code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 1: Exploring languages through word frequencies
# ## Learning Objectives:
# In this lab you will learn the following linguistic concepts and programming skills:
# * Basic text processing and regular expressions.
# * What do word frequencies tell us about a language?
# * How do different languages compare?
# * How to manipulate corpora and plot insightful graphs?
# ## About this assignment
#
# - This is a Jupyter notebook. You can execute cell blocks by pressing control-enter.
# - You will be submitting the lab1.py file on Gradescope.
# - We have provided local access to the Gradescope autograder test cases. In order to run the test cases locally, simply run <code>python run_tests.py</code> or <code>python run_tests.py -j</code> (this commands gives information about the performance on each test case in the form of a readable json object).
# ## Pre-requisites:
# For this lab, you need to make sure you have the following installed:
# * python3.6 (python2.7 should also work)
# * nltk (python package)
# * matplotlib
#
# To make sure your installation is successful, execute the block below.
import nltk
import matplotlib
import importlib
import lab1
# As a packaged solution, I would recommend installing [conda](https://docs.conda.io/en/latest/miniconda.html), and creating a conda [environment](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html) 'ece365_nlp' to use for the rest of this module.
# ## Exercise 1: Working With Text
# Before getting started, let's work with some simple text processing!
# +
text1 = "Ethics are built right into the ideals and objectives of the United Nations "
len(text1) # The length of text1 (String Length)
# -
text2 = text1.split(' ') # Return a list of the words in text2, separating by ' '.
len(text2) # Word Length
text2
# <br>
# List comprehension allows us to find specific words:
[w for w in text2 if len(w) > 3] # Words that are greater than 3 letters long in text2
[w for w in text2 if w.istitle()] # Capitalized words in text2
[w for w in text2 if w.endswith('s')] # Words in text2 that end in 's'
# <br>
# We can find unique words using 'set()'.
# +
text3 = 'To be or not to be'
text4 = text3.split(' ')
len(text4)
# -
len(set(text4))
set(text4)
len(set([w.lower() for w in text4])) # .lower converts the string to lowercase.
set([w.lower() for w in text4])
# ### Processing free-text
# +
text5 = '"Ethics are built right into the ideals and objectives of the United Nations" \
#UNSG @ NY Society for Ethical Culture bit.ly/2guVelr'
text6 = text5.split(' ')
text6
# -
# <br>
# Finding hastags:
[w for w in text6 if w.startswith('#')]
# <br>
# Finding callouts:
[w for w in text6 if w.startswith('@')]
text7 = '@UN @UN_Women "Ethics are built right into the ideals and objectives of the United Nations" \
#UNSG @ NY Society for Ethical Culture bit.ly/2guVelr'
text8 = text7.split(' ')
[w for w in text8 if w.startswith('@')]
# ### Regular Expressions
# We can use regular expressions to help us with more complex parsing. A regular expression is a special sequence of characters that helps you match or find other strings or sets of strings, using a specialized syntax held in a pattern. Regular expressions are widely used in UNIX world.
#
# For example `'@[A-Za-z0-9_]+'` will return all words that:
# * start with `'@'` and are followed by at least one:
# * capital letter (`'A-Z'`)
# * lowercase letter (`'a-z'`)
# * number (`'0-9'`)
# * or underscore (`'_'`)
# +
import re # import re - a module that provides support for regular expressions
[w for w in text8 if re.search('@[A-Za-z0-9_]+', w)]
# -
# Let's get more familiar with Regular Expression on pandas (pandas is a powerful open source data analysis Python tool.)!
# +
import pandas as pd
time_sentences = ["Monday: The doctor's appointment is at 2:45pm.",
"Tuesday: The dentist's appointment is at 11:30 am.",
"Wednesday: At 7:00pm, there is a basketball game!",
"Thursday: Be back home by 11:15 pm at the latest.",
"Friday: Take the train at 08:10 am, arrive at 09:00am."]
df = pd.DataFrame(time_sentences, columns=['text'])
df
# -
# find the number of characters for each string in df['text']
df['text'].str.len()
# find the number of tokens for each string in df['text']
df['text'].str.split().str.len()
# find which entries contain the word 'appointment'
df['text'].str.contains('appointment')
# find how many times a digit occurs in each string
df['text'].str.count(r'\d')
# find all occurances of the digits
df['text'].str.findall(r'\d')
# group and find the hours and minutes
df['text'].str.findall(r'(\d?\d):(\d\d)')
# replace weekdays with '???'
df['text'].str.replace(r'\w+day\b', '???')
# replace weekdays with 3 letter abbrevations
df['text'].str.replace(r'(\w+day\b)', lambda x: x.groups()[0][:3])
# create new columns from first match of extracted groups
df['text'].str.extract(r'(\d?\d):(\d\d)')
# extract the entire time, the hours, the minutes, and the period
df['text'].str.extractall(r'((\d?\d):(\d\d) ?([ap]m))')
# extract the entire time, the hours, the minutes, and the period with group names
df['text'].str.extractall(r'(?P<time>(?P<hour>\d?\d):(?P<minute>\d\d) ?(?P<period>[ap]m))')
# You may realize there is nothing for you to answer. This section is ungraded, but you are strongly recommended to read&understand this section before moving on.
# ## Exercise 2: Word Frequencies
# We live in a multi-lingual world. The languages we use are like English in some ways and distinct from English in many ways. In this exercise, we will explore some aspects of languages that make them different from English by the use of quantitative indices.
#
# Before we begin comparing languages, let us begin with English. How many words are there in English? Well, that depends on who we ask. The Second Edition of the 20-volume Oxford English Dictionary contains full entries for 171,476 words in current use (and 47,156 obsolete words). Looking elsewhere, Webster's Third New International Dictionary, Unabridged, together with its 1993 Addenda Section, includes some 470,000 entries. But, the number of words in the Oxford and Webster Dictionaries are not the same as the number of words in English. Why is that? First, it takes a while for dictionary publishers like Oxford University and Merriam-Webster to include new words in their dictionaries. While it may seem surprising new words are being coined at a rapid rate, a recent article in The Guardian reports that English speakers are adding new words at the rate of around 1,000 a year. Recent dictionary debutants include blog, grok, crowdfunding, hackathon, airball, e-marketing, sudoku, twerk and Brexit, many of which are words we find in use in our everyday lives. Slang and jargon could also be considered in this list. You have probably observed how some of these terms depend on where you live (e.g., 'prepone' in India means the opposite of postpone), whereas others are common in many places (e.g. the portmanteau, brunch = breakfast + lunch).
#
# A natural question that arises in this setting is, are all words equally likely, or do they occur with different frequencies? As you can expect, words occur with different frequencies, but what you would would not have expected is how skewed the word frequencies can be. That is what you will see in your first exercise.
#
# First, you will count the frequency of words from a word list derived from a large collection of words -- a 'corpus' (meaning 'a body of text'). For this part of the exercise, you will use the corpus of Reuters from which you will count the number of times each word occurs.
# For this, you will need to do some tokenization. Towards that, you will lowercase all words, remove the punctuation marks and numbers. Then you will use NLTK to get the frequency distribution of the tokenized text.
#
# Based on the frequency distribution of word that you will collect, you will answer the following questions.
#
# * What are the 10 most frequent words?
# * What are the 10 least frequent words?
# * What proportion of words have a frequency of 1? These singleton words are termed 'hapax legomena' (a sophisticated Greek name) and the numebr of singletons in a corpus is a measure of the richness of the vocabulary of that collection, giving you the rate at which new words appear in that text. If you take a very large text in a language and call it representative of that language, then the rate of singletons is a measure of its richness.
# * What are the answers to the above questions, if we consider stemming or lemmatization?
#
# **Total points: 50 points**
# ### Coding Questions
#
# a. In the lab1.py file, complete the function "get_freqs" that takes as an input the "Reuters" corpus (type str) from nltk and returns as an output a dictionary with the key being a word, and the value being the frequency of the word in the corpus.
# Make sure to lowercase all words in the corpus and to replace all punctuations and digits with a space character. This will take care of tokenization for you. To avoid confusion, the list of punctuation marks are given to you. (15 points)
# + nbgrader={"grade": false, "grade_id": "cell-053fc5a44cfd341b", "locked": false, "schema_version": 3, "solution": true, "task": false}
puncts = ['.','!','?',',',';',':','[', ']', '{', '}', '(', ')', '\'', '\"']
# +
importlib.reload(lab1)
raw_corpus = nltk.corpus.reuters.raw()
freqs = lab1.get_freqs(raw_corpus, puncts)
# -
freqs
# b. Next, complete the function called "get_top_10" that takes in the "freqs" dictionary, and returns the top 10 most frequent words as a list. (5 points)
# + nbgrader={"grade": false, "grade_id": "cell-4aec5a37bbd4cef5", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
print(lab1.get_top_10(freqs))
# + nbgrader={"grade": true, "grade_id": "cell-76c42b84f8038afc", "locked": true, "points": 20, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.get_top_10(freqs) == ['the', 'of', 'to', 'in', 'and', 'said', 'a', 'mln', 's', 'vs']
### END HIDDEN TESTS
# -
# c. Next, complete the function called "get_bottom_10" that takes in the "freqs" dictionary, and returns the top 10 least frequent words as a list. (5 points)
# + nbgrader={"grade": false, "grade_id": "cell-f7d5084d8b86ea60", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
print(lab1.get_bottom_10(freqs))
# + nbgrader={"grade": true, "grade_id": "cell-eedc6f45889eadd8", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.get_bottom_10(freqs) == ['inflict', 'sheen', 'stand-off', 'avowed', 'kilolitres', 'kilowatt/hour', 'janunary/march', 'pineapples', 'hasrul', 'paian']
### END HIDDEN TESTS
# -
# d. Complete the function called "get_percentage_singletons" which takes in the "freqs" dictionary and returns a float value of the percentage of words that appear once in the corpus. (5 points)
# + tags=["outputPrepend"]
print(freqs)
# + nbgrader={"grade": false, "grade_id": "cell-a165768ae088ea2c", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
print(lab1.get_percentage_singletons(freqs))
# + nbgrader={"grade": true, "grade_id": "cell-9e57695b3b6a675d", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert (lab1.get_percentage_singletons(freqs)>40.4)
assert (lab1.get_percentage_singletons(freqs)<40.6)
### END HIDDEN TESTS
# -
# e. The next two blocks show examples of how stemming and lemmatization are done.
from nltk.stem import PorterStemmer
porter = PorterStemmer()
print(porter.stem("cats"))
print(porter.stem("trouble"))
print(porter.stem("troubling"))
print(porter.stem("troubled"))
from nltk.stem import WordNetLemmatizer
wordnet_lemmatizer = WordNetLemmatizer()
sentence = "He was running and eating at same time. He has bad habit of swimming after playing long hours in the Sun."
for word in sentence.split():
print ("{0:20}{1:20}".format(word,wordnet_lemmatizer.lemmatize(word, pos="v")))
# f. Repeat steps b,c,d by doing stemming. You should modify the get_freqs_stemming function. (5 points)
# + nbgrader={"grade": false, "grade_id": "cell-9e9343b8ca4c2880", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
freqs_stemming = lab1.get_freqs_stemming(raw_corpus, puncts)
print(lab1.get_top_10(freqs_stemming))
print(lab1.get_bottom_10(freqs_stemming))
print(lab1.get_percentage_singletons(freqs_stemming))
# + nbgrader={"grade": true, "grade_id": "cell-d58f04c1eef6cf43", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.get_top_10(freqs_stemming) == ['the', 'of', 'to', 'in', 'and', 'said', 'a', 'mln', 'it', 's']
assert lab1.get_bottom_10(freqs_stemming) == ['inflict', 'sheen', 'stand-off', 'avow', 'kilolitr', 'kilowatt/hour', 'janunary/march', 'hasrul', 'paian', 'sawn']
assert (lab1.get_percentage_singletons(freqs_stemming)>41.9)
assert (lab1.get_percentage_singletons(freqs_stemming)<42.2)
### END HIDDEN TESTS
# -
# g. Repeat steps b,c,d by doing lemmatization. You should modify the get_freqs_lemmatized function. (5 points)
# + nbgrader={"grade": false, "grade_id": "cell-f364b1fc291d7e74", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
freqs_lemmatized = lab1.get_freqs_lemmatized(raw_corpus, puncts)
print(lab1.get_top_10(freqs_lemmatized))
print(lab1.get_bottom_10(freqs_lemmatized))
print(lab1.get_percentage_singletons(freqs_lemmatized))
# + nbgrader={"grade": true, "grade_id": "cell-cad80b5f0643a89f", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.get_top_10(freqs_lemmatized) == ['the', 'of', 'to', 'in', 'be', 'say', 'and', 'a', 'mln', 's']
assert lab1.get_bottom_10(freqs_lemmatized) == ['inflict', 'sheen', 'stand-off', 'avow', 'kilolitres', 'kilowatt/hour', 'janunary/march', 'pineapples', 'hasrul', 'paian']
assert (lab1.get_percentage_singletons(freqs_lemmatized)>41.9)
assert (lab1.get_percentage_singletons(freqs_lemmatized)<42.2)
### END HIDDEN TESTS
# -
# h. What is the vocabulary size of this corpus (i.e., raw_corpus)? How about the vocabulary size after doing stemming and lemmatization respectively? Note that we lowercase all words in the corpus and replace all punctuations and digits with empty spaces. Add your code to the size_of_raw_corpus, size_of_stemmed_raw_corpus and size_of_lemmatized_raw_corpus functions (5 points)
# + nbgrader={"grade": false, "grade_id": "cell-60cc741a45f09b60", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
print(lab1.size_of_raw_corpus(freqs)) # Vocalbulary size of raw_corpus.
print(lab1.size_of_stemmed_raw_corpus(freqs_stemming)) # Vocalbulary size of raw_corpus after stemming.
print(lab1.size_of_lemmatized_raw_corpus(freqs_lemmatized)) # Vocalbulary size of raw_corpus after lemmatization.
# + nbgrader={"grade": true, "grade_id": "cell-c3277367acf9daa0", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.size_of_raw_corpus(freqs) == 33206
assert lab1.size_of_stemmed_raw_corpus(freqs_stemming) == 25778
assert lab1.size_of_lemmatized_raw_corpus(freqs_lemmatized) == 29032
### END HIDDEN TESTS
# -
# i. Different documents, even of equal length are usually composed of different vocabularies. We will compare two documents of equal length, and see the percentage of unseen vocabulary between them.
#
# More specifically, we have document "a" to be the first 100 words of raw_corpus and document "b" to be the last 100 words of raw_corpus. How many percent of words in document "a" does NOT appear in document "b"? What if we change the document size to be 1000 (first 1000 words of raw_corpus v.s. last 1000 words of raw_corpus), 10000, 100000, 500000?
# What do you observe with the document size increasing? You may find set(a)-set(b) is a useful function. Modify the percentage_of_unseen_vocab function (5 points)
# + nbgrader={"grade": true, "grade_id": "cell-3e819a96835cdc2b", "locked": false, "points": 5, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
length = [100,1000,10000,100000,500000]
for length_i in length:
a = raw_corpus.split()[:length_i]
b = raw_corpus.split()[-length_i:]
print(lab1.percentage_of_unseen_vocab(a, b, length_i))
### Write down your observation here: (Ungraded)
# + nbgrader={"grade": true, "grade_id": "cell-3e391821c0e4a4d1", "locked": true, "points": 5, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.percentage_of_unseen_vocab(raw_corpus.split()[:100], raw_corpus.split()[-100:], 100) == 0.79
assert lab1.percentage_of_unseen_vocab(raw_corpus.split()[:1000], raw_corpus.split()[-1000:], 1000) == 0.464
assert lab1.percentage_of_unseen_vocab(raw_corpus.split()[:10000], raw_corpus.split()[-10000:], 10000) == 0.2182
assert lab1.percentage_of_unseen_vocab(raw_corpus.split()[:100000], raw_corpus.split()[-100000:], 100000) == 0.10077
assert lab1.percentage_of_unseen_vocab(raw_corpus.split()[:500000], raw_corpus.split()[-500000:], 500000) == 0.052344
### END HIDDEN TESTS
# -
# ## Exercise 3: Pareto principle
# The popular Pareto principle (also known as the 80/20 rule), states that for many events, roughly 80% of the effects come from 20% of the causes. This includes observations that found that the distribution of global income is very uneven, with the richest 20% of the world's population controlling 82.7% of the world's income. This seems to be the case with words as well.
#
# In this exercise, we observe something similar to the Pareto principle in words. By calculating what fraction of the most frequent words accounts for 80% of the total words in the corpus, you will see that a very small number of frequent words account for a large number of words.
#
# **Total points: 15 points**
#
# a. Complete the function called "frac_80_perc" which takes in "freqs" as an input, and returns a float representing the fraction of words that account for 80% of the tokens in the corpus (the expected answer is around 3% for Reuters corpus -- a News corpus). Note: you should be considering the words in decreasing order of frequency until reaching 80% of word (frequency) count. (15 points)
# + nbgrader={"grade": false, "grade_id": "cell-7d532348cdd5379a", "locked": false, "schema_version": 3, "solution": true, "task": false}
importlib.reload(lab1)
print(lab1.frac_80_perc(freqs))
# + nbgrader={"grade": true, "grade_id": "cell-d39b1cdba3cd5662", "locked": true, "points": 15, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert lab1.frac_80_perc(freqs) > 0.033
assert lab1.frac_80_perc(freqs) < 0.034
### END HIDDEN TESTS
# -
# This relation between the frequency and rank for words is called Zipf's law. It states that given a large sample of words used, the frequency of any word is inversely proportional to its rank in the frequency table. So word number n has a frequency proportional to 1/n. In order to see this, sort the words in a decreasing order of their frequencies and do a rank-frequency plot, with the words (indicated by their ranks) indicated along the x-axis and their frequencies in the y-axis.
#
# b. Accordingly, we will plot the frequency of words when ranked in decreasing order. Complete the function "plot_zipf" that takes in "freqs" as an input, and generates a plot using matplotlib. In this plot, the x-axis represents the rank of words in decreasing order of frequency, and the y-axis represents the frequency of the corresponding word. (Ungraded)
# + nbgrader={"grade": true, "grade_id": "cell-42ff4546787c495e", "locked": false, "points": 5, "schema_version": 3, "solution": true, "task": false}
import matplotlib.pyplot as plt
# -
importlib.reload(lab1)
lab1.plot_zipf(freqs)
# ## Exercise 4: Type-to-Token Ratio (TTR)
# Another way of measuring the richenss of vocabulary is by looking at the type-token distribution of words in a language. Word types are unique words in a corpus, whereas the tokens are the words in a corpus with repetition. And so, a sentence such as "I am taking this class because I love taking on challenges" has 11 tokens, but 9 types since the words "I" and "taking" are repeated twice. Accordingly, Type-to-Token Ratio (TTR) is the ratio of types to tokens, and the higher it is, the less words are repeated, and the richer is the language.
#
# **Total points: 15 points**
#
# a. In this exercise we will be exploring, for every language, the amount of "types" explored as we explore larger portions of the corpus, or tokens. We will be considering the Universal Declaration of Human Rights in 4 languages. Particularly, we will be plotting the amount of types explored per language as we explore 100 more tokens. For this exercise, complete the following function "get_TTRs" which takes in as an input a predefined set of languages, and returns as an output the dictionary TTR, which has a language as the key, and the value as a list showing the count of types as we explore 100 tokens, 200 tokens, 300 tokens, up until 1300 tokens of the respective corpus. Accordingly, each list in the dictionary should be made of 13 data points. Do not forget to lowercase, but you do not need to perform tokenization as the corpora now are actually a list of words instead of one string. (15 points)
# + nbgrader={"grade": false, "grade_id": "cell-f89f433ea4675fed", "locked": false, "schema_version": 3, "solution": true, "task": false}
from nltk.corpus import udhr
languages = ['Italian-Latin1', 'English-Latin1', 'German_Deutsch-Latin1', 'Finnish_Suomi-Latin1']
# -
importlib.reload(lab1)
TTRs = lab1.get_TTRs(languages)
print(TTRs)
# + nbgrader={"grade": true, "grade_id": "cell-65fae85d3536e61a", "locked": true, "points": 15, "schema_version": 3, "solution": false, "task": false}
### BEGIN HIDDEN TESTS
assert TTRs['Italian-Latin1'] == [64, 110, 143, 179, 221, 260, 286, 326, 355, 386, 412, 426, 451]
assert TTRs['English-Latin1'] == [57, 99, 133, 167, 207, 231, 262, 292, 318, 339, 358, 381, 403]
assert TTRs['German_Deutsch-Latin1'] == [63, 113, 155, 204, 254, 284, 324, 358, 388, 418, 446, 475, 504]
assert TTRs['Finnish_Suomi-Latin1'] == [74, 137, 192, 252, 303, 356, 406, 459, 491, 537, 586, 631, 675]
### END HIDDEN TESTS
# -
# b. Next, plot a line graph (one line for every language, four lines in total) that shows the count of types discovered on the y-axis and the amount of tokens in the corpus discovered on the x-axis, in increments of 100 tokens, up to 1300. (Ungraded)
# + nbgrader={"grade": true, "grade_id": "cell-8260219dae07a1cd", "locked": false, "points": 5, "schema_version": 3, "solution": true, "task": false}
import matplotlib.pyplot as plt
importlib.reload(lab1)
lab1.plot_TTRs(TTRs)
# -
# c. Which language has the highest TTR? What could be driving the TTR? Share your thoughts in the textbox below: (Ungraded)
# + [markdown] nbgrader={"grade": true, "grade_id": "cell-97883af5aa5d83f4", "locked": false, "points": 5, "schema_version": 3, "solution": true, "task": false}
# **Share your thoughts here:**
# -
|
ECE365/nlp/ece365sp21_nlp_lab1dist/Lab1_NB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importing modules
import networkx as nx
import matplotlib.pyplot as plt
G = nx.DiGraph()
G.add_edges_from([('A', 'D'), ('B', 'C'), ('B', 'E'), ('C', 'A'),
('D', 'C'), ('E', 'D'), ('E', 'B'), ('E', 'F'),
('E', 'C'), ('F', 'C'), ('F', 'H'), ('G', 'A'),
('G', 'C'), ('H', 'A')])
plt.figure(figsize =(10, 10))
nx.draw_networkx(G, with_labels = True)
hubs, authorities = nx.hits(G, max_iter = 50, normalized = True)
# The in-built hits function returns two dictionaries keyed by nodes
# containing hub scores and authority scores respectively.
print("Hub Scores: ", hubs)
print("Authority Scores: ", authorities)
# -
|
Semester 6/DWM/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (dataSc)
# language: python
# name: datasc
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Imports" data-toc-modified-id="Imports-1"><span class="toc-item-num">1 </span>Imports</a></span></li><li><span><a href="#Load-data" data-toc-modified-id="Load-data-2"><span class="toc-item-num">2 </span>Load data</a></span></li><li><span><a href="#Dark-custom-mpl-styles" data-toc-modified-id="Dark-custom-mpl-styles-3"><span class="toc-item-num">3 </span>Dark custom mpl styles</a></span></li><li><span><a href="#Light-custom-mpl-styles" data-toc-modified-id="Light-custom-mpl-styles-4"><span class="toc-item-num">4 </span>Light custom mpl styles</a></span></li><li><span><a href="#Default-mpl-styles" data-toc-modified-id="Default-mpl-styles-5"><span class="toc-item-num">5 </span>Default mpl styles</a></span></li></ul></div>
# -
# # Imports
# +
import numpy as np
import pandas as pd
import seaborn as sns
import bhishan
from bhishan import bp
import matplotlib.pyplot as plt
# %load_ext autoreload
# %load_ext watermark
# %autoreload 2
# %watermark -a "<NAME>" -d -v -m
# %watermark -iv
# -
# # Load data
df = sns.load_dataset('titanic')
df.head()
df.plot.scatter(x='age',y=['fare'])
# # Dark custom mpl styles
plt.style.use(bp.get_mpl_style(-1))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(-2))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(-3))
df.plot.scatter(x='age',y='fare')
# # Light custom mpl styles
plt.style.use(bp.get_mpl_style(-100))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(-200))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(-300))
df.plot.scatter(x='age',y='fare')
# # Default mpl styles
plt.style.use(bp.get_mpl_style(0))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(1))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(2))
df.plot.scatter(x='age',y='fare')
plt.style.use(bp.get_mpl_style(3))
df.plot.scatter(x='age',y='fare')
s = bp.get_mpl_style(4)
plt.style.use(s)
df.plot.scatter(x='age',y='fare')
|
examples/example_mpl_style.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (deeplearning)
# language: python
# name: deeplearning
# ---
# +
from keras import backend as K
n_cores = 10
K.set_session(K.tf.Session(config=K.tf.ConfigProto(
intra_op_parallelism_threads=n_cores,
inter_op_parallelism_threads=n_cores)))
from keras.layers import Input, Dense, Lambda
from keras.callbacks import TensorBoard
from keras.models import Model
from keras import regularizers
from keras.datasets import mnist
from IPython.display import SVG
from keras.utils.vis_utils import plot_model
from keras.losses import binary_crossentropy, kullback_leibler_divergence
import numpy as np
import time
import matplotlib.pyplot as plt
tb_session_name = "SAE"
tb_logs = "/home/nanni/tensorboard_logs"
# -
def get_tensorboard_callback():
return TensorBoard(log_dir="{}/{}__{}".format(tb_logs, tb_session_name,time.strftime('%Y_%m_%d__%H_%M')))
# ## MNIST
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test = x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
print(x_train.shape)
print(x_test.shape)
# ## Sparse AutoEncoder
# +
# this is the size of our encoded representations
encoding_dim = 32 # 32 floats -> compression of factor 24.5, assuming the input is 784 floats
# this is our input placeholder
input_img = Input(shape=(784,))
# "encoded" is the encoded representation of the input with a L1 activity regularizer
encoded = Dense(encoding_dim, activation='relu',
activity_regularizer=regularizers.l2(10e-7))(input_img)
# "decoded" is the lossy reconstruction of the input
decoded = Dense(784, activation='sigmoid')(encoded)
# this model maps an input to its reconstruction
autoencoder = Model(input_img, decoded)
# this model maps an input to its encoded representation
encoder = Model(input_img, encoded)
# create a placeholder for an encoded (32-dimensional) input
encoded_input = Input(shape=(encoding_dim,))
# retrieve the last layer of the autoencoder model
decoder_layer = autoencoder.layers[-1]
# create the decoder model
decoder = Model(encoded_input, decoder_layer(encoded_input))
# logging and compilation
autoencoder.compile(optimizer='adadelta', loss='binary_crossentropy')
# -
# ### Fitting
autoencoder.fit(x_train, x_train, verbose=0,
epochs=100,
batch_size=256,
shuffle=True,
validation_data=(x_test, x_test),
callbacks=[get_tensorboard_callback()])
# ### Encoding and Decoding
# encode and decode some digits
# note that we take them from the *test* set
encoded_imgs = encoder.predict(x_test)
decoded_imgs = decoder.predict(encoded_imgs)
n = 10 # how many digits we will display
plt.figure(figsize=(20, 4))
for i in range(n):
# display original
ax = plt.subplot(2, n, i + 1)
plt.imshow(x_test[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
# display reconstruction
ax = plt.subplot(2, n, i + 1 + n)
plt.imshow(decoded_imgs[i].reshape(28, 28))
plt.gray()
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# ## Variational AutoEncoder
# +
def sampling(args):
"""Reparameterization trick by sampling from an isotropic unit Gaussian.
# Arguments:
args (tensor): mean and log of variance of Q(z|X)
# Returns:
z (tensor): sampled latent vector
"""
z_mean, z_log_var = args
batch = K.shape(z_mean)[0]
dim = K.int_shape(z_mean)[1]
# by default, random_normal has mean=0 and std=1.0
epsilon = K.random_normal(shape=(batch, dim))
return z_mean + K.exp(0.5 * z_log_var) * epsilon
image_size = x_train.shape[1]
# network parameters
intermediate_dim = 512
batch_size = 128
latent_dim = 2
epochs = 50
# +
inputs = Input(shape=(image_size, ), name='encoder_input')
x = Dense(intermediate_dim, activation='relu')(inputs)
z_mean = Dense(latent_dim, name='z_mean')(x)
z_log_var = Dense(latent_dim, name='z_log_var')(x)
# use reparameterization trick to push the sampling out as input
# note that "output_shape" isn't necessary with the TensorFlow backend
z = Lambda(sampling, output_shape=(latent_dim,), name='z')([z_mean, z_log_var])
# instantiate encoder model
encoder = Model(inputs, [z_mean, z_log_var, z], name='encoder')
# build decoder model
latent_inputs = Input(shape=(latent_dim,), name='z_sampling')
x = Dense(intermediate_dim, activation='relu')(latent_inputs)
outputs = Dense(image_size, activation='sigmoid')(x)
# instantiate decoder model
decoder = Model(latent_inputs, outputs, name='decoder')
# instantiate VAE model
outputs = decoder(encoder(inputs)[2])
vae = Model(inputs, outputs, name='vae_mlp')
vae.summary()
# -
#
# References:
# - [<NAME>. & <NAME>. Auto-Encoding Variational Bayes](https://arxiv.org/pdf/1312.6114.pdf).
# - [<NAME>. Tutorial on Variational Autoencoders. (2016)](https://arxiv.org/pdf/1606.05908.pdf)
# $$
# D_{KL}[N(\mathbf z; \mu, \sigma^2) || N(0, 1)] = \frac{1}{2} \, \sum_j \left( \sigma^2_j + \mu^2_j - 1 - \log \sigma^2_j \right)
# $$
#
# In practice, however, it’s better to model $\Sigma(X)$ as $\log\Sigma(X)$ as it is more numerically stable to take exponent compared to computing log. Hence, our final KL divergence term is:
# $$
# D_{KL}[N(\mathbf z;\mu, \sigma^2) || N(0, 1)] = \frac{1}{2} \, \sum_j \left( \exp(\sigma^2_j) + \mu^2_j - 1 - \sigma^2_j \right)
# $$
def vae_loss(x, x_decoded):
xent_loss = binary_crossentropy(x, x_decoded)
kl_loss = 0.5 * K.mean(K.exp(z_log_var) + K.square(z_mean) - 1 - z_log_var, axis=-1)
return xent_loss + kl_loss
vae.compile(optimizer='rmsprop', loss=vae_loss)
vae.fit(x_train, x_train,
shuffle=True,
epochs=epochs,
batch_size=batch_size,
validation_data=(x_test, x_test),
callbacks=[get_tensorboard_callback()])
z_mean_test_encoded, z_log_var_test_encoded, z_test_encoded = encoder.predict(x_test, batch_size=batch_size)
plt.figure(figsize=(6, 6))
plt.scatter(z_mean_test_encoded[:, 0], z_mean_test_encoded[:, 1], c=y_test)
plt.colorbar()
plt.show()
|
notebooks/staging/AutoEncoders_in_Keras_Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab_type="code" id="RXZT2UsyIVe_" colab={"base_uri": "https://localhost:8080/", "height": 411} outputId="d8202c79-9856-45b7-f98e-846186d55fd7"
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/horse-or-human.zip \
# -O /tmp/horse-or-human.zip
# !wget --no-check-certificate \
# https://storage.googleapis.com/laurencemoroney-blog.appspot.com/validation-horse-or-human.zip \
# -O /tmp/validation-horse-or-human.zip
import os
import zipfile
local_zip = '/tmp/horse-or-human.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp/horse-or-human')
local_zip = '/tmp/validation-horse-or-human.zip'
zip_ref = zipfile.ZipFile(local_zip, 'r')
zip_ref.extractall('/tmp/validation-horse-or-human')
zip_ref.close()
# Directory with our training horse pictures
train_horse_dir = os.path.join('/tmp/horse-or-human/horses')
# Directory with our training human pictures
train_human_dir = os.path.join('/tmp/horse-or-human/humans')
# Directory with our training horse pictures
validation_horse_dir = os.path.join('/tmp/validation-horse-or-human/horses')
# Directory with our training human pictures
validation_human_dir = os.path.join('/tmp/validation-horse-or-human/humans')
# + [markdown] colab_type="text" id="5oqBkNBJmtUv"
# ## Building a Small Model from Scratch
#
# But before we continue, let's start defining the model:
#
# Step 1 will be to import tensorflow.
# + id="qvfZg3LQbD-5" colab_type="code" colab={}
import tensorflow as tf
# + [markdown] colab_type="text" id="BnhYCP4tdqjC"
# We then add convolutional layers as in the previous example, and flatten the final result to feed into the densely connected layers.
# + [markdown] id="gokG5HKpdtzm" colab_type="text"
# Finally we add the densely connected layers.
#
# Note that because we are facing a two-class classification problem, i.e. a *binary classification problem*, we will end our network with a [*sigmoid* activation](https://wikipedia.org/wiki/Sigmoid_function), so that the output of our network will be a single scalar between 0 and 1, encoding the probability that the current image is class 1 (as opposed to class 0).
# + id="PixZ2s5QbYQ3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="401b7e7e-941c-4d77-f84f-042498f07420"
model = tf.keras.models.Sequential([
# Note the input shape is the desired size of the image 300x300 with 3 bytes color
# This is the first convolution
tf.keras.layers.Conv2D(16, (3,3), activation='relu', input_shape=(300, 300, 3)),
tf.keras.layers.MaxPooling2D(2, 2),
# The second convolution
tf.keras.layers.Conv2D(32, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The third convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The fourth convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# The fifth convolution
tf.keras.layers.Conv2D(64, (3,3), activation='relu'),
tf.keras.layers.MaxPooling2D(2,2),
# Flatten the results to feed into a DNN
tf.keras.layers.Flatten(),
# 512 neuron hidden layer
tf.keras.layers.Dense(512, activation='relu'),
# Only 1 output neuron. It will contain a value from 0-1 where 0 for 1 class ('horses') and 1 for the other ('humans')
tf.keras.layers.Dense(1, activation='sigmoid')
])
# + colab_type="code" id="8DHWhFP_uhq3" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="10e49dad-be2d-42ef-e7ae-e71788396ded"
from tensorflow.keras.optimizers import RMSprop
model.compile(loss='binary_crossentropy',
optimizer=RMSprop(lr=1e-4),
metrics=['acc'])
# + colab_type="code" id="ClebU9NJg99G" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="8557f72d-0f00-4779-c8b0-4ee4aeadf9e2"
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# All images will be rescaled by 1./255
train_datagen = ImageDataGenerator(
rescale=1./255,
rotation_range=40,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
validation_datagen = ImageDataGenerator(rescale=1/255)
# Flow training images in batches of 128 using train_datagen generator
train_generator = train_datagen.flow_from_directory(
'/tmp/horse-or-human/', # This is the source directory for training images
target_size=(300, 300), # All images will be resized to 150x150
batch_size=128,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
# Flow training images in batches of 128 using train_datagen generator
validation_generator = validation_datagen.flow_from_directory(
'/tmp/validation-horse-or-human/', # This is the source directory for training images
target_size=(300, 300), # All images will be resized to 150x150
batch_size=32,
# Since we use binary_crossentropy loss, we need binary labels
class_mode='binary')
# + colab_type="code" id="Fb1_lgobv81m" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="552ec10c-79ea-43c4-feea-251ea5136b40"
history = model.fit_generator(
train_generator,
steps_per_epoch=8,
epochs=100,
verbose=1,
validation_data = validation_generator,
validation_steps=8)
# + id="7zNPRWOVJdOH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="5045526c-fe9f-4732-9609-a90e8d8be619"
import matplotlib.pyplot as plt
acc = history.history['acc']
val_acc = history.history['val_acc']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.figure()
plt.plot(epochs, loss, 'r', label='Training Loss')
plt.plot(epochs, val_loss, 'b', label='Validation Loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# + [markdown] id="MjOwWyejCdKe" colab_type="text"
# The problem is Data augmentation introduces a random element to the training images but if **validation set doesn't have the same randomness**, **the type of images too close to the images in the training set**.
|
Convolutional Neural Networks in TensorFlow/week2 Augmentation to Avoid Overfitting/Horse_or_Human_WithAugmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
skuinfo=[]
skuinfo.append('%s' % '\t'.join(["cid3","cid3n","sku_id","sku_name", "word22", "aaa"]))
print(str(skuinfo)[1:-1].split("\\t")[4:-1])
arrstr = str(skuinfo)[1:-1].split("\\t")
arrstr[4:-1].split(" ")
|
other/Untitled12.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# # Arrays
ar = np.array([1, 2])
ar
ar2 = np.array([[1,2,4],[1,4,5]])
ar2
ar3 = np.array([[[1,2,3],[3,4,5],[5,6,7]],
[[1,2,3],[3,4,5],[5,6,7]],
[[1,2,3],[3,4,5],[5,6,7]]])
ar3
ar.dtype
type(ar), type(ar2), type(ar3)
ar4 = np.array([[[[ar3, ar3, ar3, ar3], [ar3, ar3,ar3, ar3]], [[ar3, ar3,ar3, ar3], [ar3, ar3,ar3, ar3]]], [[[ar3, ar3,ar3, ar3], [ar3, ar3,ar3, ar3]], [[ar3, ar3,ar3, ar3], [ar3, ar3,ar3, ar3]]]])
ar4
ar.shape, ar2.shape, ar3.shape, ar4.shape
ar.ndim, ar2.ndim, ar3.ndim, ar4.ndim
ar.size, ar2.size, ar3.size, ar4.size
ar5 = np.array([[ar4], [ar4]])
pd.DataFrame(ar2)
# # Criando Arrays
# ### Completando com 1:
um = np.ones((1, 2, 2)) #Dá pra fazer com zeros também.
um
um.size, um.ndim
# ### Povoando com números aleatórios:
np.arange(1, 20, 2)
np.random.randint(0, 50, size = (2, 2, 3))
matriz = np.random.random((3,4))
matriz
rand = np.random.seed(seed = 10)
a, b = np.random.randint(100), np.random.randint(100)
a, b
# # Enxergando Arrays e Matrizes
a = np.random.randint(10, size = 10)
a
np.unique(a)
a.min()
matriz.min(), matriz[1].min()
matriz, matriz[1:3, 1:3],
#
# $x=\frac{10}{2}$
# Eu não sei como surgiu a linha "Type Markdown and LaTeX: 𝛼2". Mas fez eu me dar conta de que posso usar alguns comandos LateX aqui dentro, o que é foda.
ar3
ar3[1][0][1:3]
cu = pd.DataFrame({'A': range(10), 'B': range(20,30)})
cu
np.array(cu), np.array(cu)[1]
# # Manipulando Arrays
a = np.array([1, 3, 4])
a
b = np.ones(3)*2
b
a-b, abs(a-b)
a*ar3
matriz1 = np.array([[5, 6, 7], [7, 8, 9], [10, 11, 12]])
matriz1
matriz2 = np.array([[1, 2, 3], [3, 4, 5], [6, 7, 8]
])
matriz2
matriz1.shape, matriz2.shape
# #### Multiplicar usando * não é o mesmo que multiplicação matricial!
matriz1*matriz2
# #### Esse tipo de multiplicação é comutativo.
#
matriz2*matriz1
# ### Multiplicação matricial é feito por meio de dot:
np.dot(matriz1, matriz2)
np.dot(matriz2, matriz1)
ar = np.array([[1], [2], [3]])
ar1 = np.array([1, 2, 3])
ar, ar1
ar.shape, ar1.shape
np.dot(ar1, matriz1)
np.dot(matriz1, ar)
# # Reshape e Transposto
a.shape
a
# ### Mudando o formato do array:
# #### Mantendo a mesma informação!
a.reshape(3,1)
a.reshape(3,1,1)
a.reshape(3,2) #Não funciona, porque não tem informação o suficiente
a.reshape(1,3)
# ### Transposto
ar3, ar3.shape
ar3.T, ar3.T.shape
ar2, ar2.shape
ar2.T, ar2.T.shape
a
ar*ar3
ar3
ar
# ### Exemplo de multiplicação matricial na vida real
vendas = pd.DataFrame({'Almond butter': [2, 9, 11, 13, 15], 'Peanut butter': [7, 4, 14, 13, 18], 'Cashew butter': [1, 16, 18, 16, 9]}, index = ['Mon', 'Tues', 'Wed', 'Thurs', 'Fri'])
vendas
precos = pd.DataFrame({'Almond butter': [10], 'Peanut butter': [8], 'Cashew butter': [12]}, index = ['Preço'])
precos
matriz_vendas = np.array(vendas)
matriz_vendas
vetor_precos = np.array(precos)
vetor_precos
vetor_precos.shape
matriz_vendas.shape
vetor_total = np.dot( matriz_vendas, vetor_precos.T)
vetor_total
vendas['Total'] = vetor_total
vendas
# ### Ordenando Arrays
np.sort(matriz_vendas)
np.sort(vendas)
cu = np.array([3,7,1])
np.argsort(cu)
np.argmin(cu)
cu = np.array([[[2, 2]], [[2, 1]]])
cu.shape
np.argmin(cu, axis = 0)
# # Exemplo Prático: IMAGENS como arrays
# Para carregar a imagem, tem que estar na forma de MarkDown! <#img src = 'blablabla'> (Tira o #)
# <img src = 'feijao'>
from matplotlib.image import imread
feijao = imread('feijao.png')
type(feijao)
feijao.size, feijao.shape, feijao.ndim
feijao.dtype
feijao[0][0][0]
|
Numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collaboration and Competition
#
# ---
#
# In this notebook, you will learn how to use the Unity ML-Agents environment for the third project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.
#
# ### 1. Start the Environment
#
# We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
from unityagents import UnityEnvironment
import numpy as np
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/Tennis.app"`
# - **Windows** (x86): `"path/to/Tennis_Windows_x86/Tennis.exe"`
# - **Windows** (x86_64): `"path/to/Tennis_Windows_x86_64/Tennis.exe"`
# - **Linux** (x86): `"path/to/Tennis_Linux/Tennis.x86"`
# - **Linux** (x86_64): `"path/to/Tennis_Linux/Tennis.x86_64"`
# - **Linux** (x86, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86"`
# - **Linux** (x86_64, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `Tennis.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="Tennis.app")
# ```
env = UnityEnvironment(file_name="./App/Tennis.app")
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 2. Examine the State and Action Spaces
#
# In this environment, two agents control rackets to bounce a ball over a net. If an agent hits the ball over the net, it receives a reward of +0.1. If an agent lets a ball hit the ground or hits the ball out of bounds, it receives a reward of -0.01. Thus, the goal of each agent is to keep the ball in play.
#
# The observation space consists of 8 variables corresponding to the position and velocity of the ball and racket. Two continuous actions are available, corresponding to movement toward (or away from) the net, and jumping.
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
#print('The state for the first agent looks like:', states)
# -
# ### 3. Take Random Actions in the Environment
#
# In the next code cell, you will learn how to use the Python API to control the agents and receive feedback from the environment.
#
# Once this cell is executed, you will watch the agents' performance, if they select actions at random with each time step. A window should pop up that allows you to observe the agents.
#
# Of course, as part of the project, you'll have to change the code so that the agents are able to use their experiences to gradually choose better actions when interacting with the environment!
for i in range(1, 6): # play game for 5 episodes
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
states = env_info.vector_observations # get the current state (for each agent)
scores = np.zeros(num_agents) # initialize the score (for each agent)
while True:
actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
actions = np.clip(actions, -1, 1) # all actions between -1 and 1
env_info = env.step(actions)[brain_name] # send all actions to tne environment
next_states = env_info.vector_observations # get next state (for each agent)
rewards = env_info.rewards # get reward (for each agent)
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score (for each agent)
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Score (max over agents) from episode {}: {}'.format(i, np.max(scores)))
# When finished, you can close the environment.
# ### 4. It's Your Turn!
#
# Now it's your turn to train your own agent to solve the environment! When training the environment, set `train_mode=True`, so that the line for resetting the environment looks like the following:
# ```python
# env_info = env.reset(train_mode=True)[brain_name]
# ```
# +
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
def hidden_init(layer):
fan_in = layer.weight.data.size()[0]
lim = 1. / np.sqrt(fan_in)
return (-lim, lim)
class Actor(nn.Module):
"""Actor (Policy) Model."""
def __init__(self, state_size, action_size, seed, fc1_units=128, fc2_units=64):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fc1_units (int): Number of nodes in first hidden layer
fc2_units (int): Number of nodes in second hidden layer
"""
super(Actor, self).__init__()
self.seed = torch.manual_seed(seed)
self.bn0 = nn.BatchNorm1d(state_size)
self.fc1 = nn.Linear(state_size, fc1_units)
self.bn1 = nn.BatchNorm1d(fc1_units)
self.fc2 = nn.Linear(fc1_units, fc2_units)
self.bn2 = nn.BatchNorm1d(fc2_units)
self.fc3 = nn.Linear(fc2_units, action_size)
self.reset_parameters()
def reset_parameters(self):
self.fc1.weight.data.uniform_(*hidden_init(self.fc1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state):
"""Build an actor (policy) network that maps states -> actions."""
x = self.bn0(state)
x = F.relu(self.bn1(self.fc1(x)))
x = F.relu(self.bn2(self.fc2(x)))
return torch.tanh(self.fc3(x))
class Critic(nn.Module):
"""Critic (Value) Model."""
def __init__(self, state_size, action_size, seed,fcs1_units=128, fc2_units=32):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fcs1_units (int): Number of nodes in the first hidden layer
fc2_units (int): Number of nodes in the second hidden layer
fc3_units (int): Number of nodes in the third hidden layer
"""
super(Critic, self).__init__()
self.seed = torch.manual_seed(seed)
self.bn0 = nn.BatchNorm1d(state_size)
self.fcs1 = nn.Linear(state_size, fcs1_units)
self.fc2 = nn.Linear(fcs1_units+action_size, fc2_units)
self.fc3 = nn.Linear(fc2_units, 1)
self.reset_parameters()
def reset_parameters(self):
self.fcs1.weight.data.uniform_(*hidden_init(self.fcs1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state, action):
"""Build a critic (value) network that maps (state, action) pairs -> Q-values."""
state = self.bn0(state)
xs = F.relu(self.fcs1(state))
x = torch.cat((xs, action), dim=1)
x = F.relu(self.fc2(x))
return self.fc3(x)
# +
import random
import copy
from collections import namedtuple, deque
import torch.optim as optim
BUFFER_SIZE = int(1e5) # replay buffer size
BATCH_SIZE = 128 # minibatch size
GAMMA = 0.99 # discount factor
TAU = 1e-3 # for soft update of target parameters
LR_ACTOR = 1e-4 # learning rate of the actor
LR_CRITIC = 1e-3 # learning rate of the critic
WEIGHT_DECAY = 0 # L2 weight decay
#DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
DEVICE = "cpu"
class DDPGAgent():
"""Interacts with and learns from the environment."""
def __init__(self, state_size, action_size, random_seed):
"""Initialize an Agent object.
Params
======
state_size (int): dimension of each state
action_size (int): dimension of each action
random_seed (int): random seed
num_agents (int): number of agents
"""
self.state_size = state_size
self.action_size = action_size
self.seed = random.seed(random_seed)
self.device = DEVICE
self.buffer_size = BUFFER_SIZE
self.batch_size = BATCH_SIZE
self.gamma = GAMMA
self.tau = TAU
# Actor Network (w/ Target Network)
self.actor_local = Actor(state_size, action_size, random_seed).to(DEVICE)
self.actor_target = Actor(state_size, action_size, random_seed).to(DEVICE)
self.actor_optimizer = optim.Adam(self.actor_local.parameters(), lr=LR_ACTOR)
# Critic Network (w/ Target Network)
self.critic_local = Critic(state_size, action_size, random_seed).to(DEVICE)
self.critic_target = Critic(state_size, action_size, random_seed).to(DEVICE)
self.critic_optimizer = optim.Adam(self.critic_local.parameters(), lr=LR_CRITIC, weight_decay=WEIGHT_DECAY)
# Noise process
self.noise = OUNoise(action_size, random_seed)
# Replay memory
self.memory = ReplayBuffer(action_size, BUFFER_SIZE, BATCH_SIZE, random_seed, DEVICE)
def step(self, state, action, reward, next_state, done):
"""Save experience in replay memory, and use random sample from buffer to learn"""
# Save experience / reward
self.memory.add(state, action, reward, next_state, done)
# Learn, if enough samples are available in memory
if len(self.memory) > self.batch_size:
experiences = self.memory.sample()
self.learn(experiences)
def act(self, state, noise=0.0):
"""Returns actions for given state as per current policy"""
state = torch.from_numpy(state).float().to(self.device)
self.actor_local.eval()
with torch.no_grad():
action = self.actor_local(state).cpu().data.numpy()
self.actor_local.train()
action += noise*self.noise.sample()
return np.clip(action, -1, 1)
def reset(self):
self.noise.reset()
def learn(self, experiences):
"""Update policy and value parameters using given batch of experience tuples
q_targets = r + γ * critic_target(next_state, actor_target(next_state))
where:
actor_target(state) -> action
critic_target(state, action) -> Q-value
Params
======
experiences (Tuple[torch.Tensor]): tuple of (s, a, r, s', done)
gamma (float): discount factor
"""
states, actions, rewards, next_states, dones = experiences
# ---------------------------- update critic ---------------------------- #
# Get predicted next-state actions and Q values from target models
next_actions = self.actor_target(next_states)
q_targets_next = self.critic_target(next_states, next_actions)
# Compute Q targets for current states (y_i)
q_targets = rewards + (self.gamma * q_targets_next * (1 - dones))
# Compute critic loss
q_expected = self.critic_local(states, actions)
critic_loss = F.mse_loss(q_expected, q_targets)
# Minimize the loss
self.critic_optimizer.zero_grad()
critic_loss.backward()
self.critic_optimizer.step()
# ---------------------------- update actor ---------------------------- #
# Compute actor loss
predicted_actions = self.actor_local(states)
actor_loss = -self.critic_local(states, predicted_actions).mean()
# Minimize the loss
self.actor_optimizer.zero_grad()
actor_loss.backward()
self.actor_optimizer.step()
# ----------------------- update target networks ----------------------- #
self.soft_update(self.critic_local, self.critic_target, self.tau)
self.soft_update(self.actor_local, self.actor_target, self.tau)
def soft_update(self, local_model, target_model, tau):
"""Soft update model parameters
θ_target = τ*θ_local + (1 - τ)*θ_target
Params
======
local_model: PyTorch model (weights will be copied from)
target_model: PyTorch model (weights will be copied to)
tau (float): interpolation parameter
"""
for target_param, local_param in zip(target_model.parameters(), local_model.parameters()):
target_param.data.copy_(
tau * local_param.data + (1.0 - tau) * target_param.data)
def add_id(self, states):
"""Add (i+1) at the end of the states of the i-th agent as its id number."""
states_with_id = [np.concatenate([s, [i+1]]) for i,s in enumerate(states)]
return np.vstack(states_with_id)
def save_progress(self):
"""Save the most recent weights of local actor and critic."""
torch.save(self.actor_local.state_dict(), './Model_Weights/checkpoint_actor.pth')
torch.save(self.critic_local.state_dict(), './Model_Weights/checkpoint_critic.pth')
class OUNoise:
"""Ornstein-Uhlenbeck process."""
def __init__(self, size, seed, mu=0., theta=0.15, sigma=0.2):
"""Initialize parameters and noise process."""
self.mu = mu * np.ones(size)
self.theta = theta
self.sigma = sigma
self.seed = random.seed(seed)
self.reset()
def reset(self):
"""Reset the internal state (= noise) to mean (mu)."""
self.state = copy.copy(self.mu)
def sample(self):
"""Update internal state and return it as a noise sample."""
x = self.state
dx = self.theta * (self.mu - x) + self.sigma * np.random.randn(len(x))
#dx = self.theta * (self.mu - x) + self.sigma * np.random.standard_normal(len(x))
self.state = x + dx
return self.state
class ReplayBuffer:
"""Fixed-size buffer to store experience tuples."""
def __init__(self, action_size, buffer_size, batch_size, seed, device):
"""Initialize a ReplayBuffer object.
Params
======
buffer_size (int): maximum size of buffer
batch_size (int): size of each training batch
"""
self.action_size = action_size
self.memory = deque(maxlen=buffer_size) # internal memory (deque)
self.batch_size = batch_size
self.experience = namedtuple("Experience", field_names=["state", "action", "reward", "next_state", "done"])
self.seed = random.seed(seed)
self.device = device
def add(self, state, action, reward, next_state, done):
"""Add a new experience to memory."""
e = self.experience(state, action, reward, next_state, done)
self.memory.append(e)
def sample(self):
"""Randomly sample a batch of experiences from memory."""
experiences = random.sample(self.memory, k=self.batch_size)
device = self.device
states = torch.from_numpy(np.vstack([e.state for e in experiences if e is not None])).float().to(device)
actions = torch.from_numpy(np.vstack([e.action for e in experiences if e is not None])).float().to(device)
rewards = torch.from_numpy(np.vstack([e.reward for e in experiences if e is not None])).float().to(device)
next_states = torch.from_numpy(np.vstack([e.next_state for e in experiences if e is not None])).float().to(device)
dones = torch.from_numpy(np.vstack([e.done for e in experiences if e is not None]).astype(np.uint8)).float().to(device)
return (states, actions, rewards, next_states, dones)
def __len__(self):
"""Return the current size of internal memory."""
return len(self.memory)
# +
# creat a new agent to test trained model
agent_test = DDPGAgent(state_size=state_size+1, action_size=action_size, random_seed=12345)
# load trained weights
agent_test.actor_local.load_state_dict(torch.load('./Model_Weights/checkpoint_actor.pth',map_location={'cuda:0': 'cpu'}))
agent_test.critic_local.load_state_dict(torch.load('./Model_Weights/checkpoint_critic.pth',map_location={'cuda:0': 'cpu'}))
# test training result
for i in range(1, 6): # play game for 5 episodes
env_info = env.reset(train_mode=False)[brain_name] # reset the environment
states = agent_test.add_id(env_info.vector_observations) # get the current state
scores = np.zeros(num_agents) # initialize the score
while True:
actions = agent_test.act(states) # select an action
env_info = env.step(actions)[brain_name] # send all actions to the environment
next_states = agent_test.add_id(env_info.vector_observations) # get next state
rewards = env_info.rewards # get reward
dones = env_info.local_done # see if episode finished
scores += env_info.rewards # update the score
states = next_states # roll over states to next time step
if np.any(dones): # exit loop if episode finished
break
print('Score (max over agents) from episode {}: {}'.format(i, np.max(scores)))
# -
env.close()
|
Projects/p3_collab-compet/Tennis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrklees/PracticalStatistics/blob/master/3_Regression_Models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="w_d9MORIfjza" colab_type="text"
# # Regression Modelling
#
# ### Don't Forget to Run All (Ctrl+F9)
# ### Alex! Press the Record Button!!!
#
#
# By the end of the session participants will...
#
# * Review Expectation and Variance
# * Review the concept of a joint distribution
# *
#
# ## Language Repository
# These are some key terms that I will throw around a lot. You can always review their definitions here or [go back to the notebook on Statistical Testing & p-values to review.](https://colab.research.google.com/drive/15GQcjwz1TlVOOZfxiYakS1fAl3-3BiJA)
#
# **Expected Value: ** This is the average! It's the long-run average value of repetitions of the same experiment.
#
# **Variance:** This is a measure of how spread out a distribution is from its mean. The greater the variance, the futher values fall from its mean.
#
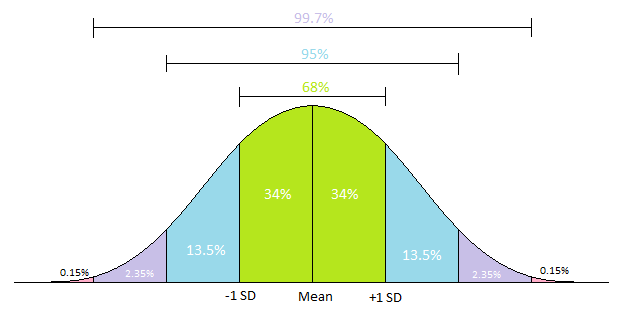
# **Standard Deviation:** This is really just the square root of the variance: $\text{std dev}(X) = \sqrt{Var(X)}$, and thus it behaves very similarly to variance. The greater the standard deviation, the futher values fall from its mean. It is often used in the context of normally distributed data because of how nicely a standard deviation partitions the normal distribution, per the obligatory graph:
#
# 
#
#
# + [markdown] id="bwTnA1BkfzOO" colab_type="text"
# ## Motivating Today with Simpson's Paradox
#
# In the last session, we learned about methods which allows us to directly compare two different variables, like focus list status and assessment scores. And, ignore the fact that I spoke at length about why you shouldn't really use those mothods, you might wonder why we don't just stop there. Why work up to more complex models?
#
# We will start off with a small example that gets at this idea. It involves a very simple table of data. Suppose we did some trial on the effectiveness of a drug on the risk of heart attacks. The data is summarized below.
#
# | | Control - Heart Attacks | Control - Total Participants | Treatment - Heart Attacks | Treatment - Total Participants+ |
# |:---:|:---:|:---:|:---:|:---:|
# |Female|1|20|3|40|
# |Male|12|40|8|20|
#
# The paradox here comes from the two stories that I can tell with this table of data.
#
# The **first story** is about a fantastic new drug! In a clinical trial of 120 participants, participants who tool the experimental drug experienced heart attacks had about a 15% reduction in heart attacks. Talk to your doctor today!
#
# However the **second story** is an investigative journlism piece about a dangerous drug put out on the market. When *looking at men and women separately* it turns that in either case it *raises your risk of heart attack* by about 25% (for men) to 50% (for women). Obviously a drug which is bad for both men and women should be banned.
#
# To be clear, these calculations are really just simple percentages. In the first story, we just pretend like the data isn't segmented. So $\frac{13}{60}=21.6\%$ participants in the control and $\frac{11}{60}=18.3\%$ in the treatment group experienced heart attacks. Of course I then presented their ratio, because it's a much more marketable number 👍.
#
# Since the second story is symmetrical, I'll just talk about women. In the control group, $\frac{1}{20}=5\%$ of women experience heart attacks where as $\frac{3}{40}=7.5\%$ women in the treatment group experienced heart attacks. A similar increase was seen for men.
#
# There is a lot within Simpson's Paradox. It was published over 60 years ago and still is the topic of some conversation. What I really want you to get from this is: this is what confounding can look like. If we just measure effects directly then there is a risk that what we measure could not only be different from the true effect, but could be in the completely wrong direction.
# + id="UsB6SDSaIf__" colab_type="code" colab={}
#@title Imports and Global Variables (run this cell first) { display-mode: "form" }
#@markdown This sets the warning status (default is `ignore`, since this notebook runs correctly)
warning_status = "ignore" #@param ["ignore", "always", "module", "once", "default", "error"]
import warnings
warnings.filterwarnings(warning_status)
with warnings.catch_warnings():
warnings.filterwarnings(warning_status, category=DeprecationWarning)
warnings.filterwarnings(warning_status, category=UserWarning)
import numpy as np
import pandas as pd
import os
#@markdown This sets the styles of the plotting (default is styled like plots from [FiveThirtyeight.com](https://fivethirtyeight.com/))
matplotlib_style = 'fivethirtyeight' #@param ['fivethirtyeight', 'bmh', 'ggplot', 'seaborn', 'default', 'Solarize_Light2', 'classic', 'dark_background', 'seaborn-colorblind', 'seaborn-notebook']
import matplotlib.pyplot as plt; plt.style.use(matplotlib_style)
# %matplotlib inline
import seaborn as sns; sns.set_context('notebook')
import statsmodels.api as sm
from statsmodels.api import OLS
# + id="fngP6ohTWJzv" colab_type="code" outputId="8ac01270-6a97-448b-83c2-519330ad5bcf" colab={"base_uri": "https://localhost:8080/", "height": 224}
#@title Read the Data from the Web { display-mode: "form" }
data_url = 'https://impactblob.blob.core.windows.net/public/anon_hmh.csv'
data = pd.read_csv(data_url)
data.head()
# + id="XwtSwre9WSzW" colab_type="code" outputId="ca78fe79-be37-4ab6-a59c-f36e56039154" colab={"base_uri": "https://localhost:8080/", "height": 365}
#@title Descriptive Stats to Reference { display-mode: "form" }
def describe_nulls(data):
desc = data.describe(include=data.dtypes.unique())
desc.loc['% Null'] = data.isna().sum() / data.shape[0]
return desc
describe_nulls(data)
# + [markdown] id="_lWAOH6QZjc3" colab_type="text"
# # Regression
#
# This is a typical section where people bring out formulas and abstract graphs to try to explain this topic... but in the flavor of practical statistics we're going to talk about regression in terms of a different conceptual model. One which will hopefully allow us to worry a little bit less about the math. The reason that I feel that I can get away with this is that *most modern statistical interfaces* allow you to operate at this level of abstraction with this kind of conceptual framework instead of having to concern yourself with the details. Of course there is some variation in the aesthetics, but largely they are all the same.
#
# ## Regression as a Framework for Prediction
#
# In our framework, regression is all about being able to take some data and make predictions about future data. In this context, we'll use a few new pieces of language with somewhat specific meanings. Let's go through the language with some explanation.
#
# **Features**: The data we use to make predictions. For a student, this might be things like grade, school, focus list student, etc... We hope that this data contains information about the **outcome** we want to predict.
#
# **Outcomes**: The variable we want to make predictions about. For example, we've made predictions about how much a student will improve on a math assessment.
#
# **Model**: A **model** is used to make predictions about **outcomes**. Generally speaking, it is simply some mathematical device which tells us how to multiply and add our data to get **predictions** about our **outcomes**.
#
# The general goal of Regression is to *fit* (or *train*) **models** on **features** where the **outcomes** are known, and then use them to make predictions on new data where the **outcomes** aren't known. In the last session we previewed a formula notation which expresses this idea. In its full form it might look like:
# $$ \text{Outcomes} \sim \text{Model(Features)}$$
# Although, we dropped the model portion, because it doesn't really tell us much. So we're just left with:
# $$ \text{Outcomes} \sim \text{Features}$$
#
# ### Fitting Model: What do we need to understand for Practical Statistics?
#
# For the purposes of Pratical Statistics, we will acknowledge that these models exist and talk *exclusively* about how to use them. Unfortunately, going much further into how a model is trained requires going into some calculus, so we will skate around the topic. Instead, since you can use software prepared by professionals you can largely trust that *as long as you specify the model correctly*, the software will not make a mistake in fitting the model correctly. What I mean by *specifying the model correctly* will be one of the key subjects we discuss.
# + id="EViiFp4wWsDg" colab_type="code" outputId="ef4c0fd8-5657-4e01-af3b-bcddb6a1e852" colab={"base_uri": "https://localhost:8080/", "height": 493}
#@title Regression Choose Your Own Adventure {run: 'auto', display-mode: "form"}
#@markdown This tool will allow you fit nearly any possible linear model from the data. Start by selecting which column will be the **Outcome**. I would recommend `MathAssess_RAWCHANGE` or `LITASSESS_RAWCHANGE`, but I've left every column available.
Outcomes = "MathAssess_RAWCHANGE" #@param ['GRADE_ID_NUMERIC', 'OFFICIALFLLIT', 'OFFICIALFLMTH', 'FL_LIT_MET_DOSAGE', 'FL_MTH_MET_DOSAGE', 'litassess_pre_value_num', 'LITASSESS_RAWCHANGE', 'LITASSESS_SRITARGET', 'mathassess_pre_value_num', 'MathAssess_RAWCHANGE', 'SMI_TARGET', 'AnonId', 'SiteId', 'SchoolId', 'att_pre_value', 'att_post_value']
#@markdown Then select which columns of data should be included in your model! Sorry for the rough interface, but Colab hasn't published a better one yet.
GRADE_ID_NUMERIC = False #@param {type:"boolean"}
OFFICIALFLLIT = False #@param {type:"boolean"}
OFFICIALFLMTH = False #@param {type:"boolean"}
FL_LIT_MET_DOSAGE = False #@param {type:"boolean"}
FL_MTH_MET_DOSAGE = False #@param {type:"boolean"}
litassess_pre_value_num = False #@param {type:"boolean"}
LITASSESS_RAWCHANGE = False #@param {type:"boolean"}
LITASSESS_SRITARGET = False #@param {type:"boolean"}
mathassess_pre_value_num = False #@param {type:"boolean"}
MathAssess_RAWCHANGE = False #@param {type:"boolean"}
SMI_TARGET = False #@param {type:"boolean"}
SiteId = False #@param {type:"boolean"}
SchoolId = False #@param {type:"boolean"}
att_pre_value = True #@param {type:"boolean"}
att_post_value = False #@param {type:"boolean"}
colnames= np.array([
'GRADE_ID_NUMERIC', 'OFFICIALFLLIT', 'OFFICIALFLMTH',
'FL_LIT_MET_DOSAGE', 'FL_MTH_MET_DOSAGE', 'litassess_pre_value_num',
'LITASSESS_RAWCHANGE', 'LITASSESS_SRITARGET',
'mathassess_pre_value_num', 'MathAssess_RAWCHANGE', 'SMI_TARGET',
'SiteId', 'SchoolId', 'att_pre_value', 'att_post_value'])
responses = np.array([GRADE_ID_NUMERIC, OFFICIALFLLIT, OFFICIALFLMTH,
FL_LIT_MET_DOSAGE, FL_MTH_MET_DOSAGE, litassess_pre_value_num,
LITASSESS_RAWCHANGE, LITASSESS_SRITARGET,
mathassess_pre_value_num, MathAssess_RAWCHANGE, SMI_TARGET,
SiteId, SchoolId, att_pre_value, att_post_value])
# Get Selected Features
Features = list(colnames[responses])
try:
assert Outcomes not in Features
model_string = Outcomes + " ~ " + ' + '.join(Features)
print(f"Current Model: {model_string}")
print("Fitting... Be aware that null data will be dropped...")
model = OLS(endog=data[Outcomes], exog=sm.add_constant(data[Features]), missing='drop').fit()
print("Done... Returning Summary...")
print(model.summary())
except AssertionError:
print("Your outcome variable is also selected as a feature you silly goose!")
# + [markdown] id="J84xOM1ntgkR" colab_type="text"
# 
# 
#
# # All Models Are Wrong
#
# Now that we have the ability to generate in fact thousands of different models, we need some method of choosing between them. It's at about this point that I should offer a piece of wisdom [attributed to statistician George Box](https://en.wikipedia.org/wiki/All_models_are_wrong): *"All models are wrong, but some are useful."* We aren't fortune tellers. As much as we would like, we cannot predict the future and our puny mathematics cannot fully represent natural processes (yet). So we start from the premise that our models are at best approximations, but thankfully the full effort of statisticians in the last century have gone into showing that we can build some good approximations and that there are some processes we can utilize to get there.
#
# ## Process for Finding Good Models
#
# Finding good models is ultimately a iterative process. Like a good scientist, we have to be consistently skeptical of our current model, knowing that there is probably a better one out there. I like thinking about this as a process, summed up in this short diagram:
#
# 
#
# The [developers of Edward call this Box's Loop](http://www.cs.columbia.edu/~blei/fogm/2015F/notes/intro.pdf), and as you can see it consists of a three step process which constantly repeats.
#
# 1. **Model:** Our model represents our beliefs about how the world works. We want to consistantly return to the question "what process generated the data that I'm observing", and try to make sure that our model is as consistent with that our beliefs about that process as possible.
# 2. **Infer:** This is a fancy word for fitting a model with data.
# 3. **Criticize:** Does out model make reasonable predictions? Is it more accurate than a coin flip?
#
# ## Model Criticism
#
# As we hinted above, one strategy for model criticism is to examine it's accuracy. What *accuracy* actually means depends on the outcome that we're trying to predict, but rest assured that there are good solutions for which ever outcome you choose.
#
# The results that we're getting from python are actually summarizing a bunch of different accuracy measurements, so let's talk a little bit about what they mean. These are the results that are drawn from the top right section of the results summary.
#
# | Measure | Example Outcome |
# |----------------|-----------|
# |R-squared| 0.01 |
# |Adj. R-squared| 0.009|
# |F-statistic|3.990|
# |Prob (F-statistic)|0.046|
# |Log-Likelihood|-7164.6|
# |AIC|1.433e+04|
# |BIC|1.434e+04|
#
# * [$R^2$ and $\text{Adj. } R^2$](https://en.wikipedia.org/wiki/Coefficient_of_determination): More formally called the coefficient of determination, this values is the percentage of the outputs variance that's explained by the features of the model. Values can be between 0 and 1, with 1 being 100% of the variance of the output being explined by the features. With $R^2 = 1$, our model should always make correct predictions. The *adjusted* value is very similar and tries to account for certain types of bias. It will typically be very close to the $R^2$ value, but is a good one to look at when comparing models.
# * [F-statistic and Prob (F-statistic)](https://en.wikipedia.org/wiki/F-test): This is a built in statistical test which essentially compares the model that you've proposed to one which has no features in it. If the Prob (F-statistic) (i.e. the p-value) is sufficiently low, then you might believe that this model with its proposed features are better than a model without them.
# * [Log-Likelihood](https://en.wikipedia.org/wiki/Maximum_likelihood_estimation): Without going into detail, when the model is fit it is trying to maximize this value. So between two models, the higher the log-likelihood the better the prediction accuracy.
# * [AIC and BIC](https://en.wikipedia.org/wiki/Akaike_information_criterion): It's not so important how these are calculated as I don't think they offer any greater interpritability, but the smaller the value the better when choosing your model.
#
#
# + [markdown] id="EnPPJmJRvKRQ" colab_type="text"
# # Mini-Hackathon Time
#
# With whatever time we have left your job is to explore the data and find the best sets of predictors for either Math or ELA assessment change.
#
# [And as usual, please leave me some feedback! It's appreciated :)](https://forms.office.com/Pages/ResponsePage.aspx?id=n4nHpSnR9kisiI-X82badMFC4tEHX8lCm8qe3Orb0kdUQkw2TkQxUFJEWjlCRElPVlJQWktVRlQ0QiQlQCN0PWcu)
|
3_Regression_Models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:azure]
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# ## Imports
from dmsbatch import create_batch_client, create_blob_client
# ## First create a batch client from the config file
client = create_batch_client('../tests/data/dmsbatch.config')
blob_client = create_blob_client('../tests/data/dmsbatch.config')
# ## Application packages
# To copy large files and programs it is best to zip (or targz) them and upload them as application packages
#
# Application packages are setup separately in either azure management apis or from the web console or cli tool
#
# These are referenced here by their name and version
# e.g. DSM2, python and other programs
app_pkgs = [('dsm2linux', '8.2.8449db2', 'DSM2-8.2.8449db2-Linux/bin')]
# ### Create or resize existing pool
# If the pool doesn't exist it will create it
# If the pool exists, it will resize to the second arg
pool_start_cmds = ['printenv',
'yum install -y glibc.i686 libstdc++.i686 glibc.x86_64 libstdc++.x86_64',# --setopt=protected_multilib=false',
'yum-config-manager --add-repo https://yum.repos.intel.com/2019/setup/intel-psxe-runtime-2019.repo',
'rpm --import https://yum.repos.intel.com/2019/setup/RPM-GPG-KEY-intel-psxe-runtime-2019',
'yum install -y intel-icc-runtime-32bit intel-ifort-runtime-32bit']
client.wrap_commands_in_shell('linux',pool_start_cmds)
client.create_pool('dsm2linuxpool',
1,
app_packages=[(app,version) for app,version,_ in app_pkgs],
vm_size='standard_f2s_v2',
tasks_per_vm=2,
os_image_data=('openlogic', 'centos', '7_8'),
start_task_cmd=client.wrap_commands_in_shell('linux',pool_start_cmds),
start_task_admin=True,
elevation_level='admin'
)
# ### Create job on pool or fail if it exists
# Jobs are containers of tasks (things that run on nodes (machines) in the pool). If this exists, the next line will fail
try:
client.create_job('dsm2linuxjobs','dsm2linuxpool')
except Exception as exc:
print('Job already exists?', exc.message)
# ### Upload input files
# Zip the input files and add them to storage container (in this case the auto storage associated with the batch account)
import os, datetime
userid = os.getlogin()
tsnow = str(datetime.datetime.now().timestamp()).split('.')[0]
task_name = f'hydro_run_{tsnow}_{userid}'
print(task_name)
local_dir = '../tests/data/dsm2v821'
input_file=blob_client.zip_and_upload('dsm2linuxjobs',f'{task_name}',local_dir,30)
input_file = client.create_input_file_spec('dsm2linuxjobs',blob_prefix=f'{task_name}/dsm2v821.zip',file_path='.')
import dmsbatch
permissions = dmsbatch.commands.azureblob.BlobPermissions.WRITE
# |helpers.azureblob.BlobPermissions.ADD|helpers.azureblob.BlobPermissions.CREATE
output_dir_sas_url = blob_client.get_container_sas_url('dsm2linuxjobs', permissions)
print(output_dir_sas_url)
std_out_files = client.create_output_file_spec('../std*.txt', output_dir_sas_url, blob_path=f'{task_name}')
output_dir = client.create_output_file_spec('**/output/*',output_dir_sas_url, blob_path=f'{task_name}')
# ### Create a task
# This uses the application package as pre -set up. If not, create one https://docs.microsoft.com/en-us/azure/batch/batch-application-packages
# tsnow = str(datetime.datetime.now().timestamp()).split('.')[0]
# task_name = f'hydro_run_{tsnow}'
set_path_string = client.set_path_to_apps(app_pkgs, ostype='linux')
cmd_string = client.wrap_cmd_with_app_path(f"""source /opt/intel/psxe_runtime/linux/bin/compilervars.sh ia32;
{set_path_string};
cd {task_name};
unzip dsm2v821.zip;
rm dsm2v821.zip;
cd study_templates/historical;
hydro hydro.inp;
qual qual_ec.inp;
""", app_pkgs, ostype='linux')
hydro_task=client.create_task(task_name, cmd_string,
resource_files=[input_file], output_files=[std_out_files, output_dir])
task_name
# ### Next submit the task and wait
#client.submit_tasks_and_wait('dsm2jobs',[hydro_task],poll_secs=120,wait_time_mins=300)
client.submit_tasks('dsm2linuxjobs',[hydro_task])
# ## Finally resize the pool to 0 to save costs
# +
#client.resize_pool('dsm2linuxpool',0)
# -
|
notebooks/sample_submit_dsm2_linux_historical.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Best Practices for Coding in Python
#
# There are certain guides to coding in Python that are expected. These can be found in more detail in [the PEP 8](https://www.python.org/dev/peps/pep-0008/) guidelines. But for the basics, read on!
#
# These coding guidelines will be enforced with a CI tool, so make sure to follow these rules!
# Python doesn't care about whether you use Tabs or Spaces, and so in general the rule is to be consistent. Since this is hard to enforce in large projects, the rule as determined is to use **spaces** over **tabs**. Use **4 spaces** every time that you are indenting.
|
IntroToPython/StyleGuide.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:TorchDetectAML]
# language: python
# name: conda-env-TorchDetectAML-py
# ---
# # Publish a Training Pipeline
# In this notebook, we will show how to automate the training/retraining of model using HyperDrive and registering best model. Once this training pipeline is published/created, it provides a REST endpoint which can be called to run this pipeline without using the Azure Machine Learning Service SDK.
#
#
# ## Imports
# +
import os
import requests
import azureml
from azureml.core import Workspace, Experiment
from azureml.core.datastore import Datastore
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.exceptions import ComputeTargetException
from azureml.data.data_reference import DataReference
from azureml.pipeline.steps import HyperDriveStep, PythonScriptStep
from azureml.pipeline.core import Pipeline, PipelineData, PipelineParameter
from azureml.core.runconfig import RunConfiguration, CondaDependencies
from azureml.train.dnn import PyTorch
from azureml.core.container_registry import ContainerRegistry
from azureml.train.hyperdrive import (
RandomParameterSampling,
BanditPolicy,
uniform,
choice,
HyperDriveConfig,
PrimaryMetricGoal,
)
from azureml.widgets import RunDetails
from dotenv import set_key, get_key, find_dotenv
from utilities import get_auth
# check core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
# -
env_path = find_dotenv(raise_error_if_not_found=True)
# ## Read in the Azure ML workspace and default datastore
# Read in the the workspace created in a previous notebook.
auth = get_auth(env_path)
ws = Workspace.from_config(auth=auth)
print(ws.name, ws.resource_group, ws.location, sep="\n")
ds = ws.get_default_datastore()
# ## Tune Model Hyperparameters
# We automatically tune hyperparameters by exploring the range of values defined for each hyperparameter. Here we use random sampling which randomly selects hyperparameter values from the defined search space. Random sampling allows for both discrete and continuous hyperparameters.
param_sampling = RandomParameterSampling(
{
"learning_rate": uniform(0.0005, 0.005),
"rpn_nms_thresh": uniform(0.3, 0.7),
"anchor_sizes": choice(
"16",
"16,32",
"16,32,64",
"16,32,64,128",
"16,32,64,128,256",
"16,32,64,128,256,512",
),
"anchor_aspect_ratios": choice(
"0.25", "0.25,0.5", "0.25,0.5,1.0", "0.25,0.5,1.0,2.0"
),
}
)
# The num epochs and maximum total run parameters deliberately have a low default value for the speed of running. In actual application, set these to higher values (i.e. num_epochs = 10, max_total_runs = 16)
# + tags=["parameters"]
# number of epochs
num_epochs = 1
# max total runs for hyperdrive
max_total_runs = 1
# -
# It is also possible to specify a maximum duration for the tuning experiment by setting `max_duration_minutes`. If both of these parameters are specified, any remaining runs are terminated once `max_duration_minutes` have passed.
# We will terminate poorly performing runs automatically with bandit early termination policy which is based on slack factor and evaluation interval. The policy terminates any run where the primary metric is not within the specified slack factor with respect to the best performing training run.
early_termination_policy = BanditPolicy(
slack_factor=0.15, evaluation_interval=2, delay_evaluation=2
)
# ## Create an estimator <a id='estimator'></a>
# Create an estimator that specifies the location of the script, sets up its fixed parameters, including the location of the data, the compute target, and specifies the packages needed to run the script. It may take a while to prepare the run environment the first time an estimator is used, but that environment will be used until the list of packages is changed.
cluster_name = get_key(env_path, 'cluster_name')
# +
try:
compute_target = ComputeTarget(workspace=ws, name=cluster_name)
print("Found existing compute target.")
except ComputeTargetException:
print("Creating a new compute target...")
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_NC6", max_nodes=8
)
# create the cluster
compute_target = ComputeTarget.create(ws, cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# use get_status() to get a detailed status for the current cluster.
print(compute_target.get_status().serialize())
# -
script_folder = "./torchdetect"
image_name = get_key(env_path, "image_name")
# point to an image in private ACR
image_registry_details = ContainerRegistry()
image_registry_details.address = get_key(env_path, "acr_server_name")
image_registry_details.username = get_key(env_path, "acr_username")
image_registry_details.password = get_key(env_path, "acr_password")
# +
estimator = PyTorch(
source_directory=script_folder,
compute_target=compute_target,
entry_script="train.py",
use_docker=True,
custom_docker_image=image_name,
image_registry_details=image_registry_details,
user_managed=True,
use_gpu=True,
)
estimator.run_config.environment.environment_variables["PYTHONPATH"] = "$PYTHONPATH:/cocoapi/PythonAPI/"
# -
# Put the estimator and the configuration information together into an HyperDrive run configuration object.
hyperdrive_config = HyperDriveConfig(
estimator=estimator,
hyperparameter_sampling=param_sampling,
policy=early_termination_policy,
primary_metric_name="mAP@IoU=0.50",
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=max_total_runs,
max_concurrent_runs=4,
)
# ## Azure Machine Learning Pipelines: Overview <a id='aml_pipeline_overview'></a>
#
# A common scenario when using machine learning components is to have a data workflow that includes the following steps:
#
# - Preparing/preprocessing a given dataset for training, followed by
# - Training a machine learning model on this data, and then
# - Deploying this trained model in a separate environment, and finally
# - Running a batch scoring task on another data set, using the trained model.
#
# Azure's Machine Learning pipelines give you a way to combine multiple steps like these into one configurable workflow, so that multiple agents/users can share and/or reuse this workflow. Machine learning pipelines thus provide a consistent, reproducible mechanism for building, evaluating, deploying, and running ML systems.
#
# To get more information about Azure machine learning pipelines, please read our [Azure Machine Learning Pipelines overview](https://aka.ms/pl-concept), or the [getting started notebook](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/machine-learning-pipelines/intro-to-pipelines/aml-pipelines-getting-started.ipynb).
#
# Let's create a data reference for the raw data to be used in HyperDrive run.
data_folder = DataReference(datastore=ds, data_reference_name="data_folder")
model_name = PipelineParameter(name="model_name", default_value="torchvision_best_model")
# ## Create AML Pipeline Tuning Step
# We create a HyperDrive step in the AML pipeline to perform a search for hyperparameters. The `tune_epochs` pipeline parameter that controls the number of epochs used in tuning deliberately has a low default value for the speed of pipeline testing.
tune_step_name="tune_model"
tune_epochs = PipelineParameter(name="tune_epochs", default_value=1) # Set to 10 when running the pipeline.
tune_step = HyperDriveStep(
name=tune_step_name,
hyperdrive_config=hyperdrive_config,
estimator_entry_script_arguments=["--data_path", data_folder,
"--workers", 8,
"--epochs", tune_epochs,
"--box_nms_thresh", 0.3,
"--box_score_thresh", 0.10],
inputs=[data_folder],
allow_reuse=False)
# ## Create AML Pipeline Register Model Step
# This Python script step registers the best model found by the HyperDrive step.
# Let's create a folder for the script.
script_folder = "./registermodel"
os.makedirs(script_folder, exist_ok=True)
# +
# %%writefile registermodel/Register_Model.py
from __future__ import print_function
import os
import json
import argparse
from azureml.core import Run
from azureml.pipeline.core import PipelineRun
from azureml.pipeline.steps import HyperDriveStepRun
import azureml.core
if __name__ == "__main__":
print("azureml.core.VERSION={}".format(azureml.core.VERSION))
parser = argparse.ArgumentParser(description="Register the model created by"
" an HyperDrive step")
parser.add_argument("--hd-step", dest="hd_step",
help="the name of the HyperDrive step")
parser.add_argument("--outputs", help="the model file outputs directory")
parser.add_argument("--model-name", dest="model_name",
help="the model file base name")
args = parser.parse_args()
model_name = args.model_name
model_file = "model_latest.pth"
model_path = os.path.join(args.outputs, model_file)
# Get the HyperDrive run.
run = Run.get_context()
print(run)
pipeline_run = PipelineRun(run.experiment, run.parent.id)
print(pipeline_run)
hd_step_run = HyperDriveStepRun(step_run=pipeline_run.find_step_run(args.hd_step)[0])
print(hd_step_run)
# Get the best run.
hd_step_run.wait_for_completion(show_output=True, wait_post_processing=True)
best_run = hd_step_run.get_best_run_by_primary_metric()
if best_run is None:
raise Exception("No best run was found")
print(best_run)
# Register the model
model = best_run.register_model(model_name=model_name, model_path=model_path)
print("Best Model registered")
# -
# Creating PythonScript Step for AML pipeline to register the best model. The `bm_steps_data` input pipeline data is only used to synchronize with the previous pipeline step.
rm_step_name = "register_model"
rm_run_config = RunConfiguration(conda_dependencies=CondaDependencies.create(
pip_packages=["azure-cli", "azureml-sdk", "azureml-pipeline"]))
rm_run_config.environment.docker.enabled = True
rm_step = PythonScriptStep(
name=rm_step_name,
script_name="Register_Model.py",
compute_target=compute_target,
source_directory=os.path.join(".", "registermodel"),
arguments=["--hd-step", tune_step_name,
"--outputs", "outputs",
"--model-name", model_name],
runconfig=rm_run_config,
allow_reuse=False)
# Let's specify to run register model step after tune model step in the pipeline.
rm_step.run_after(tune_step)
# ## Create & Run a Pipeline
# When we specify the rm_step, Pipeline walks the dependency graph to include the other steps.
experiment_name = "torchvision"
exp = Experiment(workspace=ws, name=experiment_name)
pipeline = Pipeline(workspace=ws, steps=[rm_step])
pipeline.validate()
# Run the pipeline before publishing it. Wait for the run to complete.
pipeline_run = exp.submit(pipeline, continue_on_step_failure=True)
RunDetails(pipeline_run).show()
pipeline_run.wait_for_completion(show_output=True)
# ## Publish The Pipeline
# You may read more about why to publish a pipeline and how the published pipeline can be triggered [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-ml-pipelines)
published_pipeline = pipeline.publish(name="DL HyperDrive Pipeline",
description="DL HyperDrive Pipeline",
continue_on_step_failure=True)
published_pipeline.endpoint
# ## Run published pipeline using its REST endpoint <a id='run_publish_aml_pipeline'></a>
# This step shows how to call the REST endpoint of a published pipeline to trigger the pipeline run. You can use this method in programs that do not have the Azure Machine Learning SDK installed.
aad_token = auth.get_authentication_header()
rest_endpoint = published_pipeline.endpoint
print("You can perform HTTP POST on URL {} to trigger this pipeline".format(rest_endpoint))
response = requests.post(rest_endpoint,
headers=aad_token,
json={"ExperimentName": experiment_name,
"RunSource": "SDK"})
run_id = response.json()["Id"]
print(run_id)
# You can now proceed to the next notebook to [delete the resources of this tutorial](06_TearDown.ipynb).
|
05_TrainWithAMLPipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python385jvsc74a57bd0c94be41889154d41bf43aca8d1a8d1cd64b97c119170e03e2ed46ca87183f0c5
# ---
# + [markdown] id="INl1Ce2XJJDT"
#
#
# ---
#
# ---
# # Acesso às bases de dados da disciplina
#
# Esse notebook tem como objetivo apenas testar o acesso aos conjuntos de dados que poderão ser utilizados no decorrer da disciplina disponibilizados no *GitHub* e *Kaggle*.
#
#
# ---
#
# ---
#
#
#
#
# + [markdown] id="F_CL-ht_Gp_k"
# ## Acesso pelo *Kaggle* (sem usar API tokens)
#
# Basta executar o comando "wget" no endereço do link utilizado pelo site para fazer o download da base de dados hospedada no "Kaggle".
# + [markdown] id="f9--MufIMrCY"
# ## Dogs vs Cats
#
# - Dados de treino (dataset_treino): 8.000 imagens = 4.000 imagens de cães e 4.000 imagens de gatos
#
# - Dados de validação (dataset_validação): 2.000 imagens = 1.000 imagens de cães e 1.000 imagens de gatos
#
# - Dados de teste (dataset_teste): 1.000 imagens de cães e gatos
#
# Para acessar a base de dados execute os seguintes passos:
#
# 1. Acesse o endereço: https://www.kaggle.com/mrcioleandrogonalves/dogs-vs-cats
# 2. Clique em "download" (e pode cancelar o processo assim que iniciar)
# 3. Copiar o link para este processo na aba de downloads do navegador e colar no espaço entre aspas simples no comando wget abaixo (o endereço é longo!)
# + id="NQqLSgiyoOj-" colab={"base_uri": "https://localhost:8080/"} outputId="06410a7c-0edf-4f4c-c947-fa68b37ef7af"
# !wget 'https://storage.googleapis.com/kaggle-data-sets/1167797/1956504/bundle/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=gcp-kaggle-com%40kaggle-161607.iam.gserviceaccount.com%2F20210508%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210508T140256Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=host&X-Goog-Signature=478264a77a6f73480edc73a394143170f9a08ff13285ccb2a034cdce35e01dd67c2dee97613df4049cd15c56567d42f5e7fa2d5c5f1e903eb6dd51d06e919682a6cca3c9fa61e359cfcce74b7b861a591a754fbb7155f1035667a07ea24c1c81edeba73e1679d35414235671e3849dabfb8b591cbc5c19ec60c0d5e4cc34415a4759b60cbad7ee29ecb465c5dc7a81827c6d29fb7dde6fca93b05cf279a3509419f975218e85c6ccc0636c119094a06551e16f1241fb61b7e8e77ccaa00c4d6425e13aecf73b4844c10988ea796033bebb3f663714b4b29685429317e09f406febe04786cdff134f6b7c996e89106bceb15c5df1fee0a6fb13546fb7330a08d1'
# + [markdown] id="XTsnRfWwoWkk"
# ### Renomear o arquivo (cujo nome fica longo demais)
#
# Copie o nome do arquivo baixado e cole no espaço entre aspas simples do comando abaixo.
# + id="b2QyerekokXP"
# !mv '/content/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=<EMAIL>%2F20210508%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210508T140256Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=hos' dogs_vs_cats.zip
# + [markdown] id="5qsU62Z8p9kV"
# ### Descompactar
# + id="IKoM5Y8cM4zH" colab={"base_uri": "https://localhost:8080/"} outputId="77067bc1-41d7-4d46-bd87-bbcbadaaf3b4"
# !unzip -qu dogs_vs_cats.zip -d Dogs_vs_cats
# !ls -l
# + [markdown] id="1DTipVXkPcV5"
# Abaixo seguem visualizações de algumas imagens do conjunto de dados de treinamento (dataset_treino).
# + id="9E-W4I5PPewO" colab={"base_uri": "https://localhost:8080/", "height": 516} outputId="b42f2c34-cd7b-479d-d13f-c90ddd282fa3"
from IPython.display import Image
Image(filename = 'Dogs_vs_cats/dataset_treino/cats/cat.7.jpg')
# + id="BR6iV6sUPilm" colab={"base_uri": "https://localhost:8080/", "height": 445} outputId="330c9c97-b332-4faa-f872-f9b891ef1161"
Image(filename='Dogs_vs_cats/dataset_treino/dogs/dog.13.jpg')
# + id="H8SuuVXlPtX0" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="645e5d3c-ef3c-4477-c19a-ab2637c348e3"
Image(filename='Dogs_vs_cats/dataset_treino/cats/cat.3901.jpg')
# + id="2rJJIISmPxLU" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="ea1f2a53-0d5f-4c1b-c169-fd903db9fbf9"
Image(filename='Dogs_vs_cats/dataset_treino/dogs/dog.3998.jpg')
# + [markdown] id="aQ4bPwYcPy3l"
# ## COVID-19
#
# Conjuntos de dados de imagens de tomografias computadorizadas (CT-scans: Computerized Tomography - scans) em duas classes:
#
# - Dados de treino: 1800 imagens = 900 imagens CT de pulmões com COVID e 900 imagens CT de pulmões sem COVID
#
# - Dados de validação: 600 imagens = 300 imagens CT de pulmões com COVID e 300 imagens CT de pulmões sem COVID
#
# - Dados de teste: 60 imagens = 30 imagens de pulmões com COVID e 30 imagens de pulmões sem COVID
#
# Para acessar a base de dados execute os seguintes passos:
#
# 1. Acesse o endereço: https://www.kaggle.com/mrcioleandrogonalves/covid19
# 2. Clique em "download" (e pode cancelar o processo assim que iniciar)
# 3. Copiar o link para este processo na aba de downloads do navegador e colar no espaço entre aspas simples no comando wget abaixo (o endereço é longo!)
#
# + id="1lSs9z4mpuUW" colab={"base_uri": "https://localhost:8080/"} outputId="dd33ff90-8f92-40d6-eba5-8e0245a54d2d"
# !wget 'https://storage.googleapis.com/kaggle-data-sets/1168490/1957615/bundle/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=gcp-kaggle-com%40kaggle-161607.iam.gserviceaccount.com%2F20210508%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210508T140418Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=host&X-Goog-Signature=423ae0f21142f08379617d6a9dca41b68c19c16f6d2e5e20618052cc02a0843f256cfe7e09643af4df652bcfd2a0efd53220f35a1ecf1f9f17a6eeba0753686f31caf1a1f72ec7e650ee491136a9360d7475f9cdef5909cee3760bd444ca3e9bf81adabed172c7e2e4e3ce6cc720ceaa054c345d0352258acce9d426185f3b90c816462f61158e5ec03a7bb36ac79672ec856a10914e3d98bfb4f2a94f5565bd86696cb426b909b5803340c18d702e970fb8c5b250ef8c6debb1e03de95c3e9a7ca4f88720126a0d97739bd130ff04f9b5c7538da304d7fdee4128bfe2c3a2075ebc2cfbe86a86e07909b69dbb1aabde63727b14ac23b60a49f8930f39a15679'
# + [markdown] id="X5RIBCOwqF1O"
# ### Renomear o arquivo (cujo nome fica longo demais)
#
# Copie o nome do arquivo baixado e cole no espaço entre aspas simples do comando abaixo.
# + id="1cwAerFlqHOl"
# !mv '/content/archive.zip?X-Goog-Algorithm=GOOG4-RSA-SHA256&X-Goog-Credential=gcp-kaggle-com@kaggle-161607.iam.gserviceaccount.com%2F20210508%2Fauto%2Fstorage%2Fgoog4_request&X-Goog-Date=20210508T140418Z&X-Goog-Expires=259199&X-Goog-SignedHeaders=hos' covid-19.zip
# + [markdown] id="SesQp88dqqNF"
# ### Descompactar
# + id="sF6IaM7RQ0Cl" colab={"base_uri": "https://localhost:8080/"} outputId="c7b9c101-0514-44c1-cdd4-bb2826ab6fe5"
# !unzip -qu covid-19.zip -d COVID-19
# !ls -l
# + [markdown] id="6sa5QBtnRBl0"
# Imagens COVID
# + id="lZf5ShNpREQ8" colab={"base_uri": "https://localhost:8080/", "height": 680} outputId="d6795977-82dd-442d-92f4-5d6754aeeeba"
import matplotlib.pyplot as plt
import os
arqs_img = os.listdir('COVID-19/dataset_treino/COVID')
max_arqs = len(arqs_img)
rows = 5
cols = 5
if (rows*cols < max_arqs):
fig, ax = plt.subplots(rows, cols, figsize=(12, 12))
n = 0
for i in range(rows):
for j in range(cols):
file_name = 'COVID-19/dataset_treino/COVID/'+arqs_img[n]
image = plt.imread(file_name)
ax[i, j].set_title(arqs_img[n])
ax[i, j].set_xticks([])
ax[i, j].set_yticks([])
ax[i, j].imshow(image)
n += 1
|
aprendizado-de-maquina-ii/acesso-base-de-dados.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Presentation materials
#
# This notebook can be used to create materials we specifically want to use for the presentation. E.g. charts
# +
# import required modules and set up environment
import os
# replace file path below with your own local convokit
os.chdir('/Users/marianneaubin/Documents/Classes/CS6742/cs6742-fork')
import convokit
from convokit import Corpus, Parser, EmoTracker, Transformer
import nltk
from datetime import datetime
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import csv
import matplotlib.dates as mdates
# +
# we have reddit data for 2007 to 2018
events = [['Fort Hood', '2009-11-05', '2009-11-13', 'forthood'],
['Aurora Theater', '2012-07-20', '2012-07-28', 'auroratheater'],
['Sandy Hook', '2012-12-14', '2012-12-22', 'sandyhook'],
['Washington Navy Yard', '2013-09-16' , '2013-09-24', 'wanavyyard'],
['San Bernardino', '2015-12-02' , '2015-12-10', 'sanbernandino'],
['Orlando Nightclub', '2016-06-12' , '2016-06-20', 'orlandonightclub'],
['Las Vegas', '2017-10-01' , '2017-10-09', 'lasvegas'],
['Sutherland Springs Church', '2017-11-05' , '2017-11-13', 'sutherland'],
['St<NAME>', '2018-02-14' , '2018-02-22', 'parkland'],
['Santa Fe High', '2018-05-18' , '2018-05-26', 'santafehigh'],
['Pittsburgh Synagogue', '2018-10-27' , '2018-11-04', 'pittsburgh']]
events_list = ['Fort Hood','Aurora Theater', 'Sandy Hook', 'Washington Navy Yard', 'San Bernadino',
'Orlando Nightclub', 'Las Vegas', 'Sutherland Springs Church', 'Stoneman Douglas High',
'Santa Fe High', 'Pittsburgh Synagogue']
dates_list = ['2009-11-05','2012-07-20','2012-12-14', '2013-09-16', '2015-12-02', '2016-06-12' ,
'2017-10-01' , '2017-11-05' , '2018-02-14' , '2018-05-18', '2018-10-27']
# -
events_df = pd.DataFrame()
events_df["date"] = dates_list
events_df["event"] = events_list
events_df.head()
def GenerateTimeLine(data, title="Timeline", xaxis_format="%d %b", day_interval=5, figsize=(8, 5)):
"""Generates timeline from a pandas dataframe
data : Pandas dataframe with datetimeindex
title Title of the plot
zaxis_format ="%d %b %Y" valids strdate format
dayinterval =1 default, can be anything
Insipred from Matplotlib's excellent examples
author: sukhbinder
date 5/6/2018
"""
levels = np.array([-5, 5, -3, 3, -1, 1])
fig, ax = plt.subplots(figsize=figsize)
# Create the base line
start = min(data.index)
stop = max(data.index)
ax.plot((start, stop), (0, 0), 'k', alpha=.5)
# Iterate through data annoting each one
for ii, (idate, iname) in enumerate(data.itertuples()):
level = levels[ii % 6]
vert = 'top' if level < 0 else 'bottom'
ax.scatter(idate, 0, s=100, facecolor='w', edgecolor='k', zorder=9999)
# Plot a line up to the text
ax.plot((idate, idate), (0, level), c='r', alpha=.7)
# Give the text a faint background and align it properly
ax.text(idate, level, iname,ha='right', va=vert, fontsize=14,
backgroundcolor=(1., 1., 1., .3))
ax.set(title=title)
# Set the xticks formatting
# format xaxis with days intervals
ax.get_xaxis().set_major_locator(mdates.DayLocator(interval=day_interval))
ax.get_xaxis().set_major_formatter(mdates.DateFormatter(xaxis_format))
fig.autofmt_xdate()
# Remove components for a cleaner look
plt.setp((ax.get_yticklabels() + ax.get_yticklines() +
list(ax.spines.values())), visible=False)
return ax
data = pd.read_csv(r'events.csv', parse_dates=True, index_col=0)
ax = GenerateTimeLine(events_df)
plt.show()
# +
events = [['Fort Hood', '2009-11-05', '2009-11-13', 'forthood'],
['Aurora Theater', '2012-07-20', '2012-07-28', 'auroratheater'],
['<NAME>', '2012-12-14', '2012-12-22', 'sandyhook'],
['Washington Navy Yard', '2013-09-16' , '2013-09-24', 'wanavyyard'],
['<NAME>', '2015-12-02' , '2015-12-10', 'sanbernandino'],
['Orlando Nightclub', '2016-06-12' , '2016-06-20', 'orlandonightclub'],
['Las Vegas', '2017-10-01' , '2017-10-09', 'lasvegas'],
['Sutherland Springs Church', '2017-11-05' , '2017-11-13', 'sutherland'],
['<NAME>', '2018-02-14' , '2018-02-22', 'parkland'],
['Santa Fe High', '2018-05-18' , '2018-05-26', 'santafehigh'],
['Pittsburgh Synagogue', '2018-10-27' , '2018-11-04', 'pittsburgh']]
import plotly.figure_factory as ff
df = [dict(Task='Event', Start='2009-11-05', Finish='2009-11-13' , Event='Fort Hood'),
dict(Task='Event', Start='2012-07-20', Finish='2012-07-28', Event = 'Aurora Theater'),
dict(Task='Event', Start='2012-12-14', Finish='2012-12-22', Event ='Sandy Hook' ),
dict(Task='Event', Start='2013-09-16', Finish='2013-09-24', Event ='Washington Navy Yard'),
dict(Task='Event', Start='2015-12-02', Finish='2015-12-10', Event ='San Bernardino'),
dict(Task='Event', Start='2016-06-12', Finish='2016-06-20', Event ='Orlando Nightclub'),
dict(Task='Event', Start='2017-10-01', Finish='2017-10-09', Event ='Las Vegas'),
dict(Task='Event', Start='2017-11-05', Finish='2017-11-13', Event ='Sutherland Springs Church'),
dict(Task='Event', Start='2018-02-14', Finish='2018-02-22', Event ='Stoneman Douglas High'),
dict(Task='Event', Start='2018-05-18', Finish='2018-05-26', Event ='Santa Fe High'),
dict(Task='Event', Start='2018-10-27', Finish='2018-11-04', Event ='Pittsburgh Synagogue'),
dict(Task='Event', Start='2009-03-10', Finish='2009-03-18', Event ='Geneva County')]
colors = {'Fort Hood': 'rgb(220, 0, 0)',
'Aurora Theater': 'rgb(0, 0, 0)',
'Sandy Hook': 'rgb(0, 0, 0)',
'Washington Navy Yard': 'rgb(220, 0, 0)',
'Las Vegas': 'rgb(0, 0, 0)',
'<NAME>': 'rgb(0, 0, 0)',
'Orlando Nightclub': 'rgb(0, 0, 0)',
'Sutherland Springs Church': 'rgb(0, 0, 0)',
'<NAME>': 'rgb(0, 0, 0)',
'Santa Fe High': 'rgb(0, 0, 0)',
'Pittsburgh Synagogue': 'rgb(220, 0, 0)',
'Geneva County': 'rgb(220, 0, 0)',
}
fig = ff.create_gantt(df, colors=colors, index_col='Event', group_tasks=True)
fig.show()
# -
|
examples/politicization/create_presentation_materials.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas pipelines
#
# [Method chaining](https://tomaugspurger.github.io/method-chaining) is a great way for writing pandas code as it allows us to go from:
#
# ```python
# raw_data = pd.read_parquet(...)
# data_with_types = set_dtypes(raw_data)
# data_without_outliers = remove_outliers(data_with_types)
# ```
#
# to
#
# ```python
# data = (
# pd.read_parquet(...)
# .pipe(set_dtypes)
# .pipe(remove_outliers)
# )
# ```
#
# But it does come at a cost, mostly in our ability to debug long pipelines. If there's a mistake somewhere along the way, you can only inspect the end result and lose the ability to inspect intermediate results. A mitigation for this is to add decorators to your pipeline functions that log common attributes of your dataframe on each step:
#
#
# ## Logging in method chaining
# In order to use the logging capabilitites we first need to ensure we have a proper logger configured. We do this by running `logging.basicConfig(level=logging.DEBUG)`.
#
#
from sklego.datasets import load_chicken
from sklego.pandas_utils import log_step
chickweight = load_chicken(give_pandas=True)
# +
import logging
logging.basicConfig(level=logging.DEBUG)
# -
# If we now add a `log_step` decorator to our pipeline function and execute the function, we see that we get some logging statements for free
@log_step
def set_dtypes(chickweight):
return chickweight.assign(
diet=lambda d: d['diet'].astype('category'),
chick=lambda d: d['chick'].astype('category'),
)
# + tags=[]
chickweight.pipe(set_dtypes).head()
# -
# We can choose to log at different log levels. For example if we have a `remove_outliers` function that calls different outlier removal functions for different types of outliers, we might in general be only interested in the total outliers removed. In order to get that, we set the log level for our specific implementations to `logging.DEBUG`
# +
@log_step(level=logging.DEBUG)
def remove_dead_chickens(chickweight):
dead_chickens = chickweight.groupby('chick').size().loc[lambda s: s < 12]
return chickweight.loc[lambda d: ~d['chick'].isin(dead_chickens)]
@log_step
def remove_outliers(chickweight):
return chickweight.pipe(remove_dead_chickens)
# + tags=[]
chickweight.pipe(set_dtypes).pipe(remove_outliers).head()
# -
# We can now easily switch between log levels to get the full detail or the general overview
# + tags=[]
logging.getLogger(__name__).setLevel(logging.INFO)
chickweight.pipe(set_dtypes).pipe(remove_outliers).head()
# -
# The log step function has some settings that let you tweak what exactly to log:
# - `time_taken`: log the time it took to execute the function (default True)
# - `shape`: log the output shape of the function (default True)
# - `shape_delta`: log the difference in shape between input and output (default False)
# - `names`: log the column names if the output (default False)
# - `dtypes`: log the dtypes of the columns of the output (default False)
#
# For example, if we don't care how long a function takes, but do want to see how many rows are removed if we remove dead chickens:
#
# + tags=[]
@log_step(time_taken=False, shape=False, shape_delta=True)
def remove_dead_chickens(chickweight):
dead_chickens = chickweight.groupby('chick').size().loc[lambda s: s < 12]
return chickweight.loc[lambda d: ~d['chick'].isin(dead_chickens)]
chickweight.pipe(remove_dead_chickens).head()
# -
# We can also define custom logging functions by using `log_step_extra`. This takes any number of functions (> 1) that can take the output dataframe and return some output that can be converted to a string. For example, if we want to log some arbitrary message and the number of unique chicks in our dataset, we can do:
# + tags=[]
from sklego.pandas_utils import log_step_extra
def count_unique_chicks(df, **kwargs):
return "nchicks=" + str(df["chick"].nunique())
def display_message(df, msg):
return msg
@log_step_extra(count_unique_chicks)
def start_pipe(df):
"""Get initial chick count"""
return df
@log_step_extra(count_unique_chicks, display_message, msg="without diet 1")
def remove_diet_1_chicks(df):
return df.loc[df["diet"] != 1]
chickweight.pipe(start_pipe).pipe(remove_diet_1_chicks).head()
# -
|
doc/pandas_pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="xnyZ2Wv8aVEp" colab_type="text"
# # Introduction
# + [markdown] id="sdZg5zTMaVEs" colab_type="text"
# GANs' potential is huge because they can learn to mimic any distribution of data, including speech, images, prose, etc.. GAN basically consists of two kinds of algorithms, generative and discriminative ones. The discriminative algorithm predicts a category (or label) which the feature belongs. On the contrast, the generative algorithm predicts the features given a label. These two opposite algorithms are reasons why the network's name is adversarial.
#
# For example, we train a discriminative model to recognize whether an email (content) is spam or not. That is, the discriminative model calculates a probability $p\{y|x\}$, which $y$ is the category result (Yes or No) and $x$ is the word. The probability means how possible the spam is when given a word $x$. On the other hand, the generative model cares about **how to get $x$** when given a category $y$. That is, the generative model tries to learn the key feature consisting of $y$. The generative model is also a classifier.
#
# In short, the discriminative model learns the boundary between classes and the generative model learns the distribution of individual classes. In simple, you can think of GAN as the combination of a counterfeiter and a cop in a game of cat and mouse.
# + [markdown] id="xMa2f82OaVEt" colab_type="text"
# # How GANs Work
# + [markdown] id="liM-mzrMaVEu" colab_type="text"
# The generative neural network (called generator) generates new data instances, while the other discriminative one evaluates them for authenticity. For example, the discriminative decides whether each instance of data it reviews belongs to the actual training dataset or not.
#
# In the following, we are going to use MINST dataset as the example. The generator creates new images that pass to the discriminator and it hopes those images are deemed authentic (but the truth is the generating images are fake). On the other hand, the discriminator learns to identify images from the generator as fake.
# + [markdown] id="VHNsi3DMaVEv" colab_type="text"
# ## Basic Workflow
# + [markdown] id="fzMS0CpNaVEx" colab_type="text"
# * The generator takes in random numbers and returns an image.
# * Both the generated image is fed into the discriminator alongside a stream of the images from the actual dataset.
# * The discriminator identifies both real and generative images and returns probabilities between 0 (fake) and 1 (authentic).
# + [markdown] id="p_CBfFb4aVEy" colab_type="text"
# 
#
# The above image is referred from https://skymind.ai/wiki/generative-adversarial-network-gan and is further referred from O'Reily.
# + [markdown] id="tjV4pNVeaVEz" colab_type="text"
# ## Neural Network Design
# + [markdown] id="6sh_FBwSaVE1" colab_type="text"
# The discriminator network is a standard convolutional network recognizing the images fed to it. It is a binary classifier labelling images as real or fake. The generator is an inverse convolutional network. A standard convolutional network takes an image as input and downsamples it to produce a probability. On the contrary, an inverse convolutional network takes a vector of random noises and upsamples it to an image.
#
# Both networks are trying to optimize different objective functions (or loss functions), in a zero-sum game.
#
# 
#
# The above image is referred from https://skymind.ai/wiki/generative-adversarial-network-gan.
# + [markdown] id="Qaw_t896aVE2" colab_type="text"
# # Training GAN Tips
# + [markdown] id="jEpPbErLaVE4" colab_type="text"
# * **Hold the Adversary in the constant**
#
# When you train the discriminator, hold the generator values constant; and when you train the generator, hold the discriminator values constant. This gives one a better read on the gradient it must learn by.
#
# * **Pretrain the discriminator first**
#
# Pretraining the discriminator first before pretraining the generator gives a clear gradient.
#
# * **the balance is necessary**
#
# Each side of the GAN can overpower the other one. If the discriminator is too good, it will return values closing to 0 or 1. This gives the generator little gradient changes to update. If the generator is too good, it will persistently exploit the weakness to the discriminator and leads to increase false negatives (or type II error).
#
# * **more training time**
# + [markdown] id="yzMlI88faVE5" colab_type="text"
# # MNIST example in Keras
# + id="kjo5aSLoaVE6" colab_type="code" colab={}
import keras
from keras.models import Sequential, Model
from keras.layers import Dense, LeakyReLU, BatchNormalization, Reshape, Flatten, Input
from keras.optimizers import Adam
from keras.datasets import mnist
import numpy as np
import matplotlib.pyplot as plt
# + id="GizqDd6oaVE_" colab_type="code" colab={}
class GAN():
def __init__(self):
self.img_rows = 28
self.img_cols = 28
self.channels = 1
self.img_shape = (self.img_rows, self.img_cols, self.channels)
optimizer = Adam(0.0002, 0.5)
# Build and compile the discriminator
self.discriminator = self.build_discriminator()
self.discriminator.compile(loss='binary_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# Build and compile the generator
self.generator = self.build_generator()
#self.generator.compile(loss='binary_crossentropy', optimizer=optimizer)
# The generator takes noise as input and generated imgs
z = Input(shape=(100,))
img = self.generator(z)
# For the combined model we will only train the generator
self.discriminator.trainable = False
# The valid takes generated images as input and determines validity
valid = self.discriminator(img)
# The combined model (stacked generator and discriminator) takes
# noise as input => generates images => determines validity
self.combined = Model(inputs=z, outputs=valid)
self.combined.compile(loss='binary_crossentropy', optimizer=optimizer)
def build_discriminator(self):
img_shape = (self.img_rows, self.img_cols, self.channels)
model = Sequential()
model.add(Flatten(input_shape=img_shape))
model.add(Dense(512))
model.add(LeakyReLU(alpha=0.2))
model.add(Dense(256))
model.add(LeakyReLU(alpha=0.2))
model.add(Dense(1, activation='sigmoid'))
model.summary()
img = Input(shape=img_shape)
validity = model(img)
return Model(img, validity)
def build_generator(self):
noise_shape = (100,)
model = Sequential()
model.add(Dense(256, input_shape=noise_shape))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(512))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(1024))
model.add(LeakyReLU(alpha=0.2))
model.add(BatchNormalization(momentum=0.8))
model.add(Dense(np.prod(self.img_shape), activation='tanh'))
model.add(Reshape(self.img_shape))
model.summary()
noise = Input(shape=noise_shape)
img = model(noise)
return Model(noise, img)
def train(self, epochs, batch_size=128, save_interval=50):
# Load the dataset
(X_train, _), (_, _) = mnist.load_data()
# Rescale -1 to 1
X_train = (X_train.astype(np.float32) - 127.5) / 127.5
X_train = np.expand_dims(X_train, axis=3)
half_batch = int(batch_size / 2)
for epoch in range(epochs):
# ---------------------
# Train Discriminator
# ---------------------
# Select a random half batch of images
idx = np.random.randint(0, X_train.shape[0], half_batch)
imgs = X_train[idx]
noise = np.random.normal(0, 1, (half_batch, 100))
# Generate a half batch of new images
gen_imgs = self.generator.predict(noise)
# Train the discriminator
d_loss_real = self.discriminator.train_on_batch(imgs, np.ones((half_batch, 1)))
d_loss_fake = self.discriminator.train_on_batch(gen_imgs, np.zeros((half_batch, 1)))
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# ---------------------
# Train Generator
# ---------------------
noise = np.random.normal(0, 1, (batch_size, 100))
# The generator wants the discriminator to label the generated samples
# as valid (ones)
valid_y = np.array([1] * batch_size)
# Train the generator
g_loss = self.combined.train_on_batch(noise, valid_y)
# If at save interval => save generated image samples
if epoch % save_interval == 0:
# Plot the progress
print ("%d [D loss: %f, acc.: %.2f%%] [G loss: %f]" % (epoch, d_loss[0], 100*d_loss[1], g_loss))
self.save_imgs(epoch)
def save_imgs(self, epoch):
r, c = 5, 5
noise = np.random.normal(0, 1, (r * c, 100))
gen_imgs = self.generator.predict(noise)
# Rescale images 0 - 1
gen_imgs = 0.5 * gen_imgs + 0.5
fig, axs = plt.subplots(r, c)
cnt = 0
for i in range(r):
for j in range(c):
axs[i,j].imshow(gen_imgs[cnt, :,:,0], cmap='gray')
axs[i,j].axis('off')
cnt += 1
plt.show()
def get_discriminator(self):
return self.discriminator
def get_generator(self):
return self.generator
# + id="YW-3DWTOaVFD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4ae144c4-c36c-48e4-8aca-5a9998e848e8" executionInfo={"status": "ok", "timestamp": 1581416707293, "user_tz": -480, "elapsed": 759671, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
if __name__ == '__main__':
gan = GAN()
gan.train(epochs=50001, batch_size=64, save_interval=5000)
# + [markdown] id="kZBgM8jJyJMd" colab_type="text"
# # Inference with SubModels
# + id="DegaUaOsaVFH" colab_type="code" colab={}
dis = gan.get_discriminator()
gen = gan.get_generator()
# + id="CxMIkY70jbAM" colab_type="code" colab={}
(_, _), (test_images, test_labels) = mnist.load_data()
# + id="GyhhRcZRjeYQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3706f4e5-9b48-4de2-ffa9-30b8335e43ca" executionInfo={"status": "ok", "timestamp": 1581416714878, "user_tz": -480, "elapsed": 1091, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
test_imgs = (test_images.astype(np.float32) - 127.5) / 127.5
test_imgs.shape
# + id="vJBSKVb5kIXi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9cd34ac2-9163-4076-b6ea-c3005a933197" executionInfo={"status": "ok", "timestamp": 1581416717883, "user_tz": -480, "elapsed": 2281, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
dis.inputs
# + id="YIu_SaMLkMmi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6a953c91-5d6a-45d3-93e3-ec6e16bb5d18" executionInfo={"status": "ok", "timestamp": 1581416720588, "user_tz": -480, "elapsed": 2024, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
len(np.where(dis.predict(np.expand_dims(test_imgs, axis=-1)) > 0.5)[0])
# + id="JWBzkuWSuN-u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="25b9fbce-7136-4d94-d86d-17eecef61c0f" executionInfo={"status": "ok", "timestamp": 1581416722453, "user_tz": -480, "elapsed": 1192, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
gen.inputs, gen.outputs
# + id="autDhzLFpuvZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="57dced3e-97c7-4cf3-c1ab-736a0d433f40" executionInfo={"status": "ok", "timestamp": 1581416736286, "user_tz": -480, "elapsed": 1075, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
genout = gen.predict(np.random.normal(0, 1, (1, 100)))
plt.imshow(keras.preprocessing.image.array_to_img(genout[0]), cmap=plt.cm.gray)
plt.axis("off")
plt.show()
# + id="P6tUlYx-uJCp" colab_type="code" colab={}
|
deep_learning/generative/Keras_SimpleGAN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Voting
# Why we need voting?
# - Equally treat each model, use 'voting' method to get the final result
# - Reduce the risk of simply employing one model?
# - For most cases, the variance will be smaller after voting
#
# ## Voting in Regression Model
# The average predicted value of all the models. (Without assigning weights on models)
# ## Voting in Classification Model
#
# Count what appeared 'most'. 'Most' has different definitions among hard and soft voting context.
#
# ### Hard Voting:
# Simply consider the number of labels that are predicted most.
#
# For example, 5 models, label A and B, if 3 models predicted B, the result from hard voting will be B.
#
# ### Soft Voting:
# Soft voting sum the probability from each model for each label.
# For example, label A and B, model 1: A: 80%, B: 20%; model 2: A: 65%, B: 35%; model 3: A: 10%, B: 90%
#
# We have category A: 80% + 65% + 10% = 155%
# category B: 20% + 35% + 90% = 145%
#
# Using soft voting, we will go for the prediction of A
#
# ### Comment on Soft and Hard Voting:
# soft and hard voting may generate completely different results. Soft voting generally generate more accurate result for considering the 'probability'.
#
# ## What should I consider when using voting method?
#
# ### Models
# - There shouldn't be significant difference among models performance, otherwise there will be noise in the ensembled model.
#
# - Voting among a linear model and a tree model generally will be better than voting among linear models or among tree models
#
# ### When to use hard/soft voting?
#
# - Hard voting: when there are only clear label of the prediction result
# - Soft voting: when 'score' such as probability or prediction score are available in the model (SVM, KNN, Decision Tree)
# ## Comment on Voting
# Voting treat each model EQUALLY. When a model perform well under one circumstance, and perform poor under another, voting may not be the best method to ensenble models.
|
Task7.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="iPpI7RaYoZuE"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" id="hro2InpHobKk"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="U9i2Dsh-ziXr"
# # 즉시 실행 기초
# + [markdown] id="Hndw-YcxoOJK"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/r1/tutorials/eager/eager_basics.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/r1/tutorials/eager/eager_basics.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
# </td>
# </table>
# + [markdown] id="D3iRBJ-xxfU3"
# Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도
# 불구하고 [공식 영문 문서](https://www.tensorflow.org/?hl=en)의 내용과 일치하지 않을 수 있습니다.
# 이 번역에 개선할 부분이 있다면
# [tensorflow/docs](https://github.com/tensorflow/docs) 깃헙 저장소로 풀 리퀘스트를 보내주시기 바랍니다.
# 문서 번역이나 리뷰에 참여하려면
# [<EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ko)로
# 메일을 보내주시기 바랍니다.
# + [markdown] id="6sILUVbHoSgH"
# 이 노트북은 텐서플로를 사용하기 위한 입문 튜토리얼입니다. 다음 내용을 다룹니다 :
#
# * 필요한 패키지 임포트
# * 텐서(Tensor) 생성 및 사용
# * GPU 가속기 사용
# * 데이터 세트
# + [markdown] id="z1JcS5iBXMRO"
# ## 텐서플로 임포트
#
# 시작하기 위해서 텐서플로 모듈을 임포트하고 즉시 실행(eager execution)을 활성화합니다. 즉시 실행 활성화로 텐서플로를 조금 더 대화형 프론트엔드(frontend)에 가깝게 만들어 줍니다. 세부사항은 나중에 이야기할 것입니다.
# + cellView="code" id="RlIWhyeLoYnG"
import tensorflow.compat.v1 as tf
# + [markdown] id="H9UySOPLXdaw"
# ## 텐서
#
# 텐서는 다차원 배열입니다. 넘파이(NumPy) `ndarray` 객체와 비슷하며, `Tensor` 객체는 데이터 타입과 크기를 가지고 있습니다. 또한 텐서는 GPU 같은 가속기 메모리에 상주할 수 있습니다. 텐서플로는 텐서를 생성하고 이용하는 풍부한 연산 라이브러리([tf.add](https://www.tensorflow.org/api_docs/python/tf/add), [tf.matmul](https://www.tensorflow.org/api_docs/python/tf/matmul), [tf.linalg.inv](https://www.tensorflow.org/api_docs/python/tf/linalg/inv) etc.)를 제공합니다. 이러한 연산자는 자동적으로 순수 파이썬 타입을 변환합니다. 예를 들어:
#
# + cellView="code" id="ngUe237Wt48W"
print(tf.add(1, 2))
print(tf.add([1, 2], [3, 4]))
print(tf.square(5))
print(tf.reduce_sum([1, 2, 3]))
print(tf.encode_base64("hello world"))
# 연산자의 오버로딩(overloding) 또한 지원합니다.
print(tf.square(2) + tf.square(3))
# + [markdown] id="IDY4WsYRhP81"
# 각각의 텐서는 크기와 데이터 타입을 가지고 있습니다.
# + id="srYWH1MdJNG7"
x = tf.matmul([[1]], [[2, 3]])
print(x.shape)
print(x.dtype)
# + [markdown] id="eBPw8e8vrsom"
# 넘파이 배열과 텐서플로 텐서의 가장 확연한 차이는 다음과 같습니다:
#
# 1. `텐서`는 가속기 메모리(GPU, TPU와 같은)에서 사용할 수 있습니다.
# 2. `텐서`는 불변성(immutable)을 가집니다.
# + [markdown] id="Dwi1tdW3JBw6"
# ### 넘파이 호환성
#
# 텐서와 넘파이 배열 사이의 변환은 다소 간단합니다.
#
# * 텐서플로 연산은 자동적으로 넘파이 배열을 텐서로 변환합니다.
# * 넘파이 연산은 자동적으로 텐서를 넘파이 배열로 변환합니다.
#
# 텐서는 `.numpy()` 메서드(method)를 호출하여 넘파이 배열로 변환할 수 있습니다.
# 가능한 경우, 텐서와 배열은 메모리 표현을 공유하기 때문에 이러한 변환은 일반적으로 간단(저렴)합니다. 그러나 텐서는 GPU 메모리에 저장될 수 있고, 넘파이 배열은 항상 호스트 메모리에 저장되므로, 이러한 변환이 항상 가능한 것은 아닙니다. 따라서 GPU에서 호스트 메모리로의 복사가 필요합니다.
# + id="lCUWzso6mbqR"
import numpy as np
ndarray = np.ones([3, 3])
print("텐서플로 연산은 자동적으로 넘파이 배열을 텐서로 변환합니다.")
tensor = tf.multiply(ndarray, 42)
print(tensor)
print("그리고 넘파이 연산은 자동적으로 텐서를 넘파이 배열로 변환합니다.")
print(np.add(tensor, 1))
print(".numpy() 메서드는 텐서를 넘파이 배열로 변환합니다.")
print(tensor.numpy())
# + [markdown] id="PBNP8yTRfu_X"
# ## GPU 가속기
#
# 대부분의 텐서플로 연산은 GPU를 사용하여 가속화할 수 있습니다. 어떠한 주석(annotation)도 없이, 텐서플로는 연산을 위해 자동적으로 CPU 또는 GPU를 사용할 것인지를 정합니다(그리고 필요시 텐서를 CPU 와 GPU에 복사합니다.) 연산에 의해 생성된 텐서는 전형적으로 연산이 실행된 장치의 메모리에 의해 실행됩니다. 예를 들어:
# + cellView="code" id="3Twf_Rw-gQFM"
x = tf.random_uniform([3, 3])
print("GPU 사용이 가능한가 : "),
print(tf.test.is_gpu_available())
print("텐서가 GPU #0에 있는가 : "),
print(x.device.endswith('GPU:0'))
# + [markdown] id="vpgYzgVXW2Ud"
# ### 장치 이름
#
# `Tensor.device`는 텐서를 구성하고 있는 호스트 장치의 풀네임을 제공합니다. 이러한 이름은 프로그램이 실행중인 호스트의 네트워크 주소 및 해당 호스트 내의 장치와 같은 많은 세부 정보를 인코딩하며, 이것은 텐서플로 프로그램의 분산 실행에 필요합니다. 텐서가 호스트의 `N`번째 GPU에 놓여지면 문자열은 `GPU:<N>`으로 끝납니다.
# + [markdown] id="ZWZQCimzuqyP"
# ### 명시적 장치 배치
#
# 텐서플로에서 "배치(replacement)"라는 용어는 개별 연산을 실행하기 위해 장치에 할당(배치) 하는 것입니다. 앞서 언급했듯이, 명시적 지침이 없을 경우 텐서플로는 연산을 실행하기 위한 장치를 자동으로 결정하고, 필요시 텐서를 장치에 복사합니다. 그러나 텐서플로 연산은 `tf.device`을 사용하여 특정한 장치에 명시적으로 배치할 수 있습니다.
# 예를 들어:
# + id="RjkNZTuauy-Q"
import time
def time_matmul(x):
start = time.time()
for loop in range(10):
tf.matmul(x, x)
result = time.time()-start
print("10 loops: {:0.2f}ms".format(1000*result))
# CPU에서 강제실행합니다.
print("On CPU:")
with tf.device("CPU:0"):
x = tf.random_uniform([1000, 1000])
assert x.device.endswith("CPU:0")
time_matmul(x)
# GPU #0가 이용가능시 GPU #0에서 강제실행합니다.
if tf.test.is_gpu_available():
with tf.device("GPU:0"): # 또는 GPU:1, GPU:2
x = tf.random_uniform([1000, 1000])
assert x.device.endswith("GPU:0")
time_matmul(x)
# + [markdown] id="o1K4dlhhHtQj"
# ## 데이터셋
#
# 이번 섹션에서는 모델에 데이터를 제공하기 위한 파이프라인을 구축하기 위해 [`tf.data.Dataset` API](https://www.tensorflow.org/r1/guide/datasets)를 시연해볼 것입니다. 이는 다음을 포함합니다.
#
# * 데이터셋 생성.
# * 즉시 실행 활성화를 통한 데이터셋 반복
#
# 모델을 훈련시키고 평가 루프를 제공할 간단하고 재사용 가능한 모듈로부터, 복잡한 입력 파이프라인을 구축하기위해 데이터셋 API를 사용하기를 권장합니다.
#
# 만약 텐서플로 그래프에 익숙하다면 알겠지만, 데이터셋 객체를 생성하기 위한 API는 즉시 실행이 활성화 되어도 동일하게 유지됩니다. 하지만 데이터셋의 요소를 반복하는 프로세스가 약간 더 간단해집니다.
# 또한 `tf.data.Dataset` 객체를 통하여 파이썬 반복문을 사용할 수 있으며, 명시적으로 `tf.data.Iterator` 객체를 생성할 필요가 없습니다.
# 그 결과, [텐서플로 가이드](https://www.tensorflow.org/r1/guide/datasets)의 반복자(iterator)에 관한 논의는 즉시 실행이 활성화될 때에는 신경 쓰지 않아도 됩니다.
# + [markdown] id="zI0fmOynH-Ne"
# ### 소스 Dataset 생성
#
# 굉장히 유용한 함수중 하나인 [`Dataset.from_tensors`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#from_tensors), [`Dataset.from_tensor_slices`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#from_tensor_slices)와 같은 팩토리(factory) 함수 중 하나를 사용하거나 파일로부터 읽어들이는 객체인 [`TextLineDataset`](https://www.tensorflow.org/api_docs/python/tf/data/TextLineDataset) 또는 [`TFRecordDataset`](https://www.tensorflow.org/api_docs/python/tf/data/TFRecordDataset)를 사용하여 소스 dataset을 생성하세요. 더 많은 정보를 위해서 [텐서플로 가이드](https://www.tensorflow.org/r1/guide/datasets#reading_input_data)를 참조하세요.
# + id="F04fVOHQIBiG"
ds_tensors = tf.data.Dataset.from_tensor_slices([1, 2, 3, 4, 5, 6])
# CSV 파일을 생성합니다.
import tempfile
_, filename = tempfile.mkstemp()
with open(filename, 'w') as f:
f.write("""Line 1
Line 2
Line 3
""")
ds_file = tf.data.TextLineDataset(filename)
# + [markdown] id="vbxIhC-5IPdf"
# ### 변환 적용
#
# [`맵(map)`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map), [`배치(batch)`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch), [`셔플(shuffle)`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#shuffle)과 같은 변환 함수를 사용하여 데이터셋의 레코드에 적용하세요. 세부사항은 [tf.data.Dataset을 위한 API 문서](https://www.tensorflow.org/api_docs/python/tf/data/Dataset)을 참조하세요.
# + id="uXSDZWE-ISsd"
ds_tensors = ds_tensors.map(tf.square).shuffle(2).batch(2)
ds_file = ds_file.batch(2)
# + [markdown] id="A8X1GNfoIZKJ"
# ### 반복
#
# 즉시 실행이 활성화되면 `Dataset` 객체는 반복이 가능합니다. 만약 텐서플로 그래프에서 데이터셋을 사용하는게 익숙하다면, `Dataset.make_one_shot_iterator()` 또는 `get_next()`와 같은 객체를 호출할 필요가 없는다는 것에 주목하세요.
# + id="ws-WKRk5Ic6-"
print('ds_tensors 요소:')
for x in ds_tensors:
print(x)
print('\nds_file 요소:')
for x in ds_file:
print(x)
|
site/ko/r1/tutorials/eager/eager_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import calendar
m, d, y = map(int, input().split())
i = calendar.weekday(y,m,d)
print(calendar.day_name[i].upper())
# -
|
hacker-rank/Python/Date and TIme/Calendar Module.ipynb
|
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Octave
% language: octave
% name: octave
% ---
% ## Trigonometric Fourier series examples
%
% **Example:** Find the compact trigonometric Fourier series for the exponential $e^{\tfrac{-t}{2}}$ over the interval $0 \leq t \leq \pi$
%
% **Solution**
%
% The fundamental frequency is $\omega_0=\frac{2\pi}{T_0}=\frac{2\pi}{\pi}=2$ rad/sec.
% $$g(t)\,=\,a_0\,+\,\sum_{n=1}^{\infty}\,a_n\,cos\,2nt\,+\,b_n\,sin\,2nt\;\;\;0\leq t\leq \pi$$
% $$a_0\:=\:\frac{1}{\pi}\:\int_{0}^{\pi}\:e^{\frac{-t}{2}}\:dt\:=\:0.504$$
% $$a_n\:=\:\frac{1}{\pi}\:\int_{0}^{\pi}\:e^{\frac{-t}{2}}\:\cos\,2nt\:dt\:=\:0.504\,\frac{2}{1+16n^2}$$
% $$b_n\:=\:\frac{1}{\pi}\:\int_{0}^{\pi}\:e^{\frac{-t}{2}}\:\sin\,2nt\:dt\:=\:0.504\,\frac{8n}{1+16n^2}$$
% $$g(t)\:=\:0.504\:[1\:+\:\sum_{n=1}^{\infty}\:\frac{2}{1+16n^2}\:(\cos 2nt+\:4n\,\sin 2nt)] \:\:\:, 0 \leq t \leq \pi$$
%
% For the compact Fourier series:
% $$C_0\,=\,a_0=\,0.504$$
% $$C_n\,=\,\sqrt{a_{n}^{2}+b_{n}^{2}}\,=\,0.504\,\sqrt{\frac{4}{(1+16n^2)^2}\frac{64n^2}{(1+16n^2)^2}}\,=\,0.504\,(\frac{2}{\sqrt{1+16n^2}})$$
% $$\theta _n=\tan ^{-1}(\frac{-b_n}{a_n})=tan ^{-1}(-4n)=-tan ^{-1}(4n) $$
%
% $$g(t)= 0.504+0.244\,\cos(2t-75.96^o)+0.125\,\cos(4t-82.42^o)\,+\,...\;\;\;,0\,\leq t\,\leq \pi$$
%
% ### Symbolic solution
%
% Using symbolic toolbox / package, you can find Fourier series coefficients
% +
pkg load symbolic % this loads symbolic package into octave
syms x n L k % x, n, L and k are the symbolic variables
evalin(symengine,'assume(n,Type::Integer)');
a = @(f,x,n,L) int(f*cos(n*2*pi*x/L)/L*2,x,0,L);
b = @(f,x,n,L) int(f*sin(n*2*pi*x/L)/L*2,x,0,L);
fs = @(f,x,k,L) a(f,x,0,L)/2 + symsum(a(f,x,n,L)*cos(n*2*pi*x/L) + b(f,x,n,L)*sin(n*2*pi*x/L),n,1,k);
f = exp(-x/2)
pretty(fs(f,x,2,pi))
% -
% ### FFT solution
% Using ```fft ``` command, Fourier sereis can be determined as follows
% +
% (c21.m)
% M is the number of coefficients to be computed
T0 = pi; N0 = 256; Ts = T0/N0; M = 10;
t = 0:Ts:Ts*(N0-1);
t = t';
g = exp(-t/2);
% fft(g) is the FFT
Dn = fft(g)/N0;
[Dnangle, Dnmag] = cart2pol(real(Dn), imag(Dn));
k = 0:length(Dn)-1;
k = k';
subplot(211), stem(k,fftshift(Dnmag))
subplot(212), stem(k,fftshift(Dnangle))
% +
C0 = Dnmag(1);
Cn = 2 * Dnmag(2:M);
Amplitudes = [C0; Cn];
Angles = Dnangle(1:M) * (180/pi);
disp('Amplitudes Angles')
[Amplitudes Angles]
% To plot the Fourier coefficients
k = 0 : length(Amplitudes)-1;
k = k';
subplot(211), stem(k,Amplitudes)
subplot(212), stem(k,Angles)
% -
% ### Fourier Synthesis
% Fourier synthesis is the opearion of rebuilding a signal from its harmonics.
% +
% This program synthesizes Fourier series to yield the original function
T0 = pi; % periodic time
w0 = 2; %
t = 0:T0/256:2*T0; % create a time vector of two cycles length
i = 10; % number of iterations (harmonics)
c0 = 0.504;
cn_sum = 0;
for n = 1:i
cn = 0.504*(2 / sqrt(1 + 16*n^2)) * cos(2*n.*t - atan(4*n) );
cn_sum = cn_sum + cn;
end
g = c0 + cn_sum; % synthesized signal
f = [exp(-t(1:end/2)/2) exp(-t(1:end/2+1)/2)]; % Original function with two cycles
% figure
plot(t,g); grid on;
hold on;
plot(t,f);
hold off
% -
|
Fourier_series.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About Dataset aggregate_covid_country.csv
# + active=""
# The dataset consists of 23688 rows and 5 features as explained below:
# 1. Date: date of occurance of cases.
# 2. Country: country in which cases occurs.
# 3. Confirmed: number of confirmed cases.
# 4. Recovered: number of recovered cases.
# 5. Deaths: number of deaths occur.
# Since the target variable Deaths is of continueous form, therefore it is a regression problem.
# -
# # Loading Dataset
import pandas as pd
df_covid = pd.read_csv('aggregate_covid_country.csv')
df_covid.head() #Printing first 5 rows
#Checking dimesion
df_covid.shape
# # Preparing Dataset
#Checking dataset information
df_covid.info()
# + active=""
# Remarks:
# -Total number of rows are 23688 and columns are 5.
# -3 Numerical and 2 Categorical features.
# -No null values are present.
# -
#Displaying null values using heatmap
import matplotlib.pyplot as plt
import seaborn as sns
sns.heatmap(df_covid.isnull())
plt.title('Heatmap for null values in dataset')
plt.show()
# + active=""
# Remarks:
# -No null values present.
# -
# # Label Encoding
#Checking unique values of categorical variables.
for i in df_covid.columns:
if df_covid[i].dtypes == object:
print(f'%-30s: {len(df_covid[i].unique())}'%i)
# + active=""
# Remarks:
# -Feature Date and Country both can be encoded as the unique values are fewer as compared to total records.
# -
#Appyling label encoding using LabelEncoder
from sklearn.preprocessing import LabelEncoder
le = {}
encoded_values = {}
for i in df_covid.columns:
if df_covid[i].dtypes == object:
le[i] = LabelEncoder()
encoded_values[i] = le[i].fit_transform(df_covid[i])
print(f"%-15s: {le[i].classes_}"%i)
# +
#Replacing the original values with encoded values into new dataframe
df = df_covid.copy()
for i in le.keys():
df[i] = encoded_values[i]
df.head() #Printing first 5 rows of new df
# -
# # Statisical Summary
#Checking statistical summary of dataset
df.describe()
# + active=""
# Remarks:
# -count is same for all features, no null value present.
# -mean is greater than median in Confirmed, Recovered and Deaths, data is right skewed.
# -Difference between 75th percentile and max is higher in Confirmed, Recovered and Deaths, outliers might be present.
# -
# # Exploratory Data Analysis (EDA)
#Checking Distribution of Data in features
rows = 3
cols = 2
fig, axes = plt.subplots(rows,cols,figsize=(rows*cols*3,rows*cols*3))
plt.subplots_adjust(hspace=0.5)
k=0
for i in range(rows):
for j in range(cols):
sns.distplot(df[df.columns[k]],ax=axes[i][j])
axes[i][j].set_title(f"Distribution Plot: {df.columns[k]}")
k = k+1
if k == 5:
break;
plt.show()
# + active=""
# Remarks:
# -Data is not distributed normally in any features.
# -
#Checking with box plot
rows = 3
cols = 2
fig, axes = plt.subplots(rows,cols,figsize=(rows*cols*3,rows*cols*3))
plt.subplots_adjust(hspace=0.5)
k=0
for i in range(rows):
for j in range(cols):
sns.boxplot(df[df.columns[k]],ax=axes[i][j])
axes[i][j].set_title(f"Distribution Plot: {df.columns[k]}")
k = k+1
if k == 5:
break;
plt.show()
# + active=""
# Remarks:
# -Feature Confirmed, Recovered and Deaths contains outliers.
# -
# # Bi-Variate Analysis
#Checking Feature Date and Deaths with Scatter Plot
x='Date'
y='Deaths'
sns.scatterplot(x,y,data=df)
plt.title(f"Scatter Plot: {x} vs {y}")
plt.show()
# + active=""
# Remarks:
# -As Date increases, Deaths also increases.
# -
#Checking Feature Country and Deaths with Scatter Plot
x='Country'
y='Deaths'
sns.scatterplot(x,y,data=df)
plt.title(f"Scatter Plot: {x} vs {y}")
plt.show()
# + active=""
# Remarks:
# -With certain exception Deaths rise in Country, almost other Country have same number of Deaths.
# -Country 174 has highest number of deaths.
# -
#Checking Feature Confirmed and Deaths with Scatter Plot
x='Confirmed'
y='Deaths'
sns.scatterplot(x,y,data=df)
plt.title(f"Scatter Plot: {x} vs {y}")
plt.show()
# + active=""
# Remarks:
# -As Confirmed increases, Deaths increases.
# -
#Checking Feature Recovered and Deaths with Scatter Plot
x='Recovered'
y='Deaths'
sns.scatterplot(x,y,data=df)
plt.title(f"Scatter Plot: {x} vs {y}")
plt.show()
# + active=""
# Remarks:
# -As the Recovered Increases, Deaths Decreases.
# -
# # Multi-Variate Analysis
# +
#Checking relationship between features
df_corr = df.corr()
#Displaying correlation of features using heatmap
sns.heatmap(df_corr,annot=True,fmt='.2f')
plt.show()
# + active=""
# Remarks:
# -Feature Confirmed and Recovered is highly correlated with Deaths.
# -Feature Date and Country is weakly correlated with Deaths.
# -
# # Removing Outliers
#Checking outliers with the help of zscore
from scipy.stats import zscore
import numpy as np
z = np.abs(zscore(df))
#Printing location of outliers
np.where(z>3)
#Removing outliers
df_wo = df[(z<=3).all(axis=1)]
print(f"Orignal Shape: {df.shape}")
print(f"New Shape: {df_wo.shape}")
print(f"% Loss: {(len(df)-len(df_wo))*100/len(df)}%")
# + active=""
# Remarks:
# -Since, loss of data is <5% therefore proceeding with data without outliers.
# -
# # Skewness
#Checking skewness of data
df.skew()
# + active=""
# Remarks:
# -Feature Confirmed, Recovered and Deaths have heavily skewed data and needs to be treated accordingly.
# -
# # Preparing Data for Model
#Seperating input and output variables
x = df.drop(columns=['Deaths'])
y = df['Deaths']
#Treating skewness of data
from sklearn.preprocessing import power_transform
x_t = power_transform(x)
#Scaling data for model using StandardScaler
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
scaled_x = sc.fit_transform(x_t)
# # Finding the Best Model
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
# +
#Defining function for best random_state
def get_best_rstate(r,model,x,y,test_size=0.25):
best_rState = 0
best_r2Score = 0
for i in r:
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=test_size,random_state=i)
model.fit(x_train,y_train)
predict_y = model.predict(x_test)
temp_r2Score = r2_score(y_test,predict_y)
if temp_r2Score>best_r2Score:
best_r2Score = temp_r2Score
best_rState = i
return best_rState,best_r2Score
#Defining function for best CV
def get_best_cv(model,parameters,x_train,y_train,r=range(2,20)):
best_cv = 0
best_cvScore = 0
for i in r:
gscv = GridSearchCV(model,parameters)
gscv.fit(x_train,y_train)
temp_cvScore = cross_val_score(gscv.best_estimator_,x_train,y_train,cv=i).mean()
if temp_cvScore>best_cvScore:
best_cvScore = temp_cvScore
best_cv = i
return best_cv,best_cvScore
#Defining function for building models
def build_model(models,x,y,r_range=range(100),t_size=0.25,cv_range=range(2,20)):
for i in models:
#Finding the best random_state for train test split
best_rState, best_r2Score = get_best_rstate(r_range,models[i]["name"],x,y)
#Splitting train test data with best random_state
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=t_size,random_state=best_rState)
#Hypertuning Parameters
#Finding best CV
best_cv, best_cvScore = get_best_cv(models[i]["name"],models[i]["parameters"],x_train,y_train,cv_range)
#Building final model with hypertuned parameters
gscv = GridSearchCV(models[i]["name"],models[i]["parameters"],cv=best_cv)
gscv.fit(x_train,y_train)
#Checking Final Performance of the model
predict_y = gscv.best_estimator_.predict(x_test)
r2Score = r2_score(y_test,predict_y)
mse = mean_squared_error(y_test,predict_y)
mae = mean_absolute_error(y_test,predict_y)
#Storing model specs.
models[i]["random_state"] = best_rState
models[i]["initial_r2_score"] = best_r2Score
models[i]["x_train"] = x_train
models[i]["x_test"] = x_test
models[i]["y_train"] = y_train
models[i]["y_test"] = y_test
models[i]["cv"] = best_cv
models[i]["cross_val_score"] = best_cvScore
models[i]["gscv"] = gscv
models[i]["predict_y"] = predict_y
models[i]["r2_score"] = r2Score
models[i]["mse"] = mse
models[i]["rmse"] = np.sqrt(mse)
models[i]["mae"] = mae
return models;
# -
# # Preparing List of Models and Testing for Best Performances
# +
import warnings
warnings.simplefilter('ignore')
from sklearn.linear_model import LinearRegression, Lasso, Ridge, SGDRegressor
from sklearn.ensemble import AdaBoostRegressor
#Preparing List of Models with parameters
models = {
"LinearRegression":{
"name": LinearRegression(),
"parameters":{
"fit_intercept":[True,False],
"normalize":[True,False]
}
},
"Lasso":{
"name": Lasso(),
"parameters":{
"alpha":[0.1,1.0],
"fit_intercept":[True,False],
"normalize":[True,False],
"selection":['cyclic','random']
}
},
"Ridge":{
"name": Ridge(),
"parameters":{
"alpha":[0.1,1.0],
"fit_intercept":[True,False],
"normalize":[True,False],
"solver":['auto','svd','cholesky','lsqr','sparse_cg','sag','saga']
}
},
"SGDRegressor":{
"name": SGDRegressor(),
"parameters":{
"loss":['squared_loss','huber','epsilon_insensitive','squared_epsilon_insensitive'],
"alpha":[0.00001,0.0001],
"shuffle":[True,False]
}
},
"AdaBoostRegressor":{
"name": AdaBoostRegressor(),
"parameters": {
"loss": ['linear','square','exponential']
}
}
}
#Building models
build_models = build_model(models,scaled_x,y)
# -
#Dispalying model performance
for i in build_models:
model = build_models[i]
print(f"START: {i}===================\n")
print(f"Best random_state: {model['random_state']} with best r2_score: {model['initial_r2_score']}\n")
print(f"Best CV: {model['cv']} with best cross_value_score: {model['cross_val_score']}\n")
print(f"Best params: {model['gscv'].best_params_}\n")
print(f"Final Performance:")
print(f"R2_SCORE: {round(model['r2_score']*100,2)}%\t MSE: {model['mse']}\t RMSE: {model['rmse']}\t MAE: {model['mae']}\n")
print(f"END: {i}=====================\n\n\n")
# + active=""
# Remarks:
# -From all tested model, AdaBoostRegressor performs well with an accuracy of 95.26% adn RMSE of 971.17, therefore,
# proceeding with AdaBoostRegressor
# -
# # Model Saving or Serialization
import joblib
final_model = build_models["AdaBoostRegressor"]
filename = "covid_19_project.pkl"
joblib.dump(final_model["gscv"].best_estimator_,open(filename,'wb'))
# # Conclusion
cnc = pd.DataFrame({"Original": np.array(final_model['y_test']),"Predicted": np.array(final_model['predict_y'])})
#Ploting using scatter plot
sns.scatterplot(x='Original',y='Predicted',data=cnc)
plt.title("Model Performance: AdaBoostRegressor")
plt.show()
# + active=""
# Remarks: Final model AdaBoostRegressor performs with an accuracy of 95.26% and RMSE 971.17, therefore, it can be improved further by training with more specific data.
|
covid-19-project/covid-19-project-code.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/scottwmwork/DS-Unit-2-Applied-Modeling/blob/master/module3/assignment_applied_modeling_3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nCc3XZEyG3XV" colab_type="text"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Applied Modeling, Module 3
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your work.
#
# - [ ] Continue to iterate on your project: data cleaning, exploration, feature engineering, modeling.
# - [ ] Make at least 1 partial dependence plot to explain your model.
# - [ ] Share at least 1 visualization on Slack.
#
# (If you have not yet completed an initial model yet for your portfolio project, then do today's assignment using your Tanzania Waterpumps model.)
#
# ## Stretch Goals
# - [ ] Make multiple PDPs with 1 feature in isolation.
# - [ ] Make multiple PDPs with 2 features in interaction.
# - [ ] If you log-transformed your regression target, then convert your PDP back to original units.
# - [ ] Use Plotly to make a 3D PDP.
# - [ ] Make PDPs with categorical feature(s). Use Ordinal Encoder, outside of a pipeline, to encode your data first. If there is a natural ordering, then take the time to encode it that way, instead of random integers. Then use the encoded data with pdpbox.I Get readable category names on your plot, instead of integer category codes.
#
# ## Links
# - [<NAME>: Interpretable Machine Learning — Partial Dependence Plots](https://christophm.github.io/interpretable-ml-book/pdp.html) + [animated explanation](https://twitter.com/ChristophMolnar/status/1066398522608635904)
# - [Kaggle / <NAME>: Machine Learning Explainability — Partial Dependence Plots](https://www.kaggle.com/dansbecker/partial-plots)
# - [Plotly: 3D PDP example](https://plot.ly/scikit-learn/plot-partial-dependence/#partial-dependence-of-house-value-on-median-age-and-average-occupancy)
# + id="-7OllAY-deRb" colab_type="code" outputId="9fe06b0a-3443-44f7-e9d5-23fcfe480688" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import sys
in_colab = 'google.colab' in sys.modules
if in_colab:
# Install packages in Colab
# !pip install pdpbox
# !pip install category_encoders==2.0.0
# !pip install pandas-profiling==2.3.0
# !pip install plotly==4.1.1
# + id="KQmiFQpd-av3" colab_type="code" colab={}
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/scottwmwork/datasets/master/tmdb_5000_movies.csv')
# + id="qWK807RONmgW" colab_type="code" outputId="ce91fffb-8c3b-474f-909a-5bef9129cd25" colab={"base_uri": "https://localhost:8080/", "height": 399}
df.isna().sum()
# + id="FiduBoSiEMOP" colab_type="code" colab={}
#Make Genres a compatible feature
X = df #Copy df into X (Doing this so I can easily copy the following code into my wrangle function)
import ast
import numpy as np
genre = []
for x in X['genres']:
if x == '[]':
genre.append(np.nan)
else:
temp = ast.literal_eval(x)
genre.append(temp[0]['name']) #grabs first genre in list of dictionaries
#create new column for dataframe
X['genre_first_listed'] = genre
# + id="HfX4QTt0PHUV" colab_type="code" outputId="c3e4b203-0116-4da2-dce2-684b4b6e464a" colab={"base_uri": "https://localhost:8080/", "height": 399}
X['genre_first_listed'].value_counts().unique
# + id="h82sp29YdTly" colab_type="code" colab={}
df['release_date'] = pd.to_datetime(df['release_date'],infer_datetime_format = True)
df['release_year'] = df['release_date'].dt.year
df['release_month'] = df['release_date'].dt.month
df['release_day'] = df['release_date'].dt.month
df = df.drop(columns = 'release_date')
# + id="fHIdHxE11Vwc" colab_type="code" colab={}
#Isolate the test set
test = df[df['release_year'] == 2016]
y_test = test['revenue']
X_test = test.drop(columns = 'revenue')
#Exclude test set from data
dfn = df[df['release_year'] != 2016]
# + id="PMOORiNCIr55" colab_type="code" colab={}
#Create train and validation data
from sklearn.model_selection import train_test_split
train, val = train_test_split(dfn, train_size = .80, test_size = 0.20, random_state = 42)
y_train = train.revenue
X_train = train.drop(columns = 'revenue')
y_val = val.revenue
X_val = val.drop(columns = 'revenue')
# + id="4waZ_vgWhPd7" colab_type="code" colab={}
import ast
import numpy as np
def wrangle(X):
X = X.copy()
X = X.reset_index()
#Make genres column usable
genre = []
for x in X['genres']:
if x == '[]':
genre.append(np.nan)
else:
temp = ast.literal_eval(x)
genre.append(temp[0]['name']) #grabs first genre in list of dictionaries
#Features to not include:
X = X.drop(columns = ['genres','homepage','keywords','overview','production_companies','production_countries','spoken_languages','tagline'])
#Engineer features:
#original title is same as title?
title_changed = []
for x in range(0,len(X['title'])):
if X['title'][x] == X['original_title'][x]:
title_changed.append(0)
else:
title_changed.append(1)
#length of title
length_of_title = []
for x in X['title']:
length_of_title.append(len(x))
#Add features to dataframe
X['title_changed'] = title_changed
X['length_of_title'] = length_of_title
X['genre_first_listed'] = genre
return X
# + id="5w2jQud1Effd" colab_type="code" colab={}
#Wrangle data
X_test = wrangle(X_test)
X_val = wrangle(X_val)
X_train = wrangle(X_train)
# + id="i-K4Qsh68_z9" colab_type="code" outputId="d99a8f0f-4af1-47d0-8a3e-9f2b25cc3b5b" colab={"base_uri": "https://localhost:8080/", "height": 454}
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestRegressor
import category_encoders as ce
from sklearn.tree import DecisionTreeClassifier
pipeline = make_pipeline(
# SimpleImputer(strategy = 'mean'),
ce.OneHotEncoder(use_cat_names = True),
SimpleImputer(),
# DecisionTreeClassifier(random_state = 5),
# LinearRegression()
# LogisticRegression()
RandomForestRegressor()
)
pipeline.fit(X_train,y_train)
# + id="icltzgdyGHT7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 582} outputId="2c95d3f7-9790-412c-fc30-ea4c6708b5c0"
from pdpbox.pdp import pdp_isolate, pdp_plot
feature = 'budget'
isolated = pdp_isolate(
model=pipeline,
dataset=X_val,
model_features=X_val.columns,
feature=feature
)
pdp_plot(isolated, feature_name=feature);
# + id="Veotngu9Rhfj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 582} outputId="f6dfff8b-7a21-46c1-e4aa-a1495a1ae2fb"
pdp_plot(isolated, feature_name=feature, plot_lines=True, frac_to_plot=100) # Plot 100 ICE lines
plt.xlim(0,30000000);
|
module3/assignment_applied_modeling_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # tempfile -- Generate temporary files and directories
# https://docs.python.org/3/library/tempfile.html
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Create-and-delete-a-temporary-file-with-a-context-manager" data-toc-modified-id="Create-and-delete-a-temporary-file-with-a-context-manager-1"><span class="toc-item-num">1 </span>Create and delete a temporary file with a context manager</a></span><ul class="toc-item"><li><span><a href="#Files-are-opened-in-binary-mode-by-default" data-toc-modified-id="Files-are-opened-in-binary-mode-by-default-1.1"><span class="toc-item-num">1.1 </span>Files are opened in binary mode by default</a></span></li><li><span><a href="#Opening-a-temporary-file-in-text-mode" data-toc-modified-id="Opening-a-temporary-file-in-text-mode-1.2"><span class="toc-item-num">1.2 </span>Opening a temporary file in text mode</a></span></li><li><span><a href="#Creating-a-temporary-file-with-a-visible-name" data-toc-modified-id="Creating-a-temporary-file-with-a-visible-name-1.3"><span class="toc-item-num">1.3 </span>Creating a temporary file with a visible name</a></span></li><li><span><a href="#Keeping-a-named-temporary-file-with-delete=False" data-toc-modified-id="Keeping-a-named-temporary-file-with-delete=False-1.4"><span class="toc-item-num">1.4 </span>Keeping a named temporary file with <code>delete=False</code></a></span></li></ul></li><li><span><a href="#Creating-a-temporary-directory" data-toc-modified-id="Creating-a-temporary-directory-2"><span class="toc-item-num">2 </span>Creating a temporary directory</a></span></li></ul></div>
# -
import tempfile
# ## Create and delete a temporary file with a context manager
# ### Files are opened in binary mode by default
with tempfile.TemporaryFile() as binary_temp_file:
binary_temp_file.write(b"Hello")
binary_temp_file.seek(0)
print(binary_temp_file.read())
# ### Opening a temporary file in text mode
with tempfile.TemporaryFile(mode="w+") as text_temp_file:
print("Hello", file=text_temp_file)
text_temp_file.seek(0)
print(text_temp_file.read())
# ### Creating a temporary file with a visible name
# +
import pathlib
with tempfile.NamedTemporaryFile() as named_temp_file:
named_temp_file.write("Hello world!\n".encode())
named_temp_file.flush()
file_path = pathlib.Path(named_temp_file.name)
assert file_path.exists()
with open(file_path, "r") as read_file:
print(read_file.read())
# Now the file is gone
assert not file_path.exists()
# -
# ### Keeping a named temporary file with `delete=False`
# +
with tempfile.NamedTemporaryFile(delete=False) as named_temp_file:
named_temp_file.write("Hello world!\n".encode())
file_path = pathlib.Path(named_temp_file.name)
assert file_path.exists()
# The file is still there
assert file_path.exists()
file_path.unlink()
assert not file_path.exists()
# -
# ## Creating a temporary directory
# +
with tempfile.TemporaryDirectory() as temp_dir:
assert isinstance(temp_dir, str)
path = pathlib.Path(temp_dir)
assert path.exists()
assert path.is_dir()
assert not path.exists()
assert not path.is_dir()
|
notes/2019/2019-11-27-python-stdlib-tempfile/tempfile.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import pickle
from keras.applications.vgg16 import VGG16, preprocess_input
from keras.preprocessing.image import load_img, img_to_array
from keras.models import Model
from PIL import Image
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
smpl_path = "../data/yelp_photos/photos/ZzZDx9p-AbZiMcl1OPyR-g.jpg"
Image.open(smpl_path)
img = load_img(smpl_path, target_size = (224,224))
img = img_to_array(img)
img = img.reshape((1, img.shape[0], img.shape[1], img.shape[2]))
img = preprocess_input(img)
model = VGG16()
layer_outputs = [layer.output for layer in model.layers]
activation_model = Model(inputs=model.input, outputs=layer_outputs)
activations = activation_model.predict(img)
def display_activation(activations, col_size, row_size, act_index):
activation = activations[act_index]
activation_index=0
fig, ax = plt.subplots(row_size, col_size, figsize=(row_size*2.5,col_size*1.5))
for row in range(0,row_size):
for col in range(0,col_size):
ax[row][col].imshow(activation[0, :, :, activation_index], cmap='gray')
activation_index += 1
activations[2].shape
def save_activations(activations, act_index, path):
activation = activations[act_index][0]
i = 0
for i in range(activation.shape[-1]):
fname = "%s/activation%d_pic%d.jpg"%(path,act_index,i)
plt.imsave(fname,activation[:,:,i],cmap = 'gray')
i += 1
display_activation(activations,8,8,1)
display_activation(activations,8,8,2)
display_activation(activations,16,8,4)
display_activation(activations,16,8,5)
# mkdir activation_viz/vgg_act0
save_activations(activations,0,"vgg_act0")
# mkdir activation_viz/vgg_act1
save_activations(activations,1,"activation_viz/vgg_act1")
# mkdir activation_viz/vgg_act4
save_activations(activations,4,"activation_viz/vgg_act4")
model.summary()
|
captioning/evaluation/Report-Visualizations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Mearoxas/CPEN-21A-ECE-2-1/blob/main/Loop_Statement.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8OegITMD0jTn"
# ## Application 1
# + colab={"base_uri": "https://localhost:8080/"} id="rR6UpQ6c0rAG" outputId="17910d4b-11cf-4efe-a0db-fd3e3d9ecb8d"
value = ["Hello Value"]
number = ["0","1", "2", "3","4", "5", "6","7","8", "9","10"]
i=0
while i<=10:
print("Hello Value",i)
i+=1 #Assignment operator for addition
# + [markdown] id="v-kQrtlB3RcO"
# ## Application 2
# + colab={"base_uri": "https://localhost:8080/"} id="mjzNbZ5m3WIC" outputId="58c94536-c2a7-4d31-fd71-feaca2c91294"
i=3
while i<10:
print(i)
i+=1 #Assignment operator for addition
|
Loop_Statement.ipynb
|
# ---
# jupyter:
# anaconda-cloud: {}
# jupytext:
# cell_metadata_filter: all
# notebook_metadata_filter: all,-language_info,-toc,-latex_envs
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %%
data = np.loadtxt('/home/sallen/MEOPAR/grid/bathymetry_202108.csv', delimiter=',')
#data = np.loadtxt('/Users/sallen/Documents/MEOPAR/grid/bathymetry_201702.csv', delimiter=',')
# %%
square = data[:, 1]/data[:, 0] - 9/4.
total = data[:, 1]*data[:, 0]
efficient = total/data[:, 2]
# %%
plt.plot(data[:, 2], total, 'o')
guide = 40*7
plt.xlim((guide-40, guide))
plt.grid()
plt.ylim((400, 800))
print (guide)
# %%
for i, datum in enumerate(data[:]):
if (datum[2] > guide-40*7 and datum[2] < guide):
# if (datum[2] == 335):
# if (efficient[i] > 1.9):
if (abs(square[i]) < 1):
if (total[i] >=540):
print(datum, square[i], efficient[i], total[i])
# %% [markdown]
# # 201702
# [ 15. 36. 279.] 0.1499999999999999 1.935483870967742 540.0
# [ 16. 34. 278.] -0.125 1.9568345323741008 544.0
# [ 18. 30. 276.] -0.5833333333333333 1.9565217391304348 540.0
# %% [markdown]
# * One : 4 x 9 : 30 for 36 : exactly square
# * Two : 6 x 14 : 62 for 84 : 0.08 from square
# * Three: 8 x 18 : 95 for 144 : exactly square
# * Four: 11 x 20 : 127 for 220 : 0.43 - 125 for 201803p
# * 9 x 22 : 126 for 198 : 0.19
# * Five: 12 x 23 : 157 for 276 : 0.33 - 156 for 201803p
# * 11 x 25 : 156 for 275 : 0.02
# * Six: 13 x 26 : 191 for 338 : 0.25
# * 12 x 28 : 190 for 336 : 0.08
# * Seven: 15 x 28 : 222 for 420 : 0.38
# * 14 x 29 : 219 for 406: 0.18
# * Eight: 15 x 32 : 254 for 480: 0.12
# * Nine: 15 x 37: 286 for 555 : 0.21
# * 16 x 34: 278 for 544 : 0.12
# * Ten : 16 x 39 = 317 for 624 : 0.19
# %% collapsed=true jupyter={"outputs_hidden": true}
[ 11. 33. 198.] 0.75 1.83333333333 363.0
# %% [markdown]
# 14X29 + 5 = 224 equiv 406 or 14X28 + 7 = 392
# 11X30 + 6 = 192 equiv 275
# 11X25 + 4 = 160 equiv 330
|
notebooks/Decomposition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monte Carlo Tree Search
#
# This notebook explores the game ConnectFour and MCTSPlayer in the aima3 collection.
#
# * You drop pieces from the top into one of the columns, and they land in the next available open spot.
# * You must get 4 positions in a row in this grid, either horizontally, vertically, or diagonally.
#
#
# - | 1 | 2 | 3 | 4 | 5 | 6 | 7
# --------|---|---|---|---|---|---|---
# **6** |( 1, 6) | ( 2, 6) | ( 3, 6) | (4,6) |(5,6) |(6,6) |(7,6)
# **5** |( 1, 5) | ( 2, 5) | ( 3, 5) | (4,5) |(5,5) |(6,5) |(7,5)
# **4** |( 1, 4) | ( 2, 4) | ( 3, 4) | (4,4) |(5,4) |(6,4) |(7,4)
# **3** |( 1, 3) | ( 2, 3) | ( 3, 3) | (4,3) |(5,3) |(6,3) |(7,3)
# **2** |( 1, 2) | ( 2, 2) | ( 3, 2) | (4,2) |(5,2) |(6,2) |(7,2)
# **1** |( 1, 1) | ( 2, 1) | ( 3, 1) | (4,1) |(5,1) |(6,1) |(7,1)
# There are a few pre-defined AI agents that can play these games. QueryPlayer is for humans, and MCTS is MonteCarloTreeSearch. The other are well-known search-based algorithms.
from aima3.games import (ConnectFour, RandomPlayer, QueryPlayer, players,
MCTSPlayer, MiniMaxPlayer, AlphaBetaCutoffPlayer,
AlphaBetaPlayer, GameState)
# Let's play a game:
p1 = AlphaBetaCutoffPlayer()
p2 = MCTSPlayer(n_playout=150)
p3 = RandomPlayer()
game = ConnectFour()
game.play_tournament(10, p1, p2, p3)
# To reset the tree search to get ready for a fresh game (and no stats), do this:
p2.mcts.update_with_move(-1)
# Now, let's run a get_move_probs(). This will simulate mcts.n_playout moves. Currently, playout is set to:
p2.mcts.n_playout
# So, 150 simulated steps from leafs in the tree for each move. We get back those moves, and their probabilities:
p2.mcts.get_move_probs(game.initial)
# To see these a bit more clearly, here is a little function:
def pp(moves_probs):
moves = moves_probs[0]
probs = moves_probs[1]
for i in range(len(moves)):
print("%7s" % (moves[i],), end=" | ")
print()
for i in range(len(probs)):
print("%7.3s" % (probs[i],), end=" | ")
print()
pp(p2.mcts.get_move_probs(game.initial))
# The above ran an additional 20 simulated moves in the game.
#
# Let's set up some board states, and see what MCTS rates them:
state = game.initial
state = game.result(state, (4,1))
game.display(state)
# We could step through a game like that, or just poke in the right values for a state:
moves = game.initial.moves[:]
moves.remove((1,1))
moves.remove((2,1))
moves.remove((7,1))
moves.remove((7,2))
state = GameState(to_move='X', board={(1,1): 'X', (2, 1): 'X',
(7,1): 'O', (7, 2): 'O'}, utility=0, moves=moves)
state = game.result(state, (3,1))
game.display(state)
s1 = game.result(game.initial, (1,1))
s1
game.display(s1)
state = game.string_to_state("""
. . . . . . .
. . . . . . .
. . . . . . .
. . . . . . O
. . . . . . O
X X X X . . O
""", to_move='X')
game.display(state)
game.actions(state)
# Looks good! Let's evaluate it. First, we reset:
p2.mcts.update_with_move(-1)
pp(p2.mcts.get_move_probs(state))
# What exactly does simulating a step do? One thing is that it expands the tree of options explored:
p2.mcts.root.children
# To see how deep that goes:
p2.mcts.root.depth()
# Ok, now let's watch the depth grow over time. In addition, we can watch a couple of move's as their probabilities change with deep exploration. Let's watch (4,1) and (7,3). The first move needs to be made to block X from winning. The second would be a step toward winning. Note that we set n_playout to 1 so we just explore one smove at a time.
import matplotlib.pyplot as plt
# +
state = game.initial
p2.mcts.update_with_move(-1)
p2.mcts.n_playout = 1
history = []
depth = [0]
watchlist = [(4,1), (7,1)]
cycles = 50
for i in range(cycles):
move_probs = p2.mcts.get_move_probs(state)
depth.append(p2.mcts.root.depth())
watch = []
for move in watchlist:
index = move_probs[0].index(move)
watch.append(move_probs[1][index])
history.append(watch)
plt.plot(history);
# -
plt.plot(depth);
state = game.string_to_state("""
.......
.......
.......
......O
......O
.XX...O
""", to_move='X')
game.display(state)
# +
state = GameState(to_move='X', board={(1,1): 'X', (2, 1): 'X',
(7,1): 'O', (7, 2): 'O'}, utility=0, moves=moves)
state = game.result(state, (3,1))
print("Start state:")
game.display(state)
p2.mcts.update_with_move(-1)
p2.mcts.is_selfplay = True
p2.mcts.temp = 0.7 # exploration (0, 1]
p2.mcts.n_playout = 30 ## 2 leads to 500 nodes after 50 cycles; 1 leads to 250 nodes after 50 cycles
history = []
depth = [0]
size = []
all_nodes = []
_ = p2.mcts.root.visit(lambda node: all_nodes.append(node.P))
watchlist = state.moves
cycles = 50
for i in range(cycles):
move_probs = p2.mcts.get_move_probs(state)
depth.append(p2.mcts.root.depth())
all_nodes = []
_ = p2.mcts.root.visit(lambda node: all_nodes.append(node.P))
size.append(len(all_nodes))
watch = []
for move in watchlist:
if move in move_probs[0]:
index = move_probs[0].index(move)
watch.append(move_probs[1][index])
history.append(watch)
plt.plot(history);
# -
plt.plot(depth);
plt.plot(size)
# Interesting! So we need at least a number of playouts before we can find an endstate.
p2.mcts.root.depth()
# ## Summary
#
# * updates root as it makes the move; no need to keep previous tree expansions
# * wherever you are, you keep the statistics about this root's possibilities
|
notebooks/monte_carlo_tree_search.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### MNIST Dataset
# http://yann.lecun.com/exdb/mnist/
#
# MNIST ("Modified National Institute of Standards and Technology") is the de facto “hello world” dataset of computer vision. Since its release in 1999, this classic dataset of handwritten images has served as the basis for benchmarking classification algorithms. As new machine learning techniques emerge, MNIST remains a reliable resource for researchers and learners alike.
# The MNIST database contains 60,000 training images and 10,000 testing images.
# 
import pandas as pd
import matplotlib.pyplot as plt
pd.options.display.max_columns = None
random_state = 42
# ### Load Data
# The MNIST data comes pre-loaded with sklearn. The first 60000 images are training data and next 1000 are test data
# ### Create a Validation set
# In real world ML scenarios we create separate Train, Validation and Test set. We train our model on Training set, optimize our model using validation set and evalaute on Test set so that we dont induce bias. Since we alreday have test set we need to split training set into separate traiining and validation sets. As we will see later that we can do K-fold cross valaidtion which removes the necessaity of creating Validations sets
# +
#display first 5 rows
# -
# ### Display Sample Image
#
# +
import matplotlib
def display_digit(digit):
# digit = X_train[2]
digit_image = digit.reshape(28,28)
plt.imshow(digit_image, cmap = matplotlib.cm.binary, interpolation = 'nearest')
plt.axis('off')
plt.show()
# -
# Each Image consist of 28 X 28 pixels with pixel values from 0 to 255. The pixel values represent the greyscale intensity increasing from 0 to 255. As we can see below digit 6 can be represented by pixel intensities of varying values and the region where pixel intensities has high value are assosciated with the image of 6
#
#Reshape the image data to 28 X 28
# ### Traget Value Counts
# ## Train Model Using Decision Tree
# #### Validation Set Accuracy
# ## Train Model Using Random Forest:Default
# #### Validation set accuracy
# We can see that just using default parameters, we are able to achieve better accuracy on Random Forest compared to a single Decision tree. Random Forest by default uses 10 decision trees to make predcitions and the end results is combined prediction of all trees.
# ## Train Model Using Random Forest: Tuned HyperParameters
# The Hyperparameters were manually Tuned, we will later see serach algorithms to find best hyperparameters automatically
#
# #### Validation Set Accuracy
# #### Test Set Accuracy
# ### Random Incorrect Predictions
# Lets display random 10 images in test data which were incorrectly predicted by our model.
# We can notice some of the images are difficult to identify even for humans
# +
def display_incorrect_preds(y_test, y_pred):
test_labels = pd.DataFrame()
test_labels['actual'] = y_test
test_labels['pred'] = y_pred
incorrect_pred = test_labels[test_labels['actual'] != test_labels['pred'] ]
random_incorrect_pred = incorrect_pred.sample(n= 10)
for i, row in random_incorrect_pred.iterrows():
print('Actual Value:', row['actual'], 'Predicted Value:', row['pred'])
display_digit(X_test[i])
# -
|
RandomForest/RandomForest_Exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# GENERATOR are created using yield keyword , it iterates or gives output
# when and when function is called you can see difference below
def simple():
for x in range(3):
print(x)
def simple_gen():
for x in range(3):
yield x
# -
a=simple()
type(a)
simple_gen()
g=simple_gen()
g
print(g)
print(next(g))
print(next(g))
print(next(g))
print(next(g))
def check():
i=0
while i<5:
print(i)
i+=1
tuple(check())
|
Generator/generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python38-azureml
# kernelspec:
# display_name: Python 3.8 - AzureML
# language: python
# name: python38-azureml
# ---
# # Notebook Environment Setup
#
# This notebook takes you through detailed setup of your settings for
# Azure Sentinel Notebooks and the MSTICPy library. It covers:
#
# - Setting up your Python environment for notebooks
# - Creating and editing your msticpyconfig.yaml file
# - Understanding and managing you config.json file.
#
# If you are
# using notebooks in the Azure Sentinel/Azure ML environment you can skip
# the first section "Configuring your Python Environment" entirely.
#
# The main part of this notebook involves setting up your msticpyconfig.yaml.
# While many of these settings are optional, if you do not configure
# them correctly you'll experience some loss of functionality. For
# example, using Threat Intelligence providers usually requires an
# API key. To save you having to type this in every time you look up
# an IP Address you should put this in a config file.
#
# This section takes you through creating settings for
# - Azure Sentinel workspaces
# - Threat Intelligence providers
# - Geo-location providers
# - Other data providers (e.g. Azure APIs)
# - Key Vault
# - Auto-loading options.
#
# You'll typically need the first three of these to use most
# of the notebooks fully.
#
# Section 3, "The config.json file" can also be ignored if you
# are happy using `msticpyconfig.yaml`. It is included here
# for background.
# + [markdown] toc=true
# <!DOCTYPE html PUBLIC "-//W3C//DTD XHTML 1.0 Strict//EN" "http://www.w3.org/TR/xhtml1/DTD/xhtml1-strict.dtd">
# <html xmlns="http://www.w3.org/1999/xhtml">
#
# <head>
# <meta name="generator" content="HTML Tidy for Windows (vers 14 February 2006), see www.w3.org" />
# <title></title>
# </head>
#
# <body>
# <h2>Contents</h2>
# <div class="toc">
# <ul class="toc-item">
# <li><span><a href="#Configuring-your-Python-Environment">Configuring your Python Environment</a></span>
# <ul class="toc-item">
# <li><span><a href="#Python-3.6-or-Later">Python 3.6 or Later</a></span></li>
# <li><span><a href="#Creating-a-virtual-environment">Creating a virtual
# environment</a></span></li>
# <li><span><a href="#Using-Requirements.txt">Using Requirements.txt</a></span></li>
# <li><span><a href="#Installing-in-a-Conda-Environment">Installing in a Conda Environment</a></span></li>
# <li><span><a href="#Installing-with---user-option">Installing with --user option</a></span></li>
# <li><span><a href="#Install-Packages-from-this-Notebook">Install Packages from this Notebook</a></span></li>
# </ul>
# </li>
# <li><span><a href="#MSTICPy-Configuration-File---msticpyconfig.yaml">MSTICPY Configuration File <code>msticpyconfig.yaml</code></a></span>
# <ul class="toc-item">
# <li><span><a href="#Display-your-existing-msticpyconfig.yaml">Display your existing msticpyconfig.yaml</a></span></li>
# <li><span><a href="#Import-your-Config.json-and-create-a-msticpyconfig.yaml-[Azure-Sentinel]">Import your Config.json and create a msticpyconfig.yaml [Azure Sentinel]</a></span></li>
# <li><span><a href="#Setting-the-path-to-your-msticpyconfig.yaml">Setting the path to your msticpyconfig.yaml</a></span></li>
# <li><span><a href="#Verify-(or-add)-Azure-Sentinel-Workspace-settings">Verify (or add) Azure Sentinel Workspace settings</a></span></li>
# <li><span><a href="#Adding-Threat-Intel-(TI)-Providers">Adding Threat Intel (TI) Providers</a></span></li>
# <li><span><a href="#Adding-GeoIP-Providers">Adding GeoIP Providers</a></span></li>
# <li><span><a href="#Optional-Settings-1---Azure-Data-and-Azure-Sentinel-APIs">Optional Settings 1 - Azure Data and Azure Sentinel APIs</a></span></li>
# <li><span><a href="#Optional-Settings-2---Autoload-QueryProviders">Optional Settings 2 - Autoload QueryProviders</a></span></li>
# <li><span><a href="#Optional-Settings-3---Autoloaded-Component">Optional Settings 3 - Autoloaded Component</a></span></li>
# <li><span><a href="#Save-your-file-and-add-the-MSTICPYCONFIG-environment-variable">Save your file and add the MSTICPYCONFIG environment variable</a></span></li>
# <li><span><a href="#Validating-your-msticpyconfig.yaml-settings">Validating your msticpyconfig.yaml settings</a></span></li>
# </li>
# <li><span><a href="#The-config.json-file">The <code>config.json</code> file</a></span></li>
# </ul>
# </div>
# </body>
#
# </html>
# -
# ---
#
# ## Configuring your Python Environment
# ### Python 3.6 or Later
# If you are running in Jupyterhub environment such as Azure Notebooks, Python is already installed. When using any of the sample notebooks or copies of them you only need to ensure that the Python 3.6 (or later) kernel is selected.
#
# If you are running the notebooks locally will you need to install Python 3.6 or later. The Ananconda distribution is a good starting point since it comes with many required packages already installed.
#
# ### Creating a virtual environment
# If you are running these notebooks locally, it is a good idea to create a clean Python *virtual environment*, before installing any of the packages . This will prevent installed packages conflicting with versions that you may need for other applications.
#
# For standard python use the [`venv`](https://docs.python.org/3/library/venv.html?highlight=venv) command.
# For Conda use the [`conda env`](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html) command.
# In both cases be sure to activate the environment before running jupyter using `venvpath/Scripts/activate` or `conda activate {my_env_name}`.
#
# Run this cell to view requirements.txt
# %pfile requirements.txt
# ### Installing in a Conda Environment
# Although you can use pip inside a conda environment it is usually better to try to install conda packages whenever possible.
#
# ```
# activate {my_env_name}
# conda config --append channels conda-forge
# conda install package1 package2
# ```
#
# See [Managing packages](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-pkgs.html) in Anaconda.
#
# For packages that are not available as conda packages use pip from with a Conda prompt/shell to install the remaining packages.
#
# ### Installing with --user option
# If you are using a shared installation of Python (i.e. one installed by the administrator) you will need to add the `--user` option to your `pip install` commands. E.g.
#
# ```
# pip install pkg_name --user --upgrade
# ```
#
# This will avoid permission errors by installing into your user folder.
#
# > **Note**: the use of the `--user` option is usually not required in a Conda environment
# > since the Python site packages are normally already installed in a per-user folder.
# ### Install Packages from this Notebook
# The first time this cell runs for a new Azure ML or Azure Notebooks notebook or other Python environment it will do the following things:
# 1. Check the kernel version to ensure that a Python 3.6 or later kernel is running
# 2. Check the msticpy version - if this is not installed or the version installed is less than the required version (in `REQ_MSTICPY_VER`)
# it will attempt to install a new version (you will be prompted whether you want to do this)
# The install can take several minutes depending on the versions of packages that you already have installed.
#
# > **Note:** These two steps are run from a local python module - this is available in the Azure-Sentinel-Notebooks repo.
# > If you do not have this locally, download it from [here](https://github.com/Azure/Azure-Sentinel-Notebooks/blob/master/utils/nb_check.py) and
# > put a copy in a `utils` subfolder of your current directory.
#
# 3. Once *msticpy* is installed and imported, the `init_notebook` function is run. This:
# - imports common modules used in the notebook
# - installs additional packages
# - sets some global options
#
# > **Note:** In subsequent runs, this cell should run quickly since you will already have the required packages installed.
#
#
# > **Warning:** you may see some warnings about incompatibility with certain packages. This should not affect the functionality of this notebook but you may need to upgrade the packages producing the warnings to a more recent version.
# +
from pathlib import Path
import os
import sys
import warnings
from IPython.display import display, HTML, Markdown
REQ_PYTHON_VER=(3, 6)
REQ_MSTICPY_VER=(0, 6, 0)
update_nbcheck = (
"<p style='color: orange; text-align=left'>"
"<b>Warning: we needed to update '<i>utils/nb_check.py</i>'</b><br>"
"Please restart the kernel and re-run this cell."
"</p>"
)
display(HTML("<h3>Starting Notebook setup...</h3>"))
if Path("./utils/nb_check.py").is_file():
try:
from utils.nb_check import check_versions
except ImportError as err:
# %xmode Minimal
# !curl https://raw.githubusercontent.com/Azure/Azure-Sentinel-Notebooks/master/utils/nb_check.py > ./utils/nb_check.py 2>/dev/null
display(HTML(update_nbcheck))
if "check_versions" not in globals():
raise ImportError("Old version of nb_check.py detected - see instructions below.")
# %xmode Verbose
check_versions(REQ_PYTHON_VER, REQ_MSTICPY_VER)
# If not using Azure Notebooks, install msticpy with
# # !pip install msticpy
from msticpy.nbtools import nbinit
nbinit.init_notebook(
namespace=globals(),
);
# -
# ---
#
# ## MSTICPy Configuration File - `msticpyconfig.yaml`
#
# *MSTICPy* is a Python package used in most of the Jupyter notebooks
# on Azure-Sentinel-Notebooks. It provides a lot of functionality specific
# to threat hunting and investigations, including:
# - Data querying against Azure Sentinel tables (also MDE, Splunk and other)
# - Threat Intelligence lookups using multiple TI providers (VirusTotal, AlienVault OTX and others)
# - Common enrichment functions (GeoIP, IoC extraction, WhoIs, etc.)
# - Visualization using event timelines, process trees and Geo-mapping
# - Advanced analysis such as Time Series decomposition, Anomaly detection and clustering.
#
# > **Note**: the configuration actions in this section are
# > an abbreviated version of the
# > [MPSettingsEditor notebook](https://github.com/microsoft/msticpy/blob/master/docs/notebooks/MPSettingsEditor.ipynb)<br>
# > Use this notebook for a fuller guide on how to configure your settings.<br>
# > Also, see these sections in the *MSTICPy* documentation:<br>
# > [MSTICPy Package Configuration](https://msticpy.readthedocs.io/en/latest/getting_started/msticpyconfig.html)<br>
# > [MSTICPy Settings Editor](https://msticpy.readthedocs.io/en/latest/getting_started/SettingsEditor.html)
#
#
#
# `config.json` provides some basic configuration for connecting to your Azure Sentinel workspace.
# However, there are many features that require additional configuration information. Some examples are:
# - Threat Intelligence Provider connection information
# - GeoIP connection information
# - Keyvault configuration for storing secrets remotely
# - MDE and Azure API connection information.
# - Connection information for multiple Azure Sentinel workspaces.
#
# Settings for these are stored in the `msticpyconfig.yaml` file. This file is read from the current directory or you can set an environment variable (`MSTICPYCONFIG`) pointing to its location.
# Form more information about *msticpy* configuration see [msticpy Package Configuration](https://msticpy.readthedocs.io/en/latest/getting_started/msticpyconfig.html).
#
# The most commonly-used sections are described below.
#
#
# #### Threat Intelligence Provider Setup
# For more information on the msticpy Threat Intel lookup class see the [documentation here](https://msticpy.readthedocs.io/en/latest/data_acquisition/TIProviders.html).
#
# Primary providers are used by default. Secondary providers are not run by default but can be invoked by using the `providers` parameter to `lookup_ioc()` or `lookup_iocs()`. Set the `Primary` config setting to `True` or `False` for each provider ID according to how you want to use them. The `providers` parameter should be a list of strings identifying the provider(s) to use.
#
# - The provider ID is given by the `Provider:` setting for each of the TI providers - do not alter this value.
# - Delete or comment out the section for any TI Providers that you do not wish to use.
# - For most providers you will usually need to supply an authorization (API) key and in some cases a user ID for each provider.
# - For the Azure Sentinel TI provider, you will need the workspace ID and tenant ID and will need to authenticate in order
# to access the data (although if you have an existing authenticated connection with the same workspace/tenant, this connection will be re-used).
#
# #### GeoIP Providers
# Like the TI providers these services normally need an API key to access. You can read more about configuration
# the supported providers here. [msticpy GeoIP Providers](https://msticpy.readthedocs.io/en/latest/data_acquisition/GeoIPLookups.html)
#
# #### Browshot Setup
# The functionality to screenshot a URL in msticpy.sectools.domain_utils relies on a service called BrowShot (https://browshot.com/). An API key is required to use this service and it needs to be defined in the `msticpyconfig` file as well. As this is not a threat intelligence provider it doesn't not fall under the `TIProviders` section of `msticpyconfig` but instead sits alone. See the cell below for example configuration.
# ---
#
# ### Display your existing `msticpyconfig.yaml`
#
# We'll be using some of the *MSTICPy* configuration tools:
# *MPConfigEdit* and *MPConfigFile*, so we'll import these first
from msticpy.config import MpConfigFile, MpConfigEdit, MpConfigControls
from msticpy.nbtools import nbwidgets
from msticpy.common import utility as utils
# Then run MpConfig file to view your current settings.
mpconfig = MpConfigFile()
mpconfig.load_default()
mpconfig.view_settings()
# #### If you see nothing but a pair of curly braces...
# ...in the settings view above it means
# that you probably need to create up a **msticpyconfig.yaml**
#
# If you know that you have configured a msticpyconfig file,
# you can search for this file using MpConfigFile. Click on **Load file**.
# Once you've done that go to the [Setting the path to your msticpyconfig.yaml](#Setting-the-path-to-your-msticpyconfig.yaml)
# ---
#
# ### Import your Config.json and create a msticpyconfig.yaml [Azure Sentinel]
#
# Follow these steps:
# 1. Run MpConfigFile
# 2. Locate your config.json
# - click **Load file** button
# - Browse - use the controls to navigate to find config.json
# - Search - set the starting directory to search and open the **Search** drop-down
# - When you see the file click on it and click **Select File** button (below the file browser)
# - optionally, click **View Settings** to confirm that this looks right
# 3. Convert to convert to msticpyconfig format
# - click **View Settings**
# 4. Save your `msticpyconfig.yaml` file
# - type a path into the **Current file** text box
# - Click on **Save file**
# 5. You can set this file to always load by assigning the path to an environment variable.
# See [Setting the path to your msticpyconfig.yaml](#Setting-the-path-to-your-msticpyconfig.yaml)
# ---
#
# ### Setting the path to your msticpyconfig.yaml
#
# This is a good point to set up an environment variable so that
# you can keep a single configuration file in a known location and always
# load the same settings. (Of course, you're free to use multiple configs
# if you need to use different settings for each notebook folder)
#
# - decide on a location for your `msticpyconfig.yaml` - this could be in "~/.msticpyconfig.yaml" or "%userprofile%/msticpyconfig.yaml"
# - copy the `msticpyconfig.yaml` file that you just created to this location.
# - set the `MSTICPYCONFIG` environment variable to point to that location:
#
# #### Windows
# <img src="https://github.com/Azure/Azure-Sentinel-Notebooks/raw/master/images/win_env_var.png" style="width: 500px;"/>
#
#
# #### Linux
# In your .bashrc (or somewhere else convenient) add:
#
# `export MSTICPYCONFIG=~/.msticpyconfig.yaml`
#
# #### Azure ML
#
# In Azure ML, you need to decide whether to store your `msticpyconfig.yaml` in
# the AML file store or on the Compute file system. If you have any secret
# key material in the file, we recommend storing on the Compute instance, since
# the AML file store is shared storage, whereas the Compute instance is
# accessible only by the user who created it.
#
# If you are happy to leave the file in the AML file store, you should be set.
# The nb_check.check_versions function run at the start of the notebook
# will find it there in your root folder and set the MSTICPYCONFIG environment
# variable to point to it.
#
# **Pointing to a path on a compute instance**
#
# 1. Open a terminal in AML
# <br>
# <img src="https://github.com/Azure/Azure-Sentinel-Notebooks/raw/master/images/aml_terminal.png" style="width: 300px;"/>
#
# 2. Verify your msticpyconfig.yaml is accessible
#
# Your current directory should be your AML file store home directory
# (this is mounted in the Compute Linux system) and the prompt will look
# something like the example below.
#
# If you created a `msticpyconfig.yaml` in the previous step,
# this should be visible if you type `ls`.
# ```bash
# azureuser@ianhelle-azml7:~/cloudfiles/code/Users/ianhelle$ ls msti*
# msticpyconfig.yaml
# ```
#
#
# 3. Move the file to your home folder
#
# ```bash
# mv msticpyconfig.yaml ~
# ```
#
#
# 4. Add an environment variable
# Because the Jupyter server is started before you connect its process
# will not inherit and environment variables from you .bashrc
# You can set it one of two places:
#
# - The `kernel.json` file for your Python kernel (there are kernels for
# both Python 3.6 and Python 3.8
# - Add a Python file `nbuser_settings.py` to the root of your user folder.
#
# These options are described in the following sections.
#
# **kernel.json**
#
# - Python 3.8 location: `/usr/local/share/jupyter/kernels/python38-azureml/kernel.json`
# - Python 3.6 location: `/usr/local/share/jupyter/kernels/python3-azureml/kernel.json`
#
# Make a copy of the file and open the original in an editor (you many need to use sudo to be able to overwrite this file).
# The file will look something like this
# ```json
# {
# "argv": [
# "/anaconda/envs/azureml_py38/bin/python",
# "-m",
# "ipykernel_launcher",
# "-f",
# "{connection_file}"
# ],
# "display_name": "Python 3.8 - AzureML",
# "language": "python"
# }
# ```
# Add the following line after the "language" item.
# ```json
# "env": { "MSTICPYCONFIG": "~/msticpyconfig.yaml" }
# ```
#
# Your file should look like this (remember to add a comma at the end of the
# `"language": "python"` line
# ```json
# {
# "argv": [
# "/anaconda/envs/azureml_py38/bin/python",
# "-m",
# "ipykernel_launcher",
# "-f",
# "{connection_file}"
# ],
# "display_name": "Python 3.8 - AzureML",
# "language": "python",
# "env": { "MSTICPYCONFIG": "~/msticpyconfig.yaml" }
# }
# ```
#
# If you use both kernels you will need to edit both files.
#
# **nbuser_settings.py**
#
# Create this file (you can do this from the AML workspace) in the
# root of your user folder (i.e. inside the folder with your username)
# and add the following lines
# ```python
# import os
# os.environ["MSTICPYCONFIG"] = "~/msticpyconfig.yaml"
# ```
#
# This file, if it exists, is imported by the `nb_check.check_versions`
# function at the start of the notebook. It will set the environment
# variable at the start of each notebook before any configuration is read.
# This is simpler and less intrusive than editing the kernel.json.
# However, it only works if you run `check_versions`. If you load
# a notebook without running this MSTICPy may not be able to find
# its configuration file.
#
# ---
#
# ### Verify (or add) Azure Sentinel Workspace settings
#
# If you loaded a config.json file into your msticpyconfig.yaml, you should see
# your workspace displayed when you run the following cell. If not,
# you can add one or more workspaces here. The **Name**, **WorkspaceId** and **TenantId** are
# mandatory. The other fields are helpful but not essential.
#
# Use the Help drop-down panel to find more information about adding workspaces and finding
# the correct values for your workspace.
#
# If this the workspace that you use frequently or all of the time, you may want to set this as the **default.**
# This creates a duplicate entry named "Default". This is used when you connect to AzureSentinel without
# needing to supply a workspace name. You can override this by specifying a workspace name at connect time,
# which you need to do if you are working with multiple workspaces.
#
# When you've finished, type a file name (usually "msticpyconfig.yaml") into the **Conf File** text box
# and click **Save File**,
#
# You can also try the **Validate Settings** button. This should show that you have a few missing
# sections (we'll fill these in later) but should show nothing under the the "Type Validation Results".
mpedit = MpConfigEdit(settings=mpconfig)
mpedit.set_tab("AzureSentinel")
mpedit
# ---
#
# ### Adding Threat Intel (TI) Providers
#
# You will likely want to do lookups of IP Addresses, URLs and other items to check for any Threat Intelligence reports.
# To do that you need to add the providers that you want to use. Most TI providers require that you
# have an account with them and supply an API key or other authentication items when you connect.
#
# Most providers have a free use tier (or in cases like AlienVault OTX) are entirely free.
# Free tiers for paid providers usually impose a certain number of requests that you
# can make in a given time period.
#
# For account creation, each provider does this slightly differently.
# Use the help links in the editor help to find where to go set each of these up.
#
# Assuming that you have done this, we can configure a provider. Be sure to
# store any authentication keys somewhere safe (and memorable).
#
# We are going to use [VirusTotal](https://www.virustotal.com) (VT) as an example TI Provider.
# For this you will need a VirusTotal API key from the
# [VirusTotal](https://developers.virustotal.com/v3.0/reference#getting-started) website.<br>
# We also support a range of other threat intelligence providers - you can read about this here [MSTICPy TIProviders](https://msticpy.readthedocs.io/en/latest/data_acquisition/TIProviders.html)
# <br><br>
# Taking VirusTotal as our example.
# - Click on the **TI Providers** tab
# - Select "VirusTotal" from the **New prov** drop-down list
# - Click **Add**
#
# This should show you the values that you need to provide:
# - a single item **AuthKey** (this is usually referred to as an "API Key"
#
# You can paste the key into the **Value** field and click the **Save** button.
#
# You can opt to store the VT AuthKey as an environment variable. This is a bit more secure than
# having it laying around in configuration files.
# Assuming that you have set you VT key as an environment variable
# ```bash
# set VT_KEY=<KEY> (Windows)
# export VT_KEY=<KEY> (Linux/MAC)
# ```
# Flip the **Storage** radio button to **EnvironmentVar** and type the name of the
# variable (`VT_KEY` in our example) into the value box.
#
# You can also use Azure Key Vault to store secrets like these but we will need to
# set up the Key Vault settings before this will work.
#
# Click the **Save File** button to save your changes.
mpedit.set_tab("TI Providers")
mpedit
# ---
#
# ### Adding GeoIP Providers
#
# MSTICPy supports two Geo IP providers - Maxmind GeoIPLite and IP Stack.
# The main difference between the two is that Maxmind downloads and uses a local
# database, while IPStack is a purely online solution.
#
# For either you need API keys to either download the free database from MaxMind
# or access the IPStack online lookup
#
# We'll use GeoIPLite as our example.
# You can sign up for a free account and API key at https://www.maxmind.com/en/geolite2/signup.
# You'll need the API for the following steps.
# - Select "GeoIPLite" from the **New Prov**
# - Click **Add**
# - Paste your Maxmind key into the **Value** field
#
# Set the maxmind data folder:
# - This defaults to "~/.msticpy"
# - On Windows this translates to the foldername `%USERPROFILE%/.msticpy`.
# - On Linux/Mac this translates to the folder `.msticpy` in your home folder.
# - This is where the downloaded GeopIP database will be stored.
# - Choose another folder name and location if you prefer.
#
# > **Note**: as with the TI providers you can opt to store your key
# > as an environment variable or keep it in Key Vault.
#
mpedit.set_tab("GeoIP Providers")
mpedit
# <h3 style='color:red'>Important Security Note</h3>
#
# > You might not be too comfortable leaving API keys stored in
# > text files. You can opt to have these settings stored either:
# > - as Environment Variables
# > - in Azure Key Vault
#
# To see how to do this see these resources
# - [MPSettingsEditor notebook](https://github.com/microsoft/msticpy/blob/master/docs/notebooks/MPSettingsEditor.ipynb)<br>
# - [MSTICPy Package Configuration](https://msticpy.readthedocs.io/en/latest/getting_started/msticpyconfig.html)<br>
# - [MSTICPy Settings Editor](https://msticpy.readthedocs.io/en/latest/getting_started/SettingsEditor.html)
#
# ---
#
# ### Optional Settings 1 - Azure Data and Azure Sentinel APIs
#
# ### Azure API and Azure Sentinel API
# To access Azure APIs (such as the Sentinel APIs or Azure resource APIs)
# you need to be able to use Azure Authentication.
# The setting is named "AzureCLI" for historical reasons - don't let that confuse you.
# We currently support two ways of authenticating:
# 1. Chained chained authentication (recommended)
# 2. With a client app ID and secret
#
# The former can try up to four methods of authentication:
# - Using creds set in environment variables
# - Using creds available in an AzureCLI logon
# - Using the Managed Service Identity (MSI) credentials of the machine you are
# running the notebook kernel on
# - Interactive browser logon
#
# To use chained authentication methods select the methods to want to use and leave
# the clientId/tenantiId/clientSecret fields empty.
#
mpedit.set_tab("Data Providers")
mpedit
# ---
#
# ### Optional Settings 2 - Autoload QueryProviders
#
# This section controls which, if any query providers you want to load automatically
# when you run `nbinit.init_notebook`.
#
# This can save a lot of time if you are frequently authoring new notebooks. It also
# allows the right providers to be loaded before other components that might use them such as
# - Pivot functions
# - Notebooklets
# (more about these in the next section)
#
# There are two types of provider support:
# - Azure Sentinel - here you specify both the provider name and the workspace name that you want to connect to.
# - Other providers - for other query providers, just specify the name of the provider.
#
# Available Azure Sentinel workspaces are taken from the items you configured in the **Azure Sentinel** tab.
# Other providers are taken from the list of available provider types in *MSTICPy*.
#
# There are two options for each of these:
# - **connect** - if this is True (checked) *MSTICPy* will try to authenticate to the
# provider backend immediately after loading. This assumes that you've configured
# credentials for the provider in your settings.
# Note: if this is not set it defaults to True.
# - **alias** - when MSTICPy loads a provider it assigns it to a Python variable name.
# By default this is "qry_*workspace_name*" for Azure Sentinel providers and
# "qry_*provider_name*" for other providers. If you want to use something a bit shorter
# and easier to type/remember you can add a *alias*. The variable name created
# will be "qry_*alias*"
#
# > **Note** if you lose track of which providers have been loaded by
# > this mechanism they are added to the `current_providers` attribute of
# > `msticpy`
# ```python
# import msticpy
# msticpy.current_providers
# ```
mpedit.set_tab("Autoload QueryProvs")
mpedit
# ---
#
# ### Optional Settings 3 - Autoloaded Component
# This section controls which, if other components you want to load automatically
# when you run `nbinit.init_notebook()`.
#
# This includes
# - TILookup - the Threat Intel provider library
# - GeopIP - the Geo ip provider that you want to use
# - AzureData - the module used to query details about Azure resources
# - AzureSentinelAPI - the module used to query the Azure Sentinel API
# - Notebooklets - loads notebooklets from the [msticnb package](https://msticnb.readthedocs.io/en/latest/)
# - Pivot - pivot functions
#
# These are loaded in this order, since the Pivot component needs query and other providers
# loaded in order to find the pivot functions that it will attach to entities.
# For more information see [pivot functions](https://msticpy.readthedocs.io/en/latest/data_analysis/PivotFunctions.html)
#
# Some components do not require any parameters (e.g. TILookup and Pivot). Others do support or require additional
# settings:
#
# **GeoIpLookup**
#
# You must type the name of the GeoIP provider that you want to use - either "GeoLiteLookup" or "IPStack"
#
# **AzureData** and **AzureSentinelAPI**
# - **auth_methods** - override the default settings for AzureCLI and connect using the selected methods
# - **connnect** - set to false to load but not connect
#
# **Notebooklets**
#
# This has a single parameter block **AzureSentinel**. At minumum you
# should specify the workspace name. This needs to be in the following
# format:
# ```
# workspace:WORKSPACENAME
# ```
# WORKSPACENAME must be one of the workspaces defined in the Azure Sentinel tab.
#
# You can also add addition parameters to send to the notebooklets init function:
# Specify these as addition key:value pairs, separated by newlines.
# ```
# workspace:WORKSPACENAME
# providers=["LocalData","geolitelookup"]
# ```
# See the
# [msticnb `init` documentation](https://msticnb.readthedocs.io/en/latest/msticnb.html#msticnb.data_providers.init)
# for more details
#
mpedit.set_tab("Autoload Components")
mpedit
# ---
#
# ### Save your file and add the MSTICPYCONFIG environment variable
#
# Save your file, and, if you haven't yet done so, create an
# enviroment variable to point to it. See [Setting the path to your msticpyconfig.yaml](#Setting-the-path-to-your-msticpyconfig.yaml)
# ---
#
# ### Validating your `msticpyconfig.yaml` settings
#
# MpConfigFile includes a validation function that can help
# you diagnose setup problems.
#
# You can run this interactively or from Python.
#
# The examples below assume that you have set `MSTICPYCONFIG` to point
# to you config file. If not, you will need to use the `load_from_file()`
# function (or Load File button) to load the file before validating.
mpconfig = MpConfigFile()
mpconfig.load_default()
mpconfig.validate_settings()
# To validate interactively:
mpconfig = MpConfigFile()
mpconfig.load_default()
mpconfig
# ---
#
# ## The `config.json` file
# When you start a notebook from Azure Sentinel for the first time it will create a `config.json` file in
# your notebooks folder. This should be populated with your workspace and tenant IDs needed to
# authenticate to Azure Sentinel.
#
# If you are using notebooks in a different environment you may need to create a `config.json` or `msticpyconfig.yaml` (see below)
# to supply this information to your notebook.
#
# We recommend creating a `msticpyconfig.yaml` since this can hold a wide variety
# of settings for your notebook, including multiple Azure Sentinel workspace settings.
# The config.json, in contrast, only holds settings for a single Azure Sentinel workspace.
#
# For more information see this [msticpy Package Configuration](https://msticpy.readthedocs.io/en/latest/getting_started/msticpyconfig.html)
#
# ---
#
# If you need to create or modify your config.json you can run the following cell.
#
# You will need the subscription and workspace IDs for your Azure Sentinel Workspace. These can be found here in the Azure Sentinel portal as shown below.
#
# <img src="https://github.com/Azure/Azure-Sentinel-Notebooks/raw/master/images/az_sentinel_settings1.png" width="600">
#
# <br>Copy the subscription and workspace IDs:<br>
# <img src="https://github.com/Azure/Azure-Sentinel-Notebooks/raw/master/images/az_sentinel_settings2.png" style="width: 600px;"/>
# +
import requests
import json
import ipywidgets as widgets
from pathlib import Path
from datetime import datetime
config_dict = {}
def get_tenant_for_subscription(sub_id):
aad_url = (
f"https://management.azure.com/subscriptions/{sub_id}?api-version=2016-01-01"
)
resp = requests.get(aad_url)
if resp.status_code == 401:
hdr_list = resp.headers["WWW-Authenticate"].split(",")
hdr_dict = {
item.split("=")[0].strip(): item.split("=")[1].strip() for item in hdr_list
}
return hdr_dict["Bearer authorization_uri"].strip('"').split("/")[3]
else:
return None
def save_config_json(file_path, **kwargs):
if Path(file_path).exists():
bk_file = (
str(Path(file_path))
+ ".bak"
+ datetime.now().isoformat(timespec="seconds").replace(":", "-")
)
print(f"Exising config found. Saving current config.json to {bk_file}")
Path(file_path).rename(bk_file)
with open(file_path, "w") as fp:
json.dump(kwargs, fp, indent=2)
print(f"Settings saved config to {file_path}")
def save_config(b):
tenant = input_tenant.value
if not tenant:
tenant = get_tenant_for_subscription(input_wgt["tenant"].value)
print(f"TenantID found: {tenant_id}")
save_config_json(
file_path=input_wgt["path"].value,
tenant_id=tenant,
subscription_id=input_wgt["sub_id"].value,
workspace_id=input_wgt["ws_id"].value,
workspace_name=input_wgt["workspace"].value,
resource_group=input_wgt["res_grp"].value,
)
DEFAULT_CONFIG = "./config.json"
WIDGET_DEFAULTS = {
"layout": widgets.Layout(width="95%"),
"style": {"description_width": "200px"},
}
input_wgt = {
"path": widgets.Text(
description="Path to config.json", value=DEFAULT_CONFIG, **WIDGET_DEFAULTS
),
"workspace": widgets.Text(
description="Workspace name", placeholder="Workspace name", **WIDGET_DEFAULTS
),
"sub_id": widgets.Text(
description="Azure Sentinel Subscription ID",
placeholder="for example, ef28a760-8c61-41d7-8167-5c8e5d91268b",
**WIDGET_DEFAULTS,
),
"ws_id": widgets.Text(
description="Azure Sentinel Workspace ID",
placeholder="for example, ef28a760-8c61-41d7-8167-5c8e5d91268b",
**WIDGET_DEFAULTS,
),
"res_grp": widgets.Text(
description="Resource group", placeholder="Resource group", **WIDGET_DEFAULTS
),
"tenant": widgets.Text(
description="TenantId", placeholder="Leave blank to look up", **WIDGET_DEFAULTS
),
}
if Path(DEFAULT_CONFIG).exists():
with open(DEFAULT_CONFIG, "r") as fp:
config_dict = json.load(fp)
input_wgt["path"].value = DEFAULT_CONFIG
input_wgt["sub_id"].value = config_dict.get("subscription_id", "")
input_wgt["ws_id"].value = config_dict.get("workspace_id" "")
input_wgt["workspace"].value = config_dict.get("workspace_name" "")
input_wgt["res_grp"].value = config_dict.get("resource_group" "")
input_wgt["tenant"].value = config_dict.get("tenant_id" "")
save_button = widgets.Button(description="Save config.json file")
save_button.on_click(save_config)
display(widgets.VBox([*(input_wgt.values()), save_button]))
|
ConfiguringNotebookEnvironment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import accuracy_score, mean_squared_error as MSE
from sklearn.ensemble import RandomForestRegressor
from sklearn.datasets import load_breast_cancer, load_boston
# -
data = load_breast_cancer()
X, y = data.data, data.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, stratify=y)
dt = DecisionTreeClassifier()
dt.get_params()
# +
params = {
'max_depth': np.arange(3, 7),
'min_samples_leaf': np.arange(0.04, .09, .02),
'max_features': np.arange(0.2, 0.9, .2)
}
gridsearch = GridSearchCV(estimator=dt, param_grid=params, scoring='accuracy', cv=10, n_jobs=-1)
gridsearch.fit(X_train, y_train)
# -
best_hyperparams = gridsearch.best_params_
print(f'Best params: {best_hyperparams}')
dt = DecisionTreeClassifier(max_depth=5, max_features=.4, min_samples_leaf=.04).fit(X_train, y_train)
print(f'DecisionTree with hyperparameter tuning score: {accuracy_score(y_test, dt.predict(X_test))}')
dt = DecisionTreeClassifier().fit(X_train, y_train)
print(f'DecisionTree without hyperparameter tuning score: {accuracy_score(y_test, dt.predict(X_test))}')
# +
# Import roc_auc_score from sklearn.metrics
from sklearn.metrics import roc_auc_score
# Define params_dt
params_dt = {
'max_depth': [2, 3, 4],
'min_samples_leaf': [.12, .14, .16, .18]
}
# Instantiate grid_dt
grid_dt = GridSearchCV(estimator=dt,
param_grid=params_dt,
scoring='roc_auc',
cv=5,
n_jobs=-1)
grid_dt.fit(X_train, y_train)
# Extract the best estimator
best_model = grid_dt.best_estimator_
# Predict the test set probabilities of the positive class
y_pred_proba = grid_dt.predict_proba(X_test)[:, 1]
# Compute test_roc_auc
test_roc_auc = roc_auc_score(y_test, y_pred_proba)
# Print test_roc_auc
print('Test set ROC AUC score: {:.3f}'.format(test_roc_auc))
# -
# # RandomForest Hyperparameters
#
# +
data = load_boston()
X, y = data.data, data.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2)
params_rf = {
'n_estimators': [300, 400, 500],
'max_depth': [4, 6, 8],
'min_samples_leaf': [.1, .2],
'max_features': ['log2', 'sqrt']
}
rf = RandomForestRegressor()
grid_rf = GridSearchCV(estimator=rf, param_grid=params_rf, cv=3, scoring='neg_mean_squared_error', verbose=1, n_jobs=-1, refit=True).fit(X_train, y_train)
# -
best_params = grid_rf.best_params_
print(best_params_)
best_model = grid_rf.best_estimator
y_pred = best_model.predict(X_test)
rmse = np.sqrt(MSE(y_test, y_pred))
print(f'RMSE: {rmse}')
|
datacamp_ml/ml_trees/tuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DTAT PROCESS FOR NEW DATA
#
import os
from os.path import join
AllFile = []
dest = "F:\\data\\newData"
for root, dirs, files in os.walk( dest ):
for OneFileName in files :
OneFullFileName = join( root, OneFileName )
AllFile.append(OneFullFileName)
print(AllFile)
# firstData = []
secondData = []
# for i in AllFile:
# if(len(i)==26):
# firstData.append(i)
# else:
# secondData.append(i)
for i in AllFile:
secondData.append(i)
# +
# firstData
# -
secondData
# +
# def read_dta(dta):
# """read the dta file and return numerical array"""
# with open(dta) as f:
# newData = []
# data = f.readlines()
# for i in data[6:]:
# newData.append(float(i[0:6]))
# return newData
# +
# datatest = read_dta(secondData[1])
# +
# def getPIValue(data):
# PIValue = []
# for j in range(9):
# upT = 0
# downT = 0
# for i in range(0+j*1200,1200+j*1200):
# if (data[i]>1488 and data[i]<2000) or (data[i]>2516 or data[i] <976):
# upT+=1
# else:
# downT+=1
# print(upT)
# print(downT)
# print(upT+downT)
# PIValue.append((upT-downT)/(upT+downT))
# return PIValue
# +
# firstDataPIValue = []
# for i in firstData:
# data = read_dta(i)
# PI = getPIValue(data)
# firstDataPIValue.append(PI)
# print(firstDataPIValue)
# +
# for i in range(9):
# firstDataPIValue[4][i] = -firstDataPIValue[4][i]
# for i in firstDataPIValue:
# print(i)
# +
secondPIValue=[]
for i in secondData:
temp = []
with open(i) as f:
newData = []
data = f.readlines()
for j in data[5][13:-2].split(","):
temp.append(float(j))
secondPIValue.append(temp)
print(secondPIValue)
# -
# +
# firstDataPIValue
# -
firstDataPIValue=[]
for i in secondPIValue:
firstDataPIValue.append(i)
firstDataPIValue
for i in firstDataPIValue:
print(i)
# +
# for i in range(9):
# firstDataPIValue[18][i] = -firstDataPIValue[18][i]
# -
# +
import pandas as pd
data = {i:firstDataPIValue[i] for i in range(len(firstDataPIValue))}
pdData = pd.DataFrame(data)
pdData
# -
s = pdData.T.describe()
s
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
plt.figure(figsize=(10,8), dpi=80)
name_list = ['test','test','tr','tr','test','tr','tr','test','test']
num_list = s.loc["mean"]
yerr = s.loc["std"]/np.sqrt(s.iloc[0,0])
plt.bar(range(len(num_list)), num_list,tick_label=name_list,color=['c','c','seagreen','seagreen','c','seagreen','seagreen','c','c'])
plt.xlabel("Sequence")
plt.ylabel("Performance Index")
plt.title("PI Figure")
plt.ylim(-0.6,0.8)
#plt.errorbar([i for i in range(9)],num_list,yerr=yerr,fmt="b",color="teal")
plt.errorbar([i for i in range(9)],num_list,yerr=yerr,ls='none',fillstyle='none',ms=9,mew=1.3,color='y')
plt.grid(color="grey",linewidth='0.1')
plt.show()
yerr
0.037000 -0.00250 0.284000 0.23750 0.044500 0.162000 0.135500 -0.03200 -0.00200
# +
import numpy as np
fig = plt.figure(0)
x = np.arange(10.0)
y = np.sin(np.arange(10.0) / 20.0 * np.pi)
plt.errorbar(x, y, yerr=0.1)
y = np.sin(np.arange(10.0) / 20.0 * np.pi) + 1
plt.errorbar(x, y, yerr=0.1, uplims=True)
y = np.sin(np.arange(10.0) / 20.0 * np.pi) + 2
upperlimits = np.array([1, 0] * 5)
lowerlimits = np.array([0, 1] * 5)
plt.errorbar(x, y, yerr=0.1,uplims=upperlimits, lolims=lowerlimits)
plt.xlim(-1, 10)
plt.show()
# +
x = np.linspace(0, 10, 50)
dy = 0.8
y = np.sin(x) + dy * np.random.randn(50)
plt.errorbar(x, y, yerr=dy, fmt='.k');
plt.show()
# -
fig,ax = plt.subplots()
# +
plt.bar(range(len(num_list)), num_list,yerr = [1,1,1,1,1,1,1,1,1],,tick_label=name_list,color=['c','c','seagreen','seagreen','c','seagreen','seagreen','c','c'])
plt.show()
# -
|
Analysis/FF-Analysis/data/data_analysis_for_new_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# Dependencies and Setup
import pandas as pd
# File to Load (Remember to Change These)
school_data_to_load = "Resources/schools_complete.csv"
student_data_to_load = "Resources/students_complete.csv"
# Read School and Student Data File and store into Pandas DataFrames
school_data = pd.read_csv(school_data_to_load)
student_data = pd.read_csv(student_data_to_load)
# Combine the data into a single dataset.
school_data_complete = pd.merge(student_data, school_data, how="left", on=["school_name", "school_name"])
school_data_complete.head()
# ## District Summary
#
# * Calculate the total number of schools
#
# * Calculate the total number of students
#
# * Calculate the total budget
#
# * Calculate the average math score
#
# * Calculate the average reading score
#
# * Calculate the percentage of students with a passing math score (70 or greater)
#
# * Calculate the percentage of students with a passing reading score (70 or greater)
#
# * Calculate the percentage of students who passed math **and** reading (% Overall Passing)
#
# * Create a dataframe to hold the above results
#
# * Optional: give the displayed data cleaner formatting
# +
#Calculate the total number of schools
# +
#Calculate the total number of students
# +
#Calculate the total budget
# +
#Calculate the average math score
# +
#Calculate the average reading score
# +
#Calculate the percentage of students with a passing math score (70 or greater)
# -
#Calculate the percentage of students with a passing reading score (70 or greater)
#Calculate the percentage of students who passed math and reading (% Overall Passing)
#Create a dataframe to hold the above results
#Optional: give the displayed data cleaner formatting
# ## School Summary
# * Create an overview table that summarizes key metrics about each school, including:
# * School Name
# * School Type
# * Total Students
# * Total School Budget
# * Per Student Budget
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * % Overall Passing (The percentage of students that passed math **and** reading.)
#
# * Create a dataframe to hold the above results
Create an overview table that summarizes key metrics about each school, calculated in part 1
Create a dataframe to hold the above results
# ## Top Performing Schools (By % Overall Passing)
# * Sort and display the top five performing schools by % overall passing.
# ## Bottom Performing Schools (By % Overall Passing)
# * Sort and display the five worst-performing schools by % overall passing.
# ## Math Scores by Grade
# * Create a table that lists the average Reading Score for students of each grade level (9th, 10th, 11th, 12th) at each school.
#
# * Create a pandas series for each grade. Hint: use a conditional statement.
#
# * Group each series by school
#
# * Combine the series into a dataframe
#
# * Optional: give the displayed data cleaner formatting
# ## Reading Score by Grade
# * Perform the same operations as above for reading scores
# ## Scores by School Spending
# * Create a table that breaks down school performances based on average Spending Ranges (Per Student). Use 4 reasonable bins to group school spending. Include in the table each of the following:
# * Average Math Score
# * Average Reading Score
# * % Passing Math
# * % Passing Reading
# * Overall Passing Rate (Average of the above two)
# ## Scores by School Size
# * Perform the same operations as above, based on school size.
# ## Scores by School Type
# * Perform the same operations as above, based on school type
|
PyCitySchools_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **What does this notebook do?**
# - Load the exported CGM values from NutriSense
# - Print out what days are included in the dataset
# - Pair down data to only one day, include CGM values, meals and exercise
# - Smooth CGM data and interpolate missing values
# - Calculate key metrics for that day
# - Pull in Garmin "run activities" and plot them
# - Create a chart of the glucose values and include metrics
# +
import pandas as pd
import plotly.express as px
import datetime
from datetime import date
from garminconnect import (
Garmin,
GarminConnectConnectionError,
GarminConnectTooManyRequestsError,
GarminConnectAuthenticationError,
)
# Read in CSV file
df = pd.read_csv('export.csv')
# Remove "time zone offset" from "occurred_at" column and add new "occurred_at_day" column
df['occurred_at_day'] = df['occurred_at'].apply(lambda x: x[:len(x) - 15])
df['occurred_at'] = df['occurred_at'].apply(lambda x: x[:len(x) - 6])
df.head()
# -
# Print all days with data
daysWithData = df['occurred_at_day'].unique()
print(daysWithData)
# +
# Filter down to one day, pick the second day in the dataset
df = df[df['occurred_at_day']==daysWithData[2]]
day = daysWithData[2]
# Create a datasets just with glucose measurments
gm = df[df['class']=='GlucoseMeasurement']
# Create a dataset for meals and exercise, sort it
mealsExercise = df[((df['class']=='Meal') | (df['class']=='ExerciseActivity') )]
mealsExerciseSorted = mealsExercise.sort_values(by=["occurred_at"], ascending=True)
# +
# Create a dataset with just 2 columns
gm_data = gm.filter(['occurred_at', 'value'])
# rename the columns for easier readability
gm_data.columns = ['time', 'value']
# turn time column into the index and delete time column
gm_data['time']= pd.to_datetime(gm_data['time'])
gm_data.index = gm_data['time']
del gm_data['time']
gm_data = gm_data.resample('1T').mean() # add rows for every 1 minute
gm_data = gm_data.interpolate(method='cubic') # interpolate the new 1 minute points with data
# Calculate a few metrics
threshold = 120 # this is an arbitrary threshold
above = gm_data[gm_data['value'] > threshold] # create a dataset with glucose measuremnts over threshold
minutesAboveThreshold = above.count()
print('Number of minutes above '+str(threshold)+': '+ minutesAboveThreshold.to_string(index=False))
percentageAboveThreshold = int(round(minutesAboveThreshold/(60*24)*100,0))
print("Time above Threshold = "+str(percentageAboveThreshold)+"%")
averageGlucose = int(round(gm_data['value'].mean()))
medianGlucose = int(round(gm_data['value'].median()))
print("Average Glucose = "+str(averageGlucose))
print("Median Glucose = "+str(medianGlucose))
# -
# Get Garmin Data
# This may not be so great, defaulting to simply retrieving the last 100 activities on Garmin.
# If the day that is plotted is further in the past, this may not work.
numberOfActivities = 100
try:
# Initialize Garmin client with credentials
# Put your userID and password for https://connect.garmin.com/ here
client = Garmin("USERID", "PASSWORD")
# Login to Garmin Connect portal
client.login()
# Get running activities
allActivities = client.get_activities(0,numberOfActivities) # 0=start, numberOfActivities=limit
except (GarminConnectConnectionError, GarminConnectAuthenticationError, GarminConnectTooManyRequestsError,) as err:
print("Error occured during Garmin Connect Client init: %s" % err)
quit()
except Exception:
print("Unknown error occured during Garmin Connect Client init.")
# +
fig = px.line(gm_data, y="value")
# add meals and exercise to the chart
yText = 55
eventColor = "green"
for index, row in mealsExerciseSorted.iterrows():
# If the activity has "run" in the description, don't use it as it is a duplicate from Garmin
if "run" in row['description']: continue
# Convert the time in pandas to something that we can use as an index for the x-axis placement
time = datetime.datetime.strptime(row['occurred_at'], '%Y-%m-%d %H:%M:%S')
# Pick a different color depending on the event
if (row['class'] == "Meal"): eventColor = "black"
else: eventColor = "green"
# draw a vertical line at the time of the meal/exercise
fig.add_shape(type="line", xref="x", yref="y", x0=time, y0=70, x1=time , y1=140, line_color=eventColor,)
# Alternate text placement so adjacent text doesn't overlap
if (yText == 55): yText = 60
else: yText = 55
# Add text
fig.add_annotation(text=row['description'], xref="x", yref="y", x=time, y=yText, showarrow=False, font=dict(color=eventColor))
# Add Garmin running activities
for i in range(numberOfActivities):
activity = allActivities[i]
# only activities that are of type "running"
if activity["activityType"]["typeKey"] == "running":
activityDateTime = activity['startTimeLocal']
activityDate = datetime.datetime.strptime(activityDateTime, "%Y-%m-%d %H:%M:%S")
if str(activityDate.date()) == day:
# draw a vertical line at the time of the running activity
fig.add_shape(type="line", xref="x", yref="y", x0=activityDateTime, y0=70, x1=activityDateTime , y1=140, line_color="green",)
# Add text
textDescr = str(activity['activityName']) + " " + str(int(round(activity['distance']/1000))) + "K run"
fig.add_annotation(text=textDescr, xref="x", yref="y", x=activityDateTime, y=65, showarrow=False, font=dict(color="green"))
# Draw a line at the threshold
fig.add_shape(type="line", xref="x", yref="y",
x0=gm_data.index[0], y0=threshold, x1=gm_data.index.max(), y1=threshold, line_color="red",)
# Show text box with summary values
fig.add_annotation(
text='Threshold = '+str(threshold)+
'<br>Minutes above Threshold = '+str(int(round(minutesAboveThreshold,0)))+
'<br>Time above Threshold = '+str(percentageAboveThreshold)+"%"+
'<br>Average Glucose = '+str(averageGlucose)+
'<br>Median Glucose = '+str(medianGlucose),
align='right', showarrow=False,
xref='paper', yref='paper', x=0.002, y=0.005,
bordercolor='black', borderwidth=1
)
# Set x and y axis title and unit
fig.update_xaxes(title_text=str(day), tickformat='%H:%M')
fig.update_yaxes(title_text='mg/dL')
fig.show()
# -
|
chart-one-day-with-garmin.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.1 64-bit (''class_31'': conda)'
# metadata:
# interpreter:
# hash: 500a2b2d21d08ee91c2864f8403870604df3cad1ec0e5a9d161bf391e745d6c7
# name: python3
# ---
from sklearn import datasets
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
import joblib
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Read data into DataFrame from CSV file
# cyclone_df = pd.read_csv("Cyclone_ML.csv")
cyclone_df = pd.read_csv("../data/Cyclone_ML.csv")
# Select features for machine learning and assign to X
selected_features = cyclone_df[["SURFACE_CODE", "CYC_TYPE", "LAT", "LON", "CENTRAL_PRES", "MAX_WIND_SPD", "CENTRAL_INDEX (CI)", "WAVE_HEIGHT"]]
# selected_features = cyclone_df[["CYC_TYPE", "LAT", "LON", "CENTRAL_PRES", "MAX_WIND_SPD"]]
X = selected_features
# Set y to compass direction of cyclone based on wind direction degree
y = cyclone_df["WIND_COMPASS"]
# y = cyclone_df["MAX_REP_WIND_DIR"]
print(X.shape, y.shape)
X
y
# +
# Use train_test_split to create training and testing data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# -
X_scaler = StandardScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
rf = RandomForestClassifier(n_estimators=200)
rf = rf.fit(X_train_scaled, y_train)
training_score = rf.score(X_train_scaled, y_train)
testing_score = rf.score(X_test_scaled, y_test)
print(f"Training Data Score: {training_score}")
print(f"Testing Data Score: {testing_score}")
rf.get_params()
sorted(zip(rf.feature_importances_, selected_features), reverse=True)
# Make predictions with the model
predictions = rf.predict(X_test_scaled)
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(rf, X_test_scaled, y_test, cmap="Blues")
plt.show()
plot_confusion_matrix(rf, X_train_scaled, y_train, cmap="Blues")
plt.show()
from sklearn.metrics import classification_report
print(classification_report(y_test, predictions,
target_names=["E", "N", "NE", "NW", "S", "SE", "SW", "W"]))
joblib.dump(rf, 'cyclone_RF.smd')
print("Model is saved.")
joblib.dump(rf, '../cyclone_RF.smd')
print("Model is saved.")
|
training/cyclone_model_rf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
import tensorflow as tf
from PIL import Image
import numpy as np
from wct import Transfer
# -
# # Load models
transfer = Transfer()
# # Get image features
# +
image = Image.open('deadpool.jpg')
# sizes must be divisible by 16 because of upsampling
# (only if i want to compare images later)
image = image.resize((752, 416), Image.LANCZOS)
image = np.array(image)
features = transfer.get_features(image)
# -
# # Decode and compare
# %%time
X = 4
restored_image = transfer.decode(features[X], X)
restored_image = (np.clip(restored_image, 0, 255)).astype('uint8')
Image.fromarray(np.concatenate([image, restored_image], axis=0))
|
inference/decoder_quality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bounds on Privacy Amplification by Shuffling
# Based on **Balle, Bell, Gascón, and Nissim, *The Privacy Blanket of the Shuffle Model*, CRYPTO 2019**
# This notebook illustrates the use of `amplification_bounds.py` to compute privacy bounds obtained from amplification by shuffling. We refer to the paper for details on how these bounds are obtained. The present implementation was used to produce the plots in the paper.
# ## Mechanisms
#
# Most of the bounds in `amplification_bounds.py` use properties of specific mechanisms (ie. local randomizers). The classes in `mechanisms.py` implement the calculation of these properties for well-known mechanisms.
# +
from shuffleddp.mechanisms import *
# Bounds can be obtained for generic mechanisms satisfying pure local DP
generic = LDPMechanism()
# But bounds are usually better if they incorporate information about the specific randomizer
laplace = LaplaceMechanism()
# By default Randomized Response is over a set with two elements
rr = RRMechanism()
# But the class also accepts a paratemeter 'k' to determine the size of the alphabet
rrk = RRMechanism(k=100)
mechanisms = [generic, laplace, rr, rrk]
# -
# By default all mechanisms are initialized to be 1-LDP
print(generic.get_eps0())
# This can be modified at construction or by calling set_eps0
eps0 = 4
for m in mechanisms:
m.set_eps0(eps0)
print(generic.get_eps0())
# By giving a target epsilon ('eps') after shuffling we can compute properties
# of the privacy amplification random variable for each mechanism.
# These properties are used in the amplification bounds.
eps = 0.25
{m.get_name(): (m.get_gamma(), m.get_max_l(eps), m.get_range_l(eps), m.get_var_l(eps)) for m in mechanisms}
# ## Bounds
#
# Several bounds are implements in `amplification_bounds.py`:
# 1. The closed-form bound of Erlingsson et al. [SODA'19]
# 2. A numeric bound based on Hoeffding's inequality
# 3. A numeric bound based on Bennett's inequality
# +
from shuffleddp.amplification_bounds import *
# This bound does not use infomation about the underlying mechanism
erlingsson = Erlingsson()
all_bounds = [erlingsson]
# The other two bounds can use a 'Generic' mechanism or a specific mechanism
bound_types = [Hoeffding, BennettExact]
for m in mechanisms:
for B in bound_types:
all_bounds.append(B(m))
# +
# We can fix an eps0-LDP guarantee for a mechanism and shuffle n copies of it
n = 5000
eps0 = 0.4
# Then we can get a delta parameter for a target epsilon
print(erlingsson.get_delta(0.15, eps0, n))
# Numeric bounds admit a larger range of parameters than the closed-form bound
eps = 0.005
{b.get_name(): b.get_delta(eps, eps0, n) for b in all_bounds}
# -
# Bounds also be used to get an epsilon for a fixed delta
delta = 1e-6
{b.get_name(): b.get_eps(eps0, n, delta) for b in all_bounds}
# Bounds can also be used for calibrating the eps0 parameter of a mechanism
# to achieve a target (eps,delta)-DP guarantee after shuffling
{b.get_name(): b.get_eps0(eps, n, delta) for b in all_bounds}
# ## Plots
#
# It is easier to compare the different bounds by plotting different parameters.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_panel(f, xs, bounds, with_mech=True, debug=False):
fig = plt.figure()
for b in bounds:
ys = list()
for x in xs:
if debug:
print('{}: {}'.format(b.get_name(), x))
ys.append(f(b, x))
if with_mech:
plt.plot(xs, ys, label=b.get_name())
else:
plt.plot(xs, ys, label=b.get_name(with_mech=False))
plt.legend()
# +
# How fast does eps decrease as we increase n?
eps0 = 0.5
delta = 1e-6
ns = np.geomspace(1000, 1000000, num=100, dtype=int)
def eps(bound, n):
return bound.get_eps(eps0, n, delta)
bounds = [Erlingsson(),
Hoeffding(LDPMechanism()),
BennettExact(LDPMechanism())]
plot_panel(eps, ns, bounds, with_mech=False)
plt.xlabel('$n$')
plt.ylabel('$\\varepsilon$')
plt.title('Generic, $\\varepsilon_0 = {:.1f}, \\delta = 10^{}$'.format(eps0, '{-%d}' % np.log10(1/delta)))
plt.xscale('log')
# +
# How large can we set eps0 as the number of users increase?
eps = 0.1
delta = 1e-6
ns = np.geomspace(100, 100000, num=100, dtype=int)
def eps0(bound, n):
return bound.get_eps0(eps, n, delta)
bounds = [Erlingsson(),
Hoeffding(LDPMechanism()),
Hoeffding(RRMechanism(k=2)),
Hoeffding(RRMechanism(k=100)),
Hoeffding(LaplaceMechanism())]
plot_panel(eps0, ns, bounds)
plt.xlabel('$n$')
plt.ylabel('$\\varepsilon_0$')
plt.title('$\\varepsilon = {:.1f}, \\delta = 10^{}$'.format(eps, '{-%d}' % np.log10(1/delta)))
plt.xscale('log')
# +
# How much better is the Bennett bound vs. the Hoeffding bound?
eps = 0.1
delta = 1e-6
ns = np.geomspace(100, 100000, num=100, dtype=int)
def eps0(bound, n):
return bound.get_eps0(eps, n, delta)
mechanisms = [RRMechanism(),
RRMechanism(k=100),
LaplaceMechanism()]
Bounds = [Hoeffding,
BennettExact]
bounds = []
for m in mechanisms:
for B in Bounds:
bounds.append(B(m))
plot_panel(eps0, ns, bounds, with_mech=True)
plt.xlabel('$n$')
plt.ylabel('$\\varepsilon_0$')
plt.title('$\\varepsilon = {:.1f}, \\delta = 10^{}$'.format(eps, '{-%d}' % np.log10(1/delta)))
plt.xscale('log')
# +
# What if k and delta depend on n?
eps = 0.1
ns = np.geomspace(100, 100000, num=50, dtype=int)
def eps0(bound, n):
bound.mechanism.set_k(int(n**(1/3)))
return bound.get_eps0(eps, n, 1.0/n**2)
bounds = [Hoeffding(RRMechanism()),
BennettExact(RRMechanism())]
plot_panel(eps0, ns, bounds, with_mech=False)
plt.xlabel('$n$')
plt.ylabel('$\\varepsilon_0$')
plt.title('$RR_k, \\varepsilon = {}, \\delta = n^{}, k = n^{}$'.format(eps, '{-2}', '{1/3}'))
plt.xscale('log')
|
Bounds on Privacy Amplification by Shuffling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('base')
# language: python
# name: python3
# ---
lista = [1,2,3,4,5]
n1, n2, *n = lista
print(n1,n2,n)
lista = [1,2,3,4,5]
print(*lista,sep = '-')
# +
def func(*args):
args = list(args)
args[0] = 20
print(args)
func(1,2,3,4,5,6.6)
# -
def func(*args):
print(args)
lista = [1,2,3,4,5]
func(*lista) #já mandar desempacotado
# +
def soma(*args):
print(type(args))
s = 0
for c, v in enumerate(args):
s += v
return s
valor = soma(2,3,4,5,6,7,8)
print(valor)
# -
#uso do kwargs:1 maneira
def func(*args,**kwargs):
print(args,kwargs)
print(kwargs)
print(kwargs['nome'])
lista = [1,2,3,4,5]
func(*lista,nome = 'Matheus')
#uso do kwargs: 2 maneira
def func(*args,**kwargs):
nome = kwargs.get('nome')
print(nome)
idade = kwargs.get('idade')
print(idade) #se o valor estiver ausente, retornará None.
if idade is not None: # verifiquei que idade != None também funciona.
print(f'{nome} tem {idade} anos')
else:
print('Valor não existe!')
lista = [1,2,3,4,5]
func(*lista,nome = 'Matheus',idade = 23)
|
Curso Udemy 2022/Curso_Luiz_Otavio/Aula 56 - args e kargs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework and bake-off: pragmatic color descriptions
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2020"
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [All two-word examples as a dev corpus](#All-two-word-examples-as-a-dev-corpus)
# 1. [Dev dataset](#Dev-dataset)
# 1. [Random train–test split for development](#Random-train–test-split-for-development)
# 1. [Question 1: Improve the tokenizer [1 point]](#Question-1:-Improve-the-tokenizer-[1-point])
# 1. [Use the tokenizer](#Use-the-tokenizer)
# 1. [Question 2: Improve the color representations [1 point]](#Question-2:-Improve-the-color-representations-[1-point])
# 1. [Use the color representer](#Use-the-color-representer)
# 1. [Initial model](#Initial-model)
# 1. [Question 3: GloVe embeddings [1 points]](#Question-3:-GloVe-embeddings-[1-points])
# 1. [Try the GloVe representations](#Try-the-GloVe-representations)
# 1. [Question 4: Color context [3 points]](#Question-4:-Color-context-[3-points])
# 1. [Your original system [3 points]](#Your-original-system-[3-points])
# 1. [Bakeoff [1 point]](#Bakeoff-[1-point])
# ## Overview
#
# This homework and associated bake-off are oriented toward building an effective system for generating color descriptions that are pragmatic in the sense that they would help a reader/listener figure out which color was being referred to in a shared context consisting of a target color (whose identity is known only to the describer/speaker) and a set of distractors.
#
# The notebook [colors_overview.ipynb](colors_overview.ipynb) should be studied before work on this homework begins. That notebook provides backgroud on the task, the dataset, and the modeling code that you will be using and adapting.
#
# The homework questions are more open-ended than previous ones have been. Rather than asking you to implement pre-defined functionality, they ask you to try to improve baseline components of the full system in ways that you find to be effective. As usual, this culiminates in a prompt asking you to develop a novel system for entry into the bake-off. In this case, though, the work you do for the homework will likely be directly incorporated into that system.
# ## Set-up
# See [colors_overview.ipynb](colors_overview.ipynb) for set-up in instructions and other background details.
from colors import ColorsCorpusReader
import os
from sklearn.model_selection import train_test_split
from torch_color_describer import (
ContextualColorDescriber, create_example_dataset)
import utils
from utils import START_SYMBOL, END_SYMBOL, UNK_SYMBOL
utils.fix_random_seeds(set_tensorflow=False)
COLORS_SRC_FILENAME = os.path.join(
"data", "colors", "filteredCorpus.csv")
# ## All two-word examples as a dev corpus
#
# So that you don't have to sit through excessively long training runs during development, I suggest working with the two-word-only subset of the corpus until you enter into the late stages of system testing.
dev_corpus = ColorsCorpusReader(
COLORS_SRC_FILENAME,
word_count=2,
normalize_colors=True)
dev_examples = list(dev_corpus.read())
# This subset has about one-third the examples of the full corpus:
len(dev_examples)
# We __should__ worry that it's not a fully representative sample. Most of the descriptions in the full corpus are shorter, and a large proportion are longer. So this dataset is mainly for debugging, development, and general hill-climbing. All findings should be validated on the full dataset at some point.
# ## Dev dataset
#
# The first step is to extract the raw color and raw texts from the corpus:
dev_rawcols, dev_texts = zip(*[[ex.colors, ex.contents] for ex in dev_examples])
# The raw color representations are suitable inputs to a model, but the texts are just strings, so they can't really be processed as-is. Question 1 asks you to do some tokenizing!
# ## Random train–test split for development
#
# For the sake of development runs, we create a random train–test split:
dev_rawcols_train, dev_rawcols_test, dev_texts_train, dev_texts_test = \
train_test_split(dev_rawcols, dev_texts)
# ## Question 1: Improve the tokenizer [1 point]
#
# This is the first required question – the first required modification to the default pipeline.
#
# The function `tokenize_example` simply splits its string on whitespace and adds the required start and end symbols:
# +
import nltk
tokenizer = nltk.tokenize.regexp.WordPunctTokenizer()
def strip_endings(s):
if s.endswith('er'):
s = s[-2:]
elif s.endswith('est'):
s = s[-3:]
elif s.endswith('ish'):
s = s[-3:]
return s
def tokenize_example(s):
# Improve me!
s = s.lower()
s = tokenizer.tokenize(s)
s = [strip_endings(_s) for _s in s]
return [START_SYMBOL] + s + [END_SYMBOL]
# -
tokenize_example(dev_texts_train[376])
# __Your task__: Modify `tokenize_example` so that it does something more sophisticated with the input text.
#
# __Notes__:
#
# * There are useful ideas for this in [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142)
# * There is no requirement that you do word-level tokenization. Sub-word and multi-word are options.
# * This question can interact with the size of your vocabulary (see just below), and in turn with decisions about how to use `UNK_SYMBOL`.
#
# __Important__: don't forget to add the start and end symbols, else the resulting models will definitely be terrible!
# ## Use the tokenizer
# Once the tokenizer is working, run the following cell to tokenize your inputs:
# +
dev_seqs_train = [tokenize_example(s) for s in dev_texts_train]
dev_seqs_test = [tokenize_example(s) for s in dev_texts_test]
# -
# We use only the train set to derive a vocabulary for the model:
dev_vocab = sorted({w for toks in dev_seqs_train for w in toks}) + [UNK_SYMBOL]
# It's important that the `UNK_SYMBOL` is included somewhere in this list. Test examples with word not seen in training will be mapped to `UNK_SYMBOL`. If you model's vocab is the same as your train vocab, then `UNK_SYMBOL` will never be encountered during training, so it will be a random vector at test time.
len(dev_vocab)
# ## Question 2: Improve the color representations [1 point]
#
# This is the second required pipeline improvement for the assignment.
#
# The following functions do nothing at all to the raw input colors we get from the corpus.
# +
from itertools import product
import numpy as np
idx = np.array(list(product([0,1,2], repeat=3)))
def represent_color_context(colors):
# Improve me!
return [represent_color(color) for color in colors]
def represent_color(color):
# Improve me!
# color = np.exp((idx * np.array([[color[0]/360, color[1]/200, color[2]/200]])).sum(-1) * np.pi * -2j)
color = np.exp((idx * np.array([[color[0], color[1], color[2]]])).sum(-1) * np.pi * -2j)
color = np.concatenate((color.real.flatten(), color.imag.flatten()), axis=0).flatten().tolist()
return color
# -
represent_color_context(dev_rawcols_train[0])
# __Your task__: Modify `represent_color_context` and/or `represent_color` to represent colors in a new way.
#
# __Notes__:
#
# * The Fourier-transform method of [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142) is a proven choice.
# * You are not required to keep `represent_color`. This might be unnatural if you want to perform an operation on each color trio all at once.
# * For that matter, if you want to process all of the color contexts in the entire data set all at once, that is fine too, as long as you can also perform the operation at test time with an unknown number of examples being tested.
# ## Use the color representer
# The following cell just runs your `represent_color_context` on the train and test sets:
# +
dev_cols_train = [represent_color_context(colors) for colors in dev_rawcols_train]
dev_cols_test = [represent_color_context(colors) for colors in dev_rawcols_test]
# -
# At this point, our preprocessing steps are complete, and we can fit a first model.
# ## Initial model
#
# The first model is configured right now to be a small model run for just a few iterations. It should be enough to get traction, but it's unlikely to be a great model. You are free to modify this configuration if you wish; it is here just for demonstration and testing:
dev_mod = ContextualColorDescriber(
dev_vocab,
embed_dim=10,
hidden_dim=10,
max_iter=5,
batch_size=128)
_ = dev_mod.fit(dev_cols_train, dev_seqs_train)
# As discussed in [colors_overview.ipynb](colors_overview.ipynb), our primary metric is `listener_accuracy`:
dev_mod.listener_accuracy(dev_cols_test, dev_seqs_test)
# We can also see the model's predicted sequences given color context inputs:
dev_mod.predict(dev_cols_test[:1])
dev_seqs_test[:1]
# ## Question 3: GloVe embeddings [1 points]
#
# The above model uses a random initial embedding, as configured by the decoder used by `ContextualColorDescriber`. This homework question asks you to consider using GloVe inputs.
#
# __Your task__: Complete `create_glove_embedding` so that it creates a GloVe embedding based on your model vocabulary. This isn't mean to be analytically challenging, but rather just to create a basis for you to try out other kinds of rich initialization.
GLOVE_HOME = os.path.join('data', 'glove.6B')
# +
def create_glove_embedding(vocab, glove_base_filename='glove.6B.50d.txt'):
# Use `utils.glove2dict` to read in the GloVe file:
##### YOUR CODE HERE
glovedict = utils.glove2dict(os.path.join(GLOVE_HOME, glove_base_filename))
print(list(glovedict.keys())[:100])
# Use `utils.create_pretrained_embedding` to create the embedding.
# This function will, by default, ensure that START_TOKEN,
# END_TOKEN, and UNK_TOKEN are included in the embedding.
##### YOUR CODE HERE
pretrained_embedding, pretrained_vocab = utils.create_pretrained_embedding(glovedict, vocab)
# Be sure to return the embedding you create as well as the
# vocabulary returned by `utils.create_pretrained_embedding`,
# which is likely to have been modified from the input `vocab`.
##### YOUR CODE HERE
return pretrained_embedding, pretrained_vocab
# -
# ## Try the GloVe representations
# Let's see if GloVe helped for our development data:
dev_glove_embedding, dev_glove_vocab = create_glove_embedding(dev_vocab)
# The above might dramatically change your vocabulary, depending on how many items from your vocab are in the Glove space:
len(dev_vocab)
len(dev_glove_vocab)
dev_mod_glove = ContextualColorDescriber(
dev_glove_vocab,
embedding=dev_glove_embedding,
hidden_dim=10,
max_iter=5,
batch_size=128)
_ = dev_mod_glove.fit(dev_cols_train, dev_seqs_train)
dev_mod_glove.listener_accuracy(dev_cols_test, dev_seqs_test)
# You probably saw a small boost, assuming your tokeization scheme leads to good overlap with the GloVe vocabulary. The input representations are larger than in our previous model (at least as I configured things), so we would need to do more runs with higher `max_iter` values to see whether this is worthwhile overall.
# ## Question 4: Color context [3 points]
#
# The final required homework question is the most challenging, but it should set you up to think in much more flexible ways about the underlying model we're using.
#
# The question asks you to modify various model components in `torch_color_describer.py`. The section called [Modifying the core model](colors_overview.ipynb#Modifying-the-core-model) from the core unit notebook provides a number of examples illustrating the basic techniques, so you might review that material if you get stuck here.
#
# __Your task__: [Monroe et al. 2017](https://transacl.org/ojs/index.php/tacl/article/view/1142) append the target color (the final one in the context) to each input token that gets processed by the decoder. The question asks you to subclass the `Decoder` and `EncoderDecoder` from `torch_color_describer.py` so that you can build models that do this.
# __Step 1__: Modify the `Decoder` so that the input vector to the model at each timestep is not just a token representaton `x` but the concatenation of `x` with the representation of the target color.
#
# __Notes__:
#
# * You might notice at this point that the original `Decoder.forward` method has an optional keyword argument `target_colors` that is passed to `Decoder.get_embeddings`. Because this is already in place, all you have to do is modify the `get_embeddings` method to use this argument.
#
# * The change affects the configuration of `self.rnn`, so you need to subclass the `__init__` method as well, so that its `input_size` argument accomodates the embedding as well as the color representations.
#
# * You can do the relevant operations efficiently in pure PyTorch using `repeat_interleave` and `cat`, but the important thing is to get a working implementation – you can always optimize the code later if the ideas prove useful to you.
#
# Here's skeleton code for you to flesh out:
# +
import torch
import torch.nn as nn
from torch_color_describer import Decoder
class ColorContextDecoder(Decoder):
def __init__(self, color_dim, *args, **kwargs):
self.color_dim = color_dim
super().__init__(*args, **kwargs)
# Fix the `self.rnn` attribute:
##### YOUR CODE HERE
self.rnn = nn.GRU(
input_size=self.embed_dim+self.color_dim,
hidden_size=self.hidden_dim,
batch_first=True)
def get_embeddings(self, word_seqs, target_colors=None):
"""You can assume that `target_colors` is a tensor of shape
(m, n), where m is the length of the batch (same as
`word_seqs.shape[0]`) and n is the dimensionality of the
color representations the model is using. The goal is
to attached each color vector i to each of the tokens in
the ith sequence of (the embedded version of) `word_seqs`.
"""
##### YOUR CODE HERE
word_embs = self.embedding(word_seqs)
target_colors = torch.repeat_interleave(target_colors.unsqueeze(1), word_seqs.size(1), dim=1)
return torch.cat([word_embs, target_colors], dim=-1)
# -
# __Step 2__: Modify the `EncoderDecoder`. For this, you just need to make a small change to the `forward` method: extract the target colors from `color_seqs` and feed them to the decoder.
# +
from torch_color_describer import EncoderDecoder
class ColorizedEncoderDecoder(EncoderDecoder):
def forward(self,
color_seqs,
word_seqs,
seq_lengths=None,
hidden=None,
targets=None):
if hidden is None:
hidden = self.encoder(color_seqs)
# Extract the target colors from `color_seqs` and
# feed them to the decoder, which already has a
# `target_colors` keyword.
##### YOUR CODE HERE
output, hidden = self.decoder(
word_seqs, seq_lengths=seq_lengths, hidden=hidden, target_colors=color_seqs[:,-1])
return output, hidden, targets
# -
# __Step 3__: Finally, as in the examples in [Modifying the core model](colors_overview.ipynb#Modifying-the-core-model), you need to modify the `build_graph` method of `ContextualColorDescriber` so that it uses your new `ColorContextDecoder` and `ColorizedEncoderDecoder`. Here's starter code:
# +
from torch_color_describer import Encoder
class ColorizedInputDescriber(ContextualColorDescriber):
def build_graph(self):
# We didn't modify the encoder, so this is
# just copied over from the original:
encoder = Encoder(
color_dim=self.color_dim,
hidden_dim=self.hidden_dim)
# Use your `ColorContextDecoder`, making sure
# to pass in all the keyword arguments coming
# from `ColorizedInputDescriber`:
##### YOUR CODE HERE
decoder = ColorContextDecoder(
color_dim=self.color_dim,
vocab_size=self.vocab_size,
embed_dim=self.embed_dim,
embedding=self.embedding,
hidden_dim=self.hidden_dim)
# Return a `ColorizedEncoderDecoder` that uses
# your encoder and decoder:
##### YOUR CODE HERE
return ColorizedEncoderDecoder(encoder, decoder)
# -
# That's it! Since these modifications are pretty intricate, you might want to use [a toy dataset](colors_overview.ipynb#Toy-problems-for-development-work) to debug it:
toy_color_seqs, toy_word_seqs, toy_vocab = create_example_dataset(
group_size=50, vec_dim=2)
toy_color_seqs_train, toy_color_seqs_test, toy_word_seqs_train, toy_word_seqs_test = \
train_test_split(toy_color_seqs, toy_word_seqs)
toy_mod = ColorizedInputDescriber(
toy_vocab,
embed_dim=10,
hidden_dim=10,
max_iter=100,
batch_size=128)
_ = toy_mod.fit(toy_color_seqs_train, toy_word_seqs_train)
toy_mod.listener_accuracy(toy_color_seqs_test, toy_word_seqs_test)
# If that worked, then you can now try this model on SCC problems!
# ## Your original system [3 points]
# There are many options for your original system, which consists of the full pipeline – all preprocessing and modeling steps. You are free to use any model you like, as long as you subclass `ContextualColorDescriber` in a way that allows its `listener_accuracy` method to behave in the expected way.
#
# So that we can evaluate models in a uniform way for the bake-off, we ask that you modify the function `my_original_system` below so that it accepts a trained instance of your model and does any preprocessing steps required by your model.
#
# If we seek to reproduce your results, we will rerun this entire notebook. Thus, it is fine if your `my_original_system` makes use of functions you wrote or modified above this cell.
def my_original_system(trained_model, color_seqs_test, texts_test):
"""Feel free to modify this code to accommodate the needs of
your system. Just keep in mind that it will get raw corpus
examples as inputs for the bake-off.
"""
# `word_seqs_test` is a list of strings, so tokenize each of
# its elements:
tok_seqs = [tokenize_example(s) for s in texts_test]
col_seqs = [represent_color_context(colors)
for colors in color_seqs_test]
# Return the `listener_accuracy` for your model:
return trained_model.listener_accuracy(col_seqs, tok_seqs)
# If `my_original_system` works on test sets you create from the corpus distribution, then it will works for the bake-off, so consider checking that. For example, this would check that `dev_mod` above passes muster:
my_original_system(dev_mod, dev_rawcols_test, dev_texts_test)
original_glove_embedding, original_glove_vocab = create_glove_embedding(dev_vocab, 'glove.6B.200d.txt')
original_mod = ColorizedInputDescriber(
original_glove_vocab,
embedding=original_glove_embedding,
hidden_dim=10,
max_iter=100,
batch_size=128)
_ = original_mod.fit(dev_cols_train, dev_seqs_train)
my_original_system(original_mod, dev_rawcols_test, dev_texts_test)
original_mod.listener_accuracy(dev_cols_test, dev_seqs_test)
# +
from torch_color_describer import Encoder
class LayerEncoder(nn.Module):
"""Simple Encoder model based on a GRU cell.
Parameters
----------
color_dim : int
hidden_dim : int
"""
def __init__(self, color_dim, hidden_dim, n_layers=1, take_top=True):
super().__init__()
self.color_dim = color_dim
self.hidden_dim = hidden_dim
self.take_top = take_top
self.rnn = nn.GRU(
input_size=self.color_dim,
hidden_size=self.hidden_dim,
batch_first=True,
num_layers=n_layers)
def forward(self, color_seqs):
output, hidden = self.rnn(color_seqs)
if self.take_top:
hidden = hidden[-1].unsqueeze(0)
return hidden
class LayerColorContextDecoder(Decoder):
def __init__(self, color_dim, *args, n_layers=1, **kwargs):
self.color_dim = color_dim
self.n_layers = n_layers
super().__init__(*args, **kwargs)
# Fix the `self.rnn` attribute:
##### YOUR CODE HERE
self.rnn = nn.GRU(
input_size=self.embed_dim+self.color_dim,
hidden_size=self.hidden_dim,
batch_first=True,
num_layers=self.n_layers)
def get_embeddings(self, word_seqs, target_colors=None):
"""You can assume that `target_colors` is a tensor of shape
(m, n), where m is the length of the batch (same as
`word_seqs.shape[0]`) and n is the dimensionality of the
color representations the model is using. The goal is
to attached each color vector i to each of the tokens in
the ith sequence of (the embedded version of) `word_seqs`.
"""
##### YOUR CODE HERE
word_embs = self.embedding(word_seqs)
target_colors = torch.repeat_interleave(target_colors.unsqueeze(1), word_seqs.size(1), dim=1)
return torch.cat([word_embs, target_colors], dim=-1)
class LayerColorizedInputDescriber(ContextualColorDescriber):
def __init__(self, *args, enc_layers=1, dec_layers=1, **kwargs):
self.enc_layers = enc_layers
self.dec_layers = dec_layers
super().__init__(*args, **kwargs)
def build_graph(self):
# We didn't modify the encoder, so this is
# just copied over from the original:
use_same = self.enc_layers == self.dec_layers
encoder = LayerEncoder(
color_dim=self.color_dim,
hidden_dim=self.hidden_dim,
n_layers=self.enc_layers,
take_top=not use_same)
# Use your `ColorContextDecoder`, making sure
# to pass in all the keyword arguments coming
# from `ColorizedInputDescriber`:
##### YOUR CODE HERE
decoder = LayerColorContextDecoder(
color_dim=self.color_dim,
vocab_size=self.vocab_size,
embed_dim=self.embed_dim,
embedding=self.embedding,
hidden_dim=self.hidden_dim,
n_layers=self.dec_layers if use_same else 1)
# Return a `ColorizedEncoderDecoder` that uses
# your encoder and decoder:
##### YOUR CODE HERE
return ColorizedEncoderDecoder(encoder, decoder)
# -
mod_2 = LayerColorizedInputDescriber(
original_glove_vocab,
embedding=original_glove_embedding,
hidden_dim=100,
max_iter=100,
batch_size=128,
enc_layers=3,
dec_layers=3)
_ = mod_2.fit(dev_cols_train, dev_seqs_train)
my_original_system(mod_2, dev_rawcols_test, dev_texts_test)
# In the cell below, please provide a brief technical description of your original system, so that the teaching team can gain an understanding of what it does. This will help us to understand your code and analyze all the submissions to identify patterns and strategies.
# +
# Enter your system description in this cell.
"""
For my system I tried to increase the capacity of the encoder and decoder models.
I increased the hidden size to 100. I also opted to use the 200 dimension GloVE Embeddings.
I also used 3 encoder and decoder layers where I used the hidden staet from the 3 encoders to initialize the 3 decoders as we had discussed in class.
For my Tokenization I followed the tokenization stemming and lowercasing described in Monroe et. al. To implement the word and punctuation splitting I used nltk.
For my color embeddings I also used the transform described in Monroe et. al.
"""
# My peak score was: 0.68874
# Please do not remove this comment.
# -
# ## Bakeoff [1 point]
# For the bake-off, we will release a test set. The announcement will go out on the discussion forum. You will evaluate your custom model from the previous question on these new datasets using your `my_original_system` function. Rules:
#
# 1. Only one evaluation is permitted.
# 1. No additional system tuning is permitted once the bake-off has started.
#
# The cells below this one constitute your bake-off entry.
#
# People who enter will receive the additional homework point, and people whose systems achieve the top score will receive an additional 0.5 points. We will test the top-performing systems ourselves, and only systems for which we can reproduce the reported results will win the extra 0.5 points.
#
# Late entries will be accepted, but they cannot earn the extra 0.5 points. Similarly, you cannot win the bake-off unless your homework is submitted on time.
#
# The announcement will include the details on where to submit your entry.
# +
# Enter your bake-off assessment code in this cell.
COLORS_BAKEOFF_SRC_FILENAME = os.path.join(
"data", "colors", "cs224u-colors-bakeoff-data.csv")
bakeoff_corpus = ColorsCorpusReader(COLORS_BAKEOFF_SRC_FILENAME)
# This code just extracts the colors and texts from the new corpus:
bakeoff_rawcols, bakeoff_texts = zip(*[
[ex.colors, ex.contents] for ex in bakeoff_corpus.read()])
# Original system function call; `my_mod` is your ***trained*** model:
my_original_system(mod_2, bakeoff_rawcols, bakeoff_texts)
# Please do not remove this comment.
# +
# On an otherwise blank line in this cell, please enter
# your listener_accuracy score as reported by the code
# above. Please enter only a number between 0 and 1 inclusive.
# Please do not remove this comment.
0.7434761201378631
# -
|
hw_colors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preliminaries
#
# The `pandas` library allows the user several data structures for different data manipulation tasks:
# 1. Data storage through its `Series` and `DataFrame` data structures.
# 2. Data filtering using multiple methods from the package.
# 3. Reading data from many different file formats such as `csv`, `txt`, `xlsx`, ...
#
# Below we provide a brief overview of the `pandas` functionalities needed for these exercises. The complete documentation can be found on the [`pandas` website](https://pandas.pydata.org/).
#
# ## Pandas data structures
#
# ### Series
# The Pandas Series data structure is similar to a one-dimensional array. It can store any type of data. The values are mutable but the size not.
#
# To create `Series`, we call the `pd.Series()` method and pass an array. A `Series` may also be created from a numpy array.
# +
import pandas as pd
import numpy as np
first_series = pd.Series([1,10,100,1000])
print(first_series)
teams = np.array(['PSV','Ajax','Feyenoord','Twente'])
second_series = pd.Series(teams)
print('\n')
print(second_series)
# -
# ### DataFrame
# One can think of a `DataFrame` as a table with rows and columns (2D structure). The columns can be of a different type (as opposed to `numpy` arrays) and the size of the `DataFrame` is mutable.
#
# To create `DataFrame`, we call the `pd.DataFrame()` method and we can create it from scratch or we can convert a numpy array or a list into a `DataFrame`.
# +
# DataFrame from scratch
first_dataframe = pd.DataFrame({
"Position": [1, 2, 3, 4],
"Team": ['PSV','Ajax','Feyenoord','Twente'],
"GF": [80, 75, 75, 70],
"GA": [30, 25, 40, 60],
"Points": [79, 78, 70, 66]
})
print("From scratch: \n {} \n".format(first_dataframe))
# DataFrme from a list
data = [[1, 2, 3, 4], ['PSV','Ajax','Feyenoord','Twente'],
[80, 75, 75, 70], [30, 25, 40, 60], [79, 78, 70, 66]]
columns = ["Position", "Team", "GF", "GA", "Points"]
second_dataframe = pd.DataFrame(data, index=columns)
print("From list: \n {} \n".format(second_dataframe.T)) # the '.T' operator is explained later on
# DataFrame from numpy array
data = np.array([[1, 2, 3, 4], ['PSV','Ajax','Feyenoord','Twente'],
[80, 75, 75, 70], [30, 25, 40, 60], [79, 78, 70, 66]])
columns = ["Position", "Team", "GF", "GA", "Points"]
third_dataframe = pd.DataFrame(data.T, columns=columns)
print("From numpy array: \n {} \n".format(third_dataframe))
# -
# ### DataFrame attributes
# This section gives a quick overview of some of the `pandas.DataFrame` attributes such as `T`, `index`, `columns`, `iloc`, `loc`, `shape` and `values`.
# transpose the index and columns
print(third_dataframe.T)
# index makes reference to the row labels
print(third_dataframe.index)
# columns makes reference to the column labels
print(third_dataframe.columns)
# iloc allows to access the index by integer-location (e.g. all team names, which are in the second columm)
print(third_dataframe.iloc[:,1])
# loc allows to access the index by label(s)-location (e.g. all team names, which are in the "Team" columm)
print(third_dataframe.loc[0, 'Team'])
# shape returns a tuple with the DataFrame dimension, similar to numpy
print(third_dataframe.shape)
# values return a Numpy representation of the DataFrame data
print(third_dataframe.values)
# ### DataFrame methods
# This section gives a quick overview of some of the `pandas.DataFrame` methods such as `head`, `describe`, `concat`, `groupby`,`rename`, `filter`, `drop` and `isna`. To import data from CSV or MS Excel files, we can make use of `read_csv` and `read_excel`, respectively.
# print the first few rows in your dataset with head()
print(third_dataframe.head()) # In this case, it is not very useful because we don't have thousands of rows
# get the summary statistics of the DataFrame with describe()
print(third_dataframe.describe())
# +
# concatenate (join) DataFrame objects using concat()
# first, we will split the above DataFrame in two different ones
df_a = third_dataframe.loc[[0,1],:]
df_b = third_dataframe.loc[[2,3],:]
print(df_a)
print('\n')
print(df_b)
print('\n')
# now, we concatenate both datasets
df = pd.concat([df_a, df_b])
print(df)
# +
# group the data by certain variable via groupby()
# here, we have grouped the data by goals for, which in this case is 75
group = df.groupby('GF')
print(group.get_group('75'))
# -
# rename() helps you change the column or index names
print(df.rename(columns={'Position':'Pos','Team':'Club'}))
# build a subset of rows or columns of your dataset according to labels via filter()
# here, items refer to the variable names: 'Team' and 'Points'; to select columns, we specify axis=1
print(df.filter(items=['Team', 'Points'], axis=1))
# dropping some labels
print(df.drop(columns=['GF', 'GA']))
# +
# search for NA (not available) entries in the DataFrame
print(df.isna()) # No NA values
print('\n')
# create a pandas Series with a NA value
# the Series as W (winnin matches)
tmp = pd.Series([np.NaN, 25, 24, 19], name="W")
# concatenate the Series with the DataFrame
df = pd.concat([df,tmp], axis = 1)
print(df)
print('\n')
# again, check for NA entries
print(df.isna())
# -
# ## Dataset
#
# For this week exercises we will use a dataset from the Genomics of Drug Sensitivity in Cancer (GDSC) project (https://www.cancerrxgene.org/). In this study (['Iorio et al., Cell, 2016']()), 265 compounds were tested on 1001 cancer cell lines for which different types of -omics data (RNA expression, DNA methylation, Copy Number Alteration, DNA sequencing) are available. This is a valuable resource to look for biomarkers of drugs sensitivity in order to try to understand why cancer patients responds very differently to cancer drugs and find ways to assign the optimal treatment to each patient.
#
# For this exercise we will use a subset of the data, focusing the response to the drug YM155 (Sepantronium bromide) on four cancer types, for a total of 148 cancer cell lines.
#
# | ID | Cancer type |
# |-------------|----------------------------------|
# | COAD/READ | Colorectal adenocarcinoma |
# | NB | Neuroblastoma |
# | KIRC | Kidney renal clear cell carcinoma|
# | BRCA | Breast carcinoma |
#
# We will use the RNA expression data (RMA normalised). Only genes with high variability across cell lines (variance > 5, resulting in 238 genes) have been kept.
#
# Drugs have been tested at different concentration, measuring each time the viability of the cells. Drug sensitivity is measured using the natural log of the fitted IC50 metric, which is defined as the half maximal inhibitory concentration. A lower IC50 corresponds to a more sensitive cell line because a lower amount of drug is sufficient to have a strong response, while a higher IC50 corresponds to a more resistant cell line because more drug is needed for killing the cells.
#
# Based on the IC50 metric, cells can be classified as sensitive or resistant. The classification is done by computing the $z$-score across all cell lines in the GDSC for each drug, and considering as sensitive the ones with $z$-score < 0 and resistant the ones with $z$-score > 0.
#
# The dataset is originally provided as 3 files ([original source](https://www.sciencedirect.com/science/article/pii/S0092867416307462?via%3Dihub)) :
#
# `GDSC_RNA_expression.csv`: gene expression matrix with the cell lines in the rows (148) and the genes in the columns (238).
#
# `GDSC_drug_response.csv`: vector with the cell lines response to the drug YM155 in terms of log(IC50) and as classification in sensitive or resistant.
#
# `GDSC_metadata.csv`: metadata for the 148 cell lines including name, COSMIC ID and tumor type (using the classification from ['The Cancer Genome Atlas TCGA'](https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga))
#
# For convenience, we provide the data already curated.
#
# `RNA_expression_curated.csv`: [148 cell lines , 238 genes]
#
# `drug_response_curated.csv`: [148 cell lines , YM155 drug]
#
# The curated data cam be read as `pandas` `DataFrame`s in the following way:
# +
import pandas as pd
gene_expression = pd.read_csv("./data/RNA_expression_curated.csv", sep=',', header=0, index_col=0)
drug_response = pd.read_csv("./data/drug_response_curated.csv", sep=',', header=0, index_col=0)
# -
# You can use the `DataFrame`s directly as inputs to the the `sklearn` models. The advantage over using `numpy` arrays is that the variable are annotated, i.e. each input and output has a name.
# ## Tools
# The `scikit-learn` library provides the required tools for linear regression/classification and shrinkage, as well as for logistic regression.
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn.linear_model import Lasso
from sklearn.linear_model import LogisticRegression
# Note that the notation used for the hyperparameters in the `scikit-learn` library is different from the one used in the lecture. More specifically, in the lecture $\alpha$ is the tunable parameter to select the compromise between Ridge and Lasso. Whereas, `scikit-learn` library refers to `alpha` as the tunable parameter $\lambda$. Please check the documentation for more details.
# # Exercises
#
# ## Selection of the hyperparameter
#
# Implement cross-validation (using `sklearn.grid_search.GridSearchCV`) to select the `alpha` hyperparameter of `sklearn.linear_model.Lasso`.
# +
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
import numpy as np
def scaler(data):
data = (data - np.mean(data))/np.std(data)
return data
X = scaler(gene_expression)
y = scaler(drug_response)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size = 0.1)
lasso = Lasso(tol=1e-2) #tolarance is standard 1e-4 but then it did not converge
alphas = np.logspace(-4, -0.5, 30)
tuned_parameters = [{'alpha': alphas}]
n_folds = 5
clf = GridSearchCV(lasso, tuned_parameters, cv=n_folds, refit=True)
clf.fit(X_train, y_train)
print("Best parameter: {}, with score: {}".format(clf.best_params_, round(clf.best_score_,6)))
print_grid = True
if print_grid:
print("Grid score:")
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/- %0.03f) for %r\n" % (mean, std * 2, params))
final_alpha = clf.best_params_['alpha']
# -
# ## Feature selection
#
# Look at the features selected using the hyperparameter which corresponds to the minimum cross-validation error.
#
# <p><font color='#770a0a'>Is the partition in training and validation sets playing a role in the selection of the hyperparameter? How will this affect the selection of the relevant features?</font></p>
#
# <p><font color='#770a0a'>Should the value of the intercept also be shrunk to zero with Lasso and Ridge regression? Motivate your answer.</font></p>
from sklearn.feature_selection import SelectFromModel
sel_ = SelectFromModel(clf.best_estimator_, prefit=True)
sel_.get_support()
selected_feat = X.columns[(sel_.get_support())]
print(selected_feat)
# ***Answer***: The first run had an alpha of 0.1373823795883264, the second run had an alpha of 0.1813930693911063. The selected features of the first run were 'CDH17', 'ABCB1', 'TSPAN8', 'RARRES3', 'SELENBP1', 'FABP1', 'CDX2', 'AGR3' and for the second run 'ABCB1', 'FABP1', 'CDX2'. This means different features are selected based on the train data, because the train data from the 2nd run was different than in the first run. Concluding, partition in training and validation plays a role in the selection of the hyperparameter and also in the selection of the relevant features.
# ***Answer***: No, the Lasso and Ridge regression does not affect the value of the intercept. The estimators of the other values will chance if you set the intercept to 0. With this you imply to have knowledge that these values should fit on the line X = Y, which is not the case.
# ## Bias-variance
#
# Show the effect of the regularization on the parameter estimates in terms of bias and variance. For this you can repeat the optimization 100 times using bootstrap and visualise the profile of the Lasso regression coefficient over a grid of the hyperparameter, optionally including the variability as error bars.
#
# <p><font color='#770a0a'>Based on the visual analysis of the plot, what are your observation on bias and variance in relation to model complexity? Motivate your answer.</font></p>
# +
from sklearn.utils import resample
# prepare bootstrap sample -- TESTING
# boot_trainx, boot_trainy = resample(X, y, replace=True, n_samples=int(0.9*len(X)), random_state=1) #90 PROCENT ALS SAMPLES?
# print('Bootstrap Sample: {} {}'.format(boot_trainx,boot_trainy))
#moet je nou voor al die 100 weer een gridsearch doen??
def optimization(x_train, y_train, print_result = False, print_grid = False):
lasso = Lasso(tol=1e-2) #tolarance is standard 1e-4 but then it did not converge
alphas = np.logspace(-5, -2, 20)
tuned_parameters = [{'alpha': alphas}]
n_folds = 5
clf = GridSearchCV(lasso, tuned_parameters, cv=n_folds, refit=True, scoring='neg_mean_squared_error')
clf.fit(X_train, y_train)
if print_result:
print("Best parameter: {}, with score: {}".format(clf.best_params_, round(clf.best_score_,6)))
if print_grid:
print("Grid score:")
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
clf.predict(X_test)
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/- %0.03f) for %r\n" % (mean, std * 2, params))
test_scores = clf.cv_results_['mean_test_score']
test_scores = abs(test_scores)
return test_scores, alphas
#Now for each train we do this optimalization and we add all to a dictionary
results_a1 = list()
results_a2 = list()
results_a3 = list()
results_a4 = list()
results_a5 = list()
results_a6 = list()
results_a7 = list()
results_a8 = list()
results_a9 = list()
results_a10 = list()
results_a11 = list()
results_a12 = list()
results_a13 = list()
results_a14 = list()
results_a15 = list()
results_a16 = list()
results_a17 = list()
results_a18 = list()
results_a19 = list()
results_a20 = list()
for i in range(1,101): #101
print("RUN: {}".format(i))
#make the bootstrap
boot_trainx, boot_trainy= resample(X, y, replace=True, n_samples=int(0.9*len(X)))
#perform optimization
nonzero_coef_list = list()
test_scores, alphas = optimization(boot_trainx, boot_trainy, print_result=True)
results_a1.append(test_scores[0])
results_a2.append(test_scores[1])
results_a3.append(test_scores[2])
results_a4.append(test_scores[3])
results_a5.append(test_scores[4])
results_a6.append(test_scores[5])
results_a7.append(test_scores[6])
results_a8.append(test_scores[7])
results_a9.append(test_scores[8])
results_a10.append(test_scores[9])
results_a11.append(test_scores[10])
results_a12.append(test_scores[11])
results_a13.append(test_scores[12])
results_a14.append(test_scores[13])
results_a15.append(test_scores[14])
results_a16.append(test_scores[15])
results_a17.append(test_scores[16])
results_a18.append(test_scores[17])
results_a19.append(test_scores[18])
results_a20.append(test_scores[19])
print(alphas)
# +
#Now we determine the mean and variance per alpha
import matplotlib.pyplot as plt
def create_mean_std(results):
means = list()
varsa = list()
for result in results:
means.append(np.mean(result))
varsa.append(np.std(result))
return means, varsa
result_list = [results_a1, results_a2, results_a3, results_a4, results_a5, results_a6, results_a7, results_a8, results_a9, results_a10,
results_a11, results_a12, results_a13, results_a14, results_a15, results_a16, results_a17, results_a18, results_a19, results_a20]
means, varsa= create_mean_std(result_list)
plt.errorbar(alphas, means, yerr=[varsa, varsa], fmt='ro', ecolor='k', capsize=10, elinewidth=0.5)
plt.xscale('log')
plt.xlabel('Lambda')
plt.ylabel('MSE')
plt.show()
# -
# ***Answer***: the variance decreases when lambda increases. This means that the model with the least nonzero coefficients performs the best based on the mean squared error. We expect this to a certain extend, however with the most optimal lambda the percentage of nonzero coefficients is really low, which we did not expect.
# ## Logistic regression
#
# <p><font color='#770a0a'>Write the expression of the objective function for the penalized logistic regression with $L_1$ and $L_2$ regularisation (as in Elastic net).</font></p>
# The expression of the objective function is
#
# $$ L_{enet}(\hat{\beta})= \frac{\sum_{i=1}^{n}(y_i -X_i \hat{\beta})^2}{2n} + \lambda \bigg(\frac{1-\alpha}{2}\sum_{j=1}^{m} \hat{\beta}_j^2 + \alpha \sum_{j=1}^{m} |\hat{\beta}_j|\bigg)$$
#
# with $$\frac{\sum_{i=1}^{n}(y_i -X_i \hat{\beta})^2}{2n}= RSS$$.
#
# From which follows $$\hat{\beta}^{elasticnet} = \text{argmin} \bigg(RSS+\lambda \sum_{j=1}^{p} (\alpha \beta_j^2 + (1-\alpha) |\beta_j|)\bigg)$$.
|
practicals/week_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import copy
import dgl
import matplotlib
import matplotlib.cm as cm
import torch
from functools import partial
from IPython.display import SVG, display
from rdkit import Chem
from rdkit.Chem import rdDepictor
from rdkit.Chem.Draw import rdMolDraw2D
from dgllife.data import PubChemBioAssayAromaticity
from dgllife.data import Tox21
from dgllife.model import load_pretrained
from dgllife.utils.featurizers import BaseAtomFeaturizer, ConcatFeaturizer, atom_type_one_hot, \
atom_degree_one_hot, atom_formal_charge, atom_num_radical_electrons, \
atom_hybridization_one_hot, atom_total_num_H_one_hot, BaseBondFeaturizer
# -
# Prepare the model, dataset and the features.
# +
# Prepare featurization functions for atoms and bonds
def chirality(atom):
try:
return one_hot_encoding(atom.GetProp('_CIPCode'), ['R', 'S']) + \
[atom.HasProp('_ChiralityPossible')]
except:
return [False, False] + [atom.HasProp('_ChiralityPossible')]
atom_featurizer = BaseAtomFeaturizer(featurizer_funcs={'hv': ConcatFeaturizer([
partial(atom_type_one_hot,
allowable_set=['B', 'C', 'N', 'O', 'F', 'Si', 'P', 'S', 'Cl',
'As', 'Se', 'Br', 'Te', 'I', 'At'],
encode_unknown=True),
partial(atom_degree_one_hot, allowable_set=list(range(6))),
atom_formal_charge, atom_num_radical_electrons,
partial(atom_hybridization_one_hot, encode_unknown=True),
lambda atom: [0], # A placeholder for aromatic information,
atom_total_num_H_one_hot, chirality])})
bond_featurizer = BaseBondFeaturizer({'he': lambda bond: [0 for _ in range(10)]})
#dataset = PubChemBioAssayAromaticity(node_featurizer=atom_featurizer, edge_featurizer=bond_featurizer)
dataset = Tox21(node_featurizer=atom_featurizer, edge_featurizer=bond_featurizer)
model = load_pretrained('AttentiveFP_Aromaticity').eval()
# -
def draw(mol_id, dataset, timestep):
"""Visualize the learned atom weights in readout.
Parameters
----------
mol_id : int
Index for the molecule to visualize in the dataset.
dataset
timestep : int
As the model has multiple rounds of readout, an additional
index is used to specify the round for the weights.
"""
# Get the weights from the model.
smiles, g, _, _ = dataset[mol_id]
print(smiles)
g = dgl.batch([g])
atom_feats, bond_feats = g.ndata.pop('hv'), g.edata.pop('he')
_, atom_weights = model(g, atom_feats, bond_feats, get_node_weight=True)
assert timestep < len(atom_weights), 'Unexpected id for the readout round.'
atom_weights = atom_weights[timestep]
min_value = torch.min(atom_weights)
max_value = torch.max(atom_weights)
atom_weights = (atom_weights - min_value) / (max_value - min_value)
# Conver the weights to atom colors
norm = matplotlib.colors.Normalize(vmin=0, vmax=1.28)
cmap = cm.get_cmap('Oranges')
plt_colors = cm.ScalarMappable(norm=norm, cmap=cmap)
atom_colors = {i: plt_colors.to_rgba(atom_weights[i].data.item()) for i in range(g.number_of_nodes())}
mol = Chem.MolFromSmiles(smiles)
rdDepictor.Compute2DCoords(mol)
drawer = rdMolDraw2D.MolDraw2DSVG(280,280)
drawer.SetFontSize(1)
op = drawer.drawOptions()
mol = rdMolDraw2D.PrepareMolForDrawing(mol)
drawer.DrawMolecule(mol,highlightAtoms=range(g.number_of_nodes()),highlightBonds=[],
highlightAtomColors=atom_colors)
drawer.FinishDrawing()
svg = drawer.GetDrawingText()
svg = svg.replace('svg:','')
display(SVG(svg))
draw(978, dataset, 0)
|
atom_weight_visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# Delete this cell to re-enable tracebacks
import sys
ipython = get_ipython()
def hide_traceback(exc_tuple=None, filename=None, tb_offset=None,
exception_only=False, running_compiled_code=False):
etype, value, tb = sys.exc_info()
value.__cause__ = None # suppress chained exceptions
return ipython._showtraceback(etype, value, ipython.InteractiveTB.get_exception_only(etype, value))
ipython.showtraceback = hide_traceback
# + nbsphinx="hidden"
# JSON output syntax highlighting
from __future__ import print_function
from pygments import highlight
from pygments.lexers import JsonLexer, TextLexer
from pygments.formatters import HtmlFormatter
from IPython.display import display, HTML
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
def json_print(inpt):
string = str(inpt)
formatter = HtmlFormatter()
if string[0] == '{':
lexer = JsonLexer()
else:
lexer = TextLexer()
return HTML('<style type="text/css">{}</style>{}'.format(
formatter.get_style_defs('.highlight'),
highlight(string, lexer, formatter)))
globals()['print'] = json_print
# -
# ## Checking Semantic Equivalence
#
# The [Environment](../api/stix2.environment.rst#stix2.environment.Environment) has a function for checking if two STIX Objects are semantically equivalent. For each supported object type, the algorithm checks if the values for a specific set of properties match. Then each matching property is weighted since every property doesn't represent the same level of importance for semantic equivalence. The result will be the sum of these weighted values, in the range of 0 to 100. A result of 0 means that the the two objects are not equivalent, and a result of 100 means that they are equivalent.
#
# TODO: Add a link to the committee note when it is released.
#
# There are a number of use cases for which calculating semantic equivalence may be helpful. It can be used for echo detection, in which a STIX producer who consumes content from other producers wants to make sure they are not creating content they have already seen or consuming content they have already created.
#
# Another use case for this functionality is to identify identical or near-identical content, such as a vulnerability shared under three different nicknames by three different STIX producers. A third use case involves a feed that aggregates data from multiple other sources. It will want to make sure that it is not publishing duplicate data.
#
# Below we will show examples of the semantic equivalence results of various objects. Unless otherwise specified, the ID of each object will be generated by the library, so the two objects will not have the same ID. This demonstrates that the semantic equivalence algorithm only looks at specific properties for each object type.
#
# **Please note** that you will need to install a few extra dependencies in order to use the semantic equivalence functions. You can do this using:
#
# ```pip install stix2[semantic]```
#
# ### Attack Pattern Example
#
# For Attack Patterns, the only properties that contribute to semantic equivalence are `name` and `external_references`, with weights of 30 and 70, respectively. In this example, both attack patterns have the same external reference but the second has a slightly different yet still similar name.
# +
import stix2
from stix2 import AttackPattern, Environment, MemoryStore
env = Environment(store=MemoryStore())
ap1 = AttackPattern(
name="Phishing",
external_references=[
{
"url": "https://example2",
"source_name": "some-source2",
},
],
)
ap2 = AttackPattern(
name="Spear phishing",
external_references=[
{
"url": "https://example2",
"source_name": "some-source2",
},
],
)
print(env.semantically_equivalent(ap1, ap2))
# -
# ### Campaign Example
#
# For Campaigns, the only properties that contribute to semantic equivalence are `name` and `aliases`, with weights of 60 and 40, respectively. In this example, the two campaigns have completely different names, but slightly similar descriptions. The result may be higher than expected because the Jaro-Winkler algorithm used to compare string properties looks at the edit distance of the two strings rather than just the words in them.
# +
from stix2 import Campaign
c1 = Campaign(
name="Someone Attacks Somebody",)
c2 = Campaign(
name="Another Campaign",)
print(env.semantically_equivalent(c1, c2))
# -
# ### Identity Example
#
# For Identities, the only properties that contribute to semantic equivalence are `name`, `identity_class`, and `sectors`, with weights of 60, 20, and 20, respectively. In this example, the two identities are identical, but are missing one of the contributing properties. The algorithm only compares properties that are actually present on the objects. Also note that they have completely different description properties, but because description is not one of the properties considered for semantic equivalence, this difference has no effect on the result.
# +
from stix2 import Identity
id1 = Identity(
name="<NAME>",
identity_class="individual",
description="Just some guy",
)
id2 = Identity(
name="<NAME>",
identity_class="individual",
description="A person",
)
print(env.semantically_equivalent(id1, id2))
# -
# ### Indicator Example
#
# For Indicators, the only properties that contribute to semantic equivalence are `indicator_types`, `pattern`, and `valid_from`, with weights of 15, 80, and 5, respectively. In this example, the two indicators have patterns with different hashes but the same indicator_type and valid_from. For patterns, the algorithm currently only checks if they are identical.
# +
from stix2.v21 import Indicator
ind1 = Indicator(
indicator_types=['malicious-activity'],
pattern_type="stix",
pattern="[file:hashes.MD5 = 'd41d8cd98f00b204e9800998ecf8427e']",
valid_from="2017-01-01T12:34:56Z",
)
ind2 = Indicator(
indicator_types=['malicious-activity'],
pattern_type="stix",
pattern="[file:hashes.MD5 = '79054025255fb1a26e4bc422aef54eb4']",
valid_from="2017-01-01T12:34:56Z",
)
print(env.semantically_equivalent(ind1, ind2))
# -
# If the patterns were identical the result would have been 100.
# ### Location Example
#
# For Locations, the only properties that contribute to semantic equivalence are `longitude`/`latitude`, `region`, and `country`, with weights of 34, 33, and 33, respectively. In this example, the two locations are Washington, D.C. and New York City. The algorithm computes the distance between two locations using the haversine formula and uses that to influence equivalence.
# +
from stix2 import Location
loc1 = Location(
latitude=38.889,
longitude=-77.023,
)
loc2 = Location(
latitude=40.713,
longitude=-74.006,
)
print(env.semantically_equivalent(loc1, loc2))
# -
# ### Malware Example
#
# For Malware, the only properties that contribute to semantic equivalence are `malware_types` and `name`, with weights of 20 and 80, respectively. In this example, the two malware objects only differ in the strings in their malware_types lists. For lists, the algorithm bases its calculations on the intersection of the two lists. An empty intersection will result in a 0, and a complete intersection will result in a 1 for that property.
# +
from stix2 import Malware
MALWARE_ID = "malware--9c4638ec-f1de-4ddb-abf4-1b760417654e"
mal1 = Malware(id=MALWARE_ID,
malware_types=['ransomware'],
name="Cryptolocker",
is_family=False,
)
mal2 = Malware(id=MALWARE_ID,
malware_types=['ransomware', 'dropper'],
name="Cryptolocker",
is_family=False,
)
print(env.semantically_equivalent(mal1, mal2))
# -
# ### Threat Actor Example
#
# For Threat Actors, the only properties that contribute to semantic equivalence are `threat_actor_types`, `name`, and `aliases`, with weights of 20, 60, and 20, respectively. In this example, the two threat actors have the same id properties but everything else is different. Since the id property does not factor into semantic equivalence, the result is not very high. The result is not zero because of the "Token Sort Ratio" algorithm used to compare the `name` property.
# +
from stix2 import ThreatActor
THREAT_ACTOR_ID = "threat-actor--8e2e2d2b-17d4-4cbf-938f-98ee46b3cd3f"
ta1 = ThreatActor(id=THREAT_ACTOR_ID,
threat_actor_types=["crime-syndicate"],
name="<NAME>",
aliases=["super-evil"],
)
ta2 = ThreatActor(id=THREAT_ACTOR_ID,
threat_actor_types=["spy"],
name="<NAME>",
aliases=["007"],
)
print(env.semantically_equivalent(ta1, ta2))
# -
# ### Tool Example
#
# For Tools, the only properties that contribute to semantic equivalence are `tool_types` and `name`, with weights of 20 and 80, respectively. In this example, the two tools have the same values for properties that contribute to semantic equivalence but one has an additional, non-contributing property.
# +
from stix2 import Tool
t1 = Tool(
tool_types=["remote-access"],
name="VNC",
)
t2 = Tool(
tool_types=["remote-access"],
name="VNC",
description="This is a tool"
)
print(env.semantically_equivalent(t1, t2))
# -
# ### Vulnerability Example
#
# For Vulnerabilities, the only properties that contribute to semantic equivalence are `name` and `external_references`, with weights of 30 and 70, respectively. In this example, the two vulnerabilities have the same name but one also has an external reference. The algorithm doesn't take into account any semantic equivalence contributing properties that are not present on both objects.
# +
from stix2 import Vulnerability
vuln1 = Vulnerability(
name="Heartbleed",
external_references=[
{
"url": "https://example",
"source_name": "some-source",
},
],
)
vuln2 = Vulnerability(
name="Heartbleed",
)
print(env.semantically_equivalent(vuln1, vuln2))
# -
# ### Other Examples
#
# Comparing objects of different types will result in a `ValueError`.
print(env.semantically_equivalent(ind1, vuln1))
# Some object types do not have a defined method for calculating semantic equivalence and by default will give a warning and a result of zero.
# +
from stix2 import Report
r1 = Report(
report_types=["campaign"],
name="Bad Cybercrime",
published="2016-04-06T20:03:00.000Z",
object_refs=["indicator--a740531e-63ff-4e49-a9e1-a0a3eed0e3e7"],
)
r2 = Report(
report_types=["campaign"],
name="Bad Cybercrime",
published="2016-04-06T20:03:00.000Z",
object_refs=["indicator--a740531e-63ff-4e49-a9e1-a0a3eed0e3e7"],
)
print(env.semantically_equivalent(r1, r2))
# -
# By default, comparing objects of different spec versions will result in a `ValueError`.
# +
from stix2.v20 import Identity as Identity20
id20 = Identity20(
name="<NAME>",
identity_class="individual",
)
print(env.semantically_equivalent(id2, id20))
# -
# You can optionally allow comparing across spec versions by providing a configuration dictionary using `ignore_spec_version` like in the next example:
# +
from stix2.v20 import Identity as Identity20
id20 = Identity20(
name="<NAME>",
identity_class="individual",
)
print(env.semantically_equivalent(id2, id20, **{"_internal": {"ignore_spec_version": True}}))
# -
# ### Detailed Results
#
# If your logging level is set to `DEBUG` or higher, the function will log more detailed results. These show the semantic equivalence and weighting for each property that is checked, to show how the final result was arrived at.
# +
import logging
logging.basicConfig(format='%(message)s')
logger = logging.getLogger()
logger.setLevel(logging.DEBUG)
ta3 = ThreatActor(
threat_actor_types=["crime-syndicate"],
name="Evil Org",
aliases=["super-evil"],
)
ta4 = ThreatActor(
threat_actor_types=["spy"],
name="<NAME>",
aliases=["007"],
)
print(env.semantically_equivalent(ta3, ta4))
logger.setLevel(logging.ERROR)
# -
# You can also retrieve the detailed results in a dictionary so the detailed results information can be accessed and used more programatically. The [semantically_equivalent()](../api/stix2.environment.rst#stix2.environment.Environment.semantically_equivalent) function takes an optional third argument, called `prop_scores`. This argument should be a dictionary into which the detailed debugging information will be stored.
#
# Using `prop_scores` is simple: simply pass in a dictionary to `semantically_equivalent()`, and after the function is done executing, the dictionary will have the various scores in it. Specifically, it will have the overall `matching_score` and `sum_weights`, along with the weight and contributing score for each of the semantic equivalence contributing properties.
#
# For example:
# +
ta5 = ThreatActor(
threat_actor_types=["crime-syndicate", "spy"],
name="<NAME>",
aliases=["super-evil"],
)
ta6 = ThreatActor(
threat_actor_types=["spy"],
name="<NAME>",
aliases=["007"],
)
prop_scores = {}
print("Semantic equivalence score using standard weights: %s" % (env.semantically_equivalent(ta5, ta6, prop_scores)))
print(prop_scores)
for prop in prop_scores:
if prop not in ["matching_score", "sum_weights"]:
print ("Prop: %s | weight: %s | contributing_score: %s" % (prop, prop_scores[prop]['weight'], prop_scores[prop]['contributing_score']))
else:
print ("%s: %s" % (prop, prop_scores[prop]))
# -
# ### Custom Comparisons
# If you wish, you can customize semantic equivalence comparisons. Specifically, you can do any of three things:
# - Provide custom weights for each semantic equivalence contributing property
# - Provide custom comparison functions for individual semantic equivalence contributing properties
# - Provide a custom semantic equivalence function for a specific object type
#
# #### The `weights` dictionary
# In order to do any of the aforementioned (*optional*) custom comparisons, you will need to provide a `weights` dictionary as the last parameter to the [semantically_equivalent()](../api/stix2.environment.rst#stix2.environment.Environment.semantically_equivalent) method call.
#
# The weights dictionary should contain both the weight and the comparison function for each property. You may use the default weights and functions, or provide your own.
#
# ##### Existing comparison functions
# For reference, here is a list of the comparison functions already built in the codebase (found in [stix2/environment.py](../api/stix2.environment.rst#stix2.environment.Environment)):
#
# - [custom_pattern_based](../api/stix2.environment.rst#stix2.environment.custom_pattern_based)
# - [exact_match](../api/stix2.environment.rst#stix2.environment.exact_match)
# - [partial_external_reference_based](../api/stix2.environment.rst#stix2.environment.partial_external_reference_based)
# - [partial_list_based](../api/stix2.environment.rst#stix2.environment.partial_list_based)
# - [partial_location_distance](../api/stix2.environment.rst#stix2.environment.partial_location_distance)
# - [partial_string_based](../api/stix2.environment.rst#stix2.environment.partial_string_based)
# - [partial_timestamp_based](../api/stix2.environment.rst#stix2.environment.partial_timestamp_based)
#
# For instance, if we wanted to compare two of the `ThreatActor`s from before, but use our own weights, then we could do the following:
# +
weights = {
"threat-actor": { # You must specify the object type
"name": (30, stix2.environment.partial_string_based), # Each property's value must be a tuple
"threat_actor_types": (50, stix2.environment.partial_list_based), # The 1st component must be the weight
"aliases": (20, stix2.environment.partial_list_based) # The 2nd component must be the comparison function
}
}
print("Using standard weights: %s" % (env.semantically_equivalent(ta5, ta6)))
print("Using custom weights: %s" % (env.semantically_equivalent(ta5, ta6, **weights)))
# -
# Notice how there is a difference in the semantic equivalence scores, simply due to the fact that custom weights were used.
#
# #### Custom Weights With prop_scores
# If we want to use both `prop_scores` and `weights`, then they would be the third and fourth arguments, respectively, to `sematically_equivalent()`:
prop_scores = {}
weights = {
"threat-actor": {
"name": (45, stix2.environment.partial_string_based),
"threat_actor_types": (10, stix2.environment.partial_list_based),
"aliases": (45, stix2.environment.partial_list_based),
},
}
env.semantically_equivalent(ta5, ta6, prop_scores, **weights)
print(prop_scores)
# #### Custom Semantic Equivalence Functions
# You can also write and use your own semantic equivalence functions. In the examples above, you could replace the built-in comparison functions for any or all properties. For example, here we use a custom string comparison function just for the `'name'` property:
# +
def my_string_compare(p1, p2):
if p1 == p2:
return 1
else:
return 0
weights = {
"threat-actor": {
"name": (45, my_string_compare),
"threat_actor_types": (10, stix2.environment.partial_list_based),
"aliases": (45, stix2.environment.partial_list_based),
},
}
print("Using custom string comparison: %s" % (env.semantically_equivalent(ta5, ta6, **weights)))
# -
# You can also customize the comparison of an entire object type instead of just how each property is compared. To do this, provide a `weights` dictionary to `semantically_equivalent()` and in this dictionary include a key of `"method"` whose value is your custom semantic equivalence function for that object type.
#
# If you provide your own custom semantic equivalence method, you **must also provide the weights for each of the properties** (unless, for some reason, your custom method is weights-agnostic). However, since you are writing the custom method, your weights need not necessarily follow the tuple format specified in the above code box.
#
# Note also that if you want detailed results with `prop_scores` you will need to implement that in your custom function, but you are not required to do so.
#
# In this next example we use our own custom semantic equivalence function to compare two `ThreatActor`s, and do not support `prop_scores`.
# +
def custom_semantic_equivalence_method(obj1, obj2, **weights):
sum_weights = 0
matching_score = 0
# Compare name
w = weights['name']
sum_weights += w
contributing_score = w * stix2.environment.partial_string_based(obj1['name'], obj2['name'])
matching_score += contributing_score
# Compare aliases only for spies
if 'spy' in obj1['threat_actor_types'] + obj2['threat_actor_types']:
w = weights['aliases']
sum_weights += w
contributing_score = w * stix2.environment.partial_list_based(obj1['aliases'], obj2['aliases'])
matching_score += contributing_score
return matching_score, sum_weights
weights = {
"threat-actor": {
"name": 60,
"aliases": 40,
"method": custom_semantic_equivalence_method
}
}
print("Using standard weights: %s" % (env.semantically_equivalent(ta5, ta6)))
print("Using a custom method: %s" % (env.semantically_equivalent(ta5, ta6, **weights)))
# -
# You can also write custom functions for comparing objects of your own custom types. Like in the previous example, you can use the built-in functions listed above to help with this, or write your own. In the following example we define semantic equivalence for our new `x-foobar` object type. Notice that this time we have included support for detailed results with `prop_scores`.
# +
def _x_foobar_checks(obj1, obj2, prop_scores, **weights):
matching_score = 0.0
sum_weights = 0.0
if stix2.environment.check_property_present("name", obj1, obj2):
w = weights["name"]
sum_weights += w
contributing_score = w * stix2.environment.partial_string_based(obj1["name"], obj2["name"])
matching_score += contributing_score
prop_scores["name"] = (w, contributing_score)
if stix2.environment.check_property_present("color", obj1, obj2):
w = weights["color"]
sum_weights += w
contributing_score = w * stix2.environment.partial_string_based(obj1["color"], obj2["color"])
matching_score += contributing_score
prop_scores["color"] = (w, contributing_score)
prop_scores["matching_score"] = matching_score
prop_scores["sum_weights"] = sum_weights
return matching_score, sum_weights
prop_scores = {}
weights = {
"x-foobar": {
"name": 60,
"color": 40,
"method": _x_foobar_checks,
},
"_internal": {
"ignore_spec_version": False,
},
}
foo1 = {
"type":"x-foobar",
"id":"x-foobar--0c7b5b88-8ff7-4a4d-aa9d-feb398cd0061",
"name": "Zot",
"color": "red",
}
foo2 = {
"type":"x-foobar",
"id":"x-foobar--0c7b5b88-8ff7-4a4d-aa9d-feb398cd0061",
"name": "Zot",
"color": "blue",
}
print(env.semantically_equivalent(foo1, foo2, prop_scores, **weights))
print(prop_scores)
|
docs/guide/equivalence.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np # advanced math library
import matplotlib.pyplot as plt # MATLAB like plotting routines
import random # for generating random numbers
from keras.datasets import mnist # MNIST dataset is included in Keras
from keras.models import Sequential # Model type to be used
from keras.layers.core import Dense, Dropout, Activation # Types of layers to be used in our model
from keras.utils import np_utils # NumPy related tools
from keras.preprocessing.image import ImageDataGenerator
from keras.layers import Conv2D, MaxPooling2D, ZeroPadding2D, GlobalAveragePooling2D, Flatten, Reshape
from keras.layers.normalization import BatchNormalization
import keras
# +
# The MNIST data is split between 60,000 28 x 28 pixel training images and 10,000 28 x 28 pixel images
(X_train, y_train), (X_test, y_test) = mnist.load_data()
X_train = X_train.reshape(60000, 28, 28, 1) #add an additional dimension to represent the single-channel
X_test = X_test.reshape(10000, 28, 28, 1)
X_train = X_train.astype('float32') # change integers to 32-bit floating point numbers
X_test = X_test.astype('float32')
X_train /= 255 # normalize each value for each pixel for the entire vector for each input
X_test /= 255
print("Training matrix shape", X_train.shape)
print("Testing matrix shape", X_test.shape)
# +
plt.rcParams['figure.figsize'] = (9,9) # Make the figures a bit bigger
for i in range(9):
plt.subplot(3,3,i+1)
num = random.randint(0, len(X_train))
plt.imshow(X_train[num], cmap='gray', interpolation='none')
plt.title("Class {}".format(y_train[num]))
plt.tight_layout()
# -
# +
nb_classes = 10 # number of unique digits
Y_train = np_utils.to_categorical(y_train, nb_classes)
Y_test = np_utils.to_categorical(y_test, nb_classes)
# +
# The Sequential model is a linear stack of layers and is very common.
model = Sequential() # Linear stacking of layers
# Convolution Layer 1
model.add(Conv2D(32, (3, 3), input_shape=(28,28,1))) # 32 different 3x3 kernels -- so 32 feature maps
model.add(BatchNormalization(axis=-1)) # normalize each feature map before activation
convLayer01 = Activation('relu') # activation
model.add(convLayer01)
# Convolution Layer 2
model.add(Conv2D(32, (3, 3))) # 32 different 3x3 kernels -- so 32 feature maps
model.add(BatchNormalization(axis=-1)) # normalize each feature map before activation
model.add(Activation('relu')) # activation
convLayer02 = MaxPooling2D(pool_size=(2,2)) # Pool the max values over a 2x2 kernel
model.add(convLayer02)
# Convolution Layer 3
model.add(Conv2D(64,(3, 3))) # 64 different 3x3 kernels -- so 64 feature maps
model.add(BatchNormalization(axis=-1)) # normalize each feature map before activation
convLayer03 = Activation('relu') # activation
model.add(convLayer03)
# Convolution Layer 4
model.add(Conv2D(64, (3, 3))) # 64 different 3x3 kernels -- so 64 feature maps
model.add(BatchNormalization(axis=-1)) # normalize each feature map before activation
model.add(Activation('relu')) # activation
convLayer04 = MaxPooling2D(pool_size=(2,2)) # Pool the max values over a 2x2 kernel
model.add(convLayer04)
model.add(Flatten()) # Flatten final 4x4x64 output matrix into a 1024-length vector
# Fully Connected Layer 5
model.add(Dense(512)) # 512 FCN nodes
model.add(BatchNormalization()) # normalization
model.add(Activation('relu')) # activation
# Fully Connected Layer 6
model.add(Dropout(0.2)) # 20% dropout of randomly selected nodes
model.add(Dense(10)) # final 10 FCN nodes
model.add(Activation('softmax')) # softmax activation
# -
model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# +
# data augmentation prevents overfitting by slightly changing the data randomly
# Keras has a great built-in feature to do automatic augmentation
gen = ImageDataGenerator(rotation_range=8, width_shift_range=0.08, shear_range=0.3,
height_shift_range=0.08, zoom_range=0.08)
test_gen = ImageDataGenerator()
# +
# We can then feed our augmented data in batches
# Besides loss function considerations as before, this method actually results in significant memory savings
# because we are actually LOADING the data into the network in batches before processing each batch
# Before the data was all loaded into memory, but then processed in batches.
train_generator = gen.flow(X_train, Y_train, batch_size=128)
test_generator = test_gen.flow(X_test, Y_test, batch_size=128)
# +
# We can now train our model which is fed data by our batch loader
# Steps per epoch should always be total size of the set divided by the batch size
# SIGNIFICANT MEMORY SAVINGS (important for larger, deeper networks)
history = model.fit(train_generator, steps_per_epoch=60000//128, epochs=2, verbose=1,
validation_data=test_generator, validation_steps=10000//128)
# -
score = model.evaluate(X_test, Y_test)
print('Test score:', score[0])
print('Test accuracy:', score[1])
model.save("mlmodels/kerasdnn1/")
#keras.models.load_model("mlmodels/kerasdnn1")
|
keras DNN MNIST model train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#PROGRAM TO COMPUTE FREQUENCY OF WORDS FROM THE INPUT #3
bmw = input().split()
dict = {}
for i in bmw:
i = dict.setdefault(i,bmw.count(i))
dict = sorted(dict.items())
for i in dict:
print("%s:%d"%(i[0],i[1]))
|
python4.3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
# CUDA DEVICE for keras
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
import tensorflow.keras.backend as K
from tensorflow.keras.layers import Dense, Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.activations import softmax
from tensorflow.keras.utils import model_to_dot, plot_model
# +
X = Input(shape=(256, ))
densor1 = Dense(64)
X1 = Dense(128)(X)
X2 = densor1(X1)
X3 = Dense(32)(X2)
model1 = Model(X, X3)
model2 = Model(X, X2)
# -
x_samples = np.random.randn(5, 256)
y3 = np.random.randn(5, 32)
start_lr = 1e-3
model1.compile(optimizer=Adam(start_lr, amsgrad=True), loss='mean_squared_error', metrics=['mse'])
x_samples.shape
y3.shape
model1.summary()
model2.summary()
output1 = model2.predict(x_samples)
layer_weights1 = model2.get_layer('dense_1')
layer_weights1.weights
K.eval(layer_weights1.weights[0])
K.eval(layer_weights1.weights[1])
model1.fit(x_samples, y3, epochs=100)
output2 = model2.predict(x_samples)
layer_weights2 = model2.get_layer('dense_1')
K.eval(layer_weights2.weights[0])
K.eval(layer_weights2.weights[1])
print(np.all(output1 == output2))
|
keras_migration/shareLayer-tf2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model
# ## Importing Data
# +
# Import necessary modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
import seaborn as sns
# Display 100 columns max
pd.set_option('display.max_columns', 100)
# Display 20 characters max within a cell
# pd.set_option('max_colwidth',60)
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Lasso
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import RandomizedSearchCV
from sklearn.metrics import mean_squared_error, mean_absolute_error, r2_score
import mlflow
import pickle
# +
# Import imputed data
df = pd.read_pickle("autos_golden_20190719.pkl")
# Import data without missing values (No imputation, no missing in the data)
df_clean = pd.read_pickle("autos_clean_golden_20190719.pkl")
# Import data with missing values (No imputation)
df_missing = pd.read_pickle("autos_missing_golden_20190719.pkl")
# -
df.shape, df_missing.shape
df_clean.shape
df.head()
# + [markdown] toc-hr-collapsed=false
# ## Model
# -
# Create features and target (df)
X = df.drop('price', axis = 1) # features
y = df['price'] # target
# Create features and target (df)
X_c = df_clean.drop('price', axis = 1) # features
y_c = df_clean['price'] # target
# +
# # Create features and target (array)
# X = df.drop('price', axis = 1).values # features
# y = df['price'].values # target
# -
# Train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state=57)
# Train-test split
X_train_c, X_test_c, y_train_c, y_test_c = train_test_split(X_c, y_c, test_size = 0.3, random_state=57)
# ### Model Evaluation Metrics
def eval_metrics(actual, pred):
rmse = np.sqrt(mean_squared_error(actual, pred))
mae = mean_absolute_error(actual, pred)
r2 = r2_score(actual, pred)
return rmse, mae, r2
# ### Linear Regression
# +
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(interaction_only=True,include_bias = False)
poly.fit_transform(X)
# -
X.columns
# ### Lasso
# +
# Create Alphas, and lists to store scores
alpha_space = np.linspace(1, 10)
lasso_scores = []
# Create a regressor
lasso_ = Lasso(normalize = True)
# Compute scores over range of alphas
for alpha in alpha_space:
# Specify alpha
lasso_.alpha = alpha
# Perfrom 5-fold CV
lasso_cv_scores = cross_val_score(lasso_, X_train, y_train, cv=5)
# Append the mean and std to the lists
lasso_scores.append(np.mean(lasso_cv_scores))
# Log parameter, metrics, and model to MLflow
with mlflow.start_run(run_name="Lasso CV"): # "with" is necessary for multiple runs
mlflow.log_param("alpha", alpha)
mlflow.log_metric("lasso_cv_score", np.mean(lasso_cv_scores))
# +
# #!mlflow ui
# +
# Create a function to display how cv scores changes with alphas
def display_plot(cv_scores):
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(alpha_space, cv_scores)
ax.set_ylabel('CV Score')
ax.set_xlabel('Alpha')
ax.axhline(np.max(cv_scores), linestyle='--', color='.5')
ax.set_xlim([alpha_space[0], alpha_space[-1]])
ax.set_xscale('log')
ax.axvline(x=1.4, c='b', linestyle='--', alpha=0.5)
plt.show()
display_plot(lasso_scores)
# -
# $alpha = 1.4$ seems to be a good choice
# +
# Instantiate Lasso regressor
lasso = Lasso(alpha = 1.4, normalize = True)
# Fit the regressor
lasso.fit(X_train, y_train)
# Print the R-squared
print("R^2: {0:.3f}".format(lasso.score(X_test, y_test)))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, lasso.predict(X_test))
# Log parameter, metrics, and model to MLflow
with mlflow.start_run(run_name="Lasso Regression"):
mlflow.log_param("alpha", 1.4)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
#mlflow.end_run()
# +
# Perform 5-fold cross-validation
cv_scores = cross_val_score(lasso, X_test, y_test, cv=5)
print("Average 5-Fold CV Score: {0:.3f}".format(np.mean(cv_scores)))
# -
# ### Linear Regression
# +
# Create the regressor
reg = LinearRegression()
# Fit the regressor to the training data
reg.fit(X_train, y_train)
# Print the R-squared
print("R^2: {0:.2f}".format(reg.score(X_test, y_test)))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, reg.predict(X_test))
# Log parameter, metrics, and model to MLflow
mlflow.start_run(run_name="Linear Regression")
mlflow.log_param("normalize", False)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
mlflow.end_run()
# +
# Create the regressor
reg = LinearRegression(normalize=True)
# Fit the regressor to the training data
reg.fit(X_train, y_train)
# Print the R-squared
print("R^2: {0:.2f}".format(reg.score(X_test, y_test)))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, reg.predict(X_test))
# Log parameter, metrics, and model to MLflow
mlflow.start_run(run_name="Linear Regression")
mlflow.log_param("normalize", True)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
mlflow.end_run()
# +
# Create the regressor
reg = LinearRegression()
# Fit the regressor to the training data
reg.fit(X_train_c, y_train_c)
# Print the R-squared
print("R^2: {0:.2f}".format(reg.score(X_test_c, y_test_c)))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, reg.predict(X_test))
# Log parameter, metrics, and model to MLflow
with mlflow.start_run(run_name="Linear Regression 'Dropped Missing'"):
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
# -
# ### Random Forest
# +
preds_rf = reg_rf.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds_rf))
print("RMSE: %f" % (rmse))
# +
reg_rf = RandomForestRegressor(random_state=57, n_estimators=100)
reg_rf.fit(X_train_c, y_train_c)
reg_rf.score(X_test_c, y_test_c)
#pd.DataFrame({'x':X_hp.columns.values, 'y':reg_rf.feature_importances_})
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test_c, reg_rf.predict(X_test_c))
# Log parameter, metrics, and model to MLflow
with mlflow.start_run(run_name="Random Forest 'Dropped Missing'"):
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
# +
preds_rf = reg_rf.predict(X_test_c)
rmse = np.sqrt(mean_squared_error(y_test_c, preds_rf))
print("RMSE: %f" % (rmse))
print("R2: ", r2)
# -
feature_importance = pd.DataFrame({'features':X_c.columns.values, 'importance':reg_rf.feature_importances_}).sort_values('importance', ascending=False)
feature_importance.head(10).set_index('features')
feature_importance.set_index('features')[:10].plot.bar(rot=60)
# ### XGBoost with Imputed Missing Values
# +
import xgboost as xgb
xg_reg = xgb.XGBRegressor(objective ='reg:linear', colsample_bytree = 0.3, learning_rate = 0.1,
max_depth = 5, alpha = 10, n_estimators = 100)
xg_reg.fit(X_train, y_train)
preds = xg_reg.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds))
print("RMSE: %f" % (rmse))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, xg_reg.predict(X_test))
# Log parameter, metrics, and model to MLflow
mlflow.start_run(run_name="XGBoost Imputed")
mlflow.log_param("alpha", 10)
mlflow.log_param("max_depth", 5)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
mlflow.end_run()
# -
xgb.plot_importance(xg_reg, max_num_features=20)
# ### XGBoost with Dropped Missing Values
# +
import xgboost as xgb
xg_reg = xgb.XGBRegressor(objective ='reg:linear', colsample_bytree = 0.3, learning_rate = 0.1,
max_depth = 5, alpha = 10, n_estimators = 100)
xg_reg.fit(X_train_c, y_train_c)
preds = xg_reg.predict(X_test_c)
rmse = np.sqrt(mean_squared_error(y_test_c, preds))
print("RMSE: %f" % (rmse))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test_c, xg_reg.predict(X_test_c))
# Log parameter, metrics, and model to MLflow
mlflow.start_run(run_name="XGBoost - 'Dropped Missing'")
mlflow.log_param("alpha", 10)
mlflow.log_param("max_depth", 5)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
mlflow.end_run()
# -
xgb.plot_importance(xg_reg, max_num_features=20)
# ### XGBoost with Missing Values
# +
# Create features and target (df)
X_m = df_missing.drop('price', axis = 1) # features
y_m = df_missing['price'] # target
# Train-test split
X_train_m, X_test_m, y_train_m, y_test_m = train_test_split(X_m, y_m, test_size = 0.3, random_state=57)
# +
xg_reg = xgb.XGBRegressor(objective ='reg:linear', colsample_bytree = 0.3, learning_rate = 0.1,
max_depth = 5, alpha = 10, n_estimators = 100)
xg_reg.fit(X_train_m, y_train_m)
preds = xg_reg.predict(X_test_m)
rmse = np.sqrt(mean_squared_error(y_test_m, preds))
print("RMSE: %f" % (rmse))
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test_m, xg_reg.predict(X_test_m))
# Log parameter, metrics, and model to MLflow
mlflow.start_run(run_name="XGBoost NOT Imputed")
mlflow.log_param("alpha", 10)
mlflow.log_param("max_depth", 5)
mlflow.log_metric("rmse", rmse)
mlflow.log_metric("mae", mae)
mlflow.log_metric("r2", r2)
mlflow.end_run()
# -
xgb.plot_importance(xg_reg, max_num_features=20)
# ## Save Model (4 features)
# Import imputed data
df_no_dummy = pd.read_pickle("autos_clean_no_dummy_20190805.pkl")
test_data = df_no_dummy[['hp', 'age', 'km', 'model', 'price']].copy()
# Create features and target (df)
X = pd.get_dummies(df_no_dummy[['hp', 'age', 'km', 'model']]) # features
y = df_no_dummy['price'] # target
X.head()
y.head()
# Train-test split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state=57)
# +
reg_rf = RandomForestRegressor(n_estimators=100, random_state=57)
reg_rf.fit(X_train, y_train)
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, reg_rf.predict(X_test))
print("rmse: {}, mae: {}, r2: {}".format(rmse, mae, r2))
# +
from sklearn.model_selection import RandomizedSearchCV
# Number of trees in random forest
n_estimators = [int(x) for x in np.linspace(start = 100, stop = 2000, num = 10)]
# Number of features to consider at every split
max_features = ['auto', 'sqrt']
# Maximum number of levels in tree
max_depth = [int(x) for x in np.linspace(10, 110, num = 11)]
max_depth.append(None)
# Minimum number of samples required to split a node
min_samples_split = [2, 5, 10]
# Minimum number of samples required at each leaf node
min_samples_leaf = [1, 2, 4]
# Method of selecting samples for training each tree
bootstrap = [True, False]
# Create the random grid
random_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'max_depth': max_depth,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_leaf,
'bootstrap': bootstrap}
# -
# Use the random grid to search for best hyperparameters
# First create the base model to tune
rf = RandomForestRegressor()
# Random search of parameters, using 3 fold cross validation,
# search across 100 different combinations, and use all available cores
rf_random = RandomizedSearchCV(estimator = rf, param_distributions = random_grid, n_iter = 10, cv = 3, verbose=2, random_state=42, n_jobs = -1)
# Fit the random search model
rf_random.fit(X_train, y_train)
rf_random.best_params_
rf_best = rf_random.best_estimator_
# +
# reg_rf = RandomForestRegressor(n_estimators=100, random_state=57)
# reg_rf.fit(X_train, y_train)
# Calculate evaluation metrics
rmse, mae, r2 = eval_metrics(y_test, rf_best.predict(X_test))
print("rmse: {}, mae: {}, r2: {}".format(rmse, mae, r2))
# +
# Save RandomForest model
# pickle.dump(rf_best, open('model_rf.pkl','wb'))
# -
list(X.columns)
# +
# Save column names to persist the features in deployment phase
# pickle.dump(list(X.columns), open('model_features.pkl','wb'))
# -
test_data['prediction'] = rf_best.predict(X)
test_data.sample(5)
# +
# Create subplots
fig, ax = plt.subplots(2, 4, figsize=(15, 12))
# Plot subplots
i = 0
j = 0
for model in ['A1', 'A3', 'Astra', 'Clio', 'Corsa', 'Espace', 'Insignia']:
df_no_dummy[df_no_dummy.model == model].boxplot('price', by='age', ax = ax[j, i])
ax[j, i].set_title(model)
# Next subplot
i += 1
if i == 4:
i = 0
j = 1
# -
df_no_dummy.model.value_counts()
sns.pairplot(df_no_dummy[df_no_dummy.model=='Clio'][['age', 'price']], kind='reg')
rf_best.decision_path(X)
rf_best
rf_best.fit(X_train, y_train)
# +
import graphviz
import pydot
tree = rf_best.estimators_[5]
# Export the image to a dot file
export_graphviz(tree, out_file = 'tree.dot', feature_names = X_train.columns,
rounded = True, precision = 1)
# Use dot file to create a graph
(graph, ) = pydot.graph_from_dot_file('tree.dot')
# Write graph to a png file
graph.write_png('tree.png')
# -
from statsmodels.graphics.factorplots import interaction_plot
fig = interaction_plot(df_no_dummy.age, df_no_dummy.model, df_no_dummy.price)
plt.show()
|
trgt/model-final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-west-2:236514542706:image/datascience-1.0
# ---
# ## Introduction
#
# Word2Vec is a popular algorithm used for generating dense vector representations of words in large corpora using unsupervised learning. The resulting vectors have been shown to capture semantic relationships between the corresponding words and are used extensively for many downstream natural language processing (NLP) tasks like sentiment analysis, named entity recognition and machine translation.
# SageMaker BlazingText which provides efficient implementations of Word2Vec on
#
# - single CPU instance
# - single instance with multiple GPUs - P2 or P3 instances
# - multiple CPU instances (Distributed training)
# In this notebook, we demonstrate how BlazingText can be used for distributed training of word2vec using multiple CPU instances.
# ## Setup
#
# Let's start by specifying:
# - The S3 buckets and prefixes that you want to use for saving model data and where training data is located. These should be within the same region as the Notebook Instance, training, and hosting. If you don't specify a bucket, SageMaker SDK will create a default bucket following a pre-defined naming convention in the same region.
# - The IAM role ARN used to give SageMaker access to your data. It can be fetched using the **get_execution_role** method from sagemaker python SDK.
# + isConfigCell=true
import sagemaker
from sagemaker import get_execution_role
import boto3
import json
sess = sagemaker.Session()
role = get_execution_role()
print(
role
) # This is the role that SageMaker would use to leverage AWS resources (S3, CloudWatch) on your behalf
region = boto3.Session().region_name
output_bucket = sess.default_bucket() # Replace with your own bucket name if needed
print(output_bucket)
output_prefix = "sagemaker/DEMO-blazingtext-text8" # Replace with the prefix under which you want to store the data if needed
data_bucket = f"sagemaker-sample-files" # Replace with the bucket where your data is located
data_prefix = "datasets/text/text8/text8"
# -
# ### Data Ingestion
#
# BlazingText expects a single preprocessed text file with space separated tokens and each line of the file should contain a single sentence. In this example, let us train the vectors on [text8](http://mattmahoney.net/dc/textdata.html) dataset (100 MB), which is a small (already preprocessed) version of Wikipedia dump. Data is already downloaded from [matt mahoney's website](http://mattmahoney.net/dc/text8.zip), uncompressed and stored in `data_bucket`.
# +
s3_client = boto3.client("s3")
s3_client.download_file(data_bucket, data_prefix, "text8")
s3_client.upload_file("text8", output_bucket, output_prefix + "/train")
s3_train_data = f"s3://{output_bucket}/{output_prefix}/train"
# -
# Next we need to setup an output location at S3, where the model artifact will be dumped. These artifacts are also the output of the algorithm's training job.
s3_output_location = f"s3://{output_bucket}/{output_prefix}/output"
# ## Training Setup
# Now that we are done with all the setup that is needed, we are ready to train our object detector. To begin, let us create a ``sageMaker.estimator.Estimator`` object. This estimator will launch the training job.
region_name = boto3.Session().region_name
container = sagemaker.amazon.amazon_estimator.get_image_uri(region_name, "blazingtext", "latest")
print(f"Using SageMaker BlazingText container: {container} ({region_name})")
# ## Training the BlazingText model for generating word vectors
# Similar to the original implementation of [Word2Vec](https://arxiv.org/pdf/1301.3781.pdf), SageMaker BlazingText provides an efficient implementation of the continuous bag-of-words (CBOW) and skip-gram architectures using Negative Sampling, on CPUs and additionally on GPU[s]. The GPU implementation uses highly optimized CUDA kernels. To learn more, please refer to [*BlazingText: Scaling and Accelerating Word2Vec using Multiple GPUs*](https://dl.acm.org/citation.cfm?doid=3146347.3146354). BlazingText also supports learning of subword embeddings with CBOW and skip-gram modes. This enables BlazingText to generate vectors for out-of-vocabulary (OOV) words, as demonstrated in this [notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/blazingtext_word2vec_subwords_text8/blazingtext_word2vec_subwords_text8.ipynb).
#
#
#
# Besides skip-gram and CBOW, SageMaker BlazingText also supports the "Batch Skipgram" mode, which uses efficient mini-batching and matrix-matrix operations ([BLAS Level 3 routines](https://software.intel.com/en-us/mkl-developer-reference-fortran-blas-level-3-routines)). This mode enables distributed word2vec training across multiple CPU nodes, allowing almost linear scale up of word2vec computation to process hundreds of millions of words per second. Please refer to [*Parallelizing Word2Vec in Shared and Distributed Memory*](https://arxiv.org/pdf/1604.04661.pdf) to learn more.
# BlazingText also supports a *supervised* mode for text classification. It extends the FastText text classifier to leverage GPU acceleration using custom CUDA kernels. The model can be trained on more than a billion words in a couple of minutes using a multi-core CPU or a GPU, while achieving performance on par with the state-of-the-art deep learning text classification algorithms. For more information, please refer to [algorithm documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/blazingtext.html) or [the text classification notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/blazingtext_text_classification_dbpedia/blazingtext_text_classification_dbpedia.ipynb).
# To summarize, the following modes are supported by BlazingText on different types instances:
#
# | Modes | cbow (supports subwords training) | skipgram (supports subwords training) | batch_skipgram | supervised |
# |:----------------------: |:----: |:--------: |:--------------: | :--------------: |
# | Single CPU instance | ✔ | ✔ | ✔ | ✔ |
# | Single GPU instance | ✔ | ✔ | | ✔ (Instance with 1 GPU only) |
# | Multiple CPU instances | | | ✔ | | |
#
# Now, let's define the resource configuration and hyperparameters to train word vectors on *text8* dataset, using "batch_skipgram" mode on two c4.2xlarge instances.
#
bt_model = sagemaker.estimator.Estimator(
container,
role,
instance_count=2,
instance_type="ml.c4.2xlarge",
train_volume_size=5,
train_max_run=360000,
input_mode="File",
output_path=s3_output_location,
sagemaker_session=sess,
)
# Please refer to [algorithm documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/blazingtext_hyperparameters.html) for the complete list of hyperparameters.
bt_model.set_hyperparameters(
mode="batch_skipgram",
epochs=5,
min_count=5,
sampling_threshold=0.0001,
learning_rate=0.05,
window_size=5,
vector_dim=100,
negative_samples=5,
batch_size=11, # = (2*window_size + 1) (Preferred. Used only if mode is batch_skipgram)
evaluation=True, # Perform similarity evaluation on WS-353 dataset at the end of training
subwords=False,
) # Subword embedding learning is not supported by batch_skipgram
# Now that the hyper-parameters are setup, let us prepare the handshake between our data channels and the algorithm. To do this, we need to create the `sagemaker.session.s3_input` objects from our data channels. These objects are then put in a simple dictionary, which the algorithm consumes.
train_data = sagemaker.session.s3_input(
s3_train_data,
distribution="FullyReplicated",
content_type="text/plain",
s3_data_type="S3Prefix",
)
data_channels = {"train": train_data}
# We have our `Estimator` object, we have set the hyper-parameters for this object and we have our data channels linked with the algorithm. The only remaining thing to do is to train the algorithm. The following command will train the algorithm. Training the algorithm involves a few steps. Firstly, the instance that we requested while creating the `Estimator` classes is provisioned and is setup with the appropriate libraries. Then, the data from our channels are downloaded into the instance. Once this is done, the training job begins. The provisioning and data downloading will take some time, depending on the size of the data. Therefore it might be a few minutes before we start getting training logs for our training jobs. The data logs will also print out `Spearman's Rho` on some pre-selected validation datasets after the training job has executed. This metric is a proxy for the quality of the algorithm.
#
# Once the job has finished a "Job complete" message will be printed. The trained model can be found in the S3 bucket that was setup as `output_path` in the estimator.
bt_model.fit(inputs=data_channels, logs=True)
# ## Hosting / Inference
# Once the training is done, we can deploy the trained model as an Amazon SageMaker real-time hosted endpoint. This will allow us to make predictions (or inference) from the model. Note that we don't have to host on the same type of instance that we used to train. Because instance endpoints will be up and running for long, it's advisable to choose a cheaper instance for inference.
bt_endpoint = bt_model.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
# ### Getting vector representations for words
# #### Use JSON format for inference
# The payload should contain a list of words with the key as "**instances**". BlazingText supports content-type `application/json`.
# +
words = ["awesome", "blazing"]
payload = {"instances": words}
response = bt_endpoint.predict(
json.dumps(payload),
initial_args={"ContentType": "application/json", "Accept": "application/json"},
)
vecs = json.loads(response)
print(vecs)
# -
# As expected, we get an n-dimensional vector (where n is vector_dim as specified in hyperparameters) for each of the words. If the word is not there in the training dataset, the model will return a vector of zeros.
# ### Evaluation
# Let us now download the word vectors learned by our model and visualize them using a [t-SNE](https://en.wikipedia.org/wiki/T-distributed_stochastic_neighbor_embedding) plot.
# +
s3 = boto3.resource("s3")
key = bt_model.model_data[bt_model.model_data.find("/", 5) + 1 :]
s3.Bucket(output_bucket).download_file(key, "model.tar.gz")
# -
# Uncompress `model.tar.gz` to get `vectors.txt`
# !tar -xvzf model.tar.gz
# If you set "evaluation" as "true" in the hyperparameters, then "eval.json" will be there in the model artifacts.
#
# The quality of trained model is evaluated on word similarity task. We use [WS-353](http://alfonseca.org/eng/research/wordsim353.html), which is one of the most popular test datasets used for this purpose. It contains word pairs together with human-assigned similarity judgments.
#
# The word representations are evaluated by ranking the pairs according to their cosine similarities, and measuring the Spearmans rank correlation coefficient with the human judgments.
#
# Let's look at the evaluation scores which are there in eval.json. For embeddings trained on the text8 dataset, scores above 0.65 are pretty good.
# !cat eval.json
# Now, let us do a 2D visualization of the word vectors
# +
import numpy as np
from sklearn.preprocessing import normalize
# Read the 400 most frequent word vectors. The vectors in the file are in descending order of frequency.
num_points = 400
first_line = True
index_to_word = []
with open("vectors.txt", "r") as f:
for line_num, line in enumerate(f):
if first_line:
dim = int(line.strip().split()[1])
word_vecs = np.zeros((num_points, dim), dtype=float)
first_line = False
continue
line = line.strip()
word = line.split()[0]
vec = word_vecs[line_num - 1]
for index, vec_val in enumerate(line.split()[1:]):
vec[index] = float(vec_val)
index_to_word.append(word)
if line_num >= num_points:
break
word_vecs = normalize(word_vecs, copy=False, return_norm=False)
# +
from sklearn.manifold import TSNE
tsne = TSNE(perplexity=40, n_components=2, init="pca", n_iter=10000)
two_d_embeddings = tsne.fit_transform(word_vecs[:num_points])
labels = index_to_word[:num_points]
# +
from matplotlib import pylab
# %matplotlib inline
def plot(embeddings, labels):
pylab.figure(figsize=(20, 20))
for i, label in enumerate(labels):
x, y = embeddings[i, :]
pylab.scatter(x, y)
pylab.annotate(
label, xy=(x, y), xytext=(5, 2), textcoords="offset points", ha="right", va="bottom"
)
pylab.show()
plot(two_d_embeddings, labels)
# -
# Running the code above might generate a plot like the one below. t-SNE and Word2Vec are stochastic, so although when you run the code the plot won’t look exactly like this, you can still see clusters of similar words such as below where 'british', 'american', 'french', 'english' are near the bottom-left, and 'military', 'army' and 'forces' are all together near the bottom.
# 
# ### Stop / Close the Endpoint (Optional)
# Finally, we should delete the endpoint before we close the notebook.
sess.delete_endpoint(bt_endpoint.endpoint)
|
introduction_to_amazon_algorithms/blazingtext_word2vec_text8/blazingtext_word2vec_text8.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Machine Learning Model Building Pipeline: Machine Learning Model Build
#
# In the following videos, we will take you through a practical example of each one of the steps in the Machine Learning model building pipeline, which we described in the previous lectures. There will be a notebook for each one of the Machine Learning Pipeline steps:
#
# 1. Data Analysis
# 2. Feature Engineering
# 3. Feature Selection
# 4. Model Building
#
# **This is the notebook for step 4: Building the Final Machine Learning Model**
#
# We will use the house price dataset available on [Kaggle.com](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data). See below for more details.
#
# ===================================================================================================
#
# ## Predicting Sale Price of Houses
#
# The aim of the project is to build a machine learning model to predict the sale price of homes based on different explanatory variables describing aspects of residential houses.
#
# ### Why is this important?
#
# Predicting house prices is useful to identify fruitful investments, or to determine whether the price advertised for a house is over or under-estimated.
#
# ### What is the objective of the machine learning model?
#
# We aim to minimise the difference between the real price and the price estimated by our model. We will evaluate model performance using the mean squared error (mse) and the root squared of the mean squared error (rmse).
#
# ### How do I download the dataset?
#
# To download the House Price dataset go this website:
# https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data
#
# Scroll down to the bottom of the page, and click on the link 'train.csv', and then click the 'download' blue button towards the right of the screen, to download the dataset. Rename the file as 'houseprice.csv' and save it to a directory of your choice.
#
# **Note the following:**
# - You need to be logged in to Kaggle in order to download the datasets.
# - You need to accept the terms and conditions of the competition to download the dataset
# - If you save the file to the same directory where you saved this jupyter notebook, then you can run the code as it is written here.
#
# ====================================================================================================
# ## House Prices dataset: Model building
#
# In the following cells, we will finally build our machine learning model, utilising the engineered data and the pre-selected features.
#
#
# ### Setting the seed
#
# It is important to note, that we are engineering variables and pre-processing data with the idea of deploying the model. Therefore, from now on, for each step that includes some element of randomness, it is extremely important that we **set the seed**. This way, we can obtain reproducibility between our research and our development code.
#
# This is perhaps one of the most important lessons that you need to take away from this course: **Always set the seeds**.
#
# Let's go ahead and load the dataset.
# +
# to handle datasets
import pandas as pd
import numpy as np
# for plotting
import matplotlib.pyplot as plt
# to build the model
from sklearn.linear_model import Lasso
# to evaluate the model
from sklearn.metrics import mean_squared_error, r2_score
from math import sqrt
# to visualise al the columns in the dataframe
pd.pandas.set_option('display.max_columns', None)
# +
# load the train and test set with the engineered variables
# we built and saved these datasets in a previous notebook.
# If you haven't done so, go ahead and check the previous notebooks (step 2)
# to find out how to create these datasets
X_train = pd.read_csv('xtrain.csv')
X_test = pd.read_csv('xtest.csv')
X_train.head()
# +
# capture the target (remember that is log transformed)
y_train = X_train['SalePrice']
y_test = X_test['SalePrice']
# +
# load the pre-selected features
# ==============================
# we selected the features in the previous notebook (step 3)
# if you haven't done so, go ahead and visit the previous notebook
# to find out how to select the features
features = pd.read_csv('selected_features.csv', header = None)
features = features.iloc[:,0].to_list()
# We will add one additional feature to the ones we selected in the
# previous notebook: LotFrontage
# # why?
#=====
# because it needs key feature engineering steps that we want to
# discuss further during the deployment part of the course.
features = features + ['LotFrontage']
# display final feature set
features
# +
# reduce the train and test set to the selected features
X_train = X_train[features]
X_test = X_test[features]
# -
# ### Regularised linear regression: Lasso
#
# Remember to set the seed.
# +
# set up the model
# remember to set the random_state / seed
lin_model = Lasso(alpha=0.005, random_state=0)
# train the model
lin_model.fit(X_train, y_train)
# +
# evaluate the model:
# ====================
# remember that we log transformed the output (SalePrice)
# in our feature engineering notebook (step 2).
# In order to get the true performance of the Lasso
# we need to transform both the target and the predictions
# back to the original house prices values.
# We will evaluate performance using the mean squared error and
# the root of the mean squared error and r2
# make predictions for train set
pred = lin_model.predict(X_train)
# determine mse and rmse
print('train mse: {}'.format(int(
mean_squared_error(np.exp(y_train), np.exp(pred)))))
print('train rmse: {}'.format(int(
sqrt(mean_squared_error(np.exp(y_train), np.exp(pred))))))
print('train r2: {}'.format(
r2_score(np.exp(y_train), np.exp(pred))))
print()
# make predictions for test set
pred = lin_model.predict(X_test)
# determine mse and rmse
print('test mse: {}'.format(int(
mean_squared_error(np.exp(y_test), np.exp(pred)))))
print('test rmse: {}'.format(int(
sqrt(mean_squared_error(np.exp(y_test), np.exp(pred))))))
print('test r2: {}'.format(
r2_score(np.exp(y_test), np.exp(pred))))
print()
print('Average house price: ', int(np.exp(y_train).median()))
# -
# let's evaluate our predictions respect to the real sale price
plt.scatter(y_test, lin_model.predict(X_test))
plt.xlabel('True House Price')
plt.ylabel('Predicted House Price')
plt.title('Evaluation of Lasso Predictions')
# We can see that our model is doing a pretty good job at estimating house prices.
# +
# let's evaluate the distribution of the errors:
# they should be fairly normally distributed
errors = y_test - lin_model.predict(X_test)
errors.hist(bins=30)
# -
# The distribution of the errors follows quite closely a gaussian distribution. That suggests that our model is doing a good job as well.
# ### Feature importance
# +
# Finally, just for fun, let's look at the feature importance
importance = pd.Series(np.abs(lin_model.coef_.ravel()))
importance.index = features
importance.sort_values(inplace=True, ascending=False)
importance.plot.bar(figsize=(18,6))
plt.ylabel('Lasso Coefficients')
plt.title('Feature Importance')
# -
# And that is all! Now we have our entire pipeline ready for deployment.
#
# In the next video, we will summarise which steps from the pipeline we will deploy to production.
|
Section-2-Machine-Learning-Pipeline-Overview/Machine-Learning-Pipeline-Step4-Model-Building.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Least squares fitting of models to data
# This is a quick introduction to `statsmodels` for physical scientists (e.g. physicists, astronomers) or engineers.
#
# Why is this needed?
#
# Because most of `statsmodels` was written by statisticians and they use a different terminology and sometimes methods, making it hard to know which classes and functions are relevant and what their inputs and outputs mean.
# + jupyter={"outputs_hidden": false}
import numpy as np
import pandas as pd
import statsmodels.api as sm
# -
# ## Linear models
# Assume you have data points with measurements `y` at positions `x` as well as measurement errors `y_err`.
#
# How can you use `statsmodels` to fit a straight line model to this data?
#
# For an extensive discussion see [Hogg et al. (2010), "Data analysis recipes: Fitting a model to data"](https://arxiv.org/abs/1008.4686) ... we'll use the example data given by them in Table 1.
#
# So the model is `f(x) = a * x + b` and on Figure 1 they print the result we want to reproduce ... the best-fit parameter and the parameter errors for a "standard weighted least-squares fit" for this data are:
# * `a = 2.24 +- 0.11`
# * `b = 34 +- 18`
# + jupyter={"outputs_hidden": false}
data = """
x y y_err
201 592 61
244 401 25
47 583 38
287 402 15
203 495 21
58 173 15
210 479 27
202 504 14
198 510 30
158 416 16
165 393 14
201 442 25
157 317 52
131 311 16
166 400 34
160 337 31
186 423 42
125 334 26
218 533 16
146 344 22
"""
try:
from StringIO import StringIO
except ImportError:
from io import StringIO
data = pd.read_csv(StringIO(data), delim_whitespace=True).astype(float)
# Note: for the results we compare with the paper here, they drop the first four points
data.head()
# -
# To fit a straight line use the weighted least squares class [WLS](https://www.statsmodels.org/devel/generated/statsmodels.regression.linear_model.WLS.html) ... the parameters are called:
# * `exog` = `sm.add_constant(x)`
# * `endog` = `y`
# * `weights` = `1 / sqrt(y_err)`
#
# Note that `exog` must be a 2-dimensional array with `x` as a column and an extra column of ones. Adding this column of ones means you want to fit the model `y = a * x + b`, leaving it off means you want to fit the model `y = a * x`.
#
# And you have to use the option `cov_type='fixed scale'` to tell `statsmodels` that you really have measurement errors with an absolute scale. If you do not, `statsmodels` will treat the weights as relative weights between the data points and internally re-scale them so that the best-fit model will have `chi**2 / ndf = 1`.
# + jupyter={"outputs_hidden": false}
exog = sm.add_constant(data['x'])
endog = data['y']
weights = 1. / (data['y_err'] ** 2)
wls = sm.WLS(endog, exog, weights)
results = wls.fit(cov_type='fixed scale')
print(results.summary())
# -
# ### Check against scipy.optimize.curve_fit
# + jupyter={"outputs_hidden": false}
# You can use `scipy.optimize.curve_fit` to get the best-fit parameters and parameter errors.
from scipy.optimize import curve_fit
def f(x, a, b):
return a * x + b
xdata = data['x']
ydata = data['y']
p0 = [0, 0] # initial parameter estimate
sigma = data['y_err']
popt, pcov = curve_fit(f, xdata, ydata, p0, sigma, absolute_sigma=True)
perr = np.sqrt(np.diag(pcov))
print('a = {0:10.3f} +- {1:10.3f}'.format(popt[0], perr[0]))
print('b = {0:10.3f} +- {1:10.3f}'.format(popt[1], perr[1]))
# -
# ### Check against self-written cost function
# + jupyter={"outputs_hidden": false}
# You can also use `scipy.optimize.minimize` and write your own cost function.
# This does not give you the parameter errors though ... you'd have
# to estimate the HESSE matrix separately ...
from scipy.optimize import minimize
def chi2(pars):
"""Cost function.
"""
y_model = pars[0] * data['x'] + pars[1]
chi = (data['y'] - y_model) / data['y_err']
return np.sum(chi ** 2)
result = minimize(fun=chi2, x0=[0, 0])
popt = result.x
print('a = {0:10.3f}'.format(popt[0]))
print('b = {0:10.3f}'.format(popt[1]))
# -
# ## Non-linear models
# + jupyter={"outputs_hidden": false}
# TODO: we could use the examples from here:
# http://probfit.readthedocs.org/en/latest/api.html#probfit.costfunc.Chi2Regression
|
examples/notebooks/chi2_fitting.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -all
# formats: ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction #
#
# First encountering a new dataset can sometimes feel overwhelming. You might be presented with hundreds or thousands of features without even a description to go by. Where do you even begin?
#
# A great first step is to construct a ranking with a **feature utility metric**, a function measuring associations between a feature and the target. Then you can choose a smaller set of the most useful features to develop initially and have more confidence that your time will be well spent.
#
# The metric we'll use is called "mutual information". Mutual information is a lot like correlation in that it measures a relationship between two quantities. The advantage of mutual information is that it can detect *any* kind of relationship, while correlation only detects *linear* relationships.
#
# Mutual information is a great general-purpose metric and especially useful at the start of feature development when you might not know what model you'd like to use yet. It is:
# - easy to use and interpret,
# - computationally efficient,
# - theoretically well-founded,
# - resistant to overfitting, and,
# - able to detect any kind of relationship
#
# # Mutual Information and What it Measures #
#
# Mutual information describes relationships in terms of *uncertainty*. The **mutual information** (MI) between two quantities is a measure of the extent to which knowledge of one quantity reduces uncertainty about the other. If you knew the value of a feature, how much more confident would you be about the target?
#
# Here's an example from the *Ames Housing* data. The figure shows the relationship between the exterior quality of a house and the price it sold for. Each point represents a house.
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/X12ARUK.png" width=400, alt="Four categories of ExterQual: Fair, Typical, Good, Excellent. A scatter plot of SalePrice within each category.">
# <figcaption style="textalign: center; font-style: italic"><center>Knowing the exterior quality of a house reduces uncertainty about its sale price.
# </center></figcaption>
# </figure>
#
# From the figure, we can see that knowing the value of `ExterQual` should make you more certain about the corresponding `SalePrice` -- each category of `ExterQual` tends to concentrate `SalePrice` to within a certain range. The mutual information that `ExterQual` has with `SalePrice` is the average reduction of uncertainty in `SalePrice` taken over the four values of `ExterQual`. Since `Fair` occurs less often than `Typical`, for instance, `Fair` gets less weight in the MI score.
#
# (Technical note: What we're calling uncertainty is measured using a quantity from information theory known as "entropy". The entropy of a variable means roughly: "how many yes-or-no questions you would need to describe an occurance of that variable, on average." The more questions you have to ask, the more uncertain you must be about the variable. Mutual information is how many questions you expect the feature to answer about the target.)
#
# # Interpreting Mutual Information Scores #
#
# The least possible mutual information between quantities is 0.0. When MI is zero, the quantities are independent: neither can tell you anything about the other. Conversely, in theory there's no upper bound to what MI can be. In practice though values above 2.0 or so are uncommon. (Mutual information is a logarithmic quantity, so it increases very slowly.)
#
# The next figure will give you an idea of how MI values correspond to the kind and degree of association a feature has with the target.
#
# <figure style="padding: 1em;">
# <img src="https://i.imgur.com/Dt75E1f.png" width=800, alt="">
# <figcaption style="textalign: center; font-style: italic"><center><strong>Left:</strong> Mutual information increases as the dependence between feature and target becomes tighter. <strong>Right:</strong> Mutual information can capture any kind of association (not just linear, like correlation.)
# </center></figcaption>
# </figure>
#
# Here are some things to remember when applying mutual information:
# - MI can help you to understand the *relative potential* of a feature as a predictor of the target, considered by itself.
# - It's possible for a feature to be very informative when interacting with other features, but not so informative all alone. MI *can't detect interactions* between features. It is a **univariate** metric.
# - The *actual* usefulness of a feature *depends on the model you use it with*. A feature is only useful to the extent that its relationship with the target is one your model can learn. Just because a feature has a high MI score doesn't mean your model will be able to do anything with that information. You may need to transform the feature first to expose the association.
#
# # Example - 1985 Automobiles #
#
# The [*Automobile*](https://www.kaggle.com/toramky/automobile-dataset) dataset consists of 193 cars from the 1985 model year. The goal for this dataset is to predict a car's `price` (the target) from 23 of the car's features, such as `make`, `body_style`, and `horsepower`. In this example, we'll rank the features with mutual information and investigate the results by data visualization.
#
# This hidden cell imports some libraries and loads the dataset.
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
plt.style.use("seaborn-whitegrid")
df = pd.read_csv("../input/fe-course-data/autos.csv")
df.head()
# -
# The scikit-learn algorithm for MI treats discrete features differently from continuous features. Consequently, you need to tell it which are which. As a rule of thumb, anything that *must* have a `float` dtype is *not* discrete. Categoricals (`object` or `categorial` dtype) can be treated as discrete by giving them a label encoding. (You can review label encodings in our [Categorical Variables](http://www.kaggle.com/alexisbcook/categorical-variables) lesson.)
# +
X = df.copy()
y = X.pop("price")
# Label encoding for categoricals
for colname in X.select_dtypes("object"):
X[colname], _ = X[colname].factorize()
# All discrete features should now have integer dtypes (double-check this before using MI!)
discrete_features = X.dtypes == int
# -
# Scikit-learn has two mutual information metrics in its `feature_selection` module: one for real-valued targets (`mutual_info_regression`) and one for categorical targets (`mutual_info_classif`). Our target, `price`, is real-valued. The next cell computes the MI scores for our features and wraps them up in a nice dataframe.
# +
from sklearn.feature_selection import mutual_info_regression
def make_mi_scores(X, y, discrete_features):
mi_scores = mutual_info_regression(X, y, discrete_features=discrete_features)
mi_scores = pd.Series(mi_scores, name="MI Scores", index=X.columns)
mi_scores = mi_scores.sort_values(ascending=False)
return mi_scores
mi_scores = make_mi_scores(X, y, discrete_features)
mi_scores[::3] # show a few features with their MI scores
# -
# And now a bar plot to make comparisions easier:
# +
def plot_mi_scores(scores):
scores = scores.sort_values(ascending=True)
width = np.arange(len(scores))
ticks = list(scores.index)
plt.barh(width, scores)
plt.yticks(width, ticks)
plt.title("Mutual Information Scores")
plt.figure(dpi=100, figsize=(8, 5))
plot_mi_scores(mi_scores)
# -
# Data visualization is a great follow-up to a utility ranking. Let's take a closer look at a couple of these.
#
# As we might expect, the high-scoring `curb_weight` feature exhibits a strong relationship with `price`, the target.
#
sns.relplot(x="curb_weight", y="price", data=df);
# The `fuel_type` feature has a fairly low MI score, but as we can see from the figure, it clearly separates two `price` populations with different trends within the `horsepower` feature. This indicates that `fuel_type` contributes an interaction effect and might not be unimportant after all. Before deciding a feature is unimportant from its MI score, it's good to investigate any possible interaction effects -- domain knowledge can offer a lot of guidance here.
sns.lmplot(x="horsepower", y="price", hue="fuel_type", data=df);
# Data visualization is a great addition to your feature-engineering toolbox. Along with utility metrics like mutual information, visualizations like these can help you discover important relationships in your data. Check out our [Data Visualization](https://www.kaggle.com/learn/data-visualization) course to learn more!
#
# # Your Turn #
#
# [**Rank the features**](https://www.kaggle.com/kernels/fork/14393925) of the *Ames Housing* dataset and choose your first set of features to start developing.
# ---
#
#
#
#
# *Have questions or comments? Visit the [Learn Discussion forum](https://www.kaggle.com/learn-forum/221677) to chat with other Learners.*
|
corso-data-science-2021/hands-on/04-regression-and-clustering/lessons/extra/mutual-information.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 在一个 m*n 的二维字符串数组中输出二叉树,并遵守以下规则:
# 1、行数 m 应当等于给定二叉树的高度。
# 2、列数 n 应当总是奇数。
# 3、根节点的值(以字符串格式给出)应当放在可放置的第一行正中间。
# 根节点所在的行与列会将剩余空间划分为两部分(左下部分和右下部分)。你
# 应该将左子树输出在左下部分,右子树输出在右下部分。
# 左下和右下部分应当有相同的大小。即使一个子树为空而另一个非空,你不需要为空的子树输出任何东西,但仍需要为另一个子树留出足够的空间。
# 然而,如果两个子树都为空则不需要为它们留出任何空间。
# 每个未使用的空间应包含一个空的字符串""。
# 使用相同的规则输出子树。
#
# 示例 1:
# 输入:
# 1
# /
# 2
# 输出:
# [["", "1", ""],
# ["2", "", ""]]
#
# 示例 2:
# 输入:
# 1
# / \
# 2 3
# \
# 4
# 输出:
# [["", "", "", "1", "", "", ""],
# ["", "2", "", "", "", "3", ""],
# ["", "", "4", "", "", "", ""]]
#
# 示例 3:
# 输入:
# 1
# / \
# 2 5
# /
# 3
# /
# 4
# 输出:
# [["", "", "", "", "", "", "", "1", "", "", "", "", "", "", ""]
# ["", "", "", "2", "", "", "", "", "", "", "", "5", "", "", ""]
# ["", "3", "", "", "", "", "", "", "", "", "", "", "", "", ""]
# ["4", "", "", "", "", "", "", "", "", "", "", "", "", "", ""]]
#
# 注意: 二叉树的高度在范围 [1, 10] 中。
# -
[["","","","","","","","","","","","","","","","1","","","","","","","","","","","","","","",""],
["","","","","","","","2","","","","","","","","","","","","","","","","3","","","","","","",""],
["","","","4","","","","","","","","5","","","","","","","","","","","","","","","","","","",""],
["","6","","","","7","","","","","","","","","","","","","","","","","","","","","","","","",""],
["","","8","","","","","","","","","","","","","","","","","","","","","","","","","","","",""]]
a = ["","","","","","","","1","","","","","","",""]
print(len(a))
# +
[["","","","","","","","1","","","","","","",""],
["","","2","","","","","","","","3","","","",""],
["","","","","5","","6","","","","","","7","4",""],
["","10","","12","","11","","","","","8","","","","9"]]
[["","","","","","","","1","","","","","","",""],
["","","","2","","","","","","","","3","","",""],
["","4","","","","5","","","","6","","","","7",""],
["8","","9","","10","","11","","12","","","","","",""]]
# +
from collections import deque
class Solution:
def printTree(self, root: TreeNode) -> List[List[str]]:
max_depth = self.get_depth(root)
max_len = pow(2, max_depth) - 1
res = [[''] * (pow(2, max_depth) - 1) for _ in range(max_depth)]
nodes = deque([(root, max_len//2)])
for i in range(max_depth):
for _ in range(len(nodes)):
node, idx = nodes.popleft()
if node:
res[i][idx] = str(node.val)
if node.left:
nodes.append((node.left, idx - pow(2, max_depth-i-2)))
if node.right:
nodes.append((node.right, idx + pow(2, max_depth-i-2)))
return res
def get_depth(self, root):
if not root:
return 0
return max(self.get_depth(root.left), self.get_depth(root.right)) + 1
# -
def get_height(node):
return 0 if not node else 1 + max(get_height(node.left), get_height(node.right))
def update_output(node, row, left, right):
if not node:
return
mid = (left + right) // 2
self.output[row][mid] = str(node.val)
update_output(node.left, row + 1 , left, mid - 1)
update_output(node.right, row + 1 , mid + 1, right)
height = get_height(root)
width = 2 ** height - 1
self.output = [[''] * width for i in range(height)]
update_output(node=root, row=0, left=0, right=width - 1)
return self.output
|
Tree/1021/655. Print Binary Tree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Guided Analysis - User Security Metadata (Private Preview)
#
# **Notebook Version:** 1.0
# **Python Version:** Python 3.6
# **Required Packages**: kqlmagic, validate_email, jsonpickle, azure-cli-core, Azure-Sentinel-Utilities
#
# **Platforms Supported**:
# - Azure Notebooks Free Compute
# - Azure Notebooks DSVM
# - OS Independent
#
# **Data Sources Required**:
# - Log Analytics : UserPeerAnalytics, UserAccessAnalytics
#
# **Permissions Required**:
# - **Log Analytics Read Permissions**: To connect and query the workspace you need to be assigned at least [Reader](https://docs.microsoft.com/en-us/azure/role-based-access-control/built-in-roles#reader) or [Azure Sentinel Reader](https://docs.microsoft.com/en-us/azure/role-based-access-control/built-in-roles#azure-sentinel-reader) role on the workspace.
# - **Directory Basic Read Permissions** : If you are a user who is a native member of the tenant, then by [default](https://docs.microsoft.com/en-us/azure/active-directory/fundamentals/users-default-permissions#compare-member-and-guest-default-permissions) you have permissions to read user, group and serviceprincipal information. If you are a guest user in the tenant, then you need to be assigned [Directory Reader](https://docs.microsoft.com/en-us/azure/active-directory/users-groups-roles/directory-assign-admin-roles#directory-readers) role.
#
# **Description**:
# This notebook introduces the concept of contextual security metadata that are gathered for AAD users. Here are the security metadata that are available* today
# - **UserAccessAnalytics**: The most important step of a security incident is to identify the blast radius of the user under investigation. This enrichment data calculates for a given user, the direct or transitive access/permission to resources. In Private Preview, we calculate the blast radius access graph only limited to RBAC access to subscriptions. For example, if the user under investigation is <NAME>, Access Graph displays all the Azure subscriptions that she either can access directly, via groups or serviceprincipals.
# - **UserPeerAnalytics**: Analysts frequently use the peers of a user under investigation to scope the security incident. This enrichment data, for a given user, provides a ranked list of peers. For example, if the user is <NAME>, Peer Enrichment calculates all of Jane’s peers based on her mailing list, security groups, etc and provides the top 20 of her peers. Specifically, this information is calculated using Natural Language Processing algorithms using group membership information from Azure Active Directory.
#
# *This is an Azure Sentinel **Private Preview** feature. If you are interested in the above analytics data please contact ramk at microsoft com.
#
# ## Contents:
# - [Setup](#setup)
# - [Install Packages](#install)
# - [Enter Tenant and Workspace Ids](#tenant-and-worskpace-ids)
# - [Connect to Log Analytics](#connect-to-la)
# - [Log into Azure CLI](#log-into-azure)
# - [Enter User Information](#user-input)
# - [Access Graph of the user](#access-graph)
# - [Ranked peers of the user](#user-peers)
# <a id='setup'></a>
# # Setup
# <a id='install'></a>
# ## Install Packages
# The first time this cell runs for a new Azure Notebooks project or local Python environment it will take several minutes to download and install the packages. In subsequent runs it should run quickly and confirm that package dependencies are already installed. Unless you want to upgrade the packages you can feel free to skip execution of the next cell.
print('Please wait. Installing required packages. This may take a few minutes...')
# !pip install Kqlmagic --no-cache-dir --upgrade
# !pip install validate_email --upgrade
# !pip install jsonpickle --upgrade
# !pip install azure-cli-core --upgrade
# !pip install --upgrade Azure-Sentinel-Utilities
# <a id='tenant-and-worskpace-ids'></a>
# ## Enter Tenant and Workspace Ids
# You can configure your TenantId and WorskpaceId in config.json file next to the notebook, see sample [here](https://github.com/Azure/Azure-Sentinel/blob/master/Notebooks/config.json). If config.json file is missing then you will be prompted to enter TenantId and WorkspaceId manually.
# To find your WorkspaceId go to [Log Analytics](https://portal.azure.com/#blade/HubsExtension/Resources/resourceType/Microsoft.OperationalInsights%2Fworkspaces), and look at the workspace properties to find the ID.
# +
import os.path
import SentinelUtils
tenantId = None
workspaceId = None
configFile = "config.json"
if os.path.isfile(configFile):
try:
print(f"Read Workspace configuration from local '{configFile}' file... ", end = "")
tenantId = SentinelUtils.config_reader.ConfigReader.read_config_values(configFile)[0]
workspaceId = SentinelUtils.config_reader.ConfigReader.read_config_values(configFile)[3]
print("Done!")
print(f"Tenant - '{tenantId}' and Log Analytics Workspace - '{workspaceId}' retrieved from {configFile}")
except:
pass
if not workspaceId or not tenantId:
print(f"Unable to retrive tenantId and workspaceid from '{configFile}'.")
print('Enter Azure TenantId: ')
tenantId = input().strip()
print()
print('Enter Sentinel Workspace Id: ')
workspaceId = input().strip()
print()
# -
# <a id='connect-to-la'></a>
# ## Connect to Log Analytics
# This is required to read the tables in your log analytics workspace.
# %reload_ext Kqlmagic
# %kql loganalytics://code().tenant(tenantId).workspace(workspaceId)
# <a id='log-into-azure'></a>
# ## Log into Azure CLI
# Azure CLI is used to retrieve display name and email address of users, groups and service principals from AAD.
# !az login --tenant $tenantId
# %run Entities.py
# %run GraphVis.py
# <a id='user-input'></a>
# ## Enter User Information
# +
from Utils import validatedate
from datetime import date
import ipywidgets as widgets
from IPython.display import display
print('Enter object Id or UPN or email address of the user: ')
userIdOrEmail = input().strip()
print()
if not userIdOrEmail :
raise Exception("Error: Empty Object Id or UPN or email address.")
print(f'Retrieving user "{userIdOrEmail}" from the tenant...', end = '')
user = User.getUserByIdOrEmail(userIdOrEmail)
print("Done!")
print("Name - {0}, Email - {1}, Id - {2}".format(user.name, user.email, user.objectId))
print()
print('[Optional] Enter date in format yyyy-MM-dd to retrieve analytics from that date. If you want latest, leave it empty and press enter: ')
time = input().strip()
if not time :
today = date.today()
time = today.strftime("%Y-%m-%d")
else:
validatedate(time)
# -
# <a id='access-graph'></a>
# # Access Graph of the user:
# Run this cell to visualize the access/permissions of the user in a graph. The cell queries the 'UserAccessAnalytics' table to retrieve direct/transitive RBAC access of the user to subscriptions.
# +
from IPython.display import clear_output, display, HTML
kql_query = f"""
let userId = "{user.objectId}";
let blastRadTime = todatetime('{time}');
let userSubAccess = UserAccessAnalytics
| where SourceEntityId == userId and TargetEntityType == "AzureSubscription" and TimeGenerated <= blastRadTime
| project UserId = SourceEntityId, TimeGenerated , SubscriptionName = TargetEntityName, Subscription = TargetEntityId, Role = AccessLevel, GroupId = "", ServicePrincipalId = ""
| summarize arg_max(TimeGenerated, *) by Subscription, Role;
let userGroupAccess = UserAccessAnalytics
| where SourceEntityId == userId and TargetEntityType == "Group" and TimeGenerated <= blastRadTime
| project UserId = SourceEntityId, GroupId = TargetEntityId, TimeGenerated
| summarize arg_max(TimeGenerated, *) by GroupId;
let userGroupSubAccess = userGroupAccess
| join kind = inner
UserAccessAnalytics
on $left.GroupId == $right.SourceEntityId
| where TargetEntityType == "AzureSubscription" and TimeGenerated <= blastRadTime
| project UserId, GroupId, ServicePrincipalId = "", TimeGenerated, SubscriptionName = TargetEntityName, Subscription = TargetEntityId, Role = AccessLevel
| summarize arg_max(TimeGenerated, *) by GroupId, Subscription, Role;
let userSPAccess = UserAccessAnalytics
| where SourceEntityId == userId and TargetEntityType == "ServicePrincipal" and TimeGenerated <= blastRadTime
| project UserId = SourceEntityId, ServicePrincipalId = TargetEntityId, TimeGenerated
| summarize arg_max(TimeGenerated, *) by ServicePrincipalId;
let userSPSubAccess = userSPAccess
| join kind = inner
UserAccessAnalytics
on $left.ServicePrincipalId == $right.SourceEntityId
| where TargetEntityType == "AzureSubscription" and TimeGenerated <= blastRadTime
| project UserId, GroupId = "", ServicePrincipalId, TimeGenerated, SubscriptionName = TargetEntityName, Subscription = TargetEntityId, Role = AccessLevel
| summarize arg_max(TimeGenerated, *) by ServicePrincipalId, Subscription, Role;
userGroupSubAccess
| union kind=outer userSubAccess
| union kind=outer userSPSubAccess"""
print(f"Executing Kql query to retrieve access analytics for user '{user.name}', on or before '{time}'.. ", end = '')
# %kql -query kql_query
print('Done!')
usersubMappings = _kql_raw_result_.to_dataframe()
if len(usersubMappings) == 0:
print(f"No access analytics data available for user '{user.name}', on or before '{time}'")
else:
print('Creating Graph visualization. This may take a few seconds.. ', end = '')
graph = GraphVis()
for index, row in usersubMappings.iterrows():
sub = Subscription(row['SubscriptionName'], row['Subscription'])
rbacRole = row['Role']
if row['GroupId'] == '' and row['ServicePrincipalId'] == '':
graph.addEdge(user.getNode(), sub.getNode(), rbacRole)
elif row['GroupId']:
group = Group.getGroupById(row['GroupId'])
graph.addEdge(user.getNode(), group.getNode(), "Member")
graph.addEdge(group.getNode(), sub.getNode(), rbacRole)
elif row['ServicePrincipalId']:
sp = ServicePrincipal.getServicePrincipalById(row['ServicePrincipalId'])
graph.addEdge(user.getNode(), sp.getNode(), "Owner")
graph.addEdge(sp.getNode(), sub.getNode(), rbacRole)
print('Done!')
display(HTML(graph.getHtml()))
# -
# <a id='user-peers'></a>
# ## Ranked peers of the user
# This cell queries the 'UserPeerAnalytics' table to return a ranked list of peers of the user.
# +
from IPython.display import clear_output, display, HTML
import tabulate
kql_query = f"""
let userId = "{user.objectId}";
let snapshotTime = todatetime('{time}');
UserPeerAnalytics
| where UserId == userId
| join kind = inner
(
UserPeerAnalytics
| where TimeGenerated <= snapshotTime and UserId == userId
| summarize max(TimeGenerated)
| project TimeGenerated = max_TimeGenerated
)
on TimeGenerated
| project PeerUserId, Rank
| order by Rank asc"""
print(f"Executing Kql query to retrieve peer analytics for user '{user.name}', on or before '{time}'.. ", end = '')
# %kql -query kql_query
print('Done!')
peerListDF = _kql_raw_result_.to_dataframe()
peerList = []
peerList.append(["UserName", "PeerUserName", "PeerEmail", "Rank"])
if len(peerListDF) == 0:
print(f"No peer analytics data available for user '{user.name}', on or before '{time}'")
else:
print('Retrieving user names and email addresses for peers. This may take a few seconds...', end = '')
for index, row in peerListDF.iterrows():
peerUserId = row['PeerUserId']
peerRank = row['Rank']
peerUser = User.getUserByIdOrEmail(peerUserId)
peerList.append([user.name, peerUser.name, peerUser.email, peerRank])
print('Done!')
display(HTML(tabulate.tabulate(peerList, tablefmt='html')))
# -
|
Notebooks/BehaviorAnalytics/UserSecurityMetadata/Guided Analysis - User Security Metadata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#1 print tup
Tup=4,5,6,7,8,9
Tup[0:6]
tup=(4,5,6,7),(7,6,8,9)
print("tup[0:4]:",tup[0:4])
list_1=[4,5,6,79]
List_1=tuple(list_1)
List_1
List_1=['4','5','6','79']
type("tup")
tup_1=('foo',[1,2],True)
tup_1[-2][1]=3
tup_1
A=(1,81,4,9,16,81,49,64,81,100,81,16,81)
A.count(81)
# +
#2Use the list below and make a new list B that has only odd elements of this list in it.
A=(1,4,9,16,25,36,49,64,81,100)
a1=[]
for i in A:
if i%2!=0:
a1.append(i)
print(a1)
# -
#3Suppose z = (2,3,4,5,6), which of the following is INCORRECT?
option D is incorrect
#4
A=(1,4,9,16,25,36,49,64,81,100)
A[1:-1]
A[::-1]
#5
record = ('<NAME>', 50, 123.45, (12, 18, 2012))
(a,b,c,(d,e,f))=record
print(a)
print(f)
#6
#we are also allowed to extract the values back into variables. This is called "unpacking"
A=(1,4,9,16,25,36,49,64,81,100)
(_,a,_,_,b,_,_,c,_,d)=A
print(a,b,c,d)
#7
record=('<NAME>','<EMAIL>','773-555-1212','847-555-1212')
(Name,Emailaddress,*phonenumber,)=record
print(Name,Emailaddress,phonenumber)
#10
i = 1
while i<=100:
if i % 3!=0 and i % 5!=0:
print(i)
i=i+1
#11
for i in range(4):
for j in range(4-i):
print("#",end="")
print()
#12
for i in range(1,5):
for j in range(5-i):
print(i+j,end=" ")
print()
#9
(a=int(input())
if a>0:
print("positive")
else:
print("negative")
f = [0, 1]
for i in range(2, 50):
f.append(f[i-1] + f[i-2])
print(f)
|
mounika sai python quiz 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %config IPython.matplotlib.backend = "retina"
from matplotlib import rcParams
rcParams["figure.dpi"] = 150
rcParams["savefig.dpi"] = 150
# # An introduction to TensorFlow for probabilistic modeling
#
# In this worksheet, we will go through an example of fitting a line to data using [TensorFlow](https://www.tensorflow.org/) to build the model combined with the standard astronomical Python stack.
# To run this notebook, you will need to have the standard scientific Python packages installed: NumPy, SciPy, and Matplotlib.
# You will also need TensorFlow. The installation instructions can be found [here](https://www.tensorflow.org/install/), but it's often enough to just run:
#
# ```bash
# pip install -U tensorflow
# ```
#
# (This is generally better than using conda because the conda packages were really old when I last checked.)
#
# ## Data & model
#
# To start, let's simulate some fake data.
# We'll generate the data from a linear model and we'll assume that the error bars are unknown (we'll fit for the magnitude of the uncertainty in a minute).
# +
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(123)
true_params = (-0.4, 0.03, 0.1)
x = np.sort(np.random.uniform(-1, 1, 50))
y = true_params[0] * x + true_params[1] + true_params[2] * np.random.randn(len(x))
plt.plot(x, y, ".k", label="data")
t = np.linspace(-1, 1, 5000)
plt.plot(t, true_params[0]*t+true_params[1], label="truth")
plt.legend()
plt.xlabel("x")
plt.ylabel("y");
# -
# We won't go into the mathematical details of this model here (if you're interested, check out [<NAME>, & Lang 2010](https://arxiv.org/abs/1008.4686)), but the basic idea is that we need to write down the following likelihood function for this model:
#
# $$
# \log p(y\,|\,m,\,b,\,s) = -\frac{1}{2}\,\sum_{n=1}^N \left[ \frac{(y_n - m\,x_n - b)^2}{s^2} + \log(2\,\pi\,s^2) \right]
# $$
#
# In NumPy, you might write this function as follows:
# +
def np_log_like(x, y, m, b, s):
model = m * x + b
return -0.5*np.sum(((y-model)/s)**2+np.log(2*np.pi*s**2))
print("The log likelihood computed using numpy: {0}".format(np_log_like(x, y, *true_params)))
# -
# Now your job is to fill in the following function and re-write this in TensorFlow. (The docs are [here](https://www.tensorflow.org/api_docs/python/tf).)
# +
import tensorflow as tf
print("Using TensorFlow version {0}".format(tf.__version__))
# The default type in tensorflow is float32, but we want to force float64 here.
T = tf.float64
# Convert the numpy arrays to tensors.
# This step of using the "constant" method isn't strictly necessary,
# but it will come in handy later.
x_tensor = tf.constant(x, dtype=T)
y_tensor = tf.constant(y, dtype=T)
# These are the parameters that we will fit for.
# Most tensors are immutable so we need to call them "Variable"s if we
# want to change them.
m_tensor = tf.Variable(true_params[0], dtype=T)
b_tensor = tf.Variable(true_params[1], dtype=T)
# s_tensor = tf.Variable(true_params[2], dtype=T)
# TODO: CHANGE THIS
log_s_tensor = tf.Variable(np.log(true_params[2]), dtype=T)
s_tensor = tf.exp(log_s_tensor)
# This is the function that you should write.
# Hint: in many cases you can get a long way by taking numpy code and
# converting "np" to "tf".
def tf_log_like(x, y, m, b, s):
# ADD YOUR CODE HERE...
model = m * x + b
return -0.5*tf.reduce_sum(((y-model)/s)**2+tf.log(2*np.pi*s**2))
# Now we execute this function to define the operations.
log_like = tf_log_like(x_tensor, y_tensor, m_tensor, b_tensor, s_tensor)
print("'log_like' is: {0}".format(log_like))
# And finally open a session to execute the model.
with tf.Session() as session:
# This step is needed to set up the variables.
session.run(tf.global_variables_initializer())
# And compute the log likelihood.
print("The log likelihood computed using tensorflow: {0}"
.format(session.run(log_like)))
# -
# If the value of the log likelihood of this cell isn't identical to the one above, edit your function until it is.
# After that, think a bit about what is happening at each step of the code.
# What does the value of `log_like` mean and why does it have this value?
# ## The "feed_dict"
#
# One useful concept in TensorFlow is called the `feed_dict` and it is passed to the `session.run` method as a keyword argument.
# It allows you to easily evaluate some part of your model for different values of any number of tensors without re-defining the full graph.
# For example, to compute the log likelihood for a different slope, we could use the following code:
# +
feed_dict = {
m_tensor: 0.1, # <-- NOTE: this is *not* a typo. The key should be
# *the tensor*, not a string!
}
with tf.Session() as session:
session.run(tf.global_variables_initializer())
print("The log likelihood for a slope of 0.1: {0}"
.format(session.run(log_like, feed_dict=feed_dict)))
# -
# This is generally most useful for parameters that you are fitting for, but it also works for other tensors.
# For example, we can compute the likelihood for a different dataset as follows:
# +
feed_dict = {
y_tensor: np.zeros_like(y),
}
with tf.Session() as session:
session.run(tf.global_variables_initializer())
print("The log likelihood for a dataset of zeros: {0}"
.format(session.run(log_like, feed_dict=feed_dict)))
# -
# ## Gradients & optimization
#
# One of the key benefits of TensorFlow is that it efficiently computes gradients of scalar functions with respect to any other tensors.
# These gradients can be used with many inference algorithms to improve the reliability and efficiency of fitting probabilistic models to data.
# For example, many non-linear optimization algorithms (like the ones that might be familiar from [scipy.optimize](https://docs.scipy.org/doc/scipy/reference/optimize.html)) can substantially benefit from gradients of the objective function.
# Furthermore, the performance of Markov chain Monte Carlo (MCMC) methods can be improved by using the gradients of the log probability function to generate samples (see [Radford Neal's review](http://www.mcmchandbook.net/HandbookChapter5.pdf), for example).
#
# In the following cell, use the [tf.gradients](https://www.tensorflow.org/api_docs/python/tf/gradients) function to define an operation to compute the gradient of `log_like` with respect to `m_tensor`, `b_tensor`, and `s_tensor`.
# Then, open a session, and evaluate this gradient (you should get something like: `[-48.6057, 52.6399, 122.76701]`).
# +
# ADD YOUR CODE HERE...
grad = tf.gradients(log_like, [m_tensor, b_tensor, s_tensor])
with tf.Session() as session:
session.run(tf.global_variables_initializer())
print(session.run(grad))
# -
# Now that you see how to compute these gradients, go back up to the cell where we defined our model in TensorFlow and re-write it in terms of $\log(s)$ instead of $s$. (This is generally a better parameterization for any model where a parameter must always be positive.)
# This should involve introducing a new `Variable` (`log_s_tensor` instead of `s_tensor`) and then redefining `s_tensor` in terms of `log_s_tensor`.
# With this new parameterization, how would you compute the gradient of `log_like` with respect to the new parameters?
#
# We can now use our model and one of the gradient based optimizers included with TensorFlow.
# Specifically, let's use the interface to the optimizers in [scipy.optimize](https://docs.scipy.org/doc/scipy/reference/optimize.html) because these optimizers are commonly used by astronomers.
# First, remember that (for historical reasons) most optimizers *minimize* their objective so we'll want to minimize the *negative* log likelihood in order to *maximize* the log likelihood.
# +
neg_log_like = -log_like
opt = tf.contrib.opt.ScipyOptimizerInterface(
neg_log_like, var_list=[m_tensor, b_tensor, log_s_tensor])
with tf.Session() as session:
session.run(tf.global_variables_initializer())
print("Initial log likelihood: {0}".format(session.run(log_like)))
opt.minimize(session)
print("Final log likelihood: {0}\n".format(session.run(log_like)))
params = session.run([m_tensor, b_tensor, s_tensor])
print("Final parameters: {0}".format(params))
print("True parameters: {0}".format(true_params))
# -
# ## Interfacing with Python & sampling using MCMC
#
# Another common task is to sample the posterior probability using MCMC.
# To start, we need to choose a log prior for the parameters.
# For today, let's choose an improper uniform prior on `log_s_tensor` (i.e. a constant value for all values of `log_s_tensor`) and use the prior [suggested by <NAME> on his blog](http://jakevdp.github.io/blog/2014/06/14/frequentism-and-bayesianism-4-bayesian-in-python/#Prior-on-Slope-and-Intercept) for `m_tensor` and `b_tensor`.
# First, write down the log prior from Jake's blog post and add it to `log_like` to get `log_prob` (a tensor that will be equal to the log posterior up to an additive constant).
log_prior = -1.5 * tf.log(1.0 + m_tensor**2)
log_prob = log_like + log_prior
with tf.Session() as session:
session.run(tf.global_variables_initializer())
print(session.run(log_prob))
# This worksheet comes with a helper class called `TFModel` that can be used to create a interface between a TensorFlow model and standard Python fitting packages.
# The key feature of this interface is that it wraps a tensor and exposes the value and gradient as Python methods (using the `feed_dict` as described above).
# For example, to wrap our log probability, we can use the following workflow:
# +
# First we say that we want the model to return the value and gradient
# of `log_prob` as a function of the parameters `m_tensor`, `b_tensor`,
# and `log_s_tensor`.
from helpers import TFModel
model = TFModel(log_prob, [m_tensor, b_tensor, log_s_tensor])
with tf.Session() as session:
session.run(tf.global_variables_initializer())
# Within the session, you first need to call the `setup` method.
model.setup(session)
# You can access the current parameter vector for the model.
# This will always be a flat numpy array.
params = model.current_vector()
# The value and gradient of the tensor can be evaluated for specific
# values of the parameters.
print("The value is: {0}".format(model.value(params)))
print("The gradient is: {0}".format(model.gradient(params)))
# -
# As an example, we can use this interface to use [emcee](http://emcee.readthedocs.io) to sample this posterior probability that we have defined.
# To run this cell, you will need to install emcee (the docs are [here](http://emcee.readthedocs.io)).
# +
import time
import emcee
emcee_time = time.time()
with tf.Session() as session:
session.run(tf.global_variables_initializer())
model.setup(session)
pos = model.current_vector()
pos = pos + 1e-5*np.random.randn(32, len(pos))
nwalkers, ndim = pos.shape
sampler = emcee.EnsembleSampler(nwalkers, ndim, model.value)
pos, _, _ = sampler.run_mcmc(pos, 200)
sampler.reset()
pos, _, _ = sampler.run_mcmc(pos, 2000)
emcee_time = time.time() - emcee_time
# -
# And then we can plot the results using the [corner](http://corner.readthedocs.io/) package.
import corner
truth = np.array(true_params)
truth[-1] = np.log(true_params[-1])
corner.corner(sampler.flatchain,
labels=["m", "b", "log(s)"],
truths=truth);
# ## Hamiltonian Monte Carlo (HMC)
#
# This looks pretty good, but emcee doesn't currently have support for methods like HMC that use gradient information to improve the efficiency of the inference.
# While there are several packages that exist in Python for running HMC, I haven't been completely happy with any of them so, for this tutorial, I wrote a simple Hamiltonian sampler (specifically it implements the [No U-Turns Sampler](https://arxiv.org/abs/1111.4246) as implemented in [Stan](http://mc-stan.org/) and [PyMC3](http://docs.pymc.io/)).
# This isn't the place to get into the details of a sampler like this, but to run it here, you can use the following code:
# +
import helpers
# We will cheat a little bit and use our previous chain to estimate the
# the appropriate tuning scales for the parameters.
metric = helpers.DiagonalMetric(3, np.var(sampler.flatchain, axis=0))
nuts_time = time.time()
with tf.Session() as session:
session.run(tf.global_variables_initializer())
# This method does the sampling:
nuts = helpers.tf_simple_nuts(
session,
log_prob,
[m_tensor, b_tensor, log_s_tensor],
3500, # The number of MCMC steps
1.0, # The integration step size
metric=metric # The scaling metric computed above
)
nuts_time = time.time() - nuts_time
# -
corner.corner(nuts[0],
range=[(np.min(v), np.max(v)) for v in sampler.flatchain.T],
labels=["m", "b", "log(s)"],
truths=truth);
# These contours look a little bit rattier than the ones above, but the real test is the computation time per *independent* sample (see [this blog post](http://dfm.io/posts/autocorr/) for a discussion of methods for estimating the number of independent samples).
# For this example, we can compute the number of independent samples from each chain:
# +
# First estimate the autocorrelation time of each chain
from autocorr import integrated_time
tau_emcee = integrated_time(np.swapaxes(sampler.chain, 0, 1))
tau_nuts = integrated_time(nuts[0][:, None, :])
# Then the effective numbers of samples
neff_emcee = len(sampler.flatchain) / np.mean(tau_emcee)
neff_nuts = len(nuts[0]) / np.mean(tau_nuts)
print("The effective number of samples from emcee: {0:.0f}".format(neff_emcee))
print("The effective number of samples from NUTS: {0:.0f}".format(neff_nuts))
print("The time per effective sample for emcee: {0:.1f} ms".format(1000*emcee_time / neff_emcee))
print("The time per effective sample for NUTS: {0:.1f} ms".format(1000*nuts_time / neff_nuts))
# -
# If everything went as planned, you should see that the NUTS sampler is substantially more efficient (i.e. it requires less computation time per independent sample) than emcee (but we're not considering the cost of tuning here).
# This disparity will tend to increase for problems with more parameters.
# In general, sampling a density with tens of parameters using emcee will require substantially more patience and computation power than sampling the same problem with NUTS, but NUTS is more sensitive to the specific choices of tuning parameters.
|
solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35opencv]
# language: python
# name: conda-env-py35opencv-py
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import cv2
# %matplotlib inline
# -
# ## Software and package versions
# +
print("*** VERSIONS ***")
import sys
print("Python {}".format(sys.version))
print("OpenCV {}".format(cv2.__version__))
print("Numpy {}".format(np.__version__))
import matplotlib
print("Matplotlib {}".format(matplotlib.__version__))
# -
# !jupyter --version
# !conda env list
# !conda list -n py35opencv
# +
def print_position_in_video(cap):
print(" position-ms: {}".format(cap.get(cv2.CAP_PROP_POS_MSEC)))
print("position-frames: {}".format(cap.get(cv2.CAP_PROP_POS_FRAMES)))
print(" position-ratio: {}".format(cap.get(cv2.CAP_PROP_POS_AVI_RATIO)))
def print_video_info(cap):
print(" width: {}".format(cap.get(cv2.CAP_PROP_FRAME_WIDTH)))
print(" height: {}".format(cap.get(cv2.CAP_PROP_FRAME_HEIGHT)))
# print(" fps: {}".format(cap.get(cv2.CAP_PROP_FRAME_FPS))) # This property name doesn't exist
print(" # frames: {}".format(cap.get(cv2.CAP_PROP_FRAME_COUNT)))
print(" gain: {}".format(cap.get(cv2.CAP_PROP_GAIN)))
print(" brightness: {}".format(cap.get(cv2.CAP_PROP_BRIGHTNESS)))
print_position_in_video(cap)
# -
# ## Grab first frame and plot
# +
cap = cv2.VideoCapture('vacuum - 100 ms.mp4')
print(cap.isOpened())
print_video_info(cap)
ret, image = cap.read()
print_position_in_video(cap)
fig, ax = plt.subplots(figsize=(12,8))
ax.imshow(image[:, :, ::-1]);
# -
# ## Grab particular frame, convert to grayscale, and plot
# +
# Set frame pointer to particular frame
frame_num = 300
cap.set(cv2.CAP_PROP_POS_FRAMES, frame_num)
ret, img = cap.read()
img_gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
fig, ax = plt.subplots(figsize=(12,8))
ax.imshow(img_gray, cmap='gray');
# -
print(img_gray.shape)
print(img_gray.min(), img_gray.max())
# ## Define region of interest and plot just that region
def add_box_to_image(img, xlow, ylow, xsize, ysize, linewidth=5, graylevel=128):
xhigh = xlow + xsize
yhigh = ylow + ysize
img[:, xlow:xlow+linewidth] = graylevel
img[:, xhigh:xhigh+linewidth] = graylevel
img[ylow:ylow+linewidth, :] = graylevel
img[yhigh:yhigh+linewidth, :] = graylevel
# +
xlow, xsize = 170, 195
ylow, ysize = 550, 230
temp_img = img_gray.copy()
add_box_to_image(temp_img, xlow, ylow, xsize, ysize)
fig, ax = plt.subplots(figsize=(12,8))
ax.imshow(temp_img, cmap='gray');
# +
temp_img = img_gray.copy()
roi = np.s_[ylow:ylow+ysize, xlow:xlow+xsize]
fig, ax = plt.subplots(figsize=(8,6))
ax.imshow(temp_img[roi], cmap='gray');
# -
# ## Threshold image region of interest and count pixels below threshold
# +
threshold = 50
replacement_value = 128
temp_img = img_gray.copy()
roi = np.s_[ylow:ylow+ysize, xlow:xlow+xsize]
temp_img_roi = temp_img[roi]
fig, ax = plt.subplots(figsize=(8,6))
ax.imshow(temp_img_roi, cmap='gray');
temp = temp_img_roi.copy()
num_pixels_below_threshold = (temp < threshold).sum()
temp[temp < threshold] = replacement_value
fig, ax = plt.subplots(figsize=(8,6))
ax.imshow(temp, cmap='gray')
ax.set_title("Pixels below threshold: {}".format(num_pixels_below_threshold));
# -
# ## Process entire video
# +
# Re-set frame pointer to start of video file
cap.set(cv2.CAP_PROP_POS_FRAMES, 0)
num_frames = int(cap.get(cv2.CAP_PROP_FRAME_COUNT))
num_pixels_below_threshold = np.zeros(num_frames)
xlow, xsize = 170, 195
ylow, ysize = 550, 230
roi = np.s_[ylow:ylow+ysize, xlow:xlow+xsize]
threshold = 50
for i in range(num_frames):
ret, img = cap.read()
img_gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
img_gray_roi = img_gray[roi]
num_pixels_below_threshold[i] = (img_gray_roi < threshold).sum()
frames = np.arange(num_frames)
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(frames, num_pixels_below_threshold);
# -
# ## Subtract average of initial black pixels
# +
num_pixels_below_threshold_zeroed = num_pixels_below_threshold - np.mean(num_pixels_below_threshold[:70])
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(frames, num_pixels_below_threshold_zeroed)
ax.set_xlabel("Frames")
ax.set_ylabel("Pixels Below Threshold Value of {}".format(threshold));
# -
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(frames, num_pixels_below_threshold_zeroed)
ax.set_xlim(70, 180)
ax.set_ylim(0, 4000)
ax.set_xlabel("Frames")
ax.set_ylabel("Pixels Below Threshold Value of {}".format(threshold));
# ## Automatically find periods
from scipy.signal import argrelextrema
minima = argrelextrema(num_pixels_below_threshold_zeroed, np.less, order=5)
minima = minima[0]
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(frames, num_pixels_below_threshold_zeroed)
for val in minima:
ax.axvline(val, c='k', ls='--', lw=1);
ax.set_xlabel("Frames")
ax.set_ylabel("Pixels Below Threshold Value of {}".format(threshold));
fig, ax = plt.subplots(figsize=(12,8))
ax.plot(frames, num_pixels_below_threshold_zeroed)
ax.set_xlim(70, 180)
ax.set_ylim(0, 4000)
for val in minima:
ax.axvline(val, c='k', ls='--', lw=1);
ax.set_xlabel("Frames")
ax.set_ylabel("Pixels Below Threshold Value of {}".format(threshold));
for i in range(3, 3+20):
print(i, minima[i+1] - minima[i])
# It looks like the pump period in units of frames is very steady at 15 frames.
#
# **I suggest just using the first 5 or so periods in your analysis to keep everything consistent because then you are guaranteed that the fluid is flowing through the serpentine and not the final straight section of channel.**
|
python/openCV_video_processing/analyze_video.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas with SQL
#
# The [`pandas.io.sql` module](http://pandas.pydata.org/pandas-docs/stable/user_guide/io.html#sql-queries) provides a collection of query wrappers to both facilitate data retrieval and to reduce dependency on DB-specific API. It supports multiple driver library, e.g., SQLAlchemy.
#
# ## Query
# To query, use `read_sql` with SQL statement to load data into DataFrame:
import pandas as pd
import sqlite3
conn = sqlite3.connect("sqlalchemy_example.db")
pd.read_sql("select * from address;", conn)
from sqlalchemy import create_engine
engine = create_engine('sqlite:///sqlalchemy_example.db')
pd.read_sql("select * from address;", engine)
pd.read_sql("select * from person;", engine)
df = pd.read_sql("select * from person, address where person.id==address.person_id;", engine)
df
print(type(df))
# ## Insert and Update
#
# Pandas API has `to_sql` function that allows you to easily insert your data into the database. But it is very slow!
df = pd.read_sql("select * from address;", engine)
df2 = df.append({'street_name':'Newington Road', 'street_number':'15',
'post_code':'12121', 'person_id':2}, ignore_index = True)
df2
engine = create_engine('sqlite:///sqlalchemy_example2.db')
df2.to_sql('address', con=engine, index=False, if_exists='replace')
engine = create_engine('sqlite:///sqlalchemy_example2.db')
pd.read_sql_query("select * from address;", engine)
df2
df2.to_dict(orient="records")
# +
from sqlalchemy import Column, ForeignKey, Integer, String
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy.orm import relationship, sessionmaker
from sqlalchemy import create_engine
Base = declarative_base()
class Person(Base):
__tablename__ = 'person'
# Here we define columns for the table person
# Notice that each column is also a normal Python instance attribute.
id = Column(Integer, primary_key=True)
name = Column(String(250), nullable=False)
class Address(Base):
__tablename__ = 'address'
# Here we define columns for the table address.
# Notice that each column is also a normal Python instance attribute.
id = Column(Integer, primary_key=True)
street_name = Column(String(250))
street_number = Column(String(250))
post_code = Column(String(250), nullable=False)
person_id = Column(Integer, ForeignKey('person.id'))
person = relationship(Person)
engine = create_engine('sqlite:///sqlalchemy_example3.db')
Base.metadata.create_all(engine)
Session = sessionmaker(bind=engine)
session = Session()
session.bulk_insert_mappings(Address, df2.to_dict(orient="records"))
session.commit()
session.close()
# -
# In SQLAlchemy ORM, there are bulk operations, e.g., `bulk_insert_mappings`, `bulk_update_mappings`. The purpose of these methods is to emit INSERT and UPDATE statements given dictionaries or object states with lower Python overhead. This is achieved by directly expose internal elements of the unit of work system.
# (For more details https://docs.sqlalchemy.org/en/latest/orm/persistence_techniques.html#bulk-operations)
#
# The advantage of this solution is that it is fast and it exploits ORM's advantages.
engine = create_engine('sqlite:///sqlalchemy_example3.db')
pd.read_sql_query("select * from address;", engine)
# ## Delete
# Pandas doesn't have any command for deleting rows. You can only overwrite the old table using `to_sql`, or drop the table and insert a new one using SQLAlchemy ORM.
|
01day02_database/pandas_sql.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import os
import torch
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
plt.ion() # interactive mode
# +
landmarks_frame = pd.read_csv('data/faces/face_landmarks.csv')
n = 65
img_name = landmarks_frame.iloc[n, 0]
landmarks = landmarks_frame.iloc[n, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
print('Image name: {}'.format(img_name))
print('Landmarks shape: {}'.format(landmarks.shape))
print('First 4 Landmarks: {}'.format(landmarks[:4]))
# +
def show_landmarks(image, landmarks):
"""Show image with landmarks"""
plt.imshow(image)
plt.scatter(landmarks[:, 0], landmarks[:, 1], s=10, marker='.', c='r')
plt.pause(0.001) # pause a bit so that plots are updated
plt.figure()
show_landmarks(io.imread(os.path.join('data/faces/', img_name)),
landmarks)
plt.show()
# -
|
DeepDetection/CocoTest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import pandas as pd
import numpy as np
prob = np.load("prob.npy")
prob
det = np.load("detEarning.npy")
det
sum([det[i]*((1+0.02)**(30-i)) for i in range(30)])
35*(1.2**5)
len(det)
sum(det[:45])*0.05
det
import random
# %pylab inline
import numpy as np
import pandas as pd
from scipy.interpolate import interp1d
from multiprocessing import Pool
from functools import partial
from pyswarm import pso
import warnings
warnings.filterwarnings("ignore")
np.printoptions(precision=2)
# time line
T_min = 0
T_max = 70
T_R = 45
beta = 1/(1+0.02)
# All the money amount are denoted in thousand dollars
earningShock = [0.8,1.2]
# Define transition matrix of economical states
# GOOD -> GOOD 0.8, BAD -> BAD 0.6
Ps = np.array([[0.6, 0.4],[0.2, 0.8]])
# current risk free interest rate
r_f = np.array([0.01 ,0.03])
# stock return depends on current and future econ states
r_m = np.array([[-0.2, 0.15],[-0.15, 0.2]])
# probability of survival
Pa = np.load("prob.npy")
# probability of employment transition
Pe = np.array([[[[0.3, 0.7], [0.1, 0.9]], [[0.25, 0.75], [0.05, 0.95]]],
[[[0.25, 0.75], [0.05, 0.95]], [[0.2, 0.8], [0.01, 0.99]]]])
# deterministic income
detEarning = np.load("detEarning.npy")
# tax rate
tau_L = 0.2
tau_R = 0.1
# minimum consumption
c_bar = 3
t = 30
r_bar = 0.0667
N = int(np.sum(Pa[t:]))
discouting = ((1+r_bar)**N - 1)/(((1+r_bar)**N - (1+r_bar)**(N-1)))
discouting
# Define the transtiion of state
def transition(x, a, t, s_next):
'''
Input: x current state: (w, n, s, A)
a action taken: (c, b, k)
Output: the next possible states with corresponding probabilities
'''
c, b, k = a
w, s, e, A = x
x_next = []
prob_next = []
if A == 0:
for s_next in [0, 1]:
x_next.append([0, s_next, 0, 0])
return np.array(x_next), Ps[int(s)]
else:
# A = 1, agent is still alive and for the next period
Pat = [1-Pa[t], Pa[t]]
r_bond = r_f[int(s)]
r_stock = r_m[int(s), s_next]
w_next = b*(1+r_bond) + k*(1+r_stock)
for e_next in [0,1]:
for A_next in [0,1]:
x_next.append([w_next, s_next, e_next, A_next])
prob_next.append(Pat[A_next] * Pe[int(s),s_next,int(e),e_next])
return x_next[np.random.choice(4, 1, p = prob_next)]
x_possible_next, next_prob = transition([100,0,0,1], [10,10,10], 20, 0)
x_possible_next[(np.random.choice(4, 1, p=next_prob))]
import numpy as np
ww = [1,2,3,4]
a = np.array([[w,s,e,A]for w in ww for s in [0, 1] for e in [0,1] for A in [0,1]])
a.reshape((4,2,2,2,4))
a.sum(axis = 1).reshape(4,2,2,2)
a.sum(axis = 1).reshape((4,2))
import numpy as np
a = np.array([[10, 7, 4], [3, 2, 1]])
a
np.quantile(a, [0,0.5,1], axis=1)
import numpy as np
np.random.rand()
|
20200613/test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob
import os
# +
folders=os.listdir(".")
folders.sort()
content=[]
f = open("letter.txt", "a")
for folder in folders:
if(not folder.startswith(".")):
print(folder+'\n')
f.write(folder)
f.close()
# -
f = open("letter.txt", "r")
print(f.read())
|
dataset/.ipynb_checkpoints/Untitled-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/supai-red/examples/blob/master/Brusselator.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="A7Zh6n72T36W" colab_type="text"
# ### Brusselator from [The Endeavour](https://www.johndcook.com/blog/2020/02/07/behold-the-brusselator/?utm_source=feedburner&utm_medium=email&utm_campaign=Feed%3A+TheEndeavour+%28The+Endeavour%29)
# + id="R2CEboPwSxI4" colab_type="code" colab={}
from scipy import linspace
from scipy.integrate import solve_ivp
import matplotlib.pyplot as plt
# + id="N_BSNR91S3rq" colab_type="code" colab={}
A, B = 1,3
# + id="Qzi8IsqgS5Dy" colab_type="code" colab={}
def brusselator(t, z):
x, y = z
return [A + x*x*y - (B+1)*x, B*x - x*x*y]
# + id="Ve0PaxtyTGvs" colab_type="code" colab={}
a, b = 0, 10
# + id="xDiUl_k_TICi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="7663ddbb-6636-45d5-c47d-1e7673d39f5a"
t = linspace(a, b, 1000)
# + id="T4A6mYxPTJx0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="2b7bf164-5a7d-4a84-ac67-7cce5d73edc5"
for x0 in range (0,6):
for y0 in [0, 3]:
sol = solve_ivp(brusselator, [a,b], [x0, y0], t_eval=t)
plt.plot(sol.y[0], sol.y[1], ":", color="tab:blue")
plt.show()
# + id="Qcej6lGGTwQ2" colab_type="code" colab={}
|
Brusselator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
data = pd.read_csv("../data/CAB_08_RJ.csv", low_memory = False, index_col=False)
data.shape
data.head()
data.info()
data.columns.values
data = data.drop([ 'psu_id','ahs_house_unit','house_hold_no', 'date_survey',
'test_salt_iodine', 'record_code_iodine_reason', 'sl_no', 'usual_residance_reason',
'identification_code', 'date_of_birth', 'year_of_birth', 'weight_measured',
'length_height_measured', 'length_height_code', 'haemoglobin_test', 'haemoglobin', 'bp_systolic',
'bp_systolic_2_reading', 'bp_diastolic', 'bp_diastolic_2reading', 'pulse_rate',
'pulse_rate_2_reading', 'diabetes_test', 'fasting_blood_glucose', 'fasting_blood_glucose_mg_dl',
'marital_status', 'gauna_perfor_not_perfor', 'duration_pregnanacy', 'illness_duration', 'v54'],
axis = 1)
data.shape
data.columns.values
data = data[data['age'] != 'Y']
data['age'].unique()
data = data[data['age'] != -1]
data['age'].unique()
data[['age']] = data[['age']].astype(float)
data = data.drop(data[(data['age_code'] == 'Y') & (data['age'] > 5) | (data['illness_type'] == -1)].index)
data.shape
data['illness_type'].value_counts()
data['illness_type'].value_counts() / 121.2
data['age_code'].value_counts()
data['record_code_iodine'].value_counts()
data['sex'].value_counts()
data['day_or_month_for_breast_feeding_'].value_counts()
data = data[:][data['day_or_month_for_breast_feeding_'] != '12']
data['day_or_month_for_breast_feeding_'].value_counts()
data = data.drop(data[data['age_code'] == '-1'].index)
data.shape
print(data['rural_urban'].value_counts())
print(data['stratum'].value_counts())
print(data['record_code_iodine'].value_counts())
print(data['sex'].value_counts())
print(data['usual_residance'].value_counts())
print(data['age_code'].value_counts())
print(data['month_of_birth'].value_counts())
data.shape
#droping birth month
data = data.drop('month_of_birth', axis=1)
data.shape
data.columns.unique()
data = data.drop('rural_urban',axis=1)
data.columns.values
data = data.drop(['stratum', 'usual_residance'],axis=1)
data.columns.values
data['weight_in_kg'][data[data['weight_in_kg'] == -1].index].value_counts()
print(data['record_code_iodine'][data[data['record_code_iodine'] == -1].index].value_counts())
print(data['sex'][data[data['sex'] == -1].index].value_counts())
print(data['age_code'][data[data['age_code'] == -1].index].value_counts())
print(data['age'][data[data['age'] == -1].index].value_counts())
print(data['weight_in_kg'][data[data['weight_in_kg'] == -1].index].value_counts())
print(data['length_height_cm'][data[data['length_height_cm'] == -1].index].value_counts())
print(data['haemoglobin_level'][data[data['haemoglobin_level'] == -1].index].value_counts())
print(data['first_breast_feeding'][data[data['first_breast_feeding'] == -1].index].value_counts())
print(data['is_cur_breast_feeding'][data[data['is_cur_breast_feeding'] == -1].index].value_counts())
print(data['day_or_month_for_breast_feeding_'][data[data['day_or_month_for_breast_feeding_'] == -1].index].value_counts())
print(data['day_or_month_for_breast_feeding'][data[data['day_or_month_for_breast_feeding'] == -1].index].value_counts())
print(data['water_month'][data[data['water_month'] == -1].index].value_counts())
print(data['ani_milk_month'][data[data['ani_milk_month'] == -1].index].value_counts())
print(data['semisolid_month_or_day'][data[data['semisolid_month_or_day'] == -1].index].value_counts())
print(data['solid_month'][data[data['solid_month'] == -1].index].value_counts())
print(data['vegetables_month_or_day'][data[data['vegetables_month_or_day'] == -1].index].value_counts())
data.shape
data['water_month'][data[data['water_month'] == -1].index].value_counts()
data['semisolid_month_or_day'][data[data['semisolid_month_or_day'] == -1].index].value_counts()
data['is_cur_breast_feeding'][data[(data['is_cur_breast_feeding'] == -1)].index].value_counts()
data['is_cur_breast_feeding'][data[(data['is_cur_breast_feeding'] == -1) & (data['age']>2) & (data['age_code'] == 'Y')].index].value_counts()
# data['is_cur_breast_feeding'][data[(data['age_code'] == 'Y') & (data['age'] > 2) & (data['is_cur_breast_feeding'] == -1)].index] = 2
data.loc[((data['age']>2) & (data['age_code'] == 'Y') & (data['is_cur_breast_feeding'] == -1)),'is_cur_breast_feeding'] = 2
data['is_cur_breast_feeding'][data[data['is_cur_breast_feeding'] == -1].index].value_counts()
data['is_cur_breast_feeding'] = data['is_cur_breast_feeding'].replace(-1, 1)
data['age'][data[data['is_cur_breast_feeding'] == -1].index]
data['weight_in_kg'][data[(data['weight_in_kg'] == -1)].index].value_counts()
data['age_code'][data[(data['weight_in_kg'] == -1)].index].value_counts()
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['weight_in_kg'][data[(data['age_code'] == y) & (data['age'] == i) & (data['weight_in_kg'] != -1)].index].mean()
data.loc[((data['age_code'] == y) & (data['age'] == i) & ((data['weight_in_kg'] == -1) | (data['weight_in_kg'].isnull()))), 'weight_in_kg'] = x
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['weight_in_kg'][data[(data['age_code'] == y) & (data['age'] == i) & (data['weight_in_kg'] != -1)].index].mean()
data['weight_in_kg'] = data['weight_in_kg'].fillna(x)
data['weight_in_kg'][data[(data['weight_in_kg'] == -1)].index].value_counts()
data['length_height_cm'][data[(data['length_height_cm'] == -1)].index].value_counts()
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['length_height_cm'][data[(data['age_code'] == y) & (data['age'] == i) & (data['length_height_cm'] != -1)].index].mean()
data.loc[((data['age_code'] == y) & (data['age'] == i) & ((data['length_height_cm'] == -1) | (data['length_height_cm'].isnull()))), 'length_height_cm'] = x
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['length_height_cm'][data[(data['age_code'] == y) & (data['age'] == i) & (data['length_height_cm'] != -1)].index].mean()
data['length_height_cm'] = data['length_height_cm'].fillna(x)
data['length_height_cm'][data[(data['length_height_cm'] == -1)].index].value_counts()
data['haemoglobin_level'][data[(data['haemoglobin_level'] == -1)].index].value_counts()
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['haemoglobin_level'][data[(data['age_code'] == y) & (data['age'] == i) & (data['haemoglobin_level'] != -1)].index].mean()
data.loc[((data['age_code'] == y) & (data['age'] == i) & (data['haemoglobin_level'] == -1)), 'haemoglobin_level'] = x
for y in data['age_code'].unique():
for i in data['age'][data[data['age_code'] == y].index].unique():
x = data['haemoglobin_level'][data[(data['age_code'] == y) & (data['age'] == i) & (data['haemoglobin_level'] != -1)].index].mean()
data['haemoglobin_level'] = data['haemoglobin_level'].fillna(x)
data['haemoglobin_level'][data[(data['haemoglobin_level'] == -1)].index].value_counts()
data['first_breast_feeding'][data[(data['first_breast_feeding'] == -1)].index].value_counts()
for i in data['district_code'].unique():
print(i)
x = data['district_code'][data[data['district_code']==i].index].value_counts()
print(x[i])
print((data['first_breast_feeding'][data[data['district_code']==i].index].value_counts().sort_index())*100/x[i])
print("----------------------------------------------------")
for i in data['district_code'].unique():
x = data['first_breast_feeding'][data[(data['district_code'] == i) & (data['first_breast_feeding'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data.loc[((data['district_code'] == i) & (data['first_breast_feeding'] == -1)), 'first_breast_feeding'] = y[0]
for i in data['district_code'].unique():
x = data['first_breast_feeding'][data[(data['district_code'] == i) & (data['first_breast_feeding'] != -1)].index].value_counts()
y = x[x == x.max()].index.item()
data['first_breast_feeding'] = data['first_breast_feeding'].fillna(y)
data['first_breast_feeding'][data[(data['first_breast_feeding'] == -1)].index].value_counts()
data['day_or_month_for_breast_feeding'][data[(data['day_or_month_for_breast_feeding'] == -1)].index].value_counts()
for i in data['district_code'].unique():
print(i)
x = data['district_code'][data[data['district_code']==i].index].value_counts()
print(x[i])
print((data['day_or_month_for_breast_feeding'][data[data['district_code']==i].index].value_counts().sort_index())*100/x[i])
print("----------------------------------------------------")
data.loc[(data['day_or_month_for_breast_feeding'] == -1), 'day_or_month_for_breast_feeding'] = 6
data['day_or_month_for_breast_feeding'][data[(data['day_or_month_for_breast_feeding'] == -1)].index].value_counts()
print(data['water_month'][data[data['water_month'] == -1].index].value_counts())
print(data['ani_milk_month'][data[data['ani_milk_month'] == -1].index].value_counts())
print(data['semisolid_month_or_day'][data[data['semisolid_month_or_day'] == -1].index].value_counts())
print(data['solid_month'][data[data['solid_month'] == -1].index].value_counts())
print(data['vegetables_month_or_day'][data[data['vegetables_month_or_day'] == -1].index].value_counts())
for i in data['district_code'].unique():
print(i)
x = data['water_month'][data[(data['district_code'] == i) & (data['water_month'] != -1)].index].value_counts()
print(x)
y = x[x == x.max()].index.tolist()
print(y[0])
data.loc[((data['district_code'] == i) & (data['water_month'] == -1)), 'water_month'] = y[0]
for i in data['district_code'].unique():
x = data['water_month'][data[(data['district_code'] == i) & (data['water_month'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data['water_month'] = data['water_month'].fillna(y[0])
for i in data['district_code'].unique():
print(i)
x = data['ani_milk_month'][data[(data['district_code'] == i) & (data['ani_milk_month'] != -1)].index].value_counts()
print(x)
y = x[x == x.max()].index.tolist()
print(y[0])
data.loc[((data['district_code'] == i) & (data['ani_milk_month'] == -1)), 'ani_milk_month'] = y[0]
for i in data['district_code'].unique():
x = data['ani_milk_month'][data[(data['district_code'] == i) & (data['ani_milk_month'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data['ani_milk_month'] = data['ani_milk_month'].fillna(y[0])
for i in data['district_code'].unique():
print(i)
x = data['semisolid_month_or_day'][data[(data['district_code'] == i) & (data['semisolid_month_or_day'] != -1)].index].value_counts()
print(x)
y = x[x == x.max()].index.tolist()
print(y[0])
data.loc[((data['district_code'] == i) & (data['semisolid_month_or_day'] == -1)), 'semisolid_month_or_day'] = y[0]
for i in data['district_code'].unique():
x = data['semisolid_month_or_day'][data[(data['district_code'] == i) & (data['semisolid_month_or_day'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data['semisolid_month_or_day'] = data['semisolid_month_or_day'].fillna(y[0])
for i in data['district_code'].unique():
print(i)
x = data['solid_month'][data[(data['district_code'] == i) & (data['solid_month'] != -1)].index].value_counts()
print(x)
y = x[x == x.max()].index.tolist()
print(y[0])
data.loc[((data['district_code'] == i) & (data['solid_month'] == -1)), 'solid_month'] = y[0]
for i in data['district_code'].unique():
x = data['solid_month'][data[(data['district_code'] == i) & (data['solid_month'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data['solid_month'] = data['solid_month'].fillna(y[0])
for i in data['district_code'].unique():
print(i)
x = data['vegetables_month_or_day'][data[(data['district_code'] == i) & (data['vegetables_month_or_day'] != -1)].index].value_counts()
print(x)
y = x[x == x.max()].index.tolist()
print(y[0])
data.loc[((data['district_code'] == i) & (data['vegetables_month_or_day'] == -1)), 'vegetables_month_or_day'] = y[0]
for i in data['district_code'].unique():
x = data['vegetables_month_or_day'][data[(data['district_code'] == i) & (data['vegetables_month_or_day'] != -1)].index].value_counts()
y = x[x == x.max()].index.tolist()
data['vegetables_month_or_day'] = data['vegetables_month_or_day'].fillna(y[0])
print(data['water_month'][data[data['water_month'] == -1].index].value_counts())
print(data['ani_milk_month'][data[data['ani_milk_month'] == -1].index].value_counts())
print(data['semisolid_month_or_day'][data[data['semisolid_month_or_day'] == -1].index].value_counts())
print(data['solid_month'][data[data['solid_month'] == -1].index].value_counts())
print(data['vegetables_month_or_day'][data[data['vegetables_month_or_day'] == -1].index].value_counts())
data = data.drop('district_code', axis=1)
data.columns.values
data.shape
for i in data[data['day_or_month_for_breast_feeding_'] == 'D'].index:
print(i,' ',data['day_or_month_for_breast_feeding_'][i],' ',data['day_or_month_for_breast_feeding'][i])
if data['day_or_month_for_breast_feeding'][i] < 15:
data['day_or_month_for_breast_feeding'][i] = 0
else :
data['day_or_month_for_breast_feeding'][i] = 1
print(i,' ',data['day_or_month_for_breast_feeding_'][i],' ',data['day_or_month_for_breast_feeding'][i])
for i in data[data['age_code'] == 'Y'].index:
x = data['age'][i]
print(i,' ',data['age_code'][i],' ',data['age'][i])
data['age'][i] = x*12
print(i,' ',data['age_code'][i],' ',data['age'][i])
for i in data[data['age_code'] == 'D'].index:
x = data['age'][i]
print(i,' ',data['age_code'][i],' ',data['age'][i])
if data['age'][i] < 15:
data['age'][i] = 0
else :
data['age'][i] = 1
print(i,' ',data['age_code'][i],' ',data['age'][i])
data.columns.values
data = data.drop(['age_code', 'day_or_month_for_breast_feeding_','treatment_type'] , axis=1)
data.columns.values
data.shape
for i in data.columns:
print(i)
print(data['state_code'][data[i] == -1].value_counts())
print(data['state_code'][data[i].isnull()].value_counts())
# +
# data.to_csv('../cleaned data/RJ.csv', encoding='utf-8', index=False)
|
ML/analysis scripts/analysis_RJ.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic Regression
#
# ## Binary classification problem
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from time import process_time
# %matplotlib inline
# ### 1. Visualizing the data
data = pd.read_csv("ex2data1.txt", header=None, names=['exam1', 'exam2', 'admitted'])
data.head()
# +
positive = data[data.admitted == 1]
negative = data[data.admitted == 0]
fig, ax = plt.subplots(figsize=(12, 8))
ax.scatter(positive.exam1, positive.exam2, s=50, c='b', marker='o', label='Admitted')
ax.scatter(negative.exam1, negative.exam2, s=50, c='r', marker='x', label='Not admitted')
ax.legend()
ax.set_xlabel('Exam 1 score')
ax.set_ylabel('Exam 2 score')
# -
data.insert(0, 'x0', 1)
data.head()
col = data.shape[1]
data_x = data.iloc[:, :col - 1].values
data_y = data.iloc[:, col - 1:].values.ravel()
theta = np.zeros(data_x.shape[1])
print(data_x.shape)
print(data_y.shape)
print(theta.shape)
# ### 2. Implementation
#
# #### 2.1 Sigmoid function
#
# The logistic regression hypothesis is defined as $$h_\theta(x)=g(\theta^Tx)$$where function g is the sigmoid function, and it is defined as$$g(z)=\frac{1}{1 + e^{-x}}$$
def sigmoid(z):
return 1 / (1 + np.exp(-z))
x = np.linspace(-10, 10, 100)
plt.plot(x, sigmoid(x))
# #### 2.2 Cost function
#
# The cost function in logistic regression is$$J(\theta)=-\frac{1}{m}\sum_{i=1}^m\left[y^{(i)}log \left( h_\theta(x^{(i)}) \right) + (1 - y^{(i)})log \left( 1-h_\theta(x^{(i)}) \right) \right]$$
def cost(theta, x, y):
return np.mean(-y * np.log(sigmoid(x @ theta)) - (1 - y) * np.log(1 - sigmoid(x @ theta)))
cost(theta, data_x, data_y)
# #### 2.3 Gradient
#
# And the gradient of the cost is defined as follow$$\frac{\partial J(\theta)}{\partial \theta_j}=\frac{1}{m}\sum_{i=1}^m\left(h_\theta(x^{(i)})-y^{(i)}\right)x_j^{(i)}$$Each iteration performs the update until convergence$$\theta_j=\theta_j-\alpha\frac{1}{m}\sum^m_{i=1}\left[h_{\theta}(x^{(i)})-y^{(i)}\right]x_j^{(i)}$$
def gradient(theta, x, y):
return (1 / len(x)) * x.T @ (sigmoid(x @ theta) - y)
def logistic_regression(theta, x, y, alpha, iters):
costs = []
st = process_time()
for i in range(iters):
theta = theta - alpha * gradient(theta, x, y)
this_cost = cost(theta, x, y)
costs.append(this_cost)
if i % 50000 == 0 and i != 0:
print("Index: {:>7d}, cost: {:>.6f}".format(i, this_cost))
ed = process_time()
print("\n----Train finish----")
print("Time cost:{:>.3f}".format(ed - st))
print("Final cost:{:>.3f}".format(cost(theta, data_x, data_y)))
print("theta: {}".format(theta))
return theta, costs
theta, costs = logistic_regression(theta, data_x, data_y, 0.001, 1000000)
# #### 2.4 Evaluating
#
# Decision boundry is defined in this case as$$\theta_0 + \theta_1x_1 + \theta_2x_2 = 0$$the line on the coordinate plane is $$x_2 = -\frac{\theta_0}{\theta_2} - \frac{\theta_1}{\theta_2}x_1$$
# +
parm = [-theta[0] / theta[2], -theta[1] / theta[2]]
x = np.linspace(data.exam1.min(), data.exam2.max(), 100)
y = np.dot(np.insert(x.reshape(100, 1), 0, np.ones(100), axis=1), parm)
fig, ax = plt.subplots(figsize=(12, 8))
ax.scatter(positive.exam1, positive.exam2, s=50, c='b', marker='o', label='Admitted')
ax.scatter(negative.exam1, negative.exam2, s=50, c='r', marker='x', label='Not admitted')
ax.plot(x, y, c='k', label='Decision boundry')
ax.legend()
ax.set_xlabel("Exam 1 score")
ax.set_ylabel("Exam 2 score")
# -
fig, ax = plt.subplots(figsize=(12, 8))
ax.plot(costs)
ax.set_xlabel("Iterations")
ax.set_ylabel("Cost")
|
exercise/ex2_logistic_reg/1_logistic_reg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib
# Matplotlib is an amazing visualization library in Python for 2D plots of arrays.
import matplotlib.pyplot as plt
# ## Plotting an array
x = [10,20,30]
y = [3,2,1]
plt.plot(x,y)
# ### Scatter
plt.scatter(x,y)
# ### Bar Plot
plt.bar(x,y)
# +
# Bar graphs make more sense when x-axis shows categorical data
x = ["new","old"]
y = [7,4]
plt.bar(x,y)
# +
# Be careful of repeated categories
# Height of the bar represents the max value in that category
x = ["new","old","new"]
y = [2,4,5]
plt.bar(x,y)
# -
plt.scatter(x,y)
# ## Plotting Insights from DataFrames
# ### Linear Regression
import pandas as pd
dataframe = pd.read_csv('startups.csv')
dataframe
# +
# We need a Series so use single brackets [] to index the column
x = dataframe['R&D Spend']
x
# -
y = dataframe['Profit']
y
# +
# Linear Regression
plt.scatter(x,y)
# -
x = dataframe['Marketing Spend']
plt.scatter(x,y)
|
workshop5/matplotlib.ipynb
|
# + [markdown] colab_type="text" id="a3bskVXPvchm"
# # Hello, TensorFlow
# ## A beginner-level, getting started, basic introduction to TensorFlow
# + [markdown] colab_type="text" id="Rb5rSpcZvYbX"
# TensorFlow is a general-purpose system for graph-based computation. A typical use is machine learning. In this notebook, we'll introduce the basic concepts of TensorFlow using some simple examples.
#
# TensorFlow gets its name from [tensors](https://en.wikipedia.org/wiki/Tensor), which are arrays of arbitrary dimensionality. A vector is a 1-d array and is known as a 1st-order tensor. A matrix is a 2-d array and a 2nd-order tensor. The "flow" part of the name refers to computation flowing through a graph. Training and inference in a neural network, for example, involves the propagation of matrix computations through many nodes in a computational graph.
#
# When you think of doing things in TensorFlow, you might want to think of creating tensors (like matrices), adding operations (that output other tensors), and then executing the computation (running the computational graph). In particular, it's important to realize that when you add an operation on tensors, it doesn't execute immediately. Rather, TensorFlow waits for you to define all the operations you want to perform. Then, TensorFlow optimizes the computation graph, deciding how to execute the computation, before generating the data. Because of this, a tensor in TensorFlow isn't so much holding the data as a placeholder for holding the data, waiting for the data to arrive when a computation is executed.
# + [markdown] colab_type="text" id="E8FhiMivhcYB"
# ## Adding two vectors in TensorFlow
#
# Let's start with something that should be simple. Let's add two length four vectors (two 1st-order tensors):
#
# $\begin{bmatrix} 1. & 1. & 1. & 1.\end{bmatrix} + \begin{bmatrix} 2. & 2. & 2. & 2.\end{bmatrix} = \begin{bmatrix} 3. & 3. & 3. & 3.\end{bmatrix}$
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 2922, "status": "ok", "timestamp": 1474675631337, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="2iv3XQ6k3eF1" outputId="7dbded62-91bc-4e38-9f25-53375c4c8dd8"
from __future__ import print_function
import tensorflow as tf
with tf.Session():
input1 = tf.constant([1.0, 1.0, 1.0, 1.0])
input2 = tf.constant([2.0, 2.0, 2.0, 2.0])
output = tf.add(input1, input2)
result = output.eval()
print("result: ", result)
# + [markdown] colab_type="text" id="dqLV5GXT3wLy"
# What we're doing is creating two vectors, [1.0, 1.0, 1.0, 1.0] and [2.0, 2.0, 2.0, 2.0], and then adding them. Here's equivalent code in raw Python and using numpy:
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 214, "status": "ok", "timestamp": 1474675631563, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="7DzDJ7sW79ao" outputId="588b573b-95d2-4587-849e-af6f3ec1303e"
print([x + y for x, y in zip([1.0] * 4, [2.0] * 4)])
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 340, "status": "ok", "timestamp": 1474675631948, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="MDWJf0lHAF4E" outputId="bee09475-24dd-4331-fc46-692a07dae101"
import numpy as np
x, y = np.full(4, 1.0), np.full(4, 2.0)
print("{} + {} = {}".format(x, y, x + y))
# + [markdown] colab_type="text" id="I52jQOyO8vAn"
# ## Details of adding two vectors in TensorFlow
#
# The example above of adding two vectors involves a lot more than it seems, so let's look at it in more depth.
#
# >`import tensorflow as tf`
#
# This import brings TensorFlow's public API into our IPython runtime environment.
#
# >`with tf.Session():`
#
# When you run an operation in TensorFlow, you need to do it in the context of a `Session`. A session holds the computation graph, which contains the tensors and the operations. When you create tensors and operations, they are not executed immediately, but wait for other operations and tensors to be added to the graph, only executing when finally requested to produce the results of the session. Deferring the execution like this provides additional opportunities for parallelism and optimization, as TensorFlow can decide how to combine operations and where to run them after TensorFlow knows about all the operations.
#
# >>`input1 = tf.constant([1.0, 1.0, 1.0, 1.0])`
#
# >>`input2 = tf.constant([2.0, 2.0, 2.0, 2.0])`
#
# The next two lines create tensors using a convenience function called `constant`, which is similar to numpy's `array` and numpy's `full`. If you look at the code for `constant`, you can see the details of what it is doing to create the tensor. In summary, it creates a tensor of the necessary shape and applies the constant operator to it to fill it with the provided values. The values to `constant` can be Python or numpy arrays. `constant` can take an optional shape parameter, which works similarly to numpy's `fill` if provided, and an optional name parameter, which can be used to put a more human-readable label on the operation in the TensorFlow operation graph.
#
# >>`output = tf.add(input1, input2)`
#
# You might think `add` just adds the two vectors now, but it doesn't quite do that. What it does is put the `add` operation into the computational graph. The results of the addition aren't available yet. They've been put in the computation graph, but the computation graph hasn't been executed yet.
#
# >>`result = output.eval()`
#
# >>`print result`
#
# `eval()` is also slightly more complicated than it looks. Yes, it does get the value of the vector (tensor) that results from the addition. It returns this as a numpy array, which can then be printed. But, it's important to realize it also runs the computation graph at this point, because we demanded the output from the operation node of the graph; to produce that, it had to run the computation graph. So, this is the point where the addition is actually performed, not when `add` was called, as `add` just put the addition operation into the TensorFlow computation graph.
# + [markdown] colab_type="text" id="H_5_2YY3ySr2"
# ## Multiple operations
#
# To use TensorFlow, you add operations on tensors that produce tensors to the computation graph, then execute that graph to run all those operations and calculate the values of all the tensors in the graph.
#
# Here's a simple example with two operations:
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 1203, "status": "ok", "timestamp": 1474675633108, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="-kQmn3U_yXX8" outputId="8ba14a4d-b0cd-4b90-8b95-790e77d35e70"
import tensorflow as tf
with tf.Session():
input1 = tf.constant(1.0, shape=[4])
input2 = tf.constant(2.0, shape=[4])
input3 = tf.constant(3.0, shape=[4])
output = tf.add(tf.add(input1, input2), input3)
result = output.eval()
print(result)
# + [markdown] colab_type="text" id="Hod0zvsly8YT"
# This version uses `constant` in a way similar to numpy's `fill`, specifying the optional shape and having the values copied out across it.
#
# The `add` operator supports operator overloading, so you could try writing it inline as `input1 + input2` instead as well as experimenting with other operators.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 350, "status": "ok", "timestamp": 1474675633468, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="yS2WElRfxz53" outputId="2e3efae6-3990-447c-e05d-56a9d9701e87"
with tf.Session():
input1 = tf.constant(1.0, shape=[4])
input2 = tf.constant(2.0, shape=[4])
output = input1 + input2
print(output.eval())
# + [markdown] colab_type="text" id="zszjoYUjkUNU"
# ## Adding two matrices
# + [markdown] colab_type="text" id="EWNYBCB6kbri"
# Next, let's do something very similar, adding two matrices:
#
# $\begin{bmatrix}
# 1. & 1. & 1. \\
# 1. & 1. & 1. \\
# \end{bmatrix} +
# \begin{bmatrix}
# 1. & 2. & 3. \\
# 4. & 5. & 6. \\
# \end{bmatrix} =
# \begin{bmatrix}
# 2. & 3. & 4. \\
# 5. & 6. & 7. \\
# \end{bmatrix}$
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 1327, "status": "ok", "timestamp": 1474675634683, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="tmWcCxSilYkg" outputId="8a135ccf-e706-457c-f4bc-2187039ffd92"
import tensorflow as tf
import numpy as np
with tf.Session():
input1 = tf.constant(1.0, shape=[2, 3])
input2 = tf.constant(np.reshape(np.arange(1.0, 7.0, dtype=np.float32), (2, 3)))
output = tf.add(input1, input2)
print(output.eval())
# + [markdown] colab_type="text" id="JuU3Bmglq1vd"
# Recall that you can pass numpy or Python arrays into `constant`.
#
# In this example, the matrix with values from 1 to 6 is created in numpy and passed into `constant`, but TensorFlow also has `range`, `reshape`, and `tofloat` operators. Doing this entirely within TensorFlow could be more efficient if this was a very large matrix.
#
# Try experimenting with this code a bit -- maybe modifying some of the values, using the numpy version, doing this using, adding another operation, or doing this using TensorFlow's `range` function.
# + [markdown] colab_type="text" id="gnXnpnuLrflb"
# ## Multiplying matrices
# + [markdown] colab_type="text" id="Ho-QNSOorj0y"
# Let's move on to matrix multiplication. This time, let's use a bit vector and some random values, which is a good step toward some of what we'll need to do for regression and neural networks.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 2353, "status": "ok", "timestamp": 1474675637053, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="uNqMaFR8sIY5" outputId="b630554e-68b3-4904-c07d-f28a0a41bbd2"
#@test {"output": "ignore"}
import tensorflow as tf
import numpy as np
with tf.Session():
input_features = tf.constant(np.reshape([1, 0, 0, 1], (1, 4)).astype(np.float32))
weights = tf.constant(np.random.randn(4, 2).astype(np.float32))
output = tf.matmul(input_features, weights)
print("Input:")
print(input_features.eval())
print("Weights:")
print(weights.eval())
print("Output:")
print(output.eval())
# + [markdown] colab_type="text" id="JDAVTPhb22AP"
# Above, we're taking a 1 x 4 vector [1 0 0 1] and multiplying it by a 4 by 2 matrix full of random values from a normal distribution (mean 0, stdev 1). The output is a 1 x 2 matrix.
#
# You might try modifying this example. Running the cell multiple times will generate new random weights and a new output. Or, change the input, e.g., to \[0 0 0 1]), and run the cell again. Or, try initializing the weights using the TensorFlow op, e.g., `random_normal`, instead of using numpy to generate the random weights.
#
# What we have here is the basics of a simple neural network already. If we are reading in the input features, along with some expected output, and change the weights based on the error with the output each time, that's a neural network.
# + [markdown] colab_type="text" id="XhnBjAUILuy8"
# ## Use of variables
#
# Let's look at adding two small matrices in a loop, not by creating new tensors every time, but by updating the existing values and then re-running the computation graph on the new data. This happens a lot with machine learning models, where we change some parameters each time such as gradient descent on some weights and then perform the same computations over and over again.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" executionInfo={"elapsed": 2561, "status": "ok", "timestamp": 1474675639610, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 420} id="vJ_AgZ8lLtRv" outputId="b8f19c28-a9b4-4fb3-9e90-6e432bf300a7"
#@test {"output": "ignore"}
import tensorflow as tf
import numpy as np
with tf.Session() as sess:
# Set up two variables, total and weights, that we'll change repeatedly.
total = tf.Variable(tf.zeros([1, 2]))
weights = tf.Variable(tf.random_uniform([1,2]))
# Initialize the variables we defined above.
tf.global_variables_initializer().run()
# This only adds the operators to the graph right now. The assignment
# and addition operations are not performed yet.
update_weights = tf.assign(weights, tf.random_uniform([1, 2], -1.0, 1.0))
update_total = tf.assign(total, tf.add(total, weights))
for _ in range(5):
# Actually run the operation graph, so randomly generate weights and then
# add them into the total. Order does matter here. We need to update
# the weights before updating the total.
sess.run(update_weights)
sess.run(update_total)
print(weights.eval(), total.eval())
# + [markdown] colab_type="text" id="kSYJr89aM_n0"
# This is more complicated. At a high level, we create two variables and add operations over them, then, in a loop, repeatedly execute those operations. Let's walk through it step by step.
#
# Starting off, the code creates two variables, `total` and `weights`. `total` is initialized to \[0, 0\] and `weights` is initialized to random values between -1 and 1.
#
# Next, two assignment operators are added to the graph, one that updates weights with random values from [-1, 1], the other that updates the total with the new weights. Again, the operators are not executed here. In fact, this isn't even inside the loop. We won't execute these operations until the `eval` call inside the loop.
#
# Finally, in the for loop, we run each of the operators. In each iteration of the loop, this executes the operators we added earlier, first putting random values into the weights, then updating the totals with the new weights. This call uses `eval` on the session; the code also could have called `eval` on the operators (e.g. `update_weights.eval`).
#
# It can be a little hard to wrap your head around exactly what computation is done when. The important thing to remember is that computation is only performed on demand.
#
# Variables can be useful in cases where you have a large amount of computation and data that you want to use over and over again with just a minor change to the input each time. That happens quite a bit with neural networks, for example, where you just want to update the weights each time you go through the batches of input data, then run the same operations over again.
# + [markdown] colab_type="text" id="fL3WfAbKzqr5"
# ## What's next?
#
# This has been a gentle introduction to TensorFlow, focused on what TensorFlow is and the very basics of doing anything in TensorFlow. If you'd like more, the next tutorial in the series is Getting Started with TensorFlow, also available in the [notebooks directory](..).
|
tutorials_previous/2_tensorflow_intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import PoisDenoiser
import torch as th
import numpy as np
from time import time, sleep
from PoisDenoiser.dataset_loader import BSDS500
from PoisDenoiser.utils import show_images as show
from PoisDenoiser.utils import psnr
import matplotlib.pyplot as plt
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import wavelet_transform as wt
# +
img_ind = 10
path2dataset = './DATASETS/BSDS500/BSDS500_validation_MAXVALs_01_2/'
BSDSval = BSDS500(path2dataset+'val/', get_name=True)
clear, noisy, name = BSDSval[img_ind]
print(name)
images = [clear, noisy]
titles = ['clear', 'noisy']
show(images, titles)
print('psnr : {}'.format(PoisDenoiser.utils.psnr(clear, noisy)))
# clear = clear[0]
# noisy = noisy[0]
# -
slice_ = (slice(0,1),
slice(50, 200),
slice(150,300))
clear = clear[slice_]
noisy = noisy[slice_]
# +
WT = wt.WaveletTransform('haar', 8, size=256)
psnrs_noisy, psnrs_clear = [], []
wt_noisy = WT.as_vector(WT.W(noisy))
masked = WT.as_vector(WT.masked_coefs(WT.W(noisy)))
epss = list(np.linspace(0.5, 10., 50))
for eps in epss:
thresh = masked*(np.abs(masked)>eps)
rec = WT.as_coefs(thresh)
rec[0] = WT.approx_coefs
inv = WT.W_inv(rec)
# psnrs_noisy.append(psnr(th.Tensor(inv), th.Tensor(noisy)).numpy())
psnrs_clear.append(psnr(th.Tensor(inv), th.Tensor(clear)).numpy())
# -
# plt.plot(epss, psnrs_noisy)
plt.plot(epss, psnrs_clear)
|
Wt_check.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # deep RNN
# In notebook 13, 3 layers of RNN did pretty well with 16 nodes each.
# Here we try the same design using 3 layers of RNN with 64 nodes each.
# Perhaps 16 nodes is similar to classification by 2-mers, and 64 nodes is similar to 3-mers.
# With ragged tensors, training only reached 0.5248 after an hour and would have taken 12 hours for 5 epochs.
# Thus, we try padding sequences so we can use regular tensors.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import ShuffleSplit
import tensorflow as tf
from tensorflow import keras
tf.keras.backend.set_floatx('float32') # save RAM
# -
# Load and encode the sequence data.
# +
MIN_SEQ_LEN=200
#MIN_SEQ_LEN=1000 ### for testing
MAX_SEQ_LEN=25000 ### this screens 4 outliers in the complete dataset
MAX_SEQ_LEN=15000 ### because it otherwise crashes on Mac
#MAX_SEQ_LEN=2000 ### we use this to reduce training RAM and CPU
DEFLINE='>'
ncfile='ncRNA.fasta'
pcfile='pcRNA.fasta'
#ncfile='tiny.ncRNA.fasta' # 10 seqs for faster debugging
#pcfile='tiny.pcRNA.fasta'
# Assume file was preprocessed to contain one line per seq.
# Returned structure is ndarray of ndarray i.e no python lists inside.
def load_fasta(filename):
seqs=[]
with open (filename,'r') as infile:
for line in infile:
if line[0]!=DEFLINE and len(line)>=MIN_SEQ_LEN and len(line)<=MAX_SEQ_LEN:
line=line.rstrip()
pad_len = MAX_SEQ_LEN-len(line)
pad_chars = 'N' * pad_len
line = line + pad_chars
chars=np.array(list(line))
seqs.append(chars.reshape(-1, 1)) # reshaped changes (any,) to (any,1)
nparray=np.array(seqs)
return nparray
print("Load "+ncfile)
nc_seqs = load_fasta(ncfile)
print("Load "+pcfile)
pc_seqs = load_fasta(pcfile)
encoder = OneHotEncoder(handle_unknown='ignore',sparse=False)
#seq=tf.reshape(nc_seqs[0],shape=(-1, 1)) # tensor flow version
seq=nc_seqs[0].reshape(-1, 1)
encoder.fit(seq)
print("Encoder categories")
encoder.categories_
print("Ecode the non-coding sequences")
nc_list=[]
for seq in nc_seqs:
encoded=encoder.transform(seq)
nc_list.append(encoded)
nc_all=np.array(nc_list) # .reshape(-1,1)
print("shape: "+str(nc_all.shape))
print("element 0 is a sequence: \n"+str(nc_all[0]))
print("element 0,0 is one letter: "+str(nc_all[0][0]))
print("Encode the protein-coding sequences")
pc_list=[]
for seq in pc_seqs:
encoded=encoder.transform(seq)
pc_list.append(encoded)
pc_all=np.array(pc_list) # .reshape(-1,1)
pc_all.shape
# -
# Create train and validation sets.
# +
nc_labels=np.zeros(shape=(17711))
pc_labels=np.ones(shape=(20152))
#nc_labels=np.zeros(shape=(10)) # fast debugging
#pc_labels=np.ones(shape=(10))
#nc_labeled=np.concatenate((nc_all,nc_labels),axis=1)
#pc_labeled=np.concatenate((pc_all,pc_labels),axis=1)
all_seqs=np.concatenate((nc_all,pc_all),axis=0)
all_labels=np.concatenate((nc_labels,pc_labels),axis=0)
print("shape of sequences, shape of labels")
all_seqs.shape,all_labels.shape
splitter = ShuffleSplit(n_splits=1, test_size=0.2, random_state=37863)
for train_index,test_index in splitter.split(all_seqs):
train_seqs = all_seqs[train_index]
train_labels = all_labels[train_index]
test_seqs = all_seqs[test_index]
test_labels = all_labels[test_index]
print("shape of train")
print(train_seqs.shape,train_labels.shape)
print("shape of test")
print(test_seqs.shape,test_labels.shape)
print("Convert numpy array to python 3D array")
def numpy_to_python_3D(np_seqs):
one_set = []
tlen = len(np_seqs)
for i in range(tlen): # for every sequence in set
one_seq = []
slen = len(np_seqs[i])
for j in range(slen): # for ever letter in sequence
one_letter=np_seqs[i][j]
one_seq.append(one_letter)
one_set.append(one_seq)
return one_set
train_seqs = numpy_to_python_3D(train_seqs)
test_seqs = numpy_to_python_3D(test_seqs)
train_labels = train_labels.tolist()
test_labels = test_labels.tolist()
# Now that values are shuffled, partition gives random sample.
data_size=len(train_seqs)
PARTITION=int(data_size*0.8)
print("Partition = "+str(PARTITION))
print("Partition train/validation")
X_train=train_seqs[:PARTITION]
X_valid=train_seqs[PARTITION:]
y_train=train_labels[:PARTITION]
y_valid=train_labels[PARTITION:]
print("Lengths of train,valid")
print((len(X_train),len(X_valid)))
# Free memory for the next step
nc_seqs=None
pc_seqs=None
all_seqs=None
nc_labels=None
pc_labels=None
train_seqs=None
train_labels=None
# -
print("Convert to tensors")
X_train = tf.constant(X_train) # This takes a long time and hogs memory
print(type(X_train))
print(X_train.shape)
y_train = tf.convert_to_tensor(y_train)
print(type(y_train))
print(y_train.shape)
X_valid = tf.constant(X_valid)
print(type(X_valid))
print(X_valid.shape)
y_valid = tf.convert_to_tensor(y_valid)
print(type(y_valid))
print(y_valid.shape)
# +
print("Build the model")
batch_size=None # none indicates variable length
input_features=4+1 # one hot encoding of ACGT + N
rnn2 = keras.models.Sequential([
keras.layers.SimpleRNN(64, return_sequences=True, input_shape=[MAX_SEQ_LEN,input_features]),
keras.layers.SimpleRNN(32, return_sequences=True),
keras.layers.SimpleRNN(16, return_sequences=True),
keras.layers.SimpleRNN(1),
])
print("Build the training environment")
bc=tf.keras.losses.BinaryCrossentropy(from_logits=False)
rnn2.compile(loss=bc, optimizer="Adam",metrics=["accuracy"])
rnn2.summary()
# -
print("Train the model")
history = rnn2.fit(X_train,y_train,epochs=5,validation_data=(X_valid,y_valid))
print("Visualize training history.")
pd.DataFrame(history.history).plot(figsize=(8,5))
plt.grid(True)
plt.gca().set_ylim(0,1)
plt.show()
# Resources.
# [Working with RNNs](https://keras.io/guides/working_with_rnns/).
# [Recurrent Neural Networks with Keras](https://www.tensorflow.org/guide/keras/rnn#rnns_with_listdict_inputs_or_nested_inputs).
# Function tf.convert_to_tensor [docs](https://www.tensorflow.org/api_docs/python/tf/convert_to_tensor).
# Function tf.reshape [docs](https://www.tensorflow.org/api_docs/python/tf/reshape).
# Ragged Tensors [tutorial](https://www.tensorflow.org/guide/tensor#ragged_tensors)
# and [docs](https://www.tensorflow.org/api_docs/python/tf/RaggedTensor#documenting_raggedtensor_shapes_2) and [module](https://www.tensorflow.org/api_docs/python/tf/ragged).
# Incredible speedup for convert to tensor by sirfz on [stackoverflow](https://stackoverflow.com/questions/44353509/tensorflow-tf-constant-initializer-is-very-slow).
#
#
#
|
Project/lncRNA_13b_deep_RNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="QQHZHevuXdEy"
# # **QSAR Model Building of Acetylcholinesterase Inhibitors**
# + [markdown] id="g1qtHa0zXfWM"
# # Read in data
# + id="9MdfbvFKXtXq"
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="nerGP0fCXfgP" outputId="b639892c-8f7b-4b24-d02c-628c9f1a1460"
ds = '../data/acetylcholinesterase_06_bioactivity_data_3class_pIC50_pubchem_fp.csv'
dataset = pd.read_csv(ds)
dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="tgFxx8m_YEUy" outputId="47903560-3aa4-497f-85b4-27b1361b200a"
X = dataset.drop(['pIC50'], axis=1)
X
# + colab={"base_uri": "https://localhost:8080/"} id="JDwxgKHqYmD4" outputId="472cd19d-7dab-4f16-b03d-fab52d3fc782"
Y = dataset.iloc[:,-1]
Y
# + [markdown] id="AQ9E0xUY_o_M"
# # Remove low variance features
# + colab={"base_uri": "https://localhost:8080/"} id="Qkgj-lsG_wOJ" outputId="d3d11b21-fdf3-4cfb-e0c8-0f2b5a1d779b"
from sklearn.feature_selection import VarianceThreshold
def remove_low_variance(input_data, threshold=0.1):
selection = VarianceThreshold(threshold)
selection.fit(input_data)
return input_data[input_data.columns[selection.get_support(indices=True)]]
X = remove_low_variance(X, threshold=0.1)
X
# -
X.to_csv('descriptor_list.csv', index = False)
# In the app, use the following to get this same descriptor list
# of 221 variables from the initial set of 881 variables
# +
# Xlist = list(pd.read_csv('descriptor_list.csv').columns)
# X[Xlist]
# + [markdown] id="LNohCdqQY5VZ"
# # Random Forest Regression Model
# + id="EanoyG2eX9cV"
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error, r2_score
# + colab={"base_uri": "https://localhost:8080/"} id="mLQJ2KLLY_9a" outputId="e0c22032-02af-40ca-c5c9-536982ec8627"
model = RandomForestRegressor(n_estimators=500, random_state=42)
model.fit(X, Y)
r2 = model.score(X, Y)
r2
# + [markdown] id="F5f8KGWjZRSc"
# ## Model Prediction
# + colab={"base_uri": "https://localhost:8080/"} id="MI3c8LB2ZCYW" outputId="1bc71664-c9f8-434f-a666-7f78a2d34da6"
Y_pred = model.predict(X)
Y_pred
# + [markdown] id="fXv7bcolZqa-"
# ## Model Performance
# + colab={"base_uri": "https://localhost:8080/"} id="6f13gYleZVKy" outputId="fd565d7f-26e4-45d1-89f7-55b37687746c"
print('Mean squared error (MSE): %.2f'
% mean_squared_error(Y, Y_pred))
print('Coefficient of determination (R^2): %.2f'
% r2_score(Y, Y_pred))
# + [markdown] id="uWvxj1iSaL3n"
# # Data Visualization (Experimental vs Predicted pIC50 for Training Data)
# + id="iPcFF0MjZlh8"
import matplotlib.pyplot as plt
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 351} id="QRNyIlGAaQQI" outputId="1cf12d14-4ba5-49d4-e627-902a049fba2e"
plt.figure(figsize=(5,5))
plt.scatter(x=Y, y=Y_pred, c="#7CAE00", alpha=0.3)
# Add trendline
# https://stackoverflow.com/questions/26447191/how-to-add-trendline-in-python-matplotlib-dot-scatter-graphs
z = np.polyfit(Y, Y_pred, 1)
p = np.poly1d(z)
plt.plot(Y,p(Y),"#F8766D")
plt.ylabel('Predicted pIC50')
plt.xlabel('Experimental pIC50')
# + [markdown] id="YzKTmvZrbFVI"
# # Save Model as Pickle Object
# + id="DzjpPyVyb8XO"
import pickle
# + id="b2K9ajBaaYUk"
pickle.dump(model, open('acetylcholinesterase_model.pkl', 'wb'))
|
bioactivity-prediction-app/bioactivity_prediction_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3_swc
# ---
# # Errors and Exceptions
#
# Programming is essentially making mistakes, then correcting them until the code runs correctly. You **will** encounter errors and problems in the code and, although they are always frustrating, understanding how to identify and correct (or cope with) them is a highly valuable skill.
# ## Tracebacks
#
# `Python` tries to be helpful when errors occur, and provides a *traceback*, as with the (non-working) example code below.
# +
# This code has an intentional error.
def favourite_ice_cream():
ice_creams = [
"chocolate",
"vanilla",
"strawberry"
]
print(ice_creams[3])
favourite_ice_cream()
# -
# The traceback takes you through every step leading up to the error, and the `<ipython-input-1…>` lines show where each step starts.
#
# The first step has an arrow showing that the error occurred when we called the function `favourite_ice_cream()`.
#
# The second step tells us that, when calling `favourite_ice_cream()` (as shown the `in favourite_ice_cream()` part of the header), the line with the problem was `print(ice_creams[3])`.
#
# Finally, the last line of the traceback tells us what the error was: `IndexError: list index out of range`. We have tried to use an index that is outside the length of the list.
#
# Some *traceback*s can be very long, if multiple functions were called in the lead-up to the error. Mostly, you can usually just look at the last couple of steps to work out what is wrong.
# ## Syntax Errors
#
# The error we just saw was a *logic* error. The code was written correctly, but it tried to do something that was not possible, or forbidden.
#
# *Syntax* errors occur when the code is not correctly written. For example, a pair of parentheses `()` may not be closed, a quotation mark (`"`) or colon (`:`) may be missing, or there might be a typo.
#
# `Python` provides *traceback* information for these errors, too.
def some_function()
msg = "hello, world!"
print(msg)
return msg
# `Python` tells us specifically that this is a `SyntaxError`, and points to the approximate location of the problem with a caret/hat (`^`).
#
# There is a colon missing at the end of the function declaration, so we can fix that.
def some_function():
msg = "hello, world!"
print(msg)
return msg
# Now `Python` informs us of an `IndentationError`. There were two problems!
#
# `Python` does not tell us about all the syntax errors at the same time (fixing one error may resolve the rest!), it gives up as soon as it doesn't understand the code.
# ## Other Errors
#
# * `NameErrors` occur when a variable name is not defined in scope
print(a)
for i in range(3):
count = count + i
# * `IndexError`s occur when you try to use an element that is not in a sequence
letters = ['a', 'b', 'c']
print("Letter #1 is", letters[0])
print("Letter #2 is", letters[1])
print("Letter #3 is", letters[2])
print("Letter #4 is", letters[3])
|
python/2017-05-18-standrews/python-02/errors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Comparison of different algorithms available in Scikit-Surprise Library. The ones used here are Singular Value Decomposition(SVD), Stochastic Gradient Descent(SGD) and Alterating Least Squares(ALS). The dataset used is the movie ratings dataset, which can be found in grouplens.org. The one used here is of size 1M.
# +
# %matplotlib inline
import numpy
import pandas
from surprise import Dataset, Reader, accuracy
from surprise import SVD, BaselineOnly, KNNBasic
from surprise.model_selection import cross_validate, train_test_split
# -
ratings_data = pandas.read_table('ml-1m/ratings.dat', sep = '::', names = ['user_id', 'movie_id', 'rating', 'time'])
print(ratings_data.head())
movie_data = ratings_data[['user_id', 'movie_id', 'rating']]
reader = Reader(line_format='user item rating', sep = ',')
movie_data = Dataset.load_from_df(movie_data, reader=reader)
training_set, testing_set = train_test_split(movie_data, test_size=0.2)
# Using SGD
baseline_options = {'method' : 'sgd', 'learning_rate': 0.001}
algorithm = BaselineOnly(bsl_options=baseline_options)
cross_validate(algorithm, movie_data, measures=['RMSE', 'MAE'], cv=4, verbose=True)
algorithm.fit(training_set)
predictions = algorithm.test(testing_set)
accuracy.rmse(predictions)
# Using ALS
baseline_options = {'method' : 'als', 'learning_rate': 16}
similarity_options = {'name': 'pearson_baseline'}
algorithm = KNNBasic(bsl_options=baseline_options, sim_options=similarity_options)
cross_validate(algorithm, movie_data, measures=['RMSE', 'MAE'], cv=4, verbose=True)
algorithm.fit(training_set)
predictions = algorithm.test(testing_set)
accuracy.rmse(predictions)
# Using SVD
algorithm = SVD()
cross_validate(algorithm, movie_data, measures=['RMSE', 'MAE'], cv=4, verbose=True)
algorithm.fit(training_set)
predictions = algorithm.test(testing_set)
accuracy.rmse(predictions)
|
Movie Recommendation System - 1M.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Importing the required modules/packages
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import re
import nltk
import string
import scipy as sp
import datetime
import pytz
import graphviz
import copy
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import KFold, cross_val_score
from sklearn.metrics import precision_recall_fscore_support
from sklearn.metrics import precision_recall_fscore_support as score
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import GradientBoostingClassifier
from sklearn import metrics
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn import tree
from sklearn.svm.libsvm import cross_validation
from sklearn.model_selection import cross_validate
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import HashingVectorizer
from sklearn import linear_model, decomposition
from sklearn.decomposition import PCA
from sklearn.decomposition import TruncatedSVD
from sklearn.random_projection import sparse_random_matrix
from textblob import TextBlob, Word
from nltk.stem.snowball import SnowballStemmer
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from nltk import word_tokenize
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from random import randint
## Elastic Search for Metrics
from datetime import datetime
from elasticsearch import Elasticsearch
# Naive Bayes
from sklearn.naive_bayes import MultinomialNB
# Logistic Regression
from sklearn.linear_model import LogisticRegression
# SVC
from sklearn.svm import SVC
# KNN Neighbors
from sklearn.neighbors import KNeighborsClassifier
# Decision tree
from sklearn.tree import DecisionTreeClassifier
# Random forest
from sklearn.ensemble import RandomForestClassifier
# Gradient Booster Classifier
from sklearn.ensemble import GradientBoostingClassifier
# -
# ### Loading file and looking into the dimensions of data
raw_data = pd.read_csv("SMSSpamCollection.tsv",sep='\t',names=['label','text'])
pd.set_option('display.max_colwidth',100)
raw_data.head()
print(raw_data.shape)
pd.crosstab(raw_data['label'],columns = 'label',normalize=True)
# +
vect = CountVectorizer()
X_train_dtm = vect.fit_transform(X_train)
X_test_dtm = vect.transform(X_test)
print('X_train Shape', X_train_dtm.shape)
# Last 50 features
print((vect.get_feature_names()[-50:]))
# +
## Looks like we have 7234 Vectors after Count Vectorizer. From 3900 lines of information.
# +
## Vocabulary used:
# vect.vocabulary_
print(X_test_dtm)
# +
# Create Test Train Fit
# Define X and y.
X = raw_data.text
y = raw_data.label
# Split the new DataFrame into training and testing sets.
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=99, test_size= 0.3)
# -
# # Calculate Null Accuracy
# Calculate null accuracy.
y_test_binary = np.where(y_test=='ham', 1, 0) # five stars become 1, one stars become 0
print('Percent Ham:', y_test_binary.mean())
print('Percent Spam:', 1 - y_test_binary.mean())
class LemmaTokenizer(object):
def __init__(self):
self.wnl = WordNetLemmatizer()
def __call__(self, articles):
return [self.wnl.lemmatize(t) for t in word_tokenize(articles)]
# # Function to cleanup the data through pre-processing
# # Calculate Metrics and Generate
def calculate_metrics_push_to_es(run_id_insert, algorithm_name_insert, test_parameters_insert, gs_best_parameters_pipe_spam_ham, score,test_scores_csv_means_std, y_test,y_pred):
macro_score = precision_recall_fscore_support(y_test, y_pred, average='macro')
micro_score = precision_recall_fscore_support(y_test, y_pred, average='micro')
weighted_score = precision_recall_fscore_support(y_test, y_pred, average='weighted')
macro_score_insert = {'macro_precision': macro_score[0] * 100, 'macro_recall': macro_score[1] * 100, 'macro_fscore':macro_score[2] * 100}
micro_score_insert = {'micro_precision': micro_score[0] * 100, 'micro_recall': micro_score[1] * 100, 'micro_fscore':micro_score[2] * 100}
weighted_score_insert = {'weighted_precision': weighted_score[0] * 100, 'weighted_recall': weighted_score[1] * 100, 'weighted_fscore':weighted_score[2] * 100}
score_insert = {'score': score}
print(score_insert)
## Print Accuracy of the current Test
print(algorithm_name_insert , ' pipeline test accuracy: %.3f' % score)
## Push the data to ElasticSearch
ES_Metric_Insert(run_id_insert, algorithm_name_insert, test_parameters_insert,gs_best_parameters_pipe_spam_ham, score_insert,test_scores_csv_means_std, macro_score_insert,micro_score_insert,weighted_score_insert)
return()
# # Pushing Data into Elastic Search
def ES_Metric_Insert(run_id_insert,algorithm_name, test_parameters, gs_best_parameters_pipe_spam_ham, score, test_scores_csv_means_std, macro_scores, micro_scores, weighted_scores):
es = Elasticsearch()
final_dict = {}
my_current_time = datetime.now(tz=pytz.utc)
timestamp_insert = {'timestamp': my_current_time}
author_insert = {'author': 'Rahul'}
final_dict.update(run_id_insert)
final_dict.update(timestamp_insert)
final_dict.update(author_insert)
final_dict.update(algorithm_name)
final_dict.update(test_parameters)
final_dict.update(gs_best_parameters_pipe_spam_ham)
final_dict.update(score)
final_dict.update(test_scores_csv_means_std)
final_dict.update(macro_scores)
final_dict.update(micro_scores)
final_dict.update(weighted_scores)
res = es.index(index="ml-performance-metrics", doc_type='text', body=final_dict)
es.indices.refresh(index="ml-performance-metrics")
return()
# # Processing the ML Pipeline and Calculate Metrics (using another function)
def ML_Pipeline_Processing_And_Metrics(run_id,X_train, y_train, X_test, y_test, grid_search_parameters, gs_clf_pipe_spam_ham, cv_value, classifier_name):
gs_clf_pipe_spam_ham.fit(X_train, y_train)
## Find predictions for the pipeline
y_pred = gs_clf_pipe_spam_ham.predict(X_test)
## Find score of predictions
score_pipe_spam_ham = gs_clf_pipe_spam_ham.score(X_test, y_test) * 100
## Best Grid Search Parameters selected for this case
gs_best_parameters_pipe_spam_ham = {}
for param_name in sorted(grid_search_parameters.keys()):
if param_name == 'vect__tokenizer':
gs_best_parameters_pipe_spam_ham[param_name] = 'LemmaTokenizer'
else:
gs_best_parameters_pipe_spam_ham[param_name] = gs_clf_pipe_spam_ham.best_params_[param_name]
## Setting up for reporting to Screen and ElasticSearch
## Add Run Id for each run. This helps with fishing out the correct dataset in cloud
run_id_insert = {'run_id' : run_id}
## Save Classifier name as a string
classifier_string = str(classifier_name)
classifer_name_only = classifier_string.split("(")[0]
algorithm_name_insert = {'Algorithm_Name' : classifer_name_only}
## Add Classifier Parameters to output
test_parameters_insert = {'test_parameters' : str(pipe_spam_ham)}
## Breaking test cv scores and calculating mean and standard Deviation of each.
cv_scores_df = pd.DataFrame.from_dict(gs_clf_pipe_spam_ham.cv_results_)
test_scores_csv_means_std = {}
test_scores_csv_means_std['mean_fit_time'] = cv_scores_df.loc[0 ,'mean_fit_time']
test_scores_csv_means_std['std_fit_time'] = cv_scores_df.loc[0 ,'std_fit_time']
test_scores_csv_means_std['mean_test_score'] = cv_scores_df.loc[0 ,'mean_test_score'] * 100
test_scores_csv_means_std['std_test_score'] = cv_scores_df.loc[0 ,'std_test_score']
test_scores_csv_means_std['mean_train_score'] = cv_scores_df.loc[0 ,'mean_train_score'] * 100
test_scores_csv_means_std['std_train_score'] = cv_scores_df.loc[0 ,'std_train_score']
## Send all the collected data to the metric collection and ES insert system.
calculate_metrics_push_to_es(run_id_insert, algorithm_name_insert, test_parameters_insert, gs_best_parameters_pipe_spam_ham, score_pipe_spam_ham, test_scores_csv_means_std, y_test,y_pred)
return()
# # Remove Vectorizers and ML Algorithms
# +
def remove_vectorizer_ml_algo(vector_ml_keyword):
## Remove from gridsearch
for key in grid_search_parameters.copy():
if vector_ml_keyword in key.lower():
del grid_search_parameters[key]
## Remove from spam ham pipeline
for item in pipe_spam_ham_features:
if vector_ml_keyword in item:
pipe_spam_ham_features.remove(item)
return()
# -
# # Add count vectorizer
# +
## Add Count Vectorizer and associated Features for Testing
def add_count_vectorizer(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['vect__binary'] = (False,True)
grid_search_parameters['vect__lowercase'] = (True,False)
grid_search_parameters['vect__tokenizer'] = (LemmaTokenizer(),None)
## Grid Search Parameters avialable for testing. After initial tests it looks like the above params work best. So using those.
# grid_search_parameters['vect__stop_words'] = ('english',None)
# grid_search_parameters['vect__ngram_range'] = [(1, 1),(1, 2),(1, 3), (1, 4)]
# grid_search_parameters['vect__max_df'] = (0.9,1)
# grid_search_parameters['vect__lowercase'] = (True, False)
# grid_search_parameters['vect__binary'] = (True, False)
# grid_search_parameters['vect__tokenizer'] = (LemmaTokenizer())
# grid_search_parameters['vect__min_df'] = (5,10)
pipe_spam_ham_features.append(('vect', CountVectorizer()))
return()
# -
# # Add Tf-Idf Vectorizer
# +
## Add Tf-Idf Vectorizer and associated Features for Testing
def add_tfidf_vectorizer(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['tfidf__norm'] = ('l2','l1')
grid_search_parameters['tfidf__smooth_idf'] = (True,False)
# ## Grid Search Parameters avialable for testing. After initial tests it looks like the above params work best. So using those.
# grid_search_parameters['tfidf__use_idf'] = (True, False)
# grid_search_parameters['tfidf__norm'] = ('l1','l2','max')
# grid_search_parameters['tfidf__smooth_idf'] = (True, False)
# grid_search_parameters['tfidf__sublinear_tf'] = (True, False)
pipe_spam_ham_features.append(('tfidf', TfidfVectorizer()))
return()
# -
# # TruncatedSVD
## Add Tf-Idf Vectorizer and associated Features for Testing
def add_TruncatedSVD(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['truncatedsvd__n_components'] = (500, 400, 200)
pipe_spam_ham_features.append(('truncatedsvd', TruncatedSVD()))
return()
# # Add Naive Bayes
## Add Naive Bayes Algorithm
def add_multinomialNB(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['nb__alpha'] = (1,0.9)
grid_search_parameters['nb__fit_prior'] = (True,False)
# ## Grid Search Parameters avialable for testing. After initial tests it looks like the above params work best. So using those.
# grid_search_parameters['nb__alpha'] = (0,1)
# grid_search_parameters['nb__fit_prior'] = (True, False)
pipe_spam_ham_features.append(('nb', MultinomialNB()))
return()
# # Add KNN
## Add Naive Bayes Algorithm
def add_knn(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['knn__n_neighbors'] = (1,2,3,4,5,6,7,8,9,10)
grid_search_parameters['knn__weights'] = ('uniform', 'distance')
#grid_search_parameters['knn__algorithm'] = ('ball_tree', 'kd_tree')
pipe_spam_ham_features.append(('knn', KNeighborsClassifier()))
return()
# # RandomForestClassifier
## Add Random Forest Algorithm
def add_randomforest(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['rf__n_estimators'] = (1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20)
grid_search_parameters['rf__max_depth'] = (10,100,1000,None)
pipe_spam_ham_features.append(('rf', RandomForestClassifier()))
return()
# # LogisticRegression
## Add Logistic Regression Algorithm
def add_logistic_regression(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['lr__penalty'] = ('l1','l2')
pipe_spam_ham_features.append(('lr', LogisticRegression()))
return()
# # SVC
## Add SVC Algorithm
def add_svc_regression(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['svc__C'] = (1.0,0.9,0.8)
pipe_spam_ham_features.append(('svc', SVC()))
return()
# # GradientBoostingClassifier
## Add GradientBoostingClassifier Algorithm
def add_gradient_boosting_classifer(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['gbc__n_estimators'] = (100,200,300,1000)
pipe_spam_ham_features.append(('gbc', GradientBoostingClassifier()))
return()
# # DecisionTreeClassifier
## Add DecisionTreeClassifier Algorithm
def add_decisiontree_classifer(pipe_spam_ham_features,grid_search_parameters):
grid_search_parameters['dtc__max_depth'] = (10,100,1000,None)
pipe_spam_ham_features.append(('dtc', DecisionTreeClassifier()))
return()
# # Full ML Pipeline
# +
pipe_spam_ham = []
pipe_spam_ham_features = []
grid_search_parameters = {}
list_ml_algo = {}
run_id = randint(100000, 999999)
## Cross_Val value
cv_value = 2
# Define 10 fold cross-validation
cv = KFold(n_splits=10)
# ## Addition of Count Vectorizer
#add_count_vectorizer(pipe_spam_ham_features,grid_search_parameters)
## Not using these, since the values score isn't much better than with Count Vectorizer.
#add_TruncatedSVD(pipe_spam_ham_features,grid_search_parameters)
#add_tfidf_vectorizer(pipe_spam_ham_features,grid_search_parameters)
## Create a dictionary of all available ML Algos
list_ml_algo['knn'] = 'knn'
list_ml_algo['rf'] = 'randomforest'
list_ml_algo['lr'] = 'logistic_regression'
list_ml_algo['nb'] = 'multinomialNB'
list_ml_algo['svc'] = 'svc_regression'
list_ml_algo['gbc'] = 'gradient_boosting_classifer'
list_ml_algo['dtc'] = 'decisiontree_classifer'
## Kick off the pipeline Execution:
## Iteration 1:
## No Vectorizer
count = 1
while count < 3:
if count == 1:
add_count_vectorizer(pipe_spam_ham_features,grid_search_parameters)
if count == 2:
add_tfidf_vectorizer(pipe_spam_ham_features,grid_search_parameters)
for key, values in list_ml_algo.items():
ml_algo_name = 'add_' + values
returnValueIfAny = globals()[ml_algo_name](pipe_spam_ham_features,grid_search_parameters)
## Setting up the pipeline
pipe_spam_ham = Pipeline(pipe_spam_ham_features)
classifier = str(pipe_spam_ham_features[-1:][0][1])
print(pipe_spam_ham)
print(grid_search_parameters)
## Adding the GridSearch CV
gs_clf_pipe_spam_ham = GridSearchCV(pipe_spam_ham, grid_search_parameters, n_jobs=1, cv = cv_value, return_train_score=True)
ML_Pipeline_Processing_And_Metrics(run_id,X_train, y_train, X_test, y_test,grid_search_parameters, gs_clf_pipe_spam_ham, cv_value, classifier)
remove_vectorizer_ml_algo(key)
# remove_vectorizer_ml_algo('truncatedsvd')
remove_vectorizer_ml_algo('vect')
remove_vectorizer_ml_algo('tfidf')
count += 1
## End of Program ..
# +
# imports needed and logging
import gzip
import gensim
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# build vocabulary and train model
model = gensim.models.Word2Vec(
raw_data,
size=150,
window=10,
min_count=2,
workers=10)
model.train(raw_data, total_examples=len(raw_data), epochs=10)
# -
|
Spam dectection v3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Чем думает рыба?
# !wget https://github.com/goto-ru/Unsupervised_ML/raw/20779daf2aebca80bfe38401bc87cf41fc7b493d/03_zebrafish/zebrafish.npy -O zebrafish.npy
#alternative link: https://www.dropbox.com/s/hhep0wj4c11qibu/zebrafish.npy?dl=1
# # Данные
#
# * Сейчас в вашем распоряжении - данные о мозговой активности малька рыбы вида Danio Rerio https://en.wikipedia.org/wiki/Zebrafish .
# * Мальку введено вещество, которое светится от электрической активности (например, от спайков нейронов). Мальки почти прозрачны, поэтому такое свечение видно извне.
# * Сами данные содержат 240 фотографий головной части рыбки, на которых видна мозговая активность в каждой точке. Каждая фотография имеет размер __230 x 202__ пикселей
# * Ваша задача - попытаться восстановить структуру мозга рыбки. Для этого можно попытаться найти, например, группы нейронов, реагирующих вместе или с одинаковой частотой.
# * Никакой разметки в данных нет, поэтому вам придётся использовать методы понижения размерности и кластеризации, чтобы эффективно анализировать данные.
#
# 
import numpy as np
data = np.load("zebrafish.npy")/255.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
tick0 = data[:,0]
tick0_image = tick0.reshape(230, 202)
print "размер 1 картинки:", tick0_image.shape
plt.imshow(tick0_image.T);
# +
#мини-библиотека для рисования рыбы
from zebrafish_drawing_factory import draw_component
draw_component(data[:,0])
# -
# # Временные ряды
#
# * Посмотрим на активность отдельных пикселей в течение времени:
# * Попробуйте вручную найти какие-то характерные группы нейронов
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=[10,10])
for i in range(0,240,10):
plt.plot(data[i])
# # Поищем характерные группы нейронов
#
# Давайте разложим временные ряды активности нейронов при помощи метода главных компонент.
#
# __Важно!__ в этой части задания объектом выборки является временной ряд активности 1 точки на картинке, а не картинка целиком.
# +
from sklearn.decomposition import PCA
pca = <создайте и обучите PCA с 20+ компонентами>
# -
data_pca = <преобразуйте данные в пространство главных компонент pca.transform>
# ## Визуализируем компоненты
draw_component(data_pca[:,1])
draw_component(data_pca[:,2])
# +
from zebrafish_drawing_factory import draw_components
draw_components(data_pca[:,2],data_pca[:,3])
# -
# # Поищем фичи
def extract_features(impulses):
"""given time series(array) of region activity, compute some feature representation of those time series
Ideas:
- fourier transform
- mean, variance and percentiles
- sums of every k-th element with shift b
"""
features = []<любые фичи>
return features
# +
data_features = np.array(list(map(extract_features, data)))
print "shape:",data_features.shape
# +
from sklearn.decomposition import PCA
pca = <обучи PCA>
# -
data_pca = <преобразуй в пространство PCA>
<визуализируй полученные компоненты>
draw_component(...)
draw_components(...)
# # Bonus: clustering in PCA space
# +
from sklearn.cluster import KMeans
from sklearn.mixture import GMM
<покластеризуй области изображения на основе двух полученных PCA-представлений, используй любой метод на выбор>
# -
cluster_ids = <предскажи номер кластера для каждого пикселя>
#cluster_ids должен содержать по 1 чиселке на пиксель
assert np.prod(cluster_ids.shape) == (230*202)
plt.imshow(cluster_ids.reshape(230,202),cmap='spectral')
|
Seminar1/Classwork_ru.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Install Nipype
#
# The best and most complete instruction on how to download and install Nipype can be found on the [official homepage](http://nipype.readthedocs.io/en/latest/users/install.html). Nonetheless, here's a short summary of some (but not all) approaches.
# ## 1. Install Nipype
#
# Getting Nipype to run on your system is rather straight forward. And there are multiple ways to do the installation:
#
#
# ### Using conda
#
# If you have [conda](http://conda.pydata.org/docs/index.html), [miniconda](https://conda.io/miniconda.html) or [anaconda](https://www.continuum.io/why-anaconda) on your system, than installing Nipype is just the following command:
#
# conda config --add channels conda-forge
# conda install nipype
#
#
# ### Using ``pip`` or ``easy_install``
#
# Installing Nipype via ``pip`` or ``easy_install`` is as simple as you would imagine.
#
# pip install nipype
#
# or
#
# easy_install nipype
#
#
# ### Using Debian or Ubuntu
#
# Installing Nipype on a Debian or Ubuntu system can also be done via ``apt-get``. For this use the following command:
#
# apt-get install python-nipype
#
#
# ### Using Github
#
# To make sure that you really have the newest version of Nipype on your system, you can run the pip command with a flag that points to the github repo:
#
# pip install git+https://github.com/nipy/nipype#egg=nipype
# ## 2. Install Dependencies
#
# For more information about the installation in general and to get a list of recommended software, go to the main page, under: http://nipype.readthedocs.io/en/latest/users/install.html
#
# For a more step by step installation guide for additional software dependencies like SPM, FSL, FreeSurfer and ANTs, go to the [Beginner's Guide](http://miykael.github.io/nipype-beginner-s-guide/installation.html).
#
# ## 3. Test Nipype
# +
# Import the nipype module
import nipype
# Run the test
nipype.test(doctests=False)
# -
# The test will create a lot of output, but if all goes well you will see an OK at the end:
#
# ----------------------------------------------------------------------
# Ran 2497 tests in 68.486s
#
# OK (SKIP=13)
#
# The number of tests and time will vary depending on which interfaces you have installed on your system.
#
# Don’t worry if some modules are being skipped or some side modules show up as errors or failures during the run. As long as no main modules cause any problems, you’re fine. The number of tests and time will vary depending on which interfaces you have installed on your system. But if you receive an OK, errors=0 and failures=0 then everything is ready.
|
notebooks/resources_installation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Crimes rates per Neighborhood</h2>
# <p>Using data from the WRPDC, we can gauge overall criminal activity in each Pittsburgh neighborhood. This data could potentially give us an idea of which neighborhood has the most violent and experienced residents, and can help estimate the possibility of 2 emerging tributes becoming victors.</p>
#
# <p><code>WRPDC - "Police Incident Blotter"</code>
# <br>from January 2016 - present</p>
# <a href='#'>https://data.wprdc.org/dataset/uniform-crime-reporting-data</a>
# Importing packages for data analysis
import pandas as pd
import geopandas
import numpy as np
from pprint import pprint
# %matplotlib inline
import matplotlib.pyplot as plt
# +
data = pd.read_csv("./data/crime-data.csv")
# Indexes for column names
count = 0
for i in data.columns:
print(f"column {count}: {i}")
count+=1
# +
cols = data.INCIDENTNEIGHBORHOOD.value_counts()
# Comment out to see full data
# pd.set_option("display.max_rows", None, "display.max_columns", None)
print(cols)
# -
# <h3>Cleaning our Data</h3>
# <p>Now that we have a good idea of what our data looks like, we are going to have to clean it to get rid of irrelevant entries. For instance, we can get rid of 'Outside State', 'Outside County', 'Mt. Oliver Boro', etc. as they are not really neighborhoods.</p>
# +
filters = [
'Outside State',
'Outside County',
'Outside City',
'Mt. Oliver Boro',
'Mt. Oliver Neighborhood',
'Golden Triangle/Civic Arena'
]
pattern = '|'.join(filters)
query_mask = data['INCIDENTNEIGHBORHOOD'].str.contains(pattern, na=False, case=False)==0
data = data[query_mask]
cols = data.INCIDENTNEIGHBORHOOD.value_counts() # update cols
print(cols)
# -
# Converting our data in pandas DataFrame so we can
# merge it with geopandas and map out results
crime_by_hood = pd.DataFrame({'Neighborhood':cols.index, 'num_incidents':cols.values})
crime_by_hood.head()
# <h3>Mapping our results</h3>
# <p>Using geopandas, we can map out our results in a heat map, to get a better idea of which areas of the city are "hot" and dangerous.
# Checking correct map of Pittsburgh
neighborhoods = geopandas.read_file("./data/Neighborhoods/Neighborhoods_.shp")
crime_map = neighborhoods.merge(crime_by_hood, how='left', left_on="hood", right_on='Neighborhood')
# Mapping crime rates to Pittsburgh with custom parameters
plt.style.use("default")
crime_map.plot(column='num_incidents', # set the data to be used for coloring
cmap='brg', # choose a color palette
edgecolor="white", # outline the districts in white
legend=True, # show the legend
legend_kwds={'label': "Crime Rate"}, # label the legend
figsize=(15, 10), # set the size
missing_kwds={"color": "lightgrey"} # set disctricts with no data to gray
)
# <h3>Map Results</h3>
# <p>As we can see from our data, South Side Flats and the Central Business District have far more reported crime-related incidents than any other neighborhoods. But that doesn't paint the whole picture so we should now see if we can relate the number of reported incidents with the severity of the crime committed. This will help us gauge the ruthlessness of potential tributes. For this we can take a look at the <code>OFFENSES</code> column in our dataset.
crimes = data.OFFENSES.value_counts()
crime_severity = pd.DataFrame({"Offense":crimes.index, "num_incidents":crimes.values})
crime_severity.head(10)
# <h3>Filtering Further</h3>
# <p>As we can see from the most common offenses committed, these do not really help us determine the propensity to violence needed to survive <strong>The Hoagie Games</strong>. We will need to filter our data further to find the most terrible offenses committed in our great city and correlate the results to the neighborhood most likely to commit them.
# +
# look through data to find certain codes for offenses
codes = [
'2715', # Weapons of Mass Destruction
'2706', # Terroristic threats
'4953', # Mob related activity
'8106', # Shots fired
'2702', # Aggravated assualt
'2501', # Criminal homicide
'2707', # Propulsion of missiles
]
pat = '|'.join(codes)
query_mask = data['OFFENSES'].str.contains(pat, na=False, case=False)
data = data[query_mask]
data['INCIDENTNEIGHBORHOOD'].value_counts()
# -
# <h3>Analyzing our data</h3>
# <p>By filtering out the worst crimes committed by neighborhood we come up with some interesting results. For instance, we see that Sheraden now has more violent crime than Central Business District, even though Sheraden wasn't even in our top 5 neighborhoods for crimes reported. By dividing these results by the total amount of crimes reported per neighborhood, we can standardize each neighborhoods propensity to violent actions.
# <p>We should also determine some sort of scoring system so as to give more weight to higher severity offenses. To keep things simple we can score each of our chosen offenses from 1-7 points with 7 being most severe. This is summarized below: </p>
# <ol>
# <li>4953 - Mob related activity</li>
# <li>8106 - Shots fired</li>
# <li>2702 - Aggravated assualt</li>
# <li>2706 - Terroristic threats</li>
# <li>2707 - Propulsion of missiles</li>
# <li>2501 - Criminal homicide</li>
# <li>2715 - Weapons of Mass Destruction</li>
# </ol>
# +
order = ['4953', '8106', '2702', '2706', '2707', '2501', '2715']
desc = [
'Mob related activity',
'Shots fired',
'Aggravated assualt',
'Terroristic threats',
'Propulsion of missiles',
'Criminal homicide',
'Weapons of Mass Destruction'
]
# Creating a pandas DataFrame for readability and
# for using with matplotlib later
results = pd.DataFrame({"offense": order,
"desc": desc,
"count": 0,
"weighted_sum":0,
})
#results2 = pd.DataFrame(index=order, columns=[hoods], data=data['INCIDENTNEIGHBORHOOD'].value_counts())
# Loop through the offenses and count each occurence
# and find its weighted sum
for i in data.OFFENSES:
for j in range(7):
if order[j] in i:
results.loc[j, 'count']+=1
results.loc[j, 'weighted_sum']+=1*(j+1)
break
results
# -
# <p>Here we check the total number of criminal offenses committed and how they weigh up against each other. From here we should break down the reported crimes by neighborhood and get a final result</p>
# +
def get_neighborhood_index(counter):
area = data.iloc[counter][6]
for i in range(len(hoods_as_cols)):
if area == hoods_as_cols[i]:
return hoods_as_cols[i]
hoods_as_cols = data.INCIDENTNEIGHBORHOOD.unique()
hoods_as_cols = pd.Series(hoods_as_cols)
combined_data = pd.DataFrame(data=0, index=order, columns=hoods_as_cols)
combined_data.insert(0, "desc", desc)
combined_data['total'] = 0
combined_data['weighted_total'] = 0
counter = 0
for i in data.OFFENSES:
for j in range(7):
if order[j] in i:
col = get_neighborhood_index(counter)
if col is not None:
combined_data.loc[order[j], col]+=1
combined_data.loc[order[j], "total"]+=1
counter+=1
combined_data
# -
# <h3>Finalizing our results</h3>
# <p>We now have a good idea of the amount of severe crimes committed and how they weigh up against each other. We can display the results in graphs to help get a better visual idea of the data distribution</p>
population_data = pd.read_csv("./ivan/results.csv")
population_data = population_data.sort_values(by=["Crimes per Population"], ascending=False)
population_data.head(15)
# <h3>Visualizing our Results</h3>
# <p>And finally let's graph our results so we can get a better idea of the data distribution...</p>
graph = data["INCIDENTNEIGHBORHOOD"].value_counts()
graph.plot(kind="bar", figsize=(20, 20))
population_data.sort_values(by="Crimes per Population")
population_data[["Neighborhood", "Crimes per Population"]].plot(kind="bar", figsize=(20, 20))
# <h3>Result</h3>
# <p>So after all that, it seems that <a href="#">Swisshelm Park</a> commits more crimes per population than any other Pittsburgh neighborhood. However I believe that since <a href="#">South Side Flats</a> reported a much more numerous amount of crimes, that its residents are much more prepared for a Hoagie Games scenario and therefore I will go against the data and declare <a href="#">South Side Flats</a> as our competitive choice.</p>
# <h2>Population Census Data</h2>
# <p>Using data from the WRPDC, we can gauge overall population and athletic activity in each Pittsburgh neighborhood. This data could potentially give us an idea of which neighborhood has the most most active residents, and can help estimate the possibility of an emerging tribute becoming a victor.</p>
#
# <p><code>WRPDC - "SNAP Census Data, Employment 2010"</code>
# <br>from 2010</p>
# <a href='#'>https://data.wprdc.org/dataset/pgh/resource/fd095080-d32c-4669-8b62-c80f4f32723a</a>
# <h3>Taking a look at Population Data...</h3>
import pandas as pd
df = pd.read_csv("./Sid/data.csv")
df
# ## Let's get a sense of what our dataframe looks like
df.columns
df["Total Working Population Percentage"] = df["Total Working Pop. (Age 16+) (2010)"] / (df["Population (2010)"] + df["Total Working Pop. (Age 16+) (2010)"])
df
# In the above cell, we obtain the ratio of the working population to the total population in each area. Let's see who has the highest....
df.sort_values(by="Total Working Population Percentage", ascending=False, inplace=True)
df
# Let's evaluate each area in Pittsburgh based on how how "athletic" their working population is.
#
# Let's award 10 points to those who commute to work via Taxi/Carpool/Vanpool/Other, 20 points to those who commute to work via Motorcycle, 40 points via Bicycle, and 60 points via walking.
#
# Let's see the most "athletic" regions in Pittsburgh....
# +
df["Athletic Score"] = 10 * df["Work at Home (2010)"]
+ 10 * df["Commute to Work: Other (2010)"]
+ 10 * df["Commute to Work: Taxi (2010)"]
+ 10 * df["Commute to Work: Carpool/Vanpool (2010)"]
+ 10 * df["Commute to Work: Public Transportation (2010)"]
+ 20 * df["Commute to Work: Motorcycle (2010)"]
+ 40 * df["Commute to Work: Bicycle (2010)"]
+ 60 * df["Commute to Work: Walk (2010)"]
df.sort_values(by="Athletic Score", inplace=True, ascending=False)
# df = df[["Neighborhood", "Athletic Score"]]
df.dropna(inplace=True)
df[["Neighborhood", "Athletic Score"]]
# -
# Here's a quick summary of the "Athletic Score" statistic throughout Pittsburgh ...
df["Athletic Score"].plot.box()
# It looks like there's a clear winner ...
import geopandas
# %matplotlib inline
import matplotlib.pyplot as plt
df[["Neighborhood", "Athletic Score"]]
# Fairywood blows everyone else out of the waters.... It isn't particularly close either.
# +
# # import dataset
# steps = pd.read_csv("steps.csv")
# # filter to important info
# num_steps = steps.groupby("neighborhood").sum()['number_of_steps']
# num_steps.sort_values(ascending=False)
# +
# # do the merge
# steps_map = neighborhoods.merge(num_steps, how='left', left_on='hood', right_on='neighborhood')
# # look at the head to confirm it merged correctly
# steps_map[['hood','number_of_steps','geometry']].head()
# -
neighborhoods = geopandas.read_file("./data/Neighborhoods/Neighborhoods_.shp")
result = neighborhoods.merge(df, how='left', left_on='hood', right_on='Neighborhood')
result[["Neighborhood", "Athletic Score"]]
result.plot(column='Athletic Score', # set the data to be used for coloring
cmap='OrRd', # choose a color palette
edgecolor="white", # outline the districts in white
legend=True, # show the legend
legend_kwds={'label': "Number of Steps"}, # label the legend
figsize=(15, 10), # set the size
missing_kwds={"color": "lightgrey"} # set disctricts with no data to gray
)
|
.ipynb_checkpoints/main-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 14 Linear Algebra
# ## Motivating problem: Two masses on three strings
# Two masses $M_1$ and $M_2$ are hung from a horizontal rod with length $L$ in such a way that a rope of length $L_1$ connects the left end of the rod to $M_1$, a rope of length $L_2$ connects $M_1$ and $M_2$, and a rope of length $L_3$ connects $M_2$ to the right end of the rod. The system is at rest (in equilibrium under gravity).
#
# 
#
# Find the angles that the ropes make with the rod and the tension forces in the ropes.
# In class we derived the equations that govern this problem – see [14_String_Problem_lecture_notes (PDF)](14_String_Problem_lecture_notes.pdf).
#
# We can represent the problem as system of nine coupled non-linear equations:
#
# $$
# \mathbf{f}(\mathbf{x}) = 0
# $$
#
# ### Summary of equations to be solved
# Treat $\sin\theta_i$ and $\cos\theta_i$ together with $T_i$, $1\leq i \leq 3$, as unknowns that have to simultaneously fulfill the nine equations
# \begin{align}
# -T_1 \cos\theta_1 + T_2\cos\theta_2 &= 0\\
# T_1 \sin\theta_1 - T_2\sin\theta_2 - W_1 &= 0\\
# -T_2\cos\theta_2 + T_3\cos\theta_3 &= 0\\
# T_2\sin\theta_2 + T_3\sin\theta_3 - W_2 &= 0\\
# L_1\cos\theta_1 + L_2\cos\theta_2 + L_3\cos\theta_3 - L &= 0\\
# -L_1\sin\theta_1 - L_2\sin\theta_2 + L_3\sin\theta_3 &= 0\\
# \sin^2\theta_1 + \cos^2\theta_1 - 1 &= 0\\
# \sin^2\theta_2 + \cos^2\theta_2 - 1 &= 0\\
# \sin^2\theta_3 + \cos^2\theta_3 - 1 &= 0
# \end{align}
#
# Consider the nine equations a vector function $\mathbf{f}$ that takes a 9-vector $\mathbf{x}$ of the unknowns as argument:
# \begin{align}
# \mathbf{f}(\mathbf{x}) &= 0\\
# \mathbf{x} &= \left(\begin{array}{c}
# x_0 \\ x_1 \\ x_2 \\
# x_3 \\ x_4 \\ x_5 \\
# x_6 \\ x_7 \\ x_8
# \end{array}\right)
# =
# \left(\begin{array}{c}
# \sin\theta_1 \\ \sin\theta_2 \\ \sin\theta_3 \\
# \cos\theta_1 \\ \cos\theta_2 \\ \cos\theta_3 \\
# T_1 \\ T_2 \\ T_3
# \end{array}\right) \\
# \mathbf{L} &= \left(\begin{array}{c}
# L \\ L_1 \\ L_2 \\ L_3
# \end{array}\right), \quad
# \mathbf{W} = \left(\begin{array}{c}
# W_1 \\ W_2
# \end{array}\right)
# \end{align}
# In more detail:
#
# \begin{align}
# f_1(\mathbf{x}) &= -x_6 x_3 + x_7 x_4 &= 0\\
# f_2(\mathbf{x}) &= x_6 x_0 - x_7 x_1 - W_1 & = 0\\
# \dots\\
# f_8(\mathbf{x}) &= x_2^2 + x_5^2 - 1 &=0
# \end{align}
#
#
# We generalize the *Newton-Raphson algorithm* from the [last lecture](http://asu-compmethodsphysics-phy494.github.io/ASU-PHY494//2018/03/21/12_Root_finding/) to $n$ dimensions:
# ## General Newton-Raphson algorithm
# Given a trial vector $\mathbf{x}$, the correction $\Delta\mathbf{x}$ can be derived from the Taylor expansion
# $$
# f_i(\mathbf{x} + \Delta\mathbf{x}) = f_i(\mathbf{x}) + \sum_{j=1}^{n} \left.\frac{\partial f_i}{\partial x_j}\right|_{\mathbf{x}} \, \Delta x_j + \dots
# $$
# or in full vector notation
# \begin{align}
# \mathbf{f}(\mathbf{x} + \Delta\mathbf{x}) &= \mathbf{f}(\mathbf{x}) + \left.\frac{d\mathbf{f}}{d\mathbf{x}}\right|_{\mathbf{x}} \Delta\mathbf{x} + \dots\\
# &= \mathbf{f}(\mathbf{x}) + \mathsf{J}(\mathbf{x}) \Delta\mathbf{x} + \dots
# \end{align}
# where $\mathsf{J}(\mathbf{x})$ is the *[Jacobian](http://mathworld.wolfram.com/Jacobian.html)* matrix of $\mathbf{f}$ at $\mathbf{x}$, the generalization of the derivative to multivariate vector functions.
#
# Solve
# $$
# \mathbf{f}(\mathbf{x} + \Delta\mathbf{x}) = 0
# $$
# i.e.,
# $$
# \mathsf{J}(\mathbf{x}) \Delta\mathbf{x} = -\mathbf{f}(\mathbf{x})
# $$
# for the correction $\Delta x$
# $$
# \Delta\mathbf{x} = -\mathsf{J}(\mathbf{x})^{-1} \mathbf{f}(\mathbf{x})
# $$
# which has the same form as the 1D Newton-Raphson correction $\Delta x = -f'(x)^{-1} f(x)$.
#
# These are *matrix equations* (we linearized the problem). One can either explicitly solve for the unknown vector $\Delta\mathbf{x}$ with the inverse matrix of the Jacobian or use other methods to solve the coupled system of linear equations of the general form
# $$
# \mathsf{A} \mathbf{x} = \mathbf{b}.
# $$
#
#
# ## Linear algebra with `numpy.linalg`
import numpy as np
# +
# np.linalg?
# -
# ### System of coupled linear equations
# Solve the coupled system of linear equations of the general form
# $$
# \mathsf{A} \mathbf{x} = \mathbf{b}.
# $$
A = np.array([
[1, 0, 0],
[0, 1, 0],
[0, 0, 2]
])
b = np.array([1, 0, 1])
# What does this system of equations look like?
for i in range(A.shape[0]):
terms = []
for j in range(A.shape[1]):
terms.append("{1} x[{0}]".format(j, A[i, j]))
print(" + ".join(terms), "=", b[i])
# Now solve it with `numpy.linalg.solve`:
x = np.linalg.solve(A, b)
print(x)
# Test that it satisfies the original equation:
# $$
# \mathsf{A} \mathbf{x} - \mathbf{b} = 0
# $$
np.dot(A, x) - b
# #### Activity: Solving matrix equations
# With
# $$
# \mathsf{A}_1 = \left(\begin{array}{ccc}
# +4 & -2 & +1\\
# +3 & +6 & -4\\
# +2 & +1 & +8
# \end{array}\right)
# $$
# and
# $$
# \mathbf{b}_1 = \left(\begin{array}{c}
# +12 \\ -25 \\ +32
# \end{array}\right), \quad
# \mathbf{b}_2 = \left(\begin{array}{c}
# +4 \\ -1 \\ +36
# \end{array}\right), \quad
# $$
# solve for $\mathbf{x}_i$
# $$
# \mathsf{A}_1 \mathbf{x}_i = \mathbf{b}_i
# $$
# and *check the correctness of your answer*.
A1 = np.array([
[+4, -2, +1],
[+3, +6, -4],
[+2, +1, +8]
])
b1 = np.array([+12, -25, +32])
b2 = np.array([ 4, -1, 36])
x1 = np.linalg.solve(A1, b1)
print(x1)
print(A1.dot(x1) - b1)
x2 = np.linalg.solve(A1, b2)
print(x2)
print(A1.dot(x2) - b2)
# ### Matrix inverse
# In order to solve directly we need the inverse of $\mathsf{A}$:
# $$
# \mathsf{A}\mathsf{A}^{-1} = \mathsf{A}^{-1}\mathsf{A} = \mathsf{1}
# $$
# Then
# $$
# \mathbf{x} = \mathsf{A}^{-1} \mathbf{b}
# $$
# If the inverse exists, `numpy.linalg.inv()` can calculate it:
Ainv = np.linalg.inv(A)
print(Ainv)
# Check that it behaves like an inverse:
Ainv.dot(A)
A.dot(Ainv)
# Now solve the coupled equations directly:
Ainv.dot(b)
# #### Activity: Solving coupled equations with the inverse matrix
# 1. Compute the inverse of $\mathsf{A}_1$ and *check the correctness*.
# 2. Compute $\mathbf{x}_1$ and $\mathbf{x}_2$ with $\mathsf{A}_1^{-1}$ and check the correctness of your answers.
A1_inv = np.linalg.inv(A1)
print(A1_inv)
A1.dot(A1_inv)
A1_inv.dot(A1)
x1 = A1_inv.dot(b1)
print(x1)
print(A1.dot(x1) - b1)
x2 = A1_inv.dot(b2)
print(x2)
print(A1.dot(x2) - b2)
# ### Eigenvalue problems
# The equation
# \begin{gather}
# \mathsf{A} \mathbf{x}_i = \lambda_i \mathbf{x}_i
# \end{gather}
# is the **eigenvalue problem** and a solution provides the eigenvalues $\lambda_i$ and corresponding eigenvectors $x_i$ that satisfy the equation.
# #### Example 1: Principal axes of a square
# The principle axes of the [moment of inertia tensor](https://en.wikipedia.org/wiki/Moment_of_inertia#The_inertia_tensor) are defined through the eigenvalue problem
# $$
# \mathsf{I} \mathbf{\omega}_i = \lambda_i \mathbf{\omega}_i
# $$
# The principal axes are the $\mathbf{\omega}_i$.
Isquare = np.array([[2/3, -1/4], [-1/4, 2/3]])
lambdas, omegas = np.linalg.eig(Isquare)
lambdas
omegas
# Note that the eigenvectors are `omegas[:, i]`! You can transpose so that axis 0 is the eigenvector index:
omegas.T
# Test:
# $$
# (\mathsf{I} - \lambda_i \mathsf{1}) \mathbf{\omega}_i = 0
# $$
# (The identity matrix can be generated with `np.identity(2)`.)
(Isquare - lambdas[0]*np.identity(2)).dot(omegas[:, 0])
(Isquare - lambdas[0]*np.identity(2)).dot(omegas.T[0])
(Isquare - lambdas[1]*np.identity(2)).dot(omegas.T[1])
# #### Example 2: Spin in a magnetic field
# In quantum mechanics, a spin 1/2 particle is represented by a spinor $\chi$, a 2-component vector. The Hamiltonian operator for a stationary spin 1/2 particle in a homogenous magnetic field $B_y$ is
# $$
# \mathsf{H} = -\gamma \mathsf{S}_y B_y = -\gamma B_y \frac{\hbar}{2} \mathsf{\sigma_y}
# = \hbar \omega \left( \begin{array}{cc} 0 & -i \\ i & 0 \end{array}\right)
# $$
# Determine the *eigenvalues* and *eigenstates*
# $$
# \mathsf{H} \mathbf{\chi} = E \mathbf{\chi}
# $$
# of the spin 1/2 particle.
#
# (To make this a purely numerical problem, divide through by $\hbar\omega$, i.e. calculate $E/\hbar\omega$.)
sigma_y = np.array([[0, -1j], [1j, 0]])
E, chis = np.linalg.eig(sigma_y)
print(E)
print(chis.T)
# Normalize the eigenvectors:
# $$
# \hat\chi = \frac{1}{\sqrt{\chi^\dagger \cdot \chi}} \chi
# $$
chi1 = chis.T[0]
print(chi1)
norm = np.dot(chi1.conjugate(), chi1)
chi1_hat = chi1/np.sqrt(norm)
print(chi1_hat)
norm
# ... they were already normalized.
# #### Activity: eigenvalues
# Find the eigenvalues and eigenvectors of
# $$
# \mathsf{A}_2 = \left(\begin{array}{ccc}
# -2 & +2 & -3\\
# +2 & +1 & -6\\
# -1 & -2 & +0
# \end{array}\right)
# $$
#
# Are the eigenvectors normalized?
#
# Check your results.
A2 = np.array([[-2, +2, -3],
[+2, +1, -6],
[-1, -2, +0]])
lambdas, evecsT = np.linalg.eig(A2)
evecs = evecsT.T
print(lambdas)
print(evecs)
np.linalg.norm(evecs, axis=1)
Identity = np.identity(A2.shape[0])
for evalue, evec in zip(lambdas, evecs):
print((A2 - evalue * Identity).dot(evec))
|
14_linear_algebra/14_Linear_Algebra.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook presents the results for the rank method.
#
# ### Régularité mensuelle TGV par liaisons
#
# ### https://data.sncf.com/explore/dataset/regularite-mensuelle-tgv-aqst/information/?sort=periode
# +
import pandas
import matplotlib.pyplot as plt
fichier="regularite-mensuelle-tgv-aqst.csv"
#df = pandas.read_csv(fichier,sep=";")
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
# -
# ### Mann-Whitney rank test
# +
import scipy.stats
import pandas as pd
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
signal = df[df['Gare de départ']=='PARIS EST']['Nombre de circulations prévues'].to_numpy()
def Detection_Rupture(data, debut, fin):
pv = 1
position = 0
for i in range (debut+1, fin-1):
data1 = data[debut:i]
data2 = data[i:fin]
if scipy.stats.mannwhitneyu(data1, data2).pvalue < pv:
position = i
pv = scipy.stats.mannwhitneyu(data1, data2).pvalue
return(position, pv)
def Segmentation_Binaire(data, debut, fin, alpha):
L = []
position, pvalue = Detection_Rupture(data, debut, fin)
print(pvalue)
if pvalue < alpha:
L.append(position)
L1 = Segmentation_Binaire(data, debut, position, alpha)
L2 = Segmentation_Binaire(data, position, fin, alpha)
L = L + L1 + L2
return(L)
# -
# ### Paris Est
# +
result = Segmentation_Binaire(signal, 0, len(signal), 0.01)
print("k_rupture = ", result)
print('Paris Est', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result[0])]
#l2 = [k for k in range(result[0],result[1])]
#l3 = [k for k in range(result[1],result[2])]
l4 = [k for k in range(result[0],signal.shape[0])]
plt.plot(l1,signal[:result[0]], color = 'green')
#plt.plot(l2,signal[result[0]:result[1]], color = 'blue')
#plt.plot(l3,signal[result[1]:result[2]], color = 'black')
plt.plot(l4,signal[result[0]:], color = 'yellow')
plt.axvline(x=result[0],color='red')
#plt.axvline(x=result[1],color='red')
#plt.axvline(x=result[2],color='red')
plt.show()
# +
df=df[df['Gare de départ']=='PARIS EST']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('Paris Est', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd[result1[2]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='PARIS EST']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('Paris Est', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
plt.plot(l2,dd1[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd1[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd1[result1[2]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### <NAME>
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.02)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],result1[3])]
l5 = [k for k in range(result1[3],result1[4])]
l6 = [k for k in range(result1[4],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd[result1[2]:result1[3]], color = 'purple')
plt.plot(l5,dd[result1[3]:result1[4]], color = 'orange')
plt.plot(l6,dd[result1[4]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='PARIS LYON']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
plt.plot(l2,dd1[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### <NAME>
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='PARIS MONTPARNASSE']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.02)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l6 = [k for k in range(result1[0],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l6,dd[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='PARIS MONTPARNASSE']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.001)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
plt.plot(l2,dd1[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### <NAME>
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.002)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],result1[3])]
l5 = [k for k in range(result1[3],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd[result1[2]:result1[3]], color = 'purple')
plt.plot(l5,dd[result1[3]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
plt.plot(l2,dd1[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd1[result1[1]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### <NAME>
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],result1[3])]
l5 = [k for k in range(result1[3],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd[result1[2]:result1[3]], color = 'purple')
plt.plot(l5,dd[result1[3]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
plt.plot(l2,dd1[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd1[result1[1]:result1[2]], color = 'orange')
plt.plot(l4,dd1[result1[2]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### <NAME>
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='<NAME>']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.06)
result1.sort()
print("k_rupture = ", result1)
print('<NAME>', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
l2 = [k for k in range(result1[0],result1[1])]
l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[2],result1[3])]
l5 = [k for k in range(result1[3],result1[4])]
l6 = [k for k in range(result1[4],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
plt.plot(l4,dd[result1[2]:result1[3]], color = 'purple')
plt.plot(l5,dd[result1[3]:result1[4]], color = 'orange')
plt.plot(l6,dd[result1[4]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='PARIS NORD']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('Paris Nord', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
#l2 = [k for k in range(result1[0],result1[1])]
#l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[0],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
#plt.plot(l2,dd1[result1[0]:result1[1]], color = 'blue')
#plt.plot(l3,dd1[result1[1]:result1[2]], color = 'orange')
plt.plot(l4,dd1[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# -
# ### Lille
# +
df=pandas.read_table(fichier,sep=";")
df.head(5) # afficher les 5 premières lignes
df=df.sort_values(by=[df.columns[0] , df.columns[1]])
df.head(9)
df=df[df['Gare de départ']=='LILLE']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd = df.groupby(['Année','Mois'])['Nombre de circulations prévues'].sum().to_numpy()
result1 = Segmentation_Binaire(dd, 0, len(dd), 0.005)
result1.sort()
print("k_rupture = ", result1)
print('Lille', " : ",'Nombre de circulations prévues' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
#l2 = [k for k in range(result1[0],result1[1])]
#l3 = [k for k in range(result1[1],result1[2])]
#l4 = [k for k in range(result1[2],result1[3])]
#l5 = [k for k in range(result1[3],result1[4])]
l6 = [k for k in range(result1[0],dd.shape[0])]
plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:result1[1]], color = 'blue')
#plt.plot(l3,dd[result1[1]:result1[2]], color = 'black')
#plt.plot(l4,dd[result1[2]:result1[3]], color = 'purple')
#plt.plot(l5,dd[result1[3]:result1[4]], color = 'orange')
plt.plot(l6,dd[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
# +
df=df[df['Gare de départ']=='LILLE']
df=df.sort_values(by=[df.columns[0],df.columns[1]])
df=df[['Année','Mois','Gare de départ','Nombre de circulations prévues','Nombre de trains annulés']]
dd1 = df.groupby(['Année','Mois'])['Nombre de trains annulés'].sum().to_numpy()
result1 = Segmentation_Binaire(dd1, 0, len(dd1), 0.01)
result1.sort()
print("k_rupture = ", result1)
print('Lille', " : ",'Nombre de trains annulés' )
fig = plt.figure(figsize=(20,7))
ax1 = fig.add_subplot(111)
# cacher les cadres : haut et droit
ax1.spines['right'].set_visible(False)
ax1.spines['top'].set_visible(False)
ax1.xaxis.set_ticks_position('bottom')
l1 = [k for k in range(result1[0])]
#l2 = [k for k in range(result1[0],result1[1])]
#l3 = [k for k in range(result1[1],result1[2])]
l4 = [k for k in range(result1[0],dd1.shape[0])]
plt.plot(l1,dd1[:result1[0]], color = 'green')
#plt.plot(l2,dd1[result1[0]:result1[1]], color = 'blue')
#plt.plot(l3,dd1[result1[1]:result1[2]], color = 'orange')
plt.plot(l4,dd1[result1[0]:], color = 'yellow')
#plt.plot(l1,dd[:result1[0]], color = 'green')
#plt.plot(l2,dd[result1[0]:], color = 'blue')
for i in result1 :
plt.axvline(x=i,color='red')
plt.show()
|
Rank Method.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MoghazyCoder/Machine-Learning-Tutorials/blob/master/Tutorials/Basic_Exploratory_Data_Analysis_using_Python_libraries.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="QinEMsEy2fDo" colab_type="text"
# # Data Engineering Tutorial
#
# ### In this tutorial we will discuss the basics of the following libraries:
# * Numby
# * Pandas
# * Matplotlib
# + [markdown] id="qAA9gt512fD4" colab_type="text"
# ### Numpy (Numerical Python)
# Unlike using basic python functions and loops, numpy is so fast as most of its modules are implemented in C language. Let's now have a look on some of the basic operations in Numpy.
# #### Calculating the mean:
# To calculate the mean for the following array, we need to use numpy.mean() function.<br>
# arr = [1,5,2,7,9,10]
# + id="zs0f55Ji2fD6" colab_type="code" colab={}
# Importing numpy library and giving it a name "np" for fast access
import numpy as np
test_arr = np.array([1,5,2,6,8,3])
# + id="ikcf417m2fEA" colab_type="code" colab={} outputId="f6b9f943-ada1-4608-e453-72774c18f023"
# Calculating the mean
mean = np.mean(test_arr)
print("The mean of the array = %f" % mean) # Must be 4.1667 Why wrong answer? HINT what does the %i in the format string do
# + [markdown] id="FvYZRHpn2fER" colab_type="text"
# #### Calculating the midean:
# We calculate the median manually by sorting the elements and picking the value (or mean of the values) in the middle. To calculate the median using Numpy we can use numpy.median() function.<br>
# We will use the same array arr == Note that the sorted array will be arr_sorted = [1,2,3,5,6,8] so the median must be (3+5)/2 = 4
# + id="Yhky_5Bh2fEi" colab_type="code" colab={} outputId="04eacf0b-aae2-4ff4-8020-aa1151a7705f"
# Calculating the midean
median = np.median(test_arr)
print("The median of the array = %0.2f" % median)
# + [markdown] id="nGqQNOL92fFK" colab_type="text"
# #### Calculating the Standart Deviation (std):
# One of the very important statistical terms that measures the dispersion of the data values of the dataset, can be a measure to judge if a data point (data instance/example) is an outlier or not (researchers usually consider points with std greater that 2.5/3 an outlier), So why are outliers bad?
# We calculate the STD using Numpy by using numpy.std() function.
# + id="ecZTXLCo2fFL" colab_type="code" colab={} outputId="696f73e2-15ef-4e08-cce0-6813dc193abb"
#Calculat the STD using the same array
std = np.std(test_arr)
print("The median of the array = %0.2f" % std)
# + [markdown] id="7_n6DX4H2fFR" colab_type="text"
# #### Calculating the mode:
# The most represented value. Numpy?
# + id="lJ_gZmJy2fFT" colab_type="code" colab={} outputId="b631a01a-1ed1-471f-cd9d-e90ae9d542ae"
#Calculat the mode using scipy
from scipy import stats
stats.mode([2,3,4,5])
# + [markdown] id="4_xye0eF2fFb" colab_type="text"
# ### Pandas 🐼🐼
# The very famous Python Data Analysis Library, mainly used in the EDA (Exploratory Data Analysis) stage to import data into a dataframe, explore the correlations between features, visualise the scatter plot of the data points and etc.
# But what is a dataframe?<br>
# Pandas is usually used to import data from a (CSV file)?? which is the most poplular formats for structured datasets. <br>
# Let's first start by importing the cover_type dataset from sklearn library. Cover_type dataset?? Trees?? 😂
#
#
# + id="IpowlQDV2fFd" colab_type="code" colab={} outputId="9352c6bf-44c7-4c14-a944-d2ece8f60f0d"
# Importing the data set from sklearn library
from sklearn.datasets import fetch_covtype
cov = fetch_covtype()
columns = ['Elevation', 'Aspect', 'Slope',
'Horizontal_Distance_To_Hydrology', 'Vertical_Distance_To_Hydrology',
'Horizontal_Distance_To_Roadways', 'Hillshade_9am', 'Hillshade_Noon',
'Hillshade_3pm', 'Horizontal_Distance_To_Fire_Points',
'Wilderness_Area1', 'Wilderness_Area2', 'Wilderness_Area3',
'Wilderness_Area4', 'Soil_Type1', 'Soil_Type2', 'Soil_Type3',
'Soil_Type4', 'Soil_Type5', 'Soil_Type6', 'Soil_Type7', 'Soil_Type8',
'Soil_Type9', 'Soil_Type10', 'Soil_Type11', 'Soil_Type12',
'Soil_Type13', 'Soil_Type14', 'Soil_Type15', 'Soil_Type16',
'Soil_Type17', 'Soil_Type18', 'Soil_Type19', 'Soil_Type20',
'Soil_Type21', 'Soil_Type22', 'Soil_Type23', 'Soil_Type24',
'Soil_Type25', 'Soil_Type26', 'Soil_Type27', 'Soil_Type28',
'Soil_Type29', 'Soil_Type30', 'Soil_Type31', 'Soil_Type32',
'Soil_Type33', 'Soil_Type34', 'Soil_Type35', 'Soil_Type36',
'Soil_Type37', 'Soil_Type38', 'Soil_Type39', 'Soil_Type40']
# ??? Why all of this?? np arrays doesn't have info about the features names
# + id="T4m6lGyo2fFk" colab_type="code" colab={}
import pandas as pd
# Import the data into a dataframe for exploration
data = pd.DataFrame(cov.data, columns = columns)
data['Cover_Type'] = cov.target
# + [markdown] id="hngPTZ9A2fFq" colab_type="text"
# Let's explore what a dataframe can do. We will start with the head([n_rows]) function that displays the first n_rows from the dataset as follows:
# + id="rPQFqmNk2fFr" colab_type="code" colab={} outputId="6d02176e-c250-4546-fbca-b4ab5a595603"
data.head(5) # Default?
# + id="4j81R73O2fFz" colab_type="code" colab={} outputId="d7291103-04f7-40a9-8eaf-cbc1babf9ee3"
data.Soil_Type35.value_counts()
# + [markdown] id="b3MO99VC2fF8" colab_type="text"
# Using pandas DataFrames we can select specific columns now only specific rows. Let's now start selecting and exploring some rows.
# + id="L34TldYF2fF-" colab_type="code" colab={} outputId="1cc60731-026d-4a8b-c71c-e22c26261bc3"
data['Elevation'] # Could be data.Elevation as well
# + [markdown] id="I9xTEsrM2fGG" colab_type="text"
# We can also know the frequency of each value in this column using value_counts() function
# + id="Fp6B8oMY2fGI" colab_type="code" colab={} outputId="9ce9b2be-9908-4ea8-ac5a-b4bf8cc8375c"
data.Elevation.value_counts();
data.Cover_Type.value_counts()
# + [markdown] id="OsCTfuZO2fGN" colab_type="text"
# Oh, the data was continuous, should have checked the column's value before counting its values.
# We mainly use two functions to get the basic statistical info from the data. The first one is DataFrame.info() function which returns a summary of the dataframe
# + id="rTk4yfA02fGP" colab_type="code" colab={} outputId="6435899a-8e97-48d1-ceae-15579a99e229"
data.info();
# + [markdown] id="_fac8DU82fGl" colab_type="text"
# The second function describes the statistical properties of each feature in the dataset as follows:
# + id="D1RchABs2fGn" colab_type="code" colab={} outputId="b2f07633-6bd9-41d8-ef5b-ea70387110b4"
data.describe()
# + [markdown] id="RmGYmJql2fGs" colab_type="text"
# We can visualize some of the dataset features histograms and correlations using pandas as follows. Remember how we calculated the frequency of the items in the label feature (which is the target value), we will now visualize the histogram.
# + id="nQN3U5zb2fGt" colab_type="code" colab={} outputId="2dd8985b-0778-4b14-c95d-4382505b6e19"
# Import matplotlib to show the graph
import matplotlib.pyplot as plt
# Why using bins??
data.Cover_Type.hist(bins=7)
plt.show()
# + [markdown] id="1bw_IKho2fGx" colab_type="text"
# It is very important to explore the correlation betwwen the features as explained in the lecture (Remember Naive bayes?). We will now descover the corelation between some numerical features that we have in the dataset usign Dataframe.corr(). The correlation value must be between -1 and 1 where -1 means inversly correlated and 1 means correlated. Why is this important? Feature selection and other reasons.
# + id="9PLZeB3X2fGy" colab_type="code" colab={} outputId="54b45f03-b22d-4270-81dc-4e8cbf5e4e7f"
data[['Elevation', 'Aspect', 'Slope', 'Cover_Type']].corr()
# + [markdown] id="dashhqPQ2fG1" colab_type="text"
# Looks good, what if the columns count was huge?? would it look good??
# + id="gxmuFGCe2fG2" colab_type="code" colab={} outputId="a2a5429a-94a6-477b-f82b-09bc1f1df791"
data.corr()
# + [markdown] id="jowQXY222fG7" colab_type="text"
# Really hard to read floating point number and compare them. Solutions?? Heat Map
# + id="eYF5vk3V2fG8" colab_type="code" colab={} outputId="6a21eb1a-2bd7-4ca4-f0db-0a3aef2c42ec"
import seaborn as sns
import matplotlib.pyplot as plt
corr = data[['Elevation', 'Aspect', 'Slope',
'Horizontal_Distance_To_Hydrology', 'Vertical_Distance_To_Hydrology',
'Horizontal_Distance_To_Roadways', 'Hillshade_9am', 'Hillshade_Noon',
'Hillshade_3pm', 'Horizontal_Distance_To_Fire_Points']].corr()
f, ax = plt.subplots(figsize=(25, 25))
# Color Map, not mandatory
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Heat Map
sns.heatmap(corr, cmap=cmap, vmax=1, vmin = -1, center=0,
square=True, linewidths=.5)
# + [markdown] id="fox4C2UG2fG_" colab_type="text"
# There are many other libraries that have very interesting usage in Data Science in general.
# We will now use seaborn library to visualize the scatter plot of three dataset features
# + id="GzaMtp3I2fHB" colab_type="code" colab={} outputId="867d4864-6efa-4b07-8f25-d28198f64f9f"
import seaborn as sns
Exploration_columns = data[['Horizontal_Distance_To_Hydrology', 'Vertical_Distance_To_Hydrology','Cover_Type']].head(1000)
sns.pairplot(Exploration_columns, hue='Cover_Type')
# + [markdown] id="jYKT4_Q02fHI" colab_type="text"
# Can also scatter using pandas but in a more premitive way
# + id="RfUc3AYn2fHJ" colab_type="code" colab={} outputId="0e7334dc-fd0b-426b-b346-9d57f706d176"
from pandas.plotting import scatter_matrix
# Can select multiple rows for exploration
scatter_matrix(data[['Elevation', 'Aspect', 'Slope']])
plt.show()
# + id="EEd5GgX62fHN" colab_type="code" colab={} outputId="cbc768a9-e022-417c-beda-522c943bc53d"
data.isna().sum()
# + [markdown] id="uwk69ins2fHS" colab_type="text"
# Resources:
#
# https://github.com/MoghazyCoder/a-2017/blob/master/Labs/Lab1_numstack/Lab1-numstack_solutions.ipynb
# https://www.kaggle.com/moghazy/ensemble-learning-with-feature-engineering
# https://pandas.pydata.org/
# https://docs.scipy.org/
# https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_covtype.html#sklearn.datasets.fetch_covtype
|
Tutorials/Basic_Exploratory_Data_Analysis_using_Python_libraries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.00584, "end_time": "2021-02-27T00:41:58.746499", "exception": false, "start_time": "2021-02-27T00:41:58.740659", "status": "completed"} tags=[]
# # Introduction
#
# A **model card** is a short document that provides key information about a machine learning model. Model cards increase transparency by communicating information about trained models to broad audiences.
#
# In this tutorial, you will learn about which audiences to write a model card for and which sections a model card should contain. Then, in the following **[exercise](#$NEXT_NOTEBOOK_URL$)**, you will apply what you have learned to a couple of real-world scenarios.
# + [markdown] papermill={"duration": 0.003871, "end_time": "2021-02-27T00:41:58.754812", "exception": false, "start_time": "2021-02-27T00:41:58.750941", "status": "completed"} tags=[]
# # Model cards
#
# Though AI systems are playing increasingly important roles in every industry, few people understand how these systems work. AI researchers are exploring many ways to communicate key information about models to inform people who use AI systems, people who are affected by AI systems and others.
#
# Model cards - introduced in a [2019 paper](https://arxiv.org/abs/1810.03993) - are one way for teams to communicate key information about their AI system to a broad audience. This information generally includes intended uses for the model, how the model works, and how the model performs in different situations.
#
# You can think of model cards as similar to the nutritional labels that you find on packaged foods.
#
# # Examples of model cards
#
# Before we continue, it might be useful to briefly skim some examples of model cards.
# - [Salesforce's model cards](https://blog.einstein.ai/model-cards-for-ai-model-transparency/)
# - [Open AI’s model card for GPT-3](https://github.com/openai/gpt-3/blob/master/model-card.md)
# - [Google Cloud's example model cards](https://modelcards.withgoogle.com/face-detection)
#
# # Who is the audience of your model card?
#
# A model card should strike a balance between being easy-to-understand and communicating important technical information. When writing a model card, you should consider your audience: the groups of people who are most likely to read your model card. These groups will vary according to the AI system’s purpose.
#
# For example, a model card for an AI system that helps medical professionals interpret x-rays to better diagnose musculoskeletal injuries is likely to be read by medical professionals, scientists, patients, researchers, policymakers and developers of similar AI systems. The model card may therefore assume some knowledge of health care and of AI systems.
# + [markdown] papermill={"duration": 0.003913, "end_time": "2021-02-27T00:41:58.762853", "exception": false, "start_time": "2021-02-27T00:41:58.758940", "status": "completed"} tags=[]
# # What sections should a model card contain?
#
# Per the original paper, a model card should have the following nine sections. Note that different organizations may add, subtract or rearrange model card sections according to their needs (and you may have noticed this in some of the examples above).
#
# As you read about the different sections, you're encouraged to review the two example model cards from the original paper. Before proceeding, open each of these model card examples in a new window:
# - [Model Card - Smiling Detection in Images](https://github.com/Kaggle/learntools/blob/master/notebooks/ethics/pdfs/smiling_in_images_model_card.pdf)
# - [Model Card - Toxicity in Text](https://github.com/Kaggle/learntools/blob/master/notebooks/ethics/pdfs/toxicity_in_text_model_card.pdf)
#
# ## 1. Model Details
# * Include background information, such as developer and model version.
#
# ## 2. Intended Use
# - What use cases are in scope?
# - Who are your intended users?
# - What use cases are out of scope?
#
# ## 3. Factors
# - What factors affect the impact of the model? For example, the smiling detection model's results vary by demographic factors like age, gender or ethnicity, environmental factors like lighting or rain and instrumentation like camera type.
#
# ## 4. Metrics
# - What metrics are you using to measure the performance of the model? Why did you pick those metrics?
# - For **classification systems** – in which the output is a class label – potential error types include false positive rate, false negative rate, false discovery rate, and false omission rate. The relative importance of each of these depends on the use case.
# - For **score-based analyses** – in which the output is a score or price – consider reporting model performance across groups.
#
#
# ## 5. Evaluation Data
# - Which datasets did you use to evaluate model performance? Provide the datasets if you can.
# - Why did you choose these datasets for evaluation?
# - Are the datasets representative of typical use cases, anticipated test cases and/or challenging cases?
#
# ## 6. Training Data
# - Which data was the model trained on?
#
# ## 7. Quantitative Analyses
# - How did the model perform on the metrics you chose? Break down performance by important factors and their intersections. For example, in the smiling detection example, performance is broken down by age (eg, young, old), gender (eg, female, male), and then both (eg, old-female, old-male, young-female, young-male).
#
# ## 8. Ethical Considerations
# - Describe ethical considerations related to the model, such as sensitive data used to train the model, whether the model has implications for human life, health, or safety, how risk was mitigated, and what harms may be present in model usage.
#
# ## 9. Caveats and Recommendations
# - Add anything important that you have not covered elsewhere in the model card.
# + [markdown] papermill={"duration": 0.003778, "end_time": "2021-02-27T00:41:58.770728", "exception": false, "start_time": "2021-02-27T00:41:58.766950", "status": "completed"} tags=[]
# # How can you use model cards in your organization?
#
# The use of detailed model cards can often be challenging because an organization may not want to reveal its processes, proprietary data or trade secrets. In such cases, the developer team should think about how model cards can be useful and empowering, without including sensitive information.
#
# Some teams use other formats - such as [FactSheets](https://aifs360.mybluemix.net/) - to collect and log ML model information.
# + [markdown] papermill={"duration": 0.003853, "end_time": "2021-02-27T00:41:58.779094", "exception": false, "start_time": "2021-02-27T00:41:58.775241", "status": "completed"} tags=[]
# # Your turn
#
# Apply what you've learned to **[decide how to use model cards](#$NEXT_NOTEBOOK_URL$)** in real-world scenarios.
|
notebooks/ethics/raw/tut5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
# %matplotlib inline
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
# +
pickle_file = '../dataset/arbimonTest1.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# -
sample_image = train_dataset[50]#[0]
sample_image = sample_image.reshape((100,100))
sample_image.shape
# +
import matplotlib.pyplot as plt
plt.figure()
plt.imshow(sample_image)
# +
from scipy.ndimage.interpolation import shift
sample_shifted_image = shift(sample_image, [5, 0])
plt.figure()
plt.imshow(sample_shifted_image)
# -
one_pixel_image = []
one_pixel_label = []
for i in range(len(train_dataset)):
pre_image = train_dataset[i].reshape((100,100))
shifted_image = shift(pre_image, [0,10])
one_pixel_image.append(shifted_image)
one_pixel_label.append(train_labels[i])
plt.figure()
plt.imshow(one_pixel_image[5])
|
Notebooks/Testing Data Augmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import geopandas as gpd
from lets_plot import *
LetsPlot.setup_html()
# -
# ## Prepare the dataset
# +
from sklearn.datasets import fetch_california_housing
california_housing_bunch = fetch_california_housing()
data = pd.DataFrame(california_housing_bunch.data, columns=california_housing_bunch.feature_names)
# Add $-value field to the dataframe.
# dataset.target: numpy array of shape (20640,)
# Each value corresponds to the average house value in units of 100,000.
data['Value($)'] = california_housing_bunch.target * 100000
data.head()
# +
# Draw a random sample from the data set.
# data = data.sample(n=1000)
# -
# ### Use `geopandas` to read a shape file to GeoDataFrame
CA = gpd.read_file("./ca-state-boundary/CA_State_TIGER2016.shp")
CA.head()
# Keeping in mind that our target is the housing value, fill the choropleth over the state contours using `geom_map()`function
# ### Make a plot out of polygon and points
#
# The color of the points will reflect the house age and
# the size of the points will reflect the value of the house.
# +
# The plot base
p = ggplot(data=data) + scale_color_gradient(name='House Age', low='red', high='green')
# The points layer
points = geom_point(aes(x='Longitude',
y='Latitude',
size='Value($)',
color='HouseAge'),
# data=data,
alpha=0.8)
# The map
p + geom_polygon(map=CA, fill='#F8F4F0', color='#B71234')\
+ points\
+ theme(axis_title='blank', axis_text='blank', axis_ticks='blank', axis_line='blank', axis_tooltip='blank')\
+ ggsize(600, 500)
# -
LetsPlot.set(maptiles_zxy(url='https://maps.wikimedia.org/osm-intl/{z}/{x}/{y}@2x.png'))
# ### Make a plot similar to the one above but interactive
p + geom_livemap()\
+ geom_polygon(map=CA, fill='white', color='#B71234', alpha=0.5)\
+ points
|
docs/examples/jupyter-notebooks-dev/issues/map-california-housing/map_california_housing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] colab_type="text" id="copyright-notice"
# #### Copyright 2017 Google LLC.
# + colab={"autoexec": {"wait_interval": 0, "startup": false}} cellView="both" colab_type="code" id="copyright-notice2"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="PTaAdgy3LS8W" colab_type="text"
# # 희소 데이터 및 임베딩 소개
#
# **학습 목표:**
# * 영화 리뷰 문자열 데이터를 희소 특성 벡터로 변환한다
# * 희소 특성 벡터를 사용하여 감정 분석 선형 모델을 구현한다
# * 데이터를 두 차원으로 투영하는 임베딩을 사용하여 감정 분석 DNN 모델을 구현한다
# * 임베딩을 시각화하여 단어 간의 관계에 대해 모델이 학습한 내용을 확인한다
#
# 이 실습에서는 희소 데이터에 대해 알아보고 [ACL 2011 IMDB 데이터 세트](http://ai.stanford.edu/~amaas/data/sentiment/)에서 가져온 영화 리뷰 텍스트 데이터로 임베딩을 사용해 봅니다. 이 데이터는 이미 `tf.Example` 형식으로 처리되어 있습니다.
# + [markdown] id="2AKGtmwNosU8" colab_type="text"
# ## 설정
#
# 우선 필요한 모듈을 import로 불러오고 학습 및 테스트 데이터를 다운로드합니다. [`tf.keras`](https://www.tensorflow.org/api_docs/python/tf/keras)에 포함된 파일 다운로드 및 캐싱 도구를 사용하여 데이터 세트를 검색할 수 있습니다.
# + id="jGWqDqFFL_NZ" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from __future__ import print_function
import collections
import io
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
from IPython import display
from sklearn import metrics
tf.logging.set_verbosity(tf.logging.ERROR)
train_url = 'https://download.mlcc.google.com/mledu-datasets/sparse-data-embedding/train.tfrecord'
train_path = tf.keras.utils.get_file(train_url.split('/')[-1], train_url)
test_url = 'https://download.mlcc.google.com/mledu-datasets/sparse-data-embedding/test.tfrecord'
test_path = tf.keras.utils.get_file(test_url.split('/')[-1], test_url)
# + [markdown] id="6W7aZ9qspZVj" colab_type="text"
# ## 감정 분석 모델 만들기
# + [markdown] id="jieA0k_NLS8a" colab_type="text"
# 이 데이터로 감정 분석 모델을 학습시켜 리뷰가 전반적으로 *긍정적*(라벨 1)인지 아니면 *부정적*(라벨 0)인지를 예측해 보겠습니다.
#
# 이렇게 하려면 문자열 값인 `단어`를 *어휘*, 즉 데이터에 나올 것으로 예상되는 각 단어의 목록을 사용하여 특성 벡터로 변환합니다. 이 실습을 진행하기 위해 제한된 단어 집합을 갖는 소규모 어휘를 만들었습니다. 이러한 단어는 대부분 *긍정* 또는 *부정*을 강하게 암시하는 것이 밝혀졌지만 일부분은 단순히 흥미를 위해 추가되었습니다.
#
# 어휘의 각 단어는 특성 벡터의 좌표에 매핑됩니다. 예의 문자열 값인 `단어`를 이 특성 벡터로 변환하기 위해, 예 문자열에 어휘 단어가 나오지 않으면 각 좌표의 값에 0을 입력하고 어휘 단어가 나오면 1을 입력하도록 인코딩하겠습니다. 예의 단어 중 어휘에 나오지 않는 단어는 무시됩니다.
# + [markdown] id="2HSfklfnLS8b" colab_type="text"
# **참고:** *물론 더 큰 어휘를 사용할 수도 있으며 이러한 어휘를 만드는 데 특화된 도구들이 있습니다. 뿐만 아니라 어휘에 나오지 않는 단어를 단순히 무시하지 않고 적은 수의 OOV(out-of-vocabulary) 버킷을 도입하여 해당 단어를 해시할 수 있습니다. 명시적인 어휘를 만드는 대신 각 단어를 해시하는 __특성 해싱__ 접근법을 사용할 수도 있습니다. 이 방법은 실무에는 적합하지만 해석 가능성이 사라지므로 실습 목적으로는 유용하지 않습니다. 이와 관련된 도구에 대해서는 tf.feature_column 모듈을 참조하세요.*
# + [markdown] id="Uvoa2HyDtgqe" colab_type="text"
# ## 입력 파이프라인 구축
# + [markdown] id="O20vMEOurDol" colab_type="text"
# 우선 텐서플로우 모델로 데이터를 가져오는 입력 파이프라인을 구성하겠습니다. 다음 함수를 사용하여 [TFRecord](https://www.tensorflow.org/guide/datasets#consuming_tfrecord_data) 형식인 입력 및 테스트 데이터를 파싱하고 특성과 해당 라벨로 이루어진 dict를 반환할 수 있습니다.
# + id="SxxNIEniPq2z" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
def _parse_function(record):
"""Extracts features and labels.
Args:
record: File path to a TFRecord file
Returns:
A `tuple` `(labels, features)`:
features: A dict of tensors representing the features
labels: A tensor with the corresponding labels.
"""
features = {
"terms": tf.VarLenFeature(dtype=tf.string), # terms are strings of varying lengths
"labels": tf.FixedLenFeature(shape=[1], dtype=tf.float32) # labels are 0 or 1
}
parsed_features = tf.parse_single_example(record, features)
terms = parsed_features['terms'].values
labels = parsed_features['labels']
return {'terms':terms}, labels
# + [markdown] id="SXhTeeYMrp-l" colab_type="text"
# 함수가 정상적으로 작동하는지 확인하기 위해 학습 데이터에 대한 `TFRecordDataset`를 생성하고 위 함수를 사용하여 데이터를 특성 및 라벨에 매핑합니다.
# + id="oF4YWXR0Omt0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Create the Dataset object.
ds = tf.data.TFRecordDataset(train_path)
# Map features and labels with the parse function.
ds = ds.map(_parse_function)
ds
# + [markdown] id="bUoMvK-9tVXP" colab_type="text"
# 다음 셀을 실행하여 학습 데이터 세트에서 첫 예를 확인합니다.
# + id="Z6QE2DWRUc4E" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
n = ds.make_one_shot_iterator().get_next()
sess = tf.Session()
sess.run(n)
# + [markdown] id="jBU39UeFty9S" colab_type="text"
# 이제 텐서플로우 에스티메이터 개체의 `train()` 메소드에 전달할 수 있는 정식 입력 함수를 만들겠습니다.
# + id="5_C5-ueNYIn_" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Create an input_fn that parses the tf.Examples from the given files,
# and split them into features and targets.
def _input_fn(input_filenames, num_epochs=None, shuffle=True):
# Same code as above; create a dataset and map features and labels.
ds = tf.data.TFRecordDataset(input_filenames)
ds = ds.map(_parse_function)
if shuffle:
ds = ds.shuffle(10000)
# Our feature data is variable-length, so we pad and batch
# each field of the dataset structure to whatever size is necessary.
ds = ds.padded_batch(25, ds.output_shapes)
ds = ds.repeat(num_epochs)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + [markdown] id="Y170tVlrLS8c" colab_type="text"
# ## 작업 1: 희소 입력 및 명시적 어휘와 함께 선형 모델 사용
#
# 첫 번째 모델로서 50개의 정보 단어를 사용하여 [`LinearClassifier`](https://www.tensorflow.org/api_docs/python/tf/estimator/LinearClassifier) 모델을 만들겠습니다. 처음에는 단순하게 시작하는 것이 좋습니다.
#
# 다음 코드는 단어에 대한 특성 열을 만듭니다. [`categorical_column_with_vocabulary_list`](https://www.tensorflow.org/api_docs/python/tf/feature_column/categorical_column_with_vocabulary_list) 함수는 문자열과 특성 벡터 간의 매핑을 포함하는 특성 열을 만듭니다.
# + id="B5gdxuWsvPcx" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# 50 informative terms that compose our model vocabulary.
informative_terms = ("bad", "great", "best", "worst", "fun", "beautiful",
"excellent", "poor", "boring", "awful", "terrible",
"definitely", "perfect", "liked", "worse", "waste",
"entertaining", "loved", "unfortunately", "amazing",
"enjoyed", "favorite", "horrible", "brilliant", "highly",
"simple", "annoying", "today", "hilarious", "enjoyable",
"dull", "fantastic", "poorly", "fails", "disappointing",
"disappointment", "not", "him", "her", "good", "time",
"?", ".", "!", "movie", "film", "action", "comedy",
"drama", "family")
terms_feature_column = tf.feature_column.categorical_column_with_vocabulary_list(key="terms", vocabulary_list=informative_terms)
# + [markdown] id="eTiDwyorwd3P" colab_type="text"
# 다음으로, `LinearClassifier`를 생성하고 학습 세트로 학습시킨 후 평가 세트로 평가합니다. 코드를 잘 읽어보고 실행하여 결과를 확인해 보세요.
# + id="HYKKpGLqLS8d" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
feature_columns = [ terms_feature_column ]
classifier = tf.estimator.LinearClassifier(
feature_columns=feature_columns,
optimizer=my_optimizer,
)
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] id="J0ubn9gULS8g" colab_type="text"
# ## 작업 2: 심층신경망(DNN) 모델 사용
#
# 위 모델은 선형 모델입니다. 비교적 좋은 성능을 발휘하지만, DNN 모델로 성능을 더 높일 수 있을까요?
#
# `LinearClassifier`를 [`DNNClassifier`](https://www.tensorflow.org/api_docs/python/tf/estimator/DNNClassifier)로 교체해 보겠습니다. 다음 셀을 실행하고 결과를 확인해 보세요.
# + id="jcgOPfEALS8h" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
##################### Here's what we changed ##################################
classifier = tf.estimator.DNNClassifier( #
feature_columns=[tf.feature_column.indicator_column(terms_feature_column)], #
hidden_units=[20,20], #
optimizer=my_optimizer, #
) #
###############################################################################
try:
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
except ValueError as err:
print(err)
# + [markdown] id="cZz68luxLS8j" colab_type="text"
# ## 작업 3: DNN 모델에 임베딩 사용
#
# 이 작업에서는 임베딩 열을 사용하여 DNN 모델을 구현합니다. 임베딩 열은 희소 데이터를 입력으로 취하고 저차원 밀집 벡터를 출력으로 반환합니다.
# + [markdown] id="AliRzhvJLS8k" colab_type="text"
# **참고:** *희소 데이터로 모델을 학습시킬 때 embedding_column은 일반적으로 연산 효율이 가장 높은 옵션입니다. 이 실습 끝부분의 [선택 섹션](#scrollTo=XDMlGgRfKSVz)에서 `embedding_column`과 `indicator_column`의 구현상 차이점 및 상대적인 장단점을 자세히 알아봅니다.*
# + [markdown] id="F-as3PtALS8l" colab_type="text"
# 아래 코드에서 다음을 수행합니다.
#
# * 데이터를 2개의 차원으로 투영하는 `embedding_column`을 사용하여 모델의 특성 열을 정의합니다. `embedding_column`의 함수 시그니처에 대한 자세한 내용은 [TF 문서](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column)를 참조하세요.
# * 다음과 같은 사양으로 `DNNClassifier`를 정의합니다.
# * 각각 20개 유닛을 포함하는 히든 레이어 2개
# * Adagrad 최적화, 학습률 0.1
# * `gradient_clip_norm`을 5.0으로 지정
# + [markdown] id="UlPZ-Q9bLS8m" colab_type="text"
# **참고:** *실무에서는 2보다 높은 50 또는 100차원으로 투영하게 됩니다. 그러나 여기에서는 시각화하기 쉽도록 2차원만 사용합니다.*
# + [markdown] id="mNCLhxsXyOIS" colab_type="text"
# ### 힌트
# + id="L67xYD7hLS8m" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Here's a example code snippet you might use to define the feature columns:
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
# + [markdown] id="iv1UBsJxyV37" colab_type="text"
# ### 아래 코드 완성하기
# + id="5PG_yhNGLS8u" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
########################## YOUR CODE HERE ######################################
terms_embedding_column = # Define the embedding column
feature_columns = # Define the feature columns
classifier = # Define the DNNClassifier
################################################################################
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] id="eQS5KQzBybTY" colab_type="text"
# ### 해결 방법
#
# 해결 방법을 보려면 아래를 클릭하세요.
# + id="R5xOdYeQydi5" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
########################## SOLUTION CODE ########################################
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
classifier = tf.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[20,20],
optimizer=my_optimizer
)
#################################################################################
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] id="aiHnnVtzLS8w" colab_type="text"
# ## 작업 4: 임베딩이 실제로 적용되는지 확인
#
# 위 모델에서 사용한 `embedding_column`은 제대로 작동하는 것 같지만, 내부적으로는 어떻게 사용되는지 알 수가 없습니다. 모델에서 내부적으로 임베딩을 실제로 사용하는지 확인하려면 어떻게 해야 할까요?
#
# 우선 모델의 텐서를 살펴보겠습니다.
# + id="h1jNgLdQLS8w" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier.get_variable_names()
# + [markdown] id="Sl4-VctMLS8z" colab_type="text"
# 이제 `'dnn/input_from_feature_columns/input_layer/terms_embedding/...'`이라는 임베딩 레이어가 있음을 확인할 수 있습니다. 여기에서 흥미로운 점은 이 레이어는 여타 히든 레이어와 마찬가지로 모델의 다른 부분과 함께 동시에 학습된다는 점입니다.
#
# 임베딩 레이어가 올바른 형태로 되어 있을까요? 다음 코드를 실행하여 알아보세요.
# + [markdown] id="JNFxyQUiLS80" colab_type="text"
# **참고:** *여기에서 사용하는 임베딩은 50차원 벡터를 2차원으로 투영하는 행렬입니다.*
# + id="1xMbpcEjLS80" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights').shape
# + [markdown] id="MnLCIogjLS82" colab_type="text"
# 잠시 동안 다양한 레이어와 형태를 직접 확인하여 모든 요소가 예상대로 연결되어 있는지 확인해 보세요.
# + [markdown] id="rkKAaRWDLS83" colab_type="text"
# ## 작업 5: 임베딩 조사
#
# 이제 실제 임베딩 공간을 조사하여 각 단어가 결국 어느 위치에 배치되었는지 확인해 보겠습니다. 다음을 수행하세요.
# 1. 다음 코드를 실행하여 **작업 3**에서 학습시킨 임베딩을 확인합니다. 결과가 예상과 일치하나요?
#
# 2. **작업 3**의 코드를 재실행하여 모델을 다시 학습시킨 후 아래의 임베딩 시각화를 다시 실행합니다. 무엇이 그대로인가요? 무엇이 달라졌나요?
#
# 3. 마지막으로 10단계만 사용하여 모델을 다시 학습시킵니다. 이렇게 하면 매우 열악한 모델이 만들어집니다. 아래의 임베딩 시각화를 다시 실행합니다. 이제 결과가 어떠한가요? 이유는 무엇일까요?
# + id="s4NNu7KqLS84" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
import numpy as np
import matplotlib.pyplot as plt
embedding_matrix = classifier.get_variable_value('dnn/input_from_feature_columns/input_layer/terms_embedding/embedding_weights')
for term_index in range(len(informative_terms)):
# Create a one-hot encoding for our term. It has 0s everywhere, except for
# a single 1 in the coordinate that corresponds to that term.
term_vector = np.zeros(len(informative_terms))
term_vector[term_index] = 1
# We'll now project that one-hot vector into the embedding space.
embedding_xy = np.matmul(term_vector, embedding_matrix)
plt.text(embedding_xy[0],
embedding_xy[1],
informative_terms[term_index])
# Do a little setup to make sure the plot displays nicely.
plt.rcParams["figure.figsize"] = (15, 15)
plt.xlim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.ylim(1.2 * embedding_matrix.min(), 1.2 * embedding_matrix.max())
plt.show()
# + [markdown] id="pUb3L7pqLS86" colab_type="text"
# ## 작업 6: 모델 성능 개선 시도
#
# 모델을 다듬어 성능을 높일 수 있는지 확인해 보세요. 다음과 같은 방법을 시도해 볼 수 있습니다.
#
# * **초매개변수 변경** 또는 Adam 등의 **다른 옵티마이저 사용**. 이 전략으로 향상되는 정확성은 1~2%에 불과할 수 있습니다.
# * **`informative_terms`에 더 많은 단어 추가.** 이 데이터 세트의 30,716개 단어를 모두 포함하는 전체 어휘 파일은 https://download.mlcc.google.com/mledu-datasets/sparse-data-embedding/terms.txt 입니다. 이 어휘 파일에서 단어를 더 추출할 수도 있고, `categorical_column_with_vocabulary_file` 특성 열을 통해 전체 어휘를 사용할 수도 있습니다.
# + id="6-b3BqXvLS86" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Download the vocabulary file.
terms_url = 'https://download.mlcc.google.com/mledu-datasets/sparse-data-embedding/terms.txt'
terms_path = tf.keras.utils.get_file(terms_url.split('/')[-1], terms_url)
# + id="0jbJlwW5LS8-" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# Create a feature column from "terms", using a full vocabulary file.
informative_terms = None
with io.open(terms_path, 'r', encoding='utf8') as f:
# Convert it to a set first to remove duplicates.
informative_terms = list(set(f.read().split()))
terms_feature_column = tf.feature_column.categorical_column_with_vocabulary_list(key="terms",
vocabulary_list=informative_terms)
terms_embedding_column = tf.feature_column.embedding_column(terms_feature_column, dimension=2)
feature_columns = [ terms_embedding_column ]
my_optimizer = tf.train.AdagradOptimizer(learning_rate=0.1)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
classifier = tf.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[10,10],
optimizer=my_optimizer
)
classifier.train(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([train_path]),
steps=1000)
print("Training set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
evaluation_metrics = classifier.evaluate(
input_fn=lambda: _input_fn([test_path]),
steps=1000)
print("Test set metrics:")
for m in evaluation_metrics:
print(m, evaluation_metrics[m])
print("---")
# + [markdown] id="ew3kwGM-LS9B" colab_type="text"
# ## 맺음말
#
# 임베딩을 사용한 DNN 솔루션이 원래의 선형 모델보다 우수할 수 있지만, 선형 모델도 성능이 그다지 나쁘지 않았으며 학습 속도는 상당히 더 빨랐습니다. 선형 모델의 학습 속도가 더 빠른 이유는 업데이트할 매개변수 또는 역전파할 레이어의 수가 더 적기 때문입니다.
#
# 응용 분야에 따라서는 선형 모델의 빠른 속도가 큰 장점이 될 수 있고, 선형 모델도 품질 면에서 충분하고도 남을 수 있습니다. 다른 분야에서는 DNN이 제공하는 추가적인 모델 복잡성과 용량이 더 중요할 수 있습니다. 모델 아키텍처를 정의할 때는 어떠한 모델이 적합한지 판단할 수 있도록 문제를 충분히 탐구해야 합니다.
# + [markdown] id="9MquXy9zLS9B" colab_type="text"
# ### *선택 토의:* `embedding_column`과 `indicator_column`의 상대적인 장단점
#
# 개념적으로, `LinearClassifier` 또는 `DNNClassifier`를 학습시킬 때 희소 열을 사용하려면 어댑터가 필요합니다. TF는 `embedding_column` 또는 `indicator_column`이라는 두 가지 옵션을 제공합니다.
#
# **작업 1**과 같이 LinearClassifier를 학습시킬 때는 내부적으로 `embedding_column`이 사용됩니다. **작업 2**에서 확인한 바와 같이 `DNNClassifier`를 학습시킬 때는 `embedding_column` 또는 `indicator_column` 중 하나를 명시적으로 선택해야 합니다. 이 섹션에서는 간단한 예를 살펴보면서 둘 사이의 차이점 및 상대적인 장단점에 대해 설명합니다.
# + [markdown] id="M_3XuZ_LLS9C" colab_type="text"
# `"great"`, `"beautiful"`, `"excellent"` 값을 포함하는 희소 데이터가 있다고 가정해 보겠습니다. 여기에서 사용하는 어휘 크기는 $V = 50$이므로 첫 번째 레이어의 각 유닛(뉴런)은 50개의 가중치를 갖습니다. 희소 입력의 단어 수는 $s$로 나타냅니다. 따라서 이 희소 데이터 예에서 $s = 3$입니다. 가능한 값이 $V$개인 입력 레이어에 대해 $d$개 유닛을 갖는 히든 레이어가 계산해야 하는 벡터 행렬 곱셈은 $(1 \times V) * (V \times d)$입니다. 연산 비용은 $O(V * d)$입니다. 이 비용은 해당 히든 레이어의 가중치 수에 비례하며 $s$와는 무관합니다.
#
# [`indicator_column`](https://www.tensorflow.org/api_docs/python/tf/feature_column/indicator_column)을 사용하여 입력에 원-핫 인코딩(길이가 $V$이고 존재하는 단어에 대해 1, 그렇지 않은 단어에 대해 0을 포함하는 부울 벡터)을 적용하면 곱셈과 덧셈의 많은 항에 0이 들어갑니다.
# + [markdown] id="I7mR4Wa2LS9C" colab_type="text"
# 크기가 $d$인 [`embedding_column`](https://www.tensorflow.org/api_docs/python/tf/feature_column/embedding_column)을 사용해도 정확히 같은 결과를 얻을 수 있으며, 다음과 같이 입력 예인 `"great"`, `"beautiful"`, `"excellent"`에 있는 3개 특성에 대한 임베딩만 조회하여 합산합니다. $(1 \times d) + (1 \times d) + (1 \times d)$. 존재하지 않는 특성에 대한 가중치는 벡터 행렬 곱셈에서 0이 곱해지므로 결과에 전혀 기여하지 않습니다. 존재하는 특성에 대한 가중치는 벡터 행렬 곱셈에서 1이 곱해집니다. 따라서 임베딩 조회를 통해 획득한 가중치를 더한 결과는 벡터 행렬 곱셈의 결과와 동일합니다.
#
# 임베딩을 사용할 때 임베딩 조회에 필요한 연산 비용은 $O(s * d)$로서, $s$가 $V$보다 훨씬 작은 경우 희소 데이터의 `indicator_column`에 대한 비용인 $O(V * d)$보다 훨씬 더 효율적입니다. 이러한 임베딩은 학습 중인 대상이므로 어떠한 학습 반복에서도 지금 조회 중인 현재 가중치가 됩니다.
# + [markdown] id="etZ9qf0kLS9D" colab_type="text"
# **작업 3**에서 확인한 것처럼, `DNNClassifier`를 학습시킬 때 `embedding_column`을 사용하면 모델에서 특성의 저차원 표현을 학습하는데, 이때 유사성 측정항목은 특성 벡터 간의 내적값으로 정의합니다. 이 예에서는 영화 리뷰라는 맥락에서 비슷하게 사용된 `"great"`, `"excellent"` 등의 단어는 임베딩 공간에서 서로 가까이 위치하므로 내적값이 크고, `"great"`와 `"bad"` 같이 상이한 단어는 임베딩 공간에서 멀리 떨어져 있으므로 내적값이 작습니다.
|
ml/cc/exercises/ko/intro_to_sparse_data_and_embeddings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.6 64-bit
# name: python3
# ---
# # Assignment is at the bottom, please head over there for evaluation.
# + [markdown] id="b5Vp-hSeu-Xp"
# # Python Tutorial
#
# ## Welcome to Python Tutorial
#
# Python has several built-in data types for storing different kinds of information in variables. Following are some commonly used data types:
#
# 1. Integer
# 2. Float
# 3. Boolean
# 4. None
# 5. String
# 6. List
# 7. Tuple
# 8. Dictionary
#
# Integer, float, boolean, None, and string are *primitive data types* because they represent a single value. Other data types like list, tuple, and dictionary are often called *data structures* or *containers* because they hold multiple pieces of data together.
# + [markdown] id="A1bA4mU5u-X4"
# ### Integer
#
# Integers represent positive or negative whole numbers, from negative infinity to infinity. Note that integers should not include decimal points. Integers have the type `int`.
# + id="vOnNGzRAu-X9"
current_year = 2021
# + id="bwkTG7isu-YB" outputId="8d5ac154-7fe6-4721-ac92-783119a89228"
type(current_year)
# + id="ZcRALJd4u-YF" outputId="fe1edd40-64f1-47ef-e93b-8b2dee93145a"
1+1
# + id="pW3vZNY-u-YH" outputId="b0e18e8a-8036-4cf8-8aaf-242964002210"
4*5
# + id="dYVZb-HZu-YJ" outputId="7ce885e1-f2ec-4723-cd4f-37774de37741"
2**10
# + id="e-N_gGfGu-YN" outputId="35669af4-da90-43b2-8a9a-e063194b9e6f"
10/2
# + id="ATMrjY7lu-YQ" outputId="797fe639-c4eb-4497-da0e-15393dcf2194"
10//2
# + id="JqMsn_Vfu-YV" outputId="66d54df4-5da4-4a8a-8427-548f7198bb12"
21%2
# + id="3a7lxpaQu-YW" outputId="ab084143-8b3c-46b8-9007-e7194cbedb28"
print("Hello students")
# + id="C5t_qMiiu-YX" outputId="b52123f2-f0a8-4499-91fb-43530faa6aab"
type()
# + [markdown] id="XuYvemkZu-YZ"
# A float is a number with a decimal place - very useful for representing things like weights or proportions.
#
# type() is the second built-in function we've seen (after print()), and it's another good one to remember. It's very useful to be able to ask Python "what kind of thing is this?"
#
# Floating point numbers can also be written using the scientific notation with an "e" to indicate the power of 10.
# + id="7exKkCf9u-Yc" outputId="7eb0e43a-be88-44b8-e8c7-caded26f869d"
one_hundredth = 2e-2
one_hundredth
# + id="AON7Bc8yu-Ye" outputId="aae84cb9-d185-4613-e665-993874d6a414"
type(one_hundredth)
# + id="t2NmbuGMu-Yf" outputId="2ef6d748-cfc7-4c31-f181-cd61f7a46c8b"
10%2
# + id="9Kzz7KZbu-Yg" outputId="e599ff77-9679-41a7-8da7-8b86a56fc03e"
10**2
# + id="43FaCUJsu-Yk"
##Check the Data types
# + id="PRh2x_Nyu-Yl" outputId="505d7c1a-dcc1-4189-e2d8-adce2b5e6322"
type(True)
# + id="hiJ5YReZu-Yo" outputId="4b9da6cc-65ac-43ef-be9d-cf2e50dbf189"
type("Hello")
# + [markdown] id="yvwwEv-mu-Yr"
# ### Strings
# + id="GxHVzI7bu-Ys" outputId="90494425-e8ba-4ecd-de59-debb67fcca29"
"Hello"
# + id="67aE3g-0u-Yu" outputId="012a97ed-5b4b-4bde-cd08-2bf36708e523"
'Lets upgrade2021'
# + id="3Gk7N4yMu-Yv" outputId="b11c055f-a4c7-49f8-9966-927c2f1e0340"
type("2021")
# + [markdown] id="E6j_4r19u-Yw"
# ### Variable Assigment
#
# Here we create a variable called say 'var' and assign it the value of 0 using =, which is called the assignment operator.
# + id="-0QJDF-Tu-Yw"
# syntax
# var_name=values
a=10
# + id="DRnqTBRdu-Yx" outputId="4afa42ba-6133-47ba-9c1b-8153b0ebbfbd"
type(a)
# + id="zeDWb0Gku-Yx"
a='Lets upgrade'
# + id="o9I45mpKu-Yz" outputId="4534c4d2-68a5-41fe-8b3e-6e97baa614fd"
type(a)
# + id="bry9UgXku-Y1" outputId="729e31bc-3155-4511-ba91-cd1524b0f061"
## Various Ways of Printing
print("Hello")
# + id="jI1OlFwCu-Y4"
first_name='Lakshmi'
last_name='Ravindran'
# + id="HSALrzqNu-Y4" outputId="275ca559-85b8-4614-9ab3-ca8a1d7f67ab"
print("My first name is {} and last name is {}".format(first_name,last_name))
# + id="rrexgaxIu-ZA" outputId="6b2dc811-5042-4ffe-9e9a-f6336ece8f23"
print("My First name is {first} and last name is {last}".format(last=last_name,first=first_name))
# + id="-wIo8GoDu-ZB" outputId="6bea2975-33ad-499b-b667-c69c6a3bedd4"
len('Lakshmi')
# + [markdown] id="osPJV6j0u-ZC"
# ## How to check for what datatype
# + id="DthkHlifu-ZD" outputId="16efdcea-9ba1-4c65-fb14-85a69a3c0027"
type(['1',2,3,4,5])
# + id="ZnceYMqou-ZE" outputId="dfbf898a-f54b-441e-b24c-74ce8a097417"
type(14.4)
# + id="HgttEc63u-ZF"
type("Hello")
# + id="aAYf4ftSu-ZF" outputId="0294743d-1871-4794-9b02-02d83243b5a4"
hat_height_cm = 25
my_height_cm = 190
# How tall am I, in meters, when wearing my hat?
total_height_meters = hat_height_cm + my_height_cm / 100
print("Height in meters =", total_height_meters, "?")
# + id="1ArxYa5Qu-ZG" outputId="902d1d87-9091-4067-9894-b353ffb5e98a"
total_height_meters = (hat_height_cm + my_height_cm) / 100
print("Height in meters =", total_height_meters)
# + id="xQpy9n7Du-ZH" outputId="a403e37a-fe3f-4618-ed82-286c105544b3"
help(round) ## help()
# + [markdown] id="rbEpJioYu-ZH"
# ### Built-in functions for working with numbers
#
# min and max return the minimum and maximum of their arguments, respectively...
# + id="yu9uiwcSu-ZI" outputId="ab1e2487-6be0-4c13-e6bb-9345ec874f94"
print(min(1, 2, 3))
print(max(1, 2, 3))
# + id="SywTjpaUu-ZK" outputId="55d6ccfb-a59f-465c-a099-efe14e7248cf"
print(abs(32)) ###abs returns the absolute value of an argument
print(abs(-32))
# + id="kbzg_lpyu-ZL"
## n addition to being the names of Python's two main numerical types,
#int and float can also be called as functions which convert their arguments to the corresponding type:
print(float(10))
print(int(3.33))
# They can even be called on strings!
print(int('807') + 1)
# + [markdown] id="v_Utvvs4u-ZN"
# The help() function is possibly the most important Python function you can learn. If you can remember how to use help(), you hold the key to understanding most other functions.
# + id="wWVxjGLSu-ZO" outputId="2dbdb706-7850-47d5-f051-9b1fbf37bb0c"
help(abs)
# + [markdown] id="TW8vTidSu-ZQ"
# ## Python Data Structures and Boolean
#
# - Boolean
# - Boolean and Logical Operators
# - Lists
# - Comparison operators
# - Dictionaries
# - Tuples
# - Sets
#
# + [markdown] id="70TnU14fu-ZR"
# ### Boolean Variables
#
# Boolean values are the two constant objects False and True.
#
# They are used to represent truth values (other values can also be considered
# false or true).
#
# In numeric contexts (for example, when used as the argument to an
# arithmetic operator), they behave like the integers 0 and 1, respectively.
#
# The built-in function bool() can be used to cast any value to a Boolean,
# if the value can be interpreted as a truth value
#
# They are written as False and True, respectively.
# + id="EYUqqKX1u-ZS" outputId="55b04e63-a72e-46ce-a583-2be15bf30326"
type(True)
# + id="MQd1LLcvu-ZT" outputId="6c55d213-568f-4160-b44b-ca7786d34323"
type(False)
# + id="KI3v_glnu-ZU"
my_str='lakshmi'
# + id="THEeLeShu-ZU" outputId="92bd4650-c246-4dfd-aee7-4b3307916b5a"
my_str.istitle()
# + id="-eKeguLXu-Za" outputId="b6a1c481-5e10-4617-a6df-8e6eeb99de76"
print(my_str.isalnum()) #check if it is alpha-numeric
print(my_str.isalpha()) #check if all char in the string are alphabetic
print(my_str.isdigit()) #test if string contains digits
print(my_str.istitle()) #test if string contains title words
print(my_str.isupper()) #test if string contains upper case
print(my_str.islower()) #test if string contains lower case
print(my_str.isspace()) #test if string contains spaces
print(my_str.endswith('i')) #test if string endswith a d
print(my_str.startswith('l')) #test if string startswith H
# + [markdown] id="OIrOCjZ2u-Zd"
# ### Boolean and Logical Operators
# + id="VSzKF3ISu-Ze" outputId="6275bde4-3e94-4eeb-d000-e18a0189a611"
True and True
# + id="JWZFPwRPu-Zh" outputId="ff002702-074d-4294-a3c3-4c250dff3e8a"
True and False
# + id="j7Ae_9Euu-Zi" outputId="eac27d73-78b8-4a0d-9d95-90c03048dbc3"
True or False
# + id="BVL7EdNTu-Zj" outputId="2f94ae62-e029-4269-c541-77a169fe6099"
True or True
# + [markdown] id="J7QOqZJWu-Zj"
# ### Lists
#
# A list is a data structure in Python that is a mutable, or changeable, ordered sequence of elements. Each element or value that is inside of a list is called an item. Just as strings are defined as characters between quotes, lists are defined by having values between square brackets [ ]
# + id="8U_e0EMLu-Zl" outputId="f4b77f10-ffc2-4e87-f382-aaf9cceb7656"
type([1,2,3,5,'python'])
# + id="ozG91ql5u-Zm"
lst=list()
# + id="-Gk28UFJu-Zm" outputId="01d9f6e0-128d-4480-bd81-6b14243dff84"
type(lst)
# + id="FZHN2NUau-Zn"
lst = ['Python', 'Java', 20,30, 40,50]
# + id="w3Yes0Wxu-Zn" outputId="bfc8f8d0-18de-49f7-c14a-e63dd35bb6a4"
len(lst)
# + id="EINfFf8Uu-Zo"
lst.append("Perl") ## Append
# + id="6uDlt2Qsu-Zp" outputId="8bb172d9-331d-4752-c70a-4a741db1631b"
lst
# + id="DdBgppj8u-Zr" outputId="8af698fe-bca9-4431-a8c3-1c88448c5672"
lst[6] ###Indexing in list
# + id="S-5YF0LDu-Zu" outputId="32971114-c02c-4f55-cab9-bdc96c053de1"
lst[2:6]
# + id="sHN8xyRzu-Zw"
lst.append(["Arthi", "Lakshmi"])
# + id="R0Fbev4hu-Zx" outputId="bba7d30e-05e8-4b82-dae3-68abb456fffc"
lst ##nested list
# + id="htP05XhEu-Zy" outputId="68d9d939-bac8-4047-b9f0-aabcd27396cc"
lst[7]
# + id="YiiufCxgu-Zy"
lst.insert(2, "gcp")
# + id="DB-3Gakfu-Zz" outputId="c757efca-f9a6-4ee2-f920-51a0c575dfc1"
lst
# + [markdown] id="oMfDVUP3u-Zz"
#
# ### List vs Set
#
# 1. The List is an ordered sequence. 1. The Set is an unordered sequence.
# 2. List allows duplicate elements 2. Set doesn’t allow duplicate elements.
# 3. Elements by their position can be accessed. 3. Position access to elements is not allowed.
# + [markdown] id="kAhDKqawu-Z0"
# ## Tuples
#
#
# A tuple is an ordered collection of values, similar to a list. However, it is not possible to add, remove, or modify values in a tuple. A tuple is created by enclosing values within parentheses `(` and `)`, separated by commas.
#
# > Any data structure that cannot be modified after creation is called *immutable*. You can think of tuples as immutable lists.
#
# Let's try some experiments with tuples.
# + id="QXAtnBrFu-Z0" outputId="e802ace6-9e9e-49ef-e8cc-cf7e72392a9a"
#Tuples are just immutable lists. Use () instead of []
x = (1, 2, 3)
len(x)
# + id="NVKOLWkVu-Z1" outputId="ea54061c-360f-4644-f1a1-0183ec9ad501"
y = (4, 5, 6)
y[2]
# + id="Uj3KrhWmu-Z1"
fruits = ('apple', 'cherry', 'dates')
# + id="tkKsd5_4u-Z2" outputId="2a2a0bb4-c96b-4759-ca27-39c991dc0567"
# check no. of elements
len(fruits)
# + id="BGchpVgTu-Z2" outputId="96a100b6-c429-4694-f936-89629a4525ed"
# get an element (positive index)
fruits[0]
# + id="RAqUhI8iu-Z4" outputId="70ba5ac0-81f4-428b-a09c-d4584a3ba74e"
# get an element (negative index)
fruits[-2]
# + id="FOHu3sdBu-Z5" outputId="5a663daa-7b58-46d0-d7f1-b471b024f636"
# check if it contains an element
'dates' in fruits
# + id="yCbA_Rvju-Z5" outputId="2277ddb5-4031-4d6b-ee6f-8b6336ed738f"
# try to change an element
fruits[0] = 'avocado'
# + id="JWPfePdru-Z6" outputId="e4833505-d5d2-4b93-b815-212bc03ee4e6"
# try to append an element
fruits.append('blueberry')
# + id="riKajceju-Z7" outputId="65640742-d334-4c94-ffac-42567a2cc8b6"
# try to remove an element
fruits.remove('apple')
# + [markdown] id="aM3Xe4LJu-Z8"
# You can also skip the parantheses `(` and `)` while creating a tuple. Python automatically converts comma-separated values into a tuple.
# + [markdown] id="iKrVvLqPu-Z8"
# # Extend
# + id="9XddZyEbu-Z9"
lst = [1,2,3,4,5,6]
# + id="RBy3aqCZu-Z-"
lst.extend([8,9])
# + id="LqRdQyMVu-Z-" outputId="003b0b2e-e7c5-4190-def2-433d4a1924bf"
lst
# + id="2XMQeHAvu-Z_" outputId="5bbad95d-ca6d-4dd6-95a7-f97a1189e313"
sum(lst) ###sum operation in list
# + id="WaVrvktku-aA" outputId="04adb3aa-9945-442c-ff31-b1dad348284a"
lst*2
# + id="-QH5ngnHu-aE" outputId="729548aa-6ae3-4552-fe03-b92d4f4175c6"
lst.pop()
# + id="RwJ4cwmyu-aF" outputId="afc878f5-60b7-49cb-a1d9-f009790a873d"
lst
# + id="QUCXQT0Uu-aF" outputId="b1d73a95-b395-4578-d7ed-473180344825"
lst.pop(0)
# + id="pE6SxZk8u-aG" outputId="fdbbda0f-8a65-4bfd-c155-c620526513d0"
lst
# + [markdown] id="V0HHYs8Yu-aG"
# ### count():Calculates total occurrence of given element of List
# + id="VWmaRnqRu-aG" outputId="9194c11e-9677-4994-d317-a8930be0bafd"
lst=[1,1,2,3,4,5]
lst.count(1)
# + id="-fvTQ-itu-aH" outputId="5a943b89-dd27-4f6b-fbd7-87437d125fbe"
#length:Calculates total length of List
len(lst)
# + id="LqAxwrELu-aI" outputId="0d984725-9903-4091-f064-9176ac09ff9c"
# index(): Returns the index of first occurrence. Start and End index are not necessary parameters
#lst.index(1)
lst.index(1,1,4)
# + id="HB1myA6Nu-aI"
min(lst)
# + id="IA7ltRSfu-aJ"
max(lst)
# + [markdown] id="Ov_qQqu6u-aJ"
# ## SETS
#
# A Set is an unordered collection data type that is iterable, mutable, and has no duplicate elements. Python's set class represents the mathematical notion of a set.This is based on a data structure known as a hash table
# + id="y_28lwZ8u-aJ" outputId="82dd123d-7bf2-46dd-c9d7-6d14351eef18"
## Defining an empty set
set_var= set()
print(set_var)
print(type(set_var))
# + id="4ff0ICOOu-aK"
set_var={1,2,3,4,3,1,2,2,3,4,4,1}
# + id="d8-0GrD6u-aK" outputId="8721af67-fb67-4c34-c40c-fde37a39f8ff"
set_var
# + id="rfnzoNipu-aL" outputId="2d48161f-3bd3-4968-81c3-17ccb3298798"
set_var={"Apple","Mango",'Guava'}
print(set_var)
type(set_var)
# + id="sQa2WK3au-aL"
##indexing
set_var.add("avocado")
# + id="kiEW6W-Mu-aM" outputId="5c7733ad-dc30-475e-e9e7-b464c311cda0"
set_var
# + [markdown] id="wZIC2qKau-aM"
# ### Assignment
#
# + id="gUIUGHs1u-aN"
a=10
b=20
#print(a*b)
#print(a/b)
#print(a%b)
#print((a*b)+(a/b)) ##BODMAS
# + id="I09xYzhPu-aN"
#Write a program which accept principle, rate and time from user and print the simple interest. The formula to calculate simple interest is: simple interest = principle x rate x time / 100 Solution
# + id="eDKxLBviu-aO"
principle = float(input("Enter Principle : "))
rate = float(input("Enter Rate : "))
time = float(input("Enter Time : "))
si = (principle*rate*time)/100
print("Simple Interest : ",si)
# run for principle = 1800 , rate = 12.5 , time = 12
|
Python for Data Science Zero to Hero/Day-1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Design MERFISH probes using the example inputs from <NAME>. The original MATLAB design pipeline can be found at https://github.com/ZhuangLab/MERFISH_analysis.
# # Download the input data
# # This is for the UNIX-like operating systems. If you are using Windows, just download the files accordingly.
# # !mkdir temporary_data
# # !wget http://zhuang.harvard.edu/merfish/MERFISHData/MERFISH_Examples2.zip -O temporary_data/MERFISH_Examples2.zip
# # !unzip -o temporary_data/MERFISH_Examples2.zip -d temporary_data
# # Make a path for output
# # !mkdir temporary_data/MERFISH_Examples2/outputs
# +
# Define all the input files you need in this script
import os
base_folder = r'\\10.245.74.212\Chromatin_NAS_2\Chromatin_Libraries'
rpkm_filename = os.path.join(base_folder, r'CTP-10_Aire\MERFISH_designer\Aire_smFISH\design\E-MTAB-6081.processed.3\mouse_rpkm.txt')
pool_folder = os.path.join(base_folder, 'CTP-10_Aire')
ref_folder = os.path.join(base_folder, r'CTP-10_Aire\MERFISH_designer\Aire_smFISH')
output_folder = os.path.join(ref_folder, 'outputs')
if not os.path.exists(output_folder):
os.makedirs(output_folder)
codebook_file = os.path.join(ref_folder, 'design', r'codebook.csv')
genome_folder = os.path.join(base_folder, r'Genomes\mouse\GRCm39_ensembl')
transcripts_fasta_file = os.path.join(genome_folder, r'Mus_musculus.GRCm39.cdna.all.fa')
ncRNA_file = os.path.join(genome_folder, r'Mus_musculus.GRCm39.ncrna.fa')
fpkm_tracking_file = os.path.join(ref_folder, r'aire_wt_mtec_genes_ensembl.fpkm_tracking')
readout_fasta_file = os.path.join(ref_folder, r'readouts.fasta')
forward_primer_file = os.path.join(base_folder, r'Primers\forward_primers.fasta')
reverse_primer_file = os.path.join(base_folder, r'Primers\reverse_primers.fasta')
# Define the output files
ottable_transcriptome_file = os.path.join(output_folder, r'ottable_transcriptome.pkl')
selected_primers_file = os.path.join(output_folder, r'selected_primers.csv')
probe_output_file = os.path.join(output_folder, r'designed_probes.csv')
transcript_level_report_file = os.path.join(output_folder, r'transcript_level_report.csv')
# -
print(transcripts_fasta_file)
# # Initialize data structures
# +
# Import the modules
import os
import sys
import pandas as pd
from IPython.display import display
import MERFISH_probe_design
import MERFISH_probe_design.IO.file_io as fio
import MERFISH_probe_design.probe_design.probe_dict as p_d
import MERFISH_probe_design.probe_design.OTTable_dict as ot
import MERFISH_probe_design.probe_design.readout_sequences as rs
import MERFISH_probe_design.probe_design.probe_selection as ps
import MERFISH_probe_design.probe_design.quality_check as qc
from MERFISH_probe_design.probe_design import filters
from MERFISH_probe_design.probe_design import plot
from MERFISH_probe_design.probe_design import primer_design
# -
# ## load transcriptome
from importlib import reload
reload(fio)
reload(qc)
# Load the transcriptome as a pandas data frame
transcriptome = fio.load_transcriptome(transcripts_fasta_file, None)
def find_gene_id_in_description(description_string):
import re
return re.search(r'gene:([A-Za-z0-9\.]+)', description_string)[0].split('gene:')[1]
def find_gene_name_in_description(description_string):
import re
gene_matches = re.search(r'gene_symbol:([A-Za-z0-9\.]+)', description_string)
if gene_matches is None:
print(description_string)
return float('nan')
else:
return gene_matches.group(0).split('gene_symbol:')[1]
transcriptome['gene_id'] = list(map(find_gene_id_in_description, transcriptome['description']))
transcriptome['gene_short_name'] = list(map(find_gene_name_in_description, transcriptome['description']))
# Make sure that the transcriptome data frame has the standard column names.
# The standard columns are: transcript_id, sequence, gene_id, gene_short_name and FPKM.
# Also remove the non-standard columns for clarity.
transcriptome = qc.check_and_standardize_transcriptome(transcriptome, remove_non_standard_columns=True)
transcriptome # Let's have a look at what's inside the transcriptome
# check wether this name exist
print(transcriptome['gene_id'][transcriptome['gene_short_name']=='Krt5'])
print(transcriptome['gene_id'][transcriptome['gene_short_name']=='Aire'])
# +
# Generate a codebook table for the target transcript
# Choose one of the full length protein coding transcript as the target
bit_names = ['RS0015', 'RS0083', 'RS0095', 'RS0109']
barcode_table = pd.DataFrame({'name':['Aire', 'Krt5'],
#'id':['ENSMUSG00000061527.8', 'ENSMUSG00000000731.16'],
'id':['ENSMUST00000023709.7', 'ENSMUST00000145975.8'],
'barcode_str':['1100','0011']})
barcode_table
# -
# Initialize the probe dictionary which is the carrier of the probes throught the design process.
probe_dict = p_d.init_probe_dict(['Aire', 'Krt5'], transcriptome,
'gene_short_name', K=30)
p_d.print_probe_dict(probe_dict)
# Select the transcripts that we want to target
# The target transcripts are already defined in the codebook
probe_dict = p_d.select_transcripts_by_ids(probe_dict,
transcript_ids=['ENSMUST00000023709.7', 'ENSMUST00000145975.8'])
p_d.print_probe_dict(probe_dict) # We excluded all the transcripts that are not our direct targets
# Initialize the off-target counting tables
# OTTable for rRNA/tRNAs
ncRNAs = fio.load_fasta_into_df(ncRNA_file)
ottable_rtRNAs = ot.get_OTTable_for_rtRNAs(ncRNAs, 15)
# OTTables for the genes we target
gene_ottable_dict = ot.get_gene_OTTables(transcriptome, ['Aire', 'Krt5'], 'gene_short_name', 17)
# %%time
# OTTable for the transcriptome.
ottable_transcriptome = ot.get_OTTable_for_transcriptome(transcriptome, 17)
# # Select target regions
# Calculate and plot the GC contents of the target regions
filters.calc_gc_for_probe_dict(probe_dict, column_key_seq='target_sequence', column_key_write='target_GC')
plot.plot_hist(probe_dict, column_key='target_GC')
# Filter GC cotent and plot the GC content after filtering
filters.filter_probe_dict_by_metric(probe_dict, 'target_GC', lower_bound=43, upper_bound=73)
plot.plot_hist(probe_dict, column_key='target_GC')
# Calculate and plot the melting-temperatures (Tm)
filters.calc_tm_for_probe_dict(probe_dict, Na_conc=390, fmd_percentile=30, probe_conc=5,
column_key_seq='target_sequence', column_key_write='target_Tm')
plot.plot_hist(probe_dict, column_key='target_Tm')
# Filter by Tm
# NOTE: here we used a higher upper bound for GC content and Tm than JM's original
# cutoffs. It was shown that higher Tm gives better signal-to-noise ratios in SM-FISH
filters.filter_probe_dict_by_metric(probe_dict, 'target_Tm', lower_bound=52)
plot.plot_hist(probe_dict, column_key='target_Tm')
# Calculate and plot the off-targets to rRNA/tRNAs
ot.calc_OTs(probe_dict, ottable_rtRNAs, 'target_sequence', 'target_OT_rtRNA', 15)
plot.plot_hist(probe_dict, 'target_OT_rtRNA', y_max=1000)
# Filter out probes that have any rRNA/tRNA off-targets
filters.filter_probe_dict_by_metric(probe_dict, 'target_OT_rtRNA', upper_bound=0.5)
plot.plot_hist(probe_dict, 'target_OT_rtRNA')
# +
# Get the FPKMs of the transcripts
transcript_fpkms = dict(zip(list(transcriptome['transcript_id']), list(transcriptome['FPKM'])))
# Calculate the specificities and isoform specificities of the target regions
ot.calc_specificity(probe_dict, ottable_transcriptome, gene_ottable_dict, transcript_fpkms,
'target_sequence', 'target_specificity', 'target_isospecificity', 17)
plot.plot_hist(probe_dict, 'target_specificity')
plot.plot_hist(probe_dict, 'target_isospecificity')
# +
# Filter the specificities of the target regions
# Here we allow the probes to target other isoforms
# Don't allow any gene-level off-targets
filters.filter_probe_dict_by_metric(probe_dict, 'target_specificity', lower_bound=0.99999)
#filters.filter_probe_dict_by_metric(probe_dict, 'target_isospecificity', upper_bound=0.1)
plot.plot_hist(probe_dict, 'target_specificity')
plot.plot_hist(probe_dict, 'target_isospecificity')
# -
# # Design readout sequences
# Load the readout sequences into a data frame
readout_seqs = fio.load_fasta_into_df(readout_fasta_file)
rs.append_on_bit_ids_to_readout_sequences(readout_seqs, bit_names)
readout_seqs
# +
import importlib
importlib.reload(rs)
# Add the readout sequences. Here we randomly add 2 readout sequences to each probe.
# Add an "T" between the concatenated sequences.
# Force each probe to have the same readout sequence
rs.add_readout_seqs_to_probes_random(probe_dict, readout_seqs, barcode_table, 2,
spacer='A', gene_id_key='name', each_probe_1_on_bit=True)
# -
probe_dict['Aire'].keys()
# probe_dict['Krt5']['ENSMUST00000023709.7']
# Filter out probes that have off-targets to rRNA/tRNAs
ot.calc_OTs(probe_dict, ottable_rtRNAs, 'target_readout_sequence', 'target_readout_OT_rtRNA', 15)
plot.plot_hist(probe_dict, 'target_readout_OT_rtRNA', y_max=1000)
filters.filter_probe_dict_by_metric(probe_dict, 'target_readout_OT_rtRNA', upper_bound=0.5)
plot.plot_hist(probe_dict, 'target_readout_OT_rtRNA')
# Calculate how many more off-targets to the transcriptome are introduced due to the readout sequences.
# The off-target counts are weighted down by the FPKMs of the on-target transcripts
ot.calc_OT_diffs(probe_dict, ottable_transcriptome, gene_ottable_dict, transcript_fpkms,
'target_sequence', 'target_readout_sequence', 'readout_OT_increase', 17)
plot.plot_hist(probe_dict, 'readout_OT_increase', y_max=2000)
import numpy as np
np.unique(probe_dict['Aire']['ENSMUST00000145975.8']['probe_barcode'], return_counts=True)
# Filter out the probes with extra off-targets due to the readouts
# Require the new weighted off-targets to be minor compared to the on-target weight.
filters.filter_probe_dict_by_metric(probe_dict, 'readout_OT_increase', upper_bound=0.5)
plot.plot_hist(probe_dict, 'readout_OT_increase')
np.unique(probe_dict['Aire']['ENSMUST00000145975.8']['probe_barcode'], return_counts=True)
# # Select probes
# %%time
# Select probes by a stochastic greedy algorithms that optimizes the on-bit coverage
# and minimizes the overlapping between probes.
ps.select_probes_greedy_stochastic(probe_dict, N_probes_per_transcript=48,
N_on_bits=2)
# Let's plot the probe coverage of an example transcript
seq_len = len(transcriptome[transcriptome['transcript_id'] == 'ENSMUST00000145975.8'].iloc[0]['sequence'])
plot.plot_sequence_coverage(probe_dict['Aire']['ENSMUST00000145975.8'], seq_len)
# Let's plot the probe coverage of an example transcript
seq_len = len(transcriptome[transcriptome['transcript_id'] == 'ENSMUST00000023709.7'].iloc[0]['sequence'])
plot.plot_sequence_coverage(probe_dict['Krt5']['ENSMUST00000023709.7'], seq_len)
probe_dict['Aire']['ENSMUST00000145975.8']
probe_dict['Krt5']['ENSMUST00000023709.7']
# # Quality check
# +
# %%time
# Filter out the probes that self complement or complement with other probes.
# Iterately remove the probes with high numbers of cis/trans-complementarity
# This filtering strategy is a compromise between speed and the number of probes to keep
while True:
# Make a OTTable from the reverse-complement sequences of the probes.
ottable_probes_rc = ot.get_OTTable_for_probe_dictionary(probe_dict, 'target_readout_sequence', 15, rc=True)
# The off-targets in this table indicates cis/trans-complementarity
ot.calc_OTs(probe_dict, ottable_probes_rc, 'target_readout_sequence', 'probe_cis_trans_OT', 15)
max_ot = max(plot.get_values_from_probe_dict(probe_dict, 'probe_cis_trans_OT'))
if max_ot == 0:
break
# Remove probes that have any cis/trans-complementarity
filters.filter_probe_dict_by_metric(probe_dict, 'probe_cis_trans_OT', upper_bound=max_ot - 0.5)
plot.plot_hist(probe_dict, 'probe_cis_trans_OT')
# -
# +
# Get the reverse-complementary sequences of the designed probes
p_d.get_rc_sequences(probe_dict, 'target_readout_sequence', 'target_readout_sequence_rc')
# Get a data frame of all probes
df_all_probes = p_d.probe_dict_to_df(probe_dict).sort_values(['probe_barcode', 'shift'])
# Make a column of probe names
probe_names = np.array(df_all_probes['gene_id']) + '_' + np.array(df_all_probes['transcript_id'])
probe_names = probe_names + '_shift:' + np.array(df_all_probes['shift']).astype(str)
probe_names = probe_names + '_GC:' + np.array(df_all_probes['target_GC']).astype('|S4').astype(str)
probe_names = probe_names + '_Tm:' + np.array(df_all_probes['target_Tm']).astype('|S4').astype(str)
probe_names = probe_names + '_specificity:' + np.array(df_all_probes['target_specificity']).astype('|S1').astype(str)
probe_names = probe_names + '_isospecificity:' + np.array(df_all_probes['target_isospecificity']).astype('|S4').astype(str)
probe_names = probe_names + '_' + np.array(df_all_probes['readout_names'])
df_all_probes['probe_names'] = probe_names
# Write the designed probes
df_all_probes.to_csv(probe_output_file, index=False)
# -
# generate IDT list
well_positions = []
for _col in 'ABCDEFGH':
for _row in range(12):
well_positions.append(f"{_col}{_row+1}")
# +
# export IDT format
# Get the reverse-complementary sequences of the designed probes
p_d.get_rc_sequences(probe_dict, 'target_readout_sequence', 'target_readout_sequence_rc')
# Get a data frame of all probes
idt_plate_df = pd.DataFrame({
'Well Position': well_positions,
'Name': df_all_probes['probe_names'][:96],
'Sequence': [_seq for _seq in df_all_probes['target_readout_sequence_rc'][:96]],
})
# Write the designed probes
idt_probe_output_file = os.path.join(output_folder, r'idt_order_new.xlsx')
idt_plate_df.to_excel(idt_probe_output_file, index=False)
# -
print(output_folder)
idt_df
|
Library_design/CTP-10_Aire/Merfish_design_smFISH_Aire.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="gfcHVOxHeNLI"
# ### Creation of the environment
# + id="2h1MRzBLtex2" colab={"base_uri": "https://localhost:8080/"} outputId="e81a6e74-2e57-49d2-9156-12a3871504a0"
## RUN THIS CELL
# %tensorflow_version 2.x
# !pip3 install --upgrade pip
# #!pip install -qU t5
# !pip3 install git+https://github.com/google-research/text-to-text-transfer-transformer.git #extra_id_x support
import functools
import os
import time
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
import tensorflow.compat.v1 as tf
import tensorflow_datasets as tfds
import t5
#Set the base dir(Google cloud bucket)
BASE_DIR = "gs://bucket_code_completion"
if not BASE_DIR or BASE_DIR == "gs://":
raise ValueError("You must enter a BASE_DIR.")
ON_CLOUD = True
if ON_CLOUD:
import tensorflow_gcs_config
from google.colab import auth
# Set credentials for GCS reading/writing from Colab and TPU.
TPU_TOPOLOGY = "2x2"
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection
TPU_ADDRESS = tpu.get_master()
print('Running on TPU:', TPU_ADDRESS)
except ValueError:
raise BaseException('ERROR: Not connected to a TPU runtime; please see the previous cell in this notebook for instructions!')
auth.authenticate_user()
tf.config.experimental_connect_to_host(TPU_ADDRESS)
tensorflow_gcs_config.configure_gcs_from_colab_auth()
tf.disable_v2_behavior()
# Improve logging.
from contextlib import contextmanager
import logging as py_logging
if ON_CLOUD:
tf.get_logger().propagate = False
py_logging.root.setLevel('INFO')
@contextmanager
def tf_verbosity_level(level):
og_level = tf.logging.get_verbosity()
tf.logging.set_verbosity(level)
yield
tf.logging.set_verbosity(og_level)
# + [markdown] id="mOcMhMileZQp"
# ### Loading of tsv files
# With this script you can load each tsv file for finetuning.
# Please be sure that the path to all tsv files are correct.
# For running the evaluation on a **single model** on a specific dataset, load only the tsv file you're interested in (e.g. *java_construct*)
# + id="glLJUm1dxIiH"
#Validation(train and test on the same dataset)
nq_tsv_path_java_construct = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_java_construct.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_java_construct.tsv',
}
num_nq_examples_java_construct = dict(train=750000, validation=106237)
# + id="_-B3_th9eP5y"
#Validation(train and test on the same dataset)
nq_tsv_path_android_construct = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_android_construct.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_android_construct.tsv',
}
num_nq_examples_android_construct = dict(train=750000, validation=100536)
# + id="6u7JbyjV8GN3"
#Validation(train and test on the same dataset)
nq_tsv_path_java_block = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_java_block.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_java_block.tsv',
}
num_nq_examples_java_block = dict(train=298470, validation=40008)
# + id="bmUMU-Pg8HVm"
#Validation(train and test on the same dataset)
nq_tsv_path_android_block = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_android_block.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_android_block.tsv',
}
num_nq_examples_android_block = dict(train=204580, validation=26978)
# + id="E5lRmNWG8HuD"
#Validation(train and test on the same dataset)
nq_tsv_path_java_token = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_java_token.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_java_token.tsv',
}
num_nq_examples_java_token = dict(train=750000, validation=219486)
# + id="QF9DHJxc8IGe"
#Validation(train and test on the same dataset)
nq_tsv_path_android_token = {
"train": 'gs://bucket_code_completion/T5_extension/ft_datasets/train_android_token.tsv',
"validation": 'gs://bucket_code_completion/T5_extension/ft_datasets/test_android_token.tsv',
}
num_nq_examples_android_token = dict(train=750000, validation=200504)
# + [markdown] id="Vx4k3GpDfNlv"
# ### Preprocess of the dataset
# In this step we preprocess the dataset.
# You have to change the path to vocab files (*vocab_model_path* and *vocab_path*)
# We're going to preprocess all the tsv file so that T5 can use them for finetuning.
# Please be sure to run **only the cells related to the specific model** you want to evaluate. Run the following cell and then only the group of cell related to the model (e.g. all the cell under JAVA_CONSTRUCT module)
# + id="PobLvzL18zzR"
## RUN THIS CELL
from t5.data import postprocessors as t5_postprocessors
from t5.seqio import Feature,SentencePieceVocabulary
# # Set the path of sentencepiece model and vocab files
# # Must be the same used for the pre-trained phase
vocab_model_path = 'gs://bucket_code_completion/T5_extension/code.model'
vocab_path = 'gs://bucket_code_completion/T5_extension/code.vocab'
TaskRegistry = t5.data.TaskRegistry
TfdsTask = t5.data.TfdsTask
def get_default_vocabulary():
return SentencePieceVocabulary(vocab_model_path, 100)
DEFAULT_OUTPUT_FEATURES = {
"inputs": Feature(
vocabulary=get_default_vocabulary(), add_eos=True, required=False),
"targets": Feature(
vocabulary=get_default_vocabulary(), add_eos=True)
}
# + [markdown] id="mn-DMH5FkSO2"
# JAVA CONSTRUCT
# + id="K0NTLbyXvkCs" colab={"base_uri": "https://localhost:8080/"} outputId="2b01335b-4691-4747-f345-7dbd4f83459a"
def nq_java_construct(split, shuffle_files=True):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_java_construct[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw train examples...")
for ex in tfds.as_numpy(nq_java_construct("train").take(5)):
print(ex)
# + id="4bJZPQgjxKZ1"
def java_construct_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['JAVA_CONSTRUCT:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + id="h3jAg8Zhx_Ep" colab={"base_uri": "https://localhost:8080/"} outputId="38775783-9f57-489c-f0fa-c43ae5facf1e"
t5.data.TaskRegistry.remove('java_construct')
t5.data.TaskRegistry.add(
"java_construct",
dataset_fn=nq_java_construct,
splits=["train", "validation"],
text_preprocessor=[java_construct_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_java_construct
)
# + id="e71p9JIFyYHm" colab={"base_uri": "https://localhost:8080/"} outputId="806ddc14-7709-43a9-d3e5-81ca028bd267"
nq_task = t5.data.TaskRegistry.get("java_construct")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + [markdown] id="yB-KY403kcCn"
# JAVA TOKEN
# + id="UNi7HPiOz27q" colab={"base_uri": "https://localhost:8080/"} outputId="6e3ad391-3f00-421a-e574-019bfd1c5d3d"
def nq_java_token(split, shuffle_files=False):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_java_token[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw valid examples...")
for ex in tfds.as_numpy(nq_java_token("validation").take(5)):
print(ex)
# + id="VvDAbgNY0B4Y"
def java_token_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['JAVA_TOKEN:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + id="-Mm6AQfw0INC" colab={"base_uri": "https://localhost:8080/"} outputId="604261bd-28ed-45b0-e500-79f750f67398"
t5.data.TaskRegistry.remove('java_token')
t5.data.TaskRegistry.add(
"java_token",
dataset_fn=nq_java_token,
splits=["train", "validation"],
text_preprocessor=[java_token_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_java_token
)
# + id="qnf25qt10Wkl" colab={"base_uri": "https://localhost:8080/"} outputId="de050ff6-6fa3-48b1-a538-0eeacce501ef"
nq_task = t5.data.TaskRegistry.get("java_token")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + [markdown] id="ZIe-u5l9ke6x"
# JAVA BLOCK
# + colab={"base_uri": "https://localhost:8080/"} id="yr0TT18ejMtY" outputId="c4cd2d00-28a5-499d-bca7-01c39356b3e9"
def nq_java_block(split, shuffle_files=False):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_java_block[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw valid examples...")
for ex in tfds.as_numpy(nq_java_block("validation").take(5)):
print(ex)
# + id="Hq0uLYNTjM9z"
def java_block_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['JAVA_BLOCK:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="ji4u8yhqjNER" outputId="f5aac828-6b32-4e46-a78e-b1dc86dcf0a5"
t5.data.TaskRegistry.remove('java_block')
t5.data.TaskRegistry.add(
"java_block",
dataset_fn=nq_java_block,
splits=["train", "validation"],
text_preprocessor=[java_block_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_java_block
)
# + colab={"base_uri": "https://localhost:8080/"} id="JG09lDZdjNKr" outputId="02526c96-8421-457e-c187-bc1ab6e84891"
nq_task = t5.data.TaskRegistry.get("java_block")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + [markdown] id="lz_Gxq_4khQt"
# ANDROID CONSTRUCT
# + colab={"base_uri": "https://localhost:8080/"} id="LNwnQAMVjNdy" outputId="8707493e-ecd8-45bc-bda7-8979e23afe16"
def nq_android_construct(split, shuffle_files=True):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_android_construct[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw train examples...")
for ex in tfds.as_numpy(nq_android_construct("train").take(5)):
print(ex)
# + id="5sBEViP5jNja"
def android_construct_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['ANDROID_CONSTRUCT:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="lP_VXjC6jNpp" outputId="07b4e96e-da9a-4f17-ed59-6525e3d1e6a3"
t5.data.TaskRegistry.remove('android_construct')
t5.data.TaskRegistry.add(
"android_construct",
dataset_fn=nq_android_construct,
splits=["train", "validation"],
text_preprocessor=[android_construct_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_android_construct
)
# + colab={"base_uri": "https://localhost:8080/"} id="DOWFYL7KjNwd" outputId="f386b70f-5b6e-4336-eb54-fff919f3806e"
nq_task = t5.data.TaskRegistry.get("android_construct")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + [markdown] id="ZZ9H1D-KngJy"
# ANDROID TOKEN
# + colab={"base_uri": "https://localhost:8080/"} id="Wgi_2yd-nb4Y" outputId="8c9f9b78-a438-4b83-c25f-460c346b1106"
def nq_android_token(split, shuffle_files=False):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_android_token[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw valid examples...")
for ex in tfds.as_numpy(nq_android_token("validation").take(5)):
print(ex)
# + id="WcKSDpJrnb_X"
def android_token_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['ANDROID_TOKEN:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="yvwEpdZIjN_9" outputId="acd5b454-b0d6-495f-beb3-e20611634809"
t5.data.TaskRegistry.remove('android_token')
t5.data.TaskRegistry.add(
"android_token",
dataset_fn=nq_android_token,
splits=["train", "validation"],
text_preprocessor=[android_token_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_android_token
)
# + colab={"base_uri": "https://localhost:8080/"} id="V6z53ZuUpT3f" outputId="ea9e843b-d357-4f34-bc0c-569286b4b72b"
nq_task = t5.data.TaskRegistry.get("android_token")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + [markdown] id="7q3xYhUwoBC6"
# ANDROID BLOCK
# + colab={"base_uri": "https://localhost:8080/"} id="SqvJtXAKjOFn" outputId="4ec93c6c-26af-4652-c2cf-b60bf1126c0c"
def nq_android_block(split, shuffle_files=False):
# We only have one file for each split.
del shuffle_files
# Load lines from the text file as examples.
ds = tf.data.TextLineDataset(nq_tsv_path_android_block[split])
ds = ds.map(
functools.partial(tf.io.decode_csv, record_defaults=["string","string"],
field_delim="\t", use_quote_delim=False),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds = ds.map(lambda *ex: dict(zip(["input", "output"], ex)))
return ds
print("A few raw valid examples...")
for ex in tfds.as_numpy(nq_android_block("validation").take(5)):
print(ex)
# + id="yvADtjcHoGCM"
def android_block_preprocessing(ds):
def to_inputs_and_targets(ex):
inputs = tf.strings.join(['ANDROID_BLOCK:' + ex['input']], separator=' ')
class_label = tf.strings.join([ex['output']], separator=' ')
return {'inputs': inputs, 'targets': class_label }
return ds.map(to_inputs_and_targets,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="UY6Cr6OyoGLR" outputId="45f3fe92-a51f-44e9-f123-1b109a288b28"
t5.data.TaskRegistry.remove('android_block')
t5.data.TaskRegistry.add(
"android_block",
dataset_fn=nq_android_block,
splits=["train", "validation"],
text_preprocessor=[android_block_preprocessing],
output_features = DEFAULT_OUTPUT_FEATURES,
metric_fns=[t5.evaluation.metrics.accuracy],
num_input_examples=num_nq_examples_android_block
)
# + colab={"base_uri": "https://localhost:8080/"} id="ZIpCwr6FjOMd" outputId="33fd05e3-1be7-49dd-f23a-d7003939278c"
nq_task = t5.data.TaskRegistry.get("android_block")
ds = nq_task.get_dataset(split="train", sequence_length={"inputs": 256, "targets": 256})
print("A few preprocessed training examples...")
for ex in tfds.as_numpy(ds.take(5)):
print(ex)
# + id="H0chadICjOT_"
# + [markdown] id="DCALtxexH2QO"
# ### Evaluation
# You can run the evaluation using the following cells.
# Please set the correct path of the variable *MODEL_DIR* (the path to save the pretrained model in)
#
# Change the mixture chosing the one you want to run (e.g. you can associate "all_tasks" to ["android token"] if you want to train android token)
#
# Please be sure to run only the cell under the specific model you want to train (e.g. all cells under **ANDROID TOKEN** section)
# + id="cz1a1TxFNKmx" colab={"base_uri": "https://localhost:8080/"} outputId="7c0ed97d-4bb2-49fb-ce76-643e02d7fe06"
## RUN THIS CELL
def _rate_num_input_examples(task):
if "train" in task.splits:
return float(task.num_input_examples("train"))
elif "validation" in task.splits:
return float(task.num_input_examples("validation"))
else:
raise ValueError("Task %s does not have a train or validation split." % (task.name))
t5.data.MixtureRegistry.remove("all_tasks")
t5.data.MixtureRegistry.add(
"all_tasks",
# ["java_construct", "java_token", "java_block", "android_construct", "android_token", "android_block"],
["java_construct"],
default_rate=_rate_num_input_examples
#default_rate=1.0
)
# + [markdown] id="WuMVmcwyvoAJ"
# JAVA CONSTRUCT
# + id="_3Qx699vN302" colab={"base_uri": "https://localhost:8080/"} outputId="427d781f-89fe-48ef-f05f-98564d327b5e"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/java_construct/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/java_construct_inputs.txt',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/java_construct_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + [markdown] id="iXmVMItzvuB8"
# ANDROID CONSTRUCT
# + id="DayFX1_0vs55"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/android_construct/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/android_construct_inputs.dms',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/android_construct_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + [markdown] id="bOerAopHxtOJ"
# JAVA BLOCK
# + id="CxyWWiLnvs9L"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/java_block/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/java_block_inputs.dms',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/java_block_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + [markdown] id="nd1Mvf7X1m3V"
# ANDROID BLOCK
# + id="8Vryu0xr1pfw"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/android_block/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/android_block_inputs.dms',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/android_block_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + [markdown] id="sQb3bawN2Odw"
# JAVA TOKEN
# + id="9nB1-LMK1pij"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/java_token/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/java_token_inputs.dms',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/java_token_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + [markdown] id="BrIBMbG62WFt"
# ANDROID TOKEN
# + id="D7LHIB4g2VlZ" colab={"base_uri": "https://localhost:8080/"} outputId="52ca7516-4cf2-4118-e580-4c20557a3ca3"
from mesh_tensorflow.transformer.learning_rate_schedules import slanted_triangular
MODEL_SIZE = "small"
# Set the folder where the checkpoints and all the others information will be writed
MODEL_DIR = 'gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/android_token/model'
model_parallelism, train_batch_size, keep_checkpoint_max = {
"small": (1, 256, 16),
"base": (2, 128, 8),
"large": (8, 64, 4),
"3B": (8, 16, 1),
"11B": (8, 16, 1)}[MODEL_SIZE]
tf.io.gfile.makedirs(MODEL_DIR)
model = t5.models.MtfModel(
model_dir=MODEL_DIR,
tpu=TPU_ADDRESS,
tpu_topology=TPU_TOPOLOGY,
model_parallelism=model_parallelism,
batch_size=train_batch_size,
learning_rate_schedule = slanted_triangular,
sequence_length={"inputs": 256, "targets": 256},
save_checkpoints_steps=5000,
keep_checkpoint_max=keep_checkpoint_max if ON_CLOUD else None,
iterations_per_loop=100,
)
vocabulary_predict=get_default_vocabulary()
model.predict(input_file='gs://bucket_code_completion/T5_extension/single_finetuning/predict/android_token_inputs.dms',
output_file='gs://bucket_code_completion/T5_extension/single_finetuning_no_PT/predict/android_token_predictions.dms',
checkpoint_steps=-1, beam_size=1, temperature=0.0, keep_top_k=-1, vocabulary=vocabulary_predict)
# + id="bBXTPr0ke04I"
|
Finetuning/single_task_no_pretrained/evaluate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 190514
# +
# function review
# -
def len_of_string(*args):
if args:
for i in args:
print(i, len(i))
# +
# open close read write file
# -
import os
os.listdir()
print(type(os.listdir()))
os.listdir('C:\\Users\\yijiun0407\\Documents')
os.listdir()
for file in os.listdir():
if 'help.py'==file:
open_file=open(file)
#print(open_file)
#print(type(open_file.read()))
#print('*'*60)
(open_file.readlines())
new_content=[]
for file in os.listdir():
if 'help.py'==file:
open_file=open(file)
#print(open_file)
#print(type(open_file.read()))
#print('*'*60)
reading=(open_file.readlines())
for _ in reading:
new_content.append(_.replace('\n',''))
print(new_content)
for file in os.listdir():
if 'help.py'==file:
open_file=open(file)
reading = open_file.read().splitlines()
print(reading)
break
for file in os.listdir():
if 'help.py'==file:
open_file=open(file)
reading1 = open_file.read().splitlines()
#open_file.close
#open_file=open(file)
reading2 = open_file.read().splitlines()
print(reading1)
print(reading2)
for file in os.listdir():
if 'help.py'==file:
with open(file) as rf1:
print(rf1)
print(rf1.read().splitlines())
with open(file) as rf1:
print(rf1)
print(rf1.read().splitlines())
# +
# open -->read --> close
# with open ...(auto close the file you file)
# -
files = os.listdir('C:\\Users\\yijiun0407\\Downloads\\python101_demo_data\\python101_demo_data\\data_wo_sudir')
TXT=[]
for file in files:
if file.endswith('.txt'):
TXT.append(_)
file_path ='C:\\Users\\yijiun0407\\Downloads\\python101_demo_data\\python101_demo_data\\data_wo_sudir\\'+file
print(TXT)
for file in TXT:
with open(file_path) as txt1:
print(txt1.read().splitlines()[0:5])
for root, subdirs, files in os.walk('C:\\Users\\yijiun0407\\Downloads\\python101_demo_data\\python101_demo_data\\data_wo_sudir'):
print(root,subdirs,files)
for root, subdirs, files in os.walk('C:\\Users\\yijiun0407\\Downloads\\python101_demo_data\\python101_demo_data\\data_wo_sudir'):
for file in files:
if file.endswith('txt'):
print(os.path.join(root,file))
for root, subdirs, files in os.walk('C:\\Users\\yijiun0407\\Downloads\\python101_demo_data\\python101_demo_data\\data_wo_sudir'):
for file in files:
if file.endswith('txt'):
with open (os.path.join(root,file)) as rf:
print (rf.read().splitlines()[0:5])
|
190514.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Goal of Week 1
# This week, we will:
# + Spend a little bit of time getting farmiliarized with the awesomeness of Jupyter Notebook (and also learn a few oddities that comes with it)
# + Try a few simple programs.
# + Learn the difference between Print() and Return.
# + And if we have time, we can load data from an excel/csv file.
#
# # Note
# + Remember to run each invidual cell you can hit `Ctrl+Enter`
# + If you want to add a cell above or below, you can hit `Esc + A` or `Esc + B`. Or you can go to the toolbar at the top -> Insert -> Insert Cell Above/Insert Cell Below
# + If you want to run all the cells altogether, go to the toolbar -> Cell -> Run All.
# + In the case something catastrophic happens and everything explodes, keep calmn, go to the the toolbar -> Kernal -> Restart
# ## Let's get started with our first program!
print("Hello World")
# Awesome. Our "program" prints out exactly what we want to see. Now let's try to do some simple arithmetic.
6 + 6
# As you can see, the beauty of using a notebook is that it's very interactive. Unlike when you use IDLE or an IDE like Pycharm, each of the cell in this notebook can be indepently ran on its own OR be used all together. It also shows you the result right away.
#
# Let's try some more simple operations.
6 - 4
6*6
6**6
6%4
6//4
# So that's all pretty neat. Let's see now what will happen if we want to define some variables.
a = 6
b = 10
my_sum = a+b
# If you just run the cell just like that, nothing will show up. This is because all you did was telling these variables to be something. You haven't called them out yet. Let's do that then.
a
b
my_sum
# That's neat. But let's say I want to print my result out a little bit more explicitly with some annotation of what the program is showing.
print("My result is" + my_sum)
# That's weird. It didn't work. Let's google this [TypeError: Can't convert 'int' object to str implicitly
# ] error to see what it means. I found this: http://stackoverflow.com/questions/13654168/typeerror-cant-convert-int-object-to-str-implicitly
#
# According to the top answer, "You cannot concatenate a string with an int. You would need to convert your int to string using str function, or use formatting to format your output."
#
# So let's try the top answerer's solution now. We will put the str() surround my_sum to see if it works.
print("My result is " + str(my_sum))
# Awesome. So now we know my_sum was apparently recognized and interpreted as interger in Python so when we try to add it to a word, Python couldn't figure out what we were trying to do. Now it can understand that yes, temporarily, we want to treat my_sum as a word so we can print it out.
# Since I want to remember why I put my_sum in the str() thingy. I'm going to comment about it so I can remember later. Comments in Python are preceded with `#`. Everytime Python sees this `#`, it's going to ignore the rest of the line
print("My commented result is " + str(my_sum)) # cast my_sum as a string so I can print the whole statement
# Great! Now let's go back to when we did our first program, "Hello World". What we just did was **printing** the words "Hello World". Let's see what happens when we try to do **return** instead of print.
return ('Hello World')
# You can see that if you just return ("Hello World") instead of print, Python will give you a [SyntaxError: 'return' outside function error]. Let's search what the error is.
#
# Not sure what everything means, but this might look like a problem:
# http://stackoverflow.com/questions/26824659/python-return-outside-function-syntax-error
#
# The return command is part of the function definition, so must be indented under the function introduction (which is def donuts()). If it weren't, there would be no way for the interpreter to know that the return statement wasn't just a part of your broader code.
#
# So it seems like return only works if it's called within a thing called "function". Let's try to do that.
#
def returnHelloWorld():
return ("Hello World")
returnHelloWorld()
# if you just run this cell by itself NameError: name 'printHelloWorld' is not defined
# Note that if you haven't ran the last cell, Python is going to be very confused and give a
#
# `NameError: name 'printHelloWorld' is not defined error`
#
# This is a very odd quirk of using Jupyter Notebook. The reason is each of these cells are acting indepently from each other. So if you just wrote the cell and don't run it (either using Ctrl+Enter or by navigating to Cell -> Run cells), the program will never be recognized.
# So let's run the def printHelloWorld cell and see what happens when we call it again.
returnHelloWorld()
# Awesome! Now it works!
# Now before we go into something a little bit more "advanced", this will be a really good opportunity for me to show some difference between **print** and **return**. Hopefully you will never have to run the agony of figuring out what went wrong with your functions.
# I'm going to write a new function that will print, not return, Hello World. In Python, a function starts with **def**. Not sure why they did it that way, but we are going to go with it.
#
# ```python
# def nameOfFunction(stuffYouWantToPassInIfThereIsAny):
# stuff you want to do # note how everything inside is indented
# ```
def printHelloWorld():
print('Hello World')
# Let's call printHelloWorld now
printHelloWorld()
# #### OK Quynh, this seems exactly the same as what returnHelloWorld did. What's the difference?
#
# There's a difference! I will show you. Let's say now I want to have a new variable named "my_result", and I want to assign whatever result I get from the HelloWorld function to the variable "my_result". Then, I want whatever contained in my_result and add ", I'm here" to the end of it. Let's see how it works out.
my_result1 = returnHelloWorld()
my_result1 + ", I'm here"
my_result2 = printHelloWorld()
my_result2 + ", I'm here"
# The printHelloWorld function apparently didn't work like we wanted. Let's search up this error and see what comes up.
#
# http://stackoverflow.com/questions/23372824/typeerror-unsupported-operand-types-for-nonetype-and-str
#
# And the reason for this is because when we use the print statement, it only prints out for us too see. But it never returns anything so that the world outside of our printHelloWorld function can use. So as a matter of fact, my_result2 is actually just "None", or nothing.
# Alright, that's all we need to mind about print and return. If you ever run into a problem where you don't know why your data is not showing up the way you want it too. This could be one of the reasons why.
|
media/f16-scientific-python/.ipynb_checkpoints/PythonScientificWS-Week1-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false nbgrader={"checksum": "cebdb0cf7dac64a3ae23ccb88f6fda3b", "grade": false, "grade_id": "cell-fa48e7f1b94baa5b", "locked": true, "schema_version": 1, "solution": false}
# # Assignment 1
# For this assignment you are welcomed to use other regex resources such a regex "cheat sheets" you find on the web. Feel free to share good resources with your peers in slack!
#
#
# + [markdown] deletable=false editable=false nbgrader={"checksum": "d17f561e3c6c08092810b982d085f5be", "grade": false, "grade_id": "cell-d4da7eb9acee2a6d", "locked": true, "schema_version": 1, "solution": false}
# Before start working on the problems, here is a small example to help you understand how to write your own answers. In short, the solution should be written within the function body given, and the final result should be returned. Then the autograder will try to call the function and validate your returned result accordingly.
# + deletable=false editable=false nbgrader={"checksum": "7eeb5e7d0f0e0137caed9f3b5cb925b1", "grade": false, "grade_id": "cell-4a96535829224b3f", "locked": true, "schema_version": 1, "solution": false}
def example_word_count():
# This example question requires counting words in the example_string below.
example_string = "Amy is 5 years old"
# YOUR CODE HERE.
# You should write your solution here, and return your result, you can comment out or delete the
# NotImplementedError below.
result = example_string.split(" ")
return len(result)
#raise NotImplementedError()
# -
# ## Part A
#
# Find a list of all of the names in the following string using regex.
# + deletable=false nbgrader={"checksum": "29bc8c161c0e246c1e3ef4820cc164f7", "grade": false, "grade_id": "names", "locked": false, "schema_version": 1, "solution": true}
import re
def names():
simple_string = """Amy is 5 years old, and her sister Mary is 2 years old.
Ruth and Peter, their parents, have 3 kids."""
output = re.findall('([A-Z][a-z]+)', simple_string)
print(output)
return output
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "ed5c09ac57f7d98130d5abc557f6d6c4", "grade": true, "grade_id": "correct_names", "locked": false, "points": 1, "schema_version": 1, "solution": false}
assert len(names()) == 4, "There are four names in the simple_string"
# + [markdown] deletable=false editable=false nbgrader={"checksum": "77b3d100c47e9e41d98f82dfeb7eba9c", "grade": false, "grade_id": "cell-ed64e3464ddd7ba7", "locked": true, "schema_version": 1, "solution": false}
# ## Part B
#
# The dataset file in [assets/grades.txt](assets/grades.txt) contains a line separated list of people with their grade in
# a class. Create a regex to generate a list of just those students who received a B in the course.
# + deletable=false nbgrader={"checksum": "e977a1df674e9fa684e6d172aec92824", "grade": false, "grade_id": "grades", "locked": false, "schema_version": 1, "solution": true}
import re
def grades():
with open ("assets/grades.txt", "r") as file:
grades = file.read()
#output = re.findall('.*: B' , grades)
output = re.findall('.*B' , grades)
for i in range(len(output)):
output[i] = output[i][:-3]
print(output)
return output
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "e0bcc452d60fc45259e58d3116d25477", "grade": true, "grade_id": "correct_grades", "locked": false, "points": 1, "schema_version": 1, "solution": false}
assert len(grades()) == 16
# + [markdown] deletable=false editable=false nbgrader={"checksum": "36e3e2a3a3e29fa7b823d22476392320", "grade": false, "grade_id": "cell-e253518e37d33f0c", "locked": true, "schema_version": 1, "solution": false}
# ## Part C
#
# Consider the standard web log file in [assets/logdata.txt](assets/logdata.txt). This file records the access a user makes when visiting a web page (like this one!). Each line of the log has the following items:
# * a host (e.g., '172.16.31.10')
# * a user_name (e.g., 'feest6811' **note: sometimes the user name is missing! In this case, use '-' as the value for the username.**)
# * the time a request was made (e.g., '21/Jun/2019:15:45:24 -0700')
# * the post request type (e.g., 'POST /incentivize HTTP/1.1' **note: not everything is a POST!**)
#
# Your task is to convert this into a list of dictionaries, where each dictionary looks like the following:
# ```
# example_dict = {"host":"172.16.31.10",
# "user_name":"feest6811",
# "time":"21/Jun/2019:15:45:24 -0700",
# "request":"POST /incentivize HTTP/1.1"}
# ```
# + deletable=false nbgrader={"checksum": "c04017e59e48b2f4c77bf425ed84b356", "grade": false, "grade_id": "logs", "locked": false, "schema_version": 1, "solution": true}
import re
def logs():
with open("assets/logdata.txt", "r") as file:
logdata = file.read()
y = logdata.splitlines()
#print(y[0])
name_list = list()
dictionary_iter = dict()
#for i in range(len(y)):
for i in range(2):
dictionary_iter['host'] = str(re.findall('.*[.][\d]* -',y[i]))
dictionary_iter['host'] = dictionary_iter['host'][2:-4]
#print(dictionary_iter['host'])
dictionary_iter['user_name'] = str(re.findall('- [\w]* [[]',y[i]))
if len(dictionary_iter['user_name']) == 2 :
dictionary_iter['user_name'] = '-'
else:
dictionary_iter['user_name'] = dictionary_iter['user_name'][4:-4]
#print(dictionary_iter['user_name'])
dictionary_iter['time'] = str(re.findall('[[].*[]]',y[i]))
dictionary_iter['time'] = dictionary_iter['time'][3:-3]
#print(dictionary_iter['time'])
dictionary_iter['request'] = str(re.findall('["].*["]',y[i]))
dictionary_iter['request'] = dictionary_iter['request'][3:-3]
#print(dictionary_iter['request'])
name_list.append(dictionary_iter)
name_list.append(i)
print(name_list)
return name_list
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"checksum": "1fd5f2cca190d37c667fb189352540d3", "grade": true, "grade_id": "cell-correct_logs", "locked": true, "points": 1, "schema_version": 1, "solution": false}
assert len(logs()) == 979
one_item={'host': '172.16.31.10',
'user_name': 'feest6811',
'time': '21/Jun/2019:15:45:24 -0700',
'request': 'POST /incentivize HTTP/1.1'}
assert one_item in logs(), "Sorry, this item should be in the log results, check your formating"
# -
|
week1/.ipynb_checkpoints/assignment1-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test package locally
# +
#import requests
#import os
#import pandas as pd
#import json
#import getpass
#import io
#from requests.adapters import HTTPAdapter
#from requests.packages.urllib3.util.retry import Retry
# -
import sddk
from importlib import reload # for testing purposes
# always be sure that you are using the local version
reload(sddk)
sddk.test_package()
# # Configure personal session
conf = sddk.configure()
# + [markdown] colab_type="text" id="PaFpJiRxkxee"
# # Testing write_file() and read_file()
# + [markdown] colab_type="text" id="8EBD8gTEFAfC"
# So far the function has been used with several different types of Python objects, like `str`, `list`, `dictionary`, Pandas' `dataframe` and Matplotlib's `figure`. These have been written either as `.txt`, `.json` or `.png`, `.feather`, based simply upon filename's ending chosen by the user. Here is a couple of examples of these objects in play:
# + colab={"base_uri": "https://localhost:8080/", "height": 265} colab_type="code" executionInfo={"elapsed": 862, "status": "ok", "timestamp": 1580672976544, "user": {"displayName": "Vojt\u011bch Ka\u0161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="bgoY7CirYEwh" outputId="f4692e6e-479e-4fed-d879-417162fdb41d"
### Python "str" object
string_object = "string content for testing"
### Python "list" object
list_object = ['a', 'b', 'c', 'd']
### Python "dictionary" object
dict_object = {"a" : 1, "b" : 2, "c":3 }
### Pandas dataframe object
import pandas as pd
dataframe_object = pd.DataFrame([("a1", "b1", "c1"), ("a2", "b2", "c2")], columns=["a", "b", "c"])
# -
# ## pandas.DataFrame to `.json` and back
import pandas as pd
dataframe_object = pd.DataFrame([("a1", "b1", "c1"), ("a2", "b2", "c2")], columns=["a", "b", "c"])
dataframe_object
sddk.write_file("personal_folder/simple_dataframe.json", dataframe_object, conf)
sddk.read_file("personal_folder/simple_dataframe.json", "df", conf)
# ## pandas.DataFrame to `.feather` and back
sddk.write_file("personal_folder/simple_dataframe.feather", dataframe_object, conf)
sddk.read_file("personal_folder/simple_dataframe.feather", "df", conf)
# %%time
EDH_sample = sddk.read_file("EDH_sample.feather", "df", "8fe7d59de1eafe5f8eaebc0044534606")
EDH_sample.head(5)
sddk.write_file("personal_folder/EDH_sample.feather", EDH_sample, conf)
# ## pandas.DataFrame to `.csv` and back
import pandas as pd
dataframe_object = pd.DataFrame([("a1", "b1", "c1"), ("a2", "b2", "c2")], columns=["a", "b", "c"])
dataframe_object
sddk.write_file("personal_folder/simple_dataframe.csv", dataframe_object, conf)
sddk.read_file("personal_folder/simple_dataframe.csv", "df", conf)
# ## String to .txt and back
### Python "str" object
string_object = "string content for testing"
# + colab={} colab_type="code" id="elit-u3qLjxE"
# write the file
sddk.write_file("personal_folder/test_string.txt", string_object, conf)
# -
# read it back
string_object = sddk.read_file("personal_folder/test_string.txt", "str", conf)
string_object
# ## Avoiding misspelling in filenames etc.
# The package attempts to help you avoid mispellings, so it warns if you make a nonsense.
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" executionInfo={"elapsed": 47454, "status": "ok", "timestamp": 1580671300859, "user": {"displayName": "<NAME>0161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="EWKUz2KXMpM3" outputId="bfff39fc-d28d-4b2d-fbdb-6e923f9f846c"
sddk.write_file("nonexistent_folder/textfile.wtf", string_object, conf)
# -
# ## List to .json and back
### Python "list" object
list_object = ['a', 'b', 'c', 'd']
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2456, "status": "ok", "timestamp": 1580671939538, "user": {"displayName": "Vojt\u011b<NAME>\u0161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="SZVsf69fGbi_" outputId="e8acdaca-f5e3-456f-f1e5-1f60666a6a9f"
sddk.write_file("personal_folder/simple_list.json", list_object, conf)
# -
sddk.read_file("personal_folder/simple_list.json", "list", conf)
# + [markdown] colab_type="text" id="lC1ZDRK7Gg5r"
# ## Dictionary to .json and back
#
# -
### Python "dictionary" object
dict_object = {"a" : 1, "b" : 2, "c":3 }
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2356, "status": "ok", "timestamp": 1580672989818, "user": {"displayName": "Vojt\u<NAME>\u0161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="BM6STSKuY-6r" outputId="c4a6078d-36ac-44b6-b95c-a83859a121a5"
sddk.write_file("personal_folder/simple_dict.json", dict_object, conf)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 2707, "status": "ok", "timestamp": 1580673524673, "user": {"displayName": "<NAME>161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="SX7VavEme089" outputId="ca458339-5253-4962-e3d0-8981aec72bc8"
sddk.read_file("personal_folder/simple_dict.json", "dict", conf)
# -
# ## matplotlib.Figure as `.png`
### Matplotlib figure object
import matplotlib.pyplot as plt
figure_object = plt.figure() # generate object
plt.plot(range(10)) # fill it by plotted values
sddk.write_file("personal_folder/fig.png", figure_object, conf)
# + [markdown] colab_type="text" id="AXwFDKJxsf2i"
# # Accessing shared folders
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 12754, "status": "ok", "timestamp": 1580652419176, "user": {"displayName": "Vojt\u<NAME>\u0161e", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="RX2vwK9pxJYp" outputId="f6aeaf37-7677-48d1-d59b-f4282a4db2d0"
# AS FOLDER OWNER
### configure a session with endpoint in the shared folder
### (login as group owner)
conf = sddk.configure("TEST_shared_folder", "<EMAIL>")
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" executionInfo={"elapsed": 5710, "status": "ok", "timestamp": 1579082791879, "user": {"displayName": "Vojt\u<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAxTlnyE78QMWq7qNrhY8cMOVti0SJqAGwQEptYkQ=s64", "userId": "01399835024022498543"}, "user_tz": -60} id="BTbHkjSQqnz7" outputId="337d4161-ba43-43f1-de5c-de9e66c23dd9"
import pandas as pd
dataframe_object = pd.DataFrame([("a1", "b1", "c1"), ("a2", "b2", "c2")], columns=["a", "b", "c"])
dataframe_object
# -
sddk.write_file("simple_dataframe.json", dataframe_object, conf)
sddk.read_file("simple_dataframe.json", "df", conf)
# + colab={"base_uri": "https://localhost:8080/", "height": 86} colab_type="code" executionInfo={"elapsed": 14863, "status": "ok", "timestamp": 1579689278018, "user": {"displayName": "Vojt\u011<NAME>\u0161e", "photoUrl": "<KEY>", "userId": "01399835024022498543"}, "user_tz": -60} id="l6GbkjZLqbSd" outputId="4f4971f3-dcf3-4368-856e-ba832085b62a"
# AS ORDINARY USER (case 1)
### configure a session with endpoint in the shared folder
### (login as ordinary user)
conf = sddk.configure("TEST_shared_folder", "<EMAIL>")
# + colab={} colab_type="code" id="oRI0k0jGqyDb"
### access the shared folder as an ordinary user
simple_dataframe = sddk.read_file("simple_dataframe.json", "df", conf)
simple_dataframe
# -
sddk.write_file("dataframe_from_user.json", simple_dataframe, conf)
# # Accessing files in public folders
gospels_cleaned = sddk.read_file("https://sciencedata.dk/public/3e0a55a4182de313e04523360cecd015", "dict")
c_aristotelicum = sddk.read_file("c_aristotelicum.json", "df", "31b393e2afe1ee96ce81869c7efe18cb")
c_aristotelicum.head(5)
|
.ipynb_checkpoints/sddk_playground-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/eduardojdiniz/CichyWanderers/blob/346ae7adc197457d4642c4c7fafa407ded8c1424/dataloader.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="J8FuAReHVFsX"
# # Data loader
# -
# ### Import CichyWanderers GitHub Repository
# !git clone https://github.com/eduardojdiniz/CichyWanderers
# ### Import dataloader
import CichyWanderers.dataloader as dataloader
# ### Download and create Cichy et al, 2014 dataset
data_dirpath, RDM_dict, stim_dict = dataloader.create_dataset()
# ### Dictionary structures
#
# #### `data_dirpath`
# data_dirpath: string with the path to the data folder
#
# #### `RDM_dict`
# RDM_dict: dict with keys 'MEG', 'fMRI_EVC', 'fMRI_IT'
# 'MEG' : ndarray, (16, 2, 1301, 92, 92)
# 16 subjects, 2 sessions, 1301 time points (from -100 ms to 1200 ms
# wrt to stimulus onset at 0 ms), 92 conditions by 92 conditions.
# The last 2 dimensions form representational dissimilarity matrices of
# decoding accuracies, symmetric accross the diagonal, with the diagonal
# undefined (NaN).
# 'fMRI_EVC': ndarray, (15, 92, 92)
# 15 subjects, 92 conditions by 92 conditions.
# The last 2 dimensions form a representational dissimilarity matrix of
# spearman correlation for the EVC cortex, symmetric accross the diagonal,
# with the diagonal undefined (NaN).
# 'fMRI_IT' : ndarray, (15, 92, 92)
# 15 subjects, 92 conditions by 92 conditions.
# The last 2 dimensions form a representational dissimilarity matrix of
# spearman correlation for the IT cortex, symmetric accross the diagonal,
# with the diagonal undefined (NaN).
#
# #### `stim_dict`
# stim_dict: dict with keys 'category', 'human', 'face', 'animate', 'natural', 'imagepath'
# 'category' : list[str], indicating category
# 'human' : list[int], indicating membership (0=not a member, 1=member)
# 'face' : list[int], indicating membership (0=not a member, 1=member)
# 'animate' : list[int], indicating membership (0=not a member, 1=member)
# 'natural' : list[int], indicating membership (0=not a member, 1=member)
# 'imagepath' : list[str], jpeg image filepath
#
# + [markdown] id="uM5mp1kwbJos" tags=[]
# # References
# 1. [Resolving human object recognition in space and time. Cichy et al. Nature Neuroscience 2014](https://www.nature.com/articles/nn.3635)
# 2. [Representational similarity analysis – connecting the branches of systems neuroscience. Kriegeskorte et al. Front. Syst. Neurosci., 2008](https://www.frontiersin.org/articles/10.3389/neuro.06.004.2008/full?utm_source=FWEB&utm_medium=NBLOG&utm_campaign=ECO_10YA_top-research)
#
#
#
#
|
CichyWanderers/dataloader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Nipype Plugins
#
# The workflow engine supports a plugin architecture for workflow execution. The available plugins allow local and distributed execution of workflows and debugging. Each available plugin is described below.
#
# Current plugins are available for Linear, Multiprocessing, [IPython](https://ipython.org/) distributed processing platforms and for direct processing on [SGE](http://www.oracle.com/us/products/tools/oracle-grid-engine-075549.html), [PBS](http://www.clusterresources.com/products/torque-resource-manager.php), [HTCondor](http://www.cs.wisc.edu/htcondor/), [LSF](http://www.platform.com/Products/platform-lsf), `OAR`, and [SLURM](http://slurm.schedmd.com/). We anticipate future plugins for the [Soma](http://brainvisa.info/soma/soma-workflow/) workflow.
#
# <div class="alert alert-info">
# **Note**:
# Currently, the distributed processing plugins rely on the availability of a shared filesystem across computational nodes.
# A variety of config options can control how execution behaves in this distributed context. These are listed later on in this page.
# </div>
#
# All plugins can be executed with:
#
# ```python
# workflow.run(plugin=PLUGIN_NAME, plugin_args=ARGS_DICT)
# ```
#
# Optional arguments:
#
# status_callback : a function handle
# max_jobs : maximum number of concurrent jobs
# max_tries : number of times to try submitting a job
# retry_timeout : amount of time to wait between tries
#
# <div class="alert alert-info">
# **Note**: Except for the status_callback, the remaining arguments only apply to the distributed plugins: MultiProc / IPython(X) / SGE / PBS / HTCondor / HTCondorDAGMan / LSF
# </div>
# ## Debug
#
# This plugin provides a simple mechanism to debug certain components of a workflow without executing any node.
#
# Mandatory arguments:
#
# callable : A function handle that receives as arguments a node and a graph
#
# The function callable will be called for every node from a topological sort of the execution graph.
# ## Linear
#
# This plugin runs the workflow one node at a time in a single process locally. The order of the nodes is determined by a topological sort of the workflow:
#
# ```python
# workflow.run(plugin='Linear')
# ```
# ## MultiProc
#
# Uses the [Python](http://www.python.org/) multiprocessing library to distribute jobs as new processes on a local system.
#
# Optional arguments:
#
# - `n_procs`: Number of processes to launch in parallel, if not set number of processors/threads will be automatically detected
#
# - `memory_gb`: Total memory available to be shared by all simultaneous tasks currently running, if not set it will be automatically set to 90% of system RAM.
#
# - `raise_insufficient`: Raise exception when the estimated resources of a node exceed the total amount of resources available (memory and threads), when ``False`` (default), only a warning will be issued.
#
# - `maxtasksperchild`: number of nodes to run on each process before refreshing the worker (default: 10).
#
#
# To distribute processing on a multicore machine, simply call:
#
# ```python
# workflow.run(plugin='MultiProc')
# ```
#
# This will use all available CPUs. If on the other hand, you would like to restrict the number of used resources (to say 2 CPUs), you can call:
#
# ```python
# workflow.run(plugin='MultiProc', plugin_args={'n_procs' : 2}
# ```
# ## IPython
#
# This plugin provides access to distributed computing using [IPython](https://ipython.org/) parallel machinery.
#
# <div class="alert alert-info">
# **Note**:
# Please read the [IPython](https://ipython.org/) documentation to determine how to set up your cluster for distributed processing. This typically involves calling ipcluster.
# </div>
#
# Once the clients have been started, any pipeline executed with:
#
# ```python
# workflow.run(plugin='IPython')
# ```
# ## SGE/PBS
#
# In order to use nipype with [SGE](http://www.oracle.com/us/products/tools/oracle-grid-engine-075549.html) or [PBS](http://www.clusterresources.com/products/torque-resource-manager.php) you simply need to call:
#
# ```python
# workflow.run(plugin='SGE')
# workflow.run(plugin='PBS')
# ```
#
# Optional arguments:
#
# template: custom template file to use
# qsub_args: any other command line args to be passed to qsub.
# max_jobname_len: (PBS only) maximum length of the job name. Default 15.
#
# For example, the following snippet executes the workflow on myqueue with a custom template:
#
# ```python
# workflow.run(plugin='SGE',
# plugin_args=dict(template='mytemplate.sh',
# qsub_args='-q myqueue')
# ```
#
# In addition to overall workflow configuration, you can use node level
# configuration for PBS/SGE:
#
# ```python
# node.plugin_args = {'qsub_args': '-l nodes=1:ppn=3'}
# ```
#
# this would apply only to the node and is useful in situations, where a particular node might use more resources than other nodes in a workflow.
#
# <div class="alert alert-info">
# **Note**: Setting the keyword `overwrite` would overwrite any global configuration with this local configuration:
# ```node.plugin_args = {'qsub_args': '-l nodes=1:ppn=3', 'overwrite': True}```
# </div>
# ### SGEGraph
#
# SGEGraph is an execution plugin working with Sun Grid Engine that allows for submitting the entire graph of dependent jobs at once. This way Nipype does not need to run a monitoring process - SGE takes care of this. The use of SGEGraph is preferred over SGE since the latter adds an unnecessary load on the submit machine.
#
# <div class="alert alert-info">
# **Note**: When rerunning unfinished workflows using SGEGraph you may decide not to submit jobs for Nodes that previously finished running. This can speed up execution, but new or modified inputs that would previously trigger a Node to rerun will be ignored. The following option turns on this functionality:
# ```workflow.run(plugin='SGEGraph', plugin_args = {'dont_resubmit_completed_jobs': True})```
# </div>
# ## LSF
#
# Submitting via LSF is almost identical to SGE above except for the optional arguments field:
#
# ```python
# workflow.run(plugin='LSF')
# ```
#
# Optional arguments:
#
# template: custom template file to use
# bsub_args: any other command line args to be passed to bsub.
# ## SLURM
#
# Submitting via SLURM is almost identical to SGE above except for the optional arguments field:
#
# ```python
# workflow.run(plugin='SLURM')
# ```
#
# Optional arguments:
#
# template: custom template file to use
# sbatch_args: any other command line args to be passed to bsub.
# jobid_re: regular expression for custom job submission id search
# ### SLURMGraph
#
# SLURMGraph is an execution plugin working with SLURM that allows for submitting the entire graph of dependent jobs at once. This way Nipype does not need to run a monitoring process - SLURM takes care of this. The use of SLURMGraph plugin is preferred over the vanilla SLURM plugin since the latter adds an unnecessary load on the submit machine.
#
# <div class="alert alert-info">
# **Note**: When rerunning unfinished workflows using SLURMGraph you may decide not to submit jobs for Nodes that previously finished running. This can speed up execution, but new or modified inputs that would previously trigger a Node to rerun will be ignored. The following option turns on this functionality:
# ```workflow.run(plugin='SLURMGraph', plugin_args = {'dont_resubmit_completed_jobs': True})```
# </div>
# ## HTCondor
#
# ### DAGMan
#
# With its [DAGMan](http://research.cs.wisc.edu/htcondor/dagman/dagman.html) component, [HTCondor](http://www.cs.wisc.edu/htcondor/) (previously Condor) allows for submitting the entire graphs of dependent jobs at once (similar to SGEGraph and SLURMGraph). With the ``CondorDAGMan`` plug-in, Nipype can utilize this functionality to submit complete workflows directly and in a single step. Consequently, and in contrast to other plug-ins, workflow execution returns almost instantaneously -- Nipype is only used to generate the workflow graph, while job scheduling and dependency resolution are entirely managed by [HTCondor](http://www.cs.wisc.edu/htcondor/).
#
# Please note that although [DAGMan](http://research.cs.wisc.edu/htcondor/dagman/dagman.html) supports specification of data dependencies as well as data provisioning on compute nodes this functionality is currently not supported by this plug-in. As with all other batch systems supported by Nipype, only HTCondor pools with a shared file system can be used to process Nipype workflows.
#
# Workflow execution with HTCondor DAGMan is done by calling:
#
# ```python
# workflow.run(plugin='CondorDAGMan')
# ```
#
# Job execution behavior can be tweaked with the following optional plug-in arguments. The value of most arguments can be a literal string or a filename, wherein the latter case the content of the file will be used as the argument value:
#
# - `submit_template` : submit spec template for individual jobs in a DAG (see CondorDAGManPlugin.default_submit_template for the default.
# - `initial_specs` : additional submit specs that are prepended to any job's submit file
# - `override_specs` : additional submit specs that are appended to any job's submit file
# - `wrapper_cmd` : path to an executable that will be started instead of a node script. This is useful for wrapper script that executes certain functionality prior to or after a node runs. If this option is given the wrapper command is called with the respective Python executable and the path to the node script as final arguments
# - `wrapper_args` : optional additional arguments to a wrapper command
# - `dagman_args` : arguments to be prepended to the job execution script in the dagman call
# - `block` : if True the plugin call will block until Condor has finished processing the entire workflow (default: False)
#
# Please see the [HTCondor documentation](http://research.cs.wisc.edu/htcondor/manual) for details on possible configuration options and command line arguments.
#
# Using the ``wrapper_cmd`` argument it is possible to combine Nipype workflow execution with checkpoint/migration functionality offered by, for example, [DMTCP](http://dmtcp.sourceforge.net/). This is especially useful in the case of workflows with long-running nodes, such as Freesurfer's recon-all pipeline, where Condor's job prioritization algorithm could lead to jobs being evicted from compute nodes in order to maximize overall throughput. With checkpoint/migration enabled such a job would be checkpointed prior eviction and resume work from the checkpointed state after being rescheduled -- instead of restarting from scratch.
#
# On a Debian system, executing a workflow with support for checkpoint/migration for all nodes could look like this:
#
# ```python
# # define common parameters
# dmtcp_hdr = """
# should_transfer_files = YES
# when_to_transfer_output = ON_EXIT_OR_EVICT
# kill_sig = 2
# environment = DMTCP_TMPDIR=./;JALIB_STDERR_PATH=/dev/null;DMTCP_PREFIX_ID=$(CLUSTER)_$(PROCESS)
# """
# shim_args = "--log %(basename)s.shimlog --stdout %(basename)s.shimout --stderr %(basename)s.shimerr"
# # run workflow
# workflow.run(
# plugin='CondorDAGMan',
# plugin_args=dict(initial_specs=dmtcp_hdr,
# wrapper_cmd='/usr/lib/condor/shim_dmtcp',
# wrapper_args=shim_args)
# )
# ```
# ## OAR
#
# In order to use nipype with OAR you simply need to call:
#
# ```python
# workflow.run(plugin='OAR')
# ```
#
# Optional arguments:
#
# template: custom template file to use
# oar_args: any other command line args to be passed to qsub.
# max_jobname_len: (PBS only) maximum length of the job name. Default 15.
#
# For example, the following snippet executes the workflow on myqueue with
# a custom template:
#
# ```python
# workflow.run(plugin='oar',
# plugin_args=dict(template='mytemplate.sh',
# oarsub_args='-q myqueue')
# ```
#
# In addition to overall workflow configuration, you can use node level configuration for OAR:
#
# ```python
# node.plugin_args = {'overwrite': True, 'oarsub_args': '-l "nodes=1/cores=3"'}
# ```
#
# this would apply only to the node and is useful in situations, where a particular node might use more resources than other nodes in a workflow. You need to set the 'overwrite' flag to bypass the general settings-template you defined for the other nodes.
# ### ``qsub`` emulation
#
# <div class="alert alert-info">
# **Note**: This plug-in is deprecated and users should migrate to the more robust and more versatile ``CondorDAGMan`` plug-in.
# </div>
#
# Despite the differences between HTCondor and SGE-like batch systems the plugin usage (incl. supported arguments) is almost identical. The HTCondor plugin relies on a ``qsub`` emulation script for HTCondor, called ``condor_qsub`` that can be obtained from a [Git repository on git.debian.org](http://anonscm.debian.org/gitweb/?p=pkg-exppsy/condor.git;a=blob_plain;f=debian/condor_qsub;hb=HEAD). This script is currently not shipped with a standard HTCondor distribution but is included in the HTCondor package from http://neuro.debian.net. It is sufficient to download this script and install it in any location on a system that is included in the ``PATH`` configuration.
#
# Running a workflow in a HTCondor pool is done by calling:
#
# ```python
# workflow.run(plugin='Condor')
# ```
#
# The plugin supports a limited set of qsub arguments (``qsub_args``) that cover the most common use cases. The ``condor_qsub`` emulation script translates qsub arguments into the corresponding HTCondor terminology and handles the actual job submission. For details on supported options see the manpage of ``condor_qsub``.
#
# Optional arguments:
#
# qsub_args: any other command line args to be passed to condor_qsub.
|
workshop/nipype_tutorial/notebooks/basic_plugins.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from geopy import distance
df_wst = pd.read_csv('./data/weather/dimension/weather_station.csv')
df_wst.head()
df_mst = pd.read_csv('./data/traffic/dimension/metro_station.csv')
df_mst.head()
def find_shortest_dist(input_lat, input_lon):
min_distance = float("inf")
station_id = ""
for index, row in df_wst.iterrows():
dist = distance.distance((input_lat, input_lon), (row['lat'], row['lon'])).km
if dist < min_distance:
min_distance = dist
station_id = row['weather_station_id']
return station_id, min_distance
df_mst['weather_station_id'] = df_mst.apply(lambda row: find_shortest_dist(row['lat'], row['lon'])[0], axis=1)
df_mst['distance'] = df_mst.apply(lambda row: find_shortest_dist(row['lat'], row['lon'])[1], axis=1)
df_mst = df_mst[['metro_station_id', 'weather_station_id', 'distance']]
df_mst.to_csv('/Users/allen.wang/projects/data-engineer/data/weather/dimension/metro_weather_station.csv', index=False)
df_mst
|
station_to_weather_station.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 (''env_container'': venv)'
# name: pythonjvsc74a57bd0827f31c7a17f115185da69808fd256ce792d94cfeb61ab0a6e35bd7e2999d502
# ---
# +
import pandas as pd
from tqdm import tqdm
from dash_website import DIMENSIONS, MAIN_CATEGORIES_TO_CATEGORIES
MAIN_DIMENSIONS = [
"Abdomen",
"Musculoskeletal",
"Lungs",
"Eyes",
"Heart",
"Arterial",
"Brain",
"Biochemistry",
"Hearing",
"BloodCells",
"PhysicalActivity",
]
PAIRS_MAIN_DIMENSIONS = [
[main_dim_1, main_dim_2]
for idx_dim, main_dim_1 in enumerate(MAIN_DIMENSIONS)
for main_dim_2 in MAIN_DIMENSIONS[idx_dim + 1 :]
]
PAIRS_SUBDIMENSIONS = [
["BrainMRI", "BrainCognitive"],
["EyesOCT", "EyesFundus"],
["HeartECG", "HeartMRI"],
["AbdomenLiver", "AbdomenPancreas"],
["BiochemistryBlood", "BiochemistryUrine"],
["MusculoskeletalScalars", "MusculoskeletalFullBody"],
["MusculoskeletalScalars", "MusculoskeletalSpine"],
["MusculoskeletalScalars", "MusculoskeletalHips"],
["MusculoskeletalScalars", "MusculoskeletalKnees"],
["MusculoskeletalFullBody", "MusculoskeletalSpine"],
["MusculoskeletalFullBody", "MusculoskeletalHips"],
["MusculoskeletalFullBody", "MusculoskeletalKnees"],
["MusculoskeletalSpine", "MusculoskeletalHips"],
["MusculoskeletalSpine", "MusculoskeletalKnees"],
["MusculoskeletalHips", "MusculoskeletalKnees"],
]
DIMENSIONS_TO_EXCLUDE = {
"*": [],
"*instances01": [],
"*instances1.5x": [],
"*instances23": [],
"Abdomen": ["AbdomenLiver", "AbdomenPancreas"],
"AbdomenLiver": ["Abdomen"],
"AbdomenPancreas": ["Abdomen"],
"Arterial": ["ArterialCarotids", "ArterialPulseWaveAnalysis"],
"ArterialCarotids": ["Arterial"],
"ArterialPulseWaveAnalysis": ["Arterial"],
"Biochemistry": ["BiochemistryBlood", "BiochemistryUrine"],
"BiochemistryBlood": ["Biochemistry"],
"BiochemistryUrine": ["Biochemistry"],
"Brain": ["BrainCognitive", "BrainMRI"],
"BrainCognitive": ["Brain"],
"BrainMRI": ["Brain"],
"Eyes": ["EyesAll", "EyesFundus", "EyesOCT"],
"EyesAll": ["Eyes"],
"EyesFundus": ["Eyes"],
"EyesOCT": ["Eyes"],
"Hearing": [],
"Heart": ["HeartECG", "HeartMRI"],
"HeartECG": ["Heart"],
"HeartMRI": ["Heart"],
"BloodCells": [],
"Lungs": [],
"Musculoskeletal": [
"MusculoskeletalFullBody",
"MusculoskeletalHips",
"MusculoskeletalKnees",
"MusculoskeletalScalars",
"MusculoskeletalSpine",
],
"MusculoskeletalFullBody": ["Musculoskeletal"],
"MusculoskeletalHips": ["Musculoskeletal"],
"MusculoskeletalKnees": ["Musculoskeletal"],
"MusculoskeletalScalars": ["Musculoskeletal"],
"MusculoskeletalSpine": ["Musculoskeletal"],
"PhysicalActivity": [],
}
FULL_CATEGORY = (
MAIN_CATEGORIES_TO_CATEGORIES["All"]
+ ["Phenotypic", "Genetics"]
+ [f"All_{main_category}" for main_category in MAIN_CATEGORIES_TO_CATEGORIES.keys()]
)
if __name__ == "__main__":
correlations_raw = pd.read_feather(f"../../../all_data/xwas/univariate_correlations/correlations.feather").set_index(
["dimension_1", "subdimension_1", "dimension_2", "subdimension_2", "category"]
)
correlations_raw.columns = pd.MultiIndex.from_tuples(
list(map(eval, correlations_raw.columns.tolist())), names=["subset_method", "correlation_type"]
)
correlations_raw.reset_index(inplace=True)
for index_dimension in [1, 2]:
correlations_raw[f"squeezed_dimension_{index_dimension}"] = correlations_raw[f"dimension_{index_dimension}"] + correlations_raw[f"subdimension_{index_dimension}"].replace("*", "")
correlations_raw = correlations_raw.drop(columns=["dimension_1", "subdimension_1", "dimension_2", "subdimension_2"]).set_index(["category", "squeezed_dimension_1", "squeezed_dimension_2"])
list_indexes = []
for dimension in DIMENSIONS + ["MainDimensions", "SubDimensions"]:
for category in FULL_CATEGORY:
list_indexes.append([dimension, category])
indexes = pd.MultiIndex.from_tuples(list_indexes, names=["dimension", "category"])
list_columns = []
for subset_method in ["all", "union", "intersection"]:
for correlation_type in ["pearson", "spearman"]:
for observation in ["mean", "std"]:
list_columns.append([subset_method, correlation_type, observation])
columns = pd.MultiIndex.from_tuples(list_columns, names=["subset_method", "correlation_type", "observation"])
averages_correlations = pd.DataFrame(None, index=indexes, columns=columns)
for subset_method in tqdm(["union", "intersection", "all"]):
for correlation_type in ["pearson", "spearman"]:
for category in FULL_CATEGORY:
correlations_category = correlations_raw.loc[category, (subset_method, correlation_type)]
averages_correlations.loc[
("MainDimensions", category), (subset_method, correlation_type, "mean")
] = correlations_category.loc[PAIRS_MAIN_DIMENSIONS].mean()
averages_correlations.loc[
("MainDimensions", category), (subset_method, correlation_type, "std")
] = correlations_category.loc[PAIRS_MAIN_DIMENSIONS].std()
averages_correlations.loc[
("SubDimensions", category), (subset_method, correlation_type, "mean")
] = correlations_category.loc[PAIRS_SUBDIMENSIONS].mean()
averages_correlations.loc[
("SubDimensions", category), (subset_method, correlation_type, "std")
] = correlations_category.loc[PAIRS_SUBDIMENSIONS].std()
for dimension in DIMENSIONS:
correlations_independant = correlations_category.loc[dimension].drop(
index=([dimension] + DIMENSIONS_TO_EXCLUDE[dimension])
)
averages_correlations.loc[
(dimension, category), (subset_method, correlation_type, "mean")
] = correlations_independant.mean()
averages_correlations.loc[
(dimension, category), (subset_method, correlation_type, "std")
] = correlations_independant.std()
averages_correlations.columns = map(str, averages_correlations.columns.tolist())
averages_correlations.reset_index() #.to_feather("data/xwas/univariate_correlations/averages_correlations.feather")
|
external_code/xwas/univariate/tries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from thompson_sampling_algorithm import *
# %matplotlib inline
ads_data = pd.read_csv("Ads_CTR_Optimisation.csv")
ads_data.head(5)
print('CRT by ad')
print('')
for col in ads_data.columns:
print(str(col) + " CTR: " + str(ads_data[col].sum()/len(ads_data)))
# running both implementations on data
ads_, total_result = thompson_sampling(ads_data.values)
ads_2, total_result2 = thompson_sampling_2(ads_data.values)
plt.hist(ads_)
plt.title('Thompson sampling algorithm on CRT optimization')
plt.xlabel('Ad version')
plt.ylabel('# of times ad was selected')
plt.show()
# +
# keep in mind that this count is 0 indexed, so counte item #4 = ad #5
# as you can see the original implementation does not explore all options
from collections import Counter
Counter(ads_)
# -
plt.hist(ads_2)
plt.title('Thompson sampling algorithm on CRT optimization')
plt.xlabel('Ad version')
plt.ylabel('# of times ad was selected')
plt.show()
# +
# my implementation, altough apparently less efficient, reaches similar results while testing every ad
Counter(ads_2)
# -
|
RL_for_CTR_optimization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 03: Splitting sentences and PoS annotation
# Let's start with a simple paragraph, copied from the course description:
text = """
Increasingly, customers send text to interact or leave comments,
which provides a wealth of data for text mining. That’s a great
starting point for developing custom search, content recommenders,
and even AI applications.
"""
repr(text)
# Notice how there are explicit *line breaks* in the text. Let's write some code to flow the paragraph without any line breaks:
text = " ".join(map(lambda x: x.strip(), text.split("\n"))).strip()
repr(text)
# Now we can use [TextBlob](http://textblob.readthedocs.io/) to *split* the paragraph into sentences:
# +
from textblob import TextBlob
for sent in TextBlob(text).sentences:
print("> ", sent)
# -
# Next we take a sentence and *annotate* it with part-of-speech (PoS) tags:
# +
import textblob_aptagger as tag
sent = "Increasingly, customers send text to interact or leave comments, which provides a wealth of data for text mining."
ts = tag.PerceptronTagger().tag(sent)
print(ts)
# -
# Given these annotations for part-of-speech tags, we can *lemmatize* nouns and verbs to get their root forms. This will also singularize the plural nouns:
# +
from textblob import Word
ts = [('InterAct', 'VB'), ('comments', 'NNS'), ('provides', 'VBZ'), ('mining', 'NN')]
for lex, pos in ts:
w = Word(lex.lower())
lemma = w.lemmatize(pos[0].lower())
print(lex, pos, lemma)
# -
# We can also lookup synonyms and definitions for each word, using *synsets* from [WordNet](https://wordnet.princeton.edu/):
# +
from textblob.wordnet import VERB
w = Word("comments")
for synset, definition in zip(w.get_synsets(), w.define()):
print(synset, definition)
# -
|
1j_NLP_Python/ex03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
tiles = """
abl ace adj aer ara art ase ave ban bar bas ble
bri buk che cme cnt coo dct dep der des dge
dim dle ent ent erp etm ewe gai gre gri gts ham
hit hod hyp ifi ign ile inc ing ini ino inv itc
ker ket men mic nae nch nee nin non ome ore
pal ran rdi res roo rry sch sci she sys taa tau
tcr ted tem tep ter tst ust ver was wat
"""
# +
import forge
from data import warehouse
from puzzle.puzzlepedia import prod_config
prod_config.init()
trie = warehouse.get('/words/unigram/trie')
# -
import re
from data.seek_sets import chain_seek_set
# +
def walk(seek_set, acc, targets, pos=0):
if pos >= len(targets):
yield ' '.join(acc)
return
target = targets[pos]
seek_set.set_length(target)
for result, weight in trie.walk(seek_set, exact_match=True):
#if weight < 5e4:
# break
acc.append(result)
yield from walk(seek_set[result:], acc, targets, pos+1)
acc.pop()
def process(tiles, targets):
#print(tiles + ['a'])
found = set()
for c in 'abcdefghijklmnopqrstuvwxyz':
seek_set = chain_seek_set.ChainSeekSet(inject(tiles, c), sum(targets))
for result in walk(seek_set, [], targets):
if result in found:
continue
print(result)
found.add(result)
def inject(tiles, c):
result = [c]
for t in tiles:
result.append(t)
result.append(c + t[0] + t[1] + t[2])
result.append(t[0] + c + t[1] + t[2])
result.append(t[0] + t[1] + c + t[2])
return result
def parse(s):
parts = s.split(' ')
result = []
for p in parts:
p = p.strip('’,;.‘^!-*')
if p:
result.append(int(p))
return result
# -
tiles = """
abl ace adj aer ara art ase ave ban bar bas ble
bri buk che cme cnt coo dct dep der des dge
dim dle ent ent erp etm ewe gai gre gri gts ham
hit hod hyp ifi ign ile inc ing ini ino inv itc
ker ket men mic nae nch nee nin non ome ore
pal ran rdi res roo rry sch sci she sys taa tau
tcr ted tem tep ter tst ust ver was wat
"""
tiles = """
aer ave bas
dct
dim
ini
ket
she
tep was
"""
process(tiles.split(), [10, 6])
print("""
abl ace adj aer ara art ase ave ban bar bas ble
bri buk che cme cnt coo dct dep der des dge
dim dle ent ent erp etm ewe gai gre gri gts ham
hit hod hyp ifi ign ile inc ing ini ino inv itc
ker ket men mic nae nch nee nin non ome ore
pal ran rdi res roo rry sch sci she sys taa tau
tcr ted tem tep ter tst ust ver was wat
""".replace(' ', '\t'))
|
src/puzzle/examples/puzzle_boat/4/how_big_is_your_hat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## 1. Setup Mario
# !pip install gym_super_mario_bros==7.3.0 nes_py
# +
# Import the game
# Super Mario documentation - https://pypi.org/project/gym-super-mario-bros/
import gym_super_mario_bros
# Import the Joypad wrapper
# JoypadSpace - https://github.com/Kautenja/nes-py/wiki/Wrappers
# nes-py - https://pypi.org/project/nes-py/
from nes_py.wrappers import JoypadSpace
# Import the SIMPLIFIED controls
from gym_super_mario_bros.actions import SIMPLE_MOVEMENT
# -
# Setup game
env = gym_super_mario_bros.make('SuperMarioBros-v0')
env = JoypadSpace(env, SIMPLE_MOVEMENT)
# Create a flag - restart or not
done = True
# Loop through each frame in the game
for step in range(1000):
# Start the game to begin with
if done:
# Start the gamee
env.reset()
# Do random actions
state, reward, done, info = env.step(env.action_space.sample())
# Show the game on the screen
env.render()
# Close the game
env.close()
# ## 2. Preprocess Environment
# Install pytorch
# !pip install torch==1.10.1+cu113 torchvision==0.11.2+cu113 torchaudio===0.10.1+cu113 -f https://download.pytorch.org/whl/cu113/torch_stable.html
# Install stable baselines for RL stuff
# !pip install stable-baselines3[extra]
# Import Frame Stacker Wrapper and GrayScaling Wrapper
from gym.wrappers import GrayScaleObservation
# Import Vectorization Wrappers
from stable_baselines3.common.vec_env import VecFrameStack, DummyVecEnv
# Import Matplotlib to show the impact of frame stacking
from matplotlib import pyplot as plt
# 1. Create the base environment
env = gym_super_mario_bros.make('SuperMarioBros-v0')
# 2. Simplify the controls
env = JoypadSpace(env, SIMPLE_MOVEMENT)
# 3. Grayscale
env = GrayScaleObservation(env, keep_dim=True)
# 4. Wrap inside the Dummy Environment
env = DummyVecEnv([lambda: env])
# 5. Stack the frames
env = VecFrameStack(env, 4, channels_order='last')
state = env.reset()
state, reward, done, info = env.step([5])
plt.figure(figsize=(20,16))
for idx in range(state.shape[3]):
plt.subplot(1,4,idx+1)
plt.imshow(state[0][:,:,idx])
plt.show()
# ## 3. Train the Reinforcement Learning Model
# Import os for file path management
import os
# Import PPO for algos
from stable_baselines3 import PPO
# Import Base Callback for saving models
from stable_baselines3.common.callbacks import BaseCallback
class TrainAndLoggingCallback(BaseCallback):
def __init__(self, check_freq, save_path, verbose=1):
super(TrainAndLoggingCallback, self).__init__(verbose)
self.check_freq = check_freq
self.save_path = save_path
def _init_callback(self):
if self.save_path is not None:
os.makedirs(self.save_path, exist_ok=True)
def _on_step(self):
if self.n_calls % self.check_freq == 0:
model_path = os.path.join(self.save_path, 'best_model_{}'.format(self.n_calls))
self.model.save(model_path)
return True
CHECKPOINT_DIR = './train/'
LOG_DIR = './logs/'
# Setup model saving callback
callback = TrainAndLoggingCallback(check_freq=10000, save_path=CHECKPOINT_DIR)
# This is the AI model started
model = PPO('CnnPolicy', env, verbose=1, tensorboard_log=LOG_DIR, learning_rate=0.000001,
n_steps=512)
# +
#model.load('latest_model')
# -
# Train the AI model, this is where the AI model starts to learn
model.learn(total_timesteps=100000, callback=callback)
# +
#model.save('latest_model')
# -
# ## 4. Test it Out
# Load model
model = PPO.load('./train/best_model_10000.zip')
state = env.reset()
# Start the game
state = env.reset()
# Loop through the game
while True:
action, _ = model.predict(state)
state, reward, done, info = env.step(action)
env.render()
|
mario/Mario_Reinforcement_Learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # POC-English baseline test `2018-11-08`, Link Grammar 5.5.1.
#
# Agglomerative clustering, test_grammar updated 2018-10-19, Link Grammar 5.5.1; server `172.16.58.3`.
# This notebook is shared as static [POC-English-2018-11-08.html](http://langlearn.singularitynet.io/data/clustering_2018/html/POC-English-2018-11-08.html)
# The "All tests" table is shared as 'short_table.txt' in [POC-English-2018-11-08](http://langlearn.singularitynet.io/data/clustering_2018/POC-English-2018-11-08/) directory.
# Previous tests:
# [POC-English-2018-10-23.html](http://langlearn.singularitynet.io/data/clustering_2018/html/POC-English-2018-10-23.html),
# [POC-English-2018-10-21.html](http://langlearn.singularitynet.io/data/clustering_2018/html/POC-English-2018-10-21.html),
# [POC-English-Amb-2018-08-09.html](http://langlearn.singularitynet.io/data/clustering_2018/html/POC-English-Amb-2018-08-09.html)
# ## Basic settings
import os, sys, time
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path: sys.path.append(module_path)
from src.grammar_learner.utl import UTC
from src.grammar_learner.read_files import check_dir
from src.grammar_learner.write_files import list2file
from src.grammar_learner.widgets import html_table
from src.grammar_learner.pqa_table import table_rows
tmpath = module_path + '/tmp/'
check_dir(tmpath, True, 'none')
table = []
long_table = []
start = time.time()
print(UTC(), ':: module_path =', module_path)
# ## Grammar Learner corpus-specific parameters
corpus = 'POC-English-Amb'
out_dir = module_path + '/output/POC-English-' + str(UTC())[:10]
runs = (1,1)
if runs != (1,1): out_dir += '-multi'
kwargs = {
'left_wall' : '' ,
'period' : False ,
'min_word_count': 1 ,
'min_link_count': 1 ,
'max_words' : 100000 ,
'max_features' : 100000 ,
'min_co-occurrence_count': 1 ,
'min_co-occurrence_probability': 1e-9,
'word_space' : 'vectors' ,
'clustering' : ('kmeans', 'kmeans++', 10),
'cluster_range' : (2,50,1,5) ,
'cluster_criteria': 'silhouette',
'clustering_metric': ('silhouette', 'cosine'),
'tmpath' : tmpath ,
'verbose' : 'min' ,
'template_path' : 'poc-turtle',
'linkage_limit' : 1000 ,
'categories_generalization': 'off'}
lines = [
[11, 'POC-English-Amb' , 'MST-fixed-manually' , 0 , 0 , 'none' ],
[12, 'POC-English-Amb' , 'MST-fixed-manually' , 0 , 0 , 'rules' ],
[13, 'POC-English-Amb' , 'LG-English' , 0 , 0 , 'none' ],
[14, 'POC-English-Amb' , 'LG-English' , 0 , 0 , 'rules' ],
[15, 'POC-English-Amb' , 'R=6-Weight=6:R-mst-weight=+1:R' , 0 , 0 , 'none' ],
[16, 'POC-English-Amb' , 'R=6-Weight=6:R-mst-weight=+1:R' , 0 , 0 , 'rules' ],
[17, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R' , 0 , 0 , 'none' ],
[18, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R' , 0 , 0 , 'rules' ],
[19, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R-agm-opt' , 0 , 0 , 'none' ],
[20, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R-agm-opt' , 0 , 0 , 'rules' ],
[21, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R-agm-100' , 0 , 0 , 'none' ],
[22, 'POC-English-disAmb' , 'R=6-Weight=6:R-mst-weight=+1:R-agm-100' , 0 , 0 , 'rules' ],
[23, 'POC-English-Amb' , 'R=6-Weight=1-mst-weight=+1:R' , 0 , 0 , 'none' ],
[24, 'POC-English-disAmb' , 'R=6-Weight=1-mst-weight=+1:R' , 0 , 0 , 'none' ],
[25, 'POC-English-disAmb' , 'R=6-Weight=1-mst-weight=+1:R-agm-opt' , 0 , 0 , 'none' ],
[26, 'POC-English-disAmb' , 'R=6-Weight=1-mst-weight=+1:R-agm-100' , 0 , 0 , 'none' ],
[27, 'POC-English-Amb' , 'LG-ANY-all-parses' , 0 , 0 , 'none' ],
[28, 'POC-English-disAmb' , 'LG-ANY-all-parses' , 0 , 0 , 'none' ],
[29, 'POC-English-disAmb' , 'LG-ANY-all-parses-agm-opt' , 0 , 0 , 'none' ],
[30, 'POC-English-disAmb' , 'LG-ANY-all-parses-agm-100' , 0 , 0 , 'none' ]]
rp = module_path + '/data/POC-English-Amb/MST-fixed-manually/poc-english_ex-parses-gold.txt'
cp = rp # corpus path = reference path
# # Baseline: POC-English-Amb, ...-disAmb, ...agm...
# ULL Project Plan ⇒ Parses ⇒ lines 11-30
# ## Connectors-DRK-Connectors
# %%capture
kwargs['context'] = 1
kwargs['grammar_rules'] = 1
average21, long21, header = table_rows(lines, out_dir, cp, rp, runs, **kwargs)
table.extend(average21)
long_table.extend(long21)
display(html_table([header]+average21))
# ## Connectors-DRK-Disjuncts
# %%capture
kwargs['grammar_rules'] = 2
average22, long22, header = table_rows(lines, out_dir, cp, rp, runs, **kwargs)
table.extend(average22)
long_table.extend(long22)
display(html_table([header]+average22))
# ## Disjuncts-DRK-Disjuncts
# %%capture
kwargs['context'] = 2
average23, long23, header = table_rows(lines, out_dir, cp, rp, runs, **kwargs)
table.extend(average23)
long_table.extend(long23)
display(html_table([header]+average23))
# ## Disjuncts-ILE-Disjuncts
# %%capture
kwargs['word_space'] = 'discrete'
kwargs['clustering'] = 'group'
average24, long24, header = table_rows(lines, out_dir, cp, rp, runs, **kwargs)
table.extend(average24)
long_table.extend(long24)
display(html_table([header]+average24))
# ## Disjuncts-ALE-Disjuncts
# %%capture
kwargs['word_space'] = 'sparse'
kwargs['cluster_range'] = (2,36,1,1)
kwargs['clustering'] = ('agglomerative', 'ward')
kwargs['clustering_metric'] = ('silhouette', 'cosine')
average25, long25, header = table_rows(lines, out_dir, cp, rp, runs, **kwargs)
table.extend(average25)
long_table.extend(long25)
display(html_table([header]+average25))
# # All tests (all entries for multi-test `runs > (1.1)`)
display(html_table([header]+long_table))
print(UTC(), ':: finished, elapsed', str(round((time.time()-start)/3600, 1)), 'hours')
table_str = list2file(table, out_dir+'/short_table.txt')
if runs == (1,1):
print('Results saved to', out_dir + '/short_table.txt')
else:
long_table_str = list2file(long_table, out_dir+'/long_table.txt')
print('Average results saved to', out_dir + '/short_table.txt\n'
'Detailed results for every run saved to', out_dir + '/long_table.txt')
|
notebooks/POC-English-LG.5.5.1-2018-11-08.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from demo_data import demo_data
# +
df = demo_data.pdx_coffee_locations.df
df.info()
# +
keep_cols = [c for c in df.columns if 'id' in c.lower() or c == 'SHAPE' or c == 'LOCNUM']
df = df.loc[:,keep_cols]
df
# -
|
notebooks/data_management/test_demo_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# https://news.daum.net/economic
path = 'https://news.daum.net/economic'
req = requests.get(path)
req.status_code
from bs4 import BeautifulSoup
soup = BeautifulSoup(req.content, 'html.parser')
soup
type(soup)
news = soup.select('div > strong.tit_thumb > a[href].link_txt')
news, type(news)
news[0]
news[0].text.strip()
news[0]['data-tiara-type']
news[0]['class']
news[0]['class'][0].strip()
news[0]['href']
for tag in news:
print(tag.text.strip(), tag['href'].strip())
|
scraping_bs4_daumnews.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finite Volume Discretisation
# In this notebook, we explain the discretisation process that converts an expression tree, representing a model, to a linear algebra tree that can be evaluated by the solvers.
#
# We use Finite Volumes as an example of a spatial method, since it is the default spatial method for most PyBaMM models. This is a good spatial method for battery problems as it is conservative: for lithium-ion battery models, we can be sure that the total amount of lithium in the system is constant. For more details on the Finite Volume method, see [Randall Leveque's book](https://books.google.co.uk/books/about/Finite_Volume_Methods_for_Hyperbolic_Pro.html?id=QazcnD7GUoUC&printsec=frontcover&source=kp_read_button&redir_esc=y#v=onepage&q&f=false).
#
# This notebook is structured as follows:
#
# 1. **Setting up a discretisation**. Overview of the parameters that are passed to the discretisation
# 2. **Discretisations and spatial methods**. Operations that are common to most spatial methods:
# - Discretising a spatial variable (e.g. $x$)
# - Discretising a variable (e.g. concentration)
# 3. **Example: Finite Volume operators**. Finite Volume implementation of some useful operators:
# - Gradient operator
# - Divergence operator
# - Integral operator
# 4. **Example: Discretising a simple model**. Setting up and solving a simple model, using Finite Volumes as the spatial method
#
# To find out how to implement a new spatial method, see the [tutorial](https://pybamm.readthedocs.io/en/latest/tutorials/add-spatial-method.html) in the API docs.
# ## Setting up a Discretisation
# We first import `pybamm` and some useful other modules, and change our working directory to the root of the `PyBaMM` folder:
# + tags=[]
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
import os
import matplotlib.pyplot as plt
from pprint import pprint
os.chdir(pybamm.__path__[0]+'/..')
# -
# To set up a discretisation, we must create a geometry, mesh this geometry, and then create the discretisation with the appropriate spatial method(s). The easiest way to create a geometry is to the inbuilt battery geometry:
# +
parameter_values = pybamm.ParameterValues(
values={
"Negative electrode thickness [m]": 0.3,
"Separator thickness [m]": 0.2,
"Positive electrode thickness [m]": 0.3,
}
)
geometry = pybamm.battery_geometry()
parameter_values.process_geometry(geometry)
# -
# We then use this geometry to create a mesh, which for this example consists of uniform 1D submeshes
# +
submesh_types = {
"negative electrode": pybamm.Uniform1DSubMesh,
"separator": pybamm.Uniform1DSubMesh,
"positive electrode": pybamm.Uniform1DSubMesh,
"negative particle": pybamm.Uniform1DSubMesh,
"positive particle": pybamm.Uniform1DSubMesh,
"current collector": pybamm.SubMesh0D,
}
var = pybamm.standard_spatial_vars
var_pts = {var.x_n: 15, var.x_s: 10, var.x_p: 15, var.r_n: 10, var.r_p: 10}
mesh = pybamm.Mesh(geometry, submesh_types, var_pts)
# -
# Finally, we can use the mesh to create a discretisation, using Finite Volumes as the spatial method for this example
spatial_methods = {
"macroscale": pybamm.FiniteVolume(),
"negative particle": pybamm.FiniteVolume(),
"positive particle": pybamm.FiniteVolume(),
}
disc = pybamm.Discretisation(mesh, spatial_methods)
# ## Discretisations and Spatial Methods
# ### Spatial Variables
# Spatial variables, such as $x$ and $r$, are converted to `pybamm.Vector` nodes
# + tags=[]
# Set up
macroscale = ["negative electrode", "separator", "positive electrode"]
x_var = pybamm.SpatialVariable("x", domain=macroscale)
r_var = pybamm.SpatialVariable("r", domain=["negative particle"])
# Discretise
x_disc = disc.process_symbol(x_var)
r_disc = disc.process_symbol(r_var)
print("x_disc is a {}".format(type(x_disc)))
print("r_disc is a {}".format(type(r_disc)))
# Evaluate
x = x_disc.evaluate()
r = r_disc.evaluate()
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(x, "*")
ax1.set_xlabel("index")
ax1.set_ylabel(r"$x$")
ax2.plot(r, "*")
ax2.set_xlabel("index")
ax2.set_ylabel(r"$r$")
plt.tight_layout()
plt.show()
# -
# We define `y_macroscale`, `y_microscale` and `y_scalar` for evaluation and visualisation of results below
# +
y_macroscale = x ** 3 / 3
y_microscale = np.cos(r)
y_scalar = np.array([[5]])
y = np.concatenate([y_macroscale, y_microscale, y_scalar])
# -
# ### Variables
# In this notebook, we will work with three variables `u`, `v`, `w`.
# +
u = pybamm.Variable("u", domain=macroscale) # u is a variable in the macroscale (e.g. electrolyte potential)
v = pybamm.Variable("v", domain=["negative particle"]) # v is a variable in the negative particle (e.g. particle concentration)
w = pybamm.Variable("w") # w is a variable without a domain (e.g. time, average concentration)
variables = [u,v,w]
# -
# Before discretising, trying to evaluate the variables raises a `NotImplementedError`:
# + tags=[]
try:
u.evaluate()
except NotImplementedError as e:
print(e)
# -
# For any spatial method, a `pybamm.Variable` gets converted to a `pybamm.StateVector` which, when evaluated, takes the appropriate slice of the input vector `y`.
# + tags=[]
# Pass the list of variables to the discretisation to calculate the slices to be used (order matters here!)
disc.set_variable_slices(variables)
# Discretise the variables
u_disc = disc.process_symbol(u)
v_disc = disc.process_symbol(v)
w_disc = disc.process_symbol(w)
# Print the outcome
print("Discretised u is the StateVector {}".format(u_disc))
print("Discretised v is the StateVector {}".format(v_disc))
print("Discretised w is the StateVector {}".format(w_disc))
# -
# Since the variables have been passed to `disc` in the order `[u,v,w]`, they each read the appropriate part of `y` when evaluated:
# +
x_fine = np.linspace(x[0], x[-1], 1000)
r_fine = np.linspace(r[0], r[-1], 1000)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(13,4))
ax1.plot(x_fine, x_fine**3/3, x, u_disc.evaluate(y=y), "o")
ax1.set_xlabel("x")
ax1.legend(["x^3/3", "u"], loc="best")
ax2.plot(r_fine, np.cos(r_fine), r, v_disc.evaluate(y=y), "o")
ax2.set_xlabel("r")
ax2.legend(["cos(r)", "v"], loc="best")
plt.tight_layout()
plt.show()
# + tags=[]
print("w = {}".format(w_disc.evaluate(y=y)))
# -
# ## Finite Volume Operators
# ### Gradient operator
#
# The gradient operator is converted to a Matrix-StateVector multiplication. In 1D, the gradient operator is equivalent to $\partial/\partial x$ on the macroscale and $\partial/\partial r$ on the microscale. In Finite Volumes, we take the gradient of an object on nodes (shape (n,)), which returns an object on the edges (shape (n-1,)).
# + tags=[]
grad_u = pybamm.grad(u)
grad_u_disc = disc.process_symbol(grad_u)
grad_u_disc.render()
# -
# The Matrix in `grad_u_disc` is the standard `[-1,1]` sparse matrix, divided by the step sizes `dx`:
# + tags=[]
macro_mesh = mesh.combine_submeshes(*macroscale)
print("gradient matrix is:\n")
print("1/dx *\n{}".format(macro_mesh.d_nodes[:,np.newaxis] * grad_u_disc.children[0].entries.toarray()))
# -
# When evaluated with `y_macroscale=x**3/3`, `grad_u_disc` is equal to `x**2` as expected:
# +
x_edge = macro_mesh.edges[1:-1] # note that grad_u_disc is evaluated on the node edges
fig, ax = plt.subplots()
ax.plot(x_fine, x_fine**2, x_edge, grad_u_disc.evaluate(y=y), "o")
ax.set_xlabel("x")
legend = ax.legend(["x^2", "grad(u).evaluate(y=x**3/3)"], loc="best")
plt.show()
# -
# Similary, we can create, discretise and evaluate the gradient of `v`, which is a variable in the negative particles. Note that the syntax for doing this is identical: we do not need to explicitly specify that we want the gradient in `r`, since this is inferred from the `domain` of `v`.
v.domain
# + tags=[]
grad_v = pybamm.grad(v)
grad_v_disc = disc.process_symbol(grad_v)
print("grad(v) tree is:\n")
grad_v_disc.render()
micro_mesh = mesh["negative particle"]
print("\n gradient matrix is:\n")
print("1/dr *\n{}".format(micro_mesh.d_nodes[:,np.newaxis] * grad_v_disc.children[0].entries.toarray()))
r_edge = micro_mesh.edges[1:-1] # note that grad_u_disc is evaluated on the node edges
fig, ax = plt.subplots()
ax.plot(r_fine, -np.sin(r_fine), r_edge, grad_v_disc.evaluate(y=y), "o")
ax.set_xlabel("x")
legend = ax.legend(["-sin(r)", "grad(v).evaluate(y=cos(r))"], loc="best")
plt.show()
# -
# #### Boundary conditions
# If the discretisation is provided with boundary conditions, appropriate ghost nodes are concatenated onto the variable, and a larger gradient matrix is used. The ghost nodes are chosen based on the value of the first/last node in the variable and the boundary condition.
# For a Dirichlet boundary condition $u=a$ on the left-hand boundary, we set the value of the left ghost node to be equal to
# $$2*a-u[0],$$
# where $u[0]$ is the value of $u$ in the left-most cell in the domain. Similarly, for a Dirichlet condition $u=b$ on the right-hand boundary, we set the right ghost node to be
# $$2*b-u[-1].$$
# Note also that the size of the gradient matrix is now (41,42) instead of (39,40), to account for the presence of boundary conditions in the State Vector.
# + tags=[]
disc.bcs = {u.id: {"left": (pybamm.Scalar(1), "Dirichlet"), "right": (pybamm.Scalar(2), "Dirichlet")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
u_eval = grad_u_disc.children[1].evaluate(y=y)
print("The value of u on the left-hand boundary is {}".format((u_eval[0] + u_eval[1]) / 2))
print("The value of u on the right-hand boundary is {}".format((u_eval[-2] + u_eval[-1]) / 2))
# -
# For a Neumann boundary condition $\partial u/\partial x=c$ on the left-hand boundary, we set the value of the left ghost node to be
# $$u[0] - c * dx,$$
# where $dx$ is the step size at the left-hand boundary. For a Neumann boundary condition $\partial u/\partial x=d$ on the right-hand boundary, we set the value of the right ghost node to be
# $$u[-1] + d * dx.$$
# + tags=[]
disc.bcs = {u.id: {"left": (pybamm.Scalar(3), "Neumann"), "right": (pybamm.Scalar(4), "Neumann")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
grad_u_eval = grad_u_disc.evaluate(y=y)
print("The gradient on the left-hand boundary is {}".format(grad_u_eval[0]))
print("The gradient of u on the right-hand boundary is {}".format(grad_u_eval[-1]))
# -
# We can mix the types of the boundary conditions:
# + tags=[]
disc.bcs = {u.id: {"left": (pybamm.Scalar(5), "Dirichlet"), "right": (pybamm.Scalar(6), "Neumann")}}
grad_u_disc = disc.process_symbol(grad_u)
print("The gradient object is:")
(grad_u_disc.render())
grad_u_eval = grad_u_disc.evaluate(y=y)
u_eval = grad_u_disc.children[1].evaluate(y=y)
print("The value of u on the left-hand boundary is {}".format((u_eval[0] + u_eval[1])/2))
print("The gradient on the right-hand boundary is {}".format(grad_u_eval[-1]))
# -
# Robin boundary conditions can be implemented by specifying a Neumann condition where the flux depends on the variable.
# ### Divergence operator
# Before computing the Divergence operator, we set up Neumann boundary conditions. The behaviour with Dirichlet boundary conditions is very similar.
disc.bcs = {u.id: {"left": (pybamm.Scalar(-1), "Neumann"), "right": (pybamm.Scalar(1), "Neumann")}}
# Now we can process `div(grad(u))`, converting it to a Matrix-Vector multiplication, plus a vector for the boundary conditions. Since we have Neumann boundary conditions, the divergence of an object of size (n,) has size (n+1,).
# + tags=[]
div_grad_u = pybamm.div(grad_u)
div_grad_u_disc = disc.process_symbol(div_grad_u)
div_grad_u_disc.render()
# -
# Once again, in 1D, the divergence matrix is a `[-1,1]` matrix (divided by the distance between the edges)
# + tags=[]
print("divergence matrix is:\n")
print("1/dx * \n{}".format(
macro_mesh.d_edges[:,np.newaxis] * div_grad_u_disc.children[0].entries.toarray()
))
# -
# We can simplify `div_grad_u_disc`, to collapse the two `[-1,1]` matrices into a single `[1,-2,1]` matrix. The vector of boundary conditions is also simplified.
# + tags=[]
div_grad_u_disc_simp = div_grad_u_disc.simplify()
div_grad_u_disc_simp.render()
# + tags=[]
print("laplacian matrix is:\n")
print("1/dx^2 *\n{}".format(
macro_mesh.d_edges[:,np.newaxis] ** 2 * div_grad_u_disc_simp.children[0].entries.toarray()
))
# -
# Simplifying the tree reduces the time taken to evaluate it:
import timeit
timeit.timeit('div_grad_u_disc.evaluate(y=y)', setup="from __main__ import div_grad_u_disc, y", number=10000)
timeit.timeit('div_grad_u_disc_simp.evaluate(y=y)', setup="from __main__ import div_grad_u_disc_simp, y", number=10000)
# ### Integral operator
# Finally, we can define an integral operator, which integrates the variable across the domain specified by the integration variable.
# + tags=[]
int_u = pybamm.Integral(u, x_var)
int_u_disc = disc.process_symbol(int_u)
print("int(u) = {} is approximately equal to 1/12, {}".format(int_u_disc.evaluate(y=y), 1/12))
# We divide v by r to evaluate the integral more easily
int_v_over_r2 = pybamm.Integral(v/r_var**2, r_var)
int_v_over_r2_disc = disc.process_symbol(int_v_over_r2)
print("int(v/r^2) = {} is approximately equal to 4 * pi * sin(1), {}".format(
int_v_over_r2_disc.evaluate(y=y), 4 * np.pi * np.sin(1))
)
# -
# The integral operators are also Matrix-Vector multiplications
# + tags=[]
print("int(u):\n")
int_u_disc.render()
print("\nint(v):\n")
int_v_over_r2_disc.render()
# -
int_u_disc.children[0].evaluate() / macro_mesh.d_edges
int_v_over_r2_disc.children[0].evaluate() / micro_mesh.d_edges
# ## Discretising a model
# We can now discretise a whole model. We create, and discretise, a simple model for the concentration in the electrolyte and the concentration in the particles, and discretise it with a single command:
# ```
# disc.process_model(model)
# ```
# +
model = pybamm.BaseModel()
c_e = pybamm.Variable("electrolyte concentration", domain=macroscale)
N_e = pybamm.grad(c_e)
c_s = pybamm.Variable("particle concentration", domain=["negative particle"])
N_s = pybamm.grad(c_s)
model.rhs = {c_e: pybamm.div(N_e) - 5, c_s: pybamm.div(N_s)}
model.boundary_conditions = {
c_e: {"left": (np.cos(0), "Neumann"), "right": (np.cos(10), "Neumann")},
c_s: {"left": (0, "Neumann"), "right": (-1, "Neumann")},
}
model.initial_conditions = {c_e: 1 + 0.1 * pybamm.sin(10*x_var), c_s: 1}
# Create a new discretisation and process model
disc2 = pybamm.Discretisation(mesh, spatial_methods)
disc2.process_model(model);
# -
# The initial conditions are discretised to vectors, and an array of concatenated initial conditions is created.
# +
c_e_0 = model.initial_conditions[c_e].evaluate()
c_s_0 = model.initial_conditions[c_s].evaluate()
y0 = model.concatenated_initial_conditions.evaluate()
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(13,4))
ax1.plot(x_fine, 1 + 0.1*np.sin(10*x_fine), x, c_e_0, "o")
ax1.set_xlabel("x")
ax1.legend(["1+0.1*sin(10*x)", "c_e_0"], loc="best")
ax2.plot(x_fine, np.ones_like(r_fine), r, c_s_0, "o")
ax2.set_xlabel("r")
ax2.legend(["1", "c_s_0"], loc="best")
ax3.plot(y0,"*")
ax3.set_xlabel("index")
ax3.set_ylabel("y0")
plt.tight_layout()
plt.show()
# -
# The discretised rhs can be evaluated, for example at `0,y0`:
# +
rhs_c_e = model.rhs[c_e].evaluate(0, y0)
rhs_c_s = model.rhs[c_s].evaluate(0, y0)
rhs = model.concatenated_rhs.evaluate(0, y0)
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, figsize=(13,4))
ax1.plot(x_fine, -10*np.sin(10*x_fine) - 5, x, rhs_c_e, "o")
ax1.set_xlabel("x")
ax1.set_ylabel("rhs_c_e")
ax1.legend(["1+0.1*sin(10*x)", "c_e_0"], loc="best")
ax2.plot(r, rhs_c_s, "o")
ax2.set_xlabel("r")
ax2.set_ylabel("rhs_c_s")
ax3.plot(rhs,"*")
ax3.set_xlabel("index")
ax3.set_ylabel("rhs")
plt.tight_layout()
plt.show()
# -
# The function `model.concatenated_rhs` is then passed to the solver to solve the model, with initial conditions `model.concatenated_initial_conditions`.
# ## Upwinding and downwinding
# If a system is advection-dominated (Peclet number greater than around 40), then it is important to use upwinding (if velocity is positive) or downwinding (if velocity is negative) to obtain accurate results. To see this, consider the following model (without upwinding)
# +
model = pybamm.BaseModel()
model.length_scales = {
"negative electrode": pybamm.Scalar(1),
"separator": pybamm.Scalar(1),
"positive electrode": pybamm.Scalar(1)
}
# Define concentration and velocity
c = pybamm.Variable("c", domain=["negative electrode", "separator", "positive electrode"])
v = pybamm.PrimaryBroadcastToEdges(1, ["negative electrode", "separator", "positive electrode"])
model.rhs = {c: -pybamm.div(c * v) + 1}
model.initial_conditions = {c: 0}
model.boundary_conditions = {c: {"left": (0, "Dirichlet")}}
model.variables = {"c": c}
def solve_and_plot(model):
model_disc = disc.process_model(model, inplace=False)
t_eval = [0,100]
solution = pybamm.CasadiSolver().solve(model_disc, t_eval)
# plot
plot = pybamm.QuickPlot(solution,["c"],spatial_unit="m")
plot.dynamic_plot()
solve_and_plot(model)
# -
# The concentration grows indefinitely, which is clearly an incorrect solution. Instead, we can use upwinding:
model.rhs = {c: -pybamm.div(pybamm.upwind(c) * v) + 1}
solve_and_plot(model)
# This gives the expected linear steady state from 0 to 1. Similarly, if the velocity is negative, downwinding gives accurate results
model.rhs = {c: -pybamm.div(pybamm.downwind(c) * (-v)) + 1}
model.boundary_conditions = {c: {"right": (0, "Dirichlet")}}
solve_and_plot(model)
# ## More advanced concepts
# Since this notebook is only an introduction to the discretisation, we have not covered everything. More advanced concepts, such as the ones below, can be explored by looking into the [API docs](https://pybamm.readthedocs.io/en/latest/source/spatial_methods/finite_volume.html).
#
# - Gradient and divergence of microscale variables in the P2D model
# - Indefinite integral
#
# If you would like detailed examples of these operations, please [create an issue](https://github.com/pybamm-team/PyBaMM/blob/master/CONTRIBUTING.md#a-before-you-begin) and we will be happy to help.
|
examples/notebooks/spatial_methods/finite-volumes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp core
# -
# # Core
# > Hello, details.
#hide
from nbdev.showdoc import *
def say_bye(to):
"Say hello to someone"
return (f"bye {to}")
from fastcore.test import *
test_eq(say_bye("AS"),"bye AS")
assert say_bye("A") == "bye A"
|
00_core.ipynb
|