_dir}
# !rm {work_dir}/model-ml_c5.tar.gz
# -
classifier_output_s3_location
# ## 動作確認
# ### keras model
keras_model = keras.models.load_model('classifier.h5')
np.argmax(keras_model.predict(test_X[0:1,:,:,:]))
# ### Neo Model
# +
import dlr
import numpy as np
classifier_neo = dlr.DLRModel(neo_dir, 'cpu', 0)
# -
pred_y = classifier_neo.run(test_X[0,:,:,:].reshape(1,1,28,28))[0]
np.argmax(pred_y)
# # 推論結果比較
# 微妙に差異がある
keras_model.predict(test_X[0:1,:,:,:]),classifier_neo.run(test_X[0,:,:,:].reshape(1,1,28,28))[0]
|
NeoCompileSample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Geological Timescale
# ======================
#
# pyrolite includes a simple geological timescale, based on a recent verion
# of the International Chronostratigraphic Chart [#ICS]_. The
# :class:`~pyrolite.util.time.Timescale` class can be used to look up names for
# specific geological ages, to look up times for known geological age names
# and to access a reference table for all of these.
#
# .. [#ICS] <NAME>., <NAME>., <NAME>., <NAME>., 2013.
# `The ICS International Chronostratigraphic Chart <http://www.stratigraphy.org/index.php/ics-chart-timescale>`__.
# Episodes 36, 199–204.
#
#
# First we'll create a timescale:
#
#
#
from pyrolite.util.time import Timescale
ts = Timescale()
# From this we can look up the names of ages (in million years, or Ma):
#
#
#
ts.named_age(1212.1)
# As geological age names are hierarchical, the name you give an age depends on what
# level you're looking at. By default, the timescale will return the most specific
# non-null level. The levels accessible within the timescale are listed
# as an attribute:
#
#
#
ts.levels
# These can be used to refine the output names to your desired level of specificity
# (noting that for some ages, the levels which are accessible can differ; see the chart):
#
#
#
ts.named_age(1212.1, level="Epoch")
# The timescale can also do the inverse for you, and return the timing information for a
# given named age:
#
#
ts.text2age("Holocene")
# We can use this to create a simple template to visualise the geological timescale:
#
#
#
# +
import pandas as pd
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, figsize=(5, 10))
for ix, level in enumerate(ts.levels):
ldf = ts.data.loc[ts.data.Level == level, :]
for pix, period in ldf.iterrows():
ax.bar(
ix,
period.Start - period.End,
facecolor=period.Color,
bottom=period.End,
width=1,
edgecolor="k",
)
ax.set_xticks(range(len(ts.levels)))
ax.set_xticklabels(ts.levels, rotation=60)
ax.xaxis.set_ticks_position("top")
ax.set_ylabel("Age (Ma)")
ax.invert_yaxis()
# -
# This doesn't quite look like the geological timescale you may be used to. We can improve
# on the output somewhat with a bit of customisation for the positioning. Notably, this is
# less readable, but produces something closer to what we're after. Some of this may soon
# be integrated as a :class:`~pyrolite.util.time.Timescale` method, if there's interest.
#
#
#
# +
import numpy as np
from matplotlib.patches import Rectangle
# first let's set up some x-limits for the different timescale levels
xlims = {
"Eon": (0, 1),
"Era": (1, 2),
"Period": (2, 3),
"Superepoch": (3, 4),
"Epoch": (3, 5),
"Age": (5, 7),
}
fig, ax = plt.subplots(1, figsize=(4, 10))
for ix, level in enumerate(ts.levels[::-1]):
ldf = ts.data.loc[ts.data.Level == level, :]
for pix, period in ldf.iterrows():
left, right = xlims[level]
if ix != len(ts.levels) - 1:
time = np.mean(ts.text2age(period.Name))
general = None
_ix = ix
while general is None:
try:
general = ts.named_age(time, level=ts.levels[::-1][_ix + 1])
except:
pass
_ix += 1
_l, _r = xlims[ts.levels[::-1][_ix]]
if _r > left:
left = _r
rect = Rectangle(
(left, period.End),
right - left,
period.Start - period.End,
facecolor=period.Color,
edgecolor="k",
)
ax.add_artist(rect)
ax.set_xticks([np.mean(xlims[lvl]) for lvl in ts.levels])
ax.set_xticklabels(ts.levels, rotation=60)
ax.xaxis.set_ticks_position("top")
ax.set_xlim(0, 7)
ax.set_ylabel("Age (Ma)")
ax.set_ylim(500, 0)
|
docs/source/examples/util/timescale.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="sRsAVXNVc9mk" colab_type="code" colab={}
from joblib import load
top_recs_df = load("/content/drive/My Drive/MScA/Machine Learning/ML Final Project/Data/cb_top_user_preds.joblib")
# + id="M0gsolpLd9Ju" colab_type="code" colab={}
import pandas as pd
train_df = pd.read_pickle("/content/drive/My Drive/MScA/Machine Learning/ML Final Project/Data/Final Input Data/train_df_with_valid_recommendations.pkl")
validation_df = pd.read_pickle("/content/drive/My Drive/MScA/Machine Learning/ML Final Project/Data/Final Input Data/validation_df_with_valid_recommendations.pkl")
test_df = pd.read_pickle("/content/drive/My Drive/MScA/Machine Learning/ML Final Project/Data/Final Input Data/test_df_with_valid_recommendations.pkl")
# + id="cXMTO2n7eXm_" colab_type="code" colab={}
train_df = train_df.merge(top_recs_df, how='left',on='reviewerID',right_index=False)
validation_df = train_df.merge(top_recs_df, how='left',on='reviewerID',right_index=False)
test_df = train_df.merge(top_recs_df, how='left',on='reviewerID',right_index=False)
# + id="88J0-nYhdztm" colab_type="code" colab={}
# Calculate accuracy @ 1
train_df["accuracy_at_1"] = 0
for index, row in train_df.iterrows():
train_df.loc[index, "accuracy_at_1"] = len(set(row["top_1_recommended"]).intersection(set(row["valid_recommendations"][:1])))/1
# Calculate accuracy @ 3
train_df["accuracy_at_3"] = 0
for index, row in train_df.iterrows():
train_df.loc[index, "accuracy_at_3"] = len(set(row["top_3_recommended"]).intersection(set(row["valid_recommendations"][:3])))/3
# Calculate accuracy @ 5
train_df["accuracy_at_5"] = 0
for index, row in train_df.iterrows():
train_df.loc[index, "accuracy_at_5"] = len(set(row["top_5_recommended"]).intersection(set(row["valid_recommendations"][:5])))/5
# Calculate accuracy @ 10
train_df["accuracy_at_10"] = 0
for index, row in train_df.iterrows():
train_df.loc[index, "accuracy_at_10"] = len(set(row["top_10_recommended"]).intersection(set(row["valid_recommendations"][:10])))/10
# Calculate accuracy @ 15
train_df["accuracy_at_15"] = 0
for index, row in train_df.iterrows():
train_df.loc[index, "accuracy_at_15"] = len(set(row["top_15_recommended"]).intersection(set(row["valid_recommendations"][:15])))/15
# + id="j3i0XLS9dP6x" colab_type="code" colab={}
# Calculate accuracy @ 1
validation_df["accuracy_at_1"] = 0
for index, row in validation_df.iterrows():
validation_df.loc[index, "accuracy_at_1"] = len(set(row["top_1_recommended_y"]).intersection(set(row["valid_recommendations"][:1])))/1
# Calculate accuracy @ 3
validation_df["accuracy_at_3"] = 0
for index, row in validation_df.iterrows():
validation_df.loc[index, "accuracy_at_3"] = len(set(row["top_3_recommended_y"]).intersection(set(row["valid_recommendations"][:3])))/3
# Calculate accuracy @ 5
validation_df["accuracy_at_5"] = 0
for index, row in validation_df.iterrows():
validation_df.loc[index, "accuracy_at_5"] = len(set(row["top_5_recommended_y"]).intersection(set(row["valid_recommendations"][:5])))/5
# Calculate accuracy @ 10
validation_df["accuracy_at_10"] = 0
for index, row in validation_df.iterrows():
validation_df.loc[index, "accuracy_at_10"] = len(set(row["top_10_recommended_y"]).intersection(set(row["valid_recommendations"][:10])))/10
# Calculate accuracy @ 15
validation_df["accuracy_at_15"] = 0
for index, row in validation_df.iterrows():
validation_df.loc[index, "accuracy_at_15"] = len(set(row["top_15_recommended_y"]).intersection(set(row["valid_recommendations"][:15])))/15
# + id="cVFV7Wc7oJeQ" colab_type="code" colab={}
# Calculate accuracy @ 1
test_df["accuracy_at_1"] = 0
for index, row in test_df.iterrows():
test_df.loc[index, "accuracy_at_1"] = len(set(row["top_1_recommended_y"]).intersection(set(row["valid_recommendations"][:1])))/1
# Calculate accuracy @ 3
test_df["accuracy_at_3"] = 0
for index, row in test_df.iterrows():
test_df.loc[index, "accuracy_at_3"] = len(set(row["top_3_recommended_y"]).intersection(set(row["valid_recommendations"][:3])))/3
# Calculate accuracy @ 5
test_df["accuracy_at_5"] = 0
for index, row in test_df.iterrows():
test_df.loc[index, "accuracy_at_5"] = len(set(row["top_5_recommended_y"]).intersection(set(row["valid_recommendations"][:5])))/5
# Calculate accuracy @ 10
test_df["accuracy_at_10"] = 0
for index, row in test_df.iterrows():
test_df.loc[index, "accuracy_at_10"] = len(set(row["top_10_recommended_y"]).intersection(set(row["valid_recommendations"][:10])))/10
# Calculate accuracy @ 15
test_df["accuracy_at_15"] = 0
for index, row in test_df.iterrows():
test_df.loc[index, "accuracy_at_15"] = len(set(row["top_15_recommended_y"]).intersection(set(row["valid_recommendations"][:15])))/15
# + id="eQpHoMsOm3XU" colab_type="code" colab={}
import numpy as np
MAP_train = np.mean([np.mean(train_df["accuracy_at_3"]),
np.mean(train_df["accuracy_at_5"]),
np.mean(train_df["accuracy_at_10"]),
np.mean(train_df["accuracy_at_15"])])
# + id="HSe8oAiH4fbr" colab_type="code" colab={}
MAP_val = np.mean([np.mean(validation_df["accuracy_at_3"]),
np.mean(validation_df["accuracy_at_5"]),
np.mean(validation_df["accuracy_at_10"]),
np.mean(validation_df["accuracy_at_15"])])
# + id="oj59x4_94kcJ" colab_type="code" colab={}
MAP_test = np.mean([np.mean(test_df["accuracy_at_3"]),
np.mean(test_df["accuracy_at_5"]),
np.mean(test_df["accuracy_at_10"]),
np.mean(test_df["accuracy_at_15"])])
# + id="9XghPToSykUS" colab_type="code" outputId="e93e9d41-1c61-473d-a568-9cbdc25ab06b" executionInfo={"status": "ok", "timestamp": 1591657060558, "user_tz": 300, "elapsed": 2405697, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjiHm_gl0Be0uWIlRtZbdiCSRPpl0jgDwy6bkBn=s64", "userId": "01272390123410607150"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
MAP_train
# + id="VrmuiyLQ4pOK" colab_type="code" outputId="0629a3d1-ccd1-495d-8145-0040dcadee54" executionInfo={"status": "ok", "timestamp": 1591657060558, "user_tz": 300, "elapsed": 2405685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjiHm_gl0Be0uWIlRtZbdiCSRPpl0jgDwy6bkBn=s64", "userId": "01272390123410607150"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
MAP_val
# + id="rAouRfjc4rGG" colab_type="code" outputId="69555f4c-9264-47b4-9bd4-51766531c383" executionInfo={"status": "ok", "timestamp": 1591657060559, "user_tz": 300, "elapsed": 2405675, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjiHm_gl0Be0uWIlRtZbdiCSRPpl0jgDwy6bkBn=s64", "userId": "01272390123410607150"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
MAP_test
# + id="3nXv94A97t-X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="35fc9258-c564-4c7a-a30a-84f8d643c7f7" executionInfo={"status": "ok", "timestamp": 1591657801168, "user_tz": 300, "elapsed": 435, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjiHm_gl0Be0uWIlRtZbdiCSRPpl0jgDwy6bkBn=s64", "userId": "01272390123410607150"}}
import matplotlib.pyplot as plt
# 5 values for K
x_train = [1, 4, 7, 10, 13]
x_val = [2, 5, 8, 11, 14]
# Training means
y_train = [np.mean(train_df["accuracy_at_1"]),
np.mean(train_df["accuracy_at_3"]),
np.mean(train_df["accuracy_at_5"]),
np.mean(train_df["accuracy_at_10"]),
np.mean(train_df["accuracy_at_15"])]
# Training stds
error_train = [np.std(train_df["accuracy_at_1"]),
np.std(train_df["accuracy_at_3"]),
np.std(train_df["accuracy_at_5"]),
np.std(train_df["accuracy_at_10"]),
np.std(train_df["accuracy_at_15"])]
# Validation means
y_val = [np.mean(validation_df["accuracy_at_1"]),
np.mean(validation_df["accuracy_at_3"]),
np.mean(validation_df["accuracy_at_5"]),
np.mean(validation_df["accuracy_at_10"]),
np.mean(validation_df["accuracy_at_15"])]
# Validation stds
error_val = [np.std(validation_df["accuracy_at_1"]),
np.std(validation_df["accuracy_at_3"]),
np.std(validation_df["accuracy_at_5"]),
np.std(validation_df["accuracy_at_10"]),
np.std(validation_df["accuracy_at_15"])]
# Plot errorbars
plt.bar(x = x_train, height = y_train, yerr = error_train, color = "peachpuff", label = "Train")
plt.bar(x = x_val, height = y_val, yerr = error_val, color = "orange", label = "Validation")
# Titles and axis labels
plt.title("Precision at k", fontweight = "bold", fontsize = 14)
plt.xlabel("Number of recommendations")
plt.ylabel("Precision")
plt.xticks(ticks = [1.5, 4.5, 7.5, 10.5, 13.5], labels = [1, 3, 5, 10, 15])
plt.ylim(top = 1, bottom = 0)
# Add legend
plt.legend()
# Show plot
plt.show()
# + id="pW7Hv_T9m_8X" colab_type="code" outputId="3f66cbfc-714f-4d2c-8b1c-c2fa5d6cb139" executionInfo={"status": "ok", "timestamp": 1591657804500, "user_tz": 300, "elapsed": 271, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "01272390123410607150"}} colab={"base_uri": "https://localhost:8080/", "height": 295}
# Plot MAP
plt.bar(x = 1, height = MAP_train, color = "peachpuff", label = "Train")
plt.bar(x = 2, height = MAP_val, color = "orange", label = "Validation")
# Titles and axis labels
plt.title("Mean Average Precision (MAP)", fontweight = "bold", fontsize = 14)
plt.xlabel("Set")
plt.ylabel("MAP")
plt.ylim(bottom = 0, top = 1)
plt.xticks(ticks = [1, 2], labels = ["Train", "Validation"])
# Show plot
plt.show()
|
Analysis/Content-Based/CB_Validation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import requests
import re
import json
from os import walk
from multiprocessing.pool import ThreadPool
# +
URL = "https://s3.amazonaws.com/irs-form-990/index_2017.json"
response = requests.get(URL)
with open("index_2017.json", 'wb') as f:
f.write(response.content)
# -
with open("index_2017.json") as f:
data = json.load(f)
data = data[list(data.keys())[0]]
df = pd.DataFrame.from_dict(data)
df.tail(5)
df.shape
# +
def download_url(url):
file_name_start = url.rfind('/') + 1
file_name = url[file_name_start:]
output_dir = "/home/meso/git_repo/exploring_990/data_2017/"
file_name = output_dir + file_name
r = requests.get(url, stream=True)
if r.status_code == requests.codes.ok:
with open(file_name, 'wb') as f:
f.write(r.content)
return url
urls = []
for i in range(df.shape[0]):
urls.append(data[i]['URL'])
results = ThreadPool(100).imap_unordered(download_url, urls)
# -
f = []
for (dirpath, dirnames, filenames) in walk('/home/meso/git_repo/exploring_990/data_2017/'):
f.extend(filenames)
break
u = list(df['URL'])
len(u)
# +
file_names_list = []
for i in range(len(u)):
file_loc = u[i].rfind("/") + 1
file_name = u[i][file_loc:]
file_names_list.append(file_name)
# -
print(len(file_names_list), str(":"), len(f))
diff = list(set(file_names_list) - set(f))
len(diff)
file_loc = u[i].rfind("/") + 1
url_pre = u[0][:file_loc]
urls = []
for i in range(len(diff)):
url = url_pre + diff[i]
urls.append(url)
for url in urls:
download_url(url)
f = []
for (dirpath, dirnames, filenames) in walk('/home/meso/git_repo/exploring_990/data_2017/'):
f.extend(filenames)
break
len(f)
diff = list(set(file_names_list) - set(f))
len(diff)
def list_duplicates(seq):
seen = set()
seen_add = seen.add
return [idx for idx, item in enumerate(seq) if item in seen or seen_add(item)]
print(list_duplicates(file_names_list))
dup_df = df.iloc[list_duplicates(file_names_list)]
dup_ein = pd.DataFrame(dup_df['EIN'])
dup_ein
# +
duplicates = pd.DataFrame([])
for dup in range(dup_ein.shape[0]):
ein = dup_ein.iloc[dup]['EIN']
duplicates = duplicates.append(df[df['EIN'] == ein])
# -
duplicates
|
data_download_2017.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
"""Sample notebook"""
import pandas as pd
df = pd.read_csv('./Subscription%20Usage%20Data-1557751202.csv')
# -
import numpy as np
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from sklearn.preprocessing import scale
print(df)
|
tests/client/apps/test_config/sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="MhoQ0WE77laV"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab={} colab_type="code" id="vasWnqRgy1H4"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="jYysdyb-CaWM"
# # Basic classification: Classify images of clothing
# + [markdown] colab_type="text" id="S5Uhzt6vVIB2"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="FbVhjPpzn6BM"
# This guide trains a neural network model to classify images of clothing, like sneakers and shirts. It's okay if you don't understand all the details; this is a fast-paced overview of a complete TensorFlow program with the details explained as you go.
#
# This guide uses [tf.keras](https://www.tensorflow.org/guide/keras), a high-level API to build and train models in TensorFlow.
# + colab={} colab_type="code" id="dzLKpmZICaWN"
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + [markdown] colab_type="text" id="yR0EdgrLCaWR"
# ## Import the Fashion MNIST dataset
# + [markdown] colab_type="text" id="DLdCchMdCaWQ"
# This guide uses the [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset which contains 70,000 grayscale images in 10 categories. The images show individual articles of clothing at low resolution (28 by 28 pixels), as seen here:
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
# </td></tr>
# </table>
#
# Fashion MNIST is intended as a drop-in replacement for the classic [MNIST](http://yann.lecun.com/exdb/mnist/) dataset—often used as the "Hello, World" of machine learning programs for computer vision. The MNIST dataset contains images of handwritten digits (0, 1, 2, etc.) in a format identical to that of the articles of clothing you'll use here.
#
# This guide uses Fashion MNIST for variety, and because it's a slightly more challenging problem than regular MNIST. Both datasets are relatively small and are used to verify that an algorithm works as expected. They're good starting points to test and debug code.
#
# Here, 60,000 images are used to train the network and 10,000 images to evaluate how accurately the network learned to classify images. You can access the Fashion MNIST directly from TensorFlow. Import and load the Fashion MNIST data directly from TensorFlow:
# + colab={} colab_type="code" id="7MqDQO0KCaWS"
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# + [markdown] colab_type="text" id="t9FDsUlxCaWW"
# Loading the dataset returns four NumPy arrays:
#
# * The `train_images` and `train_labels` arrays are the *training set*—the data the model uses to learn.
# * The model is tested against the *test set*, the `test_images`, and `test_labels` arrays.
#
# The images are 28x28 NumPy arrays, with pixel values ranging from 0 to 255. The *labels* are an array of integers, ranging from 0 to 9. These correspond to the *class* of clothing the image represents:
#
# <table>
# <tr>
# <th>Label</th>
# <th>Class</th>
# </tr>
# <tr>
# <td>0</td>
# <td>T-shirt/top</td>
# </tr>
# <tr>
# <td>1</td>
# <td>Trouser</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Pullover</td>
# </tr>
# <tr>
# <td>3</td>
# <td>Dress</td>
# </tr>
# <tr>
# <td>4</td>
# <td>Coat</td>
# </tr>
# <tr>
# <td>5</td>
# <td>Sandal</td>
# </tr>
# <tr>
# <td>6</td>
# <td>Shirt</td>
# </tr>
# <tr>
# <td>7</td>
# <td>Sneaker</td>
# </tr>
# <tr>
# <td>8</td>
# <td>Bag</td>
# </tr>
# <tr>
# <td>9</td>
# <td>Ankle boot</td>
# </tr>
# </table>
#
# Each image is mapped to a single label. Since the *class names* are not included with the dataset, store them here to use later when plotting the images:
# + colab={} colab_type="code" id="IjnLH5S2CaWx"
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + [markdown] colab_type="text" id="Brm0b_KACaWX"
# ## Explore the data
#
# Let's explore the format of the dataset before training the model. The following shows there are 60,000 images in the training set, with each image represented as 28 x 28 pixels:
# + colab={} colab_type="code" id="zW5k_xz1CaWX"
train_images.shape
# + [markdown] colab_type="text" id="cIAcvQqMCaWf"
# Likewise, there are 60,000 labels in the training set:
# + colab={} colab_type="code" id="TRFYHB2mCaWb"
len(train_labels)
# + [markdown] colab_type="text" id="YSlYxFuRCaWk"
# Each label is an integer between 0 and 9:
# + colab={} colab_type="code" id="XKnCTHz4CaWg"
train_labels
# + [markdown] colab_type="text" id="TMPI88iZpO2T"
# There are 10,000 images in the test set. Again, each image is represented as 28 x 28 pixels:
# + colab={} colab_type="code" id="2KFnYlcwCaWl"
test_images.shape
# + [markdown] colab_type="text" id="rd0A0Iu0CaWq"
# And the test set contains 10,000 images labels:
# + colab={} colab_type="code" id="iJmPr5-ACaWn"
len(test_labels)
# + [markdown] colab_type="text" id="ES6uQoLKCaWr"
# ## Preprocess the data
#
# The data must be preprocessed before training the network. If you inspect the first image in the training set, you will see that the pixel values fall in the range of 0 to 255:
# + colab={} colab_type="code" id="m4VEw8Ud9Quh"
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] colab_type="text" id="Wz7l27Lz9S1P"
# Scale these values to a range of 0 to 1 before feeding them to the neural network model. To do so, divide the values by 255. It's important that the *training set* and the *testing set* be preprocessed in the same way:
# + colab={} colab_type="code" id="bW5WzIPlCaWv"
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] colab_type="text" id="Ee638AlnCaWz"
# To verify that the data is in the correct format and that you're ready to build and train the network, let's display the first 25 images from the *training set* and display the class name below each image.
# + colab={} colab_type="code" id="oZTImqg_CaW1"
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# + [markdown] colab_type="text" id="59veuiEZCaW4"
# ## Build the model
#
# Building the neural network requires configuring the layers of the model, then compiling the model.
# + [markdown] colab_type="text" id="Gxg1XGm0eOBy"
# ### Set up the layers
#
# The basic building block of a neural network is the *layer*. Layers extract representations from the data fed into them. Hopefully, these representations are meaningful for the problem at hand.
#
# Most of deep learning consists of chaining together simple layers. Most layers, such as `tf.keras.layers.Dense`, have parameters that are learned during training.
# + colab={} colab_type="code" id="9ODch-OFCaW4"
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation='relu'),
keras.layers.Dense(10)
])
# + [markdown] colab_type="text" id="gut8A_7rCaW6"
# The first layer in this network, `tf.keras.layers.Flatten`, transforms the format of the images from a two-dimensional array (of 28 by 28 pixels) to a one-dimensional array (of 28 * 28 = 784 pixels). Think of this layer as unstacking rows of pixels in the image and lining them up. This layer has no parameters to learn; it only reformats the data.
#
# After the pixels are flattened, the network consists of a sequence of two `tf.keras.layers.Dense` layers. These are densely connected, or fully connected, neural layers. The first `Dense` layer has 128 nodes (or neurons). The second (and last) layer returns a logits array with length of 10. Each node contains a score that indicates the current image belongs to one of the 10 classes.
#
# ### Compile the model
#
# Before the model is ready for training, it needs a few more settings. These are added during the model's *compile* step:
#
# * *Loss function* —This measures how accurate the model is during training. You want to minimize this function to "steer" the model in the right direction.
# * *Optimizer* —This is how the model is updated based on the data it sees and its loss function.
# * *Metrics* —Used to monitor the training and testing steps. The following example uses *accuracy*, the fraction of the images that are correctly classified.
# + colab={} colab_type="code" id="Lhan11blCaW7"
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] colab_type="text" id="qKF6uW-BCaW-"
# ## Train the model
#
# Training the neural network model requires the following steps:
#
# 1. Feed the training data to the model. In this example, the training data is in the `train_images` and `train_labels` arrays.
# 2. The model learns to associate images and labels.
# 3. You ask the model to make predictions about a test set—in this example, the `test_images` array.
# 4. Verify that the predictions match the labels from the `test_labels` array.
#
# + [markdown] colab_type="text" id="Z4P4zIV7E28Z"
# ### Feed the model
#
# To start training, call the `model.fit` method—so called because it "fits" the model to the training data:
# + colab={} colab_type="code" id="xvwvpA64CaW_"
model.fit(train_images, train_labels, epochs=10)
# + [markdown] colab_type="text" id="W3ZVOhugCaXA"
# As the model trains, the loss and accuracy metrics are displayed. This model reaches an accuracy of about 0.91 (or 91%) on the training data.
# + [markdown] colab_type="text" id="wCpr6DGyE28h"
# ### Evaluate accuracy
#
# Next, compare how the model performs on the test dataset:
# + colab={} colab_type="code" id="VflXLEeECaXC"
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('\nTest accuracy:', test_acc)
# + [markdown] colab_type="text" id="yWfgsmVXCaXG"
# It turns out that the accuracy on the test dataset is a little less than the accuracy on the training dataset. This gap between training accuracy and test accuracy represents *overfitting*. Overfitting happens when a machine learning model performs worse on new, previously unseen inputs than it does on the training data. An overfitted model "memorizes" the noise and details in the training dataset to a point where it negatively impacts the performance of the model on the new data. For more information, see the following:
# * [Demonstrate overfitting](https://www.tensorflow.org/tutorials/keras/overfit_and_underfit#demonstrate_overfitting)
# * [Strategies to prevent overfitting](https://www.tensorflow.org/tutorials/keras/overfit_and_underfit#strategies_to_prevent_overfitting)
# + [markdown] colab_type="text" id="v-PyD1SYE28q"
# ### Make predictions
#
# With the model trained, you can use it to make predictions about some images.
# The model's linear outputs, [logits](https://developers.google.com/machine-learning/glossary#logits). Attach a softmax layer to convert the logits to probabilities, which are easier to interpret.
# + colab={} colab_type="code" id="DnfNA0CrQLSD"
probability_model = tf.keras.Sequential([model,
tf.keras.layers.Softmax()])
# + colab={} colab_type="code" id="Gl91RPhdCaXI"
predictions = probability_model.predict(test_images)
# + [markdown] colab_type="text" id="x9Kk1voUCaXJ"
# Here, the model has predicted the label for each image in the testing set. Let's take a look at the first prediction:
# + colab={} colab_type="code" id="3DmJEUinCaXK"
predictions[0]
# + [markdown] colab_type="text" id="-hw1hgeSCaXN"
# A prediction is an array of 10 numbers. They represent the model's "confidence" that the image corresponds to each of the 10 different articles of clothing. You can see which label has the highest confidence value:
# + colab={} colab_type="code" id="qsqenuPnCaXO"
np.argmax(predictions[0])
# + [markdown] colab_type="text" id="E51yS7iCCaXO"
# So, the model is most confident that this image is an ankle boot, or `class_names[9]`. Examining the test label shows that this classification is correct:
# + colab={} colab_type="code" id="Sd7Pgsu6CaXP"
test_labels[0]
# + [markdown] colab_type="text" id="ygh2yYC972ne"
# Graph this to look at the full set of 10 class predictions.
# + colab={} colab_type="code" id="DvYmmrpIy6Y1"
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array, true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array, true_label[i]
plt.grid(False)
plt.xticks(range(10))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + [markdown] colab_type="text" id="Zh9yABaME29S"
# ### Verify predictions
#
# With the model trained, you can use it to make predictions about some images.
# + [markdown] colab_type="text" id="d4Ov9OFDMmOD"
# Let's look at the 0th image, predictions, and prediction array. Correct prediction labels are blue and incorrect prediction labels are red. The number gives the percentage (out of 100) for the predicted label.
# + colab={} colab_type="code" id="HV5jw-5HwSmO"
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], test_labels)
plt.show()
# + colab={} colab_type="code" id="Ko-uzOufSCSe"
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], test_labels)
plt.show()
# + [markdown] colab_type="text" id="kgdvGD52CaXR"
# Let's plot several images with their predictions. Note that the model can be wrong even when very confident.
# + colab={} colab_type="code" id="hQlnbqaw2Qu_"
# Plot the first X test images, their predicted labels, and the true labels.
# Color correct predictions in blue and incorrect predictions in red.
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions[i], test_labels)
plt.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="R32zteKHCaXT"
# ## Use the trained model
#
# Finally, use the trained model to make a prediction about a single image.
# + colab={} colab_type="code" id="yRJ7JU7JCaXT"
# Grab an image from the test dataset.
img = test_images[1]
print(img.shape)
# + [markdown] colab_type="text" id="vz3bVp21CaXV"
# `tf.keras` models are optimized to make predictions on a *batch*, or collection, of examples at once. Accordingly, even though you're using a single image, you need to add it to a list:
# + colab={} colab_type="code" id="lDFh5yF_CaXW"
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
# + [markdown] colab_type="text" id="EQ5wLTkcCaXY"
# Now predict the correct label for this image:
# + colab={} colab_type="code" id="o_rzNSdrCaXY"
predictions_single = probability_model.predict(img)
print(predictions_single)
# + colab={} colab_type="code" id="6Ai-cpLjO-3A"
plot_value_array(1, predictions_single[0], test_labels)
_ = plt.xticks(range(10), class_names, rotation=45)
# + [markdown] colab_type="text" id="cU1Y2OAMCaXb"
# `keras.Model.predict` returns a list of lists—one list for each image in the batch of data. Grab the predictions for our (only) image in the batch:
# + colab={} colab_type="code" id="2tRmdq_8CaXb"
np.argmax(predictions_single[0])
# + [markdown] colab_type="text" id="YFc2HbEVCaXd"
# And the model predicts a label as expected.
|
site/en/tutorials/keras/classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import json
# %pwd
data = pd.read_json('My_Request-1589942477185.json')
data.sample(5)
test.shape
test['comments'].value_counts()
|
test_api_result_json.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# ## Course materials
#
# - https://github.com/jakevdp/WhirlwindTourOfPython
# - https://github.com/jakevdp/PythonDataScienceHandbook
# - http://r4ds.had.co.nz/index.html
# - https://github.com/amueller/introduction_to_ml_with_python
# ## Introduction to Data Science
#
# - [What is Data Science?](./02-WhatIsDataScience.ipynb)
# - [Theory of Data](./03-TheoryofData.ipynb)
# - [Programming](../Programming/01-Introduction.ipynb)
# - [Workflow](../Workflow/01-Introduction.ipynb)
# ## Explore
#
# Explore = Visualize + Transform
#
# - [Data Visualisation](../Visualize/01-Introduction.ipynb)
# - [Basic Data Transformation](../Transform/02-BasicDataTransformation.ipynb)
# - [Exploratory Data Analysis (EDA)](../Explore/02-EDA.ipynb)
# ## Wrangle
#
# Wrangle = Import + Tidy + Transform
#
# - [Import](../Import/01-Introduction.ipynb)
# - [Tidy](../Tidy/01-Introduction.ipynb)
# - [Transform](../Transform/01-Introduction.ipynb)
# ## Model
#
# - [Model](../Model/01-Introduction.ipynb)
# ## Communicate
#
# - GitHub and Notebooks
# - nbconvert
# - nbviewer
|
Content/Introduction/01-Introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # User requirements Research
# ### Research Question
# 1.Which businesses are getting top/negative reviews?
#
# 2.Which categories of business are getting top/negative reviews?
#
# 3.What are the keywords/frequent of top/negative reviewed businesses from users?
#
# 4.What are the most frequent words from users reviews system?
#
# 4.What are the relationship between rating scores,price, review count?
# ### Purpose
# 1.Understand current small business in Portland.
#
# 2.Understand socail demand and expect for different categories of small business in different location.
#
# 3.Understand users requirements for small business quality.
#
# 4.Provide business owners a reference and comparison to choose location and predict their development plan.
# ## 1.Data Collecting
# ### 1.1 Download Yelp business review datasets using API
import requests
import json
import pandas as pd
import numpy as np
import matplotlib.pylab as plt
import fiona
import descartes
from shapely.geometry import Point, Polygon
import statsmodels.formula.api as smf
import matplotlib.pylab as plt
import math
import copy
# %matplotlib inline
#Porland Zipcode
zipcode=[97201, 97202,97203, 97204, 97205, 97206, 97209, 97210,97211, 97212, 97213, 97214, 97215, 97216,97217, 97218, 97219, 97220, 97221, 97222, 97223, 97225, 97227, 97229, 97230, 97231, 97232,
97233, 97236, 97239, 97258, 97266]
HEADERS={'Authorization':'Bearer %s' % api_key}
ENDPOINT='https://api.yelp.com/v3/businesses/search'
json_total = []
for i in zipcode:
PARAMETERS={'limit':50,
'radius':1000,
'offset':50,
'location':'Porland, OR, %d' % i}
json_total.append(requests.get(url=ENDPOINT,params=PARAMETERS,headers=HEADERS))
# +
# transfer Bussiness Review json to dataframe
df_total=[]
for j in range(len(json_total)):
data=json_total[j].json()
rating=[]
latitude=[]
longitude=[]
zip_code=[]
name=[]
review_count=[]
ID=[]
price=[]
categories=[]
for i in range(len(data['businesses'])):
ID+=[data['businesses'][i]['id']]
latitude+=[data['businesses'][i]['coordinates']['latitude']]
longitude+=[data['businesses'][i]['coordinates']['longitude']]
zip_code+=[data['businesses'][i]['location']['zip_code']]
rating+=[data['businesses'][i]['rating']]
name+=[data['businesses'][i]['name']]
review_count+=[data['businesses'][i]['review_count']]
categories+=[data['businesses'][i]['categories'][0]['title']]
df_total.append(pd.DataFrame({'Business_id':ID,
'categories':categories,
'rating':rating,
'latitude':latitude,
'longitude':longitude,
'zipcode':zip_code,
'review_count':review_count},index=name))
# -
# Bussiness Review information in Portland
business= pd.concat(df_total)
business.head()
business.to_csv('business.csv')
# ### 1.2 Good/Negative Reviewed small business
rating_good=business[business['rating']>4.0]
rating_good.head()
rating_bad=business[business['rating']<3.5]
rating_bad.head()
rating_bad.to_csv('rating_bad.csv')
rating_good.to_csv('rating_good.csv')
# ### 1.3 Using API to get user reviews datasets
reviews=[]
for i in business['Business_id']:
HEADERS={'Authorization':'Bearer %s' % api_key}
ENDPOINT='https://api.yelp.com/v3/businesses/%s/reviews' % i
reviews.append(requests.get(url=ENDPOINT,headers=HEADERS))
# +
R=[]
for j in range(len(reviews)):
data=reviews[j].json()
rating=[]
text=[]
time=[]
Review_ID=[]
for i in range(len(data['reviews'])):
Review_ID+=[data['reviews'][i]['id']]
text+=[data['reviews'][i]['text']]
time+=[data['reviews'][i]['time_created']]
rating+=[data['reviews'][i]['rating']]
R.append(pd.DataFrame({'Review_ID':Review_ID,
'text':text,
'rating':rating,
'time':time}))
for i in range(len(business)):
R[i]['Business_ID']=business['Business_id'][i]
Review_Portland= pd.concat(R)
Review_Portland.head()
# -
Review_Portland.to_csv('Review_Portland.csv')
# ### 1.4 Get Good/Bad Reviews Text
review_good=[]
for i in rating_good['Business_id']:
HEADERS={'Authorization':'Bearer %s' % api_key}
ENDPOINT='https://api.yelp.com/v3/businesses/%s/reviews' % i
review_good.append(requests.get(url=ENDPOINT,headers=HEADERS))
# +
Rgood=[]
for j in range(len(review_good)):
data=review_good[j].json()
rating=[]
text=[]
time=[]
Review_ID=[]
for i in range(len(data['reviews'])):
Review_ID+=[data['reviews'][i]['id']]
text+=[data['reviews'][i]['text']]
time+=[data['reviews'][i]['time_created']]
rating+=[data['reviews'][i]['rating']]
Rgood.append(pd.DataFrame({'Review_ID':Review_ID,
'text':text,
'rating':rating,
'time':time}))
for i in range(len(rating_good)):
Rgood[i]['Business_ID']=rating_good['Business_id'][i]
Review_good= pd.concat(Rgood)
Review_good.head()
# -
review_bad=[]
for i in rating_bad['Business_id']:
HEADERS={'Authorization':'Bearer %s' % api_key}
ENDPOINT='https://api.yelp.com/v3/businesses/%s/reviews' % i
review_bad.append(requests.get(url=ENDPOINT,headers=HEADERS))
# +
Rbad=[]
for j in range(len(review_bad)):
data=review_bad[j].json()
rating=[]
text=[]
time=[]
Review_ID=[]
for i in range(len(data['reviews'])):
Review_ID+=[data['reviews'][i]['id']]
text+=[data['reviews'][i]['text']]
time+=[data['reviews'][i]['time_created']]
rating+=[data['reviews'][i]['rating']]
Rbad.append(pd.DataFrame({'Review_ID':Review_ID,
'text':text,
'rating':rating,
'time':time}))
for i in range(len(rating_bad)):
Rbad[i]['Business_ID']=rating_bad['Business_id'][i]
Review_bad= pd.concat(Rbad)
Review_bad.head()
# -
Review_bad.to_csv('Review_bad.csv')
Review_good.to_csv('Review_good.csv')
Review_pos=Review_Portland[Review_Portland['rating']>4.5]
Review_neg=Review_Portland[Review_Portland['rating']<3.0]
Review_neg.head()
Review_pos.head()
# ## 2.Data Processing
import nltk
import string
import nltk.stem
from nltk.stem import PorterStemmer
from nltk.stem import WordNetLemmatizer
from nltk.corpus import stopwords
import nltk.stem
from PIL import Image
import sys
import re
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
nltk.download('punkt')
def preprocess_data(data):
# This function should return a list of lists of preprocessed tokens for each message
preprocessed_text=[]
for i in data:
words = nltk.word_tokenize(i)
words = [word.lower() for word in words if word.isalpha()] #clean punctuation
words = [word.lower() for word in words if re.match('^[a-zA-Z]+', word)]
filtered_words = [word for word in words if word not in stopwords.words('english')] # clean stopwords
s = nltk.stem.SnowballStemmer('english') #
cleaned_words = [s.stem(word) for word in words]
preprocessed_text.append(cleaned_words)
return preprocessed_text
# +
text = Review_Portland.loc[:,'text']
good_text = Review_good.loc[:,'text']
bad_text = Review_bad.loc[:,'text']
pos_text = Review_pos.loc[:,'text']
neg_text = Review_neg.loc[:,'text']
sen=preprocess_data(text)
good_sen = preprocess_data(good_text)
bad_sen= preprocess_data(bad_text)
pos_sen= preprocess_data(pos_text)
neg_sen= preprocess_data(neg_text)
# -
# ### 2.1 Get the most frequent words of all reviewed business in City of Portland
# +
tokens = []
for i in range(len(sen)):
token = " ".join(sen[i])
tokens.append(token)
tokens = ''.join(tokens)
# Create and generate a word cloud image:
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(tokens)
# Display the generated image:
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
# ### 2.2 Get the most frequent words of Good/Negative Reviewed business
# +
good_token = []
for i in range(len(good_sen)):
token = " ".join(good_sen[i])
good_token.append(token)
good_token=' '.join(good_token)
# Create and generate a word cloud image:
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(good_token)
# Display the generated image:
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# +
bad_token = []
for i in range(len(bad_sen)):
token = " ".join(bad_sen[i])
bad_token.append(token)
bad_token=' '.join(bad_token)
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(bad_token)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
# ### 2.3 Get the most frequent words of pos/neg reviewed rating business in City of Portland
# +
pos_token = []
for i in range(len(pos_sen)):
token = " ".join(pos_sen[i])
pos_token.append(token)
pos_token=' '.join(pos_token)
# Create and generate a word cloud image:
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(pos_token)
# Display the generated image:
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# +
neg_token = []
for i in range(len(neg_sen)):
token = " ".join(neg_sen[i])
neg_token.append(token)
neg_token=' '.join(neg_token)
# Create and generate a word cloud image:
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(neg_token)
# Display the generated image:
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
# ## 3.Data Modeling and Analysis
# ### 3.1 calculate the categories in positive and negative ratings in Portland.
# ### 3.2 sentiment analysis about user reviews
# ## 4.Data Analysis and Visualization
# ### 4.1 Map the small business in Portland with different review rating scores
# ### 4.2 Explore relationship between rating scores,price, review count
|
User requirements_Portland.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Resources
# - 链接: https://docs.python.org/3/library/pathlib.html
# - 版本: 3.7.0
# - 平台: macOS <small>(不测试部分 func)</small>
# ### 0x01
# - basic usage of *pathlib*
from pathlib import Path
# +
p = Path('..')
# trimmed (res too long)
lis_dir = [x for x in p.iterdir() if x.is_dir()]
# test other funcs
p.absolute() # '..' as path
p.resolve() # path after `cd ..`
p.is_file() # nope
p.is_dir() # yep
p.stat() # display file status (system call!)
# and ..
list(
p.glob('**/*.md')
)
# navigating
np = p / 'builtin_stuff'
np
np.resolve()
# open a file!
a_file = Path('/usr/share/dict/words')
with a_file.open(mode='r') as f:
f.readline()
# -
# ### 0x02
# - brief of *PurePath*
# - It provides path-handling opts which ***don't actually access*** a filesystem.
from pathlib import (
PurePath,
PurePosixPath, PureWindowsPath
)
# +
# null path
PurePath()
PurePath('.')
# multiple abspath
PurePath('/etc', '/usr', 'share') # only the last one (/)
# multiple abspath (for windows)
PureWindowsPath('C:/Windows', 'D:/Music') # u get the last one
PureWindowsPath('C:/Windows', '/Fonts') # the '/' is a bit different !
# +
# concat
PurePath('test.txt')
PurePath('/usr', 'share/dict', 'words')
PurePath('/usr')
PurePath('/usr') / 'share'
'/usr' / PurePath('share/words')
# +
# split into parts
nix_pt = PurePath('/usr/local/bin/python3')
win_pt = PureWindowsPath('C:/Program Files/python3')
nix_pt.parts
win_pt.parts
# +
# raw path
PurePath('/usr')
PurePath('/usr').__str__()
PureWindowsPath('C:/Windows').__bytes__()
PureWindowsPath('C:/Windows')
PureWindowsPath('C:/Windows').__str__()
PureWindowsPath('C:/Windows').__bytes__()
# +
# spurious '/' and '.'
PurePath('foo//bar') # // => no need (and wrongly use)
PurePath('foo/./bar') # . => current
PurePath('foo/../bar'); print()
# two subclasses (of 'PurePath')
PurePosixPath('/etc/')
PureWindowsPath('C:/Windows')
# +
# compariosn & flavor (win/*nix)
''' case-sensitive '''
PurePosixPath('foo') == PurePosixPath('FOO')
PureWindowsPath('foo') == PureWindowsPath('FOO')
''' Windows specific '''
PureWindowsPath('FOO') in { PureWindowsPath('foo') }
PureWindowsPath('C:') < PureWindowsPath('C:/Windows') # NOT `<' in math!!
PureWindowsPath('C:') < PureWindowsPath('E:') # "C->D->E" or "d: d:file"
''' posix VS windows '''
try:
PurePosixPath('foo') != PureWindowsPath('foo') # no error
PurePosixPath('foo') < PureWindowsPath('foo') # raised by this line
except TypeError as err:
err
# -
# ### 0x03
# - more about *PurePath*
# just a shortcut (not using it all the time)
pws = PureWindowsPath
pix = PurePosixPath
# drive & root-dir
# +
pws('C:/Windows').drive, pws('C:/Windows').root
pws('C:Windows').drive, pws('C:Windows').root
pix('/etc').drive, pix('/etc').root
pws('//host/share/foo.txt').drive, \
pws('//host/share').root
# -
# anchor (concat of 'drive' and 'root' )
# +
pws('C:/Windows').anchor
pws('C:Windows').anchor
pix('/etc').anchor
pws('//host/share').anchor
# -
# parents & parent
# +
pr = PureWindowsPath('C:/foo/bar/setup.py')
pr.parents[0] # == prt.parent (first-level-parent)
pr.parents[1]
pr.parents[2]
pix('/').parent # they've gotta no parent! XDD
pix('.').parent
pix('foo/..').parent # resolve it first
# -
# relative_to
# +
rlt = PurePosixPath('/etc/passwd')
rlt.relative_to('/')
rlt.relative_to('/etc')
try:
rlt.relative_to('/usr')
except ValueError as err:
err
# -
# name & with_name
# +
# name
pix('my/path').name
pix('my/path/.zshrc').name
pws('//some/share').name
pws('//some/share/test.txt').name
# with_name
pws('C:/Downloads/hello.py').with_name('python3-webinstall.exe')
pws('C:/Downloads').with_name('python3-webinstall.exe')
try:
pws('C:/').with_name('hello.py') # at least two parts
except ValueError as err:
err
# -
# suffix, suffixes & with_suffix
# +
# suffix & suffixes
pix('my/code/run.py').suffix # .py
pix('my/code/run.py').suffixes # [.py]
pix('my/pack/mdx.tar.gz').suffix # .gz
pix('my/pack/mdx.tar.gz').suffixes # [.tar, .gz]
pix('my/pack').suffix
pix('my/pack').suffixes
# with_suffix
pix('/lib/README').with_suffix('.md')
pws('C:/Downloads/hello.py').with_suffix('.ipynb')
pix('C:/Downloads/pack.tar.gz').with_suffix('.bz2')
# -
# stem
pix('my/library.tar.gz').stem # library.tar
pix('my/library.tar').stem # library
pix('my/library').stem # library
# as_posix & as_uri
# +
# '\\' to '/'
PureWindowsPath('C:\\Windows').__str__()
PureWindowsPath('C:\\Windows').as_posix()
# rep the path as 'file' URI
PurePosixPath('/etc/passwd').as_uri() # the path must be absolute
PureWindowsPath('C:\\Windows').as_uri() # if not, raise ValueError
# -
# is_absolute & is_reserved
# +
# abs - nix
pix('/a/b').is_absolute(), \
pix('a/b').is_absolute()
# abs - win
pws('C:/a/b').is_absolute(), \
pws('/a/b').is_absolute(), \
pws('C:').is_absolute(), \
pws('//some/share').is_absolute()
# is reserved (or not)
pws('nul').is_reserved(), \
pix('nul').is_reserved() # always `False` for *nix
# -
# joinpath & match
# +
# joinpath
pix('/usr').joinpath('share', 'dict')
pws('C:\\').joinpath('Windows', 'Fonts')
pix('/usr').joinpath(pix('local', 'bin'))
# match
PurePath('a/b.py').match('*.py'), \
PurePath('/a/b/c.py').match('b/*.py'), \
PurePath('/a/b/c.py').match('a/*.py')
PurePath('/a.py').match( '/*.py'), \
PurePath('a/b.py').match('/*.py')
# match - case
PureWindowsPath('b.py').match('*.PY'), \
PurePosixPath( 'b.py').match('*.PY')
# -
# ### 0x04
# - Let's talk about <q>*concrete path*</q>!
# - It's subclasses of the pure path classes.
# - It provides methods to ***do system calls on path objects***.
from pathlib import (
Path,
PosixPath, WindowsPath
)
# create (init) a path object
# +
Path('setup.py')
try:
# u can only init the class corresponds to your system
PosixPath('/usr/local')
WindowsPath('C:/Windows/Fonts')
except NotImplementedError as err:
print('Well...', err)
# -
# cwd, home, stat, lstat
# +
Path.cwd()
Path.home()
f = Path('pathlib_offidoc.ipynb') # file -- stat
fl = Path('/Users/alex/vid') # symlink -- lstat
f
f.stat().st_size // 1024
fl
fl.lstat().st_size # show the symlink's info rather than its target's
# -
# exists, expanduser
# +
Path('..').exists()
Path('/etc').exists()
Path('/Users/alex/vid/').exists() # symlink is supported :)
PosixPath('~/vid')
PosixPath('~/vid').expanduser()
# -
# glob, rglob, owner, group
# +
# glob
sorted(
Path('../__HOWTOs__').glob('*.ipynb')
)
sorted(
Path('..').glob('**/*.md') # crt dir & all subdirs
)
# rglob
sorted(
Path('..').rglob('*.md') # just like 'glob' but add '**'
)
# -
# owner, group
Path('.').owner() # -> alex
Path('.').group() # -> staff
# is_dir, is_file, is_symlink
# +
# is_dir
Path('/Users/alex/vid/').is_dir()
Path('../file_and_directory_access/').is_dir()
# is_file
Path('pathlib_offidoc.ipynb').is_file()
# is_mount
# Path('/Users/alex/vid').is_mount() # Python 3.7 only
# is_symlink
Path('/User/alex/vid').is_symlink() # hmm..
# -
# iterdir, open
# +
# iterdir
list(
Path('/Users/alex/Movies').iterdir()
)
# open
with Path('/etc/hosts').open('r') as f:
f.readline()
# -
# read_bytes, read_text
# +
pb = Path('src/my_bin_file')
pt = Path('src/my_txt_file')
pb.write_bytes(b'hello byte') # automatically closed (hell yeah)
pb.read_bytes()
pt.write_text('hey guys!')
pt.read_text(encoding='utf-8') # same (encode) meaning as `open()`
# -
# rename, replace
# +
before = Path('src/file_to_be_renamed.txt')
target = Path('src/just_normal_file.txt')
before.open('w').write('aha gotta ya!')
before
before.rename(
target # => just_normal_file.txt
)
target.open().read()
# then `replace`
tip = '''
the same as shell's command: `mv`
if another-file => rename
if it-is-a-dir => move 'this' file to the dir
'''
# -
# resolve
# +
Path('..')
Path('..').resolve()
Path('../__HOWTOs__/').resolve(strict=True) # no idea about `strict`
# -
# symlink_to
# +
link = Path('src/link_to_home')
try:
link.symlink_to('/Users/')
except FileExistsError:
link
link.resolve()
# -
# touch, unlink
# +
a = Path('src/inst_file')
a
a.touch()
a.touch(mode=0o666, exist_ok=True)
a.unlink() # delete the file (or symlink)
|
file_and_directory_access/pathlib_offidoc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Relevancy Analysis
# <div class="alert alert-info">
#
# This tutorial is available as an IPython notebook at [Malaya/example/relevancy](https://github.com/huseinzol05/Malaya/tree/master/example/relevancy).
#
# </div>
# <div class="alert alert-warning">
#
# This module only trained on standard language structure, so it is not save to use it for local language structure.
#
# </div>
# %%time
import malaya
# ### Explanation
#
# Positive relevancy: The article or piece of text is relevant, tendency is high to become not a fake news. Can be a positive or negative sentiment.
#
# Negative relevancy: The article or piece of text is not relevant, tendency is high to become a fake news. Can be a positive or negative sentiment.
#
# **Right now relevancy module only support deep learning model**.
negative_text = 'Roti Massimo Mengandungi DNA Babi. Roti produk Massimo keluaran Syarikat The Italian Baker mengandungi DNA babi. Para pengguna dinasihatkan supaya tidak memakan produk massimo. Terdapat pelbagai produk roti keluaran syarikat lain yang boleh dimakan dan halal. Mari kita sebarkan berita ini supaya semua rakyat Malaysia sedar dengan apa yang mereka makna setiap hari. Roti tidak halal ada DNA babi jangan makan ok.'
positive_text = 'Jabatan Kemajuan Islam Malaysia memperjelaskan dakwaan sebuah mesej yang dikitar semula, yang mendakwa kononnya kod E dikaitkan dengan kandungan lemak babi sepertimana yang tular di media sosial. . Tular: November 2017 . Tular: Mei 2014 JAKIM ingin memaklumkan kepada masyarakat berhubung maklumat yang telah disebarkan secara meluas khasnya melalui media sosial berhubung kod E yang dikaitkan mempunyai lemak babi. Untuk makluman, KOD E ialah kod untuk bahan tambah (aditif) dan ianya selalu digunakan pada label makanan di negara Kesatuan Eropah. Menurut JAKIM, tidak semua nombor E yang digunakan untuk membuat sesuatu produk makanan berasaskan dari sumber yang haram. Sehubungan itu, sekiranya sesuatu produk merupakan produk tempatan dan mendapat sijil Pengesahan Halal Malaysia, maka ia boleh digunakan tanpa was-was sekalipun mempunyai kod E-kod. Tetapi sekiranya produk tersebut bukan produk tempatan serta tidak mendapat sijil pengesahan halal Malaysia walaupun menggunakan e-kod yang sama, pengguna dinasihatkan agar berhati-hati dalam memilih produk tersebut.'
# ### List available Transformer models
malaya.relevancy.available_transformer()
# Make sure you can check accuracy chart from here first before select a model, https://malaya.readthedocs.io/en/latest/Accuracy.html#relevancy
#
# **You might want to use Alxlnet, a very small size, 46.8MB, but the accuracy is still on the top notch.**
# ### Load Transformer model
#
# ```python
# def transformer(model: str = 'xlnet', quantized: bool = False, **kwargs):
# """
# Load Transformer relevancy model.
#
# Parameters
# ----------
# model : str, optional (default='bert')
# Model architecture supported. Allowed values:
#
# * ``'bert'`` - Google BERT BASE parameters.
# * ``'tiny-bert'`` - Google BERT TINY parameters.
# * ``'albert'`` - Google ALBERT BASE parameters.
# * ``'tiny-albert'`` - Google ALBERT TINY parameters.
# * ``'xlnet'`` - Google XLNET BASE parameters.
# * ``'alxlnet'`` - Malaya ALXLNET BASE parameters.
# * ``'bigbird'`` - Google BigBird BASE parameters.
# * ``'tiny-bigbird'`` - Malaya BigBird BASE parameters.
# * ``'fastformer'`` - FastFormer BASE parameters.
# * ``'tiny-fastformer'`` - FastFormer TINY parameters.
#
# quantized : bool, optional (default=False)
# if True, will load 8-bit quantized model.
# Quantized model not necessary faster, totally depends on the machine.
#
# Returns
# -------
# result: model
# List of model classes:
#
# * if `bert` in model, will return `malaya.model.bert.MulticlassBERT`.
# * if `xlnet` in model, will return `malaya.model.xlnet.MulticlassXLNET`.
# * if `bigbird` in model, will return `malaya.model.xlnet.MulticlassBigBird`.
# * if `fastformer` in model, will return `malaya.model.fastformer.MulticlassFastFormer`.
# """
# ```
model = malaya.relevancy.transformer(model = 'tiny-bigbird')
# ### Load Quantized model
#
# To load 8-bit quantized model, simply pass `quantized = True`, default is `False`.
#
# We can expect slightly accuracy drop from quantized model, and not necessary faster than normal 32-bit float model, totally depends on machine.
quantized_model = malaya.relevancy.transformer(model = 'alxlnet', quantized = True)
# #### Predict batch of strings
#
# ```python
# def predict(self, strings: List[str]):
# """
# classify list of strings.
#
# Parameters
# ----------
# strings: List[str]
#
# Returns
# -------
# result: List[str]
# """
# ```
# +
# %%time
model.predict([negative_text, positive_text])
# +
# %%time
quantized_model.predict([negative_text, positive_text])
# -
# #### Predict batch of strings with probability
#
# ```python
# def predict_proba(self, strings: List[str]):
# """
# classify list of strings and return probability.
#
# Parameters
# ----------
# strings : List[str]
#
# Returns
# -------
# result: List[dict[str, float]]
# """
# ```
# +
# %%time
model.predict_proba([negative_text, positive_text])
# +
# %%time
quantized_model.predict_proba([negative_text, positive_text])
# -
# #### Open relevancy visualization dashboard
#
# ```python
# def predict_words(
# self, string: str, method: str = 'last', visualization: bool = True
# ):
# """
# classify words.
#
# Parameters
# ----------
# string : str
# method : str, optional (default='last')
# Attention layer supported. Allowed values:
#
# * ``'last'`` - attention from last layer.
# * ``'first'`` - attention from first layer.
# * ``'mean'`` - average attentions from all layers.
# visualization: bool, optional (default=True)
# If True, it will open the visualization dashboard.
#
# Returns
# -------
# result: dict
# """
# ```
#
# Default when you call `predict_words` it will open a browser with visualization dashboard, you can disable by `visualization=False`.
#
# **This method not available for BigBird models**.
model.predict_words(negative_text)
quantized_model.predict_words(negative_text)
# +
from IPython.core.display import Image, display
display(Image('relevancy-dashboard.png', width=800))
# -
# ### Vectorize
#
# Let say you want to visualize sentence / word level in lower dimension, you can use `model.vectorize`,
#
# ```python
# def vectorize(self, strings: List[str], method: str = 'first'):
# """
# vectorize list of strings.
#
# Parameters
# ----------
# strings: List[str]
# method : str, optional (default='first')
# Vectorization layer supported. Allowed values:
#
# * ``'last'`` - vector from last sequence.
# * ``'first'`` - vector from first sequence.
# * ``'mean'`` - average vectors from all sequences.
# * ``'word'`` - average vectors based on tokens.
#
# Returns
# -------
# result: np.array
# """
# ```
# #### Sentence level
texts = [negative_text, positive_text]
r = model.vectorize(texts, method = 'first')
# +
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
tsne = TSNE().fit_transform(r)
tsne.shape
# -
plt.figure(figsize = (7, 7))
plt.scatter(tsne[:, 0], tsne[:, 1])
labels = texts
for label, x, y in zip(
labels, tsne[:, 0], tsne[:, 1]
):
label = (
'%s, %.3f' % (label[0], label[1])
if isinstance(label, list)
else label
)
plt.annotate(
label,
xy = (x, y),
xytext = (0, 0),
textcoords = 'offset points',
)
# #### Word level
r = quantized_model.vectorize(texts, method = 'word')
x, y = [], []
for row in r:
x.extend([i[0] for i in row])
y.extend([i[1] for i in row])
tsne = TSNE().fit_transform(y)
tsne.shape
plt.figure(figsize = (7, 7))
plt.scatter(tsne[:, 0], tsne[:, 1])
labels = x
for label, x, y in zip(
labels, tsne[:, 0], tsne[:, 1]
):
label = (
'%s, %.3f' % (label[0], label[1])
if isinstance(label, list)
else label
)
plt.annotate(
label,
xy = (x, y),
xytext = (0, 0),
textcoords = 'offset points',
)
# Pretty good, the model able to know cluster bottom left as positive relevancy.
# ### Stacking models
#
# More information, you can read at [https://malaya.readthedocs.io/en/latest/Stack.html](https://malaya.readthedocs.io/en/latest/Stack.html)
albert = malaya.relevancy.transformer(model = 'albert')
malaya.stack.predict_stack([albert, model], [positive_text, negative_text])
|
docs/load-relevancy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
config = tf.compat.v1.ConfigProto(
gpu_options = tf.compat.v1.GPUOptions(per_process_gpu_memory_fraction=0.8),
)
config.gpu_options.allow_growth = True
session = tf.compat.v1.Session(config=config)
tf.compat.v1.keras.backend.set_session(session)
# +
import random
import matplotlib.pyplot as plt
import numpy as np
import numpy.ma as ma
import pandas as pd
import sklearn
import tensorflow as tf
from amp.utils.basic_model_serializer import load_master_model_components
from amp.utils import basic_model_serializer
from amp.inference.filtering import amino_based_filtering
import amp.data_utils.data_loader as data_loader
from amp.data_utils.sequence import pad, to_one_hot
from amp.utils import phys_chem_propterties as phys
from keras import backend, layers
from keras.optimizers import Adam
from sklearn.model_selection import train_test_split
from keras import layers
from pathlib import Path
from keras import models as m
from tqdm import tqdm
from joblib import dump, load
from sklearn.decomposition import PCA
import os
import scipy
import modlamp.descriptors
import modlamp.analysis
import modlamp.sequences
seed = 7
MIN_LENGTH = 0
MAX_LENGTH = 25
latent_dim = 64
input_to_encoder = layers.Input(shape=(MAX_LENGTH,))
input_to_decoder = layers.Input(shape=(latent_dim+2,))
# +
def translate_generated_peptide(encoded_peptide):
alphabet = list('ACDEFGHIKLMNPQRSTVWY')
return ''.join([alphabet[el - 1] if el != 0 else "" for el in encoded_peptide.argmax(axis=1)])
def translate_peptide(encoded_peptide):
alphabet = list('ACDEFGHIKLMNPQRSTVWY')
return ''.join([alphabet[el-1] if el != 0 else "" for el in encoded_peptide])
# +
from joblib import dump, load
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
import numpy as np
from scipy.stats import wilcoxon, mannwhitneyu
sns.set_style('whitegrid', {'grid.color': '.95', 'axes.spines.right': False, 'axes.spines.top': False})
params = {'axes.labelsize': 8,'axes.titlesize':8, 'font.size': 8, 'legend.fontsize': 6, 'xtick.labelsize': 8, 'ytick.labelsize': 8}
plt.rcParams.update(params)
# +
models = [
'HydrAMP',
'PepCVAE',
'Basic',
]
model_labels = [
'HydrAMP',
'PepCVAE',
'Basic'
]
# -
hydra_color = '#B80018'
pepcvae_color = '#1d3557'
basic_color = '#B4C5E4'
# +
def calculate_length(data:list):
lengths = [len(x) for x in data]
return lengths
def calculate_molarweight(x:list):
h = modlamp.descriptors.GlobalDescriptor(data)
h.calculate_MW()
return list(h.descriptor.flatten())
def calculate_charge(data:list):
h = modlamp.analysis.GlobalAnalysis(data)
h.calc_charge()
return h.charge
def calculate_isoelectricpoint(data:list):
h = modlamp.analysis.GlobalDescriptor(data)
h.isoelectric_point()
return list(h.descriptor.flatten())
def calculate_aromaticity(data:list):
h = modlamp.analysis.GlobalDescriptor(data)
h.aromaticity()
return list(h.descriptor.flatten())
def calculate_hydrophobicity(data:list):
h = modlamp.analysis.GlobalAnalysis(data)
h.calc_H(scale='eisenberg')
return list(h.H)
def calculate_hydrophobicmoment(data:list):
h = modlamp.descriptors.PeptideDescriptor(data, 'eisenberg')
h.calculate_moment()
return list(h.descriptor.flatten())
def calculate_alphahelixpropensity(data:list):
h = modlamp.descriptors.PeptideDescriptor(data, 'levitt_alpha')
h.calculate_global()
return list(h.descriptor.flatten())
def calculate_instability_index(data:list):
h = modlamp.analysis.GlobalDescriptor(data)
h.instability_index()
return list(h.descriptor.flatten())
def calculate_hscore(data:list):
return [phys.helical_search(x) for x in data]
def calculate_hydrophobic_ratio(data:list):
h = modlamp.analysis.GlobalDescriptor(data)
h.hydrophobic_ratio()
return list(h.descriptor.flatten())
# return [phys.helical_search(x) for x in data]
def calculate_boman_index(data:list):
h = modlamp.analysis.GlobalDescriptor(data)
h.boman_index()
return list(h.descriptor.flatten())
# -
def select_peptides(results, mode):
if mode=='pos':
peptides = np.array(results[f'pos_peptides']).reshape(64, -1).T
amp = (results['pos_class_prediction'] < 0.8).reshape(64, -1)
mic = results['pos_mic_prediction'].reshape(64, -1)
combined = ma.masked_where(amp, mic)
good = combined.argmax(axis=0)
good_peptides = peptides[list(range(peptides.shape[0])), good]
good_amp = np.array(results['pos_class_prediction']).reshape(64, -1).T[list(range(peptides.shape[0])), good]
good_mic = np.array(results['pos_mic_prediction']).reshape(64, -1).T[list(range(peptides.shape[0])), good]
return pd.DataFrame.from_dict({
'sequence': good_peptides.tolist(),
'amp': good_amp.tolist(),
'mic': good_mic.tolist(),
}
)
else:
peptides = np.array(results['neg_peptides']).reshape(64, -1).T
amp = (results['neg_class_prediction'] > 0.2).reshape(64, -1)
mic = results['neg_mic_prediction'].reshape(64, -1)
combined = ma.masked_where(amp, mic)
good = combined.argmin(axis=0)
good_peptides = peptides[list(range(peptides.shape[0])), good]
good_amp = np.array(results['neg_class_prediction']).reshape(64, -1).T[list(range(peptides.shape[0])), good]
good_mic = np.array(results['neg_mic_prediction']).reshape(64, -1).T[list(range(peptides.shape[0])), good]
return pd.DataFrame.from_dict({
'sequence': good_peptides.tolist(),
'amp': good_amp.tolist(),
'mic': good_mic.tolist(),
}
)
# # Unconstrained
# +
random.seed(seed)
data_manager = data_loader.AMPDataManager(
'../data/unlabelled_positive.csv',
'../data/unlabelled_negative.csv',
min_len=MIN_LENGTH,
max_len=MAX_LENGTH)
amp_x, amp_y = data_manager.get_merged_data()
amp_x_train, amp_x_test, amp_y_train, amp_y_test = train_test_split(amp_x, amp_y, test_size=0.1, random_state=36)
amp_x_train, amp_x_val, amp_y_train, amp_y_val = train_test_split(amp_x_train, amp_y_train, test_size=0.2, random_state=36)
# Restrict the length
ecoli_df = pd.read_csv('../data/mic_data.csv')
mask = (ecoli_df['sequence'].str.len() <= MAX_LENGTH) & (ecoli_df['sequence'].str.len() >= MIN_LENGTH)
ecoli_df = ecoli_df.loc[mask]
mic_x = pad(to_one_hot(ecoli_df['sequence']))
mic_y = ecoli_df.value
mic_x_train, mic_x_test, mic_y_train, mic_y_test = train_test_split(mic_x, mic_y, test_size=0.1, random_state=36)
mic_x_train, mic_x_val, mic_y_train, mic_y_val = train_test_split(mic_x_train, mic_y_train, test_size=0.2, random_state=36)
pos = np.vstack([amp_x_test[amp_y_test == 1], mic_x_test[mic_y_test < 1.5]])
neg = np.vstack([amp_x_test[amp_y_test == 0], mic_x_test[mic_y_test > 1.5]])
positives = [translate_peptide(x) for x in pos]
negatives = [translate_peptide(x) for x in neg]
#Load Uniprot
uniprot = list(pd.read_csv('../data/Uniprot_0_25_train.csv').Sequence)
uniprot = random.sample(uniprot, 50000)
#Get random peptides
random_gen = modlamp.sequences.Random(50000, 1, 25)
random_gen.generate_sequences(proba='random')
random_peptides = random_gen.sequences
# -
hydra_results = load(f'../results/unconstrained_{models[0]}.joblib')
pepcvae_results = load(f'../results/unconstrained_{models[1]}.joblib')
basic_results = load(f'../results/unconstrained_{models[2]}.joblib')
# +
hydra_positives = select_peptides(hydra_results, 'pos').sequence.tolist()
hydra_negatives = select_peptides(hydra_results, 'neg').sequence.tolist()
pepcvae_positives = select_peptides(pepcvae_results, 'pos').sequence.tolist()
pepcvae_negatives = select_peptides(pepcvae_results, 'neg').sequence.tolist()
basic_positives = select_peptides(basic_results, 'pos').sequence.tolist()
basic_negatives = select_peptides(basic_results, 'neg').sequence.tolist()
# -
len(hydra_positives)
def calculate_physchem(peptides, datasets, n):
physchem = {}
physchem['dataset'] = []
physchem['length'] = []
physchem['charge'] = []
physchem['pi'] = []
physchem['aromacity'] = []
physchem['hydrophobicity'] = []
physchem['hm'] = []
physchem['alpha'] = []
physchem['boman'] = []
physchem['h_score'] = []
physchem['hydrophobic_ratio'] = []
physchem['instability'] = []
for dataset, name in zip(peptides, datasets):
physchem['dataset'] += (len(dataset) * [name])
physchem['length'] += calculate_length(dataset)
physchem['charge'] += calculate_charge(dataset)[0].tolist()
physchem['pi'] += calculate_isoelectricpoint(dataset)
physchem['aromacity'] += calculate_aromaticity(dataset)
physchem['hydrophobicity'] += calculate_hydrophobicity(dataset)[0].tolist()
physchem['hm'] += calculate_hydrophobicmoment(dataset)
physchem['alpha'] += calculate_alphahelixpropensity(dataset)
physchem['boman'] += calculate_boman_index(dataset)
physchem['hydrophobic_ratio'] += calculate_hydrophobic_ratio(dataset)
physchem['h_score'] += calculate_hscore(dataset)
physchem['instability'] += calculate_instability_index(dataset)
return pd.DataFrame(dict([ (k, pd.Series(v)) for k,v in physchem.items() ]))
# +
datasets = [
'Random',
'Uniprot',
'Non-AMP test data',
'Non-AMP HydrAMP',
'Non-AMP PepCVAE',
'Non-AMP Basic',
'AMP test data',
'AMP HydrAMP',
'AMP PepCVAE',
'AMP Basic',
]
peptides = [
random_peptides,
uniprot,
negatives,
hydra_negatives,
pepcvae_negatives,
basic_negatives,
positives,
hydra_positives,
pepcvae_positives,
basic_positives,
]
# -
physchem = calculate_physchem(peptides, datasets, 10000)
physchem
boxprops = dict(linewidth=0.0, color='k')
flierprops = dict(linewidth=0.5)
medianprops = dict(linewidth=0.5, color='k')
whiskerprops = dict(linewidth=0.5)
capprops = dict(linewidth=0.5)
datasets = [
['Random','Uniprot'],
['Non-AMP test data', 'AMP test data'],
['Non-AMP HydrAMP', 'AMP HydrAMP'],
['Non-AMP PepCVAE', 'AMP PepCVAE'],
['Non-AMP Basic', 'AMP Basic']
]
def wilcox(wilx_prop):
if wilx_prop > 0.05:
symbol = 'ns'
if wilx_prop <= 0.05:
symbol = '*'
if wilx_prop <= 0.01:
symbol = '**'
if wilx_prop <= 0.001:
symbol = '***'
return symbol
# +
fig, master_axes = plt.subplots(
ncols=11,
nrows=2,
figsize=(6, 5),
dpi=300,
# gridspec_kw={'width_ratios': [1, 1, 1, 1, 1, 0]}
)
palette = [
'grey',
'lightgray',
'yellow',
'violet',
'yellow',
'violet',
'yellow',
'violet',
'yellow',
'violet',
]
for prop, label, axes, in zip(
['pi', 'charge', 'hydrophobic_ratio', 'aromacity', ],
['Isoelectric point', 'Charge', 'Hydrophobic moment', 'Aromaticity',],
[master_axes[0][:5], master_axes[0][6:], master_axes[1][:5], master_axes[1][6:]] ,
):
for ind, (ax, dataset) in enumerate(zip(axes, datasets)):
data = [physchem[physchem['dataset'] == x][prop].tolist() for x in dataset]
parts = ax.boxplot(
data,
showfliers=False,
patch_artist=True,
boxprops=boxprops,
flierprops=flierprops,
medianprops=medianprops,
whiskerprops=whiskerprops,
capprops=capprops,
widths=0.4
)
if dataset == ['Random','Uniprot']:
for patch, color in zip(parts['boxes'], ['grey','lightgray']):
patch.set_facecolor(color)
else:
for patch, color in zip(parts['boxes'], ['#66BDBA', '#F7CF8B',]):
patch.set_facecolor(color)
# ax.set_ylim(axes[0].get_ylim()[0], axes[0].get_ylim()[1])
ax.spines['left'].set_visible(False)
ax.set_yticklabels([])
# ax.set_xticks([])
if dataset == ['Random', 'Uniprot']:
continue
if dataset == ['Non-AMP test data', 'AMP test data']:
wilx_prop = mannwhitneyu(
physchem[physchem.dataset == dataset[1]][prop].tolist(),
physchem[physchem.dataset == dataset[0]][prop],
alternative='greater'
)[1]
else:
wilx_prop = wilcoxon(
physchem[physchem.dataset == dataset[1]][prop].tolist(),
physchem[physchem.dataset == dataset[0]][prop],
alternative='greater'
)[1]
# print(prop, dataset, wilx_prop, symbol)
symbol = wilcox(wilx_prop)
ax.text(
x=2,
y=1.03 * parts['caps'][3].get_ydata()[1],
s=symbol,
ha='center'
)
mins = [ax.get_ylim()[0] for ax in axes]
maxs = [ax.get_ylim()[1] for ax in axes]
for ax in axes:
ax.set_ylim(min(mins), max(maxs))
axes[0].set_xticks(range(2))
axes[0].set_xticklabels([], rotation=45)
axes[1].set_xticks([1])
axes[1].set_xticklabels([], rotation=45)
axes[2].set_xticks([1])
axes[2].set_xticklabels([], rotation=45)
axes[3].set_xticks([1])
axes[3].set_xticklabels([], rotation=45)
axes[4].set_xticks([1])
axes[4].set_xticklabels([], rotation=45)
axes[0].set_ylabel(label)
fig.delaxes(master_axes[0][5])
fig.delaxes(master_axes[1][5])
for i, (label, ax) in enumerate(
zip(
['a', 'b', 'c', 'd'],
[master_axes[0][0], master_axes[0][6], master_axes[1][0], master_axes[1][6]]
)
):
ax.annotate(label, xy=(-0.05, 1.1), xycoords='axes fraction',
fontweight='bold', va='top', ha='right')
master_axes[1][0].set_xticks(range(2))
master_axes[1][0].set_xticklabels(['Random','Uniprot'], rotation=45)
master_axes[1][1].set_xticks([1])
master_axes[1][1].set_xticklabels(['Test data'], rotation=45)
master_axes[1][2].set_xticks([1])
master_axes[1][2].set_xticklabels(['HydrAMP'], rotation=45)
master_axes[1][3].set_xticks([1])
master_axes[1][3].set_xticklabels(['PepCVAE'], rotation=45)
master_axes[1][4].set_xticks([1])
master_axes[1][4].set_xticklabels(['Basic'], rotation=45)
master_axes[1][6].set_xticks(range(2))
master_axes[1][6].set_xticklabels(['Random','Uniprot'], rotation=45)
master_axes[1][7].set_xticks([1])
master_axes[1][7].set_xticklabels(['Test data'], rotation=45)
master_axes[1][8].set_xticks([1])
master_axes[1][8].set_xticklabels(['HydrAMP'], rotation=45)
master_axes[1][9].set_xticks([1])
master_axes[1][9].set_xticklabels(['PepCVAE'], rotation=45)
master_axes[1][10].set_xticks([1])
master_axes[1][10].set_xticklabels(['Basic'], rotation=45)
plt.legend(
handles=[
# mpatches.Patch(color=palette[0], label='Random'),
# mpatches.Patch(color=palette[1], label='Uniprot'),
mpatches.Patch(color='#F7CF8B', label='Positive'),
mpatches.Patch(color='#66BDBA', label='Negative'),
],
bbox_to_anchor = (-3.75, -0.5),
ncol=4,
)
# fig.tight_layout()
fig.savefig("../figures/Fig4-Wilcox.svg")
fig.savefig("../figures/Fig4-Wilcox.pdf", bbox_inches="tight")
plt.show()
# -
|
scripts/supp_fig1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Example. Study it carefully. A question from *The Accumulator Pattern with Conditions*
# For each word in words, add ‘d’ to the end of the word if the word ends in “e” to make it past tense. Otherwise, add ‘ed’ to make it past tense. Save these past tense words to a list called past_tense.
words = ["adopt", "bake", "beam", "confide", "grill", "plant", "time", "wave", "wish"]
# +
# Answer (Write your code here):
past_tense=[]
for word in words:
if word[-1]!="e":
past_tense.append(word+"ed")
else:
past_tense.append(word+"d")
print(past_tense)
# -
# ### Question 1
# *rainfall_mi* is a string that contains the average number of inches of rainfall in Michigan for every month (in inches) with every month separated by a comma. Write code to compute the number of months that have more than 3 inches of rainfall. Store the result in the variable *num_rainy_months*. In other words, count the number of items with values *> 3.0*.
#
# Hard-coded answers will receive no credit.
rainfall_mi = "1.65, 1.46, 2.05, 3.03, 3.35, 3.46, 2.83, 3.23, 3.5, 2.52, 2.8, 1.85"
# +
rainfall_mi = "1.65, 1.46, 2.05, 3.03, 3.35, 3.46, 2.83, 3.23, 3.5, 2.52, 2.8, 1.85"
rainfall_mi_split = rainfall_mi.split(",")
num_rainy_months = 0
for x in rainfall_mi_split:
x = float(x)
if x > 3.0:
num_rainy_months += 1
print("Number of months that have more than 3 inches of rainfall:-",num_rainy_months)
# -
# ### Question 2
# The variable *sentence* stores a string. Write code to determine how many words in *sentence* start and end with the same letter, including one-letter words. Store the result in the variable *same_letter_count*.
#
# Hard-coded answers will receive no credit.
sentence = "students flock to the arb for a variety of outdoor activities such as jogging and picnicking"
sentence = "students flock to the arb for a variety of outdoor activities such as jogging and picnicking"
same_letter_count = 0
sentence_split = sentence.split(' ')
for i in sentence_split:#Index Reference
if i[0] == i[-1]:
same_letter_count += 1
print(same_letter_count)
# +
### Question 3
Write code to count the number of strings in list *items* that have the character w in it. Assign that number to the variable acc_num.
HINT 1: Use the accumulation pattern!
HINT 2: the _in_ operator checks whether a substring is present in a string.
Hard-coded answers will receive no credit.
# -
items = ["whirring", "wow!", "calendar", "wry", "glass", "", "llama","tumultuous","owing"]
items = ["whirring", "wow!", "calendar", "wry", "glass", "", "llama","tumultuous","owing"]
acc_num=0
for i in items:
if "w" in i:
acc_num +=1
print(acc_num)
# ### Question 4
# Write code that counts the number of words in sentence that contain either an “a” or an “e”. Store the result in the variable num_a_or_e.
#
# Note 1: be sure to not double-count words that contain both an a and an e.
#
# HINT 1: Use the in operator.
#
# HINT 2: You can either use or or elif.
#
# Hard-coded answers will receive no credit.
sentence = "python is a high level general purpose programming language that can be applied to many different classes of problems."
sentence = "python is a high level general purpose programming language that can be applied to many different classes of problems."
num_a_or_e = 0
for i in sentence.split():
if ('a' in i) or ('e' in i):
num_a_or_e += 1
print(num_a_or_e)
# ### Question 5
# Write code that will count the number of vowels in the sentence s and assign the result to the variable num_vowels. For this problem, vowels are only a, e, i, o, and u. Hint: use the in operator with vowels.
s = "singing in the rain and playing in the rain are two entirely different situations but both can be fun"
vowels = ['a','e','i','o','u']
s = "singing in the rain and playing in the rain are two entirely different situations but both can be fun"
vowels = ['a','e','i','o','u']
num_vowels = 0
for vowels in s.split():
num_vowels += 1
print(num_vowels)
|
Jupyter Assignment 02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Nunesdejesus/Cursoemvideo-html5/blob/main/Untitled26.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="e-oaBFrAkYuZ" outputId="ceda2ffc-e00b-4e12-c757-3b41f44da895"
# %%writefile carros.csv
id,valor_venda,valor_manutencao,portas,pessoas,porta_malas
1,vhigh,med,2,2,small
2,med,vhigh,2,2,small
3,low,vhigh,2,2,small
4,low,high,2,2,small
5,low,high,2,2,small
6,low,high,4,4,big
7,low,high,4,4,big
8,low,med,2,2,small
9,low,med,2,2,small
10,low,med,2,2,small
11,low,med,4,4,big
12,low,low,2,2,small
13,low,low,4,4,small
14,low,low,4,4,med
# + colab={"base_uri": "https://localhost:8080/"} id="j6kzgXzR-BJQ" outputId="9e669974-2e9e-488b-b7b2-2754f954e135"
# %%writefile musica.txt
Roda viva
<NAME>
Tem dias que a gente se sente
Como quem partiu ou morreu
A gente etancou de repente
Ou foi o mundo então que cresceu
A gente quer ter voz ativa
No nosso destino mandar
Mas eis que chega a roda viva
E carrega o destino pra lá
Roda mundo, roda-gigante
Roda moinho, roda,pião
O tempo rodou num instante
Nas voltas do meu coração
A gente vai contra a corrente
Até não poder resistir
Na volta do barco é que sente
O quanto deixou de cumprir
Faz tempo que a gente cultiva
A mais linda roseira que há
Mas eis que chega a roda viva
E carrega a roseira pra lá
Roda mundo, roda-gigante
Roda moinho, roda,pião
# + id="zpxITXRvA1p1"
class ArquivoTexto(object):
def __init__(self, arquivo: str):
self.arquivo = arquivo
self.conteudo = self._extrair_conteudo()
self.extrair_linha = self.extrair_linha
def _extrair_conteudo(self):
conteudo = None
with open(file=self.arquivo,mode='r',encoding='utf8') as arquivo:
conteudo = arquivo.readlines()
return conteudo
def extrair_linha(self, numero_linha: int):
return self.conteudo[numero_linha-1]
def extrair_coluna(self, indice_coluna: str):
coluna = list()
for linha in self.conteudo:
conteudo_linha = linha.strip().split(sep=',')
coluna.append(conteudo[indice_coluna])
coluna.pop[-1]
return coluna
# + id="tKf4FzZWkbkl"
arquivo_texto = ArquivoTexto(arquivo='musica.txt')
# + colab={"base_uri": "https://localhost:8080/"} id="6xal28vck3ds" outputId="896a5267-a7d1-4af9-a12c-fe93f5ac9719"
numero_linha = 1
print(arquivo_texto.extrair_linha(numero_linha=numero_linha))
# + colab={"base_uri": "https://localhost:8080/"} id="qdXRNfZ3lQtB" outputId="8310deaa-00cf-487b-b2f7-70d99565139f"
numero_linha = 10
print(arquivo_texto.extrair_linha(numero_linha=numero_linha))
# + id="9onGc-EfJOke"
class ArquivoCSV(ArquivoTexto):
def __init__(self, arquivo: str):
self.arquivo = arquivo
self.conteudo = self._extrair_conteudo()
self.extrair_coluna_da_linha =self.extrair_coluna_da_linha
self.coluna = self.extrair_coluna
def _extrair_conteudo(self):
conteudo = None
with open(file=self.arquivo,mode='r',encoding='utf8') as arquivo:
conteudo = arquivo.readlines()
return conteudo
def extrair_coluna_da_linha(self,numero_linha: int,numero_coluna:int):
return self.conteudo[numero_linha-1]
def extrair_coluna(self, indice_coluna: str):
coluna = list()
for linha in self.conteudo:
conteudo_linha = linha.strip().split(sep=',')
coluna.append(conteudo[indice_coluna])
coluna.pop[-1]
return coluna
# + id="L695YWL2ytJq"
arquivo_csv = ArquivoCSV(arquivo='carros.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="NNF9Z0774F_S" outputId="e927f6e1-2784-458c-bd15-67f94fc0e5b5"
print(arquivo_csv.coluna)
# + colab={"base_uri": "https://localhost:8080/"} id="_cUfW6xlz2dt" outputId="4dc17b3a-c080-4cc8-c441-37829466530d"
numero_linha = 1
print(arquivo_csv.extrair_linha(numero_linha=numero_linha))
# + colab={"base_uri": "https://localhost:8080/"} id="d1kk2i1e0s_a" outputId="b6e7d543-08fd-4697-b653-30946d00f9c1"
numero_linha = 10
print(arquivo_csv.extrair_linha(numero_linha=numero_linha))
|
Untitled26.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# First, let's import python modules that we'll need, everything is in
# the standard library except for the astpretty module. We're using
# that to output an AST in a more readible format. Feel free to replace
# the ast.pprint calls with ast.dump calls if you want to avoid
# installing the astpretty module
#
#
#
#
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
import argparse
import ast
import collections
import copy
import sys
import timeit
import astpretty
# -
# We'll use `example.py` as the source code that we'll examine and modify. The `ast.parse` method will take a string with python code and return a corresponding `ast.AST` object. We'll use this throughout the code to load and generate an AST from `example.py`
#
# Now, let's look at using the `ast.NodeVisitor` to examine our source code. First, we'll do the functional equivalent of a hello world program by printing out any functions that we define in our source code. We use `ast.NodeVisitor` to do this by inheriting from the class and overriding the various visit_* methods. I've done this in two different ways:
#
# * override `generic_visit` and then if the node we're visiting is an `ast.FunctionDef` object, print out it's name
# * override `visit_FunctionDef` and then print out the node name since this method only gets called on FunctionDef nodes
# +
class HelloVisitor(ast.NodeVisitor):
def generic_visit(self, node):
if node.__class__ == ast.FunctionDef:
sys.stdout.write(f"Defining function: {node.name} \n")
super().generic_visit(node)
class FunctionVisitor(ast.NodeVisitor):
def visit_FunctionDef(self, node):
sys.stdout.write(f"Defining function: {node.name} \n")
# -
# Finally we'll instantiate our objects and call the `visit` methods on the objects to traverse the AST tree:
with open("example.py", "r") as source:
tree = ast.parse(source.read())
sys.stdout.write("Visiting nodes using generic_visit:\n")
hello_visitor = HelloVisitor()
hello_visitor.visit(tree)
sys.stdout.write("Visiting nodes using specific visit function:\n")
function_visitor = FunctionVisitor()
function_visitor.visit(tree)
# Now we'll do something a bit more complex with the `NodeVisitor` class. We'll do a bit of static analysis to flag variables that are defined but haven't been used. We'll do this by looking at all the `Assign` nodes to record all the variables created. Next we'll look at the `Name` nodes to record when the variables are used. By comparing the two lists, we can see when a variable has been created but hasn't been used.
#
# **Note:** we need to be careful here to respect python's scoping rules. We need to record if a variable is defined within a function so that we can catch cases where a variable is created in one scope and used in another scope. To make life (and the code) simpler, we'll ignore class definitions entirely but handling that would be analagous to handling variables within a function. Also, this misses variables used in code that is evaluated at runtime, but if you're doing that, you should be prepared for this.
class UnusedVariables(ast.NodeVisitor):
"""
Print out variables that are set but might be unused
"""
def __init__(self):
super().__init__()
self.variable_def = {'global': {'global': [],
'nonlocal': [],
'local': []}}
self.variable_used = {'global': {}}
self.stack = ["global"]
self.function_name = "global"
def check(self, node: ast.AST) -> None:
self.generic_visit(node)
for var in self.variable_def['global']['local']:
if var[0] not in self.variable_used['global']:
sys.stdout.write(f"Variable {var[0]} defined in global on line {var[2]} not used\n")
def visit_FunctionDef(self, node: ast.FunctionDef) -> None:
self.function_name = node.name
self.stack.append(node.name)
self.variable_used[node.name] = {}
self.variable_def[node.name] = {'global': [],
'nonlocal': [],
'local': []}
for arg in node.args.args:
self.variable_def[node.name]['local'].append((arg.arg, node.name, node.lineno))
super().generic_visit(node)
for var in self.variable_def[node.name]['local']:
if var[0] not in self.variable_used[node.name]:
sys.stdout.write(f"Variable {var[0]} defined in {node.name} on line {var[2]} not used\n")
del self.variable_used[node.name]
del self.variable_def[node.name]
# remove function information
self.function_name = self.stack.pop()
self.function_name = self.stack[-1]
def register_variable(self, scope: str, var_name: str, line: int) -> None:
if scope in self.variable_def[self.function_name]:
if var_name not in [x[0] for x in self.variable_def[self.function_name][scope]]:
self.variable_def[self.function_name][scope].append((var_name, self.function_name, line))
else:
self.variable_def[self.function_name][scope] = [(var_name, self.function_name, line)]
def visit_Nonlocal(self, node: ast.Nonlocal) -> None:
for name in node.names:
self.register_variable('nonlocal', name, node.lineno)
def visit_Global(self, node: ast.Global) -> None:
for name in node.names:
self.register_variable('global', name, node.lineno)
def visit_Assign(self, node: ast.Assign) -> None:
# register the LHS of assignment
if isinstance(node.targets[0], ast.Name):
self.register_variable('local', node.targets[0].id, node.lineno)
# examine the RHS of assignment
super().generic_visit(node.value)
def visit_Call(self, node: ast.Call) -> None:
# visit parameters and register them
for arg in node.args:
if isinstance(arg, ast.Name):
self.register_usage(arg.id)
def var_defined(self, scope: str, var_name: str, function: str = None) -> bool:
if function is None:
function = self.function_name
return var_name in [x[0] for x in self.variable_def[function][scope]]
def register_usage(self, var_name: str) -> None:
# if we know the var is global we can short-circuit the registration
if self.var_defined('global', var_name):
if var_name not in self.variable_used['global']:
self.variable_used['global'][var_name] = True
return
# otherwise we need to check up the stack
for function in reversed(self.stack):
# short circuit global vars
if self.var_defined('global', var_name, function):
if var_name not in self.variable_used['global']:
self.variable_used['global'][var_name] = True
return
# nonlocal variables are defined deeper in the stack
if self.var_defined('nonlocal', var_name, function):
continue
# if definition found, mark var as being used and exit
if self.var_defined('local', var_name, function):
if var_name not in self.variable_used[function]:
self.variable_used[function][var_name] = True
return
# continue going deeper in the stack to find the variable
# if we're here, we haven't found the variable anywhere
sys.stdout.write(f"In {self.function_name}, {var_name} used without being defined\n")
def visit_Name(self, node: ast.Name) -> None:
self.register_usage(node.id)
# The basic idea of the code is to run some initialization call whenever a function is defined in order to setup dictionaries to track local, nonlocal, and global variables as well as a "stack" of where we are. Then the code gets invoked whenever a `nonlocal`, `global`, function call, variable assignment, or variable reference is made. The code will then register the variable in the appropriate location in the `self.variable_def` dictionary. When a variable is used (i.e. gets referenced in a `Name` node), the `self.variable_used` dictionary gets updated. Finally, once a function body is processed, the code will go through and see which variables are not used.
#
# The code below runs this analysis on a test file(`var_assignment.py`)
with open("var_assignment.py", "r") as source:
tree = ast.parse(source.read())
check_code = UnusedVariables()
check_code.check(tree)
# Note that `__name__` is marked as being used without being defined since the code doesn't know about builtins. Also, baz is incorrectly analyzed. That's because the test_traversal function is defined before the code for file. This can be avoided by having the code do multiple analysis passes to catch forward references like this.
# ### Static Analysis
#
# Now we'll look at using static analysis to modify ASTs and possibly optimize our code. We'll do two different optimizations:
#
# * dead code elimination
# * function inlining
# #### Dead code elimination
#
# Dead code elimination involves analyzing our code and removing code that we can guarantee will never run. For example, in the following code:
#
# ```
# if True:
# a = 5
# else:
# a = 10
# ```
#
# we know that the ```a=5``` branch will always be taken. So we can effectively replace a comparison and variable assignment with a single assignment. This is potentially a next bonus by reducing our code size, and avoiding a comparison and jump.
#
# This becomes even more powerful if we use constant folding and constant propagation to evaluate code at runtime. E.g. replacing code like
#
# ```
# x = 5
# y = x + 10
# if y > 5:
# x = 20
# ```
#
# with
#
# ```
# x = 5
# y = 15
# if 15 > 5:
# x = 20
# ```
# then
# ```
# x = 5
# y = 15
# x = 20
# ```
#
# and finally
#
# ```
# y = 15
# x = 20
# ```
# Notice that in optimizing the code, running some optimizations like dead code elimination at different stages might be useful.
# #### Function inlining
#
# Function inlining is a bit more ambiguous optimization. Function inlining involves taking the body of a function, and replacing function calls with the body of the function. E.g. `a = func(a)` gets replaced with the code for `func` and instead of having `func` return a value, it is assigned to `a`. Depending on the size of the function, this may be a net benefit or may even slow down the code. This also is a bit more complicated since we need to propagate function arguments into the body and make sure that variables declared within the function are renamed so that they don't overwrite variables where the function was called.
#
# To make the code much simpler, we'll just inline functions that consist of calls to other functions.
# + [markdown] pycharm={"name": "#%%\n"}
# First we'll define a helper function to time our code fragments. The `timeit.timeit` call runs our code fragment 10,000 times and outputs the time it took to run. Since timeit runs the code in a clean namespace we need to use the globals keyword arg to pass in the AST that we want to time and then use the `compile` and `exec` calls to first compile an `ast.AST` object to executable code and then to execute it.
# -
def time_tree(tree: ast.AST) -> float:
"""
Time ast execution
:param tree: AST tree
:return: float recording number of seconds to run snippet
"""
return timeit.timeit(stmt="exec(compile(tree, 'test', mode='exec'))", number=10000, globals={'tree': tree})
# #### Dead code elimination using ASTs
#
# Now, we'll look at the code for doing dead code elimination:
class RemoveDeadCode(ast.NodeTransformer):
"""
Simplify branches that test constants to remove dead code
e.g. if true: a = 5 else a = 10 gets turned into a = 5
"""
def visit_If(self, node):
# make sure to recurse on branches
super().generic_visit(node)
if type(node.test) in [ast.Constant, ast.NameConstant]:
if node.test.value:
new_node = node.body
else:
new_node = node.orelse
if not new_node:
# if branch being used is empty, delete
return None
return new_node
else:
return node
def optimize(self, node: ast.AST):
"""
Optimize nodes
:param node: node to optimize
:return: ast tree
"""
new_node = self.visit(node)
ast.fix_missing_locations(new_node)
return new_node
# The code is fairly simple, it looks at any if statements and if the comparison in the if statement is a constant, the code will evaluate the constant and return the code in the appropriate branch. There's quite a bit more that can be done but that's left as an exercise for the reader.
# #### Function inlining
#
# As mentioned before function inlining is quite a bit more complex since we need to be careful to avoid overwriting variables and to preserve scoping rules when moving code around. To make things much more simple, we'll only inline functions where the body consists of other function calls and which does not return any values. E.g. a function like this
#
# ```
# def foo(a, b, c):
# bar(a)
# baz(a, c)
# bar(b)
#
# ```
#
# This will let us doing inlining by replacing the function call with the body after some substitution of function arguments.
#
# First, we'll write some code that'll scan the source and figure out which functions can be inlined and to cache the function body if a function can be inlined.
class CallOnlyChecker(ast.NodeVisitor):
"""
Visit and check function definitions to make sure that it can be inlined,
i.e. make sure function only consists of calls
Also cache function defs so that they can later be inlined
"""
def __init__(self):
self.__function_cache__ = collections.defaultdict(lambda: {'args': [], 'body': []})
self.__inlineable_cache__ = collections.defaultdict(lambda: False)
super().__init__()
def can_inline(self, name: str) -> bool:
"""
Check to see if function can be inlined
:param name: name of function to check
:return: True if function can be inlined, False otherwise
"""
return self.__inlineable_cache__[name]
def get_cached_function(self, name: str) -> dict:
"""
Get dictionary with function information
:param name: name of function
:return: a
"""
return copy.copy(self.__function_cache__[name])
def visit_FunctionDef(self, node: ast.FunctionDef):
"""
Examine function and cache if needed
:param node: ast.FunctionDef to analyze
:return: None
"""
for func_node in ast.iter_child_nodes(node):
if isinstance(func_node, ast.Expr) and type(func_node.value) in [ast.Str, ast.Call]:
next
elif isinstance(func_node, ast.arguments):
next
else:
self.__inlineable_cache__[node.name] = False
return
self.__inlineable_cache__[node.name] = True
self.__function_cache__[node.name] = {'args': node.args}
if isinstance(node.body[0], ast.Expr) and isinstance(node.body[0].value, ast.Str):
# skip docstring
call_list = node.body[1:]
else:
call_list = node.body
call_list = map(lambda x: x.value, call_list)
self.__function_cache__[node.name]['body'] = call_list
# The code just examines all the `FunctionDef` nodes and then looks at the function body to see if it can be inlined and caches the function body if that's the case.
#
# Now the code that does the actual inlining:
class InlineFunctions(ast.NodeTransformer):
"""Inline functions that only have calls in them"""
def __init__(self):
self.checker = CallOnlyChecker()
super().__init__()
def optimize(self, tree: ast.AST):
"""
verify and inline functions if functions only consist of calls
:param tree: ast node
:return: None
"""
# get information about ast
self.checker.visit(tree)
inlined_tree = self.visit(tree)
inlined_tree = CleanupAST().cleanup(inlined_tree)
ast.fix_missing_locations(inlined_tree)
return inlined_tree
def visit_Call(self, node):
"""
Look at function calls and replace if possible
"""
super().generic_visit(node)
if self.checker.can_inline(node.func.id):
replacement_code = self.checker.get_cached_function(node.func.id)
replacement_args = replacement_code['args']
replacement_nodes = []
replacement_table = {}
index = 0
for arg in node.args:
arg_name = replacement_args.args[index].arg
replacement_table[arg_name] = arg
index += 1
for call in replacement_code['body']:
new_args = []
for arg in call.args:
if isinstance(arg, ast.Name):
if arg.id in replacement_table:
new_args.append(replacement_table[arg.id])
else:
new_args.append(arg)
else:
new_args.append(arg)
call.args = new_args
replacement_nodes.append(call)
if len(replacement_nodes) == 1:
return replacement_nodes[0]
else:
for node in replacement_nodes:
node.lineno = 0
return replacement_nodes
else:
return node
# This code just goes through an AST and first processes it to find functions that can be inlined. Once that's done, it goes through the AST again and replaces a `Call` node with the appropriate `Call` nodes from within the function while doing substitutions between the parameters in the original `Call` node and the parameters in the function definition. There's a few corner cases this code won't cover but the existing code should illustrate the general principles
#
# There is one tricky bit here due to the way the python ASTs and the ast.NodeTransformer works. The resulting AST will have `Expr` nodes with multiple `Call` nodes which is not allowed. The following code will fix this by going through the AST and splitting these invalid `Expr` nodes into multiple `Expr` nodes.
# + pycharm={"name": "#%%\n"}
class CleanupAST(ast.NodeTransformer):
"""
Scan and cleanup ASTs to split Expr nodes with values that contain multiple
expressions
"""
def cleanBody(self, node_body: list) -> list:
"""
Clean up expr nodes in a list, splitting expr with
multiple statements in value field
:param node_body: list of ast nodes
:return: list of cleaned ast nodes
"""
new_body = []
for child_node in node_body:
if isinstance(child_node, ast.Expr) and isinstance(child_node.value, list):
if len(child_node.value) == 1:
new_body.append(ast.Expr(value=child_node.value[0]))
else:
for stmt in child_node.value:
new_body.append(ast.Expr(value=child_node.value[0]))
else:
new_body.append(child_node)
return new_body
def visit_Module(self, node: ast.Module):
"""
Clean up expr nodes in a module
:param node: ast.Module instance to cleanup
:return: cleaned up node
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_FunctionDef(self, node: ast.FunctionDef):
"""
Examine function and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_If(self, node: ast.If):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_For(self, node: ast.For):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_While(self, node: ast.While):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_With(self, node: ast.With):
"""
Examine with statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_Try(self, node: ast.Try):
"""
Examine try statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.finalbody = self.cleanBody(node.finalbody)
return node
def visit_TryExcept(self, node: ast.Try):
"""
Examine try statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def cleanup(self, tree: ast.AST) -> ast.AST:
"""
Clean up an AST tree, splitting incorrect Expr if found
:param tree: AST to clean
:return: cleaned AST
"""
new_tree = self.visit(tree)
ast.fix_missing_locations(new_tree)
return new_tree
# -
# Now, let's see the code in action and see how it affects code execution time:
# + pycharm={"name": "#%%\n"}
with open("example.py", "r") as source:
tree = ast.parse(source.read())
sys.stdout.write(f"AST code:\n")
astpretty.pprint(tree)
code_timing = time_tree(tree)
branch_transformer = RemoveDeadCode()
pruned_tree = branch_transformer.optimize(tree)
deadcode_timing = time_tree(pruned_tree)
sys.stdout.write(f"transformed AST code:\n")
astpretty.pprint(pruned_tree)
function_transformer = InlineFunctions()
inlined_tree = function_transformer.optimize(pruned_tree)
astpretty.pprint(inlined_tree)
inlined_tree = ast.fix_missing_locations(inlined_tree)
inlined_code_timing = time_tree(inlined_tree)
sys.stdout.write(f"inlined AST code:\n")
astpretty.pprint(inlined_tree)
# + pycharm={"name": "#%%\n"}
sys.stdout.write(f"Time for code: {code_timing}\n")
sys.stdout.write(f"Time for code after using RemoveDeadCode: {deadcode_timing}\n")
sys.stdout.write(f"Time for code after using RemoveDeadCode and InlineFunction: {inlined_code_timing}\n")
# -
# So the dead code eliminiation sped things up a bunch and inlining didn't do much. These numbers will probably vary a bit on different systems but the relationship should remain about the same.
# + pycharm={"name": "#%%\n"}
# -
#
# +
class CleanupAST(ast.NodeTransformer):
"""
Scan and cleanup ASTs to split Expr nodes with values that contain multiple
expressions
"""
def cleanBody(self, node_body: list) -> list:
"""
Clean up expr nodes in a list, splitting expr with
multiple statements in value field
:param node_body: list of ast nodes
:return: list of cleaned ast nodes
"""
new_body = []
for child_node in node_body:
if isinstance(child_node, ast.Expr) and isinstance(child_node.value, list):
if len(child_node.value) == 1:
new_body.append(ast.Expr(value=child_node.value[0]))
else:
for stmt in child_node.value:
new_body.append(ast.Expr(value=child_node.value[0]))
else:
new_body.append(child_node)
return new_body
def visit_Module(self, node: ast.Module):
"""
Clean up expr nodes in a module
:param node: ast.Module instance to cleanup
:return: cleaned up node
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_FunctionDef(self, node: ast.FunctionDef):
"""
Examine function and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_If(self, node: ast.If):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_For(self, node: ast.For):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_While(self, node: ast.While):
"""
Examine if statement and clean up if necessary
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def visit_With(self, node: ast.With):
"""
Examine with statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
return node
def visit_Try(self, node: ast.Try):
"""
Examine try statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.finalbody = self.cleanBody(node.finalbody)
return node
def visit_TryExcept(self, node: ast.Try):
"""
Examine try statement
:param node: ast.FunctionDef to analyze
:return: None
"""
super().generic_visit(node)
node.body = self.cleanBody(node.body)
node.orelse = self.cleanBody(node.orelse)
return node
def cleanup(self, tree: ast.AST) -> ast.AST:
"""
Clean up an AST tree, splitting incorrect Expr if found
:param tree: AST to clean
:return: cleaned AST
"""
new_tree = self.visit(tree)
ast.fix_missing_locations(new_tree)
return new_tree
# -
# Now, let's see the code in action and see how it affects code execution time:
with open("example.py", "r") as source:
tree = ast.parse(source.read())
sys.stdout.write(f"AST code:\n")
astpretty.pprint(tree)
code_timing = time_tree(tree)
branch_transformer = RemoveDeadCode()
pruned_tree = branch_transformer.optimize(tree)
deadcode_timing = time_tree(pruned_tree)
sys.stdout.write(f"transformed AST code:\n")
astpretty.pprint(pruned_tree)
function_transformer = InlineFunctions()
inlined_tree = function_transformer.optimize(pruned_tree)
astpretty.pprint(inlined_tree)
inlined_tree = ast.fix_missing_locations(inlined_tree)
inlined_code_timing = time_tree(inlined_tree)
sys.stdout.write(f"inlined AST code:\n")
astpretty.pprint(inlined_tree)
sys.stdout.write(f"Time for code: {code_timing}\n")
sys.stdout.write(f"Time for code after using RemoveDeadCode: {deadcode_timing}\n")
sys.stdout.write(f"Time for code after using RemoveDeadCode and InlineFunction: {inlined_code_timing}\n")
# So the dead code eliminiation sped things up a bunch and inlining didn't do much. These numbers will probably vary a bit on different systems but the relationship should remain about the same.
|
python/ast_examples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **The neighbour-based recommendation for XXXXXXX**
# ---
#
# Various approaches(defers by algorithms) are to be tested and based on the time-lag, one will be finalised.
# - The main objective is to increase the user engagement from current average usage of **6 minutes**.
# - As each video is of almost 15 secs, then we have to take care of first 24 videos.
# - The user will be asked to give genre preference and upon these, recommendations shall occur.
# - The first k videos will be clubbed using rules
# - most recent
# - most viewed
# - most liked
# - most commented
# - For remaining **(24-k)** videos, the following algorithm are applied
# - k-Nearest Neighbour
class nearest_neighbour_recommender():
# #!pip install sklearn,pandas
#in case you do not have the dependecies, uncomment above line and run.
"""
---------------------------------
This class will create an object of recommendation algorithm.
This object will be fitted to the data provided and then,
the required recommendations will be provided as per the query by
the object.
Parameters
==================================================================
data = The data on which recommendation algorithm will be trained.
recs = Total recommendations required
n_jobs = -1 (This is the total processors that can be used for training.)
Procedure
=================================================================
1. call preprocessing function and pass the data path.
2. fit function will train the algorithm on data
3. For recommendations, call recommend and pass the data for which
recommendation is in need """
def __init__(self, recs_reqd = 10,n_jobs =-1):
self.recs_reqd = 10
self.n_jobs = -1
def preprocessing(self,data_path):
"""
Parameters
===============================================================
data_path = path of the data stored in csv format.
Functions
===============================================================
1.This function will read the data in .CSV format
2.It will store the video_names from which recommendation
will be offered.
3.It will drop the tags,description and video_name
columns so that the remaining data is in numeric form
for feeding the algorithm.
4. The missing values are filled with MODE of the relevant columns.
5. Genres are filtered and the most relevant ones are then one-hot encoded
6. The original genre column will be dropped to maintain non-redundancy.
"""
import pandas as pd
data = pd.read_csv(data_path)
self.video_names = data.video_name
data = data.drop(['tags','description','video_name'],axis=1)
data.fillna(data.mode().iloc[0], inplace=True)
gen = [i for i in list(data['genre'].str.lower().replace('#|/|.\\','').str.get_dummies(sep=',').
rename(lambda x: '' + x, axis='columns')) if len(i) > 3 and len(i) < 30 ]
gens = data['genre'].str.lower().replace('#|/|.\\','').str.get_dummies(sep=',').rename(lambda x: '' + x, axis='columns')[gen]
self.data = pd.concat([data.drop(['genre'],axis=1),gens],axis=1)
def fit(self):
from sklearn.neighbors import NearestNeighbors
#import pandas as pd
recommendations_object = NearestNeighbors(n_neighbors=self.recs_reqd,n_jobs=self.n_jobs)
recommendations_object.fit(self.data)
self.rec_sys = recommendations_object
print('training is over. Call recommend function for getting recommendations.')
def recommend(self,data_for_recommend_path):
"""
Parameters:
=========================================================================
data_for_recommend_path = data in .CSV for which recommendation is seeked
Function:
=========================================================================
returns the indeices of videos in database recommendations
"""
recommendations = self.rec_sys.kneighbors(self.preprocessing(data_for_recommend_path),return_distance=False)
#the below returns the indices of recommended videos.
return recommendations
data_path = 'data.csv'
recommender = nearest_neighbour_recommender()
recommender.preprocessing(data_path)
recommender.fit()
recs=recommender.recommend('Tnatan.csv')
#passed the same file for testing.
recs
recs[0]
#recommendations for first video in the test file
|
The neighbour-based recommendation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='Pierian_Data_Logo.png' /></a>
# ___
# # Welcome to the Course
#
# This is the repository for the course: **Python for Data Science and Machine Learning Bootcamp**
#
# Thank you so much for enrolling in the course!
#
# ## Overview of this Repository
#
# This repository contains everything you need for the course, it has all the notebooks, links
|
Udemy/Refactored_Py_DS_ML_Bootcamp-master/00-Welcome to the Course.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# +
import sys
import datetime
sys.path.append('..')
from models import I3D_movement as I3D
from importlib import reload
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn.metrics import confusion_matrix
import seaborn as sns
import numpy as np
from tensorflow.keras.optimizers import Adam
import configparser
config = configparser.ConfigParser()
config.read('../config.ini')
EPOCHS = 100
BATCH_SIZE = 32
LEARNING_RATE = 0.001
# -
load_model_filename = config['model']['path']
# +
X_train, X_train_mvm, y_train, X_val, X_val_mvm, y_val, X_test, X_test_mvm, y_test = I3D.load_datasets()
model = I3D.define_joint_model()
from tensorflow import keras
#model.compile(loss='categorical_crossentropy', optimizer = Adam(learning_rate=LEARNING_RATE), metrics=["accuracy"])
model.compile(optimizer= 'adam' , loss= keras.losses.binary_crossentropy, metrics=['accuracy']) #error fix
train_data = I3D.DataGenerator(X_train, X_train_mvm, y_train, BATCH_SIZE, True, 10, 0, 0)
val_data = I3D.DataGenerator(X_val, X_val_mvm, y_val, BATCH_SIZE)
test_data = I3D.DataGenerator(X_test, X_test_mvm, y_test, BATCH_SIZE)
current_time = datetime.datetime.now().isoformat().replace(":", "-")
model.load_weights(load_model_filename)
# Evaluating the final model on the test set
# Model expects 32x32 images not 24x24; not sure what was done without this.
X_test_padded = np.pad(X_test, ((0,0), (0,0), (4,4), (4,4), (0,0)))
X_val_padded = np.pad(X_val, ((0,0), (0,0), (4,4), (4,4), (0,0)))
val_predictions = model.predict([X_val_padded, X_val_mvm])
predictions = model.predict([X_test_padded, X_test_mvm])
y_pred = np.argmax(predictions, axis = 1)
y_test = np.argmax(y_test, axis = 1)
# +
y_pred_conf = np.max(predictions, axis=1)
size = len(y_pred)
confusion = confusion_matrix(y_test, y_pred)
confidence_matrix = np.zeros((13,13))
for i in range(size):
confidence_matrix[y_test[i]][y_pred[i]] += y_pred_conf[i]
norm_conf_matrix = np.divide(confidence_matrix.round(decimals=4), confusion, where =confusion!=0)
norm_conf_matrix[norm_conf_matrix < 0.001] = 0
# -
fig, ax = plt.subplots(figsize=(11, 9))
labs = ["Bird","Cat","Dog","False Pos","Hedgehog","Human","Insect","Leporidae","Mustelid","Possum","Rodent","Sheep","Wallaby"]
sns.heatmap(norm_conf_matrix, annot=True,cmap="OrRd",xticklabels=labs,yticklabels=labs)
plt.show()
fig, ax = plt.subplots(figsize=(15, 12))
sns.heatmap(confusion, annot=True,cmap="OrRd",xticklabels=labs,yticklabels=labs)
plt.show()
missclassifications = y_pred != y_test
np.count_nonzero(missclassifications)
def divide_zeros(a, b):
"Divide a by b, accepting 0/0=0"
return np.divide(a, b, out=np.zeros_like(a, dtype=float), where=b!=0)
def plot_confidence_hists(predictions, missclassifications, axes=None, BINS=100, xmin=None, xmax=1, label=None, alpha=1, color=None, scale_cumulative=False):
if axes is None:
fig, axes = plt.subplots(1, 4, figsize=(16,4), dpi=100)
# All Classifications
axes[0].hist(np.max(predictions, axis=1), bins=BINS, label=label, alpha=alpha, color=color)
axes[0].set_xlabel("Confidence")
axes[0].set_ylabel("All Classifications (Count)")
axes[0].set_xlim(xmin, xmax)
axes[0].set_title("Confidence")
# Misclassifications
axes[1].hist(np.max(predictions[missclassifications], axis=1), bins=BINS, alpha=alpha, color=color)
axes[1].set_xlabel("Confidence")
axes[1].set_ylabel("Misclassifications (Count)")
axes[1].set_xlim(xmin, xmax)
axes[1].set_title("Confidence of Misclassifications")
# Fraction of classifications at a particular confidence which are misclassifications
# Basically the misclassifications' confidences normalized by confidence distribution
all_bins = np.histogram(np.max(predictions, axis=1), bins=BINS)
misclassified_bins = np.histogram(np.max(predictions[missclassifications], axis=1), bins=all_bins[1])
axes[2].hist(all_bins[1][:-1], all_bins[1], weights=divide_zeros(misclassified_bins[0], all_bins[0]), alpha=alpha, color=color)
axes[2].set_xlabel("Confidence")
axes[2].set_ylabel("Misclassifications (Fraction)")
axes[2].set_xlim(xmin, xmax)
axes[2].set_title("Fraction of Misclassifications")
sums = np.cumsum(divide_zeros(misclassified_bins[0], all_bins[0])[::-1])[::-1]
if scale_cumulative:
#print(all_bins[1])
#print(sums)
scaled_bins = (all_bins[1]-np.min(all_bins[1]))/(np.max(all_bins[1])-np.min(all_bins[1]))
else:
scaled_bins = all_bins[1]
axes[3].hist(scaled_bins[:-1], scaled_bins, weights=sums/np.sum(sums), alpha=alpha, color=color)
axes[3].set_xlabel("Confidence")
axes[3].set_ylabel("Misclassifications (Fraction)")
axes[3].set_xlim(xmin, xmax)
axes[3].set_title("Cumulative Sum of Fraction")
if axes is None:
fig.suptitle("I3D Base Model Misclassifications by Confidience")
fig.tight_layout()
plot_confidence_hists(predictions, missclassifications)
predictions[:5]
# TODO:
# * Do this for binary classification so we get the result we really care about
# * Try techniques to improve it!
# ### Temperature Scaling
from temperature_scaling.temperature_scaling import find_scaling_temperature, apply_temperature_scaling, inverse_softmax
temperature = find_scaling_temperature(tf.constant(np.where(y_val==1)[1]), inverse_softmax(val_predictions))
temperature
scaled_predictions = apply_temperature_scaling(temperature, predictions)
# ### Platt Binning
import calibration as cal
calibrator = cal.PlattBinnerMarginalCalibrator(X_val.shape[0], num_bins=100)
calibrator.train_calibration(val_predictions, np.where(y_val==1)[1])
plat_scaled_predictions = calibrator.calibrate(predictions)
cmap = plt.get_cmap('Dark2')
cmap
fig, axes = plt.subplots(3, 4, figsize=(16, 12), dpi=100)
plot_confidence_hists(predictions, missclassifications, xmax=None, axes=axes[0], label='base', color=cmap.colors[0])
plot_confidence_hists(scaled_predictions, missclassifications, xmax=None, axes=axes[1], label='temperature scaled', color=cmap.colors[1])
plot_confidence_hists(plat_scaled_predictions, missclassifications, xmax=None, axes=axes[2], label='platt binned', color=cmap.colors[2])
fig.legend()
fig.tight_layout()
fig, axes = plt.subplots(1, 4, figsize=(16, 4), dpi=100)
plot_confidence_hists(predictions, missclassifications, xmax=None, axes=axes, label='base', alpha=1, xmin=0, scale_cumulative=True, color=cmap.colors[0])
plot_confidence_hists(scaled_predictions, missclassifications, xmax=None, axes=axes, label='temperature scaled', alpha=0.8, xmin=0, scale_cumulative=True, color=cmap.colors[1])
plot_confidence_hists(plat_scaled_predictions, missclassifications, xmax=None, axes=axes, label='platt binned', alpha=0.8, xmin=0, scale_cumulative=True, color=cmap.colors[2])
fig.legend()
fig.tight_layout()
# * Platt binning is better than temperature scaling, but both help a little bit
# * Time to read the Platt paper & then try CCAC?
|
notebooks/base_model_confidence.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
# +
# %%capture
import os
import site
os.sys.path.insert(0, '/home/schirrmr/code/reversible/')
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
os.sys.path.insert(0, '/home/schirrmr/code/explaining/reversible//')
# %load_ext autoreload
# %autoreload 2
import numpy as np
import logging
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
import matplotlib
from matplotlib import pyplot as plt
from matplotlib import cm
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
matplotlib.rcParams['figure.figsize'] = (12.0, 1.0)
matplotlib.rcParams['font.size'] = 14
import seaborn
seaborn.set_style('darkgrid')
from reversible2.sliced import sliced_from_samples
from numpy.random import RandomState
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
import copy
import math
import itertools
import torch as th
from braindecode.torch_ext.util import np_to_var, var_to_np
from reversible2.splitter import SubsampleSplitter
from reversible2.view_as import ViewAs
from reversible2.affine import AdditiveBlock
from reversible2.plot import display_text, display_close
from reversible2.bhno import load_file, create_inputs
th.backends.cudnn.benchmark = True
# +
sensor_names = ['Fz',
'FC3','FC1','FCz','FC2','FC4',
'C5','C3','C1','Cz','C2','C4','C6',
'CP3','CP1','CPz','CP2','CP4',
'P1','Pz','P2',
'POz']
orig_train_cnt = load_file('/data/schirrmr/schirrmr/HGD-public/reduced/train/4.mat')
train_cnt = orig_train_cnt.reorder_channels(sensor_names)
train_inputs = create_inputs(train_cnt, final_hz=256, half_before=True)
n_split = len(train_inputs[0]) - 40
test_inputs = [t[-40:] for t in train_inputs]
train_inputs = [t[:-40] for t in train_inputs]
# -
cuda = True
if cuda:
train_inputs = [i.cuda() for i in train_inputs]
test_inputs = [i.cuda() for i in test_inputs]
# +
from reversible2.graph import Node
from reversible2.branching import CatChans, ChunkChans, Select
from reversible2.constantmemory import sequential_to_constant_memory
from reversible2.constantmemory import graph_to_constant_memory
def invert(feature_model, out):
return feature_model.invert(out)
from copy import deepcopy
from reversible2.graph import Node
from reversible2.distribution import TwoClassDist
from reversible2.wrap_invertible import WrapInvertible
from reversible2.blocks import dense_add_no_switch, conv_add_3x3_no_switch
from reversible2.rfft import RFFT, Interleave
from reversible2.util import set_random_seeds
from torch.nn import ConstantPad2d
import torch as th
from reversible2.splitter import SubsampleSplitter
set_random_seeds(2019011641, cuda)
n_chans = train_inputs[0].shape[1]
n_time = train_inputs[0].shape[2]
base_model = nn.Sequential(
SubsampleSplitter(stride=[2,1],chunk_chans_first=False),
conv_add_3x3_no_switch(2*n_chans,32),
conv_add_3x3_no_switch(2*n_chans,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 4 x 128
conv_add_3x3_no_switch(4*n_chans,32),
conv_add_3x3_no_switch(4*n_chans,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 8 x 64
conv_add_3x3_no_switch(8*n_chans,32),
conv_add_3x3_no_switch(8*n_chans,32))
base_model.cuda();
branch_1_a = nn.Sequential(
SubsampleSplitter(stride=[2,1],chunk_chans_first=False), # 8 x 32
conv_add_3x3_no_switch(8*n_chans,32),
conv_add_3x3_no_switch(8*n_chans,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True),# 16 x 16
conv_add_3x3_no_switch(16*n_chans,32),
conv_add_3x3_no_switch(16*n_chans,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True), # 32 x 8
conv_add_3x3_no_switch(32*n_chans,32),
conv_add_3x3_no_switch(32*n_chans,32),
)
branch_1_b = nn.Sequential(
*(list(deepcopy(branch_1_a).children()) + [
ViewAs((-1, 32*n_chans,n_time//64,1), (-1,(n_time // 2)*n_chans)),
dense_add_no_switch((n_time // 2)*n_chans,32),
dense_add_no_switch((n_time // 2)*n_chans,32),
dense_add_no_switch((n_time // 2)*n_chans,32),
dense_add_no_switch((n_time // 2)*n_chans,32),
]))
branch_1_a.cuda();
branch_1_b.cuda();
branch_2_a = nn.Sequential(
SubsampleSplitter(stride=[2,1], chunk_chans_first=False),# 32 x 4
conv_add_3x3_no_switch(32*n_chans,32),
conv_add_3x3_no_switch(32*n_chans,32),
SubsampleSplitter(stride=[2,1],chunk_chans_first=True),# 64 x 2
conv_add_3x3_no_switch(64*n_chans,32),
conv_add_3x3_no_switch(64*n_chans,32),
ViewAs((-1, (n_time // 4)*n_chans,1,1), (-1,(n_time // 4)*n_chans)),
dense_add_no_switch((n_time // 4)*n_chans,64),
dense_add_no_switch((n_time // 4)*n_chans,64),
dense_add_no_switch((n_time // 4)*n_chans,64),
dense_add_no_switch((n_time // 4)*n_chans,64),
)
branch_2_b = deepcopy(branch_2_a).cuda()
branch_2_a.cuda();
branch_2_b.cuda();
final_model = nn.Sequential(
dense_add_no_switch(n_time*n_chans,256),
dense_add_no_switch(n_time*n_chans,256),
dense_add_no_switch(n_time*n_chans,256),
dense_add_no_switch(n_time*n_chans,256),
RFFT(),
)
final_model.cuda();
o = Node(None, base_model)
o = Node(o, ChunkChans(2))
o1a = Node(o, Select(0))
o1b = Node(o, Select(1))
o1a = Node(o1a, branch_1_a)
o1b = Node(o1b, branch_1_b)
o2 = Node(o1a, ChunkChans(2))
o2a = Node(o2, Select(0))
o2b = Node(o2, Select(1))
o2a = Node(o2a, branch_2_a)
o2b = Node(o2b, branch_2_b)
o = Node([o1b,o2a,o2b], CatChans())
o = Node(o, final_model)
o = graph_to_constant_memory(o)
feature_model = o
if cuda:
feature_model.cuda()
feature_model.eval();
# -
# ### set feature model weights and distribution to good start parameters
# +
from reversible2.constantmemory import clear_ctx_dicts
from reversible2.distribution import TwoClassDist
feature_model.data_init(th.cat((train_inputs[0], train_inputs[1]), dim=0))
# Check that forward + inverse is really identical
t_out = feature_model(train_inputs[0][:2])
inverted = invert(feature_model, t_out)
clear_ctx_dicts(feature_model)
assert th.allclose(train_inputs[0][:2], inverted, rtol=1e-3,atol=1e-4)
device = list(feature_model.parameters())[0].device
from reversible2.ot_exact import ot_euclidean_loss_for_samples
class_dist = TwoClassDist(2, np.prod(train_inputs[0].size()[1:]) - 2)
class_dist.cuda()
for i_class in range(2):
with th.no_grad():
this_outs = feature_model(train_inputs[i_class])
mean = th.mean(this_outs, dim=0)
std = th.std(this_outs, dim=0)
class_dist.set_mean_std(i_class, mean, std)
# Just check
setted_mean, setted_std = class_dist.get_mean_std(i_class)
assert th.allclose(mean, setted_mean)
assert th.allclose(std, setted_std)
clear_ctx_dicts(feature_model)
optim_model = th.optim.Adam(feature_model.parameters(), lr=1e-3, betas=(0.9,0.999))
optim_dist = th.optim.Adam(class_dist.parameters(), lr=1e-2, betas=(0.9,0.999))
# +
# %%writefile plot.py
import torch as th
import matplotlib.pyplot as plt
import numpy as np
from reversible2.util import var_to_np
from reversible2.plot import display_close
from matplotlib.patches import Ellipse
import seaborn
def plot_outs(feature_model, train_inputs, test_inputs, class_dist):
with th.no_grad():
# Compute dist for mean/std of encodings
data_cls_dists = []
for i_class in range(len(train_inputs)):
this_class_outs = feature_model(train_inputs[i_class])[:,:2]
data_cls_dists.append(
th.distributions.MultivariateNormal(th.mean(this_class_outs, dim=0),
covariance_matrix=th.diag(th.std(this_class_outs, dim=0) ** 2)))
for setname, set_inputs in (("Train", train_inputs), ("Test", test_inputs)):
outs = [feature_model(ins) for ins in set_inputs]
c_outs = [o[:,:2] for o in outs]
c_outs_all = th.cat(c_outs)
cls_dists = []
for i_class in range(len(c_outs)):
mean, std = class_dist.get_mean_std(i_class)
cls_dists.append(
th.distributions.MultivariateNormal(mean[:2],covariance_matrix=th.diag(std[:2] ** 2)))
preds_per_class = [th.stack([cls_dists[i_cls].log_prob(c_out)
for i_cls in range(len(cls_dists))],
dim=-1) for c_out in c_outs]
pred_labels_per_class = [np.argmax(var_to_np(preds), axis=1)
for preds in preds_per_class]
labels = np.concatenate([np.ones(len(set_inputs[i_cls])) * i_cls
for i_cls in range(len(train_inputs))])
acc = np.mean(labels == np.concatenate(pred_labels_per_class))
data_preds_per_class = [th.stack([data_cls_dists[i_cls].log_prob(c_out)
for i_cls in range(len(cls_dists))],
dim=-1) for c_out in c_outs]
data_pred_labels_per_class = [np.argmax(var_to_np(data_preds), axis=1)
for data_preds in data_preds_per_class]
data_acc = np.mean(labels == np.concatenate(data_pred_labels_per_class))
print("{:s} Accuracy: {:.1f}%".format(setname, acc * 100))
fig = plt.figure(figsize=(5,5))
ax = plt.gca()
for i_class in range(len(c_outs)):
#if i_class == 0:
# continue
o = var_to_np(c_outs[i_class]).squeeze()
incorrect_pred_mask = pred_labels_per_class[i_class] != i_class
plt.scatter(o[:,0], o[:,1], s=20, alpha=0.75, label=["Right", "Rest"][i_class])
assert len(incorrect_pred_mask) == len(o)
plt.scatter(o[incorrect_pred_mask,0], o[incorrect_pred_mask,1], marker='x', color='black',
alpha=1, s=5)
means, stds = class_dist.get_mean_std(i_class)
means = var_to_np(means)[:2]
stds = var_to_np(stds)[:2]
for sigma in [0.5,1,2,3]:
ellipse = Ellipse(means, stds[0]*sigma, stds[1]*sigma)
ax.add_artist(ellipse)
ellipse.set_edgecolor(seaborn.color_palette()[i_class])
ellipse.set_facecolor("None")
for i_class in range(len(c_outs)):
o = var_to_np(c_outs[i_class]).squeeze()
plt.scatter(np.mean(o[:,0]), np.mean(o[:,1]),
color=seaborn.color_palette()[i_class+2], s=80, marker="^",
label=["Right Mean", "Rest Mean"][i_class])
plt.title("{:6s} Accuracy: {:.1f}%\n"
"From data mean/std: {:.1f}%".format(setname, acc * 100, data_acc * 100))
plt.legend(bbox_to_anchor=(1,1,0,0))
display_close(fig)
return
# +
import pandas as pd
df = pd.DataFrame()
from reversible2.training import OTTrainer
trainer = OTTrainer(feature_model, class_dist,
optim_model, optim_dist)
# +
from reversible2.constantmemory import clear_ctx_dicts
from reversible2.timer import Timer
from plot import plot_outs
from reversible2.gradient_penalty import gradient_penalty
i_start_epoch_out = 4001
n_epochs = 10001
gen_frequency = 10
for i_epoch in range(n_epochs):
epoch_row = {}
with Timer(name='EpochLoop', verbose=False) as loop_time:
gen_update = (i_epoch % gen_frequency) == (gen_frequency-1)
loss_on_outs = i_epoch >= i_start_epoch_out
result = trainer.train(train_inputs, loss_on_outs=loss_on_outs)
epoch_row.update(result)
epoch_row['runtime'] = loop_time.elapsed_secs * 1000
if i_epoch % (n_epochs // 20) != 0:
df = df.append(epoch_row, ignore_index=True)
# otherwise add ot loss in
else:
for i_class in range(len(train_inputs)):
with th.no_grad():
class_ins = train_inputs[i_class]
samples = class_dist.get_samples(i_class, len(train_inputs[i_class]) * 4)
inverted = feature_model.invert(samples)
clear_ctx_dicts(feature_model)
ot_loss_in = ot_euclidean_loss_for_samples(class_ins.view(class_ins.shape[0], -1),
inverted.view(inverted.shape[0], -1)[:(len(class_ins))])
epoch_row['ot_loss_in_{:d}'.format(i_class)] = ot_loss_in.item()
df = df.append(epoch_row, ignore_index=True)
print("Epoch {:d} of {:d}".format(i_epoch, n_epochs))
print("Loop Time: {:.0f} ms".format(loop_time.elapsed_secs * 1000))
display(df.iloc[-3:])
plot_outs(feature_model, train_inputs, test_inputs,
class_dist)
fig = plt.figure(figsize=(8,2))
plt.plot(var_to_np(th.cat((th.exp(class_dist.class_log_stds),
th.exp(class_dist.non_class_log_stds)))),
marker='o')
display_close(fig)
# +
from reversible2.plot import plot_head_signals_tight
mean_bps_per_class = []
for i_class in range(len(train_inputs)):
samples = class_dist.get_samples(i_class, 400)
inverted = feature_model.invert(samples)
mean_bps_per_class.append(
np.mean(np.abs(np.fft.rfft(var_to_np(inverted.squeeze()))), axis=0))
fig = plt.figure(figsize=(8,3))
plt.plot(np.mean(mean_bps_per_class, axis=1).T)
display_close(fig)
fig = plot_head_signals_tight(np.stack(mean_bps_per_class, axis=-1), sensor_names=sensor_names,
figsize=(20,12));
# -
fig = plot_head_signals_tight(np.log(mean_bps_per_class[0]/mean_bps_per_class[1]), sensor_names=sensor_names,
figsize=(20,12));
for i_class in range(2):
samples = class_dist.get_samples(i_class, 20)
inverted = feature_model.invert(samples)
plot_head_signals_tight(var_to_np(inverted)[0].squeeze(), sensor_names=sensor_names,
figsize=(20,12));
for i_class in range(2):
samples = class_dist.get_mean_std(i_class)[0].unsqueeze(0)
inverted = train_inputs[i_class]
plot_head_signals_tight(var_to_np(inverted)[0].squeeze(), sensor_names=sensor_names,
figsize=(20,12));
for i_class in range(2):
samples = class_dist.get_mean_std(i_class)[0].unsqueeze(0)
inverted = feature_model.invert(samples)
plot_head_signals_tight(var_to_np(inverted)[0].squeeze(), sensor_names=sensor_names,
figsize=(20,12));
inverted_per_class = []
for i_class in range(2):
samples = class_dist.get_mean_std(i_class)[0].unsqueeze(0)
inverted = feature_model.invert(samples)
inverted_per_class.append(var_to_np(inverted)[0].squeeze())
plot_head_signals_tight(np.stack(inverted_per_class, axis=-1), sensor_names=sensor_names,
figsize=(20,12));
# +
for i_class in range(len(train_inputs)):
with th.no_grad():
class_ins = train_inputs[i_class]
samples = class_dist.get_samples(i_class, len(train_inputs[i_class]) * 4)
inverted = feature_model.invert(samples)
clear_ctx_dicts(feature_model)
ot_loss_in = ot_euclidean_loss_for_samples(class_ins.view(class_ins.shape[0], -1),
inverted.view(inverted.shape[0], -1)[:(len(class_ins))])
epoch_row['ot_loss_in_{:d}'.format(i_class)] = ot_loss_in.item()
df = df.append(epoch_row, ignore_index=True)
print("Epoch {:d} of {:d}".format(i_epoch, n_epochs))
print("Loop Time: {:.0f} ms".format(loop_time.elapsed_secs * 1000))
display(df.iloc[-3:])
plot_outs(feature_model, train_inputs, test_inputs,
class_dist)
fig = plt.figure(figsize=(8,2))
plt.plot(var_to_np(th.cat((th.exp(class_dist.class_log_stds),
th.exp(class_dist.non_class_log_stds)))),
marker='o')
display_close(fig)
# + language="javascript"
# var kernel = IPython.notebook.kernel;
# var thename = window.document.getElementById("notebook_name").innerHTML;
# var command = "nbname = " + "'"+thename+"'";
# kernel.execute(command);
# -
nbname
folder_path = '/data/schirrmr/schirrmr/reversible/models/notebooks/{:s}/'.format(nbname)
os.makedirs(folder_path,exist_ok=True)
name_and_variable = [('feature_model', feature_model),
('class_dist', class_dist),
('non_class_log_stds', class_dist.non_class_log_stds),
('class_log_stds', class_dist.class_log_stds,),
('class_means', class_dist.class_means),
('non_class_means', class_dist.non_class_means),
('feature_model_params', feature_model.state_dict()),
('optim_model', optim_model.state_dict()),
('optim_dist', optim_dist.state_dict())]
for name, variable in name_and_variable:
th.save(variable, os.path.join(folder_path, name + '.pkl'))
# ls -ahl /data/schirrmr/schirrmr/reversible/models/notebooks/21ChansOT/
# +
# compute metric log stuff fft
# and maybe even inception distance
# -
# first start just starting from one class all at 0, then optimize coefficients in loss
# then other class also
# repeat again, but starting from mean of classes respectively
# also add likelihood loss on them
# then just do some perturbations and inverse loss still
# then add class differentiation training,
# just switch classes explicitly by changing other features
# make it impossible to tell real class
# -> discriminator in --äääcääääääääääääääääääääääääääääääääääääääääääääääääääääääääää--...-m-input <y
|
notebooks/bhno-with-adversary/21ChansOT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''playground'': conda)'
# language: python
# name: python36964bitplaygroundconda3a42ba2689cb4827a840ce2c3efb16bd
# ---
# ## Remove entries in DrugBank that are similar to CYP450 dataset
# +
import sys
sys.path.append("..")
import itertools
import pandas as pd
from rdkit import DataStructs
from rdkit import Chem
from rdkit.Chem.rdMolDescriptors import GetMorganFingerprintAsBitVect
from tqdm import tqdm
# -
# ### Load datasets
cyp_data_path = "../data/fromraw_cid_inchi_smiles_fp_labels_onehots.csv"
cyp_df = pd.read_csv(cyp_data_path)
drugbank_data_path = "../data/DrugBank_smiles_fp.csv"
drugbank_df = pd.read_csv(drugbank_data_path)
cyp_df.head()
drugbank_df.head()
# ### Generate ECFPs
cyp_fps = list()
for smiles in cyp_df["isomeric_SMILES"]:
mol = Chem.MolFromSmiles(smiles)
fp = GetMorganFingerprintAsBitVect(mol, 4, nBits=2048)
cyp_fps.append(fp)
drugbank_fps = list()
for smiles in drugbank_df["SMILES"]:
mol = Chem.MolFromSmiles(smiles)
fp = GetMorganFingerprintAsBitVect(mol, 4, nBits=2048)
drugbank_fps.append(fp)
# ### Find the chemicals in DrugBank with a Tanimoto score higher than 0.85 to any chemicals in CYP450 dataset
db_rm_indices = list()
for cyp_fp in tqdm(cyp_fps):
for i, db_fp in enumerate(drugbank_fps):
score = DataStructs.FingerprintSimilarity(cyp_fp, db_fp)
if score > 0.85:
db_rm_indices.append(i)
# ### Remove similar chemicals
db_rm_indices_set = set(db_rm_indices)
filtered_drugbank = drugbank_df.drop(db_rm_indices_set)
# ### Save the filtered dataset
filtered_drugbank.to_csv("../data/DrugBank_smiles_fp_fil.csv")
|
notebooks/filter_drugbank.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import os
import sys
curr_path = os.getcwd()
gerkin_path = os.path.split(curr_path)[0]
olfaction_prediction_path = os.path.split(gerkin_path)[0]
if olfaction_prediction_path not in sys.path:
sys.path.append(olfaction_prediction_path)
import opc_python
import numpy as np
from datetime import datetime
from copy import copy,deepcopy
from scipy.stats import bernoulli
import matplotlib.pyplot as plt
import pystan
from sklearn.grid_search import GridSearchCV
from sklearn.cross_validation import ShuffleSplit
from sklearn.base import BaseEstimator
from sklearn.metrics import roc_curve,auc
from opc_python.utils import loading, scoring
from opc_python.gerkin import dream,fit1,fit2,params
perceptual_headers, perceptual_obs_data = loading.load_perceptual_data('training')
y,_ = dream.make_Y_obs('training',target_dilution='high',imputer='mask')
#values = y['mean_std'][:,:21]
#values.shape
values = np.dstack([y['subject'][i] for i in range(1,50)])
values = np.ma.array(values,mask=np.isnan(values))/100
values.data[values.mask]=0
code = """
data {
int<lower=0,upper=1000> n_mol; // number of molecules
int<lower=0,upper=100> n_desc; // number of descriptors
int<lower=0,upper=100> n_sub; // number of subjects
real<lower=0,upper=1> values[n_mol,n_desc,n_sub]; // data and replicates
int<lower=0,upper=1> mask[n_mol,n_desc,n_sub]; // data and replicates
}
parameters {
vector<lower=0.01,upper=0.49>[n_desc] mu1[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0.51,upper=0.99>[n_desc] mu2[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0,upper=100>[n_desc] n1[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0,upper=100>[n_desc] n2[n_sub]; // Strength of each molecule in each descriptor.
simplex[5] state[n_sub,n_desc]; // Strength of each molecule in each descriptor.
vector<lower=0,upper=1>[n_desc] dip[n_sub];
}
transformed parameters {
vector<lower=0>[n_desc] a1[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0>[n_desc] a2[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0>[n_desc] b1[n_sub]; // Strength of each molecule in each descriptor.
vector<lower=0>[n_desc] b2[n_sub]; // Strength of each molecule in each descriptor.
for(k in 1:n_sub) {
a1[k] <- mu1[k].*n1[k];
a2[k] <- mu2[k].*n2[k];
b1[k] <- (1-mu1[k]).*n1[k];
b2[k] <- (1-mu2[k]).*n2[k];
}
}
model {
real x;
real p50;
real lp;
for(k in 1:n_sub) {
//a1[k] ~ lognormal(0,1);
//b1[k] ~ lognormal(1,1);
//a2[k] ~ lognormal(1,1);
//b2[k] ~ lognormal(0,1);
for(j in 1:n_desc) {
state[k][j][3] ~ beta(1,1000);
//dip[k][j] ~ lognormal(0,1);
}
}
for (i in 1:n_mol) {
for(j in 1:n_desc) {
for(k in 1:n_sub) {
if(!mask[i][j][k]) {
x <- values[i][j][k];
if(x==0) {
lp <- log(
state[k][j][1] +
//exp(log(state[k][j][2]) + beta_log(0,a1[k][j],b1[k][j])) +
exp(log(state[k][j][3]) + uniform_log(0,0,1))
);
}
else if(x==1) {
lp <- log(
state[k][j][5] +
//exp(log(state[k][j][4]) + beta_log(1,a2[k][j],b2[k][j])) +
exp(log(state[k][j][3]) + uniform_log(1,0,1))
);
}
else {
lp <- log(
exp(log(state[k][j][2]) + beta_log(x,a1[k][j],b1[k][j])) +
exp(log(state[k][j][3]) + uniform_log(x,0,1)) +
exp(log(state[k][j][4]) + beta_log(x,a2[k][j],b2[k][j]))
);
}
p50 <- exp(log(state[k][j][2]) + beta_log(0.5,a1[k][j],b1[k][j])) +
exp(log(state[k][j][3]) + uniform_log(0.5,0,1)) +
exp(log(state[k][j][4]) + beta_log(0.5,a2[k][j],b2[k][j]));
if(x>0.495 && x<0.505) {
//lp <- log(exp(lp) - dip[k][j]*p50);
}
else {
//lp <- log(exp(lp) + dip[k][j]*p50/100);
}
increment_log_prob(lp);
}
}
}
}
}
"""
model = pystan.StanModel(model_code=code)
data = {'n_mol': values.shape[0], # number of molecules
'n_desc': values.shape[1], # number of descriptors
'n_sub' : values.shape[2],
'values' : values.data, # data and replicates
'mask' : values.mask.astype(int),
}
results = model.optimizing(data=data, verbose=True)
results['dip'].max()
from scipy.stats import norm,beta,lognorm,gamma,chi2,uniform
fig,axes = plt.subplots(3,7,figsize=(15,10))
x = np.linspace(0,1,51)
bins = 25
sub = 2
for i,ax in enumerate(axes.flat):
a1,b1,a2,b2,s,dip = [np.array(results[key])[sub][i] for key in ['a1','b1','a2','b2','state','dip']]
if i==0 and 0:
a1=2; b1=6; a2=12; b2=4; s = np.array([0.4,0.25,0.01,1,0.09])
s /= s.sum()
y = s[1]*beta.pdf(x,a1,b1) + \
s[3]*beta.pdf(x,a2,b2) + \
s[2]*uniform.pdf(x,0,1)
y[:2] = s[0]*bins + s[2]*uniform.pdf(0,0,1)
y[-2:] = s[4]*bins + s[2]*uniform.pdf(1,0,1)
#y[(len(x)-1)/2] *= (1-dip)
ax.plot(x,y)
ax.hist(values[:,i,sub],range=(0,1),bins=bins,normed=True)
ax.set_title('%d' % i)
from scipy.stats import norm,beta,lognorm,gamma,chi2,uniform
state,a1,a2,b1,b2 = [np.array(results[x]) for x in ['state','a1','a2','b1','b2']]
sub = 4
desc = 0
if 0:
state[sub][desc] = np.array([0.4,0.25,0.01,1,0.09])
state[sub][desc] /= state[sub][desc].sum()
a1[sub][desc] = 2
a2[sub][desc] = 6
b1[sub][desc] = 12
b2[sub][desc] = 4
logp_sum = 0
xs = []
logps = []
n_mol = values.shape[0]
for i in range(n_mol):
if not values.mask[i,1,3]:
x = values[i,1,3]
if x==0:
logp = np.log(state[sub][desc][0])# +
#state[3][1][2]*uniform.pdf(0,0,1) +
#state[3][1][1]*beta.pdf(0.00001,a1[3][1],b1[3][1]))
elif x==1:
logp = np.log(state[sub][desc][4])# +
#state[3][1][2]*uniform.pdf(1,0,1) +
#state[3][1][3]*beta.pdf(0.99999,a2[3][1],b2[3][1]))
else:
logp = np.log(state[sub][desc][1]*beta.pdf(x,a1[sub][desc],b1[sub][desc]) +
state[sub][desc][2]*uniform.pdf(x,0,1) +
state[sub][desc][3]*beta.pdf(x,a2[sub][desc],b2[sub][desc]))
#print(x,logp)
logp_sum += logp
xs.append(x)
logps.append(logp)
xs = np.array(xs)
logps = np.array(logps)
print(logp_sum)
plt.scatter(xs[(xs!=0) & (xs!=1)],logps[(xs!=0) & (xs!=1)])
plt.figure()
plt.hist(logps[(xs!=0) & (xs!=1)],bins=25);
logps[(xs!=0) & (xs!=1)].sum()
CIDs = loading.get_CIDs('training')
values = np.zeros((49,len(CIDs),21,2),dtype='float')
mask = np.ones((49,len(CIDs),21,2),dtype='int')
for line in perceptual_obs_data:
CID_index = CIDs.index(int(line[0]))
subject = int(line[5])
is_replicate = line[2]
if line[4] == '1/1,000':
#dilution_index = ['1/10','1/1,000','1/100,000'].index(line[4])
for i,value in enumerate(line[6:]):
if value != 'NaN':
mask[subject-1,CID_index,i,int(is_replicate)] = 0
if float(value) < 1:
value = 0.4
elif float(value) > 99:
value = 99.6
values[subject-1,CID_index,i,int(is_replicate)] = float(value)
|
opc_python/gerkin/stan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# language: python
# name: python38264bit7e3c702f16764767914e1c3a23341f94
# ---
# # Python Learning Notes
#
# ## Objected-oriented Programming
# ## 05/29/2020 - 06/01/2020
# ### Daily Objects
#
# 1. Python-100-days: Day 8
#
# 2. *Beginning Python*: Read Chapter 3
#
# 3. *Python Programming*: Read Chapter 1
#
# ### Progress
#
# 1. Python-100-days: Day 7 (100%)
#
# 2. Python-100-days: Day 8
#
# ### Python-100-days: Object-oriented Programming (OOP)
# - Object
#
# - Everything is object
#
# - Each object has attributes and behavior
#
# - Each object is unique
#
# - Class
#
# - Use to describe objects with similar attributes
class Coding(object):
# Use __init__ to initialize objects
def __init__(self, programmer, level):
self.programmer = programmer
self.level = level
# Define actions using function
def develop(self, project):
print(f'{self.programmer} is developing {project}.')
def procrastinate(self):
if self.level <= 2:
print(f'Are you out of your mind, {self.programmer}?! Your supervisor is watching!')
else:
print(f'You can do whatever you want, {self.programmer}. You\'re the boss!')
# Create new programmer files
programmer_1 = Coding('<NAME>', 5)
programmer_2 = Coding('Unpaid Intern', 1)
# Control actions using attributes
programmer_1.develop('WeChat Mini Programs')
programmer_2.procrastinate()
# - Encapsulation
#
# - To reduce complexity, conceal all details except the main logic flow.
# +
# Exercise 1
# Define a digital clock using class
from time import sleep
class Clock(object):
"""Digital clock"""
def __init__(self, hour=0, minute=0, second=0):
"""Initial: return to zero position
:param hour = hour (protected)
:param minute = minute (protected)
:param second = second (protected)
"""
self._hour = hour
self._minute = minute
self._second = second
def operate(self):
"""Operating"""
self._second += 1
if self._second == 60:
self._minute += 1
self._second = 0
if self._minute == 60:
self._hour += 1
self._minute = 0
if self._hour == 24:
self._hour = 0
def show_time(self):
return f'{self._hour:2d}:{self._minute:2d}:{self._second:2d}'
# Test
clock = Clock(10, 59, 55)
test = 0
while test < 11:
print(clock.show_time())
sleep(1)
clock.operate()
test += 1
# +
# Exercise 2
# Calculate the distance of two points using class
class Point(object):
def __init__(self, x=0, y=0):
self.x = x
self.y = y
def distance(self, other):
dx = self.x - other.x
dy = self.y - other.y
return (dx ** 2 + dy ** 2) ** 0.5
# Test
p1 = Point()
p2 = Point(3, 4)
print(p1.distance(p2))
|
Daily Progress/Day 8 - Object-oriented programming (OOP).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import libraries
import matplotlib.pyplot as plt
#from comet_ml import Experiment
import numpy as np
import os
import pandas as pd
import random
import seaborn as sns
import scikitplot as skplt
from sklearn.metrics import accuracy_score
from sklearn.metrics import roc_auc_score
from sklearn.metrics import roc_curve
from sklearn.metrics import f1_score
import tensorflow as tf
from tensorflow.keras import backend as K
from tensorflow.keras import initializers
from transformers import DistilBertTokenizerFast
from transformers import TFDistilBertModel, DistilBertConfig
# Import matplotlib
pd.plotting.register_matplotlib_converters()
get_ipython().run_line_magic('matplotlib', 'inline')
# Load training data
X_train = pd.read_csv(
'train.csv')['Transcription']
X_valid = pd.read_csv(
'valid.csv')['Transcription']
y_train = pd.read_csv('train.csv')['Type'] == 'Truthful'
y_valid = pd.read_csv(
'valid.csv')['Type'] == 'Truthful'
# Load test data
test = pd.read_csv('test.csv')
X_test = test['Transcription']
y_test = test['Type'] == 'Truthful'
# Check data
print('Our training data has ', len(X_train.index), ' rows.')
print('Our validation data has ', len(X_valid.index), ' rows.')
print('Our test data has ', len(X_test.index), ' rows.')
# Allow us to see full text (not truncated)
pd.set_option('display.max_colwidth', None)
# +
########## Ensure reproducibility ##########
# 1. Set `PYTHONHASHSEED` environment variable at a fixed value
os.environ['PYTHONHASHSEED'] = str(42)
# 2. Set `python` built-in pseudo-random generator at a fixed value
random.seed(42)
# 3. Set `numpy` pseudo-random generator at a fixed value
np.random.seed(42)
# 4. Set `tensorflow` pseudo-random generator at a fixed value
tf.random.set_seed(seed=42)
# -
print (y_train)
# +
# Define the maximum number of words to tokenize (DistilBERT can tokenize up to 512)
MAX_LENGTH = 256
# Define function to encode text data in batches
def batch_encode(tokenizer, texts, batch_size=256, max_length=MAX_LENGTH):
"""""""""
A function that encodes a batch of texts and returns the texts'
corresponding encodings and attention masks that are ready to be fed
into a pre-trained transformer model.
Input:
- tokenizer: Tokenizer object from the PreTrainedTokenizer Class
- texts: List of strings where each string represents a text
- batch_size: Integer controlling number of texts in a batch
- max_length: Integer controlling max number of words to tokenize in a given text
Output:
- input_ids: sequence of texts encoded as a tf.Tensor object
- attention_mask: the texts' attention mask encoded as a tf.Tensor object
"""""""""
input_ids = []
attention_mask = []
for i in range(0, len(texts), batch_size):
batch = texts[i:i+batch_size]
inputs = tokenizer.batch_encode_plus(batch,
max_length=MAX_LENGTH,
padding='max_length', # implements dynamic padding
truncation=True,
pad_to_max_length=True,
return_attention_mask=True,
return_token_type_ids=False
)
input_ids.extend(inputs['input_ids'])
attention_mask.extend(inputs['attention_mask'])
return tf.convert_to_tensor(input_ids), tf.convert_to_tensor(attention_mask)
# +
tokenizer = DistilBertTokenizerFast.from_pretrained('distilbert-base-uncased')
# Encode X_train
X_train_ids, X_train_attention = batch_encode(tokenizer, X_train.tolist())
# Encode X_valid
X_valid_ids, X_valid_attention = batch_encode(tokenizer, X_valid.tolist())
# Encode X_test
X_test_ids, X_test_attention = batch_encode(tokenizer, X_test.tolist())
# +
MAX_LENGTH = 256
LAYER_DROPOUT = 0.2
LEARNING_RATE = 3e-5
RANDOM_STATE = 42
def build_model(transformer, max_length=MAX_LENGTH):
"""""""""
Template for building a model off of the BERT or DistilBERT architecture
for a binary classification task.
Input:
- transformer: a base Hugging Face transformer model object (BERT or DistilBERT)
with no added classification head attached.
- max_length: integer controlling the maximum number of encoded tokens
in a given sequence.
Output:
- model: a compiled tf.keras.Model with added classification layers
on top of the base pre-trained model architecture.
"""""""""
# Define weight initializer with a random seed to ensure reproducibility
weight_initializer = tf.keras.initializers.GlorotNormal(
seed=RANDOM_STATE)
# Define input layers
input_ids_layer = tf.keras.layers.Input(shape=(max_length,),
name='input_ids',
dtype='int32')
input_attention_layer = tf.keras.layers.Input(shape=(max_length,),
name='input_attention',
dtype='int32')
# DistilBERT outputs a tuple where the first element at index 0
# represents the hidden-state at the output of the model's last layer.
# It is a tf.Tensor of shape (batch_size, sequence_length, hidden_size=768).
last_hidden_state = transformer(
[input_ids_layer, input_attention_layer])[0]
# We only care about DistilBERT's output for the [CLS] token, which is located
# at index 0. Splicing out the [CLS] tokens gives us 2D data.
cls_token = last_hidden_state[:, 0, :]
D1 = tf.keras.layers.Dropout(LAYER_DROPOUT,
seed=RANDOM_STATE
)(cls_token)
X = tf.keras.layers.Dense(128,
activation='relu',
kernel_initializer=weight_initializer,
bias_initializer='zeros'
)(D1)
D2 = tf.keras.layers.Dropout(LAYER_DROPOUT,
seed=RANDOM_STATE
)(X)
# X = tf.keras.layers.Dense(32,
# activation='relu',
# kernel_initializer=weight_initializer,
# bias_initializer='zeros'
# )(D2)
# D3 = tf.keras.layers.Dropout(LAYER_DROPOUT,
# seed=RANDOM_STATE
# )(X)
# Define a single node that makes up the output layer (for binary classification)
output = tf.keras.layers.Dense(1,
activation='sigmoid',
kernel_initializer=weight_initializer, # CONSIDER USING CONSTRAINT
bias_initializer='zeros'
)(D2)
# Define the model
model = tf.keras.Model([input_ids_layer, input_attention_layer], output)
# Compile the model
model.compile(tf.keras.optimizers.Adam(lr=LEARNING_RATE),
loss='binary_crossentropy',
metrics=['accuracy'])
return model
# +
from transformers import TFDistilBertModel, DistilBertConfig
DISTILBERT_DROPOUT = 0.2
DISTILBERT_ATT_DROPOUT = 0.2
# The bare, pretrained DistilBERT transformer model outputting raw hidden-states
# and without any specific head on top.
config = DistilBertConfig(dropout=DISTILBERT_DROPOUT,
attention_dropout=DISTILBERT_ATT_DROPOUT,
output_hidden_states=True)
distilBERT = TFDistilBertModel.from_pretrained(
'distilbert-base-uncased', config=config)
# Freeze DistilBERT layers to preserve pre-trained weights
for layer in distilBERT.layers:
layer.trainable = False
# Build model
model = build_model(distilBERT)
# -
print(X_train_ids.shape)
print(X_train_attention.shape)
print(y_train.to_numpy().shape)
print(X_valid_ids.shape)
print(X_valid_attention.shape)
print(y_valid.to_numpy().shape)
# +
EPOCHS = 6
BATCH_SIZE = 16
NUM_STEPS = len(X_train.index) // BATCH_SIZE
print(NUM_STEPS)
# Train the model
train_history1 = model.fit(
x=[X_train_ids, X_train_attention],
y=y_train.to_numpy(),
epochs=EPOCHS,
batch_size=BATCH_SIZE,
steps_per_epoch=NUM_STEPS,
validation_data=([X_valid_ids, X_valid_attention], y_valid.to_numpy()),
verbose=2
)
# +
FT_EPOCHS = 4
BATCH_SIZE = 16
NUM_STEPS = len(X_train.index) // BATCH_SIZE
FT_LEARNING_RATE = 2e-5
# Unfreeze DistilBERT weights to enable fine-tuning
for layer in distilBERT.layers:
layer.trainable = True
# Lower the learning rate to prevent destruction of pre-trained weights
optimizer = tf.keras.optimizers.Adam(lr=FT_LEARNING_RATE)
# Recompile model after unfreezing
model.compile(optimizer=optimizer,
loss='binary_crossentropy',
metrics=['accuracy'])
# Define callbacks
early_stopping = tf.keras.callbacks.EarlyStopping(monitor='val_loss',
mode='min',
min_delta=0,
patience=0,
restore_best_weights=True)
# Train the model
train_history2 = model.fit(
x=[X_train_ids, X_train_attention],
y=y_train.to_numpy(),
epochs=FT_EPOCHS,
batch_size=BATCH_SIZE,
steps_per_epoch=NUM_STEPS,
validation_data=([X_valid_ids, X_valid_attention], y_valid.to_numpy()),
callbacks=[early_stopping],
verbose=2
)
# +
# Generate predictions
y_pred = model.predict([X_test_ids, X_test_attention])
y_pred_thresh = np.where(y_pred >=0.5, 1, 0)
# Get evaluation results
accuracy = accuracy_score(y_test, y_pred_thresh)
auc_roc = roc_auc_score(y_test, y_pred)
# Log the ROC curve
fpr, tpr, thresholds = roc_curve(y_test.to_numpy(), y_pred)
fig, ax = plt.subplots(figsize=(10, 7))
ax.plot(fpr, tpr)
ax.plot(np.linspace(0, 1, 100),
np.linspace(0, 1, 100),
label='baseline',
linestyle='--')
plt.title('Receiver Operating Characteristic Curve', fontsize=18)
plt.ylabel('TPR', fontsize=16)
plt.xlabel('FPR', fontsize=16)
plt.legend(fontsize=12)
print('Accuracy: ', accuracy)
print('ROC-AUC: ', auc_roc)
# +
# Build train_history
history_df1 = pd.DataFrame(train_history1.history)
history_df2 = pd.DataFrame(train_history2.history)
history_df = history_df1.append(history_df2, ignore_index=True)
# Plot training and validation loss over each epoch
history_df.loc[:, ['loss', 'val_loss']].plot()
plt.title(label='Training + Validation Loss Over Time', fontsize=17, pad=19)
plt.xlabel('Epoch', labelpad=14, fontsize=14)
plt.ylabel('Binary Crossentropy Loss', labelpad=16, fontsize=14)
print("Minimum Validation Loss: {:0.4f}".format(history_df['val_loss'].min()))
# Save figure
# +
# Plot confusion matrix
skplt.metrics.plot_confusion_matrix(y_test.to_list(),
y_pred_thresh.tolist(),
figsize=(6, 6),
text_fontsize=14)
plt.title(label='Test Confusion Matrix', fontsize=20, pad=17)
plt.xlabel('Predicted Label', labelpad=14)
plt.ylabel('True Label', labelpad=14)
# -
|
distilbert.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/02_Python_Datatypes/tree/main/003_Python_List_Methods)**
# </i></small></small>
# # Python List `reverse()`
#
# The **`reverse()`** method reverses the elements of the list.
#
# **Syntax**:
#
# ```python
# list.reverse()
# ```
# ## `reverse()` Parameters
#
# The **`reverse()`** method doesn't take any arguments.
# ## Return Value from `reverse()`
#
# The **`reverse()`** method doesn't return any value. It updates the existing list.
# +
# Example 1: Reverse a List
# Operating System List
systems = ['Windows', 'macOS', 'Linux']
print('Original List:', systems)
# List Reverse
systems.reverse()
# updated list
print('Updated List:', systems)
# -
# There are other several ways to reverse a list.
# +
# Example 2: Reverse a List Using Slicing Operator
# Operating System List
systems = ['Windows', 'macOS', 'Linux']
print('Original List:', systems)
# Reversing a list
#Syntax: reversed_list = systems[start:stop:step]
reversed_list = systems[::-1]
# updated list
print('Updated List:', reversed_list)
# +
# Example 3: Accessing Elements in Reversed Order
# If you need to access individual elements of a list in the reverse order,
# it's better to use reversed() function.
# Operating System List
systems = ['Windows', 'macOS', 'Linux']
# Printing Elements in Reversed Order
for o in reversed(systems):
print(o)
# -
|
003_Python_List_Methods/008_Python_List_reverse().ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import os, glob
from osgeo import gdal
import earthpy as et
import geopandas as gpd
import rioxarray as rxr
import xarray as xr
from shapely.geometry import box
from tifffile import imsave
from rioxarray.merge import merge_arrays
from rasterio.plot import plotting_extent
import earthpy.plot as ep
import rasterio
from rasterio.plot import show
# +
# data paths
nitrogen_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen')
phosphorus_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'phosphorus')
nitrogen_to_mosaic = glob.glob(os.path.join(str(nitrogen_path), "*.tif"))
phosphorus_to_mosaic = glob.glob(os.path.join(str(phosphorus_path), "*.tif"))
# crop path
rr_hu8_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'polygon',
'RR_HU8-polygon.shp')
rr_hu8_boundary = gpd.read_file(rr_hu8_path)
# -
# # Functions section
# ### rxr library
# +
# function for clipping to shp and opening with rioxarray
def open_clean_bands(band_path,
crop_bound,
valid_range=None,
variable=None):
# YOUR CODE HERE
"""Open and clean a single landsat band .
Parameters
-----------
band_path:string A path to the array to be opened
crop_bound:geopandas GeoDataFrame
A geopandas dataframe to be used to crop the raster data using rioxarray clip().
valid_range:tuple (optional)
A tuple of min and max range of values for the data. Default = None
Returns
-----------
band : xarray DataArray
An xarray DataArray clipped to a crop boundary and masked if a range is given
"""
crop_bound_box = [box(*crop_bound.bounds.loc[0])]
try:
band = rxr.open_rasterio(band_path,
masked=True,
variable=variable,
parse_coordinates=False).rio.clip(crop_bound_box,
#crs=crop_bound.crs,
all_touched=True,
from_disk=True).squeeze()
except:
raise ValueError(
"Oops - I couldn't clip your data. This may be due to a crs error.")
# Only mask the data to the valid range if a valid range tuple is provided
if valid_range is not None:
mask = ((band < valid_range[0]) | (band > valid_range[1]))
band = band.where(~xr.where(mask, True, False))
return band
# -
# ### GDAL library
# +
# function for converting to raster after using GDAL merge command
def raster2array(geotif_file):
metadata = {}
dataset = gdal.Open(geotif_file)
metadata['array_rows'] = dataset.RasterYSize
metadata['array_cols'] = dataset.RasterXSize
metadata['bands'] = dataset.RasterCount
metadata['driver'] = dataset.GetDriver().LongName
metadata['projection'] = dataset.GetProjection()
metadata['geotransform'] = dataset.GetGeoTransform()
mapinfo = dataset.GetGeoTransform()
metadata['pixelWidth'] = mapinfo[1]
metadata['pixelHeight'] = mapinfo[5]
xMin = mapinfo[0]
xMax = mapinfo[0] + dataset.RasterXSize/mapinfo[1]
yMin = mapinfo[3] + dataset.RasterYSize/mapinfo[5]
yMax = mapinfo[3]
metadata['extent'] = (xMin,xMax,yMin,yMax)
raster = dataset.GetRasterBand(1)
array_shape = raster.ReadAsArray(0,0,metadata['array_cols'],metadata['array_rows']).astype(float).shape
metadata['noDataValue'] = raster.GetNoDataValue()
metadata['scaleFactor'] = raster.GetScale()
array = np.zeros((array_shape[0],array_shape[1],dataset.RasterCount),'uint8') #pre-allocate stackedArray matrix
if metadata['bands'] == 1:
raster = dataset.GetRasterBand(1)
metadata['noDataValue'] = raster.GetNoDataValue()
metadata['scaleFactor'] = raster.GetScale()
array = dataset.GetRasterBand(1).ReadAsArray(0,0,metadata['array_cols'],metadata['array_rows']).astype(float)
array[np.where(array==metadata['noDataValue'])]=np.nan
array = array/metadata['scaleFactor']
elif metadata['bands'] > 1:
for i in range(1, dataset.RasterCount+1):
band = dataset.GetRasterBand(i).ReadAsArray(0,0,metadata['array_cols'],metadata['array_rows']).astype(float)
band[np.where(band==metadata['noDataValue'])]=np.nan
#band = band/metadata['scaleFactor']
array[...,i-1] = band
return array, metadata
# -
# ### plotting
# +
# plotting function
def plot_array(array,spatial_extent,colorlimit,ax=plt.gca(),title='',cmap_title='',colormap=''):
plot = plt.imshow(array,extent=spatial_extent,clim=colorlimit);
cbar = plt.colorbar(plot,aspect=40); plt.set_cmap(colormap);
cbar.set_label(cmap_title,rotation=90,labelpad=20);
plt.title(title); ax = plt.gca();
ax.ticklabel_format(useOffset=False, style='plain');
rotatexlabels = plt.setp(ax.get_xticklabels(),rotation=90);
# -
# # Clipping section
# ### rxr library
# +
# prep shapefile for clipping
# convert to crs of original raster
rr_hu8_boundary_crs = rr_hu8_boundary.to_crs('EPSG:32610')
# +
# run function for clipping and cleaning nitrogen tifs (outputs a list of arrays)
nitrogen_tiles = []
for image, i in zip(nitrogen_to_mosaic, range(0,10)):
tile = open_clean_bands(image,
rr_hu8_boundary_crs,
valid_range=(0,1000),
variable=None)
nitrogen_tiles.append(tile)
# -
# ### GDAL library
# +
# GDAL library uses Warp() for clipping
for image, i in zip(nitrogen_to_mosaic, range(0,10)):
nitrogen_clip_gdal = gdal.Warp("/Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-clips-gdal/nitrogen-rr-hu8-crop-gdal{}.tif".format(i),
image,
cutlineDSName='/Users/merielle/desktop/earth-analytics/python/final-project/polygon/RR_HU8-polygon.shp',
cropToCutline=True,
dstNodata = 0)
# -
# # Merging Section
# ### rxr library
# +
# rearrange list of arrays from output of open_clean_bands
# order 1
rxr_order1 = [3, 0, 1, 2, 6, 7, 4, 8, 9, 5]
rxr_tiles_order1 = [nitrogen_tiles[i] for i in rxr_order1]
# order 2
rxr_order2 = [0, 5, 6, 4, 7, 8, 1, 3, 9, 2]
rxr_tiles_order2 = [nitrogen_tiles[i] for i in rxr_order2]
# +
# merge list of arrays for mosaic
# order 1
merged_nitrogen_order1 = merge_arrays(rxr_tiles_order1)
mask = (merged_nitrogen_order1 == 0)
merged_nitrogen_mask1 = (merged_nitrogen_order1.where(~xr.where(mask, True, False)))
# order2
merged_nitrogen_order2 = merge_arrays(rxr_tiles_order2)
mask = (merged_nitrogen_order2 == 0)
merged_nitrogen_mask2 = (merged_nitrogen_order2.where(~xr.where(mask, True, False)))
# +
# plot extent
nitrogen_extent = plotting_extent(merged_nitrogen[0],
merged_nitrogen.rio.transform())
# -
# ## GDAL library
# ### plots 2&3
# +
# file paths to rxr cropped rasters
nitrogen_clipped_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen',
'clipped')
nitrogen_clipped_to_mosaic = glob.glob(os.path.join(str(nitrogen_clipped_path), "*.tif"))
# +
# rearrange rxr clipped files
# order 1
order1 = [3, 1, 0, 2, 4, 7, 6, 8, 5, 9]
nitrogen_rxr_crop_to_mosaic_order1 = [nitrogen_clipped_to_mosaic[i] for i in order1]
string_nitrogen_rxr_crop_to_mosaic_order1 = " ".join(nitrogen_rxr_crop_to_mosaic_order1)
# order 2
order2 = [9, 5, 4 ,7, 8, 2, 3, 6, 1, 0]
nitrogen_rxr_crop_to_mosaic_order2 = [nitrogen_clipped_to_mosaic[i] for i in order2]
string_nitrogen_rxr_crop_to_mosaic_order2 = " ".join(nitrogen_rxr_crop_to_mosaic_order2)
# +
# merge rxr clipped files with 2 different orderings using gdal merge
# order1
command_nitrogen1 = "gdal_merge.py -o /Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-mosaic-clipped-rxr-order1.tif -of gtiff " + string_nitrogen_rxr_crop_to_mosaic_order1
print(os.popen(command_nitrogen1).read())
# +
# order2
command_nitrogen2 = "gdal_merge.py -o /Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-mosaic-clipped-rxr-order2.tif -of gtiff " + string_nitrogen_rxr_crop_to_mosaic_order2
print(os.popen(command_nitrogen2).read())
# -
# ### plots 4&5
# +
# file paths for GDAL clipped rasters
nitrogen_crop_gdal_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-clips-gdal')
nitrogen_crop_gdal_to_mosaic = glob.glob(
os.path.join(str(nitrogen_crop_gdal_path), "*.tif"))
# +
# rearrange GDAL clipped rasters to test out different orderings
# order 1
order1 = [3, 0, 1, 2, 6, 7, 4, 8, 9, 5]
nitrogen_crop_gdal_to_mosaic_order1 = [nitrogen_crop_gdal_to_mosaic[i] for i in order1]
nitrogen_string_gdal_clip_order1 = " ".join(nitrogen_crop_gdal_to_mosaic_order1)
# order 2
order2 = [9, 5, 4 ,7, 8, 2, 3, 6, 1, 0]
nitrogen_crop_gdal_to_mosaic_order2 = [nitrogen_crop_gdal_to_mosaic[i] for i in order2]
nitrogen_string_gdal_clip_order2 = " ".join(nitrogen_crop_gdal_to_mosaic_order2)
# +
# merge rasters
# order 1
command_nitrogen1 = "gdal_merge.py -o /Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-mosaic-clipped-order1.tif -of gtiff " + nitrogen_string_gdal_clip_order1
print(os.popen(command_nitrogen2).read())
# order 2
command_nitrogen2 = "gdal_merge.py -o /Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-mosaic-clipped-order2.tif -of gtiff " + nitrogen_string_gdal_clip_order2
print(os.popen(command_nitrogen2).read())
# -
# ### convert merged rasters to arrays for plotting
# +
# file paths to merged rasters
nitrogen_mosaic_gdal_clipped_path = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-mosaic-rr-hu8-crop-gdal.tif')
nitrogen_mosaic_path_order1 = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-mosaic-clipped-order1.tif')
nitrogen_mosaic_path_order1 = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-mosaic-clipped-order2.tif')
nitrogen_mosaic_path_rxr_order1 = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-mosaic-clipped-rxr-order1.tif')
nitrogen_mosaic_path_rxr_order2 = os.path.join(et.io.HOME,
'desktop',
'earth-analytics',
'python',
'final-project',
'nitrogen-mosaic-rxr-clipped-order2.tif')
# +
# convert rasters to arrays for plotting
nitrogen_crop_array_gdal, nitrogen_crop_gdal_metadata = raster2array(
nitrogen_mosaic_gdal_clipped_path)
# +
nitrogen_crop_array_rxr_order1, nitrogen_crop_rxr_order1_metadata = raster2array(
'/Users/merielle/desktop/earth-analytics/python/final-project/nitrogen-mosaic-clipped-order1.tif')
# -
nitrogen_crop_array_gdal_order2, nitrogen_crop_gdal_order2_metadata = raster2array(
nitrogen_mosaic_path_order2)
nitrogen_crop_array, nitrogen_crop_metadata = raster2array(
nitrogen_mosaic_path_rxr_order1)
nitrogen_crop_array, nitrogen_crop_metadata = raster2array(
nitrogen_mosaic_path_rxr_order2)
# # Plotting Section
# ## plot 1: rxr clipped and merged
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(merged_nitrogen_mask2[0,:,:],
nitrogen_extent,
(0,50),
title='Nitrogen rioxarray',
cmap_title='Nitrogen',
colormap='jet')
# set no data values to black
current_cmap = plt.cm.get_cmap()
current_cmap.set_bad(color='black')
# -
# ## rxr clipped, GDAL merged
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(nitrogen_crop_array_rxr[:,:,0],
nitrogen_crop_rxr_metadata['extent'],
(0,50),
title='Nitrogen Clipped with rioxarray, merged with GDAL',
cmap_title='Nitrogen Level',
colormap='jet')
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(nitrogen_crop_array[:,:,0],
nitrogen_crop_metadata['extent'],
(0,70),
title='Nitrogen',
cmap_title='Nitrogen Level',
colormap='jet')
# -
# ## clipped and merged with GDAL
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(nitrogen_crop_array_gdal_order1[:,:,0],
nitrogen_crop_gdal_order1_metadata['extent'],
(0,100),
title='Nitrogen order1',
cmap_title='Nitrogen Level',
colormap='jet')
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(nitrogen_crop_array_gdal_order2[:,:,0],
nitrogen_crop_gdal_order2_metadata['extent'],
(0,100),
title='Nitrogen order2',
cmap_title='Nitrogen Level',
colormap='jet')
# +
fig, ax = plt.subplots(figsize=(10,10))
plot_array(nitrogen_crop_array_gdal[:,:,0],
nitrogen_crop_gdal_metadata['extent'],
(0,100),
title='Nitrogen order 3',
cmap_title='Nitrogen Level',
colormap='jet')
# -
|
notebooks/data-processing/aviris.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Magic function to set the backend of matplotlib to the 'inline' backend
# %matplotlib inline
# # The CosmicIntegrator class
# ## 1. Introduction
#
#
# Here we show how to calculate the merger rate densities and observed probabilities for a population of double compact object mergers using the CosmicIntegrator class.
#
# In reality this class is one major for loop to redo the calculation at a single redshift for different redshifts, and a range of metallicities.
#
# The CosmicIntegrator class combines all the classes/modules that we presented in the cosmic integration notes.
# In addition it stores the results into several 2D arrays for easy plotting/binning, taking into account selection effects and volumes of the different redshift shells.
#
# Here we show:
#
# - How to set up the entire pipeline. The code specifically raises warnings/statements to help to remind you. This is simply a summary of the previous notes in the Cosmic Integration section.
#
# - How to calculate the rate and plot the results using the 2D arrays in the ClassCosmicIntegrator.
#
# - How to loop over variations efficiently to create different distributions.
# ## 1.1 Paths
# +
import os
pathNoteBook = os.getcwd()
pathScripts = pathNoteBook + '/PythonScripts/'
pathData = '/path-to-data/'
# -
# ## 1.2 Imports
import numpy as np
import sys
import matplotlib.pyplot as plt
#custom scripts
sys.path.append(pathScripts)
import ClassCosmicIntegrator #this imports other routines by itself
# ## 2. Creating an instance of the CosmicIntegrator class
# The main thing that needs to be defined in this class is the universe and the
# number of shells for which we want to calculate the merger rate densities. In addition we also set the GW-detector here.
#
# To this end, the following variables need to be defined in the instance:
#
# #For Data
# pathCOMPAS = None
#
# #Defining Universe
# Cosmology = 'WMAP'
# hubbleConstant = 67.8
# omegaMatter = 0.308
# redshiftFirstSFR = 10.0
#
# #Defining integral
# minRedshift = 0.0
# maxRedshift = 2.0
# nrRedshiftBins = 20
#
# #Method for calculating redshift from age
# RedshiftTabulated = True
# RedshiftTabulatedResolution = 100000
#
# #Detector
# GWdetector_sensitivity = 'O1'
# GWdetector_snrThreshold = 8
#
# #Print steps
# verbose = False
# `pathCOMPAS`
#
# Path to the h5-data.
#
# `Cosmology`
#
# Which of the astropy cosmology settings to use. Currently it takes either 'WMAP' or you set it to 'Custom Flat'. If you do the latter you need to set the hubbleConstant and omegaMatter. In the case of 'WMAP' these are ignored and a predefined setting is used.
#
# `hubbleconstant`
#
# See Cosmology.
#
# `omegaMatter`
#
# See Cosmology.
#
# `redshiftFirstSFR`
#
# The highest redshift at which we assume the first stars are born. Any star born at a higher redshift is assigned a rate of zero.
#
# `minRedshift`
#
# The lowest limit/edge of the redshift bins for the integral.
#
# `maxRedshift`
#
# The upper limit/edge of the redshfit bins for the integral.
#
# `nrRedshiftBins`
#
# The number of bins in which to resolve the integral.
#
# `RedshiftTabulated`
#
# Boolean value used to determine the method to calculate redshift of universe based on age. The functions are slow and hence looking it up in a predefined table is quicker and still sub-percent precision. If True we use the table method. If False we use an alternative function, which might be more precise but it is order of magnitude slower.
#
# `RedshiftTabulatedResolution`
#
# The resolution of the table (in from 0- to redshift first SFR) to calculate ages. The default tabulated method has a precision of $dz = \frac{1}{10000}$.
#
# `GWdetector_sensitivity`
#
# The sensitivity curve assumed to estimate the signal-to-noise of a system.
#
# `GWdetector_snrThreshold`
#
# The single detecter signal-to-noise limit above which we define a system to be detectable.
#
# `verbose`
#
# If True progress messages will be printed during calculation.
#
# ### 2.1 Creating all the instances
#using the defaults
CI = ClassCosmicIntegrator.CosmicIntegrator(pathCOMPAS=pathData)
#
# #### 2.1.1 Class COMPAS
#Info python Submit and assumptions to recover Msun evovled
CI.COMPAS.Mlower = 5.
CI.COMPAS.Mupper = 150.
CI.COMPAS.binaryFraction = 0.7
CI.COMPAS.setGridAndMassEvolved()
# which DCOs I want
CI.COMPAS.setCOMPASDCOmask(types='BBH', pessimistic=True)
# Get me the data
CI.COMPAS.setCOMPASData()
# #### 2.1.2 Class MSSFR
#
# +
# Pick the SFR model
CI.MSSFR.SFRprescription = 'Neijssel et al. (2019)'
#metallicity
CI.MSSFR.Zprescription = 'logNormal'
CI.MSSFR.logNormalPrescription ='Neijssel Phenomenological'
#Use the grid of the simulation
CI.MSSFR.metallicityGrid = CI.COMPAS.metallicityGrid
CI.MSSFR.calculateMetallicityBinEdges()
# -
# #### 2.1.3 Class CosmicIntegrator
#
#Use the COMPAS data and redshift grid to define 2D arrays and precalculate
#values needed for quick integration. This takes a bit. That is why we do it outside
#all other functions. Say you want to calculate the rates for three different
#SFR models. These values don't change
#(they only change when the redshift shells or the data changes)
#Hence you do it once.
CI.setBirthTimesAnd2Darrays()
# ## 3. Calculate and plot
# everthing is set so now so calculate :)
CI.cosmologicalIntegration()
# The results are stored in several different 2D arrays. Shown below is directly copied from source code:
# + active=""
# #################################################
# # #
# # the eventual results of interest #
# # if we already have data we can set it #
# # else we have to manually call tis function#
# #################################################
# #Calculating birth age is simple subtraction age-merger -delaytime
# self.PerSystemPerRedshift_ageBirth = None
# #Cannot do this in redshift so convert table above to a redshift table
# self.PerSystemPerRedshift_redshiftBirth = None
# # dN Gpc-3 per year at z=redshift. #each row is redshift merger
# self.PerSystemPerRedshift_ratesIntrinsic = None
# # dN per year in detector from shell
# self.PerSystemPerRedshift_ratesObserved = None
#
# -
# These are 2D arrays where each row is a redshift and each column is a single system.
print(CI.PerSystemPerRedshift_ratesIntrinsic.shape)
print(CI.nrRedshiftBins)
print(len(CI.COMPAS.delayTimes))
# This means that each cell is the rate of a single system at a single redshift.
#
# Now you can add rows/columns according to your interest. Examples are shown below.
# ### 3.1 Example rate in shell, and rate as function of redshift
#central redshift in the shell
redshifts = CI.Shell_centerRedshift
print(redshifts)
#intrinsic merger rate of all systems at redshift of interest
# say I want to know it in the 10th shell which is centered around
#central redshift in the shell
print(CI.Shell_centerRedshift[10])
TotalIntrinsicRateAtSHell = np.sum(CI.PerSystemPerRedshift_ratesIntrinsic[10])
print(TotalIntrinsicRateAtSHell)
# +
# Plot the intrinsic rate in shell
z = CI.Shell_centerRedshift
#Now use numpy to just collapse all values
rate = np.sum(CI.PerSystemPerRedshift_ratesIntrinsic, axis=1)
fig, axes = plt.subplots(1,1)
axes.plot(z, rate)
axes.set_xlabel('redshift')
axes.set_ylabel('rate Intrinsic [dN/dGpc3/dYr]')
# -
# ### 3.2 Example mask rate by mass, and observed rate
#
# The columns in the 2D array relate to the individual systems. They are ordered the same as the systems in the COMPAS class, so we can use the parameters in the COMPAS class to create a mask for our result! (hence the lazy data).
#
# A detector looks at all redshifts simultaneously, so we sum the rate of each system from every shell in the detector frame (i.e. collapse to an array whose length is the number of systems).
ratePerSystemObserved = np.sum(CI.PerSystemPerRedshift_ratesObserved, axis=0)
print(len(ratePerSystemObserved))
# +
# plot chirp mass distribution
chirpMass = CI.COMPAS.mChirp
#for every chirpmass I know the rate at which you observe
ratePerSystemObserved = np.sum(CI.PerSystemPerRedshift_ratesObserved, axis=0)
#I only want systems with chirp masses below 10 Msun (for some reason)
mask = chirpMass < 10
binsMchirp = np.linspace(0,15,20)
dMchirp = np.diff(binsMchirp)
center = (binsMchirp[1:]+binsMchirp[:-1])/2.
yvalues, _ = np.histogram(chirpMass[mask], bins=binsMchirp, \
weights=ratePerSystemObserved[mask])
dydMchirp = np.divide(yvalues, dMchirp)
fig, axes = plt.subplots(1,1)
axes.plot(center, dydMchirp)
axes.set_xlabel('Mchirp Msun')
axes.set_ylabel('rate detect dN/dyr/dMchirp')
plt.show()
# -
# ## 4. Loop over the cosmic integrator
#
# Once the class is set, you could quickly change prescription or DCO type and recalculate without having to redo the entire instance. This especially saves time in terms of reading/setting the COMPAS class data,and preparing the 2D arrays in the CosmicIntegrator class. The exception being if you loop over the data or change the redshift shells. We provide an example in both cases.
# +
# loop over prescriptions using the already defined class
#data and redshift shells remain unchanged
SFRprescriptions = ['Madau et al. (2014)',\
'Madau et al. (2017)',\
'Strolger et al. (2004)',\
'Neijssel et al. (2019)']
for SFRprescription in SFRprescriptions:
#change SFR
CI.MSSFR.SFRprescription = SFRprescription
#calculate the result
CI.cosmologicalIntegration()
#print the total observed rate
observedRate = np.round(np.sum(CI.PerSystemPerRedshift_ratesObserved))
print('total predicted rate for %s for SFR=%s mergers per year' \
%(CI.GWdetector_sensitivity, observedRate))
# +
# loop over prescriptions using the already defined class
#careful the SFR is still the one of the previous cell!
#loop over DCO types changing the Data
#This takes longer
types = ['BBH', 'BHNS', 'BNS']
for t in types:
# which DCOs I want
CI.COMPAS.setCOMPASDCOmask(types=t, pessimistic=True)
# Get me the data
CI.COMPAS.setCOMPASData()
#the number of systems and their delay times have changed
#reset arrays results. Note that this goes considerably faster
#then the first time we do this, because we dont recalculate
#the redshift table if it is already done :) :) :)
CI.setBirthTimesAnd2Darrays()
#calculate
CI.cosmologicalIntegration()
#print the total observed rate
observedRate = np.round(np.sum(CI.PerSystemPerRedshift_ratesObserved))
print('------total predicted rate for %s =%s mergers per year' \
%(t, observedRate))
# -
# Note that his means if you want to loop over both DCO types and SFR prescriptions, it is more time-efficient to loop over types first and then SFR prescriptions.
#
# With this you should be all set - good luck!
#
# For more information on how to create plots using this pipeline see the plotting library in the postProcessing notes.
#
#
|
docs/online-docs/notebooks/cosmicIntegration/5_MergersAsFunctionOfRedhisft.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %pylab inline
import numpy as np
import pandas as pd
import seaborn as sns
sns.set_style('ticks')
sns.set_context('paper')
import warnings
warnings.filterwarnings('ignore')
# -
df = pd.read_csv('superfolders_ALL.csv')
# +
subset_df = df.loc[df['MainPlot']==1]
sns.set_context('poster')
figure(figsize=(20,9))
metrics = ['DegScore_half_life', 'DegScore_PSU_half_life', 'AUP init 14', 'AUP', 'dG(MFE)', 'CAI']
metric_labels = ['Est. half-life', 'Est. half-life (PSU)', r'$AUP_{init 14}$', r'$AUP_{CDS}$', 'dG(MFE) (kcal/mol)', 'CAI']
for i, metric in enumerate(metrics):
subplot(1,6,i+1)
sns.barplot(y='shortName', x=metric, data=subset_df, dodge=False, color='grey')
title(metric_labels[i])
xlabel(metric_labels[i])
legend([],frameon=False)
if metric=='AUP_init':
xlim([0,1])
if i!=0:
yticks([])
ylabel('')
savefig('../assets/results_barplot_24Dec2021.png',dpi=300, bbox_inches='tight')
# +
def distinct_scatter(df, x='DegScore_half_life', y='AUP init 14', hue='Source', hue_list=['Eterna','Control','Eterna-PSU'], color_list=['Blues','Oranges','Purples']):
assert len(hue_list) == len(color_list)
for typ, color in list(zip(hue_list, color_list)):
tmp = df.loc[df[hue]==typ]
# tmp['_sort'] = tmp[x]*tmp[y]
# tmp = tmp.sort_values('_sort',ascending=False)
palette = sns.color_palette(color,len(tmp))
sns.scatterplot(x=x, y=y, data=tmp, style='Designer', hue='Designer', palette=palette, linewidth=0.5, edgecolor='k', s=100)
ylabel('AUP init 14: increase translation')
ylim([0,1])
legend(bbox_to_anchor=(1,1),frameon=False)
# removed duplicates, need to duplicate controls for third plot
sns.set_context('paper')
figure(figsize=(4,25))
subplot(5,1,1)
distinct_scatter(df.loc[df.Variant=='S-2P'], x='DegScore_half_life', y='AUP init 14', hue_list=['Eterna','Control',], color_list=['rainbow_r','Greys',])
title('WT S-2P mRNAs')
subplot(5,1,2)
distinct_scatter(df.loc[df.Variant=='S-2P, Delta'][df.Nucleotide=='Unmod'], x='DegScore_half_life', y='AUP init 14',hue_list=['Eterna','Control',], color_list=['flare','Greys',])
title('Delta mRNAs')
subplot(5,1,3)
distinct_scatter(df.loc[df.Variant=='S-2P, Delta'][df.Nucleotide=='PSU'], x='DegScore_PSU_half_life', y='AUP init 14',hue_list=['Eterna-PSU','Control',], color_list=['crest','Greys'])
title('Delta PSU-stabilized mRNAs')
subplot(5,1,4)
distinct_scatter(df.loc[df.Variant=='S-2P, Omicron'][df.Nucleotide=='Unmod'], x='DegScore_half_life', y='AUP init 14',hue_list=['Eterna','Control',], color_list=['rocket','Greys',])
title('Omicron mRNAs')
subplot(5,1,5)
distinct_scatter(df.loc[df.Variant=='S-2P, Omicron'][df.Nucleotide=='PSU'], x='DegScore_PSU_half_life', y='AUP init 14',hue_list=['Eterna-PSU','Control',], color_list=['mako','Greys'])
title('Omicron PSU-stabilized mRNAs')
tight_layout()
savefig('../assets/result_scatterplots_24Dec2021.png', dpi=300, bbox_inches='tight')
savefig('../assets/result_scatterplots_24Dec2021.pdf', bbox_inches='tight')
# -
|
analysis/analysis_plotting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/cowiety/energy-forecast/blob/main/Data_cleaning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Ud7v703LISY2"
# # OpenWeather API Script
#
# + [markdown] id="XiFwoe6PIt6s"
# Import statements
# + id="MWPHtbTmhOTT"
import requests
from tkinter import *
import math
import time
import csv
import pandas as pd
# + [markdown] id="AKkcJw1SIytX"
# Global variable declaration
# + id="IbqEkWBCIwdY"
cities = {
"Toronto": [43.7001, -79.4163, 5429524],
"Ottawa": [45.4112, -75.6981, 989567],
"Hamilton": [43.2334, -79.9496, 693645],
"Kitchener": [43.4254, -80.5112, 470015],
"London": [42.9834, -81.233, 383437],
"Oshawa": [43.9001, -78.8496, 308875],
"Windsor": [42.3001, -83.0165, 287069],
"<NAME>": [43.1668, -79.2496, 229246],
"Barrie": [44.4001, -79.6663, 145614],
"Guelph": [43.5501, -80.2497, 132397],
"Kingston": [44.2298, -76.481, 117660]
}
total_pop = 9187049
api_key = "ffe15c3fc982bc7ae70c6edfa2fc779c"
# + [markdown] id="ZVsBFKUHTs1C"
# Get data for time (hour), cloudiness, temperature, humidity and wind speed for each hour of the day
# + id="leJ2dro4OTQJ"
def get_daily_weather(city):
# format city dictionary into API parameters
lat = cities[city][0]
lon = cities[city][1]
url = f"https://api.openweathermap.org/data/2.5/onecall?lat={lat}&lon={lon}&exclude=current,minutely,daily,alerts&appid={api_key}"
# pulls API request and converts to json
response = requests.get(url).json()
# initialize 4 empty lists
hours, clouds, temp, humidity, w_speed = [], [], [], [], []
# looping through next 24 hours of weather data
for i in range(0,24):
hour = time.gmtime(int(response['hourly'][i]['dt'] + response['timezone_offset']))[3]
hours.append(hour)
cloudiness = response['hourly'][i]['clouds']
clouds.append(cloudiness)
temperature = "{:.2f}".format(response['hourly'][i]['temp'] - 273.15)
temp.append(float(temperature))
humidex = response['hourly'][i]['humidity']
humidity.append(humidex)
wind = response['hourly'][i]['wind_speed']
w_speed.append(wind)
return hours, clouds, temp, humidity, w_speed
# + [markdown] id="DqHAkgVRP8zm"
# Collect the data for each city and split the data into 4 arrays by feature
# + id="n23s0zWJsVRf"
# Create file or overwrite if exists
open('results.csv', 'w+')
weighted_clouds, weighted_temp, weighted_humidity, weighted_w_speed = [], [], [], []
for city in cities:
hours, clouds, temp, humidity, w_speed = get_daily_weather(city)
# weight each city's data point according to the population of that city
clouds = [element * cities[city][2] for element in clouds]
temp = [element * cities[city][2] for element in temp]
humidity = [element * cities[city][2] for element in humidity]
w_speed = [element * cities[city][2] for element in w_speed]
# concatenate the data from each city into individual arrays for data processing later
weighted_clouds = weighted_clouds + clouds
weighted_temp = weighted_temp + temp
weighted_humidity = weighted_humidity + humidity
weighted_w_speed = weighted_w_speed + w_speed
# + [markdown] id="vBvCwnIpQTl7"
# Calculate the weighted average for each hour of the day and export to csv
# + colab={"base_uri": "https://localhost:8080/"} id="jAB3ai-K52hX" outputId="9057a5a0-a35d-4232-d2e4-2e4cfd6f2966"
# initialize 4 empty lists
ave_clouds, ave_temp, ave_humidity, ave_w_speed = [0]*24, [0]*24, [0]*24, [0]*24
# loop through the concatenated array for each hour of the day
for i in range(len(hours)):
# loop through the same concatenated array for each city
for j in range(len(cities)):
# since there are 24 hours x 11 cities in each list
# [j*24 + i - 1] collects all data for the ith hour
ave_clouds[i] = ave_clouds[i] + weighted_clouds[j*24 + i - 1]
ave_temp[i] = ave_temp[i] + weighted_temp[j*24 + i - 1]
ave_humidity[i] = ave_humidity[i] + weighted_humidity[j*24 + i - 1]
ave_w_speed[i] = ave_w_speed[i] + weighted_w_speed[j*24 + i - 1]
# format the temperature and wind speed to 3 decimal places
formatter = "{0:.3f}"
# format the data appropriately and divide each element by the total population
ave_clouds = [round(element / total_pop) for element in ave_clouds]
ave_temp = [formatter.format(element / total_pop) for element in ave_temp]
ave_humidity = [round(element / total_pop) for element in ave_humidity]
ave_w_speed = [formatter.format(element / total_pop) for element in ave_w_speed]
# format all the data into a new DataFrame
data = {'Hour': hours,
'Cloudiness (%)': ave_clouds,
'Temp (°C)': ave_temp,
'Humidity (%)': ave_humidity,
'Wind Speed (m/s)': ave_w_speed
}
df = pd.DataFrame(data)
# convert DataFrame to csv file and print to terminal
df.to_csv('results.csv', index=False)
print(df)
# + [markdown] id="Fs7a7PnbgmFJ"
# # Creating Holiday CSV File
# + [markdown] id="JOK4ajJFg3lx"
# import statements
# + id="Dvl8D5NRgt-o"
import pandas as pd #data processing, CSV I/O
import holidays
from datetime import date
# + [markdown] id="jtNZXVBo_u68"
# load holiday library into object variable, import calendar.csv into dataframe
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="vfXAZ0rZg6Et" outputId="7e3c6b16-468a-44bf-a254-e2ad83ceb898"
can_holidays = holidays.Canada()
df = pd.read_csv('calendar.csv')
df.head(10)
# + [markdown] id="JBF-toTwP5Pp"
# add a column to dataframe called 'isHoliday'
# + id="FRlWLqrohfyG"
df['isHoliday'] = range(len(df))
# + [markdown] id="bGOLC1KCP_fK"
# check if each date is a holiday and place the result in the isHoliday column
# + id="YW2qiygiP4Ky"
for row in range(len(df)-1):
df['isHoliday'][row] = date(df['year'][row], df['month'][row], df['day'][row]) in can_holidays
df.head(10)
# + [markdown] id="12wGeEGOQIw_"
# export dataframe to csv
# + id="B_H_5f2kQa97"
df.to_csv('holidays.csv', index=False)
# + [markdown] id="jwQzXzYFsFWJ"
# # Cleaning IESO Demand Data
# + [markdown] id="55T6633uGt7x"
# Produces an up-to-date csv file of Ontario hourly demand data from the previous year
# + id="CJMCGW_RsLBT"
import pandas as pd #data processing, CSV I/O
import datetime
# + [markdown] id="37yinnJMGlYt"
# Downloads CSVs from IESO website for the current and prior year
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="QItQH9kdsOW8" outputId="5b3960b1-60a8-402f-fdd3-a88dcba0da7a"
current_year = date.today().year
last_year = current_year - 1
last_year_link = 'http://reports.ieso.ca/public/Demand/PUB_Demand_{}.csv'.format(last_year)
current_year_link = 'http://reports.ieso.ca/public/Demand/PUB_Demand_{}.csv'.format(current_year)
df_1 = pd.read_csv(last_year_link, skiprows=3)
df_2 = pd.read_csv(current_year_link, skiprows=3)
df_2.head(10)
# + [markdown] id="evPgPm60G864"
# Function splits the date column into year, month and day
# + id="5BXNfDTZxr0G"
def reformat_df(df):
df['Year'] = df.Date.apply(lambda x: int(x[:4]))
df['Month'] = df.Date.apply(lambda x: int(x[5:7]))
df['Day'] = df.Date.apply(lambda x: int(x[8:]))
df['Demand'] = df['Market Demand']
df.drop(['Ontario Demand', 'Market Demand', 'Date'], axis=1, inplace = True)
# + [markdown] id="iZb-go-oHDJ6"
# Apply the previous function and remove rows that are earlier than one year ago
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="udUH9yVT1-Ns" outputId="4dd9e9ae-b1be-4157-e840-192c1d51595d"
reformat_df(df_1)
reformat_df(df_2)
first_index = df_1[(df_1.Day == date.today().day) & (df_1.Hour == (datetime.datetime.now().hour - 5)) & (df_1.Month == date.today().month)].index[0]
df_1.drop(list(range(first_index)), inplace = True)
full_year = df_1.append(df_2, ignore_index = True)
full_year.head(10)
# + [markdown] id="ULRYw3WMHOWI"
# Reformat the date columns into ISO formatting
# + colab={"base_uri": "https://localhost:8080/", "height": 833} id="uc0BzcD93l3C" outputId="c3a83787-36f3-4ce8-ad61-30ef6df5733f"
full_year['Date'] = full_year.apply(lambda row: '{:04d}-{:02d}-{:02d}'.format(row['Year'], row['Month'], row['Day']), axis = 1)
full_year.drop(['Day', 'Year', 'Month'], inplace = True, axis = 1)
full_year.head(10)
# + [markdown] id="VLuWYHDPHTcV"
# Export the dataframe into a csv file named with the current date
# + id="raxYyUzwFDmD"
full_year.to_csv('Demand Data ({:04d}-{:02d}-{:02d}).csv'.format(date.today().year, date.today().month, date.today().day), index = False)
|
Data_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Categorical variables**
#
# 1. **Nominal**(order not follows) (ex: male,female)
# 2. **Ordinal**(order follows) (ex: degree, masters, phd)
import pandas as pd
df = pd.read_csv("https://raw.githubusercontent.com/patilankita79/ML_DataPreprocessing/master/ML_DataPreprocessing/Data.csv")
df
dummies = pd.get_dummies(df['Country'])
dummies
merged = pd.concat([df,dummies], axis='columns')
merged
#no need one columns in dummies
final_df = merged.drop(['Country','France'],axis='columns')
final_df
# **Label encoding**
df
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
|
code-basics-dummy-variables-one-hot-encoding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pytorch_tabnet.tab_model import TabNetClassifier
import torch
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import roc_auc_score, accuracy_score
import pandas as pd
import numpy as np
np.random.seed(0)
import os
import wget
from pathlib import Path
from datasets import load_dataset
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# # Download census-income dataset
dataset_name = 'dataset_players_statistics'
out = Path(os.getcwd()+'/data/'+dataset_name+'.csv')
# # Load data and split
# +
train = pd.read_csv(out)
#train = train.drop(['game'], axis=1)
target = 'result'
if "Set" not in train.columns:
train["Set"] = np.random.choice(["train", "valid", "test"], p =[.8, .1, .1], size=(train.shape[0],))
train_indices = train[train.Set=="train"].index
valid_indices = train[train.Set=="valid"].index
test_indices = train[train.Set=="test"].index
train
# + [markdown] tags=[]
# # Simple preprocessing
#
# Label encode categorical features and fill empty cells.
# +
nunique = train.nunique()
types = train.dtypes
categorical_columns = []
categorical_dims = {}
for col in train.columns:
if types[col] == 'object' or nunique[col] < 200:
print(col, train[col].nunique())
l_enc = LabelEncoder()
train[col] = train[col].fillna("VV_likely")
train[col] = l_enc.fit_transform(train[col].values)
categorical_columns.append(col)
categorical_dims[col] = len(l_enc.classes_)
else:
train.fillna(train.loc[train_indices, col].mean(), inplace=True)
# -
categorical_columns
categorical_dims
# check that pipeline accepts strings
train.loc[train[target]==0, target] = "lose"
train.loc[train[target]==1, target] = "win"
# # Define categorical features for categorical embeddings
# +
unused_feat = ['Set']
features = [ col for col in train.columns if col not in unused_feat+[target]]
cat_idxs = [ i for i, f in enumerate(features) if f in categorical_columns]
cat_dims = [ categorical_dims[f] for i, f in enumerate(features) if f in categorical_columns]
# -
# # Network parameters
# +
tabnet_params = {"cat_idxs":cat_idxs,
"cat_dims":cat_dims,
"cat_emb_dim":1,
"optimizer_fn":torch.optim.Adam,
"optimizer_params":dict(lr=2e-2),
"scheduler_params":{"step_size":10, # how to use learning rate scheduler
"gamma":0.9},
"scheduler_fn":torch.optim.lr_scheduler.StepLR,
"mask_type":'sparsemax' # "sparsemax, entmax"
}
clf = TabNetClassifier(**tabnet_params
)
# -
# # Training
# +
X_train = train[features].values[train_indices]
y_train = train[target].values[train_indices]
X_valid = train[features].values[valid_indices]
y_valid = train[target].values[valid_indices]
X_test = train[features].values[test_indices]
y_test = train[target].values[test_indices]
# -
max_epochs = 1000 if not os.getenv("CI", False) else 2
# + tags=[]
# This illustrates the warm_start=False behaviour
save_history = []
for _ in range(2):
clf.fit(
X_train=X_train, y_train=y_train,
eval_set=[(X_train, y_train), (X_valid, y_valid)],
eval_name=['train', 'valid'],
eval_metric=['accuracy'],
max_epochs=max_epochs , patience=50,
batch_size=1024, virtual_batch_size=128,
num_workers=0,
weights=1,
drop_last=False
)
save_history.append(clf.history["valid_accuracy"])
assert(np.all(np.array(save_history[0]==np.array(save_history[1]))))
# -
# plot losses
plt.plot(clf.history['loss'])
# plot auc
plt.plot(clf.history['train_accuracy'])
plt.plot(clf.history['valid_accuracy'])
# plot learning rates
plt.plot(clf.history['lr'])
# ## Predictions
# +
preds = clf.predict_proba(X_test)
test_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_test ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
preds_valid = clf.predict_proba(X_valid)
valid_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_valid ], y_pred=[0 if x < 0.5 else 1 for x in preds_valid[:,1]])
print(f"BEST VALID SCORE FOR {dataset_name} : {clf.best_cost}")
print(f"FINAL TEST SCORE FOR {dataset_name} : {test_acc}")
# -
# check that best weights are used
assert np.isclose(valid_acc, np.max(clf.history['valid_accuracy']), atol=1e-6)
clf.predict(X_test)
# # Save and load Model
# save tabnet model
saving_path_name = "./tabnet_model_test_1"
saved_filepath = clf.save_model(saving_path_name)
# define new model with basic parameters and load state dict weights
loaded_clf = TabNetClassifier()
loaded_clf.load_model(saved_filepath)
loaded_preds = loaded_clf.predict_proba(X_test)
loaded_test_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_test ], y_pred=[0 if x < 0.5 else 1 for x in loaded_preds[:,1]])
print(f"FINAL TEST SCORE FOR {dataset_name} : {loaded_test_acc}")
assert(test_acc == loaded_test_acc)
loaded_clf.predict(X_test)
# # Global explainability : feat importance summing to 1
clf.feature_importances_.shape
# # Local explainability and masks
explain_matrix, masks = clf.explain(X_test)
# +
fig, axs = plt.subplots(1, 3, figsize=(20,20))
for i in range(3):
axs[i].imshow(masks[i][:50])
axs[i].set_title(f"mask {i}")
# -
# # XGB
# +
#pip install xgboost
# +
from xgboost import XGBClassifier
import xgboost as xgb
import matplotlib.pyplot as plt
clf_xgb = XGBClassifier(max_depth=8,
learning_rate=0.1,
n_estimators=1000,
verbosity=0,
silent=None,
objective='binary:logistic',
booster='gbtree',
n_jobs=-1,
nthread=None,
gamma=0,
min_child_weight=1,
max_delta_step=0,
subsample=0.7,
colsample_bytree=1,
colsample_bylevel=1,
colsample_bynode=1,
reg_alpha=0,
reg_lambda=1,
scale_pos_weight=1,
base_score=0.5,
random_state=0,
seed=None,)
clf_xgb.fit(X_train, y_train,
eval_set=[(X_valid, y_valid)],
early_stopping_rounds=200,
verbose=10)
# -
importances = clf_xgb.feature_importances_
features = list(train.columns[:-2])
importances_zip = zip(importances,features)
sorted_importances_zip = sorted(importances_zip)
fig, ax = plt.subplots(1,1, figsize=(12,28))
importances, features = zip(*sorted_importances_zip)
ax.barh(features, importances)
# +
preds = np.array(clf_xgb.predict_proba(X_valid))
valid_acc = accuracy_score(y_true= [0 if x =='lose' else 1 for x in y_valid ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(valid_acc)
preds = np.array(clf_xgb.predict_proba(X_test))
test_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_test ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(test_acc)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn import metrics
# 학습 진행
forest = RandomForestClassifier(n_estimators=1000, min_samples_leaf=3, min_samples_split=10, max_features=7)
forest.fit(X_train, y_train)
preds = np.array(forest.predict_proba(X_valid))
valid_acc = accuracy_score(y_true= [0 if x =='lose' else 1 for x in y_valid ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(valid_acc)
preds = np.array(forest.predict_proba(X_test))
test_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_test ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(test_acc)
# +
from sklearn.linear_model import LogisticRegression
logistic = LogisticRegression(max_iter=500)
logistic.fit(X_train, y_train)
preds = np.array(logistic.predict_proba(X_valid))
valid_acc = accuracy_score(y_true= [0 if x =='lose' else 1 for x in y_valid ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(valid_acc)
preds = np.array(logistic.predict_proba(X_test))
test_acc = accuracy_score(y_true=[0 if x =='lose' else 1 for x in y_test ], y_pred=[0 if x < 0.5 else 1 for x in preds[:,1]])
print(test_acc)
# -
|
other_feature_lol_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from tpot import TPOTRegressor
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import metrics
import pandas as pd
import seaborn as sns
import numpy as np
import datetime
# data
acidoCEST_ML = pd.read_csv('acido_CEST_MRI_MegaBox_01_to_08_clean.csv')
acidoCEST_ML = acidoCEST_ML.drop(['Unnamed: 0','ApproT1(sec)','Temp','FILE','Conc(mM)'], axis = 1)
print(acidoCEST_ML.shape)
acidoCEST_ML.iloc[20146,:].head(10)
acidoCEST_ML.iloc[20146,:]
Z = acidoCEST_ML.iloc[:,9::]
Z.shape
import h2o
from h2o.estimators import H2OGeneralizedLowRankEstimator
h2o.init()
data = h2o.H2OFrame(Z)
# Split the dataset into a train and valid set:
train, valid = data.split_frame(ratios=[.8], seed=1234)
train.shape
valid.shape
glrm_model = H2OGeneralizedLowRankEstimator(k=5,
loss="quadratic",
gamma_x=0.5,
gamma_y=0.5,
max_iterations=700,
recover_svd=True,
init="SVD",
transform="standardize")
glrm_model.train(training_frame=train)
glrm_model
glrm_model.scoring_history().set_index('iterations')['objective'].plot()
import plotly.express as px
len(Z.columns.tolist())
Y = glrm_model._model_json["output"]['archetypes'].as_data_frame()
X = Z @ np.linalg.pinv(Y.iloc[:,1::].values)
Y
# +
x= Y.iloc[2,1::]
y= Y.iloc[1,1::]
df = pd.DataFrame()
df['x'] = x
df['y'] = y
df['color']= [ float(q) for q in acidoCEST_ML.iloc[:,9::].columns.tolist() ]
df['h']= ['sat @ ' + str(q) + 'ppm' for q in Z.columns ]
df['size'] = 12
fig = px.scatter(df ,x='x',y='y',color='color', hover_data='h', size='size' )
fig.write_html("./CEST_projections_Y_freqs.html")
# -
from scipy.spatial.distance import pdist
from scipy.spatial import distance_matrix
# +
f, ax = plt.subplots(dpi = 200)
w = acidoCEST_ML.iloc[:,9::].columns
D = pd.DataFrame( distance_matrix( Y.iloc[:,1::].T, Y.iloc[:,1::].T), index = w, columns= w)
sns.heatmap(D,ax=ax)
# -
fig = px.imshow(D)
fig.show()
fig.write_html("./HEAT.html")
# +
df = pd.DataFrame()
df['x'] = X[0]
df['y'] = X[1]
df['size'] = 12
df['sample'] = np.arange(df.x.shape[0])
fig = px.scatter(df ,x='x',y='y', hover_name='sample')
fig.write_html("./CEST_projections_X_samples.html")
# -
from h2o.estimators import H2OKMeansEstimator
# +
# Split the dataset into a train and valid set:
train2, valid2 = h2o.H2OFrame(X).split_frame(ratios=[.8], seed=1234)
# Build and train the model:
X_kmeans = H2OKMeansEstimator(k=10,
estimate_k=True,
standardize=False,
seed=1234)
X_kmeans.train(
training_frame=train2,
validation_frame=valid2)
# Eval performance:
perf = X_kmeans.model_performance()
# Generate predictions on a validation set (if necessary):
pred = X_kmeans.predict(valid2)
# -
perf
# +
# Split the dataset into a train and valid set:
train, valid = iris.split_frame(ratios=[.8], seed=1234)
# Build and train the model:
iris_kmeans = H2OKMeansEstimator(k=10,
estimate_k=True,
standardize=False,
seed=1234)
iris_kmeans.train(x=predictors,
training_frame=train,
validation_frame=valid)
# Eval performance:
perf = iris_kmeans.model_performance()
# Generate predictions on a validation set (if necessary):
pred = iris_kmeans.predict(valid)
|
GLRM_CEST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Blood only analysis
# ---
# Evaluate the performance of different models when training on the subset of the data containing only gene expression from blood.
# + language="bash"
# PLIER_OUT='../data/blood_plier'
# SUBSET_OUT='../data/blood_expression'
# mkdir -p $PLIER_OUT
# mkdir -p $SUBSET_OUT
# -
# ## Plot Initial Data and Subset
#
# Plot the healthy data containing all samples, followed by the subset of the data containing only healthy blood samples. These plots should both help visualize (dis)similarity between studies, and get an idea of what the PLIER training data will look like.
from plotnine import *
import pandas
import umap
initial_df = pandas.read_csv('../data/plier_healthy.tsv', sep='\t')
initial_df.iloc[:5,:5]
reducer = umap.UMAP()
embedded_expression = reducer.fit_transform(initial_df.T)
embedded_df = pandas.DataFrame({'x': embedded_expression[:,0], 'y': embedded_expression[:,1]})
embedded_df['sample'] = initial_df.columns
split = embedded_df['sample'].str.split('.', expand=True)
embedded_df['sample'] = split[1]
embedded_df['study'] = split[0]
embedded_df.head()
ggplot(embedded_df, aes(x='x', y='y', color='study')) + \
geom_point(alpha=.8) + \
ggtitle('Healthy gene expression from different studies')
# ## Subset Studies
#
# From the set of all labeled data (output by download_categorized_data.ipynb), extract the subset corresponding to blood (UBERON:0000178).
# + language="bash"
# SUBSET_OUT='../data/blood_expression'
#
# python subset_studies.py ../data/plier_healthy.tsv ../data/plier_disease.tsv \
# ../data/classifier_healthy.tsv ../data/classifier_disease.tsv \
# ${SUBSET_OUT}/blood_plier ${SUBSET_OUT}/blood_classifier \
# UBERON:0000178
# -
blood_df = pandas.read_csv('../data/blood_expression/blood_plier_healthy.tsv', sep='\t')
blood_df.iloc[:5,:5]
reducer = umap.UMAP()
embedded_expression = reducer.fit_transform(blood_df.T)
embedded_df = pandas.DataFrame({'x': embedded_expression[:,0], 'y': embedded_expression[:,1]})
embedded_df['sample'] = blood_df.columns
split = embedded_df['sample'].str.split('.', expand=True)
embedded_df['sample'] = split[1]
embedded_df['study'] = split[0]
embedded_df.head()
ggplot(embedded_df, aes(x='x', y='y', color='study')) + \
geom_point(alpha=.8) + \
ggtitle('Healthy gene expression from blood samples only')
# ## Train PLIER
#
# Train PLIER for dimensionality reduction on a fraction of the studies with blood samples
# + language="bash"
# SUBSET_OUT='../data/blood_expression'
# PLIER_OUT='../data/blood_plier'
#
# # K_VALS stores the different values of k (number of PCs) to be used by PLIER.
# # This array is intentionally reverse sorted for more efficient scheduling (you want
# # the longest jobs to be loaded first)
# K_VALS=(50)
#
# # Pass each K_value to run_plier and execute the different instances in paralell
# # The code before the pipe prints all the values of K on its own line
# # The code after the pipe tells bash to run run_plier.R once for each value of K,
# # But using no more than NUM_PROCESSES threads to do so
# printf "${SUBSET_OUT}/blood_plier_healthy.tsv ${SUBSET_OUT}/blood_plier_disease.tsv $PLIER_OUT -k %s" \
# "${K_VALS[0]}" | xargs -n 5 --max-procs=4 Rscript run_plier.R
# -
# ## Train Discriminator Model
# + language="bash"
# PLIER_OUT='../data/blood_plier'
# SUBSET_OUT='../data/blood_expression'
# python create_identity_matrix.py ${PLIER_OUT}/plier_${K_VALS[0]}_Z.tsv ${PLIER_OUT}/plier_all_Z.tsv
#
# echo "All run_plier instances completed"
#
# python create_identity_matrix_df.py ../data/blood_plier/plier_50_Z.tsv ../data/blood_plier/plier_all_Z.tsv
#
# time python evaluate_models.py $PLIER_OUT ${SUBSET_OUT}/blood_classifier_healthy.tsv \
# ${SUBSET_OUT}/blood_classifier_disease.tsv \
# --out_path ../results/blood_eval_results.csv --epochs 400
# -
import plotnine
plotnine.options.figure_size = (8,6)
# +
# Load the dataframe generated by evaluate_models.py
result_df = pandas.read_csv('../results/blood_eval_results.csv', index_col=0)
result_df['acc_over_baseline'] = result_df['val_acc'] - result_df['val_baseline']
result_df.head()
# -
# ## Plot Performance
ggplot(result_df, aes(x='factor(lv_count)', y='val_acc', color='factor(lv_count)')) +\
geom_boxplot() +\
facet_grid('. ~ Model') +\
theme(axis_text_x=element_text(rotation=90, hjust=0.5)) +\
ggtitle('Tuning Set Accuracy for Blood Samples')
ggplot(result_df, aes(x='factor(lv_count)', y='acc_over_baseline', color='factor(lv_count)')) +\
geom_boxplot() +\
facet_grid('. ~ Model') +\
theme(axis_text_x=element_text(rotation=90, hjust=0.5)) +\
ggtitle('Accuracy Above Constant Baseline for Tuning Set Blood Samples')
ggplot(result_df, aes(x='factor(lv_count)', y='val_auroc', color='factor(lv_count)')) +\
geom_boxplot() +\
facet_grid('. ~ Model') +\
theme(axis_text_x=element_text(rotation=90, hjust=0.5)) +\
ggtitle('Blood Sample Tuning Set Area Under the ROC Curve')
|
brdnet/blood_only_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("/home/jovyan/TF_NEW/tf-transformers/src/")
# -
# +
import os
import tempfile
import json
import glob
import datasets
import shutil
import tensorflow as tf
from hydra import initialize, initialize_config_module, initialize_config_dir, compose
from omegaconf import OmegaConf
from tf_transformers.data import TFReader, TFWriter
from tf_transformers.models import Classification_Model
from tf_transformers.losses import cross_entropy_loss_for_classification
from model import get_model, get_tokenizer, get_optimizer, get_trainer
# -
with initialize(config_path="conf/"):
cfg = compose(config_name="config", overrides=["data.take_sample=true", "+glue=mrpc"])
print(cfg)
# +
# Steps
# 1. Download the data
# 2. Prepare TFRecords
# 3. Read TFrecords to tf.data
# 4. Train the model
# -
def get_classification_model(num_classes, return_all_layer_outputs, is_training, use_dropout):
def model_fn():
model = get_model(return_all_layer_outputs, is_training, use_dropout)
classification_model = Classification_Model(model,
num_classes,
use_all_layers=return_all_layer_outputs,
is_training=is_training,
use_dropout=use_dropout)
classification_model = classification_model.get_model()
return classification_model
return model_fn
# +
# Convert data to features using specific length
# into a temp dir (and log it as well for monitoring)
def get_dataset(data, batch_size, tokenizer, max_seq_length, mode, tfrecord_dir, take_sample=False):
if mode not in ["train", "eval"]:
raise ValueError("Inavlid mode `{}` specified. Available mode is ['train', 'eval']".format(mode))
def get_tfrecord_example(data):
result = {}
for f in data:
input_ids_s1 = [tokenizer.cls_token] + tokenizer.tokenize(f['sentence1'])[: max_seq_length-2] + [tokenizer.sep_token] # -2 to add CLS and SEP
input_ids_s1 = tokenizer.convert_tokens_to_ids(input_ids_s1)
input_type_ids_s1 = [0] * len(input_ids_s1) # 0 for s1
input_ids_s2 = tokenizer.tokenize(f['sentence2'])[: max_seq_length-1] + [tokenizer.sep_token] # -1 to add SEP
input_ids_s2 = tokenizer.convert_tokens_to_ids(input_ids_s2)
input_type_ids_s2 = [1] * len(input_ids_s2)
# concatanate two sentences
input_ids = input_ids_s1 + input_ids_s2
input_type_ids = input_type_ids_s1 + input_type_ids_s2
input_mask = [1] * len(input_ids) # 1 for s2
result = {}
result['input_ids'] = input_ids
result['input_mask'] = input_mask
result['input_type_ids'] = input_type_ids
result['labels'] = f['label']
yield result
schema = {
"input_ids": ("var_len", "int"),
"input_mask": ("var_len", "int"),
"input_type_ids": ("var_len", "int"),
"labels": ("var_len", "int"),
}
# Create a temp dir
if mode == "train":
# Write tf records
train_data_dir = os.path.join(tfrecord_dir,"train")
tfrecord_filename = 'mrpc'
tfwriter = TFWriter(schema=schema,
file_name=tfrecord_filename,
model_dir=train_data_dir,
tag='train',
overwrite=False
)
data_train = data['train']
# Take sample
if take_sample:
data_train = data_train.select(range(500))
tfwriter.process(parse_fn=get_tfrecord_example(data_train))
# Read tfrecord to dataset
schema = json.load(open("{}/schema.json".format(train_data_dir)))
stats = json.load(open('{}/stats.json'.format(train_data_dir)))
all_files = glob.glob("{}/*.tfrecord".format(train_data_dir))
tf_reader = TFReader(schema=schema,
tfrecord_files=all_files)
x_keys = ['input_ids', 'input_type_ids', 'input_mask']
y_keys = ['labels']
train_dataset = tf_reader.read_record(auto_batch=True,
keys=x_keys,
batch_size=batch_size,
x_keys = x_keys,
y_keys = y_keys,
shuffle=True,
drop_remainder=True
)
return train_dataset, stats['total_records']
if mode == "eval":
# Write tfrecords
eval_data_dir = os.path.join(tfrecord_dir,"eval")
tfrecord_filename = 'mrpc'
tfwriter = TFWriter(schema=schema,
file_name=tfrecord_filename,
model_dir=eval_data_dir,
tag='dev',
overwrite=False
)
data_eval = data['validation']
# Take sample
if take_sample:
data_eval = data_eval.select(range(500))
tfwriter.process(parse_fn=get_tfrecord_example(data_eval))
# Read tfrecord to dataset
schema = json.load(open("{}/schema.json".format(eval_data_dir)))
stats = json.load(open('{}/stats.json'.format(eval_data_dir)))
all_files = glob.glob("{}/*.tfrecord".format(eval_data_dir))
tf_reader = TFReader(schema=schema,
tfrecord_files=all_files)
x_keys = ['input_ids', 'input_type_ids', 'input_mask']
y_keys = ['labels']
eval_dataset = tf_reader.read_record(auto_batch=True,
keys=x_keys,
batch_size=batch_size,
x_keys = x_keys,
y_keys = y_keys,
shuffle=False,
drop_remainder=False
)
return eval_dataset, stats['total_records']
# -
def get_loss(loss_type):
if loss_type and loss_type == 'joint':
def loss_fn(y_true_dict, y_pred_dict):
"""Joint loss over all layers"""
loss_dict = {}
loss_holder = []
for layer_count, per_layer_output in enumerate(y_pred_dict['class_logits']):
loss = cross_entropy_loss_for_classification(
labels=tf.squeeze(y_true_dict['labels'], axis=1),
logits=per_layer_output
)
loss_dict['loss_{}'.format(layer_count + 1)] = loss
loss_holder.append(loss)
# Mean over batch
loss_dict['loss'] = tf.reduce_mean(loss_holder, axis=0)
return loss_dict
else:
def loss_fn(y_true_dict, y_pred_dict):
"""last layer loss"""
loss_dict = {}
loss = cross_entropy_loss_for_classification(
labels=tf.squeeze(y_true_dict['labels'], axis=1),
logits=y_pred_dict['class_logits']
)
loss_dict['loss'] = loss
return loss_dict
return loss_fn
cfg
# +
# Data specific configuration
max_seq_len = cfg.data.max_seq_length
take_sample = cfg.data.take_sample
max_seq_length = cfg.data.max_seq_length
train_batch_size = cfg.data.train_batch_size
eval_batch_size = cfg.data.eval_batch_size
# Trainer specifics
device = cfg.trainer.type
num_gpus = cfg.trainer.num_gpus
tpu_address = cfg.trainer.tpu_address
dtype = cfg.trainer.dtype
epochs = cfg.trainer.epochs
strategy = cfg.trainer.strategy
# Optimizer
learning_rate = cfg.optimizer.learning_rate
loss_type = cfg.optimizer.loss_type
return_all_layer_outputs = False
if loss_type and loss_type == 'joint':
return_all_layer_outputs = True
# Core data specifics
data_name = cfg_task.glue.data.name
num_classes = cfg_task.glue.data.num_classes
# Model specific
is_training = cfg.model.is_training
use_dropout = cfg.model.use_dropout
# -
# +
# Load tokenizer
tokenizer = get_tokenizer()
# Load data
data = datasets.load_dataset("glue", data_name)
tfrecord_dir = tempfile.mkdtemp()
train_dataset, total_train_examples = get_dataset(data, train_batch_size,tokenizer, max_seq_length, "train", tfrecord_dir, take_sample)
eval_dataset, total_eval_examples = get_dataset(data, eval_batch_size,tokenizer, max_seq_len, "eval", tfrecord_dir, take_sample)
# +
# Load optimizer
optimizer_fn = get_optimizer(learning_rate, total_train_examples, train_batch_size, epochs)
# Load trainer
# trainer = get_trainer(device, dtype, strategy, num_gpus, tpu_address)
# -
# Load model function
model_fn = get_classification_model(num_classes,
return_all_layer_outputs, is_training, use_dropout)
# Load loss function
train_loss_fn = get_loss(loss_type)
for (batch_inputs, batch_labels) in train_dataset.take(1):
print(batch_inputs['input_ids'].shape, batch_labels['labels'].shape)
class Callback():
def __init__(self):
pass
def call(trainer_kwargs):
for k, v in trainer_kwargs.items():
print(k, '-->', v)
callback = Callback()
# +
# coding=utf-8
# Copyright 2021 TF-Transformers Authors.
# All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
import tensorflow as tf
import tqdm
from absl import logging
from tf_transformers.core import keras_utils
from tf_transformers.core.distribute_utils import get_distribution_strategy
from tf_transformers.core.performance_utils import (
configure_optimizer,
get_tf_dtype,
is_float16,
set_mixed_precision_policy,
)
def flat_metric_dict(metric_dict):
"""Flatten the dict"""
dict_flatten = {}
dict_flatten['steps'] = list(metric_dict.keys())
for _key, value in metric_dict.items():
for sub_key, sub_value in value.items():
if sub_key not in dict_flatten:
dict_flatten[sub_key] = [sub_value]
else:
dict_flatten[sub_key].append(sub_value)
return dict_flatten
def save_model_checkpoints(model, overwrite_checkpoint_dir, model_checkpoint_dir, max_number_of_models):
# Model checkpoint
if not overwrite_checkpoint_dir:
import os
if os.path.exists(model_checkpoint_dir):
raise FileExistsError("Model directory exists")
checkpoint = tf.train.Checkpoint(model=model)
manager = tf.train.CheckpointManager(checkpoint, directory=model_checkpoint_dir, max_to_keep=max_number_of_models)
return manager
def get_loss_metric_dict(training_loss_names, validation_loss_names):
training_loss_dict_metric = {name: tf.keras.metrics.Mean(name, dtype=tf.float32) for name in training_loss_names}
training_loss_dict_metric["learning_rate"] = tf.keras.metrics.Mean(
"learning_rate", dtype=tf.float32
) # We store learning rate here and reset after every global steps
validation_loss_dict_metric = {}
if validation_loss_names:
validation_loss_dict_metric = {
name: tf.keras.metrics.Mean(name, dtype=tf.float32) for name in validation_loss_names
}
return training_loss_dict_metric, validation_loss_dict_metric
def get_and_reset_metric_from_dict(metric_dict):
if not metric_dict:
return {}
metric_result = {name: metric.result().numpy() for name, metric in metric_dict.items()}
for _name, metric in metric_dict.items():
metric.reset_states()
return metric_result
def get_tensorboard_writers(model_checkpoint_dir):
train_log_dir = model_checkpoint_dir + "/logs/train"
test_log_dir = model_checkpoint_dir + "/logs/dev"
train_summary_writer = tf.summary.create_file_writer(train_log_dir)
test_summary_writer = tf.summary.create_file_writer(test_log_dir)
return train_summary_writer, test_summary_writer
def train_and_eval(
model,
optimizer,
strategy,
epochs,
steps_per_epoch,
steps_per_call,
train_dataset_iter,
train_loss_fn,
GLOBAL_BATCH_SIZE,
training_loss_dict_metric,
validation_dataset_distributed,
validation_loss_fn,
validation_loss_dict_metric,
validation_interval_steps,
mixed_precision,
callbacks,
callbacks_interval_steps,
trainer_kwargs,
checkpoint_manager,
model_checkpoint_dir,
model_save_interval_steps,
):
def save_model(epoch_end=False):
if not epoch_end:
if model_save_interval_steps:
if global_step % model_save_interval_steps == 0:
checkpoint_manager.save()
logging.info("Model saved at step {}".format(global_step))
else:
checkpoint_manager.save()
logging.info("Model saved at epoch {}".format(epoch))
# @tf.function(experimental_relax_shapes=True)
def write_metrics(metric_dict, writer, step):
# @tf.function
def _write(step):
# other model code would go here
with writer.as_default():
for name, result in metric_dict.items():
tf.summary.scalar(name, result, step=step)
_write(step)
writer.flush()
def compute_loss(batch_labels, model_outputs):
"""Loss computation which takes care of loss reduction based on GLOBAL_BATCH_SIZE"""
per_example_loss = train_loss_fn(batch_labels, model_outputs)
per_example_loss_averaged = {}
# Inplace update
# Avergae loss per global batch size , recommended
for name, loss in per_example_loss.items():
per_example_loss_averaged[name] = tf.nn.compute_average_loss(loss, global_batch_size=GLOBAL_BATCH_SIZE)
return per_example_loss_averaged
def compute_loss_valid(batch_labels, model_outputs):
"""Validation Loss computation which takes care of loss reduction based on GLOBAL_BATCH_SIZE"""
per_example_loss = validation_loss_fn(batch_labels, model_outputs)
per_example_loss_averaged = {}
# Inplace update
# Avergae loss per global batch size , recommended
for name, loss in per_example_loss.items():
per_example_loss_averaged[name] = tf.nn.compute_average_loss(loss, global_batch_size=GLOBAL_BATCH_SIZE)
return per_example_loss_averaged
# Train Functions
@tf.function
def do_train(iterator):
"""The step function for one training step"""
def train_step(dist_inputs):
"""The computation to run on each device."""
batch_inputs, batch_labels = dist_inputs
with tf.GradientTape() as tape:
model_outputs = model(batch_inputs)
loss = compute_loss(batch_labels, model_outputs)
tf.debugging.check_numerics(loss['loss'], message='Loss value is either NaN or inf')
if isinstance(optimizer, tf.keras.mixed_precision.LossScaleOptimizer):
loss_scaled = {name: optimizer.get_scaled_loss(loss_value) for name, loss_value in loss.items()}
# TODO
# Scales down the loss for gradients to be invariant from replicas.
# loss = loss / strategy.num_replicas_in_sync
if mixed_precision:
scaled_gradients = tape.gradient(loss_scaled["loss"], model.trainable_variables)
grads = optimizer.get_unscaled_gradients(scaled_gradients)
else:
grads = tape.gradient(loss["loss"], model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# training_loss.update_state(loss * strategy.num_replicas_in_sync)
return loss
for _ in tf.range(tf.convert_to_tensor(steps_per_call)):
dist_inputs = next(iterator)
loss = strategy.run(train_step, args=(dist_inputs,))
# strategy reduce
loss = {
name: strategy.reduce(tf.distribute.ReduceOp.MEAN, loss_value, axis=None)
for name, loss_value in loss.items()
}
for name, loss_value in loss.items():
training_loss = training_loss_dict_metric[name]
training_loss.update_state(loss_value)
# Get current learning rate
if isinstance(optimizer, tf.keras.mixed_precision.LossScaleOptimizer):
current_lr = optimizer._optimizer._decayed_lr(tf.float32)
else:
current_lr = optimizer._decayed_lr(tf.float32)
training_loss_dict_metric["learning_rate"].update_state(current_lr)
# training_result = get_and_reset_metric_from_dict(training_loss_dict_metric)
# do validation
def do_validation(validation_dataset_distributed):
"""Validation step"""
@tf.function
def _validate_step(dist_inputs):
batch_inputs, batch_labels = dist_inputs
model_outputs = model(batch_inputs)
loss = compute_loss_valid(batch_labels, model_outputs)
return loss
if not epoch_end:
if (
validation_dataset_distributed
and validation_loss_fn
and validation_interval_steps
and (global_step % validation_interval_steps == 0)
):
logging.info("Validation in progress at step {} . . . .".format(global_step))
with tqdm.tqdm(validation_dataset_distributed, unit=" Val batch ") as val_batches:
for dist_inputs in val_batches:
loss = strategy.run(_validate_step, args=(dist_inputs,))
for name, loss_value in loss.items():
loss_value = strategy.reduce(tf.distribute.ReduceOp.SUM, loss_value, axis=None)
validation_loss = validation_loss_dict_metric[name]
validation_loss.update_state(loss_value)
validation_result = get_and_reset_metric_from_dict(validation_loss_dict_metric)
validation_history[global_step] = validation_result
write_metrics(validation_result, val_summary_writer, global_step)
logging.info("Validation result at step {}".format(validation_result))
print("\n")
else:
if validation_dataset_distributed and validation_loss_fn:
logging.info("Validation in progress at epoch end {} . . . .".format(epoch))
with tqdm.tqdm(validation_dataset_distributed, unit=" Val batch ") as val_batches:
for dist_inputs in val_batches:
loss = strategy.run(_validate_step, args=(dist_inputs,))
for name, loss_value in loss.items():
loss_value = strategy.reduce(tf.distribute.ReduceOp.SUM, loss_value, axis=None)
validation_loss = validation_loss_dict_metric[name]
validation_loss.update_state(loss_value)
validation_result = get_and_reset_metric_from_dict(validation_loss_dict_metric)
write_metrics(validation_result, val_summary_writer, global_step)
# validation_history[global_step] = validation_result
logging.info("Validation result at epoch {} is {}".format(epoch, validation_result))
print("\n")
def do_callbacks(callbacks):
"""Call callbacks"""
if not epoch_end:
callback_scores = None
if callbacks and callbacks_interval_steps:
logging.info("Callbacks in progress at step {} . . . .".format(global_step))
callback_scores = []
for callback, callback_steps in zip(callbacks, callbacks_interval_steps):
if callback_steps and (global_step % callback_steps == 0):
score = callback(trainer_kwargs)
callback_scores.append(score)
else:
callback_scores.append(None)
return callback_scores
else:
callback_scores = None
if callbacks:
logging.info("Callbacks in progress at epoch end {} . . . .".format(epoch))
callback_scores = []
for callback in callbacks:
score = callback(trainer_kwargs)
callback_scores.append(score)
# Try to write a callback scores (only on epoch end)
# If we are returning a dict like {'exact_match': 81} or
# {'rougue-1': 30} etc . . . .
if score and isinstance(score, dict):
write_metrics(score, val_summary_writer, epoch)
return callback_scores
# Loop starts here
# Get Tensorboard writers
train_summary_writer, val_summary_writer = get_tensorboard_writers(model_checkpoint_dir)
validation_history = {}
training_history = {}
global_step = 0
epoch_end = False
STEPS = steps_per_epoch // steps_per_call
print("STEPS", STEPS)
for epoch in range(1, epochs + 1):
# start_epoch_time = time.time()
with tqdm.trange(STEPS, unit="batch ") as tepoch:
for step in tepoch:
steps_covered = (step + 1) * steps_per_call
global_step += steps_per_call
print("Started epoch {} and step {}".format(epoch, global_step))
tepoch.set_description(
"Epoch {}/{} --- Step {}/{} --- ".format(epoch, epochs, steps_covered, steps_per_epoch)
)
# Call Train
do_train(train_dataset_iter)
print("Train done")
# Call Validation
do_validation(validation_dataset_distributed)
print("Val done")
# Call Callbacks
callback_scores = do_callbacks(callbacks)
# Train Metrics
training_result = get_and_reset_metric_from_dict(training_loss_dict_metric)
training_history[global_step] = training_result
write_metrics(training_result, train_summary_writer, global_step)
# training_result["learning_rate"] = learning_rate_holder.result().numpy()
# learning_rate_holder.reset_states()
tepoch.set_postfix(**training_result)
# Save model
save_model()
# Do after every epoch
epoch_end = True
save_model(epoch_end)
#do_validation(validation_dataset_distributed)
#callback_scores = do_callbacks(callbacks)
epoch_end = False
# Flatten the results
training_history = flat_metric_dict(training_history)
validation_history = flat_metric_dict(validation_history)
return training_history, validation_history, callback_scores
class GPUTrainer:
def __init__(
self,
distribution_strategy,
num_gpus=0,
all_reduce_alg=None,
num_packs=1,
tpu_address=None,
dtype='fp32',
loss_scale='dynamic',
):
self.distribution_strategy = get_distribution_strategy(
distribution_strategy=distribution_strategy,
num_gpus=num_gpus,
all_reduce_alg=all_reduce_alg,
num_packs=num_packs,
tpu_address=tpu_address,
)
self.num_replicas = self.distribution_strategy.num_replicas_in_sync
self._dtype = get_tf_dtype(dtype)
# Setting dtype policy
set_mixed_precision_policy(self._dtype)
self.use_float16 = is_float16(self._dtype)
self.loss_scale = loss_scale
# # TODO
# if self.use_tpu:
# params["num_replicas"] = self.distribution_strategy.num_replicas_in_sync
# else:
# logging.info("Running transformer with num_gpus = %d", num_gpus)
# Add keras utils threads
def run(
self,
model_fn,
optimizer_fn,
train_dataset,
train_loss_fn,
epochs,
steps_per_epoch,
model_checkpoint_dir,
batch_size,
training_loss_names=None,
validation_loss_names=None,
validation_dataset=None,
validation_loss_fn=None,
validation_interval_steps=None,
steps_per_call=100,
enable_xla=True,
callbacks=None,
callbacks_interval_steps=None,
overwrite_checkpoint_dir=False,
max_number_of_models=10,
model_save_interval_steps=None,
repeat_dataset=True,
latest_checkpoint=None,
):
if steps_per_epoch:
logging.info("Make sure `steps_per_epoch` should be less than or equal to number of batches in dataset.")
if callbacks:
if callbacks_interval_steps is None:
callbacks_interval_steps = [None for callback in callbacks]
assert len(callbacks) == len(callbacks_interval_steps)
# Enable XLA
keras_utils.set_session_config(enable_xla=enable_xla)
logging.info("Policy: ----> {}".format(keras_utils.get_policy_name()))
logging.info("Strategy: ---> {}".format(self.distribution_strategy))
logging.info("Num GPU Devices: ---> {}".format(self.distribution_strategy.num_replicas_in_sync))
tf.keras.backend.clear_session()
# Under Strategy Scope
with self.distribution_strategy.scope():
# Model
model = model_fn()
# Optimizer
optimizer = optimizer_fn()
optimizer = configure_optimizer(optimizer, use_float16=self.use_float16, loss_scale=self.loss_scale)
# We use this to avoid inferring names from loss functions
_training_loss_names = ['loss']
_validation_loss_names = ['loss']
if training_loss_names:
_training_loss_names += training_loss_names
if validation_loss_names:
_validation_loss_names += validation_loss_names
# Make unique names
training_loss_names = list(set(_training_loss_names))
validation_loss_names = list(set(_validation_loss_names))
# Checkpoint manager
checkpoint_manager = save_model_checkpoints(
model, overwrite_checkpoint_dir, model_checkpoint_dir, max_number_of_models
)
# Try to load latest checkpoint
model.load_checkpoint(checkpoint_dir=model_checkpoint_dir, checkpoint_path=latest_checkpoint, opt=optimizer)
# Get metric dicts before distributing the dataset
# ddistributed datasets has no attribute .take
training_loss_dict_metric, validation_loss_dict_metric = get_loss_metric_dict(
training_loss_names, validation_loss_names
)
# Distribute dataset
if not repeat_dataset:
train_dataset_distributed = self.distribution_strategy.experimental_distribute_dataset(
train_dataset.repeat(epochs + 1)
)
else:
train_dataset_distributed = self.distribution_strategy.experimental_distribute_dataset(
train_dataset.repeat()
)
validation_dataset_distributed = None
if validation_dataset:
validation_dataset_distributed = self.distribution_strategy.experimental_distribute_dataset(
validation_dataset
)
# Make train dataset iterator
train_dataset_distributed = iter(train_dataset_distributed)
history = {}
training_history, validation_history, callback_scores = train_and_eval(
model,
optimizer,
self.distribution_strategy,
epochs,
steps_per_epoch,
steps_per_call,
train_dataset_distributed,
train_loss_fn,
batch_size,
training_loss_dict_metric,
validation_dataset_distributed,
validation_loss_fn,
validation_loss_dict_metric,
validation_interval_steps,
self.use_float16,
callbacks,
callbacks_interval_steps,
locals(),
checkpoint_manager,
model_checkpoint_dir,
model_save_interval_steps,
)
history['training_history'] = training_history
history['validation_hsitory'] = validation_history
history['callbacks'] = callback_scores
# Save json
return history
# -
distribution_strategy = "mirrored"
num_gpus = 2
trainer = GPUTrainer(
distribution_strategy,
num_gpus=num_gpus,
all_reduce_alg=None,
num_packs=1,
tpu_address=None,
dtype='fp32',
loss_scale='dynamic',
)
model_checkpoint_dir = "/tmp/model_ckpt/"
history = trainer.run(
model_fn = model_fn,
optimizer_fn = optimizer_fn,
train_dataset = train_dataset,
train_loss_fn = train_loss_fn,
epochs = 2,
steps_per_epoch = 100,
model_checkpoint_dir=model_checkpoint_dir,
batch_size=train_batch_size,
training_loss_names=None,
validation_loss_names=None,
validation_dataset=eval_dataset,
validation_loss_fn=train_loss_fn,
validation_interval_steps=None,
steps_per_call=1,
enable_xla=False,
callbacks=[callback],
callbacks_interval_steps=None,
overwrite_checkpoint_dir=True,
max_number_of_models=10,
model_save_interval_steps=None,
repeat_dataset=True,
latest_checkpoint=None,
)
# Under Strategy Scope
with trainer.distribution_strategy.scope():
# Model
model = model_fn()
# Optimizer
optimizer = optimizer_fn()
optimizer = configure_optimizer(optimizer, use_float16=False, loss_scale="dynamic")
def compute_loss(batch_labels, model_outputs):
"""Loss computation which takes care of loss reduction based on GLOBAL_BATCH_SIZE"""
per_example_loss = train_loss_fn(batch_labels, model_outputs)
per_example_loss_averaged = {}
# Inplace update
# Avergae loss per global batch size , recommended
for name, loss in per_example_loss.items():
per_example_loss_averaged[name] = tf.nn.compute_average_loss(loss, global_batch_size=GLOBAL_BATCH_SIZE)
return per_example_loss_averaged
# Train Functions
@tf.function
def do_train(iterator):
"""The step function for one training step"""
def train_step(dist_inputs):
"""The computation to run on each device."""
batch_inputs, batch_labels = dist_inputs
with tf.GradientTape() as tape:
model_outputs = model(batch_inputs)
loss = compute_loss(batch_labels, model_outputs)
tf.debugging.check_numerics(loss['loss'], message='Loss value is either NaN or inf')
if isinstance(optimizer, tf.keras.mixed_precision.LossScaleOptimizer):
loss_scaled = {name: optimizer.get_scaled_loss(loss_value) for name, loss_value in loss.items()}
# TODO
# Scales down the loss for gradients to be invariant from replicas.
# loss = loss / strategy.num_replicas_in_sync
if mixed_precision:
scaled_gradients = tape.gradient(loss_scaled["loss"], model.trainable_variables)
grads = optimizer.get_unscaled_gradients(scaled_gradients)
else:
grads = tape.gradient(loss["loss"], model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# training_loss.update_state(loss * strategy.num_replicas_in_sync)
return loss
for _ in tf.range(tf.convert_to_tensor(steps_per_call)):
dist_inputs = next(iterator)
loss = strategy.run(train_step, args=(dist_inputs,))
# strategy reduce
loss = {
name: strategy.reduce(tf.distribute.ReduceOp.MEAN, loss_value, axis=None)
for name, loss_value in loss.items()
}
t_loss.update_state(loss['loss'])
# training_result = get_and_reset_metric_from_dict(training_loss_dict_metric)
train_dataset_distributed = trainer.distribution_strategy.experimental_distribute_dataset(train_dataset)
train_dataset_distributed = iter(train_dataset_distributed)
steps_per_call = 1
GLOBAL_BATCH_SIZE = 32
mixed_precision = False
strategy = trainer.distribution_strategy
t_loss = tf.keras.metrics.Mean("loss", dtype=tf.float32)
l = do_train(train_dataset_distributed)
t_loss.result()
shutil.rmtree(tfrecord_dir)
shutil.rmtree(model_checkpoint_dir)
|
research/glue/test_glue.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GLM: Robust Regression with Outlier Detection
#
# **A minimal reproducable example of Robust Regression with Outlier Detection using Hogg 2010 Signal vs Noise method.**
#
# + This is a complementary approach to the Student-T robust regression as illustrated in [<NAME>'s notebook]((GLM-robust.ipynb), that approach is also compared here.
# + This model returns a robust estimate of linear coefficients and an indication of which datapoints (if any) are outliers.
# + The likelihood evaluation is essentially a copy of eqn 17 in "Data analysis recipes: Fitting a model to data" - [Hogg 2010](http://arxiv.org/abs/1008.4686).
# + The model is adapted specifically from <NAME>' [implementation](http://www.astroml.org/book_figures/chapter8/fig_outlier_rejection.html) (3rd model tested).
# + The dataset is tiny and hardcoded into this Notebook. It contains errors in both the x and y, but we will deal here with only errors in y.
#
#
# **Note:**
#
# + Python 3.4 project using latest available [PyMC3](https://github.com/pymc-devs/pymc3)
# + Developed using [ContinuumIO Anaconda](https://www.continuum.io/downloads) distribution on a Macbook Pro 3GHz i7, 16GB RAM, OSX 10.10.5.
# + During development I've found that 3 data points are always indicated as outliers, but the remaining ordering of datapoints by decreasing outlier-hood is slightly unstable between runs: the posterior surface appears to have a small number of solutions with similar probability.
# + Finally, if runs become unstable or Theano throws weird errors, try clearing the cache `$> theano-cache clear` and rerunning the notebook.
#
#
# **Package Requirements (shown as a conda-env YAML):**
# ```
# $> less conda_env_pymc3_examples.yml
#
# name: pymc3_examples
# channels:
# - defaults
# dependencies:
# - python=3.4
# - ipython
# - ipython-notebook
# - ipython-qtconsole
# - numpy
# - scipy
# - matplotlib
# - pandas
# - seaborn
# - patsy
# - pip
#
# $> conda env create --file conda_env_pymc3_examples.yml
#
# $> source activate pymc3_examples
#
# $> pip install --process-dependency-links git+https://github.com/pymc-devs/pymc3
#
# ```
# ## Setup
# +
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import optimize
import pymc3 as pm
import theano as thno
import theano.tensor as T
# configure some basic options
sns.set(style="darkgrid", palette="muted")
pd.set_option('display.notebook_repr_html', True)
plt.rcParams['figure.figsize'] = 12, 8
np.random.seed(0)
# -
# ### Load and Prepare Data
# We'll use the Hogg 2010 data available at https://github.com/astroML/astroML/blob/master/astroML/datasets/hogg2010test.py
#
# It's a very small dataset so for convenience, it's hardcoded below
# +
#### cut & pasted directly from the fetch_hogg2010test() function
## identical to the original dataset as hardcoded in the Hogg 2010 paper
dfhogg = pd.DataFrame(np.array([[1, 201, 592, 61, 9, -0.84],
[2, 244, 401, 25, 4, 0.31],
[3, 47, 583, 38, 11, 0.64],
[4, 287, 402, 15, 7, -0.27],
[5, 203, 495, 21, 5, -0.33],
[6, 58, 173, 15, 9, 0.67],
[7, 210, 479, 27, 4, -0.02],
[8, 202, 504, 14, 4, -0.05],
[9, 198, 510, 30, 11, -0.84],
[10, 158, 416, 16, 7, -0.69],
[11, 165, 393, 14, 5, 0.30],
[12, 201, 442, 25, 5, -0.46],
[13, 157, 317, 52, 5, -0.03],
[14, 131, 311, 16, 6, 0.50],
[15, 166, 400, 34, 6, 0.73],
[16, 160, 337, 31, 5, -0.52],
[17, 186, 423, 42, 9, 0.90],
[18, 125, 334, 26, 8, 0.40],
[19, 218, 533, 16, 6, -0.78],
[20, 146, 344, 22, 5, -0.56]]),
columns=['id','x','y','sigma_y','sigma_x','rho_xy'])
## for convenience zero-base the 'id' and use as index
dfhogg['id'] = dfhogg['id'] - 1
dfhogg.set_index('id', inplace=True)
## standardize (mean center and divide by 1 sd)
dfhoggs = (dfhogg[['x','y']] - dfhogg[['x','y']].mean(0)) / dfhogg[['x','y']].std(0)
dfhoggs['sigma_y'] = dfhogg['sigma_y'] / dfhogg['y'].std(0)
dfhoggs['sigma_x'] = dfhogg['sigma_x'] / dfhogg['x'].std(0)
## create xlims ylims for plotting
xlims = (dfhoggs['x'].min() - np.ptp(dfhoggs['x'])/5
,dfhoggs['x'].max() + np.ptp(dfhoggs['x'])/5)
ylims = (dfhoggs['y'].min() - np.ptp(dfhoggs['y'])/5
,dfhoggs['y'].max() + np.ptp(dfhoggs['y'])/5)
## scatterplot the standardized data
g = sns.FacetGrid(dfhoggs, size=8)
_ = g.map(plt.errorbar, 'x', 'y', 'sigma_y', 'sigma_x', marker="o", ls='')
_ = g.axes[0][0].set_ylim(ylims)
_ = g.axes[0][0].set_xlim(xlims)
plt.subplots_adjust(top=0.92)
_ = g.fig.suptitle('Scatterplot of Hogg 2010 dataset after standardization', fontsize=16)
# -
# **Observe**:
#
# + Even judging just by eye, you can see these datapoints mostly fall on / around a straight line with positive gradient
# + It looks like a few of the datapoints may be outliers from such a line
# ## Create Conventional OLS Model
# The *linear model* is really simple and conventional:
#
# $$\bf{y} = \beta^{T} \bf{X} + \bf{\sigma}$$
#
# where:
#
# $\beta$ = coefs = $\{1, \beta_{j \in X_{j}}\}$
# $\sigma$ = the measured error in $y$ in the dataset `sigma_y`
# ### Define model
#
# **NOTE:**
# + We're using a simple linear OLS model with Normally distributed priors so that it behaves like a ridge regression
with pm.Model() as mdl_ols:
## Define weakly informative Normal priors to give Ridge regression
b0 = pm.Normal('b0_intercept', mu=0, sd=100)
b1 = pm.Normal('b1_slope', mu=0, sd=100)
## Define linear model
yest = b0 + b1 * dfhoggs['x']
## Use y error from dataset, convert into theano variable
sigma_y = thno.shared(np.asarray(dfhoggs['sigma_y'],
dtype=thno.config.floatX), name='sigma_y')
## Define Normal likelihood
likelihood = pm.Normal('likelihood', mu=yest, sd=sigma_y, observed=dfhoggs['y'])
# ### Sample
with mdl_ols:
## take samples
traces_ols = pm.sample(2000, tune=1000)
# ### View Traces
#
# **NOTE**: I'll 'burn' the traces to only retain the final 1000 samples
_ = pm.traceplot(traces_ols[-1000:], figsize=(12,len(traces_ols.varnames)*1.5),
lines={k: v['mean'] for k, v in pm.df_summary(traces_ols[-1000:]).iterrows()})
# **NOTE:** We'll illustrate this OLS fit and compare to the datapoints in the final plot
# ---
#
# ---
# ## Create Robust Model: Student-T Method
# I've added this brief section in order to directly compare the Student-T based method exampled in [<NAME>'s notebook](GLM-robust.ipynb).
#
# Instead of using a Normal distribution for the likelihood, we use a Student-T, which has fatter tails. In theory this allows outliers to have a smaller mean square error in the likelihood, and thus have less influence on the regression estimation. This method does not produce inlier / outlier flags but is simpler and faster to run than the Signal Vs Noise model below, so a comparison seems worthwhile.
#
# **Note:** we'll constrain the Student-T 'degrees of freedom' parameter `nu` to be an integer, but otherwise leave it as just another stochastic to be inferred: no need for prior knowledge.
# ### Define Model
with pm.Model() as mdl_studentt:
## Define weakly informative Normal priors to give Ridge regression
b0 = pm.Normal('b0_intercept', mu=0, sd=100)
b1 = pm.Normal('b1_slope', mu=0, sd=100)
## Define linear model
yest = b0 + b1 * dfhoggs['x']
## Use y error from dataset, convert into theano variable
sigma_y = thno.shared(np.asarray(dfhoggs['sigma_y'],
dtype=thno.config.floatX), name='sigma_y')
## define prior for Student T degrees of freedom
nu = pm.Uniform('nu', lower=1, upper=100)
## Define Student T likelihood
likelihood = pm.StudentT('likelihood', mu=yest, sd=sigma_y, nu=nu,
observed=dfhoggs['y'])
# ### Sample
with mdl_studentt:
## take samples
traces_studentt = pm.sample(2000, tune=1000)
# #### View Traces
_ = pm.traceplot(traces_studentt[-1000:],
figsize=(12,len(traces_studentt.varnames)*1.5),
lines={k: v['mean'] for k, v in pm.df_summary(traces_studentt[-1000:]).iterrows()})
# **Observe:**
#
# + Both parameters `b0` and `b1` show quite a skew to the right, possibly this is the action of a few samples regressing closer to the OLS estimate which is towards the left
# + The `nu` parameter seems very happy to stick at `nu = 1`, indicating that a fat-tailed Student-T likelihood has a better fit than a thin-tailed (Normal-like) Student-T likelihood.
# + The inference sampling also ran very quickly, almost as quickly as the conventional OLS
#
#
# **NOTE:** We'll illustrate this Student-T fit and compare to the datapoints in the final plot
# ---
#
# ---
# ## Create Robust Model with Outliers: Hogg Method
# Please read the paper (Hogg 2010) and <NAME>' code for more complete information about the modelling technique.
#
# The general idea is to create a 'mixture' model whereby datapoints can be described by either the linear model (inliers) or a modified linear model with different mean and larger variance (outliers).
#
#
# The likelihood is evaluated over a mixture of two likelihoods, one for 'inliers', one for 'outliers'. A Bernouilli distribution is used to randomly assign datapoints in N to either the inlier or outlier groups, and we sample the model as usual to infer robust model parameters and inlier / outlier flags:
#
# $$
# \mathcal{logL} = \sum_{i}^{i=N} log \left[ \frac{(1 - B_{i})}{\sqrt{2 \pi \sigma_{in}^{2}}} exp \left( - \frac{(x_{i} - \mu_{in})^{2}}{2\sigma_{in}^{2}} \right) \right] + \sum_{i}^{i=N} log \left[ \frac{B_{i}}{\sqrt{2 \pi (\sigma_{in}^{2} + \sigma_{out}^{2})}} exp \left( - \frac{(x_{i}- \mu_{out})^{2}}{2(\sigma_{in}^{2} + \sigma_{out}^{2})} \right) \right]
# $$
#
# where:
# $\bf{B}$ is Bernoulli-distibuted $B_{i} \in [0_{(inlier)},1_{(outlier)}]$
#
#
# ### Define model
def logp_signoise(yobs, is_outlier, yest_in, sigma_y_in, yest_out, sigma_y_out):
'''
Define custom loglikelihood for inliers vs outliers.
NOTE: in this particular case we don't need to use theano's @as_op
decorator because (as stated by Twiecki in conversation) that's only
required if the likelihood cannot be expressed as a theano expression.
We also now get the gradient computation for free.
'''
# likelihood for inliers
pdfs_in = T.exp(-(yobs - yest_in + 1e-4)**2 / (2 * sigma_y_in**2))
pdfs_in /= T.sqrt(2 * np.pi * sigma_y_in**2)
logL_in = T.sum(T.log(pdfs_in) * (1 - is_outlier))
# likelihood for outliers
pdfs_out = T.exp(-(yobs - yest_out + 1e-4)**2 / (2 * (sigma_y_in**2 + sigma_y_out**2)))
pdfs_out /= T.sqrt(2 * np.pi * (sigma_y_in**2 + sigma_y_out**2))
logL_out = T.sum(T.log(pdfs_out) * is_outlier)
return logL_in + logL_out
with pm.Model() as mdl_signoise:
## Define weakly informative Normal priors to give Ridge regression
b0 = pm.Normal('b0_intercept', mu=0, sd=10, testval=pm.floatX(0.1))
b1 = pm.Normal('b1_slope', mu=0, sd=10, testval=pm.floatX(1.))
## Define linear model
yest_in = b0 + b1 * dfhoggs['x']
## Define weakly informative priors for the mean and variance of outliers
yest_out = pm.Normal('yest_out', mu=0, sd=100, testval=pm.floatX(1.))
sigma_y_out = pm.HalfNormal('sigma_y_out', sd=100, testval=pm.floatX(1.))
## Define Bernoulli inlier / outlier flags according to a hyperprior
## fraction of outliers, itself constrained to [0,.5] for symmetry
frac_outliers = pm.Uniform('frac_outliers', lower=0., upper=.5)
is_outlier = pm.Bernoulli('is_outlier', p=frac_outliers, shape=dfhoggs.shape[0],
testval=np.random.rand(dfhoggs.shape[0]) < 0.2)
## Extract observed y and sigma_y from dataset, encode as theano objects
yobs = thno.shared(np.asarray(dfhoggs['y'], dtype=thno.config.floatX), name='yobs')
sigma_y_in = thno.shared(np.asarray(dfhoggs['sigma_y'], dtype=thno.config.floatX),
name='sigma_y_in')
## Use custom likelihood using DensityDist
likelihood = pm.DensityDist('likelihood', logp_signoise,
observed={'yobs': yobs, 'is_outlier': is_outlier,
'yest_in': yest_in, 'sigma_y_in': sigma_y_in,
'yest_out': yest_out, 'sigma_y_out': sigma_y_out})
# ### Sample
with mdl_signoise:
## two-step sampling to create Bernoulli inlier/outlier flags
step1 = pm.Metropolis([frac_outliers, yest_out, sigma_y_out, b0, b1])
step2 = pm.step_methods.BinaryGibbsMetropolis([is_outlier])
## take samples
traces_signoise = pm.sample(20000, step=[step1, step2], tune=10000, progressbar=True)
# ### View Traces
traces_signoise[-10000:]['b0_intercept']
_ = pm.traceplot(traces_signoise[-10000:], figsize=(12,len(traces_signoise.varnames)*1.5),
lines={k: v['mean'] for k, v in pm.df_summary(traces_signoise[-1000:]).iterrows()})
# **NOTE:**
#
# + During development I've found that 3 datapoints id=[1,2,3] are always indicated as outliers, but the remaining ordering of datapoints by decreasing outlier-hood is unstable between runs: the posterior surface appears to have a small number of solutions with very similar probability.
# + The NUTS sampler seems to work okay, and indeed it's a nice opportunity to demonstrate a custom likelihood which is possible to express as a theano function (thus allowing a gradient-based sampler like NUTS). However, with a more complicated dataset, I would spend time understanding this instability and potentially prefer using more samples under Metropolis-Hastings.
# ---
#
# ---
# ## Declare Outliers and Compare Plots
# ### View ranges for inliers / outlier predictions
# At each step of the traces, each datapoint may be either an inlier or outlier. We hope that the datapoints spend an unequal time being one state or the other, so let's take a look at the simple count of states for each of the 20 datapoints.
# +
outlier_melt = pd.melt(pd.DataFrame(traces_signoise['is_outlier', -1000:],
columns=['[{}]'.format(int(d)) for d in dfhoggs.index]),
var_name='datapoint_id', value_name='is_outlier')
ax0 = sns.pointplot(y='datapoint_id', x='is_outlier', data=outlier_melt,
kind='point', join=False, ci=None, size=4, aspect=2)
_ = ax0.vlines([0,1], 0, 19, ['b','r'], '--')
_ = ax0.set_xlim((-0.1,1.1))
_ = ax0.set_xticks(np.arange(0, 1.1, 0.1))
_ = ax0.set_xticklabels(['{:.0%}'.format(t) for t in np.arange(0,1.1,0.1)])
_ = ax0.yaxis.grid(True, linestyle='-', which='major', color='w', alpha=0.4)
_ = ax0.set_title('Prop. of the trace where datapoint is an outlier')
_ = ax0.set_xlabel('Prop. of the trace where is_outlier == 1')
# -
# **Observe**:
#
# + The plot above shows the number of samples in the traces in which each datapoint is marked as an outlier, expressed as a percentage.
# + In particular, 3 points [1, 2, 3] spend >=95% of their time as outliers
# + Contrastingly, points at the other end of the plot close to 0% are our strongest inliers.
# + For comparison, the mean posterior value of `frac_outliers` is ~0.35, corresponding to roughly 7 of the 20 datapoints. You can see these 7 datapoints in the plot above, all those with a value >50% or thereabouts.
# + However, only 3 of these points are outliers >=95% of the time.
# + See note above regarding instability between runs.
#
# The 95% cutoff we choose is subjective and arbitrary, but I prefer it for now, so let's declare these 3 to be outliers and see how it looks compared to <NAME>as' outliers, which were declared in a slightly different way as points with means above 0.68.
# ### Declare outliers
#
# **Note:**
# + I will declare outliers to be datapoints that have value == 1 at the 5-percentile cutoff, i.e. in the percentiles from 5 up to 100, their values are 1.
# + Try for yourself altering cutoff to larger values, which leads to an objective ranking of outlier-hood.
cutoff = 5
dfhoggs['outlier'] = np.percentile(traces_signoise[-1000:]['is_outlier'],cutoff, axis=0)
dfhoggs['outlier'].value_counts()
# ### Posterior Prediction Plots for OLS vs StudentT vs SignalNoise
# +
g = sns.FacetGrid(dfhoggs, size=8, hue='outlier', hue_order=[True,False],
palette='Set1', legend_out=False)
lm = lambda x, samp: samp['b0_intercept'] + samp['b1_slope'] * x
pm.plot_posterior_predictive_glm(traces_ols[-1000:],
eval=np.linspace(-3, 3, 10), lm=lm, samples=200, color='#22CC00', alpha=.2)
pm.plot_posterior_predictive_glm(traces_studentt[-1000:], lm=lm,
eval=np.linspace(-3, 3, 10), samples=200, color='#FFA500', alpha=.5)
pm.plot_posterior_predictive_glm(traces_signoise[-1000:], lm=lm,
eval=np.linspace(-3, 3, 10), samples=200, color='#357EC7', alpha=.3)
_ = g.map(plt.errorbar, 'x', 'y', 'sigma_y', 'sigma_x', marker="o", ls='').add_legend()
_ = g.axes[0][0].annotate('OLS Fit: Green\nStudent-T Fit: Orange\nSignal Vs Noise Fit: Blue',
size='x-large', xy=(1,0), xycoords='axes fraction',
xytext=(-160,10), textcoords='offset points')
_ = g.axes[0][0].set_ylim(ylims)
_ = g.axes[0][0].set_xlim(xlims)
# -
# **Observe**:
#
# + The posterior preditive fit for:
# + the **OLS model** is shown in **Green** and as expected, it doesn't appear to fit the majority of our datapoints very well, skewed by outliers
# + the **Robust Student-T model** is shown in **Orange** and does appear to fit the 'main axis' of datapoints quite well, ignoring outliers
# + the **Robust Signal vs Noise model** is shown in **Blue** and also appears to fit the 'main axis' of datapoints rather well, ignoring outliers.
#
#
# + We see that the **Robust Signal vs Noise model** also yields specific estimates of _which_ datapoints are outliers:
# + 17 'inlier' datapoints, in **Blue** and
# + 3 'outlier' datapoints shown in **Red**.
# + From a simple visual inspection, the classification seems fair, and agrees with Jake Vanderplas' findings.
#
#
# + Overall, it seems that:
# + the **Signal vs Noise model** behaves as promised, yielding a robust regression estimate and explicit labelling of inliers / outliers, but
# + the **Signal vs Noise model** is quite complex and whilst the regression seems robust and stable, the actual inlier / outlier labelling seems slightly unstable
# + if you simply want a robust regression without inlier / outlier labelling, the **Student-T model** may be a good compromise, offering a simple model, quick sampling, and a very similar estimate.
# ---
# Example originally contributed by <NAME> 2015-12-21 [github.com/jonsedar](https://github.com/jonsedar)
|
docs/source/notebooks/GLM-robust-with-outlier-detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
chemicals = pd.read_csv('chemicals.csv')
chemicals.describe()
chemicals.head()
IndusOccu = pd.read_csv('industry_occupation.csv', encoding="ISO-8859-1")
IndusOccu.describe()
IndusOccu[:100]
earnings = pd.read_csv('earnings.csv', encoding="ISO-8859-1")
earnings.head()
earnings.describe()
ChemEarning = pd.concat([chemicals, earnings], axis=1, join='outer', ignore_index=False)
ChemEarning.head()
ChemEarning.corr()
test = earnings[['wholesale_trade', 'year']]
WholeSale_Trade = test.groupby('year').size()
print(WholeSale_Trade)
#Plotting the Graph
plot_by_year = WholeSale_Trade.plot(title='Yearly Sales',xticks=(2010,2011,2012,2013,2014,2015,2016))
plot_by_year.set_xlabel('Years')
plot_by_year.set_ylabel('Wholesale Trade')
test1 = IndusOccu[['total_employed', 'year']]
import seaborn as sns
sns.set()
TotalEmpYear = test1.groupby('year').size()
print(TotalEmpYear)
#Plotting the Graph
plot_by_year = TotalEmpYear.plot(title='Yearly Employment',xticks=(2010,2011,2012,2013,2014,2015,2016))
plot_by_year.set_xlabel('Years')
plot_by_year.set_ylabel('TotalEmpYear')
import seaborn as sns
sns.set()
chemicals.chemical_species.value_counts().plot(kind='bar')
ChemEarningIndus = pd.concat([ChemEarning, IndusOccu], axis=1, join='outer', ignore_index=False)
#correlation matrix
import seaborn as sns
corrmat = ChemEarningIndus.corr()
f, ax = plt.subplots(figsize=(19, 12))
sns.heatmap(corrmat, vmax=.8, square=True);
ChemEarningIndus.head()
|
Chemical industry_occupation analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Open the file
# My BU ID is U64501194
# Therefore, I am dealing with the silver medals
# ### 1. load the "country medals" csv as a list of lines using Python and construct a sublist for you group
# +
# -*- coding: utf-8 -*-
"""
Created on Mon Nov 5 14:37:29 2018
@author: epinsky
this scripts reads your ticker file (e.g. MSFT.csv) and
constructs a list of lines
"""
import os
ticker='Country_Medals'
input_dir = os.getcwd()
ticker_file = os.path.join(input_dir, ticker + '.csv')
try:
with open(ticker_file) as f:
lines = f.read().splitlines()
print('opened file for ticker: ', ticker)
""" your code for assignment 1 goes here
"""
except Exception as e:
print(e)
print('failed to read stock data for ticker: ', ticker)
# -
lines[0:5]
# # Extract the data
# Class a data type (year, countryName, silverMetal)
class DataNode:
def __init__(self, year, countryName, silver):
self.year = year
self.countryName = countryName
self.silver = silver
def show(self):
print("{:5} {:15} {:3}".format(self.year, self.countryName, self.silver))
# +
# Split the raw data into list
table = []
for line in lines:
temp = line.split(',')
table.append(temp)
# Extract the raw data
extractData = []
for row in table:
y = row[0]
cn = row[2]
s = row[6]
dataNode = DataNode(y, cn, s)
extractData.append(dataNode)
# Remove the title from the extractData
extractData = extractData[1:]
# -
# Show the first 20 row of the extracted data
for i in range(20):
extractData[i].show()
# # Analysis
# ### 2. how many entries are there?
# How many entries
entries = len(extractData)
print("we have {} entries of data and 1 row of title".format(entries))
# Calculate the average medals for each country
# Sort the list with the key 'countryName'
sortData = sorted(extractData, key= lambda x:x.countryName)
# Show first 20 data
for node in sortData[0:20]:
node.show()
# +
# Extract the country from data
temp = []
for node in sortData:
c = node.countryName
temp.append(c)
# Processing the data into the form (countryname, list(a series of medal wins)
countrySet = sorted(set(temp))
medalList = [[] for i in range(len(countrySet))]
for node in sortData:
n = node.countryName
s = node.silver
index = countrySet.index(n)
medalList[index].append(int(s))
countryMedalData = list(zip(countrySet, medalList))
# -
# Print the first 20 countryMedalData
for data in countryMedalData[0:20]:
print(data)
# ### 3. compute the average numbers of medals per country and write this (in decreasing order) to a
# le "average medals per country.csv" for your group
#
# ### 4. which country has the highest (average) number of medals?
# ### 5. list top 10 countries by (averaged) number of medals
# Average medal for each country
averageMedal = []
for data in countryMedalData:
name = data[0]
medal = data[1]
aveMedal = sum(data[1]) // len(data[1])
averageMedal.append((name, aveMedal))
# Sorting the averageMedal in decreasing ordert
temp = sorted(averageMedal, key=lambda x:x[1], reverse = True)
print(temp)
print("\n\n\n\n{} has won the most averaged silver medal".format(temp[0][0]), )
print("Unified Team only once participated the olympic game in 1992, and the team won 38 silver metals")
# Wirte the average data into a new cvs name ""average medals per country.csv"
import csv
try:
with open('average_medals_per_country.csv', 'w', newline = '') as file:
writer = csv.writer(file)
writer.writerow(['Country Name', 'Average Silver Medal'])
for data in temp:
name = data[0]
medal = data[1]
writer.writerow([name, medal])
print('\nOutput csv SUCCESSFUL\n')
except:
print("Something wrong when output the csv file")
# List top 10 countries by (average) number of medals
for data in temp[0:10]:
print("{:25} {}".format(data[0], data[1]))
# ### 6. compute the median number of medals per country and write this (in decreasing order) to a
# le "median medals per country.csv" for your group
# +
def medianNum(inputList):
inputList = sorted(inputList)
l = len(inputList)
if l % 2 == 0:
index = l // 2
return (inputList[index] + inputList[index - 1]) // 2
else:
index = l // 2
return inputList[index]
# +
medMedal = []
for data in countryMedalData:
name = data[0]
medal = data[1]
l = len(medal)
median = medianNum(medal)
medMedal.append((name, median))
medMedal = sorted(medMedal, key=lambda d:d[1], reverse=True)
print(medMedal)
# -
# Output csv file
try:
with open('median_medals_per_country.csv', 'w', newline = '') as file:
writer = csv.writer(file)
writer.writerow(['Country Name', 'median of Silver Medal'])
for data in medMedal:
name = data[0]
m = data[1]
writer.writerow([name, m])
print('\nOutput csv SUCCESSFUL\n')
except:
print("Something wrong when output the csv file")
# ### 7. which country has the highest median number of medals?
print("{} has the highest median number of silver medals, which is {}".format(medMedal[0][0], medMedal[0][1]))
# ### 8. list top 10 countries by median number of medals
for data in medMedal[0:10]:
country = data[0]
m = data[1]
print("{:25} {:5}".format(country, m))
|
Assignment_1/Part_1_medals/Olympic Medal Code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from datetime import datetime
from datetime import timedelta
# # Basic
now = datetime.now()
# now
now.year, now.month, now.day
diff_dt = datetime(2018, 8, 31) - datetime(2018, 8, 30, 23, 0)
diff_dt.days, diff_dt.seconds, type(diff_dt), now + timedelta(1)
# ## fromat, convert
str(now), now.strftime('%Y-%m-%d %H:%M:%S')
value = '2018-08-30 18:28:47'
dt1 = datetime.strptime(value, '%Y-%m-%d %H:%M:%S')
values = ['2018/08/30', '2018/08/31']
dt2 = [datetime.strptime(x, '%Y/%m/%d') for x in values]
dt1, dt2
# ## index, split
dates = [datetime(2018, 9, x) for x in range(1, 7)]
ts = pd.Series(data = np.random.randn(6), index=dates)
dates, ts, type(ts.index), type(ts)
# [start:end:step]
ts.index[0], ts[::2], ts['2018-9-2':'2018-9-5']
ts['2018-9-3':'2018-9-5']
ts.truncate(before='2018-09-03', after='2018-09-5')
# ## range
dt3 = pd.date_range(start='15/6/2016', periods=1000, freq='W-WED')
type(dt3)
df3 = pd.DataFrame(data = np.random.randn(1000, 4), index=dt3, columns=['col' + str(x) for x in range(1, 5)])
df3[:5], df3.tail(5), df3.index.size, df3.columns.size, df3.size
df3.loc['2022-09'] # df3.loc['9-2022']
# ## duplicate indices
dt4 = pd.DatetimeIndex(data=['1/9/2018', '2/9/2018', '2/9/2018', '2/9/2018', '3/9/2018'])
df4 = pd.Series(data=np.arange(5), index=dt4)
dt4.is_unique, df4.index.is_unique
df4['1/9/2018'], df4['2/9/2018'], df4['3/9/2018']
df4
df4_grouped = df4.groupby(level=0) # index only one level
df4_grouped.count()
ts2 = ts[::2]
ts2
ts3 = ts2.resample('D')
ts3.mean()
dt5 = pd.date_range('1/9/2018', periods=12, freq='T')
s5 = pd.Series(data=np.arange(12), index=dt5)
s5
s5_1 = s5.resample(rule='5min')
s5_1.mean()
|
ML/learn/Python_For_Data_Analysis/time_series.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 4. Gaussian Mixture Models
# Gaussian Mixture Models are a form of **density estimation**. They give us an approximation of the probability distribution of our data. We want to use gaussian mixture models when we notice that our data is multimodal (meaning there are multiple modes or bumps). From probability, we can recall that the **mode** is just the most common value. For instance, a multi modal distribution can be seen below:
# +
import numpy as np
from scipy.stats import bernoulli, binom, norm
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
sns.set(style="white", palette="husl")
sns.set_context("talk")
sns.set_style("ticks")
# +
x_axis = np.arange(-15, 15, 0.001)
fig = plt.figure(figsize=(7,4))
plt.plot(x_axis, norm.pdf(x_axis, -4, 2) + norm.pdf(x_axis, 4, 2), c=sns.xkcd_rgb["green"])
plt.title("Multi Modal Gaussian", fontsize=16)
plt.show()
# -
# Multi modal gaussians can be viewed as a **linear combination** of individual gaussians:
# +
x_axis = np.arange(-15, 15, 0.001)
mu = {
"1": 2,
"2": -2
}
sigma = {
"1": 3,
"2": 1
}
weights = {
"1": 0.5,
"2": 0.5
}
gaussian_1 = norm.pdf(x_axis, mu["1"], sigma["1"])
gaussian_2 = norm.pdf(x_axis, mu["2"], sigma["2"])
fig = plt.figure(figsize=(16,3.5))
plt.subplot(1, 3, 1)
plt.plot(x_axis, gaussian_1, c=sns.xkcd_rgb["red"])
plt.plot(x_axis, gaussian_2, c=sns.xkcd_rgb["blue"])
plt.legend(["Gaussian 1", "Gaussian 2"], fontsize=12)
plt.subplot(1, 3, 2)
plt.plot(x_axis, gaussian_1, c=sns.xkcd_rgb["red"], alpha=0.5)
plt.plot(x_axis, gaussian_2, c=sns.xkcd_rgb["blue"], alpha=0.5)
plt.plot(x_axis, gaussian_1*weights["1"] + gaussian_2*weights["2"], c=sns.xkcd_rgb["purple"])
plt.legend(["Gaussian 1", "Gaussian 2", "Gaussian Mixture"], fontsize=10)
plt.subplot(1, 3, 3)
plt.plot(x_axis, gaussian_1*weights["1"] + gaussian_2*weights["2"], c=sns.xkcd_rgb["purple"])
plt.legend(["Gaussian Mixture"], fontsize=10)
fig.suptitle('Gaussian Mixture Model', fontsize=16)
plt.show()
# -
# A very important thing to note is that GMM's are a type of model known as a **generative model**. What that means is that if you were to be presented with a set of observations, $x$, that was generated from the gaussian mixture distribution, it would be generated as follows:
#
# ```
# for i in range(len(X)):
# select gaussian based on prior distribution of gaussians
# sample from selected gaussian to generate data point x
# ```
#
# This can be seen in code below:
# +
fig = plt.figure(figsize=(12,7))
# Upper left
plt.subplot(2, 2, 1)
plt.plot(x_axis, gaussian_1 + gaussian_2, c=sns.xkcd_rgb["purple"])
plt.legend(["Gaussian Mixture"], fontsize=10)
def generate_x(num_samples=100):
num_gaussians = 2
prior = np.array([0.5, 0.5]) # Uniform prior across distribution 1 and 2
X = []
for i in range(num_samples):
selected_gaussian = np.random.choice(num_gaussians, p=prior)
x = norm.rvs(mu[str(selected_gaussian + 1)], sigma[str(selected_gaussian + 1)])
X.append(x)
return X
# Upper right
plt.subplot(2, 2, 2)
plt.hist(generate_x(), bins=30, density=True, color=sns.xkcd_rgb["light purple"], alpha=0.5, edgecolor="black")
plt.plot(x_axis, gaussian_1*weights["1"] + gaussian_2*weights["2"], c=sns.xkcd_rgb["purple"])
plt.legend(["100 Samples"], fontsize=10)
# lower left
plt.subplot(2, 2, 3)
plt.hist(generate_x(500), bins=30, density=True, color=sns.xkcd_rgb["light purple"], alpha=0.5, edgecolor="black")
plt.plot(x_axis, gaussian_1*weights["1"] + gaussian_2*weights["2"], c=sns.xkcd_rgb["purple"])
plt.legend(["500 Samples"], fontsize=10)
# lower right
plt.subplot(2, 2, 4)
plt.hist(generate_x(10000), bins=50, density=True, color=sns.xkcd_rgb["light purple"], alpha=0.5, edgecolor="black")
plt.plot(x_axis, gaussian_1*weights["1"] + gaussian_2*weights["2"], c=sns.xkcd_rgb["purple"])
plt.legend(["10,000 Samples"], fontsize=10)
plt.show()
# -
# We have our data generation process encapsulated in the function `generate_x`, and we can see that was we draw more samples we approximate the shape closer and closer. Let's now take a moment to hone in on the mathematics at work here.
#
# As I stated earlier, a Gaussian mixture is just the sum of weighted gaussians. To represent these weights we will introduce a new symbol called $\pi$. $\pi_k$ is the probability that $x$ belongs to the $k$th Gaussian.
#
# $$P(x) = \pi_1 N(x \mid \mu_1, \Sigma_1) + \pi_2 N(x \mid \mu_2, \Sigma_2) + \pi_3 N(x \mid \mu_3, \Sigma_3)$$
#
# $\pi$ can be thought of as a **prior**, specifically the prior probability of a data point being generated from a particular gaussian. To get an intuitive sense of for why this is necessary, consider the following situation. We have a list of 100 heights, but we do not know whether they the gender corresponding to each particular height:
#
# |idx|height|
# |---|------|
# |1 |5' 6"|
# |2 |6' 4"|
# |. |.|
# |. |.|
# |. |.|
# |100|5' 10"|
#
# We then are presented with distributions related to each of the groups heights, seen below:
# +
x_axis = np.arange(54, 80, 0.001)
men_mean = 70
women_mean = 64
fig = plt.figure(figsize=(8,5))
plt.plot(x_axis, norm.pdf(x_axis, men_mean, 2), c=sns.xkcd_rgb["red"])
plt.plot(x_axis, norm.pdf(x_axis, women_mean, 2), c=sns.xkcd_rgb["blue"])
plt.legend(["Men", "Women"], fontsize=16)
plt.xlabel("Height (inches)")
plt.show()
# -
# So, we know that our list of heights came from the two distributions above, but what if we knew that the list contained only 10 heights that were male, and 90 that were female? If we then were given a point that 5'8" we may be far more likely to classify it is as male (especially if we already had 10 other heights in our list that were greater than 6 feet. This concept of the base probability that a height comes from a given gaussian is encapsulated via the our term $\pi$, and can be seen below when we view the mixture distribution:
# +
x_axis = np.arange(54, 80, 0.001)
men_mean = 70
women_mean = 64
men_prior = 0.1
women_prior = 0.9
fig = plt.figure(figsize=(16,3.5))
plt.subplot(1, 3, 1)
men_gaussian = norm.pdf(x_axis, men_mean, 2)
women_gaussian = norm.pdf(x_axis, women_mean, 2)
plt.plot(x_axis, men_gaussian, c=sns.xkcd_rgb["red"])
plt.plot(x_axis, women_gaussian, c=sns.xkcd_rgb["blue"])
plt.legend(["Men", "Women"], fontsize=10)
plt.title("Men & Women Distrubutions", fontsize=16)
plt.yticks(fontsize=12)
plt.xticks(fontsize=12)
plt.subplot(1, 3, 2)
men_weighted = men_gaussian * men_prior
women_weighted = women_gaussian * women_prior
plt.plot(x_axis, men_weighted, c=sns.xkcd_rgb["red"], alpha=0.5)
plt.plot(x_axis, women_weighted, c=sns.xkcd_rgb["blue"], alpha=0.5)
plt.plot(x_axis, men_weighted + women_weighted, c=sns.xkcd_rgb["purple"])
plt.legend(["Men, Weighted", "Women, weighted", "Gaussian Mixture"], fontsize=10)
plt.title("Men & Women Weighted \n Distributions, Mixture", fontsize=16)
plt.yticks(fontsize=12)
plt.xticks(fontsize=12)
plt.subplot(1, 3, 3)
plt.plot(x_axis, men_weighted + women_weighted, c=sns.xkcd_rgb["purple"])
plt.legend(["Gaussian Mixture"], fontsize=10)
plt.title("Men & Women Weighted Mixture", fontsize=16)
plt.yticks(fontsize=12)
plt.xticks(fontsize=12)
plt.show()
# -
# Mathematically, we can describe the above as follows; we had information about the prior probabilities of men and women:
#
# $$P(men) = \frac{10}{100}$$
#
# $$P(women) = \frac{90}{100}$$
#
# Our generative process must clearly account for this, and we do so by introducing the **prior**, or **base** probability, $\pi$. For those familiar with the Bayesian paradigm, that is where the term _prior_ often comes from. The prior is the original probability that you update based on new information/data. Note, if we didn't know what the prior probabilities were, but we deemed both men and women to be equally like (as in the real world), we would choose a **uniform prior**, where $P(men) = 0.5$ and $P(women) = 0.5$.
#
# We can highlight our original equation of $P(x)$ with:
#
# $$P(x) = \overbrace{\pi_1}^\text{prior} \overbrace{N(x \mid \mu_1, \Sigma_1)}^\text{likelihood} + \pi_2 N(x \mid \mu_2, \Sigma_2) + \pi_3 N(x \mid \mu_3, \Sigma_3)$$
#
# Also, we must notice that there is a constraint here that all of the $\pi$'s have to sum to 1:
#
# $$1 = \int p(x)dx = \int \pi_1 N(x | \mu_1, \Sigma_1)dx + \pi_2 N(x | \mu_2, \Sigma_2)dx$$
# $$\pi_1 1 + \pi_2 1$$
#
# This is because $\pi$ is a distribution over all of the gaussians themselves, which actually brings us to a new viewpoint. Not only can we view $\pi_k$ as the probability prior probability of belonging to (aka being generated by) gaussian $k$, but we can also think of these gaussians as **hidden states**, and $\pi_k$ is the probability of being in a specific hidden state (a specific gaussian). Now, the minute that the word hidden appears you should immediately be thinking of **latent variables**, or variables are not observed. In this case, we don't actually get to see which gaussian (men or women) generated a specific point; we merely have a list of 100 unlabeled points, each holding a specific height. The actual gaussian that each point came from is _hidden_, or _latent_. From a tabular standpoint, it looks like:
#
# |idx|height|Gaussian (latent variable: gender)|
# |---|------|---|
# |1 |5' 6"|women|
# |2 |6' 4"|men|
# |. |.|.|
# |. |.|.|
# |. |.|.|
# |100|5' 10"|men|
#
# So, in this case our latent variable (the gaussian that generated each point) is _gender_. Another way of thinking of this is that we introduced a new random variable called $Z$. $Z$ represents which gaussian the data came from. So, we can say that:
#
# $$\pi_k = P(Z = k)$$
#
# Or in english the above can be rewritten as:
#
# $$k = \{ men, women\}$$
#
# $$\pi_{men} = P(Z = men)$$
#
# $$\pi_{women} = P(Z = women)$$
#
# What we are saying is that there is some hidden cause called $Z$ that we can't measure; in this case the hidden cause is gender (from a causal standpoint we can confidently state that gender effects height):
#
# <img src="https://drive.google.com/uc?id=1eTak1mkXE33OvACQ2DEjKeo7K8qzr_e9" width="300">
#
# Each of these $Z$'s (gender) is causing a corresponding gaussian to be generated (the male gaussian or the female gaussian), and all we can see in our list of 100 heights is the combined effects of those individual $Z$'s (the mixture). This is rather important because it puts GMM's into the framework of **expectation maximization**.
#
# ### Training a GMM
# When it comes to training a GMM, it is very much like the k-means algorithm. There are two steps that mirror what we saw with k-means.
#
# 1. **Calculate Responsibilites**<br>
# Each gaussian will be partially responsible for each point. $\gamma_k^{(n)}$ is the responsibility of the $k$th gaussian for generating the $n$th point. If $\pi_k$ is large here, then it will overtake the other gaussians, and this will be approximately equal to 1.
# $$\gamma_k^{(n)} = p(z^{(n)}|x) = \frac{\overbrace{\pi_k N (x^{(n)} | \mu_k, \Sigma_k) }^\text{Responsibility of k}}{\underbrace{\sum_{j=1}^K \pi_j N (x^{(n)} | \mu_j, \Sigma_j)}_\text{Total responsibility, normalizer}}$$
#
# For example, let's say that we were looking at the 27th point, with a height of 5'9", and we are trying to figure out the responsbility for the male gaussian:
#
# $$\gamma^{27}_{male} = P(Z^{27} \mid X = 5'9") =
# \frac{\pi_{male} N (x^{(27)} | \mu_{male}, \Sigma_{male})}{\sum_{j=1}^K \pi_j N (x^{(27)} | \mu_j, \Sigma_j)}$$
#
# Where again, $\gamma^{27}_{male}$ is the probability that the 27th data point was generated via the male gaussian, given that the data point, $x$ was 5'9". Again, we can think of the responsibilities as a form of _assignment_.
#
# 2. **Calculate model parameters of the gaussians**
# Now that we have just updated our responsibilities (assignments), we want to recalculate our model parameters, $\mu$, $\Sigma$, and $\pi$. The way that this is done is also similar to k-means, where we weight each samples influence on the parameter, by the responsibility. If that responsibility is small, then that $x$ matters less in the total calculation. We first calculate the total responsibility:
#
# $$N_k = \sum_{n=1}^N \gamma_k^{(n)}$$
#
# And then we can find $\mu_k$ by multiplying each point times it's responsibility from gaussian $k$, and finally dividing by the total responsibility.
#
# $$\mu_k = \frac{1}{N_k}\sum_{n=1}^N \gamma_k^{(n)} x^{(n)}$$
#
# $\Sigma_k$ is calculated in a similar fashion:
#
# $$\Sigma_k = \frac{1}{N_k} \sum_{n=1}^N \gamma_k^{(n)} (x^{(n)} - \mu_k)(x^{(n)} - \mu_k)^T$$
#
# Finally, $\pi$ is updated to simply be the total responsibility of the $k$th gaussian, divided by the total responsibility:
#
# $$\pi_k = \frac{N_k}{N}$$
#
# It is worth noting that we do not technically need both the variable $\pi$ and $\gamma$; I define each so as to allow for clarity. However, $\gamma$ is an $NxK$ matrix, and we see that $\pi$ is entirely dependent upon the column sum of $\gamma$, divided by the total number of points $N$. Specifically, if $\gamma$ were to be:
#
# $$
# \gamma =
# \begin{bmatrix}
# 0.6 & 0.4 \\
# 0.2 & 0.8 \\
# 0.3 & 0.7 \\
# 0.1 & 0.9 \\
# 0.9 & 0.1 \\
# \end{bmatrix}
# $$
#
# Then $\pi$ would simply be the column sum, divided by the number of rows:
#
# $$
# \pi =
# \begin{bmatrix}
# 0.138 0.28
# \end{bmatrix}
# $$
# # 2. GMM vs Soft K-Means
# Lets look at both **soft k-means** and **gmm's** side by side. Recall that for k-means the algorithm looked like:
#
# **Pseudocode**<br>
# ```
# Initialize m1...mk = random points in X
# While not converged:
# Step 1: Calculate cluster responsibilities
# ```
#
# $$r_k^{(n)} = \frac{exp\Big[-\beta d(m_k, x^{(n)})\Big]}{\sum_j exp \Big[-\beta d(m_j, x^{(n)})\Big]}$$
#
# ```
# Step 2: Recalculate means
# ```
# $$m_k = \frac{\sum_n r_k^{(n)}x^{(n)}}{\sum_n r_k^{(n)}}$$
#
#
# ## 2.1 Compare Steps
# Let's now compare the two steps of each training algorithm.
#
# > 1. We can see that the first step in both is to calculate the responsibilities.
# 2. The second step in both is to calculate the model parameters.
#
# We can see now why **k-means** looks for clusters of equal weight. It is because it has no $\pi$ variable. This is equivalent to saying that $\pi$ is uniform or equal to $\frac{1}{k}$. Note, this means that GMMs are limited in the same way as K means, since you still have to choose $k$.
#
# The second thing to notice is that k-means has this $\beta$ term, whereas GMMs have the full covariance $\Sigma$. This allows GMMs to have a lot more flexibility in the shape of its distribution. With K-means, since you only have this one $\beta$ it means that all of your clusters have to be spherical. With a full covariance matrix you can have any type of elliptical shape in any orientation. Notice, finally, that the equations for the mean are exactly the same.
#
# In conclusion, we can think of soft k-means as a GMM where each cluster has the same weight, and each cluster is spherical with the same radius.
# ---
#
# # 3. Gaussian Mixture Model in Code
# Notes:
# * Each sample ($N$ total) has a responsibility associated with it, which is just really the probability the it belongs to each specific cluster, of which there are $k$ total.
# * The idea of responsibility, $\gamma$, and overall gaussian weight, $\pi$, may be slightly confusing. The best way to think of it is that the responsibility is saying:
# > * "*the probability that a specific sample belongs to a specific cluster*"
# * While the overall gaussian weight is saying:
# > * "*based on all of the responsibilities across all of the samples, which clusters are responsible for containing most of the points?*"
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import multivariate_normal
# %matplotlib inline
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import multivariate_normal
def gmm(X, K, max_iter=20, smoothing=1e-2):
N, D = X.shape # Get number of rows and columns in X
M = np.zeros((K, D)) # Set means to zeros
R = np.zeros((N, K)) # Set the responsibilities to zeros.
C = np.zeros((K, D, D)) # Covariance matrix, 3 dimensional
pi = np.ones(K) / K # Uniform distribution
# Iterate over all K gaussians
for k in range(K):
M[k] = X[np.random.choice(N)] # Set the means to random points of X
C[k] = np.diag(np.ones(D))
costs = np.zeros(max_iter)
weighted_pdfs = np.zeros((N, K)) # Store pdf values ---> Numerator of responsibility, gamma
for i in range(max_iter):
# --------------- Step 1: Calculate Responsibilities ---------------
for k in range(K): # Iterate through all K gaussians
for n in range(N): # Iterate through all N data points
weighted_pdfs[n, k] = pi[k]*multivariate_normal.pdf(X[n], M[k], C[k])
for k in range(K):
for n in range(N):
R[n, k] = weighted_pdfs[n, k] / weighted_pdfs[n, :].sum()
# ---------- Step 2: Re-Calculate parameters (pi, mu, cov) ----------
for k in range(K):
Nk = R[:, k].sum() # sum of all responsibilities for specific gaussian k
pi[k] = Nk / N
M[k] = R[:, k].dot(X) / Nk
# Regularization for covariance
C[k] = np.sum(R[n, k]*np.outer(X[n] - M[k], X[n] - M[k]) for n in range(N)) / Nk + np.eye(D)*smoothing
# Calculate log likelihood!!!
costs[i] = np.log(weighted_pdfs.sum(axis=1)).sum()
if i > 0:
if np.abs(costs[i] - costs[i - 1]) < 0.1:
break
fig, ax = plt.subplots(figsize=(8, 5))
plt.plot(costs)
plt.title("Costs")
plt.show()
random_colors = np.random.random((K, 3))
colors = R.dot(random_colors)
fig, ax = plt.subplots(figsize=(8, 5))
plt.scatter(X[:, 0], X[:, 1], c=colors)
plt.title(r"Learned Mixture/Clusters, $K = 3$")
plt.show()
print("Learned pi:\n", pi)
print("Learned means:\n", M)
print("Learned covariances:\n", C)
return R
def main():
# Create 3 Gaussian distributed clusters
D = 2
s = 4
mu1 = np.array([0, 0])
mu2 = np.array([s, s])
mu3 = np.array([0, s])
N = 2000 # Number of samples
X = np.zeros((N, D))
X[:1200, :] = np.random.randn(1200, D)*2 + mu1 # Covariance = 2
X[1200:1800, :] = np.random.randn(600, D) + mu2 # Covariance = 1
X[1800:, :] = np.random.randn(200, D)*0.5 + mu3 # Covariance = 0.5
gaussian_1 = X[:1200, :]
gaussian_2 = X[1200:1800, :]
gaussian_3 = X[1800:, :]
fig, ax = plt.subplots(figsize=(8, 5))
plt.scatter(gaussian_1[:, 0], gaussian_1[:, 1], c="red", alpha=0.5)
plt.scatter(gaussian_2[:, 0], gaussian_2[:, 1], c="blue", alpha=0.5)
plt.scatter(gaussian_3[:, 0], gaussian_3[:, 1], c="green", alpha=0.5)
plt.title(r"Original Mixture/Clusters, $K = 3$")
plt.show()
K = 3
gmm(X, K)
if __name__ == "__main__":
main()
# -
# # 4. Singular Covariance Problem
# To get a better idea for why this may be a problem, lets think about a one dimensional gaussian for a second. Imagine that all of the points are close together, so that they variance is almost 0. Or, consider what happens if you are trying to find the variance of just 1 point. In the gaussian formula, we divide by the variance. This means that if the variance is 0, then we are dividing by 0, which results in a singularity.
#
#
# You can think of inverting the matrix as taking 1 over it, since something times 1 over itself will be 1, or our identity.
# $$A * \frac{1}{A} = Identity$$
#
# So in multiple dimensions, it is the same problem. If your covariance is too small, the covariance will approach infinity. This is yet another disadvantage of falling into local minimum. How can we fix this problem?
#
# ## 4.1 Diagonal Covariance
# One solution to fix this problem is to use what is called a diagonal covariance. Diagonal covariance matrices are very useful, not only for avoiding the singularity problem, but also for speeding up our computation. When you have a diagonal covariance, it is very easy to take the inverse. You just take every element that is not 0, which will be along the diagonal, and you then invert it; i.e. you take 1 over that element.
#
# <img src="https://drive.google.com/uc?id=1g2JKWHMsdOk-q6O5IUM0PyBbFnNxWpxk">
#
# The assumption that you are making when using a diagonal covariance is that each of your dimensions is independent.
#
# $$\Sigma_{ij} = E\Big[(x_i - \mu_i)(x_j - \mu_j)\Big]$$
# And we can then write (because $x_i$ and $x_j$ are independent we can split the expectations):
# $$\Sigma_{ij} = E\Big[(x_i - \mu_i)\Big]E\Big[(x_j - \mu_j)\Big]$$
# $$\Sigma_{ij} = \Big(E(x_i) - \mu_i\Big)\Big(E(x_j) - \mu_j\Big)$$
# $$\Sigma_{ij} = (\mu_i - \mu_i)(\mu_j - \mu_j)$$
# $$\Sigma_{ij} = 0$$
#
# From the above you can see how we eventually end up at 0. You can think of the diagonal matrix as a means of regularization-you are making your model simpler by using less parameters.
#
# ## 4.2 Spherical Gaussian
# Sometimes even when you use diagonal covariance, you still get singularities. In that case you may want to use a spherical gaussian, where we use the same covariance along every dimension. This is even less computationaly expensive.
# ---
#
# <br>
# # 5. Kernel Density Estimation
# Kernel Density Estimation is just the fitting of a probability distribution with kernels. Before we really dive into this, let's first just talk about what the easiest solution to this problem is. the simplest solution is to just take the frequency estimate-a **histogram**! This would work perfectly for discrete probabilities and would be the maximum likelihood estimate.
#
# However, if our data was continuous (as the data we have been working with so far has been), then the easiest way it to use a **gaussian mixture model**! In fact, using a gaussian mixture model we are inherently doing kernel density estimation, where the gaussian is the kernel. However, we will have the problem that we can only have gaussian looking shapes in your distribution. This is the same problem we have with k-means, in we have to determine the correct number of components to use. One strategy would be to look at the distribution of the data, and to look at the number of peaks seen in the histogram.
#
# Another method is to use gaussians with all the same variance, set the number of gaussians equal to the number of points, and use the GMM algorithm on that.
#
# $$P(x) = \frac{1}{N}\sum_{i=1}^N \frac{1}{(2\pi h^2)^\frac{D}{2}} exp\Big(-\frac{||x - x_n||^2}{2h^2}\Big)$$
#
# Here we call **h** the window size, and you would need to play with h in order to find the ideal size for your distribution.
# ---
#
# <br>
# # 6. Expectation Maximization
# We are now going to take an abstract look at the Gaussian Mixture Model, and create a framework called **Expectation Maximization**, aka the **E-M Algorithm**. This algorithm generalizes some important concepts that we have talked about, such as latent variables, coordinate descent, and increasing our likelihood guarantee.
#
# ### 6.1 Maximum Likelihood
# The first important thing that we should realize is that we still start off trying to do maximum likehood. We have some data $X$ and some model $\theta$, and we are trying to maximize the probability of observing our data, $X$, given $\theta$, which generally defines are gaussian distribution.
#
# $$Maximize \rightarrow \; P(X | \theta) \; or \; log P(X | \theta) = L(\theta)$$
#
# By introducing hidden variables called $Z$, we can imagine that $P(X | \theta)$ is actually a marginalized distribution over $P(X, Z | \theta)$:
#
# $$P(X | \theta) = \sum_Z P(X, Z | \theta)$$
#
# We can expand that to get:
#
# $$P(X | \theta) = \sum_Z P(X, Z | \theta) = \sum_Z P(X | Z, \theta)P(Z|\theta)$$
#
# ### 6.2 EM Algorithm: 2 steps
# As we know, in the EM algorithm there are two main steps; one where we adjust $Z$ and one where we adjust $\theta$.
#
# #### E-step:
# > We can think of the E-step as finding the distribution of Z given X and the current setting of theta:
#
# $$P(Z | X, \theta_n)$$
#
# #### M-step:
# > Then you can think of the M-step as finding the best $\theta$ that maximizes the joint distribution of $log P(X, Z | \theta)$. However, instead of maximizing this directly, we are maximizing the expected value of this over the distribution we found in the previous step, which was the distribution $P(Z | X, \theta_n)$ :
#
# $$Maximize \rightarrow E \Big[log P(X, Z | \theta)\Big]$$
# + active=""
# <script>
# function code_toggle() {
# if (code_shown){
# $('div.input').hide('500');
# $('#toggleButton').val('Show Code')
# } else {
# $('div.input').show('500');
# $('#toggleButton').val('Hide Code')
# }
# code_shown = !code_shown
# }
#
# $( document ).ready(function(){
# code_shown=false;
# $('div.input').hide()
# });
# </script>
# <form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>
# -
|
Machine_Learning/04-Unsupervised_Learning_Cluster_Analysis-04-Cluster-Analysis-Gaussian-Mixture-Models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from evaluate import evaluate
from unet import UNet
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
import torch
from PIL import Image
from torchvision import transforms
net = UNet(n_channels=3, n_classes=2, bilinear=False)
# -
from torchsummary import summary
net.to(device='cuda')
summary(net, (3, 224, 224))
net.load_state_dict(torch.load('checkpoints/checkpoint_epoch_224x224_33.pth'))
net.eval()
def preprocess(pil_img, scale, is_mask):
w, h = pil_img.size
print(w,h)
newW, newH = int(scale * w), int(scale * h)
assert newW > 0 and newH > 0, 'Scale is too small, resized images would have no pixel'
pil_img = pil_img.resize((newW, newH))
img_ndarray = np.asarray(pil_img)
if img_ndarray.ndim == 2 and not is_mask:
img_ndarray = img_ndarray[np.newaxis, ...]
elif not is_mask:
img_ndarray = img_ndarray.transpose((2, 0, 1))
if not is_mask:
img_ndarray = img_ndarray / 255
return img_ndarray
filename = '../dfdc_deepfake_challenge/dataset_new_1/testing/crops/crops_testing/orrltuoxgt_1.png'
scale=1.0
img = Image.open(filename).resize((224,224),Image.NEAREST)
img = torch.from_numpy(preprocess(img, scale, is_mask=False))
# img = torch.as_tensor(img.copy()).float().contiguous()?\
img = img.unsqueeze(0)
img = img.to(device='cpu', dtype=torch.float32)
pred, classifi = net(img)
pred.shape
probs = F.softmax(pred, dim=1)[0]
tf = transforms.Compose([
transforms.ToPILImage(),
transforms.ToTensor()
])
full_mask = tf(probs.cpu()).squeeze()
full_mask
final = F.one_hot(full_mask.argmax(dim=0), 2).permute(2, 0, 1).numpy()
final.shape
final[0].shape
from matplotlib import pyplot as plt
plt.imshow(final[1], interpolation='nearest')
plt.show()
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img = mpimg.imread('../dfdc_deepfake_challenge/dataset_new_1/testing/crops/crops_testing/orrltuoxgt_1.png')
imgplot = plt.imshow(img)
plt.show()
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img = mpimg.imread('../dfdc_deepfake_challenge/dataset_new_1/testing/masks/masks_testing/orrltuoxgt_1.gif')
imgplot = plt.imshow(img)
plt.show()
classifi
|
Pytorch-UNet-classification-segmentation/fyp_predict.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
import pandas as pd
import numpy as np
import os
import re
import matplotlib.pyplot as plt
from pandas.plotting import scatter_matrix
from collections import defaultdict
from IPython.display import display
# %matplotlib inline
# # Read model predictions
path = "./running/external/transformer_hier++_001/"
df = pd.read_csv(os.path.join(path, "error_greedy_test.tsv"), delimiter="\t")
df_stats = pd.read_csv(os.path.join(path, "stats_greedy_test.tsv"), delimiter="\t")
df.shape, df_stats.shape
df.head()
df.describe()
# # Correlation b/w Metrics
# ## [DEFINE] Context Length + Bleu bins
df['context_length'] = df.apply(lambda r: len(r['context'].split('<br>')), 1)
df['bleu_bin'] = df.apply(lambda r: r['bleu']//10, 1)
df.sample(10)
df.iloc[2099]['generated']
df.iloc[2099]['gold']
# ## Bleu vs. Context Length/turns
#
# Observing slight decrease in bleu with context length.
# - Would be interesting to compare high context length performance across models
# - Check how many samples are availble for training for different context length
df.boxplot(column='bleu', by='context_length', figsize=(8,6))
df.boxplot(column='bleu_bin', by='context_length', figsize=(8,6))
# ## [DEFINE] num_entities
regex_entity = r'\[([a-z\_\-]+)\]'
df['gold_entities'] = df.apply(lambda r: re.findall(regex_entity, r['gold']), 1)
df['num_entities'] = df.apply(lambda r: len(r['gold_entities']), 1)
df.boxplot(column='f1_entity', by='num_entities', figsize=(8,6))
# ## [EXP] Scatter all pairs
_ = scatter_matrix(df[['bleu', 'loss', 'f1_entity', 'context_length', 'num_entities']], alpha=0.2, diagonal='kde', figsize=(12,12))
# ## [EXP] Entitywise Avg. f1-entity
entity_log = {
'entity':[],
'loss':[],
'bleu':[],
'f1_entity':[],
'context_length':[]
}
for _, row in df.iterrows():
for e in row['gold_entities']:
entity_log['entity'].append(e)
entity_log['loss'].append(row['loss'])
entity_log['bleu'].append(row['bleu'])
entity_log['f1_entity'].append(row['f1_entity'])
entity_log['context_length'].append(row['context_length'])
df_entity_log = pd.DataFrame.from_dict(entity_log)
df_entity_log.boxplot(column='f1_entity', by='entity', figsize=(15,8))
plt.xticks(rotation=90)
df_entity_log.boxplot(column='loss', by='entity', figsize=(15,8))
plt.xticks(rotation=90)
# ## [DEFINE] pred_entities
df['pred_entities'] = df.apply(lambda r: re.findall(regex_entity, r['generated']), 1)
df['num_pred_entities'] = df.apply(lambda r: len(r['pred_entities']), 1)
df.sample(4)
# # Extracting Samples
# ## [SAMPLES] By Loss
def print_row(r):
print(f"\n[CONTEXT]")
print("{}".format('\n'.join(r['context'].split('<br>')[-5:])))
print(f"\n[GOLD]: {r['gold']}")
print(f"\n[GEN]: {r['generated']}")
for _, row in df.nlargest(2, ['loss']).iterrows():
print_row(row)
for _, row in df.nsmallest(2, ['loss']).iterrows():
print_row(row)
for _, row in df.nsmallest(4, ['f1_entity']).iterrows():
print_row(row)
for _, row in df.nlargest(50, ['context_length']).sample(2).iterrows():
print_row(row)
for _, row in df.nlargest(5, ['num_entities']).iterrows():
print_row(row)
# ## Compare Target and Predicted Entities
ent_pred_ledger = defaultdict(list)
for _, row in df.iterrows():
gold = set(row['gold_entities'])
gen = set(row['pred_entities'])
for e in gold - gen:
# These didn't get predicted, lost count
ent_pred_ledger[e].append(-1)
for e in gen - gold:
# These weren't supposed to get predicted, excess count
ent_pred_ledger[e].append(+1)
for e in gen.intersection(gold):
# These are ok
ent_pred_ledger[e].append(0)
df_ent_pred_ledger = {}
for e in ent_pred_ledger:
preds = np.array(ent_pred_ledger[e])
# print(f"\n{e.upper()}")
fp = (preds == 1).sum()
fn = (preds == -1).sum()
tp = (preds == 0).sum()
# print(f" TP: {tp}, {100*tp/(fp+fn+tp):0.2F}")
# print(f" FP: {fp}, {100*fp/(fp+fn+tp):0.2F}")
# print(f" FN: {fn}, {100*fn/(fp+fn+tp):0.2F}")
df_ent_pred_ledger[e] = {
'gold_appear': fn + tp,
'tpr': 100*tp/(fp+fn+tp),
'fpr': 100*fp/(fp+fn+tp),
'fnr': 100*fn/(fp+fn+tp),
'tp': tp,
'fp': fp,
'fn': fn,
}
df_ent_pred_ledger = pd.DataFrame.from_dict(df_ent_pred_ledger, orient='index')
df_ent_pred_ledger = df_ent_pred_ledger.sort_values('gold_appear', ascending=False)
df_ent_pred_ledger.sort_values('tpr', ascending=False)
# ## Macro-Avg Entity Prediction Success at different context lengths
def proc_row(row):
gold = set(row['gold_entities'])
gen = set(row['pred_entities'])
L = len(gold)
entity_recall = len(gen.intersection(gold))/L if L > 0 else 0
entity_prec = len(gen.intersection(gold))/len(gen) if len(gen) > 0 else 0
entity_f1 = 2*entity_prec*entity_recall/(entity_recall+entity_prec) if (entity_recall+entity_prec) >0 else 0
return entity_prec, entity_recall, entity_f1
df['entity_prec'] = df.apply(lambda r: proc_row(r)[0], 1)
df['entity_recall'] = df.apply(lambda r: proc_row(r)[1], 1)
df['entity_f1'] = df.apply(lambda r: proc_row(r)[2], 1)
df[(df['num_entities'] > 0)].describe()
# ## [EXP] Entity prec/recall/f1 vs. Context Length
df.groupby('context_length')['entity_prec'].mean().plot()
plt.grid()
df.groupby('context_length')['entity_recall'].mean().plot()
df.groupby('context_length')['bleu'].mean().plot()
df.groupby('context_length')['entity_f1'].mean().plot()
plt.grid()
plt.title("Entity-F1 Score")
df.groupby('context_length')['file'].count().plot()
plt.grid()
plt.title("Number of samples in test set")
# ## [RESULT] Correctness of entities predicted
#
# Our model is more accurate in the entities being mentioned also uses appropriate number of entities in the generated response. Our model uses almost the same or sometimes less numbers of entities in its responses compared to Marco as is more accurate on avg.
df.groupby('context_length')[['num_pred_entities', 'num_entities']].sum().plot()
plt.grid()
df.loc[df['num_entities']>0, ['entity_prec']].hist(bins=20)
plt.ylim(0, 3600)
plt.title("Histogram of Entity Precision")
df.loc[df['num_entities']>0, ['entity_recall']].hist(bins=20)
plt.ylim(0, 3600)
plt.title("Histogram of Entity Recall")
# ## [EXP] People vs. Number of Entities
df.groupby('num_entities')['file'].count().plot()
df.groupby('num_entities')[['entity_recall', 'entity_prec', 'entity_f1']].mean().plot()
plt.grid()
df.groupby('num_entities')[['bleu']].mean().plot()
df.groupby('num_entities')[['num_pred_entities']].mean().plot(figsize=(5,5))
plt.grid()
plt.xlim((0, 15))
plt.ylim((0, 15))
# ## [DEFINE] Avg. Response Length
response_lengths = df.apply(lambda x: len(x["generated"].split()), 1)
print("Mean response length:", response_lengths.mean())
response_lengths.hist(bins=20)
plt.xlim(0, 50)
# # Process Dialog level Stats (Inform/Success)
# +
df_temp = df.groupby('file')[['context_length']].max().reset_index()
df_temp['file'] = df_temp.apply(lambda r: r['file']+'.json', 1)
df_combo = pd.merge(df_temp, df_stats, how='inner', on='file')
df_combo.groupby('context_length')['context_length'].count().plot()
plt.ylabel('Number of instances')
df_combo.groupby('context_length')[['success', 'match']].mean().plot()
plt.grid()
df_combo.groupby('context_length')[['success', 'match']].median().plot()
plt.grid()
# +
df_temp = df.groupby('file')[['num_entities']].sum().reset_index()
df_temp['file'] = df_temp.apply(lambda r: r['file']+'.json', 1)
df_combo = pd.merge(df_temp, df_stats, how='inner', on='file')
df_combo.groupby('num_entities')['num_entities'].count().plot()
plt.ylabel('Number of instances')
df_combo.groupby('num_entities')[['success', 'match']].mean().plot()
plt.grid()
df_combo.groupby('num_entities')[['success', 'match']].median().plot()
plt.grid()
# +
df_temp = df.groupby('file')[['num_pred_entities']].sum().reset_index()
df_temp['file'] = df_temp.apply(lambda r: r['file']+'.json', 1)
df_combo = pd.merge(df_temp, df_stats, how='inner', on='file')
df_combo.groupby('num_pred_entities')['num_pred_entities'].count().plot()
plt.grid(); plt.xlim(0, 40);
plt.ylabel('Number of instances')
df_combo.groupby('num_pred_entities')[['success', 'match']].mean().plot()
plt.grid()
df_combo.groupby('num_pred_entities')[['success', 'match']].median().plot()
plt.grid()
# -
# ## Match/Success pairwise stats
for key, g in df_stats.groupby(['match', 'success']):
print(key, g.shape[0], 'rows')
# ## Find dialogs by match/success constraints:
df2 = df.copy()
df2['file'] = df2.apply(lambda r: r['file']+'.json', 1)
df_full = df_stats.merge(df2, on=['file'])
for _, r in df_stats.iterrows():
file = r['file']
match = r['match']
success = r['success']
if match == 1 and success == 0:
piece = df2[df2['file'] == file]
showme = ['gold_entities', 'num_entities', 'pred_entities', 'num_pred_entities', 'entity_prec', 'entity_recall']
print(f"{file}, Match: {match}, Success: {success}")
for _, rc in piece.iterrows():
print('-------------------------------')
display(piece.loc[_, showme])
print('-------------------------------')
# print('[CON]', rc['context'])
print('[GOLD]', rc['gold'])
print('[GEN]', rc['generated'])
break
|
HIER/error_reader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing extracting OSM data using Osmium
# +
import os, sys, time, importlib
import osmnx
import geopandas as gpd
import pandas as pd
import networkx as nx
import numpy as np
sys.path.append("../../../GOSTNets")
import GOSTnets as gn
# pip install osmium
# import osmium, logging
# import shapely.wkb as wkblib
from shapely.geometry import LineString, Point
import time
# -
# This is a Jupyter Notebook extension which reloads all of the modules whenever you run the code
# This is optional but good if you are modifying and testing source code
# %load_ext autoreload
# %autoreload 2
from GOSTnets.load_traffic import *
# vavuniya is a city in northern Sri Lanka, and this will be a small area for testing
# set file
some_file = './colombo.osm.pbf'
vavuniya = OSM_to_network(some_file)
vavuniya.roads_raw
vavuniya.nodes_raw
vavuniya.nodes_raw.loc[vavuniya.nodes_raw.osm_id == 4082624672]
vavuniya.nodes_raw.loc[vavuniya.nodes_raw.osm_id == 7369784272]
# +
# vavuniya.nodes_raw.drop_duplicates(subset ="osm_id",
# keep = False, inplace = True)
# -
#vavuniya.nodes_raw.isnull()
np.where(pd.isnull(vavuniya.nodes_raw))
vavuniya.roads_raw.loc[vavuniya.roads_raw.stnode == 4082624672]
vavuniya.roads_raw
vavuniya.nodes_raw[vavuniya.nodes_raw['geometry'] == ''].index
vavuniya.nodes_raw.head()
vavuniya.apply_traffic_speeds_to_roads_raw("./osm/1233300-Asia-Colombo.csv", "./osm/1233302-Asia-Colombo.csv" )
vavuniya.generateRoadsGDF(verbose = True)
len(vavuniya.roads_raw)
len(vavuniya.roadsGDF)
vavuniya.nodesGDF.loc[vavuniya.nodesGDF.osm_id == 60796641]
len(vavuniya.nodesGDF)
vavuniya.nodesGDF[125693:125703]
vavuniya.nodesGDF.loc[vavuniya.nodesGDF.osm_id == 3727426631]
gdfnodes.loc[gdfnodes.node_ID == 3727426631]
vavuniya.initialReadIn()
# ### Now still need to clean graph (which includes simplifying the edges). By the way during the clean graph function it sums of the lengths if it removes uneeded nodes. Then travel times still need to be applied.
vavuniya
vavuniya.network
len(vavuniya.network.nodes)
vavuniya.network.nodes[3727426633]
vavuniya.network.nodes[3727426631]
gn.example_edge(vavuniya.network, 8)
gn.example_node(vavuniya.network, 8)
# create a node gdf from graph
gdfnodes = gn.node_gdf_from_graph(vavuniya.network)
vavuniya.network.nodes[127658543]
gdfnodes
gdfnodes.loc[gdfnodes.node_ID == 127658543]
gn.save(vavuniya.network,'vavuniya_unclean','./', pickle = True, edges = True, nodes = True)
|
Implementations/FY21/ACC_mapbox_traffic/test_integrating_traffic_into_GOSTnets2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementing Feature Union
# Using the given custom transformer, `StartingVerbExtractor`, add a feature union to your pipeline to incorporate a feature that indicates with a boolean value whether the starting token of a post is identified as a verb.
import nltk
nltk.download(['punkt', 'wordnet', 'averaged_perceptron_tagger'])
# +
import re
import numpy as np
import pandas as pd
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.metrics import confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
# -
from custom_transformer import StartingVerbExtractor
url_regex = 'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+'
# ### Build your pipeline to have this structure:
# - Pipeline
# - feature union
# - text pipeline
# - count vectorizer
# - TFIDF transformer
# - starting verb extractor
# - classifier
def model_pipeline():
pipeline = Pipeline(
[
('union', FeatureUnion(
[
(
'text', Pipeline(
[
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer())
]
)
),
(
'starting_verb',
StartingVerbExtractor()
),
]
)),
('clf', RandomForestClassifier()),
]
)
return pipeline
# ### Run program to test
# +
def load_data():
df = pd.read_csv('corporate_messaging.csv', encoding='latin-1')
df = df[(df["category:confidence"] == 1) & (df['category'] != 'Exclude')]
X = df.text.values
y = df.category.values
return X, y
def tokenize(text):
detected_urls = re.findall(url_regex, text)
for url in detected_urls:
text = text.replace(url, "urlplaceholder")
tokens = word_tokenize(text)
lemmatizer = WordNetLemmatizer()
clean_tokens = []
for tok in tokens:
clean_tok = lemmatizer.lemmatize(tok).lower().strip()
clean_tokens.append(clean_tok)
return clean_tokens
def display_results(y_test, y_pred):
labels = np.unique(y_pred)
confusion_mat = confusion_matrix(y_test, y_pred, labels=labels)
accuracy = (y_pred == y_test).mean()
print("Labels:", labels)
print("Confusion Matrix:\n", confusion_mat)
print("Accuracy:", accuracy)
def main():
X, y = load_data()
X_train, X_test, y_train, y_test = train_test_split(X, y)
model = model_pipeline()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
display_results(y_test, y_pred)
main()
# -
|
01_Data_Engineering/02_Machine_Learning_Pipelines/feature_union_practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# <b>pandas</b> is a fast, powerful, flexible and easy to use open source data analysis and manipulation tool,
# built on top of the Python programming language.
# <br>
# <br>
# <br>
# <b>Interesting Read</b> : [mlcourse.ai : EDA with Pandas](https://mlcourse.ai/articles/topic1-exploratory-data-analysis-with-pandas/)
#
# <img src='https://habrastorage.org/webt/ia/m9/zk/iam9zkyzqebnf_okxipihkgjwnw.jpeg' width='300' align='left'>
# <br>
# <br>
# <br>
# <br>
# <br>
# <b>Much of this Notebook has been adopted from `pandas` docs</b>
# + [markdown] heading_collapsed=true
# # Imports
# + hidden=true
import pandas as pd
from datetime import datetime
from pytz import all_timezones
# + hidden=true
pd.__version__
# + hidden=true
# ?pd
# + [markdown] heading_collapsed=true
# # Load and Explore the Dataset
# + hidden=true
root_path = '../'
raw_datapath = root_path+'Raw Data/'
prepared_datapath = root_path+'Prepared Data/'
britannia_datapath = raw_datapath+'BRITANNIA.NS.csv'
mpc61_datapath = raw_datapath+'MPC61.txt'
# + [markdown] heading_collapsed=true
# # Pandas DataStructures
#
# `pandas` creates and stores data in rectangular format.
#
# On a broad stroke there are majorly two forms of a datatype in `pandas` :-
# - [Series](https://pandas.pydata.org/docs/user_guide/dsintro.html#series) : "Series is a one-dimensional labeled array capable of holding any data type (integers, strings, floating point numbers, Python objects, etc.)."
# - [Dataframe](https://pandas.pydata.org/docs/user_guide/dsintro.html#dataframe) : "DataFrame is a 2-dimensional labeled data structure with columns of potentially different types. You can think of it like a spreadsheet or SQL table, or a dict of Series objects."
# + [markdown] heading_collapsed=true hidden=true
# ## Series
# + [markdown] hidden=true
# <b>Series - Generating One</b>
# + hidden=true
data_size=20
sdata=np.random.normal(20, scale=5.0, size=(data_size,))
sindex=np.arange(20)
pd_series = pd.Series(sdata,
index=sindex)
pd_series
# + [markdown] hidden=true
# <b>Series - Indicing</b>
# + hidden=true
pd_series.iloc[15]
# + hidden=true
np.arange(10)[5:]
# + [markdown] hidden=true solution2="shown" solution2_first=true
# **Q)** Get all the values in the series after indices 15, how?
# + hidden=true solution2="shown"
pd_series.loc[14:18]
# + hidden=true
#----------YOUR SOLUTION-----------#
#....
# + [markdown] hidden=true
# <b>Series - Query</b>
# + hidden=true
pd_series[pd_series>20]
# + [markdown] hidden=true
# <b>Series - Stats</b>
# + hidden=true
pd_series.describe()
# + [markdown] hidden=true
# <b>Series - To a Dictionary</b>
# + hidden=true
pd_series.to_dict()
# + [markdown] hidden=true
# <b>Series - Numpy Like Vectorized Operations</b>
# + hidden=true
pd_series*pd_series
# + [markdown] hidden=true
# <b>Series - Giving it a name</b>
# + hidden=true
pd_series.name = 'Random Normal Series'
pd_series
# + [markdown] hidden=true
# <b>Series - To a DataFrame</b>
# + hidden=true
pd_series.to_frame()
# + [markdown] heading_collapsed=true hidden=true
# ## DataFrame
# + hidden=true
# What will happen to the name that we gave to the series above?
dfdata_dict = {'one':pd_series,
'two':pd_series*2}
datadf = pd.DataFrame(dfdata_dict)
datadf.head()
# + [markdown] hidden=true
# <b>DataFrame - Creating One</b>
#
# DataFrame can be creating by feeding any of the following:
#
# - Dict of 1D ndarrays, lists, dicts, or Series
# - 2-D numpy.ndarray
# - Structured or record ndarray
# - A Series
# - Another DataFrame
# + hidden=true
countrytemp_data = {'Country':['Brazil', 'India', 'India', 'Germany', 'China', 'Zambia'],
'City':['Brasília', 'New Delhi', 'Kashmir', 'Berlin', 'Beijing', 'Lusaka'],
'AverageTemperature':[30.1, 34.3, 22.4, 19.9, 26.2, 30.3],
'Humidity':[0.65, 0.67, 0.49, 0.44, 0.45, 0.76]}
# + hidden=true
temperatureDf = pd.DataFrame(countrytemp_data,
index=['a', 'b', 'c', 'd', 'e', 'f'])
temperatureDf
# + [markdown] hidden=true
# <b>DataFrame - Indicing</b>
# + hidden=true
temperatureDf.index
# + hidden=true
temperatureDf[temperatureDf.index>'b']
# + hidden=true
temperatureDf.iloc[2:4,:]
# + hidden=true
temperatureDf.iloc[:,1:3]
# + hidden=true
temperatureDf['b':'d']
# + [markdown] hidden=true solution2="shown" solution2_first=true
# **Q)** I want the indices from 'b' to 'd', and first two columns 'Country' & 'City'
# + hidden=true solution2="shown"
temperatureDf.loc['b':'d'].iloc[:,:2]
# + hidden=true
#----------YOUR SOLUTION-----------#
#....
# + [markdown] hidden=true
# <b>DataFrame - Stats</b>
# + hidden=true
temperatureDf.describe()
# + hidden=true
temperatureDf.AverageTemperature.min()
# + hidden=true
temperatureDf.Humidity.min(), temperatureDf.Humidity.max()
# + [markdown] hidden=true
# <b>DataFrame - Plot</b>
# + hidden=true
temperatureDf
# + hidden=true
temperatureDf.plot()
# + hidden=true
temperatureDf.set_index('City').plot(subplots=True)
# + [markdown] hidden=true
# <b>DataFrame - Groupby Operations</b>
# + hidden=true
temperatureDf
# + hidden=true
temperatureDf.groupby(['Country']).size()
# + hidden=true
# The result of groupby operation is usually in Multiindex Series Format
temperatureDf.groupby(['Country']).AverageTemperature.mean()
# + hidden=true
for egidx, eg in temperatureDf.groupby(['Country']):
print(egidx)
print(eg)
print()
# + code_folding=[] hidden=true
# Why did it only show one Value?
# temperatureDf.groupby(['Country']).AverageTemperature.nth(0)
# temperatureDf.groupby(['Country']).AverageTemperature.nth(1)
temperatureDf.groupby(['Country']).AverageTemperature.nth(-1)
# + [markdown] heading_collapsed=true hidden=true
# ## Reading a file to Pandas DataFrame
# + [markdown] hidden=true
# <b>Reading a .csv file</b>
# + hidden=true
britannia_datapath
# + hidden=true
britannia_data = pd.read_csv(britannia_datapath, index_col=0, parse_dates=True)
britannia_data.head()
# + hidden=true
britannia_data.dtypes
# + hidden=true
britannia_data.index
# + [markdown] hidden=true
# <b>Reading a .txt file</b>
# + hidden=true
pd.read_fwf(mpc61_datapath)
# + hidden=true
mpc61_datapath
# + hidden=true
pd.read_csv(mpc61_datapath)
# + hidden=true
mpc61_data = pd.read_fwf(mpc61_datapath,
skiprows=range(50),
names=['RUN', 'WAFER', 'PROBE',
'MONTH', 'DAY', 'OP',
'TEMP', 'AVERAGE', 'STDDEV'])
mpc61_data.head()
# + [markdown] heading_collapsed=true
# # Pandas Datetime Operations
#
# "Pandas builds upon `dateutil`, `datetime` & `numpy.datetime64` the tools just discussed to provide a Timestamp object, which combines the ease-of-use of datetime and dateutil with the efficient storage and vectorized interface of numpy.datetime64. From a group of these Timestamp objects, Pandas can construct a DatetimeIndex that can be used to index data in a Series or DataFrame" - [Jake-Python Data Science Handbook](https://jakevdp.github.io/PythonDataScienceHandbook/03.11-working-with-time-series.html#Dates-and-times-in-pandas:-best-of-both-worlds)
# + [markdown] hidden=true
# <b>Generating a DatetimeIndex</b>
#
# Look at the `dtype` and `freq`
# + hidden=true
datetime.now().strftime('%Y-%m-%d')
# + hidden=true
current_date = pd.to_datetime(datetime.now().strftime('%Y-%m-%d'))
roll_dates = current_date+pd.to_timedelta(range(10), 'D')
current_date, type(current_date), roll_dates
# + [markdown] hidden=true
# <b>How can we set the frequency? And is it possible to infer the frequency intrinsically?</b>
# + hidden=true
pd.infer_freq(roll_dates)
# + hidden=true
roll_dates.freq='D'
roll_dates
# + hidden=true
roll_dates.freq = 'M'
# + [markdown] hidden=true
# <b>Lets make a Time Series with Dates as index</b>
# + hidden=true
sindex = ['2020-01-01', '2020-01-15', '2020-01-31',
'2020-02-01', '2020-02-15', '2020-02-28',
'2020-03-01', '2020-03-15', '2020-03-31',
'2021-01-01', '2021-01-15', '2021-01-31',
'2021-02-01', '2021-02-15', '2021-02-29',
'2021-03-01', '2021-03-15', '2021-03-31']
dt_series = pd.Series(np.arange(18), index=sindex)
dt_series
dt_series.index
# + [markdown] hidden=true
# <b>Lets try to slice the series, and retrieve data only of year `2020`</b>
# + hidden=true
dt_series['2020']
# + [markdown] hidden=true
# <b>But why, cant i index this series using just the year, why didnt pandas understand that?</b>
# + hidden=true
dt_series.index
# + hidden=true
dt_series.index = pd.to_datetime(dt_series.index)
# + [markdown] hidden=true
# So basically, pandas is trying to parse the date `2021-02-29`, but while doing so, has encountered an error saying the 29th is not in the calendar
# + hidden=true
sindex = ['2020-01-01', '2020-01-15', '2020-01-31',
'2020-02-01', '2020-02-15', '2020-02-28',
'2020-03-01', '2020-03-15', '2020-03-31',
'2021-01-01', '2021-01-15', '2021-01-31',
'2021-02-01', '2021-02-15', '2021-02-28', # Change the date
'2021-03-01', '2021-03-15', '2021-03-31']
dt_series.index = sindex
dt_series
# + [markdown] hidden=true
# <b>Once the parsing error cause has been fixed lets try to make a `DatetimeIndex` for our series</b>
# + hidden=true
dt_series.index = pd.to_datetime(dt_series.index)
dt_series.index
# + [markdown] hidden=true
# <b>Now can we filter out only `2020`</b>
# + hidden=true
dt_series['2020-02']
# + [markdown] hidden=true
# ****
# <b>[Pandas has four main concepts pertaining to Time](https://pandas.pydata.org/pandas-docs/stable/user_guide/timeseries.html#overview)</b>
#
# > <b>Where methods of Pandas shine - Human Readable Times..</b>
#
# 1.) Timestamp - Similar to datetime, its an instnace in Time eg pd.Timestamp(2020,12,7) == pd.Timestamp("7th December 2020")
#
# 2.) Timedeltas - A absolute duration of time, eg pd.Timedelta('1 day') = 24 hours
#
# 3.) Timespans - A span of time, i.e Starting from 2020-12-07, every other month.
#
# 4.) DateOffsets - To support caneldar arithmetic, similar to `dateutil.relativedelta.relativedelta`
#
#
# <table class="table">
# <colgroup>
# <col style="width: 15%">
# <col style="width: 12%">
# <col style="width: 13%">
# <col style="width: 31%">
# <col style="width: 28%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Concept</p></th>
# <th class="head"><p>Scalar Class</p></th>
# <th class="head"><p>Array Class</p></th>
# <th class="head"><p>pandas Data Type</p></th>
# <th class="head"><p>Primary Creation Method</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p>Date times</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">Timestamp</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">DatetimeIndex</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">datetime64[ns]</span></code> or <code class="docutils literal notranslate"><span class="pre">datetime64[ns,</span> <span class="pre">tz]</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">to_datetime</span></code> or <code class="docutils literal notranslate"><span class="pre">date_range</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Time deltas</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">Timedelta</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">TimedeltaIndex</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">timedelta64[ns]</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">to_timedelta</span></code> or <code class="docutils literal notranslate"><span class="pre">timedelta_range</span></code></p></td>
# </tr>
# <tr class="row-even"><td><p>Time spans</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">Period</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">PeriodIndex</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">period[freq]</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">Period</span></code> or <code class="docutils literal notranslate"><span class="pre">period_range</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Date offsets</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">DateOffset</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">None</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">None</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">DateOffset</span></code></p></td>
# </tr>
# </tbody>
# </table>
# + [markdown] hidden=true
# ****
# <b>Playing around with Timestamps</b>
#
# Its inferred format will always be in **YYYY-MM-DD HH:MM:SS...**
#
# <b>Attributes available to Pandas Timestamp object :</b>
#
# <table class="longtable table autosummary">
# <colgroup>
# <col style="width: 10%">
# <col style="width: 90%">
# </colgroup>
# <tbody>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.asm8.html#pandas.Timestamp.asm8" title="pandas.Timestamp.asm8"><code class="xref py py-obj docutils literal notranslate"><span class="pre">asm8</span></code></a></p></td>
# <td><p>Return numpy datetime64 format in nanoseconds.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.dayofweek.html#pandas.Timestamp.dayofweek" title="pandas.Timestamp.dayofweek"><code class="xref py py-obj docutils literal notranslate"><span class="pre">dayofweek</span></code></a></p></td>
# <td><p>Return day of the week.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.dayofyear.html#pandas.Timestamp.dayofyear" title="pandas.Timestamp.dayofyear"><code class="xref py py-obj docutils literal notranslate"><span class="pre">dayofyear</span></code></a></p></td>
# <td><p>Return the day of the year.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.days_in_month.html#pandas.Timestamp.days_in_month" title="pandas.Timestamp.days_in_month"><code class="xref py py-obj docutils literal notranslate"><span class="pre">days_in_month</span></code></a></p></td>
# <td><p>Return the number of days in the month.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.daysinmonth.html#pandas.Timestamp.daysinmonth" title="pandas.Timestamp.daysinmonth"><code class="xref py py-obj docutils literal notranslate"><span class="pre">daysinmonth</span></code></a></p></td>
# <td><p>Return the number of days in the month.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.freqstr.html#pandas.Timestamp.freqstr" title="pandas.Timestamp.freqstr"><code class="xref py py-obj docutils literal notranslate"><span class="pre">freqstr</span></code></a></p></td>
# <td><p>Return the total number of days in the month.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.is_leap_year.html#pandas.Timestamp.is_leap_year" title="pandas.Timestamp.is_leap_year"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_leap_year</span></code></a></p></td>
# <td><p>Return True if year is a leap year.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.is_month_end.html#pandas.Timestamp.is_month_end" title="pandas.Timestamp.is_month_end"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_month_end</span></code></a></p></td>
# <td><p>Return True if date is last day of month.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.is_month_start.html#pandas.Timestamp.is_month_start" title="pandas.Timestamp.is_month_start"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_month_start</span></code></a></p></td>
# <td><p>Return True if date is first day of month.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.is_quarter_end.html#pandas.Timestamp.is_quarter_end" title="pandas.Timestamp.is_quarter_end"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_quarter_end</span></code></a></p></td>
# <td><p>Return True if date is last day of the quarter.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.is_quarter_start.html#pandas.Timestamp.is_quarter_start" title="pandas.Timestamp.is_quarter_start"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_quarter_start</span></code></a></p></td>
# <td><p>Return True if date is first day of the quarter.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.is_year_end.html#pandas.Timestamp.is_year_end" title="pandas.Timestamp.is_year_end"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_year_end</span></code></a></p></td>
# <td><p>Return True if date is last day of the year.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.is_year_start.html#pandas.Timestamp.is_year_start" title="pandas.Timestamp.is_year_start"><code class="xref py py-obj docutils literal notranslate"><span class="pre">is_year_start</span></code></a></p></td>
# <td><p>Return True if date is first day of the year.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.quarter.html#pandas.Timestamp.quarter" title="pandas.Timestamp.quarter"><code class="xref py py-obj docutils literal notranslate"><span class="pre">quarter</span></code></a></p></td>
# <td><p>Return the quarter of the year.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.tz.html#pandas.Timestamp.tz" title="pandas.Timestamp.tz"><code class="xref py py-obj docutils literal notranslate"><span class="pre">tz</span></code></a></p></td>
# <td><p>Alias for tzinfo.</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="pandas.Timestamp.week.html#pandas.Timestamp.week" title="pandas.Timestamp.week"><code class="xref py py-obj docutils literal notranslate"><span class="pre">week</span></code></a></p></td>
# <td><p>Return the week number of the year.</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="pandas.Timestamp.weekofyear.html#pandas.Timestamp.weekofyear" title="pandas.Timestamp.weekofyear"><code class="xref py py-obj docutils literal notranslate"><span class="pre">weekofyear</span></code></a></p></td>
# <td><p>Return the week number of the year.</p></td>
# </tr>
# </tbody>
# </table>
# + hidden=true
pd.Timestamp(datetime.now())
# + hidden=true
pd.Timestamp("7th December 2020")
# + hidden=true
pd.Timestamp("December 7 2020")
# + hidden=true
pd.Timestamp("7th of December 2020 9:20:23.123123")
# + hidden=true
pd.Timestamp("7th Dec 2020 7 PM")
# + hidden=true
pd.Timestamp("7th Dec 2020 7 PM")==pd.Timestamp("7th Dec 2020 19:00")
# + hidden=true
pd.Timestamp("7/12 2020")==pd.Timestamp("2020-7/12")
# + hidden=true
pd.Timestamp("7/12/2020")==pd.Timestamp("12/7/2020")
# + hidden=true
pd.Timestamp(year=2020, month=12, day=7)
# + hidden=true
pdtstamp = pd.Timestamp("7th Dec 2020 7 PM")
pdtstamp
# + hidden=true
print('**Timestamp Attributes**')
print('------------------------')
print('Day Name : ', pdtstamp.day_name())
print('Quarter : ', pdtstamp.quarter)
print('Date : ', pdtstamp.day)
print('Month : ', pdtstamp.month)
print('Monthdays : ', pdtstamp.daysinmonth)
print('Year : ', pdtstamp.year)
print('Week yr : ', pdtstamp.weekofyear)
# + [markdown] hidden=true
# ****
# <b>Playing around with `date_range`, `bdate_range`, Time Period</b>
#
# What is the difference between TimePeriod and Timestamp?
# > Well in most of the cases, atleast Time Series problems, its necessary to have a representation of the span instead of just a point in time
#
# ****
# <b>Available `freq` or `offset_aliases` :</b>
# <table class="colwidths-given table">
# <colgroup>
# <col style="width: 16%">
# <col style="width: 16%">
# <col style="width: 68%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Date Offset</p></th>
# <th class="head"><p>Frequency String</p></th>
# <th class="head"><p>Description</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.DateOffset.html#pandas.tseries.offsets.DateOffset" title="pandas.tseries.offsets.DateOffset"><code class="xref py py-class docutils literal notranslate"><span class="pre">DateOffset</span></code></a></p></td>
# <td><p>None</p></td>
# <td><p>Generic offset class, defaults to 1 calendar day</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BDay.html#pandas.tseries.offsets.BDay" title="pandas.tseries.offsets.BDay"><code class="xref py py-class docutils literal notranslate"><span class="pre">BDay</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.BusinessDay.html#pandas.tseries.offsets.BusinessDay" title="pandas.tseries.offsets.BusinessDay"><code class="xref py py-class docutils literal notranslate"><span class="pre">BusinessDay</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'B'</span></code></p></td>
# <td><p>business day (weekday)</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.CDay.html#pandas.tseries.offsets.CDay" title="pandas.tseries.offsets.CDay"><code class="xref py py-class docutils literal notranslate"><span class="pre">CDay</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.CustomBusinessDay.html#pandas.tseries.offsets.CustomBusinessDay" title="pandas.tseries.offsets.CustomBusinessDay"><code class="xref py py-class docutils literal notranslate"><span class="pre">CustomBusinessDay</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'C'</span></code></p></td>
# <td><p>custom business day</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Week.html#pandas.tseries.offsets.Week" title="pandas.tseries.offsets.Week"><code class="xref py py-class docutils literal notranslate"><span class="pre">Week</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'W'</span></code></p></td>
# <td><p>one week, optionally anchored on a day of the week</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.WeekOfMonth.html#pandas.tseries.offsets.WeekOfMonth" title="pandas.tseries.offsets.WeekOfMonth"><code class="xref py py-class docutils literal notranslate"><span class="pre">WeekOfMonth</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'WOM'</span></code></p></td>
# <td><p>the x-th day of the y-th week of each month</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.LastWeekOfMonth.html#pandas.tseries.offsets.LastWeekOfMonth" title="pandas.tseries.offsets.LastWeekOfMonth"><code class="xref py py-class docutils literal notranslate"><span class="pre">LastWeekOfMonth</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'LWOM'</span></code></p></td>
# <td><p>the x-th day of the last week of each month</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.MonthEnd.html#pandas.tseries.offsets.MonthEnd" title="pandas.tseries.offsets.MonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">MonthEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'M'</span></code></p></td>
# <td><p>calendar month end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.MonthBegin.html#pandas.tseries.offsets.MonthBegin" title="pandas.tseries.offsets.MonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">MonthBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'MS'</span></code></p></td>
# <td><p>calendar month begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BMonthEnd.html#pandas.tseries.offsets.BMonthEnd" title="pandas.tseries.offsets.BMonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">BMonthEnd</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.BusinessMonthEnd.html#pandas.tseries.offsets.BusinessMonthEnd" title="pandas.tseries.offsets.BusinessMonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">BusinessMonthEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BM'</span></code></p></td>
# <td><p>business month end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BMonthBegin.html#pandas.tseries.offsets.BMonthBegin" title="pandas.tseries.offsets.BMonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">BMonthBegin</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.BusinessMonthBegin.html#pandas.tseries.offsets.BusinessMonthBegin" title="pandas.tseries.offsets.BusinessMonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">BusinessMonthBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BMS'</span></code></p></td>
# <td><p>business month begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.CBMonthEnd.html#pandas.tseries.offsets.CBMonthEnd" title="pandas.tseries.offsets.CBMonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">CBMonthEnd</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.CustomBusinessMonthEnd.html#pandas.tseries.offsets.CustomBusinessMonthEnd" title="pandas.tseries.offsets.CustomBusinessMonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">CustomBusinessMonthEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'CBM'</span></code></p></td>
# <td><p>custom business month end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.CBMonthBegin.html#pandas.tseries.offsets.CBMonthBegin" title="pandas.tseries.offsets.CBMonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">CBMonthBegin</span></code></a> or <a class="reference internal" href="../reference/api/pandas.tseries.offsets.CustomBusinessMonthBegin.html#pandas.tseries.offsets.CustomBusinessMonthBegin" title="pandas.tseries.offsets.CustomBusinessMonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">CustomBusinessMonthBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'CBMS'</span></code></p></td>
# <td><p>custom business month begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.SemiMonthEnd.html#pandas.tseries.offsets.SemiMonthEnd" title="pandas.tseries.offsets.SemiMonthEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">SemiMonthEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'SM'</span></code></p></td>
# <td><p>15th (or other day_of_month) and calendar month end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.SemiMonthBegin.html#pandas.tseries.offsets.SemiMonthBegin" title="pandas.tseries.offsets.SemiMonthBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">SemiMonthBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'SMS'</span></code></p></td>
# <td><p>15th (or other day_of_month) and calendar month begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.QuarterEnd.html#pandas.tseries.offsets.QuarterEnd" title="pandas.tseries.offsets.QuarterEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">QuarterEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'Q'</span></code></p></td>
# <td><p>calendar quarter end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.QuarterBegin.html#pandas.tseries.offsets.QuarterBegin" title="pandas.tseries.offsets.QuarterBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">QuarterBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'QS'</span></code></p></td>
# <td><p>calendar quarter begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BQuarterEnd.html#pandas.tseries.offsets.BQuarterEnd" title="pandas.tseries.offsets.BQuarterEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">BQuarterEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BQ</span></code></p></td>
# <td><p>business quarter end</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BQuarterBegin.html#pandas.tseries.offsets.BQuarterBegin" title="pandas.tseries.offsets.BQuarterBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">BQuarterBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BQS'</span></code></p></td>
# <td><p>business quarter begin</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.FY5253Quarter.html#pandas.tseries.offsets.FY5253Quarter" title="pandas.tseries.offsets.FY5253Quarter"><code class="xref py py-class docutils literal notranslate"><span class="pre">FY5253Quarter</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'REQ'</span></code></p></td>
# <td><p>retail (aka 52-53 week) quarter</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.YearEnd.html#pandas.tseries.offsets.YearEnd" title="pandas.tseries.offsets.YearEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">YearEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'A'</span></code></p></td>
# <td><p>calendar year end</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.YearBegin.html#pandas.tseries.offsets.YearBegin" title="pandas.tseries.offsets.YearBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">YearBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'AS'</span></code> or <code class="docutils literal notranslate"><span class="pre">'BYS'</span></code></p></td>
# <td><p>calendar year begin</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BYearEnd.html#pandas.tseries.offsets.BYearEnd" title="pandas.tseries.offsets.BYearEnd"><code class="xref py py-class docutils literal notranslate"><span class="pre">BYearEnd</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BA'</span></code></p></td>
# <td><p>business year end</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BYearBegin.html#pandas.tseries.offsets.BYearBegin" title="pandas.tseries.offsets.BYearBegin"><code class="xref py py-class docutils literal notranslate"><span class="pre">BYearBegin</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BAS'</span></code></p></td>
# <td><p>business year begin</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.FY5253.html#pandas.tseries.offsets.FY5253" title="pandas.tseries.offsets.FY5253"><code class="xref py py-class docutils literal notranslate"><span class="pre">FY5253</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'RE'</span></code></p></td>
# <td><p>retail (aka 52-53 week) year</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Easter.html#pandas.tseries.offsets.Easter" title="pandas.tseries.offsets.Easter"><code class="xref py py-class docutils literal notranslate"><span class="pre">Easter</span></code></a></p></td>
# <td><p>None</p></td>
# <td><p>Easter holiday</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.BusinessHour.html#pandas.tseries.offsets.BusinessHour" title="pandas.tseries.offsets.BusinessHour"><code class="xref py py-class docutils literal notranslate"><span class="pre">BusinessHour</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'BH'</span></code></p></td>
# <td><p>business hour</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.CustomBusinessHour.html#pandas.tseries.offsets.CustomBusinessHour" title="pandas.tseries.offsets.CustomBusinessHour"><code class="xref py py-class docutils literal notranslate"><span class="pre">CustomBusinessHour</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'CBH'</span></code></p></td>
# <td><p>custom business hour</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Day.html#pandas.tseries.offsets.Day" title="pandas.tseries.offsets.Day"><code class="xref py py-class docutils literal notranslate"><span class="pre">Day</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'D'</span></code></p></td>
# <td><p>one absolute day</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Hour.html#pandas.tseries.offsets.Hour" title="pandas.tseries.offsets.Hour"><code class="xref py py-class docutils literal notranslate"><span class="pre">Hour</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'H'</span></code></p></td>
# <td><p>one hour</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Minute.html#pandas.tseries.offsets.Minute" title="pandas.tseries.offsets.Minute"><code class="xref py py-class docutils literal notranslate"><span class="pre">Minute</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'T'</span></code> or <code class="docutils literal notranslate"><span class="pre">'min'</span></code></p></td>
# <td><p>one minute</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Second.html#pandas.tseries.offsets.Second" title="pandas.tseries.offsets.Second"><code class="xref py py-class docutils literal notranslate"><span class="pre">Second</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'S'</span></code></p></td>
# <td><p>one second</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Milli.html#pandas.tseries.offsets.Milli" title="pandas.tseries.offsets.Milli"><code class="xref py py-class docutils literal notranslate"><span class="pre">Milli</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'L'</span></code> or <code class="docutils literal notranslate"><span class="pre">'ms'</span></code></p></td>
# <td><p>one millisecond</p></td>
# </tr>
# <tr class="row-even"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Micro.html#pandas.tseries.offsets.Micro" title="pandas.tseries.offsets.Micro"><code class="xref py py-class docutils literal notranslate"><span class="pre">Micro</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'U'</span></code> or <code class="docutils literal notranslate"><span class="pre">'us'</span></code></p></td>
# <td><p>one microsecond</p></td>
# </tr>
# <tr class="row-odd"><td><p><a class="reference internal" href="../reference/api/pandas.tseries.offsets.Nano.html#pandas.tseries.offsets.Nano" title="pandas.tseries.offsets.Nano"><code class="xref py py-class docutils literal notranslate"><span class="pre">Nano</span></code></a></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">'N'</span></code></p></td>
# <td><p>one nanosecond</p></td>
# </tr>
# </tbody>
# </table>
# + hidden=true
pd.Timestamp('2020-01-01')
# + hidden=true
pd.Period('2020'), pd.Period('2020-01'), pd.Period('2020-01-01 12'), pd.Period('2020-01-01 12:8')
# + hidden=true
pd.Timestamp('2020')+1
# + hidden=true
pd.Period('2020-02', freq='A-MAR') + 1
# + hidden=true
(pd.Period('2020', freq='A-MAR') + 1).month
# + hidden=true
pd.date_range('2020-01-01', freq='B', periods=10)
# + hidden=true
pd.date_range('2020-01-01', freq='3h43min', periods=10)
# + [markdown] hidden=true
# <b>Custom Buisness days with a list of Holidays</b>
# + hidden=true
pd.bdate_range('2020-01-01', freq='B', periods=10, holidays=['2020-01-10','2020-01-13'])
# + [markdown] hidden=true solution2="shown" solution2_first=true
# **Q)** How to fix it? HINT : Check the table above.
# + hidden=true solution2="shown"
pd.bdate_range('2020-01-01', freq='C', periods=10, holidays=['2020-01-10','2020-01-13'])
# + hidden=true
#----------YOUR SOLUTION-----------#
#....
# + [markdown] hidden=true
# ****
# + [markdown] hidden=true
# <b>Notice the difference between the two :</b>
# + hidden=true
pd.date_range('2020-01-01', freq='D',periods=10)
# + hidden=true
pd.period_range('2020-01-01', freq='D', periods=10)
# + [markdown] hidden=true
# <b>Well thats not much of difference to worry about now is it? So why do we need Period?</b>
# + hidden=true
pd.date_range('2020-01-01', freq='Q-MAR', periods=10)
# + hidden=true
pd.date_range('2020-01-01', freq='QS-MAR', periods=10)
# + hidden=true
pd.period_range('2020-01-01', freq='Q-MAR', periods=10)
# + [markdown] hidden=true
# ****
# <b>Playing around with Time Deltas</b>
# + hidden=true
pd.Period('2020-01-01')+pd.Timedelta(days=3)
# + hidden=true
pd.Timestamp('2020-01-01')+pd.Timedelta(days=3)
# + hidden=true
dtrange = pd.date_range('2020-01-01', freq='D', periods=10)
dtrange
# + hidden=true
dtrange+pd.Timedelta(days=+4, hours=+2)
# + [markdown] heading_collapsed=true
# # Pandas TimeZones
# + hidden=true
all_timezones
# + hidden=true
[k for k in all_timezones if 'New' in k]
# + [markdown] hidden=true
# <b>Inherently pandas `Timestamp` doesnt assume a time-zone, unless explicity given</b>
# + hidden=true
pdtstamp = pd.Timestamp(datetime.now())
pdtstamp
# + hidden=true
# + hidden=true
print(pdtstamp.tz)
# + [markdown] hidden=true
# <b>Or, you call the tz_localize() with the `tz`</b>
# + hidden=true
pdtstamp = pd.Timestamp(datetime.now(), tz='Asia/Calcutta')
pdtstamp.tz
# + [markdown] hidden=true
# **Converting the timestamps between timezones**
# + hidden=true
pdtstamp.tz_convert('Australia/Melbourne')
# + [markdown] hidden=true solution2="shown" solution2_first=true
# **Q)** Give me the current time in Calcutta, Bangkok & New York ->
# + hidden=true solution2="shown"
current_tznaive_time = pd.Timestamp(datetime.now())
current_tzlocalised_time = current_tznaive_time.tz_localize('Asia/Calcutta')
current_tzconverted_time1 = current_tzlocalised_time.tz_convert('Asia/Bangkok')
current_tzconverted_time2 = current_tzlocalised_time.tz_convert('America/New_York')
# + hidden=true
# + hidden=true solution2="shown"
print('Current Näive Time : ', current_tznaive_time)
print('Current Localised Time (Asia/Calcutta) : ', current_tzlocalised_time)
print('Whats the time in Bangkok? -> ', current_tzconverted_time1)
print('Whats the time in New York? -> ', current_tzconverted_time2)
# + hidden=true
#----------YOUR SOLUTION-----------#
#....
# + [markdown] hidden=true
# <b>Playing with your time series data</b>
# + hidden=true
print(britannia_data.index)
britannia_data.head()
# + hidden=true
britannia_data.index = britannia_data.index.tz_localize('Asia/Calcutta')
print(britannia_data.index)
britannia_data.head()
# + hidden=true
britannia_data['Date(SNG)'] = britannia_data.index.tz_convert('Asia/Singapore')
britannia_data.head()
# + hidden=true
britannia_data.drop('Date(SNG)', axis=1, inplace =True)
# -
# # Pandas Resampling
#
# ***NOTE : Do not confuse this with Undersampling and Oversampling***
#
# <img src='https://raw.githubusercontent.com/rafjaa/machine_learning_fecib/master/src/static/img/resampling.png'>
britannia_data.head()
# <b>Lower Granularity</b>
britannia_data.asfreq('1min', method='bfill')['1996-01-02']
# <b>Higher Granularity</b>
britannia_data.resample('3D', label='left').min()
# <b>Can you see the problem here if we do this to our dataset??</b>
#
# - Some dates have been added, although they might had been off days or holidays even in 'B' frequency
#
# - Values assumed by the features are now defined by us, and no longer can be said they are the tru value
# # Random Testing Space
|
Notebooks/02-Pandas Introduction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic regression: Predict fuel efficiency
#
# https://www.tensorflow.org/tutorials/keras/regression
# ## Import Keras
pip install -q seaborn
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
# Make numpy printouts easier to read.
np.set_printoptions(precision=3, suppress=True)
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
print(tf.__version__)
# -
# ## The Auto MPG Dataset
# +
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data'
column_names = ['MPG', 'Cylinders', 'Displacement', 'Horsepower', 'Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(url, names=column_names,
na_values='?', comment='\t',
sep=' ', skipinitialspace=True)
# -
dataset = raw_dataset.copy()
dataset.tail()
# ### Data Cleaning
dataset.isna().sum()
# Drop the NAN row.
dataset = dataset.dropna()
# One-hot coding. The "Origin" column is really categorical, not numeric. So convert that to a one-hot.
dataset['Origin'] = dataset['Origin'].map({1: 'USA', 2: 'Europe', 3: 'Japan'})
dataset = pd.get_dummies(dataset, prefix='', prefix_sep='')
dataset.tail()
# ### Data Split
# +
train_dataset = dataset.sample(frac=0.8, random_state=0)
test_dataset = dataset.drop(train_dataset.index)
train_features = train_dataset.copy()
test_features = test_dataset.copy()
train_labels = train_features.pop('MPG')
test_labels = test_features.pop('MPG')
# -
# ### Data Inspection
# Quick look data joint distribution
sns.pairplot(train_dataset[['MPG', 'Cylinders', 'Displacement', 'Weight']], diag_kind='kde')
train_dataset.describe().transpose()
# ### Normalization
# It is good practice to normalize features that use different scales and ranges.
#
# One reason this is important is because the features are multiplied by the model weights. So the scale of the outputs and the scale of the gradients are affected by the scale of the inputs.
#
# Although a model might converge without feature normalization, normalization makes training much more stable.
#
# Reason is about the optimizer gradient.
train_dataset.describe().transpose()[['mean', 'std']]
normalizer = preprocessing.Normalization()
normalizer.adapt(np.array(train_features))
print(normalizer.mean.numpy())
# +
first = np.array(train_features[:1])
with np.printoptions(precision=2, suppress=True):
print('First example:', first)
print()
print('Normalized:', normalizer(first).numpy())
# -
# ## Linear regression model
# ### One Variable Model
# Start with one variable model. Predict MPG from Horsepower. `y = weight * x + bias`
horsepower = np.array(train_features['Horsepower'])
horsepower_normalizer = preprocessing.Normalization(input_shape=[1,])
horsepower_normalizer.adapt(horsepower)
# +
horsepower_model = tf.keras.Sequential([
horsepower_normalizer,
layers.Dense(units=1)
])
horsepower_model.summary()
# -
# Take a glance of untrained model with first 10 results.
horsepower_model.predict(horsepower[:10])
horsepower_model.compile(
optimizer=tf.optimizers.Adam(learning_rate=0.1),
loss='mean_absolute_error'
)
# +
# %%time
epochs = 100
history = horsepower_model.fit(
train_features['Horsepower'],
train_labels,
epochs=epochs,
verbose=0,
validation_split=0.2
)
# -
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
def plot_loss(history):
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
plt.ylim([0, 10])
plt.xlabel('Epoch')
plt.ylabel('Error [MPG]')
plt.legend()
plt.grid(True)
plot_loss(history)
# Collect the result on the test set, for later:
test_results = {}
test_results['horsepower_model'] = horsepower_model.evaluate(
test_features['Horsepower'],
test_labels,
verbose=0
)
# #### Draw the model input output relationship
x = tf.linspace(0.0, 250, 251)
y = horsepower_model.predict(x)
def plot_horsepower(x, y):
plt.scatter(train_features['Horsepower'], train_labels, label='Data')
plt.plot(x, y, color='k', label='Predictions')
plt.xlabel('Horsepower')
plt.ylabel('MPG')
plt.legend()
plot_horsepower(x, y)
# ### Multiple Inputs Model
linear_model = tf.keras.Sequential([
normalizer, # use fully parameters normalizer input layer
layers.Dense(units=1)
])
linear_model.predict(train_features[:10])
linear_model.layers[1].kernel
linear_model.compile(
optimizer=tf.optimizers.Adam(learning_rate=0.1),
loss='mean_absolute_error'
)
# +
# %%time
epochs = 100
history = linear_model.fit(
train_features,
train_labels,
epochs=epochs,
verbose=0,
validation_split=0.2
)
# -
plot_loss(history)
test_results['linear_model'] = linear_model.evaluate(test_features, test_labels, verbose=0)
# ## DNN Regression
def build_and_compile_model(norm):
model = keras.Sequential([
norm,
layers.Dense(128, activation='relu'),
layers.Dense(128, activation='relu'),
layers.Dense(1)
])
model.compile(
loss='mean_absolute_error',
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001)
)
return model
# ### One Variable
dnn_horsepower_model = build_and_compile_model(horsepower_normalizer)
dnn_horsepower_model.summary()
# +
# %%time
epochs=100
history = dnn_horsepower_model.fit(
train_features['Horsepower'],
train_labels,
verbose=0,
epochs=epochs,
validation_split=0.2
)
# -
plot_loss(history)
x = tf.linspace(0.0, 250, 251)
y = dnn_horsepower_model.predict(x)
plot_horsepower(x, y)
test_results['dnn_horsepower_model'] = dnn_horsepower_model.evaluate(
test_features['Horsepower'],
test_labels,
verbose=0
)
# ### Full Parameters Model
dnn_model = build_and_compile_model(normalizer)
dnn_model.summary()
# +
# %%time
epochs=100
history = dnn_model.fit(
train_features,
train_labels,
verbose=0,
epochs=epochs,
validation_split=0.2
)
# -
plot_loss(history)
test_results['dnn_model'] = dnn_model.evaluate(test_features, test_labels, verbose=0)
# ## Performance Summary
pd.DataFrame(test_results, index=['Mean absolute error [MPG]']).T
# ### Predictions
# +
test_predictions = dnn_model.predict(test_features).flatten()
a = plt.axes(aspect='equal')
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
lims = [0, 50]
plt.xlim(lims)
plt.ylim(lims)
_ = plt.plot(lims, lims)
# -
error = test_predictions - test_labels
plt.hist(error, bins=25)
plt.xlabel('Prediction Error [MPG]')
_ = plt.ylabel('Count')
# ### Model Benchmark
dnn_model.save('dnn_model')
# +
reloaded = tf.keras.models.load_model('dnn_model')
test_results['reloaded'] = reloaded.evaluate(
test_features, test_labels, verbose=0)
# -
pd.DataFrame(test_results, index=['Mean absolute error [MPG]']).T
|
src/tensorflow_lr/ML_basics_with_keras/tensorflow_basic_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--NAVIGATION-->
# < [ipycanvas: Interactive Canvas](04.04-ipycanvas.ipynb) | [Contents](00.00-index.ipynb) |
# # ipyvolume: 3D plotting in the notebook
#
# ## https://github.com/maartenbreddels/ipyvolume
#
# IPyvolume is a Python library to visualize 3d volumes and glyphs (e.g. 3d scatter plots), in the Jupyter notebook, with minimal configuration and effort. It is currently pre-1.0, so use at own risk. IPyvolume’s volshow is to 3d arrays what matplotlib’s imshow is to 2d arrays.
#
# - MIT Licensed
#
# By <NAME>
#
# **Installation:**
#
# ```bash
# conda install -c conda-forge ipyvolume
# ```
# # 3-D Scatter Plots
import ipyvolume as ipv
import numpy as np
import ipywidgets as widgets
x, y, z = np.random.random((3, 10000))
ipv.figure()
scatter = ipv.scatter(x, y, z, size=1, marker="sphere")
ipv.show()
x, y, z, u, v, w = np.random.random((6, 1000))*2-1
selected = np.random.randint(0, 1000, 100)
fig = ipv.figure()
quiver = ipv.quiver(x, y, z, u, v, w, size=5, size_selected=8, selected=selected)
ipv.show()
size = widgets.FloatSlider(min=0, max=30, step=0.1)
size_selected = widgets.FloatSlider(min=0, max=30, step=0.1)
color = widgets.ColorPicker()
color_selected = widgets.ColorPicker()
widgets.jslink((quiver, 'size'), (size, 'value'))
widgets.jslink((quiver, 'size_selected'), (size_selected, 'value'))
widgets.jslink((quiver, 'color'), (color, 'value'))
widgets.jslink((quiver, 'color_selected'), (color_selected, 'value'))
widgets.VBox([size, size_selected, color, color_selected])
# # Animations
# create 2d grids: x, y, and r
u = np.linspace(-10, 10, 25)
x, y = np.meshgrid(u, u)
r = np.sqrt(x**2+y**2)
print("x,y and z are of shape", x.shape)
# and turn them into 1d
x = x.flatten()
y = y.flatten()
r = r.flatten()
print("and flattened of shape", x.shape)
# create a sequence of 15 time elements
time = np.linspace(0, np.pi*2, 15)
z = np.array([(np.cos(r + t) * np.exp(-r/5)) for t in time])
print("z is of shape", z.shape)
# draw the scatter plot, and add controls with animate_glyphs
ipv.figure()
s = ipv.scatter(x, z, y, marker="sphere")
ipv.animation_control(s, interval=200)
ipv.ylim(-3,3)
ipv.show()
# Now also include, color, which containts rgb values
color = np.array([[np.cos(r + t), 1-np.abs(z[i]), 0.1+z[i]*0] for i, t in enumerate(time)])
size = (z+1)
print("color is of shape", color.shape)
color = np.transpose(color, (0, 2, 1)) # flip the last axes
ipv.figure()
s = ipv.scatter(x, z, y, color=color, size=size, marker="sphere")
ipv.animation_control(s, interval=200)
ipv.ylim(-3, 3)
ipv.show()
# <!--NAVIGATION-->
# < [ipycanvas: Interactive Canvas](04.04-ipycanvas.ipynb) | [Contents](00.00-index.ipynb) |
|
notebooks/04.Widget-libraries/04.05-ipyvolume.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pcsilcan/dm/blob/master/20202/dm_20202_0901_apriori.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="RpRqVz6tom-6" outputId="6650d82b-7155-48ee-a4d4-8aef667ddcb6" colab={"base_uri": "https://localhost:8080/", "height": 54}
T = [[c for c in 'MONKEY'],
[c for c in 'DONKEY'],
[c for c in 'MAKE'],
[c for c in 'MUCKY'],
[c for c in 'COOKIE']]
print(T)
# + id="6F9f5jbUpCzO" outputId="25762a8c-a8e8-4c1f-a5b5-a2e69e71b1e8" colab={"base_uri": "https://localhost:8080/", "height": 34}
I = set()
for Ti in T:
for i in Ti:
I.add(i)
print(I)
# + id="SOCgVFbiqU2n" outputId="0789bc0c-eae0-4ad7-b03a-f6988728d254" colab={"base_uri": "https://localhost:8080/", "height": 34}
def calcSup(itemset):
cont = 0
for Ti in T:
cont += 1 if set(itemset).issubset(Ti) else 0
return cont / len(T)
calcSup(['M'])
# + id="J4J5Wqj4pLAY" outputId="ce9abb65-b6a4-4c33-85c0-b41ea79eefda" colab={"base_uri": "https://localhost:8080/", "height": 67}
import itertools as it
minsup = 0.6
Ck = [[i] for i in I]
L = []
k = 1
while len(Ck) > 0:
L.append([])
for itemset in Ck:
if calcSup(itemset) >= minsup:
L[k-1].append(itemset)
temp = set()
for itemset in L[k-1]:
for i in itemset:
temp.add(i)
k += 1
Ck = [list(x) for x in it.combinations(list(temp), k)]
for Li in L:
print(Li)
# + id="jlQc16ZOtpTR"
def calcConf(X, Y):
return calcSup(X + Y) / calcSup(X)
# + id="TK-aNimlooq0" outputId="72348552-1460-4355-c1d8-dc192af899c2" colab={"base_uri": "https://localhost:8080/", "height": 151}
minconf = 0.8
for k in range(1, len(L)):
for itemset in L[k]:
for j in range(1, k + 1):
ante = [set(x) for x in it.combinations(itemset, j)]
setit = set(itemset)
for setX in ante:
X = list(setX)
Y = list(setit - setX)
if calcConf(X, Y) > minconf:
print(X, "->", Y)
# + id="9klOJxqAsDTl"
|
20202/dm_20202_0901_apriori.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing Open Source Baseball Data
#
# **This notebook performs the same data analysis as 01_Intro.ipynb, except that it uses Postgres as the data source instead of CSV files.**
#
# This notebook examines:
# * how game length in minutes has been increasing over the years
# * how the number of pitchers used per game has been increasing over the years
# * the relationship between game length and pitcher count
#
# This notebook also examines:
# * how much the use of the DH increases run scoring
# ## Setup
# [Preliminaries](#Preliminaries)
# [Imports and Setup](#Imports-and-Setup)
# [Load Data](#Load-the-Data)
#
# ## Baseball Analysis/Questions
# [Are Baseball Games Getting Longer?](#Are-Baseball-Games-Getting-Longer-?)
# [Are Games Longer due to Using More Pitchers?](#Are-Games-Longer-Due-to-Using-More-Pitchers?)
# [Are Games that Use a DH Higher Scoring?](#Are-Games-that-Use-a-DH-Higher-Scoring?)
# ## Preliminaries
#
# This notebook assumes that the Lahman and Retrosheet data sets have been downloaded and wrangled using the scripts in the `../download_scripts` directory of this repo.
#
# Furthermore the postgres_load_data.py script has been run to load the wrangled data into Postgres.
#
# For these notebooks, Retrosheet data from 1955 through 2019 inclusive is used.
# ## MLB Data Dictionary Summary
#
# The most complete data dictionary is from Lahman and has been copied to:
# https://github.com/sdiehl28/baseball-analytics/blob/master/data/lahman/readme2017.txt
#
# Although the data says 2017, is applies to 2018 and 2019 Lahman data as well.
#
# The Retrosheet tables are similar to the Lahman tables, so reading the above is helpful for understanding both data sets.
# ### Most Used Tables
#
# A player may play for team A, be traded to team B, and then be traded back to team A, all in the same year. Lahman would set stint equal to 1, 2, 3, respectively in this scenario.
#
# A player may field at any of the nine fielding positions (pos): P, C, 1B, 2B, 3B, SS, LF, CF, RF
#
# **Lahman Tables**
# * lahman_batting
# * pkey: player_id, year_id, stint
# * lahman_pitching
# * pkey: player_id, year_id, stint
# * lahman_fielding
# * pkey: player_id, year_id, stint, pos
# * lahman_people
# * pkey: player_id
# * fkey: retro_id -- the Retrosheet player_id
# * lahman_teams
# * pkey: team_id, year_id
# * fkey: team_id_retro, year_id -- the Retrosheet team_id, year_id
#
# **Retrosheet Tables**
# * retro_batting
# * pkey: player_id, game_id
# * retro_pitching
# * pkey: player_id, game_id
# * retro_fielding
# * pkey: player_id, game_id, pos
# * retro_team_game
# * pkey: team_id, game_id
# * retro game
# * pkey: game_id
# ## Imports and Setup
import os
import pandas as pd
import numpy as np
from pathlib import Path
import re
from scipy.stats import linregress
from sqlalchemy.engine import create_engine
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import matplotlib as mpl
mpl.rcParams['figure.dpi'] = 100 # increase dpi, will make figures larger and clearer
pd.set_option("display.max_columns", 50)
# +
import sys
# import data_helper.py from download_scripts directory
sys.path.append('../download_scripts')
import data_helper as dh
# -
# ## Connect to Postgres
# Modify as needed for your DBMS.
# +
# Get the user and password from the environment (rather than hardcoding it in the notebook)
import os
db_user = os.environ.get('DB_USER')
db_pass = <PASSWORD>('DB_PASS')
# avoid putting passwords directly in code
connect_str = f'postgresql://{db_user}:{db_pass}@localhost:5432/baseball'
# -
# ### Database Connections
#
# engine = create_engine(connect_str)
#
# Using engine.execute(query) will:
# 1. cause a DB connection object to be allocated for use
# 2. will use that connection object to execute the query
# 3. will commit any data changes
# 4. will release that connection object back to the open connection pool
#
# For transaction processing, using the Python DB API, with SQL Alchemy, use:
# ```connection = create_engine(connect_str).connect()```
# https://stackoverflow.com/questions/34322471/sqlalchemy-engine-connection-and-session-difference#answer-42772654
engine = create_engine(connect_str)
type(engine)
# # Are Baseball Games Getting Longer ?
sql = """
SELECT DATE_PART('year', game_start) AS year, AVG(minutes_game_ct) AS mean_game_len
FROM retro_game
GROUP BY year
ORDER BY year
"""
mean_game_len = pd.read_sql(sql, engine)
mean_game_len.head()
# For nicely annotated plots, matplotlib is often better.
# For quick data exploration, seaborn is often better.
# +
# plot the data and the 3 year moving average
plt.style.use('seaborn-darkgrid')
fig, ax = plt.subplots()
ax.set_title('Game Length vs Year')
ax.set_xlabel('Year')
ax.set_ylabel('Game Length in Minutes')
ax.set_title('Game Length vs Year')
ax.plot('year', 'mean_game_len', data = mean_game_len, label='Mean Length per Year')
df_smoothed = mean_game_len.set_index('year').rolling(3).mean().reset_index()
ax.plot('year', 'mean_game_len', data=df_smoothed, label='3 Yr Moving Avg')
ax.legend();
# -
# ## Summary
# Baseball games are taking longer to complete. From the mid 70s through 2019, the game length has increased by about 40 minutes.
#
# The drop in game time from 2000 to about 2005 might be due to a MLB rule change. For the 2020 season, MLB has changed the rules to speed up games. It will be interesting to see if the average game length for 2020 is lower than the 3 year moving average.
# # Are Games Longer Due to Using More Pitchers?
sql = """
SELECT rtg.year, AVG(pitcher_ct) AS mean_pitcher_ct
FROM retro_game rg JOIN retro_team_game rtg
ON rg.game_id = rtg.game_id
GROUP BY year
ORDER BY year
"""
mean_pitcher_ct = pd.read_sql(sql, engine)
mean_pitcher_ct.head()
# +
fig, ax = plt.subplots()
ax.set_title('Pitcher Count per Game vs Year')
ax.set_xlabel('Year')
ax.set_ylabel('Pitcher Count')
ax.plot('year', 'mean_pitcher_ct', data=mean_pitcher_ct, label='Mean Pitchers per Game per Year')
df_ma2 = mean_pitcher_ct.set_index('year').rolling(3).mean().reset_index()
ax.plot('year', 'mean_pitcher_ct', data=df_ma2, label='3 Yr Moving Avg')
ax.legend();
# -
# The number of pitchers per game has been steadily increasing since the late 70s.
# +
# show game length and pitcher count on same plot
# compute 3 year moving averages
df_ma = mean_game_len.set_index('year').rolling(3).mean().reset_index()
df_ma2 = mean_pitcher_ct.set_index('year').rolling(3).mean().reset_index()
x = df_ma['year']
y1 = df_ma['mean_game_len']
y2 = df_ma2['mean_pitcher_ct']
plt.style.use('default')
fig, ax1 = plt.subplots()
ax1.set_title('Game Length and Pitchers vs Year')
ax1.set_xlabel('Year')
color = 'tab:red'
ax1.set_ylabel('Minutes Per Game', color=color)
lns1 = ax1.plot(x, y1, label='3 Yr MA of Game Length', color=color)
ax1.tick_params(axis='y', labelcolor=color)
# instantiate a second axes that shares the same x-axis
ax2 = ax1.twinx()
color = 'tab:blue'
ax2.set_ylabel('Pitchers Per Game', color=color)
lns2 = ax2.plot(x, y2, label='3 Yr MA of Picther Count', color=color)
ax2.tick_params(axis='y', labelcolor=color)
# create the legend
lns = lns1 + lns2
labs = [l.get_label() for l in lns]
ax1.legend(lns, labs, loc=0)
fig.tight_layout()
# -
# Both have been increasing since the late 70s.
#
# How correlated is the unsmoothed ungrouped data?
sql = """
SELECT CORR(pitcher_ct, minutes_game_ct) AS pearson_r, COUNT(*) as count
FROM retro_team_game rtg JOIN retro_game rg
ON rtg.game_id = rg.game_id
"""
pd.read_sql(sql, engine).round(3)
# compute mean_game_len per pitcher count
sql = """
SELECT pitcher_ct, AVG(minutes_game_ct) AS mean_game_len
FROM retro_team_game rtg JOIN retro_game rg
ON rtg.game_id = rg.game_id
GROUP BY pitcher_ct
"""
mean_pitcher_ct = pd.read_sql(sql, engine)
mean_pitcher_ct
# the relationship looks quite linear
sns.lmplot('pitcher_ct', 'mean_game_len', data=mean_pitcher_ct);
sql = """
SELECT pitcher_ct, minutes_game_ct
FROM retro_team_game rtg JOIN retro_game rg
ON rtg.game_id = rg.game_id
"""
df = pd.read_sql(sql, engine)
# get some additional stats from scipy's linear regression
# note this uses *all* the games, not just the 13 points above
linregress(df['pitcher_ct'], df['minutes_game_ct'])
# From the Linear Regression:
# * r = .618
# * r^2 = .382 => 38% of the variance is explained by the number of pitchers using a linear model
# * p-value = 0.0 => statistically significant
# * slope = 12.4 => each additional pitcher adds 12.4 minutes to game length
# The above is for the period 1955 through 2019. Perhaps the relationship is different using just the last 3 years of data?
sql = """
SELECT pitcher_ct, minutes_game_ct
FROM retro_team_game rtg JOIN retro_game rg
ON rtg.game_id = rg.game_id
WHERE year >= 2017
"""
df_3years = pd.read_sql(sql, engine)
linregress(df_3years['pitcher_ct'], df_3years['minutes_game_ct'])
# The values are nearly the same.
# ## Summary
# Game length and pitchers per game have both increased significantly since the late 70s.
#
# There is a statistically significant linear relationship between the number of pitchers used in a game, and the total time of the game. Each additional pitcher is associated with a increase in game time of about 12 minutes.
#
# The increase in the number of pitchers explains about 38% of the variance, so there are other factors involved in game length.
#
# The new MLB rules for 2020 to speed up the game are clues as to what MLB thinks is causing the game to take so long. The new rules are:
# * a pitcher must pitch to at least 3 batters or until the end of an inning (unless injured)
# * the time between innings is reduced by 5 seconds for most games (and 25 seconds for nationally televised games)
# * the number of meetings on the mound is reduced from 6 to 5 per game
# * instant reply will be fed more quickly to the dug out so the decision to challenge a call can be made more quickly
# # Are Games that Use a DH Higher Scoring?
sql = """
SELECT CAST( MIN ( DATE_PART('year', game_start)) AS integer)
FROM retro_game
WHERE dh = TRUE
"""
dh_min = pd.read_sql(sql, engine)
dh_first_year = dh_min.iloc[0,0]
dh_first_year
# sort by the retro_team_game's primary key, so that the results are repeatable
sql = f"""
SELECT rg.game_id, rtg.team_id, dh, r
FROM retro_team_game rtg JOIN retro_game rg
ON rtg.game_id = rg.game_id
WHERE year >= {dh_first_year}
ORDER BY rg.game_id, rtg.team_id
"""
game_dh = pd.read_sql(sql, engine)
game_dh.head()
dh_df = game_dh.groupby('dh')['r'].agg(['mean', 'count'])
dh_df
delta = dh_df.loc[True, 'mean'] - dh_df.loc[False, 'mean']
delta
# Given the large number of games, this looks significant. Run the <a href="https://en.wikipedia.org/wiki/Resampling_(statistics)#Monte_Carlo_testing">Monte Carlo</a> version of the <a href="https://en.wikipedia.org/wiki/Resampling_(statistics)#Permutation_tests">Permutation Test</a> to see how likely this large of a difference is.
dh_r = game_dh.query('dh == True')['r']
no_dh_r = game_dh.query('dh == False')['r']
dh_r.agg('mean')
no_dh_r.agg('mean')
def perm_test(x, y):
pooled = np.hstack([x, y])
np.random.shuffle(pooled)
x_sample = pooled[:len(x)]
y_sample = pooled[len(x):]
return x_sample.mean() - y_sample.mean()
# set the random seed, so that the results are repeatable
np.random.seed(100)
N = 1000
result_array = np.empty(N)
for i in range(N):
result_array[i] = perm_test(dh_r, no_dh_r)
result_array.min(), result_array.max()
(result_array >= delta).sum()
# There are no random permutations which show anywhere near as large of a run difference as was observed by partitioning games into "DH" and "no DH".
# seaborn catplot: group by values of 'x' and display the mean of 'y'
plt.style.use('seaborn-darkgrid')
sns.catplot(x="dh", y="r", data=game_dh, kind="point", height=3, aspect=1.77)
plt.title('Runs by DH');
# ## Summary
# On average, about 1/4 of a run per team per game more is scored when the Designated Hitter is used. This difference is much greater than could be observed by chance alone.
#
# The DH is better at producing runs than the pitcher. (It would be interesting to compare the wOBA Sabermetric for pitchers vs the DH.)
#
# That said, the DH may not be responsible the entire 1/4 run difference. There may be additional factors because the DH is only used in the American League and perhaps there is something else different about hitting in the American League. Perhaps it's easier to score runs in American league parks. Perhaps the National League has much better pitchers. These are not likely to be as significant to run scoring as the use of the DH, but further analysis would be needed to say if the entire 1/4 run per team per game could be attributed to the use of the DH.
#
# As the DH is only used in games that are played in American League parks, it would be more accurate to summarize the above as:
# * games played in American League parks have about 1/4 run per team per game more than games played in National League parks
|
baseball_jupyter_nb/01_Intro_SQL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://github.com/jtsulliv/ML-from-scratch/blob/master/Neural-Networks/perceptron.ipynb
# https://datascience.stackexchange.com/questions/21929/perceptron-weight-vector-update
# https://www.freecodecamp.org/news/building-a-3-layer-neural-network-from-scratch-99239c4af5d3/
# https://towardsdatascience.com/6-steps-to-write-any-machine-learning-algorithm-from-scratch-perceptron-case-study-335f638a70f3
# -
import numpy as np
import matplotlib.pyplot as plt
from past.builtins import xrange
# +
# NAND gate features
# note: x0 is a dummy variable for the bias term
# x0 x1 x2
x = [[1., 0., 0.],
[1., 0., 1.],
[1., 1., 0.],
[1., 1., 1.]]
# Desired outputs
y = [1.,
1.,
1.,
0.]
# +
#
# x: feature data
# y: outputs
# z: threshold
# eta: learning rate
# t: number of iterations
def perceptron_train(x, y, z, eta, t):
# Initializing parameters for the Perceptron
w = np.zeros(len(x[0])) # weights
n = 0
# Initializing additional parameters to compute SSE
yhat_vec = np.ones(len(y)) # vector for predictions
errors = np.ones(len(y)) # vector for errors (actual - predictions)
J = [] # vector for the SSE cost function
while n < t:
for i in xrange(0, len(x)):
# summation step
f = np.dot(x[i], w)
# activation function
if f > z:
yhat = 1.
else:
yhat = 0.
yhat_vec[i] = yhat
# updating the weights
for j in xrange(0, len(w)):
w[j] = w[j] + eta*(y[i]-yhat)*x[i][j]
n += 1
# computing the sum-of-squared errors
for i in xrange(0,len(y)):
errors[i] = (y[i]-yhat_vec[i])**2
# print("E",y[i]-yhat_vec[i])
# print("SSE",((y[i]-yhat_vec[i])**2))
J.append(0.5*np.sum(errors))
# function returns the weight vector, and sum-of-squared errors + y_hatvec (predictions)
return w, J,yhat_vec
# +
ytrue-ypred = error
# 1 - 0 = 1
# 1 ---> 1
0 - 1 = -1
# -1 --> 1
squared error
0-1=-1
1-0=1
0 -1= -1
0-1 =1
1-0= 1
# -
z = 0.0 # threshold
eta = 0.1 # learning rate
t = 50 # number of iterations
# +
print ("The weights are:")
print (perceptron_train(x, y, z, eta, t)[0], "\n")
print ("The sum-of-squared erros are:")
print (perceptron_train(x, y, z, eta, t)[1])
# +
J = perceptron_train(x, y, z, eta, t)[1] # pulling out the sum-of-squared errors from the tuple
epoch = np.linspace(1,len(J),len(J))
#https://stackoverflow.com/questions/4752626/epoch-vs-iteration-when-training-neural-networks
# %matplotlib inline
plt.plot(epoch, J)
plt.xlabel('Epoch')
plt.ylabel('Sum-of-Squared Error')
plt.title('Perceptron Convergence')
# +
# eta = 0.5 # new learning rate
# z = 0.0
# print ("The weights are:")
# print (perceptron_train(x, y, z, eta, t)[0], "\n")
# J = perceptron_train(x, y, z, eta, t)[1]
# epoch = np.linspace(1,len(J),len(J))
# print ("The sum-of-squared erros are:")
# print (J)
# # %matplotlib inline
# plt.plot(epoch, J)
# plt.xlabel('Epoch')
# plt.ylabel('Sum-of-Squared Error')
# plt.title('Perceptron Convergence')
# +
eta = 0.1
z = 0.5 # new threshold
print ("The weights are:")
print (perceptron_train(x, y, z, eta, t)[0], "\n")
J = perceptron_train(x, y, z, eta, t)[1]
epoch = np.linspace(1,len(J),len(J))
print ("The sum-of-squared erros are:")
print (J)
# %matplotlib inline
plt.plot(epoch, J)
plt.xlabel('Epoch')
plt.ylabel('Sum-of-Squared Error')
plt.title('Perceptron Convergence')
# +
# eta = 0.1
# z = 0.5 # new threshold
# print ("The weights are:")
# print (perceptron_train(x, y, z, eta, t)[0], "\n")
# J = perceptron_train(x, y, z, eta, t)[1]
# epoch = np.linspace(1,len(J),len(J))
# print ("The sum-of-squared erros are:")
# print (J)
# # %matplotlib inline
# plt.plot(epoch, J)
# plt.xlabel('Epoch')
# plt.ylabel('Sum-of-Squared Error')
# plt.title('Perceptron Convergence')
# +
# prediction/output verification
# -
y_pred = perceptron_train(x, y, z, .233, 26)[2]
y_pred
y
y == y_pred
# +
t = 26
eta = .233
y_pred = perceptron_train(x, y, z, eta, t)[2]
print(y == y_pred )
print ("The weights are:")
print (perceptron_train(x, y, z, eta, t)[0], "\n")
J = perceptron_train(x, y, z, eta, t)[1]
epoch = np.linspace(1,len(J),len(J))
print ("The sum-of-squared erros are:")
print (J)
# %matplotlib inline
plt.plot(epoch, J)
plt.xlabel('Epoch')
plt.ylabel('Sum-of-Squared Error')
plt.title('Perceptron Convergence')
# +
# http://homepages.gold.ac.uk/nikolaev/311perc.htm
# perceptron rule
# gradient descent rule
# delta rule
# verify layer/weight updates per iteration/epoch at every step then compare output to non-python implt. in script
# formula to check if dataset/input/features are non-convergence state/linearly seperatable
# i.e perceptron convergence theorem
# +
# from sklearn.metrics import accuracy_score
# w = perceptron_train(x, y, z, eta, t)[0]
# def perceptron_test(x, w, z, eta, t):
# y_pred = []
# for i in xrange(0, len(x-1)):
# f = np.dot(x[i], w)
# # activation function
# if f > z:
# yhat = 1
# else:
# yhat = 0
# y_pred.append(yhat)
# return y_pred
# y_pred = perceptron_test(x, w, z, eta, t)
# print(accuracy_score(y_test, y_pred))
# -
|
nn_p2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Intermediate Lesson on Geospatial Data
#
# ## Spatial Queries
#
# <strong>Lesson Developers:</strong> <NAME>, <NAME>, <NAME>
#
# #### Part 5 of 5
# + hide_input=true init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
# sys.path.append('supplementary')
import hourofci
try:
import os
os.chdir('supplementary')
except:
pass
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# Now when we look at the results we might not see any spatial patterns as such. But this is where spatial data and spatial database shines.
#
# To show the spatial distribution we use a particular type of map called **Choropleth map**.
#
# **Below is a choropleth map showing spatial distribution of earthquakes**
# -
from ipyleaflet import Map,DrawControl,GeoJSON,LayerGroup
import spatialite
import pandas as pd
import geopandas as gpd
import json
import time
from branca.colormap import linear
import matplotlib.pyplot as plt
from ipyleaflet import Choropleth
disp = widgets.Output()
db = spatialite.connect('databases/spatialDB.sqlite')
stateDataSql = f"""SELECT u.stusps,count(*) as total_earthquakes from us_states u,earthquakes s
where st_contains(u.geom,s.geometry) and s.rowid in(SELECT ROWID
FROM SpatialIndex
WHERE f_table_name = 'earthquakes'
AND search_frame = u.geom) group by u.stusps"""
dat = pd.read_sql_query(stateDataSql,db)
stateSql = """SELECT stusps,st_asbinary(geom) as geom FROM us_states"""
#df=pd.read_sql_query(sql,db)
df = gpd.read_postgis(stateSql,db)
dat = df[['stusps']].merge(dat,on='stusps',how='left').fillna(0)
dat = dict(zip(dat['stusps'].tolist(), dat['total_earthquakes'].tolist()))
jsondata = json.loads(df.to_json())
for feature in jsondata['features']:
feature['id'] = feature['properties']['stusps']
layer = Choropleth(
geo_data=jsondata,
choro_data=dat,
colormap=linear.OrRd_03,
border_color='black',
style={'fillOpacity': 0.8, 'dashArray': '5, 5'})
sMap= Map(center=(41.482222, -81.669722), zoom=3,prefer_canvas =True)
sMap.add_layer(layer)
sMap
# + [markdown] slideshow={"slide_type": "slide"}
# ### Intersection Query
# -
# #### Intersects
# First we look at how to use intersect function to check whether two geometries intersect.
#
# The function **st_intersects(geometry A,geometry B)** returns true if geometry A and geometry B intersect or touch at atleast a single point
#
# <img src = "supplementary/images/intersect.png" width = "500px">
# + [markdown] slideshow={"slide_type": "slide"}
# A real world example would be to identify the houses that fall with in a hazard zone. We would not only want the houses that are with in the hazard zone but also the houses that has some portion of it inside the hazard zone
#
# <img src = "supplementary/images/hazard_zones.png" width = "400px">
#
# Let's look at an interactive example. You can use either the line or the rectangle tool to draw your own geometry. The geometries (in this cases us_states) that intersects with the geometry you have drawn will be highlighted in green color.
# + slideshow={"slide_type": "slide"}
from ipyleaflet import Map,DrawControl,GeoJSON,LayerGroup
import geojson
import json
from shapely.geometry import shape
import geopandas as gpd
import numpy as np
import time
import spatialite
matching=[]
def handle_draw(target, action, geo_json):
global matching
draw_group.clear_layers()
geo_dat = GeoJSON(data = geo_json)
draw_group.add_layer(geo_dat)
dc.clear()
g1 = geojson.loads(json.dumps(geo_json['geometry']))
g2 = shape(g1)
sql = f"SELECT stusps from us_states u where st_intersects(u.geom,ST_TRANSFORM(ST_GeomFromText('{g2.wkt}',4326),4269))"
pdf = pd.read_sql_query(sql, db)
matching = pdf.values
for layer in stateGroup.layers:
layer.style={'weight':2+np.random.random_sample()}
def styleChange(feature):
props=feature['properties']
if props['stusps'] in matching:
return {'opacity': 1, 'fillOpacity': 0, 'weight': 1,'color':'green'}
else:
return {'opacity': 0.3, 'fillOpacity': 0, 'weight': 1,'color':'red'}
sMap= Map(center=(44.967243, -103.771556), zoom=8,prefer_canvas =True)
sMap.fit_bounds(((24.1, -126.1), (49.9, -64.4)))
dc = DrawControl(
marker={},
rectangle={"shapeOptions": {"color": "#0000FF",'fillOpacity':0}},
circle={},
circlemarker={},
polygon={}
)
dc.on_draw(handle_draw)
sMap.add_control(dc)
draw_group = LayerGroup()
sMap.add_layer(draw_group)
stateGroup = LayerGroup()
sMap.add_layer(stateGroup)
db = spatialite.connect('databases/spatialDB.sqlite')
stateGeomSql = f"SELECT stusps,ST_AsBinary(geom) as geom FROM us_states;"
gdf = gpd.GeoDataFrame.from_postgis(stateGeomSql, db,crs = 'EPSG:4269').to_crs('EPSG:4326')
sMap.zoom = 6
geo_data = GeoJSON(data = json.loads(gdf.to_json()),style={'opacity': 0.3, 'fillOpacity': 0, 'weight': 1,'color':'red'},style_callback=styleChange)
stateGroup.add_layer(geo_data)
sMap
# + [markdown] slideshow={"slide_type": "slide"}
# #### Intersection
# While intersects check whether two geometries intersect, intersection returns the geometry shared by the two geometries.
#
# The function **st_intersection(geometry A,geometry B)** returns the geometry shared by geometry A and geometry B
#
# <img src = "supplementary/images/intersection.png" width = "500px">
# + [markdown] slideshow={"slide_type": "slide"}
# Now we will look at a concrete example of using st_intersection
#
# **Find total length of subway line in each neighborhood**
#
# <img src = "supplementary/images/intersection_example.png" width = "500px">
# + [markdown] slideshow={"slide_type": "slide"}
# Here we will use one more function **st_length(geometry)** for calculating the length of a geometry
#
# The function **st_length(geometry A)** returns the length of geometry A
#
# Following are the tables involved in this query:
# -
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select boroname,name,geom as geometry from nyc_neighborhoods limit 5',db)
table2 = pd.read_sql_query('select pk_uid,geometry from nyc_subway_lines limit 5',db)
table1_disp = widgets.Output()
table2_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_NEIGHBORHOODS</center></b>")
table2_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_SUBWAY_LINES</center></b>")
with table1_disp:
display(table1)
with table2_disp:
display(table2)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0')),VBox([table2_header,table2_disp])])
out
# + [markdown] slideshow={"slide_type": "slide"}
# The required query
#
# ```sql
# SELECT u.boroname,sum(ST_Length(st_intersection(u.geom,s.geometry))) as total_length from nyc_neighborhoods u,nyc_subway_lines s
# where st_intersects(u.geom,s.geometry) group by u.boroname
# ```
# -
disp = widgets.Output()
db = spatialite.connect('databases/spatialDB.sqlite')
stateGeomSql = f"""SELECT u.boroname,sum(ST_Length(st_intersection(u.geom,s.geometry))) as total_length from nyc_neighborhoods u,nyc_subway_lines s
where st_intersects(u.geom,s.geometry) and s.rowid in(SELECT ROWID
FROM SpatialIndex
WHERE f_table_name = 'nyc_subway_lines'
AND search_frame = u.geom) group by u.boroname"""
data = pd.read_sql_query(stateGeomSql,con=db)
with disp:
display(data)
disp
# + [markdown] slideshow={"slide_type": "slide"}
# ### Within a Distance Queries
#
# With in distance queries are used to find out geometrical objects that are with in a specific distance of a particular geometrical object.
#
#
# <img src = "supplementary/images/withindistance.png" width = "800px">
#
# + [markdown] slideshow={"slide_type": "slide"}
#
# <img src = "supplementary/images/distance_within_example.png" width = "600px">
# + [markdown] slideshow={"slide_type": "slide"}
# #### Buffer
#
# <img src = "supplementary/images/buffer.png" width = "600px">
#
# The function **st_buffer(geometry A,distance)** encircles geometry A at a specified **distance** and returns a geometry object that is the buffer that surrounds the source object (A).
#
# Lets look at a concrete example
#
# **Number of homicides with in 100 meter radius of NYC substations**
#
# The required tables are the following:
# + slideshow={"slide_type": "subslide"}
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select pk_uid,name,geom as geometry from nyc_substations limit 5',db)
table2 = pd.read_sql_query('select pk_uid,weapon,year,geom as geometry from nyc_homicides limit 5',db)
table1_disp = widgets.Output()
table2_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_SUBSTATIONS</center></b>")
table2_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_HOMICIDES</center></b>")
with table1_disp:
display(table1)
with table2_disp:
display(table2)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0')),VBox([table2_header,table2_disp])])
out
# + [markdown] slideshow={"slide_type": "slide"}
# And the query is
#
# ```sql
# SELECT u.name,count(*) as total_homicides from nyc_substations u,
# nyc_homicides s where st_contains(st_buffer(u.geom,100),s.geom) group by u.name
# ```
# -
disp = widgets.Output()
db = spatialite.connect('databases/spatialDB.sqlite')
stateGeomSql = f"""SELECT u.name,count(*) as total_homicides from (select name,st_buffer(geom,100) as geom from nyc_substations) u,
nyc_homicides s where st_contains(u.geom,s.geom) and s.rowid in(SELECT ROWID
FROM SpatialIndex
WHERE f_table_name = 'nyc_homicides'
and f_geometry_column = 'geom'
AND search_frame = u.geom) group by u.name order by count(*) desc"""
data = pd.read_sql_query(stateGeomSql,con=db)
with disp:
display(data)
disp
# + [markdown] slideshow={"slide_type": "slide"}
# Let's look at another example
#
# **Number of shooting incidents with in 50 meter of schools**
#
# and the tables are
# -
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
import spatialite
import pandas as pd
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select pk_uid,schoolname,sch_type,geometry from nyc_schools limit 5',db)
table2 = pd.read_sql_query('select pk_uid,geometry from nyc_shooting limit 5',db)
table1_disp = widgets.Output()
table2_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_SCHOOLS</center></b>")
table2_header = widgets.HTML(value = f"<b><font color='red'><center>NYC_SHOOTING</center></b>")
with table1_disp:
display(table1)
with table2_disp:
display(table2)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0')),VBox([table2_header,table2_disp])])
out
# + [markdown] slideshow={"slide_type": "slide"}
# And the query
#
# ```sql
# SELECT a.schoolname,count(*) as total_shooting_incidents from nyc_shooting u,nyc_schools a
# where st_contains(st_buffer(a.geom,50),u.geometry) group by a.schoolname
# ```
# -
disp = widgets.Output()
db = spatialite.connect('databases/spatialDB.sqlite')
stateGeomSql = f"""WITH a AS (select schoolname,st_buffer(geometry,50) as geom from nyc_schools)
SELECT a.schoolname,count(*) as total_shooting_incidents from nyc_shooting u,a
where st_contains(a.geom,u.geometry) and u.rowid in(SELECT ROWID
FROM SpatialIndex
WHERE f_table_name = 'nyc_shooting'
and f_geometry_column = 'geometry'
AND search_frame = a.geom) group by a.schoolname order by count(*) desc"""
data = pd.read_sql_query(stateGeomSql,con=db)
with disp:
display(data)
disp
# + [markdown] slideshow={"slide_type": "slide"}
# Let's look at another example
#
# **How many hospitals are there with in a specific distance of where you are**
#
# The required tables
# -
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
import spatialite
import pandas as pd
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select pk_uid,geom as geometry from hospitals limit 5',db)
table1_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>HOSPITALS</center></b>")
with table1_disp:
display(table1)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0'))])
out
# + [markdown] slideshow={"slide_type": "slide"}
# Let's look at an interactive example. Double click anywhere on the map and it will be selected as the current location. Based on the slider value selected (default is 3000 meter). You can change the slider to change the buffer value
# +
from ipyleaflet import Map, DrawControl,GeoData,LayerGroup,Polygon,GeoJSON,Marker
from ipywidgets import Button, HBox, VBox,widgets,Layout,GridspecLayout,IntSlider,HTML
from IPython.display import display
import spatialite
import pandas as pd
import geopandas as gpd
import json
import time
db = spatialite.connect('databases/spatialDB.sqlite')
coords=None
def handle_click(**kwargs):
global coords
if kwargs.get('type') == 'dblclick':
layer_group.clear_layers()
coords = kwargs.get('coordinates')
layer_group.add_layer(Marker(location=coords))
findHospitals()
def findHospitals():
global coords
if coords is not None:
stateGeomSql = f"""SELECT st_asbinary(u.geom) as geom from hospitals u
where st_contains(st_buffer(st_transform(MakePoint({coords[1]},{coords[0]},4326),3857),{radiusSlider.value}),
st_transform(u.geom,3857))"""
gdf = gpd.GeoDataFrame.from_postgis(stateGeomSql, db,crs = 'EPSG:4326')
if len(gdf)!=0:
geo_data = GeoData(geo_dataframe = gdf,
style={'color': 'black', 'radius':8, 'fillColor': '#3366cc', 'opacity':0.5, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
point_style={'radius': 5, 'color': 'red', 'fillOpacity': 0.8, 'fillColor': 'blue', 'weight': 3},
name = 'Release')
layer_group.add_layer(geo_data)
center = [gdf.centroid.y.values[0],gdf.centroid.x.values[0]]
sMap.center = center
sMap.zoom = 12
def radiusChanged(slider):
findHospitals()
sMap= Map(center=(41.482222, -81.669722), zoom=15,prefer_canvas =True)
radiusSlider = widgets.IntSlider(
value=3000,
min=0,
max=100000,
step=500,
description='Radius:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
radiusSlider.observe(radiusChanged, 'value')
layer_group = LayerGroup()
sMap.add_layer(layer_group)
sMap.on_interaction(handle_click)
filterParams=HBox([sMap,VBox([radiusSlider])])
filterParams
# + [markdown] slideshow={"slide_type": "slide"}
# ### Distance Function
#
# The distance function is used to find the distance between two geometries
#
# The function **st_distance(geometry A,geometry B)** returns the distance between geometry A and geometry B
#
# <img src = "supplementary/images/distance_restaurant_example.png" width = "500px">
#
# Let's look at an example
#
# **Show five nearest tourist place from your current location**
#
# The required tables
# + slideshow={"slide_type": "subslide"}
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
import spatialite
import pandas as pd
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select pk_uid,geom as geometry from toUrism limit 5',db)
table1_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>TOURISM</center></b>")
with table1_disp:
display(table1)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0'))])
out
# + slideshow={"slide_type": "slide"}
from ipyleaflet import Map, DrawControl,GeoData,LayerGroup,Polygon,GeoJSON,Marker
from ipywidgets import Button, HBox, VBox,widgets,Layout,GridspecLayout,IntSlider,HTML
from IPython.display import display
import spatialite
import pandas as pd
import geopandas as gpd
import json
import time
db = spatialite.connect('databases/spatialDB.sqlite')
coords=None
def handle_click(**kwargs):
global coords
if kwargs.get('type') == 'dblclick':
layer_group.clear_layers()
coords = kwargs.get('coordinates')
layer_group.add_layer(Marker(location=coords))
findNearestTouristPlaces()
def findNearestTouristPlaces():
global coords
if coords is not None:
stateGeomSql = f"""SELECT u.pk_uid,st_asbinary(u.geom) as geom,st_distance(st_transform(u.geom,3857),
st_transform(MakePoint({coords[1]},{coords[0]},4326),3857)) as
dist_to_tourist from tourism u order by dist_to_tourist asc limit 5"""
gdf = gpd.GeoDataFrame.from_postgis(stateGeomSql, db,crs = 'EPSG:4326')
geo_data = GeoData(geo_dataframe = gdf,
style={'color': 'black', 'radius':8, 'fillColor': '#3366cc', 'opacity':0.5, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
point_style={'radius': 5, 'color': 'red', 'fillOpacity': 0.8, 'fillColor': 'blue', 'weight': 3},
name = 'Release')
layer_group.add_layer(geo_data)
center = [gdf.centroid.y.values[0],gdf.centroid.x.values[0]]
sMap.center = center
sMap.zoom = 12
sMap= Map(center=(41.482222, -81.669722), zoom=15,prefer_canvas =True)
layer_group = LayerGroup()
sMap.add_layer(layer_group)
sMap.on_interaction(handle_click)
sMap
# + [markdown] slideshow={"slide_type": "slide"}
# And one final example
#
# **Show five nearest gas stations from your current location**
#
# The required table
# -
from ipywidgets import HBox, VBox,widgets,Layout,HTML
from IPython.display import display
import spatialite
import pandas as pd
db = spatialite.connect('databases/spatialDB.sqlite')
table1 = pd.read_sql_query('select pk_uid,geom from gas_stationS limit 5',db)
table1_disp = widgets.Output()
table1_header = widgets.HTML(value = f"<b><font color='red'><center>GAS_STATIONS</center></b>")
with table1_disp:
display(table1)
out=HBox([VBox([table1_header,table1_disp],layout = Layout(margin='0 100px 0 0'))])
out
# + slideshow={"slide_type": "slide"}
from ipyleaflet import Map, DrawControl,GeoData,LayerGroup,Polygon,GeoJSON,Marker
from ipywidgets import Button, HBox, VBox,widgets,Layout,GridspecLayout,IntSlider,HTML
from IPython.display import display
import spatialite
import pandas as pd
import geopandas as gpd
import json
import time
db = spatialite.connect('databases/spatialDB.sqlite')
coords=None
def handle_click(**kwargs):
global coords
if kwargs.get('type') == 'dblclick':
layer_group.clear_layers()
coords = kwargs.get('coordinates')
layer_group.add_layer(Marker(location=coords))
findNearestGasStation()
def findNearestGasStation():
global coords
if coords is not None:
stateGeomSql = f"""SELECT u.pk_uid,st_asbinary(u.geom) as geom,st_distance(u.geom,
MakePoint({coords[1]},{coords[0]},4326)) as
dist_to_gas_station from gas_stations u order by dist_to_gas_station asc limit 5"""
gdf = gpd.GeoDataFrame.from_postgis(stateGeomSql, db,crs = 'EPSG:4326')
geo_data = GeoData(geo_dataframe = gdf,
style={'color': 'black', 'radius':8, 'fillColor': '#3366cc', 'opacity':0.5, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
point_style={'radius': 5, 'color': 'red', 'fillOpacity': 0.8, 'fillColor': 'blue', 'weight': 3},
name = 'Release')
layer_group.add_layer(geo_data)
center = [gdf.centroid.y.values[0],gdf.centroid.x.values[0]]
sMap.center = center
sMap.zoom = 12
sMap= Map(center=(41.482222, -81.669722), zoom=15,prefer_canvas =True)
layer_group = LayerGroup()
sMap.add_layer(layer_group)
sMap.on_interaction(handle_click)
sMap
# + [markdown] slideshow={"slide_type": "slide"}
#
# # Congratulations!
#
#
# **You have finished an Hour of CI!**
#
#
# But, before you go ...
#
# 1. Please fill out a very brief questionnaire to provide feedback and help us improve the Hour of CI lessons. It is fast and your feedback is very important to let us know what you learned and how we can improve the lessons in the future.
# 2. If you would like a certificate, then please type your name below and click "Create Certificate" and you will be presented with a PDF certificate.
#
# <font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="https://forms.gle/JUUBm76rLB8iYppN7">Take the questionnaire and provide feedback</a></font>
#
#
# + hide_input=true slideshow={"slide_type": "-"} tags=["Hide", "Init"]
# This code cell loads the Interact Textbox that will ask users for their name
# Once they click "Create Certificate" then it will add their name to the certificate template
# And present them a PDF certificate
from PIL import Image
from PIL import ImageFont
from PIL import ImageDraw
from ipywidgets import interact
def make_cert(learner_name, lesson_name):
cert_filename = 'hourofci_certificate.pdf'
img = Image.open("../../../supplementary/hci-certificate-template.jpg")
draw = ImageDraw.Draw(img)
cert_font = ImageFont.load_default()
cert_font = ImageFont.truetype('../../../supplementary/cruft.ttf', 150)
cert_fontsm = ImageFont.truetype('../../../supplementary/cruft.ttf', 80)
w,h = cert_font.getsize(learner_name)
draw.text( xy = (1650-w/2,1100-h/2), text = learner_name, fill=(0,0,0),font=cert_font)
w,h = cert_fontsm.getsize(lesson_name)
draw.text( xy = (1650-w/2,1100-h/2 + 750), text = lesson_name, fill=(0,0,0),font=cert_fontsm)
img.save(cert_filename, "PDF", resolution=100.0)
return cert_filename
interact_cert=interact.options(manual=True, manual_name="Create Certificate")
@interact_cert(name="Your Name")
def f(name):
print("Congratulations",name)
filename = make_cert(name, 'Intermediate Geospatial Data')
print("Download your certificate by clicking the link below.")
# -
# <font size="+1"><a style="background-color:blue;color:white;padding:12px;margin:10px;font-weight:bold;" href="supplementary/hourofci_certificate.pdf?download=1" download="supplementary/hourofci_certificate.pdf">Download your certificate</a></font>
|
intermediate-lessons/geospatial-data/gd-5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Negative slice silently ignored when using ix
# See: <https://github.com/pydata/pandas/issues/2600>
import pandas as pd
pd.__version__
df = pd.DataFrame({'foo': range(5), 'bar': range(5)})
df.ix[2:4] # works as expected
df.ix[-2:] # always returns full DataFrame...
|
notebooks/2016-03-17-negative-slicing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# +
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to change the path if needed.)
school_data_to_load = "../Resources/schools_complete.csv"
student_data_to_load = "../Resources/students_complete.csv"
# Read the School Data and Student Data and store into a Pandas DataFrame
school_data_df = pd.read_csv(school_data_to_load)
student_data_df = pd.read_csv(student_data_to_load)
# Cleaning Student Names and Replacing Substrings in a Python String
# Add each prefix and suffix to remove to a list.
prefixes_suffixes = ["Dr. ", "Mr. ","Ms. ", "Mrs. ", "Miss ", " MD", " DDS", " DVM", " PhD"]
# Iterate through the words in the "prefixes_suffixes" list and replace them with an empty space, "".
for word in prefixes_suffixes:
student_data_df["student_name"] = student_data_df["student_name"].str.replace(word,"")
# Check names.
student_data_df.head(10)
# -
# ## Deliverable 1: Replace the reading and math scores.
#
# ### Replace the 9th grade reading and math scores at Thomas High School with NaN.
# Install numpy using conda install numpy or pip install numpy.
# Step 1. Import numpy as np.
import numpy as np
# Step 2. Use the loc method on the student_data_df to select all the reading scores from the 9th grade at Thomas High School and replace them with NaN.
student_data_df.loc[(student_data_df["grade"] == "9th") & (student_data_df["school_name"] == "Thomas High School"), "reading_score"] = np.nan
# Step 3. Refactor the code in Step 2 to replace the math scores with NaN.
student_data_df.loc[(student_data_df["grade"] == "9th") & (student_data_df["school_name"] == "Thomas High School"), "math_score"] = np.nan
# Step 4. Check thestudent data for NaN's.
student_data_df
# # <span style="color:red">**/\*\*Module 4 Work Below\*\*/**</span>
# ## <span style="color:blue">*Load and Read CSV Files*</span>
# add pandas dependency
import pandas as pd
# files to load
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
school_data_df = pd.read_csv(school_data_to_load)
school_data_df
# read student data file and store it in a pandas DataFrame
student_data_df = pd.read_csv(student_data_to_load)
student_data_df
# ## <span style="color:blue">*Find Missing Values*</span>
# determine if there are any missing values in the school data
school_data_df.count()
# determine if there are any missing values in the student data
student_data_df.count()
# determine if there are any missing values in the school data
school_data_df.isnull()
# determine if there are any missing values in the student data
student_data_df.isnull().sum()
# Determine if there are not any missing values in the school data.
school_data_df.notnull()
# Determine if there are not any missing values in the student data.
student_data_df.notnull().sum()
#
# ## <span style="color:blue">*Determine Data Types*</span>
# determine data types for the school DataFrame
school_data_df.dtypes
# ## <span style="color:blue">*Merge DataFrames*</span>
# add each prefix and suffix to remove from a list
prefixes_suffixes = ["Dr. ", "Mr. ","Ms. ", "Mrs. ", "Miss ", " MD", " DDS", " DVM", " PhD"]
# iterate through the words in the "prefixes_suffixes" list and replace tham with an empty space ''
for word in prefixes_suffixes:
student_data_df["student_name"] = student_data_df["student_name"].str.replace(word, "")
student_data_df
# combine the data into a single dataset
school_data_complete_df = pd.merge(student_data_df, school_data_df, on=["school_name", "school_name"])
school_data_complete_df
# ## <span style="color:blue">*Get the Number of Students*</span>
# get the total number of students
student_count = school_data_complete_df.count()
student_count
student_count = school_data_complete_df["Student ID"].count()
# calculate total number of schools
school_count = school_data_df["school_name"].count()
school_count
# calculate the total number of schools using school_data_complete_df
school_count_2 = school_data_complete_df["school_name"].unique()
len(school_count_2)
# ## <span style="color:blue">*Get the total budget*</span>
# calculate the total budget
total_budget = school_data_df["budget"].sum()
total_budget
# ## <span style="color:blue">*Get the Average Score*</span>
# calculate the average reading score
average_reading_score = school_data_complete_df["reading_score"].mean()
average_reading_score
# calculate the average reading score
average_math_score = school_data_complete_df["math_score"].mean()
average_math_score
# ## <span style="color:blue">*Get the Passing Percentages*</span>
passing_math = school_data_complete_df["math_score"] >= 70
passing_math
passing_reading = school_data_complete_df["reading_score"] >= 70
passing_reading
passing_math = school_data_complete_df[school_data_complete_df["math_score"] >= 70]
passing_math
passing_reading = school_data_complete_df[school_data_complete_df["reading_score"] >= 70]
passing_reading
# +
#calculate number of students passing math
passing_math_count = passing_math["student_name"].count()
# calculate the number of students passing reading
passing_reading_count = passing_reading["student_name"].count()
# -
print(passing_math_count)
print(passing_reading_count)
# +
# calculate the percent that passed math
passing_math_percentage = passing_math_count / float(student_count) * 100
# calculate percent that passed reading
passing_reading_percentage = passing_reading_count / float(student_count) * 100
# -
print(passing_math_percentage)
print(passing_reading_percentage)
# +
# Calculate the students who passed both math and reading
passing_math_reading = school_data_complete_df[(school_data_complete_df["math_score"] >= 70) & (school_data_complete_df["reading_score"] >= 70)]
passing_math_reading
# -
# Calculate the number of students who passed both math and reading
overall_passing_math_reading_count = passing_math_reading["student_name"].count()
overall_passing_math_reading_count
# Calculate the overall passing percentage.
overall_passing_percentage = overall_passing_math_reading_count / student_count * 100
overall_passing_percentage
# ## <span style="color:blue">*Create District Summary DataFrame*</span>
# adding a list of values with keys to create a new dataframe
district_summary_df = pd.DataFrame(
[{"Total Schools":school_count,
"Total Students":student_count,
"Total Budget": total_budget,
"Average Math Score": average_math_score,
"Average Reading Score": average_reading_score,
"% Passing Math": passing_math_percentage,
"% Passing Reading": passing_reading_percentage,
"% Overall Passing": overall_passing_percentage}])
district_summary_df
# ## <span style="color:blue">*Format Columns*</span>
# format the "Total Students" to have the comma for the thousands separator
district_summary_df["Total Students"] = district_summary_df["Total Students"].map("{:,}".format)
district_summary_df
# Format "Total Budget" to have the comma for a thousands separator, a decimal separator, and a "$".
district_summary_df["Total Budget"] = district_summary_df["Total Budget"].map("${:,.2f}".format)
district_summary_df
# +
# Format the columns.
district_summary_df["Average Math Score"] = district_summary_df["Average Math Score"].map("{:.1f}".format)
district_summary_df["Average Reading Score"] = district_summary_df["Average Reading Score"].map("{:.1f}".format)
district_summary_df["% Passing Math"] = district_summary_df["% Passing Math"].map("{:.0f}".format)
district_summary_df["% Passing Reading"] = district_summary_df["% Passing Reading"].map("{:.0f}".format)
district_summary_df["% Overall Passing"] = district_summary_df["% Overall Passing"].map("{:.0f}".format)
district_summary_df
# -
# ## <span style="color:blue">*Reorder Columns*</span>
# +
# reorder columns in the order you want them to appear
new_column_order = ["Total Schools", "Total Students", "Total Budget","Average Math Score", "Average Reading Score", "% Passing Math", "% Passing Reading", "% Overall Passing"]
# assign district summary df the new column order
district_summary_df = district_summary_df[new_column_order]
district_summary_df
# -
# # <span style="color:purple">**Generate School Summaries**</span>
# ## <span style="color:blue">*Set index to School Name*</span>
# determine the school type
per_school_types = school_data_df.set_index(["school_name"])["type"]
per_school_types
# add the per_school_types into a DataFrame for testing
df = pd.DataFrame(per_school_types)
# ## <span style="color:blue">*Get the Student Count per school*</span>
# calculate the total student count
per_school_counts = school_data_df.set_index(["school_name"])["size"]
per_school_counts
# calculate the total student counts using student_data_complete_df
per_school_counts = school_data_complete_df['school_name'].value_counts()
per_school_counts
# ## <span style="color:blue">*Get the Budget per student*</span>
# calculate the total school budget
per_school_budget = school_data_df.set_index(["school_name"])['budget']
per_school_budget
# calculate the per capita spending
per_school_capita = per_school_budget / per_school_counts
per_school_capita
# calculate the math scores
student_school_math = student_data_df.set_index(["school_name"])["math_score"]
student_school_math
# calculate the average math scores
per_school_averages = school_data_complete_df.groupby(["school_name"]).mean()
per_school_averages
# +
# calculate the average math scores
per_school_math = school_data_complete_df.groupby(["school_name"]).mean()["math_score"]
# calculate the average reading scores
per_school_reading = school_data_complete_df.groupby(["school_name"]).mean()['reading_score']
# -
per_school_math
per_school_reading
# +
# calculate the passing scores by creating a new dataframe
per_school_passing_math = school_data_complete_df[(school_data_complete_df["math_score"]) >= 70]
per_school_passing_reading = school_data_complete_df[(school_data_complete_df["reading_score"]) >= 70]
# -
per_school_passing_math
# +
per_school_passing_math = per_school_passing_math.groupby(["school_name"]).count()["student_name"]
per_school_passing_reading = per_school_passing_reading.groupby(["school_name"]).count()["student_name"]
# -
per_school_passing_math
# calculate the percentage of passing math and reading scores per school
per_school_passing_math = per_school_passing_math / per_school_counts * 100
per_school_passing_reading = per_school_passing_reading / per_school_counts * 100
per_school_passing_reading
# calculate the students who passed both math and reading
per_passing_math_reading = school_data_complete_df[(school_data_complete_df["math_score"] >= 70) & (school_data_complete_df["reading_score"] >= 70)]
per_passing_math_reading
# calculate the number of students who passed both math and reading
per_passing_math_reading = per_passing_math_reading.groupby(["school_name"]).count()["student_name"]
per_passing_math_reading
# calculate overall passing percentage
per_overall_passing_percentage = per_passing_math_reading / per_school_counts * 100
per_overall_passing_percentage
# +
# Adding a list of values with keys to create a new DataFrame.
per_school_summary_df = pd.DataFrame({
"School Type": per_school_types,
"Total Students": per_school_counts,
"Total School Budget": per_school_budget,
"Per Student Budget": per_school_capita,
"Average Math Score": per_school_math,
"Average Reading Score": per_school_reading,
"% Passing Math": per_school_passing_math,
"% Passing Reading": per_school_passing_reading,
"% Overall Passing": per_overall_passing_percentage})
per_school_summary_df.head()
# +
# Format the Total School Budget and the Per Student Budget columns.
per_school_summary_df["Total School Budget"] = per_school_summary_df["Total School Budget"].map("${:,.2f}".format)
per_school_summary_df["Per Student Budget"] = per_school_summary_df["Per Student Budget"].map("${:,.2f}".format)
# Display the data frame
per_school_summary_df
# +
# Reorder the columns in the order you want them to appear.
new_column_order = ["School Type", "Total Students", "Total School Budget", "Per Student Budget", "Average Math Score", "Average Reading Score", "% Passing Math", "% Passing Reading", "% Overall Passing"]
# Assign district summary df the new column order.
per_school_summary_df = per_school_summary_df[new_column_order]
per_school_summary_df.head()
# -
# # <span style="color:purple">**High and Low Performing Schools**</span>
# +
# sort and show the top 5 schools
top_schools = per_school_summary_df.sort_values(["% Overall Passing"], ascending = False)
top_schools.head()
# -
# sort and show top five lowest performing schools
bottom_schools = per_school_summary_df.sort_values(["% Overall Passing"], ascending=True)
bottom_schools.head()
# # <span style="color:purple">**Average Math and Reading Scores by Grade**</span>
# create a grade level DataFrames
ninth_graders = school_data_complete_df[(school_data_complete_df["grade"] == "9th")]
tenth_graders = school_data_complete_df[(school_data_complete_df["grade"] == "10th")]
eleventh_graders = school_data_complete_df[(school_data_complete_df["grade"] == "11th")]
twelfth_graders = school_data_complete_df[(school_data_complete_df["grade"] == "12th")]
ninth_graders
# Group each grade level DataFrame by the school name for the average math score
ninth_grade_math_scores = ninth_graders.groupby(["school_name"]).mean()["math_score"]
tenth_grade_math_scores = tenth_graders.groupby(["school_name"]).mean()["math_score"]
eleventh_grade_math_scores = eleventh_graders.groupby(["school_name"]).mean()["math_score"]
twelfth_grade_math_scores = twelfth_graders.groupby(["school_name"]).mean()["math_score"]
ninth_grade_math_scores
eleventh_grade_math_scores
# Group each grade level DataFrame by the school name for the average reading score
ninth_grade_reading_scores = ninth_graders.groupby(["school_name"]).mean()["reading_score"]
tenth_grade_reading_scores = tenth_graders.groupby(["school_name"]).mean()["reading_score"]
eleventh_grade_reading_scores = eleventh_graders.groupby(["school_name"]).mean()["reading_score"]
twelfth_grade_reading_scores = twelfth_graders.groupby(["school_name"]).mean()["reading_score"]
ninth_grade_reading_scores
twelfth_grade_reading_scores
# combine each grade level Series for average math scores by school into a single DataFrame
math_scores_by_grade = pd.DataFrame({
"9th":ninth_grade_math_scores,
"10th":tenth_grade_math_scores,
"11th":eleventh_grade_math_scores,
"12th":twelfth_grade_math_scores
})
math_scores_by_grade.head()
# +
# Combine each grade level Series for average reading scores by school into a single DataFrame
reading_scores_by_grade = pd.DataFrame({
"9th": ninth_grade_reading_scores,
"10th": tenth_grade_reading_scores,
"11th": eleventh_grade_reading_scores,
"12th": twelfth_grade_reading_scores})
reading_scores_by_grade.head()
# +
# Format each grade column
math_scores_by_grade["9th"] = math_scores_by_grade["9th"].map("{:.1f}".format)
math_scores_by_grade["10th"] = math_scores_by_grade["10th"].map("{:.1f}".format)
math_scores_by_grade["11th"] = math_scores_by_grade["11th"].map("{:.1f}".format)
math_scores_by_grade["12th"] = math_scores_by_grade["12th"].map("{:.1f}".format)
# Make sure the columns are in the correct order
math_scores_by_grade = math_scores_by_grade[
["9th", "10th", "11th", "12th"]]
# Remove the index name
math_scores_by_grade.index.name = None
# Display the DataFrame
math_scores_by_grade.head()
# +
# Format each grade column
reading_scores_by_grade["9th"] = reading_scores_by_grade["9th"].map("{:,.1f}".format)
reading_scores_by_grade["10th"] = reading_scores_by_grade["10th"].map("{:,.1f}".format)
reading_scores_by_grade["11th"] = reading_scores_by_grade["11th"].map("{:,.1f}".format)
reading_scores_by_grade["12th"] = reading_scores_by_grade["12th"].map("{:,.1f}".format)
# Make sure the columns are in the correct order
reading_scores_by_grade = reading_scores_by_grade[
["9th", "10th", "11th", "12th"]]
# Remove the index name
reading_scores_by_grade.index.name = None
# Display the data frame
reading_scores_by_grade.head()
# -
# # <span style="color:purple">**Group Scores by School Spending per Student**</span>
# Get the descriptive statistics for the per_school_capita
per_school_capita.describe()
# cut the per_school_capita into the spending ranges
spending_bins = [0,585,630,645,675]
group_names = ["<$584", "$585-629", "$630-644", "$645-675"]
per_school_capita.groupby(pd.cut(per_school_capita, spending_bins)).count()
# +
# Categorize spending based on the bins
per_school_summary_df["Spending Ranges (Per Student)"] = pd.cut(per_school_capita, spending_bins, labels=group_names)
per_school_summary_df
# -
# Calculate averages for the desired columns
spending_math_scores = per_school_summary_df.groupby(["Spending Ranges (Per Student)"]).mean()["Average Math Score"]
spending_reading_scores = per_school_summary_df.groupby(["Spending Ranges (Per Student)"]).mean()["Average Reading Score"]
spending_passing_math = per_school_summary_df.groupby(["Spending Ranges (Per Student)"]).mean()["% Passing Math"]
spending_passing_reading = per_school_summary_df.groupby(["Spending Ranges (Per Student)"]).mean()["% Passing Reading"]
overall_passing_spending = per_school_summary_df.groupby(["Spending Ranges (Per Student)"]).mean()["% Overall Passing"]
overall_passing_spending
# +
# Assemble into DataFrame
spending_summary_df = pd.DataFrame({
"Average Math Score" : spending_math_scores,
"Average Reading Score": spending_reading_scores,
"% Passing Math": spending_passing_math,
"% Passing Reading": spending_passing_reading,
"% Overall Passing": overall_passing_spending})
spending_summary_df
# +
# Formatting
spending_summary_df["Average Math Score"] = spending_summary_df["Average Math Score"].map("{:.1f}".format)
spending_summary_df["Average Reading Score"] = spending_summary_df["Average Reading Score"].map("{:.1f}".format)
spending_summary_df["% Passing Math"] = spending_summary_df["% Passing Math"].map("{:.0f}".format)
spending_summary_df["% Passing Reading"] = spending_summary_df["% Passing Reading"].map("{:.0f}".format)
spending_summary_df["% Overall Passing"] = spending_summary_df["% Overall Passing"].map("{:.0f}".format)
spending_summary_df
# -
# # <span style="color:purple">**Group Scores by School Size**</span>
# Establish the bins
size_bins = [0, 1000, 2000, 5000]
group_names = ["Small (<1000)", "Medium (1000-2000)", "Large (2000-5000)"]
# categorize spending based on bins
per_school_summary_df["School Size"] = pd.cut(per_school_summary_df["Total Students"], size_bins, labels=group_names)
per_school_summary_df.head()
# Calculate averages for the desired columns
size_math_scores = per_school_summary_df.groupby(["School Size"]).mean()["Average Math Score"]
size_reading_scores = per_school_summary_df.groupby(["School Size"]).mean()["Average Reading Score"]
size_passing_math = per_school_summary_df.groupby(["School Size"]).mean()["% Passing Math"]
size_passing_reading = per_school_summary_df.groupby(["School Size"]).mean()["% Passing Reading"]
size_overall_passing = per_school_summary_df.groupby(["School Size"]).mean()["% Overall Passing"]
# +
# Assemble into dataframe
size_summary_df = pd.DataFrame({
"Average Math Score" : size_math_scores,
"Average Reading Score": size_reading_scores,
"% Passing Math": size_passing_math,
"% Passing Reading": size_passing_reading,
"% Overall Passing": size_overall_passing})
size_summary_df
# +
# Formatting
size_summary_df["Average Math Score"] = size_summary_df["Average Math Score"].map("{:.1f}".format)
size_summary_df["Average Reading Score"] = size_summary_df["Average Reading Score"].map("{:.1f}".format)
size_summary_df["% Passing Math"] = size_summary_df["% Passing Math"].map("{:.0f}".format)
size_summary_df["% Passing Reading"] = size_summary_df["% Passing Reading"].map("{:.0f}".format)
size_summary_df["% Overall Passing"] = size_summary_df["% Overall Passing"].map("{:.0f}".format)
size_summary_df
# -
# # <span style="color:purple">**Group Scores by School Type**</span>
# +
# Calculate averages for the desired columns.
type_math_scores = per_school_summary_df.groupby(["School Type"]).mean()["Average Math Score"]
type_reading_scores = per_school_summary_df.groupby(["School Type"]).mean()["Average Reading Score"]
type_passing_math = per_school_summary_df.groupby(["School Type"]).mean()["% Passing Math"]
type_passing_reading = per_school_summary_df.groupby(["School Type"]).mean()["% Passing Reading"]
type_overall_passing = per_school_summary_df.groupby(["School Type"]).mean()["% Overall Passing"]
# +
# Assemble into DataFrame
type_summary_df = pd.DataFrame({
"Average Math Score" : type_math_scores,
"Average Reading Score": type_reading_scores,
"% Passing Math": type_passing_math,
"% Passing Reading": type_passing_reading,
"% Overall Passing": type_overall_passing})
type_summary_df
# +
# Formatting
type_summary_df["Average Math Score"] = type_summary_df["Average Math Score"].map("{:.1f}".format)
type_summary_df["Average Reading Score"] = type_summary_df["Average Reading Score"].map("{:.1f}".format)
type_summary_df["% Passing Math"] = type_summary_df["% Passing Math"].map("{:.0f}".format)
type_summary_df["% Passing Reading"] = type_summary_df["% Passing Reading"].map("{:.0f}".format)
type_summary_df["% Overall Passing"] = type_summary_df["% Overall Passing"].map("{:.0f}".format)
type_summary_df
# -
|
Challenge/PyCitySchools_Challenge_testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Automated Machine Learning
# _**Regression with Aml Compute**_
#
# ## Contents
# 1. [Introduction](#Introduction)
# 1. [Setup](#Setup)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Results](#Results)
# 1. [Test](#Test)
#
# ## Introduction
# In this example we use the Hardware Performance Dataset to showcase how you can use AutoML for a simple regression problem. The Regression goal is to predict the performance of certain combinations of hardware parts.
# After training AutoML models for this regression data set, we show how you can compute model explanations on your remote compute using a sample explainer script.
#
# If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../../../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
#
# An Enterprise workspace is required for this notebook. To learn more about creating an Enterprise workspace or upgrading to an Enterprise workspace from the Azure portal, please visit our [Workspace page.](https://docs.microsoft.com/azure/machine-learning/service/concept-workspace#upgrade)
#
# In this notebook you will learn how to:
# 1. Create an `Experiment` in an existing `Workspace`.
# 2. Instantiating AutoMLConfig with FeaturizationConfig for customization
# 3. Train the model using remote compute.
# 4. Explore the results and featurization transparency options
# 5. Setup remote compute for computing the model explanations for a given AutoML model.
# 6. Start an AzureML experiment on your remote compute to compute explanations for an AutoML model.
# 7. Download the feature importance for engineered features and visualize the explanations for engineered features on azure portal.
# 8. Download the feature importance for raw features and visualize the explanations for raw features on azure portal.
#
# ## Setup
#
# As part of the setup you have already created an Azure ML `Workspace` object. For Automated ML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
# +
import logging
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
import azureml.dataprep as dprep
from azureml.automl.core.featurization import FeaturizationConfig
from azureml.train.automl import AutoMLConfig
from azureml.core.dataset import Dataset
# -
# This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
print("This notebook was created using version 1.9.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
# +
ws = Workspace.from_config()
# Choose a name for the experiment.
experiment_name = 'automl-regression-hardware-explain'
experiment = Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ### Create or Attach existing AmlCompute
# You will need to create a [compute target](https://docs.microsoft.com/azure/machine-learning/service/concept-azure-machine-learning-architecture#compute-target) for your AutoML run. In this tutorial, you create `AmlCompute` as your training compute resource.
#
# **Creation of AmlCompute takes approximately 5 minutes.** If the AmlCompute with that name is already in your workspace this code will skip the creation process.
#
# As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your cluster.
amlcompute_cluster_name = "hardware-cluster"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=amlcompute_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_D2_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, amlcompute_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# ### Setup Training and Test Data for AutoML experiment
#
# Load the hardware dataset from a csv file containing both training features and labels. The features are inputs to the model, while the training labels represent the expected output of the model. Next, we'll split the data using random_split and extract the training data for the model. We also register the datasets in your workspace using a name so that these datasets may be accessed from the remote compute.
# +
data = 'https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/machineData.csv'
dataset = Dataset.Tabular.from_delimited_files(data)
# Split the dataset into train and test datasets
train_data, test_data = dataset.random_split(percentage=0.8, seed=223)
# Register the train dataset with your workspace
train_data.register(workspace = ws, name = 'machineData_train_dataset',
description = 'hardware performance training data',
create_new_version=True)
# Register the test dataset with your workspace
test_data.register(workspace = ws, name = 'machineData_test_dataset', description = 'hardware performance test data', create_new_version=True)
label ="ERP"
train_data.to_pandas_dataframe().head()
# -
# ## Train
#
# Instantiate an `AutoMLConfig` object to specify the settings and data used to run the experiment.
#
# |Property|Description|
# |-|-|
# |**task**|classification, regression or forecasting|
# |**primary_metric**|This is the metric that you want to optimize. Regression supports the following primary metrics: <br><i>spearman_correlation</i><br><i>normalized_root_mean_squared_error</i><br><i>r2_score</i><br><i>normalized_mean_absolute_error</i>|
# |**experiment_timeout_hours**| Maximum amount of time in hours that all iterations combined can take before the experiment terminates.|
# |**enable_early_stopping**| Flag to enble early termination if the score is not improving in the short term.|
# |**featurization**| 'auto' / 'off' / FeaturizationConfig Indicator for whether featurization step should be done automatically or not, or whether customized featurization should be used. Setting this enables AutoML to perform featurization on the input to handle *missing data*, and to perform some common *feature extraction*. Note: If the input data is sparse, featurization cannot be turned on.|
# |**n_cross_validations**|Number of cross validation splits.|
# |**training_data**|(sparse) array-like, shape = [n_samples, n_features]|
# |**label_column_name**|(sparse) array-like, shape = [n_samples, ], targets values.|
# ## Customization
#
# This step requires an Enterprise workspace to gain access to this feature. To learn more about creating an Enterprise workspace or upgrading to an Enterprise workspace from the Azure portal, please visit our [Workspace page.](https://docs.microsoft.com/azure/machine-learning/service/concept-workspace#upgrade).
#
# Supported customization includes:
# 1. Column purpose update: Override feature type for the specified column.
# 2. Transformer parameter update: Update parameters for the specified transformer. Currently supports Imputer and HashOneHotEncoder.
# 3. Drop columns: Columns to drop from being featurized.
# 4. Block transformers: Allow/Block transformers to be used on featurization process.
# Create FeaturizationConfig object using API calls
# + tags=["sample-featurizationconfig-remarks2"]
featurization_config = FeaturizationConfig()
featurization_config.blocked_transformers = ['LabelEncoder']
#featurization_config.drop_columns = ['MMIN']
featurization_config.add_column_purpose('MYCT', 'Numeric')
featurization_config.add_column_purpose('VendorName', 'CategoricalHash')
#default strategy mean, add transformer param for for 3 columns
featurization_config.add_transformer_params('Imputer', ['CACH'], {"strategy": "median"})
featurization_config.add_transformer_params('Imputer', ['CHMIN'], {"strategy": "median"})
featurization_config.add_transformer_params('Imputer', ['PRP'], {"strategy": "most_frequent"})
#featurization_config.add_transformer_params('HashOneHotEncoder', [], {"number_of_bits": 3})
# + tags=["sample-featurizationconfig-remarks3"]
automl_settings = {
"enable_early_stopping": True,
"experiment_timeout_hours" : 0.25,
"max_concurrent_iterations": 4,
"max_cores_per_iteration": -1,
"n_cross_validations": 5,
"primary_metric": 'normalized_root_mean_squared_error',
"verbosity": logging.INFO
}
automl_config = AutoMLConfig(task = 'regression',
debug_log = 'automl_errors.log',
compute_target=compute_target,
featurization=featurization_config,
training_data = train_data,
label_column_name = label,
**automl_settings
)
# -
# Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while.
# In this example, we specify `show_output = True` to print currently running iterations to the console.
remote_run = experiment.submit(automl_config, show_output = False)
remote_run
# Run the following cell to access previous runs. Uncomment the cell below and update the run_id.
# +
#from azureml.train.automl.run import AutoMLRun
#remote_run = AutoMLRun(experiment=experiment, run_id='<run_ID_goes_here')
#remote_run
# -
remote_run.wait_for_completion()
best_run, fitted_model = remote_run.get_output()
best_run_customized, fitted_model_customized = remote_run.get_output()
# ## Transparency
#
# View updated featurization summary
custom_featurizer = fitted_model_customized.named_steps['datatransformer']
custom_featurizer.get_featurization_summary()
# is_user_friendly=False allows for more detailed summary for transforms being applied
custom_featurizer.get_featurization_summary(is_user_friendly=False)
custom_featurizer.get_stats_feature_type_summary()
# ## Results
# #### Widget for Monitoring Runs
#
# The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
# ## Explanations
# This step requires an Enterprise workspace to gain access to this feature. To learn more about creating an Enterprise workspace or upgrading to an Enterprise workspace from the Azure portal, please visit our [Workspace page.](https://docs.microsoft.com/azure/machine-learning/service/concept-workspace#upgrade).
# This section will walk you through the workflow to compute model explanations for an AutoML model on your remote compute.
#
# ### Retrieve any AutoML Model for explanations
#
# Below we select the some AutoML pipeline from our iterations. The `get_output` method returns the a AutoML run and the fitted model for the last invocation. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
automl_run, fitted_model = remote_run.get_output(metric='r2_score')
# ### Setup model explanation run on the remote compute
# The following section provides details on how to setup an AzureML experiment to run model explanations for an AutoML model on your remote compute.
# #### Sample script used for computing explanations
# View the sample script for computing the model explanations for your AutoML model on remote compute.
with open('train_explainer.py', 'r') as cefr:
print(cefr.read())
# #### Substitute values in your sample script
# The following cell shows how you change the values in the sample script so that you can change the sample script according to your experiment and dataset.
# +
import shutil
import os
# create script folder
script_folder = './sample_projects/automl-regression-hardware'
if not os.path.exists(script_folder):
os.makedirs(script_folder)
# Copy the sample script to script folder.
shutil.copy('train_explainer.py', script_folder)
# Create the explainer script that will run on the remote compute.
script_file_name = script_folder + '/train_explainer.py'
# Open the sample script for modification
with open(script_file_name, 'r') as cefr:
content = cefr.read()
# Replace the values in train_explainer.py file with the appropriate values
content = content.replace('<<experiment_name>>', automl_run.experiment.name) # your experiment name.
content = content.replace('<<run_id>>', automl_run.id) # Run-id of the AutoML run for which you want to explain the model.
content = content.replace('<<target_column_name>>', 'ERP') # Your target column name
content = content.replace('<<task>>', 'regression') # Training task type
# Name of your training dataset register with your workspace
content = content.replace('<<train_dataset_name>>', 'machineData_train_dataset')
# Name of your test dataset register with your workspace
content = content.replace('<<test_dataset_name>>', 'machineData_test_dataset')
# Write sample file into your script folder.
with open(script_file_name, 'w') as cefw:
cefw.write(content)
# -
# #### Create conda configuration for model explanations experiment from automl_run object
# +
from azureml.core.runconfig import RunConfiguration
from azureml.core.conda_dependencies import CondaDependencies
import pkg_resources
# create a new RunConfig object
conda_run_config = RunConfiguration(framework="python")
# Set compute target to AmlCompute
conda_run_config.target = compute_target
conda_run_config.environment.docker.enabled = True
# specify CondaDependencies obj
conda_run_config.environment.python.conda_dependencies = automl_run.get_environment().python.conda_dependencies
# -
# #### Submit the experiment for model explanations
# Submit the experiment with the above `run_config` and the sample script for computing explanations.
# +
# Now submit a run on AmlCompute for model explanations
from azureml.core.script_run_config import ScriptRunConfig
script_run_config = ScriptRunConfig(source_directory=script_folder,
script='train_explainer.py',
run_config=conda_run_config)
run = experiment.submit(script_run_config)
# Show run details
run
# -
# %%time
# Shows output of the run on stdout.
run.wait_for_completion(show_output=True)
# ### Feature importance and visualizing explanation dashboard
# In this section we describe how you can download the explanation results from the explanations experiment and visualize the feature importance for your AutoML model on the azure portal.
# #### Download engineered feature importance from artifact store
# You can use *ExplanationClient* to download the engineered feature explanations from the artifact store of the *automl_run*. You can also use azure portal url to view the dash board visualization of the feature importance values of the engineered features.
from azureml.explain.model._internal.explanation_client import ExplanationClient
client = ExplanationClient.from_run(automl_run)
engineered_explanations = client.download_model_explanation(raw=False, comment='engineered explanations')
print(engineered_explanations.get_feature_importance_dict())
print("You can visualize the engineered explanations under the 'Explanations (preview)' tab in the AutoML run at:-\n" + automl_run.get_portal_url())
# #### Download raw feature importance from artifact store
# You can use *ExplanationClient* to download the raw feature explanations from the artifact store of the *automl_run*. You can also use azure portal url to view the dash board visualization of the feature importance values of the raw features.
raw_explanations = client.download_model_explanation(raw=True, comment='raw explanations')
print(raw_explanations.get_feature_importance_dict())
print("You can visualize the raw explanations under the 'Explanations (preview)' tab in the AutoML run at:-\n" + automl_run.get_portal_url())
# ## Operationailze
# In this section we will show how you can operationalize an AutoML model and the explainer which was used to compute the explanations in the previous section.
#
# ### Register the AutoML model and the scoring explainer
# We use the *TreeScoringExplainer* from *azureml.explain.model* package to create the scoring explainer which will be used to compute the raw and engineered feature importances at the inference time.
# In the cell below, we register the AutoML model and the scoring explainer with the Model Management Service.
# Register trained automl model present in the 'outputs' folder in the artifacts
original_model = automl_run.register_model(model_name='automl_model',
model_path='outputs/model.pkl')
scoring_explainer_model = automl_run.register_model(model_name='scoring_explainer',
model_path='outputs/scoring_explainer.pkl')
# ### Create the conda dependencies for setting up the service
# We need to create the conda dependencies comprising of the *azureml-explain-model*, *azureml-train-automl* and *azureml-defaults* packages.
# +
conda_dep = automl_run.get_environment().python.conda_dependencies
with open("myenv.yml","w") as f:
f.write(conda_dep.serialize_to_string())
with open("myenv.yml","r") as f:
print(f.read())
# -
# ### View your scoring file
with open("score_explain.py","r") as f:
print(f.read())
# ### Deploy the service
# In the cell below, we deploy the service using the conda file and the scoring file from the previous steps.
# +
from azureml.core.webservice import Webservice
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice
from azureml.core.model import Model
from azureml.core.environment import Environment
aciconfig = AciWebservice.deploy_configuration(cpu_cores=1,
memory_gb=1,
tags={"data": "Machine Data",
"method" : "local_explanation"},
description='Get local explanations for Machine test data')
myenv = Environment.from_conda_specification(name="myenv", file_path="myenv.yml")
inference_config = InferenceConfig(entry_script="score_explain.py", environment=myenv)
# Use configs and models generated above
service = Model.deploy(ws, 'model-scoring', [scoring_explainer_model, original_model], inference_config, aciconfig)
service.wait_for_deployment(show_output=True)
# -
# ### View the service logs
service.get_logs()
# ### Inference using some test data
# Inference using some test data to see the predicted value from autml model, view the engineered feature importance for the predicted value and raw feature importance for the predicted value.
if service.state == 'Healthy':
X_test = test_data.drop_columns([label]).to_pandas_dataframe()
# Serialize the first row of the test data into json
X_test_json = X_test[:1].to_json(orient='records')
print(X_test_json)
# Call the service to get the predictions and the engineered and raw explanations
output = service.run(X_test_json)
# Print the predicted value
print(output['predictions'])
# Print the engineered feature importances for the predicted value
print(output['engineered_local_importance_values'])
# Print the raw feature importances for the predicted value
print(output['raw_local_importance_values'])
# ### Delete the service
# Delete the service once you have finished inferencing.
service.delete()
# ## Test
# +
# preview the first 3 rows of the dataset
test_data = test_data.to_pandas_dataframe()
y_test = test_data['ERP'].fillna(0)
test_data = test_data.drop('ERP', 1)
test_data = test_data.fillna(0)
train_data = train_data.to_pandas_dataframe()
y_train = train_data['ERP'].fillna(0)
train_data = train_data.drop('ERP', 1)
train_data = train_data.fillna(0)
# +
y_pred_train = fitted_model.predict(train_data)
y_residual_train = y_train - y_pred_train
y_pred_test = fitted_model.predict(test_data)
y_residual_test = y_test - y_pred_test
# +
# %matplotlib inline
from sklearn.metrics import mean_squared_error, r2_score
# Set up a multi-plot chart.
f, (a0, a1) = plt.subplots(1, 2, gridspec_kw = {'width_ratios':[1, 1], 'wspace':0, 'hspace': 0})
f.suptitle('Regression Residual Values', fontsize = 18)
f.set_figheight(6)
f.set_figwidth(16)
# Plot residual values of training set.
a0.axis([0, 360, -100, 100])
a0.plot(y_residual_train, 'bo', alpha = 0.5)
a0.plot([-10,360],[0,0], 'r-', lw = 3)
a0.text(16,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_train, y_pred_train))), fontsize = 12)
a0.text(16,140,'R2 score = {0:.2f}'.format(r2_score(y_train, y_pred_train)),fontsize = 12)
a0.set_xlabel('Training samples', fontsize = 12)
a0.set_ylabel('Residual Values', fontsize = 12)
# Plot residual values of test set.
a1.axis([0, 90, -100, 100])
a1.plot(y_residual_test, 'bo', alpha = 0.5)
a1.plot([-10,360],[0,0], 'r-', lw = 3)
a1.text(5,170,'RMSE = {0:.2f}'.format(np.sqrt(mean_squared_error(y_test, y_pred_test))), fontsize = 12)
a1.text(5,140,'R2 score = {0:.2f}'.format(r2_score(y_test, y_pred_test)),fontsize = 12)
a1.set_xlabel('Test samples', fontsize = 12)
a1.set_yticklabels([])
plt.show()
# -
# %matplotlib inline
test_pred = plt.scatter(y_test, y_pred_test, color='')
test_test = plt.scatter(y_test, y_test, color='g')
plt.legend((test_pred, test_test), ('prediction', 'truth'), loc='upper left', fontsize=8)
plt.show()
|
how-to-use-azureml/automated-machine-learning/regression-explanation-featurization/auto-ml-regression-explanation-featurization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Exploring CSW access in Python using OWSLib with NODC Geoportal
from IPython.core.display import HTML
HTML('<iframe src=http://www.nodc.noaa.gov/geoportal/ width=900 height=280></iframe>')
from owslib.csw import CatalogueServiceWeb
# +
# connect to CSW, explore it's properties
endpoint = 'http://www.ngdc.noaa.gov/geoportal/csw' # NGDC Geoportal
#endpoint = 'http://data.nodc.noaa.gov/geoportal/csw' # NODC Geoportal: collection level
#endpoint = 'http://geodiscover.cgdi.ca/wes/serviceManagerCSW/csw' # NRCAN CUSTOM
#endpoint = 'http://geoport.whoi.edu/gi-cat/services/cswiso' # USGS Woods Hole GI_CAT
#endpoint = 'http://cida.usgs.gov/gdp/geonetwork/srv/en/csw' # USGS CIDA Geonetwork
#endpoint = 'http://www.nodc.noaa.gov/geoportal/csw' # NODC Geoportal: granule level
csw = CatalogueServiceWeb(endpoint,timeout=30)
csw.version
# -
[op.name for op in csw.operations]
#bbox=[-141,42,-52,84]
bbox=[-71.5, 39.5, -63.0, 46]
csw.getrecords(keywords=['temperature'],bbox=bbox,maxrecords=20)
#csw.getrecords(keywords=['sea_water_temperature'],maxrecords=20)
csw.results
for rec,item in csw.records.iteritems():
print item.title
print(csw.records.keys())
# choose a sample record
a=csw.records['satellite.G1.ssta.1day']
print a.title
# unfortunately the "uris" property is empty
print a.uris
# yet I can see the URIs here:
print a.xml
# lets look at the references
a.references
# get specific ServiceType URL from records
def service_urls(records,service_string='urn:x-esri:specification:ServiceType:OPeNDAP'):
urls=[]
for key,rec in records.iteritems():
#create a generator object, and iterate through it until the match is found
#if not found, gets the default value (here "none")
url = next((d['url'] for d in rec.references if d['scheme'] == service_string), None)
if url is not None:
urls.append(url)
return urls
dap_urls = service_urls(csw.records,service_string='urn:x-esri:specification:ServiceType:OPeNDAP')
print dap_urls
# find all the WMS ServiceType URLs
wms_urls = service_urls(csw.records,service_string='urn:x-esri:specification:ServiceType:WMS')
print wms_urls
a.uris
type a.uris
type(a.uris)
|
CSW/CSW Testing NODC Geoportal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# 
# -
# # <center> Algoritmi e Programmazione per l'Analisi dei Dati </center>
# ## <center> Project Assignment </center>
#
# #### <center> 2019-2020 </center>
# ##### <center> <NAME> </center>
# + [markdown] slideshow={"slide_type": "subslide"}
# #### <div style="text-align: center"> Abstract </div>
# -
# <center> The study consists, firstly, in handling with NetworkX graphs, while a second part is dedicated to implement an algorithm for constructing Eulerian paths on graphs, another one for computing degree distribution of graphs and eventually one for estimating the power law exponent. </center>
# +
__AUTHORS__ = {'gb': ("<NAME>",
"<EMAIL>",)}
__KEYWORDS__ = ['Python', 'Italian province', 'NetworkX', 'graphs', 'Eulerian path', 'degree distribution', 'power law']
# -
# <center><img src='img/python_logo.svg'></center>
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction
# -
# We will use Covid-19 data with respect to our country, Italy. Data are available at [Protezione Civile GitHub repository](https://github.com/pcm-dpc/COVID-19) and, especially, we will use JSON province data.
# + [markdown] slideshow={"slide_type": "fragment"}
# * *Bachelor degree*: Economia e Commercio, UniPi
# * *Master degree*: Statistica e Data Science, UniFi
# * *GitHub repository*: https://github.com/guidobei
# * *Computer*: HP Notebook
# * *Operating system*: Windows 10 Home
# * *Processor*: Intel Core i5-6200U CPU @ 2.30GHz 2.40GHz
# * *Installed RAM*: 4,00 GB
# * *System type*: 64-bit operating system, x64-based processor
# * *Python release*: Python 3.8.3
# * *IDE*: IDLE
# * *Environment*: Python Virtual Environment
# * *Libraries installed*:
# * `jupyter`
# * `numpy`
# * `pandas`
# * `matplotlib`
# * `networkx`
# + [markdown] slideshow={"slide_type": "subslide"}
# In the following slides we will see:
# + [markdown] slideshow={"slide_type": "fragment"}
# * An Italian province graph `P`
# + [markdown] slideshow={"slide_type": "fragment"}
# * A graph `R` with 2000 random nodes;
# + [markdown] slideshow={"slide_type": "fragment"}
# * `P` and `R` graphs in a weighted version;
# + [markdown] slideshow={"slide_type": "fragment"}
# * The *Seven Bridges of Königsberg* problem;
# + [markdown] slideshow={"slide_type": "fragment"}
# * An algorithm for constructing `Eulerian path` on graphs;
# + [markdown] slideshow={"slide_type": "fragment"}
# * An algorithm for computing `degree distribution` of graphs;
# + [markdown] slideshow={"slide_type": "fragment"}
# * An algorithm for estimating the `power law exponent`.
# + [markdown] slideshow={"slide_type": "subslide"}
# The following modules will be used:
# + slideshow={"slide_type": "fragment"}
import random
import numpy as np
import pandas as pd
import json
import matplotlib as mpl
import matplotlib.pyplot as plt
import networkx as nx
# -
# And the following file is loaded:
# + slideshow={"slide_type": "fragment"}
with open('data\dpc-covid19-ita-province.json') as f:
d = json.load(f)
# + [markdown] slideshow={"slide_type": "slide"}
# # Italian province graph
# + [markdown] slideshow={"slide_type": "fragment"}
# The scope of this chapter is to build the undirected graph `P` of Italian provinces, using `NetworkX`, in the most possible efficent way.
#
# Each node corresponds to a city and two cities *a* and *b* are connected by an edge if the following holds: if (*x*, *y*) is the position of *a*, where *x* is the longitude and *y* is the longitude, then *b* is in position (*z*, *w*) with *z* in [*x-d*, *x+d*] and *w* in [*y-d*, *y+d*], with *d* = 0.8.
# + [markdown] slideshow={"slide_type": "fragment"}
# * `Data cleaning` --> a dictionary of provinces is created, identified by their abbreviations, i.e. `{'LU': (43.84432283, 10.50151366), 'FI': (43.76923077, 11.25588885), 'NA': (40.83956555, 14.25084984), ...}` with 107 provinces;`
# + slideshow={"slide_type": "skip"}
def data_cleaning(ls):
'''Funzione che ritorna una dizionario di dati puliti, nel caso in cui venga passata una lista di dizionari.
Il dizionario ritornato ha come chiavi le sigle delle province, e come valori le tuple (latitudine, longitudine).'''
cleaned = {} # creo anche un dizionario con il duplice scopo di accelerare i tempi di controllo quando faccio il while se una provincia è già stata memorizzata e per memorizzare le posizioni
i = 0
while i < len(ls) and ls[i].get('sigla_provincia') not in cleaned: # faccio un while in modo che si fermi al primo doppione
if ls[i].get('sigla_provincia') != '':
cleaned[ls[i].get('sigla_provincia')] = (ls[i].get('lat'), ls[i].get('long'))
i += 1
return cleaned
# + [markdown] slideshow={"slide_type": "subslide"}
# # Simulated cities graph
# + [markdown] slideshow={"slide_type": "fragment"}
# We then generate 2000 pairs of double (*x*, *y*) with *x* in [*30*, *50*) and *y* in [*10*, *20*) and the algorithm is repeated, in order to build the undirected graph `R` where each pair is a node and two nodes are connected with the same rule reported above, except for the distance *d* = 0.08.
# + [markdown] slideshow={"slide_type": "fragment"}
# * `Data generating` --> a dictionary of fake cities is created, identified by a number, i.e. `{26: (43.84432283, 10.50151366), 567: (43.76923077, 11.25588885), 1994: (40.83956555, 14.25084984), ...}` with 2000 cities;
# + slideshow={"slide_type": "skip"}
def data_generating(n):
'''Funzione che ritorna un dizionari di dati casuali di n città che hanno latitudine nell'intervallo [30, 50] e
longitudine in [10, 20].
Il dizionario ritornato ha come chiavi il numero identificativo della città e come valori le tuple (latitudine,
longitudine).'''
casual = {i: (30 + 20 * random.random(), 10 + 10 * random.random()) for i in range(n)}
return casual
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The algorithm
# -
# Hereafter we present the algorithm, step by step:
# + [markdown] slideshow={"slide_type": "fragment"}
# * The dictionary got from the above processes is split into three lists, for efficient purposes: namely one with province `abbreviations`, one with `latitudes` and one with `longitudes`.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Lists are sorted by `latitude` or `longitude`, depending on their variances:
# * if the `latitude` variance of all the nodes is greater than `longitude` one, `latitude` list is sorted and the `longitude` list follows;
# * if the `longitude` variance of all the nodes is greater than `latitude` one, `longitude` list is sorted and the `latitude` list follows;
# * **it is important to maintain the corrispondences amongst the three lists throughout the elaboration**.
# + [markdown] slideshow={"slide_type": "fragment"}
# * Iteration is performed on the sorted list for city `i` and node is created;
# + [markdown] slideshow={"slide_type": "fragment"}
# * Another iteration is performed for node `j` starting from the following city: `j=i+1`;
# + [markdown] slideshow={"slide_type": "subslide"}
# * If the one-dimensionally distance between city `j` and city `i` is less or equal to 0.8 (0.08), the other dimension distance is checked: if also the latter one holds, the two nodes are connected and an edged is traced (with *Euclidean distance* as weight when requested).
# + [markdown] slideshow={"slide_type": "fragment"}
# * Iteration on `j` stops once a node `j` with distance from node `i` greater than 0.8 (0.08) is reached, as the following ones will necessarily have a greater distance.
# + [markdown] slideshow={"slide_type": "fragment"}
# * New iteration is then started on `i = i + 1` moving forward the above steps.
# + [markdown] slideshow={"slide_type": "fragment"}
# * The whole process stops once every city `i` is visited.
# + slideshow={"slide_type": "skip"}
def graph(diz, distanza):
'''Funzione che restituisce in tempo lineare un grafo quando le viene passata un dizionario ed una distanza per
impostare gli archi.
Il dizionario passato deve avere la sigla delle provincia come chiave e la tupla (latitudine, longitudine) come valori
corrispondenti. Se si intende lavorare con il file .json fornito dal dipartimento della Protezione Civile è
sufficiente pulire i dati con la funzione `data_cleaning(ls)`; se altrimenti si vuole procedere con una simulazione è
necessario ricorrere alla funzione `casual_data(n)`.
I vertici sono rappresentati dalle città e si traccia un arco tra di essi se ciò che segue è verificato: sia (x,y) la
posizione di a, allora b è in posizione (z,w), con z in [x-d, x+d] e w in [y-d, y+d], con d = distanza.'''
nomi = list(diz.keys())
lat = [avalue[0] for avalue in diz.values()]
long = [avalue[1] for avalue in diz.values()]
var_lat = np.var(lat)
var_long = np.var(long)
G = nx.Graph()
if var_lat > var_long:
tuples = zip(*sorted(zip(lat, nomi, long)))
lat, nomi, long = [list(tuple) for tuple in tuples]
edges = []
for i in range(len(nomi)):
nome_i = nomi[i]
G.add_node(nome_i, pos=(long[i], lat[i]))
j = i + 1
dist_lat = 0
while dist_lat <= distanza and j < len(nomi):
dist_lat = lat[j] - lat[i] # non prendo il valore assoluto perché è un valore già positivo
dist_long = abs(long[j] - long[i])
nome_j = nomi[j]
if dist_long <= distanza:
edges.append((nome_i, nome_j))
j += 1
else:
tuples = zip(*sorted(zip(long, nomi, lat)))
long, nomi, lat = [list(tuple) for tuple in tuples]
edges = []
for i in range(len(nomi)):
nome_i = nomi[i]
G.add_node(nome_i, pos=(long[i], lat[i]))
j = i + 1
dist_long = 0
while dist_long <= distanza and j < len(nomi):
dist_long = long[j] - long[i] # non prendo il valore assoluto perché è un valore già positivo
dist_lat = abs(lat[j] - lat[i])
nome_j = nomi[j]
if dist_lat <= distanza:
edges.append((nome_i, nome_j))
j += 1
G.add_edges_from(edges)
return G
# + slideshow={"slide_type": "skip"}
def weighted_graph(diz, distanza):
'''Funzione che restituisce in tempo lineare un grafo pesato quando le viene passata un dizionario ed una distanza per
impostare gli archi.
Il dizionario passato deve avere la sigla delle provincia come chiave e la tupla (latitudine, longitudine) come valori
corrispondenti. Se si intende lavorare con il file .json fornito dal dipartimento della Protezione Civile è
sufficiente pulire i dati con la funzione `data_cleaning(ls)`; se altrimenti si vuole procedere con una simulazione è
necessario ricorrere alla funzione `casual_data(n)`.
I vertici sono rappresentati dalle città e si traccia un arco tra di essi se ciò che segue è verificato: sia (x, y) la
posizione di a, allora b è in posizione (z,w), con z in [x-d, x+d] e w in [y-d, y+d], con d = distanza.'''
nomi = list(diz.keys())
lat = [avalue[0] for avalue in diz.values()]
long = [avalue[1] for avalue in diz.values()]
var_lat = np.var(lat)
var_long = np.var(long)
G = nx.Graph()
if var_lat > var_long:
tuples = zip(*sorted(zip(lat, nomi, long)))
lat, nomi, long = [list(tuple) for tuple in tuples]
edges = []
for i in range(len(nomi)):
nome_i = nomi[i]
G.add_node(nome_i, pos=(long[i], lat[i]))
j = i + 1
dist_lat = 0
while dist_lat <= distanza and j < len(nomi):
dist_lat = lat[j] - lat[i] # non prendo il valore assoluto perché è un valore già positivo
dist_long = abs(long[j] - long[i])
nome_j = nomi[j]
if dist_long <= distanza:
Eucl_dist = (dist_lat ** 2 + dist_long ** 2) ** (1/2)
edges.append((nome_i, nome_j, Eucl_dist))
j += 1
else:
tuples = zip(*sorted(zip(long, nomi, lat)))
long, nomi, lat = [list(tuple) for tuple in tuples]
edges = []
for i in range(len(nomi)):
nome_i = nomi[i]
G.add_node(nome_i, pos=(long[i], lat[i]))
j = i + 1
dist_long = 0
while dist_long <= distanza and j < len(nomi):
dist_long = long[j] - long[i] # non prendo il valore assoluto perché è un valore già positivo
dist_lat = abs(lat[j] - lat[i])
nome_j = nomi[j]
if dist_lat <= distanza:
Eucl_dist = (dist_long ** 2 + dist_lat ** 2) ** (1/2)
edges.append((nome_i, nome_j, Eucl_dist))
j += 1
G.add_weighted_edges_from(edges)
return G
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center> The Italian province graph P </center>
# + slideshow={"slide_type": "skip"}
dati_reali = data_cleaning(d)
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center> The simulated province graph R </center>
# + slideshow={"slide_type": "skip"}
dati_casuali = data_generating(2000)
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Time complexity analysis
# -
# The algorithm implemented in the previous slide is the most efficient we have found for *sparse* graphs: it takes linear time because, when we iterate on a node and since there are few edges, not every other node is scanned. Time complexity is $O(|V|)$, where $|V|$ is the number of nodes.
#
# However, a sorting algorithm has been run whose its time complexity is $O(|V|\log |V|)$.
#
# The expedient of computing the variance before deciding the dimension to be sorted guarantees the robustness of this method: if data had about the same values for a dimension, i.e. `longitude`, an eventual sorting by `longitude` would lead the algorithm to check each node with every node, and time complexity would be $O(V^2)$, the same as checking node by node with a two-nested for loop.
# + slideshow={"slide_type": "skip"}
P = graph(dati_reali, 0.8)
R = graph(dati_casuali, 0.08)
# + slideshow={"slide_type": "fragment"}
# %timeit P = graph(dati_reali, 0.8)
# %timeit R = graph(dati_casuali, 0.08)
# + [markdown] slideshow={"slide_type": "slide"}
# # Eulerian Path
# -
# In graph theory, an **Eulerian path** is a trail in a finite graph that visits every edge exactly once (allowing for revisiting vertices) and the graph is said to be *semi-Eulerian*. Similarly, an **Eulerian circuit** is an **Eulerian path** that starts and ends on the same vertex, and the graph is said to be *Eulerian*.
#
# They were first discussed by <NAME> (1707-1783) while solving the famous Seven Bridges of Königsberg problem in 1736.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Seven Bridges of Königsberg
# -
# In the 18th century, Königsberg in Prussia (now Kaliningrad, Russia), flowed through by the Pregel River and its tributaries, used to have two extensive islands that are connected to each other and to the two main areas of the city by seven bridges.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# An urban legend says that, on Sundays, the wealthy citizens of Königsberg used to walk around their city trying in vain to follow a path that crosses each bridge once and only once.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Euler's approach
# + [markdown] slideshow={"slide_type": "-"}
# The resolution of the problem was entrusted to the best living mathematician Leonard Euler, who showed that the hypothesized walk was not possible.
# + [markdown] slideshow={"slide_type": "fragment"}
# * **Abstraction of the map** --- by removing all the contingent aspects with the exception of urban areas delimited by the river arms and bridges that connect them.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# * **Modelling as a graph** --- by replacing each urban area with a point, now called `vertex`, and each bridge with a segment, called `edge`, inventing the modern graph theory.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# * **Attributing a number to each node** --- by computing how many edges a vertex is touched by, called `degree`.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Euler's theorem and solution
# + [markdown] slideshow={"slide_type": "-"}
# Before reaching a conclusion, Euler hypothesized different situations of zones and bridges: with four nodes and four bridges it is possible to start, for example, from A, and to go back through all the bridges once and only once. The degree of each node is an even number. If instead we start from A to get to D, each node is of equal degree with the exception of two nodes, of odd degree (1). Based on these observations, Euler enunciated the following theorem:
# > *Any connected graph is walkable if and only if it has all the nodes of even degree (**Eulerian circuit**), or two of them are of odd degree (**Eulerian path**); to traverse a "possible" graph with two nodes of odd degree, it is necessary to start from one of them, and it will end on the other odd node.*
# + [markdown] slideshow={"slide_type": "fragment"}
# * **Eulerian path**: two nodes of odd degree and all the other ones of even degree.
# + [markdown] slideshow={"slide_type": "fragment"}
# * **Eulerian circuit**: every node of even degree.
# + [markdown] slideshow={"slide_type": "subslide"}
# Hence, the *Seven Bridges of Königsberg* problem has no solution, because every node has an odd degree.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Fleury's algorithm
# -
# **Fleury's algorithm** (1883) is an elegant but inefficient algorithm that construct **Eulerian paths** and **Eulerian circuits**.
# + [markdown] slideshow={"slide_type": "subslide"}
# Here we present the algorithm, step by step:
# + [markdown] slideshow={"slide_type": "fragment"}
# * A graph is checked to have all edges in the same component and all nodes of even degree or exactly two of them of odd degree.
# + [markdown] slideshow={"slide_type": "fragment"}
# * The algorithm starts at a vertex of odd degree, or, if the graph has none, it starts with an arbitrarily chosen vertex.
#
# + [markdown] slideshow={"slide_type": "fragment"}
# * An edge whose removal would not disconnect the graph, called `bridge`, is chosen.
# + [markdown] slideshow={"slide_type": "fragment"}
# * To detect a `bridge`, we set up an ancillary algorithm that uses `DFS`:
# * let *G* be the graph and (*u*, *v*) the interested edge;
# * let *i* be the number of nodes reachable in *G* starting from *u*;
# * let us remove edge (*u*, *v*): a new graph *G'* is obtained;
# * let *j* be the number of nodes reachable in *G'* starting from *u*;
# * if *i* = *j* or *j* = 0, then edge(*u*, *v*) is not a `bridge`;
# * otherwise, edge(*u*, *v*) is a `bridge`.
# + [markdown] slideshow={"slide_type": "fragment"}
# * The process move on to the other endpoint of that edge and deletes the edge.
# + [markdown] slideshow={"slide_type": "subslide"}
# * The above steps are repeated until the algorithm stops once there are no edges left, and the sequence forms an **Eulerian cycle** if the starting point is the same as the ending point, or an **Eulerian path** if the starting point is different from the ending point.
# + slideshow={"slide_type": "skip"}
def eulerian_path(G):
"""Funzione che prende in input un grafo e che stabilisce, in primis, se questo è Euleriano, e, se questo è verificato,
ritorna il cammino euleriano.
La funzione è implementata secondo l'algoritmo di Fleury."""
CC = list(nx.connected_components(G))
CC_count = 0
for c in CC:
if len(c) != 1:
CC_count += 1
if CC_count != 1:
return 'This graph is not Eulerian nor semi-Eulerian.'
edges = G.edges
degree = G.degree
nodes = list(G.nodes)
odd_degree_nodes = 0
odd_list = []
for anode in nodes:
if degree[anode] % 2 != 0:
odd_degree_nodes += 1
odd_list.append(anode)
edge_path = []
if odd_degree_nodes == 2:
next_node = odd_list[random.randint(0, len(odd_list)-1)]
while edges:
current_node = next_node
node_edges = list(edges(current_node))
arandom = random.randint(0, len(node_edges)-1)
edge = node_edges[arandom]
if not is_bridge(edge, G):
edge_path.append(current_node)
next_node = edge[1]
G.remove_edge(*edge)
edge_path.append(next_node)
return edge_path
elif odd_degree_nodes == 0:
next_node = nodes[random.randint(0, len(nodes)-1)]
while edges:
current_node = next_node
node_edges = list(edges(current_node))
arandom = random.randint(0, len(node_edges)-1)
edge = node_edges[arandom]
if not is_bridge(edge, G):
edge_path.append(current_node)
next_node = edge[1]
G.remove_edge(*edge)
edge_path.append(next_node)
return edge_path
else:
return 'This graph is not Eulerian nor semi-Eulerian.'
def is_bridge(edge, G):
"""Funzione ausiliaria all'algoritmo di Fleury che stabilisce se un arco è un ponte oppure no. Prende in input l'arco
stesso ed il grafo interessato."""
H = G.copy()
next_node = edge[0]
i = dfs(next_node, H) - 1
H.remove_edge(*edge)
j = dfs(next_node, H) - 1
if i == j or j == 0:
return False
return True
def dfs(next_node, G):
'''Funzione di Depth First Search (DFS).'''
nodes = list(G.nodes())
colors = {anode: 'white' for anode in nodes}
return visit(next_node, colors, G)
def visit(current, colors, G, count=0):
'''Funzione ausiliara alla DFS.'''
count += 1
colors[current] = 'grey'
adj = list(G.adj[current])
for anode in adj:
if colors[anode] == 'white':
count = visit(anode, colors, G, count)
return count
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Applications
# -
# Our algorithm cannot be applied to our `P` and `R` graphs, as, firstly, they are not connected graphs.
# + slideshow={"slide_type": "fragment"}
P = graph(dati_reali, 0.8)
eulerian_path(P)
# + slideshow={"slide_type": "fragment"}
R = graph(dati_casuali, 0.08)
eulerian_path(R)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Eulerian path
# -
# Let's create an Eulerian graph as example, with exactly two odd nodes.
# + slideshow={"slide_type": "fragment"}
K = nx.Graph()
K.add_nodes_from([0, 1, 2, 3, 4, 5])
K.add_edges_from([(0, 1), (0, 5), (1, 2), (1, 4), (2, 3), (2, 4), (2, 5), (3, 4)])
nx.draw_shell(K, with_labels=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's start our algorithm from an odd vertex, i.e. *1*, and let's remove edge (*1*, *4*), which is not a *bridge*.
# + slideshow={"slide_type": "fragment"}
K.remove_edge(*(1, 4))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(4, 3))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(3, 2))
nx.draw_shell(K, with_labels=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# At this point, we should be careful. If we choose edge (*2*, *4*), we would get stuck in *4* and we would not be able to visit the other nodes. Edge (*2*, *4*) is so a *bridge*.
#
# We then move on vertex *5*.
# + slideshow={"slide_type": "fragment"}
K.remove_edge(*(2, 5))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(5, 0))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(0, 1))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(1, 2))
nx.draw_shell(K, with_labels=True)
# + slideshow={"slide_type": "subslide"}
K.remove_edge(*(2, 4))
nx.draw_shell(K, with_labels=True)
# + [markdown] slideshow={"slide_type": "fragment"}
# The process is complete and we got an **Eulerian path** $[1, 4, 3, 2, 5, 0, 1, 2, 4]$
# + slideshow={"slide_type": "subslide"}
K = nx.Graph()
K.add_nodes_from([0, 1, 2, 3, 4, 5])
K.add_edges_from([(0, 1), (0, 5), (1, 2), (1, 4), (2, 3), (2, 4), (2, 5), (3, 4)])
nx.draw_shell(K, with_labels=True)
eulerian_path(K)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Eulerian circuit: *House of St. Niklas*
# -
# Let's now see a graph that has an **Eulerian circuit**.
# + slideshow={"slide_type": "fragment"}
Q = nx.Graph()
Q.add_nodes_from([0, 1, 2, 3, 4, 5])
Q.add_edges_from([(0, 1), (0, 3), (1, 2), (1, 3), (1, 4), (2, 4), (3, 4), (3, 5), (4, 5)])
pos = {0: (0, 0), 1: (0.5, 0), 2: (1, 0), 3: (0, 4), 4: (1, 4), 5: (0.5, 8)}
nx.draw(Q, pos=pos, with_labels=True)
# + [markdown] slideshow={"slide_type": "subslide"}
# In this case, we can choose any vertex as starting point, since all the nodes are of even degree: we would never get stuck.
# + slideshow={"slide_type": "fragment"}
Q.remove_edge(*(0, 1))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(1, 2))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(2, 4))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(4, 1))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(1, 3))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(3, 4))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(4, 5))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(5, 3))
nx.draw(Q, pos=pos, with_labels=True)
# + slideshow={"slide_type": "subslide"}
Q.remove_edge(*(3, 0))
nx.draw(Q, pos=pos, with_labels=True)
# + [markdown] slideshow={"slide_type": "fragment"}
# The process is complete and we got an **Eulerian circuit** $[0, 1, 2, 4, 1, 3, 4, 5, 3, 0]$
# + slideshow={"slide_type": "subslide"}
Q = nx.Graph()
Q.add_nodes_from([0, 1, 2, 3, 4, 5])
Q.add_edges_from([(0, 1), (0, 3), (1, 2), (1, 3), (1, 4), (2, 4), (3, 4), (3, 5), (4, 5)])
pos = {0: (0, 0), 1: (0.5, 0), 2: (1, 0), 3: (0, 4), 4: (1, 4), 5: (0.5, 8)}
nx.draw(Q, pos=pos, with_labels=True)
eulerian_path(Q)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Time complexity analysis
# -
# While the graph traversal in **Fleury's algorithm** is linear in the number of edges, i.e. $O(|E|)$, an algorithm for detecting *bridges* has to be taken into account: this is linear in the number of edges as well, since a `DFS` is run.
#
# **Fleury's algorithm** will then have a time complexity of $O(|E|^2)$.
# + [markdown] slideshow={"slide_type": "slide"}
# # Centrality
# -
# In graph theory, indicators of **centrality** identify the most important vertices within a graph.
#
# **Centrality** indices are answers to the question "What characterizes an important vertex?". The answer is given in terms of a real-valued function on the vertices of a graph, where the values produced are expected to provide a ranking which identifies the most important nodes.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Degree centrality
# -
# Historically first and conceptually simplest meausure of centrality is **degree centrality**, which is defined as the number of links incident upon a node (i.e., the number of edges leaving a vertex).
# + [markdown] slideshow={"slide_type": "fragment"}
# A related index is the **degree distribution** that is the probability distribution of these degrees over the whole network.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Degree distribution
# -
# The **degree distribution** $P(d)$ of a graph is then defined to be the fraction of nodes in the network with degree $d$. Thus if there are $V$ nodes in total in a graph and $n_k$ of them have degree $d$, we have $P(d)$ = $n_d/V$.
# + slideshow={"slide_type": "skip"}
def degree_distribution(G):
'''Funzione che prende in input un grafo e ritorna un vocabolario che ha come chiavi i gradi e come valori le
frequenze.'''
degree = dict(G.degree)
degree_list = list(degree.values())
degree_voc = {adegree: None for adegree in degree_list} # faccio un vocabolario in modo da eliminare tutti i doppioni (mi costa O(n))
degree_list_ = [adegree for adegree in degree_voc] # riconverto tutto in lista
degree_list_.sort() # ordino la lista (mi costa(O(n*log(n)))
n = len(degree)
freq = [0]*(max(degree_list_)+1)
for i in degree_list:
freq[i] = freq[i] + 1
distr_dict = {}
for i in degree_list_:
distr_dict[i] = freq[i] / n
return distr_dict
def freq(G, k):
'''Funzione che prende in input un grafo ed un grado e ritorna la rispettiva frequenza'''
return degree_distribution(G).get(k)
# + slideshow={"slide_type": "subslide"}
degree_distribution(P)
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + slideshow={"slide_type": "subslide"}
degree_distribution(R)
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Power Law
#
# In social network, it seems that the degree distribution $P(d)$ follows a power law: $P(d)$ ∝ $\frac{1}{d^γ}$, for some constant $γ$, typically $1 ≤ γ ≤ 3$.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Power Law Exponent estimation
# + [markdown] slideshow={"slide_type": "-"}
# We would like to estimate the exponent $γ$ for our two graphs `P` and `R`.
# + [markdown] slideshow={"slide_type": "fragment"}
# There are many ways of estimating the value the **power law exponent**. We will used one based on the method of maximum likelihood.
# + [markdown] slideshow={"slide_type": "fragment"}
# Clauset et al. (2009) proposed the MLE for a discrete dataset, much more complicated than the continuos case, showing that the appropriate estimator for $α$ is given by the solution to the transcendental equation $\frac{ζ′(\hat{α}, x_{min})}{ζ(\hat{α}, x_{min})} = -\frac{1}{n}\sum_{i=1}^n \ln x_i$. In practice, evaluation of $\hat{α}$ requires us to solve this equation numerically. Alternatively, one can estimate $α$ by direct numerical maximization of the likelihood function itself, or equivalently of its logarithm (which is usually simpler): $L(α) = −n \ln ζ(α, x_{min}) − α\sum_{i=1}^n\ln x_i$,
#
# where $ζ(α, x_{min}) = \sum_{n=0}^\infty(n + x_min)^{-α} $
# + [markdown] slideshow={"slide_type": "subslide"}
# We then fix an $x_{min}$, greater than $0$, we calculate each likelihood score for any $α$ in $[1.06, 7.00]$ and we finally take the $α$ which has release the maximum score.
# + slideshow={"slide_type": "skip"}
def exponent_power_law(G, xmin=1):
"""Funzione che prende in input un grafo ed un x minimo (grado minimo da cui si vuole partire per la stima) e ritorna
una stima dell'esponente della power law.
xmin > 0, default=1."""
deg = list(dict(G.degree).values())
x = []
for el in deg:
if el >= xmin:
x.append(el)
n = len(x)
intervallo = []
for i in range(1, 1001):
q = 1 + 6*i/1000
intervallo.append(q)
LL = {}
for alfa in intervallo:
zeta = 0
for i in range(10000):
zeta = zeta + (i + xmin) ** (-alfa)
likelihood = -n*zeta - alfa*sum(np.log(x))
LL[alfa] = likelihood
alfa_hat = max(LL, key=LL.get)
return alfa_hat
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's see `P`.
# + slideshow={"slide_type": "fragment"}
exponent_power_law(P, 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's fit a power law starting from 7, degree from which the frequency seemed to decrease.
# + slideshow={"slide_type": "fragment"}
exponent_power_law(P, 7)
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# And now let's get done with `R`.
# + slideshow={"slide_type": "fragment"}
exponent_power_law(R, 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# 
|
slides.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import numpy as np
import xarray as xr
import matplotlib
import matplotlib.pyplot as plt
sys.path.append("..")
from forcing import Forcing
from Layer import LayerModel
from plotfunctions import prettyplot,quickread
np.seterr(all='ignore')
# %matplotlib notebook
# %matplotlib inline
# %config InlineBackend.print_figure_kwargs={'bbox_inches':None}
# %load_ext autoreload
# %autoreload 2
# +
# %%time
ds = quickread('Thwaites_e')
ds = Forcing(ds).tanh(ztcl=-550,Tdeep=0.5)
layer = LayerModel(ds)
ds = layer.compute(days=12)
prettyplot(ds,figsize=(10,10))
|
src/notebooks/Calc_layer_realistic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="CPQA-QCdiov-"
# # ME-314 - Final Project - <NAME>
#
# *In this project, I simulated the dynamics of a planar jack-in-a-box system. I simulated and animated the dynamics of the multi-body system in time, with external forces applied on the box and impact accrues between the two rigid bodies (the jack and the box’s walls).*
#
# *Both bodies in the planar systems are in gravity (in the -y direction of the world frame). In addition, two external forces are applied to the box – the first is an equal and opposite force to gravity (to ensure the box stays at the same position), and the second is a torque to rotate the box. The applied torque has a sinusoidal form to make the box rotate back and forth in a constant frequency.*
#
# ***
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 703, "status": "ok", "timestamp": 1605068634047, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ginp48i0jmOxCe1Ash9fdfU0b4Pe6DGLT9uqf_M=s64", "userId": "16296401493550679771"}, "user_tz": 360} id="1vKvR7-iiqev" outputId="753acb4d-cbc2-481b-ae37-64ec1ef337be"
import numpy as np
import sympy as sym
from sympy.abc import t
from sympy import sin, cos, symbols, Function, Inverse, Eq, Matrix, Rational, simplify, lambdify, solve
from math import pi
import matplotlib.pyplot as plt
# + id="uZ6LtTewiowO"
def mat_inverse(g):
''' A functions for ''invers'' operations.
'''
R = Matrix([[g[0,0], g[0,1], g[0,2]],
[g[1,0], g[1,1], g[1,2]],
[g[2,0], g[2,1], g[2,2]]])
R_inv = R.T
p = Matrix([g[0,3], g[1,3], g[2,3]])
p_inv = -R_inv * p
g_inv = Matrix([[R_inv[0,0], R_inv[0,1], R_inv[0,2], p_inv[0]],
[R_inv[1,0], R_inv[1,1], R_inv[1,2], p_inv[1]],
[R_inv[2,0], R_inv[2,1], R_inv[2,2], p_inv[2]],
[ 0, 0, 0, 1]])
return g_inv
def unhat(g):
''' A functions for ''unhat'' operations.
'''
V = Matrix([0, 0, 0, 0, 0, 0])
V[0, 0] = g[0, 3]
V[1, 0] = g[1, 3]
V[2, 0] = g[2, 3]
V[3, 0] = g[2, 1]
V[4, 0] = g[0, 2]
V[5, 0] = g[1, 0]
return V
def get_se3_np(x, y, theta):
"""
This function return SE(3) given x, y, theta
"""
return np.array([[np.cos(theta), -np.sin(theta), 0, x],
[np.sin(theta), np.cos(theta), 0, y],
[ 0, 0, 1, 0],
[ 0, 0, 0, 1]])
def integrate(f, xt, dt, time):
"""
This function takes in an initial condition x(t) and a timestep dt,
as well as a dynamical system f(x) that outputs a vector of the
same dimension as x(t). It outputs a vector x(t+dt) at the future
time step.
Parameters
============
dyn: Python function
derivate of the system at a given step x(t),
it can considered as \dot{x}(t) = func(x(t))
x0: NumPy array
current step x(t)
dt:
step size for integration
time:
step time
Return
============
new_x:
value of x(t+dt) integrated from x(t)
"""
k1 = dt * f(xt, time)
k2 = dt * f(xt+k1/2., time)
k3 = dt * f(xt+k2/2., time)
k4 = dt * f(xt+k3, time)
new_xt = xt + (1/6.) * (k1+2.0*k2+2.0*k3+k4)
return new_xt
def simulate_impact(f, x0, tspan, dt, integrate):
"""
This function takes in an initial condition x0, a timestep dt,
a time span tspan consisting of a list [min_time, max_time],
as well as a dynamical system f(x) that outputs a vector of the
same dimension as x0. It outputs a full trajectory simulated
over the time span of dimensions (xvec_size, time_vec_size).
Parameters
============
f: Python function
derivate of the system at a given step x(t),
it can considered as \dot{x}(t) = func(x(t))
x0: NumPy array
initial conditions
tspan: Python list
tspan = [min_time, max_time], it defines the start and end
time of simulation
dt:
time step for numerical integration
integrate: Python function
numerical integration method used in this simulation
Return
============
x_traj:
simulated trajectory of x(t) from t=0 to tf
"""
N = int((max(tspan)-min(tspan))/dt)
x = np.copy(x0)
tvec = np.linspace(min(tspan),max(tspan), N)
xtraj = np.zeros((len(x0), N))
time = 0
for i in range(N):
time = time + dt
(impact, impact_num) = impact_condition(x, phi_func, 1e-1)
if impact is True:
x = impact_update(x, impact_eqns_list[impact_num], dum_list)
xtraj[:, i]=integrate(f, x, dt, time)
else:
xtraj[:, i]=integrate(f, x, dt, time)
x = np.copy(xtraj[:,i])
return xtraj
# +
# Box parameters:
l_box, M_box = 6, 100
j_box = 4*M_box*l_box**2
# Jack parameters:
l_jack, m_jack = 1, 1
j_jack = 4*m_jack*l_jack**2
# General parameters:
# g = 0 # Without gravity
g = 9.81 # With gravity
# Define box and jack's variable:
x_box = Function(r'x_box')(t)
y_box = Function(r'y_box')(t)
theta_box = Function(r'\theta_box')(t)
x_jack = Function(r'x_jack')(t)
y_jack = Function(r'y_jack')(t)
theta_jack = Function(r'\theta_jack')(t)
# Define the external force:
theta_d_box = sin(2*pi*t/5)
k = 30000
F_y_box = 4*M_box*g
F_theta_box = k*(theta_d_box)
F = Matrix([0, F_y_box, F_theta_box, 0, 0, 0])
# Define the configuration vector:
q = Matrix([x_box, y_box, theta_box, x_jack, y_jack, theta_jack])
qdot = q.diff(t)
qddot = qdot.diff(t)
# Finding the homogeneous representations of the rigid bodies:
g_w_b = Matrix([[cos(theta_box), -sin(theta_box), 0, x_box], [sin(theta_box), cos(theta_box), 0, y_box], [0, 0, 1, 0], [0, 0, 0, 1]])
g_b_b1 = Matrix([[1, 0, 0, l_box], [0, 1, 0, l_box], [0, 0, 1, 0], [0, 0, 0, 1]])
g_b_b2 = Matrix([[1, 0, 0, 0], [0, 1, 0, -l_box], [0, 0, 1, 0], [0, 0, 0, 1]])
g_b_b3 = Matrix([[1, 0, 0, -l_box], [0, 1, 0, 0], [0, 0, 1, 0], [0, 0, 0, 1]])
g_b_b4 = Matrix([[1, 0, 0, 0], [0, 1, 0, l_box], [0, 0, 1, 0], [0, 0, 0, 1]])
g_w_j = Matrix([[cos(theta_jack), -sin(theta_jack), 0, x_jack], [sin(theta_jack), cos(theta_jack), 0, y_jack], [0, 0, 1, 0], [0, 0, 0, 1]])
g_j_j1 = Matrix([[1, 0, 0, l_jack], [0, 1, 0, 0], [0, 0, 1, 0], [0, 0, 0, 1]])
g_j_j2 = Matrix([[1, 0, 0, 0], [0, 1, 0, -l_jack], [0, 0, 1, 0], [0, 0, 0, 1]])
g_j_j3 = Matrix([[1, 0, 0, -l_jack], [0, 1, 0, 0], [0, 0, 1, 0], [0, 0, 0, 1]])
g_j_j4 = Matrix([[1, 0, 0, 0], [0, 1, 0, l_jack], [0, 0, 1, 0], [0, 0, 0, 1]])
g_w_b1 = g_w_b * g_b_b1
g_w_b2 = g_w_b * g_b_b2
g_w_b3 = g_w_b * g_b_b3
g_w_b4 = g_w_b * g_b_b4
g_w_j1 = g_w_j * g_j_j1
g_w_j2 = g_w_j * g_j_j2
g_w_j3 = g_w_j * g_j_j3
g_w_j4 = g_w_j * g_j_j4
g_b1_j1 = mat_inverse(g_w_b1) * g_w_j1
g_b1_j2 = mat_inverse(g_w_b1) * g_w_j2
g_b1_j3 = mat_inverse(g_w_b1) * g_w_j3
g_b1_j4 = mat_inverse(g_w_b1) * g_w_j4
g_b2_j1 = mat_inverse(g_w_b2) * g_w_j1
g_b2_j2 = mat_inverse(g_w_b2) * g_w_j2
g_b2_j3 = mat_inverse(g_w_b2) * g_w_j3
g_b2_j4 = mat_inverse(g_w_b2) * g_w_j4
g_b3_j1 = mat_inverse(g_w_b3) * g_w_j1
g_b3_j2 = mat_inverse(g_w_b3) * g_w_j2
g_b3_j3 = mat_inverse(g_w_b3) * g_w_j3
g_b3_j4 = mat_inverse(g_w_b3) * g_w_j4
g_b4_j1 = mat_inverse(g_w_b4) * g_w_j1
g_b4_j2 = mat_inverse(g_w_b4) * g_w_j2
g_b4_j3 = mat_inverse(g_w_b4) * g_w_j3
g_b4_j4 = mat_inverse(g_w_b4) * g_w_j4
# Calculate the rigid body velocities for the box and the jack:
V_box = unhat(mat_inverse(g_w_b) * g_w_b.diff(t))
V_jack = unhat(mat_inverse(g_w_j) * g_w_j.diff(t))
# Calculate the body inertia matrix:
I_box = Matrix([[4*M_box, 0, 0, 0, 0, 0], [0, 4*M_box, 0, 0, 0, 0], [0 ,0 ,4*M_box ,0 ,0 ,0], [0, 0, 0, 0, 0, 0], [0, 0, 0, 0, 0, 0], [0, 0, 0, 0, 0, j_box]])
I_jack = Matrix([[4*m_jack, 0, 0, 0, 0, 0], [0, 4*m_jack, 0, 0, 0, 0], [0, 0, 4*m_jack, 0, 0, 0], [0, 0, 0, 0, 0, 0], [0, 0, 0 ,0, 0, 0], [0, 0, 0, 0, 0, j_jack]])
# Calculate the Lagrangian:
KE = simplify(0.5*(V_box.T)*I_box*V_box + 0.5*(V_jack.T)*I_jack*V_jack)[0]
PE = simplify(g*(4*M_box*y_box + 4*m_jack*y_jack))
L = simplify(KE - PE)
# Compute the Euler-Lagrange Equations:
dL_dq = simplify(Matrix([L]).jacobian(q).T)
dL_dqdot = simplify(Matrix([L]).jacobian(qdot).T)
ddL_dqdot_dt = simplify(dL_dqdot.diff(t))
lhs = simplify(ddL_dqdot_dt - dL_dq)
rhs = simplify(F)
EL_Eqs = simplify(Eq(lhs, rhs))
display(EL_Eqs)
# Solve the Euler-Lagrange Equations:
EL_solns = solve(EL_Eqs, qddot, dict=True)
# +
x_box_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[0]])
y_box_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[1]])
theta_box_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[2]])
x_jack_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[3]])
y_jack_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[4]])
theta_jack_ddot_func = lambdify([q[0], q[1], q[2], q[3], q[4], q[5], qdot[0], qdot[1], qdot[2], qdot[3], qdot[4], qdot[5], t], EL_solns[0][qddot[5]])
def dynamics(s, t):
sdot = np.array([
s[6],
s[7],
s[8],
s[9],
s[10],
s[11],
x_box_ddot_func(*s, t),
y_box_ddot_func(*s, t),
theta_box_ddot_func(*s, t),
x_jack_ddot_func(*s, t),
y_jack_ddot_func(*s, t),
theta_jack_ddot_func(*s, t)
])
return sdot
# +
# Define acceleration matrix:
qddot_Matrix = Matrix([qdot[0], EL_solns[0][qddot[0]],
qdot[1], EL_solns[0][qddot[1]],
qdot[2], EL_solns[0][qddot[2]],
qdot[3], EL_solns[0][qddot[3]],
qdot[4], EL_solns[0][qddot[4]],
qdot[5], EL_solns[0][qddot[5]]])
# Define dummy symbols:
x_b_l, y_b_l, theta_b_l, x_j_l, y_j_l, theta_j_l, x_b_ldot, y_b_ldot, theta_b_ldot, x_j_ldot, y_j_ldot, theta_j_ldot = symbols('x_box_l, y_box_l, theta_box_l, x_jack_l, y_jack_l, theta_jack_l, x_box_ldot, y_box_ldot, theta_box_ldot, x_jack_ldot, y_jack_ldot, theta_jack_ldot')
dummy_dict = {q[0]:x_b_l, q[1]:y_b_l, q[2]:theta_b_l,
q[3]:x_j_l, q[4]:y_j_l, q[5]:theta_j_l,
qdot[0]:x_b_ldot, qdot[1]:y_b_ldot, qdot[2]:theta_b_ldot,
qdot[3]:x_j_ldot, qdot[4]:y_j_ldot, qdot[5]:theta_j_ldot}
qddot_d = qddot_Matrix.subs(dummy_dict)
qddot_lambdify = lambdify([x_b_l, x_b_ldot ,y_b_l, y_b_ldot, theta_b_l, theta_b_ldot,
x_j_l, x_j_ldot ,y_j_l, y_j_ldot, theta_j_l, theta_j_ldot, t], qddot_d)
# +
r_jack_hat = Matrix([x_jack, y_jack, theta_jack, 1])
# Define impact constraint for wall 1:
phi_b1_j1 = (g_b1_j1[3]).subs(dummy_dict)
phi_b1_j2 = (g_b1_j2[3]).subs(dummy_dict)
phi_b1_j3 = (g_b1_j3[3]).subs(dummy_dict)
phi_b1_j4 = (g_b1_j4[3]).subs(dummy_dict)
# Define impact constraint for wall 2:
phi_b2_j1 = (g_b2_j1[7]).subs(dummy_dict)
phi_b2_j2 = (g_b2_j2[7]).subs(dummy_dict)
phi_b2_j3 = (g_b2_j3[7]).subs(dummy_dict)
phi_b2_j4 = (g_b2_j4[7]).subs(dummy_dict)
# Define impact constraint for wall 3:
phi_b3_j1 = (g_b3_j1[3]).subs(dummy_dict)
phi_b3_j2 = (g_b3_j2[3]).subs(dummy_dict)
phi_b3_j3 = (g_b3_j3[3]).subs(dummy_dict)
phi_b3_j4 = (g_b3_j4[3]).subs(dummy_dict)
# Define impact constraint for wall 4:
phi_b4_j1 = (g_b4_j1[7]).subs(dummy_dict)
phi_b4_j2 = (g_b4_j2[7]).subs(dummy_dict)
phi_b4_j3 = (g_b4_j3[7]).subs(dummy_dict)
phi_b4_j4 = (g_b4_j4[7]).subs(dummy_dict)
# Define impact constraint:
phi_dum = simplify(Matrix([[phi_b1_j1], [phi_b1_j2], [phi_b1_j3], [phi_b1_j4], # all jack frames in the box1 frame
[phi_b2_j1], [phi_b2_j2], [phi_b2_j3], [phi_b2_j4], # all jack frames in the box2 frame
[phi_b3_j1], [phi_b3_j2], [phi_b3_j3], [phi_b3_j4], # all jack frames in the box3 frame
[phi_b4_j1], [phi_b4_j2], [phi_b4_j3], [phi_b4_j4]])) # all jack frames in the box4 frame
# Compute the Hamiltonian:
H = simplify((dL_dqdot.T * qdot)[0] - L)
# Compute expressions:
H_dum = H.subs(dummy_dict)
dL_dqdot_dum = dL_dqdot.subs(dummy_dict)
dPhidq_dum = phi_dum.jacobian([x_b_l, y_b_l, theta_b_l, x_j_l, y_j_l, theta_j_l])
# Define dummy symbols for tau+:
lamb = symbols(r'lambda')
x_b_dot_Plus, y_b_dot_Plus, theta_b_dot_Plus, x_j_dot_Plus, y_j_dot_Plus, theta_j_dot_Plus = symbols(r'x_box_dot_+, y_box_dot_+, theta_box_dot_+, x_jack_dot_+, y_jack_dot_+, theta_jack_dot_+')
impact_dict = {x_b_ldot:x_b_dot_Plus, y_b_ldot:y_b_dot_Plus, theta_b_ldot:theta_b_dot_Plus,
x_j_ldot:x_j_dot_Plus, y_j_ldot:y_j_dot_Plus, theta_j_ldot:theta_j_dot_Plus}
# Evaluate expressions at tau+:
dL_dqdot_dumPlus = simplify(dL_dqdot_dum.subs(impact_dict))
dPhidq_dumPlus = simplify(dPhidq_dum.subs(impact_dict))
H_dumPlus = simplify(H_dum.subs(impact_dict))
impact_eqns_list = []
# Define equations
lhs = Matrix([dL_dqdot_dumPlus[0] - dL_dqdot_dum[0],
dL_dqdot_dumPlus[1] - dL_dqdot_dum[1],
dL_dqdot_dumPlus[2] - dL_dqdot_dum[2],
dL_dqdot_dumPlus[3] - dL_dqdot_dum[3],
dL_dqdot_dumPlus[4] - dL_dqdot_dum[4],
dL_dqdot_dumPlus[5] - dL_dqdot_dum[5],
H_dumPlus - H_dum])
for i in range(phi_dum.shape[0]):
rhs = Matrix([lamb*dPhidq_dum[i,0],
lamb*dPhidq_dum[i,1],
lamb*dPhidq_dum[i,2],
lamb*dPhidq_dum[i,3],
lamb*dPhidq_dum[i,4],
lamb*dPhidq_dum[i,5],
0])
impact_eqns_list.append(simplify(Eq(lhs, rhs)))
# +
dum_list = [x_b_dot_Plus, y_b_dot_Plus, theta_b_dot_Plus,
x_j_dot_Plus, y_j_dot_Plus, theta_j_dot_Plus]
def impact_update(s, impact_eqns, dum_list):
""" This function updated the system after impact.
It returns the uptadet s array after impact.
"""
subs_dict = {x_b_l:s[0], y_b_l:s[1], theta_b_l:s[2],
x_j_l:s[3], y_j_l:s[4], theta_j_l:s[5],
x_b_ldot:s[6], y_b_ldot:s[7], theta_b_ldot:s[8],
x_j_ldot:s[9], y_j_ldot:s[10], theta_j_ldot:s[11]}
new_impact_eqns = impact_eqns.subs(subs_dict)
impact_solns = solve(new_impact_eqns, [x_b_dot_Plus, y_b_dot_Plus, theta_b_dot_Plus,
x_j_dot_Plus, y_j_dot_Plus, theta_j_dot_Plus,
lamb], dict=True)
if len(impact_solns) == 1:
print("Damn, only one solution")
else:
for sol in impact_solns:
lamb_sol = sol[lamb]
if abs(lamb_sol) < 1e-06:
pass
else:
return np.array([
s[0], #q will be the same after impact
s[1],
s[2],
s[3],
s[4],
s[5],
float(sym.N(sol[dum_list[0]])), #q_dot will change after impact
float(sym.N(sol[dum_list[1]])),
float(sym.N(sol[dum_list[2]])),
float(sym.N(sol[dum_list[3]])),
float(sym.N(sol[dum_list[4]])),
float(sym.N(sol[dum_list[5]])),
])
# +
phi_func = lambdify([x_b_l, y_b_l, theta_b_l,
x_j_l, y_j_l, theta_j_l,
x_b_ldot, y_b_ldot, theta_b_ldot,
x_j_ldot, y_j_ldot, theta_j_ldot],
phi_dum)
def impact_condition(s, phi_func, threshold = 1e-1):
""" This function checks the systems for impact.
In the case of an impact (abs(phi_val) < threshold),
the function returns True and the row number of the
phi matrix in which the impact accured.
"""
phi_val = phi_func(*s)
for i in range(phi_val.shape[0]):
if (phi_val[i] > -threshold) and (phi_val[i] < threshold):
return (True, i)
return (False, None)
# print('term impact condition function:', impact_condition(s_test, phi_func, 0.01))
# +
# Simulate the motion:
tspan = [0, 10]
dt = 0.01
s0 = np.array([0, 0, 0, 0, 0, 0, 0, 0, -1.6, 0, 0, 0])
N = int((max(tspan) - min(tspan))/dt)
tvec = np.linspace(min(tspan), max(tspan), N)
traj = simulate_impact(dynamics, s0, tspan, dt, integrate)
plt.figure()
plt.plot(tvec, traj[0], label='x_box')
plt.plot(tvec, traj[1], label='y_box')
plt.plot(tvec, traj[2], label='theta_box')
plt.title('Box Motion Simulation')
plt.xlabel('t')
plt.ylabel('state')
plt.legend(loc="best")
plt.show()
plt.figure()
plt.plot(tvec, traj[3], label='x_jack')
plt.plot(tvec, traj[4], label='y_jack')
plt.plot(tvec, traj[5], label='theta_jack')
plt.title('Jack Motion Simulation')
plt.xlabel('t')
plt.ylabel('state')
plt.legend(loc="best")
plt.show()
plt.figure()
plt.plot(tvec, traj[6], label='x_box_dot')
plt.plot(tvec, traj[7], label='y_box_dot')
plt.plot(tvec, traj[8], label='theta_box_dot')
plt.title('Box Velocity Simulation')
plt.xlabel('t')
plt.ylabel('state')
plt.legend(loc="best")
plt.show()
plt.figure()
plt.plot(tvec, traj[9], label='x_jack_dot')
plt.plot(tvec, traj[10], label='y_jack_dot')
plt.plot(tvec, traj[11], label='theta_jack_dot')
plt.title('Jack Velocity Simulation')
plt.xlabel('t')
plt.ylabel('state')
plt.legend(loc="best")
plt.show()
# +
def animate_jack_in_a_box(config_array,l=1,w=0.2,T=10):
"""
Function to generate web-based animation of the system
Parameters:
================================================
config_array:
trajectory of theta1 and theta2, should be a NumPy array with
shape of (2,N)
L1:
length of the first pendulum
L2:
length of the second pendulum
T:
length/seconds of animation duration
Returns: None
"""
################################
# Imports required for animation. (leave this part)
from plotly.offline import init_notebook_mode, iplot
from IPython.display import display, HTML
import plotly.graph_objects as go
#######################
# Browser configuration. (leave this part)
def configure_plotly_browser_state():
import IPython
display(IPython.core.display.HTML('''
<script src="/static/components/requirejs/require.js"></script>
<script>
requirejs.config({
paths: {
base: '/static/base',
plotly: 'https://cdn.plot.ly/plotly-1.5.1.min.js?noext',
},
});
</script>
'''))
configure_plotly_browser_state()
init_notebook_mode(connected=False)
###############################################
# Getting data from pendulum angle trajectories.
N = len(config_array[0])
x_box_array = config_array[0]
y_box_array = config_array[1]
theta_box_array = config_array[2]
x_jack_array = config_array[3]
y_jack_array = config_array[4]
theta_jack_array = config_array[5]
b1_x_array = np.zeros(N, dtype=np.float32)
b1_y_array = np.zeros(N, dtype=np.float32)
b2_x_array = np.zeros(N, dtype=np.float32)
b2_y_array = np.zeros(N, dtype=np.float32)
b3_x_array = np.zeros(N, dtype=np.float32)
b3_y_array = np.zeros(N, dtype=np.float32)
b4_x_array = np.zeros(N, dtype=np.float32)
b4_y_array = np.zeros(N, dtype=np.float32)
j_x_array = np.zeros(N, dtype=np.float32)
j_y_array = np.zeros(N, dtype=np.float32)
j1_x_array = np.zeros(N, dtype=np.float32)
j1_y_array = np.zeros(N, dtype=np.float32)
j2_x_array = np.zeros(N, dtype=np.float32)
j2_y_array = np.zeros(N, dtype=np.float32)
j3_x_array = np.zeros(N, dtype=np.float32)
j3_y_array = np.zeros(N, dtype=np.float32)
j4_x_array = np.zeros(N, dtype=np.float32)
j4_y_array = np.zeros(N, dtype=np.float32)
for t in range(N):
g_w_b = get_se3_np(x_box_array[t], y_box_array[t], theta_box_array[t])
g_w_j = get_se3_np(x_jack_array[t], y_jack_array[t], theta_jack_array[t])
b1 = g_w_b.dot(np.array([l_box, l_box, 0, 1]))
b1_x_array[t] = b1[0]
b1_y_array[t] = b1[1]
b2 = g_w_b.dot(np.array([l_box, -l_box, 0, 1]))
b2_x_array[t] = b2[0]
b2_y_array[t] = b2[1]
b3 = g_w_b.dot(np.array([-l_box, -l_box, 0, 1]))
b3_x_array[t] = b3[0]
b3_y_array[t] = b3[1]
b4 = g_w_b.dot(np.array([-l_box, l_box, 0, 1]))
b4_x_array[t] = b4[0]
b4_y_array[t] = b4[1]
j = g_w_j.dot(np.array([0, 0, 0, 1]))
j_x_array[t] = j[0]
j_y_array[t] = j[1]
j1 = g_w_j.dot(np.array([l_jack, 0, 0, 1]))
j1_x_array[t] = j1[0]
j1_y_array[t] = j1[1]
j2 = g_w_j.dot(np.array([0, -l_jack, 0, 1]))
j2_x_array[t] = j2[0]
j2_y_array[t] = j2[1]
j3 = g_w_j.dot(np.array([-l_jack, 0, 0, 1]))
j3_x_array[t] = j3[0]
j3_y_array[t] = j3[1]
j4 = g_w_j.dot(np.array([0, l_jack, 0, 1]))
j4_x_array[t] = j4[0]
j4_y_array[t] = j4[1]
####################################
# Axis limits.
xm = -13
xM = 13
ym = -13
yM = 13
###########################
# Defining data dictionary.
data=[dict(name = 'Box'),
dict(name = 'Jack'),
dict(name = 'Mass1_Jack'),
]
################################
# Preparing simulation layout.
layout=dict(xaxis=dict(range=[xm, xM], autorange=False, zeroline=False,dtick=1),
yaxis=dict(range=[ym, yM], autorange=False, zeroline=False,scaleanchor = "x",dtick=1),
title='Jack in a Box Simulation',
hovermode='closest',
updatemenus= [{'type': 'buttons',
'buttons': [{'label': 'Play','method': 'animate',
'args': [None, {'frame': {'duration': T, 'redraw': False}}]},
{'args': [[None], {'frame': {'duration': T, 'redraw': False}, 'mode': 'immediate',
'transition': {'duration': 0}}],'label': 'Pause','method': 'animate'}
]
}]
)
########################################
# Defining the frames of the simulation.
frames=[dict(data=[
dict(x=[b1_x_array[k],b2_x_array[k],b3_x_array[k],b4_x_array[k],b1_x_array[k]],
y=[b1_y_array[k],b2_y_array[k],b3_y_array[k],b4_y_array[k],b1_y_array[k]],
mode='lines',
line=dict(color='blue', width=3)
),
dict(x=[j1_x_array[k],j3_x_array[k],j_x_array[k],j2_x_array[k],j4_x_array[k]],
y=[j1_y_array[k],j3_y_array[k],j_y_array[k],j2_y_array[k],j4_y_array[k]],
mode='lines',
line=dict(color='green', width=3)
),
go.Scatter(
x=[j1_x_array[k],j2_x_array[k],j3_x_array[k],j4_x_array[k]],
y=[j1_y_array[k],j2_y_array[k],j3_y_array[k],j4_y_array[k]],
mode="markers",
marker=dict(color='darkgreen', size=6)),
]) for k in range(N)]
#######################################
# Putting it all together and plotting.
figure1=dict(data=data, layout=layout, frames=frames)
iplot(figure1)
##############
# The animation:
animate_jack_in_a_box(traj)
# -
|
code/final project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # Optical Character Recognition
#
# 
#
# A common computer vision challenge is to detect and interpret text in an image. This kind of processing is often referred to as *optical character recognition* (OCR).
#
# ## Use the Computer Vision Service to Read Text in an Image
#
# The **Computer Vision** cognitive service provides support for OCR tasks, including:
#
# - An **OCR** API that you can use to read text in multiple languages. This API can be used synchronously, and works well when you need to detect and read a small amount of text in an image.
# - A **Read** API that is optimized for larger documents. This API is used asynchronously, and can be used for both printed and handwritten text.
#
# You can use this service by creating either a **Computer Vision** resource or a **Cognitive Services** resource.
#
# If you haven't already done so, create a **Cognitive Services** resource in your Azure subscription.
#
# 1. In another browser tab, open the Azure portal at https://portal.azure.com, and sign in with your Microsoft account.
# 2. Click the **+Create a resource** button, search for *Cognitive Services*, and create a **Cognitive Services** resource with the following settings:
# - **Name**: *Enter a unique name*.
# - **Subscription**: *Your Azure subscription*.
# - **Location**: *Any available location*.
# - **Pricing tier**: S0
# - **Resource group**: *Create a resource group with a unique name*.
# 3. Wait for deployment to complete. Then go to your cognitive services resource, and on the **Overview** page, click the link to manage the keys for the service. You will need the endpoint and keys to connect to your cognitive services resource from client applications.
#
# ### Get the Key and Endpoint for your Cognitive Services resource
#
# To use your cognitive services resource, client applications need its endpoint and authentication key:
#
# 1. In the Azure portal, on the **Keys and Endpoint** page for your cognitive service resource, copy the **Key1** for your resource and paste it in the code below, replacing **YOUR_COG_KEY**.
# 2. Copy the **endpoint** for your resource and and paste it in the code below, replacing **YOUR_COG_ENDPOINT**.
# 3. Run the code in the cell below by clicking the **Run cell** (▷) button (to the left of the cell).
# + gather={"logged": 1599694246277}
cog_key = 'YOUR_COG_KEY'
cog_endpoint = 'YOUR_COG_ENDPOINT'
print('Ready to use cognitive services at {} using key {}'.format(cog_endpoint, cog_key))
# + [markdown] nteract={"transient": {"deleting": false}}
# Now that you've set up the key and endpoint, you can use your computer vision service resource to extract text from an image.
#
# To do this from Python, you'll need to run the following cell to install the Azure Cognitive Services Computer Vision package.
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# ! pip install azure-cognitiveservices-vision-computervision
# -
# Now you're ready to use the Computer Vision service to read the text in an image.
#
# Let's start with the **OCR** API, which enables you to synchronously analyze an image and read any text it contains. In this case, you have an adventising image for the fictional Northwind Traders retail company that includes some text. Run the cell below to read it.
# + gather={"logged": 1599694257280}
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw
import os
# %matplotlib inline
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Read the image file
image_path = os.path.join('data', 'ocr', 'advert.jpg')
image_stream = open(image_path, "rb")
# Use the Computer Vision service to find text in the image
read_results = computervision_client.recognize_printed_text_in_stream(image_stream)
# Process the text line by line
for region in read_results.regions:
for line in region.lines:
# Read the words in the line of text
line_text = ''
for word in line.words:
line_text += word.text + ' '
print(line_text.rstrip())
# Open image to display it.
fig = plt.figure(figsize=(7, 7))
img = Image.open(image_path)
draw = ImageDraw.Draw(img)
plt.axis('off')
plt.imshow(img)
# -
# The text found in the image is organized into a hierarchical structure of regions, lines, and words, and the code reads these to retrieve the results.
#
# In the results, view the text that was read above the image.
#
# ## Display bounding boxes
#
# The results also include *bounding box* coordinates for the lines of text and individual words found in the image. Run the cell below to see the bounding boxes for the lines of text in the advertising image you retrieved above.
# + gather={"logged": 1599694266106}
# Open image to display it.
fig = plt.figure(figsize=(7, 7))
img = Image.open(image_path)
draw = ImageDraw.Draw(img)
# Process the text line by line
for region in read_results.regions:
for line in region.lines:
# Show the position of the line of text
l,t,w,h = list(map(int, line.bounding_box.split(',')))
draw.rectangle(((l,t), (l+w, t+h)), outline='magenta', width=5)
# Read the words in the line of text
line_text = ''
for word in line.words:
line_text += word.text + ' '
print(line_text.rstrip())
# Show the image with the text locations highlighted
plt.axis('off')
plt.imshow(img)
# -
# In the result, the bounding box for each line of text is shown as a rectangle on the image.
#
# ## Use the Read API
#
# The OCR API you used previously works well for images with a small amount of text. When you need to read larger bodies of text, such as scanned documents, you can use the **Read** API. This requires a multi-step process:
#
# 1. Submit an image to the Computer Vision service to be read and analyzed asynchronously.
# 2. Wait for the analysis operation to complete.
# 3. Retrieve the results of the analysis.
#
# Run the following cell to use this process to read the text in a scanned letter to the manager of a Northwind Traders store.
# + gather={"logged": 1599694312346}
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from azure.cognitiveservices.vision.computervision.models import OperationStatusCodes
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image
import time
import os
# %matplotlib inline
# Read the image file
image_path = os.path.join('data', 'ocr', 'letter.jpg')
image_stream = open(image_path, "rb")
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Submit a request to read printed text in the image and get the operation ID
read_operation = computervision_client.read_in_stream(image_stream,
raw=True)
operation_location = read_operation.headers["Operation-Location"]
operation_id = operation_location.split("/")[-1]
# Wait for the asynchronous operation to complete
while True:
read_results = computervision_client.get_read_result(operation_id)
if read_results.status not in [OperationStatusCodes.running]:
break
time.sleep(1)
# If the operation was successfuly, process the text line by line
if read_results.status == OperationStatusCodes.succeeded:
for result in read_results.analyze_result.read_results:
for line in result.lines:
print(line.text)
# Open image and display it.
print('\n')
fig = plt.figure(figsize=(12,12))
img = Image.open(image_path)
plt.axis('off')
plt.imshow(img)
# -
# Review the results. There's a full transcription of the letter, which consists mostly of printed text with a handwritten signature. The original image of the letter is shown beneath the OCR results (you may need to scroll to see it).
#
# ## Read handwritten text
#
# In the previous example, the request to analyze the image specified a text recognition mode that optimized the operation for *printed* text. Note that despite this, the handwritten signature was read.
#
# This ability to read handwritten text is extremely useful. For example, suppose you've written a note containing a shopping list, and want to use an app on your phone to read the note and transcribe the text it contains.
#
# Run the cell below to see an example of a read operation for a handwritten shopping list.
# + gather={"logged": 1599694340593}
from azure.cognitiveservices.vision.computervision import ComputerVisionClient
from azure.cognitiveservices.vision.computervision.models import OperationStatusCodes
from msrest.authentication import CognitiveServicesCredentials
import matplotlib.pyplot as plt
from PIL import Image
import time
import os
# %matplotlib inline
# Read the image file
image_path = os.path.join('data', 'ocr', 'note.jpg')
image_stream = open(image_path, "rb")
# Get a client for the computer vision service
computervision_client = ComputerVisionClient(cog_endpoint, CognitiveServicesCredentials(cog_key))
# Submit a request to read printed text in the image and get the operation ID
read_operation = computervision_client.read_in_stream(image_stream,
raw=True)
operation_location = read_operation.headers["Operation-Location"]
operation_id = operation_location.split("/")[-1]
# Wait for the asynchronous operation to complete
while True:
read_results = computervision_client.get_read_result(operation_id)
if read_results.status not in [OperationStatusCodes.running]:
break
time.sleep(1)
# If the operation was successfuly, process the text line by line
if read_results.status == OperationStatusCodes.succeeded:
for result in read_results.analyze_result.read_results:
for line in result.lines:
print(line.text)
# Open image and display it.
print('\n')
fig = plt.figure(figsize=(12,12))
img = Image.open(image_path)
plt.axis('off')
plt.imshow(img)
# -
# ## More Information
#
# For more information about using the Computer Vision service for OCR, see [the Computer Vision documentation](https://docs.microsoft.com/en-us/azure/cognitive-services/computer-vision/concept-recognizing-text)
|
01e - Optical Character Recognition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/twwhatever/cs101/blob/master/ml/backprop/pytorch/Backprop.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="RXrsZaLlcvll" colab_type="text"
# # Backprop example
#
# End-to-end illustration of backpropagation using PyTorch.
#
# + id="1IioYVkRdDIZ" colab_type="code" colab={}
import torch
# + [markdown] id="4PVRiUyHdJcn" colab_type="text"
# We will use the x-or dataset as an example
# + id="_FR3QCyxdGSz" colab_type="code" colab={}
x = torch.tensor(
[
[0, 0],
[0, 1],
[1, 0],
[1, 1],
],
dtype=torch.float32,
)
y = torch.tensor(
[
0,
1,
1,
0,
],
dtype=torch.float32,
)
# + [markdown] id="kbi7TBDQdnOF" colab_type="text"
# We'll set up a two-layer network. PyTorch defines a bunch of ready-made layers and architectures, but for this example we'll explicitly build each of the parameters.
#
# For exposition purposes, we're setting the weights to a known-good initial point.
# + id="Mi52iR9MeUec" colab_type="code" colab={}
# Weights for layer 1
w1 = torch.tensor([[0.1, 0.3], [0.2, 0.1]], requires_grad=True)
# bias for layer 1
b1 = torch.tensor([0.2, -0.2], requires_grad=True)
# Weights for layer 2
w2 = torch.tensor([[0.1], [-0.4]], requires_grad=True)
# bias for layer 2
b2 = torch.tensor([0.0], requires_grad=True)
# + [markdown] id="LjsV3u95foRQ" colab_type="text"
# The first thing we need to do is compute the forward pass through the network and the loss. We'll use MSE (actually, it's square) as the loss for simplicity.
# + id="YHYURPTdfm8h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="fa78a9c0-09a4-495b-ab9f-dd2495c34f3c"
o1 = torch.relu(torch.matmul(x[0], w1) + b1)
print(o1)
o2 = torch.sigmoid(torch.matmul(o1, w2) + b2)
print(o2)
# MSE loss
loss = (y[0] - o2) ** 2
print(loss)
# + [markdown] id="9lxkQF4vi1GL" colab_type="text"
# The forward pass tracks operations on the parameters. For example, you can see the `.grad_fn` attribute in the tensors above.
#
# Now we use the loss to compute the backward pass.
# + id="E4a-9Aqvh9dT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="5e651bcd-758b-4a31-8eb4-beca16f0443e"
loss.backward()
print(w2.grad, b2.grad)
print(w1.grad, b1.grad)
# + [markdown] id="Cef8cO5ZkVhA" colab_type="text"
# The backward pass has computed the partial derivatives of the loss with respect to each parameter and stored the results in the `.grad` attribute for each parameter.
#
# Now we just need to perform the gradient step.
# + id="95Nk_SrijfiA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="6db4bdaf-9a2a-4c92-cbf1-d92b0b7d942d"
# learning rate
n = 0.01
# We don't want to track updates from gradient descent!
with torch.no_grad():
w1 -= n * w1.grad
b1 -= n * b1.grad
print(w1, b1)
w2 -= n * w2.grad
b2 -= n * b2.grad
print(w2, b2)
# + [markdown] id="fgyteV1Lmncw" colab_type="text"
# Now that we've gone through the basics, we'll write some convenience functions that allow us to illustrate a full training run.
# + id="279R-lXQkzTm" colab_type="code" colab={}
# Forward pass
def forward(x):
ol1 = torch.relu(torch.matmul(x, w1) + b1)
ol2 = torch.sigmoid(torch.matmul(ol1, w2) + b2)
return ol2
# Gradient update
def step(ts):
with torch.no_grad():
for t in ts:
t -= n * t.grad
# Zero gradients
def zero_grad(ts):
for t in ts:
if t.grad is not None:
t.grad.data.zero_()
# + id="kNhTtakUn61J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="e408a00c-e63d-46e2-fd4c-948d8e8718a3"
EPOCHS = 5000
for epoch in range(EPOCHS):
# "batch size" of 1
for i in range(4):
zero_grad([w1, b1, w2, b2])
loss = (y[i] - forward(x[i])) ** 2
loss.backward()
step([w1, b1, w2, b2])
# report loss on full dataset every 1000 epochs
if epoch % 1000 == 0:
loss = torch.mean((y - forward(x).squeeze()) ** 2)
print(f"EPOCH {epoch + 1} complete, loss {loss.item()}")
print(w1, b1)
print(w2, b2)
for i in range(4):
print(f"{y[i], forward(x[i])}")
|
ml/backprop/pytorch/Backprop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../fasp/runner/credits/images/FASPNotebook10.jpg" style="float: right;">
#
# ### GECCO PhenoPackets
#
# This notebook runs the same work as FASPScript10. In addition to showing how the three FASP steps can be run from a notebook, it uses the Seven Bridges WES Client in place of direct use of the Seven Bridges CGC API.
# +
# IMPORTS
from fasp.runner import FASPRunner
# The implementations we're using
from fasp.search import DiscoverySearchClient
from fasp.loc import DRSMetaResolver
from fasp.workflow import sbcgcWESClient
# +
pp_dbgap_join = """SELECT sp.dbGaP_Subject_ID, 'sbcgc:'||sb_drs_id
FROM dbgap_demo.scr_gecco_susceptibility.subject_phenotypes_multi sp
join dbgap_demo.scr_gecco_susceptibility.sample_multi sm on sm.dbgap_subject_id = sp.dbgap_subject_id
join dbgap_demo.scr_gecco_susceptibility.sb_drs_index di on di.sample_id = sm.sample_id
join sample_phenopackets.ga4gh_tables.gecco_phenopackets pp on pp.id = sm.biosample_accession
where json_extract_scalar(pp.phenopacket, '$.subject.sex') = 'MALE' and file_type = 'cram' limit 3 """
# Step 1 - Discovery
# query for relevant DRS objects
searchClient = DiscoverySearchClient('https://ga4gh-search-adapter-presto-public.prod.dnastack.com/')
# -
# Step 2 - DRS - a metaresolver will deal with which drs server is required
drsClient = DRSMetaResolver()
# Step 3 - set up a class that run a compute for us
faspRunner = FASPRunner()
settings = faspRunner.settings
wesClient = sbcgcWESClient(settings['SevenBridgesProject'])
# +
faspRunner.configure(searchClient, drsClient, wesClient)
faspRunner.runQuery(pp_dbgap_join, 'Phenopacket Gecco')
# -
|
notebooks/FASPNotebook10-PhenoPackets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
OABC is any quadrilateral in 3D space.
P is the midpoint of OA, Q is the midpoint of AB, R is the midpoint of BC and S is the midpoint of OC.
Prove that PQ is parallel to SR
#Define a coordinate system
from sympy.vector import CoordSys3D
Sys = CoordSys3D('Sys')
# Define point O to be Sys’ origin. We can do this without loss of generality
O = Sys.origin
# +
#Define point A with respect to O
from sympy import symbols
a1, a2, a3 = symbols('a1 a2 a3')
A = O.locate_new('A', a1*Sys.i + a2*Sys.j + a3*Sys.k)
# -
A
# +
# Similarly define points B and C
b1, b2, b3 = symbols('b1 b2 b3')
B = O.locate_new('B', b1*Sys.i + b2*Sys.j + b3*Sys.k)
c1, c2, c3 = symbols('c1 c2 c3')
C = O.locate_new('C', c1*Sys.i + c2*Sys.j + c3*Sys.k)
# -
B
C
# P is the midpoint of OA. Lets locate it with respect to O (you could also define it with respect to A).
P = O.locate_new('P', A.position_wrt(O) + (O.position_wrt(A) / 2))
P
# Similarly define points Q, R and S as per the problem definitions.
Q = A.locate_new('Q', B.position_wrt(A) / 2)
R = B.locate_new('R', C.position_wrt(B) / 2)
S = O.locate_new('R', C.position_wrt(O) / 2)
Q
R
S
# Now compute the vectors in the directions specified by PQ and SR.
PQ = Q.position_wrt(P)
SR = R.position_wrt(S)
PQ
SR
# Compute cross product
PQ.cross(SR)
# +
# Since the cross product is a zero vector, the two vectors have to be parallel, thus proving that PQ || SR.
|
Personal_Projects/vector math calculus 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pywikibot
import mwparserfromhell
import pprint
pp = pprint.PrettyPrinter(indent=4)
# -
category = "1985_video_games"
# ?pywikibot.Category
# +
site = pywikibot.Site('en', 'wikipedia') # any site will work, this is just an example
categoryPage = pywikibot.Category(site, title=category)
# -
articles = categoryPage.articles()
for article in articles:
print(article.title())
str('2')
|
notebooks/Category Scraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import psychrnn
from psychrnn.tasks import rdm as rd
from psychrnn.backend.models.basic import Basic
import tensorflow as tf
from matplotlib import pyplot as plt
# %matplotlib inline
rdm = rd.RDM(dt = 10, tau = 100, T = 2000, N_batch = 128)
gen = rdm.batch_generator()
params = rdm.__dict__
params['name'] = 'model'
params['N_rec'] = 50
model = Basic(params)
model.build()
model.train(gen)
x,_,_ = next(gen)
plt.plot(model.test(x)[0][0,:,:])
model.destruct()
# -
|
psychrnn/notebooks/Minimal_Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: threesix
# language: python
# name: threesix
# ---
# # Introduction
# A lesson code cell
2 + 2
# A lesson markdown cell
# > An empty cell below
|
dscreate/tests/files/no_solution_cells_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Converter-file" data-toc-modified-id="Converter-file-1"><span class="toc-item-num">1 </span>Converter file</a></span><ul class="toc-item"><li><span><a href="#Latest-and-Greatest" data-toc-modified-id="Latest-and-Greatest-1.1"><span class="toc-item-num">1.1 </span>Latest and Greatest</a></span></li></ul></li><li><span><a href="#Testing" data-toc-modified-id="Testing-2"><span class="toc-item-num">2 </span>Testing</a></span></li><li><span><a href="#New-Feature-Development" data-toc-modified-id="New-Feature-Development-3"><span class="toc-item-num">3 </span>New Feature Development</a></span></li><li><span><a href="#Morphology" data-toc-modified-id="Morphology-4"><span class="toc-item-num">4 </span>Morphology</a></span><ul class="toc-item"><li><span><a href="#MorphGNT:sblgnt-Data-Description" data-toc-modified-id="MorphGNT:sblgnt-Data-Description-4.1"><span class="toc-item-num">4.1 </span>MorphGNT:sblgnt Data Description</a></span></li></ul></li><li><span><a href="#ABC-dictionary-order" data-toc-modified-id="ABC-dictionary-order-5"><span class="toc-item-num">5 </span>ABC dictionary order</a></span></li><li><span><a href="#Word-Frequency" data-toc-modified-id="Word-Frequency-6"><span class="toc-item-num">6 </span>Word Frequency</a></span></li><li><span><a href="#English-Dictionary" data-toc-modified-id="English-Dictionary-7"><span class="toc-item-num">7 </span>English Dictionary</a></span></li></ul></div>
# -
# # Converter file
# ## Latest and Greatest
# +
import os
import re
import collections
import json
import csv
# from glob import glob
from tf.fabric import Fabric
from tf.convert.walker import CV
# from tf.compose import modify
source_dirs = 'input' # "input" is the name of the input folder that contains the source file
output_dirs = 'output' # "output" is the name of the output folder to which the finished TF files will be dumped into
bo2book = {line.split()[0]:line.split()[1] for line in '''
OTt4 Old_Testament
'''.split('\n') if line} # "OT" is the name of the file in the input folder AND "split()" splits at space
# patts = {'section': re.compile('(\d*):(\d*)\.(\d*)')}
def director(cv):
'''
Walks through LXX and triggers
slot and node creation events.
'''
# process books in order
for bo, book in bo2book.items():
book_loc = os.path.join(source_dirs, f'{bo}.txt')
print(f'\thandling {book_loc}...')
with open(book_loc, 'r', encoding="utf8") as infile:
text = [w for w in infile.read().split('\n') if w]
this_book = cv.node('book')
# keep track of when to trigger paragraph, chapter, and verse objects
# para_track = 1 # keep counts of paragraphs
prev_book = "Gen" # start at Genesis
prev_chap = 1 # start at 1
prev_verse = 1 # start at 1
prev_subverse = ''
wrdnum = 0 # start at 0
this_chap = cv.node('chapter')
# this_para = cv.node('paragraph')
this_verse = cv.node('verse')
this_subverse = cv.node('subverse')
# iterate through words and construct objects
for word in text:
wrdnum += 1
data = word.split('\t')
# word_data, lemmas = data[:7], data[7:]
word_data = data[:26] #the number here is the amount of columns
morphology = ' '.join(data[26:]) #the number here is the amount of columns
# segment out word data
# bo_code, ref, brake, ketiv, qere, morph, strongs = word_data
orig_order, book, chapter, verse, subverse, word, lex_utf8, g_cons_utf8, translit_SBL, lemma_gloss, strong, sp, morphology, case, nu, gn, degree, tense, voice, mood, ps, lemma_translit, abc_order, freq_lemma, BOL_lexeme_dict, BOL_gloss = word_data
# if chapter == "Prolog":
# chapter = 0
subverse == ""
#try:
# verse = int(verse)
#except ValueError:
# subverse = verse[-1:]
# verse = verse[:-1]
if verse == "":
print(f'{orig_order}: {verse} {subverse}')
# strongs_lemma, anlex_lemma = ' '.join(lemmas).split('!') # reconstitute lemmas and split on !
# chapt, verse, wrdnum = [int(v) for v in patts['section'].match(ref).groups()]
# -- handle TF events --
# detect book boundary
if prev_book != book:
# end subverse
cv.feature(this_subverse, subverse=prev_subverse)
cv.terminate(this_subverse)
# end verse
cv.feature(this_verse, verse=prev_verse)
cv.terminate(this_verse)
# end chapter
cv.feature(this_chap, chapter=prev_chap)
cv.terminate(this_chap)
# end book
cv.feature(this_book, book=prev_book)
cv.terminate(this_book)
# new book, chapter, verse, and subverse begin
this_book = cv.node('book')
prev_book = book
this_chap = cv.node('chapter')
prev_chap = chapter
this_verse = cv.node('verse')
prev_verse = verse
this_subverse = cv.node('subverse')
prev_subverse = subverse
wrdnum = 1
# detect chapter boundary
elif prev_chap != chapter:
# end subverse
cv.feature(this_subverse, subverse=prev_subverse)
cv.terminate(this_subverse)
# end verse
cv.feature(this_verse, verse=prev_verse)
cv.terminate(this_verse)
# end chapter
cv.feature(this_chap, chapter=prev_chap)
cv.terminate(this_chap)
# new chapter, verse, and subverse begin
this_chap = cv.node('chapter')
prev_chap = chapter
this_verse = cv.node('verse')
prev_verse = verse
this_subverse = cv.node('subverse')
prev_subverse = subverse
wrdnum = 1
# detect verse boundary
elif prev_verse != verse:
# end subverse
cv.feature(this_subverse, subverse=prev_subverse)
cv.terminate(this_subverse)
# end verse
cv.feature(this_verse, verse=prev_verse)
cv.terminate(this_verse)
# new verse and subverse begin
this_verse = cv.node('verse')
prev_verse = verse
this_subverse = cv.node('subverse')
prev_subverse = subverse
wrdnum = 1
# detect subverse boundary
elif prev_subverse != subverse:
cv.feature(this_subverse, subverse=prev_subverse)
cv.terminate(this_subverse)
this_subverse = cv.node('subverse')
prev_subverse = subverse
# detect paragraph boundary
# if brake == 'P':
# cv.feature(this_para, para=para_track)
# cv.terminate(this_para)
# this_para = cv.node('paragraph') # start a new paragraph
# para_track += 1 # count paragraphs in the book
# make word object
this_word = cv.slot()
cv.feature(this_word,
orig_order=orig_order,
book=book,
chapter=chapter,
verse=verse,
subverse=subverse,
word=word,
lex_utf8=lex_utf8,
g_cons_utf8=g_cons_utf8,
translit_SBL=translit_SBL,
lemma_gloss=lemma_gloss,
strong=strong,
sp=sp,
morphology=morphology,
case=case,
nu=nu,
gn=gn,
degree=degree,
tense=tense,
voice=voice,
mood=mood,
ps=ps,
lemma_translit=lemma_translit,
abc_order=abc_order,
freq_lemma=freq_lemma,
BOL_lexeme_dict=BOL_lexeme_dict,
BOL_gloss=BOL_gloss,
# ketiv=ketiv,
# qere=qere,
# strongs=strongs,
# str_lem=strongs_lemma.strip(),
# anlex_lem=anlex_lemma.strip()
)
cv.terminate(this_word)
# end book and its objects
# - end subverse
cv.feature(this_subverse, subverse=prev_subverse)
cv.terminate(this_subverse)
# - end verse
cv.feature(this_verse, verse=prev_verse)
cv.terminate(this_verse)
# - end paragraph
# cv.feature(this_para, para=para_track)
# cv.terminate(this_para)
# - end chapter
cv.feature(this_chap, chapter=prev_chap)
cv.terminate(this_chap)
# - end book
cv.feature(this_book, book=prev_book)
cv.terminate(this_book)
slotType = 'word'
otext = {'fmt:text-orig-full':'{word} ',
'sectionTypes':'book,chapter,verse',
'sectionFeatures':'book,chapter,verse'}
generic = {'Name': 'LXX',
'Version': '1935',
'Author': 'Rahlfs',
'Editors': 'CCAT, <NAME>',
'Converter': '<NAME>, <NAME>',
'Source:':'https://github.com/eliranwong/LXX-Rahlfs-1935',
'Note':'?'}
intFeatures = {'chapter', 'verse'}
featureMeta = {
'orig_order': {'description': 'original word order in corpus'},
'book': {'description': 'book'},
'chapter': {'description': 'chapter'},
'verse': {'description': 'verse'},
'subverse': {'description': 'subverse'},
'word': {'description': 'text realized word'},
'lex_utf8': {'description': 'normalized word'},
'g_cons_utf8': {'description': 'word without accents'},
'translit_SBL': {'description': 'SBL transliteration'},
'lemma_gloss': {'description': 'English gloss'},
'strong': {'description': 'Strong numbers'},
'sp': {'description': 'part of speech'},
'morphology': {'description': 'morphology'},
'case': {'description': 'case'},
'nu': {'description': 'number'},
'gn': {'description': 'gender'},
'degree': {'description': 'degree'},
'tense': {'description': 'tense'},
'voice': {'description': 'voice'},
'mood': {'description': 'mood'},
'ps': {'description': 'person'},
'lemma_translit': {'description': 'lemma transliteration'},
'abc_order': {'description': 'dictionary order'},
'freq_lemma': {'description': 'frequency of word in corpus'},
'BOL_lexeme_dict': {'description': 'BOL dictionary form of lemma'},
'BOL_gloss': {'description': 'BOL English gloss'},
# 'para': {'description': 'A paragraph number'},
# 'ketiv': {'descrption': 'The text as it is written in the printed Tischendorf'},
# 'qere': {'description': 'The text as the editor thinks it should have been'},
# 'strongs': {'description': 'A word\'s number in Strongs'},
# 'str_lem': {'description': 'Word lemma that corresponds to The NEW Strong\'sComplete Dictionary of Bible Words'},
# 'anlex_lem': {'description': 'Word lemma that corresponds to Friberg, Friberg and Miller\'s ANLEX'}
}
# configure metadata/output
version = '1935'
generic['Version'] = version
output = os.path.join(output_dirs, version)
print(f'Processing Version {version}')
output_dir = output_dirs.format(version=version)
TF = Fabric(locations=output_dir, silent=True)
cv = CV(TF)
cv.walk(director,
slotType,
otext=otext,
generic=generic,
intFeatures=intFeatures,
featureMeta=featureMeta,
warn=True,
force=False,)
# +
# First, I have to laod different modules that I use for analyzing the data and for plotting:
import sys, os, collections
import pandas as pd
import numpy as np
import re
import csv
import seaborn as sns
import matplotlib.pyplot as plt; plt.rcdefaults()
from matplotlib.pyplot import figure
from collections import Counter
# Second, I have to load the Text Fabric app
from tf.fabric import Fabric
from tf.app import use
# -
featureadd=pd.read_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_CCATLXX/LXX_source_v1.3.xlsx',sheet_name='FULL_data')
pd.set_option('display.max_columns', 50)
featureadd.head(10)
from unidecode import unidecode
featureadd['lemma_translit']=featureadd['lex_utf8'].apply(unidecode)
featureadd.head(5)
ABC1=featureadd[['orig_order','lex_utf8']]
ABC1.head(5)
ABC1.describe()
ABCdict = ABC1.drop_duplicates(['lex_utf8']).sort_values(by='lex_utf8', ascending=[True])
ABCdict.head(10)
ABCdict.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_CCATLXX/feature-dev/ABC1order.xlsx')
ABC2=pd.read_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_CCATLXX/feature-dev/ABC2order.xlsx')
pd.set_option('display.max_columns', 50)
ABC2.head(10)
ABC2=ABC2.drop(['orig_order'], axis=1)
ABC2.head()
featureadd.describe()
featureadd=pd.merge (featureadd, ABC2,
on='lex_utf8',
how='outer')
featureadd.head(5)
featureadd.describe()
featureaddstage2 = featureadd
featureaddstage2.head(5)
featureaddstage2.describe()
featureaddstage2["freq_lemma"]=featureaddstage2.groupby(["lex_utf8"])["lex_utf8"].transform("count")
featureaddstage2.head(5)
featureaddstage2.describe()
featureaddstage2.sort_values(['orig_order'], ascending=True).head(10)
featureaddstage2.describe()
BOLgreekDICT=pd.read_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/NA1904_dictionary_v1.0.xlsx')
pd.set_option('display.max_columns', 50)
BOLgreekDICT.head(10)
BOLgreekDICT=BOLgreekDICT[['Lexeme','Lexeme_dict', 'gloss']]
BOLgreekDICT.head(10)
BOLgreekDICT = BOLgreekDICT.rename({'Lexeme':'lex_utf8', 'Lexeme_dict':'BOL_lexeme_dict', 'gloss':'BOL_gloss'}, axis=1)
BOLgreekDICT.head(5)
BOLgreekDICT.describe()
featureaddstage3=featureaddstage2
featureaddstage3.describe()
featureaddstage4=pd.merge (featureaddstage3, BOLgreekDICT,
on='lex_utf8',
how='left')
featureaddstage4.head(5)
featureaddstage4.describe()
featureaddstage4 = featureaddstage4.drop_duplicates(['orig_order']).sort_values(by='orig_order', ascending=[True])
featureaddstage4.head(10)
featureaddstage4.describe()
featureaddstage4.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_CCATLXX/LXX_source_v1.4.xlsx')
# # Testing
# %load_ext autoreload
# %autoreload 2
# +
# First, I have to laod different modules that I use for analyzing the data and for plotting:
import sys, os, collections
import pandas as pd
import numpy as np
import re
import csv
import seaborn as sns
import matplotlib.pyplot as plt; plt.rcdefaults()
from matplotlib.pyplot import figure
from collections import Counter
# Second, I have to load the Text Fabric app
from tf.fabric import Fabric
from tf.app import use
# -
#LXX = use('CCATLXX/tf/1994_v1', hoist=globals())
LXX = use('CCATLXX/tf/1994_v2', hoist=globals())
LXX = use('D:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_CCATLXX/CCATLXX/tf/1994_v2', hoist=globals())
Search0 = '''
book book=Gen
chapter chapter=1
verse verse=1
word
'''
Search0 = LXX.search(Search0)
LXX.show(Search0, start=1, end=1, condensed=True, colorMap={1:'pink'}, extraFeatures={'orig_order','book','chapter','verse','subverse','word','lex_utf8','g_cons_utf8','translit_SBL','lemma_gloss','strong','sp','morphology','case','nu','gn','degree','tense','voice','mood','ps','lemma_translit','abc_order','freq_lemma','BOL_lexeme_dict','BOL_gloss'})
Search1 = '''
verse book=Gen chapter=1 verse=1
word
'''
Search1 = LXX.search(Search1)
LXX.show(Search1, start=1, end=1, condensed=True, colorMap={1:'pink'}, extraFeatures={'subverse'})
Search2 = '''
book book=Gen
chapter chapter=1
verse verse=1
word word* lex_utf8 g_cons_utf8 morphology* translit_SBL
'''
Search2 = LXX.search(Search2)
LXX.show(Search2, start=1, end=1, condensed=True, colorMap={1:'pink'}, extraFeatures={'subverse'})
Search3 = '''
book book=Num
chapter chapter=21
verse verse=3
'''
Search3 = LXX.search(Search3)
LXX.show(Search3, start=1, end=1, condensed=True, colorMap={1:'pink'}, extraFeatures={'word', 'lemma_translit', 'case'})
Eisakouw = '''
book book#Esth
verse
word lemma_translit=epakouo
word lemma_translit=theos|kurios
word lemma_translit=phone case=Gen
'''
Eisakouw = LXX.search(Eisakouw)
LXX.show(Eisakouw, start=1, end=100, condensed=True, colorMap={1:'pink'}, extraFeatures={'word', 'lemma_translit', 'case', 'lex_utf8', 'BOL_lexeme_dict'})
# # New Feature Development
# # Morphology
# ## MorphGNT:sblgnt Data Description
# Here: https://github.com/morphgnt/sblgnt
#
translitadd['lemma']=translitadd[0]
translitadd.head(5)
translitadd=translitadd[['lemma']]
translitadd.head(5)
translitadd['orig_order'] = translitadd.index +1
translitadd.head(5)
from unidecode import unidecode
s = "βίβλος"
s = unidecode(s)
print(s)
translitadd['translit'] = translitadd['lemma'].apply(unidecode)
translitadd.head(5)
translitadd['translit'].to_csv('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_translit.tf', index=None)
# # ABC dictionary order
#
ABC1=pd.read_csv('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_copy.tf',header=None, delimiter='\t',encoding='utf-8')
pd.set_option('display.max_columns', 50)
ABC1.head(10)
ABC1['lemma']=lemma[0]
ABC1.head(5)
ABC1['orig_order'] = ABC1.index +1
ABC1.head(5)
ABC1=ABC1[['orig_order','lemma']]
ABC1.head(5)
ABC1.describe()
ABCdict = ABC1.drop_duplicates(['lemma']).sort_values(by='lemma', ascending=[True])
ABCdict.head(10)
ABCdict.describe()
ABC1.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/ABC1order.xlsx', encoding='utf-8')
# Now I am ordering the word alphabetically iwth libreoffice writer since I cannot do that in pandas (yet?).
#
ABC2=pd.read_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/ABC2order.xlsx')
pd.set_option('display.max_columns', 50)
ABC2.head(10)
# Now we merge the ABCorder dataframe with the original lemma DF.
lemma_ABC=pd.merge (ABC1, ABC2,
on='lemma',
how='outer')
lemma_ABC.head(5)
lemma_ABC.describe()
lemma_ABC.sort_values(['orig_order_x'], ascending=True).head(10)
lemma_ABC.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_abc.xlsx')
# # Word Frequency
frequencyadd=pd.read_csv('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_copy.tf',header=None, delimiter='\t',encoding='utf-8')
pd.set_option('display.max_columns', 50)
frequencyadd.head(20)
frequencyadd['orig_order'] = frequencyadd.index +1
frequencyadd['lemma']=frequencyadd[0]
frequencyadd=frequencyadd[['orig_order','lemma']]
frequencyadd.head(5)
frequencyadd["freq_lemma"]=frequencyadd.groupby(["lemma"])["lemma"].transform("count")
#("count") is actually utilizing the 'count' function!
frequencyadd.head(20)
frequencyadd.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_freq.xlsx')
# # English Dictionary
# Lets first load the NA1904 BibleOL dictionary:
BOLgreekDICT=pd.read_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/NA1904_dictionary_v1.0.xlsx')
pd.set_option('display.max_columns', 50)
BOLgreekDICT.head(20)
BOLgreekDICT=BOLgreekDICT[['Lexeme','Lexeme_dict', 'Strong\'s number', 'gloss']]
BOLgreekDICT.head(10)
BOLgreekDICT.describe()
# Lets load the SBLGNT lemmas
SBLGNTlemmas=pd.read_csv('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/lemma_copy.tf',header=None, delimiter='\t',encoding='utf-8')
pd.set_option('display.max_columns', 50)
SBLGNTlemmas.head(2)
SBLGNTlemmas['orig_order']=SBLGNTlemmas.index +1
SBLGNTlemmas['Lexeme']=SBLGNTlemmas[0]
SBLGNTlemmas=SBLGNTlemmas[['orig_order','Lexeme']]
SBLGNTlemmas.head(5)
SBLGNTlemmas.describe()
# Now lets try a merge of the two files
SBLGNTglosses=pd.merge (SBLGNTlemmas,BOLgreekDICT,
on='Lexeme',
how='outer')
SBLGNTglosses.head(5)
SBLGNTglosses.describe()
SBLGNTglosses.head(20)
SBLGNTglosses.to_excel('d:/OneDrive/1200_AUS-research/Fabric-TEXT/0_data_SBLGNT/feature-dev/SBLGNTglosses.xlsx')
|
programs/CCATLXX TF feature production.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="0Hk4wNbNytnB"
# # ResNet-101 on CIFAR-10
#
#
#
#
# + [markdown] id="LFpHDxd6HHLy"
# ### Imports
# + id="lEpof2LG7NA3"
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchvision import datasets
from torchvision import transforms
from torch.utils.data import DataLoader
if torch.cuda.is_available():
torch.backends.cudnn.deterministic = True
# + [markdown] id="PXz_dHMTHHL0"
# ### Settings and Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="9zWGjESt2QYa" outputId="949a0c95-cb44-49e7-ac68-8e6e2df70a26"
# Device
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# Hyperparameters
random_seed = 1
learning_rate = 0.001
num_epochs = 10
batch_size = 128
torch.manual_seed(random_seed)
# Architecture
num_features = 784
num_classes = 10
# Data
train_dataset = datasets.CIFAR10(root='data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = datasets.CIFAR10(root='data',
train=False,
transform=transforms.ToTensor())
train_loader = DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# Checking the dataset
for images, labels in train_loader:
print('Image batch dimensions:', images.shape)
print('Image label dimensions:', labels.shape)
break
# + [markdown] id="F1a6OD4-HHL3"
# ### Model
# + id="8PPnruRVXvOP"
def conv3x3(in_planes, out_planes, stride=1):
"""3x3 convolution with padding"""
return nn.Conv2d(in_planes, out_planes, kernel_size=3, stride=stride,
padding=1, bias=False)
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv2d(inplanes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3, stride=stride,
padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv3 = nn.Conv2d(planes, planes * 4, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm2d(planes * 4)
self.relu = nn.ReLU(inplace=True)
self.downsample = downsample
self.stride = stride
def forward(self, x):
residual = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
out = self.relu(out)
out = self.conv3(out)
out = self.bn3(out)
if self.downsample is not None:
residual = self.downsample(x)
out += residual
out = self.relu(out)
return out
class ResNet(nn.Module):
def __init__(self, block, layers, num_classes, grayscale):
self.inplanes = 64
if grayscale:
in_dim = 1
else:
in_dim = 3
super(ResNet, self).__init__()
self.conv1 = nn.Conv2d(in_dim, 64, kernel_size=7, stride=2, padding=3,
bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.relu = nn.ReLU(inplace=True)
self.maxpool = nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
self.layer1 = self._make_layer(block, 64, layers[0])
self.layer2 = self._make_layer(block, 128, layers[1], stride=2)
self.layer3 = self._make_layer(block, 256, layers[2], stride=2)
self.layer4 = self._make_layer(block, 512, layers[3], stride=2)
self.avgpool = nn.AvgPool2d(7, stride=1, padding=2)
self.fc = nn.Linear(2048, num_classes)
for m in self.modules():
if isinstance(m, nn.Conv2d):
n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
m.weight.data.normal_(0, (2. / n)**.5)
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
def _make_layer(self, block, planes, blocks, stride=1):
downsample = None
if stride != 1 or self.inplanes != planes * block.expansion:
downsample = nn.Sequential(
nn.Conv2d(self.inplanes, planes * block.expansion,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(planes * block.expansion))
layers = []
layers.append(block(self.inplanes, planes, stride, downsample))
self.inplanes = planes * block.expansion
for i in range(1, blocks):
layers.append(block(self.inplanes, planes))
return nn.Sequential(*layers)
def forward(self, x):
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = x.view(x.size(0), -1)
logits = self.fc(x)
probas = F.softmax(logits, dim=1)
return logits, probas
def ResNet101(num_classes):
model = ResNet(block=Bottleneck,
layers=[3, 4, 23, 3],
num_classes=num_classes,
grayscale=False)
return model
# + id="n1UTmg_vo1T_"
model = ResNet101(num_classes)
model.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# + [markdown] id="rzFIKC5mSCNs"
# ### Training
# + colab={"base_uri": "https://localhost:8080/"} id="Pnl1_8H7SHK9" outputId="e30a613e-4fa1-48b5-959a-ec824b8d4f9d"
def compute_accuracy(model, data_loader):
correct_pred, num_examples = 0, 0
for i, (features, targets) in enumerate(data_loader):
features = features.to(device)
targets = targets.to(device)
logits, probas = model(features)
_, predicted_labels = torch.max(probas, 1)
num_examples += targets.size(0)
correct_pred += (predicted_labels == targets).sum()
return correct_pred.float()/num_examples * 100
for epoch in range(num_epochs):
model.train()
for batch_idx, (features, targets) in enumerate(train_loader):
features = features.to(device)
targets = targets.to(device)
# Forward and Backprop
logits, probas = model(features)
cost = F.cross_entropy(logits, targets)
optimizer.zero_grad()
cost.backward()
# update model paramets
optimizer.step()
# Logging
if not batch_idx % 50:
print ('Epoch: %03d/%03d | Batch %04d/%04d | Cost: %.4f'
%(epoch+1, num_epochs, batch_idx,
len(train_loader), cost))
model.eval()
with torch.set_grad_enabled(False):
print('Epoch: %03d/%03d | Train: %.3f%% ' %(
epoch+1, num_epochs,
compute_accuracy(model, train_loader)))
# + [markdown] id="7sQBgaPtHHL7"
# ### Evaluation
# + colab={"base_uri": "https://localhost:8080/"} id="B0MYkRbBQs5_" outputId="66388472-3aaf-49c7-c428-071d1cfbdcee"
with torch.set_grad_enabled(False):
print('Test accuracy: %.2f%%' % (compute_accuracy(model, test_loader)))
|
pytorch/12_resnet101_cifar10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# ## algorithm
def eratosthenes(n):
n = (n + 1) >> 1
p = np.ones(n, dtype=np.int8)
i, j = 1, 3
while i < n:
if p[i]:
p[j * j >> 1::j] = 0
i, j = i + 1, j + 2
return p.sum()
# ## run
for j in range(1, 8):
print(10 ** j, eratosthenes(10 ** j))
|
100days/day 05 - eratosthenes sieve.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # [Histogram](https://plotly.com/python/histograms/)
#
# ## 1. importar las librerías + csv con los datos de la encuesta.
# +
# importar librerias
import pandas as pd
import plotly.express as px
from dash import Dash, dcc, html, Input, Output
#crear un dataframe con toda la informacion de la encuesta
df_csv = pd.read_csv ('survey/survey_results_public2021.csv', index_col = [0]) # El indice sera la columna con el ID de la respuesta
df_csv #mostrar df ()
# -
# ## 2. Preprocesar datos.
#
# Tratar las columnas/conjunto de datos para comenzar a crear los gráficos. En este caso Age1stcode
df_csv['Age1stCode'].value_counts()
# Para lidiar con rangos de edades, algunos de los cuales tienen texto, se va a calcular una nueva columna con la media de todos ellos.
#
# +
#se hace una copia del df.
df= df_csv.copy()
#normalizar todos los datos.
df = df[df['Age1stCode'].notna()] #eliminar los nulos
df.loc[df["Age1stCode"] == "Younger than 5 years", "Age1stCode"] = "04 - 04 years" #ya hay un 05 anyos en el df.
df.loc[df["Age1stCode"] == "Older than 64 years", "Age1stCode"] = "65 - 65 years"
df.loc[df["Age1stCode"] == "5 - 10 years", "Age1stCode"] = "05 - 10 years"
#primero se seleccionan los digitos del string (la columna del df es string) y el resultado se convierte a entero
df['min'] = df.Age1stCode.astype(str).str[:2].astype(int) #la edad minima del rango es el primer numero
df['max'] = df.Age1stCode.astype(str).str[5:7].astype(int) # el maximo es el segundo numero
#una vez ya se tiene la edad minima y la maxima, se calcula la media de ambas columnas.
df['media'] = df[['min', 'max']].mean(axis=1)
# -
# ## 3. Grafico.
#
# En este caso, un diagrama de barras.
# +
app = Dash(__name__)
server = app.server #heroku
app.layout = html.Div([
html.H1("Tipo de desarrollador", style={'text-align': 'center'}), #cabecero h1. Header
#primera mini prueba con un menu desplegable.
dcc.Dropdown(id="select_opt",
options=[ #el usuario va a ver las label.
{"label": "#", "value": "numero"},
{"label": "%", "value": "porcentaje"}],
multi=False,
value="numero",
style={'width': "40%"}
),
dcc.Graph(id='my_survey', figure={}) # graph container
])
# -
@app.callback(
Output(component_id='my_survey', component_property='figure'),
Input(component_id='select_opt', component_property='value'))
def update_graph(option_slctd):
#filtered_df = df[df.year == selected_year]
fig = px.histogram(df, x="media",
title='Histograma de edad',
labels={'media':'media', 'count':'total'}, # can specify one label per df column
opacity=0.8,
color_discrete_sequence=['indianred'] # color of histogram bars
)
# no implementado la opcion con el porcentaje
return fig
# ## 4. run server
app.run_server(debug=True, use_reloader=False)
|
.ipynb_checkpoints/Histogram-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# loan data mining
# part 5
# Grid Search CV
import numpy as np
import pandas as pd
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import GridSearchCV
# -
# use train data
train=pd.read_csv('loantrain.csv')
test=pd.read_csv('loantest.csv')
train=train.drop('custid',axis=1)
test=test.drop('custid',axis=1)
train = train.replace(np.nan, 0)
test = test.replace(np.nan, 0)
Xtrain = train.drop('status',axis=1)
ytrain = train['status']
Xtest = test.drop('status',axis=1)
ytest = test['status']
from sklearn.ensemble import GradientBoostingClassifier
#Gradient Boosting Classifier
GBC = GradientBoostingClassifier()
gb_param_grid = {'loss' : ["deviance"],
'n_estimators' : [100,200,300],
'learning_rate': [0.1, 0.05, 0.01],
'max_depth': [4, 8],
'min_samples_leaf': [100,150],
'max_features': [0.3, 0.1]
}
modelgsGBC = GridSearchCV(GBC,param_grid = gb_param_grid, cv=5,
scoring="accuracy", n_jobs= -1, verbose = 1)
modelgsGBC.fit(Xtrain,ytrain)
# +
# Logistic Regression
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
logreg = LogisticRegression()
scores = cross_val_score(logreg, Xtrain, ytrain, cv=5)
scores
# +
# Random Forests
from sklearn.ensemble import RandomForestClassifier
from sklearn import metrics
# refer to: https://blog.csdn.net/qq_35040963/article/details/88832030
rf0 = RandomForestClassifier(oob_score=True, random_state=2019)
rf0.fit(Xtrain,ytrain.astype('int'))
print (rf0.oob_score_)
# -
y_predprob = rf0.predict_proba(Xtest)[:,1]
print( "AUC Score (Train): %f" % metrics.roc_auc_score(ytest, y_predprob))
# GridSearch for n_estimators
param_test1 = {'n_estimators':[50,120,160,200,250]}
gsearch1 = GridSearchCV(estimator = RandomForestClassifier(min_samples_split=100,
min_samples_leaf=20,max_depth=8,max_features='sqrt' ,random_state=10),
param_grid = param_test1, scoring='roc_auc',cv=5)
gsearch1.fit(Xtrain,ytrain)
print( gsearch1.best_params_, gsearch1.best_score_)
param_test2 = {'max_depth':[1,2,3,5,7,9,11,13]}#, 'min_samples_split':[100,120,150,180,200,300]}
gsearch2 = GridSearchCV(estimator = RandomForestClassifier(n_estimators=50, min_samples_split=100,
min_samples_leaf=20,max_features='sqrt' ,oob_score=True, random_state=10),
param_grid = param_test2, scoring='roc_auc',iid=False, cv=5)
gsearch2.fit(Xtrain,ytrain)
print( gsearch2.best_params_, gsearch2.best_score_)
rf1 = RandomForestClassifier(n_estimators= 160, max_depth=7, min_samples_split=100, min_samples_leaf=20,max_features='sqrt' ,oob_score=True, random_state=2019)
rf1.fit(Xtrain,ytrain)
print( rf1.oob_score_)
y_predprob = rf1.predict_proba(Xtest)[:,1]
print( "AUC Score (Train): %f" % metrics.roc_auc_score(ytest, y_predprob))
param_test3 = {'min_samples_split':[80,100,120], 'min_samples_leaf':[10,20,30,50]}
gsearch3 = GridSearchCV(estimator = RandomForestClassifier(n_estimators= 160, max_depth=7,
max_features='sqrt' ,oob_score=True, random_state=2017),
param_grid = param_test3, scoring='roc_auc',iid=False, cv=5)
gsearch3.fit(Xtrain,ytrain)
print( gsearch3.best_params_, gsearch3.best_score_)
param_test4 = {'max_features':[3,5,7,9]}
gsearch4 = GridSearchCV(estimator = RandomForestClassifier(n_estimators= 160, max_depth=7, min_samples_split=80,
min_samples_leaf=30 ,oob_score=True, random_state=2019),
param_grid = param_test4, scoring='roc_auc',iid=False, cv=5)
gsearch4.fit(Xtrain,ytrain)
print( gsearch4.best_params_, gsearch4.best_score_)
# the optimal RF
rf2 = RandomForestClassifier(n_estimators= 160, max_depth=7, min_samples_split=80,
min_samples_leaf=30,max_features=7,oob_score=True, random_state=2019)
rf2.fit(Xtrain,ytrain)
print (rf2.oob_score_)
# SVM
from sklearn.metrics import classification_report
from sklearn.svm import SVC
tuned_parameters = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
scores = ['precision', 'recall']
# +
clf = GridSearchCV(SVC(), tuned_parameters, cv=5
)
clf.fit(Xtrain, ytrain)
print("Best parameters set found on development set:")
print(clf.best_params_)
# -
y_pred=clf.predict(X_test)
print(classification_report(y_test, y_pred))
|
loan5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0-dev
# language: julia
# name: julia-0.5
# ---
# # Querying the database of Chicago employees
# ## The database
# We start with loading a sample database. Our sample database is derived from the dataset of all employees of the city of Chicago ([source](https://data.cityofchicago.org/Administration-Finance/Current-Employee-Names-Salaries-and-Position-Title/xzkq-xp2w)).
# +
include("../citydb.jl")
using RBT
setdb(citydb)
# -
# We can execute a query using `@query()` command:
@query(6*(3+4))
# ## Traversing the database structure
# *Find the names of all departments.*
@query(department.name)
# *Find the names of all employees.*
@query(department.employee.name)
# We are not longer restricted by the hierarchical structure of the database, so we can query employees directly.
@query(employee.name)
# We can traverse the schema in any direction, for instance, from employees to their departments.
@query(employee.department.name)
# *Show the list of all salaries.*
@query(employee.salary)
# If the traversal ends at an entity class, an array of records is generated.
@query(employee)
# Which fields are selected depends on the path to the class.
@query(department.employee)
# ## Summarizing data
# *Find the number of departments.*
@query(count(department))
# *Find the number of employees for each department.*
@query(department.count(employee))
# *Find the total number of employees.*
@query(count(department.employee))
# Again, we can query `employee` directly.
@query(count(employee))
# *Find the top salary among all employees.*
@query(max(employee.salary))
# *Find the maximum number of employees per department.*
@query(max(department.count(employee)))
# ## Selecting output columns
# *For each department, find the number of employees.*
@query(department:select(name,count(employee)))
# The `:select` notation is a syntax sugar for regular function call where the first argument is placed before the function name (postfix notation).
@query(select(department,name,count(employee)))
# It is easy to add new columns to the output. Let us add *the top salary per department.*
@query(
department
:select(
name,
count(employee),
max(employee.salary)))
# ## Filtering data
# *Find the employees with salary greater than $200k.*
@query(
employee
:filter(salary>200000)
:select(name,surname,position,salary))
# You can apply `:filter()` on any selected column.
@query(
employee
:select(name,surname,position,salary)
:filter(salary>200000))
# *Find the number of employees with salary in the range from \$100k to \$200k.*
@query(
employee
:filter((salary>100000)&(salary<=200000))
:count)
# *Find the departments with more than 1000 employees.*
@query(
department
:filter(count(employee)>1000)
.name)
# *Find the number of departments with more than 1000 employees.*
@query(
count(
department
:filter(count(employee)>1000)))
# *For each department, find the number of employees with salary higher than $100k.*
@query(
department
:select(
name,
count(employee:filter(salary>100000))))
# *For each department with the number of employees less than 1000, find the employees with salary higher than $125k.*
@query(
department
:filter(count(employee)<1000)
:select(
name,
employee
:filter(salary>125000)
:select(name,surname,position)))
# ## Sorting
#
# We use the `:sort` combinator to sort an array of values.
#
# *List the names of departments in alphabetical order.*
@query(department.name:sort)
# We can also specify the attribute by which the elements of the array are to be sorted.
#
# *Show the employees sorted by salary.*
@query(employee:sort(salary))
# Use `:desc` indicator to reverse the order.
@query(employee:sort(salary:desc))
# It is possible to specify several sorting keys.
@query(
employee
:sort(
salary:desc,
surname:asc,
name:asc))
# `:sort` can be used together with `:select` and `:filter`.
@query(
department
:select(name, size => count(employee))
:filter(size>1000)
:sort(size:desc))
# Use `:define` to name a commonly used expression.
@query(
department
:define(size => count(employee))
:select(name, size)
:filter(size>1000)
:sort(size:desc))
# ## Limiting
#
# Use combinators `:first`, `:last`, `:take` to limit the size of the output array. Use `:reverse` to reverse the output array.
#
# *The first employee.*
@query(first(employee))
# *The name of the first employee.*
@query(first(employee).name)
# *The department with the largest number of employees.*
@query(
department
:select(name, size => count(employee))
:sort(size:desc)
:first)
# Same query without `:sort`.
#
# *The department with the largest number of employees.*
@query(
department
:select(name, size => count(employee))
:first(size))
# *Last employee.*
@query(employee:last)
# *The department with the largest number of employees.*
@query(
department
:select(name, size => count(employee))
:sort(size)
:last)
# Same query could be written without `:sort`.
#
# *The department with the largest number of employees.*
@query(
department
:select(name, size => count(employee))
:last(size:desc))
# *Show first 5 employees.*
@query(employee:take(5))
# *Skip first 10 employees, show next 5.*
@query(employee:skip(10):take(5))
# *Show last ten employees.*
@query(employee:skip(-10))
# *Show approximately half of employees.*
@query(employee:take(count(employee)/2))
# *Reverse the order of departments.*
@query(department:reverse)
# ## Identity
#
# An identity of a database record is a value that identifies the record among all the entities of the same class. Use `id` attribute to find the identity of the input record.
@query(department:select(id,name))
# Use `:get` combinator to find the record by its identity.
@query(department:get(5))
# If a record is not found, `null` value is returned.
@query(department:get(-1))
# You can use brackets instead of `:get`.
@query(
department[5]
:select(id, name, count(employee)))
# *Show all employees of a selected department.*
@query(department[5].employee)
# ## Hierarchical queries
# (Note: the data on organizational structure is not available, so the output is largely meaningless).
#
# *Find the employees who earn more than their manager.*
@query(
employee
:filter(salary>managed_by.salary)
:select(name, surname, position, managed_by, salary-managed_by.salary))
# *For all employees in a certain department, list their seniors and the number of their subordinates.*
@query(
department[26].employee
:select(
name,
surname,
position,
connect(managed_by),
count(connect(manages))))
# *List employees of a certain department in hierarchical order.*
@query(
department[26].employee
:sort_connect(managed_by)
:select(
depth(managed_by),
name,
surname,
position))
# ## Grouping
# Use `:unique` combinator to generate all unique values that appear in a sequence.
#
# *List all distinct positions.*
@query(employee.position:unique)
# *Find the number of distinct positions for each department.*
@query(
department
:select(
name,
count(unique(employee.position)),
count(employee)))
# We can also list distinct positions using `:group` combinator. With each position, we get a list of employees having this position.
@query(employee:group(position))
# For each row generated by `employee:group(position)`, combinator `employee` will give you employees that have this position.
#
# *For each position, find the number of employees.*
@query(
employee
:group(position)
:select(position, size => count(employee))
:sort(size:desc))
# *Find positions provided by no less than 5 departments.*
@query(
employee
:group(position)
:define(department => unique(employee.department))
:filter(count(department)>=5)
:select(position, department)
:sort(count(department):desc))
# *Find the popular names of Chicago employees.*
@query(
employee
:group(name)
:select(name, size => count(employee))
:sort(size:desc))
# *Find the top salary by the first name, but only if there are at least 10 employees having this name.*
@query(
employee
:group(name)
:filter(count(employee)>=10)
:select(name, max_salary => max(employee.salary))
:sort(max_salary:desc))
# *Find the number of employees for each department and salary bracket.*
@query(
employee
:group(department, salary_bracket => salary/10000*10000 :desc)
:select(department, salary_bracket, salary_bracket+9999, count(employee)))
# To generate totals on each dimension, use `:group_cube`.
#
# *Find the number of employees for each department and salary bracket, including totals.*
@query(
employee
:group_cube(department, salary_bracket => salary/10000*10000 :desc)
:select(department, salary_bracket, salary_bracket+9999, count(employee)))
# Add `:dataframe` to present this data in tabular form.
@query(
employee
:group_cube(
department,
salary_bracket => salary/10000*10000 :desc)
:select(
department,
low => salary_bracket,
high => salary_bracket+9999,
size => count(employee))
:dataframe)
# You can specify dimensions separately using `:partition`.
#
# *Find the number of positions, the number of employees and the highest salary for the first 3 departments.*
@query(
employee
:partition(department:take(3))
:select(department.name, count(unique(employee.position)), count(employee), max(employee.salary)))
# Similar to `:group_cube`, `:partition_cube` adds totals.
#
# *Find the numbers of positions and employees, the highest salary and the most popular position for the first 3 departments, and include the totals.*
@query(
employee
:partition_cube(department:take(3))
:select(
department.name,
num_pos => count(unique(employee.position)),
num_empl => count(employee),
max_salary => max(employee.salary),
pop_position => employee:group(position):first(count(employee)).position)
:dataframe)
# You can use an array constructor or `range()` combinator to specify the dimensions.
@query(range(0, 60000, max(employee.salary)))
# *For the given departments, employee's names and salary brackets, find the number of employees, the number of different positions and the most popular position.*
@query(
employee
:define(salary_bracket => salary/60000*60000)
:partition(
department:take(3),
name => ["ANTHONY", "BRIAN"],
salary_bracket => range(0, 60000, max(employee.salary)))
:select(
dept => department.name,
name,
low => salary_bracket,
high => salary_bracket+59999,
pop_position => employee:group(position):first(count(employee)).position,
num_pos => count(unique(employee.position)),
num_empl => count(employee))
:dataframe)
# ## Output formatting
#
# The output can be produced in the form of a JSON value or a `DataFrame` object.
@query(department:json)
# Selector items become fields of the JSON dictionary.
@query(
department
:select(
name,
size => count(employee),
head => employee:first(salary))
:json)
# You can pass a list of output fields to the `json` combinator.
@query(
department
:json(
name,
size => count(employee),
head => employee:first(salary)))
# Use `:dataframe` combinator to generate `DataFrame` output.
@query(employee:dataframe)
@query(
department
:select(
name,
size => count(employee),
max_salary => max(employee.salary))
:dataframe)
# You can pass a list of output fields to the `dataframe` combinator.
@query(
department
:dataframe(
name,
size => count(employee),
max_salary => max(employee.salary)))
# ## Cartesian product and tagged union
#
# Use `:mix` to generate a Cartesian product.
#
# *Multiplication table.*
@query(
mix(a => range(2,1,10),
b => range(2,1,10),
c => range(2,1,10))
:filter((a <= b) & (b <= c))
:select(a, b, c, (a*b)*c))
# *All pairs of departments with approximately equal number of employees.*
@query(
mix(department, department)
:filter((left.id != right.id) & (left.count(employee)/10 == right.count(employee)/10))
:select(left.name, left.count(employee), right.name, right.count(employee)))
# Use `pack()` combinator to generate a tagged union.
@query(
pack(
a => range(1,1,5),
z => range(95,1,99)))
# Values generated by `pack()` don't have to have the same type.
@query(pack(employee, department))
# You can extract tagged values using combinators named after the tags.
@query(pack(employee, department).employee)
@query(pack(employee, department).department)
@query(pack(employee, department):select(employee.position, department.name))
# Use `unlink` to create an unconditional link to an entity class.
#
# *Find the employees with salary within 50% of the top salary.*
@query(
employee
:take(500)
:filter(salary > max(unlink(employee).salary)/2))
# Use `link` to create a link on an arbitrary condition.
#
# *For a given employee, find all his namesakes.*
@query(
employee[10]
:define(namesake => link((left.id!=right.id)&(left.name==right.name), employee))
:select(name, count(namesake), namesake:take(3)))
# ## Queries with parameteres
#
# *Find all employees with the given name and position.*
@query(
employee:filter((position==POSITION) & (name==NAME)),
POSITION="POLICE OFFICER",
NAME="CHARLES")
# *Find all departments bigger than the given size.*
@query(
department
:filter(count(employee)>SIZE)
:select(name, count(employee)-SIZE),
SIZE=1000)
# *Find all employees in the given departments.*
@query(
employee:filter(department.name in DEPTS),
DEPTS=["POLICE", "FIRE"])
# Parameters could also be calculated dynamically using combinator `given`.
#
# *Find the highest paid employee.*
@query(
employee
:filter(salary==MAX_SALARY)
:given(MAX_SALARY => max(employee.salary)))
# *Find the highest paid employee in each department.*
@query(
department
:select(
name,
employee
:filter(salary==MAX_SALARY)
:given(MAX_SALARY => max(employee.salary))))
# ## Navigating the context
#
# Sometimes you may want to refer to other values in the output. Combinators `before`, `after` and `around` allow you to establish a link between output values.
#
# Combinator `before` refers to the set of previous values in the output.
@query(
department
:dataframe(
name,
past_names => before.name))
# Similarly, combinator `after` refers to all the subsequent values.
@query(
department
:dataframe(
name,
next_name => first(after).name))
# You can use the `before` combinator to number output records.
@query(
department
:select(1+count(before), name))
# A variant of `before` called `and_before` includes the current record in the set. You can also use `and_before` to calculate running totals.
@query(
department
:define(size => count(employee))
:dataframe(
name,
size,
total => sum(and_before.size)))
# Combinator `and_around` let you refer to the full set of the output values.
#
# *Find the departments with the largest number of employees.*
@query(
department
:define(size => count(employee))
:filter(size == max(and_around.size))
:select(name, size))
# Combinator `around` can also give you output values that have the same property as the current value.
#
# *For each employee in a certain department, find how much does their salary differ from the top salary for their position.*
@query(
employee
:filter(department.name == DEPT)
:dataframe(
name,
surname,
position,
salary,
salary_diff => max(and_around(position).salary)-salary),
DEPT="TREASURER")
# By default, context extends to all values produced by the combinator while executing the query. You can limit the scope of the context using the `frame` combinator.
#
# *Find the highest paid employee in each department.*
@query(
department
:select(
name,
employee
:filter(salary==max(and_around.salary))
:frame))
# ## Compiling and executing queries
# You can compile and execute queries separately. To compile a query, use the `RBT.prepare()` function.
q1 = RBT.@prepare(department.employee.name)
q2 = RBT.@prepare(count(department))
q3 = RBT.@prepare(department:select(name,count(employee)))
q4 = RBT.@prepare(count(employee:filter((salary>100000)&(salary<200000))))
# You can also prepare a query with parameters.
q5 = RBT.@prepare(X*(Y*Z), X=Int, Y=Nullable{Int}, Z=Vector{Int})
q6 = RBT.@prepare(employee:filter((name == NAME) & (salary > MIN_SALARY)), NAME=UTF8String, MIN_SALARY=Int)
# Queries know their parameters.
RBT.params(q1)
RBT.params(q5)
RBT.params(q6)
# To execute a query, call the compiled query as a function.
q1()
q2()
q3()
q4()
q5(X=5, Y=Nullable(4), Z=[3,2,1])
q6(NAME="CHARLES", MIN_SALARY=100000)
|
jl/demo/citydb-with-rabbit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dlnd
# language: python
# name: dlnd
# ---
# # TensorFlow 2.0
# +
import os
from glob import glob
from datetime import datetime
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras import datasets
import matplotlib.pyplot as plt
# %load_ext tensorboard
# %matplotlib inline
# -
# ## Hyperparameter Tunning
# +
num_epochs = 10
batch_size = 32
learning_rate = 0.001
dropout_rate = 0.5
input_shape = (32, 32, 3)
num_classes = 10
# -
# ## Build Model
# +
inputs = layers.Input(input_shape)
net = layers.Conv2D(32, (3, 3), padding='SAME')(inputs)
net = layers.Activation('relu')(net)
net = layers.Conv2D(32, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.MaxPooling2D(pool_size=(2, 2))(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Conv2D(64, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.Conv2D(64, (3, 3), padding='SAME')(net)
net = layers.Activation('relu')(net)
net = layers.MaxPooling2D(pool_size=(2, 2))(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Flatten()(net)
net = layers.Dense(512)(net)
net = layers.Activation('relu')(net)
net = layers.Dropout(dropout_rate)(net)
net = layers.Dense(num_classes)(net)
net = layers.Activation('softmax')(net)
model = tf.keras.Model(inputs=inputs, outputs=net, name='Basic_CNN')
# -
# Model is the full model w/o custom layers
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate), # Optimization
loss='sparse_categorical_crossentropy', # Loss Function
metrics=['accuracy']) # Metrics / Accuracy
# # Data Preprocess
train_paths = glob('../../dataset/cifar/cifar/train/*.png')[:100]
test_paths = glob('../../dataset/cifar/cifar/test/*.png')[:100]
def get_class_name(path):
return path.split('_')[-1].replace('.png', '')
train_labels = [get_class_name(path) for path in train_paths]
class_names = np.unique(train_labels)
def get_label(path):
fname = tf.strings.split(path, '_')[-1]
lbl_name = tf.strings.regex_replace(fname, '.png', '')
onehot = tf.cast(lbl_name == class_names, tf.uint8)
return tf.argmax(onehot) # 이번에는 onehot이 아닌 label 번호로
def load_image_label(path):
gfile = tf.io.read_file(path)
image = tf.io.decode_image(gfile)
image = tf.cast(image, tf.float32) / 255. # rescale
label = get_label(path)
return image, label
def image_preprocess(image, label):
image = tf.image.random_flip_up_down(image)
image = tf.image.random_flip_left_right(image)
return image, label
AUTOTUNE = tf.data.experimental.AUTOTUNE
train_dataset = tf.data.Dataset.from_tensor_slices(train_paths)
train_dataset = train_dataset.map(load_image_label, num_parallel_calls=AUTOTUNE)
train_dataset = train_dataset.map(image_preprocess, num_parallel_calls=AUTOTUNE)
train_dataset = train_dataset.batch(batch_size)
train_dataset = train_dataset.shuffle(buffer_size=len(train_paths))
train_dataset = train_dataset.repeat()
test_dataset = tf.data.Dataset.from_tensor_slices(test_paths)
test_dataset = test_dataset.map(load_image_label, num_parallel_calls=AUTOTUNE)
test_dataset = test_dataset.batch(batch_size)
test_dataset = test_dataset.repeat()
# # Callbacks
logdir = os.path.join('logs', datetime.now().strftime("%Y%m%d-%H%M%S"))
tensorboard = tf.keras.callbacks.TensorBoard(
log_dir=logdir,
write_graph=True,
write_images=True,
histogram_freq=1
)
# https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/r2/image_summaries.ipynb#scrollTo=IJNpyVyxbVtT
# %tensorboard --logdir logs --port 8008
# ## Training
# http://localhost:6006
# +
steps_per_epoch = len(train_paths) // batch_size
validation_steps = len(test_paths) // batch_size
model.fit_generator(
train_dataset,
steps_per_epoch=steps_per_epoch,
validation_data=test_dataset,
validation_steps=validation_steps,
epochs=num_epochs,
callbacks=[tensorboard]
)
# -
# https://www.tensorflow.org/tensorboard/r2/image_summaries#setup
# ### LambdaCallback
# +
# https://www.tensorflow.org/tensorboard/r2/image_summaries
import sklearn.metrics
import itertools
import io
file_writer_cm = tf.summary.create_file_writer(logdir + '/cm')
def plot_to_image(figure):
"""Converts the matplotlib plot specified by 'figure' to a PNG image and
returns it. The supplied figure is closed and inaccessible after this call."""
# Save the plot to a PNG in memory.
buf = io.BytesIO()
plt.savefig(buf, format='png')
# Closing the figure prevents it from being displayed directly inside
# the notebook.
plt.close(figure)
buf.seek(0)
# Convert PNG buffer to TF image
image = tf.image.decode_png(buf.getvalue(), channels=4)
# Add the batch dimension
image = tf.expand_dims(image, 0)
return image
def plot_confusion_matrix(cm, class_names):
"""
Returns a matplotlib figure containing the plotted confusion matrix.
Args:
cm (array, shape = [n, n]): a confusion matrix of integer classes
class_names (array, shape = [n]): String names of the integer classes
"""
figure = plt.figure(figsize=(8, 8))
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
plt.title("Confusion matrix")
plt.colorbar()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names, rotation=45)
plt.yticks(tick_marks, class_names)
# Normalize the confusion matrix.
cm = np.around(cm.astype('float') / cm.sum(axis=1)[:, np.newaxis], decimals=2)
# Use white text if squares are dark; otherwise black.
threshold = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
color = "white" if cm[i, j] > threshold else "black"
plt.text(j, i, cm[i, j], horizontalalignment="center", color=color)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
return figure
# +
test_images, test_labels = next(iter(test_dataset)) # Confusion Matrix 그릴 때 필요한 Test Image
def log_confusion_matrix(epoch, logs):
# Use the model to predict the values from the validation dataset.
test_pred_raw = model.predict(test_images)
test_pred = np.argmax(test_pred_raw, axis=1)
# Calculate the confusion matrix.
cm = sklearn.metrics.confusion_matrix(test_labels, test_pred)
# Log the confusion matrix as an image summary.
figure = plot_confusion_matrix(cm, class_names=class_names)
cm_image = plot_to_image(figure)
# Log the confusion matrix as an image summary.
with file_writer_cm.as_default():
tf.summary.image("Confusion Matrix", cm_image, step=epoch)
# -
# Define the per-epoch callback.
cm_callback = tf.keras.callbacks.LambdaCallback(on_epoch_end=log_confusion_matrix)
# +
steps_per_epoch = len(train_paths) // batch_size
validation_steps = len(test_paths) // batch_size
model.fit_generator(
train_dataset,
steps_per_epoch=steps_per_epoch,
validation_data=test_dataset,
validation_steps=validation_steps,
epochs=num_epochs,
callbacks=[tensorboard, cm_callback]
)
# -
# ## Expert
# +
loss_object = tf.keras.losses.SparseCategoricalCrossentropy()
optimizer = tf.keras.optimizers.Adam()
# +
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='train_accuracy')
test_loss = tf.keras.metrics.Mean(name='test_loss')
test_accuracy = tf.keras.metrics.SparseCategoricalAccuracy(name='test_accuracy')
# -
@tf.function
def train_step(images, labels):
with tf.GradientTape() as tape:
predictions = model(images)
loss = loss_object(labels, predictions) # Loss 계산
gradients = tape.gradient(loss, model.trainable_variables) # 모델의 trainable_variable을 하여금 loss를 통해 기울기를 얻음
optimizer.apply_gradients(zip(gradients, model.trainable_variables)) # 구한 최적화된 값을 variable에 적용
train_loss(loss)
train_accuracy(labels, predictions)
@tf.function
def test_step(images, labels):
predictions = model(images)
t_loss = loss_object(labels, predictions)
test_loss(t_loss)
test_accuracy(labels, predictions)
logdir = os.path.join('logs', datetime.now().strftime("%Y%m%d-%H%M%S"))
file_writer = tf.summary.create_file_writer(logdir)
for epoch in range(num_epochs):
for step, (images, labels) in enumerate(train_dataset):
train_step(images, labels)
with file_writer.as_default():
tf.summary.image('input_image', images, step=step+(step*epoch))
tf.summary.scalar('train_loss', train_loss.result(), step=step+(step*epoch))
tf.summary.scalar('train_accuracy', train_accuracy.result(), step=step+(step*epoch))
for test_images, test_labels in test_dataset:
test_step(test_images, test_labels)
template = 'Epoch {}, Loss: {}, Accuracy: {}, Test Loss: {}, Test Accuracy: {}'
print (template.format(epoch+1,
train_loss.result(),
train_accuracy.result()*100,
test_loss.result(),
test_accuracy.result()*100))
|
DL_TF20/Part 16 - callbacks - tensorboard-Antonio.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Implementing a Learning Rate Finder from Scratch
# > Choosing the right learning rate is important when training Deep Learning models. Let's implement a Learning Rate Finder from scratch, taking inspiration from Leslie Smith's LR range test--a nifty technique to find optimal learning rates.
#
# - toc: false
# - badges: true
# - comments: true
# - categories: [machine-learning]
# - image: images/lrf-from-scratch.png
# - keywords: machine learning, ml, learning rate, learning rate finder, lr finder, lr_finder, lr_find, deep neural network, neural network, convolutional neural network, beginner concepts, fastai, fast.ai, pytorch
# + id="SXzEGY2xrly7" colab={"base_uri": "https://localhost:8080/"} outputId="9aa2ba2a-e274-46ab-adbc-5a45e4f0357c"
#hide
# !pip install -Uq fastbook
from fastbook import *
# + [markdown] id="g9zzkgKSnGXB"
# ## Introduction
#
# In this post we will implement a **learning rate finder** from scratch. A learning rate finder helps us find sensible learning rates for our models to train with, including minimum and maximum values to use in a **cyclical learning rate** policy. Both concepts were invented by <NAME> and I suggest you check out his [paper](https://arxiv.org/pdf/1506.01186.pdf){% fn 1 %}!
#
# We'll implement the learning rate finder (and cyclical learning rates in a future post) into our Deep Neural Network created from scratch in the 2-part series [Implementing a Deep Neural Network from Scratch]({% post_url 2020-12-28-Implementing-a-Deep-Neural-Network-from-Scratch-Part-1 %}). Check that out first if you haven't read it already!
# + [markdown] id="v12m5gkk4iAQ"
# ## Understanding the Learning Rate
#
# Before we start, what _is_ the learning rate? The learning rate is just a value we multiply our gradients by in Stochastic Gradient Descent before updating our parameters with those values. Think of it like a "weight" that reduces the impact of each step change so as to ensure we are not over-shooting our loss function. If you imagine our loss function like a parabola, and our parameters starting somehwere along the parabola, descending along by a specific amount will bring us further "down" in the parabola, to it's minimum point eventually. If the step amount is too big however, we risk overshooting that minimum point. That's where the learning rate comes into play: it helps us achieve very small steps if desired.
#
# To refresh your memory, here is what happens in the optimization step of Stochastic Gradient Descent ([Part 1]({% post_url 2020-12-28-Implementing-a-Deep-Neural-Network-from-Scratch-Part-1 %}) of our DNN series explains this formula in more detail, so read that first):
#
# $ w := w - \eta \nabla Q({w}) $
#
# Our parameter $w$ is updated by subtracting its gradient calculated with respect to the loss function $\nabla Q({w})$ after multiplying it by a weighing factor $\eta$. That weighing factor $\eta$ is our learning rate!
#
# It's even easier in code:
#
# ```python
# # part of the .step() method in our SGD_Optimizer
# for p in self.parameters: p.data -= p.grad.data * self.lr
# ```
#
# This is outlined in the `.step` method of our optimizer (check the setup code in the next section).
# As we saw towards the end of [Part 2]({% post_url 2021-01-01-Implementing-a-Deep-Neural-Network-from-Scratch-Part-2 %}) of our Implementing a Deep Neural Network from Scratch series, the learning rate has a _big_ impact on training for our model: the lower the learning rate, the more epochs required to reach a given accuracy, the higher the learning rate, the higher risk of overshooting (or never reaching) the minimum loss. There's a lot more factors at play that we cover in that series, but for now let's just consider the learning rate.
#
# Another aspect of learning rates is that a single value is rarely optimal for the duration of the training. We could say that its efficacy **degrades over time** (time measured in batches/epochs). That's why common techniques include decreasing the learning rate by a step-wise fixed amount or by an exponentially decreasing amount during the course of the training. The logic being that as the model is further into training and is approaching the minimum loss, it needs less pronounced updates (steps), and therefore would benefit from smaller increments.
#
#
# ## The Learning Rate Finder
#
# The **learning rate finder**, or more appropriately **learning rate _range_ finder**, is a method outlined in a [paper](https://arxiv.org/pdf/1506.01186.pdf) by <NAME> written in 2015[^1]. The paper introduces the concept of **cyclical learning rates** (i.e. repeatedly cycling between learning rates inbetween a set minimum and a maximum has shown to be effective for training--a method we will implement from scratch in a future post!) whereby the minimum and maximum values to cycle through are found by a function that Leslie Smith defines as the "_LR range test_". Here's what the author himself has to say about it:
#
# > There is a simple way to estimate reasonable minimum
# and maximum boundary values with one training run of the
# network for a few epochs. It is a “LR range test”; run your
# model for several epochs while letting the learning rate increase linearly between low and high LR values. This test
# is enormously valuable whenever you are facing a new architecture or dataset.{% fn 1 %}
#
#
# So where does the learning rate finder come into play? Well, it helps us find how learning rates affect our training loss, helping us spot a "sweet spot" of ranges that maximize the loss. That extremes of that range will also be minimum and maximum values to use in the cyclical learning rates policy <NAME> outlines in his paper -- before we can implement cyclical learning rates, we need to implement a learning rate range finder!
#
# To give you an idea of what we're trying to create, let's see [fast.ai](https://fastai1.fast.ai/callbacks.lr_finder.html)'s implementation, probably one of the first frameworks to implement this LR range test. fast.ai offers a convenient helper function called the Learning Rate Finder (`Learner.lr_finder()`) that helps us see the effect a variety of learning rates have on our model's training performance as well as suggest a `min_grad_lr` where the gradient of the training loss is steepest:
#
# 
#
# Let's implement something similar!
# + [markdown] id="-ObvXMp1DmJi"
# ## Getting Started
#
# Below is a recap of all the preparatory steps to setup our data pipeline. In order, we'll download the data, generate a list of file paths, create training and validation tensor stacks from the file paths, convert those tensors from rank-2 (2D matrix) tensors (i.e. size: 28, 28) to rank-1 (1D vector) tensors (i.e. size: 784). We'll then generate labels corresponding to the digit index and merge the input tensors and labels into datasets to create DataLoader. This will provide us with minibatches of sample data to run our SGD along.
# + id="yvUq2zzr0yrm" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="9251f17e-3255-4436-e20b-d994f3b1c70e"
#collapse
# Requirements
# # !pip install -Uq fastbook # includes all the common imports (plt, fastai, pytorch, etc)
# Download the data
path = untar_data(URLs.MNIST)
# Import the paths of our training and testing images
training = { f'{num}' : (path/f'training/{num}').ls().sorted() for num in range(10) }
testing = { f'{num}' : (path/f'testing/{num}').ls().sorted() for num in range(10) }
# Prepare training tensor stacks
training_tensors = [
torch.stack([
tensor(Image.open(digit)).float()/255 for digit in training[f'{num}']
]) for num in range(10)
]
validation_tensors = [
torch.stack([
tensor(Image.open(digit)).float()/255 for digit in testing[f'{num}']
]) for num in range(10)
]
# Convert our 2D image tensors (28, 28) into 1D vectors
train_x = torch.cat(training_tensors).view(-1, 28*28)
valid_x = torch.cat(validation_tensors).view(-1, 28*28)
# Generate our labels based on the digit each image represents
train_y = torch.from_numpy(np.concatenate([[i]*len(training[f'{i}']) for i in range(10)]))
valid_y = torch.from_numpy(np.concatenate([[i]*len(testing[f'{i}']) for i in range(10)]))
# Create datasets to feed into the dataloaders
dset = list(zip(train_x, train_y))
dset_valid = list(zip(valid_x, valid_y))
# Setup our dataloders
dl = DataLoader(dset, batch_size=256, shuffle=True)
valid_dl = DataLoader(dset_valid, batch_size=256, shuffle=True)
# + [markdown] id="1cByk3hO4CT1"
# We'll also import the code that we created in our series on [Implementing a Deep Neural Network from Scratch](https://muttoni.github.io/blog/machine-learning/2020/12/28/Implementing-a-Deep-Neural-Network-from-Scratch-Part-1.html) in Python. The code we'll need is our general purpose `LinearModel`, our SGD optimizer `SGD_Optimizer` and our beloved `DeepClassifier`. Feel free to toggle the code below to see how each one works.
# + id="FMYHyLoCHRzz"
#collapse
# General purpose Linear Model
class LinearModel:
def __init__(self, inputs, outputs,):
self.input_size = inputs
self.output_size = outputs
self.weights, self.bias = self._init_params()
def parameters(self):
return self.weights, self.bias
def model(self, x):
return x@self.weights + self.bias
def _init_params(self):
weights = (torch.randn(self.input_size, self.output_size)).requires_grad_()
bias = (torch.randn(self.output_size)).requires_grad_()
return weights, bias
# + id="LwoRgv3Ly5dH"
#collapse
# General purpose SGD Optimizer
class SGD_Optimizer:
def __init__(self, parameters, lr):
self.parameters = list(parameters)
self.lr = lr
def step(self):
for p in self.parameters: p.data -= p.grad.data * self.lr
for p in self.parameters: p.grad = None
# + id="lOjVl-QG91xy"
#collapse
# General Purpose Classifier
class DeepClassifier:
"""
A multi-layer Neural Network using ReLU activations and SGD
params: layers to use (LinearModels)
methods: fit(train_dl, valid_dl, epochs, lr) and predict(image_tensor)
"""
def __init__(self, *layers):
self.accuracy_scores = []
self.training_losses = []
self.layers = layers
def fit(self, **kwargs):
self.train_dl = kwargs.get('train_dl')
self.valid_dl = kwargs.get('valid_dl')
self.epochs = kwargs.get('epochs', 5)
self.lr = kwargs.get('lr', 0.1)
self.verbose = kwargs.get('verbose', True)
self.optimizers = []
self.epoch_losses = []
self.last_layer_index = len(self.layers)-1
for layer in self.layers:
self.optimizers.append(SGD_Optimizer(layer.parameters(),self.lr))
for i in range(self.epochs):
for xb, yb in self.train_dl:
preds = self._forward(xb)
self._backward(preds, yb)
self._validate_epoch(i)
def predict(self, image_tensor):
probabilities = self._forward(image_tensor).softmax(dim=1)
_, prediction = probabilities.max(-1)
return prediction, probabilities
def _forward(self, xb):
res = xb
for layer_idx, layer in enumerate(self.layers):
if layer_idx != self.last_layer_index:
res = layer.model(res)
res = self._ReLU(res)
else:
res = layer.model(res)
return res
def _backward(self, preds, yb):
loss = self._loss_function(preds, yb)
self.epoch_losses.append(loss)
loss.backward()
for opt in self.optimizers: opt.step()
def _batch_accuracy(self, xb, yb):
predictions = xb.softmax(dim=1)
_, max_indices = xb.max(-1)
corrects = max_indices == yb
return corrects.float().mean()
def _validate_epoch(self, i):
accs = [self._batch_accuracy(self._forward(xb), yb) for xb, yb in self.valid_dl]
score = round(torch.stack(accs).mean().item(), 4)
self.accuracy_scores.append(score)
epoch_loss = round(torch.stack(self.epoch_losses).mean().item(), 4)
self.epoch_losses = []
self.training_losses.append(epoch_loss)
self._print(f'Epoch #{i}', 'Loss:', epoch_loss, 'Accuracy:', score)
def _loss_function(self, predictions, targets):
log_sm_preds = torch.log_softmax(predictions, dim=1)
idx = range(len(predictions))
results = -log_sm_preds[idx, targets]
return results.mean()
def _ReLU(self, x):
return x.max(tensor(0.0))
def _print(self, *args):
if self.verbose:
print(*args)
# + [markdown] id="AOJMZTysGwAZ"
# **Note**: If you're looking for a walkthrough of the code above, make sure you read the 2 part series on implementing a deep neural network from scratch! I would especially focus on the second part. Here are the links: [Part 1]({% post_url 2020-12-28-Implementing-a-Deep-Neural-Network-from-Scratch-Part-1 %}), [Part 2]({% post_url 2021-01-01-Implementing-a-Deep-Neural-Network-from-Scratch-Part-2 %}).
# + [markdown] id="4B4ottpekwH3"
# ## Requirements
#
# Now that our code and model is setup, we need to recap what we need to add to our DeepClassifier in order to properly support a `lr_finder` method.
#
# **Our goal**: a method called `lr_finder` that when called performs a round of training (aka fitting) for a predetermined number of epochs, starting with a very small learning rate, increasing it exponentially every minibatch (why exponentially? So that we have an equal representation of small learning rate values vs large ones -- more on this later). We can sort out specifics later.
#
#
# >Warning: The `lr_finder` can't start with random parameters, it should start with the current parameter state of the model, _without affecting it_ during training! So we'll need to **clone** our parameters.
#
#
# We'll also need to update our code to keep track of training loss at every minibatch (across epochs), and store the respective learning rates used for each minibatch.
#
#
# + [markdown] id="F2djWMPLYEeX"
# Now, before we continue, I want to make a big **disclaimer**: the additions we'll make today are going to be "patches" added on to a codebase that is, to put it lightly, fragile. The code above was created for demonstration purposes only, and essentially contains the bare essentials to make a deep neural network classifier work properly. The sole use of this code, for me, is to tinker with it and try out ideas and concepts as I come across them.
#
# While I'm tempted to re-write everything from scratch and build it into a more modular/generalized framework, it would introduce abstractions that for the purposes of our learning process would make "getting" the concepts more difficult, as effective abstractions inevitably hide away the lower level workings of a piece of code.
#
# So I apologize for the entropy that we are about to introduce into this already "entropic" code. We might look into making it more robust in a future post. In the meantime, you are more than welcome to take this and refactor it to your liking!
#
# With the disclaimer out of the way, let's recap how we normally used this DeepClassifier. We would instantiate the class and pass in the desired layers. Like so:
# + id="vEf-fLkbuJOi"
my_nn = DeepClassifier( # example usage
LinearModel(28*28, 20), # a layer with 28*28 inputs (pixels), 20 activations
LinearModel(20, 10) # a layer with 20 inputs and 10 activations/outputs
)
# + [markdown] id="b02o2OtWamky"
# Each layer contains the input and output parameter counts. Every layer's autput is ReLU'd automatically, except for the last one that uses Cross-Entropy Loss based on log and softmax. Read [Part 2]({% post_url 2021-01-01-Implementing-a-Deep-Neural-Network-from-Scratch-Part-2 %}) of the DNN from scratch series to see exactly how this works.
#
# We then call the fit method and pass in the data, the learning rate and the epochs to train for:
# + id="DRoFx8wPb5-L"
my_nn.fit(
train_dl=train_dl,
valid_dl=valid_dl,
lr=0.1,
epochs=100
)
# -
# >Note: Note the limitations here: the learning rate is fixed--the very purpose of a learning rate finder is that it helps us map the effect of a _range_ of learning rates. Furthermore, the data is linked with the `fit` method, meaning that if we don't call `.fit`, we won't have any data inside our classifier.
#
#
# So we must change our code to allow for data to be fed in at the instantiation stage of the DeepClassifier class. We'll also need to move some of the properties that we used to create in our `fit` method, at the instantiation step. First however, let's tweak our dependencies LinearModel and SGD_Optimizer.
# + [markdown] id="vBr7VEYaWYil"
# ### SGD_Optimizer
#
# Given our learning rate needs to change over the course of each batch, we'll first need to be able to pass in a custom learning rate to our SGD optimizer's `step` method (where the updates are multiplied by the learning rate, as seen above). In our original code, the learning rate was set on instantiation and never touched again. We'll change it so the `step` method accepts a keyworded argument called `lr`, otherwise it defaults to its parameter `self.lr`.
# + id="AVI1Hm1NWXFO"
# General purpose SGD Optimizer
class SGD_Optimizer:
def __init__(self, parameters, lr):
self.parameters = list(parameters)
self.lr = lr
def step(self, **kwargs): # add **kawrgs
lr = kwargs.get('lr', self.lr) # get 'lr' if set, otherwise set to self.lr
for p in self.parameters: p.data -= p.grad.data * lr
for p in self.parameters: p.grad = None
# + [markdown] id="8Av5I_TbgLaK"
# ### LinearModel
#
# + [markdown] id="Gm8WqrCYhYh4"
# In our `LinearModel` we need to introduce some changes. We need two new methods that our `lr_finder` can use to copy and set parameters so as not to overwrite the actual model parameters. We'll introduce two new methods: `copy_parameters` (note the `parameters` method already acts as a `get`), and a `set_parameters`. These are quite self-explanatory and the only tricky thing is to make sure to clone and detach any copies from the gradient calculations, as well as reset gradient tracking when setting the parameters. See the code below:
# + id="Mr9_ctdakTX3"
#collapse_show
# General purpose Linear Model
class LinearModel:
def __init__(self, inputs, outputs,):
self.input_size = inputs
self.output_size = outputs
self.weights, self.bias = self._init_params()
def parameters(self):
return self.weights, self.bias
def copy_parameters(self):
return self.weights.clone().detach(), self.bias.clone().detach()
def set_parameters(self, parameters):
self.weights, self.bias = parameters
self.weights.requires_grad_()
self.bias.requires_grad_()
def model(self, x):
return x@self.weights + self.bias
def _init_params(self):
weights = (torch.randn(self.input_size, self.output_size)).requires_grad_()
bias = (torch.randn(self.output_size)).requires_grad_()
return weights, bias
# + [markdown] id="v11rUllyjf00"
# ## DeepClassifier
# + [markdown] id="2yGPj68TkRap"
# ### Prep Work
# + [markdown] id="km49Aae1i2jA"
# Now we can proceed with updating DeepClassifier. We'll start by by changing our ``__init__`` method as follows:
# + id="ZEjxnx-Oeb-n"
def __init__(self, *layers, **kwargs): # added **kwargs
self.layers = layers
self.accuracy_scores = []
self.training_losses = []
# Moved all of the following from the 'fit' method
self.optimizers = []
self.epoch_losses = []
self.train_dl = kwargs.get('train_dl', None)
self.valid_dl = kwargs.get('valid_dl', None)
self.lr = kwargs.get('lr', 1e-2)
self.last_layer_index = len(self.layers)-1
# We'll use this Boolean to determine whether to save
# the individual batch losses in a list, instead of
# averaging them at every epoch. We need to do this
# as we'll need to plot them against each learning rate.
self.save_batch_losses = False
# if save_batch_losses is True, we save them here
self.batch_losses = []
# + [markdown] id="Zrv0LsGpfaE4"
# We essentially moved a bunch of stuff from the `fit` method to the `__init__` method so as to be able to access them from our upcoming lr_finder method. You can read the code above to see what changed.
# + [markdown] id="e6W16RtsjqDk"
# The backward method will need to accept an optional `lr` and pass it on to the step method of our optimizers. This allows us to feed in a dynamic learning rate. at each step function, which is exactly what we need.
#
# We also have a `self.save_batch_losses` that flags whether our individual batch losses (that usually get aggregated and averaged per epoch) should be persisted at batch granularity in `self.batch_losses`. This is necessary with the lr_finder because we need to plot individual batch losses with individual learning rates that changed every batch.
# + id="2bxkme0xjo1r"
def _backward(self, preds, yb, **kwargs):
lr = kwargs.get('lr', self.lr)
loss = self._loss_function(preds, yb)
if self.save_batch_losses:
self.batch_losses.append(loss.item())
else:
self.epoch_losses.append(loss)
loss.backward()
for opt in self.optimizers: opt.step(lr=lr)
# + [markdown] id="PlZ3CNBukJrV"
# ### Implementing lr_finder()
#
# Believe it or not, those are all the changes we need to make lr_finder work! Now we are to implement the new method. Here it is in all its glory, and we'll go line by line.
# + id="3lJPXroJkeMs"
#collapse_show
def lr_finder(self):
base_lr = 1e-6
max_lr = 1e+1
epochs = 3
current_lr = base_lr
self.old_params = [layer.copy_parameters() for layer in self.layers]
self.batch_losses = []
self.lr_finder_lrs = []
self.save_batch_losses = True
batch_size = self.train_dl.bs
samples = len(self.train_dl.dataset)
iters = epochs * round(samples / batch_size)
step_size = abs(math.log(max_lr/base_lr) / iters)
print(batch_size, samples, iters, step_size)
for layer in self.layers:
self.optimizers.append(SGD_Optimizer(layer.parameters(), base_lr))
while current_lr <= max_lr:
for xb, yb in self.train_dl:
if current_lr > max_lr:
break
preds = self._forward(xb)
self._backward(preds, yb, lr=current_lr)
self.lr_finder_lrs.append(current_lr)
current_lr = math.e**(math.log(current_lr)+step_size*2)
# clean up
self.save_batch_losses = False
self.optimizers = []
for i in range(len(self.old_params)):
self.layers[i].set_parameters(self.old_params[i])
# + [markdown] id="Hau60uEJtr5i"
# Let's look at the code in detail:
# + [markdown] id="f6gsKKn5kkDL"
# ```python
# base_lr = 1e-7
# max_lr = 1e+1
# epochs = 3
# current_lr = base_lr
# ```
#
# Our brand new lr_finder method accepts a `base_lr` learning rate to start the range of experimentation. It will stop when it reaches `max_lr`. `epochs` is an arbitrary number that we set to determine how many epochs to run the finder for. We also set a `current_lr` to the minimum lr we want to test and this variable will be incremented at every step. The first three parameters are critical as they determine how many steps will be carried out, as the steps are determined by batch size, total sample size and epochs, like so:
#
# ```python
# batch_size = self.train_dl.bs
# samples = len(self.train_dl.dataset)
# iters = epochs * round(samples / batch_size)
# step_size = abs(math.log(max_lr/base_lr) / iters)
# ```
#
# In my particular implementation, the step size `step_size` is determined by the range of learning rates to try out, divided by the iterations the model will go through (number of epochs multiplied by the number of batches in our sample).
#
# Now you may wonder why we are dividing the maximum learning rate by the minimum learning rate, instead of subtracting. The reason is: logs! Since we are dealing with logs, rather than do: $\frac{max\_lr - min\_lr}{iters}$, we do: $\frac{log(\frac{max\_lr}{min\_lr})}{iters}$. Rather than divide the learning rates linearly, which, when plotted on log scale, has the effect of condensing the majority of the learning rates towards the higher end, I opted to increment our step size exponentially. To do this we need to take the log so that we deal with exponents and can later increment the next learning rate by $e^{current\_lr+step\_size}$. This will ensure that when we plot our learning rates on a log scale, our iterations will be evenly spaced out.
#
# ```python
# self.old_params = [layer.copy_parameters() for layer in self.layers]
# self.lr_finder_lrs = []
# self.save_batch_losses = True
# ```
#
# Next we save the old parameters so we can reset our model later. We create a list to store our learning rates and set the flag `save_batch_losses` to True, so that our _backward function knows to save them in our `batch_losses` list. The two lists `lr_finder_lrs` and `batch_losses` are the two lists that we will plot!
#
# ```python
#
# for layer in self.layers:
# self.optimizers.append(SGD_Optimizer(layer.parameters(), base_lr))
#
# while current_lr <= max_lr:
# for xb, yb in self.train_dl:
# if current_lr > max_lr:
# break
# preds = self._forward(xb)
# self._backward(preds, yb, lr=current_lr)
# self.lr_finder_lrs.append(current_lr)
# current_lr = math.e**(math.log(current_lr)+step_size)
#
# ```
#
# Next we proceed as if we were a `fit` method, by initiating optimizers for each layer and going through our training cycle. As you can see, the loop is a little different, where we don't worry about epochs, we just check that the `current_lr` is less than or equal to the `max_lr`. We then go through each batch, apply `_forward`, then apply `_backward` and then make sure to save our current learning rate and update the learning rate for the next batch. Since our step_size is exponent-based (i.e. we logged it at the beginning, giving us the exponent of $e$ that will give us that value), we need to increment the exponent of $e$ by that step size. We do this by taking the log of our current learning rate and incrementing that by the step_size which is already logarithmic.
#
# The formula is as follows:
# $ current\_lr = e^{log(current\_lr) + step\_size}$
#
# Once our training rounds are complete, the last thing left to do is cleanup!
#
# ```python
# self.save_batch_losses = False
# self.optimizers = []
# for i in range(len(self.old_params)):
# self.layers[i].set_parameters(self.old_params[i])
# ```
#
# We reset our flag `save_batch_losses`, reset our optimizers and copy back the saved parameters into the model. Again, the reason we want to save the parameters is so that we can apply the lr_finder at any stage of our fitting cycles without it affecting our parameter state.
#
# Here is the updated DeepClassifier code:
# + id="4JGND4BGn2xV"
#collapse_show
class DeepClassifier:
"""
A multi-layer Neural Network using ReLU activations and SGD
params: layers to use (LinearModels)
methods:
- fit(train_dl, valid_dl, epochs, lr)
- predict(image_tensor)
- lr_finder()
"""
def __init__(self, *layers, **kwargs):
self.layers = layers
self.accuracy_scores = []
self.training_losses = []
self.optimizers = []
self.epoch_losses = []
self.batch_losses = []
self.train_dl = kwargs.get('train_dl', None)
self.valid_dl = kwargs.get('valid_dl', None)
self.last_layer_index = len(self.layers)-1
self.save_batch_losses = False
self.lr = 0.1
def fit(self, **kwargs):
self.train_dl = kwargs.get('train_dl', self.train_dl)
self.valid_dl = kwargs.get('valid_dl', self.valid_dl)
self.epochs = kwargs.get('epochs', 5)
self.lr = kwargs.get('lr', 0.1)
self.verbose = kwargs.get('verbose', True)
self.optimizers = []
for layer in self.layers:
self.optimizers.append(SGD_Optimizer(layer.parameters(),self.lr))
for i in range(self.epochs):
for xb, yb in self.train_dl:
preds = self._forward(xb)
self._backward(preds, yb)
self._validate_epoch(i)
def predict(self, image_tensor):
probabilities = self._forward(image_tensor).softmax(dim=1)
_, prediction = probabilities.max(-1)
return prediction, probabilities
def lr_finder(self):
base_lr = 1e-6
max_lr = 1e+1
epochs = 3
current_lr = base_lr
self.old_params = [layer.copy_parameters() for layer in self.layers]
self.lr_finder_lrs = []
self.save_batch_losses = True
batch_size = self.train_dl.bs
samples = len(self.train_dl.dataset)
iters = epochs * round(samples / batch_size)
step_size = abs(math.log(max_lr/base_lr) / iters)
for layer in self.layers:
self.optimizers.append(SGD_Optimizer(layer.parameters(), base_lr))
while current_lr <= max_lr:
for xb, yb in self.train_dl:
if current_lr > max_lr:
break
preds = self._forward(xb)
self._backward(preds, yb, lr=current_lr)
self.lr_finder_lrs.append(current_lr)
current_lr = math.e**(math.log(current_lr)+step_size)
# clean up
self.save_batch_losses = False
self.optimizers = []
for i in range(len(self.old_params)):
self.layers[i].set_parameters(self.old_params[i])
def _forward(self, xb):
res = xb
for layer_idx, layer in enumerate(self.layers):
if layer_idx != self.last_layer_index:
res = layer.model(res)
res = self._ReLU(res)
else:
res = layer.model(res)
return res
def _backward(self, preds, yb, **kwargs):
lr = kwargs.get('lr', self.lr)
loss = self._loss_function(preds, yb)
if self.save_batch_losses:
self.batch_losses.append(loss.item())
else:
self.epoch_losses.append(loss)
loss.backward()
for opt in self.optimizers: opt.step(lr=lr)
def _batch_accuracy(self, xb, yb):
predictions = xb.softmax(dim=1)
_, max_indices = xb.max(-1)
corrects = max_indices == yb
return corrects.float().mean()
def _validate_epoch(self, i):
accs = [self._batch_accuracy(self._forward(xb), yb) for xb, yb in self.valid_dl]
score = round(torch.stack(accs).mean().item(), 4)
self.accuracy_scores.append(score)
epoch_loss = round(torch.stack(self.epoch_losses).mean().item(), 4)
self.epoch_losses = []
self.training_losses.append(epoch_loss)
self._print(f'Epoch #{i}', 'Loss:', epoch_loss, 'Accuracy:', score)
def _loss_function(self, predictions, targets):
log_sm_preds = torch.log_softmax(predictions, dim=1)
idx = range(len(predictions))
results = -log_sm_preds[idx, targets]
return results.mean()
def _ReLU(self, x):
return x.max(tensor(0.0))
def _print(self, *args):
if self.verbose:
print(*args)
# + [markdown] id="85wnm7aasG4V"
# ## Demo
# + [markdown] id="Lx4O7rIyrS5H"
# Now let's try it out! We instantiate a new `DeepClassifier` by specifying the layers and passing in the data so we can run our `lr_finder` before running `fit` (where in our old code we would normally feed in our dataset).
# + id="zL2Ue1-39QLO"
my_nn = DeepClassifier(
LinearModel(28*28, 20),
LinearModel(20, 10),
train_dl=dl,
valid_dl=valid_dl
)
my_nn.lr_finder()
# + [markdown] id="f6RDNF1-re06"
# Now let's create a quick function to plot the results (this can easily be turned into a method as well, but beyond the scope of this post:
# + id="LR8EKiKvM-Ig"
def plot_lr_loss(my_nn):
import matplotlib.pyplot as plt
x = my_nn.lr_finder_lrs
y = my_nn.batch_losses
fig, ax = plt.subplots()
ax.plot(x, y, 'g-')
ax.set_xlabel('Learning Rates')
ax.set_ylabel('Training Loss', color='g')
plt.title(f"Results with lr={my_nn.lr}")
plt.xscale('log')
plt.show()
# -
plot_lr_loss(my_nn)
# + [markdown] colab={"base_uri": "https://localhost:8080/", "height": 299} id="ZZbCuwS_NS7h" outputId="2d56572d-2abc-4874-b001-37b38ee41c96"
# 
# + [markdown] id="8bSxSjC-sCyZ"
# ## Analyzing the lr_finder plot
#
# As rules of thumb, fastai recommends picking the LR where the slope is steepest, or pick the end of the slop minimum and divide it by 10. In our case it seems our point of steepest slope is between $10^{-3}$ and $10^{-2}$. We definitely don't want to pick anything beyond 5e-1 as it seems the loss is flattening before picking up again. We also probably don't want to pick anything before $10^{-4}$ as the loss stays flat meaning it would take a lot of epochs before we see any significant improvement. If I were to pick a learning rate I would probably go with $10^{-2}$ for the first couple cycles, and then move to a finer one (e.g. $10^{-3}$).
#
# ## Conclusion
#
# Learning rates are a critical part of training a neural network using SGD and a good learning rate will not only help you get closest to the minimum loss, but will also speed up your training (i.e. less epochs to reach a specific accuracy).
#
# Experiment with lr_finder yourself! Suggestions: try adding weights to our step_size increments (e.g. *2) and try incrementing it linearly instead of logarithmically and see how it affects the plot.
#
# If you have suggestions for what to implement next, comment below! Hope you found this useful and all the best in your machine learning journey.
#
# **References**
#
# {{ '<NAME>. (v6 2017, v1 2015). Cyclical Learning Rates for Training Neural Networks [Link](https://arxiv.org/abs/1506.01186)' | fndetail: 1 }}
#
|
_notebooks/2021-01-08-Implementing-a-Learning-Rate-Finder-from-Scratch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from nltk.stem.snowball import SnowballStemmer
from gensim import models, corpora
def load_data(input_file):
data = []
with open(input_file, 'r') as f:
for line in f.readlines():
data.append(line.strip())
return data
def process(input_text):
tokenizer = RegexpTokenizer(r'\w+')
tokens = tokenizer.tokenize(input_text.lower())
stop_words = stopwords.words('english')
tokens = [x for x in tokens if not x in stop_words]
stemmer = SnowballStemmer('english')
tokens_stemmed = [stemmer.stem(x) for x in tokens]
return tokens_stemmed
data = load_data('data.txt')
tokens = [process(x) for x in data]
dict_tokens = corpora.Dictionary(tokens)
doc_term_mat = [dict_tokens.doc2bow(token) for token in tokens]
num_topics = 2
ldamodel = models.ldamodel.LdaModel(doc_term_mat,
num_topics=num_topics, id2word=dict_tokens, passes=25)
num_words = 5
print('Top ' + str(num_words) + ' contributing words to each topic:')
for n,values in ldamodel.show_topics(num_topics=num_topics, num_words=num_words,
formatted=False):
print('\nTopic', n)
for word,weight in values:
print(word, '==>', str(round(float(weight) * 100, 2)) + '%')
|
artificial-intelligence-with-python-ja-master/Chapter 10/topic_modeler.ipynb
|