code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Investigate the impact of miscentering on shear profiles
#
# _the LSST-DESC CLMM team_
#
# This notebook demonstrates the impact of taking wrong cluster centers to construct and derive the mass from reduced shear profiles with`CLMM`. This notebook is based on notebook "demo_dataops_functionality.ipynb".
#
# +
import matplotlib.pyplot as plt
import clmm
import clmm.dataops
from clmm.dataops import compute_tangential_and_cross_components, make_radial_profile, make_bins
from clmm.galaxycluster import GalaxyCluster
import clmm.utils as u
from clmm import Cosmology
from clmm.support import mock_data as mock
import clmm.galaxycluster as gc
from numpy import random
import numpy as np
import clmm.dataops as da
from clmm.support.sampler import fitters
from astropy.coordinates import SkyCoord
import astropy.units as u
# -
# Make sure we know which version we're using
clmm.__version__
# ### Define cosmology object
mock_cosmo = Cosmology(H0 = 70.0, Omega_dm0 = 0.27 - 0.045, Omega_b0 = 0.045, Omega_k0 = 0.0)
np.random.seed(11)
# ## 1. Generate cluster objects from mock data
# In this example, the mock data only include galaxies drawn from redshift distribution.
# Define toy cluster parameters for mock data generation
# +
cosmo = mock_cosmo
cluster_id = "Awesome_cluster"
cluster_m = 1.e15
cluster_z = 0.3
concentration = 4
ngal_density = 50 #gal/arcmin2
cluster_ra = 20.
cluster_dec = 40.
zsrc_min = cluster_z + 0.1 # we only want to draw background galaxies
field_size = 20 #Mpc
ideal_data_z = mock.generate_galaxy_catalog(cluster_m,
cluster_z,
concentration,
cosmo,
'chang13',delta_so=200,
massdef="mean",
zsrc_min=zsrc_min,
ngal_density=ngal_density,
cluster_ra=cluster_ra,
cluster_dec = cluster_dec,
field_size=field_size)
# -
# We want to load this mock data into several CLMM cluster objects, with cluster centers located in a 0.4 x 0.4 degrees windows around the original cluster position `(cluster_ra, cluster_dec)`. The user can change the number of cluster centers if desired. We set the first center to `(cluster_ra,cluster_dec)` for comparison reasons, which corresponds to a == 0 in the cell below.
# +
center_number = 5
cluster_list = []
coord = []
for a in range (0, center_number):
if a == 0:
cluster_ra_new = cluster_ra
cluster_dec_new = cluster_dec
else:
cluster_ra_new = random.uniform(cluster_ra - 0.2, cluster_ra + 0.2)
cluster_dec_new = random.uniform(cluster_dec - 0.2, cluster_dec + 0.2)
cl = clmm.GalaxyCluster(cluster_id, cluster_ra_new, cluster_dec_new,
cluster_z, ideal_data_z)
print("Cluster info = ID:", cl.unique_id, "; ra:", "%.2f" %cl.ra,
"; dec:", "%.2f" %cl.dec, "; z_l :", cl.z)
print("The number of source galaxies is :", len(cl.galcat))
cluster_list.append(cl)
coord.append(SkyCoord(cl.ra*u.deg, cl.dec*u.deg))
# -
#Offset of the different cluster centers from the position 0,0 (in degree)
offset = [coord[0].separation(coord[i]).value for i in range(5)]
# ## 2. Basic checks and plots
# - galaxy positions
# - redshift distribution
#
# For a better visualization, we plot all the different cluster centers, represented by the red dots.
# +
f, ax = plt.subplots(1, 2, figsize=(12, 4))
for cl in cluster_list:
ax[0].scatter(cl.galcat['ra'], cl.galcat['dec'], color='blue', s=1, alpha=0.3)
ax[0].plot(cl.ra, cl.dec, 'ro')
ax[0].set_ylabel('dec', fontsize="large")
ax[0].set_xlabel('ra', fontsize="large")
hist = ax[1].hist(cl.galcat['z'], bins=20)[0]
ax[1].axvline(cl.z, c='r', ls='--')
ax[1].set_xlabel('$z_{source}$', fontsize="large")
xt = {t:f'{t}' for t in ax[1].get_xticks() if t!=0}
xt[cl.z] ='$z_{cl}$'
xto = sorted(list(xt.keys())+[cl.z])
ax[1].set_xticks(xto)
ax[1].set_xticklabels(xt[t] for t in xto)
ax[1].get_xticklabels()[xto.index(cl.z)].set_color('red')
plt.xlim(0, max(xto))
plt.show()
# -
# ## 3. Compute the center effect on the shear profiles
#
# Next, we generate the profiles for all the Cluster objects and save the profiles into a list. We also save the `gt`, `gx`, and `radius` columns of each `profile` into lists, so we can make a plot of these components.
#
# +
bin_edges = make_bins(0.3, 6, 10) # We want to specify the same bins for all the centers.
profile_list = []
for cl in cluster_list:
theta, e_t, e_x = compute_tangential_and_cross_components(
ra_lens=cl.ra, dec_lens=cl.dec,
ra_source=cl.galcat['ra'], dec_source=cl.galcat['dec'],
shear1=cl.galcat['e1'], shear2=cl.galcat['e2'])
cl.compute_tangential_and_cross_components(add=True)
cl.make_radial_profile("Mpc", cosmo=cosmo,bins=bin_edges, include_empty_bins=False)
profile_list.append(cl.profile)
# +
fig = plt.figure(figsize=(10, 6))
for a in range (0, len(profile_list)):
fig.gca().errorbar(profile_list[a]['radius'],profile_list[a]['gt'],profile_list[a]['gt_err'],linestyle='-',
marker='o', label =f'offset = {"{:.2f}".format(offset[a])}°')
plt.xlabel('log(radius)', size=14)
plt.ylabel('gt', size=14)
plt.legend()
plt.show()
# +
fig2 = plt.figure(figsize=(10, 6))
for a in range (0, len(profile_list)):
fig2.gca().errorbar(profile_list[a]['radius'],profile_list[a]['gx'],profile_list[a]['gx_err'],linestyle='-',
marker='o', label =f'offset = {"{:.2f}".format(offset[a])}°')
plt.xlabel('log(radius)', size=14)
plt.ylabel('gx', size=14)
plt.legend(loc=4)
plt.show()
# -
# Since we consider GalaxyCluster objects with no shape noise or photo-z errors, the center (0,0) gives the expected result `gx = 0`, by construction. For the other cluster centers, we can see that the cross shear term average to zero as expected, but the profiles are noisier.
# ## 4. Compute the center effect by fitting the Halo mass
#
# In this last step, we compute the fitting Halo mass with the `nfw` model and, using a plot, compare the impact of the Cluster centers on the weak lensing mass.
from clmm.support.sampler import samplers
# The function below defines the Halo model. For further information, check the notebook "Example2_Fit_Halo_Mass_to_Shear_Catalog.ipynb"
logm_0 = random.uniform(13., 17., 1)[0]
def shear_profile_model(r, logm, z_src):
m = 10.**logm
gt_model = clmm.compute_reduced_tangential_shear(r,
m, concentration,
cluster_z, z_src, cosmo,
delta_mdef=200,
halo_profile_model='nfw')
return gt_model
# +
for a in range(0, len(cluster_list)):
popt,pcov = fitters['curve_fit'](lambda r, logm:shear_profile_model(r, logm, profile_list[a]['z']),
profile_list[a]['radius'],
profile_list[a]['gt'],
profile_list[a]['gt_err'], bounds=[13.,17.])
m_est1 = 10.**popt[0]
m_est_err1 = m_est1 * np.sqrt(pcov[0][0]) * np.log(10) # convert the error on logm to error on m
print(f"The fitted mass is :","%.2e"%m_est1, "for the offset distance:", "%.2f"%offset[a],"}°")
plt.errorbar(offset[a], m_est1, yerr = m_est_err1, fmt='.', color='black', markersize=10)#, label=f'Offset:{"{:.2f}".format(offset[a])}°')
plt.xlabel('offset [deg]', size=12)
plt.ylabel('fitted mass $M_{200,m}$ [M$_{\odot}$]', size=12)
plt.yscale('log')
plt.ylim([5.e12,2.e15])
plt.axhline(cluster_m, label = "input mass")
plt.legend(loc="best")
plt.show()
# -
# We can see that for cluster centers differing from (0,0), we have a negative effect on the lensing mass, which increases with the offset distance.
|
examples/Example1_Fit_shear_profile_with_miscentering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: 2db524e06e9f5f4ffedc911c917cb75e12dbc923643829bf417064a77eb14d37
# name: python3
# ---
# # Regressão Linear Simples
# +
# !pip install numpy
# !pip install matplotlib
# !pip install pandas
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# -
salarios = pd.read_csv('salary_data.csv')
salarios.head()
# !pip install sklearn
# +
X = salarios.iloc[:, :-1].values
y = salarios.iloc[:, -1].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.3, random_state=42)
# -
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
lr.fit(X_train, y_train)
# ## Visualização dos desempenhos em treinamento
_ = plt.scatter(X_train, y_train, color="red")
plt.ylabel('Salário')
plt.xlabel('Experiência')
_ = plt.scatter(X_test, y_test, color="red")
plt.plot(X_test, lr.predict(X_test), color="blue")
plt.title('Salário vs Experiência (Test)')
plt.ylabel('Salário')
plt.xlabel('Experiência')
|
aprendizado-de-maquina-i/regressao/regressao_linear_simples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 2
# *This notebook includes both coding and written questions. Please hand in this notebook file with all the outputs and your answers to the written questions.*
#
# This assignment covers Canny edge detector and Hough transform.
# +
# Setup
import numpy as np
import matplotlib.pyplot as plt
from time import time
from skimage import io
from __future__ import print_function
# %matplotlib inline
plt.rcParams['figure.figsize'] = (15.0, 12.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# for auto-reloading extenrnal modules
# %load_ext autoreload
# %autoreload 2
# -
# ## Part 1: Canny Edge Detector (85 points)
# In this part, you are going to implment Canny edge detector. The Canny edge detection algorithm can be broken down in to five steps:
# 1. Smoothing
# 2. Finding gradients
# 3. Non-maximum suppression
# 4. Double thresholding
# 5. Edge tracking by hysterisis
# ### 1.1 Smoothing (10 points)
# #### Implementation (5 points)
# We first smooth the input image by convolving it with a Gaussian kernel. The equation for a Gaussian kernel of size $(2k+1)\times(2k+1)$ is given by:
#
# $$h_{ij}=\frac{1}{2\pi\sigma^2}\exp{\Bigl(-\frac{(i-k)^2+(j-k)^2}{2\sigma^2}\Bigr)}, 0\leq i,j < 2k+1$$
#
# Implement **`gaussian_kernel`** in `edge.py` and run the code below.
# +
from edge import conv, gaussian_kernel
# Define 3x3 Gaussian kernel with std = 1
kernel = gaussian_kernel(3, 1)
kernel_test = np.array(
[[ 0.05854983, 0.09653235, 0.05854983],
[ 0.09653235, 0.15915494, 0.09653235],
[ 0.05854983, 0.09653235, 0.05854983]]
)
# Test Gaussian kernel
if not np.allclose(kernel, kernel_test):
print('Incorrect values! Please check your implementation.')
# +
# Test with different kernel_size and sigma
kernel_size = 5
sigma = 1.4
# Load image
img = io.imread('iguana.png', as_grey=True)
# Define 5x5 Gaussian kernel with std = sigma
kernel = gaussian_kernel(kernel_size, sigma)
# Convolve image with kernel to achieve smoothed effect
smoothed = conv(img, kernel)
plt.subplot(1,2,1)
plt.imshow(img)
plt.title('Original image')
plt.axis('off')
plt.subplot(1,2,2)
plt.imshow(smoothed)
plt.title('Smoothed image')
plt.axis('off')
plt.show()
# -
# #### Question (5 points)
# What is the effect of the kernel_size and sigma?
# **Your Answer:** Write your solution in this markdown cell.
# ### 1.2 Finding gradients (15 points)
# The gradient of a 2D scalar function $I:\mathbb{R}^2\rightarrow{\mathbb{R}}$ in Cartesian coordinate is defined by:
#
# $$\nabla{I(x,y)}=\bigl[\frac{\partial{I}}{\partial{x}},\frac{\partial{I}}{\partial{y}}\bigr],$$
#
# where
#
# $$
# \frac{\partial{I(x,y)}}{\partial{x}}=\lim_{\Delta{x}\to{0}}\frac{I(x+\Delta{x},y)-I(x,y)}{\Delta{x}} \\
# \frac{\partial{I(x,y)}}{\partial{y}}=\lim_{\Delta{y}\to{0}}\frac{I(x,y+\Delta{y})-I(x,y)}{\Delta{y}}.
# $$
#
# In case of images, we can approximate the partial derivatives by taking differences at one pixel intervals:
#
# $$
# \frac{\partial{I(x,y)}}{\partial{x}}\approx{\frac{I(x+1,y)-I(x-1,y)}{2}} \\
# \frac{\partial{I(x,y)}}{\partial{y}}\approx{\frac{I(x,y+1)-I(x,y-1)}{2}}
# $$
#
# Note that the partial derivatives can be computed by convolving the image $I$ with some appropriate kernels $D_x$ and $D_y$:
#
# $$
# \frac{\partial{I}}{\partial{x}}\approx{I*D_x}=G_x \\
# \frac{\partial{I}}{\partial{y}}\approx{I*D_y}=G_y
# $$
# #### Implementation (5 points)
# Find the kernels $D_x$ and $D_y$ and implement **`partial_x`** and **`partial_y`** using `conv` defined in `edge.py`.
#
# *-Hint: Remeber that convolution flips the kernel.*
# +
from edge import partial_x, partial_y
# Test input
I = np.array(
[[0, 0, 0],
[0, 1, 0],
[0, 0, 0]]
)
# Expected outputs
I_x_test = np.array(
[[ 0, 0, 0],
[ 0.5, 0, -0.5],
[ 0, 0, 0]]
)
I_y_test = np.array(
[[ 0, 0.5, 0],
[ 0, 0, 0],
[ 0, -0.5, 0]]
)
# Compute partial derivatives
I_x = partial_x(I)
I_y = partial_y(I)
# Test correctness of partial_x and partial_y
if not np.all(I_x == I_x_test):
print('partial_x incorrect')
if not np.all(I_y == I_y_test):
print('partial_y incorrect')
# +
# Compute partial derivatives of smoothed image
Gx = partial_x(smoothed)
Gy = partial_y(smoothed)
plt.subplot(1,2,1)
plt.imshow(Gx)
plt.title('Derivative in x direction')
plt.axis('off')
plt.subplot(1,2,2)
plt.imshow(Gy)
plt.title('Derivative in y direction')
plt.axis('off')
plt.show()
# -
# #### Question (5 points)
# What is the reason for performing smoothing prior to computing the gradients?
# **Your Answer:** Write your solution in this markdown cell.
# #### Implementation (5 points)
# Now, we can compute the magnitude and direction of gradient with the two partial derivatives:
#
# $$
# G = \sqrt{G_{x}^{2}+G_{y}^{2}} \\
# \Theta = arctan\bigl(\frac{G_{y}}{G_{x}}\bigr)
# $$
#
# Implement **`gradient`** in `edge.py` which takes in an image and outputs $G$ and $\Theta$.
#
# *-Hint: Use np.arctan2 to compute $\Theta$.*
# +
from edge import gradient
G, theta = gradient(smoothed)
if not np.all(G >= 0):
print('Magnitude of gradients should be non-negative.')
if not np.all((theta >= 0) * (theta < 360)):
print('Direction of gradients should be in range 0 <= theta < 360')
plt.imshow(G)
plt.title('Gradient magnitude')
plt.axis('off')
plt.show()
# -
# ### 1.3 Non-maximum suppression (15 points)
# You should be able to note that the edges extracted from the gradient of the smoothed image is quite thick and blurry. The purpose of this step is to convert the "blurred" edges into "sharp" edges. Basically, this is done by preserving all local maxima in the gradient image and discarding everything else. The algorithm is for each pixel (x,y) in the gradient image:
# 1. Round the gradient direction $\Theta[y,x]$ to the nearest 45 degrees, corresponding to the use of an 8-connected neighbourhood.
#
# 2. Compare the edge strength of the current pixel with the edge strength of the pixel in the positive and negative gradient direction. For example, if the gradient direction is south (theta=90), compare with the pixels to the north and south.
#
# 3. If the edge strength of the current pixel is the largest; preserve the value of the edge strength. If not, suppress (i.e. remove) the value.
#
# Implement **`non_maximum_suppression`** in `edge.py`
# +
from edge import non_maximum_suppression
# Test input
g = np.array(
[[0.4, 0.5, 0.6],
[0.3, 0.5, 0.7],
[0.4, 0.5, 0.6]]
)
# Print out non-maximum suppressed output
# varying theta
for angle in range(0, 180, 45):
print('Thetas:', angle)
t = np.ones((3, 3)) * angle # Initialize theta
print(non_maximum_suppression(g, t))
# -
nms = non_maximum_suppression(G, theta)
plt.imshow(nms)
plt.title('Non-maximum suppressed')
plt.axis('off')
plt.show()
# ### 1.4 Double Thresholding (20 points)
#
# The edge-pixels remaining after the non-maximum suppression step are (still) marked with their strength pixel-by-pixel. Many of these will probably be true edges in the image, but some may be caused by noise or color variations, for instance, due to rough surfaces. The simplest way to discern between these would be to use a threshold, so that only edges stronger that a certain value would be preserved. The Canny edge detection algorithm uses double thresholding. Edge pixels stronger than the high threshold are marked as strong; edge pixels weaker than the low threshold are suppressed and edge pixels between the two thresholds are marked as weak.
#
# Implement **`double_thresholding`** in `edge.py`
# +
from edge import double_thresholding
low_threshold = 0.02
high_threshold = 0.03
strong_edges, weak_edges = double_thresholding(nms, high_threshold, low_threshold)
assert(np.sum(strong_edges & weak_edges) == 0)
edges=strong_edges * 1.0 + weak_edges * 0.5
plt.subplot(1,2,1)
plt.imshow(strong_edges)
plt.title('Strong Edges')
plt.axis('off')
plt.subplot(1,2,2)
plt.imshow(edges)
plt.title('Strong+Weak Edges')
plt.axis('off')
plt.show()
# -
# ### 1.5 Edge tracking (15 points)
#
# Strong edges are interpreted as “certain edges”, and can immediately be included in the final edge image. Weak edges are included if and only if they are connected to strong edges. The logic is of course that noise and other small variations are unlikely to result in a strong edge (with proper adjustment of the threshold levels). Thus strong edges will (almost) only be due to true edges in the original image. The weak edges can either be due to true edges or noise/color variations. The latter type will probably be distributed in dependently of edges on the entire image, and thus only a small amount will be located adjacent to strong edges. Weak edges due to true edges are much more likely to be connected directly to strong edges.
#
# Implement **`link_edges`** in `edge.py`
# +
from edge import get_neighbors, link_edges
test_strong = np.array(
[[1, 0, 0, 0],
[0, 0, 0, 0],
[0, 0, 0, 0],
[0, 0, 0, 1]]
)
test_weak = np.array(
[[0, 0, 0, 1],
[0, 1, 0, 0],
[1, 0, 0, 0],
[0, 0, 1, 0]]
)
test_linked = link_edges(test_strong, test_weak)
plt.subplot(1, 3, 1)
plt.imshow(test_strong)
plt.title('Strong edges')
plt.subplot(1, 3, 2)
plt.imshow(test_weak)
plt.title('Weak edges')
plt.subplot(1, 3, 3)
plt.imshow(test_linked)
plt.title('Linked edges')
plt.show()
# +
edges = link_edges(strong_edges, weak_edges)
plt.imshow(edges)
plt.axis('off')
plt.show()
# -
# ### 1.6 Canny edge detector
# Implement **`canny`** in `edge.py` using the functions you have implemented so far. Test edge detector with different parameters.
#
# Here is an example of the output:
#
# 
# +
from edge import canny
# Load image
img = io.imread('iguana.png', as_grey=True)
# Run Canny edge detector
edges = canny(img, kernel_size=5, sigma=1.4, high=0.03, low=0.02)
print (edges.shape)
plt.imshow(edges)
plt.axis('off')
plt.show()
# -
# ### 1.7 Question (10 points)
#
# 
# **(a)** Suppose that the Canny edge detector successfully detects an edge in an image. The edge (see the figure above) is then rotated by θ, where the relationship between a point on the original edge $(x, y)$ and a point on the rotated edge $(x', y')$ is defined as
#
# $$
# x'=x\cos{\theta}\\
# y'=x\sin{\theta}
# $$
#
# Will the rotated edge be detected using the same Canny edge detector? Provide either a mathematical proof or a counter example.
#
# *-Hint: The detection of an edge by the Canny edge detector depends only on the magnitude of its derivative. The derivative at point (x, y) is determined by its components along the x and y directions. Think about how these magnitudes have changed because of the rotation.*
# **Your Answer:** Write your solution in this markdown cell.
# **(b)** After running the Canny edge detector on an image, you notice that long edges are broken into short segments separated by gaps. In addition, some spurious edges appear. For each of the two thresholds (low and high) used in hysteresis thresholding, explain how you would adjust the threshold (up or down) to address both problems. Assume that a setting exists for the two thresholds that produces the desired result. Briefly explain your answer.
# **Your Answer:** Write your solution in this markdown cell.
# ### Extra Credit: Optimizing Edge Detector
# One way of evaluating an edge detector is to compare detected edges with manually specified ground truth edges. Here, we use precision, recall and F1 score as evaluation metrics. We provide you 40 images of objects with ground truth edge annotations. Run the code below to compute precision, recall and F1 score over the entire set of images. Then, tweak the parameters of the Canny edge detector to get as high F1 score as possible. You should be able to achieve F1 score higher than 0.31 by carefully setting the parameters.
# +
from os import listdir
from itertools import product
# Define parameters to test
sigmas = []
highs = []
lows = []
for sigma, high, low in product(sigmas, highs, lows):
print("sigma={}, high={}, low={}".format(sigma, high, low))
n_detected = 0.0
n_gt = 0.0
n_correct = 0.0
for img_file in listdir('images/objects'):
img = io.imread('images/objects/'+img_file, as_grey=True)
gt = io.imread('images/gt/'+img_file+'.gtf.pgm', as_grey=True)
mask = (gt != 5) # 'don't' care region
gt = (gt == 0) # binary image of GT edges
edges = canny(img, kernel_size=5, sigma=sigma, high=high, low=low)
edges = edges * mask
n_detected += np.sum(edges)
n_gt += np.sum(gt)
n_correct += np.sum(edges * gt)
p_total = n_correct / n_detected
r_total = n_correct / n_gt
f1 = 2 * (p_total * r_total) / (p_total + r_total)
print('Total precision={:.4f}, Total recall={:.4f}'.format(p_total, r_total))
print('F1 score={:.4f}'.format(f1))
# -
# ## Part2: Lane Detection (15 points)
#
# In this section we will implement a simple lane detection application using Canny edge detector and Hough transform.
# Here are some example images of how your final lane detector will look like.
# <img src="lane1.png" width="400">
# <img src="lane2.png" width="400">
#
# The algorithm can broken down into the following steps:
# 1. Detect edges using the Canny edge detector.
# 2. Extract the edges in the region of interest (a triangle covering the bottom corners and the center of the image).
# 3. Run Hough transform to detect lanes.
#
#
# ### 2.1 Edge detection
# Lanes on the roads are usually thin and long lines with bright colors. Our edge detection algorithm by itself should be able to find the lanes pretty well. Run the code cell below to load the example image and detect edges from the image.
# +
from edge import canny
# Load image
img = io.imread('road.jpg', as_grey=True)
# Run Canny edge detector
edges = canny(img, kernel_size=5, sigma=1.4, high=0.03, low=0.02)
plt.subplot(211)
plt.imshow(img)
plt.axis('off')
plt.title('Input Image')
plt.subplot(212)
plt.imshow(edges)
plt.axis('off')
plt.title('Edges')
plt.show()
# -
# ### 2.2 Extracting region of interest (ROI)
# We can see that the Canny edge detector could find the edges of the lanes. However, we can also see that there are edges of other objects that we are not interested in. Given the position and orientation of the camera, we know that the lanes will be located in the lower half of the image. The code below defines a binary mask for the ROI and extract the edges within the region.
# +
H, W = img.shape
# Generate mask for ROI (Region of Interest)
mask = np.zeros((H, W))
for i in range(H):
for j in range(W):
if i > (H / W) * j and i > -(H / W) * j + H:
mask[i, j] = 1
# Extract edges in ROI
roi = edges * mask
plt.subplot(1,2,1)
plt.imshow(mask)
plt.title('Mask')
plt.axis('off')
plt.subplot(1,2,2)
plt.imshow(roi)
plt.title('Edges in ROI')
plt.axis('off')
plt.show()
# -
# ### 2.3 Fitting lines using Hough transform (15 points)
# The output from the edge detector is still a collection of connected points. However, it would be more natural to represent a lane as a line parameterized as $y = ax + b$, with a slope $a$ and y-intercept $b$. We will use Hough transform to find parameterized lines that represent the detected edges.
#
# In general, a straight line $y = ax + b$ can be represented as a point $(a, b)$ in the parameter space. However, this cannot represent vertical lines as the slope parameter will be unbounded. Alternatively, we parameterize a line using $\theta\in{[-\pi, \pi]}$ and $\rho\in{\mathbb{R}}$ as follows:
#
# $$
# \rho = x\cdot{cos\theta} + y\cdot{sin\theta}
# $$
#
# Using this parameterization, we can map everypoint in $xy$-space to a sine-like line in $\theta\rho$-space (or Hough space). We then accumulate the parameterized points in the Hough space and choose points (in Hough space) with highest accumulated values. A point in Hough space then can be transformed back into a line in $xy$-space.
#
# *See [notes](http://web.ipac.caltech.edu/staff/fmasci/home/astro_refs/HoughTrans_lines_09.pdf) on Hough transform.*
#
# Implement **`hough_transform`** in `edge.py`.
# +
from edge import hough_transform
# Perform Hough transform on the ROI
acc, rhos, thetas = hough_transform(roi)
# Coordinates for right lane
xs_right = []
ys_right = []
# Coordinates for left lane
xs_left = []
ys_left = []
for i in range(20):
idx = np.argmax(acc)
r_idx = idx // acc.shape[1]
t_idx = idx % acc.shape[1]
acc[r_idx, t_idx] = 0 # Zero out the max value in accumulator
rho = rhos[r_idx]
theta = thetas[t_idx]
# Transform a point in Hough space to a line in xy-space.
a = - (np.cos(theta)/np.sin(theta)) # slope of the line
b = (rho/np.sin(theta)) # y-intersect of the line
# Break if both right and left lanes are detected
if xs_right and xs_left:
break
if a < 0: # Left lane
if xs_left:
continue
xs = xs_left
ys = ys_left
else: # Right Lane
if xs_right:
continue
xs = xs_right
ys = ys_right
for x in range(img.shape[1]):
y = a * x + b
if y > img.shape[0] * 0.6 and y < img.shape[0]:
xs.append(x)
ys.append(int(round(y)))
plt.imshow(img)
plt.plot(xs_left, ys_left, linewidth=5.0)
plt.plot(xs_right, ys_right, linewidth=5.0)
plt.axis('off')
plt.show()
# -
|
fall_2017/hw2_release/hw2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "My Linear Regressions Are Looking Irregular"
# > "An introductory guide to regularizing linear regressions."
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [linear regression, regularization]
# - hide: false
# - search_exclude: true
# Adapted from [General Assembly's *Data Science Immersive*](https://generalassemb.ly/education/data-science-immersive/).
# ## Theory
# ### Why do we need to regularize our linear regressions? Overfitting.
# What is overfitting?
# - Overfitting means building a model that matches the training data "too closely."
# - The model ends up fitting to noise rather than signal.
# - The **bias is too low** and the **variance is too high**.
#
# What can cause overfitting?
# - Irrelevant features are included in the model.
# - The number of features is close to the number of observations.
# - The features are correlated to each other.
# - The coefficients of the features are large.
# What is regularization and how does it help?
# - Regularization helps against overfitting by imposing a penalty that decreases the coefficients of the features.
# - Regularization lowers the fit for the training data, but it improves the fit for the test data.
#
# ### The two most common types of regularization for linear regressions are *Lasso* and *Ridge*. *Elastic Net* is a mixture of the two.
# - Lasso uses the L1 regularization method, Ridge uses L2, and Elastic Net uses both.
# - We will be regularizing linear regressions in this post, but other types of regressions can be regularized with L1 and L2 as well.
# ## Examples
# ### What kind of situations would call for a regularized linear regression?
# - Predicting home prices based on number of bedrooms upstairs, number of bedrooms downstairs, and total number of bedrooms.
# - Predicting a wine's rating based on its fixed acidity, volatile acidity, citric acid, residual sugar, chloride, free sulfur dioxide, total sulfur dioxide, density, pH, sulphates, and percent alcohol.
# - Predicting the ratio of scores for the teams in a basketball game based on the player stats and the team stats.
# - Estimating a country's life expectancy based on gross domestic product per person, infant mortality, and region of the world.
# ## Code
# ### How to run the Ridge, Lasso, and Elastic Net regressions in Python, using example values.
#
# ### *Preparation for Running the Regressions*
# - Start by importing the Pandas and NumPy libraries, in order to create DataFrames and arrays, as well as the different types of linear regressions we will use, Ridge, Lasso, and Elastic Net:
import pandas as pd
import numpy as np
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LinearRegression, Ridge, RidgeCV, Lasso, LassoCV, ElasticNet, ElasticNetCV
# - Store your training dataset's features in a Pandas DataFrame as X
# - Store your training dataset's target variable in an array as y.
# - This is the R<sup>2</sup> score from **cross-validation** for a linear regression for the sample dataset:
lr_model = LinearRegression()
cross_val_score(lr_model, X_overfit, y, cv = 5).mean()
# > 0.1913277098938003
# - The R<sup>2</sup> score is very low. Is the model overfitted, and will regularization improve it?
#
# ### *Running a Ridge Regression*
# - Ridge regression is **more computationally efficient than Lasso**.
# - Ridge regression usually uses larger $\alpha$ (values for the penalty) than the Lasso regression.
#
# Pick a list of $\alpha$-values, instantiate the model, and fit the regression. Save the optimal model and check its R<sup>2</sup> score from cross-validation:
alpha_list = np.logspace(0, 5, 200)
ridge_model = RidgeCV(alphas = alpha_list, store_cv_values = True) # store_cv_values picks best alpha value
ridge_model = ridge_model.fit(X_overfit, y)
ridge_model.alpha_
# > 821.434358491943
ridge_opt_model = Ridge(alpha = ridge_model.alpha_)
cross_val_score(ridge_opt_model, X_overfit, y, cv = 5).mean()
# > 0.2221069221200395
# This R<sup>2</sup> score is an improvement over the linear regression.
# ### *Running a Lasso Regression*
# - If group of predictors are highly correlated, the Lasso regression picks only one of them and shrinks the others to zero.
# - The Lasso regression is better with smaller $\alpha$ (values for the penalty) than the Ridge regression.
#
# Pick a list of $\alpha$-values, instantiate the model, and fit the regression. Save the optimal model and check its R<sup>2</sup> score from cross-validation:
l_alpha_list = np.arange(0.001, 0.15, 0.0025)
lasso_model = LassoCV(alphas = l_alpha_list, cv = 5)
lasso_model = lasso_model.fit(X_overfit, y)
lasso_model.alpha_
# > 0.011
lasso_model = Lasso(alpha = lasso_model.alpha_)
cross_val_score(lasso_model, X_overfit, y, cv = 5).mean()
# > 0.2617417795359766
# This R<sup>2</sup> score is an improvement over the Ridge regression.
#
# ### *Running an Elastic Net Regression*
# - The Elastic Net regression adds a larger penalty than the Ridge and Lasso regressions.
# - Elastic Net should only be used if the training accuracy is much higher than the test accuracy or if the independent variables are highly correlated.
#
# Pick a list of $\alpha$-values, instantiate the model, and fit the regression. Save the optimal model and check its R<sup>2</sup> score from cross-validation:
enet_alphas = np.arange(0.5, 1.0, 0.005)
enet_model = ElasticNetCV(alphas = enet_alphas)
enet_model = enet_model.fit(X_overfit, y)
enet_model.alpha_
# > 0.5
enet_model = ElasticNet(alpha = enet_model.alpha_)
cross_val_score(enet_model, X_overfit, y, cv = 5).mean()
# > 0.07522288223865307
# This R<sup>2</sup> score is worse than the linear regression, the Ridge regression, and the Lasso regression, so this model penalized our feature coefficients too much.
#
# ### *Conclusions from the Code*
# - For this dataset, the best model was Lasso regression. Compare all of the models with ranges of parameters to determine the best-fitting model.
# - Don't forget to perform feature engineering before modeling, and to re-optimize your model if you go back and engineer features. Feature engineering will affect the optimal model and its optimal parameters.
|
_notebooks/2020-04-01-My-Linear-Regressions-Are-Looking-Irregular.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 3 - NumPy
# ### NAME: <NAME>
# ### STUDENT ID: 3031835795
# Do not delete this cell.
import numpy as np
# ### Since we will run autograder on your notebook, for each question please do not reuse variable names that have been used in previous questions.
# ## NumPy Introduction
#
# __1a) Create two numpy arrays called a and b where a should be all integers between 25-34 (inclusive), and b should be ten evenly spaced numbers between 1-6 (inclusive). Print a and b.__
a = np.array(range(25,35))
b = np.array(np.linspace(1,6,10))
print(a)
print(b)
# __1b) Cube (i.e. raise to the power of 3) all the elements in both a and b (element-wise). Store the results in two new arrays called cubed_a and cubed_b. Print cubed_a and cubed_b.__
cubed_a = a**3
cubed_b = b**3
print(cubed_a)
print(cubed_b)
# __1c) Add the two cubed arrays (e.g., [1,2] + [3,4] = [4,6]) and store the result in an array called c. Print c.__
c = cubed_a + cubed_b
print(c)
# __1d) Sum the elements with even indices of c and store the result in a variable called d. Print d.__
#Assuming the 0th index is even
d_array = [c[i] for i in range(len(c)) if i % 2 == 0]
d = sum(d_array)
print(d)
# __1e) Take the element-wise square root of the c and store the result in an array called e. Print e.__
e = np.sqrt(c)
print(e)
# __1f) Append b to a, reshape the appended array so that it is a 4x5, 2d array and store the results in a variable called m. Print m.__
#
m = np.reshape(np.append(a, b),(4,5))
print(m)
# __1g) Extract the third and the fourth column of the m matrix. Store the resulting 4x2 matrix in a new variable called m2. Print m2.__
#
# +
#Assume the third and the fourth column are 1th indexed
m2 = m[:,[2,3]]
print(m2)
# -
# __1h) Take the dot product of m and m2. Store the results in a matrix called m3. Print m3.
# Note that Dot product of two matrices A$\cdot$B =A<sup>T</sup>B__
m3 = m.T @ m2
print(m3)
# __1i) Round the m3 matrix to three decimal points. Store the result in m4. Print m4.__
#
m4 = np.round(m3, 3)
print(m4)
# ## NumPy and Masks
# __2a) Create an array called f where the values are cosine(x) for x from 0 to pi with 50 equally spaced values (inclusive). Print f.__
f = np.cos(np.linspace(0, np.pi, 50))
print(f)
# __2b) Use a 'mask' to get an array that is True when f >= 1/2 and False when f < 1/2. Store the result in an array called g. Print g.__
g = f >= 0.5
print(g)
# __2c) Create an array called h that has only those values where f>= 1/2. Print h.__
h = f[g]
print(h)
# ## NumPy and 2 Variable Prediction
# __Let 'x' be the number of miles a person drives per day and 'y' be the dollars spent on buying car fuel (per day).__
#
# __We have created 2 numpy arrays each of size 100 that represent x and y.
# x (number of miles) ranges from 1 to 10 with a uniform noise of (0,1/2)
# y (money spent in dollars) will be from 1 to 20 with a uniform noise (0,1)__
#
# +
# seed the random number generator with a fixed value
# Do not delete this cell
np.random.seed(500)
x=np.linspace(1,10,100)+ np.random.uniform(low=0,high=.5,size=100)
y=np.linspace(1,20,100)+ np.random.uniform(low=0,high=1,size=100)
print ('x = ', x)
print ('y = ', y)
# -
# __3a) Find Expected value of x and the expected value of y. Store the results in two variables called ex and ey.__
# +
ex = np.mean(x)
ey = np.mean(y)
print(ex)
print(ey)
# -
# __3b) Find variance of distributions of x and y. Store the results in two variables called varx and vary.__
varx = np.var(x)
vary = np.var(y)
print(varx)
print(vary)
# __3c) Find co-variance of x and y. Store the result in a variable called cov.__
cov = np.mean(x*y) - ex*ey
print(cov)
# __3d) Assuming that number of dollars spent in car fuel is only dependant on the miles driven, by a linear relationship.
# Write code that uses a linear predictor to calculate a predicted value of y for each x
# i.e. y_predicted = f(x) = y0 + kx. Store your predicitions in an array called y_pred. Print y_pred.__
# +
y_0 = ey - cov/np.var(x)*ex
slope = cov/np.var(x)
y_pred = slope*x + y_0
print(y_pred)
# -
# __3e) Put the prediction error into an array called y_error. Print y_error.__
# +
#I assume the error is y_pred-y_actual
y_error = y_pred - y
print(y_error)
# -
# __3f) Write code that calculates the root mean square error (RMSE), that is root of average of y_error squared. Store the result in a variable called rmse. Print rmse.__
rmse = np.sqrt(np.mean(y_error**2))
print(rmse)
# ## Congratulations on completing hw3! Don't forget to click Kernel -> Restart & Run All, and submit this notebook on Gradescope.
|
homeworks/hw03/hw3_numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hamilton-Jacobi Reachibility
# In the following we will analyse the Hamilton-Jacobi Reachibility for a 1-dimensional coupled space, with state $\vec{x}$ and dynamics $\frac{d}{dt} \vec{x}$:
#
# \begin{equation}
# \vec{x} = \begin{pmatrix}x\\v_x\end{pmatrix} = \begin{pmatrix}x_{robot} - x_{pedestrian}\\v_x\end{pmatrix}
# \end{equation}
#
# \begin{equation}
# \frac{d}{dt}\vec{x} = \begin{pmatrix}v_x - d\\u\end{pmatrix}
# \end{equation}
#
# with $u$ being the control input applied by the robot and $d$ the disturbance, the pedestrian with single integrator dynamics. In order to be provably safe we look for the tube in that the robot's state must be in at every point in time so that no matter what the pedestrian does they both will not collide (collision is modelled here as being less or equal than one meter apart from each other).
#
# The Hamilton-Jacobi Reachibility computes the values of a min-max game, in which the robot chooses its actions in order to avoid some set (here a tube around $\Delta x = 0$) as best as possible while the the disturbance does the oppositie, i.e. the disturbance (aka the pedestrian) tries to push the system in the avoid set.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.ioff()
from IPython.display import HTML
import seaborn as sns
# +
import numpy as np
import torch
from mantrap.agents import DoubleIntegratorDTAgent
from mantrap.environment import PotentialFieldEnvironment
from mantrap.modules import HJReachabilityModule
from mantrap.solver import SGradSolver
env = PotentialFieldEnvironment(DoubleIntegratorDTAgent, ego_position=torch.zeros(2), dt=0.4)
env.add_ado(position=torch.tensor([8, 0]), num_modes=1)
env.add_ado(position=torch.tensor([-8, 0]), num_modes=1)
module = HJReachabilityModule(env=env, t_horizon=3, weight=1.0)
# -
# ### Value Function and Gradient Plot (2D slices)
# +
N = 100
plt.close('all')
fig = plt.figure(figsize=(13, 5))
ticks = np.arange(0, N, step=N//10)
# Draw x-vx two-dimensional slice of value function at y = vy = 0.
x_linspace = np.linspace(*env.x_axis, num=N)
v_linspace = np.linspace(*env.ego.speed_limits, num=N)
x_grid, v_grid = np.meshgrid(x_linspace, v_linspace)
x_grid = x_grid.flatten()
v_grid = v_grid.flatten()
xyv_grid = np.stack((x_grid, np.zeros(N**2), v_grid, np.zeros(N**2))).T
value_function = module.value_function(xyv_grid)
plt.subplot(1, 2, 1)
im = plt.imshow(value_function.reshape(N, N))
plt.xlabel("dx[m]")
plt.ylabel("vx[m/s]")
plt.xticks(ticks, np.round(x_linspace, 1)[ticks])
plt.yticks(ticks, np.round(v_linspace, 1)[ticks])
plt.title("value function")
plt.colorbar(im)
# Draw x-y two-dimensional slice of value function at vx = vy = 0.
y_linspace = np.linspace(*env.y_axis, num=N)
x_grid, y_grid = np.meshgrid(x_linspace, y_linspace)
x_grid = x_grid.flatten()
y_grid = y_grid.flatten()
xyv_grid = np.stack((x_grid, y_grid, np.zeros(N**2), np.zeros(N**2))).T
value_function = module.value_function(xyv_grid)
plt.subplot(1, 2, 2)
im = plt.imshow(value_function.reshape(N, N))
plt.xlabel("dx[m]")
plt.ylabel("dy[m]")
plt.xticks(ticks, np.round(x_linspace, 1)[ticks])
plt.yticks(ticks, np.round(y_linspace, 1)[ticks])
plt.title("value function")
plt.colorbar(im)
plt.show()
# -
# ### HJ-Constraint in Optimization
# +
env = PotentialFieldEnvironment(DoubleIntegratorDTAgent, ego_position=torch.zeros(2), dt=0.4)
env.add_ado(position=torch.tensor([3, 0]), num_modes=1)
env.add_ado(position=torch.tensor([-3, 0]), num_modes=1)
solver = SGradSolver(env, goal=torch.rand(2), t_planning=3, modules=[HJReachabilityModule])
HTML(solver.visualize_heat_map(propagation="constant"))
# -
# ### Value Function Approximation
# + pycharm={"name": "#%%\n"}
import os
import mantrap
import numpy as np
import pandas as pd
import torch
t_horizon = 5
env = mantrap.environment.Trajectron(mantrap.agents.DoubleIntegratorDTAgent,
ego_position=torch.zeros(2))
_ = env.add_ado(position=torch.rand(2) * 4, velocity=torch.rand(2))
# + pycharm={"name": "#%%\n"}
file_baseline = "2D_large.mat"
module_baseline = mantrap.modules.HJReachabilityModule(env, t_horizon=t_horizon, data_file=file_baseline)
# Loading accuartely computed value function and compute meshgrid containing every computed grid coordinate.
_, _, grid_size_by_dim, _, (grid_min, grid_max) = mantrap.modules.HJReachabilityModule.unpack_mat_file(
mat_file_path=os.path.join(mantrap.utility.io.build_os_path("third_party/reachability"), file_baseline)
)
x_grid, y_grid, vx_grid, vy_grid = np.meshgrid(*(np.linspace(grid_min[i], grid_max[i], num=grid_size_by_dim[i]) \
for i in range(4)))
mesh = np.stack((x_grid.flatten(), y_grid.flatten(), vx_grid.flatten(), vy_grid.flatten())).T
# In order to reduce the number of evaluation points, sample 1000 random points.
idx = np.random.randint(low=0, high=mesh.shape[0] - 1, size=1000)
mesh = mesh[idx, :]
print(f"Meshgrid with {mesh.shape[0]} elements ready for evaluation !")
# + pycharm={"name": "#%%\n"}
# Evaluate value function error of approximation.
models = {"small": "2D_small.mat", "interior": "2D_interior.mat", "medium": "2D_medium.mat"}
results_df = pd.DataFrame(columns=["model", "error", "x", "y", "vx", "vy"])
for label, file_name in models.items():
print(label, file_name)
module = mantrap.modules.HJReachabilityModule(env, t_horizon=t_horizon, data_file=file_name)
for x_rel in mesh:
error = module.value_function(x_rel) - module_baseline.value_function(x_rel)
results_df = results_df.append({"model": label, "error": float(error),
"x": x_rel[0], "y": x_rel[1], "vx": x_rel[2], "vy": x_rel[3]},
ignore_index=True)
# + pycharm={"name": "#%%\n"}
results_df["log_error"] = np.log(np.abs(results_df["error"]) + 1e-10) # avoiding nan values (log(0)/log(-x))
results_df
# + pycharm={"name": "#%%\n"}
# Plot results as bar plot.
ax = sns.boxplot(x="log_error", y="model", data=results_df)
plt.show()
# + pycharm={"name": "#%%\n"}
ax = sns.pairplot(data=results_df, hue="model", y_vars=["error"], x_vars=["x", "y", "vx", "vy"], diag_kind="kde")
plt.show()
# -
# ax = sns.pairplot(data=results_df, hue="model", y_vars=["error"], x_vars=["x", "y", "vx", "vy"], diag_kind="kde")
# plt.show()
# +
import os
import mantrap
import numpy as np
import pandas as pd
import torch
t_horizon = 5
env = mantrap.environment.Trajectron(mantrap.agents.DoubleIntegratorDTAgent,
ego_position=torch.zeros(2))
_ = env.add_ado(position=torch.rand(2) * 4, velocity=torch.rand(2))
# +
file_baseline = "2D_large.mat"
module_baseline = mantrap.modules.HJReachabilityModule(env, t_horizon=t_horizon, data_file=file_baseline)
# Loading accuartely computed value function and compute meshgrid containing every computed grid coordinate.
_, _, grid_size_by_dim, _, (grid_min, grid_max) = mantrap.modules.HJReachabilityModule.unpack_mat_file(
mat_file_path=os.path.join(mantrap.utility.io.build_os_path("third_party/reachability"), file_baseline)
)
x_grid, y_grid, vx_grid, vy_grid = np.meshgrid(*(np.linspace(grid_min[i], grid_max[i], num=grid_size_by_dim[i]) \
for i in range(4)))
mesh = np.stack((x_grid.flatten(), y_grid.flatten(), vx_grid.flatten(), vy_grid.flatten())).T
# In order to reduce the number of evaluation points, sample 1000 random points.
idx = np.random.randint(low=0, high=mesh.shape[0] - 1, size=1000)
mesh = mesh[idx, :]
print(f"Meshgrid with {mesh.shape[0]} elements ready for evaluation !")
# +
# Evaluate value function error of approximation.
models = {"small": "2D_small.mat", "interior": "2D_interior.mat", "medium": "2D_medium.mat"}
results_df = pd.DataFrame(columns=["model", "error", "x", "y", "vx", "vy"])
for label, file_name in models.items():
print(label, file_name)
module = mantrap.modules.HJReachabilityModule(env, t_horizon=t_horizon, data_file=file_name)
for x_rel in mesh:
error = module.value_function(x_rel) - module_baseline.value_function(x_rel)
results_df = results_df.append({"model": label, "error": float(error),
"x": x_rel[0], "y": x_rel[1], "vx": x_rel[2], "vy": x_rel[3]},
ignore_index=True)
# -
results_df["log_error"] = np.log(np.abs(results_df["error"]) + 1e-10) # avoiding nan values (log(0)/log(-x))
results_df
# Plot results as bar plot.
ax = sns.boxplot(x="log_error", y="model", data=results_df)
plt.show()
ax = sns.pairplot(data=results_df, hue="model", y_vars=["error"], x_vars=["x", "y", "vx", "vy"], diag_kind="kde")
plt.show()
|
examples/modules/module_hjr.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Python - part 2
# Author: <NAME>. This work is licensed under a CC Attribution 3.0 Unported license (http://creativecommons.org/licenses/by/3.0/).
# Original material, "Introduction to Python programming", was created by <NAME> under the CC Attribution 3.0 Unported license (http://creativecommons.org/licenses/by/3.0/) and can be found at https://github.com/jrjohansson/scientific-python-lectures.
# ### Functions
# Functions are reusable blocks of code. A function in Python is defined using the keyword `def` followed by: a function name, a list of parameters within parentheses `()`, and a colon `:`. The following code, with one additional level of indentation, is the function body.
def func0():
print("test")
func0()
# Optionally, we can define a so called "docstring", which is a description of the function. The docstring should follow directly after the function definition, before the code of the function body.
def func1(s):
"""
Print a string 's' and tell how many characters it has
"""
print(s + " has " + str(len(s)) + " characters")
help(func1)
# Functions that returns a value use the `return` keyword:
def square(x):
"""
Return the square of x.
"""
return x ** 2
square(4)
# We can return multiple values from a function using tuples (see above):
def powers(x):
"""
Return a few powers of x.
"""
return x ** 2, x ** 3, x ** 4
powers(3)
# #### Local and global variables
# According to where they are declared, variables can be accessed from anywhere in a program (**global**) or only inside the functions which declared them (**local**). Global variables can be referenced within functions, but they can be modified only if they are declared global inside functions:
# +
x = 1
def addone(number):
return number + x
print(addone(1))
# +
def edit_global_wrong():
x = 10
edit_global_wrong()
print(x)
# +
def edit_global_correct():
global x
x = 10
edit_global_correct()
print(x)
# -
# In contrast, local variables exist only in the functions that declare them:
# +
def somefun():
y = 5
print(y)
print(y)
# -
# #### Passing by value and by reference
# Can you modify the value of a variable inside a function? Most languages distinguish “passing by value” and “passing by reference”. **In Python, parameters of functions are passed by value**.
#
# If the value passed in a function is **immutable**, the function does not modify the caller’s variable. If the value is **mutable**, the function may modify the caller’s variable. Let's see some examples:
# +
# swap two variables
def swap(a, b):
temp = a
a = b
b = temp
print(a)
x = 10
y = 5
swap(x,y)
print(x)
# +
# modify a list
def add_element(local_list):
local_list.append(15)
list = [5,10]
add_element(list)
print(list)
# -
# ## Modules
# Most of the functionality in Python is provided by *modules*. The Python Standard Library is a large collection of modules that provides *cross-platform* implementations of common facilities such as access to the operating system, file I/O, string management, network communication, and much more.
# To use a module in a Python program it first has to be imported. A module can be imported using the `import` statement. For example, to import the module `math`, which contains many standard mathematical functions, we can do:
import math
# This includes the whole module and makes it available for use later in the program. For example, we can do:
# +
import math
x = math.cos(2 * math.pi)
print(x)
# -
# To know more about a module and its contents, use the `help` function:
help(math)
# A complete lists of standard modules for Python 3 are available at http://docs.python.org/3/library/.
# #### Creating modules, main function and modules import
# If we want to write larger and well organized programs, where some objects are defined and reused several times, we have to create our own modules.
#
# Furthermore, sometimes we want code to be executed when a module is run directly, but not when it is imported by another module.
#
# Let us define the following module `main.py`:
# +
def print_b():
print ('b')
def print_a():
print ('a')
# print_b() runs on import
print_b()
if __name__ == '__main__':
# print_a() is only executed when the module is run directly.
print_a()
# -
# To test how this works, create two scripts in the same folder: main.py (aforementioned code) and main2.py, containing:
import main
# You will see that the output is only b, because `print_a()` is called only when the module is executed directly.
# # EXERCISE 3:
# Given an unordered list of numbers, without using any package or built-in function, define functions to (and print results):
# - swap the values of two elements in the list
# - order ascendently the list
# - find mean and median of the list
input = [30,10,40,20,50]
# # EXERCISE 4:
# Given a list of 2-dimensional tuples, without using any package or built-in function, define functions to (and print results):
# - find the Euclidean distance between two tuples t1 and t2
# - find the Euclidean distance between all tuples of the list
# - to compute the coordinates of the centroid of the list of tuples
# TIP: you can use the math module (use `import math`)
input = [(1.0,1.0), (2.0,2.0), (3.0,3.0), (4.0,4.0)]
# This notebook can be found at:
|
notebook/intro-to-python2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import mplfinance as mpf
df = pd.read_csv('C:\\Users\\KIIT\\Desktop\\AMZN.csv')
df
df.info()
df.describe()
df.columns
df.Date = pd.to_datetime(df.Date)
df.info()
df = df.set_index('Date')
df
mpf.plot(df)
mpf.plot(df, type='line',volume=True)
mpf.plot(df['2020-03'],volume=True)
mpf.plot(df['2020-03'],type='candle',volume=True)
mpf.plot(df['2020-03':'2020-07'],type='candle',mav=(20),volume=True)
mpf.plot(df['2020-03':'2020-07'],type='candle',mav=(20),volume=True,tight_layout=True)
mpf.plot(df['2020-03':'2020-07'],figratio=(20,12),title='Amazon Price 2019/20',
type='candle',mav=(20),volume=True,
tight_layout=True)
mpf.plot(df['2020-03':'2020-07'],figratio=(20,12),title='Amazon Price 2019/20',
type='candle',mav=(20),volume=True,
tight_layout=True,style='yahoo')
# ## We analysed Amazon price dataset and made candlestick chart from month of march 2020 to july 2020.
|
Amazon price Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="YX1hxqUQn47M"
# ## PyTorch/XLA ResNet50/CIFAR10 (GPU or TPU)
#
# + [markdown] id="pLQPoJ6Fn8wF"
# ### [RUNME] Install Colab compatible PyTorch/XLA wheels and dependencies
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="O53lrJMDn9Rd" outputId="0645a7e9-9c26-4f6b-ef22-41ccda94997a"
# !pip install cloud-tpu-client==0.10 https://storage.googleapis.com/tpu-pytorch/wheels/torch_xla-1.8.1-cp37-cp37m-linux_x86_64.whl
# + [markdown] id="IednejwkIW-K"
# Only run the below commented cell if you would like a nightly release
# + id="xiFzLg5gy7l6"
# PyTorch/XLA GPU Setup (only if GPU runtime)
import os
from datetime import datetime
if os.environ.get('COLAB_GPU', '0') == '1':
os.environ['GPU_NUM_DEVICES'] = '1'
os.environ['XLA_FLAGS'] = '--xla_gpu_cuda_data_dir=/usr/local/cuda/'
# + [markdown] id="rroH9yiAn-XE"
# ### Define Parameters
#
#
# + id="cMojPWZUqr2s"
# Result Visualization Helper
from matplotlib import pyplot as plt
M, N = 4, 6
RESULT_IMG_PATH = '/tmp/test_result.jpg'
CIFAR10_LABELS = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
def plot_results(images, labels, preds):
images, labels, preds = images[:M*N], labels[:M*N], preds[:M*N]
inv_norm = transforms.Normalize(
mean=(-0.4914/0.2023, -0.4822/0.1994, -0.4465/0.2010),
std=(1/0.2023, 1/0.1994, 1/0.2010))
num_images = images.shape[0]
fig, axes = plt.subplots(M, N, figsize=(16, 9))
fig.suptitle('Correct / Predicted Labels (Red text for incorrect ones)')
for i, ax in enumerate(fig.axes):
ax.axis('off')
if i >= num_images:
continue
img, label, prediction = images[i], labels[i], preds[i]
img = inv_norm(img)
img = img.permute(1, 2, 0) # (C, M, N) -> (M, N, C)
label, prediction = label.item(), prediction.item()
if label == prediction:
ax.set_title(u'\u2713', color='blue', fontsize=22)
else:
ax.set_title(
'X {}/{}'.format(CIFAR10_LABELS[label],
CIFAR10_LABELS[prediction]), color='red')
ax.imshow(img)
plt.savefig(RESULT_IMG_PATH, transparent=True)
# + id="iMdPRFXIn_jH"
# Define Parameters
FLAGS = {}
FLAGS['data_dir'] = "/tmp/cifar"
FLAGS['batch_size'] = 128
FLAGS['num_workers'] = 1
FLAGS['learning_rate'] = 0.01
FLAGS['momentum'] = 0.9
FLAGS['num_epochs'] = 350
FLAGS['num_cores'] =8 if os.environ.get('TPU_NAME', None) else 1
FLAGS['log_steps'] = 20
FLAGS['metrics_debug'] = False
# + colab={"base_uri": "https://localhost:8080/"} id="OTc37n5z4J9o" outputId="d225bfaf-53a3-43d6-e605-4a9506814e17"
print(FLAGS)
# + id="Micd3xZvoA-c"
import numpy as np
import os
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.optim import lr_scheduler
import torch_xla
import torch_xla.core.xla_model as xm
import torch_xla.debug.metrics as met
import torch_xla.distributed.parallel_loader as pl
import torch_xla.distributed.xla_multiprocessing as xmp
import torch_xla.utils.utils as xu
import torchvision
from torchvision import datasets, transforms
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, in_planes, planes, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = nn.Conv2d(
in_planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3,
stride=1, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, in_planes, planes, stride=1):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3,
stride=stride, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv3 = nn.Conv2d(planes, self.expansion *
planes, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm2d(self.expansion*planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = F.relu(self.bn2(self.conv2(out)))
out = self.bn3(self.conv3(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class ResNet(nn.Module):
def __init__(self, block, num_blocks, num_classes=10):
super(ResNet, self).__init__()
self.in_planes = 64
self.conv1 = nn.Conv2d(3, 64, kernel_size=3,
stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.layer1 = self._make_layer(block, 64, num_blocks[0], stride=1)
self.layer2 = self._make_layer(block, 128, num_blocks[1], stride=2)
self.layer3 = self._make_layer(block, 256, num_blocks[2], stride=2)
self.layer4 = self._make_layer(block, 512, num_blocks[3], stride=2)
self.linear = nn.Linear(512*block.expansion, num_classes)
def _make_layer(self, block, planes, num_blocks, stride):
strides = [stride] + [1]*(num_blocks-1)
layers = []
for stride in strides:
layers.append(block(self.in_planes, planes, stride))
self.in_planes = planes * block.expansion
return nn.Sequential(*layers)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
def ResNet18():
return ResNet(BasicBlock, [2, 2, 2, 2])
def ResNet34():
return ResNet(BasicBlock, [3, 4, 6, 3])
def ResNet50():
return ResNet(Bottleneck, [3, 4, 6, 3])
def ResNet101():
return ResNet(Bottleneck, [3, 4, 23, 3])
def ResNet152():
return ResNet(Bottleneck, [3, 8, 36, 3])
modelToTrain = ResNet50()
# + id="8vMl96KLoCq8"
SERIAL_EXEC = xmp.MpSerialExecutor()
# Only instantiate model weights once in memory.
WRAPPED_MODEL = xmp.MpModelWrapper(modelToTrain)
def train_resnet():
torch.manual_seed(1)
def get_dataset():
norm = transforms.Normalize(
mean=(0.4914, 0.4822, 0.4465), std=(0.2023, 0.1994, 0.2010))
transform_train = transforms.Compose([
transforms.ColorJitter(brightness=0.1, contrast=0.1, saturation=0.1, hue=0.1),
transforms.RandomHorizontalFlip(),
transforms.RandomCrop(32, padding=4),
transforms.RandomAffine(degrees=5),
transforms.ToTensor(),
norm,
])
transform_test = transforms.Compose([
transforms.ToTensor(),
norm,
])
train_dataset = datasets.CIFAR10(
root=FLAGS['data_dir'],
train=True,
download=True,
transform=transform_train)
test_dataset = datasets.CIFAR10(
root=FLAGS['data_dir'],
train=False,
download=True,
transform=transform_test)
return train_dataset, test_dataset
# Using the serial executor avoids multiple processes
# to download the same data.
train_dataset, test_dataset = SERIAL_EXEC.run(get_dataset)
train_sampler = torch.utils.data.distributed.DistributedSampler(
train_dataset,
num_replicas=xm.xrt_world_size(),
rank=xm.get_ordinal(),
shuffle=True)
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=FLAGS['batch_size'],
sampler=train_sampler,
num_workers=FLAGS['num_workers'],
drop_last=True)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=FLAGS['batch_size'],
shuffle=False,
num_workers=FLAGS['num_workers'],
drop_last=True)
# Scale learning rate to num cores
learning_rate = FLAGS['learning_rate'] * xm.xrt_world_size()
# Get loss function, optimizer, and model
device = xm.xla_device()
model = WRAPPED_MODEL.to(device)
optimizer = optim.SGD(model.parameters(), lr=learning_rate, momentum=FLAGS['momentum'], weight_decay=5e-4)
scheduler = lr_scheduler.CosineAnnealingLR(optimizer, T_max=350)
loss_fn = nn.CrossEntropyLoss()
def train_loop_fn(loader):
tracker = xm.RateTracker()
model.train()
for x, (data, target) in enumerate(loader):
optimizer.zero_grad()
output = model(data)
loss = loss_fn(output, target)
loss.backward()
xm.optimizer_step(optimizer)
tracker.add(FLAGS['batch_size'])
# if x % FLAGS['log_steps'] == 0:
# print('[xla:{}]({}) Loss={:.5f} Rate={:.2f} GlobalRate={:.2f} Time={}'.format(
# xm.get_ordinal(), x, loss.item(), tracker.rate(),
# tracker.global_rate(), time.asctime()), flush=True)
def test_loop_fn(loader):
total_samples = 0
correct = 0
model.eval()
data, pred, target = None, None, None
for data, target in loader:
output = model(data)
pred = output.max(1, keepdim=True)[1]
correct += pred.eq(target.view_as(pred)).sum().item()
total_samples += data.size()[0]
accuracy = 100.0 * correct / total_samples
print('[xla:{}] Accuracy={:.2f}%'.format(
xm.get_ordinal(), accuracy), flush=True)
return accuracy, data, pred, target
# Train and eval loops
accuracy = 0.0
data, pred, target = None, None, None
for epoch in range(1, FLAGS['num_epochs'] + 1):
para_loader = pl.ParallelLoader(train_loader, [device])
train_loop_fn(para_loader.per_device_loader(device))
xm.master_print("{}({:.0f}), Finished training epoch {}".format(datetime.now().strftime("%H:%M:%S"), time.time(), epoch))
scheduler.step()
para_loader = pl.ParallelLoader(test_loader, [device])
accuracy, data, pred, target = test_loop_fn(para_loader.per_device_loader(device))
if FLAGS['metrics_debug']:
xm.master_print(met.metrics_report(), flush=True)
return accuracy, data, pred, target
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["3ed5e4d997f34e6db84a248a7e24c160", "8a82aa441dd2461289aac342ecfb2f71", "62a5eef180c44eb2abcd7870488cc440", "6f5b9f7319f247a39223f7d9032bd0ee", "2842e31e03ec46b4ade0441ed814e786", "20b3704d7f1e400ea9f9dc1326244c6d", "1d139abaf9434f358e1423e0d96c8db4", "86d276374f384969a4290398afef442b"]} id="_2nL4HmloEyl" outputId="32a8ee79-4659-4ef5-e7a2-b6bedf368860"
# Start training processes
def _mp_fn(rank, flags):
global FLAGS
FLAGS = flags
torch.set_default_tensor_type('torch.FloatTensor')
accuracy, data, pred, target = train_resnet()
if rank == 0:
# Retrieve tensors that are on TPU core 0 and plot.
plot_results(data.cpu(), pred.cpu(), target.cpu())
print("{}({:.0f}), Staring training".format(datetime.now().strftime("%H:%M:%S"), time.time()))
xmp.spawn(_mp_fn, args=(FLAGS,), nprocs=FLAGS['num_cores'], start_method='fork')
# + id="Rj-Xj2W9E1YT"
model_path = "result1.pth"
torch.save(modelToTrain.state_dict(), model_path)
# + id="Uw2GtnFCKg_C"
model_path = "result2.pth"
torch.save(model.state_dict(), model_path)
|
Assignment5/question5.1_TPU1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
# # Logistic Regression Project
#
# In this project we will be working with a fake advertising data set, indicating whether or not a particular internet user clicked on an Advertisement on a company website. We will try to create a model that will predict whether or not they will click on an ad based off the features of that user.
#
# This data set contains the following features:
#
# * 'Daily Time Spent on Site': consumer time on site in minutes
# * 'Age': cutomer age in years
# * 'Area Income': Avg. Income of geographical area of consumer
# * 'Daily Internet Usage': Avg. minutes a day consumer is on the internet
# * 'Ad Topic Line': Headline of the advertisement
# * 'City': City of consumer
# * 'Male': Whether or not consumer was male
# * 'Country': Country of consumer
# * 'Timestamp': Time at which consumer clicked on Ad or closed window
# * 'Clicked on Ad': 0 or 1 indicated clicking on Ad
#
# ## Import Libraries
#
# **Import a few libraries you think you'll need (Or just import them as you go along!)**
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# ## Get the Data
# **Read in the advertising.csv file and set it to a data frame called ad_data.**
ad_data = pd.read_csv('advertising.csv')
# **Check the head of ad_data**
ad_data.head()
# ** Use info and describe() on ad_data**
ad_data.info()
ad_data.describe()
# ## Exploratory Data Analysis
#
# Let's use seaborn to explore the data!
#
# Try recreating the plots shown below!
#
# ** Create a histogram of the Age**
sns.set_style('whitegrid')
ad_data['Age'].hist(bins=30)
plt.xlabel('Age')
# **Create a jointplot showing Area Income versus Age.**
sns.jointplot(x='Age',y='Area Income',data=ad_data)
# **Create a jointplot showing the kde distributions of Daily Time spent on site vs. Age.**
sns.jointplot(x='Age',y='Daily Time Spent on Site',data=ad_data,color='red',kind='kde');
# ** Create a jointplot of 'Daily Time Spent on Site' vs. 'Daily Internet Usage'**
sns.jointplot(x='Daily Time Spent on Site',y='Daily Internet Usage',data=ad_data,color='green')
# ** Finally, create a pairplot with the hue defined by the 'Clicked on Ad' column feature.**
sns.pairplot(ad_data,hue='Clicked on Ad',palette='bwr')
# # Logistic Regression
#
# Now it's time to do a train test split, and train our model!
#
# You'll have the freedom here to choose columns that you want to train on!
# ** Split the data into training set and testing set using train_test_split**
from sklearn.model_selection import train_test_split
X = ad_data[['Daily Time Spent on Site', 'Age', 'Area Income','Daily Internet Usage', 'Male']]
y = ad_data['Clicked on Ad']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# ** Train and fit a logistic regression model on the training set.**
from sklearn.linear_model import LogisticRegression
logmodel = LogisticRegression()
logmodel.fit(X_train,y_train)
# ## Predictions and Evaluations
# ** Now predict values for the testing data.**
predictions = logmodel.predict(X_test)
# ** Create a classification report for the model.**
from sklearn.metrics import classification_report
print(classification_report(y_test,predictions))
# # Reference
# _______
# http://www.pieriandata.com
|
Logistic Regression Advertising.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="lQBaanaBhuh3" colab_type="text"
# XGBoost(eXtra Gradient Boost) 주요 장점
# - 뛰어난 예측 성능
# - GBM 대비 빠른 수행 시간
# - CPU 병렬 처리, GPU 지원
# - 다양한 성능 향상 기능
# - 규제(Regularization) 기능 탑재
# - Tree Pruning (가지치기)
# - 다양한 편의 기능
# - 조기 중단(Early Stopping)
# - 자체 내장된 교차 검증
# - 결손값 자체 처리
# - 조기 중단 기능 (Early Stopping)
# - 특정 반복 횟수 만큼 더 이상 비용 함수가 감소하지 않으면 지정된 반복횟수를 다 완료하지 않고 수행을 종료할 수 있음
# - 학습을 위한 시간을 단축 시킬 수 있음
# - 반복 횟수를 너무 단축할 경우 예측 성능 최적화가 안된 상태에서 학습이 종료 될 수 있음
# - 주요 파라미터
# - early_stopping_rounds : 더 이상 비용 평가 지표가 감소하지 않는 최대 반복횟수
# - eval_metric : 반복 수행 시 사용하는 비용 평가 지표
# - eval_set : 평가를 수행하는 별도의 검증 데이터 세트. 일반적으로 검증 데이터 세트에서 반복적으로 비용 감소 성능 평가
# + [markdown] id="5dMfEL0UwcK-" colab_type="text"
# Python wrapper XGBoost
# + id="DplQtea7mOpX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 196} outputId="08046dda-6e11-4bec-b792-67319740a64b"
import xgboost as xgb
from xgboost import plot_importance
import pandas as pd
import numpy as np
from sklearn.datasets import load_breast_cancer
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
dataset = load_breast_cancer()
X_features = dataset.data
y_label = dataset.target
cancer_df = pd.DataFrame(data=X_features, columns=dataset.feature_names)
cancer_df['target'] = y_label
cancer_df.head(3)
# + id="EaWPdyrqnl_K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="d7dbb3ef-b6c5-4866-c97d-610a38448ceb"
print(dataset.target_names)
print(cancer_df['target'].value_counts())
# + id="JjgoqY7Ir11X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f900121f-d622-438a-93a5-ad651fc56900"
X_train, X_test, y_train, y_test = train_test_split(X_features, y_label, test_size=0.2, random_state=156)
print(X_train.shape, X_test.shape)
# + id="V0VjezT-tMo5" colab_type="code" colab={}
dtrain = xgb.DMatrix(data=X_train, label=y_train)
dtest = xgb.DMatrix(data=X_test, label=y_test)
# + id="XbiLtCnGtWaX" colab_type="code" colab={}
params = {
'max_depth':3,
'eta':0.1, # learning rate
'objective':'binary:logistic', # 결정함수
'eval_metric':'logloss', # 손실 측정
'early_stoppings':100 }
num_rounds = 400
# + id="iVcnrrpgttwf" colab_type="code" colab={}
# train data set : 'train', evaluation(test) data set : 'eval'
wlist = [(dtrain, 'train'), (dtest, 'eval')]
# 하이퍼 파라미터와 early stopping 파라미터를 train() 함수의 파라미터로 전달
xgb_model = xgb.train(params = params, dtrain=dtrain, num_boost_round=num_rounds, evals=wlist)
# + id="eAfVAJJJut3E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="03f16061-f68f-4d6d-e970-6a50337d7b43"
pred_probs = xgb_model.predict(dtest)
print('predict() 수행 결과값을 10개만 표시, 예측 확률 값으로 표시됨')
print(np.round(pred_probs[:10], 3))
# 예측 확률이 0.5 보다 크면 1, 그렇지 않으면 0으로 예측값 결정하여 List 객체인 preds에 저장
preds = [1 if x > 0.5 else 0 for x in pred_probs]
print('예측값 10개만 표시:', preds[:10])
# + id="NpVyIe7PvND7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="a69de845-6ac5-43d7-f7cd-5771ede22af0"
from sklearn.metrics import confusion_matrix, accuracy_score, precision_score, recall_score
from sklearn.metrics import f1_score, roc_auc_score
def get_clf_eval(y_test, pred):
confusion = confusion_matrix(y_test, pred)
accuracy = accuracy_score(y_test, pred)
precision = precision_score(y_test, pred)
recall = recall_score(y_test, pred)
f1 = f1_score(y_test, pred)
roc_auc = roc_auc_score(y_test, pred)
print('오차 행렬')
print(confusion)
print('정확도 : {0:.4f}, 정밀도 : {1:.4f}, 재현율 : {2:.4f}, F1 : {3:.4f},\
AUC : {4:.4f}'.format(accuracy, precision, recall, f1, roc_auc))
get_clf_eval(y_test, preds)
# + id="DcnVBB5QwIf5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="1b035d4e-b935-476a-c760-d9662ca50b6d"
import matplotlib.pyplot as plt
# %matplotlib inline
fig, ax = plt.subplots(figsize=(10, 12))
plot_importance(xgb_model, ax=ax)
# + [markdown] id="hXjQqsPmwgLy" colab_type="text"
# sklearn Wrapper XGBoost 적용
# + id="A2rLOeoPwmhc" colab_type="code" colab={}
from xgboost import XGBClassifier
evals = [(X_test, y_test)]
xgb_wrapper = XGBClassifier(n_estimators=400, learning_rate=0.1, max_depth=3)
xgb_wrapper.fit(X_train, y_train, early_stopping_rounds=400, eval_set=evals, eval_metric='logloss', verbose=True)
w_preds = xgb_wrapper.predict(X_test)
# + id="HK2p_8YbxA8k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="6df6c55e-ebd6-4f94-a6a1-a9f4050ff6af"
get_clf_eval(y_test, w_preds)
# + id="BHYFrA7xx7ZB" colab_type="code" colab={}
from xgboost import XGBClassifier
xgb_wrapper = XGBClassifier(n_estimators=400, learning_rate=0.1, max_depth=3)
evals = [(X_test, y_test)]
xgb_wrapper.fit(X_train, y_train, early_stopping_rounds=100, eval_metric='logloss', eval_set=evals, verbose=True)
ws100_preds = xgb_wrapper.predict(X_test)
# + id="csRKcCA-yljq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="04354c52-bd3a-4377-843b-81a95413f706"
get_clf_eval(y_test, ws100_preds)
# + colab_type="code" id="Io9xe4gQzkb5" colab={}
# early_stopping_rounds를 10으로 설정하고 재학습.
xgb_wrapper.fit(X_train, y_train, early_stopping_rounds=10, eval_metric='logloss', eval_set=evals, verbose=True)
ws10_preds = xgb_wrapper.predict(X_test)
get_clf_eval(y_test, ws10_preds)
# + id="mjRVKUJkzNdb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="386a7216-78bb-446a-b0d6-82236c258149"
from xgboost import plot_importance
import matplotlib.pyplot as plt
# %matplotlib inline
fig, ax = plt.subplots(figsize=(10, 12))
plot_importance(xgb_wrapper, ax=ax)
|
ml/sklearn/ensemble/XGBoost.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extract Data from SQL Databases
#
# In this workbook, you'll gain experience extracting data from SQL databases. This is an overview of Python tools and assumes you already have experience writing SQL queries.
# # Pandas and sqlite3
#
# You can use Pandas to open a SQL database or to run a SQL query against a database. __There is more than one way to do this depending on the type of SQL database you are working with__: the [sqlite3 library](https://www.sqlite.org/about.html) or the [sqlalchemy library](https://www.sqlalchemy.org/). *The sqlite3 library can handle sqlite databases. SQLAlchemy can handle sqlite3 as well as other types of SQL databases like MySQL and PostgreSQL.*
#
# In the same folder as this Jupyter notebook, **there is a _SQLite database_ file called "population_data.db".** SQLite is a database engine meant for single applications. The entire database is contained in one file. You can read more about SQLite [here](https://www.sqlite.org/whentouse.html).
#
# In this example, the "population_data.db" database contains only one table called "population_data". Run the code in the following cells to see how to use a SQLite database with pandas. (If you're curious how the data was converted from a csv file to a database, go to File->Open and click on create_db.py).
# # SQLite3 and Pandas Example
# +
import sqlite3
import pandas as pd
# connect to the database
conn = sqlite3.connect('population_data.db')
# run a query
pd.read_sql('SELECT * FROM population_data', conn)
# -
pd.read_sql('SELECT "Country_Name", "Country_Code", "1960" FROM population_data', conn)
# # SQLAlchemy and Pandas example
#
# If you are working with a different type of database such as MySQL or PostgreSQL, you can use the SQLAlchemy library with pandas. Here are the instructions for connecting to [different types of databases using SQLAlchemy](http://docs.sqlalchemy.org/en/latest/core/engines.html).
#
# Run the code below to see how to connect to the population_data.db database.
# +
###
# create a database engine
# to find the correct file path, use the python os library:
# import os
# print(os.getcwd())
#
###
import os
print(os.getcwd())
# -
from sqlalchemy import create_engine
'''
# Unix/Mac - 4 initial slashes in total
engine = create_engine('sqlite:////absolute/path/to/foo.db')
# Windows
engine = create_engine('sqlite:///C:\\path\\to\\foo.db')
# Windows alternative using raw string
engine = create_engine(r'sqlite:///C:\path\to\foo.db')
'''
# +
## https://stackoverflow.com/questions/19260067/sqlalchemy-engine-absolute-path-url-in-windows
file = "sqlite:///C:\\Users\\DJ\\Documents\\git\\DSND_Term2\\lessons\\5 ETLPipelines\\3_sql_exercise\\population_data.db"
# -
file = "sqlite:///C:/Users/DJ/Documents/git/DSND_Term2/lessons/5 ETLPipelines/3_sql_exercise/population_data.db"
engine = create_engine(file)
pd.read_sql("SELECT * FROM population_data", engine)
# # Exercise
#
# Connect to the population_data.db SQLite database, and answer the following questions:
# 1. Write a query that finds the change in population from 1960 to 1961 in Aruba
# 2. Write a query that finds the population of Belgium and also Luxembourg in 1975. The output should have two rows.
#
# There is a solution if you go to File->Open->3_sql_exercise_solution.ipynb
# +
import sqlite3
import pandas as pd
# connect to the database
conn = sqlite3.connect('population_data.db')
# run a query
pd.read_sql('SELECT Country_Name, ("1961" - "1960") as diff_1961_1960 FROM population_data where Country_Name = "Aruba"', conn)
# -
# run a query
pd.read_sql('SELECT Country_Name, "1975" FROM population_data where Country_Name in ("Belgium", "Luxembourg")', conn)
|
lessons/5 ETLPipelines/3_sql_exercise/3_sql_exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Truncation Error vs Rounding Error
#
# Copyright (C) 2010-2020 <NAME><br>
# Copyright (C) 2020 <NAME>
#
# <details>
# <summary>MIT License</summary>
# Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN
# THE SOFTWARE.
# </details>
# In this notebook, we'll investigate two common sources of error: Truncation error and rounding error.
# + jupyter={"outputs_hidden": false}
import numpy as np
import matplotlib.pyplot as pt
# -
# **Task:** Approximate a function (here: a parabola, by a line)
# + jupyter={"outputs_hidden": false}
center = -1
width = 6
def f(x):
return - x**2 + 3*x
def df(x):
return -2*x + 3
grid = np.linspace(center-width/2, center+width/2, 100)
fx = f(grid)
pt.plot(grid, fx)
pt.plot(grid, f(center) + df(center) * (grid-center))
pt.xlim([grid[0], grid[-1]])
pt.ylim([np.min(fx), np.max(fx)])
# -
# * What's the error we see?
# * What if we make `width` smaller?
# + jupyter={"outputs_hidden": true}
|
demos/error_and_fp/Truncation vs Rounding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Trapezoid rule: f(x)=e2xcos(10x)
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# defining my function:
def func(x):
a=2
b=10
return np.e**(a*x) * np.cos(b*x)
# find the integral
def func_integral(x):
a=2
b=10
return (np.e**(a*x))/52 *(5*np.sin(b*x)+np.cos(b*x))
def trapezoid_core(f,x,h):
return 0.5*h*(f(x+h)+f(x))
def trapezoid_method(f,a,b,N):
#f===function to integrate
#a===lower limit of integration
#b===upper limit of integration
#N===number of function evaluations to use
#define x values to perform trap rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
#define the value of the integral
Fint = 0.0
#perform the integral using the trapezoid method
for i in range(0,len(x)-1,1):
Fint += trapezoid_core(f,x[i],h)
#return the answer
return Fint
def simpson_core(f,x,h):
return h*( f(x) + 4*f(x+h) + f(x+2*h))/3
def simpsons_method(f,a,b,N):
#f===function to integrate
#a===lower limit of integration
#b===upper limit of integration
#N===number of function evaluations to use
#note the number of chunks will be N-1
#if N odd --> then we dont need to adjust the last segment
#define x values to perform simps rule
x = np.linspace(a,b,N)
h = x[1]-x[0]
#define the value of the integral
Fint = 0.0
#perform the integral using the trapezoid method
for i in range(0,len(x)-2,2):
Fint += simpson_core(f,x[i],h)
#apply simps rule over last interval if N even
if ((N%2)==0):
Fint +=simpson_core(f,x[-2], 0.5*h)
#return the answer
return Fint
def romberg_core(f,a,b,i):
#we need the difference b-a
h = b-a
#and the increment between new fuc evals
dh = h/2.**(i)
#we need the cofactor
K = h/2.**(i+1)
#and the function evals
M = 0.0
for j in range(2**i):
M += f(a + 0.5*dh + j*dh)
#return the answer
return K*M
def romberg_integration(f,a,b,tol):
#define an iteration variable
i = 0
#define a max # of iterations
imax=1000
#define an error estimate, set to a large value
delta = 100.0*np.fabs(tol)
#set an array of integral answers
I = np.zeros(imax,dtype=float)
#get the zeroth romber iteration
I[0] = 0.5*(b-a)*(f(a)+f(b))
#iterate by 1
i += 1
while(delta>tol):
#find this romberg iteration
I[i] = 0.5*I[i-1]+romberg_core(f,a,b,i)
#compute new fractional error estimate
delta=np.fabs( (I[i]-I[i-1])/I[i])
print(i,I[i],I[i-1],delta)
if(delta>tol):
i+=1
if(i>imax):
print("max iterations reached")
raise StopIteration('stopping iterations after',i)
return I[i]
# +
Answer = func_integral(np.pi)-func_integral(0)
print(Answer)
print("Trapezoid")
N = 2
tolerance = True
while(tolerance):
if(np.absolute(trapezoid_method(func,0,np.pi,N)-Answer)<1.0e-6):
tolerance= False
N+=16
print(N)
print(trapezoid_method(func,0,np.pi,N))
print("Simpson's Method")
N = 3
tolerance = True
while(tolerance):
if(np.absolute(simpsons_method(func,0,np.pi,N)-Answer)<1.0e-6):
tolerance= False
N+=1
print(N)
print(simpsons_method(func,0,np.pi,N))
print("Romberg")
tolerance = 1.0e-6
RI=romberg_integration(func,0,np.pi,tolerance)
print(RI, (RI-Answer)/Answer, tolerance)
# -
# ### It takes 23 iterations or Romberg integration to achieve the accuracy requested. It takes 29,682 intervals to achieve the level of accuracy. The takes 646 intervals to achieve the level of accuracy.
#
|
hw_4_astr_119.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbsphinx="hidden"
# # The Discrete-Time Fourier Transform
#
# *This Jupyter notebook is part of a [collection of notebooks](../index.ipynb) in the bachelors module Signals and Systems, Comunications Engineering, Universität Rostock. Please direct questions and suggestions to [<EMAIL>](mailto:<EMAIL>).*
# -
# ## Properties
#
# The discrete-time Fourier transform (DTFT) has a number of specific properties that are reviewed in the following.
# ### Invertibility
#
# For many types of signals it is possible to recover the discrete signal $x[k]$ from its DTFT $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$
#
# \begin{equation}
# x[k] = \mathcal{F}_*^{-1} \left\{ \mathcal{F}_* \{ x[k] \} \right\}
# \end{equation}
#
# A sufficient condition for the theorem to hold is that both the signal $x[k]$ and its DTFT are absolutely summable/integrable. For this type of signals, above relation can be proven by applying the definition of the DTFT and its inverse and rearranging terms.
# **Example**
#
# The invertibility of the DTFT is illustrated at the example of the [complex exponential signal](../discrete_signals/standard_signals.ipynb#Complex-Exponential-Signal) $x[k] = e^{j \Omega_0 k}$ [whose DTFT is given as](definition.ipynb#Transformation-of-the-Exponential-Signal) $X(j \omega) = {\bot \!\! \bot \!\! \bot} ( \frac{\Omega - \Omega_0}{2 \pi} )$. Note that the signal nor its spectrum are absolutely integrable. However, the invertibilty still holds as is shown by evaluating the [integral of the inverse DTFT](definition.ipynb#Definition). Since the integration is only performed in the range $\Omega = -\pi$ to $\pi$, it is sufficient to consider a single Dirac impulse $2 \pi \cdot \delta(\Omega - \Omega_0)$ instead of the Dirac comb for the computation.
# +
# %matplotlib inline
import sympy as sym
sym.init_printing()
k = sym.symbols('k', integer=True)
W, W0 = sym.symbols('Omega Omega0', real=True)
X = 2*sym.pi*sym.DiracDelta(W - W0)
x = 1/(2*sym.pi) * sym.integrate(X * sym.exp(sym.I*W*k), (W, -sym.pi, sym.pi))
x
# -
# This result includes the restriction of the normalized angular frequency to $-\pi < \Omega_0 < \pi$ due to the usage of a single Dirac impulse instead of the Dirac comb. The result is specialized to $\Omega_0 = \frac{1}{2}$ in order to show that above result indeed constitutes a complex exponential signal.
x.subs(W0, sym.S.Half)
# ### Linearity
#
# The DTFT is a linear operation. For two signals $x_1[k]$ and $x_2[k]$ with transforms $X_1(e^{j \Omega}) = \mathcal{F}_* \{ x_1[k] \}$ and $X_2(e^{j \Omega}) = \mathcal{F}_* \{ x_2[k] \}$ the following holds
#
# \begin{equation}
# \mathcal{F}_* \{ A \cdot x_1[k] + B \cdot x_2[k] \} = A \cdot X_1(e^{j \Omega}) + B \cdot X_2(e^{j \Omega})
# \end{equation}
#
# with $A, B \in \mathbb{C}$. The DTFT of a weighted superposition of discrete signals is equal to the weighted superposition of the individual DTFTs. This property is useful to derive the DTFT of signals that can be expressed as superposition of other signals for which the DTFT is known or can be calculated easier. Linearity holds also for the inverse DTFT.
# #### Transformation of the cosine and sine signal
#
# The DTFT of $\cos(\Omega_0 k)$ and $\sin(\Omega_0 k)$ is derived by expressing both as harmonic exponential signals using [Euler's formula](https://en.wikipedia.org/wiki/Euler's_formula)
#
# \begin{align}
# \cos(\Omega_0 k) &= \frac{1}{2} \left( e^{-j \Omega_0 k} + e^{j \Omega_0 k} \right) \\
# \sin(\Omega_0 k) &= \frac{j}{2} \left( e^{-j \Omega_0 k} - e^{j \Omega_0 k} \right)
# \end{align}
#
# together with the DTFT $\mathcal{F}_* \{ e^{j \Omega_0 k} \} = {\bot \!\! \bot \!\! \bot} ( \frac{\Omega - \Omega_0}{2 \pi} )$ of the complex exponential signal yields
#
# \begin{align}
# \mathcal{F} \{ \cos(\Omega_0 k) \} &= \frac{1}{2} \left[ {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega + \Omega_0}{2 \pi} \right) + {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega - \Omega_0}{2 \pi} \right) \right] \\
# \mathcal{F} \{ \sin(\Omega_0 k) \} &= \frac{j}{2} \left[ {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega + \Omega_0}{2 \pi} \right) - {\bot \!\! \bot \!\! \bot} \left( \frac{\Omega - \Omega_0}{2 \pi} \right) \right]
# \end{align}
# ### Symmetries
#
# In order to investigate the symmetries of the DTFT $X(e^{j \Omega}) = \mathcal{F}_* \{ x[k] \}$ of a signal $x[k]$, first the case of a real valued signal $x[k] \in \mathbb{R}$ is considered. The results are then generalized to complex signals $x[k] \in \mathbb{C}$.
# #### Real valued signals
#
# Decomposing a real valued signal $x[k] \in \mathbb{R}$ into its even and odd part $x[k] = x_\text{e}[k] + x_\text{o}[k]$ and introducing these into the definition of the DTFT yields
#
# \begin{align}
# X(e^{j \Omega}) &= \sum_{k = -\infty}^{\infty} \left( x_\text{e}[k] + x_\text{o}[k] \right) e^{-j \Omega k} \\
# &= \sum_{k = -\infty}^{\infty} \left( x_\text{e}[k] + x_\text{o}[k] \right) \cdot \left( \cos[\Omega k] - j \sin[\Omega k] \right) \\
# &= \underbrace{\sum_{k = -\infty}^{\infty} x_\text{e}[k] \cos[\Omega k]}_{X_\text{e}(e^{j \Omega})} +
# j \underbrace{\sum_{k = -\infty}^{\infty} - x_\text{o}[k] \sin[\Omega k] }_{X_\text{o}(e^{j \Omega})}
# \end{align}
#
# For the last equality the fact was exploited that an infinite series with symmetric limits is zero for odd functions. In order to conclude on the symmetry of $X(e^{j \Omega})$ its behavior for a reverse of the sign of $\Omega$ has to be investigated. Due to the symmetry properties of $\cos[\Omega k]$ and $\sin[\Omega k]$, it follows that the DTFT of the
#
# * even part $x_\text{e}[k]$ is real valued with even symmetry $X_\text{e}(e^{j \Omega}) = X_\text{e}(e^{-j \Omega})$
# * odd part $x_\text{o}[k]$ is imaginary with odd symmetry $X_\text{o}(e^{j \Omega}) = - X_\text{o}(e^{-j \Omega})$
#
# Combining this, it can be concluded that the DTFT $X(e^{j \Omega})$ of a real-valued signal $x[k] \in \mathbb{R}$ shows complex conjugate symmetry
#
# \begin{equation}
# X(e^{j \Omega}) = X^*(e^{- j \Omega})
# \end{equation}
# #### Complex Signals
#
# By following the same procedure as above for an imaginary signal, the symmetries of the DTFT of the even and odd part of an imaginary signal can be derived. The results can be combined, by decomposing a complex signal $x[k] \in \mathbb{C}$ and its DTFT into its even and odd part for both the real and imaginary part. This results in the following symmetry relations
#
# 
#
# The transformation symbols $\circ \!\! - \!\! \bullet$ illustrate which part of the signal $x[k]$ is related to which part of its spectrum $X(e^{j \Omega})$. For instance, the odd part of the real part $\Re \{ x_\text{o} [k] \}$ results in an imaginary spectrum with odd symmetry $\Im \{ X_\text{o} (e^{j \Omega}) \}$.
# + [markdown] nbsphinx="hidden"
# **Copyright**
#
# The notebooks are provided as [Open Educational Resource](https://de.wikipedia.org/wiki/Open_Educational_Resources). Feel free to use the notebooks for your own educational purposes. The text is licensed under [Creative Commons Attribution 4.0](https://creativecommons.org/licenses/by/4.0/), the code of the IPython examples under the [MIT license](https://opensource.org/licenses/MIT). Please attribute the work as follows: *Lecture Notes on Signals and Systems* by <NAME>.
|
discrete_time_fourier_transform/properties.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=["hide-input"]
from IPython.core.display import HTML as Center
Center(""" <style>
.output_png {
display: table-cell;
text-align: center;
vertical-align: middle;
}
</style> """)
# -
# # Introduction
#
# ## Symmetries
#
# **Symmetries** provide one of the key ideas in modern mathematics and physics. They are naturally associated with the notion of **groups**. A typical problem in this context is to construct a system that behaves in a certain way under the action of a symmetry, for example, a system that is invariant under a given symmetry.
#
# To tackle this kind of problems, we need to know how the symmetries act on the building blocks of systems and these blocks are often the elements of vector spaces, i.e. vectors. This leads to a natural question of how to classify such actions of groups on vector spaces. The mathematical theory that deals with this question is the **representation theory**.
#
# $$
# \begin{array}{cc}
# \mathrm{Physics}&&\mathrm{Mathematics}\\
# \mathrm{symmetry}&\leftrightarrow&\mathrm{group}
# \end{array}
# $$
#
# Many systems in classical and quantum physics are invariant under some symmetry operations. We distinguish two types of symmetries: **discrete** and **continuous** symmetries.
#
# In the first cathegory we have for example
# * reflection (parity) symmetry
# * discrete rotations
# * time reversal
# * CPT symmetry of the Standard Model
#
# Continuous symmetries include:
# * time translations
# * space translations
# * spatial rotations
# * Poincare transformations
# * gauge symmetries of quuntum field theories
#
# Each symmetry can be described mathematically using groups and their representations.
#
# ## First look at groups and representations
#
# Consider the following square in 2D space:
# + tags=["hide-input"]
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.patches import Polygon
pts = np.array([[0,1], [1,0], [0,-1],[-1,0]])
p = Polygon(pts, closed=True)
ax = plt.gca()
ax.set_aspect('equal', 'box')
ax.grid()
ax.add_patch(p)
ax.set_xlim(-1.5,1.5)
ax.set_ylim(-1.5,1.5)
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.annotate("$x_2=(0,1)$", xy=(0, 1), xytext=(0.25, 1.25),
arrowprops=dict(arrowstyle="->"))
ax.annotate("$x_1=(1,0)$", xy=(1,0), xytext=(1.25, 0.25),
arrowprops=dict(arrowstyle="->"))
ax.annotate("$x_3=(-1,0)$", xy=(-1,0), xytext=(-1.5, -0.5),
arrowprops=dict(arrowstyle="->"))
ax.annotate("$x_4=(0,-1)$", xy=(0,-1), xytext=(0.25, -1.25),
arrowprops=dict(arrowstyle="->"))
plt.show()
# -
# It is symmetric under rotations and reflections. For example: the rotation by $90^\circ$ clockwise does not change the shape of the square. However, it **permutes** its vertices in the following way:
#
# $$x_1\to x_4\to x_3 \to x_2 \to x_1$$
#
# We associate a *permutation* to this symmetry: $(1432)$.
#
# Another example: lreflection with respect to the $x$-axis. It permutes vertices:
#
# $$x_2 \to x_4 \to x_2,$$
#
# and $x_1$ and $x_3$ are unchanged.
# We associate another permutation to this symmetry: $(1)(3)(24)=(24)$.
#
# There are 8 such symmetries in total (four rotations and four reflections):
# * $e_1=id$ - identity
# * $e_2=(1432)$ - rotation by $90^\circ$ clockwise
# * $e_3=(13)(24)$ - rotation by $180^\circ$ clockwise
# * $e_4=(1234)$ - rotation by $270^\circ$ clockwise
# * $e_5=(24)$ - $x$-axis reflection
# * $e_6=(13)$ - $y$-axis reflection
# * $e_7=(14)(23)$ - reflection with respect to the top-left to bottom-right diagonal
# * $e_8=(12)(34)$ - reflection with respect to the top-right to bottom-left diagonal
#
# Two symmetries applied one after another is also a symmetry: $e_2$ followed by $e_2$ is $e_3$. We indicate it as
#
# $$e_2 \star e_2 =e_3$$
#
# where $\star$ is a multiplication of group elements (binary operation). This can be done using vertex permutations:
#
# $$(1432)\circ(1432) =(13)(24)$$
#
# where we used the symbol $\circ$ to indicate multiplication (composition) of permutations.
#
# One can construct a table collecting all results of multiplications of two symmetries - **Cayley table**.
#
# This group is called the **dihedral group $D_4$**
#
# $$
# \begin{array}{c||c|c|c|c|c|c|c|c}
# \star&e_1&e_2&e_3&e_4&e_5&e_6&e_7&e_8\\
# \hline
# \hline
# e_1&e_1&e_2&e_3&e_4&e_5&e_6&e_7&e_8\\
# \hline
# e_2&e_2&e_3&e_4&e_1&e_7&e_8&e_6&e_5\\
# \hline
# e_3&e_3&e_4&e_1&e_2&e_6&e_5&e_8&e_7\\
# \hline
# e_4&e_4&e_1&e_2&e_3&e_8&e_7&e_5&e_6\\
# \hline
# e_5&e_5&e_8&e_6&e_7&e_1&e_3&e_4&e_2\\
# \hline
# e_6&e_6&e_7&e_5&e_8&e_3&e_1&e_2&e_4\\
# \hline
# e_7&e_7&e_5&e_8&e_6&e_2&e_4&e_1&e_3\\
# \hline
# e_8&e_8&e_6&e_7&e_5&e_4&e_2&e_3&e_1
# \end{array}
# $$
#
# Some important observations from this table:
# * each element appears only once in each column and in each row
# * the element $e_1$ acts as the identity element, namely
#
# $$e_1 \star e_i=e_i\star e_1=e_i$$
#
# * every element has its inverse, i.e. for each $e_i$ there exists $e_j$ such that
#
# $$e_i\star e_j=e_j\star e_i=e_1$$
#
# These are common properties for all groups.
#
# Groups are very abstract mathematical objects that are defined as a set with binary operation of its elements satisfying a series of conditions. We are interested in representing these elements in such a way that we can see how group elements act on the square. We can do it by associating $2\times 2$ matrices to each group element:
#
# $$
# \begin{align*}
# &e_1 \to \begin{pmatrix}1&0\\0&1\end{pmatrix}\qquad & e_2 \to \begin{pmatrix}0&1\\-1&0\end{pmatrix}\qquad
# &e_3 \to \begin{pmatrix}-1&0\\0&-1\end{pmatrix}\qquad & e_4 \to \begin{pmatrix}0&-1\\1&0\end{pmatrix}\\
# &e_5 \to \begin{pmatrix}1&0\\0&-1\end{pmatrix}\qquad & e_6 \to \begin{pmatrix}1&0\\0&-1\end{pmatrix}\qquad
# &e_7 \to \begin{pmatrix}0&-1\\-1&0\end{pmatrix}\qquad & e_8 \to \begin{pmatrix}0&1\\1&0\end{pmatrix}\\
# \end{align*}
# $$
#
# These matrices satisfy the same multiplication table as the group elements. We call these set of matrices a representation of group $D_4$. This is not the only representation and we will discuss many more of them in this module.
|
_build/jupyter_execute/Lectures/Lecture1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dexafrica/TravelGAN/blob/development/workspace4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="9WAmjQYVbcs4" colab_type="code" colab={}
self.g_lr
self.hidden_size
# + [markdown] id="y_vCRyFkKwtt" colab_type="text"
# **Data Processing**
# + id="FiL6dZv7K0iF" colab_type="code" colab={}
pip install pandasql s2sphere pyproj nltk
# + id="9YqNOfzNLRr-" colab_type="code" colab={}
import pandas as pd
import pandasql as psql
from pyproj import Proj, transform
import numpy as np
from numpy import array
from sklearn.preprocessing import MinMaxScaler
from datetime import datetime, timedelta
# + id="8C89Z_77LTlW" colab_type="code" colab={}
# mount the google drive
from google.colab import drive
drive.mount('/content/drive')
# + id="a-lIruucLbEH" colab_type="code" colab={}
data_path = 'drive/My Drive/ThesisProposal/trips_taz.csv'
train_data = pd.read_csv(data_path)
# + id="ipo-Jym0MwHt" colab_type="code" colab={}
train_data.head()
# + id="opVmmPGLqoAc" colab_type="code" colab={}
cols = ['HID', 'ORIG', 'DEST', 'D_HREDE', 'D_MOTIF']
test_data = train_data[cols]
test_data['DEP_TIME'] = test_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[:2] + ':' + test_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[2:] + ':00'
test_data['DEP_TIME'] = pd.to_datetime( pd.to_timedelta(test_data['DEP_TIME']).dt.round('5min') ).dt.strftime("%H:%M:%S")
test_data['D_MOTIF'] = test_data['D_MOTIF'].astype(str)
test_data['END_TIME'] = test_data['DEP_TIME'] + timedelta(hours=1)
test_origs = test_data.groupby('HID')['ORIG'].apply(' '.join).reset_index(name='ORIGS')
test_times = test_data.groupby('HID')['DEP_TIME'].apply(' '.join).reset_index(name='TIMES')
test_purpose = test_data.groupby('HID')['D_MOTIF'].apply(' '.join).reset_index(name='PURPOSE')
end_rec = test_data.groupby(['HID']).last().reset_index()
first_rec = test_data.groupby(['HID']).first().reset_index()
# + id="l6NGR88WJjzK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e789713a-9e5b-41b3-8cb9-6dd800e4daac"
full_trip = test_origs['ORIGS'] + ' ' + end_rec['DEST']
full_times = test_times['TIMES'] + ' ' + pd.to_datetime(end_rec['END_TIME']).dt.strftime("%H:%M:%S")
full_purpose = test_purpose['PURPOSE'] + ' ' + first_rec['D_MOTIF']
# + id="U1QP5wYcTwYj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 468} outputId="52e76946-2694-4595-b015-dd3fb9e93bab"
a = full_trip[0:5].str.split(' ')
b = full_times[0:5].str.split(' ')
c = full_purpose[0:5].str.split(' ')
d = test_origs['HID']
for i in range(len(a)):
print (d[i])
for j in range(len(a[i])):
mlist = [a[i][j], b[i][j], c[i][j]]
print (mlist)
# print(a[i][j])
# print(b[i][j])
# print(c[i][j])
# for col in row:
# + id="Ionq7U0QBjqQ" colab_type="code" colab={}
end_dest = test_data.groupby(['HID']).last().reset_index()
# test_data['END_TIME'] = (pd.to_datetime( pd.to_timedelta(test_data['DEP_TIME']) + timedelta(hours=1)).dt.round('5min') ).dt.strftime("%H:%M:%S")
# + id="p0v0-TMrC7WI" colab_type="code" colab={}
df_test = pd.merge(test_data, last_records, on='HID', how='outer')
# df_outer['TRAVEL'] = df_outer['ORIG'] + ' ' + df_outer['TRIPS']
# + id="yvhmPXMxDgmb" colab_type="code" colab={}
full_trip = test_origs['ORIGS'] + ' ' + end_dest['DEST']
full_times = test_times['TIMES'] + ' ' + end_dest['END_TIME']
# + id="oyugxRSCF66z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="1615e60b-1d19-4f1e-fcef-29cd66a486af"
test_origs['ORIGS']
# + id="AKpJ6YVDGQQW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c27f1458-2ad7-49c2-80b9-a544336bf5d3"
full_times[0]
# + id="mAJlnWf6Buol" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="8dc764f3-613b-4dcf-9989-e0d3fc15d162"
last_records
# + id="JZNBOPDNvKyI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="8935c212-8e68-4896-d3a9-b8dd22fc583d"
test_times
# + id="hv7F68afveiG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 416} outputId="fdef867a-dcb3-47af-f7a1-b5a17e6eab9b"
test_origs
# + id="gaQ1VzM2riky" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="5bfd8972-7985-4136-d8d5-ba9f769c7e23"
test_data[['ORIG', 'D_HREDE']].values
# + [markdown] id="Dbn-YwJNP4--" colab_type="text"
# **Normalize numeric variables**
# + id="6jxwpL-gQBIl" colab_type="code" colab={}
mm_scaler = preprocessing.MinMaxScaler()
X_train_minmax = mm_scaler.fit_transform(X_train)
mm_scaler.transform(X_test)
# + id="m-QLbvjqRIzn" colab_type="code" colab={}
scaler_olng=scaler_olat=scaler_dlng=scaler_dlat = MinMaxScaler()
# + id="QfvrG8JmSN-Z" colab_type="code" colab={}
sc_olat = scaler_olat.fit_transform(train_data['S_ORIG_LAT'].values.reshape(-1,1))
sc_olng = scaler_olng.fit_transform(train_data['S_ORIG_LON'].values.reshape(-1,1))
sc_dlat = scaler_dlat.fit_transform(train_data['S_DEST_LAT'].values.reshape(-1,1))
sc_dlng = scaler_dlng.fit_transform(train_data['S_DEST_LON'].values.reshape(-1,1))
# + id="zZiE8DlfU6Re" colab_type="code" colab={}
df_hid = train_data['HID']
num_cols = np.concatenate((sc_olat, sc_olng, sc_dlat, sc_dlng), axis=1)
normalized_df = pd.DataFrame(num_cols, columns=["Orig_Lat", "Orig_Lng", "Dest_Lat", "Dest_Lng"])
df = pd.concat([df_hid, normalized_df], axis=1)
# + id="7BHeAdCAcGRT" colab_type="code" colab={}
_t = df[['Orig_Lat', 'Orig_Lng', 'Dest_Lat', 'Dest_Lng']].values.tolist()
# + id="LKvldwyVXAdC" colab_type="code" outputId="72401059-d9e8-401a-d18d-7d6aafb09da8" colab={"base_uri": "https://localhost:8080/", "height": 34}
normalized_df.columns
# + id="9nqUz_8CrXFU" colab_type="code" colab={}
_s = df.groupby('HID')['Orig_Lat', 'Orig_Lng', 'Dest_Lat', 'Dest_Lng'].apply(pd.Series.tolist).tolist()
# + id="tWL4vHLtrmhb" colab_type="code" colab={}
from keras.preprocessing.sequence import pad_sequences
trips_seq = pad_sequences(_s, maxlen=15)
# + id="A-jNFC9kLdPO" colab_type="code" colab={}
train_data['D_HREDE'] = train_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[:2] + ':' + train_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[2:] + ':00'
train_data['DEP_TIME'] = pd.to_datetime( pd.to_timedelta(train_data['D_HREDE']).dt.round('5min') ).dt.strftime("%H:%M:%S")
train_data['TRIP'] = train_data[['ORIG', 'DEST', 'DEP_TIME']].values.tolist()
# + id="QGkAZBCOLl2H" colab_type="code" outputId="68faf952-9a3d-45dd-e8ef-a95381d98ea7" colab={"base_uri": "https://localhost:8080/", "height": 52}
train_data['D_HREDE'] = train_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[:2] + ':' + train_data['D_HREDE'].map(int).astype(str).str.zfill(4).str[2:] + ':00'
train_data['DEP_TIME'] = pd.to_datetime( pd.to_timedelta(train_data['D_HREDE']).dt.round('5min') ).dt.strftime("%H:%M:%S")
trips_data = train_data.groupby('HID')['DEST'].apply(' '.join).reset_index(name='TRIPS')
first_records = train_data.groupby(['HID']).first().reset_index()
df_outer = pd.merge(trips_data, first_records, on='HID', how='outer')
df_outer['TRAVEL'] = df_outer['ORIG'] + ' ' + df_outer['TRIPS']
cols = ['P_SEXE', 'P_AGE', 'P_STATUT', 'P_PERMIS', 'P_MOBIL', 'D_HREDE', 'ZT03', 'TRAVEL']
trips_data = df_outer[cols]
# + id="7_i23q6VpRzl" colab_type="code" colab={}
trips_data
# + id="lC7_LXPXSetx" colab_type="code" colab={}
# df_outer['TRAVEL'].to_csv('drive/My Drive/ThesisProposal/export_trip_101119.csv', index = None, header=True)
# SEQUENCE LENGTH CLEANUP
df_outer['LEN'] = len(df_outer['TRAVEL'].str.split(' '))
dt = df_outer.copy()
dt[['LEN']] = dt.apply(lambda row:int(len(row['TRAVEL'].split(' '))), axis=1).apply(pd.Series)
# Delete sequences of length = 2
trips_data = dt[dt.LEN != 2]
# + id="Bgk-ayLHhaR0" colab_type="code" colab={}
trips_data['TRAVEL'].to_csv('drive/My Drive/ThesisProposal/export_trip_251119.csv', index = None, header=True)
# + id="Ad9Ht1lntt5F" colab_type="code" outputId="ca6e8d9f-f091-436a-a1e2-3cbb71c23b51" colab={"base_uri": "https://localhost:8080/", "height": 34}
trips_data.shape
# + [markdown] id="U6MEqgDo87Zw" colab_type="text"
# Prefix Tree
# + id="P183k9ID8-a8" colab_type="code" colab={}
from typing import Tuple
class TrieNode(object):
"""
Our trie node implementation. Very basic. but does the job
"""
def __init__(self, loc: str, trip_data=None, epsilon=None):
self.loc = loc
self.children = []
# Is it the last character of the word.`
self.trip_ended = False
# How many times this character appeared in the addition process
self.counter = 1
# dataset of trip chain
self.trip_data = trip_data
# trajectories in the database
self.trajectories = self.add_trajectories()
# set privacy budget, epsilon
self.epsilon = epsilon
# set height of prefix tree, h
self.h = 10
# set privacy budget allocation
self.budget_allocation = self.get_privacy_allocation()
# set noisy counter for nodes
self.noisy_counter = 1
# location taxononmy tree
# self.location_tree = self.location_taxonomy()
# set level of node
self.level = 0
def location_taxonomy(self):
m = self.trip_data.str.split(' ').tolist()
m = [i for j in m for i in j]
n = np.unique(np.array(m))
return n
def add_trajectories(self):
traj = None
if self.trip_data is not None:
# traj = self.trip_data.tolist()
traj = self.trip_data
return traj
return None
def get_privacy_allocation(self):
priv_alloc = None
if self.epsilon is not None:
priv_alloc = self.epsilon / self.h
return priv_alloc
def get_noisy_count(in_count):
noise_count = in_count
def add(root, trip: str):
"""
Adding a word in the trie structure
"""
node = root
# print(trip)
for loc in trip:
# print (loc)
found_in_child = False
# Search for the character in the children of the present `node`
for child in node.children:
if child.loc == loc:
# We found it, increase the counter by 1 to keep track that another
# word has it as well
child.counter += 1
# And point the node to the child that contains this char
node = child
found_in_child = True
break
# We did not find it so add a new chlid
if not found_in_child:
new_node = TrieNode(loc)
node.children.append(new_node)
if node == root:
new_node.level = 1
# And then point node to the new child
node = new_node
# Everything finished. Mark it as the end of a word.
node.trip_ended = True
def traverse(root, prefix):
result = []
if root.isEnd:
result.append(prefix[:])
for c,n in root.children.items():
prefix.append(c)
self.traverse(n, prefix, result)
prefix.pop(-1)
return [''.join(r) for r in result]
def find_prefix(root, prefix: str) -> Tuple[bool, int]:
"""
Check and return
1. If the prefix exists in any of the words we added so far
2. If yes then how may words actually have the prefix
"""
node = root
# If the root node has no children, then return False.
# Because it means we are trying to search in an empty trie
if not root.children:
return False, 0
for loc in prefix:
char_not_found = True
# Search through all the children of the present `node`
for child in node.children:
if child.loc == loc:
# We found the char existing in the child.
char_not_found = False
# Assign node as the child containing the char and break
node = child
break
# Return False anyway when we did not find a char.
if char_not_found:
return False, 0
# Well, we are here means we have found the prefix. Return true to indicate that
# And also the counter of the last node. This indicates how many words have this
# prefix
# return True, node.counter
return node
# + id="5txIyXXqbpY1" colab_type="code" colab={}
# + id="oTN4q81A4gxm" colab_type="code" colab={}
# SEQUENCE LENGTH CLEANUP
df_outer['LEN'] = len(df_outer['TRAVEL'].str.split(' '))
dt = df_outer.copy()
dt[['LEN']] = dt.apply(lambda row:int(len(row['TRAVEL'].split(' '))), axis=1).apply(pd.Series)
# Delete sequences of length = 2
trips_data = dt[dt.LEN != 2]
# + id="W7yZfGkMjbQb" colab_type="code" colab={}
# ADD DATA TO PREFIX TREE
m = trips_data['TRAVEL']
root = TrieNode('*', epsilon=0.1, trip_data=m)
# add nodes to PT
for index, row in trips_data.head(100).iterrows():
add(root, row['TRAVEL'].split(' '))
# + id="SM1AebkF5o1Q" colab_type="code" colab={}
#ADD NOISE TO ROOT COUNTER NODES
# Gets random laplacian noise for all values
location = 1.0
scale = 1.0
noise = np.random.laplace(location,scale, len(root.children))
index = 0
for child in root.children:
child.noisy_counter = int(child.counter + noise[index])
print(child.loc, child.counter, child.noisy_counter, [a.loc for a in child.children])
index = index + 1
# + id="J0zO1w2yYW-L" colab_type="code" colab={}
# Get empty nodes
empty_nodes = []
threshold = 3
for child in root.children:
if child.noisy_counter >= threshold:
empty_nodes.append(child)
child.noisy_counter = int(child.counter + noise[index])
print(child.loc, child.counter, child.noisy_counter, [a.loc for a in child.children])
# Get non-empty nodes
# + id="ugSbKmHim6l2" colab_type="code" outputId="799e0bde-bfc7-41e6-fcca-ffb01ba6c573" colab={"base_uri": "https://localhost:8080/", "height": 69}
v = np.random.exponential(1, 43 )
np.random.laplace(scale=10, size=43)
np.random.exponential(100,10)
# + id="VLt25qBHKdwQ" colab_type="code" colab={}
v
# + id="pV6BLRAiA4gL" colab_type="code" colab={}
root = TrieNode('*')
for index, row in trips_data.head(100).iterrows():
add(root, row['TRAVEL'].split(' '))
# + id="XqM1PslXkW73" colab_type="code" colab={}
for index, row in trips_data.head(100).iterrows():
add(root, row['TRAVEL'].split(' '))
# + id="dwq7LX22lVJP" colab_type="code" colab={}
st = '4cc91ea026fa376b 4cc91a5cdca68651 4cc91a43c5089bad 4cc90f48bf459b03 4cc91ea026fa376b 4cc8d8d2dcb61c63 4cc91ea026fa376b'
st = st.split(' ')
# print(find_prefix(root, st))
_s = find_prefix(root, st)
# + id="_gZEK9Mv1d4K" colab_type="code" outputId="a69bf438-163d-4ce3-bc28-e698cb93e97b" colab={"base_uri": "https://localhost:8080/", "height": 797}
[[child.loc, child.counter] for child in root.children]
# + id="W67fGvXbJ46W" colab_type="code" colab={}
# for r in root.children:
index = 0
for child in root.children:
child.noisy_counter = child.counter + v[index]
print(child.loc, child.counter, child.noisy_counter, child.level, [a.loc for a in child.children])
index = index + 1
# + id="gZ_96Sqz1sBF" colab_type="code" colab={}
trips_data['TRAVEL'].head(20)
# + id="LJ0aXveLk8b_" colab_type="code" outputId="22f1fb29-c733-4941-c358-4414a5bafde4" colab={"base_uri": "https://localhost:8080/", "height": 69}
root.location_tree
# + id="kxgjORV7b3oo" colab_type="code" colab={}
# for r in root.children:
node = root
for child in node.children:
print(child.loc, child.counter, [[a.loc, a.counter] for a in child.children])
# + id="x7vqgRwbDf_3" colab_type="code" colab={}
# + id="u37rw0v0neGj" colab_type="code" colab={}
# + id="kGjG0ILtkNFr" colab_type="code" outputId="c05009bb-73e0-4e8e-9663-c9fea9601d9d" colab={"base_uri": "https://localhost:8080/", "height": 69}
r = root.location_tree
r
# + id="JiI6kIRbRVdE" colab_type="code" colab={}
n = m.str.split(' ')
# + id="Dnbte7TsTRUs" colab_type="code" colab={}
import numpy as np
# function to get unique values
def unique(list1):
x = np.array(list1)
return np.unique(x)
# + id="tHG10oRWT2dx" colab_type="code" colab={}
a = trips_data['TRAVEL'].str.split(' ').tolist()
a = [i for j in a for i in j]
# + id="NW-RPEqbUNn-" colab_type="code" colab={}
b = unique(a)
# + id="0Mpjy_LOUeia" colab_type="code" outputId="487b8f84-fb35-4b2c-fb72-68a6f329cc08" colab={"base_uri": "https://localhost:8080/", "height": 34}
b[0]
# + id="ZCL7i6y7TTN_" colab_type="code" outputId="c63bd472-1481-4f29-8bbe-9941ebda8057" colab={"base_uri": "https://localhost:8080/", "height": 156}
list1 = [10, 20, 10, 30, 40, 40]
print("the unique values from 1st list is")
unique(n)
# + id="pgIJ_kX5SHch" colab_type="code" colab={}
n.tolist().unique()
# + id="u3tGPA1EPDGW" colab_type="code" outputId="3e13484b-61b5-4d64-ca1e-252bf0f383fb" colab={"base_uri": "https://localhost:8080/", "height": 34}
s = trips_data['TRAVEL'].head(4)
t = s.str.split(' ')
v = t[0]
v
# + id="E34l-QEUrcRA" colab_type="code" colab={}
b = root.trajectories
c = b.str.split(' ')
# + id="TimLG8L-rou-" colab_type="code" colab={}
f = root.trajectories[1]
[f.split(' ') for i in f]
# + id="lhpEINMapN7T" colab_type="code" outputId="d087a812-1012-4ef0-ac0b-322fc6f0c00e" colab={"base_uri": "https://localhost:8080/", "height": 104}
s
# + id="2su79h3_CpqU" colab_type="code" outputId="d222ab96-cc42-4f77-f119-972475158fcb" colab={"base_uri": "https://localhost:8080/", "height": 34}
st = '4cc8de61f2e92d05'
st = st.split(' ')
print(find_prefix(root, st))
# print(traverse(root, st))
# + id="nm_OOrEh-XfB" colab_type="code" colab={}
m = trips_data['TRAVEL'].tolist()
# + id="4TJbPc9LiUJA" colab_type="code" outputId="a7a83252-6e09-4c8d-a4a9-3351b3aba4a0" colab={"base_uri": "https://localhost:8080/", "height": 34}
m[1]
# + id="cKr9EQIw9u7J" colab_type="code" outputId="a1425b1f-4a9d-4d4b-9ae7-2be72a7b5654" colab={"base_uri": "https://localhost:8080/", "height": 34}
print(find_prefix(root, '4cc8de61f2e92d05 4cc8de67553ddeaf 4cc8de61f2e92d05 4cc8dea4fa05383f 4cc8de61f2e92d05'))
# + id="7ktrBZd15fQd" colab_type="code" outputId="d1d06125-e1e1-4d32-ef83-9671b670bada" colab={"base_uri": "https://localhost:8080/", "height": 34}
trips_data['TRAVEL'][1]
# + id="X5itg5c7iZe4" colab_type="code" colab={}
df_outer['LEN'] = len(df_outer['TRAVEL'].str.split(''))
# + id="7nuk1_rMly3X" colab_type="code" colab={}
# dt[['LEN']] = dt.apply(lambda row:str(getCellId(row['ORIG_LON'], row['ORIG_LAT']).to_token()), axis=1).apply(pd.Series)
dt = df_outer.copy()
dt[['LEN']] = dt.apply(lambda row:int(len(row['TRAVEL'].split(' '))), axis=1).apply(pd.Series)
# + id="s4WsK-NHmjuJ" colab_type="code" colab={}
# SELECT HOME BASED TRIPS
dt_hbt = dt[dt.LEN != 2]
# + id="civMCi3PpiL7" colab_type="code" colab={}
dt_hbt = dt_hbt[cols]
# + id="zEM2Xfc4kUNL" colab_type="code" colab={}
# + [markdown] id="zJPPVyBRX-vJ" colab_type="text"
# # **Trie data traversal**
# + id="Jerf4c1BYFiZ" colab_type="code" colab={}
def list_words(trie):
my_list = []
for k,v in trie.items():
if k != '_':
for el in list_words(v):
my_list.append(k+el)
else:
my_list.append('')
return my_list
# + [markdown] id="BH0nFNtDOHE3" colab_type="text"
# # **Laplace noise**
# + id="N6K8LaOeOJp9" colab_type="code" colab={}
from math import log
from random import random
def exp_sample(mean):
return -mean*log(random())
def laplace(scale):
e1 = exp_sample(scale)
e2 = exp_sample(scale)
return e1 - e2
# + id="whIRJYzIQorA" colab_type="code" outputId="4a83d8a5-72e9-4145-e070-8d2b2ce204fa" colab={"base_uri": "https://localhost:8080/", "height": 166}
laplace(10)
# + id="hofVSmzVwpUy" colab_type="code" outputId="b3ef2504-1b88-4287-e145-07342a8588a9" colab={"base_uri": "https://localhost:8080/", "height": 72}
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data', sep=r'\s*,\s*',na_values="?")
datacount = df["United-States"].value_counts()
# + id="zduErs33yBkB" colab_type="code" outputId="693634e3-11f1-4e37-9929-b1452d5d906d" colab={"base_uri": "https://localhost:8080/", "height": 745}
datacount
# + id="Hh40g6CaxxG2" colab_type="code" outputId="9dc4d5c0-57b3-4b3e-88be-d0e5cf44beef" colab={"base_uri": "https://localhost:8080/", "height": 416}
datacount.plot(kind="bar", color = 'r')
# + id="zuZ_auSAxAeg" colab_type="code" outputId="a45a6f9a-afba-423b-e678-ee8b586e1799" colab={"base_uri": "https://localhost:8080/", "height": 34}
Exponential_noise = np.random.exponential(1)
print ("Exponentially generated noise:", Exponential_noise)
# + id="YG3iL8rOxmXp" colab_type="code" outputId="22516b4c-7bd0-4214-9f36-e06407a21697" colab={"base_uri": "https://localhost:8080/", "height": 416}
noisydata = datacount + Exponential_noise
noisydata.plot(kind="bar", color = 'r')
# + id="b4peA2zwx7su" colab_type="code" outputId="879a1ce6-92af-4468-a987-725e75dd6f18" colab={"base_uri": "https://localhost:8080/", "height": 745}
noisydata
# + id="f-l8-e3_Yvna" colab_type="code" colab={}
# Tokenize Travel Chain
from keras.preprocessing.text import Tokenizer
tr_token = Tokenizer(num_words=None)
tr_token.fit_on_texts(trips_data['TRAVEL'])
tr_tokens = tr_token.texts_to_sequences(trips_data['TRAVEL'])
trips_data['TRIPS'] = tr_tokens
# + id="026YI_uVaWVu" colab_type="code" colab={}
# Pad sequences
from keras.preprocessing.sequence import pad_sequences
trips_seq = pad_sequences(trips_data['TRIPS'],padding='post', maxlen=15)
# + id="vCLFEV7IoGkG" colab_type="code" outputId="edc66891-bde3-4253-ad1c-9831a63bbb4c" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(tr_token.word_index)
# + id="L5a-HABIsRon" colab_type="code" colab={}
df_outer['TRAVEL'].to_csv('drive/My Drive/ThesisProposal/export_trip_101119.csv', index = None, header=True)
# + [markdown] colab_type="text" id="7vEUyxfrtalL"
# **Pre-Training**
# + id="_qo2AARPmy0_" colab_type="code" colab={}
import tensorflow as tf
from keras.optimizers import Adam
from keras.layers import Input, Dropout, Concatenate, Embedding, Dense, LSTM, Lambda
from keras.models import Model, Sequential
import keras.backend as K
_session = tf.Session()
K.set_session(_session)
# + id="QLqjpnxY3qB0" colab_type="code" colab={}
# + [markdown] id="D9o1CWHNRJ3v" colab_type="text"
# **Reinforcement Learning**
# + id="bhjjHBLCZgJt" colab_type="code" colab={}
# from models import Generator, GeneratorPretraining, Discriminator
# from utils import DiscriminatorGenerator
import keras.backend as K
import numpy as np
class Agent ( object ):
'''
On each step, Agent act on state.
Then Environment return next state, reward, and so on.
'''
def __init__ ( self , sess , batch_size , vocab_dim , embedding_dim , hidden_size , lr = 1e-3 ):
self .sess = sess
self .num_actions = vocab_dim
self .B = batch_size
self .V = vocab_dim
self .E = embedding_dim
self .H = hidden_size
self .lr = lr
self .eps = 0.1
self .generator = Generator (sess, B, V, E, H, lr)
def act ( self , state , epsilon = 0 , deterministic = False ):
'''
# Arguments:
state: numpy array, dtype = int, shape = (B, t)
epsilon: float, 0 <= epsilon <= 1,
if epsilon is 1, the Agent will act completely random.
# Returns:
action: numpy array, dtype = int, shape = (B, 1)
'''
trip = state[:, - 1 ].reshape ([ - 1 , 1 ])
return self._act_on_trip (trip, epsilon = epsilon, deterministic = deterministic)
def _act_on_trip ( self , trip , epsilon = 0 , deterministic = False , PAD = 0 , EOS = 2 ):
'''
# Arguments:
trip: numpy array, dtype = int, shape = (B, 1),
trip indicates current trip.
epsilon: float, 0 <= epsilon <= 1,
if epsilon is 1, the Agent will act completely random.
# Returns:
action: numpy array, dtype = int, shape = (B, 1)
'''
action = None
is_PAD = trip == PAD
is_EOS = trip == EOS
is_end = is_PAD.astype (np.int) + is_EOS.astype (np.int)
is_end = 1 - is_end
is_end = is_end.reshape ([ self .B, 1 ])
if np.random.rand () <= epsilon:
action = np.random.randint ( low = 0 , high = self.num_actions, size = ( self.B, 1 ))
elif not deterministic:
probs = self .generator.predict (trip)
action = self .generator.sampling_trip(probs).reshape ([ self.B, 1 ]) ###### cross check
else :
probs = self .generator.predict (trip) # (B, T)
action = np.argmax (probs, axis = - 1 ).reshape ([ self.B, 1 ])
return action * is_end
def reset ( self ):
self.generator.reset_rnn_state ()
def save ( self , path ):
self.generator.save (path)
def load ( self , path ):
self.generator.load (path)
class Environment ( object ):
'''
On each step, Agent act on state.
Then Environment return next state, reward, and so on.
'''
def __init__ ( self , discriminator , data_generator , g_beta , n_sample = 16 ):
'''
Environment class for Reinforced Learning
# Arguments:
discriminator: keras model
data_generator: SeqGAN.models.GeneratorPretrainingGenerator
g_beta: SeqGAN.rl.Agent, copy of Agent
params of g_beta.generator should be updated with those of original
generator on regular occasions.
# Optional Arguments
n_sample: int, default is 16, the number of Monte Carlo search sample
'''
self.data_generator = data_generator
self.B = data_generator.B
# self.t = data_generator.T
self.T = data_generator.T
self.n_sample = n_sample
self.BOS = data_generator.BOS #Begin of sentence
self.discriminator = discriminator
self.g_beta = g_beta
self.reset ()
def get_state ( self ):
if self.t == 1 :
return self._state
else:
return self._state [:, 1 :] # Exclude BOS
def reset ( self ):
self.t = 1
self._state = np.zeros ([ self.B, 1 ], dtype = np.int32)
self._state [:, 0 ] = self.BOS
self.g_beta.reset ()
def step ( self , action ):
'''
Step t-> t + 1 and returns a result of the Agent action.
# Arguments:
action: numpy array, dtype = int, shape = (B, 1),
state is Y_0: t-1, and action is y_t
# Returns:
next_state: numpy array, dtype = int, shape = (B, t)
reward: numpy array, dtype = float, shape = (B, 1)
is_episode_end: bool
info: dict
'''
self.t = self.t + 1
reward = self.Q (action, self .n_sample)
is_episode_end = self.T > self.T
self ._append_state (action)
next_state = self.get_state ()
info = None
return [next_state, reward, is_episode_end, info]
def render ( self , head = 1 ):
for i in range (head):
ids = self .get_state () [i]
words = [ self .data_generator.id2word [ id ] for id in ids.tolist ()]
print ( ' ' .join (words))
print ( ' - ' * 80 )
def Q ( self , action , n_sample = 16 ):
'''
State-Action value function using Rollout policy
# Arguments:
action: numpy array, dtype = int, shape = (B, 1)
# Optional Arguments:
n_sample: int, default is 16, number of samples for Monte Calro Search
# Returns:
reward: numpy array, dtype = float, shape = (B,), State-Action value
# Requires:
t, T: used to define time range.
state: determined texts, Y [0: t-1], used for Rollout.
action: next words, y [t], used for sentence Y [0: t].
g_beta: Rollout policy.
'''
h, c = self.g_beta.generator.get_rnn_state ()
reward = np.zeros([self.B, 1 ])
if self.t == 2 :
Y_base = self._state # Initial case
else:
Y_base = self.get_state () # (B, t-1)
if self.t >= self.T + 1 :
Y = self._append_state (action, state = Y_base)
return self.discriminator.predict (Y)
# Rollout
for idx_sample in range (n_sample):
Y = Y_base
self.g_beta.generator.set_rnn_state (h, c)
y_t = self.g_beta.act (Y, epsilon = self .g_beta.eps)
Y = self._append_state (y_t, state = Y)
for tau in range ( self.t + 1, self.T):
y_tau = self .g_beta.act (Y, epsilon = self.g_beta.eps)
Y = self ._append_state (y_tau, state = Y)
reward += self .discriminator.predict (Y) / n_sample
return reward
def _append_state ( self , trip , state = None ):
'''
# Arguments:
trip: numpy array, dtype = int, shape = (B, 1)
'''
trip = trip.reshape ( - 1 , 1 )
if state is None :
self ._state = np.concatenate ([ self._state, trip], axis = -1 )
else :
return np.concatenate ([state, trip], axis = -1 )
# + id="3OIhKufH2OMB" colab_type="code" colab={}
seqgan = SeqGAN()
# + id="XFKUEuhjg_IL" colab_type="code" colab={}
class Trainer():
def __init__ (self, vocab_size, g_data, d_data = None):
self.batch_size = 64
self.seq_len = 64
self.vocab_size = vocab_size
self.n_sample = 16
self.d_dropout = 0.01
self.g_lr = 1e-3
self.d_lr = 1e-3
self.g_E, self.d_E = 50
self.g_H, self.d_H = 50
self.g_data = g_data
self.d_data = d_data
self.agent = Agent(_session, self.batch_size, self.vocab_size, self.g_E, self.g_H, self.g_lr)
self.g_beta = Agent(_session, self.batch_size, self.vocab_size, self.g_E, self.g_H, self.g_lr )
self.critic = Discriminator(self.vocab_size, self.d_E, dropout = self.d_dropout)
self.env = Environment(self.critic, self.g_data, self.g_beta, n_sample=self.n_sample)
self.generator_pre = GeneratorPreTraining(self.vocab_size, self.g_E, self.g_H)
# + [markdown] id="Nb-uKXfAgxY5" colab_type="text"
# # **Generative Model**
# + id="2AANwNCqg4Yl" colab_type="code" colab={}
class SeqGAN():
def __init__(self):
self.seq_length = 15
self.seq_shape = (self.seq_length, 1)
self.latent_dim = 100
self.disc_loss = []
self.gen_loss =[]
self.vocab_size = 1000
self.data_shape = (39,)
self.batch_size = 64
self.hidden_size = 64
self.embedding_size = 64
optimizer = Adam(0.0002, 0.5)
# Build and compile the discriminator
self.discriminator = self.build_discriminator()
self.discriminator.compile(loss='binary_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# The generator takes noise as input and generates note sequences
z = Input(shape=(self.latent_dim,))
generated_seq = self.generator(z)
# For the combined model we will only train the generator
self.discriminator.trainable = False
# The discriminator takes generated images as input and determines validity
validity = self.discriminator(generated_seq)
# The combined model (stacked generator and discriminator)
# Trains the generator to fool the discriminator
self.combined = Model(z, validity)
self.combined.compile(loss='binary_crossentropy', optimizer=optimizer)
def pretrain_generator (self):
model = Sequential()
model.add(Embedding(self.vocab_size, 64, mask_zero=True, name='Embedding'))
model.add(LSTM(64, return_sequences=True, name='LSTM'))
model.add(TimeDistributed(Dense(self.vocab_size, activation='softmax')))
model.summary()
data_input = Input(shape=(self.data_shape), dtype='int32', name='Input')
output = model(data_input)
return Model(data_input, output)
def build_generator(self):
model = Sequential()
model.add(Embedding(self.vocab_size, 64, mask_zero=True))
model.add(LSTM(64, dropout_U = 0.2, dropout_W = 0.2, return_sequences=True))
model.add(Dense(self.vocab_size, activation='softmax'))
data_input = Input(shape=(self.data_shape), dtype='int32', name='Input')
out = model(data_input)
return Model(data_input, out)
def reset_rnn_state (self):
self.h = np.zeros([self.batch_size, self.hidden_size])
self.c = np.zeros([self.batch_size, self.hidden_size])
def set_rnn_state(self, h, c):
self.h = h
self.c = c
def get_rnn_state(self):
return self.h, self.c
def predict(self, state, stateful = True):
feed_dict = {
self.state_in: state,
self.h_in: self.h,
self.c_in: self.c
}
prob, next_h, next_c = self.sess.run(
[self.prob, self.next_h, self.next_c], feed_dict
)
if stateful:
self.h = next_h
self.c = next_c
return prob
else:
return prob, next_h, next_c
def update(self, state, action, reward, h=None, c=None, stateful=True):
if h is None:
h = self.h
if c is None:
c= self.c
state = state[:, -1].reshape(-1, 1)
reward = reward.reshape(-1)
feed_dict = {
self.state_in: state,
self.h_in: h,
self.c_in: c,
self.action: to_categorical(action, self.vocal_size),
self.reward: reward
}
_, loss, next_h, next_c = self.sess.run(
[self.minimize, self.loss, self.next_h, self.next_c], feed_dict
)
if stateful:
self.h = next_h,
self.c = next_c,
return loss
else:
return loss, next_h, next_c
def sampling_trip(self, prob):
action = np.zeros((self.batch_size,), dtype='int32')
for i in range(self.batch_size):
p = prob[i]
action = np.random.choice(self.vocab_size, p = p)
return action
def sampling_trip_chain(self, T, BOS=1):
self.reset_rnn.state()
action = np.zeros([self.batch_size, 1], dtype='int32')
action[:, 0] = BOS
actions = action
for _ in range(T):
prob = self.predict(action)
action = self.sampling_trip(prob).reshape(-1, 1)
actions = np.concatenate([actions, action], axis=-1)
actions = actions[:, 1:]
self.reset_rnn_state()
return actions
def generate_trips(self, T, g_data, num, output_file):
trip_chains = []
for _ in range(num // self.batch_size + 1):
actions = self.sampling_trip_chain(T)
actions_list = actions.tolist()
for chain_id in actions_list:
chain = [g_data.id2word[action] for action in chain_id]
trip_chains.append(chain)
output_str = ' '
for i in range(num):
output_str += ' '.join(trip_chains[i] + '\n')
with io.open(output_file, 'w', encoding='utf-8') as f:
f.write(output_str)
def save(self, path):
weights = []
for layer in self.layers:
w = layer.get_weights()
weights.append(w)
with io.open(path, 'wb') as f:
pickle.dump(weights, f)
def load(self, path):
with io.open(path, 'rb') as f:
weights = pickle.load(f)
for layer, w in zip(self.layers, weights):
layer.set_weights(w)
def Highway (self, x , num_layers = 1 , activation = 'relu' , name_prefix = '' ):
input_size = K.int_shape (x) [ 1 ]
for i in range (num_layers):
gate_ratio_name = '{}Highway/Gate_ratio_{}'.format (name_prefix, i)
fc_name = '{}Highway/FC_{}'.format (name_prefix, i)
gate_name = '{}Highway/Gate_{}'.format (name_prefix, i)
gate_ratio = Dense (input_size, activation = 'sigmoid' , name = gate_ratio_name) (x)
fc = Dense (input_size, activation = activation, name = fc_name) (x)
x = Lambda ( lambda args : args [ 0 ] * args [ 2 ] + args [ 1 ] * ( 1 - args [ 2 ]), name = gate_name) ([fc, x, gate_ratio])
return x
def pre_train_generator (self , g_epochs = 3, g_pre_path = None, lr = 1e-3 ):
if g_pre_path is None:
self.g_pre_path = os.path.join (self.top, 'data', 'save', 'generator_pre.hdf5' )
else :
self.g_pre_path = g_pre_path
g_adam = Adam (lr)
self.generator_pre.compile (g_adam, 'categorical_crossentropy' )
print (' Generator pre-training ' )
self.generator_pre.summary ()
self.generator_pre.fit_generator (
self.g_data,
steps_per_epoch = None,
epochs = g_epochs)
self .generator_pre.save_weights (self.g_pre_path)
self .reflect_pre_train()
def pre_train_discriminator (self, d_epochs = 1, d_pre_path = None, lr = 1e-3 ):
if d_pre_path is None:
self.d_pre_path = os.path.join (self.top, 'data', 'save', 'discriminator_pre.hdf5' )
else:
self.d_pre_path = d_pre_path
print ('Start Generating sentences ')
self.agent.generator.generate_samples (self.T, self.g_data,
self.generate_samples, self.path_neg)
self .d_data = DiscriminatorGenerator (
path_pos = self.path_pos,
path_neg = self.path_neg,
B = self.B,
shuffle = True )
d_adam = Adam (lr)
self.discriminator.compile (d_adam, 'binary_crossentropy' )
self.discriminator.summary ()
print('Discriminator pre-training ' )
self.discriminator.fit_generator (
self.d_data,
steps_per_epoch = None ,
epochs = d_epochs)
self.discriminator.save (self.d_pre_path)
def build_discriminator(self, dropout=0.1):
data_input = Input( shape=(None,), dtype='int32', name='Input')
out = Embedding(self.vocab_size, self.embedding_size, mask_zero = True, name='Embedding') (data_input)
out = LSTM (self.hidden_size) (out)
out = self.Highway (out, num_layers = 1)
out = Dropout (dropout, name='Dropout') (out)
output = Dense(1, activation='sigmoid') (out)
return Model(data_input, output)
def Highway (self, x , num_layers = 1 , activation = 'relu' , name_prefix = '' ):
input_size = K.int_shape (x) [ 1 ]
for i in range (num_layers):
gate_ratio_name = '{}Highway/Gate_ratio_{}'.format (name_prefix, i)
fc_name = '{}Highway/FC_{}'.format (name_prefix, i)
gate_name = '{}Highway/Gate_{}'.format (name_prefix, i)
gate_ratio = Dense (input_size, activation = 'sigmoid' , name = gate_ratio_name) (x)
fc = Dense (input_size, activation = activation, name = fc_name) (x)
x = Lambda ( lambda args : args [ 0 ] * args [ 2 ] + args [ 1 ] * ( 1 - args [ 2 ]), name = gate_name) ([fc, x, gate_ratio])
return x
def train(self, train_data, epochs, batch_size=128, sample_interval=50, g_epochs=1):
# Load and convert the data
# x_train, y_train = prepare_data(data)
X_train = train_data
# Adversarial ground truths
real = np.ones((batch_size, 1))
fake = np.zeros((batch_size, 1))
d_adam = Adam(self.d_lr)
self.discriminator.compile(d_adam, 'binary_crossentropy')
# Training the model
for epoch in range(epochs):
# Generator training
for _ in range(g_epochs):
rewards = np.zeros([self.batch_size, self.T])
self.agent.reset()
self.env.reset()
for t in range(self.T):
state = self.env.get_state()
action = self.agent.act(state, epsilon=0.0)
next_state, reward, is_episode_end, info = self.env.step(action)
self.agent.generator.update(state, action, reward)
rewards[:, t] = reward.reshape([self.B,])
if is_episode_end:
if verbose:
print('Reward: {: .3f}, Episode end'.format(np.average(rewards)))
self.env.render(head = head)
break
# Discriminator training
for _ in range(d_epochs):
self.agent.generator.generate_samples(
self.T,
self.g_data,
self.generate_samples,
self.path_neg)
self.d_data = DiscriminatorGenerator(
path_pos = self.path_pos,
path_neg = self.path_neg,
batch_size = self.batch_size,
shuffle = True
)
self.discriminator.fit_generator(
self.d_data,
steps_per_epoch = None,
epochs = d_epochs
)
# Update env.g_beta to agent
self.agent_save (g_weights_path)
self.g_beta.load(g_weights_path)
self.discriminator.save(d_weights_path)
self.epsilon = max(self.epsilon * (1-float(step) / steps * 4), 1e-4)
# Training the discriminator
# Select a random batch of note sequences
idx = np.random.randint(0, X_train.shape[0], batch_size)
real_seqs = X_train[idx]
# Noise
noise = np.random.normal(0, 1, (batch_size, self.latent_dim))
# Generate a batch of new sequences
gen_seqs = self.generator.predict(noise)
# Train the discriminator
d_loss_real = self.discriminator.train_on_batch(real_seqs, real)
d_loss_fake = self.discriminator.train_on_batch(gen_seqs, fake)
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# Train the Generator
noise = np.random.normal(0, 1, (batch_size, self.latent_dim))
# Train the generator (to have the discriminator label samples as real)
g_loss = self. combined.train_on_batch(noise, real)
self.discriminator.save('drive/My Drive/ThesisProposal/gan-mix-discriminator.h5')
self.generator.save('drive/My Drive/ThesisProposal/gan-mix-generator.h5')
def save_model( self, g_path, d_path ):
self.agent.save (g_path)
self.discriminator.save (d_path)
def load_model(self, g_path, d_path ):
self.agent.load (g_path)
self.g_beta.load (g_path)
self.discriminator.load_weights (d_path)
# + [markdown] id="ubuuJguuDdez" colab_type="text"
#
# + id="0hOZvR1MDgxj" colab_type="code" colab={}
https://github.com/cs109/a-2017/tree/master/Lectures
https://harvard-iacs.github.io/2019-CS109B/pages/schedule.html
|
workspace4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
from six.moves import range
from PIL import Image
import sys
dir_path = '/home/avoyd/GANtor-Arts-Center/src/code/main.py'
sys.path.append(dir_path)
sys.path.append('/home/avoyd/GANtor-Arts-Center/src/code/')
import torch.backends.cudnn as cudnn
import torch
import torch.nn as nn
from torch.autograd import Variable
from torch.nn import functional as F
import torch.optim as optim
import os
import time
import numpy as np
import torchfile
from miscc.config import cfg, cfg_from_file
from miscc.utils import mkdir_p
from miscc.utils import weights_init
from miscc.utils import save_img_results, save_model
from miscc.utils import KL_loss
from miscc.utils import compute_discriminator_loss, compute_generator_loss
from tensorboard import summary
from tensorboardX import FileWriter
import torchvision
import torchvision.utils as vutils
from matplotlib import pyplot as plt
import torch.utils.data
from torchvision.models.inception import inception_v3
import numpy as np
from scipy.stats import entropy
import torchvision.transforms as transforms
import datetime
import dateutil
import dateutil.tz
from PIL import Image
import csv
import matplotlib.pyplot as plt
import numpy as np
from miscc.datasets import TextDataset
from miscc.utils import mkdir_p
from trainer import GANTrainer
# -
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# device = torch.device(0)
print(device)
print(torch.cuda.is_available())
# +
def initialize_inception_wikiart(load_path, num_classes, use_pretrained=True):
# Initialize these variables which will be set in this if statement. Each of these
# variables is model specific.
model_ft = torchvision.models.inception_v3(pretrained=use_pretrained)
#set_parameter_requires_grad(model_ft, feature_extract)
# Handle the auxilary net
num_ftrs = model_ft.AuxLogits.fc.in_features
model_ft.AuxLogits.fc = nn.Linear(num_ftrs, num_classes)
# Handle the primary net
num_ftrs = model_ft.fc.in_features
model_ft.fc = nn.Linear(num_ftrs,num_classes)
if load_path != None:
model_ft.load_state_dict(torch.load(load_path))
return model_ft
# +
def inception_score(inception_model, imgs, num_classes=27, cuda=True, batch_size=32, resize=False, splits=1,
gpu_list=None):
"""Computes the inception score of the generated images imgs
imgs -- Torch dataset of (3xHxW) numpy images normalized in the range [-1, 1]
cuda -- whether or not to run on GPU
batch_size -- batch size for feeding into Inception v3
splits -- number of splits
"""
N = len(imgs)
assert batch_size > 0
assert N > batch_size
# Set up dtype
if cuda:
dtype = torch.cuda.FloatTensor
else:
if torch.cuda.is_available():
print("WARNING: You have a CUDA device, so you should probably set cuda=True")
dtype = torch.FloatTensor
# Set up dataloader
dataloader = torch.utils.data.DataLoader(imgs, batch_size=batch_size, num_workers=8, drop_last=True)
# # Load inception model
# inception_model = inception_v3(pretrained=True, transform_input=False).type(dtype)
# inception_model.eval();
up = nn.Upsample(size=(299, 299), mode='bilinear').type(dtype)
def get_pred(x):
if resize:
x = up(x)
if gpu_list != None:
#Parallelize across >1 GPUs
x = nn.parallel.data_parallel(inception_model, x, GPU_list)
else:
x = inception_model(x)
return F.softmax(x, dim=1).data.cpu().numpy()
# Get predictions
preds = np.zeros((N, num_classes))
for i, batch in enumerate(dataloader, 0):
# batch = torch.tensor(batch)
batch = batch[0].type(dtype)
batchv = Variable(batch)
batch_size_i = batch.size()[0]
preds[i*batch_size:i*batch_size + batch_size_i] = get_pred(batchv)
# Now compute the mean kl-div
split_scores = []
for k in range(splits):
part = preds[k * (N // splits): (k+1) * (N // splits), :]
py = np.mean(part, axis=0)
scores = []
for i in range(part.shape[0]):
pyx = part[i, :]
scores.append(entropy(pyx, py))
split_scores.append(np.exp(np.mean(scores)))
return np.mean(split_scores), np.std(split_scores)
if __name__ == '__main__':
# Specify classification category and image size
category = "genre"
num_classes = 27 if category == "style" else 10
image_size = 256 # 64 for S1 outs, 256 for S2 outs
# gen_sample_dir = './baseline_generated/{}{}/'.format(category, image_size)
gen_sample_dir = './v2_generated/{}{}/'.format(category, image_size)
batch_size = 32
#Trained for 5 eps on style
# inception_path = './ft_wikiart/ft_{}_5eps.pth'.format(category)
#Trained for 15 eps on genre
inception_path = './ft_wikiart/ft_style_5eps.pth' if category == "style" else './ft_wikiart/ft_genre_15eps.pth'
print("Loading inception v3 weights from {}".format(inception_path))
# image_transform = transforms.Compose(
# [transforms.CenterCrop(GAN_inp_size),
# transforms.ToTensor(),
# transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])])
image_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
gen_sample_dataset = torchvision.datasets.ImageFolder(gen_sample_dir, transform=image_transform)
#Model expects input size of 299
inception_model = initialize_inception_wikiart(inception_path, num_classes)
inception_model.to(device)
inception_model.eval()
inception_model = nn.DataParallel(inception_model)
print ("Calculating Inception Score...")
score_mean, score_std = inception_score(inception_model, gen_sample_dataset,
num_classes=num_classes, cuda=True, batch_size=batch_size, resize=True, splits=10)
print("Inception score, mean and std: {} +- {}".format(score_mean, score_std))
# +
############IGNORE################################
# class IgnoreLabelDataset(torch.utils.data.Dataset):
# def __init__(self, orig):
# self.orig = orig
# def __getitem__(self, index):
# return self.orig[index][0]
# def __len__(self):
# return len(self.orig)
# # cifar = dset.CIFAR10(root='data/', download=True,
# # transform=transforms.Compose([
# # transforms.Scale(32),
# # transforms.ToTensor(),
# # transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
# # ])
# )
print(score_mean, score_std)
# +
## Unused func
# def initialize_Generator(
# config_path='../cfg/wikiart_s2.yml',
# S1_path = '../../../results/wikiart_stageI_2019_05_14_23_03_43/Model/netG_epoch_60.pth',
# S2_path = '../../../results/wikiart_stageII_2019_05_15_03_54_54/Model/netG_epoch_45.pth'
# ):
# cfg_from_file(config_path)
# from model import STAGE1_G, STAGE2_G, STAGE2_D
# Stage1_G = STAGE1_G()
# netG = STAGE2_G(Stage1_G)
# netG.apply(weights_init)
# stage_1_file = S1_path
# stage_2_file = S2_path
# state_dict = torch.load(stage_2_file, map_location=lambda storage, loc: storage)
# netG.load_state_dict(state_dict)
# print('Load from: ', stage_2_file)
# state_dict = torch.load(stage_1_file,map_location=lambda storage, loc: storage)
# netG.STAGE1_G.load_state_dict(state_dict)
# print('Load from: ', stage_1_file)
# if cfg.CUDA:
# netG.cuda()
# print(netG)
# return netG
|
src/code/inception/.ipynb_checkpoints/Inception Score-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 사전작업
# ## 라이브러리 로드
import numpy as np
import pandas as pd
import warnings
import gc
from tqdm import tqdm_notebook
import lightgbm as lgb
from scipy.sparse import vstack, csr_matrix, save_npz, load_npz
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.model_selection import StratifiedKFold
warnings.filterwarnings("ignore")
gc.enable()
pd.set_option('max_rows', 150)
pd.set_option('max_colwidth', 500)
pd.set_option('max_columns', 500)
# + hide_input=true
dtypes = {
'MachineIdentifier': 'object',
'ProductName': 'category',
'EngineVersion': 'category',
'AppVersion': 'category',
'AvSigVersion': 'category',
'IsBeta': 'int8',
'RtpStateBitfield': 'float16',
'IsSxsPassiveMode': 'int8',
'DefaultBrowsersIdentifier': 'float16',
'AVProductStatesIdentifier': 'float32',
'AVProductsInstalled': 'float16',
'AVProductsEnabled': 'float16',
'HasTpm': 'int8',
'CountryIdentifier': 'int16',
'CityIdentifier': 'float32',
'OrganizationIdentifier': 'float16',
'GeoNameIdentifier': 'float16',
'LocaleEnglishNameIdentifier': 'int8',
'Platform': 'category',
'Processor': 'category',
'OsVer': 'category',
'OsBuild': 'int16',
'OsSuite': 'int16',
'OsPlatformSubRelease': 'category',
'OsBuildLab': 'category',
'SkuEdition': 'category',
'IsProtected': 'float16',
'AutoSampleOptIn': 'int8',
'PuaMode': 'category',
'SMode': 'float16',
'IeVerIdentifier': 'float16',
'SmartScreen': 'category',
'Firewall': 'float16',
'UacLuaenable': 'float32',
'Census_MDC2FormFactor': 'category',
'Census_DeviceFamily': 'category',
'Census_OEMNameIdentifier': 'float16',
'Census_OEMModelIdentifier': 'float32',
'Census_ProcessorCoreCount': 'float16',
'Census_ProcessorManufacturerIdentifier': 'float16',
'Census_ProcessorModelIdentifier': 'float16',
'Census_ProcessorClass': 'category',
'Census_PrimaryDiskTotalCapacity': 'float32',
'Census_PrimaryDiskTypeName': 'category',
'Census_SystemVolumeTotalCapacity': 'float32',
'Census_HasOpticalDiskDrive': 'int8',
'Census_TotalPhysicalRAM': 'float32',
'Census_ChassisTypeName': 'category',
'Census_InternalPrimaryDiagonalDisplaySizeInInches': 'float16',
'Census_InternalPrimaryDisplayResolutionHorizontal': 'float16',
'Census_InternalPrimaryDisplayResolutionVertical': 'float16',
'Census_PowerPlatformRoleName': 'category',
'Census_InternalBatteryType': 'category',
'Census_InternalBatteryNumberOfCharges': 'float32',
'Census_OSVersion': 'category',
'Census_OSArchitecture': 'category',
'Census_OSBranch': 'category',
'Census_OSBuildNumber': 'int16',
'Census_OSBuildRevision': 'int32',
'Census_OSEdition': 'category',
'Census_OSSkuName': 'category',
'Census_OSInstallTypeName': 'category',
'Census_OSInstallLanguageIdentifier': 'float16',
'Census_OSUILocaleIdentifier': 'int16',
'Census_OSWUAutoUpdateOptionsName': 'category',
'Census_IsPortableOperatingSystem': 'int8',
'Census_GenuineStateName': 'category',
'Census_ActivationChannel': 'category',
'Census_IsFlightingInternal': 'float16',
'Census_IsFlightsDisabled': 'float16',
'Census_FlightRing': 'category',
'Census_ThresholdOptIn': 'float16',
'Census_FirmwareManufacturerIdentifier': 'float16',
'Census_FirmwareVersionIdentifier': 'float32',
'Census_IsSecureBootEnabled': 'int8',
'Census_IsWIMBootEnabled': 'float16',
'Census_IsVirtualDevice': 'float16',
'Census_IsTouchEnabled': 'int8',
'Census_IsPenCapable': 'int8',
'Census_IsAlwaysOnAlwaysConnectedCapable': 'float16',
'Wdft_IsGamer': 'float16',
'Wdft_RegionIdentifier': 'float16',
'HasDetections': 'int8'
}
# -
# ## 데이터 로드
# %%time
train = pd.read_csv('./data/train.csv', dtype=dtypes)
test = pd.read_csv('./data/test.csv', dtype=dtypes)
# ## train, test 합치기
test['HasDetections'] = np.nan
data = train.append(test)
data.reset_index(drop=True, inplace=True)
data = data.reset_index().drop(['MachineIdentifier'], axis=1).rename(columns={'index':'MachineIdentifier'})
del train, test
gc.collect()
# ## Get Date - AvSigVersion
datedict = np.load('./data/AvSigVersionTimestamps.npy')
datedict = datedict[()]
data['Date'] = data['AvSigVersion'].map(datedict)
data['Date_YMD'] = pd.to_datetime(data['Date'].astype(str).str.slice(0, 10))
del datedict
test = data[data.HasDetections.isna()].groupby(['Date_YMD']).HasDetections.agg(['size']).reset_index()
test[test['Date_YMD'] >= '2018-10-25']['size'].sum()
str2 = 'dndnkfdn(dff)'
str2.replace('(dff)', '')
# ## Train, Test split
train = data[~data.HasDetections.isna()]
test = data[data.HasDetections.isna()]
del data
gc.collect()
# +
train.MachineIdentifier = range(len(train))
train.reset_index(drop=True, inplace=True)
test.MachineIdentifier = range(len(test))
test.reset_index(drop=True, inplace=True)
# -
debug = False
if debug:
train = train[:10000]
test = test[:10000]
print('Transform all features to category.\n')
for usecol in tqdm_notebook(train.columns.tolist()[1:-1]):
train[usecol] = train[usecol].astype('str')
test[usecol] = test[usecol].astype('str')
#Fit LabelEncoder
le = LabelEncoder().fit(
np.unique(train[usecol].unique().tolist()+
test[usecol].unique().tolist()))
#At the end 0 will be used for dropped values
train[usecol] = le.transform(train[usecol])+1
test[usecol] = le.transform(test[usecol])+1
agg_tr = (train
.groupby([usecol])
.aggregate({'MachineIdentifier':'count'})
.reset_index()
.rename({'MachineIdentifier':'Train'}, axis=1))
agg_te = (test
.groupby([usecol])
.aggregate({'MachineIdentifier':'count'})
.reset_index()
.rename({'MachineIdentifier':'Test'}, axis=1))
agg = pd.merge(agg_tr, agg_te, on=usecol, how='outer').replace(np.nan, 0)
#Select values with more than 1000 observations
agg = agg[(agg['Train'] > 1000)].reset_index(drop=True)
agg['Total'] = agg['Train'] + agg['Test']
#Drop unbalanced values
agg = agg[(agg['Train'] / agg['Total'] > 0.2) & (agg['Train'] / agg['Total'] < 0.8)]
agg[usecol+'Copy'] = agg[usecol]
train[usecol] = (pd.merge(train[[usecol]],
agg[[usecol, usecol+'Copy']],
on=usecol, how='left')[usecol+'Copy']
.replace(np.nan, 0).astype('int').astype('category'))
test[usecol] = (pd.merge(test[[usecol]],
agg[[usecol, usecol+'Copy']],
on=usecol, how='left')[usecol+'Copy']
.replace(np.nan, 0).astype('int').astype('category'))
del le, agg_tr, agg_te, agg, usecol
gc.collect()
train.shape
y_train = np.array(train['HasDetections'])
train_ids = train.index
test_ids = test.index
del train['HasDetections'], train['MachineIdentifier'], test['MachineIdentifier'], test['HasDetections']
gc.collect()
# +
print("If you don't want use Sparse Matrix choose Kernel Version 2 to get simple solution.\n")
print('--------------------------------------------------------------------------------------------------------')
print('Transform Data to Sparse Matrix.')
print('Sparse Matrix can be used to fit a lot of models, eg. XGBoost, LightGBM, Random Forest, K-Means and etc.')
print('To concatenate Sparse Matrices by column use hstack()')
print('Read more about Sparse Matrix https://docs.scipy.org/doc/scipy/reference/sparse.html')
print('Good Luck!')
print('--------------------------------------------------------------------------------------------------------')
# -
#Fit OneHotEncoder
ohe = OneHotEncoder(categories='auto', sparse=True, dtype='uint8').fit(train)
#Transform data using small groups to reduce memory usage
m = 100000
train = vstack([ohe.transform(train[i*m:(i+1)*m]) for i in range(train.shape[0] // m + 1)])
test = vstack([ohe.transform(test[i*m:(i+1)*m]) for i in range(test.shape[0] // m + 1)])
train.shape
save_npz('./data_temp/train.npz', train, compressed=True)
save_npz('./data_temp/test.npz', test, compressed=True)
del ohe, train, test
gc.collect()
train = load_npz('./data_temp/train.npz')
test = load_npz('./data_temp/test.npz')
test = csr_matrix(test, dtype='float32')
# +
skf = StratifiedKFold(n_splits=3, shuffle=True, random_state=42)
skf.get_n_splits(train_ids, y_train)
lgb_test_result = np.zeros(test_ids.shape[0])
lgb_train_result = np.zeros(train_ids.shape[0])
counter = 0
# +
print('\nLightGBM\n')
for train_index, test_index in skf.split(train_ids, y_train):
print('Fold {}\n'.format(counter + 1))
X_fit = vstack([train[train_index[i*m:(i+1)*m]] for i in range(train_index.shape[0] // m + 1)])
X_val = vstack([train[test_index[i*m:(i+1)*m]] for i in range(test_index.shape[0] // m + 1)])
X_fit, X_val = csr_matrix(X_fit, dtype='float32'), csr_matrix(X_val, dtype='float32')
y_fit, y_val = y_train[train_index], y_train[test_index]
gc.collect()
lgb_model = lgb.LGBMClassifier(max_depth=-1,
n_estimators=1000,
learning_rate=0.1,
num_leaves=2**5-1,
objective='binary',
boosting_type='gbdt',
# overfitting handling
# max_bin=120,
# lambda_l1=6,
# lambda_l2=2,
save_binary=True,
feature_fraction=0.8,
feature_fraction_seed=42,
n_jobs=-1)
print("fitting")
lgb_model.fit(X_fit, y_fit, eval_metric='auc',
eval_set=[(X_val, y_val)],
verbose=200, early_stopping_rounds=100)
del X_fit, X_val, y_fit, y_val, train_index, test_index
gc.collect()
print("predicting")
lgb_test_result += lgb_model.predict_proba(test)[:,1]
counter += 1
gc.collect()
# -
submission = pd.read_csv('./data/sample_submission.csv')
submission.head(3)
submission.HasDetections = lgb_test_result / counter
submission.head(3)
submission.to_csv('./data/sub_lgb_base_open_kernel.csv', index=False)
param = {
'objective': 'binary',
'boosting_type': 'gbdt',
'learning_rate': 0.05,
'max_depth': -1,
'num_leaves': 31,
'min_data_in_leaf': 20,
'min_sum_hessian_in_leaf': 0.0025,
'max_bin': 120,
'lambda_l1': 5,
'lambda_l2': 2,
'min_gain_to_split': 0.65,
'save_binary': True,
'bagging_fraction': 1.0,
'bagging_freq': 5,
'feature_fraction': 0.05,
'seed': 42,
'feature_fraction_seed': 42,
'bagging_seed': 42,
'drop_seed': 42,
'data_random_seed': 42,
'verbose': 1,
'metric': 'auc'
}
max_depth=-1,
n_estimators=1000,
learning_rate=0.1,
num_leaves=2**5-1,
objective='binary',
boosting_type='gbdt',
save_binary=True,
feature_fraction=0.8,
feature_fraction_seed=42,
n_jobs=-1
|
Microsoft Malware Prediction/code/preprocessing/03-1 FE - Private 1 Public 0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chavgova/My-AI/blob/master/emotion_recognition_03.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="eFA7SmfR-yKp" colab_type="text"
# IMPORT
# + id="8-N_FynR2O4E" colab_type="code" colab={}
#this is the copy of another projecct and ill make changes to see how i can make it better
import librosa
import librosa.display
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from matplotlib.pyplot import specgram
import keras
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import Dense, Embedding
from keras.layers import LSTM
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.layers import Input, Flatten, Dropout, Activation
from keras.layers import Conv1D, MaxPooling1D, AveragePooling1D
from keras.models import Model
from keras.callbacks import ModelCheckpoint
from sklearn.metrics import confusion_matrix
from keras import regularizers
import os
# + id="ACpe5DYcCxWD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="e6277e56-9455-4db4-e315-f2a2a4b7ebb6"
from google.colab import drive
import os
path = '/content/drive/My Drive/My_AI/RawData'
mylist = []
mylist = os.listdir(path)
print(mylist)
# + id="Ut4HPDFDZXch" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="37649a62-f276-45a2-8fff-aa7d691c28da"
print(mylist[50])
# + id="mVJN1WAQ5s_r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="741b9db8-e672-481e-ff22-15ca3b1b483c"
print(mylist[18][6:-16])
# + [markdown] id="ERpg_GG5--ee" colab_type="text"
# LABLES
# + id="L4vr7Ssh-jrl" colab_type="code" colab={}
feeling_list=[]
for item in mylist:
if int(item[18:-4])%2==0: #female
if item[6:-16]=='01':
feeling_list.append('female_neutral')
elif item[6:-16]=='02':
feeling_list.append('female_calm')
elif item[6:-16]=='03':
feeling_list.append('female_happy')
elif item[6:-16]=='04':
feeling_list.append('female_sad')
elif item[6:-16]=='05':
feeling_list.append('female_angry')
elif item[6:-16]=='06':
feeling_list.append('female_fearful')
elif item[6:-16]=='07':
feeling_list.append('female_disgust')
elif item[6:-16]=='08':
feeling_list.append('female_surprised')
else:
if item[6:-16]=='01':
feeling_list.append('male_neutral')
elif item[6:-16]=='02':
feeling_list.append('male_calm')
elif item[6:-16]=='03':
feeling_list.append('male_happy')
elif item[6:-16]=='04':
feeling_list.append('male_sad')
elif item[6:-16]=='05':
feeling_list.append('male_angry')
elif item[6:-16]=='06':
feeling_list.append('male_fearful')
elif item[6:-16]=='07':
feeling_list.append('male_disgust')
elif item[6:-16]=='08':
feeling_list.append('male_surprised')
# + id="TCiFI41--eBm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 343} outputId="2699b614-6746-4d17-fe50-7b46eb6513a9"
import pandas as pd
labels = pd.DataFrame(feeling_list)
labels[:10] #print
# + [markdown] id="pRYX3XLu_Fd8" colab_type="text"
# Getting the features of audio files using librosa
# + id="Ntnk_M9jZlub" colab_type="code" colab={}
import librosa
import numpy as np
def extract_feature(my_file, **kwargs):
mfcc = kwargs.get("mfcc")
chroma = kwargs.get("chroma")
mel = kwargs.get("mel")
contrast = kwargs.get("contrast")
tonnetz = kwargs.get("tonnetz")
X, sample_rate = librosa.core.load(my_file)
if chroma or contrast:
stft = np.abs(librosa.stft(X))
result = np.array([])
if mfcc:
mfccs = np.mean(librosa.feature.mfcc(y=X, sr=sample_rate, n_mfcc=40).T, axis=0)
result = np.hstack((result, mfccs))
if chroma:
chroma = np.mean(librosa.feature.chroma_stft(S=stft, sr=sample_rate).T,axis=0)
result = np.hstack((result, chroma))
if mel:
mel = np.mean(librosa.feature.melspectrogram(X, sr=sample_rate).T,axis=0)
result = np.hstack((result, mel))
if contrast:
contrast = np.mean(librosa.feature.spectral_contrast(S=stft, sr=sample_rate).T,axis=0)
result = np.hstack((result, contrast))
if tonnetz:
tonnetz = np.mean(librosa.feature.tonnetz(y=librosa.effects.harmonic(X), sr=sample_rate).T,axis=0)
result = np.hstack((result, tonnetz))
return result
f = os.fspath('/content/drive/My Drive/My_AI/RawData/03-01-08-01-01-02-01.wav')
a = extract_feature(f, mel=True, mfcc=True, contrast=True, chroma=True, tonnetz=True)
#print(a, a.shape)
# + id="ZwW0DUUU_GbG" colab_type="code" colab={}
df = pd.DataFrame(columns=['all_features'])
bookmark=0
#mylist = mylist[:100]
for index,y in enumerate(mylist):
all_features_ndarray = extract_feature('/content/drive/My Drive/My_AI/RawData/'+y, mel=True, mfcc=True, contrast=True, chroma=True, tonnetz=True)
df.loc[bookmark] = [all_features_ndarray]
bookmark=bookmark+1
#df[:5] #print
# + id="hR-qAHPe_W6J" colab_type="code" colab={}
df3 = pd.DataFrame(df['all_features'].values.tolist())
# + id="_yqMEDwR_W0h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 560} outputId="78d5deaa-d1db-455c-f436-f945b6ba7526"
newdf = pd.concat([df3,labels], axis=1)
rnewdf = newdf.rename(index=str, columns={"0": "label"})
rnewdf[:10] #print
# + id="EjFS7uNn_hta" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 560} outputId="d551c6c6-55ea-4879-e12d-2a4395febd22"
from sklearn.utils import shuffle
rnewdf = shuffle(newdf)
rnewdf[:10] #print
# + id="K9GjC-P6_kp9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="f4bc14b2-591e-4eb8-db02-9b35856104d2"
'''
labels = rnewdf.pop(-1)
print(labels)
ds_norm=(rnewdf-rnewdf.min())/(rnewdf.max()-rnewdf.min())
ds_norm['labels'] = labels
ds_norm=ds_norm.fillna(0)
'''
# + [markdown] id="PJpwFs_b5Ps3" colab_type="text"
#
# Dividing the data into test and train
# + id="2pNznFkU_kjc" colab_type="code" colab={}
newdf1 = np.random.rand(len(rnewdf)) < 0.8
train = rnewdf[newdf1]
test = rnewdf[~newdf1]
# + id="6K8aywep5T6w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 560} outputId="8bb926d9-df9e-47be-cc62-e0034448c9db"
train[250:260]
# + id="qPL9iaKR5XFB" colab_type="code" colab={}
trainfeatures = train.iloc[:, :-1]
trainlabel = train.iloc[:, -1:]
# + id="ql_03gWC5eYp" colab_type="code" colab={}
testfeatures = test.iloc[:, :-1]
testlabel = test.iloc[:, -1:]
# + id="1YwvIzsI5nU4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="98c01641-f8a7-477f-c003-3d19c238a073"
from keras.utils import np_utils
from sklearn.preprocessing import LabelEncoder
X_train = np.array(trainfeatures)
y_train = np.array(trainlabel)
X_test = np.array(testfeatures)
y_test = np.array(testlabel)
lb = LabelEncoder()
y_train = np_utils.to_categorical(lb.fit_transform(y_train))
y_test = np_utils.to_categorical(lb.fit_transform(y_test))
# + id="OhC3quOd5p3Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="edc16016-3316-4aea-d30b-8c6edcac79c8"
y_train
# + id="Wbaw6I3G5twE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7bb6c77c-01f0-4eb4-e96f-24c80aafdd1e"
X_train.shape
# + [markdown] id="KIVYxQMq5yRs" colab_type="text"
# Changing dimension for CNN model
# + id="bBHW97HY5x-8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 840} outputId="3ed6d81d-67bb-4128-beb4-280fdb7b207a"
x_traincnn =np.expand_dims(X_train, axis=2)
x_testcnn= np.expand_dims(X_test, axis=2)
print(x_testcnn)
# + id="7CO4IkGp52Rn" colab_type="code" colab={}
model = Sequential()
model.add(Conv1D(256, 5,padding='same',
input_shape=(193,1)))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same'))
model.add(Activation('relu'))
model.add(Dropout(0.1))
model.add(MaxPooling1D(pool_size=(8)))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Conv1D(128, 5,padding='same',))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(16))
model.add(Activation('softmax'))
opt = keras.optimizers.RMSprop(lr=0.00001, decay=1e-6)
# + id="fThlzaPU5_ui" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 756} outputId="e1cf0be4-64a0-4e39-ae47-4e8226fe73b4"
model.summary()
# + id="-J_L_NQb6C9b" colab_type="code" colab={}
model.compile(loss='categorical_crossentropy', optimizer=opt,metrics=['accuracy'])
# + [markdown] id="16X3w8Sc6HC8" colab_type="text"
# Removed the whole training part for avoiding unnecessary long epochs list
# + id="3jpf-3Zh6IxI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="e12ac657-87d7-4848-cbe3-b985a62ba627"
cnnhistory=model.fit(x_traincnn, y_train, batch_size=32, epochs=250, validation_data=(x_testcnn, y_test))
# + id="cfN9RhED6Lrs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="dfe7c9e1-3722-4d72-9baf-f503765ca188"
plt.figure()
plt.plot(cnnhistory.history['loss'])
plt.plot(cnnhistory.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.grid(True)
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + id="Bcx0kn54SrWo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="d97405d9-0602-4b2c-e318-026884eb1c4d"
tf.keras.utils.plot_model(
model,
to_file="model.png",
show_shapes=False,
show_layer_names=True,
rankdir="TB",
expand_nested=False,
dpi=96,
)
dot_img_file = '/content/drive/My Drive/My_AI/img.png'
tf.keras.utils.plot_model(model, to_file=dot_img_file, show_shapes=True)
# + [markdown] id="TgoIRnXx5Kwj" colab_type="text"
# SAVING THE MODEL
# + id="k10VeAGW5KKl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7be8e4dc-6b9d-4a70-d153-17db35ca17b2"
model_name = 'Emotion_Voice_Detection_CNN.h5'
path = '/content/drive/My Drive/My_AI/'
save_dir = os.path.join(os.getcwd(), 'saved_models')
# Save model and weights
if not os.path.isdir(save_dir):
os.makedirs(save_dir)
model_path = os.path.join(save_dir, model_name)
model.save(model_path)
print('Saved trained model at %s ' % model_path)
# + id="JHI_Y6tg5aqe" colab_type="code" colab={}
import json
model_json = model.to_json()
with open("/content/drive/My Drive/My_AI/Voice-Emotion-Detector-master/model.json", "w") as json_file:
json_file.write(model_json)
# + [markdown] id="cAXCT5O15bl1" colab_type="text"
# LOADING THE MODEL
# + id="iBPoY_-45agM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="23c5b4c2-5134-4367-d283-079879352ab5"
# loading json and creating model
from keras.models import model_from_json
json_file = open('/content/drive/My Drive/My_AI/Voice-Emotion-Detector-master/model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("/content/saved_models/Emotion_Voice_Detection_CNN.h5")
print("Loaded model from disk")
# evaluate loaded model on test data
loaded_model.compile(loss='categorical_crossentropy', optimizer=opt, metrics=['accuracy'])
score = loaded_model.evaluate(x_testcnn, y_test, verbose=0)
print("%s: %.2f%%" % (loaded_model.metrics_names[1], score[1]*100))
# + [markdown] id="qoFefCox6jRe" colab_type="text"
# Predicting emotions on the test data
# + id="soFuVUkh6kbW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 360} outputId="7498feac-6331-4ff2-afb6-d2c360ec430e"
preds = loaded_model.predict(x_testcnn, batch_size=32, verbose=1)
preds1=preds.argmax(axis=1)
abc = preds1.astype(int).flatten()
predictions = (lb.inverse_transform((abc)))
preddf = pd.DataFrame({'predictedvalues': predictions})
actual=y_test.argmax(axis=1)
abc123 = actual.astype(int).flatten()
actualvalues = (lb.inverse_transform((abc123)))
actualdf = pd.DataFrame({'actualvalues': actualvalues})
finaldf = actualdf.join(preddf)
finaldf[10:20]
# + id="QjtRhwAxeZ2j" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="b146ac28-1c28-4423-ba5e-8cd0a24eeccf"
finaldf.groupby('actualvalues').count()
# + id="QZsfui1hwknw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 550} outputId="f489daa3-52bf-401c-f9e4-7ab96d00fec0"
finaldf.groupby('predictedvalues').count()
|
Male&Female Previous Experiments/emotion_recognition_03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import geemap
import ee
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
print(classifier.explain())
test = composite.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_pov(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_pov)
chart = ui.Chart.array.values({
'array': ee.Array(accuracies),
'axis': 0,
'xLabels': numTreesList
}).setOptions({
'title': 'Hyperparameter Tuning for the numberOfTrees Parameters',
'vAxis': '{title': 'Validation Accuracy'},
'hAxis': '{title': 'Number of Tress', 'gridlines': '{count': 15}}
})
print(chart)
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_xry(numTrees):
return bagFractionList.map(function(bagFraction) {
classifier = ee.Classifier.smileRandomForest({
'numberOfTrees': numTrees,
'bagFraction': bagFraction
}) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,
'numberOfTrees': numTrees,
'bagFraction': bagFraction})
})
accuracies = numTreesList.map(func_xry
).flatten()
).flatten()
resultFc = ee.FeatureCollection(accuracies)
# Export the result as CSV
Export.table.toDrive({
'collection': resultFc,
'description': 'Multiple_Parameter_Tuning_Results',
'folder': 'earthengine',
'fileNamePrefix': 'numtrees_bagfraction',
'fileFormat': 'CSV'})
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
print(classifier.explain())
test = composite.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_pov(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_pov)
chart = ui.Chart.array.values({
'array': ee.Array(accuracies),
'axis': 0,
'xLabels': numTreesList
}).setOptions({
'title': 'Hyperparameter Tuning for the numberOfTrees Parameters',
'vAxis': '{title': 'Validation Accuracy'},
'hAxis': '{title': 'Number of Tress', 'gridlines': '{count': 15}}
})
print(chart)
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_xry(numTrees):
return bagFractionList.map(function(bagFraction) {
classifier = ee.Classifier.smileRandomForest({
'numberOfTrees': numTrees,
'bagFraction': bagFraction
}) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,
'numberOfTrees': numTrees,
'bagFraction': bagFraction})
})
accuracies = numTreesList.map(func_xry
).flatten()
).flatten()
resultFc = ee.FeatureCollection(accuracies)
# Export the result as CSV
Export.table.toDrive({
'collection': resultFc,
'description': 'Multiple_Parameter_Tuning_Results',
'folder': 'earthengine',
'fileNamePrefix': 'numtrees_bagfraction',
'fileFormat': 'CSV'})
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
print(classifier.explain())
test = composite.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_dmn(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_dmn)
chart = ui.Chart.array.values({
'array': ee.Array(accuracies),
'axis': 0,
'xLabels': numTreesList
}).setOptions({
'title': 'Hyperparameter Tuning for the numberOfTrees Parameters',
'vAxis': '{title': 'Validation Accuracy'},
'hAxis': '{title': 'Number of Tress', 'gridlines': '{count': 15}}
})
print(chart)
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_fdj(numTrees):
return bagFractionList.map(function(bagFraction) {
classifier = ee.Classifier.smileRandomForest({
'numberOfTrees': numTrees,
'bagFraction': bagFraction
}) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,
'numberOfTrees': numTrees,
'bagFraction': bagFraction})
})
accuracies = numTreesList.map(func_fdj
).flatten()
).flatten()
resultFc = ee.FeatureCollection(accuracies)
# Export the result as CSV
Export.table.toDrive({
'collection': resultFc,
'description': 'Multiple_Parameter_Tuning_Results',
'folder': 'earthengine',
'fileNamePrefix': 'numtrees_bagfraction',
'fileFormat': 'CSV'})
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.select('B.*')
composite = filtered.median().clip(boundary)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'tileScale': 16,
'scale': 10,
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.select('B.*')
composite = filtered.median().clip(boundary)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'tileScale': 16,
'scale': 10,
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
print(classifier.explain())
test = composite.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_dmn(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_dmn)
chart = ui.Chart.array.values({
'array': ee.Array(accuracies),
'axis': 0,
'xLabels': numTreesList
}).setOptions({
'title': 'Hyperparameter Tuning for the numberOfTrees Parameters',
'vAxis': '{title': 'Validation Accuracy'},
'hAxis': '{title': 'Number of Tress', 'gridlines': '{count': 15}}
})
print(chart)
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_fdj(numTrees):
return bagFractionList.map(function(bagFraction) {
classifier = ee.Classifier.smileRandomForest({
'numberOfTrees': numTrees,
'bagFraction': bagFraction
}) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,
'numberOfTrees': numTrees,
'bagFraction': bagFraction})
})
accuracies = numTreesList.map(func_fdj
).flatten()
).flatten()
resultFc = ee.FeatureCollection(accuracies)
# Export the result as CSV
Export.table.toDrive({
'collection': resultFc,
'description': 'Multiple_Parameter_Tuning_Results',
'folder': 'earthengine',
'fileNamePrefix': 'numtrees_bagfraction',
'fileFormat': 'CSV'})
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.select('B.*')
composite = filtered.median().clip(boundary)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'tileScale': 16,
'scale': 10,
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
# The function also divides ths image by 10000 to ensure
# the pixels values are between 0 and 1
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
# We divide the elevation and slope with maximum values in the region
# to ensure the values are between 0 and 1
# A more robust technique for image normalization is provided in the course supplement
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
# The function also divides ths image by 10000 to ensure
# the pixels values are between 0 and 1
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
# We divide the elevation and slope with maximum values in the region
# to ensure the values are between 0 and 1
# A more robust technique for image normalization is provided in the course supplement
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
# The function also divides ths image by 10000 to ensure
# the pixels values are between 0 and 1
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
# We divide the elevation and slope with maximum values in the region
# to ensure the values are between 0 and 1
# A more robust technique for image normalization is provided in the course supplement
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
# The function also divides ths image by 10000 to ensure
# the pixels values are between 0 and 1
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
# We divide the elevation and slope with maximum values in the region
# to ensure the values are between 0 and 1
# A more robust technique for image normalization is provided in the course supplement
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.select('B.*')
composite = filtered.median().clip(boundary)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
# Overlay the point on the image to get training data.
training = composite.sampleRegions({
'collection': trainingGcp,
'properties': ['landcover'],
'scale': 10,
'tileScale': 16
})
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train({
'features': training,
'classProperty': 'landcover',
'inputProperties': composite.bandNames()
})
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions({
'collection': validationGcp,
'properties': ['landcover'],
'tileScale': 16,
'scale': 10,
})
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix)
print('Test Accuracy', testConfusionMatrix.accuracy())
# -
Map=geemap.Map()
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(bangalore))
composite = filtered.median().clip(bangalore)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# +
geometry = ee.Geometry.Point([77.60412933051538, 12.952912912328241])
s2 = ee.ImageCollection("COPERNICUS/S2")
# Computing stats on a list
myList = ee.List.sequence(1, 10)
print(myList)
# Use a reducer to compute min and max in the list
mean = myList.reduce(ee.Reducer.mean())
print(mean)
# Apply a reducer on a image collection
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2020-01-01')) \
.filter(ee.Filter.bounds(geometry)) \
.select('B.*')
print(filtered.size())
collMean = filtered.reduce(ee.Reducer.mean())
print('Reducer on Collection', collMean)
image = ee.Image(filtered.first())
# If we want to compute min and max for each band, use reduceRegion instead
stats = image.reduceRegion({
'reducer': ee.Reducer.mean(),
'geometry': image.geometry(),
'scale': 100,
'maxPixels': 1e10
})
print(stats)
# Result of reduceRegion is a dictionary.
# We can extract the values using .get() function
print('Average value in B4', stats.get('B4'))
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(bangalore))
composite = filtered.median().clip(bangalore)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# +
geometry = ee.Geometry.Point([77.60412933051538, 12.952912912328241])
s2 = ee.ImageCollection("COPERNICUS/S2")
# Computing stats on a list
myList = ee.List.sequence(1, 10)
print(myList)
# Use a reducer to compute min and max in the list
mean = myList.reduce(ee.Reducer.mean())
print(mean)
# Apply a reducer on a image collection
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2020-01-01')) \
.filter(ee.Filter.bounds(geometry)) \
.select('B.*')
print(filtered.size())
collMean = filtered.reduce(ee.Reducer.mean())
print('Reducer on Collection', collMean)
image = ee.Image(filtered.first())
# If we want to compute min and max for each band, use reduceRegion instead
stats = image.reduceRegion({
'reducer': ee.Reducer.mean(),
'geometry': image.geometry(),
'scale': 100,
'maxPixels': 1e10
})
print(stats)
# Result of reduceRegion is a dictionary.
# We can extract the values using .get() function
print('Average value in B4', stats.get('B4'))
# -
Map=geemap.Map()
Map
roi=Map.draw_last_feature
# +
#*
# Function to mask clouds using the Sentinel-2 QA band
# @param {ee.Image} image Sentinel-2 image
# @return {ee.Image} cloud masked Sentinel-2 image
#
def maskS2clouds(image):
qa = image.select('QA60')
# Bits 10 and 11 are clouds and cirrus, respectively.
cloudBitMask = 1 << 10
cirrusBitMask = 1 << 11
# Both flags should be set to zero, indicating clear conditions.
mask = qa.bitwiseAnd(cloudBitMask).eq(0) \
.And(qa.bitwiseAnd(cirrusBitMask).eq(0))
return image.updateMask(mask).divide(10000)
dataset = ee.ImageCollection('COPERNICUS/S2_SR')\
.filterBounds(roi.geometry())\
.filterDate('2020-01-01', '2020-01-30') \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE',20)) \
.map(maskS2clouds)
visualization = {
'min': 0.0,
'max': 0.3,
'bands': ['B4', 'B3', 'B2'],
}
lat,log=roi.geometry().coordinates().getInfo()
Map.setCenter(lat, log, 12)
Map.addLayer(dataset.mean(), visualization, 'RGB')
# -
Map=geemap.Map()
s2= ee.ImageCollection('COPERNICUS/S2')
roi=Map.draw_last_feature
roi.getInfo()
# +
dataset=ee.ImageCollection('COPERNICUS/S2_SR')\
.filterBounds(roi.geometry())\
.filter(ee.Filter.date('2001-06-05','2021-06-05'))\
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE',20))\
.filter(ee.Filter.eq('SPACECRAFT_NAME','Sentinel-2A'))
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
lat,log=roi.geometry().coordinates().getInfo()
Map.setCenter(lat, log, 12)
mosaic = dataset.mosaic()
medianComposite = dataset.median();
Map.addLayer(dataset, rgbVis, 'Filtered Collection');
Map.addLayer(mosaic, rgbVis, 'Mosaic');
Map.addLayer(medianComposite, rgbVis, 'Median Composite')
# -
Map=geemap.Map()
Map
# +
states = ee.FeatureCollection("FAO/GAUL_SIMPLIFIED_500m/2015/level2")
MadhyaPradesh = states.filter(ee.Filter.eq('ADM2_NAME', 'Varanasi'))
visParams = {'color': 'red'}
Map.addLayer(MadhyaPradesh, visParams, 'Karnataka Districts')
# -
filtered2020 = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2020-01-01', '2021-01-01'))\
.filter(ee.Filter.bounds(roi.geometry()))
image2020=filtered2020.median()
Map.addLayer(image2020,rgbVis,'2020')
# # PART2
Map=geemap.Map()
Map
from datetime import datetime
now=datetime.now()
now=ee.Date(now)
past=now.advance(-1,'year')
roi=Map.draw_last_feature
lat,long=roi.geometry().coordinates().getInfo()
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR");
admin2 = ee.FeatureCollection("FAO/GAUL_SIMPLIFIED_500m/2015/level2")
gwalior = admin2.filter(ee.Filter.eq('ADM2_NAME', 'Gwalior'))
geometry = gwalior.geometry()
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.bounds(geometry)).filterDate('2019-01-01', '2020-01-01')
image = filtered.median();
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi']);
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi']);
savi = image.expression(
'1.5 * ((NIR - RED) / (NIR + RED + 0.5))', {
'NIR': image.select('B8').multiply(0.0001),
'RED': image.select('B4').multiply(0.0001),
}).rename('savi');
ndbi = image.expression(
'(SWIR1 - NIR) /(SWIR1 + NIR)', {
'SWIR1': image.select('B11').multiply(0.0001),
'NIR': image.select('B8').multiply(0.0001),
}).rename('ndbi');
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
ndviVis = {"min":0, "max":1, "palette": ['white', 'green']}
ndwiVis = {"min":0, "max":0.5, "palette": ['white', 'blue']}
ndbiVis = {"min":-1, "max":1, "palette":['Red','black']}
Map.set_center(lat, long,12)
Map.addLayer(image.clip(geometry), rgbVis, 'Image');
Map.addLayer(mndwi.clip(geometry), ndwiVis, 'mndwi')
Map.addLayer(savi.clip(geometry), ndviVis, 'savi')
Map.addLayer(ndvi.clip(geometry), ndviVis, 'ndvi')
Map.addLayer(ndbi.clip(geometry),ndbiVis, 'ndbi')
# -
Map=geemap.Map()
Map
# +
s2 = ee.ImageCollection("COPERNICUS/S2");
admin1 = ee.FeatureCollection("FAO/GAUL_SIMPLIFIED_500m/2015/level1");
MadhyaPradesh= admin1.filter(ee.Filter.eq('ADM1_NAME', 'Madhya Pradesh'))
geometry = MadhyaPradesh.geometry()
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2020-01-01'))\
.filter(ee.Filter.bounds(geometry))
composite = filtered.median().clip(geometry)
Map.addLayer(composite, rgbVis, 'MadhyaPradesh Composite')
# -
Map=geemap.Map()
Map
# +
s2 = ee.ImageCollection("COPERNICUS/S2");
admin1 = ee.FeatureCollection("FAO/GAUL_SIMPLIFIED_500m/2015/level1");
madhyapradesh = admin1.filter(ee.Filter.eq('ADM1_NAME', 'Madhya Pradesh'))
geometry = madhyapradesh.geometry()
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2020-01-01'))\
.filter(ee.Filter.bounds(geometry))
composite = filtered.median().clip(geometry)
Map.addLayer(composite, rgbVis, 'MadhyaPradesh Composite')
#// Write a function that computes NDVI for an image and adds it as a band
def addNDVI(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename('ndvi');
ndwi = image.normalizedDifference(['B3', 'B8']).rename('ndwi');
return image.addBands(ndvi).addBands(ndwi);
withNdvi = filtered.map(addNDVI);
composite = withNdvi.median()
ndviComposite = composite.select('ndwi').clip(madhyapradesh)
palette = [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718',
'74A901', '66A000', '529400', '3E8601', '207401', '056201',
'004C00', '023B01', '012E01', '011D01', '011301'];
ndviVis = {"min":0, "max":0.5, "palette": ["white","blue"] }
Map.addLayer(ndviComposite, ndviVis, 'ndwi')
# -
# # CLOUD MASKING
Map=geemap.Map()
Map
# +
image = ee.Image('COPERNICUS/S2/20190703T050701_20190703T052312_T43PGP')
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
Map.centerObject(image)
Map.addLayer(image, rgbVis, 'Full Image')
#// Write a function for Cloud masking
def maskS2clouds(image):
qa = image.select('QA60')
cloudBitMask = 1 << 10;
cirrusBitMask = 1 << 11;
mask = qa.bitwiseAnd(cloudBitMask).eq(0).And(qa.bitwiseAnd(cirrusBitMask).eq(0))
return image.updateMask(mask).select("B.*").copyProperties(image, ["system:time_start"])
maskedImage = ee.Image(maskS2clouds(image))
Map.addLayer(maskedImage, rgbVis, 'Masked Image')
# -
Map=geemap.Map()
Map
# +
imageSR = ee.Image('COPERNICUS/S2_SR/20190703T050701_20190703T052312_T43PGP')
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
Map.centerObject(imageSR)
#Map.addLayer(imageSR, rgbVis, 'SR Image')
#// Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB');
snowProb = image.select('MSK_SNWPRB');
cloud = cloudProb.lt(5);
snow = snowProb.lt(5);
scl = image.select('SCL');
shadow = scl.eq(3); #// 3 = cloud shadow
cirrus = scl.eq(10); #// 10 = cirrus
#// Cloud probability less than 5% or cloud shadow classification
mask = (cloud.And(snow)).And(cirrus.neq(1)).And(shadow.neq(1));
return image.updateMask(mask).select("B.*").copyProperties(image, ["system:time_start"])
#// Exercise
#// Apply the above cloud masking function to SR image
#// Add the masked image to the map
maskedImageSR=ee.Image(maskCloudAndShadowsSR(imageSR))
Map.addLayer(imageSR,rgbVis,'MASKED')
#// Hint: After adding the masked image to the map, turn-off
#// the original image layer to see the result of the masking function
# -
# # REDUCERS
# +
geometry = ee.Geometry.Point([77.60412933051538, 12.952912912328241])
s2 = ee.ImageCollection("COPERNICUS/S2");
#// Computing stats on a list
myList = ee.List.sequence(1, 10);
#print(myList)
#// Use a reducer to compute min and max in the list
mean = myList.reduce(ee.Reducer.mean());
print(mean.min());
'''#// Apply a reducer on a image collection
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2020-01-01'))\
.filter(ee.Filter.bounds(geometry))\
.select('B.*')
print(filtered.size())
collMean = filtered.reduce(ee.Reducer.mean());
print('Reducer on Collection', collMean);
image = ee.Image(filtered.first())
#// If we want to compute min and max for each band, use reduceRegion instead
stats = image.reduceRegion({
"reducer": ee.Reducer.mean(),
"geometry": image.geometry(),
"scale": 100,
"maxPixels": 1e10
})
print(stats);
#// Result of reduceRegion is a dictionary.
#// We can extract the values using .get() function
print('Average value in B4', stats.get('B4'))'''
# +
geometry = ee.Geometry.Point([77.60412933051538, 12.952912912328241])
s2 = ee.ImageCollection("COPERNICUS/S2")
# Computing stats on a list
myList = ee.List.sequence(1, 10)
print(myList.getInfo())
# Use a reducer to compute min and max in the list
mean = myList.reduce(ee.Reducer.mean())
print(mean.getInfo())
# Apply a reducer on a image collection
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2020-01-01')) \
.filter(ee.Filter.bounds(geometry)) \
.select('B.*')
print(filtered.size().getInfo())
collMean = filtered.reduce(ee.Reducer.mean())
print('Reducer on Collection', collMean.getInfo())
image = ee.Image(filtered.first())
# If we want to compute min and max for each band, use reduceRegion instead
stats = image.reduceRegion(
reducer= ee.Reducer.mean(),
geometry= image.geometry(),
scale= 100,
maxPixels= 1e10
)
#print((stats.get('B4')).getInfo())
# Result of reduceRegion is a dictionary.
# We can extract the values using .get() function
print('Average value in B4', stats.get('B4').getInfo())
# -
Map=geemap.Map()
Map
roi=Map.draw_last_feature
print(composite.geometry().getInfo())
# +
s2= ee.ImageCollection('COPERNICUS/S2')
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2020-01-01'))\
.filter(ee.Filter.bounds(roi.geometry()))
def addNDVI(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename('ndvi');
ndwi = image.normalizedDifference(['B3', 'B8']).rename('ndwi');
return image.addBands(ndvi).addBands(ndwi);
withNdvi = filtered.map(addNDVI);
composite = withNdvi.median()
ndviComposite = composite.select('ndvi').clip(roi)
palette = [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718',
'74A901', '66A000', '529400', '3E8601', '207401', '056201',
'004C00', '023B01', '012E01', '011D01', '011301'];
ndviVis = {"min":0, "max":0.5, "palette": palette }
Map.addLayer(ndviComposite, ndviVis, 'ndwi')
stats =composite.reduceRegion(
reducer= ee.Reducer.mean(),
geometry= roi.geometry(),
scale= 100,
maxPixels= 1e10
)
print('Average value in NDVI', stats.get('ndwi').getInfo())
# -
# # roi=Map.draw_last_feature
geometry=roi.geometry()
# +
#geometry = ee.Geometry.Polygon([[
## [82.60642647743225, 27.16350437805251],
## [82.60984897613525, 27.1618529901377],
# [82.61088967323303, 27.163695288375266],
# [82.60757446289062, 27.16517483230927]
#]]);
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
image = ee.Image('COPERNICUS/S2/20190223T050811_20190223T051829_T44RPR')
Map.addLayer(image, rgbVis, 'Image')
Map.addLayer(geometry, {"color": 'red'}, 'Farm')
Map.centerObject(geometry)
ndwi = image.normalizedDifference(['B3', 'B8']).rename('ndwi');
image=image.addBands(ndwi)
#// Exercise
#// Compute the average NDVI for the farm from the given image
#// Hint: Use the reduceRegion() function
stats = image.reduceRegion(
reducer= ee.Reducer.mean(),
geometry= image.geometry(),
scale= 100,
maxPixels= 1e10
)
print('Average value in NDVI', stats.get('ndwi').getInfo())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR");
Map.centerObject(geometry)
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
filtered = s2.filter(ee.Filter.date('2017-01-01', '2018-01-01'))\
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.bounds(geometry))
#// Write a function for Cloud masking
def maskS2clouds(image):
qa = image.select('QA60')
cloudBitMask = 1 << 10;
cirrusBitMask = 1 << 11;
mask = qa.bitwiseAnd(cloudBitMask).eq(0).And(qa.bitwiseAnd(cirrusBitMask).eq(0))
return image.updateMask(mask).divide(10000).select("B.*").copyProperties(image, ["system:time_start"])
filtered = filtered.map(maskS2clouds)
#// Write a function that computes NDVI for an image and adds it as a band
def addNDVI(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename('ndvi');
return image.addBands(ndvi);
withNdvi = filtered.map(addNDVI);
# -
# # SUPERVISED CLASSIFICATION
Map=geemap.Map()
Map
# +
bangalore = ee.FeatureCollection("users/ujavalgandhi/public/bangalore_boundary")
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
#// The following collections were created using the #
#// Drawing Tools in the code editor
urban = ee.FeatureCollection("users/ujavalgandhi/e2e/urban_gcps")
bare = ee.FeatureCollection("users/ujavalgandhi/e2e/bare_gcps")
water = ee.FeatureCollection("users/ujavalgandhi/e2e/water_gcps")
vegetation = ee.FeatureCollection("users/ujavalgandhi/e2e/vegetation_gcps")
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2019-12-31'))\
.filter(ee.Filter.bounds(bangalore))\
.select('B.*')
composite = filtered.median().clip(bangalore)
#// Display the input composite.
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
Map.addLayer(composite, rgbVis, 'image');
gcps = urban.merge(bare).merge(water).merge(vegetation)
#// Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= gcps,
properties= ['landcover'],
scale=10
);
#// Train a classifier.
classifier = ee.Classifier.smileRandomForest(50).train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
);
#// // Classify the image.
classified = composite.classify(classifier);
Map.addLayer(classified, {"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']}, '2019');
#// Display the GCPs
#// We use the style() function to style the GCPs
palette = ee.List(['gray','brown','blue','green'])
landcover = ee.List([0, 1, 2, 3])
def fun1(lc):
color = palette.get(landcover.indexOf(lc));
markerStyle = { "color": 'white', "pointShape": 'diamond', "pointSize": 4, "width": 1, "fillColor": color}
def fun2(point):
return point.set('style', markerStyle)
return gcps.filter(ee.Filter.eq('landcover', lc)).map(fun2)
gcpsStyled = ee.FeatureCollection(landcover.map(fun1)).flatten();
Map.addLayer(gcpsStyled.style(styleProperty="style"), {}, 'GCPs')
Map.centerObject(gcpsStyled)
# -
Map=geemap.Map()
Map
s2= ee.ImageCollection("COPERNICUS/S2_SR")
urbanAreas = ee.FeatureCollection("users/ujavalgandhi/e2e/ne_10m_urban_areas")
city = urbanAreas.filter(ee.Filter.eq('system:index', '00000000000000002bf8'))
geometry = city.geometry()
Map.centerObject(geometry)
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2019-12-31'))\
.filter(ee.Filter.bounds(geometry))\
.select('B.*')
# +
composite = filtered.median().clip(geometry)
#// Display the input composite.
rgbVis = {"min": 0.0, "max": 3000, "bands": ['B4', 'B3', 'B2']};
Map.addLayer(composite, rgbVis, 'image');
# -
F=Map.draw_features
def func1(geom,style):
return ee.Feature(geom.geometry(),{"landmark":style})
#FU=ee.FeatureCollection([func1(t,0) for t in F])
# +
#FB=ee.FeatureCollection([func1(t,1) for t in F])
# +
#FW=ee.FeatureCollection([func1(t,2) for t in F])
# +
#FV=ee.FeatureCollection([func1(t,3) for t in F])
# -
gcps = FU.merge(FB).merge(FW).merge(FV)
# +
training = composite.sampleRegions(
collection= gcps,
properties= ['landmark'],
scale= 200,
tileScale= 16
);
print(training)
#// // Train a classifier.
classifier = ee.Classifier.smileRandomForest(50).train(
features= training,
classProperty= 'landmark',
inputProperties= composite.bandNames()
);
#// // // Classify the image.
classified = composite.classify(classifier);
Map.addLayer(classified, {"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']}, '2019');
# -
Map=geemap.Map()
Map
# +
bangalore = ee.FeatureCollection("users/ujavalgandhi/public/bangalore_boundary")
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
#// The following collections were created using the #
#// Drawing Tools in the code editor
urban = ee.FeatureCollection("users/ujavalgandhi/e2e/urban_gcps")
bare = ee.FeatureCollection("users/ujavalgandhi/e2e/bare_gcps")
water = ee.FeatureCollection("users/ujavalgandhi/e2e/water_gcps")
vegetation = ee.FeatureCollection("users/ujavalgandhi/e2e/vegetation_gcps")
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2019-12-31'))\
.filter(ee.Filter.bounds(bangalore))\
.select('B.*')
composite = filtered.median().clip(bangalore)
#// Display the input composite.
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
Map.addLayer(composite, rgbVis, 'image');
gcps = urban.merge(bare).merge(water).merge(vegetation)
#// Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= gcps,
properties= ['landcover'],
scale=10
);
#// Train a classifier.
classifier = ee.Classifier.smileRandomForest(50).train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
);
#// // Classify the image.
classified = composite.classify(classifier);
Map.addLayer(classified, {"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']}, '2019');
#// Display the GCPs
#// We use the style() function to style the GCPs
palette = ee.List(['gray','brown','blue','green'])
landcover = ee.List([0, 1, 2, 3])
def fun1(lc):
color = palette.get(landcover.indexOf(lc));
markerStyle = { "color": 'white', "pointShape": 'diamond', "pointSize": 4, "width": 1, "fillColor": color}
def fun2(point):
return point.set('style', markerStyle)
return gcps.filter(ee.Filter.eq('landcover', lc)).map(fun2)
gcpsStyled = ee.FeatureCollection(landcover.map(fun1)).flatten();
Map.addLayer(gcpsStyled.style(styleProperty="style"), {}, 'GCPs')
Map.centerObject(gcpsStyled)
# -
Map=geemap.Map()
Map
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR");
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7");
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps");
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
rgbVis = {
"min": 0.0,
"max": 3000,
"bands": ['B4', 'B3', 'B2'],
};
filtered = s2.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2019-12-31'))\
.filter(ee.Filter.bounds(boundary))\
.select('B.*')
composite = filtered.median().clip(boundary)
#// Display the input composite.
Map.addLayer(composite, rgbVis, 'image');
#// Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= gcp,
properties= ['landcover'],
scale= 11,
);
#// Train a classifier.
classifier = ee.Classifier.smileRandomForest(50).train(
features= training,
classProperty= 'landcover',
inputProperties=composite.bandNames()
);
#// Classify the image.
classified = composite.classify(classifier);
Map.addLayer(classified, {"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']}, '2019');
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
# The function also divides ths image by 10000 to ensure
# the pixels values are between 0 and 1
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
# We divide the elevation and slope with maximum values in the region
# to ensure the values are between 0 and 1
# A more robust technique for image normalization is provided in the course supplement
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= trainingGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
)
#print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions(
collection= validationGcp,
properties= ['landcover'],
scale=10,
tileScale= 16
)
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix.getInfo())
print('Test Accuracy', testConfusionMatrix.accuracy().getInfo())
# -
Map=geemao.Map()
Map
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.select('B.*')
composite = filtered.median().clip(boundary)
# Display the input composite.
Map.addLayer(composite, rgbVis, 'image')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
# Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= trainingGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train(
features= training,
classProperty='landcover',
inputProperties=composite.bandNames()
)
# Classify the image.
classified = composite.classify(classifier)
Map.addLayer(classified, {'min': 0, 'max': 3, 'palette': ['gray', 'brown', 'blue', 'green']}, '2019')
#**************************************************************************
# Accuracy Assessment
#**************************************************************************
# Use classification map to assess accuracy using the validation fraction
# of the overall training set created above.
test = classified.sampleRegions(
collection= validationGcp,
properties= ['landcover'],
tileScale= 16,
scale= 10,
)
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
# Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix.getInfo())
print('Test Accuracy', testConfusionMatrix.accuracy().getInfo())
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= trainingGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
)
#print(training)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
#print(classifier.explain())
test = composite.sampleRegions(
collection= validationGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
)
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_dmn(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_dmn)
print(ee.List(accuracies.getInfo()))
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_fdj(numTrees):
def func2:
classifier = ee.Classifier.smileRandomForest(
numberOfTrees= numTrees,
bagFraction= bagFraction
) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,'numberOfTrees': numTrees,'bagFraction': bagFraction})
return bagFractionList.map(function(bagFraction)
classifier = ee.Classifier.smileRandomForest(
numberOfTrees= numTrees,
bagFraction= bagFraction
) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,
'numberOfTrees': numTrees,
'bagFraction': bagFraction})
})
accuracies = numTreesList.map(func_fdj).flatten()
resultFc = ee.FeatureCollection(accuracies)
print(accuracies.getInfo())
# -
print((accuracies.getInfo()))
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7")
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps")
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2")
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
}
# Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB')
snowProb = image.select('MSK_SNWPRB')
cloud = cloudProb.lt(10)
scl = image.select('SCL')
shadow = scl.eq(3); # 3 = cloud shadow
cirrus = scl.eq(10); # 10 = cirrus
# Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1))
return image.updateMask(mask).divide(10000)
filtered = s2 \
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30)) \
.filter(ee.Filter.date('2019-01-01', '2019-12-31')) \
.filter(ee.Filter.bounds(boundary)) \
.map(maskCloudAndShadowsSR) \
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi'])
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi'])
mndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi'])
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'), #swir1
'Y': image.select('B4'), #red
'A': image.select('B8'), # nir
'B': image.select('B2'), # blue
}).rename('bsi')
return image.addBands(ndvi).addBands(ndbi).addBands(mndwi).addBands(bsi)
composite = addIndices(composite)
elev = alos.select('AVE_DSM').divide(2000).rename('elev')
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope')
composite = composite.addBands(elev).addBands(slope)
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2}
Map.addLayer(composite, visParams, 'RGB')
# Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
# This being a simpler classification, we take 60% points
# for validation. Normal recommended ratio is
# 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6))
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6))
Map.addLayer(validationGcp)
# Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= trainingGcp,
properties= ['landcover'],
scale= 10,
tileScale=16
)
# Train a classifier.
classifier = ee.Classifier.smileRandomForest(50) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
#**************************************************************************
# Hyperparameter Tuning
#**************************************************************************
# Run .explain() to see what the classifer looks like
print(classifier.explain())
test = composite.sampleRegions(
collection= validationGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
)
# Tune the numberOfTrees parameter.
numTreesList = ee.List.sequence(10, 150, 10)
def func_pov(numTrees):
classifier = ee.Classifier.smileRandomForest(numTrees) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
return test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
accuracies = numTreesList.map(func_pov)
print(accuracies.getInfo())
# Tuning Multiple Parameters
# We can tune many parameters together using
# nested map() functions
# Let's tune 2 parameters
# numTrees and bagFraction
numTreesList = ee.List.sequence(10, 150, 10)
bagFractionList = ee.List.sequence(0.1, 0.9, 0.1)
def func_xry(numTrees):
def func1(bagFraction):
classifier = ee.Classifier.smileRandomForest(
numberOfTrees= numTrees,
bagFraction= bagFraction
) \
.train(
features= training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
)
# Here we are classifying a table instead of an image
# Classifiers work on both images and tables
accuracy = test \
.classify(classifier) \
.errorMatrix('landcover', 'classification') \
.accuracy()
return ee.Feature({}, {'accuracy': accuracy,'numberOfTrees': numTrees,'bagFraction': bagFraction})
return bagFractionList.map(func1)
accuracies = numTreesList.map(func_xry).flatten()
resultFc = ee.FeatureCollection(accuracies)
# -
Map=geemap.Map()
Map
# +
s2 = ee.ImageCollection("COPERNICUS/S2_SR");
basin = ee.FeatureCollection("WWF/HydroSHEDS/v1/Basins/hybas_7");
gcp = ee.FeatureCollection("users/ujavalgandhi/e2e/arkavathy_gcps");
alos = ee.Image("JAXA/ALOS/AW3D30/V2_2");
arkavathy = basin.filter(ee.Filter.eq('HYBAS_ID', 4071139640))
boundary = arkavathy.geometry()
s2 = ee.ImageCollection("COPERNICUS/S2_SR")
rgbVis = {
'min': 0.0,
'max': 3000,
'bands': ['B4', 'B3', 'B2'],
};
#// Function to remove cloud and snow pixels from Sentinel-2 SR image
def maskCloudAndShadowsSR(image):
cloudProb = image.select('MSK_CLDPRB');
snowProb = image.select('MSK_SNWPRB');
cloud = cloudProb.lt(10);
scl = image.select('SCL');
shadow = scl.eq(3); #// 3 = cloud shadow
cirrus = scl.eq(10); #// 10 = cirrus
#// Cloud probability less than 10% or cloud shadow classification
mask = cloud.And(cirrus.neq(1)).And(shadow.neq(1));
return image.updateMask(mask).divide(10000);
filtered = s2\
.filter(ee.Filter.lt('CLOUDY_PIXEL_PERCENTAGE', 30))\
.filter(ee.Filter.date('2019-01-01', '2019-12-31'))\
.filter(ee.Filter.bounds(boundary))\
.map(maskCloudAndShadowsSR)\
.select('B.*')
composite = filtered.median().clip(boundary)
def addIndices(image):
ndvi = image.normalizedDifference(['B8', 'B4']).rename(['ndvi']);
ndbi = image.normalizedDifference(['B11', 'B8']).rename(['ndbi']);
ndwi = image.normalizedDifference(['B3', 'B11']).rename(['mndwi']);
bsi = image.expression(
'(( X + Y ) - (A + B)) /(( X + Y ) + (A + B)) ', {
'X': image.select('B11'),# //swir1
'Y': image.select('B4'), #//red
'A': image.select('B8'), #// nir
'B': image.select('B2'), #// blue
}).rename('bsi');
return image.addBands(ndvi).addBands(ndbi).addBands(ndwi).addBands(bsi)
composite = addIndices(composite);
elev = alos.select('AVE_DSM').divide(2000).rename('elev');
slope = ee.Terrain.slope(alos.select('AVE_DSM')).divide(30).rename('slope');
composite = composite.addBands(elev).addBands(slope);
visParams = {'bands': ['B4', 'B3', 'B2'], 'min': 0, 'max': 0.3, 'gamma': 1.2};
Map.addLayer(composite, visParams, 'RGB');
#// Add a random column and split the GCPs into training and validation set
gcp = gcp.randomColumn()
#// This being a simpler classification, we take 60% points
#// for validation. Normal recommended ratio is
#// 70% training, 30% validation
trainingGcp = gcp.filter(ee.Filter.lt('random', 0.6));
validationGcp = gcp.filter(ee.Filter.gte('random', 0.6));
Map.addLayer(validationGcp)
#// Overlay the point on the image to get training data.
training = composite.sampleRegions(
collection= trainingGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
);
#print(training)
#// Train a classifier.
classifier = ee.Classifier.smileRandomForest(10).train(
features=training,
classProperty= 'landcover',
inputProperties= composite.bandNames()
);
#// Classify the image.
classified = composite.classify(classifier);
Map.addLayer(classified, {"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']}, '2019');
#// Use classification map to assess accuracy using the validation fraction
#// of the overall training set created above.
test = classified.sampleRegions(
collection= validationGcp,
properties= ['landcover'],
scale= 10,
tileScale= 16
);
testConfusionMatrix = test.errorMatrix('landcover', 'classification')
#// Printing of confusion matrix may time out. Alternatively, you can export it as CSV
print('Confusion Matrix', testConfusionMatrix.getInfo());
print('Test Accuracy', testConfusionMatrix.accuracy().getInfo());
#//**************************************************************************
#// Exporting Results
#//**************************************************************************
#// Create a Feature with null geometry and the value we want to export.
#// Use .array() to convert Confusion Matrix to an Array so it can be
#// exported in a CSV file
#fc = ee.FeatureCollection([
# ee.Feature(null, {
# 'accuracy': testConfusionMatrix.accuracy(),
# 'matrix': testConfusionMatrix.array()
# })
# ])
#print(fc)
# +
classified = ee.Image("users/ujavalgandhi/e2e/bangalore_classified");
bangalore = ee.FeatureCollection("users/ujavalgandhi/public/bangalore_boundary");
admin2 = ee.FeatureCollection("FAO/GAUL_SIMPLIFIED_500m/2015/level2");
Map.addLayer(bangalore, {"color": 'blue'}, 'Bangalore City')
Map.addLayer(classified,
{"min": 0, "max": 3, "palette": ['gray', 'brown', 'blue', 'green']},
'Classified Image 2019');
#// Calling .geometry() on a feature collection gives the
#// dissolved geometry of all features in the collection
#// .area() function calculates the area in square meters
cityArea = bangalore.geometry().area()
#// We can cast the result to a ee.Number() and calculate the
#// area in square kilometers
cityAreaSqKm = ee.Number(cityArea).divide(1e6).round()
print(cityAreaSqKm.getInfo())
#// Area Calculation for Images
vegetation = classified.eq(3)
#// If the image contains values 0 or 1, we can calculate the
#// total area using reduceRegion() function
#// The result of .eq() operation is a binary image with pixels
#// values of 1 where the condition matched and 0 where it didn't
Map.addLayer(vegetation, {"min":0, "max":1, "palette": ['white', 'green']}, 'Green Cover')
#// Since our image has only 0 and 1 pixel values, the vegetation
#// pixels will have values equal to their area
areaImage = vegetation.multiply(ee.Image.pixelArea())
#// Now that each pixel for vegetation class in the image has the value
#// equal to its area, we can sum up all the values in the region
#// to get the total green cover.
area = areaImage.reduceRegion(
reducer= ee.Reducer.sum(),
geometry= bangalore.geometry(),
scale= 10,
maxPixels= 1e10
)
#// The result of the reduceRegion() function is a dictionary with the key
#// being the band name. We can extract the area number and convert it to
#// square kilometers
vegetationAreaSqKm = ee.Number(area.get('classification')).divide(1e6).round()
print(vegetationAreaSqKm.getInfo())
print("Green Area %")
print(vegetationAreaSqKm.getInfo()/cityAreaSqKm.getInfo()*100)
# -
|
EXAMPLES/notebooks/Intermediate_important.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: myenv
# ---
import numpy as np
from tqdm import tqdm
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def bce(y_true, y_pred):
def safe_log(x):
return np.log(x) if x != 0 else 0
bce_loss = 0
for cur_y_true, cur_y_pred in zip(y_true, y_pred):
bce_loss += cur_y_true*safe_log(cur_y_pred) + (1 - cur_y_true)*safe_log(1 - cur_y_pred)
return -bce_loss / len(y_true)
bce([1, 0], [0.3, 0.7])
class LogisticRegression:
def __init__(self, lr=0.01, epochs=1000):
self.epochs = epochs
self.lr = lr
self.W = 0
self.loss = []
def initialize(self, n_features):
self.W = np.random.normal(0, 1, size=(n_features, 1))
self.W = np.squeeze(self.W, axis=1)
def sigmoid(self, z):
return 1 / (1 + np.exp(-z))
def bce(self, y_true, y_pred):
def safe_log(x):
return np.log(x) if x != 0 else 0
bce_loss = 0
for cur_y_true, cur_y_pred in zip(y_true, y_pred):
bce_loss += cur_y_true*safe_log(cur_y_pred) + (1 - cur_y_true)*safe_log(1 - cur_y_pred)
return -bce_loss / len(y_true)
def gradient(self, X, y, n_samples):
y_pred = self.sigmoid(np.dot(X, self.W))
d_w = np.dot(X.T, y_pred - y) / n_samples
return d_w
def fit(self, X, y):
losses = []
# Load sample and features
n_samples, n_features = X.shape
# Init weights
self.initialize(n_features)
# Calculate gradient descent per epoch
for _ in tqdm(range(self.epochs)):
y_pred = self.sigmoid(np.dot(X, self.W))
d_w = self.gradient(X, y, n_samples)
self.W -= self.lr * d_w
loss = self.bce(y, y_pred)
losses.append(loss)
print('Loss: ', loss)
def predict(self, X):
return [1 if i > 0.5 else 0 for i in self.sigmoid(np.dot(X, self.W))]
import pandas as pd
import sklearn.datasets
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import classification_report
cancer = sklearn.datasets.load_breast_cancer()
data = pd.DataFrame(cancer.data, columns = cancer.feature_names)
data["label"] = cancer.target
data.head()
# +
X = data.iloc[:,:-1] #all rows, all columns except the last
y = data.iloc[:,-1] # all rows, only the last column
X = MinMaxScaler().fit_transform(X)
X_train, X_test, Y_train, Y_test = train_test_split(X,y, test_size = 0.30, random_state = 1)
# -
model = LogisticRegression(lr=0.1, epochs=20000)
model.fit(X_train, Y_train)
Y_pred = model.predict(X_test)
print(classification_report(Y_test, Y_pred))
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(random_state=0).fit(X_train, Y_train)
Y_pred_sk = clf.predict(X_test)
print(classification_report(Y_test, Y_pred_sk))
|
regression/logistic_regresion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''pytorch'': conda)'
# name: python37664bitpytorchconda0cdad03962454fdfb22b6d3ea1ad8fae
# ---
# http://preview.d2l.ai/d2l-en/PR-1089/chapter_appendix-mathematics-for-deep-learning/information-theory.html
# +
import torch
from torch.nn import NLLLoss
def nansum(x):
# Define nansum, as pytorch doesn't offer it inbuilt.
return x[~torch.isnan(x)].sum()
def self_information(p):
return -torch.log2(torch.tensor(p)).item()
self_information(1 / 64)
# +
def entropy(p):
entropy = - p * torch.log2(p)
# Operator nansum will sum up the non-nan number
out = nansum(entropy)
return out
entropy(torch.tensor([0.1, 0.5, 0.1, 0.3]))
# +
def joint_entropy(p_xy):
joint_ent = -p_xy * torch.log2(p_xy)
# nansum will sum up the non-nan number
out = nansum(joint_ent)
return out
joint_entropy(torch.tensor([[0.1, 0.5], [0.1, 0.3]]))
# +
def conditional_entropy(p_xy, p_x):
p_y_given_x = p_xy/p_x
cond_ent = -p_xy * torch.log2(p_y_given_x)
# nansum will sum up the non-nan number
out = nansum(cond_ent)
return out
conditional_entropy(torch.tensor([[0.1, 0.5], [0.2, 0.3]]),
torch.tensor([0.2, 0.8]))
# +
def mutual_information(p_xy, p_x, p_y):
p = p_xy / (p_x * p_y)
mutual = p_xy * torch.log2(p)
# Operator nansum will sum up the non-nan number
out = nansum(mutual)
return out
mutual_information(torch.tensor([[0.1, 0.5], [0.1, 0.3]]),
torch.tensor([0.2, 0.8]), torch.tensor([[0.75, 0.25]]))
# -
def kl_divergence(p, q):
kl = p * torch.log2(p / q)
out = nansum(kl)
return out.abs().item()
# +
torch.manual_seed(1)
tensor_len = 10000
p = torch.normal(0, 1, (tensor_len, ))
q1 = torch.normal(-1, 1, (tensor_len, ))
q2 = torch.normal(1, 1, (tensor_len, ))
p = torch.sort(p)[0]
q1 = torch.sort(q1)[0]
q2 = torch.sort(q2)[0]
# +
kl_pq1 = kl_divergence(p, q1)
kl_pq2 = kl_divergence(p, q2)
similar_percentage = abs(kl_pq1 - kl_pq2) / ((kl_pq1 + kl_pq2) / 2) * 100
kl_pq1, kl_pq2, similar_percentage
# +
kl_q2p = kl_divergence(q2, p)
differ_percentage = abs(kl_q2p - kl_pq2) / ((kl_q2p + kl_pq2) / 2) * 100
kl_q2p, differ_percentage
# -
def cross_entropy(y_hat, y):
ce = -torch.log(y_hat[range(len(y_hat)), y])
return ce.mean()
# +
labels = torch.tensor([0, 2])
preds = torch.tensor([[0.3, 0.6, 0.1], [0.2, 0.3, 0.5]])
cross_entropy(preds, labels)
# -
# Implementation of CrossEntropy loss in pytorch combines nn.LogSoftmax() and
# nn.NLLLoss()
nll_loss = NLLLoss()
loss = nll_loss(torch.log(preds), labels)
loss
|
Ch18_math-for-DL/information-theory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # SoftMax Distributions for Human-Robot Interaction
#
# This iPython Notebook is a living document that details the development and usage of SoftMax distributions -- a powerful tool to probabilistically decompose state spaces.
#
# **DISCLAIMER**: This document's current status is **in development**. That means code might be broken, text may have typos, and math might be missing/wrong. Read this as ongoing research notes until this disclaimer is removed.
#
# **NOTE**: While you can read this document as-is using nbviewer, the best way to read it is by downloading the `.ipynb` files and associated code from [the project's GitHub repository](https://github.com/COHRINT/cops_and_robots), then running `ipython notebook` locally.
#
# The notebook is publically available at http://nbviewer.ipython.org/github/COHRINT/cops_and_robots/blob/master/notebooks/softmax/Main.ipynb
#
# ## [Chapter 1 - Introduction to SoftMax Distributions](01_intro.ipynb)
# We look at the basics of why you'd want to use SoftMax distributions, how to make them, and what their general basic properties are.
#
# ## [Chapter 2 - Using Normals Instead of Weights](02_from_normals.ipynb)
# Instead of defining weights by hand, we show other ways to create SoftMax distributions: using vector normals.
#
# ## [Chapter 3 - Building SoftMax Distributions from Templates](03_from_templates.ipynb)
# We can abstract the SoftMax distribution creation even further using templates.
#
# ## [Chapter 4 - Subclassing and Superclassing: Multimodal SoftMax](04_mms.ipynb)
# SoftMax distributions can be overly simplistic for some models, so we investigate ways to modify SoftMax distributions for general, non-symmetric cases.
#
# ## [Chapter 5 - Learning SoftMax Distributions from Data](05_from_data.ipynb)
# TODO
#
# ## [Chapter 6 - Shaping with Class Boundary Priors](06_priors.ipynb)
# TODO
#
# ## [Chapter 7 - Using Symmetry](07_symmetry.ipynb)
# TODO
#
# ## [Chapter 8 - N-Dimensional SoftMax](08_n_dimensions.ipynb)
# TODO
# +
from IPython.core.display import HTML
# Borrowed style from Probabilistic Programming and Bayesian Methods for Hackers
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
|
resources/notebooks/softmax/Main.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pickle
import gzip
import utils
import loompy
import pandas as pd
import numpy as np
import scipy.sparse as sp
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# %%time
data = pd.read_table("data/GSE63472_P14Retina_merged_digital_expression.txt.gz", index_col=0)
data.head()
# %%time
cluster_ids = pd.read_table("data/retina_clusteridentities.txt", header=None, index_col=0, squeeze=True)
cluster_ids.head()
# Reorder
cluster_ids = cluster_ids[data.columns.values]
data.shape
# Only use cells where metadata is available
ind = data.columns.isin(cluster_ids.index)
data = data.loc[:, ind]
data.shape, cluster_ids.shape
mask = ~cluster_ids.isna()
data = data.loc[:, mask.values]
cluster_ids = cluster_ids[mask]
assert not cluster_ids.isna().any(), "Did not properly remove cells with NaN label"
data.shape, cluster_ids.shape
# %%time
counts = sp.csr_matrix(data.values)
counts
# %%time
cpm_counts = utils.calculate_cpm(counts, axis=0)
log_counts = utils.log_normalize(cpm_counts)
# +
cell_types = cluster_ids.astype(object)
cell_types.loc[cell_types == 1] = "Horizontal cells"
cell_types.loc[cell_types == 2] = "Retinal ganglion cells"
cell_types.loc[cell_types.isin(range(3, 24))] = "Amacrine cells"
cell_types.loc[cell_types == 24] = "Rods"
cell_types.loc[cell_types == 25] = "Cones"
cell_types.loc[cell_types.isin(range(26, 34))] = "Bipolar cells"
cell_types.loc[cell_types == 34] = "Muller glia"
cell_types.loc[cell_types == 35] = "Astrocytes"
cell_types.loc[cell_types == 36] = "Fibroblasts"
cell_types.loc[cell_types == 37] = "Vascular endothelium"
cell_types.loc[cell_types == 38] = "Pericytes"
cell_types.loc[cell_types == 39] = "Microglia"
cell_types.value_counts()
# -
# ## Preprocess data set
# ### Dropout based feature selection
# %time gene_mask = utils.select_genes(counts.T, n=3000, threshold=0)
x = log_counts.T[:, gene_mask].toarray()
x.shape
# ### Standardize data
x -= x.mean(axis=0)
x /= x.std(axis=0)
# ### PCA preprocessing
# %%time
U, S, V = np.linalg.svd(x, full_matrices=False)
U[:, np.sum(V, axis=1) < 0] *= -1
x_reduced = np.dot(U, np.diag(S))
x_reduced = x_reduced[:, np.argsort(S)[::-1]][:, :50]
x_reduced.shape
cell_types.shape
# ## Write data
data_dict = {"pca_50": x_reduced,
"CellType1": cell_types.values.astype(str),
"CellType2": cluster_ids.values.astype(str)}
# %%time
with gzip.open("data/macosko_2015.pkl.gz", "wb") as f:
pickle.dump(data_dict, f)
|
examples/prepare_macosko_2015.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="EU3xHI_MKl7v"
#if you use colab
from google.colab import drive
drive.mount('/content/drive')
# + id="5rJ-SgSuKvbu"
#change the module_dir your file located
module_dir = ("/content/drive/My Drive/AI project/Final work/")
import sys
sys.path.append(module_dir)
# + id="spqzr75Uj5b_"
# #!pip3 install tensorflow
# #!pip3 install pandas
# #!pip3 install numpy
# #!pip3 install opencv-python
# #!pip3 install matplotlib
# + id="K6P-KEl3UkTT"
from classifier import styler, styler_url,random_img
# + id="iB8RlAvwK1o2"
option = 'url input' #@param ["upload","url input"] {type:"string"}
if option == 'url input':
url = input('pls input url here\n')
result = styler_url(url, module_dir)
elif option == 'upload':
from google.colab import files
print("pls upload image with jpg, jpeg, png formate\n")
img = files.upload()
result = styler(img, module_dir)
print("style: ", result)
img_path = random_img(result, module_dir)
import cv2
import matplotlib.pyplot as plt
img = cv2.imread(img_path)
plt.imshow(cv2.cvtColor(img,cv2.COLOR_BGR2RGB))
|
Final Work/UserInterphase.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] iooxa={"id": {"block": "h6m9YbHtjm7lMhF3zATA", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# # 3D Magnetic Inversion of Raglan Data with Lp-norm
#
# In this notebook, we inverted the magnetic data acquired at [Raglan deposit](https://en.wikipedia.org/wiki/Raglan_Mine) located in Northern Quebec, Canada to obtain a three-dimensional susceptibility model of the subsurface. Magnetic module of an open-source geophysics software, [SimPEG ](https://www.simpeg.xyz) was used for this inversion.
#
# About 20 years ago, this data was inverted using the [MAG3D code](https://mag3d.readthedocs.io/en/latest/content/overview.html) developed by [UBC-GIF](https://gif.eos.ubc.ca/about) group, and a 3D susceptibility model was obtained. This was sort of the first time that field magnetic data was inverted in 3D, and made a significant impact on locating drilling location for a mineral exploration.
#
# In the [previous notebook](./1-magnetic-inversion-raglan-reproduce), we inverted these magnetic data using an L2-norm inversion. To promote the compactness of the target body, we used the Lp-norm inversion approach (Founier and Oldenburg, 2019).
# + [markdown] iooxa={"id": {"block": "xSOil0mBA9bWRmvStN41", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# <center>
# <img src="./figures/mag3disco.gif", class="center"></img>
# </center>
# + [markdown] iooxa={"id": {"block": "rh4isubZDWAIl9CGuTHm", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Import modules
#
#
#
# + iooxa={"id": {"block": "Dv1sFnCMjAak6eeNfTDV", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# %matplotlib inline
import matplotlib
import os
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import tarfile
from discretize import TensorMesh
from discretize.utils import mesh_builder_xyz
from SimPEG.potential_fields import magnetics
from SimPEG import dask
from SimPEG.utils import plot2Ddata, surface2ind_topo
from SimPEG import (
maps,
data,
inverse_problem,
data_misfit,
regularization,
optimization,
directives,
inversion,
utils,
)
import pandas as pd
from ipywidgets import widgets, interact
# + [markdown] iooxa={"id": {"block": "Yv24shLad5ZjXMm0aQlU", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Load Data and Plot
# + iooxa={"id": {"block": "korKsPB9v4Yugw9fFSZX", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
def read_ubc_magnetic_data(data_filename):
with open(data_filename, 'r') as f:
lines = f.readlines()
tmp = np.array(lines[0].split()[:3]).astype(float)
n_data = int(float(lines[2].split()[0]))
meta_data = {}
meta_data['inclination'] = float(tmp[0])
meta_data['declination'] = float(tmp[1])
meta_data['b0'] = float(tmp[2])
meta_data['n_data'] = n_data
data = np.zeros((n_data, 5), order='F')
for i_data in range(n_data):
data[i_data,:] = np.array(lines[3+i_data].split()).astype(float)
df = pd.DataFrame(data=data, columns=['x', 'y', 'z', 'data', 'data_error'])
return df, meta_data
# + iooxa={"id": {"block": "rMcHA8kIQWWj9HIGWNK9", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
data_filename = "./data/Raglan_1997/obs.mag"
df, meta_data = read_ubc_magnetic_data(data_filename)
# + iooxa={"id": {"block": "XOiiqBjRIICg98V2ZO83", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "qrnOaXVwP2Tw1rDgeVUB", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
meta_data
# + iooxa={"id": {"block": "6kxgIwhyE2xrH13C8k4d", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "ph0hlnhwbZpEtCLXojCI", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
df.head(3)
# + iooxa={"id": {"block": "vVCY1DmVrHu4P4nKOwF8", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "g2j93s9wm5GwIO2JeCQs", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# Down sample the data
matplotlib.rcParams['font.size'] = 14
nskip = 2
receiver_locations = df[['x', 'y', 'z']].values[::nskip,:]
xyz_topo = np.c_[receiver_locations[:,:2], np.zeros(receiver_locations.shape[0])]
dobs = df['data'].values[::nskip]
# Plot
fig = plt.figure(figsize=(12, 10))
vmin, vmax = np.percentile(dobs, 0.5), np.percentile(dobs, 99.5)
tmp = np.clip(dobs, vmin, vmax)
ax1 = fig.add_axes([0.1, 0.1, 0.75, 0.85])
plot2Ddata(
receiver_locations,
tmp,
ax=ax1,
ncontour=30,
clim=(vmin-5, vmax+5),
contourOpts={"cmap": "Spectral_r"},
)
ax1.set_title("TMI Anomaly")
ax1.set_xlabel("x (m)")
ax1.set_ylabel("y (m)")
ax2 = fig.add_axes([0.9, 0.25, 0.05, 0.5])
norm = mpl.colors.Normalize(vmin=vmin, vmax=vmax)
cbar = mpl.colorbar.ColorbarBase(
ax2, norm=norm, orientation="vertical", cmap=mpl.cm.Spectral_r
)
cbar.set_label("$nT$", rotation=270, labelpad=15, size=12)
plt.show()
# + [markdown] iooxa={"id": {"block": "u7sQlIJtgfFAj8oXcClo", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Assign Uncertainty
#
# Inversion with SimPEG requires that we define data error i.e., standard deviation on the observed data.
# This represents our estimate of the noise in our data.
# For this magnetic inversion, 2% relative error and a noise floor of 2 nT are assigned.
# + iooxa={"id": {"block": "nj91uOIFTa8McscCJuNI", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
standard_deviation = 0.02 * abs(dobs) + 2
# + [markdown] iooxa={"id": {"block": "Bfu9nCkVK0R3GFTBfD5H", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Defining the Survey
#
# Here, we define a survey object that will be used for the simulation.
# The user needs an (N, 3) array to define
# the xyz locations of the observation locations and the list of field components
# which are to be modeled and the properties of the Earth's field.
# + iooxa={"id": {"block": "zP3YD0OkTc9ENhPjbTWz", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# Define the component(s) of the field we are inverting as a list. Here we will
# Invert total magnetic intensity data.
components = ["tmi"]
# Use the observation locations and components to define the receivers. To
# simulate data, the receivers must be defined as a list.
receiver_list = magnetics.receivers.Point(receiver_locations, components=components)
receiver_list = [receiver_list]
# Define the inducing field H0 = (intensity [nT], inclination [deg], declination [deg])
inclination = meta_data['inclination']
declination = meta_data['declination']
strength = meta_data['b0']
inducing_field = (strength, inclination, declination)
source_field = magnetics.sources.SourceField(
receiver_list=receiver_list, parameters=inducing_field
)
# Define the survey
survey = magnetics.survey.Survey(source_field)
# + [markdown] iooxa={"id": {"block": "NFI3ARAHrmk0TLnNxdgp", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Defining the Data
#
# Here is where we define the data that is inverted. The data is defined by
# the survey, the observation values and the standard deviations.
#
#
#
# + iooxa={"id": {"block": "17kdgUcgQfIkaHteoGly", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
data_object = data.Data(survey, dobs=dobs, standard_deviation=standard_deviation)
# + [markdown] iooxa={"id": {"block": "uGWOcn1YVf3AmFR4Ny87", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Defining a Tensor Mesh
#
# Here, we create the tensor mesh that will be used to invert TMI data.
# If desired, we could define an OcTree mesh.
# + iooxa={"id": {"block": "Emx4JRxt5izoIIwU8ina", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
dx = 100
dy = 100
dz = 100
depth_core = 1000
padding_distance_x_left = 1000
padding_distance_x_right = 1000
padding_distance_y_left = 1000
padding_distance_y_right= 1000
padding_distance_z_lower = 1000
padding_distance_z_upper = 0
mesh = mesh_builder_xyz(
xyz=xyz_topo,
h=[dx, dy, dz],
depth_core=depth_core,
padding_distance=[
[padding_distance_x_left, padding_distance_x_right],
[padding_distance_y_left, padding_distance_y_right],
[padding_distance_z_lower, padding_distance_z_upper]
]
)
# + iooxa={"id": {"block": "3N6xgKtbGfC20FDVYFsp", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "lDXlJFe7PROuBdgMqRww", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
mesh.plotSlice(np.ones(mesh.nC)*np.nan, ax=ax, grid=True)
ax.plot(receiver_locations[:,0], receiver_locations[:,1], 'r.')
ax.set_xlabel("Easting (m)")
ax.set_ylabel("Northing (m)")
ax.set_aspect(1)
# + iooxa={"id": {"block": "PketRpNq9eDeCz2Hg8Sd", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "eIu7BaHX4EFP1qGDV5Fu", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
fig, ax = plt.subplots(1, 1, figsize=(10, 10))
mesh.plotSlice(np.ones(mesh.nC)*np.nan, ax=ax, grid=True, normal='Y')
ax.set_xlabel("Easting (m)")
ax.set_ylabel("Depth (m)")
ax.set_aspect(1)
# + [markdown] iooxa={"id": {"block": "pJTzXhrGHqthw9ntNRv2", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Starting/Reference Model and Mapping on Tensor Mesh
#
# Here, we would create starting and/or reference models for the inversion as
# well as the mapping from the model space to the active cells. Starting and
# reference models can be a constant background value or contain a-priori
# structures. Here, the background is 1e-4 SI.
#
#
#
# + iooxa={"id": {"block": "M57lXY0THBaap8W2wzSw", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# Define background susceptibility model in SI. Don't make this 0!
# Otherwise the gradient for the 1st iteration is zero and the inversion will
# not converge.
background_susceptibility = 1e-4
# Find the indecies of the active cells in forward model (ones below surface)
ind_active = surface2ind_topo(mesh, np.c_[receiver_locations[:,:2], np.zeros(survey.nD)])
# Define mapping from model to active cells
nC = int(ind_active.sum())
model_map = maps.IdentityMap(nP=nC) # model consists of a value for each cell
# Define starting model
starting_model = background_susceptibility * np.ones(nC)
reference_model = np.zeros(nC)
# + [markdown] iooxa={"id": {"block": "jvQIJqPdFKO9TlRzPJsE", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Define the Physics
#
# Here, we define the physics of the magnetics problem by using the simulation
# class.
#
#
#
# + iooxa={"id": {"block": "b8u2dHOg51oNDo5o4xvF", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# Define the problem. Define the cells below topography and the mapping
simulation = magnetics.simulation.Simulation3DIntegral(
survey=survey,
mesh=mesh,
modelType="susceptibility",
chiMap=model_map,
actInd=ind_active,
)
# + [markdown] iooxa={"id": {"block": "k7pl5S6JfwtM3TGGA87G", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Define Inverse Problem
#
# The inverse problem is defined by 3 things:
#
# 1) Data Misfit: a measure of how well our recovered model explains the field data
# 2) Regularization: constraints placed on the recovered model and a priori information
# 3) Optimization: the numerical approach used to solve the inverse problem
#
#
#
# + iooxa={"id": {"block": "xYq92mR9VSr3IkNnB0B1", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# Define the data misfit. Here the data misfit is the L2 norm of the weighted
# residual between the observed data and the data predicted for a given model.
# Within the data misfit, the residual between predicted and observed data are
# normalized by the data's standard deviation.
dmis = data_misfit.L2DataMisfit(data=data_object, simulation=simulation)
# Define the regularization (model objective function)
reg = regularization.Sparse(
mesh,
indActive=ind_active,
mapping=model_map,
mref=reference_model,
gradientType="total",
alpha_s=1,
alpha_x=1,
alpha_y=1,
alpha_z=1,
)
# Define sparse and blocky norms ps, px, py, pz
ps = 0
px = 2
py = 2
pz = 2
reg.norms = np.c_[ps, px, py, pz]
# Define how the optimization problem is solved. Here we will use a projected
# Gauss-Newton approach that employs the conjugate gradient solver.
opt = optimization.ProjectedGNCG(
maxIter=100, lower=0.0, upper=np.Inf, maxIterLS=20, maxIterCG=30, tolCG=1e-3
)
# Here we define the inverse problem that is to be solved
inv_prob = inverse_problem.BaseInvProblem(dmis, reg, opt)
# + [markdown] iooxa={"id": {"block": "9dnfccgTdxMgTgEUhWyX", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Define Inversion Directives
#
# Here we define any directiveas that are carried out during the inversion. This
# includes the cooling schedule for the trade-off parameter (beta), stopping
# criteria for the inversion and saving inversion results at each iteration.
#
#
#
# + iooxa={"id": {"block": "xoattZV3IOqjWDTFQ5vA", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
# Defining a starting value for the trade-off parameter (beta) between the data
# misfit and the regularization.
starting_beta = directives.BetaEstimate_ByEig(beta0_ratio=1)
beta_schedule = directives.BetaSchedule(coolingFactor=2, coolingRate=1)
# Options for outputting recovered models and predicted data as a dictionary
save_dictionary = directives.SaveOutputDictEveryIteration()
# Defines the directives for the IRLS regularization. This includes setting
# the cooling schedule for the trade-off parameter.
update_IRLS = directives.Update_IRLS(
f_min_change=1e-3, max_irls_iterations=20, coolEpsFact=1.5, beta_tol=1e-2,
chifact_target=1, chifact_start=1
)
# Updating the preconditionner if it is model dependent.
update_jacobi = directives.UpdatePreconditioner()
# Setting a stopping criteria for the inversion.
target_misfit = directives.TargetMisfit(chifact=1)
# Add sensitivity weights
sensitivity_weights = directives.UpdateSensitivityWeights(everyIter=False)
opt.remember('xc')
# The directives are defined as a list.
directives_list = [
sensitivity_weights,
starting_beta,
save_dictionary,
update_IRLS,
update_jacobi,
]
# + [markdown] iooxa={"id": {"block": "QAEzeEMBS4UYigFzuPac", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Running the Inversion
#
# To define the inversion object, we need to define the inversion problem and
# the set of directives. We can then run the inversion.
#
#
#
# + iooxa={"id": {"block": "gQ5328059eGkAsDbWips", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "f1fmFdNpovSwRy4Sve6F", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# Here we combine the inverse problem and the set of directives
inv = inversion.BaseInversion(inv_prob, directives_list)
# Print target misfit to compare with convergence
# print("Target misfit is " + str(target_misfit.target))
# Run the inversion
recovered_model = inv.run(starting_model)
# + iooxa={"id": {"block": "s1NBa8bHKt5ID5psRCtC", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
def plot_tikhonov_curve(iteration, scale):
phi_d = []
phi_m = []
beta = []
iterations = np.arange(len(save_dictionary.outDict)) + 1
for kk in iterations:
phi_d.append(save_dictionary.outDict[kk]['phi_d'])
phi_m.append(save_dictionary.outDict[kk]['phi_m'])
beta.append(save_dictionary.outDict[kk]['beta'])
fig, axs = plt.subplots(1, 2, figsize=(12,5))
axs[0].plot(phi_m ,phi_d, 'k.-')
axs[0].plot(phi_m[iteration-1] ,phi_d[iteration-1], 'go', ms=10)
axs[0].set_xlabel("$\phi_m$")
axs[0].set_ylabel("$\phi_d$")
axs[0].grid(True)
axs[1].plot(iterations, phi_d, 'k.-')
axs[1].plot(iterations[iteration-1], phi_d[iteration-1], 'go', ms=10)
ax_1 = axs[1].twinx()
ax_1.plot(iterations, phi_m, 'r.-')
ax_1.plot(iterations[iteration-1], phi_m[iteration-1], 'go', ms=10)
axs[1].set_ylabel("$\phi_d$")
ax_1.set_ylabel("$\phi_m$")
axs[1].set_xlabel("Iterations")
axs[1].grid(True)
axs[0].set_title(
"$\phi_d$={:.1e}, $\phi_m$={:.1e}, $\\beta$={:.1e}".format(phi_d[iteration-1], phi_m[iteration-1], beta[iteration-1]),
fontsize = 14
)
axs[1].set_title("Target misfit={:.0f}".format(survey.nD/2))
for ii, ax in enumerate(axs):
if ii == 0:
ax.set_xscale(scale)
ax.set_yscale(scale)
xlim = ax.get_xlim()
ax.hlines(survey.nD/2, xlim[0], xlim[1], linestyle='--', label='$\phi_d^{*}$')
ax.set_xlim(xlim)
axs[0].legend()
plt.tight_layout()
# + iooxa={"id": {"block": "qVnPWo578COas4ItZ6jR", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "JsoHKBWN62IFY8vMpwF2", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
interact(
plot_tikhonov_curve,
iteration=widgets.IntSlider(min=1, max=len(save_dictionary.outDict), step=1, continuous_update=False),
scale=widgets.RadioButtons(options=["linear", "log"])
)
# + iooxa={"id": {"block": "1uBaWvM9cFr371dHqT8o", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
susceptibility_model = save_dictionary.outDict[31]['m']
# + iooxa={"id": {"block": "am3p4LsPAGjCvlEssNOk", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
def plot_model_histogram(iteration, yscale):
out = plt.hist(save_dictionary.outDict[iteration]['m'], bins=np.linspace(0, 0.1))
plt.xlabel('Susceptibility (SI)')
plt.yscale(yscale)
plt.ylabel('Counts')
# plt.ylim(10, 1e5)
# + iooxa={"id": {"block": "GiHN32hUsqD73U53As9y", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "yCQZOejMhi7rP8n6SKSC", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
interact(
plot_model_histogram,
iteration=widgets.IntSlider(min=1, max=len(save_dictionary.outDict), step=1),
yscale=widgets.RadioButtons(options=["linear", "log"])
)
# + iooxa={"id": {"block": "wfKAKue4ZpRyEaI4pDOM", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
def plot_dobs_vs_dpred(iteration):
# Predicted data with final recovered model
dpred = save_dictionary.outDict[iteration]['dpred']
# Observed data | Predicted data | Normalized data misfit
data_array = np.c_[dobs, dpred, (dobs - dpred) / standard_deviation]
vmin, vmax = dobs.min(), dobs.max()
fig = plt.figure(figsize=(17, 4))
plot_title = ["Observed", "Predicted", "Normalized Misfit"]
plot_units = ["nT", "nT", ""]
ax1 = 3 * [None]
ax2 = 3 * [None]
norm = 3 * [None]
cbar = 3 * [None]
cplot = 3 * [None]
v_lim = [(vmin, vmax), (vmin, vmax),(-3,3)]
for ii in range(0, 3):
ax1[ii] = fig.add_axes([0.33 * ii + 0.03, 0.11, 0.25, 0.84])
cplot[ii] = plot2Ddata(
receiver_list[0].locations,
data_array[:, ii],
ax=ax1[ii],
ncontour=30,
clim=v_lim[ii],
contourOpts={"cmap": "Spectral_r"},
)
ax1[ii].set_title(plot_title[ii])
ax1[ii].set_xlabel("x (m)")
ax1[ii].set_ylabel("y (m)")
ax2[ii] = fig.add_axes([0.33 * ii + 0.27, 0.11, 0.01, 0.84])
norm[ii] = mpl.colors.Normalize(vmin=v_lim[ii][0], vmax=v_lim[ii][1])
cbar[ii] = mpl.colorbar.ColorbarBase(
ax2[ii], norm=norm[ii], orientation="vertical", cmap=mpl.cm.Spectral_r
)
cbar[ii].set_label(plot_units[ii], rotation=270, labelpad=15, size=12)
for ax in ax1[1:]:
ax.set_ylabel("")
ax.set_yticklabels([])
plt.show()
# + iooxa={"id": {"block": "e9SIBXX14ow1fa2Ug2S8", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "28DF3QQTrsk5RNq7Xnv8", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
interact(plot_dobs_vs_dpred, iteration=widgets.IntSlider(min=1, max=len(save_dictionary.outDict), step=1, value=1))
# + iooxa={"id": {"block": "rFCb20u7KpIf4VSmnEBa", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "RUiWSHv62CDX0G4XFeRH", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
def plot_recovered_model(iteration, xslice, yslice, zslice, vmax):
fig = plt.figure(figsize=(10, 10))
mesh.plot_3d_slicer(
save_dictionary.outDict[iteration]['m'], clim=(0, vmax),
xslice=xslice,
yslice=yslice,
zslice=zslice,
fig=fig,
pcolor_opts={'cmap':'Spectral_r'}
)
interact(
plot_recovered_model,
iteration=widgets.IntSlider(min=1, max=len(save_dictionary.outDict), value=0),
xslice=widgets.FloatText(value=2000, step=100),
yslice=widgets.FloatText(value=41000, step=100),
zslice=widgets.FloatText(value=-800, step=100),
vmax=widgets.FloatText(value=0.07),
)
# + [markdown] iooxa={"id": {"block": "mYiSLbeGieb8NTsKbWqW", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ## Comparing the historic model with the recovered model
#
# + iooxa={"id": {"block": "ALFEYmYgU5pSpAq3bjoT", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
from discretize.utils import ExtractCoreMesh
zmin, zmax = -1500, 0
ymin, ymax = receiver_locations[:,1].min(), receiver_locations[:,1].max()
xmin, xmax = receiver_locations[:,0].min(), receiver_locations[:,0].max()
xyzlim = np.array([[xmin, xmax],[ymin, ymax], [zmin, zmax]])
inds_core, mesh_core = ExtractCoreMesh(xyzlim, mesh)
# + iooxa={"id": {"block": "KURVN5vpHplvEsqxTZz8", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": null}
import pyvista as pv
def plot_3d_with_pyvista(model, notebook=True, threshold=0.04):
pv.set_plot_theme("document")
# Get the PyVista dataset of the inverted model
dataset = mesh_core.to_vtk({'susceptibility':model})
# Create the rendering scene
p = pv.Plotter(notebook=notebook)
# add a grid axes
p.show_grid()
# Extract volumetric threshold
threshed = dataset.threshold(threshold, invert=False)
# Add spatially referenced data to the scene
dparams = dict(
show_edges=False,
cmap="Spectral_r",
clim=[0, 0.07],
stitle='Susceptibility (SI)',
)
p.add_mesh(threshed, **dparams)
p.set_scale(1,1,1)
cpos = [(-5248.506818695238, 35263.832232792156, 4945.734122744097),
(2140.1554568144284, 40814.32410594353, -1198.9698078219635),
(0.4274014723619113, 0.35262874486945933, 0.8324547733749025)]
p.camera_position = cpos
p.show(window_size=[1024, 768])
# + [markdown] iooxa={"id": {"block": "C4qI6WHmxq7cc5LAbOmz", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ### Recovered susceptiblity model from L2-norm inversion
# + iooxa={"id": {"block": "m5WoFEunUeIefQ3nxsp4", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "5AgnYefnFLeN9mUHP3kV", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
plot_3d_with_pyvista(inv_prob.l2model[inds_core], notebook=True, threshold=0.03)
# + [markdown] iooxa={"id": {"block": "Qz9yky7qUFvKfbVwyWLb", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
# ### Recovered susceptiblity model from Lp-norm inversion
# + iooxa={"id": {"block": "ZTgDJB0thZrQDM2jidBS", "project": "FesuZy9nERkRH9r67vBe", "version": 1}, "outputId": {"block": "q1OOmsjAY081euRb95nt", "project": "FesuZy9nERkRH9r67vBe", "version": 1}}
plot_3d_with_pyvista(susceptibility_model[inds_core], notebook=True, threshold=0.07)
# + iooxa
|
3-magnetic-inversion-raglan-lp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''AImusic'': conda)'
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import warnings
warnings.filterwarnings("ignore")
# -
tf.test.is_gpu_available()
data = pd.read_table("dataset.txt",index_col=0)
data
x = data['Education']
y = data['Income']
plt.scatter(x,y)
model = tf.keras.Sequential()
model.add(tf.keras.layers.Dense(1,input_shape=(1,)))
model.summary()
model.compile(optimizer="adam",
loss="mse" )
# + tags=[]
model.fit(x,y,epochs=5000)
# + tags=[]
z = model.predict(x)
z
# -
plt.scatter(x,y)
plt.plot(x,z)
plt.show()
|
docs/jupyter/LineRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Import raw data
import graphlab as gl
gl.canvas.set_target('ipynb')
sf = gl.SFrame('https://static.turi.com/datasets/extended-bakery/bakery_train.sf')
sf
sf['Item'].show()
# ## Find Patterns
model = gl.frequent_pattern_mining.create(sf,
item='Item',
features = ['Receipt', 'StoreNum'],
min_length=2)
patterns = model.get_frequent_patterns()
patterns.print_rows(max_column_width=100)
model.save('pattern_mining_model.gl')
# ## Making predictions!
new_data = gl.SFrame({'Receipt': [1356]*2,
'StoreNum': [2]*2,
'Item': ['CherryTart', 'ApplePie']})
model.predict(new_data)
model.predict_topk(new_data)
# ## Extract features
pattern_sf = model.extract_features(sf)
pattern_sf
# ## Cluster in Employee Space
emps = sf.groupby(['StoreNum','Receipt'], {
'EmpId': gl.aggregate.SELECT_ONE('EmpId')})
emps
emp_space = emps.join(pattern_sf)\
.groupby('EmpId', {'all_features': gl.aggregate.SUM('extracted_features')})
emp_space
cl_model = gl.kmeans.create(emp_space,
features = ['all_features'],
num_clusters=3)
emp_space['cluster_id'] = cl_model['cluster_id']['cluster_id']
emp_space
emp_space.show(x='cluster_id', y = 'StoreNum', view='Bar Chart')
|
webinars/pattern-mining/demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Hc0XEe7S6Ft4" colab_type="text"
# # Seq2Seq with Attention for Korean-English Neural Machine Translation
# - Network architecture based on this [paper](https://arxiv.org/abs/1409.0473)
# - Fit to run on Google Colaboratory
# + id="jCfquy219jiJ" colab_type="code" colab={}
import os
import io
import tarfile
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchtext
from torchtext.data import Dataset
from torchtext.data import Example
from torchtext.data import Field
from torchtext.data import BucketIterator
# + [markdown] id="jlASYXQobfgp" colab_type="text"
# # 1. Upload Data to Colab Workspace
#
# 로컬에 존재하는 다음 3개의 데이터를 가상 머신에 업로드. 파일의 원본은 [여기](https://github.com/jungyeul/korean-parallel-corpora/tree/master/korean-english-news-v1/)에서도 확인
#
# - korean-english-park.train.tar.gz
# - korean-english-park.dev.tar.gz
# - korean.english-park.test.tar.gz
#
#
# + id="Jkgihb-qb5Iz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 144} outputId="413ab44e-002b-4b2c-8e06-67053a6efb14" executionInfo={"status": "ok", "timestamp": 1564618973556, "user_tz": -540, "elapsed": 6206, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# 현재 작업경로를 확인 & 'data' 폴더 생성
# !echo 'Current working directory:' ${PWD}
# !mkdir -p data/
# !ls -al
# + id="e8aYPNtsdAeG" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 148} outputId="3fb49427-4e93-4376-e309-e993e9874e97" executionInfo={"status": "ok", "timestamp": 1564619081580, "user_tz": -540, "elapsed": 92109, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# 로컬의 데이터 업로드
from google.colab import files
uploaded = files.upload()
# + id="A_3Z9eMddr8E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="f24aaa8f-dcd2-4150-b076-5e2add7956bb" executionInfo={"status": "ok", "timestamp": 1564619117329, "user_tz": -540, "elapsed": 3611, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# 'data' 폴더 하위로 이동, 잘 옮겨졌는지 확인
# !mv *.tar.gz data/
# !ls -al data/
# + [markdown] id="Prea2yTCfwKe" colab_type="text"
# # 2. Check Packages
# + [markdown] id="xcuXRzDdkl0M" colab_type="text"
# ## KoNLPy (설치 필요)
# + id="c8IEEmXEecvQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="73894605-92f0-4fb8-9cb5-9dad2a377c08" executionInfo={"status": "ok", "timestamp": 1564619212775, "user_tz": -540, "elapsed": 89594, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Java 1.8 & KoNLPy 설치
# !apt-get update
# !apt-get install g++ openjdk-8-jdk python-dev python3-dev
# !pip3 install JPype1-py3
# !pip3 install konlpy
# + id="7-fVO8z6iatj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 237} outputId="8a724e68-09ca-44a7-da06-9c1ac781cabd" executionInfo={"status": "ok", "timestamp": 1564619879065, "user_tz": -540, "elapsed": 7857, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
from konlpy.tag import Okt
ko_tokens = Okt().pos('트위터 데이터로 학습한 형태소 분석기가 잘 실행이 되는지 확인해볼까요?') # list of (word, POS TAG) tuples
ko_tokens = [t[0] for t in ko_tokens] # Only get words
print(ko_tokens)
del ko_tokens # 필요 없으니까 삭제
# + [markdown] id="Rk3DQJRdkqmc" colab_type="text"
# ## Spacy (이미 설치되어 있음)
# + id="ub_8K6GxkU_V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 199} outputId="9042ae99-523b-4d0b-cf54-5bcf22e62e1d" executionInfo={"status": "ok", "timestamp": 1564619886004, "user_tz": -540, "elapsed": 4101, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# 설치가 되어있는지 확인
# !pip show spacy
# + id="KIB9ZBpWln-G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 92} outputId="058fd67c-17f1-453c-b25f-09cd6e2feca9" executionInfo={"status": "ok", "timestamp": 1564619894215, "user_tz": -540, "elapsed": 6354, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# 설치가 되어있는지 확인 (없다면 자동설치됨)
# !python -m spacy download en_core_web_sm
# + id="18MJjo21kyG1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4eec21f5-d56a-4be8-ef88-73913126abe8" executionInfo={"status": "ok", "timestamp": 1564619896670, "user_tz": -540, "elapsed": 1488, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
import spacy
spacy_en = spacy.load('en_core_web_sm')
en_tokens = [t.text for t in spacy_en.tokenizer('Check that spacy tokenizer works.')]
print(en_tokens)
del en_tokens # 필요 없으니까 삭제
# + [markdown] id="skY4owhASmK5" colab_type="text"
# # 3. Define Tokenizing Functions
# 문장을 받아 그보다 작은 어절 혹은 형태소 단위의 리스트로 반환해주는 함수를 각 언어에 대해 작성
# - Korean: konlpy.tag.Okt() <- Twitter()에서 명칭변경
# - English: spacy.tokenizer
# + [markdown] id="0R17vYJMT8mq" colab_type="text"
# ## Korean Tokenizer
# + id="XoJWSFgLTTjG" colab_type="code" colab={}
#from konlpy.tag import Okt
class KoTokenizer(object):
"""For Korean."""
def __init__(self):
self.tokenizer = Okt()
def tokenize(self, text):
tokens = self.tokenizer.pos(text)
tokens = [t[0] for t in tokens]
return tokens
# + id="MLilyD9p1dwQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e0ea394f-1546-4179-87cd-74bd8fc402bb" executionInfo={"status": "ok", "timestamp": 1564619906657, "user_tz": -540, "elapsed": 585, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Usage example
print(KoTokenizer().tokenize('전처리는 언제나 지겨워요.'))
# + [markdown] id="-HF7H_h_UAXb" colab_type="text"
# ## English Tokenizer
# + id="Nh5klj0N0VQz" colab_type="code" colab={}
#import spacy
class EnTokenizer(object):
"""For English."""
def __init__(self):
self.spacy_en = spacy.load('en_core_web_sm')
def tokenize(self, text):
tokens = [t.text for t in self.spacy_en.tokenizer(text)]
return tokens
# + id="xH9qI4OH1x0t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="62f9bd75-4990-400c-be03-2eacdb3ba36c" executionInfo={"status": "ok", "timestamp": 1564619924860, "user_tz": -540, "elapsed": 913, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Usage example
print(EnTokenizer().tokenize("What I cannot create, I don't understand."))
# + [markdown] id="tkprvaRwR2HB" colab_type="text"
# # 4. Data Preprocessing
# + [markdown] id="31Rm1iulT4Wb" colab_type="text"
# ## Load data
# + id="tVGEo34ClGkr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 144} outputId="b15a075f-cd4b-496e-c6dc-2869a92c28be" executionInfo={"status": "ok", "timestamp": 1564619930898, "user_tz": -540, "elapsed": 3603, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Current working directory & list of files
# !echo 'Current working directory:' ${PWD}
# !ls -al
# + id="5SOUbMf59YWo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 108} outputId="e6e535b0-7c0e-4162-ec18-9df11ec6ae20" executionInfo={"status": "ok", "timestamp": 1564619938544, "user_tz": -540, "elapsed": 581, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
DATA_DIR = './data/'
print('Data directory exists:', os.path.isdir(DATA_DIR))
print('List of files:')
print(*os.listdir(DATA_DIR), sep='\n')
# + id="CBmf2ZXQ9w5m" colab_type="code" colab={}
def get_data_from_tar_gz(filename):
"""
Retrieve contents from a `tar.gz` file without extraction.
Arguments:
filename: path to `tar.gz` file.
Returns:
dict, (name, content) pairs
"""
assert os.path.exists(filename)
out = {}
with tarfile.open(filename, 'r:gz') as tar:
for member in tar.getmembers():
lang = member.name.split('.')[-1] # ex) korean-english-park.train.ko -> ko
f = tar.extractfile(member)
if f is not None:
content = f.read().decode('utf-8')
content = content.splitlines()
out[lang] = content
assert isinstance(out, dict)
return out
# + id="mlZutqFNBtED" colab_type="code" colab={}
# Each 'xxx_data' is a dictionary with keys; 'ko', 'en'
train_dict= get_data_from_tar_gz(os.path.join(DATA_DIR, 'korean-english-park.train.tar.gz')) # train
dev_dict = get_data_from_tar_gz(os.path.join(DATA_DIR, 'korean-english-park.dev.tar.gz')) # dev
test_dict = get_data_from_tar_gz(os.path.join(DATA_DIR, 'korean-english-park.test.tar.gz')) # test
# + id="kkSYlbgEFcpp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="323e3517-623f-4fc2-c52d-6bb0682814ab" executionInfo={"status": "ok", "timestamp": 1564619955012, "user_tz": -540, "elapsed": 600, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Some samples (ko)
train_dict['ko'][100:105]
# + id="7n-b6F_8HWF1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="29557f2e-be45-4fa6-f67c-a27c6a38b24c" executionInfo={"status": "ok", "timestamp": 1564619956865, "user_tz": -540, "elapsed": 595, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Some samples (en)
train_dict['en'][100:105]
# + [markdown] id="E4J3HQQ3LGQ-" colab_type="text"
# ## Define Datasets
# + id="5qlAw_oDGYCV" colab_type="code" colab={}
#from torchtext.data import Dataset
#from torchtext.data import Example
class KoEnTranslationDataset(Dataset):
"""A dataset for Korean-English Neural Machine Translation."""
@staticmethod
def sort_key(ex):
return torchtext.data.interleave_keys(len(ex.src), len(ex.trg))
def __init__(self, data_dict, field_dict, source_lang='ko', max_samples=None, **kwargs):
"""
Only 'ko' and 'en' supported for `language`
Arguments:
data_dict: dict of (`language`, text) pairs.
field_dict: dict of (`language`, Field instance) pairs.
source_lang: str, default 'ko'.
Other kwargs are passed to the constructor of `torchtext.data.Dataset`.
"""
if not all(k in ['ko', 'en'] for k in data_dict.keys()):
raise KeyError("Check data keys.")
if not all(k in ['ko', 'en'] for k in field_dict.keys()):
raise KeyError("Check field keys.")
if source_lang == 'ko':
fields = [('src', field_dict['ko']), ('trg', field_dict['en'])]
src_data = data_dict['ko']
trg_data = data_dict['en']
elif source_lang == 'en':
fields = [('src', field_dict['en']), ('trg', field_dict['ko'])]
src_data = data_dict['en']
trg_data = data_dict['ko']
else:
raise NotImplementedError
if not len(src_data) == len(trg_data):
raise ValueError('Inconsistent number of instances between two languages.')
examples = []
for i, (src_line, trg_line) in enumerate(zip(src_data, trg_data)):
src_line = src_line.strip()
trg_line = trg_line.strip()
if src_line != '' and trg_line != '':
examples.append(
torchtext.data.Example.fromlist(
[src_line, trg_line], fields
)
)
i += 1
if max_samples is not None:
if i >= max_samples:
break
super(KoEnTranslationDataset, self).__init__(examples, fields, **kwargs)
# + [markdown] id="xui8jchJJThJ" colab_type="text"
# ## Define Fields
# - Instantiate tokenizers; one for each language.
# - The 'tokenize' argument of `Field` requires a tokenizing function.
# + id="NFVGDL5fOVUn" colab_type="code" colab={}
#from torchtext.data import Field
ko_tokenizer = KoTokenizer() # korean tokenizer
en_tokenizer = EnTokenizer() # english tokenizer
# Field instance for korean
KOREAN = Field(
init_token='<sos>',
eos_token='<eos>',
tokenize=ko_tokenizer.tokenize,
batch_first=True,
lower=False
)
# Field instance for english
ENGLISH = Field(
init_token='<sos>',
eos_token='<eos>',
tokenize=en_tokenizer.tokenize,
batch_first=True,
lower=True
)
# Store Field instances in a dictionary
field_dict = {
'ko': KOREAN,
'en': ENGLISH,
}
# + [markdown] id="hk2n8MRxLNys" colab_type="text"
# ## Instantiate datasets
# - one for each set (train, dev, test)
# + id="p-4k377GoFki" colab_type="code" colab={}
# 학습시간 단축을 위해 학습 데이터 줄이기
MAX_TRAIN_SAMPLES = 10000
# + id="9RYxWABtPxc7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="1e0fda13-e40b-4a2d-8064-ed05807746e9" executionInfo={"status": "ok", "timestamp": 1564620242131, "user_tz": -540, "elapsed": 52651, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Instantiate with data
train_set = KoEnTranslationDataset(train_dict, field_dict, max_samples=MAX_TRAIN_SAMPLES)
print('Train set ready.')
print('#. examples:', len(train_set.examples))
dev_set = KoEnTranslationDataset(dev_dict, field_dict)
print('Dev set ready...')
print('#. examples:', len(dev_set.examples))
test_set = KoEnTranslationDataset(test_dict, field_dict)
print('Test set ready...')
print('#. examples:', len(test_set.examples))
# + id="q8frH5waRAfa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="ce409644-246f-4cdc-c110-2e914cfc823a" executionInfo={"status": "ok", "timestamp": 1564620244390, "user_tz": -540, "elapsed": 568, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Training example (KO, source language)
train_set.examples[50].src
# + id="1OukZ7MqRCff" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="87e0c0a6-221f-4286-c6aa-4c85372368c3" executionInfo={"status": "ok", "timestamp": 1564620246770, "user_tz": -540, "elapsed": 596, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Training example (EN, target language)
train_set.examples[50].trg
# + [markdown] id="g9mfm9nyzA8f" colab_type="text"
# ## Build Vocabulary
# - 각 언어별 생성: `Field`의 인스턴스를 활용
# - 최소 빈도수(`MIN_FREQ`) 값을 작게 하면 vocabulary의 크기가 커짐.
# - 최소 빈도수(`MIN_FREQ`) 값을 크게 하면 vocabulary의 크기가 작아짐.
#
# + id="tqTH_3EelkFC" colab_type="code" colab={}
MIN_FREQ = 2 # TODO: try different values
# + id="gB3ssFibRLOi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="919a60a0-b082-4db9-d061-eda64151e122" executionInfo={"status": "ok", "timestamp": 1564630186266, "user_tz": -540, "elapsed": 740, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Build vocab for Korean
KOREAN.build_vocab(train_set, dev_set, test_set, min_freq=MIN_FREQ) # ko
print('Size of source vocab (ko):', len(KOREAN.vocab))
# + id="myIIOwR29spe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="110454b2-450d-46a1-9c01-5483e32e3171" executionInfo={"status": "ok", "timestamp": 1564630190549, "user_tz": -540, "elapsed": 574, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Check indices of some important tokens
tokens = ['<unk>', '<pad>', '<sos>', '<eos>']
for token in tokens:
print(f"{token} -> {KOREAN.vocab.stoi[token]}")
# + id="YYV-xvYnzL-k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a9902faf-6e3d-47e5-8ce8-d3374f0a3b99" executionInfo={"status": "ok", "timestamp": 1564630200499, "user_tz": -540, "elapsed": 568, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Build vocab for English
ENGLISH.build_vocab(train_set, dev_set, test_set, min_freq=MIN_FREQ) # en
print('Size of target vocab (en):', len(ENGLISH.vocab))
# + id="ygH06w7p-XFS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="d<PASSWORD>8e10-6<PASSWORD>" executionInfo={"status": "ok", "timestamp": 1564630204973, "user_tz": -540, "elapsed": 470, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Check indices of some important tokens
tokens = ['<unk>', '<pad>', '<sos>', '<eos>']
for token in tokens:
print(f"{token} -> {KOREAN.vocab.stoi[token]}")
# + [markdown] id="rGZS2-xBz8I-" colab_type="text"
# ## Configure Device
# - *'런타임' -> '런타임 유형변경'* 에서 하드웨어 가속기로 **GPU** 선택
# + id="39RfXwWszTi8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b58ac17b-a0d7-4507-f183-5a12c1a9e56e" executionInfo={"status": "ok", "timestamp": 1564630207583, "user_tz": -540, "elapsed": 540, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print('Device to use:', device)
# + [markdown] id="VGVUyRY_0f7s" colab_type="text"
# ## Create Data Iterators
# - 데이터를 미니배치(mini-batch) 단위로 반환해주는 역할
# - `train_set`, `dev_set`, `test_set`에 대해 개별적으로 정의해야 함
# - `BATCH_SIZE`를 정의해주어야 함
# - `torchtext.data.BucketIterator`는 하나의 미니배치를 서로 비슷한 길이의 관측치들로 구성함
# - [Bucketing](https://medium.com/@rashmi.margani/how-to-speed-up-the-training-of-the-sequence-model-using-bucketing-techniques-9e302b0fd976)의 효과: 하나의 미니배치 내 padding을 최소화하여 연산의 낭비를 줄여줌
#
# + id="ZmBkViElnEAj" colab_type="code" colab={}
BATCH_SIZE = 128
# + id="PoX4TwE5m6si" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="195fd9a4-24d5-4576-898a-26e0ad9831d6" executionInfo={"status": "ok", "timestamp": 1564630211789, "user_tz": -540, "elapsed": 588, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
#from torchtext.data import BucketIterator
# Train iterator
train_iterator = BucketIterator(
train_set,
batch_size=BATCH_SIZE,
train=True,
shuffle=True,
device=device
)
print(f'Number of minibatches per epoch: {len(train_iterator)}')
# + id="Pc0B3HS7nBkl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="261e7a0e-2898-4381-9aad-ef88845d6272" executionInfo={"status": "ok", "timestamp": 1564630219540, "user_tz": -540, "elapsed": 546, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
#from torchtext.data import BucketIterator
# Dev iterator
dev_iterator = BucketIterator(
dev_set,
batch_size=100,
train=False,
shuffle=False,
device=device
)
print(f'Number of minibatches per epoch: {len(dev_iterator)}')
# + id="3YNd4heenVGn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="cab434a8-32a8-4b7a-e1ff-38bea095f522" executionInfo={"status": "ok", "timestamp": 1564630221806, "user_tz": -540, "elapsed": 578, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
#from torchtext.data import BucketIterator
# Test iterator
test_iterator = BucketIterator(
test_set,
batch_size=200,
train=False,
shuffle=False,
device=device
)
print(f'Number of minibatches per epoch: {len(test_iterator)}')
# + id="Ivq4PK8O1qG2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="41695b12-cb89-4335-e505-b043d51dfff4" executionInfo={"status": "ok", "timestamp": 1564630223143, "user_tz": -540, "elapsed": 738, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
train_batch = next(iter(train_iterator))
print('a batch of source examples has shape:', train_batch.src.size()) # (b, s)
print('a batch of target examples has shape:', train_batch.trg.size()) # (b, s)
# + id="otfd7srHBec9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2897bda0-bafd-41c8-a0f9-1579a876f642" executionInfo={"status": "ok", "timestamp": 1564630224739, "user_tz": -540, "elapsed": 529, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Checking first sample in mini-batch (KO, source lang)
ko_indices = train_batch.src[0]
ko_tokens = [KOREAN.vocab.itos[i] for i in ko_indices]
for t, i in zip(ko_tokens, ko_indices):
print(f"{t} ({i})")
del ko_indices, ko_tokens
# + id="cizBVHogBhbY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 945} outputId="48ea21cd-52d2-401c-a2a9-6304d3f64fa2" executionInfo={"status": "ok", "timestamp": 1564630229897, "user_tz": -540, "elapsed": 613, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Checking first sample in mini-batch (EN, target lang)
en_indices = train_batch.trg[0]
en_tokens = [ENGLISH.vocab.itos[i] for i in en_indices]
for t, i in zip(en_tokens, en_indices):
print(f"{t} ({i})")
del en_indices, en_tokens
# + id="vUC250YvB4cS" colab_type="code" colab={}
del train_batch # 더 이상 필요 없으니까 삭제
# + [markdown] id="E4h6GTTJ1_vs" colab_type="text"
# # 5. Building Seq2Seq Model
# + [markdown] id="k_Eq8AyXMmpH" colab_type="text"
# ## Hyperparameters
# + id="MWed4n21MkGN" colab_type="code" colab={}
# Hyperparameters
INPUT_DIM = len(KOREAN.vocab)
OUTPUT_DIM = len(ENGLISH.vocab)
ENC_EMB_DIM = DEC_EMB_DIM = 100
ENC_HID_DIM = DEC_HID_DIM = 60
USE_BIDIRECTIONAL = False
# + [markdown] id="hWK3vdrW2Io3" colab_type="text"
# ## Encoder
# + id="iPIPAhi01ruC" colab_type="code" colab={}
class Encoder(nn.Module):
"""
Learns an embedding for the source text.
Arguments:
input_dim: int, size of input language vocabulary.
emb_dim: int, size of embedding layer output.
enc_hid_dim: int, size of encoder hidden state.
dec_hid_dim: int, size of decoder hidden state.
bidirectional: uses bidirectional RNNs if True. default is False.
"""
def __init__(self, input_dim, emb_dim, enc_hid_dim, dec_hid_dim, bidirectional=False):
super(Encoder, self).__init__()
self.input_dim = input_dim
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.bidirectional = bidirectional
self.embedding = nn.Embedding(
num_embeddings=self.input_dim,
embedding_dim=self.emb_dim
)
self.rnn = nn.GRU(
input_size=self.emb_dim,
hidden_size=self.enc_hid_dim,
bidirectional=self.bidirectional,
batch_first=True
)
self.rnn_output_dim = self.enc_hid_dim
if self.bidirectional:
self.rnn_output_dim *= 2
self.fc = nn.Linear(self.rnn_output_dim, self.dec_hid_dim)
self.dropout = nn.Dropout(.2)
def forward(self, src):
"""
Arguments:
src: 2d tensor of shape (batch_size, input_seq_len)
Returns:
outputs: 3d tensor of shape (batch_size, input_seq_len, num_directions * enc_h)
hidden: 2d tensor of shape (b, dec_h). This tensor will be used as the initial
hidden state value of the decoder (h0 of decoder).
"""
assert len(src.size()) == 2, 'Input requires dimension (batch_size, seq_len).'
# Shape: (b, s, h)
embedded = self.embedding(src)
embedded = self.dropout(embedded)
outputs, hidden = self.rnn(embedded)
if self.bidirectional:
# (2, b, enc_h) -> (b, 2 * enc_h)
hidden = torch.cat((hidden[-2, :, :], hidden[-1, :, :]), dim=1)
else:
# (1, b, enc_h) -> (b, enc_h)
hidden = hidden.squeeze(0)
# (b, num_directions * enc_h) -> (b, dec_h)
hidden = self.fc(hidden)
hidden = torch.tanh(hidden)
return outputs, hidden
# + [markdown] id="iJtFNEYp-ett" colab_type="text"
# ## Attention
# + id="EdjDNWD1gnS_" colab_type="code" colab={}
class Attention(nn.Module):
def __init__(self, enc_hid_dim, dec_hid_dim, encoder_is_bidirectional=False):
super(Attention, self).__init__()
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.encoder_is_bidirectional = encoder_is_bidirectional
self.attention_input_dim = enc_hid_dim + dec_hid_dim
if self.encoder_is_bidirectional:
self.attention_input_dim += enc_hid_dim # 2 * h_enc + h_dec
self.linear = nn.Linear(self.attention_input_dim, dec_hid_dim)
self.v = nn.Parameter(torch.rand(dec_hid_dim))
def forward(self, hidden, encoder_outputs):
"""
Arguments:
hidden: 2d tensor with shape (batch_size, dec_hid_dim).
encoder_outputs: 3d tensor with shape (batch_size, input_seq_len, enc_hid_dim).
if encoder is bidirectional, expects (batch_size, input_seq_len, 2 * enc_hid_dim).
"""
# Shape check
assert hidden.dim() == 2
assert encoder_outputs.dim() == 3
batch_size, seq_len, _ = encoder_outputs.size()
# (b, dec_h) -> (b, s, dec_h)
hidden = hidden.unsqueeze(1).expand(-1, seq_len, -1)
# concat; shape results in (b, s, enc_h + dec_h).
# if encoder is bidirectional, (b, s, 2 * h_enc + h_dec).
concat = torch.cat((hidden, encoder_outputs), dim=2)
# concat; shape is (b, s, dec_h)
concat = self.linear(concat)
concat = torch.tanh(concat)
# tile v; (dec_h, ) -> (b, dec_h, 1)
v = self.v.repeat(batch_size, 1).unsqueeze(2)
# attn; (b, s, dec_h) @ (b, dec_h, 1) -> (b, s, 1) -> (b, s)
attn_scores = torch.bmm(concat, v).squeeze(-1)
assert attn_scores.dim() == 2 # Final shape check: (b, s)
return F.softmax(attn_scores, dim=1)
# + [markdown] id="6-d9zt0SF9Ff" colab_type="text"
# ## Decoder
# + id="IcG1tbhDC8-1" colab_type="code" colab={}
class Decoder(nn.Module):
"""
Unlike the encoder, a single forward pass of
a `Decoder` instance is defined for only a single timestep.
Arguments:
output_dim: int,
emb_dim: int,
enc_hid_dim: int,
dec_hid_dim: int,
attention_module: torch.nn.Module,
encoder_is_bidirectional: False
"""
def __init__(self, output_dim, emb_dim, enc_hid_dim, dec_hid_dim, attention_module, encoder_is_bidirectional=False):
super(Decoder, self).__init__()
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.output_dim = output_dim
self.encoder_is_bidirectional = encoder_is_bidirectional
if isinstance(attention_module, nn.Module):
self.attention_module = attention_module
else:
raise ValueError
self.rnn_input_dim = enc_hid_dim + emb_dim # enc_h + dec_emb_dim
if self.encoder_is_bidirectional:
self.rnn_input_dim += enc_hid_dim # 2 * enc_h + dec_emb_dim
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.GRU(
input_size=self.rnn_input_dim,
hidden_size=dec_hid_dim,
bidirectional=False,
batch_first=True,
)
out_input_dim = 2 * dec_hid_dim + emb_dim # hidden + dec_hidden_dim + dec_emb_dim
self.out = nn.Linear(out_input_dim, output_dim)
self.dropout = nn.Dropout(.2)
def forward(self, inp, hidden, encoder_outputs):
"""
Arguments:
inp: 1d tensor with shape (batch_size, )
hidden: 2d tensor with shape (batch_size, dec_hid_dim).
This `hidden` tensor is the hidden state vector from the previous timestep.
encoder_outputs: 3d tensor with shape (batch_size, seq_len, enc_hid_dim).
If encoder_is_bidirectional is True, expects shape (batch_size, seq_len, 2 * enc_hid_dim).
"""
assert inp.dim() == 1
assert hidden.dim() == 2
assert encoder_outputs.dim() == 3
# (batch_size, ) -> (batch_size, 1)
inp = inp.unsqueeze(1)
# (batch_size, 1) -> (batch_size, 1, emb_dim)
embedded = self.embedding(inp)
embedded = self.dropout(embedded)
# attention probabilities; (batch_size, seq_len)
attn_probs = self.attention_module(hidden, encoder_outputs)
# (batch_size, 1, seq_len)
attn_probs = attn_probs.unsqueeze(1)
# (b, 1, s) @ (b, s, enc_hid_dim) -> (b, 1, enc_hid_dim)
weighted = torch.bmm(attn_probs, encoder_outputs)
# (batch_size, 1, emb_dim + enc_hid_dim)
rnn_input = torch.cat((embedded, weighted), dim=2)
# output; (batch_size, 1, dec_hid_dim)
# new_hidden; (1, batch_size, dec_hid_dim)
output, new_hidden = self.rnn(rnn_input, hidden.unsqueeze(0))
embedded = embedded.squeeze(1) # (b, 1, emb) -> (b, emb)
output = output.squeeze(1) # (b, 1, dec_h) -> (b, dec_h)
weighted = weighted.squeeze(1) # (b, 1, dec_h) -> (b, dec_h)
# output; (batch_size, emb + 2 * dec_h) -> (batch_size, output_dim)
output = self.out(torch.cat((output, weighted, embedded), dim=1))
return output, new_hidden.squeeze(0)
# + [markdown] id="1FfbPnfr3IgQ" colab_type="text"
# ## Seq2Seq
# + id="in1zWsUvQZdv" colab_type="code" colab={}
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, device):
super(Seq2Seq, self).__init__()
self.encoder = encoder
self.decoder = decoder
self.device = device
def forward(self, src, trg, teacher_forcing_ratio=.5):
batch_size, max_seq_len = trg.size()
trg_vocab_size = self.decoder.output_dim
# An empty tesnor to store decoder outputs (time index first for indexing)
outputs_shape = (max_seq_len, batch_size, trg_vocab_size)
outputs = torch.zeros(outputs_shape).to(self.device)
encoder_outputs, hidden = self.encoder(src)
# first input to the decoder is '<sos>'
# trg; shape (batch_size, seq_len)
initial_dec_input = output = trg[:, 0] # get first timestep token
for t in range(1, max_seq_len):
output, hidden = self.decoder(output, hidden, encoder_outputs)
outputs[t] = output # Save output for timestep t, for 1 <= t <= max_len
top1_val, top1_idx = output.max(dim=1)
teacher_force = torch.rand(1).item() >= teacher_forcing_ratio
output = trg[:, t] if teacher_force else top1_idx
# Switch batch and time dimensions for consistency (batch_first=True)
outputs = outputs.permute(1, 0, 2) # (s, b, trg_vocab) -> (b, s, trg_vocab)
return outputs
# + [markdown] id="AmnGbjjJ3GrO" colab_type="text"
# ## Build Model
# + id="v1lU0L8UOBB7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="9dc3b6ce-519e-4b12-fdcc-fe3071481ed6" executionInfo={"status": "ok", "timestamp": 1564630235006, "user_tz": -540, "elapsed": 547, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Define encoder
enc = Encoder(
input_dim=INPUT_DIM,
emb_dim=ENC_EMB_DIM,
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
bidirectional=USE_BIDIRECTIONAL
)
print(enc)
# + id="ut6eTqZnORIX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="168f407e-b940-4017-9dd8-9db9751d0473" executionInfo={"status": "ok", "timestamp": 1564630235243, "user_tz": -540, "elapsed": 617, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Define attention layer
attn = Attention(
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
encoder_is_bidirectional=USE_BIDIRECTIONAL
)
print(attn)
# + id="RHmtjt6sOMp6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 181} outputId="95c4e7d0-5f87-4cc2-c7d7-a3429f885a2f" executionInfo={"status": "ok", "timestamp": 1564630243954, "user_tz": -540, "elapsed": 707, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Define decoder
dec = Decoder(
output_dim=OUTPUT_DIM,
emb_dim=DEC_EMB_DIM,
enc_hid_dim=ENC_HID_DIM,
dec_hid_dim=DEC_HID_DIM,
attention_module=attn,
encoder_is_bidirectional=USE_BIDIRECTIONAL
)
print(dec)
# + id="pvUGmhhZOP2d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 326} outputId="74775ab7-c198-4153-d688-c1cd87b90f55" executionInfo={"status": "ok", "timestamp": 1564630245643, "user_tz": -540, "elapsed": 560, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
model = Seq2Seq(enc, dec, device).to(device)
print(model)
# + id="VKaAzVGAOwjw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a9745408-0de9-4053-bc49-01c19acf79e2" executionInfo={"status": "ok", "timestamp": 1564630247860, "user_tz": -540, "elapsed": 543, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters.')
# + [markdown] id="MxqW8sfvPV2z" colab_type="text"
# # 6. Train
# + [markdown] id="Sg3cOfxAPYzV" colab_type="text"
# ## Optimizer
# - Use `optim.Adam` or `optim.RMSprop`.
# + id="J0cJYGriPXc8" colab_type="code" colab={}
optimizer = optim.Adam(model.parameters(), lr=0.001)
#optimizer = optim.RMSprop(model.parameters(), lr=0.01)
# + [markdown] id="wo2OgmdYPjs5" colab_type="text"
# ## Loss function
# + id="zN84sIssPklu" colab_type="code" colab={}
# Padding indices should not be considered when loss is calculated.
PAD_IDX = ENGLISH.vocab.stoi['<pad>']
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
# + [markdown] id="-sdvuh9VP1nC" colab_type="text"
# ## Train function
# + id="N7jbrmD6P2cc" colab_type="code" colab={}
def train(seq2seq_model, iterator, optimizer, criterion, grad_clip=1.0):
seq2seq_model.train()
epoch_loss = .0
for i, batch in enumerate(iterator):
print('.', end='')
src = batch.src
trg = batch.trg
optimizer.zero_grad()
decoder_outputs = seq2seq_model(src, trg, teacher_forcing_ratio=.5)
seq_len, batch_size, trg_vocab_size = decoder_outputs.size() # (b, s, trg_vocab)
# (b-1, s, trg_vocab)
decoder_outputs = decoder_outputs[:, 1:, :]
# ((b-1) * s, trg_vocab)
decoder_outputs = decoder_outputs.contiguous().view(-1, trg_vocab_size)
# ((b-1) * s, )
trg = trg[:, 1:].contiguous().view(-1)
loss = criterion(decoder_outputs, trg)
loss.backward()
# Gradient clipping; remedy for exploding gradients
torch.nn.utils.clip_grad_norm_(seq2seq_model.parameters(), grad_clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="3lSnoKXuWS20" colab_type="text"
# ## Evaluate function
# + id="UOlqFlRTSjR1" colab_type="code" colab={}
def evaluate(seq2seq_model, iterator, criterion):
seq2seq_model.eval()
epoch_loss = 0.
with torch.no_grad():
for i, batch in enumerate(iterator):
print('.', end='')
src = batch.src
trg = batch.trg
decoder_outputs = seq2seq_model(src, trg, teacher_forcing_ratio=0.)
seq_len, batch_size, trg_vocab_size = decoder_outputs.size() # (b, s, trg_vocab)
# (b-1, s, trg_vocab)
decoder_outputs = decoder_outputs[:, 1:, :]
# ((b-1) * s, trg_vocab)
decoder_outputs = decoder_outputs.contiguous().view(-1, trg_vocab_size)
# ((b-1) * s, )
trg = trg[:, 1:].contiguous().view(-1)
loss = criterion(decoder_outputs, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + [markdown] id="K4tw3PJUWVUw" colab_type="text"
# ## Epoch time measure function
# + id="2gI5DjKwPQhZ" colab_type="code" colab={}
def epoch_time(start_time, end_time):
"""Returns elapsed time in mins & secs."""
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + [markdown] id="UtMu_ZQREe1v" colab_type="text"
# ## Train for multiple epochs
# + id="xx4zXJ4JR5lv" colab_type="code" colab={}
NUM_EPOCHS = 50
# + id="kOdqo0ieR9zs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="c7c4258e-5851-442d-919d-038712805442" executionInfo={"status": "ok", "timestamp": 1564634438628, "user_tz": -540, "elapsed": 1628, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
import time
import math
best_dev_loss = float('inf')
for epoch in range(NUM_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion)
dev_loss = evaluate(model, dev_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if dev_loss < best_dev_loss:
best_dev_loss = dev_loss
torch.save(model.state_dict(), './best_model.pt')
print("\n")
print(f"Epoch: {epoch + 1:>02d} | Time: {epoch_mins}m {epoch_secs}s")
print(f"Train Loss: {train_loss:>.4f} | Train Perplexity: {math.exp(train_loss):7.3f}")
print(f"Dev Loss: {dev_loss:>.4f} | Dev Perplexity: {math.exp(dev_loss):7.3f}")
# + [markdown] id="UcQ-BE01W-c-" colab_type="text"
# ## Save last model (overfitted)
# + id="ig5cGvhwXC-b" colab_type="code" colab={}
torch.save(model.state_dict(), './last_model.pt')
# + [markdown] id="F6xofPzNVckv" colab_type="text"
# # 7. Test
# + [markdown] id="r-NR8RlmRQGt" colab_type="text"
# ## Function to convert indices to original text strings
# + id="VhD2yDYOFarQ" colab_type="code" colab={}
def indices_to_text(src_or_trg, lang_field):
assert src_or_trg.dim() == 1, f'{src_or_trg.dim()}' #(seq_len, )
assert isinstance(lang_field, torchtext.data.Field)
assert hasattr(lang_field, 'vocab')
return [lang_field.vocab.itos[t] for t in src_or_trg]
# + [markdown] id="Bbff5Y3LRVQk" colab_type="text"
# ## Function to make predictions
# - Returns a list of examples, where each example is a (src, trg, prediction) tuple.
# + id="OaPhaF_CH6qr" colab_type="code" colab={}
def predict(seq2seq_model, iterator):
seq2seq_model.eval()
out = []
with torch.no_grad():
for i, batch in enumerate(iterator):
src = batch.src
trg = batch.trg
decoder_outputs = seq2seq_model(src, trg, teacher_forcing_ratio=0.)
seq_len, batch_size, trg_vocab_size = decoder_outputs.size() # (b, s, trg_vocab)
# Discard initial decoder input (index = 0)
#decoder_outputs = decoder_outputs[:, 1:, :]
decoder_predictions = decoder_outputs.argmax(dim=-1) # (b, s)
for i, pred in enumerate(decoder_predictions):
out.append((src[i], trg[i], pred))
return out
# + [markdown] id="5UL-AIFORkzq" colab_type="text"
# ## Load best model
# + id="Ep3O8b5ufRtb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="95950354-f108-41ad-cbe1-98f6e6f1d669" executionInfo={"status": "ok", "timestamp": 1564634729498, "user_tz": -540, "elapsed": 1975, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# !ls -al
# + id="rTvAUJTnRFqS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b9ead01a-ddac-40a5-f0b3-0e69da27c7a5" executionInfo={"status": "ok", "timestamp": 1564634730828, "user_tz": -540, "elapsed": 713, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# Load model
model.load_state_dict(torch.load('./best_model.pt'))
# + [markdown] id="rA4amsXERoAI" colab_type="text"
# ## Make predictions
# + id="hnU9YnbYH0RL" colab_type="code" colab={}
# Make prediction
test_predictions = predict(model, dev_iterator)
# + id="yNx9vWU2JgEm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 929} outputId="60eb9a6b-99eb-4281-f596-665056c2a3b3" executionInfo={"status": "ok", "timestamp": 1564634734678, "user_tz": -540, "elapsed": 609, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
for i, prediction in enumerate(test_predictions):
src, trg, pred = prediction
src_text = indices_to_text(src, lang_field=KOREAN)
trg_text = indices_to_text(trg, lang_field=ENGLISH)
pred_text = indices_to_text(pred, lang_field=ENGLISH)
print('source:\n', src_text)
print('target:\n', trg_text)
print('prediction:\n', pred_text)
print('-' * 160)
if i > 5:
break
# + [markdown] id="de77guAiCPgZ" colab_type="text"
# # 8. Download Model
# + id="5yYiqOcuY530" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 163} outputId="7e7825e0-17cb-4e69-96e9-af36d81fa0cb" executionInfo={"status": "ok", "timestamp": 1564634740999, "user_tz": -540, "elapsed": 2515, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
# !ls -al
# + id="DptY_D9qCVTN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1814eefa-3927-43c5-8b16-a33713baaf3c" executionInfo={"status": "ok", "timestamp": 1564634832598, "user_tz": -540, "elapsed": 42356, "user": {"displayName": "\u00ad\uac15\ud604\uad6c[ \ub300\ud559\uc6d0\uc11d\u00b7\ubc15\uc0ac\ud1b5\ud569\uacfc\uc815\uc218\ub8cc\uc5f0\uad6c(\uc7ac\ud559) / \uc0b0\uc5c5\uacbd\uc601\uacf5\ud559\uacfc ]", "photoUrl": "", "userId": "05809067100169424638"}}
from google.colab import files
print('Downloading models...') # Known bug; if using Firefox, a print statement in the same cell is necessary.
files.download('./best_model.pt')
files.download('./last_model.pt')
# + [markdown] id="HuIsoKdxgP0u" colab_type="text"
# # 9. Discussions
# + id="AymW4-HXf8qO" colab_type="code" colab={}
|
colab/NMT-Seq2SeqWithAttention.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reading outputs from E+
# some initial set up
# if you have not installed epp, and only downloaded it
# you will need the following lines
import sys
# pathnameto_eppy = 'c:/eppy'
pathnameto_eppy = '../'
sys.path.append(pathnameto_eppy)
# ## Using titletable() to get at the tables
# So far we have been making changes to the IDF input file.
# How about looking at the outputs.
#
# Energyplus makes nice htmlout files that look like this.
import ex_inits #no need to know this code, it just shows the image below
for_images = ex_inits
for_images.display_png(for_images.html_snippet1) #display the image below
# If you look at the clipping of the html file above, you see tables with data in them. Eppy has functions that let you access of these tables and get the data from any of it's cells.
#
# Let us say you want to find the "Net Site Energy".
#
# This is in table "Site and Source Energy".
#
# The number you want is in the third row, second column and it's value is "47694.47"
#
# Let us use eppy to extract this number
#
# +
from eppy.results import readhtml # the eppy module with functions to read the html
fname = "../eppy/resources/outputfiles/V_7_2/5ZoneCAVtoVAVWarmestTempFlowTable_ABUPS.html" # the html file you want to read
filehandle = open(fname, 'r').read()
htables = readhtml.titletable(filehandle) # reads the tables with their titles
# -
# If you open the python file readhtml.py and look at the function titletable, you can see the function documentation.
#
# It says the following
"""return a list of [(title, table), .....]
title = previous item with a <b> tag
table = rows -> [[cell1, cell2, ..], [cell1, cell2, ..], ..]"""
# The documentation says that it returns a list.
# Let us take a look inside this list.
# Let us look at the first item in the list.
firstitem = htables[0]
print(firstitem)
# Ughh !!! that is ugly. Hard to see what it is.
# Let us use a python module to print it pretty
import pprint
pp = pprint.PrettyPrinter()
pp.pprint(firstitem)
# Nice. that is a little clearer
firstitem_title = firstitem[0]
pp.pprint(firstitem_title)
firstitem_table = firstitem[1]
pp.pprint(firstitem_table)
# How do we get to value of "Net Site Energy".
# We know it is in the third row, second column of the table.
#
# Easy.
thirdrow = firstitem_table[2] # we start counting with 0. So 0, 1, 2 is third row
print(thirdrow)
thirdrow_secondcolumn = thirdrow[1]
thirdrow_secondcolumn
# the text from the html table is in unicode.
# That is why you see that weird 'u' letter.
#
# Let us convert it to a floating point number
net_site_energy = float(thirdrow_secondcolumn)
net_site_energy
# Let us have a little fun with the tables.
#
# Get the titles of all the tables
alltitles = [htable[0] for htable in htables]
alltitles
# Now let us grab the tables with the titles "Building Area" and "Site to Source Energy Conversion Factors"
# twotables = [htable for htable in htables if htable[0] in ["Building Area", "Site to Source Energy Conversion Factors"]]
# twotables
# Let us leave readtables for now.
#
# It gives us the basic functionality to read any of the tables in the html output file.
# ## Fast HTML table file read
# The function`readhtml.titletable()` will be slow with extremeley large files. If you are dealing with a very large file use the following functions
from eppy.results import fasthtml
fname = "../eppy/resources/outputfiles/V_7_2/5ZoneCAVtoVAVWarmestTempFlowTable_ABUPS.html" # the html file you want to read
filehandle = open(fname, 'r') # get a file handle to the html file
firsttable = fasthtml.tablebyindex(filehandle, 0)
pp.pprint(firstitem)
filehandle = open(fname, 'r') # get a file handle to the html file
namedtable = fasthtml.tablebyname(filehandle, "Site and Source Energy")
pp.pprint(namedtable)
# - You can read only one table at a time
# - You need to open the file each time you call the function. The function will close the file.
# ## Using lines_table() to get at the tables
# We have been using titletable() to get at the tables. There is a constraint using function titletable(). Titletable() assumes that there is a unique title (in HTML bold) just above the table. It is assumed that this title will adequetly describe the table. This is true in most cases and titletable() is perfectly good to use. Unfortuntely there are some tables that do not follow this rule. The snippet below shows one of them.
import ex_inits #no need to know this code, it just shows the image below
for_images = ex_inits
for_images.display_png(for_images.html_snippet2) # display the image below
# Notice that the HTML snippet shows a table with three lines above it. The first two lines have information that describe the table. We need to look at both those lines to understand what the table contains. So we need a different function that will capture all those lines before the table. The funtion lines_table() described below will do this.
#
# +
from eppy.results import readhtml # the eppy module with functions to read the html
fname = "../eppy/resources/outputfiles/V_8_1/ASHRAE30pct.PI.Final11_OfficeMedium_STD2010_Chicago-baseTable.html" # the html file you want to read
filehandle = open(fname, 'r').read() # get a file handle to the html file
ltables = readhtml.lines_table(filehandle) # reads the tables with their titles
# -
# The html snippet shown above is the last table in HTML file we just opened. We have used lines_table() to read the tables into the variable ltables. We can get to the last table by ltable[-1]. Let us print it and see what we have.
#
import pprint
pp = pprint.PrettyPrinter()
pp.pprint(ltables[-1])
# We can see that ltables has captured all the lines before the table. Let us make our code more explicit to see this
# +
last_ltable = ltables[-1]
lines_before_table = last_ltable[0]
table_itself = last_ltable[-1]
pp.pprint(lines_before_table)
# -
# We found this table the easy way this time, because we knew it was the last one. How do we find it if we don't know where it is in the file ? Python comes to our rescue :-) Let assume that we want to find the table that has the following two lines before it.
#
# - Report: FANGER DURING COOLING AND ADAPTIVE COMFORT
# - For: PERIMETER_MID_ZN_4
#
line1 = 'Report: FANGER DURING COOLING AND ADAPTIVE COMFORT'
line2 = 'For: PERIMETER_MID_ZN_4'
#
# check if those two lines are before the table
line1 in lines_before_table and line2 in lines_before_table
# find all the tables where those two lines are before the table
[ltable for ltable in ltables
if line1 in ltable[0] and line2 in ltable[0]]
# That worked !
#
# What if you want to find the words "FANGER" and "PERIMETER_MID_ZN_4" before the table. The following code will do it.
#
# +
# sample code to illustrate what we are going to do
last_ltable = ltables[-1]
lines_before_table = last_ltable[0]
table_itself = last_ltable[-1]
# join lines_before_table into a paragraph of text
justtext = '\n'.join(lines_before_table)
print(justtext)
# -
"FANGER" in justtext and "PERIMETER_MID_ZN_4" in justtext
# Let us combine the this trick to find the table
[ltable for ltable in ltables
if "FANGER" in '\n'.join(ltable[0]) and "PERIMETER_MID_ZN_4" in '\n'.join(ltable[0])]
# ## Extracting data from the tables
# The tables in the HTML page in general have text in the top header row. The first vertical row has text. The remaining cells have numbers. We can identify the numbers we need by looking at the labelin the top row and the label in the first column. Let us construct a simple example and explore this.
# ignore the following three lines. I am using them to construct the table below
from IPython.display import HTML
atablestring = '<TABLE cellpadding="4" style="border: 1px solid #000000; border-collapse: collapse;" border="1">\n <TR>\n <TD> </TD>\n <TD>a b</TD>\n <TD>b c</TD>\n <TD>c d</TD>\n </TR>\n <TR>\n <TD>x y</TD>\n <TD>1</TD>\n <TD>2</TD>\n <TD>3</TD>\n </TR>\n <TR>\n <TD>y z</TD>\n <TD>4</TD>\n <TD>5</TD>\n <TD>6</TD>\n </TR>\n <TR>\n <TD>z z</TD>\n <TD>7</TD>\n <TD>8</TD>\n <TD>9</TD>\n </TR>\n</TABLE>'
HTML(atablestring)
# This table is actually in the follwoing form:
atable = [["", "a b", "b c", "c d"],
["x y", 1, 2, 3 ],
["y z", 4, 5, 6 ],
["z z", 7, 8, 9 ],]
# We can see the labels in the table. So we an look at row "x y" and column "c d". The value there is 3
# right now we can get to it by saying atable[1][3]
print(atable[1][3])
# readhtml has some functions that will let us address the values by the labels. We use a structure from python called named tuples to do this. The only limitation is that the labels have to be letters or digits. Named tuples does not allow spaces in the labels. We could replace the space with an underscore ' _ '. So "a b" will become "a_b". So we can look for row "x_y" and column "c_d". Let us try this out.
from eppy.results import readhtml
h_table = readhtml.named_grid_h(atable)
print(h_table.x_y.c_d)
# We can still get to the value by index
print(h_table[0][2])
# Note that we used atable[1][3], but here we used h_table[0][2]. That is because h_table does not count the rows and columns where the labels are.
# We can also do the following:
print(h_table.x_y[2])
# or
print(h_table[0].c_d)
# Wow … that is pretty cool. What if we want to just check what the labels are ?
print(h_table._fields)
# That gives us the horizontal lables. How about the vertical labels ?
h_table.x_y._fields
# There you go !!!
# How about if I want to use the labels differently ? Say I want to refer to the row first and then to the column. That woul be saying table.c_d.x_y. We can do that by using a different function
v_table = readhtml.named_grid_v(atable)
print(v_table.c_d.x_y)
# And we can do the following
print(v_table[2][0])
print(v_table.c_d[0])
print(v_table[2].x_y)
# Let us try to get the numbers in the first column and then get their sum
v_table.a_b
# Look like we got the right column. But not in the right format. We really need a list of numbers
[cell for cell in v_table.a_b]
# That looks like waht we wanted. Now let us get the sum
values_in_first_column = [cell for cell in v_table.a_b]
print(values_in_first_column)
print(sum(values_in_first_column)) # sum is a builtin function that will sum a list
# To get the first row we use the variable h_table
values_in_first_row = [cell for cell in h_table.x_y]
print(values_in_first_row)
print(sum(values_in_first_row))
# ## Fast HTML table file read
# To read the html table files you would usually use the functions described in [Reading outputs from E+](./Outputs_Tutorial.html). For instance you would use the functions as shown below.
# +
from eppy.results import readhtml # the eppy module with functions to read the html
import pprint
pp = pprint.PrettyPrinter()
fname = "../eppy/resources/outputfiles/V_7_2/5ZoneCAVtoVAVWarmestTempFlowTable_ABUPS.html" # the html file you want to read
html_doc = open(fname, 'r').read()
htables = readhtml.titletable(html_doc) # reads the tables with their titles
firstitem = htables[0]
pp.pprint(firstitem)
# -
# `titletable` reads all the tables in the HTML file. With large E+ models, this file can be extremeely large and `titletable` will load all the tables into memory. This can take several minutes. If you are trying to get one table or one value from a table, waiting several minutes for you reseult can be exessive.
#
# If you know which table you are looking for, there is a faster way of doing this. We used index=0 in the above example to get the first table. If you know the index of the file you are looking for, you can use a faster function to get the table as shown below
from eppy.results import fasthtml
fname = "../eppy/resources/outputfiles/V_7_2/5ZoneCAVtoVAVWarmestTempFlowTable_ABUPS.html" # the html file you want to read
filehandle = open(fname, 'r') # get a file handle to the html file
firsttable = fasthtml.tablebyindex(filehandle, 0)
pp.pprint(firstitem)
# You can also get the table if you know the title of the table. This is the **bold** text just before the table in the HTML file. The title of our table is **Site and Source Energy**. The function `tablebyname` will get us the table.
filehandle = open(fname, 'r') # get a file handle to the html file
namedtable = fasthtml.tablebyname(filehandle, "Site and Source Energy")
pp.pprint(namedtable)
# Couple of things to note here:
#
# - We have to open the file again using `filehandle = open(fname, 'r')`
# - This is because both `tablebyname` and `tablebyindex` will close the file once they are done
# - Some tables do not have a **bold title** just before the table. `tablebyname` will not work for those functions
|
docs/Outputs_Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Federated FedProx PyTorch MNIST Tutorial
# The only difference between this notebook and Federated_Pytorch_MNIST_Tutorial.ipynb is overriding of the `train_epoch` function in model definition. [See details](#FedProx)
#
#Install dependencies if not already installed
# !pip install torch torchvision
# +
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
import torchvision.transforms as transforms
import openfl.native as fx
from openfl.federated import FederatedModel,FederatedDataSet
# -
# After importing the required packages, the next step is setting Up our openfl workspace. To do this, simply run the `fx.init()` command as follows:
#Setup default workspace, logging, etc.
fx.init('torch_cnn_mnist')
# Now we are ready to define our dataset and model to perform federated learning on. The dataset should be composed of a numpy arrayWe start with a simple fully connected model that is trained on the MNIST dataset.
# +
def one_hot(labels, classes):
return np.eye(classes)[labels]
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.MNIST(root='./data', train=True,
download=True, transform=transform)
train_images,train_labels = trainset.train_data, np.array(trainset.train_labels)
train_images = torch.from_numpy(np.expand_dims(train_images, axis=1)).float()
train_labels = one_hot(train_labels,10)
validset = torchvision.datasets.MNIST(root='./data', train=False,
download=True, transform=transform)
valid_images,valid_labels = validset.test_data, np.array(validset.test_labels)
valid_images = torch.from_numpy(np.expand_dims(valid_images, axis=1)).float()
valid_labels = one_hot(valid_labels,10)
# -
# # FedProx
from openfl.utilities.optimizers.torch import FedProxOptimizer
# +
feature_shape = train_images.shape[1]
classes = 10
fl_data = FederatedDataSet(train_images,train_labels,valid_images,valid_labels,batch_size=32,num_classes=classes)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(1, 16, 3)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(16, 32, 3)
self.fc1 = nn.Linear(32 * 5 * 5, 32)
self.fc2 = nn.Linear(32, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(x.size(0),-1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return F.log_softmax(x, dim=1)
def train_epoch(self, batch_generator):
from openfl.federated.task import PyTorchTaskRunner
self.optimizer.set_old_weights([p for p in self.parameters()])
return PyTorchTaskRunner.train_epoch(self, batch_generator)
optimizer = lambda x: FedProxOptimizer(x, lr=1e-3, mu=0.1)
def cross_entropy(output, target):
"""Binary cross-entropy metric
"""
return F.binary_cross_entropy_with_logits(input=output,target=target.float())
# +
#Create a federated model using the pytorch class, lambda optimizer function, and loss function
fl_model = FederatedModel(build_model=Net,optimizer=optimizer,loss_fn=cross_entropy,data_loader=fl_data)
# -
# The `FederatedModel` object is a wrapper around your Keras, Tensorflow or PyTorch model that makes it compatible with openfl. It provides built in federated training and validation functions that we will see used below. Using it's `setup` function, collaborator models and datasets can be automatically defined for the experiment.
collaborator_models = fl_model.setup(num_collaborators=2)
collaborators = {'one':collaborator_models[0],'two':collaborator_models[1]}#, 'three':collaborator_models[2]}
# +
#Original MNIST dataset
print(f'Original training data size: {len(train_images)}')
print(f'Original validation data size: {len(valid_images)}\n')
#Collaborator one's data
print(f'Collaborator one\'s training data size: {len(collaborator_models[0].data_loader.X_train)}')
print(f'Collaborator one\'s validation data size: {len(collaborator_models[0].data_loader.X_valid)}\n')
#Collaborator two's data
print(f'Collaborator two\'s training data size: {len(collaborator_models[1].data_loader.X_train)}')
print(f'Collaborator two\'s validation data size: {len(collaborator_models[1].data_loader.X_valid)}\n')
#Collaborator three's data
#print(f'Collaborator three\'s training data size: {len(collaborator_models[2].data_loader.X_train)}')
#print(f'Collaborator three\'s validation data size: {len(collaborator_models[2].data_loader.X_valid)}')
# -
# We can see the current plan values by running the `fx.get_plan()` function
#Get the current values of the plan. Each of these can be overridden
import json
print(json.dumps(fx.get_plan(), indent=4, sort_keys=True))
# Now we are ready to run our experiment. If we want to pass in custom plan settings, we can easily do that with the `override_config` parameter
#Run experiment, return trained FederatedModel
final_fl_model = fx.run_experiment(collaborators,{'aggregator.settings.rounds_to_train':5})
#Save final model
final_fl_model.save_native('final_pytorch_model')
|
openfl-tutorials/Federated_FedProx_PyTorch_MNIST_Tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Publishing SDs, Shapefiles and CSVs
#
# Publishing your data can be accomplished in two simple steps:
# 1. Add the local data as an item to the portal
# 2. Call the publish() method on the item
#
# This sample notebook shows how different types of GIS datasets can be added to the GIS, and published as web layers.
from IPython.display import display
from arcgis.gis import GIS
import os
gis = GIS('Home')
# In order to have the codes below run smoothly, use the pre-requisite cells as in cell[2] to delete existing .zip, .sd, or services from the `gis` content, and in cell [3] to delete existing folder.
# +
def delete_existing_items(item_types, name_list):
for current_item_type in item_types:
for file_name in name_list:
search_result = gis.content.search(query=file_name, item_type=current_item_type)
if len(search_result) > 0:
for item in search_result:
item.delete()
print("Deleted existing " + current_item_type + ": ", item)
item_types = ["Service Definition", "Feature Layer Collection", "Map Service"]
name_list = ["Nursing_home_locations", "NewPy_WTL_test_SingleLayerBuildCache"]
delete_existing_items(item_types, name_list)
item_types = ["Shapefile", "Feature Layer Collection"]
name_list = ["power_pedestals_2012"]
delete_existing_items(item_types, name_list)
item_types = ["CSV", "Feature Layer Collection"]
name_list = ["Chennai_precipitation"]
delete_existing_items(item_types, name_list)
# +
def delete_existing_folder(folder_name):
try:
return gis.content.delete_folder(folder=folder_name)
except:
return False
my_folder_name = "Rainfall Data"
delete_existing_folder(my_folder_name) # returns True if folder exists, or False if non-exist
# -
#
# # Publish all the service definition files in a folder
#
# The sample below lists all the service definition (.sd) files in a data directory and publishes them as web layers. To publish a service definition file, we first add the .sd file to the Portal, and then call the publish() method:
# +
# path relative to this notebook
data_dir = "data/publishing_sd_shapefiles_and_csv/"
#Get list of all files
file_list = os.listdir(data_dir)
#Filter and get only .sd files
sd_file_list = [x for x in file_list if x.endswith(".sd")]
print("Number of .sd files found: " + str(len(sd_file_list)))
# -
# Loop through each file and publish it as a service
for current_sd_file in sd_file_list:
item = gis.content.add({}, data_dir + current_sd_file) # .sd file is uploaded and a .sd file item is created
if "BuildCache" not in current_sd_file:
published_item = item.publish() # .sd file item is published and a web layer item is created
else:
published_item = item.publish(build_initial_cache=True) # publish as hosted tile layer with "build cache" enabled
display(published_item)
# In the example shown above, one .sd file produced a web feature layer and another produced a web tile layer
# # Publish a feature service from a shapefile and update the item information
#
# To publish a shapefile, we first add the zipped shapefile to the Portal as an item, then call publish() method on the item to create a web layer. Often times, your shape files or service definitions may not contain the metadata you want to show on the portal item. This sample demonstrates how you can update those properties after publishing a web layer.
data = "data/power_pedestals_2012.zip"
shpfile = gis.content.add({}, data)
shpfile
published_service = shpfile.publish()
display(published_service)
# The web layer item has minimal information and a default thumbnail.
#
# ### Update the layer item's metadata
# To update the metadata and set the thumbnail, use the update() method on the web layer's item obtained during publishing.
thumbnail_path = "data/power_pedestals_thumbnail.PNG"
item_properties = {"snippet":"""This dataset was collected from Utah DOT open data portal.
Source URL: <a href="http://udot.uplan.opendata.arcgis.com/
datasets/a627bb128ac44767832402f7f9bde909_10">http://udot.uplan.opendata.arcgis.com/
datasets/a627bb128ac44767832402f7f9bde909_10</a>""",
"title":"Locations of power pedestals collected in 2012",
"tags":"opendata"}
published_service.update(item_properties, thumbnail=thumbnail_path)
display(published_service)
# # Publish a CSV file and move it into a folder
#
# To publish a CSV file, we first add the .csv file to the Portal, and then call the publish() method to publish it as a layer. Once published, we create a destination folder on the server and then move the published items into that folder
csv_file = 'data/Chennai_precipitation.csv'
csv_item = gis.content.add({}, csv_file)
display(csv_item)
# The csv file used in this sample has a column titled `LOCATION` containing place names in text. During the publishing process we specify this column as an `address_fields` parameter. The server geocodes these locations to create point features for the web layer.
csv_lyr = csv_item.publish(None, {"Address":"LOCATION"})
display(csv_lyr)
# ### Create a new folder for the items
# The `create_folder()` from `GIS.content` can be used to create a new folder. Once created, the `move()` of the `Item` can be used to move the items into the folder.
# create a new folder called 'Rainfall Data'
new_folder_details = gis.content.create_folder(my_folder_name)
print(new_folder_details)
# +
# move both the csv_item and csv_lyr items into this new folder
csv_item.move(new_folder_details) # Here you could either pass name of the folder or the dictionary
# returned from create_folder() or folders property on a User object
csv_lyr.move(new_folder_details)
# -
# Now that the items are moved, we can request for the item's `ownerFolder` property and ensure it matches the `id` of the folder created in the previous step
print(csv_lyr.ownerFolder)
|
samples/05_content_publishers/publishing_sd_shapefiles_and_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Import exiobase3 in brightway2
# by <NAME>, Jan19
import pandas as pd
import numpy as np
from pyxlsb import open_workbook as open_xlsb
from brightway2 import *
projects.set_current('test_exio3_import_simple')
bw2setup() # need to import biosphere3
# +
#this takes some minutes, but seems the only way to import xlsb files in python
HIO_Arr = []
with open_xlsb("exiobase_3.3.15_hsut_2011/Exiobase_MR_HIOT_2011_v3_3_15_by_prod_tech.xlsb") as wb:
# Read the sheet to array first and convert to pandas first for quick access
with wb.get_sheet(2) as sheet:
for row in sheet.rows(sparse = True):
#print(row[0])
vals = [item.v for item in row]
HIO_Arr.append(vals)
HIO = pd.DataFrame(HIO_Arr)
# +
# create some unique activity IDs ('country-code1-code2', e.g. 'AU-c01.i-P_CATL')
exio_IDs = HIO.iloc[4:, 0].values + "-" + HIO.iloc[4:, 2].values + "-" + HIO.iloc[4:, 3].values
n_act = len(exio_IDs)
# other metadata
exio_countries = HIO.iloc[4:, 0].values # lists
exio_names = HIO.iloc[4:, 1].values
exio_codes1 = HIO.iloc[4:, 2].values
exio_codes2 = HIO.iloc[4:, 3].values
exio_units = HIO.iloc[4:, 4].values
# just the simplest IO table with only column names and row names, used just below for filtering of zero values
exio_data = HIO.iloc[4:, 5:] # pandas dataframe
exio_data.columns = exio_IDs
exio_data.index = exio_IDs
exio_data.head()
# -
# filter out the zero values in the database
d = {c : exio_data.loc[exio_data[c] !=0 , c] for c in exio_IDs } # creates a dict with name, df with values != 0
db_name = 'exio3HIO'
if db_name in databases:
del databases[db_name] #just to make sure there isnt another one
# +
# Create the database as a dict (didn't take long).
data = [{
'name': exio_names[i],
'code': exio_IDs[i],
'key': (db_name, exio_IDs[i]),
'exiobase_code1': exio_codes1[i],
'exiobase_code2': exio_codes2[i],
'database': db_name,
'location': exio_countries[i],
'unit': exio_units[i],
'type': 'process',
'exchanges': [{
'input': (db_name, d[exio_IDs[i]].index[j]),
'amount': d[exio_IDs[i]][j],
'type': 'technosphere'
}
for j in range(d[exio_IDs[i]].shape[0])]
} for i in range(n_act)]
exio_d = {obj['key']: obj for obj in data}
# -
# This takes time! 15 min on my mac
# write the entire IO table
exio3 = Database(db_name)
exio3.write(exio_d)
# check if it's alright
myact = exio3.get('AU-c01.i-P_CATL')
for exc in list(myact.exchanges())[0:4]:
print(exc)
# seems alright
# +
# Import principal production vector from xlsb file (fast)
PPV_Arr = []
with open_xlsb("exiobase_3.3.15_hsut_2011/Exiobase_MR_HIOT_2011_v3_3_15_by_prod_tech.xlsb") as wb:
# Read the sheet to array first and convert to pandas first for quick access
with wb.get_sheet(3) as sheet:
for row in sheet.rows(sparse = True):
#print(row[0])
vals = [item.v for item in row]
PPV_Arr.append(vals)
PPV = pd.DataFrame(PPV_Arr)
p_prod = PPV.iloc[5,1:] # just the numeric values
p_prod.index = exio_IDs
# -
# set the principal production as reference flow for each activity (type = production)
for act in exio3:
exc = act.new_exchange(input = (db_name, act['code']), amount = p_prod[act['code']], unit = act['unit'], type = 'production')
exc.save()
# check if it's alright
myact = exio3.get('AU-c01.i-P_CATL')
for exc in list(myact.exchanges()):
if exc['type'] == 'production':
print(exc)
# seems alright
# +
# import data from the extensions file, but just the emissions for this test
ArrExt_emiss = []
with open_xlsb('exiobase_3.3.15_hsut_2011/MR_HSUT_2011_v3_3_15_extensions.xlsb') as wb:
# Read the sheet to array first and convert to pandas first for quick access
with wb.get_sheet('Emiss_act') as sheet:
for row in sheet.rows(sparse = True):
#print(row[0])
vals = [item.v for item in row]
ArrExt_emiss.append(vals)
Ext_emiss = pd.DataFrame(ArrExt_emiss)
Ext_emiss.head(6)
# -
# I have only manually found these three ones for now, will have to do the same work for the remaining ones
emission_codes = {'Carbon dioxide, fossil': 'aa7cac3a-3625-41d4-bc54-33e2cf11ec46',
'N2O':'afd6d670-bbb0-4625-9730-04088a5b035e',
'CH4':'70ef743b-3ed5-4a6d-b192-fb6d62378555'}
# Again some formatting necessary for the next step
Ext_emiss_s = Ext_emiss.iloc[4:7,3:]
Ext_emiss_s.columns = [exio_IDs]
Ext_emiss_s.index = list(emission_codes.values())
Ext_emiss_s.head()
# Add biosphere3 flows to the database
for act in exio3:
for i in range(Ext_emiss_s[act['code']].shape[0]):
exc = act.new_exchange(input = ('biosphere3', Ext_emiss_s[act['code']].index[i]), amount = float(Ext_emiss_s[act['code']].iloc[i].values)*1000, type = 'biosphere')
exc.save()
# check if it's alright
myact = exio3.get('AU-c01.i-P_CATL')
for exc in list(myact.exchanges()):
if exc['type'] == 'biosphere':
print(exc)
# Seems alright
# now try some calculations
functional_unit = {exio3.get('AU-c01.i-P_CATL'): 1}
mymethod = ('IPCC 2013', 'climate change', 'GWP 100a')
lca = LCA(functional_unit, mymethod)
lca.lci()
lca.lcia()
print(lca.score)
inventory = pd.DataFrame(lca.inventory.toarray())
sum(inventory.iloc[0,:].values) # 4.7 tons of CO2 emissions for 1 ton of cattle farming in Australia
# +
# end
# -
backup_project_directory('test_exio3_import_simple')
|
Import_exio_clean.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''earthy_test'': conda)'
# name: python3
# ---
# # W2 - Accessibility (Distance Fields)
#
# In this workshop we will learn the foundations to quantitatively approach spatial accessibility. We will learn about distance fields, construct a euclidean distance field, and construct a manifold distance field.
# ## 0. Initialization
#
# ### 0.1 Importing the packages
# +
import os
import topogenesis as tg
import pyvista as pv
import trimesh as tm
import numpy as np
# pv.set_jupyter_backend("ipyvtklink")
# convert mesh to pv_mesh
def tri_to_pv(tri_mesh):
faces = np.pad(tri_mesh.faces, ((0, 0),(1,0)), 'constant', constant_values=3)
pv_mesh = pv.PolyData(tri_mesh.vertices, faces)
return pv_mesh
# -
# ### 0.2 import meshes
# +
envelope_path = os.path.relpath('../data/compulsory_envelope.obj')
context_path = os.path.relpath('../data/immediate_context.obj')
# load the mesh from file
envelope_mesh = tm.load(envelope_path)
context_mesh = tm.load(context_path)
# Check if the mesh is watertight
print(envelope_mesh.is_watertight)
print(context_mesh.is_watertight)
# +
# initiating the plotter
p = pv.Plotter(notebook=True)
# adding the meshes
p.add_mesh(tri_to_pv(envelope_mesh), color='#abd8ff')
p.add_mesh(tri_to_pv(context_mesh), color='#aaaaaa')
# plotting
# p.show()
# -
# ### 0.3 Importing the Envelope Lattice
# loading the lattice from csv
lattice_path = os.path.relpath('../data/voxelized_envelope.csv')
envelope_lattice = tg.lattice_from_csv(lattice_path)
# +
# initiating the plotter
p = pv.Plotter()
# fast visualization of the lattice
envelope_lattice.fast_vis(p)
# adding the meshes
p.add_mesh(tri_to_pv(context_mesh), color='#aaaaaa')
# plotting
# p.show()
# -
# ### 0.4 Importing the Street Points
# import the streetnetwork as a point cloud
street_pc = tg.cloud_from_csv("../data/main_street_points.csv")
# +
# initiating the plotter
p = pv.Plotter()
# fast visualization of the lattice
envelope_lattice.fast_vis(p)
# fast visualization of the point cloud
street_pc.fast_notebook_vis(p)
# adding the meshes
p.add_mesh(tri_to_pv(context_mesh), color='#aaaaaa')
# plotting
# p.show()
# -
# ## 1. Euclidean Distance Lattice
#
# ### 1.1 Distance Matrix
# +
# extracting the centroid of all voxels
env_cens = envelope_lattice.centroids_threshold(-1)
# initializing the distance matrix
dist_m = []
# for each voxel ...
for voxel_cen in env_cens:
# initializing the distance vector (per each voxel)
dist_v = []
# for each street point ...
for street_point in street_pc:
# find the difference vector
diff = voxel_cen - street_point
# raise the components to the power of two
diff_p2 = diff**2
# sum the components
diff_p2s = diff_p2.sum()
# compute the square root
dist = diff_p2s**0.5
# add the distance to the distance vector
dist_v.append(dist)
# add the distance vector to the distance matrix
dist_m.append(dist_v)
# change the distance matrix type, from list to array
dist_m = np.array(dist_m)
"""
st_m_shape = (env_cens.shape[0], street_pc.shape[0], 3)
st_m = np.broadcast_to(street_pc, st_m_shape)
ep_m_shape = (street_pc.shape[0], env_cens.shape[0], 3)
ep_m = np.broadcast_to(env_cens, ep_m_shape).transpose((1,0,2))
dist_m = np.linalg.norm(st_m - ep_m, axis=2)
"""
# -
# ### 1.2 Distance to Closest Street Point
# find the distance to the closest street point for each voxel
min_dist = dist_m.min(axis=1)
# convert the minimum distance list to a lattice
street_eu_distance_lattice = tg.to_lattice(min_dist.reshape(envelope_lattice.shape), envelope_lattice)
# zero the value of the exterior voxels
envelope_eu_dist_lattice = street_eu_distance_lattice * envelope_lattice
# +
# initiating the plotter
p = pv.Plotter()
l = envelope_eu_dist_lattice * envelope_lattice
# remapping
l = 250 * (l - l.min()) / l.max()
# Create the spatial reference
grid = pv.UniformGrid()
# Set the grid dimensions: shape because we want to inject our values
grid.dimensions = l.shape
# The bottom left corner of the data set
grid.origin = l.minbound
# These are the cell sizes along each axis
grid.spacing = l.unit
# Add the data values to the cell data
grid.point_arrays["Distance"] = l.flatten(order="F") # Flatten the Lattice
# adding the meshes
p.add_mesh(tri_to_pv(context_mesh), opacity=0.1, style='wireframe')
# fast visualization of the point cloud
street_pc.fast_notebook_vis(p)
# adding the volume
opacity = np.array([0,0.6,0.6,0.6,0.6,0.6,0.6]) * 1.5
p.add_volume(grid, cmap="coolwarm", opacity=opacity, shade=True, show_scalar_bar=False)
# plotting
p.show(use_ipyvtk=True)
# -
# ## 2 Manifold Distance Lattice
#
# ### 2.1 Selecting the Closest Voxels
# selecting the closest voxels by setting a threshold
street_connection_lattice = (0 < envelope_eu_dist_lattice) * (envelope_eu_dist_lattice < 12)
# ### 2.2. The Stencil
# +
# creating neighborhood definition
stencil = tg.create_stencil("von_neumann", 1, 1)
# setting the center to zero
stencil.set_index([0,0,0], 0)
print(stencil)
# -
# ### 2.3 Initializing the Manifold Distance Lattice
# +
# retrieve the neighbour list of each cell
neighs = street_connection_lattice.find_neighbours(stencil)
# set the maximum distance to sum of the size of the lattice in all dimensions.
max_dist = np.sum(street_connection_lattice.shape)
# initialize the street network distance lattice with all the street cells as 0, and all other cells as maximum distance possible
mn_dist_lattice = 1 - street_connection_lattice
mn_dist_lattice[mn_dist_lattice==1] = max_dist
# flatten the distance lattice for easy access
mn_dist_lattice_flat = mn_dist_lattice.flatten()
# flatten the envelope lattice
env_lat_flat = envelope_lattice.flatten()
# -
# ### 2.4 Breadth-First Traversal
# +
# main loop for breath-first traversal
for i in range(1, max_dist):
# find the neighbours of the previous step
next_step = neighs[mn_dist_lattice_flat == i - 1]
# find the unique neighbours
next_unq_step = np.unique(next_step.flatten())
# check if the neighbours of the next step are inside the envelope
validity_condition = env_lat_flat[next_unq_step]
# select the valid neighbours
next_valid_step = next_unq_step[validity_condition]
# make a copy of the lattice to prevent overwriting in the memory
mn_nex_dist_lattice_flat = np.copy(mn_dist_lattice_flat)
# set the next step cells to the current distance
mn_nex_dist_lattice_flat[next_valid_step] = i
# find the minimum of the current distance and previous distances to avoid overwriting previous steps
mn_dist_lattice_flat = np.minimum(mn_dist_lattice_flat, mn_nex_dist_lattice_flat)
# check how many of the cells have not been traversed yet
filled_check = mn_dist_lattice_flat * env_lat_flat == max_dist
# if all the cells have been traversed, break the loop
if filled_check.sum() == 0:
print(i)
break
# reshape and construct a lattice from the street network distance list
mn_dist_lattice = mn_dist_lattice_flat.reshape(mn_dist_lattice.shape)
# +
# the commented code here is the equivalent of the same process but without the assumption that we are working on a regular grid. Producing the distance lattice through this algorithm will take several hours. (Q+) Can you show why these two approaches are equivalent? Can you explain why the second algorithm takes more time?
"""
# find the number of all voxels
vox_count = street_connection_lattice.size
# initialize the adjacency matrix
adj_mtrx = np.zeros((vox_count,vox_count))
# Finding the index of the available voxels in avail_lattice
avail_index = np.array(np.where(street_connection_lattice == 1)).T
# fill the adjacency matrix using the list of all neighbours
for vox_loc in avail_index:
# find the 1D id
vox_id = np.ravel_multi_index(vox_loc, street_connection_lattice.shape)
# retrieve the list of neighbours of the voxel based on the stencil
vox_neighs = street_connection_lattice.find_neighbours_masked(stencil, loc = vox_loc)
# iterating over the neighbours
for neigh in vox_neighs:
# setting the entry to one
adj_mtrx[vox_id, neigh] = 1.0
# construct the graph
g = nx.from_numpy_array(adj_mtrx)
# compute the distance of all voxels to all voxels using floyd warshal algorithm
dist_mtrx = nx.floyd_warshall_numpy(g)
"""
# +
# set the lattice to be visualized
l = mn_dist_lattice * envelope_lattice
# remapping
l = 250 * (l - l.min()) / l.max()
# initiating the plotter
p = pv.Plotter(notebook=True)
# Create the spatial reference
grid = pv.UniformGrid()
# Set the grid dimensions: shape because we want to inject our values
grid.dimensions = l.shape
# The bottom left corner of the data set
grid.origin = l.minbound
# These are the cell sizes along each axis
grid.spacing = l.unit
# Add the data values to the cell data
grid.point_arrays["Street Access"] = l.flatten(order="F") # Flatten the Lattice
# adding the volume
opacity = np.array([0,0.6,0.6,0.6,0.6,0.6,0.6]) * 1.5
p.add_volume(grid, cmap="coolwarm", opacity=opacity, shade=True, show_scalar_bar=False)
# plotting
# p.show(use_ipyvtk=True)
# -
#remap
str_acc_lattice = mn_dist_lattice * envelope_lattice * 1.0
str_acc_lattice /= str_acc_lattice.max()
str_acc_lattice = 1 - str_acc_lattice
# ## 3 Save Lattice into a CSV
# +
# save the sun access latice to csv
csv_path = os.path.relpath('../data/str_acc.csv')
str_acc_lattice.to_csv(csv_path)
# -
# ## Credits
__author__ = "<NAME>"
__license__ = "MIT"
__version__ = "1.0"
__url__ = "https://github.com/shervinazadi/earthy_workshops"
__summary__ = "Earthy Design Studio"
|
notebooks/w2_distance_fields.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Adding New Workloads
#
# Adding new workloads into benchmark-wrapper is a fairly straightforward process, but requires a bit of work. This page tracks all the changes that a user needs to make when adding in a new benchmark.
#
# <div class="alert alert-info">
#
# benchmark-wrapper is currently undergoing a re-write, meaning a lot of change is happening pretty quickly. The information on this page is relevant to the new modifications which change the way in which benchmarks should be added. It may not be consistent to the way in which existing benchmarks are developed.
# </div>
#
# <div class="alert alert-warning">
#
# This page is written within a Jupyter Notebook, however running benchmark-wrapper from within a Jupyter Notebook is not tested nor supported.
# </div>
#
# ## Step Zero: Prep
#
# A Benchmark within benchmark-wrapper is essentially just a Python module which handles setting up, running, parsing and tearing down a benchmark. To create our benchmark, we need to understand the following items:
#
# 1. What is the human-readable, camel-case-able string name for our benchmark?
# 1. What arguments does the benchmark wrapper need from the user?
# 1. Are there any setup tasks that our benchmark wrapper needs to perform?
# 1. How do we run our benchmark?
# 1. Are there any cleanup tasks that our benchmark wrapper needs to perform?
# 1. What data should the benchmark export?
#
# In this example, we'll create a new benchmark wrapper that does a ping test against a list of given hostnames and IPs:
#
# 1. We'll call it ``pingtest``
# 1. We need to know which hosts the user wants to ping and how many pings the user wants to perform.
# 1. We need to verify that the arguments that the user gave us are valid. We'll also create a temp file to show that the benchmark is running.
# 1. We can run our ping tests using the ``ping`` shell command.
# 1. We need to clean up our 'I-am-running' temp file.
# 1. For each pinged host, we want to output a single result detailing the result of the ping session (RTT information, packet loss %, IP resolution, errors).
#
# ## Step One: Initialize
#
# To begin, let's create the required files for our benchmark by creating a new Python package under ``snafu/benchmarks``:
#
# ```text
# snafu/benchmarks/pingtest/
# ├── __init__.py
# └── pingtest.py
# ```
#
# Inside ``pingtest.py``, we'll create our initial ``Benchmark`` subclass. Inside this subclass, there are a few
# class variables which we need to set:
#
# 1. ``tool_name``: This is the camel-case name for our benchmark.
# 1. ``args``: These are the arguments which our Benchmark will pull from the user through the CLI, OS environment, and/or from a configuration file (CLI is preferred over the OS environment, which is preferred over the configuration file). In the background, snafu uses [configargparse](https://pypi.org/project/ConfigArgParse/) to do the dirty-work, which is a helpful wrapper around Python's own [argparse](https://docs.python.org/3/library/argparse.html). The ``args`` class variable should be set to a tuple of ``snafu.config.ConfigArgument``s, which take in arguments just like ``configargparse.ArgumentParser.add_argument``. If you have used argparse in the past, this should look super familiar.
# 1. ``metadata``: This is an iterable of strings which represent the metadata that will be exported with the Benchmark's results. Each string corresponds to the attribute name that the argument is stored under by ``configargparse``. For instance, creating a new argument under the ``args`` class variable with ``"--my-metadata", dest="mmd"`` would result in the attribute name being ``mmd``. Adding ``mmd`` under ``metadata`` will in turn cause the value for the ``--my-metadata`` argument to be exported as metadata. Benchmarks will by default specify ``cluster_name``, ``user`` and ``uuid`` arguments as metadata, but if you want to use your own set of metadata keys it can be set here.
#
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""Ping hosts and export results."""
from snafu.config import ConfigArgument
from snafu.benchmarks import Benchmark
class PingTest(Benchmark):
"""Wrapper for the Ping Test benchmark."""
tool_name = "pingtest"
args = (
ConfigArgument(
"--host",
help="Host(s) to ping. Can give more than one host by separating them with "
"spaces on the CLI and in config files, or by giving them in a "
"pythonic-list format through the OS environment",
dest="host",
nargs="+",
env_var="HOST",
type=str,
required=True,
),
ConfigArgument(
"--count",
help="Number of pings to perform per sample.",
dest="count",
env_var="COUNT",
default=1,
type=int,
),
ConfigArgument(
"--samples",
help="Number of samples to perform.",
dest="samples",
env_var="SAMPLES",
default=1,
type=int,
),
ConfigArgument(
"--htlhcdtwy",
help="Has The Large Hadron Collider Destroyed The World Yet?",
dest="htlhcdtwy",
env_var="HTLHCDTWY",
default="no",
type=str,
choices=["yes", "no"]
),
)
# don't care about Cluster Name, but the Hadron Collider is serious business
metadata = ("user", "uuid", "htlhcdtwy")
def setup(self):
"""Setup the Ping Test Benchmark."""
pass
def collect(self):
"""Run the Ping Test Benchmark and collect results."""
pass
def cleanup(self):
"""Cleanup the Ping Test Benchmark."""
pass
# -
# Let's check that we're ready to move on by trying to parse some configuration parameters. Let's load up Python!
#
# benchmark-wrapper includes a special variable called ``snafu.registry.TOOLS`` which will map a benchmark's
# camel-case string name to its wrapper class. Let's use this to create an instance of our benchmark and
# parse some configuration.
# +
from snafu.registry import TOOLS
from pprint import pprint
pingtest = TOOLS["pingtest"]()
# Set some config parameters
# Config file
# !echo "samples: 3" > my_config.yaml
# !echo "count: 5" >> my_config.yaml
# OS ENV
import os
os.environ["HOST"] = "[www.google.com,www.bing.com]"
# Parse arguments and print result
# Since we aren't running within the main script (run_snafu.py),
# need to add the config option manually
pingtest.config.parser.add_argument("--config", is_config_file=True)
pingtest.config.parse_args(
"--config my_config.yaml --labels=notebook=true --uuid 1337 --user snafu "
"--htlhcdtwy=no".split(" ")
)
pprint(vars(pingtest.config.params))
del pingtest
# !rm my_config.yaml
# -
# Now that we have our configuration all ready to go, let's start filling in our benchmark.
# ## Step Two: Setup Method
#
# Each benchmark is expected to have a ``setup`` method, which will return ``True`` if setup tasks completed successfully and otherwise return ``False``.
#
# For our use case, let's write a file to ``/tmp`` that can signal other programs that our benchmark is running. We'll also check if our temporary file exists before writing it, which would indicate that something is wrong.
#
# <div class="alert alert-info">
# Remeber, we are still working in our module found at snafu/benchmarks/pingtest.py
# </div>
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""Ping hosts and export results."""
from snafu.config import ConfigArgument
from snafu.benchmarks import Benchmark
# We'll also import this helpful function from the config module
import os
from snafu.config import check_file
class PingTest(Benchmark):
"""Wrapper for the Ping Test benchmark."""
tool_name = "pingtest"
args = (
ConfigArgument(
"--host",
help="Host(s) to ping. Can give more than one host by separating them with "
"spaces on the CLI and in config files, or by giving them in a "
"pythonic-list format through the OS environment",
dest="host",
nargs="+",
env_var="HOST",
type=str,
required=True,
),
ConfigArgument(
"--count",
help="Number of pings to perform per sample.",
dest="count",
env_var="COUNT",
default=1,
type=int,
),
ConfigArgument(
"--samples",
help="Number of samples to perform.",
dest="samples",
env_var="SAMPLES",
default=1,
type=int,
),
ConfigArgument(
"--htlhcdtwy",
help="Has The Large Hadron Collider Destroyed The World Yet?",
dest="htlhcdtwy",
env_var="HTLHCDTWY",
default="no",
type=str,
choices=["yes", "no"]
),
)
# don't care about Cluster Name, but the Hadron Collider is serious business
metadata = ("user", "uuid", "htlhcdtwy")
TMP_FILE_PATH = "/tmp/snafu-pingtest"
def setup(self) -> bool:
"""
Setup the Ping Test Benchmark.
This method creates a temporary file at ``/tmp/snafu-pingtest`` to let others
know that the benchmark is currently running.
Returns
-------
bool
True if the temporary file was created successfully, othewise False. Will
also return False if the temporary file already exists.
"""
if check_file(self.TMP_FILE_PATH):
# The benchmark base class exposes a logger at self.logger which we can use
self.logger.critical(
f"Temporary file located at {self.TMP_FILE_PATH} already exists."
)
return False
try:
tmp_file = open(self.TMP_FILE_PATH, "x")
tmp_file.close()
except Exception as e:
self.logger.critical(
f"Unable to create temporary file at {self.TMP_FILE_PATH}: {e}",
exc_info=True
)
return False
else:
self.logger.info(
f"Successfully created temp file at {self.TMP_FILE_PATH}"
)
return True
def collect(self):
"""Run the Ping Test Benchmark and collect results."""
pass
def cleanup(self):
"""Cleanup the Ping Test Benchmark."""
pass
# -
# Let's test it out and make sure our setup method works properly:
# +
from snafu.registry import TOOLS
pingtest = TOOLS["pingtest"]()
# !rm -f /tmp/snafu-pingtest
# No file exists
print(f"Setup result is: {pingtest.setup()}")
# File exists
print(f"Setup result is: {pingtest.setup()}")
# Create failure in open
# !rm -f /tmp/snafu-pingtest
open_bak = open
open = lambda file, mode: int("I'm a string")
print(f"Setup result is: {pingtest.setup()}")
# Cleanup
open = open_bak
# !rm -f /tmp/snafu-pingtest
del pingtest
# -
# ## Step Three: Cleanup Method
#
# Now let's go ahead and populate our cleanup method. The ``cleanup`` method has the same usage as the setup method: return ``True`` if the cleanup was successfull, otherwise ``False``. For the ping test benchmark, we just need to remove our temporary file:
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""Ping hosts and export results."""
import os
from snafu.config import ConfigArgument, check_file
from snafu.benchmarks import Benchmark
class PingTest(Benchmark):
"""Wrapper for the Ping Test benchmark."""
tool_name = "pingtest"
args = (
ConfigArgument(
"--host",
help="Host(s) to ping. Can give more than one host by separating them with "
"spaces on the CLI and in config files, or by giving them in a "
"pythonic-list format through the OS environment",
dest="host",
nargs="+",
env_var="HOST",
type=str,
required=True,
),
ConfigArgument(
"--count",
help="Number of pings to perform per sample.",
dest="count",
env_var="COUNT",
default=1,
type=int,
),
ConfigArgument(
"--samples",
help="Number of samples to perform.",
dest="samples",
env_var="SAMPLES",
default=1,
type=int,
),
ConfigArgument(
"--htlhcdtwy",
help="Has The Large Hadron Collider Destroyed The World Yet?",
dest="htlhcdtwy",
env_var="HTLHCDTWY",
default="no",
type=str,
choices=["yes", "no"]
),
)
# don't care about Cluster Name, but the Hadron Collider is serious business
metadata = ("user", "uuid", "htlhcdtwy")
TMP_FILE_PATH = "/tmp/snafu-pingtest"
def setup(self) -> bool:
"""
Setup the Ping Test Benchmark.
This method creates a temporary file at ``/tmp/snafu-pingtest`` to let others
know that the benchmark is currently running.
Returns
-------
bool
True if the temporary file was created successfully, othewise False. Will
also return False if the temporary file already exists.
"""
if check_file(self.TMP_FILE_PATH):
# The benchmark base class exposes a logger at self.logger which we can use
self.logger.critical(
f"Temporary file located at {self.TMP_FILE_PATH} already exists."
)
return False
try:
tmp_file = open(self.TMP_FILE_PATH, "x")
tmp_file.close()
except Exception as e:
self.logger.critical(
f"Unable to create temporary file at {self.TMP_FILE_PATH}: {e}",
exc_info=True
)
return False
else:
self.logger.info(
f"Successfully created temp file at {self.TMP_FILE_PATH}"
)
return True
def collect(self):
"""Run the Ping Test Benchmark and collect results."""
pass
def cleanup(self) -> bool:
"""
Cleanup the Ping Test Benchmark.
This method removes the temporary file at ``/tmp/snafu-pingtest`` to let others
know that the benchmark has finished running.
Returns
-------
bool
True if the temporary file was deleted successfully, otherwise False.
"""
try:
os.remove(self.TMP_FILE_PATH)
except Exception as e:
self.logger.critical(
f"Unable to remove temporary file at {self.TMP_FILE_PATH}: {e}",
exc_info=True
)
return False
else:
self.logger.info(
f"Successfully removed temp file at {self.TMP_FILE_PATH}"
)
return True
# -
# And again, some quick tests just to verify it works as expected:
# +
from snafu.registry import TOOLS
pingtest = TOOLS["pingtest"]()
# !rm -f /tmp/snafu-pingtest
# No file exists, so should error
print(f"Cleanup result is {pingtest.cleanup()}")
# Create the file using setup(), then cleanup()
print(f"Setup result is {pingtest.setup()}")
print(f"Cleanup result is {pingtest.cleanup()}")
# Cleanup
del pingtest
# -
# Now we have our setup and cleanup methods good to go, let's get to the fun part.
# ## Part Four: Collect Method
#
# The collect method is an iterable that returns a special dataclass that is shipped with benchmark-wrapper, called a ``BenchmarkResult``. BenchmarkResult holds important information about a benchmark's resulting data, such as the configuration, metadata, labels and numerical data. It also understands how to prepare itself for export. All Benchmarks are expected to return their results using this dataclass in order to support a common interface for data exporters, reduce code reuse, and reduce extra overhead.
#
# The base Benchmark class includes a helpful method called ``create_new_result``, which we will use in the example below.
#
# For our ping test benchmark, our collect method needs to run the ping command, parse its output, and yield a new BenchmarkResult. To help prevent the collect method itself from becoming super large and out of control, we'll create some new methods in our wrapper class that the collect method will call to help do its thing. Benchmarks can have any number of additional methods, just as long as they have setup, collect and cleanup.
#
# One last note here before the code: benchmark-wrapper ships with another helpful module called ``process``, which contains functions and classes to facilitate running subprocesses. In particular, we'll be using the ``LiveProcess`` wrapper.
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
"""Ping hosts and export results."""
import os
from snafu.config import ConfigArgument, check_file
from snafu.benchmarks import Benchmark
# Grab the LiveProcess class, the BenchmarkResult class, stuff
# for type hints, and dataclasses for storing our ping results
from snafu.process import LiveProcess, ProcessRun
from snafu.benchmarks import BenchmarkResult
from typing import Iterable, Optional
from dataclasses import dataclass, asdict
# Also shlex for creating our ping command
import shlex
# And finally subprocess to help with running Ping
import subprocess
@dataclass
class PingResult:
ip: Optional[str] = None
success: Optional[bool] = None
fail_msg: Optional[str] = None
host: Optional[str] = None
transmitted: Optional[int] = None
received: Optional[int] = None
packet_loss: Optional[float] = None
packet_bytes: Optional[int] = None
time_ms: Optional[float] = None
rtt_min_ms: Optional[float] = None
rtt_avg_ms: Optional[float] = None
rtt_max_ms: Optional[float] = None
rtt_mdev_ms: Optional[float] = None
class PingTest(Benchmark):
"""Wrapper for the Ping Test benchmark."""
tool_name = "pingtest"
args = (
ConfigArgument(
"--host",
help="Host(s) to ping. Can give more than one host by separating them with "
"spaces on the CLI and in config files, or by giving them in a "
"pythonic-list format through the OS environment",
dest="host",
nargs="+",
env_var="HOST",
type=str,
required=True,
),
ConfigArgument(
"--count",
help="Number of pings to perform per sample.",
dest="count",
env_var="COUNT",
default=1,
type=int,
),
ConfigArgument(
"--samples",
help="Number of samples to perform.",
dest="samples",
env_var="SAMPLES",
default=1,
type=int,
),
ConfigArgument(
"--htlhcdtwy",
help="Has The Large Hadron Collider Destroyed The World Yet?",
dest="htlhcdtwy",
env_var="HTLHCDTWY",
default="no",
type=str,
choices=["yes", "no"]
),
)
# don't care about Cluster Name, but the Hadron Collider is serious business
metadata = ("user", "uuid", "htlhcdtwy")
TMP_FILE_PATH = "/tmp/snafu-pingtest"
def setup(self) -> bool:
"""
Setup the Ping Test Benchmark.
This method creates a temporary file at ``/tmp/snafu-pingtest`` to let others
know that the benchmark is currently running.
Returns
-------
bool
True if the temporary file was created successfully, othewise False. Will
also return False if the temporary file already exists.
"""
if check_file(self.TMP_FILE_PATH):
# The benchmark base class exposes a logger at self.logger which we can use
self.logger.critical(
f"Temporary file located at {self.TMP_FILE_PATH} already exists."
)
return False
try:
tmp_file = open(self.TMP_FILE_PATH, "x")
tmp_file.close()
except Exception as e:
self.logger.critical(
f"Unable to create temporary file at {self.TMP_FILE_PATH}: {e}",
exc_info=True
)
return False
else:
self.logger.info(
f"Successfully created temp file at {self.TMP_FILE_PATH}"
)
return True
@staticmethod
def parse_host_line(host_line: str, store: PingResult) -> None:
"""
Parse the host line of ping stdout.
Expected format is: ``PING host (ip) data_bytes(ICMP_data_bytes) ...``.
Parameters
----------
host_line : str
Host line from ping to parse
store : PingResult
PingResult instance to store parsed variables into
"""
words = host_line.split(" ")
host = words[1]
ip = words[2].strip("()")
data_size = words[3]
if "(" in data_size:
data_size = data_size.split("(")[1].strip(")")
data_size = int(data_size)
if host == ip:
host = None # user pinged an IP rather than a host
store.host = host
store.ip = ip
store.packet_bytes = data_size
@staticmethod
def parse_packet_stats(packet_line: str, store: PingResult) -> None:
"""
Parse the packet statistics line of ping stdout.
Expected format is:
``A packets transmitted, B received, C% packet loss, time Dms``
Parameters
----------
packet_line : str
Packet statistics line to parse from ping
store : PingResult
PingResult instance to store parsed variables into
"""
sections = [sec.strip().split(" ") for sec in packet_line.split(",")]
store.transmitted = int(sections[0][0])
store.received = int(sections[1][0])
store.packet_loss = float(sections[2][0].strip("%"))
store.time_ms = int(sections[3][1].strip("ms"))
@staticmethod
def parse_rtt_stats(rtt_line: str, store: PingResult) -> None:
"""
Parse the RTT statistics line of ping stdout.
Expected format is: ``rtt min/avg/max/mdev = A/B/C/D ms``
Parameters
----------
rtt_line : str
RTT statistics line to parse from ping
store : PingResult
PingResult instance to store parsed variables into
"""
rtt_min, rtt_avg, rtt_max, rtt_mdev = map(
float, rtt_line.split("=")[1].strip(" ms").split("/")
)
store.rtt_min_ms = rtt_min
store.rtt_avg_ms = rtt_avg
store.rtt_max_ms = rtt_max
store.rtt_mdev_ms = rtt_mdev
def parse_stdout(self, stdout: str) -> PingResult:
"""
Parse the stdout of the ping command.
Tested against ping from iputils 20210202 on Fedora Linux 34
Returns
-------
PingResult
"""
# We really only care about the first line, and the last two lines
lines = stdout.strip().split("\n")
# Check if we got an error
if len(lines) == 1:
msg = lines[0]
return PingResult(
fail_msg=msg,
success=False
)
host_info = lines[0]
packet_info = lines[-2]
rtt_info = lines[-1]
result = PingResult(success=True)
self.parse_host_line(host_info, result)
self.parse_packet_stats(packet_info, result)
self.parse_rtt_stats(rtt_info, result)
return result
def ping_host(self, host: str) -> Iterable[BenchmarkResult]:
"""
Run the ping test benchmark against the given host.
Parameters
----------
host : str
Host to ping
Returns
-------
iterable
Iterable of BenchmarkResults
"""
self.logger.info(f"Running ping test against host {host}")
cmd = shlex.split(f"ping -c {self.config.count} {host}")
self.logger.debug(f"Using command: {cmd}")
# A config instance allows for accessing params directly,
# therefore self.config.samples == self.config.params.samples
for sample_num in range(self.config.samples):
self.logger.info(f"Collecting sample {sample_num}")
# We'll use the LiveProcess context manager to run ping
# LiveProcess will expose the Popen object at 'process',
# create a queue with lines from stdout at 'stdout',
# and create a snafu.process.ProcessRun instance at `attempt`
# Here we will tell LiveProcess to send stdout and stderr
# to the same pipe
with LiveProcess(cmd, stdout=subprocess.PIPE, stderr=subprocess.STDOUT) as lp:
# if process hasn't finished
while lp.process.poll() is None:
# if we have a new line in stdout to get
if not lp.stdout.empty():
self.logger.debug(lp.stdout.get())
# when we get here, the process has finished
process_run: ProcessRun = lp.attempt
self.logger.debug(f"Got process run: {vars(process_run)}")
result: PingResult = self.parse_stdout(process_run.stdout)
# manually set host if we fail, since it won't always be parsable
# through stdout
if result.success is False:
result.host = host
self.logger.info(f"Got sample: {vars(result)}")
yield self.create_new_result(
# We use vars here because create_new_result expects
# dict objects, not dataclasses
data=vars(result),
config={"samples": self.config.samples, "count": self.config.count},
# tag is a method for labeling results for exporters
# right now it specifies the ES index to export to
tag="jupyter"
)
plural = "s" if self.config.samples > 1 else ""
self.logger.info(
f"Finised collecting {self.config.samples} sample{plural} against {host}"
)
def collect(self) -> Iterable[BenchmarkResult]:
"""
Run the Ping Test Benchmark and collect results.
"""
self.logger.info("Running pings and collecting results.")
self.logger.debug(f"Using config: {vars(self.config.params)}")
if isinstance(self.config.host, str):
yield from self.ping_host(self.config.host)
else:
for host in self.config.host:
yield from self.ping_host(host)
self.logger.info("Finished")
def cleanup(self) -> bool:
"""
Cleanup the Ping Test Benchmark.
This method removes the temporary file at ``/tmp/snafu-pingtest`` to let others
know that the benchmark has finished running.
Returns
-------
bool
True if the temporary file was deleted successfully, otherwise False.
"""
try:
os.remove(self.TMP_FILE_PATH)
except Exception as e:
self.logger.critical(
f"Unable to remove temporary file at {self.TMP_FILE_PATH}: {e}",
exc_info=True
)
return False
else:
self.logger.info(
f"Successfully removed temp file at {self.TMP_FILE_PATH}"
)
return True
# -
# We have finished our new ping test benchmark! Let's try it out!
# +
from snafu.registry import TOOLS
from pprint import pprint
import logging
pingtest = TOOLS["pingtest"]()
# All Benchmark loggers work under the "snafu" logger
logger = logging.getLogger("snafu")
if not logger.hasHandlers():
logger.addHandler(logging.StreamHandler())
logger.setLevel(logging.DEBUG)
# !rm -rf /tmp/snafu-pingtest
# !rm -f my_config.yaml
# Set some config parameters
# Config file
# !echo "samples: 1" > my_config.yaml
# !echo "count: 5" >> my_config.yaml
# OS ENV
import os
os.environ["HOST"] = "[www.google.com,www.bing.com,www.idontexist.heythere]"
# Parse arguments and print result
# Since we aren't running within the main script (run_snafu.py),
# need to add the config option manually
pingtest.config.parser.add_argument("--config", is_config_file=True)
pingtest.config.parse_args(
"--config my_config.yaml --labels=notebook=true --uuid 1337 --user snafu "
"--htlhcdtwy=no".split(" ")
)
# The base benchmark class includes a run method that runs setup -> collect -> cleanup
results = list(pingtest.run())
# !rm -rf /tmp/snafu-pingtest
# !rm -f my_config.yaml
# -
print(f"Got {len(results)} results")
for result in results[:5]:
pprint(vars(result))
# And that's that! As soon as you have your benchmark working that way you'd like, submit a PR and we'll give it a LGTM.
|
docs/source/contributing/adding_workloads.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Managing Workforce projects
#
# [ArcGIS Workforce](http://doc.arcgis.com/en/workforce/) is a mobile solution that uses the power of location-based decision making for better field workforce coordination and teamwork. It is composed of a web app used by project administrators and dispatchers in the office, and a mobile app used by mobile workers on their devices. Organizations using ArcGIS Workforce get these benefits:
#
# - Everything you need on one device. Mobile workers can easily view and process work assignments, provide updates on work status, and inform others of their location, all from one device.
#
# - Greater agility—Using real-time and location-based information, dispatchers can assign and prioritize fieldwork on the fly and ensure that work is assigned to the right people at the right time.
#
# - Increased productivity—Replace time-consuming and error-prone manual workforce management processes, reduce downtime, and keep projects on schedule.
#
# Please note that working with an offline-enabled Workforce project ensures you have a version of the Python API > 1.8.2.
# ## Contents of a Workforce project
#
# An offline-enabled (Version 2) workforce project is composed of two web maps and a feature service. The feature service contains two layers and three tables:
#
# **Layers**
# - **Assignments** - contains a record for each assignment. It stores information such as status, location, and description, among others.
# - **Workers** - contains a record for each mobile worker along with their contact number and job title.
#
# **Tables**
# - **Dispatchers** - contains a record for each dispatcher within the project and includes information such as name and contact number.
# - **Assignment Types** - contains a record for each type of assignment that can be added to the project.
# - **Assignment Integrations** - contains a record for each URL an assignment can link to in order to perform external actions such as navigation and survey collection
#
# **Web Maps**
# - **Dispatcher web map** - for back-office dispatchers. It shows the assignments and worker locations
# - **Worker web map** - This map is what a field worker uses on their iOS or Android device.
#
# The webmaps and feature service are connected via a "WorkforceMap2FeatureService" relationship. To learn more about those details, refer [here](https://doc.arcgis.com/en/workforce/android-phone/help/workforce-schema.htm). Previously, Workforce used a "classic" project consisting of four feature layers and two web maps; these can still be managed and created using the Python API.
# ## What can you do with this module?
# Using the `workforce` module under the `arcgis.apps` module, you can automate the following tasks:
#
# ### Projects
# - Creating Projects
# - Managing Projects created either through Python or the web app
# - Deleting Projects
#
# ### Workers and Dispatchers
# - Adding Dispatchers and Workers to a Project
# - Deleting Dispatchers and Workers from a Project
# - Updating Workers and Dispatchers in a Project
# - Searching Workers and Dispatchers in a Project
#
# ### Assignments
# - Adding Assignments to a Project
# - Deleting Assignments from a Project
# - Updating Assignments in a Project
# - Assigning Assignments in a Project
# - Searching Assignments in a Project
# - Adding/Removing/Downloading Attachments
#
# ### Integrations
# - Adding, Updating, and Deleting Integrations
# ## Get Started
#
# A user must be authenticated with a GIS in order to fetch or create a Project. The workforce functionality is available in `arcgis.apps.workforce` module.
# +
from arcgis.gis import GIS
from arcgis.apps import workforce
gis = GIS('home')
# -
# ### Workforce Project
# A project is created using its corresponding item or created via the `create_project` function. This can either be a version 1 "Classic" workforce project or a version 2 (recommended) "Offline" workforce project. While the functions are the same no matter if you're using a Classic or Offline Workforce project, you want to ensure the item you pass into the `arcgis.apps.workforce.Project` class is correct - a Workforce Project item for classic and a hosted feature service for offline.
## These create instances of arcgis.apps.workforce.Project
old_workforce_project = arcgis.apps.workforce.create_project('old project', major_version=1)
new_workforce_project = arcgis.apps.workforce.create_project('new project', major_version=2)
# +
## This instantiates a project that previously existed for a classic project
workforce_project_item = gis.content.get("c63e3d46af7d4204b66b18d43a188c2e")
workforce_project_item
# This instantiates a project that previously existed for an offline-enabled project.
fs_item_id = "04b66b18d43a1804b66b1a188c2e"
fs_item = gis.content.get(fs_item_id)
offline_project = arcgis.apps.workforce.Project(fs_item)
# -
project = workforce.Project(item=workforce_project_item)
project
# ### Assignments
# `Assignment` objects are accessed by using the `assignments` property off the `Project` object. Assignments can be added, updated, or deleted. Additionally, attachments can be added, deleted, or downloaded by using the `attachments` property off the `Assignment` object.
# Search all assignments
assignments = project.assignments.search()
assignments
# +
# View first assignment
assignment = assignments[0]
print(f"Status: {assignment.status}")
print(f"Description: {assignment.description}")
print(f"Priority: {assignment.priority}")
print(f"Assigned To: {assignment.worker.name}")
print(f"Type: {assignment.assignment_type}")
# Update the description of the assignment
assignment.update(description="You need to do an inspection here")
print("--------------------")
print(f"Updated Description: {project.assignments.search()[0].description}")
# Download the assignment using the AssignmentAttachmentManager
assignment.attachments.download()
# -
# ### Assignment Types
# Assignment types are created when the workforce project is originally created. All assignments fall under one or the other assignment types. You can access these types by calling the `assignment_types` property off the `Project` item. Assignment types can be added, updated, or deleted.
# +
# List all assignment types
assignment_types = project.assignment_types.search()
for at in assignment_types:
print(f"Type: {at.name}")
# Add a new assignment type
project.assignment_types.add(name="Repair")
# Confirm that it was added
print("--------------------")
assignment_types = project.assignment_types.search()
for at in assignment_types:
print(f"Type: {at.name}")
# -
# ### Workers
# `Worker` objects are accessed by using `workers` property off the `Project` object. Workers can be added, updated, or deleted.
# +
# Search all workers and print details about first worker
workers = project.workers.search()
worker = workers[0]
print(f"Name: {worker.name}")
print(f"Number: {worker.contact_number}")
# Update the workers contact number
worker.update(contact_number="123-456-7890")
print("--------------------")
print(f"Number: {project.workers.search()[0].contact_number}")
# -
# **Note**: Workers and Dispatchers should be named users in your GIS.
# Add a new worker
project.workers.add(name="Demo User",
user_id="demouser_nitro",
contact_number="123-987-4560")
# ### Dispatchers
# `Dispatcher` objects are accessed by using `dispatchers` property off the `Project` object. Dispatchers can be added, updated, or deleted.
# +
# Search for all dispatchers and print details about first dispatcher
dispatchers = project.dispatchers.search()
dispatcher = dispatchers[0]
print(f"Name: {dispatcher.name}")
print(f"Number: {dispatcher.contact_number}")
# Update the dispatchers contact number
dispatcher.update(contact_number="123-456-7890")
print("--------------------")
print(f"Number: {project.dispatchers.search()[0].contact_number}")
# -
# ### Integrations
# `Integration` objects are accessed by using `integrations` property off the `Project` object. Integrations can be added, updated, or deleted.
## Search for all integrations
integrations = project.integrations.search()
integration = integrations[0]
integration.update(integration_id="arcgis-explorer")
# ### Web maps
# The dispatcher and worker `WebMap` objects can be accessed using the corresponding properties as shown in the following code snippet. Using the `WebMap` object, additional layers could be added or removed from either maps.
# Worker webmap
worker_webmap = project.worker_webmap
type(worker_webmap)
worker_webmap
# 
# Dispatcher webmap
dispatcher_webmap = project.dispatcher_webmap
dispatcher_webmap
# 
# ## Putting it all together
# In the following snippet, a new assignment will be created at the ESRI campus. Assignments (as well as workers, assignment types, dispatchers, and integrations) are all validated prior to upload. The ensures the integrity of the workforce project.
# +
# Add a new assignment and assign it to demouser
from datetime import datetime
demouser = project.workers.get(user_id='demouser_nitro')
dispatcher = project.dispatchers.get(user_id='workforce_scripts')
repair = project.assignment_types.get(name="Repair")
# Use the geocoder to find the location of ESRI
from arcgis.geocoding import geocode
geometry = geocode("ESRI, Redlands, CA", out_sr=102100)[0]['location']
# Add a new assignment
project.assignments.add(assignment_type=repair,
status="assigned",
assigned_date=datetime.now(),
worker=demouser,
dispatcher=dispatcher,
location="ESRI, Redlands, CA",
geometry=geometry)
# -
# ### Reset the Demo Project
# The following code resets the project to the intial state for this demo. It also highlights how batch methods can be used to add/update/delete many workforce items at once. This is useful when processing large amounts of assignments, workers, or dispatchers. In general, batch methods make fewer calls to the backend server thus reducing the overall time of the script.
# +
# Use batch functions to process multiple items at a time
project.assignment_types.batch_delete([project.assignment_types.get(name="Repair")])
project.workers.batch_delete([project.workers.get(user_id="demouser_nitro")])
# Reset the assignment description
a = project.assignments.get(object_id=1)
a.update(description="Do some work at the ESRI R&D Center")
# Reset the worker using batch update
w1 = project.workers.get(object_id=1)
w1.contact_number = None
project.workers.batch_update([w1])
# Reset the dispatcher using batch update
d1 = project.dispatchers.get(object_id=1)
d1.contact_number = "987-654-3210"
project.dispatchers.batch_update([d1])
# Reset integration using batch update
i1 = project.integrations.get(integration_id="arcgis-explorer")[0]
i1.integration_id = "default-navigator"
project.integrations.batch_update([i1])
|
guide/13-managing-arcgis-applications/managing-workforce-projects.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Erik-Silver/daa_2021_1/blob/master/19octubre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="LhmMFmqHSG1a" outputId="feff0987-59c0-4229-a093-57de7324f70d" colab={"base_uri": "https://localhost:8080/", "height": 510}
def ejemplo4(n):
count = 0
i = n
print(f"Nivel 1: {i}")
while i >= 1:
count += 1
print(f"Nivel 2, contador: {count}")
i = i //2
basura = 3+2
return count
prueba = ejemplo4(10)
print(f"10 - Procesos: {prueba}")
prueba = ejemplo4(4)
print(f"10 - Procesos: {prueba}")
prueba = ejemplo4(8)
print(f"10 - Procesos: {prueba}")
prueba = ejemplo4(5)
print(f"10 - Procesos: {prueba}")
prueba = ejemplo4(20)
print(f"10 - Procesos: {prueba}")
# + id="_tjll5HH4Kjl"
|
19octubre.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
tips = sns.load_dataset('tips')
# ### style
sns.countplot(x='sex',data=tips)
sns.set_style('white')
sns.countplot(x='sex',data=tips)
sns.set_style('ticks')
sns.countplot(x='sex',data=tips,palette='deep')
sns.countplot(x='sex',data=tips)
sns.despine()
sns.countplot(x='sex',data=tips)
sns.despine(left=True)
# Non Grid Plot
plt.figure(figsize=(12,3))
sns.countplot(x='sex',data=tips)
# Grid Type Plot
sns.lmplot(x='total_bill',y='tip',size=2,aspect=4,data=tips)
sns.set_context('poster',font_scale=4)
sns.countplot(x='sex',data=tips,palette='coolwarm')
|
Seaborn/Style and Color.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vdnew/Performance-Metric-without-any-library/blob/main/5_Performance_metrics_Instructions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="s0Ej_bXyQvnV"
# # Compute performance metrics for the given Y and Y_score without sklearn
# + id="4CHb6NE7Qvnc"
import numpy as np
import pandas as pd
# other than these two you should not import any other packages
# + [markdown] id="KbsWXuDaQvnq"
# <pre>
# <font color='red'><b>A.</b></font> Compute performance metrics for the given data <strong>5_a.csv</strong>
# <b>Note 1:</b> in this data you can see number of positive points >> number of negatives points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_a.csv</b>
# <b>Note 3:</b> you need to derive the class labels from given score</pre> $y^{pred}= \text{[0 if y_score < 0.5 else 1]}$
#
# <pre>
# <ol>
# <li> Compute Confusion Matrix </li>
# <li> Compute F1 Score </li>
# <li> Compute AUC Score, you need to compute different thresholds and for each threshold compute tpr,fpr and then use numpy.trapz(tpr_array, fpr_array) <a href='https://stackoverflow.com/q/53603376/4084039'>https://stackoverflow.com/q/53603376/4084039</a>, <a href='https://stackoverflow.com/a/39678975/4084039'>https://stackoverflow.com/a/39678975/4084039</a> Note: it should be numpy.trapz(tpr_array, fpr_array) not numpy.trapz(fpr_array, tpr_array)</li>
# <li> Compute Accuracy Score </li>
# </ol>
# </pre>
# + id="WaFLW7oBQvnt"
# write your code here
data_a = pd.read_csv("/content/5_a.csv")
data_a.shape
# + id="nQtNj6VD5AjH"
# predict defination
def predict(data, y, thresh_hold):
'''
Argument:
data, DataFrame
y = 'y' column in dataframe
thresh_hold = Thresh hold value for predict method
AIM: predict the value as '1' if y value is more than 0.5 else it will be '0'
Output:
will return y_pred list as return type with classified value either '1' or '0'
'''
#initialize y_pred list
y_pred = []
# check label with thresh_hold value (0.5)
for label in data[y]:
if label<thresh_hold:
y_pred.append(0)
else:
y_pred.append(1)
return y_pred
# confusion matrix calculation method
def calculate_confusion(data, y, y_pred):
'''
Argument:
data = DataFrame
y = 'y' column in dataframe
y_pred = 'y_pred' column in dataframe
AIM: calculate confusion matrix: TP, TN, FN, FP
Output: dictionary with TN, TP, FN, FP as key and value pair
'''
# initialize variables
tp = 0
tn = 0
fn = 0
fp = 0
# confuntion matrix tp, tn, fn, fp calculation logic
for value1, value2 in enumerate(data['y']):
if (data.y_pred[value1]==1) and data.y[value1] == 1:
tp = tp + 1
if (data.y_pred[value1]==0) and data.y[value1] == 0:
tn = tn + 1
if (data.y_pred[value1]==0) and data.y[value1] == 1:
fn = fn + 1
if (data.y_pred[value1]==1) and data.y[value1] == 0:
fp = fp + 1
return { 'tn': tn, 'tp': tp, 'fn': fn, 'fp': fp}
# + id="A03BLmCr7XCK"
# Thresh hold value declation
thresh_hold = 0.5
# predicting the value and storing in dataframe as y_pred column
data_a['y_pred'] = predict(data_a, 'proba', thresh_hold)
# calculating confunsion matrix
confusion_matrix = calculate_confusion(data_a, 'y', 'y_pred')
# + id="3BBrDuyZ7zi_"
# confusion matrix values
print("the confusion matrix is:", confusion_matrix)
# + id="4U1pZQGw78lx"
# F1 score calculation
x = data_a.y.value_counts()
P = x[1]
# precision calculation = TP / (TP + FP)
precision = confusion_matrix['tp']/(confusion_matrix['tp']+confusion_matrix['fp'])
# recall calculation = TP / P
recall = confusion_matrix['tp']/P
# F1 score calculation with formula
F1 = 2* precision * recall / (precision + recall)
# display result of F1 score
print("the F1 score is:", F1)
# + id="dKBZAI808_Zs"
# Accuracy calculation with formula
Acc = (confusion_matrix['tp']+confusion_matrix['tn'])/data_a.shape[0]
print("The accuracy is:",Acc)
# + id="OHqhDiwB-BHK"
# AUC score calculation defination
from tqdm import tqdm_notebook
def auc(data):
'''
Argument:
data: DataFrame
AIM: This method to calculate the AUC (Area under the curve)
with confuntion matrix, it will calculate tpr and fpr.
Output: Definite integral as approximated by trapezoidal rule.
'''
# variable initialization
s = data['y'].value_counts()
P = s[1]
N = s[0]
tpr = []
fpr = []
#calculating auc
for element in tqdm_notebook(data['proba']):
data['y_pred'] = predict(data, 'proba', element)
confusion_matrix= calculate_confusion(data, 'y', 'y_pred')
tpr.append(confusion_matrix['tp']/P)
fpr.append(confusion_matrix['fp']/N)
data.drop(columns=['y_pred'])
return np.trapz(tpr, fpr)
# + id="EgA1H81j_BsS"
# sort the value by 'proba' value
data_a = data_a.sort_values(by='proba',ascending=False)
# droping 'y_pred' column
data_a.drop(columns=['y_pred'])
# + id="adDQSVfqAV8J"
# calculate AUC score
AUC_score=auc(data_a)
# display auc score
print("The AUC Score is :",AUC_score)
# + [markdown] id="V5KZem1BQvn2"
# <pre>
# <font color='red'><b>B.</b></font> Compute performance metrics for the given data <strong>5_b.csv</strong>
# <b>Note 1:</b> in this data you can see number of positive points << number of negatives points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_b.csv</b>
# <b>Note 3:</b> you need to derive the class labels from given score</pre> $y^{pred}= \text{[0 if y_score < 0.5 else 1]}$
#
# <pre>
# <ol>
# <li> Compute Confusion Matrix </li>
# <li> Compute F1 Score </li>
# <li> Compute AUC Score, you need to compute different thresholds and for each threshold compute tpr,fpr and then use numpy.trapz(tpr_array, fpr_array) <a href='https://stackoverflow.com/q/53603376/4084039'>https://stackoverflow.com/q/53603376/4084039</a>, <a href='https://stackoverflow.com/a/39678975/4084039'>https://stackoverflow.com/a/39678975/4084039</a></li>
# <li> Compute Accuracy Score </li>
# </ol>
# </pre>
# + id="U2sKlq0YQvn5"
# write your code
# reading dataframe b
data_b = pd.read_csv("/content/5_b.csv")
data_b.head()
# + id="tA_iWVxdjwch"
# thresh hold value intialization
thresh_hold = 0.5
# predicting y_pred using predict method
data_b['y_pred'] = predict(data_b, 'proba', thresh_hold)
# calculating confusion matrix
confusion_matrix_b = calculate_confusion(data_b, 'y', 'y_pred')
# + id="4Z3b_2e1kNtY"
# confusion matrix result display
print("The confusion matrix is :",confusion_matrix_b)
# + id="bwPi7FcXkYP0"
# F1 score calculation
x = data_b.y.value_counts()
P = x[1]
# precision calculation = TP / (TP + FP)
precision_b = confusion_matrix_b['tp']/(confusion_matrix_b['tp']+confusion_matrix_b['fp'])
# recall calculation = Tp / P
recall_b = confusion_matrix_b['tp'] / P
#F1 calculation
f1_b = 2*precision_b*recall_b / (precision_b + recall_b)
print("The F1 Score is :", f1_b)
# + id="IevUsvNUlF1t"
# Accuracy calculation
Acc_b = (confusion_matrix_b['tp'])+confusion_matrix_b['tn']/data_b.shape[0]
print("The accuracy is :", Acc_b)
# + id="nS_wXn_ClZfF"
# Auc score calculation
data_b = data_b.sort_values(by='proba', ascending=False)
print("The Accuracy is :", Acc_b)
# + [markdown] id="GiPGonTzQvoB"
# <font color='red'><b>C.</b></font> Compute the best threshold (similarly to ROC curve computation) of probability which gives lowest values of metric <b>A</b> for the given data <strong>5_c.csv</strong>
# <br>
#
# you will be predicting label of a data points like this: $y^{pred}= \text{[0 if y_score < threshold else 1]}$
#
# $ A = 500 \times \text{number of false negative} + 100 \times \text{numebr of false positive}$
#
# <pre>
# <b>Note 1:</b> in this data you can see number of negative points > number of positive points
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_c.csv</b>
# </pre>
# + id="x5HIJzq1QvoE"
# write your code
# reading dataframe c
data_c = pd.read_csv("/content/5_c.csv")
data_c.shape
# + id="l2klV05mI0Z3"
# min metric calculation method
def Min_MetricCalculation(data):
'''
Argument:
data: DataFrame
AIM:
Min matrix calculation
Output:
metric as dictionary
'''
# initialize variables
s = data['y'].value_counts()
P = s[1]
N = s[0]
tpr = []
fpr = []
metric = {}
for element in tqdm_notebook(data['prob']):
data['y_pred'] = predict(data, 'prob', element)
confusion_matrix_c = calculate_confusion(data, 'y', 'y_pred')
metric_value = (500*confusion_matrix_c['fn'])+(100*confusion_matrix_c['fp'])
metric[element] = metric_value
data.drop(columns=['y_pred'])
return (metric)
# + id="f8C-QqYBNGWF"
# sorting dataframe values by prob column
data_c = data_c.sort_values(by='prob', ascending=False)
# calculating min matrix
result = Min_MetricCalculation(data_c)
# + id="YUvBTl1zNRSH"
temp = min(result.values())
res = [key for key in result if result[key] == temp]
print("The key:value pair for min value of the specified metric is :", res, temp)
# + [markdown] id="sD4CcgjXQvoL"
# <pre>
# <font color='red'><b>D.</b></font> Compute performance metrics(for regression) for the given data <strong>5_d.csv</strong>
# <b>Note 2:</b> use pandas or numpy to read the data from <b>5_d.csv</b>
# <b>Note 1:</b> <b>5_d.csv</b> will having two columns Y and predicted_Y both are real valued features
# <ol>
# <li> Compute Mean Square Error </li>
# <li> Compute MAPE: https://www.youtube.com/watch?v=ly6ztgIkUxk</li>
# <li> Compute R^2 error: https://en.wikipedia.org/wiki/Coefficient_of_determination#Definitions </li>
# </ol>
# </pre>
# + id="2JuA0zUnQ5ox"
# reading dataframe d
data_d = pd.read_csv("/content/5_d.csv")
data_d.head()
# + [markdown] id="2STAIj1Gxxbh"
#
# + id="FEMNoPmRQ9NL"
# error calculation method
def Error_Calculation(data, column1, column2):
value = []
for index, (value1, value2) in enumerate(zip(data[column1], data[column2])):
value.append(value1-value2)
return value
# absolute error calculation method
def absolute_error(data,column):
val=[]
for index,value in enumerate(data[column]):
val.append(abs(value))
return val
# mean sqaure error calculation method
def mean_sqaure_error(data, column):
return ss_res(data,column)/len(data[column])
# method for mape
def mape(data, column1, column2):
value = sum(data[column1]/sum(data[column2]))
return value
# method for ss_res
def ss_res(data, column):
value = 0
for index, value in enumerate(data[column]):
value = value + (value*value)
return value
# method for ss_tot
def ss_tot(data, column):
value = 0
mean_value = data_d['y'].mean()
for index, value in enumerate(data[column]):
value = value + (value-mean_value)*(value-mean_value)
return value
# + id="kjXXO2vmTyCv"
# calculating the error
data_d['error'] = Error_Calculation(data_d, 'y', 'pred')
# calculating the absolute error
data_d['abs_error'] = absolute_error(data_d,'error')
# + id="m6nhXSvkUGgE"
MSE = mean_sqaure_error(data_d, 'error')
print("The Mean sqaured error is :", MSE)
# + id="74gvbGhaVaw9"
# calculating mape
MAPE = mape(data_d, 'abs_error', 'y')
print("The MAPE value is :", MAPE)
# + id="tTZjf9hFVzg5"
# calculating the co-efficient of determination with formula and methods
SS_RES = ss_res(data_d, 'error')
SS_TOT = ss_tot(data_d, 'y')
R_square = 1 - (SS_RES/ SS_TOT)
print("The Co-efficient of determination value is :", R_square)
# + id="1L5HsPPGWBiR"
# Reference
# https://towardsdatascience.com/understanding-confusion-matrix-a9ad42dcfd62 - for different way to understand
#
|
5_Performance_metrics_Instructions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pythonで遊ぼう
#
# ## 目的
#
# - Pythonで遊べる環境を構築する.
# - Pythonと機械学習ライブラリをわかった気になる.
# ## 準備
#
# `PowerShell`を開いて,以下のコマンドを打ってください.
#
# ```
# > jupyter notebook
# ```
#
# エラーが出たら教えてください.無事Notebookが開けたら,以下のセルを選択して`Shift + Enter`を押してください.おまじないです.
# Notebook上できれいに図を表示するためのおまじない.
# %matplotlib inline
# %config InlineBackend.figure_formats = {'png', 'retina'}
# ## Jupyter notebook
#
# 本資料はJupyter notebookで作成しています.Jupyter notebookは,コメントを残しながらプログラミングできるのでとても便利です.
#
# - `In []`と書かれたブロックで`Shift + Enter`すると,Pythonプログラムを実行することができます.
print('Hello world!')
# - ブロックを選択して`Enter`キーを押すと,ブロックの中身を編集できます.`esc`キーを押すと選択を解除できます.
# - 選択を解除した状態で`m`キーを押すと,マークダウンブロックになります.このブロックのように本文が編集できます.
# - 選択を解除した状態で`y`キーを押すと,コマンドブロックになります.ブロックの左側に`In []`が付き,Pythonスクリプトを実行可能になります.
# +
# このブロックを,マークダウンブロックと,コマンドブロックで切り替えて遊んでください.マークダウンブロックだと大見出し,コマンドブロックだとコメントになるはずです.
# -
# - 選択を解除した状態で`a`キーを押すと,上にブロックを追加できます.`b`キーを押すと下にブロックを追加できます.
# - 選択を解除した状態で`d`キーを二連打すると,ブロックを削除できます.`z`キーを押すと,削除したブロックを復元できます.
# - 選択を解除した状態で`s`キーを押すと,Notebookを保存できます.
# - 選択を解除した状態で`h`キーを押すと,ヘルプを見ることができます.
# ## 目次
#
# 本Notebookでは,Pythonの基本的な動かし方と,データ分析によく用いるライブラリ(徐々に応用的なライブラリに進みます)をざっくり説明します.細かいところは飛ばすので,気になる場合はググってください.
#
# - Python:基本の基本
# - Numpy:数値計算ライブラリ
# - Matplotlib:グラフ描画ライブラリ
# - Pandas:データ処理ライブラリ
# - Scikit learn:機械学習モデル全般のライブラリ
# - Light GBM:勾配ブースティングモデルのライブラリ
# ## Python:基本の基本
# ### 四則演算
1 + 2
1 - 2
1 * 2
5 / 3
# 割り切り
5 // 3
# あまり
5 % 3
# 累乗
5 ** 3
# ### 代入
a = 1
b = 2
a + b
# aの値を変更
a = 5
a + b
# ### リスト
a = [0, 1, 2, 3, 4]
a
# 0番目の要素を抽出
a[0]
# 最後の要素を抽出
a[-1]
# 1番目から2番目(3番目ではない)までの要素を抽出
a[1:3]
# リストの長さ
len(a)
# リストに値を追加
a.append(10)
a
# ### ループ
for i in [0, 1, 2, 3, 4, 5]:
# iを表示
print(i)
# range()を使うと同じことができる.
for i in range(6):
print(i)
# ### 条件分岐
a = 1
if a<5:
print('aは5より小さい')
else:
print('aは5以上')
# aの値を変更.
a = 10
if a<5:
print('aは5より小さい')
else:
print('aは5以上')
# ### ループと条件分岐
# 3の倍数のときだけアホになる
for i in range(15):
if i%3==0: # iを3で割ったあまりが0のとき
print('なんだっけ?')
else:
print(i)
# ## Numpy:数値計算ライブラリ
import numpy as np # numpyというライブラリを,npという名前で使います
# ### ベクトル
a = np.array([0, 1, 2, 3]) # np.array(リスト)と書く
a
a + 1
a - 2
a * 3
a / 4
b = np.array([2, 3, 4, 5])
a + b
a - b
a * b
a / b
np.dot(a, b) # 内積
# ### 行列
# リストを入れ子にすると行列が作れる.
# 入れ子の数はいくらでも増やせる.
a = np.array([
[0, 1, 2],
[2, 3, 4]])
a
a + 2
a - 2
# ### 便利関数
a
a.max()
# 縦方向の最大値
a.max(axis=0)
# 横方向の最大値
a.max(axis=1)
# 最小値
a.min()
# 平均値
a.mean()
# 標準偏差
a.std()
# ## Matplotlib:グラフ描画ライブラリ
import matplotlib.pyplot as plt # matplotlibのpyplotをpltと呼んで使います
x = [0, 1, 2, 3, 4]
y = [0, 3, 4, 2, 1]
plt.plot(x, y, '-') # 折れ線グラフ
plt.plot(x, y, 'o') # 散布図
plt.bar(x, y) # 棒グラフ
plt.hist(x) # ヒストグラム
# グラフの細かい設定等は,ググってください.
# ### Pandas:データ処理ライブラリ
#
# エクセルのデータ分析的なことはだいたいできます.Pandasは内部でNumpyを使っています.
import pandas as pd # pandasをpdと呼んで使えるようにする.
# data/rawにあるboston.csvを読み込む
data = pd.read_csv('data/raw/boston.csv')
data # とりあえず全体を表示
# `boston.csv`は次のようなデータで,練習問題としてよく使われます.
#
# - CRIM:人口 1 人当たりの犯罪発生数
# - ZN:25,000 平方フィート以上の住居区画の占める割合
# - INDUS:小売業以外の商業が占める面積の割合
# - CHAS:チャールズ川によるダミー変数 (1: 川の周辺, 0: それ以外)
# - NOX:NOx の濃度
# - RM:住居の平均部屋数
# - AGE:1940 年より前に建てられた物件の割合
# - DIS:5 つのボストン市の雇用施設からの距離 (重み付け済)
# - RAD:環状高速道路へのアクセスしやすさ
# - TAX:$10,000 ドルあたりの不動産税率の総計
# - PTRATIO:町毎の児童と教師の比率
# - B:町毎の黒人 (Bk) の比率を次の式で表したもの。 1000(Bk – 0.63)^2
# - LSTAT:給与の低い職業に従事する人口の割合 (%)
# - PRICE:住宅価格
#
# ここでは,`CRIM`から`LSTAT`の情報を使って,`PRICE`を予測する問題に挑戦します.
# ### 外観
data.shape # 大きさを確認
data.head(3) # 最初の3行だけ表示
data.tail(3) # 最後の3行だけ表示
data.describe() # 各種指標で要約
data.corr() # 相関係数
# ### ソート
# PRICEが小さい順にソート
data.sort_values('PRICE')
# ### 抽出
data['CRIM'] # CRIM列だけ抜き出す
data.iloc[5] # 5行目だけ表示
data.iloc[:, 0:5]
# [行数,列数]を指定できる.「:」は全部,を意味する.
# 上記は,0-4列目までの全行を抽出する.
# 行名一覧
data.columns
# 住宅価格が10以上のものだけ抽出
data[data['PRICE']>20]
# 住宅価格が20以上で,かつCRIMEが3.5以上のものを抽出
data[(data['PRICE']>=20)&(data['CRIM']>=3.5)]
# ### Numpyに変換
data.values
# ### プロット
# CRIMとPRICEの散布図を書く.
# alphaで透明度を指定できる.
plt.plot(data['CRIM'], data['PRICE'], 'o', alpha=.5)
# 住宅価格のヒストグラム
plt.hist(data['PRICE'])
temp = data.corr()['PRICE'] # 相関行列のPRICE行のみ抽出
plt.plot(temp)
plt.xticks(rotation=90) # x軸名を90度回転してみやすく.
# RM(住居の平均部屋数)が最も相関が大きいことがわかります.
# ## Scikit-learn:機械学習ライブラリ
#
# 機械学習系のモデルを簡単に利用できます.今回は,線形回帰モデル(LinearRegressor)を使います.
#
# $ y = a_1 x_1 + a_2 x_2 + \dots + a_k x_k $
from sklearn.linear_model import LinearRegression
# sklearn(scikit-learn)のLinearRegression(線形回帰モデル)を利用します.
# データ整形
data = pd.read_csv('data/raw/boston.csv')
x = data.iloc[:, :-1] # -1番目のカラムまで(PRICE以外)抽出
y = data.iloc[:, -1] # PRICEのみ抽出
x.head()
y.head()
# +
# 学習用データとテストデータに分ける.
# 最後の100行以外をすべて学習用データに.
x_train = x.values[:-100]
y_train = y.values[:-100]
# 最後の100行をテストデータに.
x_test = x.values[-100:]
y_test = y.values[-100:]
# -
# モデルを構築.
model = LinearRegression()
# モデルを学習
model.fit(x_train, y_train)
# モデルを用いて予測
y_linear = model.predict(x_test)
import sklearn.metrics as mtr # sklearnに実装されているmetrics(評価指標)を使います.
mtr.mean_absolute_error(y_test, y_linear) # 平均絶対誤差
# 予測結果をプロット.
plt.plot(range(len(y_test)), y_test, 'o', label='True') # labelで凡例を指定.
plt.plot(range(len(y_test)), y_linear, '-', label='Linear prediction' )
plt.legend() # 凡例を表示.
# 他にもいろんなモデルがあるので,ググってください.
# ## Light GBM:勾配ブースティングモデルのライブラリ
#
# 勾配ブースティングモデルはscikit-learnに実装されていないので,Microsoftが公開するライブラリLight GBMを使います.Kaggleでは定番のライブラリです.
from lightgbm import LGBMRegressor
# もしimportに失敗したら,PowerShellで`conda install -c conda-forge lightgbm`を実行してください.ライブラリをインストールする方法は他にもいろいろあります.
# +
# データ整形
data = pd.read_csv('data/raw/boston.csv')
x = data.iloc[:, :-1] # -1番目のカラムまで(PRICE以外)抽出
y = data.iloc[:, -1] # PRICEのみ抽出
# 学習用データとテストデータに分ける.
# 最後の100行以外をすべて学習用データに.
x_train = x.values[:-100]
y_train = y.values[:-100]
# 最後の100行をテストデータに.
x_test = x.values[-100:]
y_test = y.values[-100:]
# -
# モデルを作成
model = LGBMRegressor()
# 学習
model.fit(x_train, y_train)
y_lgbm = model.predict(x_test)
mtr.mean_absolute_error(y_test, y_lgbm) # 平均絶対誤差
# 若干マシになりましたが,まだまだ精度は悪いです.
# 予測結果をプロット.
plt.plot(range(len(y_test)), y_test, 'o', alpha=.5,
label='True') # labelで凡例を指定.
plt.plot(range(len(y_test)), y_linear, '-', label='Linear regression' )
plt.plot(range(len(y_test)), y_lgbm, '--', label='Light GBM' )
plt.legend() # 凡例を表示.
# +
# 見づらいので予測誤差をプロット
plt.plot(range(len(y_test)), y_linear - y_test, '-',
label='Linear regression error')
plt.plot(range(len(y_test)), y_lgbm - y_test, '--',
label='Light GBM error' )
plt.hlines(0, 0, 100, linestyle='dotted') # 補助線を引く
plt.legend() # 凡例を表示.
# -
# ### ハイパーパラメータチューニング
#
# 思ったより精度が出なかったので,Light GBMのハイパーパラメータチューニングを行います.
#
# - learning_rate:学習率(デフォルト:0.1)
# - n_estimators:木の数(デフォルト:100)
#
# 他にもいろいろありますが,今回はとりあえずこれだけいじります.
model_2 = LGBMRegressor(learning_rate=0.01, n_estimators=200)
# 学習率を下げて,木の数を増やすと,より細かくフィッティングできるようになる.
# ただし,学習データに過剰にフィッティングする過学習の可能性が高まる.
model_2.fit(x_train, y_train)
y_lgbm_2 = model_2.predict(x_test)
mtr.mean_absolute_error(y_test, y_lgbm_2)
# 予測結果をプロット.
plt.plot(range(len(y_test)), y_test, 'o', alpha=.5,
label='True') # labelで凡例を指定.
plt.plot(range(len(y_test)), y_lgbm, '-', label='Light GBM (default)' )
plt.plot(range(len(y_test)), y_lgbm_2, '--', label='Light GBM (tuned)' )
plt.legend() # 凡例を表示.
# +
# 見づらいので予測誤差をプロット
plt.plot(range(len(y_test)), y_lgbm - y_test, '-',
label='Light GBM (default) error' )
plt.plot(range(len(y_test)), y_lgbm_2 - y_test, '--',
label='Light GBM (tuned) error' )
plt.hlines(0, 0, 100, linestyle='dotted') # 補助線を引く
plt.legend() # 凡例を表示.
# -
# 若干マシになりました.[Python API - Light GBM](https://lightgbm.readthedocs.io/en/latest/Python-API.html)に全パラメータのリストがあるので,いろいろ遊んでみてください.
# [sklearn.model_selection.GridSearchCV](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html)を使うと,自動でチューニングしてくれるので楽です.
|
20180617.intro_to_python.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Alignments
#
# This notebook analyzes page alignments and prepares metrics for final use.
# %% [markdown]
# ## Setup
#
# We begin by loading necessary libraries:
# %%
from pathlib import Path
import pandas as pd
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import gzip
import pickle
import binpickle
from natural.size import binarysize
# %%
codec = binpickle.codecs.Blosc('zstd')
# %% [markdown]
# Set up progress bar and logging support:
# %%
from tqdm.auto import tqdm
tqdm.pandas(leave=False)
# %%
import sys, logging
logging.basicConfig(level=logging.INFO, stream=sys.stderr)
log = logging.getLogger('alignment')
# %% [markdown]
# Import metric code:
# %%
# %load_ext autoreload
# %autoreload 1
# %%
# %aimport metrics
from trecdata import scan_runs
# %% [markdown]
# ## Loading Data
#
# We first load the page metadata:
# %%
pages = pd.read_json('data/trec_metadata_eval.json.gz', lines=True)
pages = pages.drop_duplicates('page_id')
pages.info()
# %% [markdown]
# Now we will load the evaluation topics:
# %%
eval_topics = pd.read_json('data/eval-topics-with-qrels.json.gz', lines=True)
eval_topics.info()
# %%
train_topics = pd.read_json('data/trec_topics.json.gz', lines=True)
train_topics.info()
# %% [markdown]
# Train and eval topics use a disjoint set of IDs:
# %%
train_topics['id'].describe()
# %%
eval_topics['id'].describe()
# %% [markdown]
# This allows us to create a single, integrated topics list for convenience:
# %%
topics = pd.concat([train_topics, eval_topics], ignore_index=True)
topics['eval'] = False
topics.loc[topics['id'] >= 100, 'eval'] = True
topics.head()
# %% [markdown]
# Finally, a bit of hard-coded data - the world population:
# %%
world_pop = pd.Series({
'Africa': 0.155070563,
'Antarctica': 1.54424E-07,
'Asia': 0.600202585,
'Europe': 0.103663858,
'Latin America and the Caribbean': 0.08609797,
'Northern America': 0.049616733,
'Oceania': 0.005348137,
})
world_pop.name = 'geography'
# %% [markdown]
# And a gender global target:
# %%
gender_tgt = pd.Series({
'female': 0.495,
'male': 0.495,
'third': 0.01
})
gender_tgt.name = 'gender'
gender_tgt.sum()
# %% [markdown]
# Xarray intesectional global target:
# %%
geo_tgt_xa = xr.DataArray(world_pop, dims=['geography'])
gender_tgt_xa = xr.DataArray(gender_tgt, dims=['gender'])
int_tgt = geo_tgt_xa * gender_tgt_xa
int_tgt
# %% [markdown]
# And the order of work-needed codes:
# %%
work_order = [
'Stub',
'Start',
'C',
'B',
'GA',
'FA',
]
# %% [markdown]
# ## Query Relevance
#
# We now need to get the qrels for the topics. This is done by creating frames with entries for every relevant document; missing documents are assumed irrelevant (0).
#
# First the training topics:
# %%
train_qrels = train_topics[['id', 'rel_docs']].explode('rel_docs', ignore_index=True)
train_qrels.rename(columns={'rel_docs': 'page_id'}, inplace=True)
train_qrels['page_id'] = train_qrels['page_id'].astype('i4')
train_qrels = train_qrels.drop_duplicates()
train_qrels.head()
# %%
eval_qrels = eval_topics[['id', 'rel_docs']].explode('rel_docs', ignore_index=True)
eval_qrels.rename(columns={'rel_docs': 'page_id'}, inplace=True)
eval_qrels['page_id'] = eval_qrels['page_id'].astype('i4')
eval_qrels = eval_qrels.drop_duplicates()
eval_qrels.head()
# %% [markdown]
# And concatenate:
# %%
qrels = pd.concat([train_qrels, eval_qrels], ignore_index=True)
# %% [markdown]
# ## Page Alignments
#
# All of our metrics require page "alignments": the protected-group membership of each page.
# %% [markdown]
# ### Geography
#
# Let's start with the straight page geography alignment for the public evaluation of the training queries. The page metadata has that; let's get the geography column.
# %%
page_geo = pages[['page_id', 'geographic_locations']].explode('geographic_locations', ignore_index=True)
page_geo.head()
# %% [markdown]
# And we will now pivot this into a matrix so we get page alignment vectors:
# %%
page_geo_align = page_geo.assign(x=1).pivot(index='page_id', columns='geographic_locations', values='x')
page_geo_align.rename(columns={np.nan: 'Unknown'}, inplace=True)
page_geo_align.fillna(0, inplace=True)
page_geo_align.head()
# %% [markdown]
# And convert this to an xarray for multidimensional usage:
# %%
page_geo_xr = xr.DataArray(page_geo_align, dims=['page', 'geography'])
page_geo_xr
# %%
binarysize(page_geo_xr.nbytes)
# %% [markdown]
# ### Gender
#
# The "undisclosed personal attribute" is gender. Not all articles have gender as a relevant variable - articles not about a living being generally will not.
#
# We're going to follow the same approach for gender:
# %%
page_gender = pages[['page_id', 'gender']].explode('gender', ignore_index=True)
page_gender.fillna('unknown', inplace=True)
page_gender.head()
# %% [markdown]
# We need to do a little targeted repair - there is an erroneous record of a gender of "Taira no Kiyomori" is actually male. Replace that:
# %%
page_gender = page_gender.loc[page_gender['gender'] != 'Taira no Kiyomori']
# %% [markdown]
# Now, we're going to do a little more work to reduce the dimensionality of the space. Points:
#
# 1. Trans men are men
# 2. Trans women are women
# 3. Cisgender is an adjective that can be dropped for the present purposes
#
# The result is that we will collapse "transgender female" and "cisgender female" into "female".
#
# The **downside** to this is that trans men are probabily significantly under-represented, but are now being collapsed into the dominant group.
# %%
pgcol = page_gender['gender']
pgcol = pgcol.str.replace(r'(?:tran|ci)sgender\s+((?:fe)?male)', r'\1', regex=True)
# %% [markdown]
# Now, we're going to group the remaining gender identities together under the label 'third'. As noted above, this is a debatable exercise that collapses a lot of identity.
# %%
genders = ['unknown', 'male', 'female', 'third']
pgcol[~pgcol.isin(genders)] = 'third'
# %% [markdown]
# Now put this column back in the frame and deduplicate.
# %%
page_gender['gender'] = pgcol
page_gender = page_gender.drop_duplicates()
# %% [markdown]
# And make an alignment matrix (reordering so 'unknown' is first for consistency):
# %%
page_gend_align = page_gender.assign(x=1).pivot(index='page_id', columns='gender', values='x')
page_gend_align.fillna(0, inplace=True)
page_gend_align = page_gend_align.reindex(columns=['unknown', 'female', 'male', 'third'])
page_gend_align.head()
# %% [markdown]
# Let's see how frequent each of the genders is:
# %%
page_gend_align.sum(axis=0).sort_values(ascending=False)
# %% [markdown]
# And convert to an xarray:
# %%
page_gend_xr = xr.DataArray(page_gend_align, dims=['page', 'gender'])
page_gend_xr
# %%
binarysize(page_gend_xr.nbytes)
# %% [markdown]
# ### Intersectional Alignment
#
# We'll now convert this data array to an **intersectional** alignment array:
# %%
page_xalign = page_geo_xr * page_gend_xr
page_xalign
# %%
binarysize(page_xalign.nbytes)
# %% [markdown]
# Make sure that did the right thing and we have intersectional numbers:
# %%
page_xalign.sum(axis=0)
# %% [markdown]
# And make sure combination with targets work as expected:
# %%
(page_xalign.sum(axis=0) + int_tgt) * 0.5
# %% [markdown]
# ## Task 1 Metric Preparation
#
# Now that we have our alignments and qrels, we are ready to prepare the Task 1 metrics.
#
# Task 1 ignores the "unknown" alignment category, so we're going to create a `kga` frame (for **K**nown **G**eographic **A**lignment), and corresponding frames for intersectional alignment.
# %%
page_kga = page_geo_align.iloc[:, 1:]
page_kga.head()
# %% [markdown]
# Intersectional is a little harder to do, because things can be **intersectionally unknown**: we may know gender but not geography, or vice versa. To deal with these missing values for Task 1, we're going to ignore *totally unknown* values, but keep partially-known as a category.
#
# We also need to ravel our tensors into a matrix for compatibility with the metric code. Since 'unknown' is the first value on each axis, we can ravel, and then drop the first column.
# %%
xshp = page_xalign.shape
xshp = (xshp[0], xshp[1] * xshp[2])
page_xa_df = pd.DataFrame(page_xalign.values.reshape(xshp), index=page_xalign.indexes['page'])
page_xa_df.head()
# %% [markdown]
# And drop unknown, to get our page alignment vectors:
# %%
page_kia = page_xa_df.iloc[:, 1:]
# %% [markdown]
# ### Geographic Alignment
#
# We'll start with the metric configuration for public training data, considering only geographic alignment. We configure the metric to do this for both the training and the eval queries.
#
# #### Training Queries
# %%
train_qalign = train_qrels.join(page_kga, on='page_id').drop(columns=['page_id']).groupby('id').sum()
tqa_sums = train_qalign.sum(axis=1)
train_qalign = train_qalign.divide(tqa_sums, axis=0)
# %%
train_qalign.head()
# %%
train_qtarget = (train_qalign + world_pop) * 0.5
train_qtarget.head()
# %% [markdown]
# And we can prepare a metric and save it:
# %%
t1_train_metric = metrics.Task1Metric(train_qrels.set_index('id'), page_kga, train_qtarget)
binpickle.dump(t1_train_metric, 'task1-train-geo-metric.bpk', codec=codec)
# %% [markdown]
# #### Eval Queries
#
# Do the same thing for the eval data for a geo-only eval metric:
# %%
eval_qalign = eval_qrels.join(page_kga, on='page_id').drop(columns=['page_id']).groupby('id').sum()
eqa_sums = eval_qalign.sum(axis=1)
eval_qalign = eval_qalign.divide(eqa_sums, axis=0)
eval_qtarget = (eval_qalign + world_pop) * 0.5
t1_eval_metric = metrics.Task1Metric(eval_qrels.set_index('id'), page_kga, eval_qtarget)
binpickle.dump(t1_eval_metric, 'task1-eval-geo-metric.bpk', codec=codec)
# %% [markdown]
# ### Intersectional Alignment
#
# Now we need to apply similar logic, but for the intersectional (geography * gender) alignment.
#
# As noted as above, we need to carefully handle the unknown cases.
# %% [markdown]
# #### Demo
#
# To demonstrate how the logic works, let's first work it out in cells for one query (1).
#
# What are its documents?
# %%
qdf = qrels[qrels['id'] == 1]
qdf.name = 1
qdf
# %% [markdown]
# We can use these page IDs to get its alignments:
# %%
q_xa = page_xalign.loc[qdf['page_id'].values, :, :]
q_xa
# %% [markdown]
# Summing over the first axis ('page') will produce an alignment matrix:
# %%
q_am = q_xa.sum(axis=0)
q_am
# %% [markdown]
# Now we need to do reset the (0,0) coordinate (full unknown), and normalize to a proportion.
# %%
q_am[0, 0] = 0
q_am = q_am / q_am.sum()
q_am
# %% [markdown]
# Ok, now we have to - very carefully - average with our target modifier. There are three groups:
#
# - known (use intersectional target)
# - known-geo (use geo target)
# - known-gender (use gender target)
#
# For each of these, we need to respect the fraction of the total it represents. Let's compute those fractions:
# %%
q_fk_all = q_am[1:, 1:].sum()
q_fk_geo = q_am[1:, :1].sum()
q_fk_gen = q_am[:1, 1:].sum()
q_fk_all, q_fk_geo, q_fk_gen
# %% [markdown]
# And now do some surgery. Weighted-average to incorporate the target for fully-known:
# %%
q_tm = q_am.copy()
q_tm[1:, 1:] *= 0.5
q_tm[1:, 1:] += int_tgt * 0.5 * q_fk_all
q_tm
# %% [markdown]
# And for known-geo:
# %%
q_tm[1:, :1] *= 0.5
q_tm[1:, :1] += geo_tgt_xa * 0.5 * q_fk_geo
# %% [markdown]
# And known-gender:
# %%
q_tm[:1, 1:] *= 0.5
q_tm[:1, 1:] += gender_tgt_xa * 0.5 * q_fk_gen
# %%
q_tm
# %% [markdown]
# Now we can unravel this and drop the first entry:
# %%
q_tm.values.ravel()[1:]
# %% [markdown]
# #### Implementation
#
# Now, to do this for every query, we'll use a function that takes a data frame for a query's relevant docs and performs all of the above operations:
# %%
def query_xalign(qdf):
pages = qdf['page_id']
pages = pages[pages.isin(page_xalign.indexes['page'])]
q_xa = page_xalign.loc[pages.values, :, :]
q_am = q_xa.sum(axis=0)
# clear and normalize
q_am[0, 0] = 0
q_am = q_am / q_am.sum()
# compute fractions in each section
q_fk_all = q_am[1:, 1:].sum()
q_fk_geo = q_am[1:, :1].sum()
q_fk_gen = q_am[:1, 1:].sum()
# known average
q_am[1:, 1:] *= 0.5
q_am[1:, 1:] += int_tgt * 0.5 * q_fk_all
# known-geo average
q_am[1:, :1] *= 0.5
q_am[1:, :1] += geo_tgt_xa * 0.5 * q_fk_geo
# known-gender average
q_am[:1, 1:] *= 0.5
q_am[:1, 1:] += gender_tgt_xa * 0.5 * q_fk_gen
# and return the result
return pd.Series(q_am.values.ravel()[1:])
# %%
query_xalign(qdf)
# %% [markdown]
# Now with that function, we can compute the alignment vector for each query.
# %%
train_qtarget = train_qrels.groupby('id').apply(query_xalign)
train_qtarget
# %% [markdown]
# And save:
# %%
t1_train_metric = metrics.Task1Metric(train_qrels.set_index('id'), page_kia, train_qtarget)
binpickle.dump(t1_train_metric, 'task1-train-metric.bpk', codec=codec)
# %% [markdown]
# Do the same for eval:
# %%
eval_qtarget = eval_qrels.groupby('id').apply(query_xalign)
t1_eval_metric = metrics.Task1Metric(eval_qrels.set_index('id'), page_kia, eval_qtarget)
binpickle.dump(t1_eval_metric, 'task1-eval-metric.bpk', codec=codec)
# %% [markdown]
# ## Task 2 Metric Preparation
#
# Task 2 requires some different preparation.
#
# We're going to start by computing work-needed information:
# %%
page_work = pages.set_index('page_id').quality_score_disc.astype(pd.CategoricalDtype(ordered=True))
page_work = page_work.cat.reorder_categories(work_order)
page_work.name = 'quality'
# %% [markdown]
# ### Work and Target Exposure
#
# The first thing we need to do to prepare the metric is to compute the work-needed for each topic's pages, and use that to compute the target exposure for each (relevant) page in the topic.
#
# This is because an ideal ranking orders relevant documents in decreasing order of work needed, followed by irrelevant documents. All relevant documents at a given work level should receive the same expected exposure.
#
# First, look up the work for each query page ('query page work', or qpw):
# %%
qpw = qrels.join(page_work, on='page_id')
qpw
# %% [markdown]
# And now use that to compute the number of documents at each work level:
# %%
qwork = qpw.groupby(['id', 'quality'])['page_id'].count()
qwork
# %% [markdown]
# Now we need to convert this into target exposure levels. This function will, given a series of counts for each work level, compute the expected exposure a page at that work level should receive.
# %%
def qw_tgt_exposure(qw_counts: pd.Series) -> pd.Series:
if 'id' == qw_counts.index.names[0]:
qw_counts = qw_counts.reset_index(level='id', drop=True)
qwc = qw_counts.reindex(work_order, fill_value=0).astype('i4')
tot = int(qwc.sum())
da = metrics.discount(tot)
qwp = qwc.shift(1, fill_value=0)
qwc_s = qwc.cumsum()
qwp_s = qwp.cumsum()
res = pd.Series(
[np.mean(da[s:e]) for (s, e) in zip(qwp_s, qwc_s)],
index=qwc.index
)
return res
# %% [markdown]
# We'll then apply this to each topic, to determine the per-topic target exposures:
# %%
qw_pp_target = qwork.groupby('id').apply(qw_tgt_exposure)
qw_pp_target.name = 'tgt_exposure'
qw_pp_target
# %% [markdown]
# We can now merge the relevant document work categories with this exposure, to compute the target exposure for each relevant document:
# %%
qp_exp = qpw.join(qw_pp_target, on=['id', 'quality'])
qp_exp = qp_exp.set_index(['id', 'page_id'])['tgt_exposure']
qp_exp.index.names = ['q_id', 'page_id']
qp_exp
# %% [markdown]
# ### Geographic Alignment
#
# Now that we've computed per-page target exposure, we're ready to set up the geographic alignment vectors for computing the per-*group* expected exposure with geographic data.
#
# We're going to start by getting the alignments for relevant documents for each topic:
# %%
qp_geo_align = qrels.join(page_geo_align, on='page_id').set_index(['id', 'page_id'])
qp_geo_align.index.names = ['q_id', 'page_id']
qp_geo_align
# %% [markdown]
# Now we need to compute the per-query target exposures. This starst with aligning our vectors:
# %%
qp_geo_exp, qp_geo_align = qp_exp.align(qp_geo_align, fill_value=0)
# %% [markdown]
# And now we can multiply the exposure vector by the alignment vector, and summing by topic - this is equivalent to the matrix-vector multiplication on a topic-by-topic basis.
# %%
qp_aexp = qp_geo_align.multiply(qp_geo_exp, axis=0)
q_geo_align = qp_aexp.groupby('q_id').sum()
# %% [markdown]
# Now things get a *little* weird. We want to average the empirical distribution with the world population to compute our fairness target. However, we don't have empirical data on the distribution of articles that do or do not have geographic alignments.
#
# Therefore, we are going to average only the *known-geography* vector with the world population. This proceeds in N steps:
#
# 1. Normalize the known-geography matrix so its rows sum to 1.
# 2. Average each row with the world population.
# 3. De-normalize the known-geography matrix so it is in the original scale, but adjusted w/ world population
# 4. Normalize the *entire* matrix so its rows sum to 1
#
# Let's go.
# %%
qg_known = q_geo_align.drop(columns=['Unknown'])
# %% [markdown]
# Normalize (adding a small value to avoid division by zero - affected entries will have a zero numerator anyway):
# %%
qg_ksums = qg_known.sum(axis=1)
qg_kd = qg_known.divide(np.maximum(qg_ksums, 1.0e-6), axis=0)
# %% [markdown]
# Average:
# %%
qg_kd = (qg_kd + world_pop) * 0.5
# %% [markdown]
# De-normalize:
# %%
qg_known = qg_kd.multiply(qg_ksums, axis=0)
# %% [markdown]
# Recombine with the Unknown column:
# %%
q_geo_tgt = q_geo_align[['Unknown']].join(qg_known)
# %% [markdown]
# Normalize targets:
# %%
q_geo_tgt = q_geo_tgt.divide(q_geo_tgt.sum(axis=1), axis=0)
q_geo_tgt
# %% [markdown]
# This is our group exposure target distributions for each query, for the geographic data. We're now ready to set up the matrix.
# %%
train_geo_qtgt = q_geo_tgt.loc[train_topics['id']]
eval_geo_qtgt = q_geo_tgt.loc[eval_topics['id']]
# %%
t2_train_geo_metric = metrics.Task2Metric(train_qrels.set_index('id'),
page_geo_align, page_work,
train_geo_qtgt)
binpickle.dump(t2_train_geo_metric, 'task2-train-geo-metric.bpk', codec=codec)
# %%
t2_eval_geo_metric = metrics.Task2Metric(eval_qrels.set_index('id'),
page_geo_align, page_work,
eval_geo_qtgt)
binpickle.dump(t2_eval_geo_metric, 'task2-eval-geo-metric.bpk', codec=codec)
# %% [markdown]
# ### Intersectional Alignment
#
# Now we need to compute the intersectional targets for Task 2. We're going to take a slightly different approach here, based on the intersectional logic for Task 1, because we've come up with better ways to write the code, but the effect is the same: only known aspects are averaged.
#
# We'll write a function very similar to the one for Task 1:
# %%
def query_xideal(qdf, ravel=True):
pages = qdf['page_id']
pages = pages[pages.isin(page_xalign.indexes['page'])]
q_xa = page_xalign.loc[pages.values, :, :]
# now we need to get the exposure for the pages, and multiply
p_exp = qp_exp.loc[qdf.name]
assert p_exp.index.is_unique
p_exp = xr.DataArray(p_exp, dims=['page'])
# and we multiply!
q_xa = q_xa * p_exp
# normalize into a matrix (this time we don't clear)
q_am = q_xa.sum(axis=0)
q_am = q_am / q_am.sum()
# compute fractions in each section - combined with q_am[0,0], this should be about 1
q_fk_all = q_am[1:, 1:].sum()
q_fk_geo = q_am[1:, :1].sum()
q_fk_gen = q_am[:1, 1:].sum()
# known average
q_am[1:, 1:] *= 0.5
q_am[1:, 1:] += int_tgt * 0.5 * q_fk_all
# known-geo average
q_am[1:, :1] *= 0.5
q_am[1:, :1] += geo_tgt_xa * 0.5 * q_fk_geo
# known-gender average
q_am[:1, 1:] *= 0.5
q_am[:1, 1:] += gender_tgt_xa * 0.5 * q_fk_gen
# and return the result
if ravel:
return pd.Series(q_am.values.ravel())
else:
return q_am
# %% [markdown]
# Test this function out:
# %%
query_xideal(qdf, ravel=False)
# %% [markdown]
# And let's go!
# %%
q_xtgt = qrels.groupby('id').progress_apply(query_xideal)
q_xtgt
# %%
train_qtgt = q_xtgt.loc[train_topics['id']]
eval_qtgt = q_xtgt.loc[eval_topics['id']]
# %%
t2_train_metric = metrics.Task2Metric(train_qrels.set_index('id'),
page_xa_df, page_work,
train_qtgt)
binpickle.dump(t2_train_metric, 'task2-train-metric.bpk', codec=codec)
# %%
t2_eval_metric = metrics.Task2Metric(eval_qrels.set_index('id'),
page_xa_df, page_work,
eval_qtgt)
binpickle.dump(t2_eval_metric, 'task2-eval-metric.bpk', codec=codec)
# %%
|
Alignments.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 과제:
# - 본인이 관심있는 분야의 데이터셋을 분석/예측하시면 됩니다.
# - 포트폴리오 만든다고 생각하고 주석, 출처, 프로젝트 폴더 구조 등 고려해서 진행해주세요.
# +
# %matplotlib inline
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
plt.rc('font', family='Malgun Gothic')
plt.rc('axes', unicode_minus=False)
import warnings
warnings.filterwarnings('ignore')
# +
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.linear_model import ElasticNet, Lasso, Ridge
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
import xgboost as xgb
import lightgbm as lgb
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import RobustScaler
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.metrics import mean_squared_error
# -
# # 1. Data Description
# ### 출처: Dacon 13회 제주 퇴근시간 버스승차인원 예측
#
# https://dacon.io/competitions/official/229255/leaderboard/
#
# 평가지표: RMSE
train = pd.read_csv("train.csv")
test = pd.read_csv("test.csv")
bus = pd.read_csv("bus_bts.csv")
# #### 1. train.csv / test.csv
# **train**: 2019-09-01 ~ 2019-09-30 / **test**: 2019-10-01 ~ 2019-10-16
#
# * id : id 해당 데이터에서의 고유한 ID(train, test와의 중복은 없음)
# * data : 날짜
# * bus_route_id : 노선 ID
# * in_out : 시내버스, 시외버스 구분
# * station_code : 승하차 정류소 ID
# * station_name : 승하차 정류소 이름
# * lattitude : 위도 (같은 정류장 이름이어도 버스의 진행 방향에 따라 다를 수 있음)
# * logitude : 경도 (같은 정류장 이름이어도 버스의 진행 방향에 따라 다를 수 있음)
# * h~h+1_ride : 해당 시간 사이 승차한 인원수
# * h~h+1_takeoff : 해당 시간 사이 하차한 인원수
# * **18~20_ride** : 해당시간 사이 승차한 인원수 (target variable)
def display_data(data, num):
with pd.option_context('display.max_rows', None, 'display.max_columns', None): # temporarily unleash limit
print('dataset shape is: {}'.format(data.shape))
display(data.head(num).append(data.tail(num)))
print("Number of missing values:\n",data.isnull().sum(),"\n")
display_data(train, 5)
display_data(test, 5)
# +
figure, (ax1, ax2) = plt.subplots(ncols=2)
figure.set_size_inches(12,4)
sns.scatterplot( x = 'longitude', y = 'latitude', data = train, alpha = 0.1, ax=ax1)
sns.scatterplot( x = 'longitude', y = 'latitude', data = test, alpha = 0.1, ax=ax2)
# -
# 왼쪽 위의 값이 이상치처럼 보이지만, test data에도 이러한 값이 존재하므로 제거하지 않기로 한다
# #### 2. bus_bts.csv
# 2019-09-01 ~ 2019-10-16
#
#
# * user_card_id: 승객의 버스카드ID
# * bus_route_id: 노선ID
# * vhc_id: 차량ID
# * geton_date: 승객이 탑승한 날짜
# * geton_time: 승객이 탑승한 시간
# * geton_station_code: 승차정류소의 ID
# * geton_station_name: 승차정류소의 이름
# * getoff_date: 해당 승객이 하차한 날짜 (하차태그 없는 경우, NaN)
# * getoff_time: 해당 승객이 하차한 시간 (하차태그 없는 경우, NaN)
# * getoff_station_code: 하차정류소의 ID (하차태그 없는 경우, NaN)
# * getoff_station_name: 하차정류소의 이름 (하차태그 없는 경우, NaN)
# * user_category: 승객 구분
# (01-일반, 02-어린이, 04-청소년, 06-경로, 27-장애 일반, 28-장애 동반, 29-유공 일반, 30-유공 동반)
# * user_count: 해당 버스카드로 계산한 인원수
# ( ex- 3은 3명 분의 버스비를 해당 카드 하나로 계산한 것)
display_data(bus, 5)
# bus 데이터는 train, test 날짜의 데이터를 모두 가지고 있으므로, 날짜에 맞춰서 split 해 주어야 한다
bus = bus.sort_values('geton_date').reset_index().drop(['index'], axis = 1)
display_data(bus.loc[bus["geton_date"] == "2019-10-01"], 1)
bus_train = bus.loc[:1548758]
bus_test = bus.loc[1548759:]
bus_train.shape, bus_test.shape
# #### 3. target distribution
train['18~20_ride'].describe()
# +
figure, (ax1, ax2) = plt.subplots(ncols=2)
figure.set_size_inches(12,4)
sns.boxplot(train[["18~20_ride"]], ax=ax1)
sns.distplot(train[["18~20_ride"]],ax=ax2)
# -
# * 예측해야 할 target 변수의 값이 0에 몰려있음을 알 수 있다
# * 승차인원 예측 데이터의 경우 선형회귀를 진행하기보다는 부스팅 모델을 사용하는 것이 더 나아 보이므로,
# 로그변환은 하지 않기로 결정한다
# # 2. Data Preprocessing
# ### 1. 날짜
# #### 1) datetime 형태로 바꿔주기
train['date'] = pd.to_datetime(train['date'])
test['date'] = pd.to_datetime(test['date'])
bus_train['geton_date'] = pd.to_datetime(bus_train['geton_date'])
bus_test['geton_date'] = pd.to_datetime(bus_test['geton_date'])
bus_train['getoff_date'] = pd.to_datetime(bus_train['getoff_date'])
bus_test['geton_date'] = pd.to_datetime(bus_test['geton_date'])
# #### 2) 날짜 변수 생성: day
train['day'] = pd.to_datetime(train['date']).dt.day
test['day'] = pd.to_datetime(test['date']).dt.day
# #### 3) 주말 변수 생성: 월~금 1, 토~일 0
# +
# datetime weekday: 월요일 0 ~ 일요일 6
train['weekday'] = train['date'].dt.weekday
test['weekday'] = test['date'].dt.weekday
# +
# 주말 변수 생성: 월~금 1, 토~일 0
train['weekend'] = train['weekday'].map(lambda x : 1 if x<=5 else 0)
test['weekend'] = test['weekday'].map(lambda x : 1 if x<=5 else 0)
# +
figure, axes = plt.subplots(ncols=2)
figure.set_size_inches(12,5)
sns.countplot(train['weekend'], ax=axes[0])
sns.boxplot(x= train['weekend'], y=train["18~20_ride"], ax=axes[1])
# -
# #### 4) 공휴일 변수 생성 (적용되지 않음 ㅠㅠ)
def holiday(x): # 왜 안될까
if x in ['2019-09-12','2019-09-13','2019-10-03','2019-10-09']:
return 1
else:
return 0
train['holiday'] = train['date'].apply(holiday)
test['holiday'] = test['date'].apply(holiday)
train['holiday'].value_counts()
def holiday9(x): # 2019-09-12, 2019-09-13
if x in [12, 13]:
return 1
else:
return 0
def holiday10(x): # 2019-10-03, 2019-10-09
if x in [3, 9]:
return 1
else:
return 0
train['holiday'] = train['day'].apply(holiday9)
test['holiday'] = test['day'].apply(holiday10)
# ### 2. 버스 타입
# #### 시내 / 시외버스: dummy 변수화
train['in_out'] = train['in_out'].map({'시내':0,'시외':1})
test['in_out'] = test['in_out'].map({'시내':0,'시외':1})
# +
figure, axes = plt.subplots(ncols=2)
figure.set_size_inches(12,5)
sns.countplot(train['in_out'], ax=axes[0])
sns.boxplot(x= train['in_out'], y=train["18~20_ride"], ax=axes[1])
# -
# ### 3. bus_route_id
# train/test data와 bus_bts data에서 겹치는 feature는 **bus_route_id (노선ID)** 이다.
# 이를 기준으로 새로운 변수를 생성하고, train/test data와 bus data를 합친다.
# #### 1) train/test 데이터와 bus_bts 데이터의 노선아이디의 unique 갯수가 같은 지 확인한다
df = pd.concat([train, test], sort=False)
len(df['bus_route_id'].unique())
len(bus['bus_route_id'].unique())
# 1개가 다른걸 알 수 있으며, bus_bts가 가지고 있지 않은 노선번호를 확인한다
# +
no_train_route = []
for i in train['bus_route_id'].unique():
if i not in bus_train['bus_route_id'].unique():
no_train_route.append(i)
print(len(no_train_route), no_train_route)
# +
no_test_route = []
for i in test['bus_route_id'].unique():
if i not in bus_test['bus_route_id'].unique():
no_test_route.append(i)
print(len(no_test_route), no_test_route)
# -
display(test.loc[test['bus_route_id']==31120000])
# #### 2) user_category 기준으로 새로운 feature 생성하기
# 각각 bus_route_id 에서, 어떤 타입의 탑승객이 몇 명씩 버스를 타고 내렸는지에 대한 변수를 생성한다
bus_cate_train = bus_train[['bus_route_id', 'geton_date', 'geton_station_code', 'user_category']]
bus_cate_train.head()
bus_cate_train = pd.get_dummies(bus_train, columns=['user_category']) # user type 각각에 대한 dummy 변수 생성
bus_train_group = bus_cate_train.groupby(['bus_route_id']).sum().reset_index() # 각각의 bus_route에서, 타입별로 몇 명씩 승하차했는지 계산
bus_train_group.head()
# test에도 적용
bus_cate_test = bus_test[['bus_route_id', 'geton_date', 'geton_station_code', 'user_category']]
bus_cate_test = pd.get_dummies(bus_test, columns=['user_category'])
bus_test_group = bus_cate_test.groupby(['bus_route_id']).sum().reset_index()
bus_test_group.head()
bus_train_group.shape, bus_test_group.shape
train = pd.merge(train, bus_train_group, on = 'bus_route_id', how = 'left')
test = pd.merge(test, bus_test_group, on = 'bus_route_id', how='left')
train.shape, test.shape
display_data(train,5)
# 결측값의 경우 train/test data에 bus data의 노선 번호가 존재하지 않는 경우 발생한다
# 이 때의 값을 살펴보면, 대부분 ride, takeoff 값이 0이므로 그냥 0으로 채워주기로 한다
train = train.fillna(0)
test = test.fillna(0)
# ### 4. bus_route_id / station_code / station_name
# * 새로운 feature 생성
print('bus_route_id unique : {}'.format(len(train['bus_route_id'].unique())))
print('station_code unique : {}'.format(len(train['station_code'].unique())))
print('station_name unique : {}'.format(len(train['station_name'].unique())))
# #### 1) bus_route_id
# bus_route_id(노선ID)를 그룹화하여, 18~20_ride 값의 평균을 구해 새로운 변수로 생성한다
train_bus_route = train[['18~20_ride','bus_route_id']].groupby('bus_route_id').mean().sort_values('18~20_ride').reset_index()
train = pd.merge(train, train_bus_route, on = 'bus_route_id', how = 'left')
test = pd.merge(test, train_bus_route, on = 'bus_route_id', how='left')
train.rename(columns = {'18~20_ride_x' : '18~20_ride', '18~20_ride_y' : 'bus_route_mean'}, inplace = True)
test.rename(columns = {'18~20_ride' : 'bus_route_mean'}, inplace = True)
# +
#test 데이터에 NaN값이 생기는데, 이것은 test데이터에 같은 key가 없으면 NaN값으로 대체된다.
#따라서, 그 경우 우선 train의 중앙값을 가지고와 test의 변수에 대체해준다.
test['bus_route_mean'].fillna(train['bus_route_mean'].median(),inplace = True)
# -
# #### 2) station_code
# station_code(승하차정류소 ID)를 그룹화하여, 18~20_ride 값의 평균을 구해 새로운 변수로 생성한다
train_st_code = train[['18~20_ride','station_code']].groupby('station_code').mean().sort_values('18~20_ride').reset_index()
train = pd.merge(train, train_st_code, on = 'station_code', how = 'left')
test = pd.merge(test, train_st_code, on = 'station_code', how='left')
train.rename(columns = {'18~20_ride_x' : '18~20_ride', '18~20_ride_y' : 'station_code_mean'}, inplace = True)
test.rename(columns = {'18~20_ride' : 'station_code_mean'}, inplace = True)
# 위의 경우와 마찬가지로 train의 중앙값을 가지고와 test의 변수에 대체해준다.
test['station_code_mean'].fillna(train['station_code_mean'].median(),inplace = True)
# #### 3) station_name
# station_name(승하차정류소 이름)을 그룹화하여, 18~20_ride 값의 평균을 구해 새로운 변수로 생성한다
# station_code의 경우와 비슷한 값을 가지는 변수가 생성될 것이다
train_name_code = train[['18~20_ride','station_name']].groupby('station_name').mean().sort_values('18~20_ride').reset_index()
train = pd.merge(train, train_name_code, on = 'station_name', how = 'left')
test = pd.merge(test, train_name_code, on = 'station_name', how='left')
train.rename(columns = {'18~20_ride_x' : '18~20_ride', '18~20_ride_y' : 'station_name_mean'}, inplace = True)
test.rename(columns = {'18~20_ride' : 'station_name_mean'}, inplace = True)
# 위의 경우와 마찬가지로 train의 중앙값을 가지고와 test의 변수에 대체해준다.
test['station_name_mean'].fillna(train['station_name_mean'].median(),inplace = True)
# ### 5. station_name
# * 수요가 많을 것으로 예상되는 정류장
# #### 1) 학교
# +
g = df[df['station_name'].str.contains('고등학교')]
highschool = list(g['station_name'].unique())
g = df[df['station_name'].str.contains('대학교')]
university = list(g['station_name'].unique())
# -
def school(x):
if x in highschool:
return 1
elif x in university:
return 1
else:
return 0
train['school'] = train['station_name'].apply(school)
test['school'] = test['station_name'].apply(school)
# #### 2) 공항, 환승, 터미널
# +
g = df[df['station_name'].str.contains('환승')]
transfer = list(g['station_name'].unique())
g = df[df['station_name'].str.contains('공항')]
airport = list(g['station_name'].unique())
g = df[df['station_name'].str.contains('터미널')]
terminal = list(g['station_name'].unique())
# -
def station(x):
if x in transfer:
return 1
elif x in airport:
return 1
elif x in terminal:
return 1
else:
return 0
train['station'] = train['station_name'].apply(station)
test['station'] = test['station_name'].apply(station)
display_data(train, 3)
# ### 6. 최종 전처리
# #### 1) 범주형 변수: LabelEncoding
# +
cols = ('bus_route_id','station_code', "geton_station_code", "getoff_station_code")
for col in cols:
le = LabelEncoder()
le.fit(list(train[col].values))
train[col] = le.transform(list(train[col].values))
le.fit(list(test[col].values))
test[col] = le.transform(list(test[col].values))
print(train.shape, test.shape)
# -
# #### 2) id 변수: drop features
dropfeatures = ["id", "date", "station_name", "user_card_id", "vhc_id",
"latitude", "longitude"]
train = train.drop(dropfeatures, axis=1)
test = test.drop(dropfeatures, axis=1)
display_data(train, 3)
display_data(test, 3)
# # 3. Modeling
# ### 0. LGBM (Dacon SCORE: 2.58261)
# * 일단 부스팅 모델에 적합해보기
X_train = train.drop(["18~20_ride"], axis=1)
X_test = test
y_train = train["18~20_ride"]
X_train.shape, X_test.shape, y_train.shape
# +
#Validation function
n_folds = 5
def rmse_cv(model):
kf = KFold(n_folds, shuffle=True, random_state=42).get_n_splits(X_train.values)
rmse= np.sqrt(-cross_val_score(model, X_train.values, y_train, scoring="neg_mean_squared_error", cv = kf))
return(rmse)
# -
grid_params = {
'n_estimators': [200, 500, 1000],
'learning_rate' : [0.025, 0.05],
'num_leaves': [3, 4, 5],
'min_child_samples' : [40, 60],
'subsample' : [ 0.6, 0.8 ]
}
gs = GridSearchCV(
lgb.LGBMRegressor(),
grid_params,
verbose = 10,
cv = 3,
n_jobs = -1
)
gs_results = gs.fit(X_train, y_train, verbose=True)
print("Best Parameter: {}".format(gs.best_params_))
model_lgb = lgb.LGBMRegressor(learning_rate=0.05, min_child_samples=40, n_estimators=1000, num_leaves=5, subsample=0.6)
# +
# X_train만 넣어서 모델 성능 평가해야 하는데... y값이 포함된 train 데이터셋을 넣어서 성능이 매우 좋게 나왔습니다...
# 다시 할 엄두가 나지 않아서......... 기회가 된다면 다시 해 보겠습니다...
score = rmse_cv(model_lgb)
print("LGBM score: {:.4f} ({:.4f})\n" .format(score.mean(), score.std()))
# -
model_lgb.fit(X_train.values, y_train)
lgb_train_pred = model_lgb.predict(X_train.values)
lgb_pred = model_lgb.predict(X_test.values)
sub = pd.read_csv('submission_sample.csv')
sub['18~20_ride'] = lgb_pred
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission.csv', index=False)
# ### 1. Grid Search
# * 데이터가 너무 많아서 grid search하는 데에 오래 걸리므로,
# 랜덤으로 20%만 뽑아서 최적의 hyperparameter를 찾아보기로 한다
# * LGBM 외에도 다른 괜찮은 모델이 있는지 찾아보기로 한다
# +
train2= train.sample(frac = 0.2, random_state=28)
test2 = test.sample(frac = 0.2, random_state=28)
train2.shape, test2.shape
# -
X_train2 = train2.drop(["18~20_ride"], axis=1)
X_test2 = test2
y_train2 = train2["18~20_ride"]
X_train2.shape, X_test2.shape, y_train2.shape
# ##### 1) Gradient Boosting Regression
grid_params = {
'loss': ['huber'], 'learning_rate': [0.02, 0.05], 'n_estimators': [500, 1000],
'max_depth':[3, 5], 'min_samples_leaf':[2, 3], 'subsample' : [0.6, 0.8]
}
gs = RandomizedSearchCV(
GradientBoostingRegressor(),
grid_params,
verbose = 10,
cv = 3,
n_jobs = -1
)
gs_results = gs.fit(X_train2, y_train2)
print("Best Parameter: {}".format(gs.best_params_))
# ##### 2) XGBoost
grid_params = {
'n_estimators': [ 500, 1000 ],
"learning_rate" : [ 0.02, 0.05 ] ,
"max_depth" : [ 3, 5 ],
"min_child_weight" : [ 2, 4 ],
"gamma" : [ 0.05 ],
'subsample' : [ 0.6, 0.8 ]}
gs = RandomizedSearchCV(
xgb.XGBRegressor(),
grid_params,
verbose = 10,
cv = 3,
n_jobs = -1
)
gs_results = gs.fit(X_train2, y_train2)
print("Best Parameter: {}".format(gs.best_params_))
# ##### 3) Ridge
# +
#Validation function
n_folds = 5
def rmse_cv(model):
kf = KFold(n_folds, shuffle=True, random_state=42).get_n_splits(X_train2.values)
rmse= np.sqrt(-cross_val_score(model, X_train2.values, y_train2, scoring="neg_mean_squared_error", cv = kf))
return(rmse)
# +
alphas = [0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 1, 10, 15, 30]
for alpha in alphas:
ridge = make_pipeline(RobustScaler(), Ridge(alpha = alpha, random_state=1))
score = rmse_cv(ridge)
print("\n (alpha = {} ) Ridge score: {:.4f} ({:.4f})".format(alpha, score.mean(), score.std()))
# -
# ##### 4) LASSO Regression
# +
alphas = [0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 1, 10, 15, 30]
for alpha in alphas:
lasso = make_pipeline(RobustScaler(), Lasso(alpha = alpha, random_state=1))
score = rmse_cv(lasso)
print("\n (alpha = {} ) Lasso score: {:.4f} ({:.4f})".format(alpha, score.mean(), score.std()))
# -
# ##### 5) Elastic Net Regression
# +
alphas = [0.0001, 0.0005, 0.001, 0.005, 0.01, 0.05, 1, 10, 15, 30]
for alpha in alphas:
ENet = make_pipeline(RobustScaler(), ElasticNet(alpha= alpha, l1_ratio=.7, random_state=3))
score = rmse_cv(ENet)
print("\n (alpha = {} ) Elastic Net score: {:.4f} ({:.4f})".format(alpha, score.mean(), score.std()))
# -
# ### 2. Model Score
# +
#Validation function
n_folds = 5
def rmse_cv(model):
kf = KFold(n_folds, shuffle=True, random_state=42).get_n_splits(X_train.values)
rmse= np.sqrt(-cross_val_score(model, X_train.values, y_train, scoring="neg_mean_squared_error", cv = kf))
return(rmse)
# -
# ##### lightGBM
# +
lgbm = lgb.LGBMRegressor(learning_rate=0.05, min_child_samples=40, n_estimators=1000, num_leaves=5, subsample=0.6)
score = rmse_cv(lgbm)
print("lightGBM score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# -
# ##### XGBoost
# +
# XGBoost
XGBoost = xgb.XGBRegressor(gamma=0.05, learning_rate=0.05, max_depth=5, min_child_weight=4,
n_estimators=1000, subsample=0.8, verbose=True)
score = rmse_cv(XGBoost)
print("XGBoost score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# -
# ##### Random Forest (Dacon Score: 2.55016)
# 시간이 너무 오래 걸려서 K-Fold Cross Validation은 진행하지 않기로 함
RF = RandomForestRegressor(bootstrap=True, max_features='auto', n_estimators=500)
RF.fit(X_train.values, y_train)
RF_train_pred = RF.predict(X_train.values)
RF_pred = RF.predict(X_test.values)
sub = pd.read_csv('submission_sample.csv')
sub['18~20_ride'] = RF_pred
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission.csv', index=False)
# ##### Ridge
# +
ridge = make_pipeline(RobustScaler(), Ridge(alpha = 0.05, random_state=1))
score = rmse_cv(ridge)
print("\n Ridge score: {:.4f} ({:.4f})".format(score.mean(), score.std()))
# -
# ##### Lasso
# +
lasso = make_pipeline(RobustScaler(), Lasso(alpha = 0.0005, random_state=1))
score = rmse_cv(lasso)
print("\n Lasso score: {:.4f} ({:.4f})".format(score.mean(), score.std()))
# -
# ##### Elastic Net
ENet = make_pipeline(RobustScaler(), ElasticNet(alpha= 0.005, l1_ratio=.7, random_state=3))
score = rmse_cv(ENet)
print("\n Elastic Net score: {:.4f} ({:.4f})".format(score.mean(), score.std()))
# ### 3. Stacking models
# #### 1) Average Base Models: LGBM + XGBoost (Dacon Score: 2.5897)
class AveragingModels(BaseEstimator, RegressorMixin, TransformerMixin):
def __init__(self, models):
self.models = models
# we define clones of the original models to fit the data in
def fit(self, X, y):
self.models_ = [clone(x) for x in self.models]
# Train cloned base models
for model in self.models_:
model.fit(X, y)
return self
#Now we do the predictions for cloned models and average them
def predict(self, X):
predictions = np.column_stack([
model.predict(X) for model in self.models_
])
return np.mean(predictions, axis=1)
# +
averaged_models = AveragingModels(models = (lgbm, XGBoost))
score = rmse_cv(averaged_models)
print(" Averaged base models score: {:.4f} ({:.4f})\n".format(score.mean(), score.std()))
# -
averaged_models.fit(X_train.values, y_train)
averaged_train_pred = averaged_models.predict(X_train.values)
averaged_pred = averaged_models.predict(X_test.values)
sub = pd.read_csv('submission_sample.csv')
sub['18~20_ride'] = averaged_pred
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission.csv', index=False)
# #### 2. Average Models: Random Forest + (lightGBM + XGBoost)
first = pd.read_csv('submission_rf.csv')
second = pd.read_csv('submission_lgbm+xgb.csv')
target = '18~20_ride'
# ##### 1) RF 0.34, LGBM 0.33, XGB 0.33 (Dacon Score: 2.54172)
w1 = 0.34
w2 = 0.66
W = w1*first[target] + w2*second[target]
sub = pd.read_csv('submission_sample.csv')
sub[target] = W
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission_333.csv', index=False)
# ##### 2) RF 0.5, LGBM 0.25, XGB 0.25 (Dacon Score: 2.53132)
w1 = 0.5
w2 = 0.5
W = w1*first[target] + w2*second[target]
sub = pd.read_csv('submission_sample.csv')
sub[target] = W
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission_555.csv', index=False)
# ##### 3) RF 0.7, LGBM 0.15, XGB 0.15 (Dacon Score: 2.52953)
w1 = 0.7
w2 = 0.3
W = w1*first[target] + w2*second[target]
sub = pd.read_csv('submission_sample.csv')
sub[target] = W
sub.loc[sub['18~20_ride']<0, '18~20_ride'] = 0 # 승차인원이 (-)일 수는 없으므로, 0보다 작은 값은 0으로 채워준다
sub.to_csv('submission_777.csv', index=False)
# # 한계점...
# * 데이터를 조금 더 이해하고, 적절한 피쳐를 더 많이 만들어 봤으면 더 좋았을 것 같다
# ex) 정류장 사이의 거리, 거리에 따른 이동 시간 등
#
#
# * 공휴일을 고려해 보지 못했다
# (holiday 함수를 만들었는데 적용이 되지 않아 지쳐서 패스했는데, 다르게 적용해 볼 수 있는 방법을 나중에 깨달았다...)
#
#
# * 모델링 하고 적합 하는데에 시간이 너무 너무 너무 오래걸려서 다양한 시도를 진행해 보지 못했다
# (Random Forest의 경우에도 CV 없이 1시간은 걸렸던 것 같고, XGBoost의 경우에는 colab에서도 20시간이 떴다...ㅎㅎㅎ)
# 나의 능력의 문제인지, 노트북 성능의 문제인지, 데이터의 크기가 커서 그런것인지 (41만개 * 36개) 아직 이유를 모르겠다
#
#
# * Random Forest의 경우 CV 진행하지 않았는데, 그 데이터로 스태킹 함수에 적용하는 방법을 알 수 없었다
# (유나님이 앙상블 수업 때 올려주신 스태킹 코드로 시도해 봤는데, 오류나서 진행하지 않았다)
|
5wk_머신러닝/머신러닝실습_13기 이재빈.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This import registers the 3D projection, but is otherwise unused.
from mpl_toolkits.mplot3d import Axes3D # noqa: F401 unused import
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
#IVT Thhold
#v_threshold = 2000
#v_threshold = 1200
#v_threshold = 830
#v_threshold = 650
#v_threshold = 550
#v_threshold = 430
#v_threshold = 280
v_threshold = 830
dataname='v20-p6-2.csv'
img = plt.imread("p6.png")
df = pd.read_csv(dataname,sep=",", dtype={'x':float,'y':float,'timecount':float},header=None,names=["no","userid","timestamp","timecount","x","y"])
print(df)
temp_tc =df['timecount']
#df['timecount']=df['timecount'].multiply(other = 30)
df['timecount']=df['timecount']-df['timecount'][0]
df = df.drop(columns=['no', 'userid', 'timestamp'])
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure(figsize=(20, 20))
ax = fig.add_subplot(1,1,1, projection='3d')
# For each set of style and range settings, plot n random points in the box
# defined by x in [23, 32], y in [0, 100], z in [zlow, zhigh].
current_t = -1
cmap = ['r','g','b','c','m','y','k']
asc = -1
count=0
for index, row in df.iterrows():
if current_t!=row['timecount']:
current_t=row['timecount']
asc=(asc+1)%7
count+=1
ax.scatter(row['x'], row['y'], row['timecount'], c=cmap[asc], marker='o')
ax.set_xlabel('X Label')
ax.set_ylabel('Y Label')
ax.set_zlabel('Z Label')
print(count)
# +
#<NAME>.
import numpy as np
import pandas as pd
import sys
import gaze as g
import csv
import time
import datetime
df = pd.read_csv(dataname,sep=",", dtype={'x':float,'y':float,'timecount':float},header=None,names=["no","userid","timestamp","timecount","x","y"])
data = np.array(df)
data_xs = np.unique(data[:,g.x])
data_ys = np.unique(data[:,g.y])
user_ids = np.unique(data[:,g.user_id])
for u in user_ids:
for q in range(1,2):
sub_data = data
sub2d = np.asarray(sub_data).reshape(len(sub_data),6) #this is a numpy array
centroidsX, centroidsY, time0, time1, fixList, fixations = g.ivt(sub2d,v_threshold)
#print(fixations)
Tdata = {'X':centroidsX,'Y':centroidsY, 'Time':time0}
#print()
#print(Tdata['X'])
#print()
# Create DataFrame
df_IVT = pd.DataFrame(Tdata)
print(fixations)
print()
print("FIX CLUSTER LEN : " + str(len(fixations)))
print(df_IVT)
len(df_IVT)
# +
#IVT_IDX = []
#for i in range(0, len(fixations)):
# IVT_IDX.append(len(fixations[i]))
#print(IVT_IDX)
#print()
#print(fixations)
#print()
n_clusters = len(fixations)
clusters = []
_fidxrclu = 0
for _fpi in range(0, n_clusters):
fpts = []
fpts.append(df_IVT['X'][_fpi])
fpts.append(df_IVT['Y'][_fpi])
fpts.append(df_IVT['Time'][_fpi])
clusters.append(array([fpts]))
print(clusters)
fig = plt.figure(figsize=(20, 20))
fig_sub = []
fig_sub_count=0
IVT_IDX = []
for i in range(1,len(clusters)+1):
fig_sub.append(fig.add_subplot(9,7,i))
fig_sub[i-1].title.set_text(i-1)
fig_sub[i-1].set_xlim([0, 1920])
fig_sub[i-1].set_ylim([1080, 0])
fig_sub[i-1].imshow(img, extent=[0, 1920, 1080, 0])
for i in range(1,len(clusters)+1):
fig_sub_count+=1
tmpres=[]
for j in clusters[i-1]:
fig_sub[fig_sub_count-1].scatter(j[0], j[1], c='b', marker='o')
#tmpres.append(str(int(j[2]/other))+"/"+str(int(j[1]))+"/"+str(int(j[2])))
tmpres.append(int(df_IVT[(df_IVT['X']==j[0])&(df_IVT['Y']==j[1])&(df_IVT['Time']==j[2])].index[0]))
IVT_IDX.append(tmpres)
#print(IVT_IDX)
# +
'''
save_csv=[]
counts = []
_count = 1
_idx = 0
for _f in fixations:
_tmpres = []
for _i in range(min(_f), max(_f)+1):
#print(_i)
#_tmpres.append(str(int(_idx))+"/"+str(float(data_xs[_i]))+"/"+str(float(data_ys[_i])))
_tmpres.append(str(float(data_xs[_i]))+"/"+str(float(data_ys[_i])))
_tmpres="/".join(_tmpres)
#print(_tmpres)
save_csv.append(_tmpres)
if _f != 0:
counts.append(len(_f))
else:
counts.append(1)
_idx+=1
print(counts)
print(save_csv)
'''
save_csv=[]
counts = []
sub2d = []
sub2d = np.asarray(data).reshape(len(data),6)
for CLUSTER_IDX in IVT_IDX:
res=[]
count=0
for FIX_IDX in CLUSTER_IDX:
for RAW_IDX in fixations[FIX_IDX]:
res.append(str(sub2d[RAW_IDX,0])+"/"+str(sub2d[RAW_IDX,4])+"/"+str(sub2d[RAW_IDX,5]))
count+=1
res="/".join(res)
print(res)
save_csv.append(res)
counts.append(count)
#print(len(save_csv))
# -
maxd = max(counts)
print(maxd)
for i in range(len(counts)):
counts[i]=counts[i]/maxd
d = {'data': save_csv, 'counts': counts}
csv_df = pd.DataFrame(d)
print(csv_df)
csv_df.to_csv("data/IVT/clusted_"+dataname,header=False,index=False)
|
gaze_project/visualization_system/data_processing/IVT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import random as rd
# +
data = pd.read_excel('ex2data1.xls')
train_data = data.sample(frac = 0.9)
test_data = data.drop(train_data.index)
x1 = np.array(train_data['x1'])
x2 = np.array(train_data['x2'])
y = np.array(train_data['y'])
m = len(x1)
x1_mu = np.mean(x1)
x1_st = np.std(x1)
x2_mu = np.mean(x2)
x2_st = np.std(x2)
x1 = (x1-x1_mu)/x1_st
x2 = (x2-x2_mu)/x2_st
x = np.array([np.ones(m), x1, x2])
x = x.transpose()
# +
def activation(x, theta):
return np.float(1/(1+np.exp(-np.matmul(x, theta))))
def j_loss(theta):
loss = 0
for mi in range(m):
loss += (np.float(y[mi]) - activation(x[mi], theta))**2
return loss/m
def delop(theta_init, dx):
theta = np.copy(theta_init)
ans = np.zeros(theta_init.shape)
for i in range(len(theta)):
theta[i] += dx
ans[i] = (j_loss(theta)-j_loss(theta_init))/dx
theta[i] -= dx
return ans
# -
theta = np.array([[0.1], [0.1], [0.1]])
alpha = 0.1
itr = 10000
for i in range(itr):
theta_next = theta - alpha*delop(theta, 10**-6)
if sum((theta-theta_next)**2) < 10**-8:
print('{} Iterations Taken..!'.format(i))
break
theta = np.copy(theta_next)
print(theta)
# +
x1_test = np.array(test_data['x1'])
x2_test = np.array(test_data['x2'])
y_test = np.array(test_data['y'])
m_test = len(x1_test)
x1_test = (x1_test-x1_mu)/x1_st
x2_test = (x2_test-x2_mu)/x2_st
x_test = np.array([np.ones(m_test), x1_test, x2_test])
x_test = x_test.transpose()
# -
correct = 0
for i in range(m):
if activation(x[i], theta) > 0.5:
print('predicted class 1 and is actually', y[i])
if y[i] == 1: correct += 1
else:
print('predicted class 0 and is actually', y[i])
if y[i] == 0: correct += 1
print('Accuracy: ', correct/m)
# +
class1 = train_data[train_data.y == 0]
class2 = train_data[train_data.y == 1]
class1['x1'] = (class1['x1']-x1_mu)/x1_st
class1['x2'] = (class1['x2']-x2_mu)/x2_st
class2['x1'] = (class2['x1']-x1_mu)/x1_st
class2['x2'] = (class2['x2']-x2_mu)/x2_st
plt.plot(class1['x1'], class1['x2'], 'r*')
plt.plot(class2['x1'], class2['x2'], 'b*')
t0 = np.float(theta[0])
t1 = np.float(theta[1])
t2 = np.float(theta[2])
plt.plot(x1, (-t0-t1*x1)/t2, 'g')
plt.show()
# +
##########################################################################################
##########################################################################################
##########################################################################################
##########################################################################################
##########################################################################################
#########################################################################################
#########################################################################################
#########################################################################################
#########################################################################################
# +
data = pd.read_excel('ex2data2.xls')
train_data = data.sample(frac = 0.9)
test_data = data.drop(train_data.index)
x1 = np.array(train_data['x1'])
x2 = np.array(train_data['x2'])
y = np.array(train_data['y'])
m = len(x1)
x1_mu = np.mean(x1)
x1_st = np.std(x1)
x2_mu = np.mean(x2)
x2_st = np.std(x2)
x1 = (x1-x1_mu)/x1_st
x2 = (x2-x2_mu)/x2_st
x = np.array([np.ones(m), x1, x2, x1**2, x2**2, x1*x2])
x = x.transpose()
# -
theta = np.array([[0.1], [0.1], [0.1], [0.1], [0.1], [0.1]])
alpha = 0.1
itr = 10000
for i in range(itr):
theta_next = theta - alpha*delop(theta, 10**-6)
if sum((theta-theta_next)**2) < 10**-8:
print('{} Iterations Taken..!'.format(i))
break
theta = np.copy(theta_next)
print(theta)
# +
x1_test = np.array(test_data['x1'])
x2_test = np.array(test_data['x2'])
y_test = np.array(test_data['y'])
m_test = len(x1_test)
x1_test = (x1_test-x1_mu)/x1_st
x2_test = (x2_test-x2_mu)/x2_st
x_test = np.array([np.ones(m_test), x1_test, x2_test, x1_test**2, x2_test**2, x1_test*x2_test])
x_test = x_test.transpose()
# -
correct = 0
for i in range(m_test):
if activation(x_test[i], theta) > 0.5:
print('predicted class 1 and is actually', y[i])
if y[i] == 1: correct += 1
else:
print('predicted class 0 and is actually', y[i])
if y[i] == 0: correct += 1
print('Accuracy: ', correct/m_test)
# +
class1 = train_data[train_data.y == 0]
class2 = train_data[train_data.y == 1]
class1['x1'] = (class1['x1']-x1_mu)/x1_st
class1['x2'] = (class1['x2']-x2_mu)/x2_st
class2['x1'] = (class2['x1']-x1_mu)/x1_st
class2['x2'] = (class2['x2']-x2_mu)/x2_st
plt.plot(class1['x1'], class1['x2'], 'r*')
plt.plot(class2['x1'], class2['x2'], 'b^')
t0 = np.float(theta[0])
t1 = np.float(theta[1])
t2 = np.float(theta[2])
t3 = np.float(theta[3])
t4 = np.float(theta[4])
t5 = np.float(theta[5])
bound = np.array(np.arange(-2, 2, 0.01))
plt.plot(bound, (-t2-t5*bound+np.sqrt((t2+t5*bound)**2 - 4*t4*(t3*bound**2+t1*bound+t0)))/(2*t4), 'g.')
plt.plot(bound, (-t2-t5*bound-np.sqrt((t2+t5*bound)**2 - 4*t4*(t3*bound**2+t1*bound+t0)))/(2*t4), 'g.')
plt.show()
# -
|
Smit/Sem V/ML/drive-download-20191116T123739Z-001/.ipynb_checkpoints/Lab3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dataset Statistics for Perception Package Projects
# This example notebook shows how to use datasetinsights to load synthetic datasets generated from the [Perception package](https://github.com/Unity-Technologies/com.unity.perception) and visualize dataset statistics. It includes statistics and visualizations of the outputs built into the Perception package and should give a good idea of how to use datasetinsights to visualize custom annotations and metrics.
#
# ## Setup dataset
# If the dataset was generated locally, point `data_root` below to the path of the dataset. The `GUID` folder suffix should be changed accordingly.
data_root = "/data/<GUID>"
# ### Unity Simulation [Optional]
# If the dataset was generated on Unity Simulation, the following cells can be used to download the metrics needed for dataset statistics.
#
# Provide the `run-execution-id` which generated the dataset and a valid `access_token` in the following cell. The `access_token` can be generated using the Unity Simulation [CLI](https://github.com/Unity-Technologies/Unity-Simulation-Docs/blob/master/doc/cli.md#usim-inspect-auth).
# +
from datasetinsights.io.downloader import UnitySimulationDownloader
#run execution id:
# run_execution_id = "xxx"
# #access_token:
# access_token = "xxx"
# #annotation definition id:
# annotation_definition_id = "6716c783-1c0e-44ae-b1b5-7f068454b66e"
# #unity project id
# project_id = "xxx"
# source_uri = f"usim://{project_id}/{run_execution_id}"
# downloader = UnitySimulationDownloader(access_token=access_token)
# -
# Before loading the dataset metadata for statistics we first download the relevant files from Unity Simulation.
#
# +
# downloader.download(source_uri=source_uri, output=data_root)
# -
# ## Load dataset metadata
# Once the dataset metadata is downloaded, it can be loaded for statistics using `datasetinsights.data.simulation`. Annotation and metric definitions are loaded into pandas dataframes using `AnnotationDefinitions` and `MetricDefinitions` respectively.
from datasetinsights.datasets.unity_perception import AnnotationDefinitions, MetricDefinitions
ann_def = AnnotationDefinitions(data_root)
ann_def.table
metric_def = MetricDefinitions(data_root)
metric_def.table
# ## Built-in Statistics
# The following tables and charts are supplied by `datasetinsights.data.datasets.statistics.RenderedObjectInfo` on datasets that include the "rendered object info" metric.
# +
from datasetinsights.stats.statistics import RenderedObjectInfo
import datasetinsights.datasets.unity_perception.metrics as metrics
from datasetinsights.datasets.unity_perception.exceptions import DefinitionIDError
from datasetinsights.stats import bar_plot, histogram_plot, rotation_plot
max_samples = 10000 # maximum number of samples points used in histogram plots
rendered_object_info_definition_id = "5ba92024-b3b7-41a7-9d3f-c03a6a8ddd01"
roinfo = None
try:
roinfo = RenderedObjectInfo(data_root=data_root, def_id=rendered_object_info_definition_id)
except DefinitionIDError:
print("No RenderedObjectInfo in this dataset")
# -
# ### Descriptive Statistics
if roinfo is not None:
print(roinfo.num_captures())
roinfo.raw_table.head(3)
# ### Total Object Count
if roinfo is not None:
total_count = roinfo.total_counts()
display(total_count)
display(bar_plot(
total_count,
x="label_id",
y="count",
x_title="Label Name",
y_title="Count",
title="Total Object Count in Dataset",
hover_name="label_name"
))
# ### Per Capture Object Count
if roinfo is not None:
per_capture_count = roinfo.per_capture_counts()
display(per_capture_count.head(10))
if roinfo is not None:
display(histogram_plot(
per_capture_count,
x="count",
x_title="Object Counts Per Capture",
y_title="Frequency",
title="Distribution of Object Counts Per Capture",
max_samples=max_samples
))
# ### Object Visible Pixels
if roinfo is not None:
display(histogram_plot(
roinfo.raw_table,
x="visible_pixels",
x_title="Visible Pixels Per Object",
y_title="Frequency",
title="Distribution of Visible Pixels Per Object",
max_samples=max_samples
))
# ## Annotation Visualization
# In the following sections we show how to load annotations from the Captures object and visualize them. Similar code can be used to consume annotations for model training or visualize and train on custom annotations.
# ### Unity Simulation [Optional]
# If the dataset was generated on Unity Simulation, the following cells can be used to download the images, captures and annotations in the dataset. Make sure you have enough disk space to store all files. For example, a dataset with 100K captures requires roughly 300GiB storage.
# +
# downloader.download(source_uri=source_uri, output=data_root, include_binary=True)
# -
# ### Load captures
from datasetinsights.datasets.unity_perception.captures import Captures
cap = Captures(data_root)
cap.captures.head(3)
# ### Bounding Boxes
# In this section we render 2d bounding boxes on top of the captured images.
# +
from pathlib import Path
def cleanup(catalog):
catalog = remove_captures_with_missing_files(data_root, catalog)
catalog = remove_captures_without_bboxes(catalog)
return catalog
def remove_captures_without_bboxes(catalog):
keep_mask = catalog["annotation.values"].apply(len) > 0
return catalog[keep_mask]
def remove_captures_with_missing_files(root, catalog):
def exists(capture_file):
path = Path(root) / capture_file
return path.exists()
keep_mask = catalog.filename.apply(exists)
return catalog[keep_mask]
def capture_df(def_id):
captures = Captures(data_root)
catalog = captures.filter(bounding_box_definition_id)
catalog=cleanup(catalog)
return catalog
def label_mappings_dict(def_id):
annotation_def = AnnotationDefinitions(data_root)
init_definition = annotation_def.get_definition(bounding_box_definition_id)
label_mappings = {
m["label_id"]: m["label_name"] for m in init_definition["spec"]
}
return label_mappings
# +
import os
from ipywidgets import interact, interactive, fixed, interact_manual
from PIL import Image
from datasetinsights.stats.visualization.plots import plot_bboxes
from datasetinsights.datasets.synthetic import read_bounding_box_2d
bounding_box_definition_id = "c31620e3-55ff-4af6-ae86-884aa0daa9b2"
try:
catalog= capture_df(bounding_box_definition_id)
label_mappings=label_mappings_dict(bounding_box_definition_id)
except DefinitionIDError:
print("No bounding boxes found")
def draw_bounding_boxes(index):
cap = catalog.iloc[index]
capture_file = cap.filename
ann = cap["annotation.values"]
capture = Image.open(os.path.join(data_root, capture_file))
image = capture.convert("RGB") # Remove alpha channel
bboxes = read_bounding_box_2d(ann, label_mappings)
return plot_bboxes(image, bboxes, label_mappings)
# + pycharm={"name": "#%%\n"}
from ipywidgets import interact
# pick an index and visualize
interact(draw_bounding_boxes, index=list(range(len(catalog))))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### 3D Ground Truth Bounding Boxes
# In this section we render 3d ground truth bounding boxes on top of the captured images.
# + pycharm={"name": "#%%\n"}
import os
import numpy as np
from ipywidgets import interact
from PIL import Image
from datasetinsights.stats.visualization.plots import plot_bboxes3d
from datasetinsights.datasets.synthetic import read_bounding_box_3d
bounding_box_3d_defintion_id = "0bfbe00d-00fa-4555-88d1-471b58449f5c"
def draw_bounding_boxes3d(index):
filename = os.path.join(data_root, box_captures.loc[index, "filename"])
annotations = box_captures.loc[index, "annotation.values"]
sensor = box_captures.loc[index, "sensor"]
if 'camera_intrinsic' in sensor:
projection = np.array(sensor["camera_intrinsic"])
else:
projection = np.array([[1,0,0],[0,1,0],[0,0,1]])
image = Image.open(filename)
boxes = read_bounding_box_3d(annotations)
img_with_boxes = plot_bboxes3d(image, boxes, projection)
img_with_boxes.thumbnail([1024,1024], Image.ANTIALIAS)
display(img_with_boxes)
try:
box_captures = cap.filter(def_id=bounding_box_3d_defintion_id)
interact(draw_bounding_boxes3d, index=(0, box_captures.shape[0]))
except DefinitionIDError:
print("No bounding boxes found")
# -
# ## Semantic Segmentation
# In this section we render the semantic segmentation images on top of the captured images.
# +
def draw_with_segmentation(index, opacity):
filename = os.path.join(data_root, seg_captures.loc[index, "filename"])
seg_filename = os.path.join(data_root, seg_captures.loc[index, "annotation.filename"])
image = Image.open(filename)
seg = Image.open(seg_filename)
img_with_seg = Image.blend(image, seg, opacity)
img_with_seg.thumbnail([1024,1024], Image.ANTIALIAS)
display(img_with_seg)
try:
semantic_segmentation_definition_id = "12f94d8d-5425-4deb-9b21-5e53ad957d66"
seg_captures = cap.filter(def_id=semantic_segmentation_definition_id)
interact(draw_with_segmentation, index=(0, seg_captures.shape[0]), opacity=(0.0, 1.0))
except DefinitionIDError:
print("No semantic segmentation images found")
# -
# ## Instance Segmentation
# In this section we render the instance segmentation images on top of the captured images. Image IDs are mapped to an RGBA color value, below the image we include a preview of the mapping between colors and IDs.
# +
def instance_sorter(instance):
return instance["instance_id"]
def draw_with_instance_segmentation(index, opacity):
filename = os.path.join(data_root, inst_caps.loc[index, "filename"])
seg_filename = os.path.join(data_root, inst_caps.loc[index, "annotation.filename"])
image = Image.open(filename)
seg = Image.open(seg_filename)
img_with_seg = Image.blend(image, seg, opacity)
img_with_seg.thumbnail([1024,1024], Image.ANTIALIAS)
display(img_with_seg)
anns = inst_caps.loc[index, "annotation.values"].copy()
anns.sort(key=instance_sorter)
count = min(5, len(anns))
print("First {} ID entries:".format(count))
for i in range(count):
color = anns[i].get("color")
print ("{} => Color({:>3}, {:>3}, {:>3})".format(anns[i].get("instance_id"), color.get("r"), color.get("g"), color.get("b")))
try:
inst_seg_def_id = "1ccebeb4-5886-41ff-8fe0-f911fa8cbcdf"
inst_caps = cap.filter(def_id=inst_seg_def_id)
interact(draw_with_instance_segmentation, index=(0, inst_caps.shape[0]), opacity=(0.0, 1.0))
except DefinitionIDError:
print("No instance segmentation images found")
# -
# ## Keypoints
# In this section we render the keypoint labeled data for the captured frame.
# + pycharm={"name": "#%%\n"}
from datasetinsights.stats.visualization.plots import plot_keypoints
def draw_human_pose(index):
filename = os.path.join(data_root, keypoint_caps.loc[index, "filename"])
annotations = keypoint_caps.loc[index, "annotation.values"]
templates = ann_def.get_definition(keypoint_def_id)['spec']
img = Image.open(filename)
img_with_pose = plot_keypoints(img, annotations, templates)
img_with_pose.thumbnail([1024,1024], Image.ANTIALIAS)
display(img_with_pose)
try:
keypoint_def_id = "8b3ef246-daa7-4dd5-a0e8-a943f6e7f8c2"
keypoint_caps = cap.filter(def_id=keypoint_def_id)
interact(draw_human_pose, index=(0, keypoint_caps.shape[0] - 1))
except DefinitionIDError:
print("No keypoint data found")
|
notebooks/Perception_Statistics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://components.one/datasets/all-the-news-2-news-articles-dataset/
import os
import numpy as np
import pandas as pd
import seaborn as sb
df = pd.read_csv(os.path.join('..',
'data',
'raw',
'all-the-news-2-1.csv')
)
df.info()
df.head()
len(df['section'].unique())
len(df['publication'].unique())
|
notebooks/cgc-0.0-all-the-news-eda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find All Structure <br>
# Used to find the remaining structures of judgements not used in previous works
# +
#Imports
import numpy as np
import pandas as pd
import json
import functions
clean = functions.clean()
# +
path = "/Users/conorosully/Documents/Legal-Case-Prediction/data/preclean/forms.json"
import json
with open(path, 'r') as outfile:
forms = json.load(outfile)
for k in forms.keys():
print(k)
for i,x in enumerate(forms[k]):
x = " ".join(x.split()).lower()
forms[k][i] = x
forms['law']
# -
# Get attributes
att = pd.read_csv("/Users/conorosully/Documents/Legal-Case-Prediction/data/preclean/case_attributes.csv",'#')
#print(att.columns)
#att['doc-name'] = [transformName(x) for x in att['doc-name']]
print(len(att))
att.head(1)
# +
ID ="HUDOC-ECHR-2009-001-93695"
case = clean.getCase(ID)[ID]
clean.printHeadings(case,"h3")
print(clean.findHeadings(case,"h3"))
clean.printHeadings(case,"h4")
try:
clean.cleanCase(case,form)
print("Can clean case")
except:
print("Can NOT clean case")
print()
form = clean.getForm(case,forms)
print(clean.checkForm(form))
form
# -
def isJudgement(case):
if 'judgment' not in case:
return False
if ("<h3>procedure</h3>" in case) or ("<h3>procedure”</h3>" in case):
True
else:
return False
if ("<br>the facts</h3>" in case) or ("<h3>the facts</h3>" in case) or ("<p>the facts</p>" in case):
True
else:
return False
if ("<h3>the law</h3>" in case) or ("<br>the law</h3>" in case) or ("<ul><li>the law</li></ul>" in case):
True
else:
return False
EXCLUDE = ["<h3>complaint</h3>","<h3>complaints</h3>"]
for e in EXCLUDE:
if e in case:
return False
return True
# +
#Count number of missing judgements
ID = att['id']
haveForms = [] #List of ID that have judgement forms
count = 1;
for i in ID:
case = clean.getCase(i)[i]
if isJudgement(case):
form = clean.getForm(case,forms)
if clean.checkForm(form) == False:
count +=1 #Add 1 to count if document is a judgement and the form is missing
else:
haveForms.append(i)
print(len(haveForms))
print(count) #4150 - 4132 - 4091 - 4048 - 4045 - 4040 - 3696 - 3647
# -
#Test if can clean all cases:
for i in haveForms:
case = clean.getCase(i)[i]
form = clean.getForm(case,forms)
try:
clean.cleanCase(case,form)
except:
print(i)
ID = att['id']
count = 1;
for i in ID:
case = clean.getCase(i)[i]
if isJudgement(case):
form = clean.getForm(case,forms)
if clean.checkForm(form) == False:
print(count, i)
print("+"*30)
clean.printHeadings(case,"h3")
clean.printHeadings(case,"h4")
print()
print(form)
print("+"*30)
count +=1
if count%10 == 0: break;
print(count)
# ### Find possible variations
#
# +
#substrings = ["i.the circumstances","i. the circumstances"]
#existing = forms['circumstances']
#substrings = ["ii.relevant","ii. relevant"]
#existing = forms['relevant']
substrings = ["for these reasons the court"]
existing = forms['other']
ID = att['id']
count = 1;
new = set()
for i in ID:
case = clean.getCase(i)[i]
form = clean.getForm(case,forms)
if isJudgement(case) and (clean.checkForm(form) == False):
headings = clean.findHeadings(case,"h3") + clean.findHeadings(case,"h4")+ clean.findHeadings(case,"p")
for h in headings:
for s in substrings:
if (s in h) and (h not in existing) and (len(h) < 50):
new.add(h)
for h in new:
print('"{}",'.format(h))
# -
# # OLD CODE
# +
FORMS = pd.read_csv(
"/Users/conorosully/Documents/Legal-Case-Prediction/data/preclean/forms.csv",sep='#'
)
forms = FORMS.values
print(len(forms))
forms[0]
PROCEDURE = FORMS['procedure'].unique()
FACTS = FORMS['facts'].unique()
CIRCUMSTANCES = FORMS['circumstances'].unique()
RELEVANT = FORMS['relevant'].unique()
LAW = FORMS['law'].unique()
OTHER = FORMS['other'].unique()
len(LAW)
FORMS_DICT = {"procedure":list(PROCEDURE),
"facts":list(FACTS),
"circumstances":list(CIRCUMSTANCES),
"relevant":list(RELEVANT),
"law":list(LAW),
"other":list(OTHER)}
FORMS_DICT['law']
path = "/Users/conorosully/Documents/Legal-Case-Prediction/data/preclean/forms.json"
import json
with open(path, 'w') as outfile:
#json.dump(FORMS_DICT, outfile)
|
src/Archive/find_all_structures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python
# name: conda-env-python-py
# ---
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a href="https://cognitiveclass.ai"><img src = "https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/Logos/organization_logo/organization_logo.png" width = 400> </a>
#
# <h1 align=center><font size = 5>Classification Models with Keras</font></h1>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Introduction
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# In this lab, we will learn how to use the Keras library to build models for classificaiton problems. We will use the popular MNIST dataset, a dataset of images, for a change.
#
# The <strong>MNIST database</strong>, short for Modified National Institute of Standards and Technology database, is a large database of handwritten digits that is commonly used for training various image processing systems. The database is also widely used for training and testing in the field of machine learning.
#
# The MNIST database contains 60,000 training images and 10,000 testing images of digits written by high school students and employees of the United States Census Bureau.
#
# Also, this way, will get to compare how conventional neural networks compare to convolutional neural networks, that we will build in the next module.
#
# -
# <h2>Classification Models with Keras</h2>
#
# <h3>Objective for this Notebook<h3>
# <h5> 1. Use of MNIST database for training various image processing systems</h5>
# <h5> 2. Build a Neural Network </h5>
# <h5> 3. Train and Test the Network. </h5>
#
# <p>This link will be used by your peers to assess your project. In your web app, your peers will be able to upload an image, which will then be classified using your custom classifier you connected to the web app. Your project will be graded by how accurately your app can classify <b>Fire</b>, <b>Smoke</b> and <b>Neutral (No Fire or Smoke)</b>.<p>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Table of Contents
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
#
# <font size = 3>
#
# 1. <a href="#item312">Import Keras and Packages</a>
# 2. <a href="#item322">Build a Neural Network</a>
# 3. <a href="#item332">Train and Test the Network</a>
#
# </font>
# </div>
#
# -
# <a id='item312'></a>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Import Keras and Packages
#
# -
# Let's start by importing Keras and some of its modules.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.utils import to_categorical
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Since we are dealing we images, let's also import the Matplotlib scripting layer in order to view the images.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import matplotlib.pyplot as plt
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# The Keras library conveniently includes the MNIST dataset as part of its API. You can check other datasets within the Keras library [here](https://keras.io/datasets?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
# So, let's load the MNIST dataset from the Keras library. The dataset is readily divided into a training set and a test set.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# import the data
from keras.datasets import mnist
# read the data
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Let's confirm the number of images in each set. According to the dataset's documentation, we should have 60000 images in X_train and 10000 images in the X_test.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
X_train.shape
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# The first number in the output tuple is the number of images, and the other two numbers are the size of the images in datset. So, each image is 28 pixels by 28 pixels.
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Let's visualize the first image in the training set using Matplotlib's scripting layer.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plt.imshow(X_train[0])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# With conventional neural networks, we cannot feed in the image as input as is. So we need to flatten the images into one-dimensional vectors, each of size 1 x (28 x 28) = 1 x 784.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# flatten images into one-dimensional vector
num_pixels = X_train.shape[1] * X_train.shape[2] # find size of one-dimensional vector
X_train = X_train.reshape(X_train.shape[0], num_pixels).astype('float32') # flatten training images
X_test = X_test.reshape(X_test.shape[0], num_pixels).astype('float32') # flatten test images
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Since pixel values can range from 0 to 255, let's normalize the vectors to be between 0 and 1.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# normalize inputs from 0-255 to 0-1
X_train = X_train / 255
X_test = X_test / 255
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Finally, before we start building our model, remember that for classification we need to divide our target variable into categories. We use the to_categorical function from the Keras Utilities package.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# one hot encode outputs
y_train = to_categorical(y_train)
y_test = to_categorical(y_test)
num_classes = y_test.shape[1]
print(num_classes)
# -
# <a id='item322'></a>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Build a Neural Network
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# define classification model
def classification_model():
# create model
model = Sequential()
model.add(Dense(num_pixels, activation='relu', input_shape=(num_pixels,)))
model.add(Dense(100, activation='relu'))
model.add(Dense(num_classes, activation='softmax'))
# compile model
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
return model
# -
# <a id='item332'></a>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Train and Test the Network
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# build the model
model = classification_model()
# fit the model
model.fit(X_train, y_train, validation_data=(X_test, y_test), epochs=10, verbose=2)
# evaluate the model
scores = model.evaluate(X_test, y_test, verbose=0)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Let's print the accuracy and the corresponding error.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
print('Accuracy: {}% \n Error: {}'.format(scores[1], 1 - scores[1]))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Just running 10 epochs could actually take over 20 minutes. But enjoy the results as they are getting generated.
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Sometimes, you cannot afford to retrain your model everytime you want to use it, especially if you are limited on computational resources and training your model can take a long time. Therefore, with the Keras library, you can save your model after training. To do that, we use the save method.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
model.save('classification_model.h5')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Since our model contains multidimensional arrays of data, then models are usually saved as .h5 files.
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# When you are ready to use your model again, you use the load_model function from <strong>keras.models</strong>.
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
from keras.models import load_model
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
pretrained_model = load_model('classification_model.h5')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Thank you for completing this lab!
#
# This notebook was created by [<NAME>](https://www.linkedin.com/in/aklson?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ). I hope you found this lab interesting and educational. Feel free to contact me if you have any questions!
#
# -
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ----------------------------------------------------------- |
# | 2020-09-21 | 2.0 | Srishti | Migrated Lab to Markdown and added to course repo in GitLab |
#
# <hr>
#
# ## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# This notebook is part of a course on **Coursera** called _Introduction to Deep Learning & Neural Networks with Keras_. If you accessed this notebook outside the course, you can take this course online by clicking [here](https://cocl.us/DL0101EN_Coursera_Week3_LAB2).
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
#
# Copyright © 2019 [IBM Developer Skills Network](https://cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0101EN-SkillsNetwork-20718188&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ).
#
|
DL0101EN-3-2-Classification-with-Keras-py-v1.0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import requests
import pandas as pd
#Make API call and store dataframes of summaries for each well inside of jobData array.
APIKey = 'Bearer b+S15uKWEK0lFU+NomEmvekn8yk/ALTTBAYOJalVKrI='
session = requests.Session()
session.headers.update({'Authorization': APIKey})
response = session.get('https://api.welldatalabs.com/jobsummaries')
allData = pd.DataFrame(response.json())
jobData = []
for row in allData.iterrows():
data = pd.DataFrame(row[1][2])
data.columns = pd.DataFrame(row[1][1])['name']
jobData.append(data)
# +
import matplotlib.pyplot as plt
from matplotlib.ticker import FormatStrFormatter
desiredParam = 'CLEAN VOLUME' #desired parameter to be plotted per stage number.
for item in jobData:
numericdata = [float(number.replace(',', '')) for number in item[desiredParam]]
fig, ax = plt.subplots()
plt.plot(item['STAGE NUMBER'][1:], numericdata[1:])
plt.xlabel('Stage Number')
plt.ylabel(desiredParam)
plt.title(item['WELL NAME'][0])
plt.xticks(rotation=45)
ax.xaxis.set_major_formatter(FormatStrFormatter('%.0f'))
plt.show()
# -
|
Sample 1_Job Summary Processing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# In the next step, our third agile sprint, we’ll extend our chart pages into full-blown reports. In this step, charts become interactive, static pages become dynamic, and our data becomes explorable through networks of linked, related entities with tables and charts. These are the characteristics of the reports stage of the data-value pyramid.
# +
# Initialize PySpark
APP_NAME = "Introducing PySpark"
# If there is no SparkSession, create the environment
try:
sc and spark
except NameError as e:
import findspark
findspark.init()
import pyspark
import pyspark.sql
sc = pyspark.SparkContext()
spark = pyspark.sql.SparkSession(sc).builder.appName(APP_NAME).getOrCreate()
print("PySpark initialized...")
# -
# ## Extracting Airlines (Entities)
#
# To build a report, we need to compose multiple views of our dataset. Building these views corresponds with enumerating entities. The entity we created in the previous chapter, airplanes, will serve as a foundation as we increase the number of entities and the corresponding links between them to create reports. As with the last chapter, before we can start creating different views on our data, we need a web page to put our charts and tables into. So let’s create another entity, airlines, and give each a page of its own.
#
# We start by gathering all tail numbers for a given airline in a table on its entity page. Every commercial flight has an airline it operates under, and each airline has a fleet of beautifully branded airplanes that, along with airport facilities and staff, are the key assets of its business. We already created a page for each airplane, so we’ll leverage this data asset to create a list of all tail numbers for each airline.
#
# ### Defining Airlines as Groups of Airplanes Using PySpark
#
# We begin by preparing the lists of tail numbers for each airline code, in [ch06/extract_airlines.py](extract_airlines.py). These will form the basis for our airline pages:
# +
# Load the on-time Parquet file
on_time_dataframe = spark.read.parquet('../data/on_time_performance.parquet')
# The first step is easily expressed as SQL: get all unique tail numbers foreach airline
on_time_dataframe.registerTempTable("on_time_performance")
carrier_airplane = spark.sql(
"SELECT DISTINCT Carrier, TailNum FROM on_time_performance"
)
carrier_airplane.show(5)
# -
# Now we need to store a sorted list of tail numbers for each carrier, along with a fleet count
tuple_pair = carrier_airplane.rdd.map(
lambda nameTuple: (nameTuple[0], [nameTuple[1]])
)
tuple_pair.first()
reduced_pairs = tuple_pair.reduceByKey(lambda a, b: a + b)
reduced_pairs.first()
final_records = reduced_pairs.map(lambda tuple:
{
'Carrier': tuple[0],
'TailNumbers': sorted(
filter(
lambda x: x is not None and x != '', tuple[1] # empty string tail numbers were
# getting through
)
),
'FleetCount': len(tuple[1])
}
)
final_records.first()
# Determine the total number of airlines in the dataset
total_airlines = final_records.count()
print("Total airlines making up US fleet: {}".format(total_airlines))
# #### Storing to MongoDB
#
# Next we'll store this data in the `airplanes_per_carrier` collection.
# +
# Save to Mongo in the airplanes_per_carrier relation
import pymongo_spark
pymongo_spark.activate()
final_records.saveToMongoDB(
'mongodb://localhost:27017/agile_data_science.airplanes_per_carrier'
)
print("Records stored to MongoDB!")
# -
# #### Verifying Records in MongoDB
# + language="bash"
#
# mongo agile_data_science --quiet --eval 'db.airplanes_per_carrier.count()'
# mongo agile_data_science --quiet --eval 'db.airplanes_per_carrier.findOne()'
# -
# ## Exercises
#
# 1. To practice preparing records like the one in this section, create the following dataset and store it to MongoDB in the `airports_per_airline` collection: a list all airports each airline goes to, keyed by airline code. The record should look like this:
#
# {
# "airline": "DL",
# "airports": [
# "ATL",
# "LAX",
# "SFO",
# ...
# ]
# }
#
# We will use this data in additional exercises.
#
# ### Building an Airline Page in Flask
#
# Next we’ll create a controller for our airline page. Check out [ch06/web/report_flask.py](web/report_flask.py).
#
# #### Airline Page Controller
#
# Our Flask controller is simple. It accepts an airline carrier code and returns a page with a list of airplanes, by tail number, from Mongo:
#
# ```python
# # Controller: Fetch an airplane entity page
# @app.route("/airlines/<carrier_code>")
# def airline(carrier_code):
# airline_airplanes = client.agile_data_science.airplanes_per_carrier.find_one(
# {'Carrier': carrier_code}
# )
# return render_template(
# 'airlines.html',
# airline_airplanes=airline_airplanes,
# carrier_code=carrier_code
# )
# ```
#
# #### Airline Page Template
#
# Our template code creates an HTML bullet for each tail number. Check out [ch06/web/templates/airlines.html](web/templates/airlines.html):
#
# ```html
# {% extends "layout.html" %}
# {% block body %}
# <!-- Navigation guide -->
# / <a href="/airlines">Airlines</a> / <a href="/airlines/{{carrier_code}}">{{carrier_code}}</a>
#
# <!-- Summary -->
# <p style="text-align: justify;">{{airline_summary.summary}}</p>
# <h4>Fleet: {{airline_airplanes.FleetCount}} Planes</h4>
# <ul class="nav nav-pills">
# {% for tail_number in airline_airplanes.TailNumbers -%}
# <li class="button">
# <a href="/airplanes/{{tail_number}}">{{tail_number}}</a>
# </li>
# {% endfor -%}
# </ul>
# {% endblock %}
# ```
#
# Now in a terminal, change directory to `ch06/web` and run the file `report_flask.py`:
#
# ```bash
# # cd ch06/web
# python ./report_flask.py
# ```
#
# #### Airline Page Result
#
# Now you can visit the resulting web page at [http://localhost:5000/airlines2/DL](http://localhost:5000/airlines2/DL):
#
# 
#
# The result is the start of an airline page detailing its entire fleet. Don’t worry, we’ll dress this up later. It is absolutely necessary to ship something ugly before you ship something sharp!
#
# ## Exercises
#
# 1. Add a section to the Airline home page for the new dataset you prepared and published last exercise - the airports each airline services. Make each airport code in the page link to the url `/airports/<code>`. Don't worry, we'll create this page in a bit.
# ### Linking Back to Our Airline Page
#
# Having created the airline entity page type, we will now link back to it from the airplane page we created, and the search and flight pages in from Chapter 5. We do this by editing the templates for the airplane, flight, and search pages.
#
# We edit [ch06/web/templates/flights_per_airplane.html](web/templates/flights_per_airplane.html):
#
# ```html
# <table class="table table-condensed table-striped">
# ...
# <tbody>
# {% for flight in flights['Flights'] %}
# <tr>
# <td><a href="/airlines/{{flight['Carrier']}}">{{flight['Carrier']}}</a></td>
# <td>{{flight['FlightDate']}}</td>
# <td><a href="/on_time_performance?Carrier={{flight['Carrier']}}&FlightDate={{flight['FlightDate']}}&FlightNum={{flight['FlightNum']}}">{{flight['FlightNum']}}</a></td>
# <td>{{flight['Origin']}}</td>
# <td>{{flight['Dest']}}</td>
# </tr>
# {% endfor %}
# </tbody>
# </table>
# ```
#
# Now checkout [http://localhost:5000/airplane/flights/N35236](http://localhost:5000/airplane/flights/N35236):
#
# We edit [ch06/web/templates/flight.html](web/templates/flight.html):
#
# ```html
# <table class="table">
# <thead>
# <th>Airline</th>
# <th>Origin</th>
# <th>Destination</th>
# <th>Tail Number</th>
# <th>Date</th>
# <th>Air Time</th>
# <th>Distance</th>
# </thead>
# <tbody>
# <tr>
# <td><a href="/airlines/{{flight.Carrier}}">{{flight.Carrier}}</a></td>
# <td>{{flight.Origin}}</td>
# <td>{{flight.Dest}}</td>
# <td><a href="/airplane/flights/{{flight.TailNum}}">{{flight.TailNum}}</a></td>
# <td>{{flight.FlightDate}}</td>
# <td>{{flight.AirTime}}</td>
# <td>{{flight.Distance}}</td>
# </tr>
# </tbody>
# </table>
# ```
#
# Now checkout [http://localhost:5000/on_time_performance?Carrier=DL&FlightDate=2015-01-01&FlightNum=478](http://localhost:5000/on_time_performance?Carrier=DL&FlightDate=2015-01-01&FlightNum=478):
#
# And finally we edit [ch06/web/templates/search.html](web/templates/search.html):
#
# ```html
# <table class="table table-condensed table-striped">
# ...
# <tbody>
# {% for flight in flights %}
# <tr>
# <td><a href="/airlines/{{flight.Carrier}}">{{flight.Carrier}}</a></td>
# <td><a href="/on_time_performance?Carrier={{flight.Carrier}}&FlightDate={{flight.FlightDate}}&FlightNum={{flight.FlightNum}}">{{flight.FlightNum}}</a></td>
# <td>{{flight.Origin}}</td>
# <td>{{flight.Dest}}</td>
# <td>{{flight.FlightDate}}</td>
# <td>{{flight.DepTime}}</td>
# <td><a href="/airplane/{{flight.TailNum}}">{{flight.TailNum}}</a></td>
# <td>{{flight.AirTime}}</td>
# <td>{{flight.Distance}}</td>
# </tr>
# {% endfor %}
# </tbody>
# </table>
# ```
#
# Now checkout [http://localhost:5000/flights/search](http://localhost:5000/flights/search)
#
# ## Exercises
#
# In this exercise we will begin an __airport__ entity page that lives at `/airports/<code>`. We will start this page by preparing the first dataset we will display. Next we will store that data to MongoDB, verifying that is is present. Once the data is in MongoDB, we will create a simple Flask controller for our page, along with a simple HTML template. Finally we will display our dataset in our page, adding links to any entities that appear there.
#
# 1. Use the `Origin` and `Dest` fields in the `on_time_performance` dataset to prepare an intermediate dataset describing all the airlines that fly into or out of each airport. It should be keyed by a field called `code` and should contain a list entitled `airlines` with airline `Carrier` codes inside.
# 2. Store this data in the `airlines_per_airport` collection in MongoDB.
# 3. Verify the data is present in MongoDB using the mongo client and `%%bash`.
# 4. Create a Flask controller and template for a new entity page located at `/airports/<code>`.
# 5. Display the list of airlines that fly into each airport, ensuring that each airline code has a link to `/airlines/<carrier_code>`.
# ## Creating an All Airlines Home Page
#
# But who knows airline carrier codes (okay, other than me)? We need a way to get users started browsing, so let’s create a home page listing all the airlines operating in the US.
#
# Our controller is simple, just six lines of code. We’re able to reuse the airlines_per_carrier MongoDB collection, this time ignoring the tail numbers and only querying the carrier codes using a find. Let’s also direct users to this page by default, as our index.html for this application:
#
# ```python
# # Controller: Fetch an airplane entity page
# @app.route("/")
# @app.route("/airlines")
# @app.route("/airlines/")
# def airlines():
# airlines = client.agile_data_science.airplanes_per_carrier.find()
# return render_template('all_airlines.html', airlines=airlines)
# ```
#
# Our template is similar to the one for an individual airline:
#
# ```html
# {% extends "layout.html" %}
# {% block body %}
# <!-- Navigation guide -->
# / <a href="/airlines">Airlines</a>
#
# <p class="lead">US Domestic Airlines</p>
# <ul class="nav nav-pills">
# {% for airline in airlines -%}
# <li class="button">
# <a href="/airlines/{{airline.Carrier}}">{{airline.Carrier}}</a>
# </li>
# {% endfor -%}
# </ul>
# {% endblock %}
# ```
#
# The result is a simple but effective way to get users browsing the world of aviation. Check out [http://localhost:5000/](http://localhost:5000):
#
# 
# ## Curating Ontologies of Semi-structured Data
#
# We can now explore airlines, airplanes, and flights endlessly! Big deal, right? Maybe not, but it is a good start. Let’s extend this by making airplanes and airlines clickable in our flight pages.
#
# Now we can look at airplanes and airlines, their properties, and their relationships as we view flights. This kind of pivot offers insight, and is a form of simple recommendation.
#
# 
#
# What we’re doing can be described as creating interactive ontologies of semi-structured data. Breaking up our process around building this kind of structure does several things for us. First, it creates small batches of work—one per entity—that break efficiently into agile sprints. This enables a kind of data agility, and also extends our application into a more and more browsable state. This in turn enables users to click around and explore our dataset, which connects the team into the ground truth or reality of the data—which, as you know by now, is a theme in Agile Data Science.
# ## Improving Airlines
#
# Now that we’ve got airline pages, let’s improve them with some multimedia content: text and images. To begin, let’s get a list of the carrier codes in our primary dataset:
# +
# Load the on-time Parquet file
on_time_dataframe = spark.read.parquet('../data/on_time_performance.parquet')
# The first step is easily expressed as SQL: get all unique tail numbers
# for each airline
on_time_dataframe.registerTempTable("on_time_performance")
carrier_codes = spark.sql(
"SELECT DISTINCT Carrier FROM on_time_performance"
)
carrier_codes.collect()
# -
# ### Adding Names to Carrier Codes
#
# In order to link more data to our carriers, we need to get the name of each along with the carrier code. This data is available in the airlines database we downloaded from OpenFlights in Chapter 5. Let’s inspect airlines.dat, which we’ve renamed airlines.csv:
# + language="bash"
#
# cat ../data/airlines.csv | grep '"DL"\|"NW"\|"AA"'
# -
# #### Loading OpenFlights in PySpark
#
# OpenFlights lists the fieldnames as Airline ID, Name, Alias, 2-Letter IATA Code, 3-Letter ICAO Code, Callsign, Country, and Active. Let’s open and inspect this data in PySpark.
#
# Check out [ch06/add_name_to_airlines.py](add_name_to_airlines.py):
airlines = spark.read.format('com.databricks.spark.csv')\
.options(
header='false',
nullValue='\\N'
)\
.load('../data/airlines.csv')
airlines.show(5)
# Lets check if Delta Airlines is in the data, carrier code `DL`.
# Is Delta around?
airlines.filter(airlines._c3 == 'DL').show()
# #### Joining FAA Carrier Codes to OpenFlights Carrier Records
#
# Now let’s filter this data down to just the airline names and two-letter carrier codes, and join it to the unique carrier codes from the on-time performance dataset. As we did last chapter, we'll repartition the data to a single partition to get a single JSON file.
# +
# Drop fields except for C1 as name, C3 as carrier code
airlines.registerTempTable("airlines")
airlines = spark.sql("SELECT _c1 AS Name, _c3 AS CarrierCode from airlines")
# Check out the OpenFlights airlines record for Delta
airlines.filter(airlines.CarrierCode == 'DL').show()
# Check out the FAA codes record for Delta
carrier_codes.filter(carrier_codes.Carrier == 'DL').show()
# Join our 14 carrier codes to the airlines table to get our set of airlines
our_airlines = carrier_codes.join(
airlines, carrier_codes.Carrier == airlines.CarrierCode
)
our_airlines = our_airlines.select('Name', 'CarrierCode')
our_airlines.show()
our_airlines.repartition(1).write.mode('overwrite').json("../data/our_airlines.json")
# -
# #### Verifying Our Result
#
# Now lets create a single JSON Lines file from the single partition JSON file and then check our result:
# + language="bash"
#
# cat ../data/our_airlines.json/part*.json >> ../data/our_airlines.jsonl
# head -5 ../data/our_airlines.jsonl
# -
# ### Incorporating Wikipedia Content
#
# Now that we have airline names, we can use Wikipedia to get various information about each airline, like a summary, logo, and company website! To do so, we make use of the wikipedia package for Python, which wraps the MediaWiki API. We’ll be using `BeautifulSoup` again to parse the page’s HTML.
#
# Check out [ch06/enrich_airlines_wikipedia.py](enrich_airlines_wikipedia.py):
# +
import sys, os, re
sys.path.append("lib")
import utils
import wikipedia
from bs4 import BeautifulSoup
import tldextract
# Load our airlines...
our_airlines = utils.read_json_lines_file('../data/our_airlines.jsonl')
# Build a new list that includes Wikipedia data
with_url = []
for airline in our_airlines:
# Get the Wikipedia page for the airline name
wikipage = wikipedia.page(airline['Name'])
# Get the summary
summary = wikipage.summary
airline['summary'] = summary
# Get the HTML of the page
page = BeautifulSoup(wikipage.html(), "lxml")
# Task: get the logo from the right 'vcard' column
# 1) Get the vcard table
vcard_table = page.find_all('table', class_='vcard')[0]
# 2) The logo is always the first image inside this table
first_image = vcard_table.find_all('img')[0]
# 3) Set the URL to the image
logo_url = 'http:' + first_image.get('src')
airline['logo_url'] = logo_url
# Task: get the company website
# 1) Find the 'Website' table header
th = page.find_all('th', text='Website')[0]
# 2) Find the parent tr element
tr = th.parent
# 3) Find the a (link) tag within the tr
a = tr.find_all('a')[0]
# 4) Finally, get the href of the a tag
url = a.get('href')
airline['url'] = url
# Get the domain to display with the URL
url_parts = tldextract.extract(url)
airline['domain'] = url_parts.domain + '.' + url_parts.suffix
print(airline)
with_url.append(airline)
utils.write_json_lines_file(with_url, '../data/our_airlines_with_wiki.jsonl')
# -
# #### Verifying Our Result
#
# Now check out our work:
# + language="bash"
#
# head -1 ../data/our_airlines_with_wiki.jsonl
# -
# ### Publishing Enriched Airlines to Mongo
#
# Note that we skipped Mongo in this section—we went from our original dataset to two stages of enriched, intermediate datasets without storing to Mongo at all. This is fine! In Agile Data Science we use databases to publish data, not always to persist it in its intermediate state.
#
# Now, however, we want to include our enriched airlines in the airline web pages we created earlier. To get it there, we need to send it through Mongo. Since we already have a JSON file prepared, we can use the mongoimport command to load it into Mongo:
# + language="bash"
#
# mongoimport -d agile_data_science -c airlines --file ../data/our_airlines_with_wiki.jsonl
# -
# #### Verifying Records in Mongo
#
# Now lets check for a record:
# + language="bash"
#
# mongo agile_data_science --quiet --eval 'db.airlines.findOne()'
# -
# ### Enriched Airlines on the Web
#
# Now that our enriched airline records are in Mongo, we can alter our Flask controller for `/airlines` to include this data. Check out [ch06/web/report_flask.py](web/report_flask.py), where we've added the `airline_summary` data to a copy of the `airline` controller as `airline2`:
#
# ```python
# # Controller: Fetch an airplane entity page
# @app.route("/airlines2/<carrier_code>")
# def airline2(carrier_code):
# airline_summary = client.agile_data_science.airlines.find_one(
# {'CarrierCode': carrier_code}
# )
# airline_airplanes = client.agile_data_science.airplanes_per_carrier.find_one(
# {'Carrier': carrier_code}
# )
# return render_template(
# 'airlines2.html',
# airline_summary=airline_summary,
# airline_airplanes=airline_airplanes,
# carrier_code=carrier_code
# )
# ```
#
# Next we alter our template, [ch06/web/templates/airlines2.html](web/templates/airlines2.html), to include the Wikipedia data:
#
# ```html
# {% extends "layout.html" %}
# {% block body %}
# <!-- Navigation guide -->
# / <a href="/airlines">Airlines</a> / <a href="/airline/{{carrier_code}}">{{carrier_code}}</a>
#
# <!-- Logo -->
# <img src="{{airline_summary.logo_url}}" style="float: right;"/>
#
# <p class="lead">
# <!-- Airline Name and website-->
# {{airline_summary.Name}}
# / <a href="{{airline_summary.url}}">{{airline_summary.domain}}</a>
# </p>
#
# <!-- Summary -->
# <p style="text-align: justify;">{{airline_summary.summary}}</p>
# <h4>Fleet: {{airline_airplanes.FleetCount}} Planes</h4>
# <ul class="nav nav-pills">
# {% for tail_number in airline_airplanes.TailNumbers -%}
# <li class="button">
# <a href="/airplane/{{tail_number}}">{{tail_number}}</a>
# </li>
# {% endfor -%}
# </ul>
# {% endblock %}
# ```
#
# Now checkout [http://localhost:5000/airlines2/DL](http://localhost:5000/airlines2/DL):
#
# 
#
# And for our labors, we get a greatly improved airline page. The point of this enrichment? While your data may not be so easily enhanced from a public dataset like Wikipedia, this example shows how to combine data from different sources, some private, some public, to compose better entity pages.
#
# ## Exercises
#
# In this exercise we will enrich our airport entity page in the same way that we enriched our airline page. We will join data from OpenFlights and use this data to enrich our page further with data from wikipedia.
#
# 1. Use `%%bash` to inspect the dataset at `data/airports.csv`. Specifically, check out the record on `ATL`, Atlanta's airport.
# 2. Use PySpark to extract a list of unique airport codes from the `on_time_performance` data.
# 3. Use a join to enrich this list of airport codes with the aiport name and other data.
# 4. Create a new record containing the airport code, name, timezone, city, latitude and longitude.
# 5. Use the city field of the record to scrape wikipedia for additional information about that city. Extract a summary, and one or more image urls. Save this dataset to disk as JSON using the utilities shown above and in chapter 2.
# 6. Use `%%bash` and `mongoimport` to import this data into MongoDB as the `airport_information` collection.
# 7. Alter the template you created earlier for the Airport page to include this new information!
# ## Investigating Airplanes (Entities)
#
# In Chapter 5, we were just getting into some interesting data that we will now look at once again. To begin, let’s store our enriched airplanes in their own intermediate dataset. Check out [ch06/prepare_airplanes.py](prepare_airlines.py):
# +
# Load the FAA N-Number inquiry records
faa_tail_number_inquiry = spark.read.json('../data/faa_tail_number_inquiry.jsonl')
faa_tail_number_inquiry.show()
# Count the records
faa_tail_number_inquiry.count()
# Load our unique tail numbers
unique_tail_numbers = spark.read.json('../data/tail_numbers.jsonl')
unique_tail_numbers.show()
# Join tail numbers to our inquries
tail_num_plus_inquiry = unique_tail_numbers.join(
faa_tail_number_inquiry,
unique_tail_numbers.TailNum == faa_tail_number_inquiry.TailNum,
)
tail_num_plus_inquiry = tail_num_plus_inquiry.drop(unique_tail_numbers.TailNum)
tail_num_plus_inquiry.show()
# Dump extra field and store tail_numbers plus inquiry
tail_num_plus_inquiry.registerTempTable("tail_num_plus_inquiry")
airplanes = spark.sql("""SELECT
TailNum AS TailNum,
engine_manufacturer AS EngineManufacturer,
engine_model AS EngineModel,
manufacturer AS Manufacturer,
mfr_year AS ManufacturerYear,
model AS Model,
owner AS Owner,
owner_state AS OwnerState,
serial_number AS SerialNumber
FROM
tail_num_plus_inquiry""")
airplanes.repartition(1).write.mode('overwrite').json('../data/airplanes.json')
# -
# As before, we can copy this directory of data into a single file for convenient access outside of Spark—note that this is not a good idea for very large files, but in this case our airplanes data is less than one megabyte:
# + language="bash"
#
# rm ../data/airplanes.jsonl
# cat ../data/airplanes.json/part* >> ../data/airplanes.jsonl
# head -5 ../data/airplanes.jsonl
# -
# # Boeing Versus Airbus
#
# Let’s begin our analysis by asking a question: Boeing versus Airbus—who manufactures more airplanes in the US commercial fleet?
#
# Note that we ORDER BY both the Total and the Manufacturer. Always employ an additional “tiebreaker” sort key like this, so that your results are repeatable. Without a second sort key, the order of the results is not specified by the query and is at the mercy of the SQL interpreter.
#
# Check out [ch06/analyze_airplanes.py](analyze_airplanes.py):
# +
airplanes = spark.read.json('../data/airplanes.jsonl')
airplanes.registerTempTable("airplanes")
manufacturer_counts = spark.sql("""SELECT
Manufacturer,
COUNT(*) AS Total
FROM
airplanes
GROUP BY
Manufacturer
ORDER BY
Total DESC"""
)
manufacturer_counts.show(10) # show top 30
# -
# Interesting, Boeing planes outnumber Airbus planes by 4 to 1! I had no idea; I thought it was much closer than this. However, what I really want to know is who has what share of the market (without having to compute a ratio in my head). In other words, I’d like to see this data as a percentage.
#
# ## SQL Subqueries Versus Dataflow Programming
#
# This is a good way to illustrate the difference between SQL subqueries and dataflow programming. SQL is declarative, in that you specify what you want without saying how to get it. Imperative dataflow programming, on the other hand, involves the step-wise computation of data that you link and compose into dataflows.
#
# First we’ll implement the percentage totals using imperative dataflows, and then we’ll do so using declarative SQL subqueries. You’ll see that in this case subqueries are more convenient, but there is a limit to the utility of subqueries—they can get obscure fast. It is better to create a series of simple SQL or dataflow statements that compose into the computation you desire rather than to try to specify it all in one large, deeply nested subquery.
#
# ### Dataflow Programming Without Subqueries
#
# Subqueries weren’t supported in Spark SQL until 2.0. Instead, given our manufacturer airplane counts, we would need to calculate the total airplane count, join that to our existing totals, and then divide the manufacturer subtotals by the overall total. We’ll reuse the manufacturer_counts relation we computed in the previous program listing:
# How many airplanes total?
total_airplanes = spark.sql(
"""SELECT
COUNT(*) AS OverallTotal
FROM airplanes"""
)
print("Total airplanes: {:,}".format(total_airplanes.collect()[0].OverallTotal))
mfr_with_totals = manufacturer_counts.crossJoin(total_airplanes)
mfr_with_totals = mfr_with_totals.rdd.map(
lambda x: {
'Manufacturer': x.Manufacturer,
'Total': x.Total,
'Percentage': round(
(
float(x.Total)/float(x.OverallTotal)
) * 100,
2
)
}
)
mfr_with_totals.toDF().show()
# This is clearly an out-of-the-way method of calculating percentage totals, but it illustrates how dataflow programming works in more complex examples as well.
# ### Subqueries in Spark SQL
#
# Subqueries are handy, and computing the percentage share of the aircraft manufacturers is easy using them:
relative_manufacturer_counts = spark.sql("""SELECT
Manufacturer,
ROUND(
100 * (
COUNT(*)/(SELECT COUNT(*) FROM airplanes)
),
2
) AS Percentage,
COUNT(*) AS Total
FROM
airplanes
GROUP BY
Manufacturer
ORDER BY
Total DESC, Manufacturer"""
)
relative_manufacturer_counts.show(20)
# ## Creating an Airplanes Home Page
#
# Now I want to see this data as a chart on a web page, which means we need somewhere to put the chart. This is a good time to create an `/airplanes` home page—a page that analyzes the fleet as a whole.
#
# Let’s create a Flask controller for `/airplanes`. Check out `ch06/web/report_flask.py`, which simply loads the data from Mongo and passes it to a template, `all_airplanes.html`:
#
# ```python
# @app.route("/airplanes1")
# @app.route("/airplanes1/")
# def airplanes():
# mfr_chart = client.agile_data_science.manufacturer_totals.find_one()
# return render_template('all_airplanes.html',mfr_chart=mfr_chart)
# ```
#
# To start, the template looks like this:
#
# ```html
# {% extends "layout.html" %}
# {% block body %}
# <!-- Navigation guide -->
# / <a href="/airplanes">Airplanes</a>
#
# <p class="lead">
# <!-- Airline name and website-->
# US Commercial Fleet
# </p>
# {% endblock %}
# ```
#
# The result looks like this:
#
# 
#
# ### Adding Search to the Airplanes Page
#
# The `/airplanes` page is a great place to implement search for the airplane records we’ve created. To do this, first we’ll need to create a new search index and then index our airplane documents in Elasticsearch via PySpark:
# + language="bash"
#
# curl -XPUT 'http://localhost:9200/agile_data_science_airplanes/' -d '{
# "settings" : {
# "index" : {
# "number_of_shards" : 1,
# "number_of_replicas" : 1
# }
# }
# }'
# +
# Load our airplanes
airplanes = spark.read.json("../data/airplanes.json")
airplanes.show(10)
# Save the DataFrame to Elasticsearch
a = airplanes.rdd.map(lambda x: ('ignored_key', x.asDict()))
a.saveAsNewAPIHadoopFile(
path='-',
outputFormatClass="org.elasticsearch.hadoop.mr.EsOutputFormat",
keyClass="org.apache.hadoop.io.NullWritable",
valueClass="org.elasticsearch.hadoop.mr.LinkedMapWritable",
conf={ "es.resource" : "agile_data_science/airplanes" })
print("Records saved to Elasticsearch!")
# -
# We can verify our documents are there with a quick search:
# + language="bash"
#
# curl -XGET 'http://localhost:9200/agile_data_science/airplanes/_search?q=*' | jq ''
# -
# Now we can add search capability to our `/airplanes` controller. Recall that we we did this in Chapter 5 for flight search, and that it took several pages of code. This time, we are going to build a reusable component for searching and displaying records in a Flask application.
#
# ### Code Versus Configuration
#
# There are varying opinions on how much code duplication is acceptable before factoring it out, and this varies greatly by context and across programming disciplines. Data science has a higher tolerance for ugly, duplicated code than does general software engineering. This is for the simple reason that most code a data scientist writes is discarded immediately after it is run. Most data scientists are doing well to commit all their code to a repository at all (an absolute must!). But when code persists and we share the result, as in our application, cleanliness starts to matter.
#
# This is the second time we’ve been tasked with implementing search, and when repeating oneself at length, it is a good time to take pause and see if one can’t reduce some of the redundant code necessary to reproduce a feature. (Because most code is throwaway in data science, it is important to wait until you repeat yourself to factor out duplicate code and generalize capabilities, as we have done here.)
#
# Being tasked with generalizing code, we have to split our code into two elements: algorithms and configuration. Algorithms define the behavior of what we’re building; configuration defines what an instance of that algorithm is like.
#
# To take one extreme, it is possible in programming to remove all redundancy in code, to the point that everything becomes a configuration file or an algorithm implementing the behavior of a configuration. This extreme, however, is not maintainable and isn’t compatible with a data scientist’s workload, where we have to remember lots of things at once and can’t dive deep into each component just to edit or make use of it. As usual in Agile Data Science, we choose the middle path, where we remove the worst of the redundancy without reducing everything to its most generalized, reusable form.
#
# ### Configuring a Search Widget
#
# Our search configuration is simple. It lays out the fields we will search and display, and an optional label we would like to use in the user interface:
#
# ```python
# search_config = [
# {'field': 'TailNum', 'label': 'Tail Number'},
# {'field': 'Owner', 'sort_order': 0},
# {'field': 'OwnerState', 'label': 'Owner State'},
# {'field': 'Manufacturer', 'sort_order': 1},
# {'field': 'Model', 'sort_order': 2},
# {'field': 'ManufacturerYear', 'label': 'MFR Year'},
# {'field': 'SerialNumber', 'label': 'Serial Number'},
# {'field': 'EngineManufacturer', 'label': 'Engine MFR', 'sort_order': 3},
# {'field': 'EngineModel', 'label': 'Engine Model', 'sort_order': 4}
# ]
# ```
#
# Our pagination works as before, but we’ve got a new configuration item for `AIRPLANE_RECORDS_PER_PAGE`:
#
# ```python
# # Pagination parameters
# start = request.args.get('start') or 0
# start = int(start)
# end = request.args.get('end') or config.AIRPLANE_RECORDS_PER_PAGE
# end = int(end)
#
# # Navigation path and offset setup
# nav_path = search_helpers.strip_place(request.url)
# nav_offsets = search_helpers.get_navigation_offsets(
# start,
# end,
# config.AIRPLANE_RECORDS_PER_PAGE
# )
# ```
#
#
# With our search config in hand, we need only define the base of the Elasticsearch query and flesh it out based on the search arguments we receive. Our base query looks like this:
#
# ```python
# # Build the base of our elasticsearch query
# query = {
# 'query': {
# 'bool': {
# 'must': []}
# },
# 'from': start,
# 'size': config.AIRPLANE_RECORDS_PER_PAGE
# }
# ```
#
# And we parameterize it like so:
#
# ```python
# arg_dict = {}
# for item in search_config:
# field = item['field']
# value = request.args.get(field)
# arg_dict[field] = value
# if value:
# query['query']['bool']['must'].append({'match': {field: value}})
# ```
#
# We submit the query as before:
#
# ```python
# # Query elasticsearch, process to get records and count
# results = elastic.search(query, index='agile_data_science_airplanes')
# airplanes, airplane_count = search_helpers.process_search(results)
# ```
#
# In our call to render our template, we now include the search_config and arg_dict, which will generate our content in the template:
#
# ```python
# # Persist search parameters in the form template
# return render_template(
# 'all_airplanes.html',
# search_config=search_config,
# args=arg_dict,
# airplanes=airplanes,
# airplane_count=airplane_count,
# nav_path=nav_path,
# nav_offsets=nav_offsets,
# )
# ```
#
# Our template, `all_airplanes.html`, is derived from `search.html` from chapter 5. Using `search_config` and the request arguments, we programmatically build all the content we manually specified before in search.html. We can reuse this code now to re-create any search controller:
#
# ```html
# {% extends "layout.html" %}
# {% block body %}
# <!-- Navigation guide -->
# / <a href="/airplanes">Airplanes</a>
#
# <p class="lead" style="margin: 10px; margin-left: 0px;">
# <!-- Airline Name and website-->
# US Commercial Fleet
# </p>
#
# <!-- Chart of fleet manufacturers -->
# <div>
# <p style="margin: 0px;">Airplanes by Manufacturer</p>
# <div id="chart"><svg class="chart"></svg></div>
# </div>
# <script src="/static/airplanes.js"></script>
#
# <!-- Generate form from search_config and request args -->
# <form action="/airplanes" method="get">
# {% for item in search_config %}
# {% if 'label' in item %}
# <label for="{{item['field']}}">{{item['label']}}</label>
# {% else %}
# <label for="{{item['field']}}">{{item['field']}}</label>
# {% endif %}
# <input name="{{item['field']}}" style="width: 36px; margin-right: 10px;" value="{{args[item['field']] if args[item['field']] else ''}}"></input>
# {% endfor %}
# <button type="submit" class="btn btn-xs btn-default" style="height: 25px">Submit</button>
# </form>
#
# <table class="table table-condensed table-striped">
# <!-- Create table header, based on search_config -->
# <thead>
# {% for item in search_config %}
# {% if 'label' in item %}
# <th>{{item['label']}}</th>
# {% else %}
# <th>{{item['field']}}</th>
# {% endif %}
# {% endfor %}
# </thead>
#
# <!-- Create table content, based on airplanes for each <tr> and search_config for each <td> -->
# <tbody>
# {% for airplane in airplanes %}
# <tr>
# {% for item in search_config %}
# <td>{{airplane[item['field']]}}</td>
# {% endfor %}
# </tr>
# {% endfor %}
# </tbody>
# </table>
#
# <!-- css for x axis in chart -->
# <style>
# .axis text {
# font: 8px sans-serif;
# }
#
# .axis path,
# .axis line {
# fill: none;
# stroke: #000;
# shape-rendering: crispEdges;
# }
#
# .bar {
# fill: #ff6600;
# }
# </style>
#
# {% import "macros.jnj" as common %}
# {% if nav_offsets and nav_path -%}
# {{ common.display_nav(nav_offsets, nav_path, airplane_count)|safe }}
# {% endif -%}
# {% endblock %}
# ```
#
# Check out the resulting search at [http://localhost:5000/airplanes](http://localhost:5000/airplanes)
# ## Creating a Manufacturers Bar Chart
#
# Now that we’ve got a place to put our chart, let’s get down to creating it!
#
# Continuing with our script, [ch06/analyze_airplanes.py](analyze_airplanes.py), we store the data for the chart in Mongo:
# +
#
# Now get these things on the web
#
relative_manufacturer_counts_dicts = relative_manufacturer_counts.rdd.map(lambda row: row.asDict())
grouped_manufacturer_counts = relative_manufacturer_counts_dicts.groupBy(lambda x: 1)
# Save to Mongo in the airplanes_per_carrier relation
import pymongo_spark
pymongo_spark.activate()
grouped_manufacturer_counts.saveToMongoDB(
'mongodb://localhost:27017/agile_data_science.airplane_manufacturer_totals'
)
print("Records stored to MongoDB!")
# -
# Next, check that the data is in Mongo:
# + language="bash"
#
# mongo agile_data_science --quiet --eval 'db.airplane_manufacturer_totals.findOne()'
# -
# The rest is similar to the bar chart from Chapter 5. We add a controller to [ch06/web/report_flask.py](web/report_flask.py) where we grab the chart from Mongo, and return it as JSON:
#
# ```python
# @app.route("/airplanes/chart/manufacturers.json")
# @app.route("/airplanes/chart/manufacturers.json")
# def airplane_manufacturers_chart():
# mfr_chart = client.agile_data_science.airplane_manufacturer_totals.find_one()
# return json.dumps(mfr_chart)
# ```
#
# Then we edit the [ch06/web/templates/all_airplanes.html](web/templates/all_airplanes.html) template to call [ch06/web/static/airplanes.js](web/static/airplanes.js), which draws the chart.
#
# This time we want x- and y-axes for our bar chart, so we’re going to draw from an example that includes them. <NAME>’s example Bar Chart IIIc is concise and straightforward. Let’s begin by titling our page and calling our chart script, `airplanes.js`:
#
# ```html
# <!-- Chart of fleet manufacturers -->
# <div>
# <p style="margin: 0px;">Airplanes by Manufacturer</p>
# <div id="chart"><svg class="chart"></svg></div>
# </div>
# <script src="/static/airplanes.js"></script>
# ```
#
# [ch06/web/static/airplanes.js](web/static/airplanes.js) has a few changes to make the example work for our chart’s data. Aside from plugging in the Total and Manufacturer field names and passing through the data.data field, we haven’t changed anything except the dimensions of the chart:
#
# ```javascript
# var margin = {top: 20, right: 30, bottom: 30, left: 40},
# width = 900 - margin.left - margin.right,
# height = 300 - margin.top - margin.bottom;
#
# var x = d3.scale.ordinal()
# .rangeRoundBands([0, width], .1);
# var y = d3.scale.linear()
# .range([height, 0]);
#
# var xAxis = d3.svg.axis()
# .scale(x)
# .orient("bottom")
# .tickFormat(function(d) {
# return truncate(d, 14);
# });
# var yAxis = d3.svg.axis()
# .scale(y)
# .orient("left");
#
# var chart = d3.select(".chart")
# .attr("width", width + margin.left + margin.right)
# .attr("height", height + margin.top + margin.bottom)
# .append("g")
# .attr("transform", "translate(" + margin.left + "," + margin.top + ")");
#
# d3.json("/airplanes/chart/manufacturers.json", function(error, data) {
# var data = data.data;
#
# x.domain(data.map(function(d) { return d.Manufacturer; }));
# y.domain([0, d3.max(data, function(d) { return d.Total; })]);
#
# chart.append("g")
# .attr("class", "x axis")
# .attr("transform", "translate(0," + height + ")")
# .call(xAxis);
#
# chart.append("g")
# .attr("class", "y axis")
# .call(yAxis);
#
# chart.selectAll(".bar")
# .data(data)
# .enter().append("rect")
# .attr("class", "bar")
# .attr("x", function(d) { return x(d.Manufacturer); })
# .attr("y", function(d) { return y(d.Total); })
# .attr("height", function(d) { return height - y(d.Total); })
# .attr("width", x.rangeBand());
# });
#
# function truncate(d, l) {
# if(d.length > l)
# return d.substring(0,l)+'...';
# else
# return d;
# }
# ```
#
# And for our trouble, we get a beautiful chart at [http://localhost:5000/airplanes](http://localhost:5000/airplanes):
#
# 
#
# ## Exercises
#
# In this exercise we will create a `/airports` home page describing America's airports.
#
# 1. Begin with a simple Flask controller and HTML template for the page, and verify that it works.
# 2. Prepare a dataset for a chart for the page. Create a bar chart showing the number of flights out of the top 20 most busy airports in the US in 2015. Label the chart with full airport names, not just codes.
# 3. Publish this dataset to MongoDB and verify it is there.
# 4. Implement a controller serving the data for the chart as JSON.
# 5. Implement the bar chart in d3.js
#
# ## Iterating on the Manufacturers Bar Chart
#
# Wait a minute, something is wrong! Remember when we said iteration is essential? Let’s debug this chart. We need to infer from the chart what might be going on. Why are the bars so thin? Why are they shoved to the left?
#
# Recall that we sorted the data by `Total` in descending order:
#
# ```python
# relative_manufacturer_counts = spark.sql("""SELECT
# Manufacturer,
# ROUND(
# 100 * (
# COUNT(*)/(SELECT COUNT(*) FROM airplanes)
# ),
# 2
# ) AS Percentage,
# COUNT(*) AS Total
# FROM
# airplanes
# GROUP BY
# Manufacturer
# ORDER BY
# Total DESC, Manufacturer
# LIMIT 20"""
# )
# ```
#
# This means that the largest values are on the left, and the smallest values are on the right… so what must be happening is that there are simply too many small values to make the chart readable! We can improve the chart by removing some of these smaller values, since they are insignificant. Note that this won’t always be the case, so think carefully before discarding data!
#
# We can fix up our chart by recomputing the data using a SQL LIMIT command. First, we need to drop the stale data from Mongo:
# + language="bash"
#
# mongo agile_data_science --quiet --eval 'db.airplane_manufacturer_totals.drop()'
# -
relative_manufacturer_counts = spark.sql("""SELECT
Manufacturer,
COUNT(*) AS Total,
ROUND(
100 * (
COUNT(*)/(SELECT COUNT(*) FROM airplanes)
),
2
) AS PercentageTotal
FROM
airplanes
GROUP BY
Manufacturer
ORDER BY
Total DESC, Manufacturer
LIMIT 10""")
relative_manufacturer_counts.show()
# +
#
# Now get these things on the web
#
relative_manufacturer_counts_dicts = relative_manufacturer_counts.rdd.map(lambda row: row.asDict())
grouped_manufacturer_counts = relative_manufacturer_counts_dicts.groupBy(lambda x: 1)
# Save to Mongo in the airplanes_per_carrier relation
import pymongo_spark
pymongo_spark.activate()
grouped_manufacturer_counts.saveToMongoDB(
'mongodb://localhost:27017/agile_data_science.airplane_manufacturer_totals'
)
print("Records stored to MongoDB!")
# -
# Now check out [http://localhost:5000/airplanes](http://localhost:5000/airplanes) again!
#
# 
#
# Running our new script and pushing our new data to Mongo results in something that clearly shows the trend of Boeing dominating the market, trailed by several other manufacturers. Note that we also created a function called truncate to shorten long manufacturer names in the x-axis, so these labels do not overlap. We call this function from the tickFormat method on our xAxis object:
#
# ```javascript
# function truncate(d, l) {
# if(d.length > l)
# return d.substring(0,l)+'...';
# else
# return d;
# }
# ```
#
# ## Entity Resolution: Another Chart Iteration
#
# However, there is another problem with the chart—the column names are duplicated, which is distorting the values of manufacturers Airbus, McDonnel Douglas, and Embraer. We need to iterate yet again! This time we’ll be tackling entity resolution.
#
# ### Entity Resolution in 30 Seconds
#
# The problem we have encountered is that there are several forms of the manufacturers’ names used in the registrations of various airplanes. Addressing this problem is called entity resolution, which is defined in a tutorial by <NAME> and <NAME> as “[the] problem of identifying and linking/grouping different manifestations of the same real world object.” Entity resolution is the process by which AIRBUS is identified as the same thing as `AIRBUS INDUSTRIE`.
#
# There are many methods of entity resolution, including complicated means employing statistical inference. We will only explore a simple heuristic-based approach, because it turns out that in this case that is simply good enough. Don’t allow your curiosity to distract you into employing machine learning and statistical techniques whenever you can. Get curious about results, instead.
#
# ### Resolving Manufacturers in PySpark
#
# Let’s begin by inspecting the different ways the Manufacturer field appears in the airplane records. We can use SQL to `SELECT DISTINCT(Manufacturer) AS Manufacturer` and then see similar records next to one another with `ORDER BY Manufacturer`. Then we need only print the data in a left-justified manner, and see what we’ve got!
#
# This results in a list that allows us to easily visualize the variety of values of Manufacturer:
# +
airplanes = spark.read.json('../data/airplanes.json')
airplanes.registerTempTable("airplanes")
manufacturer_variety = spark.sql(
"""SELECT
DISTINCT(Manufacturer) AS Manufacturer
FROM
airplanes
ORDER BY
Manufacturer"""
)
manufacturer_variety_local = manufacturer_variety.collect()
# We need to print these left justified
for mfr in manufacturer_variety_local:
print(mfr.Manufacturer)
# -
# It turns out that we don’t have very much variety at all: only 35 distinct values. Reconciling the Manufacturer field of these records could be done manually, with a simple table elaborating the matches in two columns. One column would contain the raw value, and the other would contain the value to map to (the “standard” you have chosen). Against this table you can then LEFT JOIN and, if there is a match, replace the value of the field, in order to get a common identifier between records.
#
# If you encounter 35 values for a field in your work, do yourself a favor: make the table manually as CSV and load it in Spark and do the join. Here we will go further to illustrate how to create such a mapping table in an automated way, and how to JOIN it and effect the mapping. We do this to give you experience in how to problem solve and “munge” your way out of these situations when you can without having to turn to more complex (and thus time-consuming) statistical techniques.
#
# A more sophisticated approach would be to inspect the data and see if we can infer a rule to use to decide if records are identical. In looking at our duplicates, it seems that whenever there is a duplicate, there is a lot of overlap at the start of the strings. This is common among company names in the wild, where trailing symbols like “Incorporated” are shortened to “Inc,” “Inc.,” “INC,” “Corp,” etc. We might then formulate a strategy: if fields between records contain more than N characters in common at the start of the string, they are identical. We would choose the longest common substring as the “standard” value among those records, and use this rule to create our mapping table.
#
# To employ this strategy, we need to compare all unique values of Manufacturer with one another. This is feasible with 35 unique values, but keep in mind that this may not always be the case when resolving entities. Sometimes it is impossible to compare all records with one another, because the square of the number of unique records is too big, even for Spark! In this case we’re only resolving one field, which keeps the cardinality low by enabling us to use the unique values of just that one field. When records have numerous fields that identify them, the number of unique records explodes. That situation is (thankfully) beyond the scope of this book, but I’ve had good experiences with the Swoosh algorithms, which are implemented in the SERF project from Stanford.
#
# Check out [ch06/resolve_airplane_manufacturers.py](resolve_airplane_manufacturers.py). Here we prepare a mapping table for similar `Manufacturer` values, using the assumption that strings whose beginnings overlap by more than five characters are the same. Note that this assumption is naive and would not work for most datasets. Nonetheless, it shows how you can munge your way out of sticky situations by learning your dataset and actually looking at the data, record by sorted, unique record.
#
# Continuing from the last code example where we computed manufacturer_variety, check out the inline comments that describe the computation here:
def longest_common_beginning(s1, s2):
"""Detect the longest common beginning string in a pair of strings"""
if s1 == s2:
return s1
min_length = min(len(s1), len(s2))
i = 0
while i < min_length:
if s1[i] == s2[i]:
i += 1
else:
break
return s1[0:i]
def compare_manufacturers(mfrs):
"""Compare two manufacturers, returning a tuple describing the result"""
mfr1 = mfrs[0]
mfr2 = mfrs[1]
lcb = longest_common_beginning(mfr1, mfr2)
lcb = lcb.strip() # remove extra spaces
len_lcb = len(lcb)
record = {
'mfr1': mfr1,
'mfr2': mfr2,
'lcb': lcb,
'len_lcb': len_lcb,
'eq': mfr1 == mfr2
}
return record
# +
# Pair every unique instance of Manufacturer field with every other for comparison
comparison_pairs = manufacturer_variety.crossJoin(manufacturer_variety)
# Do the comparisons
comparisons = comparison_pairs.rdd.map(compare_manufacturers)
# Matches have > 5 starting chars in common
matches = comparisons.filter(lambda f: f['eq'] == False and f['len_lcb'] > 5)
matches.take(5)
# +
#
# Now we create a mapping of duplicate keys from their raw value to the one we're going to use
#
# 1) Group the matches by the longest common beginning ('lcb')
common_lcbs = matches.groupBy(lambda x: x['lcb'])
# 2) Emit the raw value for each side of the match along with the key, our 'lcb'
mfr1_map = common_lcbs.map(lambda x: [(y['mfr1'], x[0]) for y in x[1]]).flatMap(lambda x: x)
mfr2_map = common_lcbs.map(lambda x: [(y['mfr2'], x[0]) for y in x[1]]).flatMap(lambda x: x)
# 3) Combine the two sides of the comparison's records
map_with_dupes = mfr1_map.union(mfr2_map)
# 4) Remove duplicates
mfr_dedupe_mapping = map_with_dupes.distinct()
# 5) Convert mapping to dataframe to join to airplanes dataframe
mapping_dataframe = mfr_dedupe_mapping.toDF()
# 6) Give the mapping column names
mapping_dataframe.registerTempTable("mapping_dataframe")
mapping_dataframe = spark.sql(
"SELECT _1 AS Raw, _2 AS NewManufacturer FROM mapping_dataframe"
)
# JOIN our mapping left outer...
airplanes_w_mapping = airplanes.join(
mapping_dataframe,
on=airplanes.Manufacturer == mapping_dataframe.Raw,
how='left_outer'
)
# Now replace Manufacturer with NewManufacturer where needed
airplanes_w_mapping.registerTempTable("airplanes_w_mapping")
resolved_airplanes = spark.sql("""SELECT
TailNum,
SerialNumber,
Owner,
OwnerState,
IF(NewManufacturer IS NOT null,NewManufacturer,Manufacturer) AS Manufacturer,
Model,
ManufacturerYear,
EngineManufacturer,
EngineModel
FROM
airplanes_w_mapping""")
# Store for later use, in place of airplanes.json
resolved_airplanes.repartition(1).write.mode("overwrite").json("../data/resolved_airplanes.json")
# -
# Now we can employ the mapping table we have created. Note that this table could have been prepared manually, given the small number of records, and in that case this is the point at which you would load the mapping table as CSV (and run the next code block):
# +
# JOIN our mapping left outer...
airplanes_w_mapping = airplanes.join(
mapping_dataframe,
on=airplanes.Manufacturer == mapping_dataframe.Raw,
how='left_outer'
)
# Now replace Manufacturer with NewManufacturer where needed
airplanes_w_mapping.registerTempTable("airplanes_w_mapping")
resolved_airplanes = spark.sql("""SELECT
TailNum,
SerialNumber,
Owner,
OwnerState,
IF(NewManufacturer IS NOT null,NewManufacturer,Manufacturer) AS Manufacturer,
Model,
ManufacturerYear,
EngineManufacturer,
EngineModel
FROM
airplanes_w_mapping""")
# Store for later use, in place of airplanes.json
resolved_airplanes.repartition(1).write.mode("overwrite").json("../data/resolved_airplanes.json")
# -
# Again, for convenience, let’s create a single JSON Lines file and check out the result:
# + language="bash"
#
# cat ../data/resolved_airplanes.json/part* >> ../data/resolved_airplanes.jsonl
# head -5 ../data/resolved_airplanes.jsonl
# -
# ### Cleaning out MongoDB
#
# Now we need to drop our original collection and re-calculate the data for the chart.
# + language="bash"
#
# mongo agile_data_science --quiet --eval 'db.airplane_manufacturer_totals.drop()'
# -
# We need to run the code below, which is just a copy of the original [ch06/analyze_airplanes.py](analyze_airplanes.py) with the new path for our resolved airplanes plugged in.
# +
airplanes = spark.read.json('../data/resolved_airplanes.json')
airplanes.registerTempTable("airplanes")
#
# Same with sub-queries
#
relative_manufacturer_counts = spark.sql("""SELECT
Manufacturer,
COUNT(*) AS Total,
ROUND(
100 * (
COUNT(*)/(SELECT COUNT(*) FROM airplanes)
),
2
) AS PercentageTotal
FROM
airplanes
GROUP BY
Manufacturer
ORDER BY
Total DESC, Manufacturer
LIMIT 10"""
)
relative_manufacturer_counts.show(30) # show top 30
#
# Now get these things on the web
#
relative_manufacturer_counts_dict = relative_manufacturer_counts.rdd.map(lambda row: row.asDict())
grouped_manufacturer_counts = relative_manufacturer_counts_dict.groupBy(lambda x: 1)
# Save to Mongo in the airplanes_per_carrier relation
import pymongo_spark
pymongo_spark.activate()
grouped_manufacturer_counts.saveToMongoDB(
'mongodb://localhost:27017/agile_data_science.airplane_manufacturer_totals'
)
print("Data stored to MongoDB!")
# -
# Once you’ve done that, check out [http://localhost:5000/airplanes](http://localhost:5000/airplanes) to see the updated chart:
#
# 
#
# The chart has changed quite a bit now that we’ve deduplicated manufacturers! Airbus isn’t so far behind as we had thought. Now I’m wondering precisely how much market share each manufacturer has using the new airplanes dataframe.
#
# It turns out that Boeing has 49% of the market, versus Airbus with 23.4%. Go Boeing! (Or, in case you’re in Europe... go Airbus!)
#
# ## Cleanliness: Benefits of Entity Resolution
#
# Raw data is always dirty. Once you dive in and start working with data and look at it in raw form, when you visualize it in web pages in tables and charts and make it searchable, problems with the data emerge. Resolving these problems as you work with the data enables you to see trends clearly, without distortion. As your visualizations benefit, so will your models. This “cleaning” sets you up for success in building effective statistical models in the next level of the data-value pyramid: predictions.
#
# # Conclusion
#
# Here’s a summary of what we’ve done so far:
#
# 1. Create interesting, interconnected records. The bar for “interesting” is initially low. We will improve it over time based on user feedback, traffic analysis, and noodling.
#
# Store these records as objects in a document store, like so:
#
# key => {property1, property2, links => [key1, key2, key3]}
#
# 2. Split records as properties increase and become complex to avoid deep nesting, or go at it as a document. Both approaches are valid if they fit your data.
#
# 3. Use a lightweight web framework like Flask or Sinatra to emit the key/value data as JSON, or use a document store that returns JSON in the first place.
#
# In the next chapter, we’ll take what we’ve learned about our data to make a prediction with lots of practical relevance: will our flight be late? And if so, by how much?
|
ch06/Exploring_Data_with_Reports.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Normal shading with textures
#
# This example shows how to:
# - modify predefined materials
# - use displacement map
# - apply large textures to material
#
# Tkinter GUI window is launched from the notebook in this example. This allows re-running code cells and see results without scrolling back to an inlined figure.
#
# 
import numpy as np
from plotoptix import TkOptiX
from plotoptix.materials import m_clear_glass, m_plastic # predefined materials
from plotoptix.utils import map_to_colors, make_color_2d, read_image, simplex
# Make some data first.
m = 300
r = 0.08 * np.random.rand(m) + 0.01
p = 3 * (np.random.rand(m,3) - 0.5)
p[:,0] *= 0.8
p[:,2] *= 1.5
p[:,1] = r[:]
# Setup the raytracer using Tkinter GUI as the output target.
rt = TkOptiX()
rt.set_param(min_accumulation_step=4, # set more accumulation frames to get rid of the noise
max_accumulation_frames=512,
light_shading="Hard") # use ligth shading best for caustics
rt.set_uint("path_seg_range", 5, 10) # more path segments to allow multiple reflections
# Only *diffuse* material is available by default. Other materials need to be configured before using.
# +
import copy
m_beads = copy.deepcopy(m_plastic) # a new material based on predefined properties
rt.setup_material("plastic", m_plastic)
rt.setup_material("glass", m_clear_glass)
rt.setup_material("beads", m_beads)
# -
# Add objects to the scene.
#
# **Note 1:** particles with ``geom="ParticleSetTextured"`` geometry can have 3D orientation, provided with ``u`` and ``v`` arguments, required for applying textures. 3D oreintation is randomized if these vectors are omitted.
#
# **Note 2:** ``geom_attr="DisplacedSurface"`` is used to allow for displacement mapping.
# +
rt.set_data("plane", geom="Parallelograms", mat="plastic",
pos=[-3, 0, -3], u=[6, 0, 0], v=[0, 0, 6], c=0.8)
rt.set_data("sphere", geom="ParticleSetTextured", mat="glass", geom_attr="DisplacedSurface",
pos=[0.0, 0.7, 0.0], u=[-1, 0, 0], r=0.4, c=10)
rt.set_data("particles", geom="ParticleSetTextured", mat="beads",
pos=p, r=r, c=0.95)
# -
# Setup a good point of view, set background and lights.
# +
rt.setup_camera("cam1", cam_type="DoF",
eye=[-2.1, 2.4, 0], target=[0, 0, 0], up=[0.28, 0.96, 0.05],
aperture_radius=0.01, fov=30, focal_scale=0.91)
rt.setup_light("light1", pos=[4, 5.1, 3], color=[12, 11, 10], radius=1.9)
rt.setup_light("light2", pos=[-1.5, 3, -2], color=[8, 9, 10], radius=0.2)
rt.set_background(0)
rt.set_ambient(0)
exposure = 0.4; gamma = 2.2
rt.set_float("tonemap_exposure", exposure)
rt.set_float("tonemap_gamma", gamma)
rt.set_float("denoiser_blend", 0.25)
rt.add_postproc("Denoiser") # apply gamma correction postprocessing stage, or
#rt.add_postproc("Gamma") # use AI denoiser (exposure and gamma are applied as well)
# -
# Open the GUI.
rt.start()
# Make a gradient texture, use it with the *glass* material:
y = np.linspace(0, 1, 100)
M = np.stack((y,y)).T
# +
M1 = map_to_colors(-M, "Purples")
M1 = make_color_2d(M1, gamma=gamma, channel_order="RGBA")
rt.set_texture_2d("purple", 10*M1)
m_clear_glass["ColorTextures"] = [ "purple" ]
m_clear_glass["VarFloat3"]["refraction_index"] = [1.4, 1.4, 1.4]
rt.update_material("glass", m_clear_glass, refresh=True)
# -
# Make a gradient with another color map for the plastic particles.
# +
M2 = map_to_colors(M, "RdYlBu")
M2 = make_color_2d(M2, gamma=gamma, channel_order="RGBA")
rt.set_texture_2d("redyellowblue", M2)
m_beads["ColorTextures"] = [ "redyellowblue" ]
rt.update_material("beads", m_beads, refresh=True)
# -
# Calculate displacement map over 2D plane using simplex noise:
# +
nn = 2000
x = np.linspace(0, 50, nn)
z = np.linspace(0, 50, nn)
X, Z = np.meshgrid(x, z)
XZ = np.stack((X.flatten(), Z.flatten(), np.full(nn**2, 1.0, dtype=np.float32))).T.reshape(nn, nn, 3)
XZ = np.ascontiguousarray(XZ, dtype=np.float32)
Y = simplex(XZ)
Y = np.sin(10 * Y)
# -
# Use the displacement map for the shading normal tilt. Displacement is relative to the object size, it should be a small value to look like a wrinkles on the plane. Shading modulation is implemented as a ``Float2`` texture calculated from the displacement, and accompanied with an inverse aspect ratio, ``normaltilt_iar``, parameter. Use ``set_normal_tilt()`` method to set these two parameters.
rt.set_normal_tilt("plastic", 0.00008*Y, refresh=True)
# OK, to make it more interesting, let's add some lines along wrinkles. An RGBA texture is needed for this.
Ym = 0.5 * (np.copy(Y) + 1)
m = (Ym > 0.45) & (Ym < 0.55)
Ym[m] = 0.0
Ym[~m] = 0.95
# +
M3 = make_color_2d(Ym, channel_order="RGBA")
rt.set_texture_2d("ripples", M3)
m_plastic["ColorTextures"] = [ "ripples" ]
rt.update_material("plastic", m_plastic, refresh=True)
# -
# Surface of spheres can be actually displaced, which gives accurate look of a strongly modified object, not only an impression of inclined surface.
# +
#D = read_image("data/blur.png", normalized=True).mean(axis=2)
#rt.set_displacement("sphere", 0.7 + 0.3*D, refresh=True)
rt.load_displacement("sphere", "data/blur.png", prescale=0.3, baseline=0.7, refresh=True)
# -
# Close GUI window, release resources.
rt.close()
|
examples/1_basics/8_shading_with_textures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Reading netflow day 3 into a txt file
# -
import time
import bz2
import pandas as pd
# Reading in the File
start = time.time()
data = []
with bz2.open("netflow_day-03.bz2", "rt") as bz_file:
for line in bz_file:
row = line.split(",")
row[10] = row[10].replace('\n', '')
data.append(row)
end = time.time()
print(end - start)
import csv
with open("netflow_day-03.csv", "wb") as f:
writer = csv.writer(f)
writer.writerows(data)
# +
############## Don't actually need this use different file to clean and convert to cleaned csv ######################
# -
# Converting the File to a pandas df
start = time.time()
header = ["Time", "Duration", "SrcDevice", "DstDevice", "Protocol", "SrcPort", "DstPort", "SrcPackets",
"DstPackets", "SrcBytes", "DstBytes"]
df = pd.DataFrame.from_records(data, columns = header)
end = time.time()
print(end - start)
|
Catherine/SIEDS_Files/Reading_In_Day3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# # Case 3: Generate Pseudo-Data with your model
using BOMBs
# Just to print the nice BOMBS logo
printLogo()
# First we need to generate the model (the package will not do it automatically)
# +
# Generate the empty dictionary with the model_def structure
model_def = defModStruct()
# Fill the different elements of the dictionary with your model specifications. See above how ^
model_def["NameF"] = ["PLacExample"];
model_def["nStat"] = [4];
model_def["nPar"] = [9];
model_def["nInp"] = [1];
model_def["stName"] = ["Cit_mrna","Cit_foldedP","Cit_fluo","Cit_AU"];
model_def["inpName"] = ["IPTG"];
model_def["parName"] = ["alpha1","Vm1","h1","Km1","d1",
"alpha2","d2","Kf","sc_molec"];
model_def["eqns"] = ["dCit_mrna=alpha1+Vm1*(IPTG^h1/(Km1^h1+IPTG^h1))-d1*Cit_mrna",
"dCit_foldedP=alpha2*Cit_mrna-(d2+Kf)*Cit_foldedP",
"dCit_fluo=Kf*Cit_foldedP-d2*Cit_fluo",
"dCit_AU = sc_molec*dCit_fluo"];
model_def["Y0eqs"] = ["Cit_mrna = (alpha1 + Vm1*(IPTG^h1/(Km1^h1+IPTG^h1)))/d1;",
"Cit_foldedP = (alpha2*Cit_mrna)/(Kf+d2);",
"Cit_fluo = (Kf*Cit_foldedP)/d2;",
"Cit_AU= sc_molec*Cit_fluo;"];
model_def["Y0Sim"] = [];
model_def["tols"] = [1e-9, 1e-9];
# Generate the scripts with the ODEs and the step-wise implementation
model_def = GenerateModel(model_def);
# -
# ### This function gives you information about the structure necessary to generate the model and how to call it.
# If you do not know how to call a specific section, just call the function with no input (infoAll()) to get the information.
infoAll("pseudodata")
# # Pseudo-Data Generation Example 1
# Here we code each element of the exeriment
pseudo_def = defPseudoDatStruct();
# +
pseudo_def["Nexp"] = [2];
pseudo_def["finalTime"] = [1439, 1439];
pseudo_def["switchT"] = [[0, 500, 1000, 1439],[0, 1439]];
pseudo_def["y0"] = [[10,10,10,10],[10,10,10,10]];
pseudo_def["preInd"] = [[0],[0]];
pseudo_def["uInd"] = [[5, 100, 1],[7.5]];
pseudo_def["theta"] = [0.000377125304442752*60, 0.00738924359598526*60, 1.53333782244337, 5.01927275636639,
0.00118831480244382*60, 0.0461264539194078*60, 0.000475563708997018*60, 0.000301803966012407*60, 68.8669567134881]; # Could be the directory and file name of a csv.
pseudo_def["tsamps"] = [collect(0:5:1439),collect(0:10:1439)];
pseudo_def["plot"] = [true];
pseudo_def["flag"] = ["Example1"];
pseudo_def["Obs"] = ["Cit_foldedP", "3*Cit_AU"];
pseudo_def["NoiseType"] = [];
pseudo_def["Noise"] = [0.1,0.2];
# -
# A print for the path where the results, CSVs and plots are saved will be shown. This will be in your working directory inside a new directory called Results.
pseudo_res, model_def, pseudo_def = GenPseudoDat(model_def, pseudo_def);
# ### Let's see the results structure
#
# The results are a dictionary 4 different entries: Sims (simulations), SimsObs (Observables Simulations) <br>
# PDat (Mean of the pseudo-data) and PError (Error of the pseudo-data)<br>
# <br>
# Contents of Sims are the same as for the simulation results from the ModelSim section
pseudo_res
# For the other 3 entries, the structure of the same, however note that the keys of the dictionary now start with PD
#
pseudo_res["PData"]
# And the contents for all of these are similar to the Model Simulation results. However, the second dimension of the resultant arrays is not the number of states, but the number of observables selected.
pseudo_res["PData"]["PDExp_1"]
# This observables can be the model states by itself or operations done with those!
#
pseudo_res["SimsObs"]["PDExp_1"][:,2,:] == pseudo_res["Sims"]["Exp_1"][:,4,:].*3
using Images # Package not called in BOMBS, this is just to display the generated PNGs
# Now we can plot the pseudo-data to see how did they go:
#
exp1 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",1,"_",pseudo_def["flag"],".png"))
exp2 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",2,"_",pseudo_def["flag"],".png"))
# If you would like to load the file with the results, just use the JLD pakage like this:
#
using JLD
using Dates
JLD.load(string(pseudo_def["savepath"], "\\",model_def["NameF"], "_", today(),"_PseudoDataResults_",pseudo_def["flag"],".jld"))
# ### CSVs with the pseu-data are also generated under a folder called PseudoDataFiles.
# For each experiment we have 3 different files:
# - (ModelName)_EXP(i)_(tag)_Simulations: A file with the simulation results
# - (ModelName)_EXP(i)_(tag)_Observables: A file with the observables
# - (ModelName)_EXP(i)_(tag)_Events_Inputs: A file with the input structure
#
# Let's load them for the first expreiment so we can have a look:
using CSV
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Simulations.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Observables.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Events_Inputs.csv"))
# # Pseudo-Data Generation Example 2
# This example showins how to introduce the experiment information using CSV files. In this case it also takes multiple theta vectors (3).
pseudo_def = defPseudoDatStructFiles();
# +
pseudo_def["ObservablesFile"] = ["DynStim_3_Observables.csv", "DynStim_4_Observables.csv"];
pseudo_def["EventInputsFile"] = ["DynStim_3_Events_Inputs.csv", "DynStim_4_Events_Inputs.csv"];
pseudo_def["theta"] = ["TestThetaPLac.csv"];
pseudo_def["MainDir"] = ["SimulationCSVs"];
pseudo_def["flag"] = "Example2";
pseudo_def["plot"] = "Yes";
pseudo_def["Obs"] = ["Cit_foldedP", "3*Cit_AU"];
pseudo_def["Noise"] = [0.1, 0.2];
# -
pseudo_res, model_def, pseudo_def = GenPseudoDat(model_def, pseudo_def);
# ### Let's see the results structure
#
# The results are a dictionary 4 different entries: Sims (simulations), SimsObs (Observables Simulations),
# PDat (Mean of the pseudo-data) and PError (Error of the pseudo-data) <br>
# <br>
# Contents of Sims are the same as for the simulation results from the ModelSim section
pseudo_res
# For the other 3 entries, the structure of the same, however note that the keys of the dictionary now start with PD
#
pseudo_res["PData"]
# And the contents for all of these are similar to the Model Simulation results. However, the second dimension of
# the resultant arrays is not the number of states, but the number of observables selected.
# Note that the 3rd dimension of the array is 3 (the number of theta vectors introduced)
pseudo_res["PData"]["PDExp_1"]
# This observables can be the model states by itself or operations done with those!
pseudo_res["SimsObs"]["PDExp_1"][:,2,:] == pseudo_res["Sims"]["Exp_1"][:,4,:].*3
# ### Now we can plot the pseudo-data to see how did they go:
#
# Note that only 1 pseudo-data trace has been ploted. This is because the average between all the traces (3 in this case)
# is given (to simplify plot generation). If you would like a plot for each theta, then you will have to do it
# yourself :( <br>
# <br>
# This is only done for the plots, in the CSVs you will see an entry for each theta
#
exp1 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",1,"_",pseudo_def["flag"],".png"))
exp2 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",2,"_",pseudo_def["flag"],".png"))
# If you would like to load the file with the results, just use the JLD pakage like this:
#
JLD.load(string(pseudo_def["savepath"], "\\",model_def["NameF"], "_", today(),"_PseudoDataResults_",pseudo_def["flag"],".jld"))
# ### CSVs with the pseu-data are also generated under a folder called PseudoDataFiles.
# For each experiment we have 3 different files:
# - (ModelName)_EXP(i)_(tag)_Simulations: A file with the simulation results
# - (ModelName)_EXP(i)_(tag)_Observables: A file with the observables
# - (ModelName)_EXP(i)_(tag)_Events_Inputs: A file with the input structure
#
# Let's load them for the first expreiment so we can have a look:
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Simulations.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Observables.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Events_Inputs.csv"))
# # Pseudo-Data Generation Example 3
# This is the same as example 1 but setting the experimental noise to be homoscedastic
pseudo_def = defPseudoDatStruct();
# +
pseudo_def["Nexp"] = [2];
pseudo_def["finalTime"] = [1439, 1439];
pseudo_def["switchT"] = [[0, 500, 1000, 1439],[0, 1439]];
pseudo_def["y0"] = [[10,10,10,10],[10,10,10,10]];
pseudo_def["preInd"] = [[0],[0]];
pseudo_def["uInd"] = [[5, 100, 1],[7.5]];
pseudo_def["theta"] = [0.000377125304442752*60, 0.00738924359598526*60, 1.53333782244337, 5.01927275636639,
0.00118831480244382*60, 0.0461264539194078*60, 0.000475563708997018*60, 0.000301803966012407*60, 68.8669567134881]; # Could be the directory and file name of a csv.
pseudo_def["tsamps"] = [collect(0:5:1439),collect(0:10:1439)];
pseudo_def["plot"] = [true];
pseudo_def["flag"] = ["Example3"];
pseudo_def["Obs"] = ["Cit_foldedP", "3*Cit_AU"];
pseudo_def["NoiseType"] = ["Homo"];
pseudo_def["Noise"] = [40,2000];
# -
# A print for the path where the results, CSVs and plots are saved will be shown. This will be in your working directory inside a new directory called Results.
pseudo_res, model_def, pseudo_def = GenPseudoDat(model_def, pseudo_def);
# ### Let's see the results structure
#
# The results are a dictionary 4 different entries: Sims (simulations), SimsObs (Observables Simulations) <br>
# PDat (Mean of the pseudo-data) and PError (Error of the pseudo-data)<br>
# <br>
# Contents of Sims are the same as for the simulation results from the ModelSim section
pseudo_res
# For the other 3 entries, the structure of the same, however note that the keys of the dictionary now start with PD
#
pseudo_res["PData"]
# And the contents for all of these are similar to the Model Simulation results. However, the second dimension of the resultant arrays is not the number of states, but the number of observables selected.
pseudo_res["PData"]["PDExp_1"]
# This observables can be the model states by itself or operations done with those!
#
pseudo_res["SimsObs"]["PDExp_1"][:,2,:] == pseudo_res["Sims"]["Exp_1"][:,4,:].*3
using Images # Package not called in BOMBS, this is just to display the generated PNGs
# Now we can plot the pseudo-data to see how did they go:
#
exp1 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",1,"_",pseudo_def["flag"],".png"))
exp2 = load(string(pseudo_def["savepath"], "\\PlotPseudoDat_Exp",2,"_",pseudo_def["flag"],".png"))
# If you would like to load the file with the results, just use the JLD pakage like this:
#
using JLD
using Dates
JLD.load(string(pseudo_def["savepath"], "\\",model_def["NameF"], "_", today(),"_PseudoDataResults_",pseudo_def["flag"],".jld"))
# ### CSVs with the pseu-data are also generated under a folder called PseudoDataFiles.
# For each experiment we have 3 different files:
# - (ModelName)_EXP(i)_(tag)_Simulations: A file with the simulation results
# - (ModelName)_EXP(i)_(tag)_Observables: A file with the observables
# - (ModelName)_EXP(i)_(tag)_Events_Inputs: A file with the input structure
#
# Let's load them for the first expreiment so we can have a look:
using CSV
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Simulations.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Observables.csv"))
CSV.read(string(pseudo_def["savepath"], "\\PseudoDataFiles\\", model_def["NameF"], "_Exp",1,"_",
pseudo_def["flag"],"_Events_Inputs.csv"))
# ### This is the end of the section. Look at the cool logo to check the package version ;)
#
versionBOMBS()
|
Examples/3_GeneratePseudoData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # XML/HTML
# ---
# ## XML
# ---
# ```xml
# <library>
# <metadata></metadata>
# <books>
# <book>
# <authors>
# <author>
# <firstname><NAME></firstname>
# <country></country>
# </author>
# <author></author>
# </authors>
# <title>Learning Python</title>
# <isbn></isbn>
# </book>
# <book></book>
# </books>
# </library>
# ```
# - Sections of a xml document are called _element_
# - Here _library_, _metadata_ are called tags of xml document (`tag` is a type of `element`)
# - _books_ tag contains multiple children tag called _book_ (An `element` can contain multiple child elements)
# - `Learning Python` is text element.
# DTD - Document Type Definition
#
# defines the structure of a xml document
# ## HTML Basics
# ---
#
# - HyperText Markup Language
# - HTML is special case of XML, where tags and attributes are predefined.
#
# *WebBrowser gets HTML as text from webserver, Creates a structure using HTML. The structure object is called DOM (Document Object Model) Tree.*
#
# *Browser then applies CSS (Cascading Style Sheet) to the DOM tree.*
#
# *Then Finally runs Javascript using embedded VM (Virtual Machine).*
# ```html
# <html>
# <head>
# <title></title>
# <meta>
# </head>
# <body>
# <article></article>
# <div id="section" class="red-text" data-type="meta">
# <span></span>
# <p></p>
# </div>
# <table>
# <thead>
# <tr>
# <th>S No</th>
# <th>Name</th>
# <th>Action</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>1</td>
# <td>Apple</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Microsoft</td>
# </tr>
# <tr>
# <td colspan="2"></td>
# </tr>
# </tbody>
# </table>
# </body>
# </html>
# ```
# - _id_ and _class_ are builtin attributes, used to define semantics of the element,
# - Also used by CSS to apply rules.
# - _id_ are unique in a document, i.e `section` should not be used elsewhere.
# - _class_ are used to identify similar kind of elements, or group certain properties
# - those attributes starting with `data-` are custom attributes i.e we are allowed to define such attributes
# - these attributes are used by many javascript frameworks as well for applying some behavior to the element
# *Go through w3schools html tutorial, [w3schools.com](s3schools.com) for complete HTML reference.*
|
References/A.HTML Basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf_gpu
# language: python
# name: tf_gpu
# ---
# ---
# # Clustering and Classification using Knowledge Graph Embeddings
# ---
#
# In this tutorial we will explore how to use the knowledge embeddings generated by a graph of international football matches (since the 19th century) in clustering and classification tasks. Knowledge graph embeddings are typically used for missing link prediction and knowledge discovery, but they can also be used for entity clustering, entity disambiguation, and other downstream tasks. The embeddings are a form of representation learning that allow linear algebra and machine learning to be applied to knowledge graphs, which otherwise would be difficult to do.
#
#
# We will cover in this tutorial:
#
# 1. Creating the knowledge graph (i.e. triples) from a tabular dataset of football matches
# 2. Training the ComplEx embedding model on those triples
# 3. Evaluating the quality of the embeddings on a validation set
# 4. Clustering the embeddings, comparing to the natural clusters formed by the geographical continents
# 5. Applying the embeddings as features in classification task, to predict match results
# 6. Evaluating the predictive model on a out-of-time test set, comparing to a simple baseline
#
# We will show that knowledge embedding clusters manage to capture implicit geographical information from the graph and that they can be a useful feature source for a downstream machine learning classification task, significantly increasing accuracy from the baseline.
#
# ---
# ## Requirements
#
# A Python environment with the AmpliGraph library installed. Please follow the [install guide](http://docs.ampligraph.org/en/latest/install.html).
#
# Some sanity check:
# +
import numpy as np
import pandas as pd
import ampligraph
ampligraph.__version__
# -
# ## Dataset
#
# We will use the [International football results from 1872 to 2019](https://www.kaggle.com/martj42/international-football-results-from-1872-to-2017) available on Kaggle (public domain). It contains over 40 thousand international football matches. Each row contains the following information:
# 1. Match date
# 2. Home team name
# 3. Away team name
# 4. Home score (goals including extra time)
# 5. Away score (goals including extra time)
# 6. Tournament (whether it was a friendly match or part of a tournament)
# 7. City where match took place
# 8. Country where match took place
# 9. Whether match was on neutral grounds
#
# This dataset comes in a tabular format, therefore we will need to construct the knowledge graph ourselves.
import requests
url = 'https://ampligraph.s3-eu-west-1.amazonaws.com/datasets/football_graph.csv'
open('football_results.csv', 'wb').write(requests.get(url).content)
df = pd.read_csv("football_results.csv").sort_values("date")
df.isna().sum()
# Dropping matches with unknown score:
df = df.dropna()
# The training set will be from 1872 to 2014, while the test set will be from 2014 to present date. Note that a temporal test set makes any machine learning task harder compared to a random shuffle.
df["train"] = df.date < "2014-01-01"
df.train.value_counts()
# ## Knowledge graph creation
# We are going to create a knowledge graph from scratch based on the match information. The idea is that each match is an entity that will be connected to its participating teams, geography, characteristics, and results.
#
# The objective is to generate a new representation of the dataset where each data point is an triple in the form:
#
# <subject, predicate, object>
#
# First we need to create the entities (subjects and objects) that will form the graph. We make sure teams and geographical information result in different entities (e.g. the Brazilian team and the corresponding country will be different).
# Entities naming
df["match_id"] = df.index.values.astype(str)
df["match_id"] = "Match" + df.match_id
df["city_id"] = "City" + df.city.str.title().str.replace(" ", "")
df["country_id"] = "Country" + df.country.str.title().str.replace(" ", "")
df["home_team_id"] = "Team" + df.home_team.str.title().str.replace(" ", "")
df["away_team_id"] = "Team" + df.away_team.str.title().str.replace(" ", "")
df["tournament_id"] = "Tournament" + df.tournament.str.title().str.replace(" ", "")
df["neutral"] = df.neutral.astype(str)
# Then, we create the actual triples based on the relationship between the entities. We do it only for the triples in the training set (before 2014).
triples = []
for _, row in df[df["train"]].iterrows():
# Home and away information
home_team = (row["home_team_id"], "isHomeTeamIn", row["match_id"])
away_team = (row["away_team_id"], "isAwayTeamIn", row["match_id"])
# Match results
if row["home_score"] > row["away_score"]:
score_home = (row["home_team_id"], "winnerOf", row["match_id"])
score_away = (row["away_team_id"], "loserOf", row["match_id"])
elif row["home_score"] < row["away_score"]:
score_away = (row["away_team_id"], "winnerOf", row["match_id"])
score_home = (row["home_team_id"], "loserOf", row["match_id"])
else:
score_home = (row["home_team_id"], "draws", row["match_id"])
score_away = (row["away_team_id"], "draws", row["match_id"])
home_score = (row["match_id"], "homeScores", np.clip(int(row["home_score"]), 0, 5))
away_score = (row["match_id"], "awayScores", np.clip(int(row["away_score"]), 0, 5))
# Match characteristics
tournament = (row["match_id"], "inTournament", row["tournament_id"])
city = (row["match_id"], "inCity", row["city_id"])
country = (row["match_id"], "inCountry", row["country_id"])
neutral = (row["match_id"], "isNeutral", row["neutral"])
year = (row["match_id"], "atYear", row["date"][:4])
triples.extend((home_team, away_team, score_home, score_away,
tournament, city, country, neutral, year, home_score, away_score))
# Note that we treat some literals (year, neutral match, home score, away score) as discrete entities and they will be part of the final knowledge graph used to generate the embeddings. We limit the number of score entities by clipping the score to be at most 5.
# Below we can see visualise a subset of the graph related to the infamous [Maracanazo](https://en.wikipedia.org/wiki/Uruguay_v_Brazil_(1950_FIFA_World_Cup)):
#
# 
# The whole graph related to this match can be summarised by the triples below:
triples_df = pd.DataFrame(triples, columns=["subject", "predicate", "object"])
triples_df[(triples_df.subject=="Match3129") | (triples_df.object=="Match3129")]
# ## Training knowledge graph embeddings
#
# We split our training dataset further into training and validation, where the new training set will be used to the knowledge embedding training and the validation set will be used in its evaluation. The test set will be used to evaluate the performance of the classification algorithm built on top of the embeddings.
#
# What differs from the standard method of randomly sampling N points to make up our validation set is that our data points are two entities linked by some relationship, and we need to take care to ensure that all entities are represented in train and validation sets by at least one triple.
#
# To accomplish this, AmpliGraph provides the [`train_test_split_no_unseen`](https://docs.ampligraph.org/en/latest/generated/ampligraph.evaluation.train_test_split_no_unseen.html#train-test-split-no-unseen) function.
# +
from ampligraph.evaluation import train_test_split_no_unseen
X_train, X_valid = train_test_split_no_unseen(np.array(triples), test_size=10000)
# -
print('Train set size: ', X_train.shape)
print('Test set size: ', X_valid.shape)
# AmpliGraph has implemented [several Knowledge Graph Embedding models](https://docs.ampligraph.org/en/latest/ampligraph.latent_features.html#knowledge-graph-embedding-models) (TransE, ComplEx, DistMult, HolE), but to begin with we're just going to use the [ComplEx](https://docs.ampligraph.org/en/latest/generated/ampligraph.latent_features.ComplEx.html#ampligraph.latent_features.ComplEx) model, which is known to bring state-of-the-art predictive power.
#
# The hyper-parameter choice was based on the [best results](https://docs.ampligraph.org/en/latest/experiments.html) we have found so far for the ComplEx model applied to some benchmark datasets used in the knowledge graph embeddings community. This tutorial does not cover [hyper-parameter tuning](https://docs.ampligraph.org/en/latest/examples.html#model-selection).
#
# +
from ampligraph.latent_features import ComplEx
model = ComplEx(batches_count=50,
epochs=300,
k=100,
eta=20,
optimizer='adam',
optimizer_params={'lr':1e-4},
loss='multiclass_nll',
regularizer='LP',
regularizer_params={'p':3, 'lambda':1e-5},
seed=0,
verbose=True)
# -
# Lets go through the parameters to understand what's going on:
#
# - **`batches_count`** : the number of batches in which the training set is split during the training loop. If you are having into low memory issues than settings this to a higher number may help.
# - **`epochs`** : the number of epochs to train the model for.
# - **`k`**: the dimensionality of the embedding space.
# - **`eta`** ($\\eta$) : the number of negative, or false triples that must be generated at training runtime for each positive, or true triple.
# - **`optimizer`** : the Adam optimizer, with a learning rate of 1e-4 set via the *optimizer_params* kwarg.
# - **`loss`** : pairwise loss, with a margin of 0.5 set via the *loss_params* kwarg.
# - **`regularizer`** : $L_p$ regularization with $p=3$, i.e. l3 regularization. $\\lambda$ = 1e-5, set via the *regularizer_params* kwarg.
# - **`seed`** : random seed, used for reproducibility.
# - **`verbose`** - displays a progress bar.
# Training should take around 10 minutes on a modern GPU:
# +
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.ERROR)
model.fit(X_train)
# -
# ## Evaluating knowledge embeddings
#
# AmpliGraph aims to follow scikit-learn's ease-of-use design philosophy and simplify everything down to **`fit`**, **`evaluate`**, and **`predict`** functions.
#
# However, there are some knowledge graph specific steps we must take to ensure our model can be trained and evaluated correctly. The first of these is defining the filter that will be used to ensure that no negative statements generated by the corruption procedure are actually positives. This is simply done by concatenating our train and test sets. Now when negative triples are generated by the corruption strategy, we can check that they aren't actually true statements.
filter_triples = np.concatenate((X_train, X_valid))
# For this we'll use the `evaluate_performance` function:
#
# - **`X`** - the data to evaluate on. We're going to use our test set to evaluate.
# - **`model`** - the model we previously trained.
# - **`filter_triples`** - will filter out the false negatives generated by the corruption strategy.
# - **`use_default_protocol`** - specifies whether to use the default corruption protocol. If True, then subj and obj are corrupted separately during evaluation.
# - **`verbose`** - displays a progress bar.
# +
from ampligraph.evaluation import evaluate_performance
ranks = evaluate_performance(X_valid,
model=model,
filter_triples=filter_triples,
use_default_protocol=True,
verbose=True)
# -
# We're going to use the mrr_score (mean reciprocal rank) and hits_at_n_score functions.
#
# - **mrr_score**: The function computes the mean of the reciprocal of elements of a vector of rankings ranks.
# - **hits_at_n_score**: The function computes how many elements of a vector of rankings ranks make it to the top n positions.
# +
from ampligraph.evaluation import mr_score, mrr_score, hits_at_n_score
mr = mr_score(ranks)
mrr = mrr_score(ranks)
print("MRR: %.2f" % (mrr))
print("MR: %.2f" % (mr))
hits_10 = hits_at_n_score(ranks, n=10)
print("Hits@10: %.2f" % (hits_10))
hits_3 = hits_at_n_score(ranks, n=3)
print("Hits@3: %.2f" % (hits_3))
hits_1 = hits_at_n_score(ranks, n=1)
print("Hits@1: %.2f" % (hits_1))
# -
# We can interpret these results by stating that the model will rank the correct entity within the top-3 possibilities 29% of the time.
#
# By themselves, these metrics are not enough to conclude the usefulness of the embeddings in a downstream task, but they suggest that the embeddings have learned a reasonable representation enough to consider using them in more tasks.
# ## Clustering and embedding visualization
# To evaluate the subjective quality of the embeddings, we can visualise the embeddings on 2D space and also cluster them on the original space. We can compare the clustered embeddings with natural clusters, in this case the continent where the team is from, so that we have a ground truth to evaluate the clustering quality both qualitatively and quantitatively.
#
# Requirements:
#
# * seaborn
# * adjustText
# * incf.countryutils
#
# For seaborn and adjustText, simply install them with `pip install seaborn adjustText`.
#
# For incf.countryutils, do the following steps:
# ```bash
# git clone https://github.com/wyldebeast-wunderliebe/incf.countryutils.git
# # cd incf.countryutils
# pip install .```
from sklearn.decomposition import PCA
import matplotlib.pyplot as plt
import seaborn as sns
from adjustText import adjust_text
from incf.countryutils import transformations
# %matplotlib inline
# We create a map from the team ID (e.g. "TeamBrazil") to the team name (e.g. "Brazil") for visualization purposes.
id_to_name_map = {**dict(zip(df.home_team_id, df.home_team)), **dict(zip(df.away_team_id, df.away_team))}
# We now create a dictionary with the embeddings of all teams:
teams = pd.concat((df.home_team_id[df["train"]], df.away_team_id[df["train"]])).unique()
team_embeddings = dict(zip(teams, model.get_embeddings(teams)))
# We use PCA to project the embeddings from the 200 space into 2D space:
embeddings_2d = PCA(n_components=2).fit_transform(np.array([i for i in team_embeddings.values()]))
# We will cluster the teams embeddings on its original 200-dimensional space using the `find_clusters` in our discovery API:
# +
from ampligraph.discovery import find_clusters
from sklearn.cluster import KMeans
clustering_algorithm = KMeans(n_clusters=6, n_init=50, max_iter=500, random_state=0)
clusters = find_clusters(teams, model, clustering_algorithm, mode='entity')
# -
# This helper function uses the `incf.countryutils` library to translate country names to their corresponding continents.
def cn_to_ctn(country):
try:
return transformations.cn_to_ctn(id_to_name_map[country])
except KeyError:
return "unk"
# This dataframe contains for each team their projected embeddings to 2D space via PCA, their continent and the KMeans cluster. This will be used alongisde Seaborn to make the visualizations.
plot_df = pd.DataFrame({"teams": teams,
"embedding1": embeddings_2d[:, 0],
"embedding2": embeddings_2d[:, 1],
"continent": pd.Series(teams).apply(cn_to_ctn),
"cluster": "cluster" + pd.Series(clusters).astype(str)})
# We plot the results on a 2D scatter plot, coloring the teams by the continent or cluster and also displaying some individual team names.
#
# We always display the names of the top 20 teams (according to [FIFA rankings](https://en.wikipedia.org/wiki/FIFA_World_Rankings)) and a random subset of the rest.
# +
top20teams = ["TeamBelgium", "TeamFrance", "TeamBrazil", "TeamEngland", "TeamPortugal", "TeamCroatia", "TeamSpain",
"TeamUruguay", "TeamSwitzerland", "TeamDenmark", "TeamArgentina", "TeamGermany", "TeamColombia",
"TeamItaly", "TeamNetherlands", "TeamChile", "TeamSweden", "TeamMexico", "TeamPoland", "TeamIran"]
def plot_clusters(hue):
np.random.seed(0)
plt.figure(figsize=(12, 12))
plt.title("{} embeddings".format(hue).capitalize())
ax = sns.scatterplot(data=plot_df[plot_df.continent!="unk"], x="embedding1", y="embedding2", hue=hue)
texts = []
for i, point in plot_df.iterrows():
if point["teams"] in top20teams or np.random.random() < 0.1:
texts.append(plt.text(point['embedding1']+0.02, point['embedding2']+0.01, str(point["teams"])))
adjust_text(texts)
# -
# The first visualisation of the 2D embeddings shows the natural geographical clusters (continents), which can be seen as a form of the ground truth:
plot_clusters("continent")
# We can see above that the embeddings learned geographical similarities even though this information was not explicit on the original dataset.
#
# Now we plot the same 2D embeddings but with the clusters found by K-Means:
plot_clusters("cluster")
# We can see that K-Means found very similar cluster to the natural geographical clusters by the continents. This shows that on the 200-dimensional embedding space, similar teams appear close together, which can be captured by a clustering algorithm.
#
# Our evaluation of the clusters can be more objective by using a metric such as the [adjusted Rand score](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.adjusted_rand_score.html), which varies from -1 to 1, where 0 is random labelling and 1 is a perfect match:
from sklearn import metrics
metrics.adjusted_rand_score(plot_df.continent, plot_df.cluster)
# ## Classification
#
# We will use the knowledge embeddings to predict future matches as a classification problem.
#
# We can model it as a multiclass problem with three classes: home team wins, home team loses, draw.
#
# The embeddings are used directly as features to a XGBoost classifier.
#
# First we need to determine the target:
df["results"] = (df.home_score > df.away_score).astype(int) + \
(df.home_score == df.away_score).astype(int)*2 + \
(df.home_score < df.away_score).astype(int)*3 - 1
df.results.value_counts(normalize=True)
# Now we create a function that extracts the features (knowledge embeddings for home and away teams) and the target for a particular subset of the dataset:
def get_features_target(mask):
def get_embeddings(team):
return team_embeddings.get(team, np.full(200, np.nan))
X = np.hstack((np.vstack(df[mask].home_team_id.apply(get_embeddings).values),
np.vstack(df[mask].away_team_id.apply(get_embeddings).values)))
y = df[mask].results.values
return X, y
clf_X_train, y_train = get_features_target((df["train"]))
clf_X_test, y_test = get_features_target((~df["train"]))
clf_X_train.shape, clf_X_test.shape
# Note that we have 200 features by team because the ComplEx model uses imaginary and real number for its embeddings, so we have twice as many parameters as defined by `k=100` in its model definition.
#
# We also have some missing information from the embeddings of the entities (i.e. teams) that only appear in the test set, which are unlikely to be correctly classified:
np.isnan(clf_X_test).sum()/clf_X_test.shape[1]
# First install xgboost with `pip install xgboost`.
from xgboost import XGBClassifier
# Create a multiclass model with 500 estimators:
clf_model = XGBClassifier(n_estimators=500, max_depth=5, objective="multi:softmax")
# Fit the model using all of the training samples:
clf_model.fit(clf_X_train, y_train)
# The baseline accuracy for this problem is 47%, as that is the frequency of the most frequent class (home team wins):
df[~df["train"]].results.value_counts(normalize=True)
metrics.accuracy_score(y_test, clf_model.predict(clf_X_test))
# In conclusion, while the baseline for this classification problem was 47%, with just the knowledge embeddings alone we were able to build a classifier that achieves **54%** accuracy.
#
# As future work, we could add more features to the model (not embeddings related) and tune the model hyper-parameters.
|
docs/tutorials/ClusteringAndClassificationWithEmbeddings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://qworld.net" target="_blank" align="left"><img src="../qworld/images/header.jpg" align="left"></a>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\stateplus}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\stateminus}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\I}{ \mymatrix{rr}{1 & 0 \\ 0 & 1} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# $ \newcommand{\pstate}[1]{ \lceil \mspace{-1mu} #1 \mspace{-1.5mu} \rfloor } $
# $ \newcommand{\greenbit}[1] {\mathbf{{\color{green}#1}}} $
# $ \newcommand{\bluebit}[1] {\mathbf{{\color{blue}#1}}} $
# $ \newcommand{\redbit}[1] {\mathbf{{\color{red}#1}}} $
# $ \newcommand{\brownbit}[1] {\mathbf{{\color{brown}#1}}} $
# $ \newcommand{\blackbit}[1] {\mathbf{{\color{black}#1}}} $
# <font style="font-size:28px;" align="left"><b>Grover's Search: Implementation </b></font>
# <br>
# _prepared by <NAME> and <NAME>_
# <br><br>
# [<img src="../qworld/images/watch_lecture.jpg" align="left">](https://youtu.be/RHR01e7oN9U)
# <br><br><br>
# Now we will consider how to implement Grover's search. Let's recall the whole algorithm.
#
# We are given $N=2^n$ elements, and one element is marked. The task is to find this marked element.
#
# We are going to use $n$ qubits. At the beginning we apply Hadamard to each qubit, so we put our quantum state into superposition. The amplitude of each basis state $ \ket{0 \cdots 0}, \ldots, \ket{1 \cdots 1} $ is set to $ \frac{1}{\sqrt{N}} $. After that we iterate the following algorithm for several times:
# <ul>
# <li>Make a query: apply a query oracle operator to qubits - it flips the sign of the amplitude of the state that corresponds to the marked element.</li>
# <li>Inversion: apply a diffusion matrix - the amplitude of each state is reflected over the mean of all amplitudes.</li>
# </ul>
#
# Let's check how we can implement the query and inversion operations.
# <h3>Query operation</h3>
#
# <h4>Oracle</h4>
#
# Suppose that there exists a function $f:\{0,1\}^n \rightarrow \{0,1\}$ with the following properties:
#
# \begin{align*}
# f(x)&=1 &\mbox{ if $x$ is marked}\\
# f(x)&=0 &\mbox{ otherwise}
# \end{align*}
#
# Grover's algorithm does not actually search a list of elements, but given function $f$ with the above properties, it finds the element $x$ such that $f(x)=1$.
#
# <h3>Task 1 (Discussion)</h3>
#
# Consider the following function $f:\{0,1\}^2 \rightarrow \{0,1\}$. Which element is marked? How many queries should you make to $f$ to find out the marked element in the worst case?
# $$
# f: \begin{array}{c|c} \mathbf{In} & \mathbf{Out} \\ \hline \ket{00} & 0 \\ \ket{01} & 0 \\ \ket{10} & 0 \\ \ket{11} & 1 \end{array}
# $$
#
# $f$ is often called as the <font color="blue">oracle</font> or blackbox. Even though $f$ might not be reversible, it can be implemented in a reversible manner by using the following idea.
#
# <img src="images/foperator.png" width="20%" align="center">
#
# Here $U_f$, the corresponding quantum operator is defined as follows, where $\oplus$ denotes bitwise addition modulo 2 (XOR).
#
# $$U_f: \ket{x}\ket{y} \mapsto \ket{x}\ket{y \oplus f(x)} $$
#
# Note that this mapping is reversible. When $\ket{y}=\ket{0}$, you get exactly $f(x)$ in the output qubit.
# <h4>Sign flip </h4>
#
# In the rest of the discussion, we will assume that we are given a quantum circuit implementing the operator $U_f$. An operator which flips the sign of the amplitude of the state corresponding to the marked element $x$ can be constructed using <font color="blue">phase-kickback</font>.
#
# Now let's assume that $\ket{y}=\ket{-} = \frac{1}{\sqrt{2}} (\ket{0} - \ket{1})$ and investigate the effect of the operator $U_f$.
# \begin{align*}
# U_f \ket{x}\ket{-} &= U_f \ket{x} \frac{1}{\sqrt{2}} \mypar{ \ket{0}-\ket{1} }\\
# &= \frac{1}{\sqrt{2}} (U_f\ket{x}\ket{0}-U_f\ket{x}\ket{1}) \\
# &= \frac{1}{\sqrt{2}} (\ket{x}\ket{f(x)\oplus 0} - \ket{x}\ket{f(x)\oplus 1}) \\
# &= \ket{x} \frac{1}{\sqrt{2}}\mypar{ \ket{f(x)}-\ket{f(x)\oplus 1} } \\
# &= \ket{x} (-1)^{f(x)} \frac{1}{\sqrt{2}} \mypar{ \ket{0}-\ket{1} }\\
# &= (-1)^{f(x)} \ket{x} \ket{-}
# \end{align*}
# We have the following transformation:
# $$ \ket{x}\ket{-} \xrightarrow{U_f} (-1)^{f(x)} \ket{x}\ket{-} $$
# When $f(x)=1$, we see that a phase of -1 is kicked back to the front of the first register. Hence by preparing the output register in state $\ket{-}$ and applying $U_f$, we obtain the sign flip effect.
#
# Note that the even if we don't know anything about $f$ (that's why it is called a blackbox), we are able to flip the sign of the amplitude of the marked element by making a query to $f$ by setting output qubit to $\ket{-}$,
#
#
# <h3>Task 2</h3>
#
# Let $N=4$. Implement the query phase and check the unitary matrix for the query operator. Note that we are interested in the top-left $4 \times 4$ part of the matrix since the remaining parts are due to the ancilla qubit.
# You are given a function $f$ and its corresponding quantum operator $U_f$. First run the following cell to load operator $U_f$. Then you can make queries to $f$ by applying the operator $U_f$ via the following command:
#
# <pre>Uf(circuit,qreg).
# %run quantum.py
# Now use phase kickback to flip the sign of the marked element:
#
# <ul>
# <li>Set output qubit (qreg[2]) to $\ket{-}$ by applying X and H.</li>
# <li>Apply operator $U_f$
# <li>Set output qubit (qreg[2]) back.</li>
# </ul>
#
# (Can you guess the marked element by looking at the unitary matrix?)
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg = QuantumRegister(3)
#No need to define classical register as we are not measuring
mycircuit = QuantumCircuit(qreg)
#
#Your code here
#
job = execute(mycircuit,Aer.get_backend('unitary_simulator'))
u=job.result().get_unitary(mycircuit,decimals=3)
#We are interested in the top-left 4x4 part
for i in range(4):
s=""
for j in range(4):
val = str(u[i][j].real)
while(len(val)<5): val = " "+val
s = s + val
print(s)
mycircuit.draw(output='mpl')
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task2">click for our solution</a>
# <hr>
#
# In Task 2, we see that the query operator looks like an identity operator with one of the entries being equal to -1, which is responsible for the sign flip.
#
# $\mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 1 & 0 \\ 0 & 0 & 0 & -1}$.
#
# By looking at the matrix, we can guess that the marked element is $\ket{11}$. Note that in general one is not able to peek at the matrix corresponding to the query operator and therefore it is not possible to find out the marked element after a single query.
# <h3>Inversion operator </h3>
#
# [<img src="../qworld/images/watch_lecture.jpg" align="left">](https://youtu.be/L9W2f74s7gs)
# <br><br><br>
# To implement the inversion (diffusion) operation, we will need additional (ancilla) qubit. This is how we implement the inversion operator:
#
# <ul>
# <li>Set the ancilla qubit to $\ket{-}$ by applying X and H.</li>
# <li>Apply H to all qubits other than the ancilla.</li>
# <li>Apply X to all qubits other than the ancilla.</li>
# <li>Apply multiple controlled NOT operator, where the ancilla qubit is target and all other qubits are used for controlling.</li>
# <li>Apply X to the ancilla qubit.</li>
# <li>Apply X to all qubits other than the ancilla.</li>
# <li>Apply H to all qubits other than the ancilla.</li>
# <li>Set ancilla qubit back by applying X and H..</li>
# </ul>
# <h4>Why these gates are chosen? (Optional) </h4>
#
# Now let's try to understand why these gates are chosen. Let's recall the inversion operator:
#
# $$ 2 \mymatrix{ccc}{
# \frac{1}{N} & \cdots & \frac{1}{N} \\
# \vdots & \ddots & \vdots \\
# \frac{1}{N} & \cdots & \frac{1}{N} \\
# }
# - I . $$
#
#
# This operator is also called the <font color="blue"> diffusion operator</font>.
#
# Recall that the diffusion operator can be expressed as $D=2\ket{u}\bra{u}-I$ where $\ket{u}=H^{\otimes n}\ket{0^n}$ is the equal superposition vector. We will simply denote $\ket{0^n}$ by $\ket{\mathbf{0}}$.
#
# - To start with let's express $D$ as follows:
#
# \begin{align*}
# D=2\ket{u}\bra{u}-I &= 2H^{\otimes n}\ket{\mathbf{0}}\bra{\mathbf{0}}H^{\otimes n}-I \\
# &= 2H^{\otimes n}\ket{\mathbf{0}}\bra{\mathbf{0}}H^{\otimes n}-H^{\otimes n}H^{\otimes n}\\
# &=H^{\otimes n} (2\ket{\mathbf{0}}\bra{\mathbf{0}}H^{\otimes n}-H^{\otimes n}) \\
# &=H^{\otimes n} (2\ket{\mathbf{0}}\bra{\mathbf{0}}-I)H^{\otimes n}
# \end{align*}
#
# <font color="blue"><i>Looking at this expression, it is clear why we have H gates at the beginning and at the end</i>
# - Now let us look at the effect of applying $2\ket{\mathbf{0}}\bra{\mathbf{0}}-I$ to any arbitrary state.
#
# $(2\ket{\mathbf{0}}\bra{\mathbf{0}}-I) \ket{x} = 2\ket{\mathbf{0}}\braket{\mathbf{0}}{x}-\ket{x} .$
#
# If $\ket{x}=\ket{\mathbf{0}}$, since $\braket{\mathbf{0}}{\mathbf{0}}=1$, then $2\ket{\mathbf{0}}\braket{\mathbf{0}}{\mathbf{0}}-\ket{\mathbf{0}} = 2\ket{\mathbf{0}}-\ket{\mathbf{0}} = \ket{\mathbf{0}}$.
#
# If $\ket{x}\neq \ket{\mathbf{0}}$, since $\braket{\mathbf{0}}{x}=0$, then $2\ket{\mathbf{0}}\braket{\mathbf{0}}{x}-\ket{x}= 2\ket{\mathbf{0}}\cdot 0 -\ket{x} = -\ket{x}$.
#
# Hence, the effect of $2\ket{\mathbf{0}}\bra{\mathbf{0}}-I$ is flipping the amplitude of any state except $\ket{\mathbf{0}}$.
# - Now let's see how we can implement this operator. Let's define function $g$ as follows and let $U_g$ be the corresponding operator.
#
# \begin{align*}
# g(x)&=0 &\mbox{ if $x$ is $\ket{\mathbf{0}}$ }\\
# g(x)&=1 &\mbox{ otherwise},
# \end{align*}
#
#
#
# Let's set ancilla qubit to state $\ket{-}$ and apply operator $U_g$.
# \begin{align*}
# U_g \ket{x}\ket{-} &= (-1)^{g(x)} \ket{x} \ket{-}.
# \end{align*}
#
#
# Note that $U_g$ flips the amplitudes of the states other than $\ket{\mathbf{0}}$ and exactly implements $2\ket{\mathbf{0}}\bra{\mathbf{0}}-I$.
# - How to implement $U_g$?
#
# Consider $g'$ which is exactly the opposite of $g$.
#
# \begin{align*}
# g'(x)&=1 &\mbox{ if $x$ is $\ket{\mathbf{0}}$ }\\
# g'(x)&=0 &\mbox{ otherwise},
# \end{align*}
#
# To implement $U_g'$, we should output 1 when $x$ is $\ket{\bf 0}$, that is we should apply $X$ gate to the output qubit when all input qubits are in state 0.
#
# <font color="blue"> <i> That's why we first apply $X$ gate to the control qubits and then apply multiple controlled NOT gate where the target is the ancilla qubit. Afterwards, we set the control qubits back (Recall the multiple controlled constructions notebook.). </i>
# We have just obtained the opposite effect of $g$: the output qubit is set to 1 if all input qubits are in state 0.
#
# <font color="blue"><i>Therefore we apply $X$-gate to the ancila qubit so that the output is 1 when $x$ is not $\ket{\bf{0}}$ and the output is 0 otherwise. </i>
# <h3>Task 3</h3>
#
# Let $N=4$. Implement the inversion operator and check whether you obtain the following matrix:
#
# $\mymatrix{cccc}{-0.5 & 0.5 & 0.5 & 0.5 \\ 0.5 & -0.5 & 0.5 & 0.5 \\ 0.5 & 0.5 & -0.5 & 0.5 \\ 0.5 & 0.5 & 0.5 & -0.5}$.
# +
def inversion(circuit,quantum_reg):
#don't implement the first and last steps in which ancilla qubit is set
#
# your code is here
#
# -
# Below you can check the matrix of your inversion operator and how the circuit looks like. We are interested in the top-left $4 \times 4$ part of the matrix, the remaining parts are because we used ancilla qubit.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg1 = QuantumRegister(3)
mycircuit1 = QuantumCircuit(qreg1)
#set ancilla qubit
mycircuit1.x(qreg1[2])
mycircuit1.h(qreg1[2])
inversion(mycircuit1,qreg1)
#set ancilla qubit back
mycircuit1.h(qreg1[2])
mycircuit1.x(qreg1[2])
job = execute(mycircuit1,Aer.get_backend('unitary_simulator'))
u=job.result().get_unitary(mycircuit1,decimals=3)
for i in range(4):
s=""
for j in range(4):
val = str(u[i][j].real)
while(len(val)<5): val = " "+val
s = s + val
print(s)
mycircuit1.draw(output='mpl')
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task3">click for our solution</a>
# <h3>Task 4: Testing Grover's search</h3>
#
# Now we are ready to test our operations and run Grover's search. Suppose that there are 4 elements in the list and try to find the marked element.
#
# You are given the operator $U_f$. First run the following cell to load it. You can access it via <pre>Uf(circuit,qreg).</pre>
# qreg[2] is the ancilla qubit and it is shared by the query and the inversion operators.
#
# Which state do you observe the most?
# %run quantum.py
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg = QuantumRegister(3)
creg = ClassicalRegister(2)
mycircuit = QuantumCircuit(qreg,creg)
#Grover
#initial step - equal superposition
#
#your code here
#
#set ancilla
#
#your code here
#
#change the number of iterations
iterations=1
#Grover's iterations.
#
#Query operator
#
#Inversion operator
#
#set ancilla
#
#your code here
#
mycircuit.measure(qreg[0],creg[0])
mycircuit.measure(qreg[1],creg[1])
job = execute(mycircuit,Aer.get_backend('qasm_simulator'),shots=10000)
counts = job.result().get_counts(mycircuit)
# print the outcome
for outcome in counts:
print(outcome,"is observed",counts[outcome],"times")
mycircuit.draw(output='mpl')
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task4">click for our solution</a>
# <h3>Task 5 (Optional, challenging)</h3>
#
# Implement the inversion operation for $n=3$ ($N=8$). This time you will need 5 qubits - 3 for the operation, 1 for ancilla, and one more qubit to implement not gate controlled by three qubits.
#
# In the implementation the ancilla qubit will be qubit 3, while qubits for control are 0, 1 and 2; qubit 4 is used for the multiple control operation. As a result you should obtain the following values in the top-left $8 \times 8$ entries:
#
# $\mymatrix{cccccccc}{-0.75 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 \\ 0.25 & -0.75 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 \\ 0.25 & 0.25 & -0.75 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 \\ 0.25 & 0.25 & 0.25 & -0.75 & 0.25 & 0.25 & 0.25 & 0.25 \\ 0.25 & 0.25 & 0.25 & 0.25 & -0.75 & 0.25 & 0.25 & 0.25 \\ 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & -0.75 & 0.25 & 0.25 \\ 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & -0.75 & 0.25 \\ 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & 0.25 & -0.75}$.
# +
def big_inversion(circuit,quantum_reg):
#
# your code is here
#
# -
# Below you can check the matrix of your inversion operator. We are interested in the top-left $8 \times 8$ part of the matrix, the remaining parts are because of additional qubits.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
big_qreg2 = QuantumRegister(5)
big_mycircuit2 = QuantumCircuit(big_qreg2)
#set ancilla
big_mycircuit2.x(big_qreg2[3])
big_mycircuit2.h(big_qreg2[3])
big_inversion(big_mycircuit2,big_qreg2)
#set ancilla back
big_mycircuit2.h(big_qreg2[3])
big_mycircuit2.x(big_qreg2[3])
job = execute(big_mycircuit2,Aer.get_backend('unitary_simulator'))
u=job.result().get_unitary(big_mycircuit2,decimals=3)
for i in range(8):
s=""
for j in range(8):
val = str(u[i][j].real)
while(len(val)<6): val = " "+val
s = s + val
print(s)
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task5">click for our solution</a>
# <h3>Task 6: Testing Grover's search for 8 elements (Optional, challenging)</h3>
#
# Now we will test Grover's search on 8 elements.
#
# You are given the operator $U_{f_8}$. First run the following cell to load it. You can access it via:
#
# <pre>Uf_8(circuit,qreg)</pre>
#
# Which state do you observe the most?
# %run quantum.py
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg8 = QuantumRegister(5)
creg8 = ClassicalRegister(3)
mycircuit8 = QuantumCircuit(qreg8,creg8)
#
#Your code here
#
job = execute(mycircuit8,Aer.get_backend('qasm_simulator'),shots=10000)
counts8 = job.result().get_counts(mycircuit8)
# print the reverse of the outcome
for outcome in counts8:
print(outcome,"is observed",counts8[outcome],"times")
mycircuit8.draw(output='mpl')
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task6">click for our solution</a>
# <h3>Task 7 (optional)</h3>
#
# Do you remember the optimal number of iterations to find the marked element with Grover's search? You are given the following code which implements Grover's search. (<i>giant_oracle</i> and <i>giant_diffusion</i> are already implemented.) There is a single marked element. First, determine the total number of elements in the search space. After that, try to find the optimal number of iterations. You can check your guess by changing the value of <i>iteration_count</i>. Please also check less optimal options like <i>iteration_count-1</i>, <i>iteration_count+1</i>.
#
# Try also calling <i>giant_oracle2</i> which marks 2 elements and try to find out the optimal number of iterations.
# +
# %run quantum.py
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg12 = QuantumRegister(19)
creg12 = ClassicalRegister(10)
mycircuit12 = QuantumCircuit(qreg12,creg12)
for i in range(10):
mycircuit12.h(qreg12[i])
mycircuit12.x(qreg12[10])
mycircuit12.h(qreg12[10])
#number of iterations - change this value
iteration_count = 1
for i in range(iteration_count):
giant_oracle2(mycircuit12,qreg12)
giant_diffusion(mycircuit12,qreg12)
mycircuit12.h(qreg12[10])
mycircuit12.x(qreg12[10])
for i in range(10):
mycircuit12.measure(qreg12[i],creg12[i])
job = execute(mycircuit12,Aer.get_backend('qasm_simulator'),shots=100000)
counts12 = job.result().get_counts(mycircuit12)
# print the reverse of the outcome
for outcome in counts12:
print(outcome,"is observed",counts12[outcome],"times")
# -
# <h3>Designing oracle functions</h3>
#
# Oracle function $f$ depends on the problem you want to solve. You can model many different problems (such as graph coloring, traveling salesman and many more) as a search problem. Elements in your search space correspond to quantum states. Instead of searching the whole space, you design $f$ so that it checks whether an element in the search space is the actual solution and marks it by outputing 1. Then you can use Grover's search to find the solution.
# Now we will design a very simple oracle function, which simply marks one of the elements. Suppose that $N=4$. Our elements in the search space are $\ket{00}$, $\ket{01}$, $\ket{10}$, $\ket{11}$ and suppose that we want to mark the element $\ket{11}$. This means that the function should output 1 if the input qubits are in state 1. This can be simply accomplished with the following function:
def oracle_11(circuit,qreg):
circuit1.ccx(qreg[0],qreg[1],qreg[2])
# Here qreg[0] and qreg[1] are the input qubits and qreg[2] is the output qubit. Let's check the matrix corresponding to the query phase.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg1 = QuantumRegister(3)
circuit1 = QuantumCircuit(qreg1)
# prepare ancilla qubit
circuit1.x(qreg1[2])
circuit1.h(qreg1[2])
#call the oracle
oracle_11(circuit1,qreg1)
# put ancilla qubit back into state |0>
circuit1.h(qreg1[2])
circuit1.x(qreg1[2])
job = execute(circuit1,Aer.get_backend('unitary_simulator'))
u=job.result().get_unitary(circuit1,decimals=3)
for i in range(4):
s=""
for j in range(4):
val = str(round(u[i][j].real,3))
while(len(val)<5): val = " "+val
s = s + val
print(s)
circuit1.draw(output='mpl')
# -
# Let's check another example, now our marked element is $\ket{01}$.
def oracle_01(circuit,qreg):
circuit.x(qreg[1])
circuit.ccx(qreg[0],qreg[1],qreg[2])
circuit.x(qreg[1])
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
qreg1 = QuantumRegister(3)
circuit1 = QuantumCircuit(qreg1)
# prepare ancilla qubit
circuit1.x(qreg1[2])
circuit1.h(qreg1[2])
#call the oracle
oracle_01(circuit1,qreg1)
# put ancilla qubit back into state |0>
circuit1.h(qreg1[2])
circuit1.x(qreg1[2])
job = execute(circuit1,Aer.get_backend('unitary_simulator'))
u=job.result().get_unitary(circuit1,decimals=3)
for i in range(4):
s=""
for j in range(4):
val = str(round(u[i][j].real,3))
while(len(val)<5): val = " "+val
s = s + val
print(s)
circuit1.draw(output='mpl')
# -
# <h3>Task 8</h3>
#
# Implement an oracle function which marks the element $\ket{00}$. Run Grover's search with the oracle you have implemented.
def oracle_00(circuit,qreg):
#Your code here
#
#
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
#Grover's search
# -
# <a href="Q92_Grovers_Search_Implementation_Solutions.ipynb#task8">click for our solution</a>
# <h3>Task 9 (Optional)</h3>
#
# Let $N=8$. Implement an oracle function which marks the elements $\ket{001}$ and $\ket{111}$. Run Grover's search with the oracle you have implemented.
def oracle_001_111(circuit,qreg):
#Your code here
#
#
# Further links:
#
# http://quantumgazette.blogspot.com/2017/12/grovers-algorithm-for-unstructured.html
#
# http://twistedoakstudios.com/blog/Post2644_grovers-quantum-search-algorithm
|
quantum-with-qiskit/Q92_Grovers_Search_Implementation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sklearn as skl
import pandas as pd
import numpy as np
from sklearn.cluster import KMeans
from sklearn.preprocessing import scale
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix,classification_report
df = pd.read_csv('C:/Users/joaqu/OneDrive/Escritorio/MovieClassifier/R_scripts/RDA_objects/tf134wide.csv')
df = pd.DataFrame(df)
df
targets = df['CLUSTER']
df = df.drop(columns=['CLUSTER'])
df = df.drop(columns=['Unnamed: 0'])
features_names= df.columns
features_names
targets = np.array(targets)
features_names = np.array(features_names)
data = np.array(df)
targets.shape
features_names.shape
data.shape
for n,i in enumerate(targets):
if i == 1:
targets[n] = 0
elif i == 3:
targets[n] = 1
elif i == 4:
targets[n] = 2
pd.DataFrame(targets).describe()
#insertar los features en una variable. Por convencion se denomina X a los features e Y a los target (o responses)
X = scale(data)
y = targets
targets
clustering = KMeans(n_clusters = 3, random_state=1)
clustering.fit(X)
# +
color_theme = np.array(['blue', 'red', 'yellow', 'orange', 'green'])
plt.scatter(x=df.shrek, y=df.girl, c=color_theme[clustering.labels_])
clustering.labels_.size
# -
print(classification_report(y,clustering.labels_))
clustering.labels_
targets
cl = pd.DataFrame(clustering.labels_)
cl.describe()
t =pd.DataFrame(targets)
t.describe()
|
R_scripts/KMEANS_MOVIES.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Created aggregated checklist
import pandas as pd
from collections import OrderedDict
data = pd.read_table('../data/interim/verified-checklist.tsv', dtype=object)
data.head()
# ## Retrieve valid records only
#
# Only records that contain ok, so no wrong matches, no matches or unverified synonyms.
valid_records = data[data['nameMatchValidation'].str.contains('^ok', regex=True, na=False)]
valid_records['index'].count()
# ## Aggregate and sort on gbifapi_acceptedName & collect unique values for some columns
def split_delimited_values(series, delimiter='|'):
# Splits values (of a series) containing a delimiter in multiple values
# Is a bit of a hack
# Only split if the series contains more than NaN values, otherwise, just return series back
if len(series.dropna()) > 0:
series = series.str.lower().str.split(delimiter).apply(pd.Series, 1).stack()
series = series.str.strip()
series.index = series.index.droplevel(-1)
return series
def get_unique_values(series):
# dropna(): remove NaN values
# tuple(): will collect all values of a series as a tuple
# set(): will only keep (unordered) unique values
# Returns a set
# It would have been easier to just return a list(), so we can sort it too,
# but then we get 'Function does not reduce' (http://stackoverflow.com/a/37955931)
# so have to use tuple()
return set(tuple(series.dropna()))
# Default dict syntax for aggregation does not preserve column order, which is why we use OrderedDict
unique_values_per_column = OrderedDict([
('kingdom', lambda x: get_unique_values(x)),
('datasetName', lambda x: get_unique_values(x)),
('euConcernStatus', lambda x: get_unique_values(x)),
('firstObservationYearBE', lambda x: get_unique_values(x)),
('firstObservationYearFL', lambda x: get_unique_values(x)),
('invasionStage', lambda x: get_unique_values(x)),
('habitat', lambda x: get_unique_values(x)),
('nativeRange', lambda x: get_unique_values(x)),
('introductionPathway', lambda x: get_unique_values(split_delimited_values(x))), # Can contain |
('presenceBE', lambda x: get_unique_values(x)),
('presenceFL', lambda x: get_unique_values(x)),
('presenceWA', lambda x: get_unique_values(x)),
('presenceBR', lambda x: get_unique_values(x)),
('gbifapi_scientificName', lambda x: get_unique_values(x)),
('index', lambda x: get_unique_values(x))
])
aggregated_records = valid_records.groupby(['gbifapi_acceptedScientificName','gbifapi_acceptedKey']).agg(unique_values_per_column).reset_index()
aggregated_records = aggregated_records.sort_values(by='gbifapi_acceptedScientificName')
aggregated_records.head()
# ## Sort and concatenate unique values per column
def sort_and_concatenate(aSet, sortAs=str):
# sortAs allows to sort more specific
# {9, 200, 12} will be sorted as:
# '12, 200, 9' with default str
# '9, 12, 200' with int
sortedList = sorted(aSet, key=sortAs)
concatenatedList = ' | '.join(str(i) for i in sortedList)
return concatenatedList # a string
aggregated_records['kingdom'] = aggregated_records['kingdom'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['datasetName'] = aggregated_records['datasetName'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['euConcernStatus'] = aggregated_records['euConcernStatus'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['firstObservationYearBE'] = aggregated_records['firstObservationYearBE'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['firstObservationYearFL'] = aggregated_records['firstObservationYearFL'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['invasionStage'] = aggregated_records['invasionStage'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['habitat'] = aggregated_records['habitat'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['nativeRange'] = aggregated_records['nativeRange'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['introductionPathway'] = aggregated_records['introductionPathway'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['presenceBE'] = aggregated_records['presenceBE'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['presenceFL'] = aggregated_records['presenceFL'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['presenceWA'] = aggregated_records['presenceWA'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['presenceBR'] = aggregated_records['presenceBR'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['gbifapi_scientificName'] = aggregated_records['gbifapi_scientificName'].apply(lambda x: sort_and_concatenate(x))
aggregated_records['index'] = aggregated_records['index'].apply(lambda x: sort_and_concatenate(x, int))
aggregated_records.head()
# ## Write aggregated records to file
aggregated_records.to_csv('../data/processed/aggregated-checklist.tsv', sep='\t', index=False)
|
notebooks/1-peterdesmet-create-aggregated-checklist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alphoenixbiz/Machine-Learning-Using-Python/blob/master/Logistic%20Regression/Example%200/Logistic_Regression_example0.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="3UnpwOomMKp_" colab_type="code" colab={}
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
# %matplotlib inline
# + id="8cu_rC2cMWTN" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="qPwTmjD7Mh-1" colab_type="code" colab={}
# !wget https://raw.githubusercontent.com/alphoenixbiz/Machine-Learning-Using-Python/master/Logistic%20Regression/Example%200/insurance_data.csv -P "gdrive/My Drive/Colab Notebooks/"
# + id="Mctl840VNH8C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="ae244ab0-790d-47ed-8ef1-78bdf57e2085"
df = pd.read_csv("gdrive/My Drive/Colab Notebooks/insurance_data.csv")
df.head()
# + id="qGgokjJCO22l" colab_type="code" colab={}
X= df[['age']]
y= df.bought_insurance
# + id="7UHsc8pzNmjn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="59d8d989-d813-49e8-af76-a83cfcb537be"
plt.scatter(X,y,marker='+',color='red')
# + id="2jJ8zBWnN6kU" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# + id="gx9wX9cYOFQl" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.1)
# + id="eb2tsBh1PZtr" colab_type="code" colab={}
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
# + id="cab7KFdUPjEn" colab_type="code" colab={}
model.fit(X_train, y_train)
# + id="ZqgWkigMUF4T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="3cb25ecb-36b2-44d5-b53f-97f1b31fb8fd"
X_test
# + id="ltA-KiJQPy-m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3f2d4495-5606-4fcd-fc1d-97824692f3ac"
y_predicted = model.predict(X_test)
y_predicted
# + id="lQ_7v94hRD-H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="af66c0c2-4c83-4b6f-a30d-3ab90cad4df6"
y_predicted_probability = model.predict_proba(X_test)
y_predicted_probability
# + id="6lowPn-9R4dQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d0312d2f-d778-48d4-e5c5-0d8ea0a546ab"
y_predicted_probability_y_equals_1 = model.predict_proba(X_test)
y_predicted_probability_y_equals_1[:,1]
# + id="OaL-12LIRmJJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="18b5b64b-7e3a-4e06-d6be-424ab270cca2"
model.score(X_test,y_test)
|
Logistic Regression/Example 0/Logistic_Regression_example0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from analyze_results import aggregate_runs_rejection_table, format_arc_table_results, default_methods, collect_configs, aggregate_runs_de, calc_rejection_curve_auc_seq
from pathlib import Path
import pandas as pd
from ue4nlp.ue_scores import *
def create_section_col(runs_dir, de):
runs_dir = Path(runs_dir) #/ 'results'
if not runs_dir.is_dir():
raise ValueError()
print(runs_dir)
default_methods = {
"bald": bald,
"sampled_max_prob": sampled_max_prob,
"variance": probability_variance,
"var_ratio": var_ratio,
"entropy": mean_entropy,
}
agg_res = aggregate_runs_rejection_table(runs_dir, methods=default_methods, de=de)
if agg_res.empty:
print('Broken')
raise ValueError()
improvement = format_arc_table_results(agg_res, baseline_col='max_prob')
improvement = improvement.loc[['max_prob'] + list(default_methods.keys())]
improvement.index = ['max_prob (baseline)'] + list(improvement.index[1:])
return improvement
def build_eval_table(dataset_paths, de = False):
BASELINE_INDEX = 'baseline (max_prob)'
columns = []
names = []
for name, paths in dataset_paths.items():
method_results = []
baseline = None
for path, dropout_type, layer in paths:
method_batch = create_section_col(path, de)
method_batch.index = [f'{dropout_type}|{layer}|{e}' for e in method_batch.index]
method_results.append(method_batch)
col = pd.concat(method_results, axis=0)
columns.append(col)
names.append(name)
name_map = {i : n for i,n in enumerate(names)}
return pd.concat(columns, axis=1).rename(columns=name_map)
# +
tasks = ['symptoms']
mc_types = ['MC']
path = "../../sbermed_ue/uncertainty-estimation/workdir/results/mc_symptoms_train_63/"
eval_table_sy = build_eval_table(
{
'symptoms' : [
(f'{path}/{tasks[0]}/{mc_types[0]}/last/10', 'MC', 'last'),
(f'{path}/{tasks[0]}/{mc_types[0]}/all/10', 'MC', 'all'),
]
},
de = False
)
eval_table_sy
# +
tasks = ['symptoms']
mc_types = ['MC']
path = "../../sbermed_ue/uncertainty-estimation/workdir/results/mc_symptoms_train_63/"
eval_table_sde = build_eval_table(
{
'symptoms' : [
(f'{path}/{tasks[0]}/{mc_types[0]}/last/10', '-', '-'),
]
},
de=True
)
eval_table_sde
# -
eval_table_sde.index = ['DeepEnsemble|' + idx.split('|')[-1] for idx in eval_table_sde.index]
res = pd.concat([eval_table_sde, eval_table_sy])
res = res[res.columns[:-1]]
res.index = ['max_prob (baseline)'] + list(res.index[1:])
res.iloc[[0, 8, 14, 2]]
# +
tasks = ['amazon']
mc_types = ['mahalanobis']
path = "../workdir/run_glue_for_model_series/electra_raw_sn/"
eval_table = build_eval_table(
{
'Amazon' : [
(f'{path}/{tasks[0]}/0.0/{mc_types[0]}', 'MD SN', '-'),
]
},
de = False
)
eval_table
# -
|
src/exps_notebooks/uncertainty_ratio_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
# # Analiza danych i uczenie maszynowe w Python
#
# Autor notebooka: <NAME>.
#
# ## SVM
#
# [Support Vector Machine (SVM)](https://en.wikipedia.org/wiki/Support_vector_machine) jest algorytmem regresji i klasyfikacji, który jest zbliżony do metod liniowych, ale może wykorzystywać funkcje jądrowe (kernel functions) do mapowania zależności nieliniowych, do przestrzeni liniowej. Takie rozwiązanie nazywa się trikiem jądrowym ([kernel trick](https://en.wikipedia.org/wiki/Kernel_method)).
#
# 
#
# W tym przykładzie skupimy się na regresji używając SVM.
#
# Wygenerujmy najpierw nielinowe dane.
# +
import numpy as np
from sklearn.svm import SVR
import matplotlib.pyplot as plt
# %matplotlib inline
# Dane przykładowe w postaci sinusa z szumem
X = np.sort(5 * np.random.rand(40, 1), axis=0)
y = np.sin(X).ravel()
y[::5] += 3 * (0.5 - np.random.rand(8))
# -
# Nauczmy teraz modele SVM z różnymi funkcjami jądrowymi.
svr_rbf = SVR(kernel='rbf', C=1e3, gamma=0.1)
svr_lin = SVR(kernel='linear', C=1e3)
svr_poly = SVR(kernel='poly', C=1e3, degree=2)
y_rbf = svr_rbf.fit(X, y).predict(X)
y_lin = svr_lin.fit(X, y).predict(X)
y_poly = svr_poly.fit(X, y).predict(X)
# Możemy wyniki przedstawić na wspólnym wykresie.
lw = 2
plt.figure(figsize=(10, 8))
plt.scatter(X, y, color='darkorange', label='x')
plt.plot(X, y_rbf, color='navy', lw=lw, label='Model RBF')
plt.plot(X, y_lin, color='c', lw=lw, label='Model liniowy')
plt.plot(X, y_poly, color='cornflowerblue', lw=lw, label='Model polimianowy')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Support Vector Regression')
plt.legend()
plt.show()
# +
# # %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn import datasets
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from sklearn.svm import SVR
COLUMNS = ['age', 'sex', 'bmi', 'bp', 's1', 's2', 's3', 's4', 's5', 's6']
# Przygotowujemy zbiór danych
diabetes = datasets.load_diabetes()
dataframe = pd.DataFrame(diabetes.data, columns=diabetes.feature_names).assign(target=diabetes.target)
# Dzielimy na zbiór danych treningowych i testowych
dane_treningowe = dataframe.iloc[:-20, :]
dane_testowe = dataframe.iloc[-20:, :]
# Wybór modelu
model = SVR(kernel='linear', C=1e3)
# Nauka modelu
model.fit(dane_treningowe[COLUMNS], dane_treningowe['target'])
dane_testowe = dane_testowe.assign(predict=lambda df: model.predict(df[COLUMNS]))
# Do wyświetlania
wspolczynniki = model.coef_
blad_sredniokwadratowy = mean_squared_error(dane_testowe['target'], model.predict(dane_testowe[COLUMNS]))
metryka_r2_wariancji = r2_score(dane_testowe['target'], dane_testowe['predict'])
print(f'Współczynniki: \n{wspolczynniki}')
print(f'Błąd średniokwadratowy: {blad_sredniokwadratowy:.2f}')
print(f'Metryka R2 (wariancji): {metryka_r2_wariancji:.2f}')
# Wyświetlanie wykresu
svr_rbf = SVR(kernel='rbf', C=1e3, gamma=0.1)
svr_lin = SVR(kernel='linear', C=1e3)
svr_poly = SVR(kernel='poly', C=1e3, degree=2)
y_rbf = svr_rbf.fit(dane_treningowe, dane_testowe).predict(dane_treningowe)
y_lin = svr_lin.fit(dane_treningowe, dane_testowe).predict(dane_treningowe)
y_poly = svr_poly.fit(dane_treningowe, dane_testowe).predict(dane_treningowe)
lw = 2
plt.figure(figsize=(10, 8))
plt.scatter(dane_treningowe, dane_testowe, color='darkorange', label='x')
plt.plot(dane_treningowe, y_rbf, color='navy', lw=lw, label='Model RBF')
plt.plot(dane_treningowe, y_lin, color='c', lw=lw, label='Model liniowy')
plt.plot(dane_treningowe, y_poly, color='cornflowerblue', lw=lw, label='Model polimianowy')
plt.xlabel('x')
plt.ylabel('y')
plt.title('Support Vector Regression')
plt.legend()
plt.show()
# -
# ## Zadanie
#
# 1. Wykorzystaj model liniowy SVM do przykładu z cukrzycą; czy jest lepszy?
# 1. Wykorzystaj model nieliniowy SVM do przykładu z cukrzycą; czy jest lepszy?
|
machine-learning/contrib/ml/2_svm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""Example of the optimization process for SGOOP. Here we find the weights of cos(phi),sin(phi),cos(psi),sin(psi)
for alanine dipeptide in vacuum from an unbiased trajectory. We use the cos and sin of dihedral angles to avoid
inaccuracy caused by the discontinuity of angles (pi = -pi)."""
import sgoop
import scipy.optimize as opt
import numpy as np
import time
import matplotlib.pyplot as plt
import kinetics as kin
np.set_printoptions(formatter={'float': '{: 0.3f}'.format})
# -
def opti_func(rc):
# function to be used in basinhopping. records function evaluations and returns -spectral gap
global nfev
nfev +=1
return -sgoop.rc_eval(rc)
def biased_opti_func(rc,old_rc):
# multidimensional equivalent of the optimization function
global nfev
nfev +=1
return -sgoop.biased_eval(rc,old_rc)
def print_fun(x, f, accepted):
# printing progress during optimization
global now,last,nfev,lastf
now=time.time()
print(x,end=' ')
if accepted == 1:
print("with spectral gap %.4f accepted after %3i runs (%.3f)" % (-f, nfev-lastf, now-last))
else:
print("with spectral gap %.4f declined after %3i runs (%.3f)" % (-f, nfev-lastf, now-last))
last=now
lastf=nfev
def temp_print_fun(x, f, accepted):
# printing progress while determining the temperature
global now,last,nfev,lastf,f_array,f_i
now=time.time()
print(x,end=' ')
if accepted == 1:
print("with spectral gap %.4f accepted after %3i runs (%.3f)" % (-f, nfev-lastf, now-last))
else:
print("with spectral gap %.4f declined after %3i runs (%.3f)" % (-f, nfev-lastf, now-last))
last=now
lastf=nfev
f_array[f_i]=-f
f_i+=1
def sgoophopping(step=1,iterations=3):
# basinhopping for 1 dimensional SGOOP
global last,nfev,lastf,f_array,f_i
f_array=np.zeros(50)
f_i=0
print('---------------------------------------------------------------------------')
print('BEGINNING',sgoop.wells,'WELL TEMPERATURE OPTIMIZATION')
print('---------------------------------------------------------------------------')
# Finding temperature value
start = time.time()
last = start
lastf = nfev = 0
minimizer_kwargs = {"options": {"maxiter":10}}
ret = opt.basinhopping(opti_func,step/10*np.ones(np.shape(sgoop.data_array)[1]),niter=50,T=.01,stepsize=step, minimizer_kwargs=minimizer_kwargs, callback=temp_print_fun)
T_new=np.mean(np.abs(f_array[0:-1]-f_array[1:]))
replicas=[None]*iterations
for i in range(iterations):
print('---------------------------------------------------------------------------')
print('BEGINNING',sgoop.wells,'WELL RC OPTIMIZATION #',i+1)
print('---------------------------------------------------------------------------')
start = time.time()
last = start
lastf = nfev = 0
minimizer_kwargs = {"options": {"maxiter":10}}
ret = opt.basinhopping(opti_func,step/10*np.ones(np.shape(sgoop.data_array)[1]),niter=200,T=T_new,stepsize=step, minimizer_kwargs=minimizer_kwargs, callback=print_fun)
replicas[i]=[ret.x,-ret.fun]
replicas=np.asarray(replicas)
soln=replicas[:,0][np.argmax(replicas[:,1])]
soln/=soln[np.argmax(np.abs(soln))]
p=sgoop.md_prob(soln)
wells_found=kin.find_wells(p)
return wells_found,soln
def biashopping(old_rc,step=1,iterations=3):
# basinhopping for multidimensional SGOOP
global last,nfev,lastf,f_array,f_i
f_array=np.zeros(50)
f_i=0
print('---------------------------------------------------------------------------')
print('BEGINNING',sgoop.wells,'WELL TEMPERATURE OPTIMIZATION')
print('---------------------------------------------------------------------------')
# Finding temperature value
start = time.time()
last = start
lastf = nfev = 0
minimizer_kwargs = {"options": {"maxiter":10},"args":old_rc}
ret = opt.basinhopping(biased_opti_func,step/10*np.ones(np.shape(sgoop.data_array)[1]),niter=50,T=.01,stepsize=step, minimizer_kwargs=minimizer_kwargs, callback=temp_print_fun)
T_new=np.mean(np.abs(f_array[0:-1]-f_array[1:]))
replicas=[None]*iterations
for i in range(iterations):
print('---------------------------------------------------------------------------')
print('BEGINNING',sgoop.wells,'WELL RC OPTIMIZATION #',i+1)
print('---------------------------------------------------------------------------')
start = time.time()
last = start
lastf = nfev = 0
minimizer_kwargs = {"options": {"maxiter":10},"args":old_rc}
ret = opt.basinhopping(biased_opti_func,step/10*np.ones(np.shape(sgoop.data_array)[1]),niter=200,T=T_new,stepsize=step, minimizer_kwargs=minimizer_kwargs, callback=print_fun)
replicas[i]=[ret.x,-ret.fun]
replicas=np.asarray(replicas)
soln=replicas[:,0][np.argmax(replicas[:,1])]
soln/=soln[np.argmax(np.abs(soln))]
p=sgoop.md_prob(soln)
wells_found=kin.find_wells(p)
return wells_found,soln
# +
sgoop.wells = 2
nfev=0
solutions=[]
wells_found,soln=sgoophopping(iterations=3) # calculating 2 well solution
p=sgoop.md_prob(soln)
plt.figure()
plt.plot(-np.ma.log(p))
plt.show()
while sgoop.wells == wells_found: # iteration for > 2 well solutions, continues while self consistent
print('---------------------------------------------------------------------------')
print('Optimized reaction coordinate:',soln)
print('This solution has',wells_found,'wells and is therefore accepted.')
print('---------------------------------------------------------------------------')
solutions.append([soln,sgoop.wells])
sgoop.wells+=1
wells_found,soln=sgoophopping(iterations=3)
p=sgoop.md_prob(soln)
plt.figure()
plt.plot(-np.ma.log(p))
plt.show()
else:
print('---------------------------------------------------------------------------')
print('This solution has',wells_found,'wells and is therefore rejected.')
print('---------------------------------------------------------------------------')
# -
rc = solutions[-1][0]
std = np.std(np.dot(sgoop.data_array,rc))
print('Optimized RC:')
print(rc)
print('Standard deviation (sigma in metad):')
print(std)
|
SGOOP Tutorial/Unbiased Loop Automated.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Load and serve a DistilBERT model from Mxnet on the fly
# !pip install mxnet gluonnlp pixiedust
# ## Train a model locally or remote
# +
import gluonnlp as nlp; import mxnet as mx;
# load distilbert
model, vocab = nlp.model.get_model('distilbert_6_768_12', dataset_name='distilbert_book_corpus_wiki_en_uncased');
# tokenize then transform
tokenizer = nlp.data.BERTTokenizer(vocab, lower=True);
transform = nlp.data.BERTSentenceTransform(tokenizer, max_seq_length=512, pair=False, pad=False);
sample = transform(['Hello world!']);
words, valid_len = mx.nd.array([sample[0]]), mx.nd.array([sample[1]])
model(words, valid_len) #if you want to save the model files, hybridize first
# If you want to save this model and upload as a file to S3 you will have to hybridize() it first before serializing.
# If you want to load the model internally using some hub, don't pass in a model or use Model = None
# model.hybridize()
# model(words, valid_len)
# # !mkdir mxnetmodel
# model.export(path='./mxnetmodel/')
# -
# ## Step 1 : Write a model transform script
#
# #### Make sure you have a ...
#
# - "load_model" function
# - input args are model path
# - returns loaded model object
# - model name is the same as what you saved the model file as (see above step)
# <br><br>
# - "predict" function
# - input args are the loaded model object and a payload
# - returns the result of model.predict
# - make sure you format it as a single (or multiple) string return inside a list for real time (for mini batch)
# - from a client, a list or string or np.array that is sent for prediction is interpreted as bytes. Do what you have to for converting back to list or string or np.array
# - return the error for debugging
#
# +
# %%writefile modelscript_mxnet.py
import gluonnlp as nlp; import mxnet as mx;
from joblib import load
import numpy as np
import os
import json
#Return loaded model
def load_model(modelpath):
model, vocab = nlp.model.get_model('distilbert_6_768_12', dataset_name='distilbert_book_corpus_wiki_en_uncased');
print("loaded")
return {'model':model,'vocab':vocab}
# return prediction based on loaded model (from the step above) and an input payload
def predict(modeldict, payload):
#set_trace()
model = modeldict['model']
vocab = modeldict['vocab']
tokenizer = nlp.data.BERTTokenizer(vocab, lower=True);
transform = nlp.data.BERTSentenceTransform(tokenizer, max_seq_length=512, pair=False, pad=False);
try:
# Local
if type(payload) == str:
sample = transform(payload);
elif type(payload) == bytes :
sample = transform(str(payload.decode()));
# Remote, standard payload comes in as a list of json strings with 'body' key
elif type(payload)==list:
sample = transform(payload[0]['body'].decode());
else:
return [json.dumps({'response':"Provide string or bytes string",
'payload':str(payload),
'type':str(type(payload))})]
words, valid_len = mx.nd.array([sample[0]]), mx.nd.array([sample[1]])
out = model(words, valid_len)
out = json.dumps({'output':out.asnumpy().tolist()})
except Exception as e:
out = str(e) #useful for debugging!
return [out]
# -
# ## Does this work locally? (not "_in a container locally_", but _actually_ in local)
from modelscript_mxnet import *
model = load_model('') # path doesn't matter here since we're loading the model directly in the script
predict(model,'Hello World!')[0]
# ### ok great! Now let's install ezsmdeploy
# !pip install ezsmdeploy
import ezsmdeploy
# #### If you have been running other inference containers in local mode, stop existing containers to avoid conflict
# !docker container stop $(docker container ls -aq) >/dev/null
# ## Deploy locally
ez = ezsmdeploy.Deploy(model = None, #loading distilbert model in script from hub
script = 'modelscript_mxnet.py',
requirements = ['pyarrow','mxnet', 'gluonnlp','numpy','joblib'], #or pass in the path to requirements.txt
instance_type = 'local',
wait = True)
# ## Test containerized version locally
# Since you are downloading this model from a hub, the first time you invoke it will be slow, so invoke again to get an inference without all of the container logs
out = ez.predictor.predict('Hello World').decode()
out
# ## Deploy on SageMaker
# !./src/build-docker.sh
ezonsm = ezsmdeploy.Deploy(model = None, #loading distilbert model in script from hub
script = 'modelscript_mxnet.py',
requirements = ['pyarrow','mxnet', 'gluonnlp','numpy','joblib'], #or pass in the path to requirements.txt
instance_type = 'ml.m4.xlarge',
wait = True)
out = ezonsm.predictor.predict('Hello World').decode()
out
ezonsm.predictor.delete_endpoint()
|
notebooks/.ipynb_checkpoints/Using ezsmdeploy for mxnet deployments-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="A4uesPTMKoSJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 551} outputId="243b7a50-ab1c-4077-a3b1-4cd641bdeae8" executionInfo={"status": "ok", "timestamp": 1583419852419, "user_tz": -60, "elapsed": 14168, "user": {"displayName": "<NAME>15bkiewicz", "photoUrl": "", "userId": "13490438346564972102"}}
# !pip install --upgrade tables
# !pip install eli5
# !pip install xgboost
# + id="DCLmaVY-LSmk" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn .ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
# + id="q0a-8EeKPm_u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="21e5d7a9-6780-46a5-cbb0-f23a8abbaf18" executionInfo={"status": "ok", "timestamp": 1583422050978, "user_tz": -60, "elapsed": 555, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
# cd '/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_car'
# + id="VFsqA-PqPp2B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f60c9fcf-913c-4b89-bcb4-cc03a0840907" executionInfo={"status": "ok", "timestamp": 1583422055089, "user_tz": -60, "elapsed": 2569, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df = pd.read_hdf('data/car.h5')
df.shape
# + [markdown] id="Y-n75VcYP38j" colab_type="text"
# ## Feature Engineering
# + id="hXiqwYBSPvYW" colab_type="code" colab={}
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
# + id="4f6IwoF8QAWF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f4c7cacd-b314-4793-8bcd-da73405c575f" executionInfo={"status": "ok", "timestamp": 1583422063423, "user_tz": -60, "elapsed": 627, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
# + id="RANN1NNmTCVP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="19c01405-3638-45ee-9195-1c05a2986801" executionInfo={"status": "ok", "timestamp": 1583422073544, "user_tz": -60, "elapsed": 5485, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
x = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, x, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="6GwaUH-3QEbq" colab_type="code" colab={}
def run_model(model, feats):
x = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, x, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + [markdown] id="ZNIyyp4HRphJ" colab_type="text"
# ## DecissionTree
# + id="KTn8ybgcQf93" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="529b2dd7-5c04-4eea-d729-9f378caec3d1" executionInfo={"status": "ok", "timestamp": 1583422160482, "user_tz": -60, "elapsed": 5181, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
run_model( DecisionTreeRegressor(max_depth=5), cat_feats )
# + [markdown] id="nHvtXNgyUN8u" colab_type="text"
# ## Random Forest
# + id="XZaHeQHhT__C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8dd2b24c-51ed-4993-c651-e10c9dc662aa" executionInfo={"status": "ok", "timestamp": 1583422491376, "user_tz": -60, "elapsed": 124617, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model( model, cat_feats )
# + id="EkHnZbxTU-Lh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="859f4c1c-73dc-4390-e6f9-8516290f486a" executionInfo={"status": "ok", "timestamp": 1583422596877, "user_tz": -60, "elapsed": 59940, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
xgb_params = {
'max_depth':5,
'n_estimators':50,
'learning_rate':0.1,
'seed':0
}
run_model(xgb.XGBRegressor(**xgb_params), cat_feats )
# + id="LqqwMhmyVnuw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 417} outputId="4a4b0679-2132-405e-c344-c0042b034574" executionInfo={"status": "ok", "timestamp": 1583422997069, "user_tz": -60, "elapsed": 366047, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(x,y)
imp = PermutationImportance(m, random_state=0).fit(x,y)
eli5.show_weights(imp, feature_names=cat_feats)
# + id="xVxre7QuV-sX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e52c4db2-2e3b-4bee-fa2e-97efc62210ca" executionInfo={"status": "ok", "timestamp": 1583423003076, "user_tz": -60, "elapsed": 647, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
len(cat_feats)
# + id="n0WMkg1dXtrb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="87d08d25-b969-44c4-fbbd-9d4665d1dbb0" executionInfo={"status": "ok", "timestamp": 1583423090515, "user_tz": -60, "elapsed": 536, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
feats = ['param_napęd_cat','param_rok-produkcji_cat','param_stan_cat','param_skrzynia-biegów_cat','param_faktura-vat_cat','param_moc_cat','param_marka-pojazdu_cat','feature_kamera-cofania_cat','param_typ_cat','param_pojemność-skokowa_cat','seller_name_cat','feature_wspomaganie-kierownicy_cat','param_model-pojazdu_cat','param_wersja_cat','param_kod-silnika_cat','feature_system-start-stop_cat','feature_asystent-pasa-ruchu_cat','feature_czujniki-parkowania-przednie_cat','feature_łopatki-zmiany-biegów_cat','feature_regulowane-zawieszenie_cat']
len(feats)
# + id="l5GngNhAYEkd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="4f696eb5-dd88-454a-ef19-80eae67c5a23" executionInfo={"status": "ok", "timestamp": 1583423695469, "user_tz": -60, "elapsed": 61125, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
feats = ['param_rok-produkcji__cat','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), cat_feats )
# + id="fTUOjh_JWvjU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="06d018d7-1bb1-4bab-8b03-fea6b60b7e77" executionInfo={"status": "ok", "timestamp": 1583423729833, "user_tz": -60, "elapsed": 568, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_napęd'].unique()
# + id="UjvU-EkMXTpl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 201} outputId="f4cfea9a-36c0-40f6-b0a6-04450263ed3c" executionInfo={"status": "ok", "timestamp": 1583423731598, "user_tz": -60, "elapsed": 552, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_rok-produkcji'].unique()
# + id="LFYzFeoMaOYo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="61057246-79e6-4b14-b04e-a2cf1936ae8e" executionInfo={"status": "ok", "timestamp": 1583424004561, "user_tz": -60, "elapsed": 13801, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x:-1 if str(x)=='None' else int(x))
feats = ['param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats )
# + id="wb7ucykMbMo9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="caba6569-cf02-4780-f206-8dc35f3b19b5" executionInfo={"status": "ok", "timestamp": 1583424020011, "user_tz": -60, "elapsed": 768, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_moc'].unique()
# + id="EGgI2hEGbQ5i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 237} outputId="97d6442f-60af-4443-f691-cc87232c30fe" executionInfo={"status": "ok", "timestamp": 1583424043822, "user_tz": -60, "elapsed": 558, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
# + id="s-12BsgpbXeb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="b4fe39f0-fb2e-4b3c-b912-ca5b455bfdb6" executionInfo={"status": "ok", "timestamp": 1583424322366, "user_tz": -60, "elapsed": 12966, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x:-1 if str(x)=='None' else int(x))
feats = ['param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats )
# + id="ZsM3MvKPcYdc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="c9a55047-22fc-42d0-b102-cc72b639aa73" executionInfo={"status": "ok", "timestamp": 1583424326764, "user_tz": -60, "elapsed": 636, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_pojemność-skokowa'].unique()
# + id="lZmiqJWlcbui" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="de611181-e292-4edf-dbca-fe0a4e61ed01" executionInfo={"status": "ok", "timestamp": 1583424674023, "user_tz": -60, "elapsed": 12967, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13490438346564972102"}}
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int( str(x).split('cm')[0].replace(' ', '')))
feats = ['param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa','seller_name__cat','feature_wspomaganie-kierownicy__cat', 'param_model-pojazdu__cat','param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats )
# + id="xkRRpxqrdqV6" colab_type="code" colab={}
|
day4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Use scikit-learn and AI lifecycle capabilities to predict Boston house prices with `ibm-watson-machine-learning`
#
# This notebook contains steps and code to demonstrate support of AI Lifecycle features in Watson Machine Learning Service. It contains steps and code to work with [ibm-watson-machine-learning](https://pypi.python.org/pypi/ibm-watson-machine-learning) library available in PyPI repository. It also introduces commands for getting model and training data, persisting model, deploying model, scoring it, updating the model and redeploying it.
#
# Some familiarity with Python is helpful. This notebook uses Python 3.
# ## Learning goals
#
# The learning goals of this notebook are:
#
# - Download an externally trained scikit-learn model with dataset.
# - Persist an external model in Watson Machine Learning repository.
# - Deploy model for online scoring using client library.
# - Score sample records using client library.
# - Update previously persisted model.
# - Redeploy model in-place.
# - Scale deployment.
#
#
# ## Contents
#
# This notebook contains the following parts:
#
# 1. [Setup](#setup)
# 2. [Download externally created scikit model and data](#download)
# 3. [Persist externally created scikit model](#persistence)
# 4. [Deploy and score in a Cloud](#scoring)
# 5. [Persist new version of the model](#update_model)
# 6. [Redeploy new version of the model](#redeploy)
# 7. [Deployment scaling](#scale)
# 8. [Clean up](#cleanup)
# 9. [Summary and next steps](#summary)
# <a id="setup"></a>
# ## 1. Set up the environment
#
# Before you use the sample code in this notebook, you must perform the following setup tasks:
#
# - Create a <a href="https://console.ng.bluemix.net/catalog/services/ibm-watson-machine-learning/" target="_blank" rel="noopener no referrer">Watson Machine Learning (WML) Service</a> instance (a free plan is offered and information about how to create the instance can be found <a href="https://dataplatform.cloud.ibm.com/docs/content/wsj/analyze-data/ml-service-instance.html?context=analytics" target="_blank" rel="noopener no referrer">here</a>).
# ### Connection to WML
#
# Authenticate the Watson Machine Learning service on IBM Cloud. You need to provide platform `api_key` and instance `location`.
#
# You can use [IBM Cloud CLI](https://cloud.ibm.com/docs/cli/index.html) to retrieve platform API Key and instance location.
#
# API Key can be generated in the following way:
# ```
# ibmcloud login
# ibmcloud iam api-key-create API_KEY_NAME
# ```
#
# In result, get the value of `api_key` from the output.
#
#
# Location of your WML instance can be retrieved in the following way:
# ```
# ibmcloud login --apikey API_KEY -a https://cloud.ibm.com
# ibmcloud resource service-instance WML_INSTANCE_NAME
# ```
#
# In result, get the value of `location` from the output.
# **Tip**: Your `Cloud API key` can be generated by going to the [**Users** section of the Cloud console](https://cloud.ibm.com/iam#/users). From that page, click your name, scroll down to the **API Keys** section, and click **Create an IBM Cloud API key**. Give your key a name and click **Create**, then copy the created key and paste it below. You can also get a service specific url by going to the [**Endpoint URLs** section of the Watson Machine Learning docs](https://cloud.ibm.com/apidocs/machine-learning). You can check your instance location in your <a href="https://console.ng.bluemix.net/catalog/services/ibm-watson-machine-learning/" target="_blank" rel="noopener no referrer">Watson Machine Learning (WML) Service</a> instance details.
#
# You can also get service specific apikey by going to the [**Service IDs** section of the Cloud Console](https://cloud.ibm.com/iam/serviceids). From that page, click **Create**, then copy the created key and paste it below.
#
# **Action**: Enter your `api_key` and `location` in the following cell.
api_key = 'PASTE YOUR PLATFORM API KEY HERE'
location = 'PASTE YOUR INSTANCE LOCATION HERE'
wml_credentials = {
"apikey": api_key,
"url": 'https://' + location + '.ml.cloud.ibm.com'
}
# ### Install and import the `ibm-watson-machine-learning` package
# **Note:** `ibm-watson-machine-learning` documentation can be found <a href="http://ibm-wml-api-pyclient.mybluemix.net/" target="_blank" rel="noopener no referrer">here</a>.
# !pip install -U ibm_watson_machine_learning
# +
from ibm_watson_machine_learning import APIClient
client = APIClient(wml_credentials)
# -
# ### Working with spaces
#
# First of all, you need to create a space that will be used for your work. If you do not have space already created, you can use [Deployment Spaces Dashboard](https://dataplatform.cloud.ibm.com/ml-runtime/spaces?context=cpdaas) to create one.
#
# - Click New Deployment Space
# - Create an empty space
# - Select Cloud Object Storage
# - Select Watson Machine Learning instance and press Create
# - Copy `space_id` and paste it below
#
# **Tip**: You can also use SDK to prepare the space for your work. More information can be found [here](https://github.com/IBM/watson-machine-learning-samples/blob/master/cloud/notebooks/python_sdk/instance-management/Space%20management.ipynb).
#
# **Action**: Assign space ID below
space_id = 'PASTE YOUR SPACE ID HERE'
# You can use `list` method to print all existing spaces.
client.spaces.list(limit=10)
# To be able to interact with all resources available in Watson Machine Learning, you need to set **space** which you will be using.
client.set.default_space(space_id)
# + [markdown] pycharm={"name": "#%% md\n"}
# <a id="download"></a>
# ## 2. Download externally created scikit model and data
# In this section, you will download externally created scikit models and data used for training it.
# + pycharm={"is_executing": false, "name": "#%%\n"}
import os
import wget
data_dir = 'BOSTON_HOUSE_PRICES_DATA'
if not os.path.isdir(data_dir):
os.mkdir(data_dir)
model_path = os.path.join(data_dir, 'boston_house_prices_model.tar.gz')
new_model_path = os.path.join(data_dir, 'new_boston_house_prices_model.tar.gz')
if not os.path.isfile(model_path):
wget.download("https://github.com/IBM/watson-machine-learning-samples/raw/master/cloud/models/scikit/boston_house_price/model/boston_house_prices_model.tar.gz", out=data_dir)
if not os.path.isfile(new_model_path):
wget.download("https://github.com/IBM/watson-machine-learning-samples/raw/master/cloud/models/scikit/boston_house_price/model/new_boston_house_prices_model.tar.gz", out=data_dir)
# +
from sklearn import datasets
import pandas as pd
boston_data = datasets.load_boston()
boston_df = pd.DataFrame(boston_data.data)
boston_df.columns = boston_data.feature_names
boston_df['PRICE'] = boston_data.target
# -
train_df = boston_df
test_df = boston_df.drop(['PRICE'], axis=1)
# <a id="persistence"></a>
# ## 3. Persist externally created scikit model
# In this section, you will learn how to store your model in Watson Machine Learning repository by using the Watson Machine Learning Client.
# ### 3.1: Publish model
# #### Publish model in Watson Machine Learning repository on Cloud.
# Define model name, autor name and email.
# + pycharm={"is_executing": false, "name": "#%%\n"}
sofware_spec_uid = client.software_specifications.get_id_by_name("default_py3.7")
# + pycharm={"is_executing": false, "name": "#%%\n"}
metadata = {
client.repository.ModelMetaNames.NAME: 'External scikit model',
client.repository.ModelMetaNames.TYPE: 'scikit-learn_0.23',
client.repository.ModelMetaNames.SOFTWARE_SPEC_UID: sofware_spec_uid
}
published_model = client.repository.store_model(
model=model_path,
meta_props=metadata,
training_data=train_df)
# -
# ### 3.2: Get model details
# + pycharm={"is_executing": false, "name": "#%%\n"}
import json
published_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(published_model_uid)
print(json.dumps(model_details, indent=2))
# -
# ### 3.3 Get all models
# + pycharm={"is_executing": false, "name": "#%%\n"}
models_details = client.repository.list_models(limit=10)
# -
# <a id="scoring"></a>
# ## 4. Deploy and score in a Cloud
# In this section you will learn how to create online scoring and to score a new data record by using the Watson Machine Learning Client.
# ### 4.1: Create model deployment
# #### Create online deployment for published model
# + pycharm={"is_executing": false, "name": "#%%\n"}
metadata = {
client.deployments.ConfigurationMetaNames.NAME: "Deployment of external scikit model",
client.deployments.ConfigurationMetaNames.ONLINE: {}
}
created_deployment = client.deployments.create(published_model_uid, meta_props=metadata)
# -
# **Note**: Here we use deployment url saved in published_model object. In next section, we show how to retrive deployment url from Watson Machine Learning instance.
deployment_uid = client.deployments.get_uid(created_deployment)
# Now you can print an online scoring endpoint.
# + pycharm={"is_executing": false, "name": "#%%\n"}
scoring_endpoint = client.deployments.get_scoring_href(created_deployment)
print(scoring_endpoint)
# -
# You can also list existing deployments.
client.deployments.list(limit=10)
# ### 4.2: Get deployment details
# + pycharm={"is_executing": false, "name": "#%%\n"}
print(json.dumps(client.deployments.get_details(deployment_uid), indent=2))
# -
# ### 4.3: Score
# You can use below method to do test scoring request against deployed model.
# **Action**: Prepare scoring payload with records to score.
score_0 = list(test_df.iloc[0])
score_1 = list(test_df.iloc[1])
# + pycharm={"is_executing": false, "name": "#%%\n"}
scoring_payload = {"input_data": [{"values": [score_0, score_1]}]}
# -
# Use ``client.deployments.score()`` method to run scoring.
# + pycharm={"is_executing": false, "name": "#%%\n"}
predictions = client.deployments.score(deployment_uid, scoring_payload)
# + pycharm={"is_executing": false, "name": "#%%\n"}
print(json.dumps(predictions, indent=2))
# + [markdown] pycharm={"name": "#%% md\n"}
# <a id="update_model"></a>
# ## 5. Persist new version of the model
# -
# In this section, you'll learn how to store new version of your model in Watson Machine Learning repository by using the Watson Machine Learning Client.
# ### 5.1: Publish new version of the model
# Save the current model version.
print(json.dumps(client.repository.create_model_revision(published_model_uid), indent=2))
# Define new model name and update model content.
# + pycharm={"is_executing": false, "name": "#%%\n"}
metadata = {
client.repository.ModelMetaNames.NAME: 'External scikit model - updated'
}
published_model = client.repository.update_model(
model_uid=published_model_uid,
update_model=new_model_path,
updated_meta_props=metadata
)
# -
# Save new model revision of the updated model.
new_model_revision = client.repository.create_model_revision(published_model_uid)
print(json.dumps(new_model_revision, indent=2))
# **Note:** Model revisions can be identified by model `id` and `rev` number.
#
# Get model `rev` number from creation details:
rev_id = new_model_revision['metadata'].get('rev')
# You can list existing revisions of the model.
client.repository.list_models_revisions(published_model_uid)
# ### 5.2: Get model details
# + pycharm={"name": "#%%\n"}
import json
published_model_uid = client.repository.get_model_uid(published_model)
model_details = client.repository.get_details(published_model_uid)
print(json.dumps(model_details, indent=2))
# -
# <a id="redeploy"></a>
# ## 6. Redeploy new version of the model
# In this section, you'll learn how to redeploy new version of the model by using the Watson Machine Learning Client.
# ### 6.1 Redeploy model
# +
metadata = {
client.deployments.ConfigurationMetaNames.ASSET: {
"id": published_model_uid,
"rev": rev_id
}
}
updated_deployment = client.deployments.update(deployment_uid=deployment_uid, changes=metadata)
# -
# Wait for the deployment update:
# +
import time
status = None
while status not in ['ready', 'failed']:
print('.', end=' ')
time.sleep(2)
deployment_details = client.deployments.get_details(deployment_uid)
status = deployment_details['entity']['status'].get('state')
print("\nDeployment update finished with status: ", status)
# -
# ### 6.2 Get updated deployment details
print(json.dumps(client.deployments.get_details(deployment_uid), indent=2))
# <a id="scale"></a>
# ## 7. Deployment scaling
# In this section, you'll learn how to scale your deployment by creating more copies of stored model with Watson Machine Learning Client.
# This feature is for providing High-Availability and to support higher throughput
# ### 7.1 Scale deployment
#
# In this example, 2 deployment copies will be made.
metadata = {
client.deployments.ConfigurationMetaNames.NAME: "Deployment of external scikit model - scaling",
client.deployments.ConfigurationMetaNames.HARDWARE_SPEC: {
"name": "S",
"num_nodes": 2
}
}
scaled_deployment = client.deployments.update(deployment_uid, metadata)
# ### 7.2 Get scaled deployment details
print(json.dumps(client.deployments.get_details(deployment_uid), indent=2))
# ### 7.3 Score updated deployment
# You can use below method to do test scoring request against deployed model.
#
# **Action**: Prepare scoring payload with records to score.
score_0 = list(test_df.iloc[0])
score_1 = list(test_df.iloc[1])
scoring_payload = {"input_data": [{"values": [score_0, score_1]}]}
# Use client.deployments.score() method to run scoring.
predictions = client.deployments.score(deployment_uid, scoring_payload)
print(json.dumps(predictions, indent=2))
# <a id="cleanup"></a>
# ## 8. Clean up
# If you want to clean up all created assets:
# - experiments
# - trainings
# - pipelines
# - model definitions
# - models
# - functions
# - deployments
#
# please follow up this sample [notebook](https://github.com/IBM/watson-machine-learning-samples/blob/master/cloud/notebooks/python_sdk/instance-management/Machine%20Learning%20artifacts%20management.ipynb).
# <a id="summary"></a>
# ## 9. Summary and next steps
# You successfully completed this notebook! You learned how to use scikit-learn machine learning as well as Watson Machine Learning for model creation and deployment. Check out our _[Online Documentation](https://dataplatform.cloud.ibm.com/docs/content/wsj/getting-started/welcome-main.html?context=analytics?pos=2)_ for more samples, tutorials, documentation, how-tos, and blog posts.
# ### Authors
#
# **<NAME>**, Software Engineer
# **<NAME>**, Intern
# Copyright © 2020, 2021 IBM. This notebook and its source code are released under the terms of the MIT License.
|
cloud/notebooks/python_sdk/lifecycle-management/Use scikit-learn and AI lifecycle capabilities to predict Boston house prices.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Video Games NLP Exploratory Analysis
# Data from
# > Justifying recommendations using distantly-labeled reviews and fined-grained aspects
# <NAME>, <NAME>, <NAME>
# Empirical Methods in Natural Language Processing (EMNLP), 2019
# [https://nijianmo.github.io/amazon/index.html#files](https://nijianmo.github.io/amazon/index.html#files)
# ## Purpose
#
# The purpose of this notebook is to read in and explore text reviews for video games from Amazon.com. These reviews were collected from 1996-2014. The dataset is labeled with star ratings and contains text reviews for purchased video games.
#
# This particular analysis will explore the data, look for interesting features about the text, and describe basic properties of it.
# import packages
import numpy as np
import pandas as pd
# read in data
vg = pd.read_json('../Amazon_Data/Video_Games_5.json.gz', lines=True, compression='gzip')
vg.head()
# choose only select columns and clean up the datatypes and missing values
vg = vg.loc[:,['overall', 'reviewText']]
vg = vg.dropna(how='any')
vg.loc[:,'overall'] = vg.overall.astype('int16')
# inspect the df info
vg.info()
# check out the distribution of ratings
vg.overall.value_counts()
# The classes are heavily imbalanced, with far more positive ratings than negative. This analysis is not seeking to predict sentiment, therefore, the data will not be upsampled/downsampled right now.
#
# The entire text data will be analyzed for interesting properties.
# ### Text Exploration
# import stop words from spacy
from spacy.lang.en.stop_words import STOP_WORDS
# generate and edit stop words
stops = list(STOP_WORDS)
print(stops)
# Preliminary analysis revealed some character conversion isses. Adding additional stop words.
# add additional stopwords
issue_words = ['\ufeff1', '\x16', '...', '\x10once', 's', '1', '2', '3', '4', '5']
stops.extend(issue_words)
print(stops)
# import additional nlp tools
import string
import spacy
from spacy.lang.en import English
# generate puntuations string
punctuations = string.punctuation
punctuations
# define custom tokenizers
parser = English()
def my_tokenizer(sentence):
mytokens = parser(sentence)
mytokens = [word.lemma_.lower().strip() if word.lemma_ != "-PRON-"
else word.lower_ for word in mytokens]
mytokens = [word for word in mytokens if word not in stops and word not in punctuations]
return mytokens
# test the tokenizer with basic text
s = 'This video game was crazy! I though it was terrific!'
print(my_tokenizer(s))
# create word_list function
def get_word_list(text, word_list):
for sentence in text:
for word in my_tokenizer(sentence):
word_list.append(word)
# generate word_list
word_list = []
get_word_list(vg.reviewText, word_list)
# inspect the first 20 words
word_list[:20]
# check for anomalous words that start with a randomly chosen special character
for word in word_list:
if word.startswith('\\'):
print(word)
# Custom tokenizer only removes tokens that contain individual puntuation marks, not words or tokens that consist of multiple punctuation marks or punctuation marks that have other characters in the token.
# +
# define remove chars function
import re
def rmv_spec_chars(sentence):
# completely remove most punctuation
sentence = re.sub("[\\\[\]@_!#$%^&*()<>?/\|}{~:']", '', sentence)
# replace hypens with spaces to split those words
sentence = re.sub('-', ' ', sentence)
return sentence
# -
# test the function
rmv_spec_chars("This 'is' going * to be! a test \/(--)\/ train-test-split")
# test the function with the tokenizer
my_tokenizer(rmv_spec_chars("This 'is' going * to be! a test \/(--)\/ train-test-split"))
# The above tokens are what we want, so going to process the text again.
# edit tokenizer to deal with tokens with special characters
def my_tokenizer(sentence):
sentence = rmv_spec_chars(sentence)
mytokens = parser(sentence)
mytokens = [word.lemma_.lower().strip() if word.lemma_ != "-PRON-"
else word.lower_ for word in mytokens]
mytokens = [word for word in mytokens if word not in stops and word not in punctuations]
return mytokens
# get a list of all words each time they occur in the reviews
# using new tokenizer in the get_word_list function
word_list = []
get_word_list(vg.reviewText, word_list)
# check for anomalous words that start with a randomly chosen special character
for word in word_list:
if word.startswith('\\'):
print(word)
# No output from function above means we at least took care of the issue from before.
# display the total number of words included in the vocabulary
# stop words and special characters (emojis) have been removed
# this includes the total number of words, not the unique words
print('The number of words in the reviews is {}'.format(len(word_list)))
# #### Generating summary stats for the words and appearances
# +
from collections import Counter, defaultdict
# get a count of every word
token_counts = Counter(word_list)
# -
# find the number of unique words in the reviews
# stop words excluded
print('The total number of unique words is {}'.format(len(token_counts)))
# inspect the 20 most common words
token_counts.most_common(20)
# #### Generate some basic visualizations for the number of word appearances
# +
import matplotlib.pyplot as plt
# create histogram of word appearances
# most words occur infrequently, so threshold for num influences this plot greatly
values = []
for tup in token_counts.most_common():
word, num = tup
if num > 10000:
values.append(num)
_ = plt.hist(values, bins=100)
_ = plt.title('Histogram for Number of Appearances')
_ = plt.xlabel('Number of Times a Word Appears')
_ = plt.ylabel('Number of Words')
# -
# This histogram shows that most words appear infrequently, but there are a few words that appear way more than the others.
# +
import seaborn as sns
# plot the 20 most common words
words = []
values = []
for tup in token_counts.most_common(20):
word, num = tup
words.append(word)
values.append(num)
_ = sns.barplot(words, values, palette='muted')
_ = plt.xticks(rotation=90)
_ = plt.title('20 Most Common Words')
_ = plt.xlabel('Word')
_ = plt.ylabel('Occurrences')
# -
# Next, we will analyze the number of reviews in which each word appears.
# +
# get a count of the number of reviews where a word appears
def get_num_docs(text_series):
"""Take a text series and and return a default dict."""
# initialize default dict
num_docs = defaultdict(int)
# iterate through and populate the default dict
for text in text_series:
ls = []
for word in my_tokenizer(text):
if word not in ls:
ls.append(word)
for x in ls:
num_docs[x] += 1
# return the default dict
return num_docs
# -
# get the number of docs in which each word appears
num_docs = get_num_docs(vg.reviewText)
# store a list of the number of appearances for each word
apps = []
for key, val in num_docs.items():
apps.append(val)
# plot the cdf for the number of reviews where each word appears
_ = plt.hist(apps, cumulative=True, histtype='step', density=True, bins=np.arange(100))
_ = plt.xlim(0,25)
_ = plt.xlabel('Number of Docs')
_ = plt.ylabel('Cumulative Density')
_ = plt.title('Density of Words Appearing \nin at Least X Reviews')
# So about 60% of the words in the vocabulary appear in only one review! When creating the sentiment analysis model, it could be useful to set a minimum appearance of 2.
# The next visualization for the words will be a wordcloud using the WordCloud class.
# import wordcloud
from wordcloud import WordCloud
# initialize wordcloud object
wc = WordCloud(background_color='white', stopwords=stops, max_words=200,
max_font_size=40, scale=3, random_state=42)
# generate the wordcloud
wc.generate(' '.join(word_list))
# +
# show the wordcloud
fig = plt.figure(1, figsize=(12, 12))
plt.axis('off')
plt.imshow(wc)
plt.show()
# -
# ## Summary of Exploratory Analysis
# The text data had some very interesting features. It was surprising to find many special characters and emojis in the data. For this analysis, these were removed, and the analysis was performed on the clean text. I think this is a very important result, given that this was not much of an issue when I was originally working with book reviews.
#
# This shows the importance of understanding the particular dataset for the problem. While the book reviews came from the same source (Amazon for the years 1996-2014), the content was different enough to require different preprocessing techniques.
#
# Plotting word counts showed that over half of the words only appeared in one review. The most common words were plotted, and unsurprisingly, the most common word was game. If using standard models to predict sentiment of reviews, it would be recommended to include only words that appear in two or more reviews.
#
# For the Flair sentiment analysis model, see the 2_Video_Games_Flair_Model.ipynb file.
|
1_Video_Games_Exploratory_NLP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.mx)
# # Palabras reservadas, nombres y el espacio de nombres.
# ## Palabras reservadas de Python.
#
# Las palabras reservadas (keywords) corresponden a los nombres de las declaraciones que el intérprete de Python incluye por defecto. No se deben utilizar dichas palabras para asignar nombres a otros objetos.
#
# El listado de palabras reservadas puede ser consultado ingresando ```help('keywords')``` desde la interfaz interactiva.
# **Ejemplo:**
# * La siguiente celda desplegará las palabras reservadas del intérpete de Python que usa la notebook.
help("keywords")
# ### Palabras reservadas en Python 2.7.
#
# Las palabras reservadas en Python 2.7 son la siguientes.
# ```
# and elif if print
# as else import raise
# assert except in return
# break exec is try
# class finally lambda while
# continue for not with
# def from or yield
# del global pass
# ```
# ### El módulo ```__builtins__```.
#
# Python es un lenguaje modular. Es decir, que tiene la capacidad de cargar piezas de código cuyos objetos se añaden al espacio de nombres del intéreprete.
#
# El módulo ```__builtins__``` es cargado automáticamente por el intérprete cada vez que este es inicializado.
#
# No se recomienda usar los nombres definidos en este módulo.
# **Ejemplo:**
# * La siguiente celda mostrará los nombres de los elementos del módulo ```__builtins__```.
dir(__builtins__)
# ## El espacio de nombres (namespace).
#
# El espacio de nombres (namespace) contiene una relación de los objetos existentes en la memoria del sistema y los nombres a los que están ligados.
#
# * Python es un lenguaje de muy alto nivel en el que todos sus elementos son objetos, incluyendo los tipos de datos básicos de tal forma que los términos "valor" y "objeto" son sinónimos.
# * Si un objeto no está ligado al menos a un nombre, dicho objeto es desechado por el intérprete.
# * Un objeto puede tener más de un nombre.
# ## El operador de asignación ```=```.
#
# Para relacionar a un nombre con un objeto, se utiliza el el operador de asignación ```=``` con la siguiente sintaxis:
#
# ```
# <nombre> = <objeto>
# ```
#
# **Ejemplo:**
#
# * Las siguientes celdas asignarán un nombre a un objeto de Python. Por el momento no se discutirá el qué tipo de objeto del que se trata.
saludo = 'Hola'
matriz = [["autobús", "diesel", True], ["automóvil", "gasolina", True]]
numero = 23.45
# ## Despliegue de valores ligados a un nombre en el entorno interactivo.
#
# En el entorno interactivo es posible desplegar el contenido de un objeto tan sólo con ingresar su nombre.
#
# Si se hace referencia a un nombre que no se encuentra en el espacio de nombres, se desencadenará un error de tipo ```NameError```.
# * Cada una de las celdas siguientes regresará el objeto que corresponde a cada nombre asignado.
saludo
matriz
numero
# * El nombre ```indefinido``` no se encuentra en el espacio de nombres y por lo tanto se desencadenará un error de tipo ```NameError```.
indefinido
# ### Asignación de múltiples nombres a igual número de objetos.
#
# Es posible asignar a varios nombres un número igual de objetos usando un sólo operador de asignación mediante la siguiente sintaxis:
#
# ```
# <nombre 1>, <nombre 2>, <nombre 3>, ..., <nombre n> = <objeto 1>, <objeto 2>, <objeto 3>, ...,<objeto n>```
# **Ejemplo:**
#
# * La siguiente celda contiene una línea que asignará 4 nombres a igual número de objetos.
entero, flotante, complejo, booleano = 12, 4.5, (12.3 + 23j), True
entero
complejo
flotante
booleano
# ### Sintaxis para la elaboración de nombres en Python 3.
#
# * Python 3 acepta el uso de [*Unicode*](https://es.wikipedia.org/wiki/Unicode), por lo que es posible utilizar cualquier caracter alfabético, incluso aquellos que no pertencen al alfabeto occidental para la elaboración de nombres.
# * Los nombres pueden empezar con un guión bajo ```_``` o un caracter alfabético.
# * Después del primer caracter, se pueden utilizar caracteres alfabéticos, números y/o guiones bajos.
# * No se permiten caracteres distintos a los alfabéticos o que pudieran confundirse con operadores como ```|```, ```~```, ```#```, ```-```, etc.
# * Se pueden utilizar mayúsculas, pero cabe señalar que Python es sensible a mayúsculas.
#
# **Nota:** Python 2 sólo soporta nombres con caracteres alfabéticos del código [*ASCII*](https://es.wikipedia.org/wiki/ASCII).
# **Ejemplos:**
#
# * Las siguientes celdas contienen asignaciones con nombres que se apegan a la sintaxis correcta, incluso usando el alfabeto hebreo.
_saludo = 'Hola'
número = 23
Numero = 45.32
# * A continuación se ejemplifica el usos de caracteres de alfabetos distintos a los occidentales utilizando el alfabeto hebreo, el cual se escribe de derecha a izquierda.
יהוה = "Dios"
# * Las siguientes celdas desplegarán el valor del objeto ligado a los nombres definidos previamente en esta notebook.
saludo
_saludo
número
Numero
יהוה
# * En la siguiente celda se realizará una asignación utilizando un nombre incorrecto, desencandeado un error de tipo ```SyntaxError```.
1error = "Ups"
# ## La función ```id()```.
#
# Cada objeto cuenta con un *número identificador*, el cual corresponde a la posición en la que se encuentra almacenado en la memoria.
#
# La función ```id()``` permite acceder al número identificador de cada objeto usando su nombre como argumento.
#
# La sintaxis es la siguiente:
#
# ```
# id(<nombre>)
# ```
# **Ejemplo:**
#
# * A continuación se le asignará el nombre ```saludo``` al objeto ```"Hola"``` y se obtendrá su número identificador mediante la función ```id()```.
saludo = "Hola"
id(saludo)
# ## Asignación de múltiples nombres al mismo objeto.
#
# Python es un lenguaje que gestiona de forma automática el uso de la memoria y trata de optimizar su uso, por lo que en caso de que se defina un objeto idéntico a otro ya existente, no creará uno nuevo, sino que ligará el nuevo nombre al objeto existente.
#
# Un objeto puede estar relacionado con más de un nombre.
# **Ejemplo:**
#
# * En la siguientes celdas se le asignarán dos nombres al objeto con valor igual a ```45```.
numero = 45
otro_numero = 45
# * Aún cuando se definen de forma separada, el resultado de la función ```id()``` es el mismo para cada nombre.
id(numero)
id(otro_numero)
# * Cuando a un nombre se le liga con otro objeto, el número identificador ligado a dicho nombre corresponde al nuevo objeto.
# **Ejemplo:**
# * La siguiente celda le asignará al objeto ```25``` el nombre de ```otro_numero```, el cual fue definido en celdas previas.
otro_numero = 25
# * Por lo tanto, ahora el número identificador de ```numero```y ```otro_numero``` son distintos.
id(otro_numero)
id(numero)
# ### Asignación de múltiples nombres a un objeto.
#
# Para asignarle más de un nombre al mismo objeto, sólo es necesario referenciar un nombre existente al nuevo nombre.
#
# ```
# <nombre 1> = <nombre 2>
# ```
# **Ejemplo:**
#
# * Las siguientes celdas le asignarán el nombre ```lista_1``` y ```lista_2``` al mismo objeto.
lista_1 = [1, 2, 3, 4, 5]
lista_2 = lista_1
# * Se puede observar que el número identificador corresponde al mismo objeto, aún cuando tiene otro nombre.
id(lista_1)
id(lista_2)
# ## Eliminación de nombres mediante la declaración ```del```.
#
# La declaración ```del``` funciona de la siguiente manera:
#
# * Desliga al nombre de un objeto en el espacio de nombres.
# * Elimina al nombre del espacio de nombres.
# * En caso de que el objeto no esté ligado a otro nombre en el espacio de nombres, el intérprete de Python podría desecharlo de forma automática.
#
# El modo en el que un objeto puede ser destruido varía dependiendo del tipo de objeto.
# **Ejemplo:**
# * Se creará el objeto ```"Juan"``` al cual se le asignarán los nombres ```nombre``` y ```otro_nombre```.
nombre = "Juan"
id(nombre)
otro_nombre = "Juan"
id(otro_nombre)
# * Se eliminará ```nombre``` del espacio de nombres.
del nombre
# * Debido a que ```nombre``` fue eliminado del espacio de nombres, la siguiente celda regresará un erro de tipo ```NameError```.
nombre
# * El objeto ```"Juan"``` sigue existiendo en la memoria, debido a que aún existe un nombre al que está ligado.
otro_nombre
# ### Despliegue del contendio del espacio de nombres.
#
# La función ```dir()``` despliega un listado del contenido del espacio de nombres de un objeto que es ingresado como argumento.
#
# ```
# dir(<objeto>)
# ```
#
# Cuando se usa la función ```dir()``` sin argumentos, esta regresa el listado de nombres del espacio de nombres principal.
# **Ejemplos:**
# * La siguiente celda despelgará el listado de nombres de esta notebook.
#
# *Se puede apreciar que el listado incluye una serie de nombre especiales y todos los nombres que se han definido a lo largo de este capítulo.
dir()
# * La siguiente celda elimina el nombre ```saludo```, el cual fue definido previamente.
del saludo
# * El resultado es que el nombre ```saludo``` ya no será enlistado al ejecutar la función ```dir()```.
dir()
# * Lo mismo sucede al eliminar el nombre ```יהוה```.
del יהוה
dir()
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2019.</p>
|
03_palabras_reservadas_y_nombres.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regularization for linear regressor models
# +
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression, Ridge, Lasso, LogisticRegression
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.datasets import load_boston, load_iris, load_wine, load_digits, \
load_breast_cancer, load_diabetes
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, precision_score, recall_score
import matplotlib.pyplot as plt
# %config InlineBackend.figure_format = 'retina'
def score(X, y, model, n=10, random_state=None):
train_scores = []
test_scores = []
for i in range(n):
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.2, random_state=random_state)
model.fit(X_train, y_train)
train_score = model.score(X_train, y_train)
train_scores.append(train_score)
test_score = model.score(X_test, y_test)
test_scores.append(test_score)
return np.mean(train_scores), np.mean(test_scores)
# -
# ## Synthetic data set
#
# Let's start with a simple linear regression model of $y$ on $x$ for synthetic data set, just to get a baseline. Since this is a toy example, we won't bother with separating a test set out. Even the training error is going to fall apart for this example.
n = 10
df = pd.DataFrame()
df['x'] = np.linspace(0,10,num=n)
df['y'] = df['x'] + np.random.normal(0,1,size=n)
df
# ### Base model
X, y = df.drop('y',axis=1), df['y']
lm = LinearRegression()
lm.fit(X, y)
beta0 = lm.intercept_
beta1 = lm.coef_[0]
score(X, y, lm) # bounces around a lot depending on data set and test set
# The model produces a nice regression line through the data points as we would expect.
plt.scatter(df['x'], df['y'], s=95)
plt.plot(df['x'], df['x']*lm.coef_[0] + lm.intercept_, c='orange')
plt.xlabel("x", fontsize=12)
plt.ylabel("y", fontsize=12)
plt.title(f"$\\beta_1$ = {beta1:.3f}, $\\beta_0$ = {lm.intercept_:.2f}", fontsize=14)
plt.savefig("/Users/parrt/Desktop/ols.svg")
# ### Adding an outlier
#
# Let's add an outlier, making the last y value 100
y.iloc[-1] = 100
y
X, y = df.drop('y',axis=1), df['y']
lm = LinearRegression()
lm.fit(X, y)
beta0 = lm.intercept_
beta1 = lm.coef_[0]
lm.score(X, y) # bounces around depending on test set
plt.scatter(X, y, s=95)
plt.plot(X, X*beta1 + lm.intercept_, c='orange')
plt.xlabel("x", fontsize=12)
plt.ylabel("y", fontsize=12)
plt.title(f"$\\beta_1$ = {beta1:.3f}, $\\beta_0$ = {lm.intercept_:.2f}", fontsize=14)
plt.savefig("/Users/parrt/Desktop/outlier.svg")
# The $\beta_1$ coefficient jumps from .99 to 5.35 and the Y intercept goes from 0.54 to -12.39. These are radically different coefficients that arise simply because one value is an outlier.
# ### L1 (Lasso) Regularization
#
# If we constrain the $\beta_1$ coefficients strongly, then we can prevent the outlier from bending the regression line upwards.
# +
X, y = df.drop('y',axis=1), df['y']
y.iloc[-1] = 100
lm = Lasso(alpha=45)
lm.fit(X, y)
beta0 = lm.intercept_
beta1 = lm.coef_[0]
print("R^2 training, test scores", score(X, y, lm))
plt.scatter(X, y, s=95)
plt.plot(X, X*beta1 + lm.intercept_, c='orange')
plt.xlabel("x", fontsize=12)
plt.ylabel("y", fontsize=12)
plt.title(f"$\\beta_1$ = {beta1:.3f}, $\\beta_0$ = {lm.intercept_:.2f}", fontsize=14)
plt.savefig("/Users/parrt/Desktop/lasso.svg")
plt.show()
# -
# The 0.11 score is not that great but we have sacrificed a bit of accuracy for a much more reasonable coefficient. The $\beta_1$ is now 0.93, very close to the 0.99 we get without the outlier.
#
# What is wrong with the $y$-intercept? There is a trade-off here between bias and generality. We have sacrificed accuracy a bit (line is above most of the points) but gained and generality because the trendline is right (we ignore the outlier).
# ## Ames housing data set
# ### Prep data set
#
# Log into Kaggle and get [House Prices: Advanced Regression Techniques](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data). A useful discussion on this data set is [here](https://nycdatascience.com/blog/student-works/machine-learning-project/).
df_ames = pd.read_csv("../../data/ames.csv")
df_ames.head()
# +
cols_with_missing = df_ames.columns[df_ames.isnull().any()]
cols = set(df_ames.columns) - set(cols_with_missing)
X = df_ames[cols].drop('SalePrice', axis=1)
y = df_ames['SalePrice']
X.head()
# -
X = pd.get_dummies(X)
X.shape
# ### Getting a baseline model
lm = LinearRegression()
score(X, y, lm, n=30)
coef = np.clip(lm.coef_,-1e10,1e8) #Clip so we can display
np.max(lm.coef_) #Wow, look at the size of this coefficient
np.std(lm.coef_)
import matplotlib as mpl
plt.figure(figsize=(6,3.5))
plt.bar(range(len(coef)),coef)
plt.xlabel("Regression coefficient $\\beta_i$ for $i>0$", fontsize=12)
plt.ylabel("Regression coefficient value\nclipped to 1e8 magnitude", fontsize=12)
plt.gca().yaxis.set_major_formatter(mpl.ticker.StrMethodFormatter('{x:.1}'))
#plt.title(lm)
plt.show()
print("R^2 training, test scores", score(X, y, lm, n=30))
# Wow, look at the size of those coefficients. They are huge and all over the place, which is what we'd expect given such a terrible test R^2 score. The coefficients are so bad that prediction suffers.
# ## Normalization
#
# We did not normalize our variable for the synthetic data set because we only had one variable. Normally, we have to convert all X variables to standard scores so they are all in the same range and zero centered. If the variables are all in different ranges, regularization will squash some coefficients more than the others because all regularization does is constrain coefficients.
from pandas.api.types import is_numeric_dtype
def normalize(X): # well, I'm creating standard variables here (u-x)/sigma
for colname in X.columns:
if is_numeric_dtype(X[colname]):
u = np.mean(X[colname])
s = np.std(X[colname])
X[colname] = (X[colname] - u) / s
X = df_ames[cols].drop('SalePrice', axis=1)
y = df_ames['SalePrice']
normalize(X) # do this before getting dummy variables
X.head(2)
X = pd.get_dummies(X) # make sure to normalize before you get the dummy variables
lm = LinearRegression()
score(X, y, lm, n=30)
import matplotlib as mpl
plt.bar(range(len(coef)),coef)
plt.xlabel("Regression coefficient $\\beta_i$ for $i>0$", fontsize=12)
plt.ylabel("Regression coefficient value\nclipped to 1e8 magnitude", fontsize=12)
plt.gca().yaxis.set_major_formatter(mpl.ticker.StrMethodFormatter('{x:.1}'))
plt.tight_layout()
plt.savefig("/Users/parrt/Desktop/ames.png", dpi=200)
plt.show()
# ### L1 regularization with normalization
lm = Lasso(alpha=10, tol=0.1)
score(X, y, lm, n=10, random_state=42)
print(f"std(coef)={np.std(lm.coef_):.1f}, num 0 coeff={len(np.where(lm.coef_<0.1)[0])}")
plt.bar(range(len(lm.coef_)),lm.coef_)
plt.xlabel("Regression coefficient $\\beta_i$ for $i>0$", fontsize=12)
plt.ylabel("Regression coefficient value", fontsize=12)
plt.ylim(-30_000, 30_000)
plt.tight_layout()
plt.savefig("/Users/parrt/Desktop/ames-L1.png", dpi=200)
plt.show()
# ### L2 regularization with normalization
lm = Ridge(alpha=10)
score(X, y, lm, n=50)
print(f"std(coef)={np.std(lm.coef_):.1f}, num 0 coeff={len(np.where(lm.coef_<0.1)[0])}")
plt.bar(range(len(lm.coef_)),lm.coef_)
plt.xlabel("Regression coefficient $\\beta_i$ for $i>0$", fontsize=12)
plt.ylabel("Regression coefficient value", fontsize=12)
plt.ylim(-30_000, 30_000)
plt.show()
|
notebooks/linear-models/regressor-regularization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# +
# 백(bag)
# -
from collections import Counter
자루 = ['빨간색 공', '초록색 공', '파란색 공', '파란색 공', '파란색 공']
print(Counter(자루))
자루.append('초록색 공')
print(Counter(자루))
# +
# 집합(set)
# -
집합A = { 1, 2, 3, 4, 5 }
집합B = { 1, 3, 5, 7, 11 }
print(f'합집합: 집합A + 집합B = {집합A | 집합B}')
print(f'교집합: 집합A & 집합B = {집합A & 집합B}')
print(f'차집합: 집합A - 집합B = {집합A - 집합B}')
print(f'차집합: 집합B - 집합A = {집합B - 집합A}')
# +
# 리스트(list)
# -
리스트 = [ 1, 2, 3 ]
print(f'리스트: {리스트}')
리스트.append(3)
print(f'3 추가 후: {리스트}')
리스트.append(2)
print(f'2 추가 후: {리스트}')
리스트.append(1)
print(f'1 추가 후: {리스트}')
# +
# %load_ext autoreload
# %autoreload 2
from cotylab.data_structure.linked_list import LinkedList
from cotylab.data_structure.linked_list import DoubledLinkedList
from cotylab.data_structure.linked_list import CircularLinkedList
단순연결리스트 = LinkedList([1, 2, 3, 3, 2, 1])
이중연결리스트 = DoubledLinkedList([1, 2, 3, 3, 2, 1])
원형연결리스트 = CircularLinkedList([1, 2, 3, 3, 2, 1])
print(단순연결리스트)
print(이중연결리스트)
print(원형연결리스트)
# +
단순연결리스트.go_to_tail()
원형연결리스트.go_to_tail()
print(f'단순연결리스트 꼬리에서 다음요소: {단순연결리스트.next()}')
print(f'원형연결리스트 꼬리에서 다음요소: {원형연결리스트.next()}')
이중연결리스트.go_to_tail()
print(이중연결리스트.current)
print(이중연결리스트.prev())
print(이중연결리스트.prev())
# -
|
notebooks/coding_basic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda pytorch_p37
# language: python
# name: pytorch_p37
# ---
# +
import sys
if 'google.colab' in sys.modules:
import os
os.system('apt-get install -y xvfb')
os.system('wget https://raw.githubusercontent.com/yandexdataschool/Practical_RL/master/xvfb -O ../xvfb')
os.system('apt-get install -y python-opengl ffmpeg')
os.system('pip install pyglet==1.2.4')
os.system('python -m pip install -U pygame --user')
print('setup complete')
# XVFB will be launched if you run on a server
import os
if type(os.environ.get("DISPLAY")) is not str or len(os.environ.get("DISPLAY")) == 0:
# !bash ../xvfb start
os.environ['DISPLAY'] = ':1'
# -
# # Implementing Advantage-Actor Critic (A2C)
# In this notebook you will implement Advantage Actor Critic algorithm that trains on a batch of Atari 2600 environments running in parallel.
#
# Firstly, we will use environment wrappers implemented in file `atari_wrappers.py`. These wrappers preprocess observations (resize, grayscal, take max between frames, skip frames and stack them together) and rewards. Some of the wrappers help to reset the environment and pass `done` flag equal to `True` when agent dies.
# File `env_batch.py` includes implementation of `ParallelEnvBatch` class that allows to run multiple environments in parallel. To create an environment we can use `nature_dqn_env` function. Note that if you are using
# PyTorch and not using `tensorboardX` you will need to implement a wrapper that will log **raw** total rewards that the *unwrapped* environment returns and redefine the implemention of `nature_dqn_env` function here.
#
#
# +
import numpy as np
from atari_wrappers import nature_dqn_env, NumpySummaries
nenvs = 8
env = nature_dqn_env("SpaceInvadersNoFrameskip-v4", nenvs=nenvs,
summaries='Numpy')
obs = env.reset()
n_actions = env.action_space.n
print('num of actions: {}'.format(n_actions))
assert obs.shape == (8, 84, 84, 4)
assert obs.dtype == np.uint8
# +
# for evaluating the agent
single_env = nature_dqn_env("SpaceInvadersNoFrameskip-v4", nenvs=1,
summaries='Numpy')
print(single_env.reset())
# -
import gym
one_env = gym.make('SpaceInvadersNoFrameskip-v4')
one_obs = one_env.reset()
print(one_obs.shape)
# Next, we will need to implement a model that predicts logits and values. It is suggested that you use the same model as in [Nature DQN paper](https://web.stanford.edu/class/psych209/Readings/MnihEtAlHassibis15NatureControlDeepRL.pdf) with a modification that instead of having a single output layer, it will have two output layers taking as input the output of the last hidden layer. **Note** that this model is different from the model you used in homework where you implemented DQN. You can use your favorite deep learning framework here. We suggest that you use orthogonal initialization with parameter $\sqrt{2}$ for kernels and initialize biases with zeros.
# +
# import tensorflow as torch
# import torch as tf
import torch.nn as nn
import torch.nn.functional as F
def conv2d_size_out(size, kernel_size, stride):
"""
common use case:
cur_layer_img_w = conv2d_size_out(cur_layer_img_w, kernel_size, stride)
cur_layer_img_h = conv2d_size_out(cur_layer_img_h, kernel_size, stride)
to understand the shape for dense layer's input
"""
return (size - (kernel_size - 1) - 1) // stride + 1
class DQN(nn.Module):
def __init__(self, obs_dim, n_actions):
"""
obs_dim: image dimension of the observation
The input image must have 2d dim
obs_dim x obs_dim
"""
super().__init__()
kernel_size = 3
stride = 2
self.conv1 = nn.Conv2d(4, 16, kernel_size, stride)
out_size = conv2d_size_out(obs_dim, kernel_size, stride)
self.conv2 = nn.Conv2d(16, 32, kernel_size, stride)
out_size = conv2d_size_out(out_size, kernel_size, stride)
self.conv3 = nn.Conv2d(32, 64, kernel_size, stride)
out_size = conv2d_size_out(out_size, kernel_size, stride)
# size of the output tensor after convolution batch_size x 64 x out_size x out_size
self.linear = nn.Linear(64*out_size*out_size, 256)
# value head
self.value = nn.Linear(256, 1)
# policy head
self.policy = nn.Linear(256, n_actions)
def forward(self, state_t):
"""
takes agent's observation (tensor), returns qvalues (tensor)
:param state_t: a batch of 4-frame buffers, shape = [batch_size, 4, h, w]
"""
# Use your network to compute qvalues for given state
# qvalues = <YOUR CODE>
#print('== type of input tensor ==', type(state_t))
#print(state_t.shape)
t = self.conv1(state_t)
t = F.relu(t)
t = self.conv2(t)
t = F.relu(t)
t = self.conv3(t)
t = F.relu(t)
t = t.view(state_t.shape[0], -1)
t = self.linear(t)
t = F.relu(t)
values = self.value(t)
# log proba of actions
logits = F.log_softmax(self.policy(t), dim=1)
return values, logits
# -
# You will also need to define and use a policy that wraps the model. While the model computes logits for all actions, the policy will sample actions and also compute their log probabilities. `policy.act` should return a dictionary of all the arrays that are needed to interact with an environment and train the model.
# Note that actions must be an `np.ndarray` while the other
# tensors need to have the type determined by your deep learning framework.
# +
import torch
from torch.distributions import Categorical
device = 'cuda' if torch.cuda.is_available() else 'cpu'
class Policy:
def __init__(self, model):
self.model = model
self.model.to(device)
def parse_state(self, inputs):
#inputs = torch.Tensor(inputs, device=device)
#inputs = inputs.permute(0, 3, 1, 2).contiguous()
# to float
inputs = inputs.astype(np.float32)
# depth major
inputs = inputs.transpose(0, 3, 1, 2)
# normalize the inputs
mean = np.mean(inputs, axis=(2, 3), keepdims=True)
std = np.std(inputs, axis=(2, 3), keepdims=True)
inputs = (inputs- mean) / std
# to torch tensor
inputs = torch.Tensor(inputs, device=device)
return inputs
def act(self, inputs):
# input dim (8, 84, 84, 4)
# 8: number of parallel env
# 4: last 4 frames to make POMDP a MDP
# 84: frame dim
# Should return a dict containing keys ['actions', 'logits', 'log_probs', 'values'].
# inputs are batched input from different environment
# each action in actions repr the action taken in one env
# Same thing with log probability
# convert input to depth major
inputs = self.parse_state(inputs)
# value estimates of states and
# log proba of actions at the states
values, logits = self.model(inputs)
# sample from log probabilities
m = Categorical(logits=logits)
# sample an action for each env
actions = m.sample()
return {
'actions': actions,
'log_probs': m.log_prob(actions),
'estimated_values': values
}
def choose_action(self, inputs):
"""Choose action greedily in a testing env"""
inputs = self.parse_state(inputs)
with torch.no_grad():
_, logits = self.model(inputs)
actions = torch.argmax(logits, dim=-1)
return actions
def get_value(self, inputs):
"""estimate state value without keeping the computation graph"""
inputs = self.parse_state(inputs)
with torch.no_grad():
v, _ = self.model(inputs)
return v
# -
# Next will pass the environment and policy to a runner that collects partial trajectories from the environment.
# The class that does is is already implemented for you.
# This runner interacts with the environment for a given number of steps and returns a dictionary containing
# keys
#
# * 'observations'
# * 'rewards'
# * 'resets'
# * 'actions'
# * all other keys that you defined in `Policy`
#
# under each of these keys there is a python `list` of interactions with the environment of specified length $T$ — the size of partial trajectory.
# To train the part of the model that predicts state values you will need to compute the value targets.
# Any callable could be passed to `EnvRunner` to be applied to each partial trajectory after it is collected.
# Thus, we can implement and use `ComputeValueTargets` callable.
# The formula for the value targets is simple:
#
# $$
# \hat v(s_t) = \left( \sum_{t'=0}^{T - 1} \gamma^{t'}r_{t+t'} \right) + \gamma^T \hat{v}(s_{t+T}),
# $$
#
# In implementation, however, do not forget to use
# `trajectory['resets']` flags to check if you need to add the value targets at the next step when
# computing value targets for the current step. You can access `trajectory['state']['latest_observation']`
# to get last observations in partial trajectory — $s_{t+T}$.
class ComputeTargetValues:
"""Explicitly compute target value along the trajectory
in the forward view
"""
def __init__(self, policy, gamma=0.99):
self.policy = policy
self.gamma = gamma
def __call__(self, trajectory):
# agent takes `nsteps` actions
nsteps = len(trajectory['actions'])
# target values forward view
target_values = []
r = self.policy.get_value(
trajectory['state']['latest_observation']).squeeze(dim=1)
for i in range(nsteps-1, -1, -1):
immediate_reward = trajectory['rewards'][i]
#print(immediate_reward)
immediate_reward = torch.Tensor(immediate_reward,
device=device)
r = immediate_reward + self.gamma * r * trajectory['resets'][i]
target_values.append(r)
# reverse discounted rewards
trajectory['target_values'] = [
r for r in reversed(target_values)]
return
# +
# get one trajectory
import runners
model = DQN(obs_dim=84, n_actions=6)
policy = Policy(model)
runner = runners.EnvRunner(
env, policy, nsteps=5,
transforms=[
ComputeTargetValues(policy),
])
trajectory = runner.get_next()
# -
# After computing value targets we will transform lists of interactions into tensors
# with the first dimension `batch_size` which is equal to `T * nenvs`, i.e. you essentially need
# to flatten the first two dimensions.
class MergeTimeBatch:
""" Merges first two axes typically representing time and env batch. """
def __call__(self, trajectory):
# Modify trajectory inplace.
#<TODO: implement>
pass
# +
def f(trajectory):
# agent takes `nsteps` actions
nsteps = len(trajectory['actions'])
# discounted rewards forward view
discounted_rewards = []
r = policy.get_value(
trajectory['state']['latest_observation']).squeeze(dim=1)
print(r.shape)
for i in range(nsteps-1, -1, -1):
immediate_reward = trajectory['rewards'][i]
print('== reward shape ==', immediate_reward.shape)
print('== reset shape ==', trajectory['resets'][i].shape)
immediate_reward = torch.Tensor(immediate_reward,
device=device)
r = immediate_reward + 0.99 * r * trajectory['resets'][i]
discounted_rewards.append(r)
print('== target reward shape ==', r.shape)
# reverse discounted rewards
trajectory['discounted_rewards'] = [
r for r in reversed(discounted_rewards)]
return
# -
# Now is the time to implement the advantage actor critic algorithm itself. You can look into your lecture,
# [Mnih et al. 2016](https://arxiv.org/abs/1602.01783) paper, and [lecture](https://www.youtube.com/watch?v=Tol_jw5hWnI&list=PLkFD6_40KJIxJMR-j5A1mkxK26gh_qg37&index=20) by <NAME>.
# Policy objective along one trajectory
#
# $$
# \frac{1}{T}\sum_{i=t_1}^{T} \log(\pi(a_i | s_i)) (R_i - V(s_i, \theta_v))
# $$
#
# The length of the trajectory is $T$.
#
# Maximize likelihood of selected actions based on its advantage.
#
# For distributed RL, just take average among all workers
# +
# policy loss along one trajectory
# estimated value into one tensor of shape (n_step, n_workers)
def transform_estimated_values(estimated_values):
"""
estimated_values: estimated values from env runner
"""
batched = [x.squeeze(1) for x in estimated_values]
return torch.vstack(batched)
# -
transform_estimated_values(trajectory['estimated_values'])
# transform target values r + \gamma v(next state) into shape (n_steps, n_workers)
trajectory['target_values']
# +
def transform_target_values(target_values):
"""transform target values r + \gamma v(next state) into shape (n_steps, n_workers)
target_values: target values along the trajectory for all workers from env runner
"""
return torch.vstack(target_values)
transform_target_values(trajectory['target_values'])
# -
trajectory['log_probs']
# +
def transform_log_probs(log_probs):
"""
log_probs: log probabilities of actions for all workers from env runner
"""
return torch.vstack(log_probs)
transform_log_probs(trajectory['log_probs'])
# +
# Compute policy loss
target_values = transform_target_values(trajectory['target_values'])
estimated_values = transform_estimated_values(trajectory['estimated_values'])
# advantages
adv = target_values - estimated_values
# log probabilities
log_probs = transform_log_probs(trajectory['log_probs'])
# policy loss
policy_loss = -torch.mean(log_probs * adv)
print(policy_loss)
# -
# Loss for critic
#
# Want to minimize the difference between estimated values and target values (just like q-learning)
#
# $$
# \frac{1}{T} \sum_{i=t_1}^T (R_i - V(s_i, \theta))^2
# $$
#
# +
# Compute Value loss
target_values = transform_target_values(trajectory['target_values'])
estimated_values = transform_estimated_values(trajectory['estimated_values'])
value_loss = torch.mean((target_values - estimated_values)**2)
print(value_loss)
# +
# Package everything in an object
# -
class A2C:
def __init__(self,
policy,
optimizer,
value_loss_coef=0.25,
entropy_coef=0.01,
max_grad_norm=0.5):
self.policy = policy
self.optimizer = optimizer
self.value_loss_coef = value_loss_coef
self.entropy_coef = entropy_coef
self.max_grad_norm = max_grad_norm
def policy_loss(self, trajectory):
# You will need to compute advantages here.
target_values = transform_target_values(trajectory['target_values'])
estimated_values = transform_estimated_values(trajectory['estimated_values'])
# advantages
adv = target_values - estimated_values
# log probabilities
log_probs = transform_log_probs(trajectory['log_probs'])
# policy loss
policy_loss = -torch.mean(log_probs * adv)
return policy_loss
def value_loss(self, trajectory):
target_values = transform_target_values(trajectory['target_values'])
estimated_values = transform_estimated_values(trajectory['estimated_values'])
value_loss = torch.mean((target_values - estimated_values)**2)
return value_loss
def loss(self, trajectory):
return self.policy_loss(trajectory) + self.value_loss(trajectory)
def step(self, trajectory):
loss = self.loss(trajectory)
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
return loss.detach().cpu().numpy().item()
# +
# evaluate the agent
def evaluate(agent, env, n_games=1):
"""Plays an a game from start till done, returns per-game rewards
agent: an agent with .choose_action interface
test
env: a testing env (no parallelism)
"""
game_rewards = []
for _ in range(n_games):
state = env.reset()
total_reward = 0
while True:
action = agent.choose_action(state)
state, reward, done, info = env.step(action)
total_reward += reward.item()
if done:
break
game_rewards.append(total_reward)
return game_rewards
model = DQN(obs_dim=84, n_actions=6)
policy = Policy(model)
single_env = nature_dqn_env("SpaceInvadersNoFrameskip-v4", nenvs=1,
summaries='Numpy')
game_rewards = evaluate(policy, single_env, n_games=10)
print('Sampled game rewards: {}'.format(game_rewards))
# -
# Now you can train your model. With reasonable hyperparameters training on a single GTX1080 for 10 million steps across all batched environments (which translates to about 5 hours of wall clock time)
# it should be possible to achieve *average raw reward over last 100 episodes* (the average is taken over 100 last
# episodes in each environment in the batch) of about 600. You should plot this quantity with respect to
# `runner.step_var` — the number of interactions with all environments. It is highly
# encouraged to also provide plots of the following quantities (these are useful for debugging as well):
#
# * [Coefficient of Determination](https://en.wikipedia.org/wiki/Coefficient_of_determination) between
# value targets and value predictions
# * Entropy of the policy $\pi$
# * Value loss
# * Policy loss
# * Value targets
# * Value predictions
# * Gradient norm
# * Advantages
# * A2C loss
#
# For optimization we suggest you use RMSProp with learning rate starting from 7e-4 and linearly decayed to 0, smoothing constant (alpha in PyTorch and decay in TensorFlow) equal to 0.99 and epsilon equal to 1e-5.
# +
#https://stackoverflow.com/questions/58686400/can-not-get-pytorch-working-with-tensorboard
# +
from torch.utils.tensorboard import SummaryWriter
model = DQN(obs_dim=84, n_actions=6)
policy = Policy(model)
env = nature_dqn_env("SpaceInvadersNoFrameskip-v4", nenvs=nenvs,
summaries='Numpy')
test_env = nature_dqn_env("SpaceInvadersNoFrameskip-v4", nenvs=1,
summaries='Numpy')
env.reset()
test_env.reset()
runner = runners.EnvRunner(
env, policy, nsteps=5,
transforms=[
ComputeTargetValues(policy),
])
optimizer = torch.optim.RMSprop(policy.model.parameters(),
lr=7e-4,
alpha=0.99,
eps=1e-5)
a2c = A2C(policy, optimizer)
writer = SummaryWriter(log_dir='a2c_log')
n_iters = 1000*100
losses = []
rewards = []
for i in range(1, n_iters+1):
trajectory = runner.get_next()
loss = a2c.step(trajectory)
if i % 1000 == 0:
print('loss at iter {} is: {}'.format(i, loss))
# look at agent's performance on test env
game_rewards = evaluate(policy, test_env, n_games=3)
avg_reward = sum(game_rewards) / len(game_rewards)
print('reward at iter {} is: {}'.format(i, avg_reward))
losses.append(loss)
rewards.append(avg_reward)
writer.add_scalar('loss', loss, i)
writer.add_scalar('rewards', avg_reward, i)
|
week06_policy_based/a2c-optional.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/probml/JSL/blob/main/JSL_notebook.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="rX9X_KFwEtoK" outputId="d4a89034-65ee-4ceb-fb52-863be8ae9225"
# !pip install --upgrade git+https://github.com/google/flax.git
# !pip install --upgrade tensorflow-probability
# !pip install git+git://github.com/blackjax-devs/blackjax.git
# !pip install git+git://github.com/deepmind/distrax.git
# !pip install superimport
# !pip install fire
# + id="ye9LqR2JE-bD"
import tensorflow as tf
import tensorflow_probability as tfp
# + id="nzlaz5G4IyTp"
# !rm -rf JSL
# + colab={"base_uri": "https://localhost:8080/"} id="asDC0nQvFXee" outputId="b1768f77-4efa-4269-854b-8faf6ba08877"
# !git clone https://github.com/probml/JSL.git
# #!pip install git+git://github.com/probml/jsl
# + colab={"base_uri": "https://localhost:8080/"} id="RHRJ6MA-I6c9" outputId="0fb1f63a-a547-4f42-a04b-bd7fb1638632"
# %cd /content/JSL/
# !pip install -e .
# + colab={"base_uri": "https://localhost:8080/"} id="YhvXkCeiI4aX" outputId="ff3a0825-c33e-495c-d8f2-c2404f0c1b4d"
# #%run JSL/jsl/demos/kf_tracking_demo.py
# #%run jsl.demos.kf_tracking_demo
# + colab={"base_uri": "https://localhost:8080/", "height": 796} id="RMKqBKfDFbcp" outputId="fc2931de-1b4b-4c9b-964c-35a9dab44ed2"
from jsl.demos import kf_tracking_demo as demo
figures = demo.main()
#print(figures)
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="_b8lNuou9RaT" outputId="089f6972-5b68-48f2-98ee-cf914fb8a06e"
from jsl.demos import hmm_casino_demo as demo
demo()
# + id="CWEhPo9YJuut"
|
JSL_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='top'></a>
# # How to slice and dice the data
# Below are a series of examples on how to slice and dice the data that is stored in the *.sqlite* file generated by the [MorningStar.com](https://www.morningstar.com) web scraper.
#
# ##### NOTE:
# - The data used in the code below come from the *.sqlite* file that is automatically generated by the web scraper once it has been installed and ran locally on your machine. See [README]() for instructions on how to run install and run the scraper.
# - Navigation links only when using [Jupyter notebook](https://jupyter.org/).
#
#
# **Content**
#
# 1. [Required modules and matplotlib backend](#modules)
# 1. [Creating a master (bridge table) DataFrame instance using the DataFrames class](#master)
# 1. [Methods for creating DataFrame instances](#methods)
# 1. `quoteheader` - [MorningStar (MS) Quote Header](#quote)
# 1. `valuation` - [MS Valuation table with Price Ratios (P/E, P/S, P/B, P/C) for the past 10 yrs](#val)
# 1. `keyratios` - [MS Ratio - Key Financial Ratios & Values](#keyratios)
# 1. `finhealth` - [MS Ratio - Financial Health](#finhealth)
# 1. `profitability` - [MS Ratio - Profitability](#prof)
# 1. `growth` - [MS Ratio - Growth](#growth)
# 1. `cfhealth` - [MS Ratio - Cash Flow Health](#cfh)
# 1. `efficiency` - [MS Ratio - Efficiency](#eff)
# 1. `annualIS` - [MS Annual Income Statements](#isa)
# 1. `quarterlyIS` - [MS Quarterly Income Statements](#isq)
# 1. `annualBS` - [MS Annual Balance Sheets](#bsa)
# 1. `quarterlyBS` - [MS Quarterly Balance Sheets](#bsq)
# 1. `annualCF` - [MS Annual Cash Flow Statements](#cfa)
# 1. `quarterlyCF` - [MS Quarterly Cash Flow Statements](#cfq)
# 1. `insider_trades` - [Insider Transactions](#it)
# 1. [Performing statistical analysis](#stats)
# 1. [Count of database records](#stats)
# 1. [Last updated dates](#lastupdate)
# 1. [Number of records by security type](#type)
# 1. [Number of records by country, based on the location of exchanges](#country)
# 1. [Number of records per exchange](#exchange)
# 1. [Number of stocks by sector](#sector)
# 1. [Number of stocks by industry](#industry)
# 1. [Mean price ratios (P/E, P/S, P/B, P/CF) of stocks by sectors](#meanpr)
# 1. [Applying various criteria to filter common stocks](#value) *(in progress)*
# 1. [Additional sample / test code](#additional) *(in progress)*
# <a id="modules"></a>
# # Required modules and matplotlib backend
# +
# %matplotlib notebook
import matplotlib.pyplot as plt
import matplotlib
import pandas as pd
import numpy as np
import dataframes #module containing class used to create DataFrame objects from SQLite database file
import datetime as DT
import re
# Reload in case changes have been made to module file
from importlib import reload
reload(dataframes);
# -
# [return to the top](#top)
# <a id="master"></a>
# # Creating the master DataFrame instance
# The DataFrames class is part of the [dataframes module](dataframes.py)
db_file_name = 'mstables' # SQLite database file that contains the data to be analyzed
df = dataframes.DataFrames('db/{}.sqlite'.format(db_file_name))
# ## Creating Master DataFrame instance from reference tables
# Merge `df.master` (*Master* bridge table) with other reference tables (e.g. `df.tickers`, `df.exchanges`, etc.) and filter out inactive / invalid records.
# ### DataFrame Instance
# Create `df_master`
df_master = df.master.copy()
# Apply the following filter:
# <br>
# - $lastdate < cutoff\_date$
# <br>
# - $cutoff\_date = date one\ week\ prior\ to\ last\ date\ the\ database\ was\ updated$
# <br>
# <br>
# This filter ensure that only active records are included in the master dataframe, excluding inactive MorningStar records which are no longer being updated on a regular basis. These inactive records are typically symbols that are no longer active in their exchange.
# +
cutoff_days = 10
df_updated_ct = df_master[['update_date', 'ticker']].groupby('update_date').count().sort_index()
cutoff_date = df_updated_ct[df_updated_ct['ticker'] > 100].index[0] - DT.timedelta(days=cutoff_days)
df_master = df_master.where(df_master['lastdate'] >= cutoff_date).dropna(axis=0, how='all')
# -
# #### DataFrame Size
msg = 'DataFrame df_master contains {:,.0f} records and {:,.0f} columns.'
print(msg.format(df_master.shape[0], df_master.shape[1]))
# #### DataFrame Columns
df_master.columns.values
# <br></br>
# [return to the top](#top)
# <a id='methods'></a>
# # Creating DataFrame instances with dataframes methods
# Class DataFrames from [dataframe.py](dataframe.py) contains the following methods that return a pd.DataFrame object for the specified database table:
#
# - `quoteheader` - [MorningStar (MS) Quote Header](#quote)
# - `valuation` - [MS Valuation table with Price Ratios (P/E, P/S, P/B, P/C) for the past 10 yrs](#val)
# - `keyratios` - [MS Ratio - Key Financial Ratios & Values](#keyratios)
# - `finhealth` - [MS Ratio - Financial Health](#finhealth)
# - `profitability` - [MS Ratio - Profitability](#prof)
# - `growth` - [MS Ratio - Growth](#growth)
# - `cfhealth` - [MS Ratio - Cash Flow Health](#cfh)
# - `efficiency` - [MS Ratio - Efficiency](#eff)
# - `annualIS` - [MS Annual Income Statements](#isa)
# - `quarterlyIS` - [MS Quarterly Income Statements](#isq)
# - `annualBS` - [MS Annual Balance Sheets](#bsa)
# - `quarterlyBS` - [MS Quarterly Balance Sheets](#bsq)
# - `annualCF` - [MS Annual Cash Flow Statements](#cfa)
# - `quarterlyCF` - [MS Quarterly Cash Flow Statements](#cfq)
# - `insider_trades` - [Insider Transactions](#it)
#
# <a id='quote'></a>
# ### Quote Header
# ##### DataFrame Instance
df_quote = df.quoteheader()
df_quote.head()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_quote)))
# <a id='val'></a>
# [return to the top](#top)
# ### Price Ratios (P/E, P/S, P/B, P/C)
# ##### DataFrame Instance
df_vals = df.valuation().reset_index()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_vals)))
# ##### DataFrame Columns
df_vals.columns
# <a id='keyratios'></a>
# [return to the top](#top)
# ### Key Ratios
# ##### DataFrame Instance
df_keyratios = df.keyratios()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_keyratios)))
# ##### DataFrame Columns
df_labels_keyratios = (df_keyratios
.loc[0, [col for col in df_keyratios.columns if 'Y' not in col and col.startswith('i')]]
.replace(df.colheaders['header']))
df_labels_keyratios
# <a id='finhealth'></a>
# [return to the top](#top)
# ### Financial Health
# ##### DataFrame Instance
df_finhealth = df.finhealth()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_finhealth)))
# ##### DataFrame Columns
df_labels_finhealth = (df_finhealth.loc[0, [col for col in df_finhealth.columns
if 'Y' not in col and '_id' not in col]]
.replace(df.colheaders['header']))
df_labels_finhealth
# <a id='prof'></a>
# [return to the top](#top)
# ### Profitability
# ##### DataFrame Instance
df_profitab = df.profitability()
df_profitab.head()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_profitab)))
# ##### DataFrame Columns
df_labels_profitab = (df_profitab.loc[0, [col for col in df_profitab.columns if 'Y' not in col and '_id' not in col]]
.replace(df.colheaders['header']))
df_labels_profitab
# <a id='growth'></a>
# [return to the top](#top)
# ### Growth
# ##### DataFrame Instance
df_growth = df.growth()
df_growth.head()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_growth)))
# ##### DataFrame Columns
df_labels_growth = (df_growth.loc[0, [col for col in df_growth.columns
if 'Y' not in col and '_id' not in col]].replace(df.colheaders['header']))
df_labels_growth
# <a id='cfh'></a>
# [return to the top](#top)
# ### Cash Flow Health
# ##### DataFrame Instance
df_cfhealth = df.cfhealth()
df_cfhealth.head()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_cfhealth)))
# ##### DataFrame Columns
df_labels_cfhealth = df_cfhealth.loc[0, [col for col in df_cfhealth.columns if 'Y' not in col
and '_id' not in col]].replace(df.colheaders['header'])
df_labels_cfhealth
# <a id='eff'></a>
# [return to the top](#top)
# ### Efficiency
# ##### DataFrame Instance
df_efficiency = df.efficiency()
df_efficiency.head()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_efficiency)))
# ##### DataFrame Columns
# Financial Health DataFrame Columns
(df_efficiency.loc[0, [col for col in df_efficiency.columns if 'Y' not in col and '_id' not in col]]
.replace(df.colheaders['header']))
# <a id='isa'></a>
# [return to the top](#top)
# ### Annual Income Statement
# ##### DataFrame Instance
df_annualIS = df.annualIS()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_annualIS)))
# ##### DataFrame Columns
# +
labels = [col for col in df_annualIS if 'label' in col]
labels = [[label, header] for label in labels
for header in df_annualIS[label].unique().tolist() if pd.notna(header)]
df_labels_aIS = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_aIS['value'] = df_labels_aIS['value'].replace(df.colheaders['header'])
df_labels_aIS[df_labels_aIS['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_aIS.values.tolist(), df_labels_aIS.index)))
# -
# <a id='isq'></a>
# [return to the top](#top)
# ### Quarterly Income Statements
# ##### DataFrame Instance
df_quarterlyIS = df.quarterlyIS()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_quarterlyIS)))
# ##### DataFrame Columns
# +
labels = [col for col in df_annualIS if 'label' in col]
labels = [[label, header] for label in labels
for header in df_annualIS[label].unique().tolist() if pd.notna(header)]
df_labels_aIS = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_aIS['value'] = df_labels_aIS['value'].replace(df.colheaders['header'])
df_labels_aIS[df_labels_aIS['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_aIS.values.tolist(), df_labels_aIS.index)))
# -
# <a id='bsa'></a>
# [return to the top](#top)
# ### Annual Balance Sheet
# ##### DataFrame Instance
df_annualBS = df.annualBS()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_annualBS)))
# ##### DataFrame Columns
# +
labels = [col for col in df_annualBS if 'label' in col]
labels = [[label, header] for label in labels
for header in df_annualBS[label].unique().tolist() if pd.notna(header)]
df_labels_aBS = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_aBS['value'] = df_labels_aBS['value'].replace(df.colheaders['header'])
df_labels_aBS[df_labels_aBS['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_aBS.values.tolist(), df_labels_aBS.index)))
# -
# <a id='bsq'></a>
# [return to the top](#top)
# ### Quarterly Balance Sheet
# ##### DataFrame Instance
df_quarterlyBS = df.quarterlyBS()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_quarterlyBS)))
# ##### DataFrame Columns
# +
labels = [col for col in df_quarterlyBS if 'label' in col]
labels = [[label, header] for label in labels
for header in df_quarterlyBS[label].unique().tolist() if pd.notna(header)]
df_labels_qBS = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_qBS['value'] = df_labels_qBS['value'].replace(df.colheaders['header'])
df_labels_qBS[df_labels_qBS['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_qBS.values.tolist(), df_labels_qBS.index)))
# -
# <a id='cfa'></a>
# [return to the top](#top)
# ### Annual Cash Flow Statement
# ##### DataFrame Instance
df_annualCF = df.annualCF()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_annualCF)))
# ##### DataFrame Columns
# +
labels = [col for col in df_annualCF if 'label' in col]
labels = [[label, header] for label in labels
for header in df_annualCF[label].unique().tolist() if pd.notna(header)]
df_labels_aCF = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_aCF['value'] = df_labels_aCF['value'].replace(df.colheaders['header'])
df_labels_aCF[df_labels_aCF['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_aCF.values.tolist(), df_labels_aCF.index)))
# -
# <a id='cfq'></a>
# [return to the top](#top)
# ### Quarterly Cash Flow Statement
# ##### DataFrame Instance
df_quarterlyCF = df.quarterlyCF()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_quarterlyCF)))
# ##### DataFrame Columns
# +
labels = [col for col in df_quarterlyCF if 'label' in col]
labels = [[label, header] for label in labels
for header in df_quarterlyCF[label].unique().tolist() if pd.notna(header)]
df_labels_qCF = (pd.DataFrame(labels, columns=['header', 'value']).set_index('header').astype('int'))
df_labels_qCF['value'] = df_labels_qCF['value'].replace(df.colheaders['header'])
df_labels_qCF[df_labels_qCF['value'].astype('str').str.contains('ncome')].sort_values(by='value')
sorted(list(zip(df_labels_qCF.values.tolist(), df_labels_qCF.index)))
# -
# <a id='it'></a>
# [return to the top](#top)
# ### Insider Transactions
# ##### DataFrame Instance
df_insidertrades = df.insider_trades()
# ##### DataFrame Length
print('DataFrame contains {:,.0f} records.'.format(len(df_insidertrades)))
# <a id="stats"></a>
# [return to the top](#top)
# # Performing statistical analysis
# ### Count of database records
# **1.** Total number of records **before** merging reference tables (length of `df.master0`)
print('DataFrame df.master contains {:,.0f} records.'.format(len(df.master0)))
# **2.** Total number of records **after** merging reference tables (length of `df.master`)
print('DataFrame df_master0 contains {:,.0f} records.'.format(len(df.master)))
# **3.** Total number of records **after** filtering out inactive records (length of `df_master`)
print('DataFrame df_master contains {:,.0f} records.'.format(len(df_master)))
# <a id="lastupdate"></a>
# [return to the top](#top)
# ### Last updated dates
# List of dates (as a pd.Series object) of when the database records were last updated.
# The values indicate the number of records updated on each date.
(df_master[['update_date', 'ticker']].groupby(by='update_date').count().sort_index(ascending=False)
.rename(columns={'ticker':'ticker_count'}))
# <a id="type"></a>
# [return to the top](#top)
# ### Number of records by Security Type
(df_master[['security_type', 'ticker']].groupby(by='security_type').count()
.rename(columns={'ticker':'ticker_count'}))
# <a id="country"></a>
# [return to the top](#top)
# ### Number of records by Country, based on the location of exchanges
(df_master[['country', 'country_c3', 'ticker']]
.groupby(by=['country', 'country_c3']).count().rename(columns={'ticker':'ticker_count'})
)
# <a id="exchange"></a>
# [return to the top](#top)
# ### Number of records per exchange
# Where $ticker\_count > 100$
cols = ['country', 'country_c3', 'exchange', 'exchange_sym', 'ticker']
df_exchanges = df_master[cols].groupby(by=cols[:-1]).count().rename(columns={'ticker':'ticker_count'})
df_exchanges[df_exchanges['ticker_count'] > 100].sort_values(by='ticker_count', ascending=False)
#
# [return to the top](#top)
# ### Number of Stocks by Country of Exchange
(df_master
.where(df_master['security_type'] == 'Stock').dropna(axis=0, how='all')[['country', 'country_c3', 'ticker']]
.groupby(by=['country', 'country_c3']).count().rename(columns={'ticker':'ticker_count'})
.sort_values(by='ticker_count', ascending=False))
# <a id="sector"></a>
# [return to the top](#top)
# ### Number of stocks by sector
(df_master
.where((df_master['security_type'] == 'Stock') & (df_master['sector'] != '—')).dropna(axis=0, how='all')
.groupby(by='sector').count()
.rename(columns={'ticker':'stock_count'}))['stock_count'].sort_values(ascending=False)
# <a id="industry"></a>
# [return to the top](#top)
# ### Number of stocks by industry
(df_master[['sector', 'industry', 'ticker']]
.where((df_master['security_type'] == 'Stock') & (df_master['industry'] != '—')).dropna(axis=0, how='all')
.groupby(by=['sector', 'industry']).count().rename(columns={'ticker':'stock_count'}))
# <a id="meanpr"></a>
# [return to the top](#top)
# ### Mean price ratios (P/E, P/S, P/B, P/CF) of stocks by sectors
# First, merge `df_master` and `df_vals` and remove outliers where Price Ratio > 10,000
# +
df_valuation = (df_master
.where((df_master['security_type'] == 'Stock') & (df_master['sector'] != '—'))
.dropna(axis=0, how='all')
.merge(df_vals, on=['ticker_id', 'exchange_id'])
.drop(['ticker_id', 'exchange_id'], axis=1))
cols = list(filter(lambda col: col.startswith('P'), df_valuation))
df0 = df_valuation.copy()
for col in cols:
df0 = df[(df[col] < 10000) | df[col].isna()]
print('There are {:,.0f} Stock records that fit this criteria.'.format(len(df)))
# -
# #### Mean TTM Price Ratios for all stocks:
# +
df_val_mean = (df[['sector', 'company']].groupby('sector').count()
.rename(columns={'company':'count'})
.merge(df.groupby('sector').mean().round(4), on='sector')
.sort_values(by='PE_TTM', ascending=False))
df_val_mean[['count', 'PE_Forward', 'PE_TTM', 'PB_TTM', 'PS_TTM', 'PC_TTM']]
# -
# #### Mean TTM Price Ratios for USA stocks:
# +
df_valuation_USA = df[df['country_c3'] == 'USA']
print('There are {:,.0f} Stock records that fit this criteria.'.format(len(df_valuation_USA)))
df_val_mean_USA = (df_valuation_USA[['sector', 'company']].groupby('sector').count()
.rename(columns={'company':'count'})
.merge(df_valuation_USA.groupby('sector').mean().round(4), on='sector')
.sort_values(by='PE_TTM', ascending=False))
df_val_mean_USA[['count', 'PE_Forward', 'PE_TTM', 'PB_TTM', 'PS_TTM', 'PC_TTM']]
# -
# #### Mean TTM Price Ratios for DEU (Germany) stocks:
# +
df_valuation_DEU = df[df['country_c3'] == 'DEU']
print('There are {:,.0f} Stock records that fit this criteria.'.format(len(df_valuation_DEU)))
df_val_mean_DEU = (df_valuation_DEU[['sector', 'company']].groupby('sector').count()
.rename(columns={'company':'count'})
.merge(df_valuation_DEU.groupby('sector').mean().round(4), on='sector')
.sort_values(by='PE_TTM', ascending=False))
df_val_mean_DEU[['count', 'PE_Forward', 'PE_TTM', 'PB_TTM', 'PS_TTM', 'PC_TTM']]
# -
# #### Mean TTM Price Ratios for S&P 500 stocks:
url = r'https://en.wikipedia.org/wiki/List_of_S%26P_500_companies'
tables = pd.read_html(url)
df_sp500 = tables[0]
df_sp500.columns = df_sp500.iloc[0]
df_sp500 = df_sp500.drop(0, axis=0).set_index('Symbol').join(df.set_index('ticker'))
df_sp500 = df_sp500[df_sp500['country_c3'] == 'USA'].fillna(0)
# +
print('There are {:,.0f} Stock records that fit this criteria.'.format(len(df_sp500)))
df_val_mean_sp500 = (df_sp500[['sector', 'company']].groupby('sector').count()
.rename(columns={'company':'count'})
.merge(df_sp500.groupby('sector').mean().round(4), on='sector')
.sort_values(by='PE_TTM', ascending=False))
df_val_mean_sp500[['count', 'PE_Forward', 'PE_TTM', 'PB_TTM', 'PS_TTM', 'PC_TTM']]
# -
# [return to the top](#top)
# #### Plot of TTM P/E by Sectors
# *All Stocks*
# +
fig_pe, (ax_pe, ax_pe_usa, ax_pe_deu) = plt.subplots(3, 1, sharex=True, sharey=True, figsize=(8, 6))
# All Stocks
pe = df_val_mean['PE_TTM']
x = [x*3 for x in range(len(pe))]
y = pe
bars = ax_pe.bar(x, y, width=2)
for bar in bars:
ax_pe.text(bar.get_x()+1, bar.get_height()+1.5, '{:.1f}'.format(bar.get_height()),
color='black', ha='center', fontsize=9)
ax_pe.get_children()[22].set_color(None)
ax_pe.get_children()[23].set_color(None)
ax_pe.get_children()[25].set_color(None)
ax_pe.set_title('All Stocks', loc='left', fontsize=9, fontweight='bold')
# USA
pe_usa = df_val_mean_USA['PE_TTM']
x = [x*3 for x in range(len(pe_usa))]
y = pe_usa
bars = ax_pe_usa.bar(x, y, width=2)
for bar in bars:
ax_pe_usa.text(bar.get_x()+1, bar.get_height()+1.5, '{:.1f}'.format(bar.get_height()),
color='black', ha='center', fontsize=9)
ax_pe_usa.get_children()[22].set_color(None)
ax_pe_usa.get_children()[23].set_color(None)
ax_pe_usa.get_children()[25].set_color(None)
ax_pe_usa.set_title('USA', loc='left', fontsize=9, fontweight='bold')
# DEU
pe_deu = df_val_mean_DEU['PE_TTM']
x = [x*3 for x in range(len(pe_deu))]
y = pe_deu
bars = ax_pe_deu.bar(x, y, width=2)
for bar in bars:
ax_pe_deu.text(bar.get_x()+1, bar.get_height()+1.5, '{:.1f}'.format(bar.get_height()),
color='black', ha='center', fontsize=9)
ax_pe_deu.get_children()[22].set_color(None)
ax_pe_deu.get_children()[23].set_color(None)
ax_pe_deu.get_children()[25].set_color(None)
ax_pe_deu.set_title('DEU', loc='left', fontsize=9, fontweight='bold')
# Plot adjustments
plt.xticks(ticks=x, labels=y.index.tolist(), fontsize=9)
plt.axis([-3, len(x)*3, 0, 100])
plt.suptitle('Average TTM P/E of Stocks by Sector for key regions', fontsize=11, fontweight='bold')
plt.yticks([])
plt.subplots_adjust(bottom=0.3, hspace=1)
for tick in ax_pe_deu.xaxis.get_ticklabels():
tick.set_rotation(90)
# -
# [return to the top](#top)
# #### P/E by Sector for past 10 yrs for US Stocks
# +
cols = list(filter(lambda col: col.startswith('PE_'), df.columns))[1:]
fig, axs = plt.subplots(11, 1, figsize=(7, 6), sharex=True)
sectors = df_val_mean_USA.index.values
x = list(range(11))
for sector, ax in zip(sectors, axs):
y = df_val_mean_USA[cols].loc[sector].values
p0 = ax.bar(x, y, width=0.5)
#ax.set_title(sector, loc='left', fontsize=9, fontweight='bold')
ax.spines['left'].set_color(None)
ax.spines['right'].set_color(None)
ax.spines['top'].set_color(None)
for pt in list(zip(x, y)):
if pt[1] > 0:
ax.text(pt[0], pt[1] + max(y)*0.05, '{:.0f}'.format(pt[1]), ha='center', fontsize=8)
ax.set_yticks([])
ax.axis([-5.5, 11, 0, max(y)+10])
ax.set_xlim(-5.5, 11)
ax.text(-5.5, max(y)+10, sector, fontweight='bold', fontsize=8)
ax0 = ax
plt.suptitle('US Stocks P/E by Sector for past 10 yrs', fontweight='bold', fontsize=11)
plt.subplots_adjust(top=0.91, bottom=0.08, hspace=0.4)
_ = plt.xticks(ticks=x, labels=list(map(lambda col: col[3:], cols)), fontsize=8, fontweight='bold')
for label in ax.xaxis.get_ticklabels():
label.set_rotation(45)
# -
# [return to the top](#top)
# #### P/S by Sector for past 10 yrs for US Stocks
# +
cols = list(filter(lambda col: col.startswith('PS_'), df.columns))
fig, axs = plt.subplots(11, 1, figsize=(7, 6), sharex=True)
sectors = df_val_mean_USA.index.values
x = list(range(11))
for sector, ax in zip(sectors, axs):
y = df_val_mean_USA[cols].loc[sector].values
p0 = ax.bar(x, y, width=0.5)
#ax.set_title(sector, loc='left', fontsize=9, fontweight='bold')
ax.spines['left'].set_color(None)
ax.spines['right'].set_color(None)
ax.spines['top'].set_color(None)
for pt in list(zip(x, y)):
if pt[1] > 0:
ax.text(pt[0], pt[1] + max(y)*0.05, '{:.0f}'.format(pt[1]), ha='center', fontsize=8)
ax.set_yticks([])
ax.axis([-5.5, 11, 0, max(y)+10])
ax.set_xlim(-5.5, 11)
ax.text(-5.5, max(y)+10, sector, fontweight='bold', fontsize=8)
ax0 = ax
plt.suptitle('US Stocks P/S by Sector for past 10 yrs', fontweight='bold', fontsize=11)
plt.subplots_adjust(top=0.91, bottom=0.08, hspace=0.4)
_ = plt.xticks(ticks=x, labels=list(map(lambda col: col[3:], cols)), fontsize=8, fontweight='bold')
for label in ax.xaxis.get_ticklabels():
label.set_rotation(45)
# -
# [return to the top](#top)
# ### Stocks in the Cannabis Industry
# Using stocks listed on [marijuanaindex.com](http://marijuanaindex.com/stock-quotes/north-american-marijuana-index/) under North America
# +
import json
with open('input/pot_stocks.json') as file:
pot_symbols = json.loads(file.read())
pot_stocks = (pd.DataFrame(pot_symbols, columns=['ticker', 'country_c3'])
.merge(df_master, how='left', on=['ticker', 'country_c3']).drop('country', axis=1)
.rename(columns={'country_c3':'country', 'exchange_sym':'exch'}))
pot_stocks = (pot_stocks.where(((pot_stocks['country'] == 'USA') |
(pot_stocks['country'] == 'CAN')) &
(pot_stocks['sector'] != '—'))
.dropna(axis=0, how='all').sort_values(by='company'))
# +
msg = 'Below are the {} stocks listed on marijuanaindex.com for North America.'
print(msg.format(len(pot_stocks['company'].unique())))
pot_stocks[['country', 'ticker', 'exch', 'company', 'sector', 'industry']]
# -
# <a id="value"></a>
# [return to the top](#top)
#
# # Applying various criteria to filter common stocks
# Below is a list of different rules that can be applied to the data to screen stocks (development of *italicized rules* is still in progress)
#
# - **[Rule 0](#rule99): CAGR > 7% for past 7 years**
# - **[Rule 1](#rule1): No earnings deficit (loss) for past 5 or 7 years**
# - **[Rule 2](#rule2): Uniterrupted and increasing Dividends for past 5 yrs**
# - **[Rule 3](#rule3): P/E Ratio of 25 or less for the past 7 yrs and less then 20 for TTM**
# - **[Rule 4](#rule4): Growth for the past year**
# - **[Rule 5](#rule5): Current Ratio > 1.2**
# - **[Rule 6](#rule6): Debt/Equity < 1.0**
# - **[Rule 7](#rule7): Return on Equity > 10%**
# - **[Rule X](#rulex): Stocks with insider buys in the past 3 months**
#
# [Merge DataFrames](#mergerules) to screen stocks
# [return to the top](#top)
# <a id="rule99"></a>
# ## Rule 0. CAGR > 7% for past 5 years
# Column labels in `df_keyratios`:
df_labels_keyratios
# ### Revenue CAGR:
# +
iid = 'i0'
field = 'Rev'
label = 'CAGR_{}'.format(field)
df_rule0_Rev = (df_keyratios.where(df_keyratios['Y9'] > pd.to_datetime('2018-04-01')).dropna(axis=0, how='all'))
df_rule0_Rev[label] = 100 * ((df_rule0_Rev['{}_Y9'.format(iid)] / df_rule0_Rev['{}_Y4'.format(iid)]) ** (1/5) - 1)
cols = ['ticker_id', 'exchange_id', label] #, '{}_Y4'.format(iid), '{}_Y9'.format(iid)]
df_rule0_Rev = (df_rule0_Rev.where(df_rule0_Rev[label] >= 7).dropna(axis=0, how='all')
.sort_values(by=label, ascending=False))[cols]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule0_Rev)))
# -
# ### Operating Income CAGR:
# +
iid = 'i2'
field = 'OpeInc'
label = 'CAGR_{}'.format(field)
df_rule0_OpeInc = (df_keyratios.where(df_keyratios['Y9'] > pd.to_datetime('2018-04-01')).dropna(axis=0, how='all'))
df_rule0_OpeInc[label] = 100 * (
(df_rule0_OpeInc['{}_Y9'.format(iid)] / df_rule0_OpeInc['{}_Y4'.format(iid)]) ** (1/5) - 1)
cols = ['ticker_id', 'exchange_id', label]
df_rule0_OpeInc = (df_rule0_OpeInc.where(df_rule0_OpeInc[label] >= 7).dropna(axis=0, how='all')
.sort_values(by=label, ascending=False))[cols]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule0_OpeInc)))
# -
# ### Operating Cash Flow CAGR:
# +
iid = 'i9'
field = 'OpeCF'
label = 'CAGR_{}'.format(field)
df_rule0_OpeCF = (df_keyratios.where(df_keyratios['Y9'] > pd.to_datetime('2018-04-01')).dropna(axis=0, how='all'))
df_rule0_OpeCF[label] = 100 * (
(df_rule0_OpeCF['{}_Y9'.format(iid)] / df_rule0_OpeCF['{}_Y4'.format(iid)]) ** (1/5) - 1)
cols = ['ticker_id', 'exchange_id', label]
df_rule0_OpeCF = (df_rule0_OpeCF.where(df_rule0_OpeCF[label] >= 7).dropna(axis=0, how='all')
.sort_values(by=label, ascending=False))[cols]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule0_OpeCF)))
# -
# ### Free Cash Flow CAGR:
# +
iid = 'i11'
field = 'FreeCF'
label = 'CAGR_{}'.format(field)
df_rule0_FreeCF = (df_keyratios.where(df_keyratios['Y9'] > pd.to_datetime('2018-04-01')).dropna(axis=0, how='all'))
df_rule0_FreeCF[label] = 100 * (
(df_rule0_FreeCF['{}_Y9'.format(iid)] / df_rule0_FreeCF['{}_Y4'.format(iid)]) ** (1/5) - 1)
cols = ['ticker_id', 'exchange_id', label]
df_rule0_FreeCF = (df_rule0_FreeCF.where(df_rule0_FreeCF[label] >= 7).dropna(axis=0, how='all')
.sort_values(by=label, ascending=False))[cols]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule0_FreeCF)))
# -
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule1"></a>
# ### Rule 1. No earnings deficit (loss) for past 5 or 7 years
# Criteria: *"Find companies with positive earnings per share growth during the past five years with no earnings deficits. Earnings need to be higher in the most recent year than five years ago. Avoiding companies with earnings deficits during the past five years will help you stay clear of high-risk companies."* [(Source)](https://cabotwealth.com/daily/value-investing/benjamin-grahams-value-stock-criteria/)
#
# #### 5 Years: (PENDING CORRECTION OF CODE)
# *a. Identify Net Income column labels in* `df_annualIS`
ilabel = 'Net income'
df_labels = df_labels_aIS[df_labels_aIS['value'] == ilabel].sort_values(by='value')
df_labels
# *b. Get column headers for 'Net income' values for the past 5 yrs*
# +
# i_ids = [(label[-3:] + '_') for label in df_labels.index]
# def get_icols(col):
# for i_id in i_ids:
# if i_id in col:
# return True
# return False
# main_cols = ['ticker_id', 'exchange_id', 'country', 'exchange_sym', 'ticker', 'company', 'sector', 'industry',
# 'stock_type', 'style', 'Year_Y_6', 'Year_Y_5', 'Year_Y_4', 'Year_Y_3', 'Year_Y_2', 'Year_Y_1']
# data_cols = sorted(list(filter(get_icols, df_annualIS.columns)), key=lambda r: (r[-1], r[5:8]), reverse=True)
# -
# *c. Create 'Net Income' DataFrame*
# +
# df_annualIS1 = df_master.merge(df_annualIS, on=['ticker_id', 'exchange_id'])
# df_netinc5 = (df_annualIS1
# .where((df_annualIS1['security_type'] == 'Stock') &
# (df_annualIS1['Year_Y_5'] > pd.to_datetime('2018-01')))
# .dropna(axis=0, how='all')
# .drop(['country'], axis=1)
# .rename(columns={'country_c3':'country'})
# )[main_cols + data_cols]
# np_netinc = df_netinc5[data_cols].values
# netinc_cols = [('NetIncome_Y' + data_cols[i * 3][-1], (i * 3, i * 3 + 1, i * 3 + 2))
# for i in range(int(len(data_cols)/3))]
# vals = []
# for row in np_netinc:
# row_vals = []
# for i in range(len(netinc_cols)):
# val = None
# for col in netinc_cols[i][1]:
# if not np.isnan(row[col]):
# val = row[col]
# break
# row_vals.append(val)
# vals.append(row_vals)
# df_netinc_vals = pd.DataFrame(vals, columns=list(zip(*netinc_cols))[0])
# df_netinc5 = df_netinc5[main_cols].join(df_netinc_vals)
# +
# df_rule1_5 = df_netinc5.where((df_netinc5['NetIncome_Y6'] > 0) &
# ((df_netinc5['NetIncome_Y5'] > 0) | (df_netinc5['NetIncome_Y5'].isna() & df_netinc5['NetIncome_Y4'].isna() & df_netinc5['NetIncome_Y3'].isna() & df_netinc5['NetIncome_Y2'].isna() & df_netinc5['NetIncome_Y1'].isna())) &
# ((df_netinc5['NetIncome_Y4'] > 0) | (df_netinc5['NetIncome_Y4'].isna() & df_netinc5['NetIncome_Y3'].isna() & df_netinc5['NetIncome_Y2'].isna() & df_netinc5['NetIncome_Y1'].isna())) &
# ((df_netinc5['NetIncome_Y3'] > 0) | (df_netinc5['NetIncome_Y3'].isna() & df_netinc5['NetIncome_Y2'].isna() & df_netinc5['NetIncome_Y1'].isna())) &
# ((df_netinc5['NetIncome_Y2'] > 0) | (df_netinc5['NetIncome_Y2'].isna() & df_netinc5['NetIncome_Y1'].isna())) &
# ((df_netinc5['NetIncome_Y1'] > 0) | (df_netinc5['NetIncome_Y1'].isna()))
# ).dropna(axis=0, how='all')
# df_rule1_5 = df_rule1_5[['ticker_id', 'exchange_id'] + df_rule1_5.columns.values.tolist()[-12:]]
# df_rule1_5.columns = [re.sub('Year_Y_', 'r1_Y', col) for col in df_rule1_5.columns]
# print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule1_5)))
# -
# #### 5 Years:
# +
cols = ['ticker_id', 'exchange_id'] + \
[col for col in df_keyratios.columns if col.startswith('i4_') or col.startswith('Y')]
df_rule1_5 = (df_keyratios
.where((df_keyratios['Y9'] >= pd.to_datetime('2018-04-01')) &
(df_keyratios['i4_Y10'] > 0) &
((df_keyratios['i4_Y9'] > 0) | (df_keyratios['i4_Y9'].isna() & df_keyratios['i4_Y8'].isna() & df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna())) &
((df_keyratios['i4_Y8'] > 0) | (df_keyratios['i4_Y8'].isna() & df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna())) &
((df_keyratios['i4_Y7'] > 0) | (df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna())) &
((df_keyratios['i4_Y6'] > 0) | (df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna())) &
((df_keyratios['i4_Y5'] > 0) | (df_keyratios['i4_Y5'].isna())))
.dropna(axis=0, how='all'))[cols]
df_rule1_5.columns = [re.sub('i4_', 'NetIncome_', col) for col in df_rule1_5.columns]
df_rule1_5.columns = [re.sub('^Y', 'r1_Y', col) for col in df_rule1_5.columns]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule1_5)))
# -
# #### 7 Years:
# +
cols = ['ticker_id', 'exchange_id'] + \
[col for col in df_keyratios.columns if col.startswith('i4_') or col.startswith('Y')]
df_rule1_7 = (df_keyratios
.where((df_keyratios['Y9'] >= pd.to_datetime('2018-04-01')) &
(df_keyratios['i4_Y10'] > 0) &
((df_keyratios['i4_Y9'] > 0) | (df_keyratios['i4_Y9'].isna() & df_keyratios['i4_Y8'].isna() & df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna() & df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y8'] > 0) | (df_keyratios['i4_Y8'].isna() & df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna() & df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y7'] > 0) | (df_keyratios['i4_Y7'].isna() & df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna() & df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y6'] > 0) | (df_keyratios['i4_Y6'].isna() & df_keyratios['i4_Y5'].isna() & df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y5'] > 0) | (df_keyratios['i4_Y5'].isna() & df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y4'] > 0) | (df_keyratios['i4_Y4'].isna() & df_keyratios['i4_Y3'].isna())) &
((df_keyratios['i4_Y3'] > 0) | (df_keyratios['i4_Y3'].isna())))
.dropna(axis=0, how='all'))[cols]
df_rule1_7.columns = [re.sub('i4_', 'NetIncome_', col) for col in df_rule1_7.columns]
df_rule1_7.columns = [re.sub('^Y', 'r1_Y', col) for col in df_rule1_7.columns]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule1_7)))
# -
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule2"></a>
# ### Rule 2. Uniterrupted and increasing *Dividends* for past 7 yrs
df_labels_keyratios
icol = df_labels_keyratios[df_labels_keyratios.str.contains('Dividends')].index[0]
icol
main_cols = ['ticker_id', 'exchange_id',
#'country_c3', 'exchange_sym', 'ticker', 'company',
#'sector', 'industry', 'stock_type', 'style',
'Y10', 'Y9', 'Y8', 'Y7', 'Y6', 'Y5']
icols = sorted([col for col in df_keyratios.columns if icol + '_' in col],
key=lambda col: int(col[4:]), reverse=True)[:8]
icols
# +
df_rule2 = (df_keyratios
.where((df_keyratios['Y9'] > pd.to_datetime('2018-04-01')) &
(df_keyratios['i6_Y10'] >= df_keyratios['i6_Y9']) &
((df_keyratios['i6_Y9'] >= df_keyratios['i6_Y8']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna() & df_keyratios['i6_Y4'].isna() & df_keyratios['i6_Y5'].isna() & df_keyratios['i6_Y6'].isna() & df_keyratios['i6_Y7'].isna() & df_keyratios['i6_Y8'].isna())) &
((df_keyratios['i6_Y8'] >= df_keyratios['i6_Y7']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna() & df_keyratios['i6_Y4'].isna() & df_keyratios['i6_Y5'].isna() & df_keyratios['i6_Y6'].isna() & df_keyratios['i6_Y7'].isna())) &
((df_keyratios['i6_Y7'] >= df_keyratios['i6_Y6']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna() & df_keyratios['i6_Y4'].isna() & df_keyratios['i6_Y5'].isna() & df_keyratios['i6_Y6'].isna())) &
((df_keyratios['i6_Y6'] >= df_keyratios['i6_Y5']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna() & df_keyratios['i6_Y4'].isna() & df_keyratios['i6_Y5'].isna())) &
((df_keyratios['i6_Y5'] >= df_keyratios['i6_Y4']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna() & df_keyratios['i6_Y4'].isna())) &
((df_keyratios['i6_Y4'] >= df_keyratios['i6_Y3']) | (df_keyratios['i6_Y2'].isna() & df_keyratios['i6_Y3'].isna())) &
((df_keyratios['i6_Y3'] >= df_keyratios['i6_Y2']) | (df_keyratios['i6_Y2'].isna())))
.dropna(axis=0, how='all').sort_values(by='Y9', ascending=False))[main_cols + icols]
df_rule2.columns = main_cols + [col.replace('i6', 'Dividend') for col in icols]
df_rule2.columns = [re.sub('^Y', 'r2_Y', col) for col in df_rule2.columns]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule2)))
# -
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule3"></a>
# ### Rule 3. P/E Ratio of 25 or less for the past 7 yrs and less then 20 for TTM
pe_cols = [col for col in df_vals.columns if 'PE_' in col]
pe_cols = ['ticker_id', 'exchange_id'] + [pe_cols[len(pe_cols)-i-1] for i in range(len(pe_cols))][:8]
pe_cols
# +
df_rule3 = (df_vals[pe_cols]
.where((df_vals['PE_TTM'] <= 10) &
(df_vals['PE_2018'] <= 25) &
((df_vals['PE_2017'] <= 25) | (df_vals['PE_2012'].isna() & df_vals['PE_2013'].isna() & df_vals['PE_2014'].isna() & df_vals['PE_2015'].isna() & df_vals['PE_2016'].isna() & df_vals['PE_2017'].isna())) &
((df_vals['PE_2016'] <= 25) | (df_vals['PE_2012'].isna() & df_vals['PE_2013'].isna() & df_vals['PE_2014'].isna() & df_vals['PE_2015'].isna() & df_vals['PE_2016'].isna())) &
((df_vals['PE_2015'] <= 25) | (df_vals['PE_2012'].isna() & df_vals['PE_2013'].isna() & df_vals['PE_2014'].isna() & df_vals['PE_2015'].isna())) &
((df_vals['PE_2014'] <= 25) | (df_vals['PE_2012'].isna() & df_vals['PE_2013'].isna() & df_vals['PE_2014'].isna())) &
((df_vals['PE_2013'] <= 25) | (df_vals['PE_2012'].isna() & df_vals['PE_2013'].isna())) &
((df_vals['PE_2012'] <= 25) | (df_vals['PE_2012'].isna())))
.dropna(axis=0, how='all').sort_values(by='PE_TTM'))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule3)))
# -
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule4"></a>
# ## Rule 4. Growth for the past year
df_labels_growth
# ### Revenue
iid = 'i28'; label = '{}_gr_Y9'.format(iid); col = 'Rev';
cols = ['ticker_id', 'exchange_id', 'gr_Y9', label]
df_rule4_Rev = (df_growth[cols]
.where((df_growth[label] > 0) & (df_growth['gr_Y9'] > pd.to_datetime('2018-04-01')))
.dropna(axis=0, how='all').sort_values(by='gr_Y9')
.rename(columns={label:'{}_Growth_Y9'.format(col)}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule4_Rev)))
# ### Operating Income
iid = 'i32'; label = '{}_gr_Y9'.format(iid); col = 'OpeInc';
cols = ['ticker_id', 'exchange_id', 'gr_Y9', label]
df_rule4_OpeInc = (df_growth[cols]
.where((df_growth[label] > 0) & (df_growth['gr_Y9'] > pd.to_datetime('2018-04-01')))
.dropna(axis=0, how='all').sort_values(by='gr_Y9')
.rename(columns={label:'{}_Growth_Y9'.format(col)}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule4_OpeInc)))
# ### Net Income
iid = 'i81'; label = '{}_gr_Y9'.format(iid); col = 'NetInc';
cols = ['ticker_id', 'exchange_id', 'gr_Y9', label]
df_rule4_NetInc = (df_growth[cols]
.where((df_growth[label] > 0) & (df_growth['gr_Y9'] > pd.to_datetime('2018-04-01')))
.dropna(axis=0, how='all').sort_values(by='gr_Y9')
.rename(columns={label:'{}_Growth_Y9'.format(col)}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule4_NetInc)))
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule5"></a>
# ### Rule 5. Current Ratio > 1.2
df_labels_finhealth[-5:]
col = 'i65_lfh_Y10'
df_rule5 = (df_finhealth[['ticker_id', 'exchange_id', col]]
.where((df_finhealth[col] > 1.2) | (df_finhealth[col].isna()))
.dropna(axis=0, how='all')
.rename(columns={col:'current_ratio'}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule5)))
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule6"></a>
# ### Rule 6. Debt/Equity < 1.5
col = 'i68_lfh_Y10'
df_rule6 = (df_finhealth[['ticker_id', 'exchange_id', col]]
.where((df_finhealth[col] < 1.5) | (df_finhealth[col].isna()))
.dropna(axis=0, how='all')
.rename(columns={col:'debt2equity'}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule6)))
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule7"></a>
# ### Rule 7. Return on Equity > 10%
df_labels_profitab[-9:]
col = 'i26_pr_pro_Y10'
df_rule7 = (df_profitab[['ticker_id', 'exchange_id', col]]
.where((df_profitab[col] > 10) | (df_profitab[col].isna()))
.dropna(axis=0, how='all')
.rename(columns={col:'return_on_equity'}))
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule7)))
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rule7"></a>
# ### Rule 8. P/B < 1.0
df_rule8 = df_vals.where(df_vals['PB_TTM'] <= 1).dropna(axis=0, how='all')
print('Total of {:,.0f} records meet this criterium.'.format(len(df_rule8)))
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="rulex"></a>
# ### Rule X. Stocks with insider buys in the past 3 months
# +
datefilter = pd.to_datetime(DT.date.today()-DT.timedelta(days=90))
df_insiderbuys0 = (df_insidertrades
.where((df_insidertrades['type'] == 'Buy') & (df_insidertrades['date'] >= datefilter))
.dropna(axis=0, how='all').groupby(['ticker_id', 'exchange_id']).sum())
df_insiderbuys = (df_master.set_index(['ticker_id', 'exchange_id'])
.join(df_insiderbuys0, how='inner').reset_index()
.groupby(['company', 'sector', 'industry']).mean().round(1)
.sort_values(by='value', ascending=False)
)[['openprice', 'yield', 'quantity', 'value']]
print('Total of {:,.0f} records meet this criterium.'.format(len(df_insiderbuys)))
# -
with open('doc/df_insiderbuys.csv', 'w') as file:
file.write(df_insiderbuys.to_csv())
# [return to top of this section](#value),
# [return to the top](#top)
# <a id="mergerules"></a>
# ### Merging DataFrames
df_master.columns.values
# +
df_rules = (df_master[df_master['security_type'] == 'Stock']
#.merge(df_rule0_Rev, on=['ticker_id', 'exchange_id']) # CAGR > 7% for past 5 yrs - Revenue
#.merge(df_rule0_OpeInc, on=['ticker_id', 'exchange_id']) # CAGR > 7% for past 5 yrs - Ope. Income
#.merge(df_rule0_OpeCF, on=['ticker_id', 'exchange_id']) # CAGR > 7% for past 5 yrs - Ope. Cash Flow
#.merge(df_rule0_FreeCF, on=['ticker_id', 'exchange_id']) # CAGR > 7% for past 5 yrs - Free Cash Flow
.merge(df_rule1_5, on=['ticker_id', 'exchange_id']) # No earnings deficit for past 5 yrs
.merge(df_rule2, on=['ticker_id', 'exchange_id']) # Uniterrupted Dividends for past 7 yrs
.merge(df_rule3, on=['ticker_id', 'exchange_id']) # P/E Ratio of 10 or less for past 7 yrs
.merge(df_rule4_Rev, on=['ticker_id', 'exchange_id']) # Growth for the past year - Revenue
.merge(df_rule4_OpeInc, on=['ticker_id', 'exchange_id']) # Growth for the past year - Ope. Income
.merge(df_rule4_NetInc, on=['ticker_id', 'exchange_id']) # Growth for the past year - Net Income
.merge(df_rule5, on=['ticker_id', 'exchange_id']) # Current Ratio > 1.2
.merge(df_rule6, on=['ticker_id', 'exchange_id']) # Debt/Equity < 1.0
.merge(df_rule7, on=['ticker_id', 'exchange_id']) # Return on Equity > 10%
.merge(df_rule8, on=['ticker_id', 'exchange_id']) # P/B < 1.0
#.merge(df_insiderbuys0, on=['ticker_id', 'exchange_id']) # Insider buys in the past 3 months
#.merge(df_vals[['ticker_id', 'exchange_id', 'PB_TTM', 'PS_TTM', 'PC_TTM']],
# on=['ticker_id', 'exchange_id'])
#.groupby(['company', 'exchange_sym', 'ticker', 'sector', 'industry']).mean().round(1)
)[['company', 'exchange_sym', 'ticker', 'sector', 'industry',
'lastprice', 'yield', '_52wk_hi', '_52wk_lo',
'PE_TTM_x', 'PB_TTM', 'PS_TTM', 'PC_TTM', 'current_ratio', 'debt2equity', 'return_on_equity',
#'CAGR_Rev', 'CAGR_OpeInc', 'CAGR_OpeCF', 'CAGR_FreeCF',
'Rev_Growth_Y9', 'OpeInc_Growth_Y9', 'NetInc_Growth_Y9'#, 'value'
]]
total_companies = df_master[df_master['security_type'] == 'Stock']#.groupby('company').count()
msg = 'A total of {:,.0f} stocks meet these criteria out of {:,.0f} (as of {})'
print(msg.format(len(df_rules), len(total_companies), DT.date.today()))
# -
df_rules
with open('doc/df_rules.csv', 'w') as file:
file.write(df_rules.to_csv())
# <a id="additional"></a>
# [return to the top](#top)
#
# ## Additional sample / test code
import requests
from io import StringIO
from datetime import datetime
url = 'http://performance.morningstar.com/perform/Performance/stock/exportStockPrice.action?t={}:{}&pd=1yr&freq=d&pg=0&culture=en-US'
req = requests.get(url.format('xnas', 'aaoi'))
|
data_overview.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Artificial Intelligence Nanodegree
#
# ## Convolutional Neural Networks
#
# ---
#
# In this notebook, we train an MLP to classify images from the MNIST database.
#
# ### 1. Load MNIST Database
# +
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("The MNIST database has a training set of %d examples." % len(X_train))
print("The MNIST database has a test set of %d examples." % len(X_test))
# -
# ### 2. Visualize the First Six Training Images
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.cm as cm
import numpy as np
# plot first six training images
fig = plt.figure(figsize=(20,20))
for i in range(6):
ax = fig.add_subplot(1, 6, i+1, xticks=[], yticks=[])
ax.imshow(X_train[i], cmap='gray')
ax.set_title(str(y_train[i]))
# -
# ### 3. View an Image in More Detail
# +
def visualize_input(img, ax):
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
ax.annotate(str(round(img[x][y],2)), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
visualize_input(X_train[0], ax)
# -
# ### 4. Rescale the Images by Dividing Every Pixel in Every Image by 255
# + jupyter={"outputs_hidden": true}
# rescale [0,255] --> [0,1]
X_train = X_train.astype('float32')/255
X_test = X_test.astype('float32')/255
# -
# ### 5. Encode Categorical Integer Labels Using a One-Hot Scheme
# +
from keras.utils import np_utils
# print first ten (integer-valued) training labels
print('Integer-valued labels:')
print(y_train[:10])
# one-hot encode the labels
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
# print first ten (one-hot) training labels
print('One-hot labels:')
print(y_train[:10])
# -
# ### 6. Define the Model Architecture
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# -
# ### 7. Compile the Model
# + jupyter={"outputs_hidden": true}
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# -
# ### 8. Calculate the Classification Accuracy on the Test Set (Before Training)
# +
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
# -
# ### 9. Train the Model
# +
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=10,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# -
# ### 10. Load the Model with the Best Classification Accuracy on the Validation Set
# + jupyter={"outputs_hidden": true}
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# -
# ### 11. Calculate the Classification Accuracy on the Test Set
# +
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=10,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-regular: %.4f%%' % accuracy)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(784, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=10,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-extra nodes: %.4f%%' % accuracy)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=10,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-extra layers: %.4f%%' % accuracy)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dense(512, activation='relu'))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=10,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-NO DROPOUT: %.4f%%' % accuracy)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='sigmoid'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='sigmoid'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='rmsprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=15,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracyno relu/sigmoid instead: %.4f%%' % accuracy)
# +
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("The MNIST database has a training set of %d examples." % len(X_train))
print("The MNIST database has a test set of %d examples." % len(X_test))
# rescale [0,255] --> [0,1]
X_train = X_train.astype('float32')/255
X_test = X_test.astype('float32')/255
from keras.utils import np_utils
# print first ten (integer-valued) training labels
print('Integer-valued labels:')
print(y_train[:10])
# one-hot encode the labels
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
# print first ten (one-hot) training labels
#print('One-hot labels:')
#print(y_train[:10])
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='SGD',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=13,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-optimizer-SGD: %.4f%%' % accuracy)
# +
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("The MNIST database has a training set of %d examples." % len(X_train))
print("The MNIST database has a test set of %d examples." % len(X_test))
# rescale [0,255] --> [0,1]
X_train = X_train.astype('float32')/255
X_test = X_test.astype('float32')/255
from keras.utils import np_utils
# print first ten (integer-valued) training labels
print('Integer-valued labels:')
print(y_train[:10])
# one-hot encode the labels
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
# print first ten (one-hot) training labels
#print('One-hot labels:')
#print(y_train[:10])
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='RMSprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=12800, epochs=20,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-different batch size 128 to 12800: %.4f%%' % accuracy)
# +
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("The MNIST database has a training set of %d examples." % len(X_train))
print("The MNIST database has a test set of %d examples." % len(X_test))
# rescale [0,255] --> [0,1]
X_train = X_train.astype('float32')/255
X_test = X_test.astype('float32')/255
from keras.utils import np_utils
# print first ten (integer-valued) training labels
print('Integer-valued labels:')
print(y_train[:10])
# one-hot encode the labels
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
# print first ten (one-hot) training labels
#print('One-hot labels:')
#print(y_train[:10])
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='RMSprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=32, epochs=8,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-different batch size 128 to 32: %.4f%%' % accuracy)
# +
from keras.datasets import mnist
# use Keras to import pre-shuffled MNIST database
(X_train, y_train), (X_test, y_test) = mnist.load_data()
print("The MNIST database has a training set of %d examples." % len(X_train))
print("The MNIST database has a test set of %d examples." % len(X_test))
# rescale [0,255] --> [0,1]
# no normailization X_train = X_train.astype('float32')/255
# no normailization X_test = X_test.astype('float32')/255
from keras.utils import np_utils
# print first ten (integer-valued) training labels
print('Integer-valued labels:')
print(y_train[:10])
# one-hot encode the labels
y_train = np_utils.to_categorical(y_train, 10)
y_test = np_utils.to_categorical(y_test, 10)
# print first ten (one-hot) training labels
#print('One-hot labels:')
#print(y_train[:10])
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
# define the model
model = Sequential()
model.add(Flatten(input_shape=X_train.shape[1:]))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
# summarize the model
model.summary()
# compile the model
model.compile(loss='categorical_crossentropy', optimizer='RMSprop',
metrics=['accuracy'])
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy: %.4f%%' % accuracy)
from keras.callbacks import ModelCheckpoint
# train the model
checkpointer = ModelCheckpoint(filepath='mnist.model.best.hdf5',
verbose=1, save_best_only=True)
hist = model.fit(X_train, y_train, batch_size=128, epochs=20,
validation_split=0.2, callbacks=[checkpointer],
verbose=1, shuffle=True)
# load the weights that yielded the best validation accuracy
model.load_weights('mnist.model.best.hdf5')
# evaluate test accuracy
score = model.evaluate(X_test, y_test, verbose=0)
accuracy = 100*score[1]
# print test accuracy
print('Test accuracy-no normalization: %.4f%%' % accuracy)
|
mnist-mlp/mnist_mlp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# language: python
# name: python38564bit02a66c47ce504b05b2ef5646cfed96c2
# ---
import warnings
warnings.simplefilter('ignore', FutureWarning)
warnings.simplefilter('ignore', UserWarning)
# # Imbalanced data
# Imbalanced data occurs in classification when the number of instances in each class are not the same. Some care is required to learn to predict the *rare* classes effectively.
#
# There is no one-size-fits-all approach to handling imbalanced data. A reasonable strategy is to consider this as a model selection problem, and use cross-validation to find an approach that works well for your data sets. We will show how to do this in the hyper-parameter optimization notebook.
#
# **Warning**: Like most things in ML, techniques should not be applied blindly, but considered carefully with the problem goal in mind. In many cases, there is a decision-theoretic problem of assigning the appropriate costs to minority and majority case mistakes that requires domain knowledge to model correctly. As you will see in this example, blind application of a technique does not necessarily improve performance.
# ## Simulate an imbalanced data set
import pandas as pd
import numpy as np
X_train = pd.read_csv('data/X_train.csv')
X_test = pd.read_csv('data/X_test.csv')
y_train = pd.read_csv('data/y_train.csv')
y_test = pd.read_csv('data/y_test.csv')
X = pd.concat([X_train, X_test])
y = pd.concat([y_train, y_test]).squeeze()
y.value_counts()
np.random.seed(0)
idx = (
(y == 0) |
((y == 1) & (np.random.uniform(0, 1, y.shape) < 0.2))
).squeeze()
X_im, y_im = X.loc[idx, :], y[idx]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_im, y_im, random_state=0)
y_test.value_counts(), y_train.value_counts()
# ## Collect more data
#
# This is the best but often impractical solution. Synthetic data generation may also be an option.
# ## Use evaluation metrics that are less sensitive to imbalance
#
# For example, the `F1` score (harmonic mean of precision and recall) is less sensitive than the accuracy score.
from sklearn.linear_model import LogisticRegression
from sklearn.utils import class_weight
from sklearn.metrics import roc_auc_score, confusion_matrix
from sklearn.dummy import DummyClassifier
clf = DummyClassifier(strategy='prior')
clf.fit(X_train, y_train)
clf.score(X_test, y_test)
from sklearn.metrics import accuracy_score, f1_score, balanced_accuracy_score
accuracy_score(clf.predict(X_test), y_test)
f1_score(clf.predict(X_test), y_test)
lr = LogisticRegression()
lr.fit(X_train, y_train)
accuracy_score(lr.predict(X_test), y_test)
balanced_accuracy_score(lr.predict(X_test), y_test)
f1_score(lr.predict(X_test), y_test)
# ## Over-sample the minority class
#
# There are many ways to over-sample the minority class. A popular algorithm is known as SMOTE (Synthetic Minority Oversampling Technique)
#
# 
# ! python3 -m pip install --quiet imbalanced-learn
import imblearn
X_train_resampled, y_train_resampled = \
imblearn.over_sampling.SMOTE().fit_resample(X_train, y_train)
X_train.shape
X_train_resampled.shape
y_train.value_counts()
# ### Evaluate if this helps
lr = LogisticRegression()
lr.fit(X_train, y_train)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
lr.fit(X_train_resampled, y_train_resampled)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
# ## Under-sample the majority class
#
# Tomek pairs are nearest neighbor pairs of instances where the classes are different. Under-sampling is done by removing the majority member of the pair.
#
# 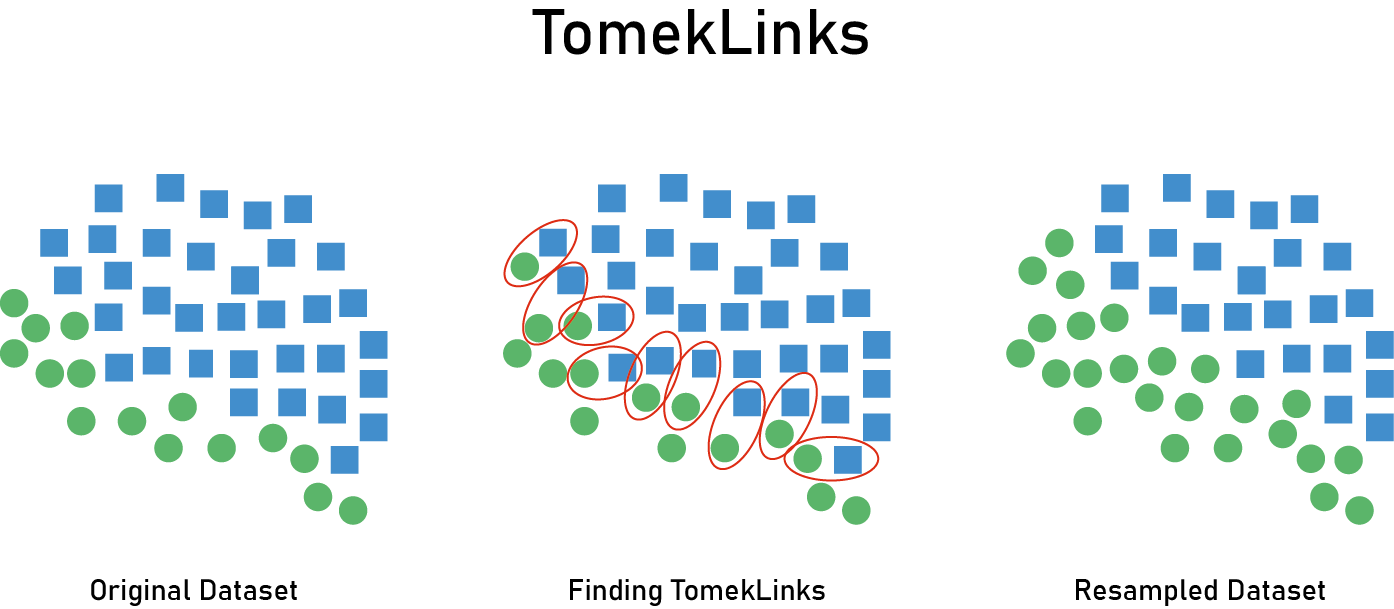
X_train_resampled, y_train_resampled = \
imblearn.under_sampling.TomekLinks().fit_resample(X_train, y_train)
X_train.shape
X_train_resampled.shape
y_train.value_counts()
y_train_resampled.value_counts()
# ### Evaluate if this helps
lr = LogisticRegression()
lr.fit(X_train, y_train)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
lr.fit(X_train_resampled, y_train_resampled)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
# ## Combine over- and under-sampling
#
# For example, over-sample using SMOTE then clean using Tomek.
X_train_resampled, y_train_resampled = \
imblearn.combine.SMOTETomek().fit_resample(X_train, y_train)
X_train.shape
X_train_resampled.shape
y_train.value_counts()
y_train_resampled.value_counts()
# ### Evaluate if this helps
lr = LogisticRegression()
lr.fit(X_train, y_train)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
lr.fit(X_train_resampled, y_train_resampled)
f1_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
# ## Use class weights to adjust the loss function
#
# We make prediction errors in the minority class more costly than prediction errors in the majority class.
wts = class_weight.compute_class_weight('balanced', np.unique(y_train), y_train)
wts
# You can then pass in the class weights. Note that there are several alternative ways to calculate possible class weights to use, and you can also do a GridSearch on weights.
# This is actually built-in to most classifiers. The defaults are equal weights to each class.
lr = LogisticRegression(class_weight=wts)
lr.fit(X_train, y_train)
lr.class_weight
f1_score(lr.predict(X_test), y_test)
roc_auc_score(lr.predict(X_test), y_test)
confusion_matrix(lr.predict(X_test), y_test)
lr_balanced = LogisticRegression(class_weight='balabced')
lr_balanced.class_weight
lr_balanced.fit(X_train, y_train)
roc_auc_score(lr_balanced.predict(X_test), y_test)
confusion_matrix(lr_balanced.predict(X_test), y_test)
f1_score(lr_balanced.predict(X_test), y_test)
# ## Use a classifier that is less sensitive to imbalance
#
# Boosted trees are generally good because of their sequential nature.
from catboost import CatBoostClassifier
cb = CatBoostClassifier()
cb.fit(X_train, y_train, verbose=0)
f1_score(cb.predict(X_test), y_test)
confusion_matrix(cb.predict(X_test), y_test)
# ### Imbalanced learn has classifiers that balance the data automatically
from imblearn.ensemble import BalancedRandomForestClassifier
brf = BalancedRandomForestClassifier()
brf.fit(X_train, y_train)
confusion_matrix(brf.predict(X_test), y_test)
f1_score(brf.predict(X_test), y_test)
|
notebooks/B07_Imbalnaced_Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + editable=false deletable=false dc={"key": "4"} run_control={"frozen": true} tags=["context"]
# ## 1. United Nations life expectancy data
# <p>Life expectancy at birth is a measure of the average a living being is expected to live. It takes into account several demographic factors like gender, country, or year of birth.</p>
# <p>Life expectancy at birth can vary along time or between countries because of many causes: the evolution of medicine, the degree of development of countries, or the effect of armed conflicts. Life expectancy varies between gender, as well. The data shows that women live longer that men. Why? Several potential factors, including biological reasons and the theory that women tend to be more health conscious.</p>
# <p>Let's create some plots to explore the inequalities about life expectancy at birth around the world. We will use a dataset from the United Nations Statistics Division, which is available <a href="http://data.un.org/Data.aspx?d=GenderStat&f=inID:37&c=1,2,3,4,5,6&s=crEngName:asc,sgvEngName:asc,timeEngName:desc&v=1">here</a>.</p>
# + dc={"key": "4"} tags=["sample_code"]
# This sets plot images to a nice size
options(repr.plot.width = 6, repr.plot.height = 6)
# Loading packages
library(dplyr)
library(tidyr)
library(ggplot2)
# Loading data
life_expectancy <- read.csv("datasets/UNdata.csv")
# Taking a look at the first few rows
head(life_expectancy)
# + editable=false deletable=false dc={"key": "11"} run_control={"frozen": true} tags=["context"]
# ## 2. Life expectancy of men vs. women by country
# <p>Let's manipulate the data to make our exploration easier. We will build the dataset for our first plot in which we will represent the average life expectancy of men and women across countries for the last period recorded in our data (2000-2005).</p>
# + dc={"key": "11"} tags=["sample_code"]
# Subsetting and reshaping the life expectancy data
subdata <- life_expectancy %>%
filter(Year == "2000-2005") %>%
select(Country.or.Area, Subgroup, Value) %>%
spread(Subgroup, Value)
# Taking a look at the first few rows
head(subdata)
# + editable=false deletable=false dc={"key": "18"} run_control={"frozen": true} tags=["context"]
# ## 3. Visualize I
# <p>A scatter plot is a useful way to visualize the relationship between two variables. It is a simple plot in which points are arranged on two axes, each of which represents one of those variables. </p>
# <p>Let's create a scatter plot using <code>ggplot2</code> to represent life expectancy of males (on the x-axis) against females (on the y-axis). We will create a straightforward plot in this task, without many details. We will take care of these kinds of things shortly.</p>
# + dc={"key": "18"} tags=["sample_code"]
# Plotting male and female life expectancy
ggplot(subdata, aes(x = Male, y = Female)) +
geom_point()
# + editable=false deletable=false dc={"key": "25"} run_control={"frozen": true} tags=["context"]
# ## 4. Reference lines I
# <p>A good plot must be easy to understand. There are many tools in <code>ggplot2</code> to achieve this goal and we will explore some of them now. Starting from the previous plot, let's set the same limits for both axes as well as place a diagonal line for reference. After doing this, the difference between men and women across countries will be easier to interpret.</p>
# <p>After completing this task, we will see how most of the points are arranged above the diagonal and how there is a significant dispersion among them. What does this all mean?</p>
# + dc={"key": "25"} tags=["sample_code"]
# Adding an abline and changing the scale of axes of the previous plots
ggplot(subdata, aes(x = Male, y = Female)) +
geom_point() +
geom_abline(intercept = 0, slope = 1, linetype = 2) +
scale_x_continuous(limits=c(35,85))+
scale_y_continuous(limits=c(35,85))
# + editable=false deletable=false dc={"key": "32"} run_control={"frozen": true} tags=["context"]
# ## 5. Plot titles and axis labels
# <p>A key point to make a plot understandable is placing clear labels on it. Let's add titles, axis labels, and a caption to refer to the source of data. Let's also change the appearance to make it clearer.</p>
# + dc={"key": "32"} tags=["sample_code"]
# Adding labels to previous plot
ggplot(subdata, aes(x=Male, y=Female))+
geom_point(colour="white", fill="chartreuse3", shape=21, alpha=.55, size=5)+
geom_abline(intercept = 0, slope = 1, linetype=2)+
scale_x_continuous(limits=c(35,85))+
scale_y_continuous(limits=c(35,85))+
labs(title="Life Expectancy at Birth by Country",
subtitle="Years. Period: 2000-2005. Average.",
caption="Source: United Nations Statistics Division",
x="Males",
y="Females")
# + editable=false deletable=false dc={"key": "39"} run_control={"frozen": true} tags=["context"]
# ## 6. Highlighting remarkable countries I
# <p>Now, we will label some points of our plot with the name of its corresponding country. We want to draw attention to some special countries where the gap in life expectancy between men and women is significantly high. These will be the final touches on this first plot.</p>
# + dc={"key": "39"} tags=["sample_code"]
# Subseting data to obtain countries of interest
top_male <- subdata %>% arrange(Male-Female) %>% head(3)
top_female <- subdata %>% arrange(Female-Male) %>% head(3)
# Adding text to the previous plot to label countries of interest
ggplot(subdata, aes(x=Male, y=Female, label=Country.or.Area))+
geom_point(colour="white", fill="chartreuse3", shape=21, alpha=.55, size=5)+
geom_abline(intercept = 0, slope = 1, linetype=2)+
scale_x_continuous(limits=c(35,85))+
scale_y_continuous(limits=c(35,85))+
labs(title="Life Expectancy at Birth by Country",
subtitle="Years. Period: 2000-2005. Average.",
caption="Source: United Nations Statistics Division",
x="Males",
y="Females") +
geom_text(data=top_male, size = 3) +
geom_text(data=top_female, size = 3) +
theme_bw()
# + editable=false deletable=false dc={"key": "46"} run_control={"frozen": true} tags=["context"]
# ## 7. How has life expectancy by gender evolved?
# <p>Since our data contains historical information, let's see now how life expectancy has evolved in recent years. Our second plot will represent the difference between men and women across countries between two periods: 2000-2005 and 1985-1990.</p>
# <p>Let's start building a dataset called <code>subdata2</code> for our second plot. </p>
# + dc={"key": "46"} tags=["sample_code"]
# Subsetting, mutating and reshaping the life expectancy data
subdata2 <- life_expectancy %>%
filter(Year %in% c("1985-1990", "2000-2005")) %>%
mutate(Sub_Year=paste(Subgroup, Year, sep="_")) %>%
mutate(Sub_Year=gsub("-", "_", Sub_Year)) %>%
select(-Subgroup, -Year) %>%
spread(Sub_Year, Value) %>%
mutate(
diff_Female = Female_2000_2005 - Female_1985_1990,
diff_Male = Male_2000_2005 - Male_1985_1990
)
# Taking a look at the first few rows
head(subdata2)
# + editable=false deletable=false dc={"key": "53"} run_control={"frozen": true} tags=["context"]
# ## 8. Visualize II
# <p>Now let's create our second plot in which we will represent average life expectancy differences between "1985-1990" and "2000-2005" for men and women.</p>
# + dc={"key": "53"} tags=["sample_code"]
# Doing a nice first version of the plot with abline, scaling axis and adding labels
ggplot(subdata2, aes(x=diff_Male, y=diff_Female, label=Country.or.Area))+
geom_point(colour="white", fill="chartreuse3", shape=21, alpha=.55, size=5)+
geom_abline(intercept = 0, slope = 1, linetype=2)+
scale_x_continuous(limits = c(-25, 25)) +
scale_y_continuous(limits = c(-25, 25)) +
labs(title="Life Expectancy at Birth by Country in Years",
subtitle="Difference between 1985-1990 and 2000-2005. Average.",
caption="Source: United Nations Statistics Division",
x="Males",
y="Females")+
theme_bw()
# + editable=false deletable=false dc={"key": "60"} run_control={"frozen": true} tags=["context"]
# ## 9. Reference lines II
# <p>Adding reference lines can make plots easier to understand. We already added a diagonal line to visualize differences between men and women more clearly. Now we will add two more lines to help to identify in which countries people increased or decreased their life expectancy in the period analyzed.</p>
# + dc={"key": "60"} tags=["sample_code"]
# Adding an hline and vline to previous plots
ggplot(subdata2, aes(x=diff_Male, y=diff_Female, label=Country.or.Area))+
geom_point(colour="white", fill="chartreuse3", shape=21, alpha=.55, size=5)+
geom_abline(intercept = 0, slope = 1, linetype=2)+
scale_x_continuous(limits=c(-25,25))+
scale_y_continuous(limits=c(-25,25))+
geom_hline(yintercept = 0, linetype = 2) +
geom_vline(xintercept = 0, linetype = 2) +
labs(title="Life Expectancy at Birth by Country",
subtitle="Years. Difference between 1985-1990 and 2000-2005. Average.",
caption="Source: United Nations Statistics Division",
x="Males",
y="Females")+
theme_bw()
# + editable=false deletable=false dc={"key": "67"} run_control={"frozen": true} tags=["context"]
# ## 10. Highlighting remarkable countries II
# <p>As we did in the first plot, let's label some points. Concretely, we will point those three where the aggregated average life expectancy for men and women increased most and those three where decreased most in the period.</p>
# + dc={"key": "67"} tags=["sample_code"]
# Subseting data to obtain countries of interest
top <- subdata2 %>% arrange(diff_Male+diff_Female) %>% head(3)
bottom <- subdata2 %>% arrange(-(diff_Male+diff_Female)) %>% head(3)
# Adding text to the previous plot to label countries of interest
ggplot(subdata2, aes(x=diff_Male, y=diff_Female, label=Country.or.Area), guide=FALSE)+
geom_point(colour="white", fill="chartreuse3", shape=21, alpha=.55, size=5)+
geom_abline(intercept = 0, slope = 1, linetype=2)+
scale_x_continuous(limits=c(-25,25))+
scale_y_continuous(limits=c(-25,25))+
geom_hline(yintercept=0, linetype=2)+
geom_vline(xintercept=0, linetype=2)+
labs(title="Life Expectancy at Birth by Country",
subtitle="Years. Difference between 1985-1990 and 2000-2005. Average.",
caption="Source: United Nations Statistics Division",
x="Males",
y="Females")+
geom_text(data=top, size=3)+
geom_text(data=bottom, size=3)+
theme_bw()
|
Visualizing Inequalities in Life Expectancy/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Swing Trade
# ### An automatic swing trading strategy implementation.
# ## Price Zone Definition and Transition
# <img src="state_flowchart.png" width=640>
#
# ## Directional Reversal (Swing)
#
# <img src="state_reversal.png" width=640>
#
#
# ## Grid-Based Oscillatory Trading
#
# The oscillatory order price is based on the last successful order price as well as the trailing price from the last highest/lowest
# order price, i.e.,
# $$P_{last\_order} - P_{last} \geq P_h \space\space and \space\space P_{last}-P_{lowest} \geq P_t \space,\space or$$
# $$P_{last} - P_{last\_order} \geq P_h \space\space and \space\space P_{highest}-P_{last} \geq P_t \ .$$
# The order quantity is determined by
# $$Q_{order} = m*Q_a + k*Q_{offset} $$
# where $m$ is the number of grids that the market traverses from $P_{last_order}$ to $P_{last}$, determined by
# $$m = \lfloor \lvert P_{last} - P_{last_order} \rvert /P_h \rfloor$$
# and $k$ is the oscillatory quantity offset coefficient, which increases by 1 when the accumulated profit in the current trading zone
# exceeds the pre-defined threshold $G_{k\_th}$ determined by
# $$G_{k\_th} = (P_H-P_L) * (N_{grids} * Q_a + k * Q_{offset}).$$
# <img src="grid_osc.png" width=640>
#
# ## Zone Switch
#
# Switching between the trading zones Net, Inc, Osc, Dec happens when market price crosses more than one grid of a neighbor zone,
# $$ p_{last} > P_{high\_bound} + P_h \space\space or \space\space p_{last} < P_{low\_bound} - P_h .$$
# <img src="zone_switch.png" width=640>
#
# ## State Transition Implementation
#
#
# <img src="state_transition.png" width=640>
#
# ## Adaptive Orders
#
# ### Order Modes from More Passive to More Aggressive:
# * PATIENT: order price is the limit price specified by user.
# * ACCELERATED: order price is the more favor one between the last price with 1 payup tick and the midpoint of bid/ask.
# * URGENT: order price is the less favor one between the last price with 1 payup tick and the midpoint of bid/ask.
# * PANIC: order price is the market price.
|
doc/swing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Nanodegree Engenheiro de Machine Learning
# ## Modelp de Avaliação e Validação
# ## Projeto 1: Estimando Preços dos Imóveis de Boston
#
# Bem-vindo ao primeiro projeto do Nanodegree de Engenheiro de Machine Learning! Neste Notebook, alguns templates de código estão sendo fornecidos para você, e você irá precisar implementar funcionalidades adicionais para completar este projeto com sucesso. Você não vai precisar modificar o código que foi incluído além do que está sendo pedido. Seções que começam com **'Implementação'** no cabeçalho indicam que o bloco de código seguinte vai exigir que você providencie funcionalidade adicional. Instruções serão fornecidas para cada seção e as especificidades da implementação são marcadas no bloco de código com o comando 'TODO'. Não esqueça de ler as instruções atentamente!
#
# Além do código implementado, haverá questões relacionadas com o projeto e sua implementação que você deve responder. Cada seção em que há uma questão para você responder, ela será precedida por **'Questão X'** no cabeçalho. Leia cada questão cuidadosamente e dê respostas completas no seguinte box de texto que contém **'Resposta: '**. O projeto enviado será avaliado com base nas respostas para cada uma das questões e a implementação que você nos forneceu.
#
# >**Nota:** Células de Código e de Markdown podem ser executadas utilizando o atalho de teclado **Shift + Enter**. Além disso, as células Markdown podem ser editadas ao clicar normalmente duas vezes na célula para entrar no modo de edição.
# ## Começando
# Neste projeto, você irá avaliar o desempenho e o poder de estimativa de um modelo que foi treinado e testado em dados coletados dos imóveis dos subúrbios de Boston, Massachusetts. Um modelo preparado para esses dados e visto como *bem ajustado* pode ser então utilizado para certas estimativas sobre um imóvel – em particular, seu valor monetário. Esse modelo seria de grande valor para alguém como um agente mobiliário, que poderia fazer uso dessas informações diariamente.
#
# O conjunto de dados para este projeto se origina do [repositório de Machine Learning da UCI](https://archive.ics.uci.edu/ml/datasets/Housing). Os dados de imóveis de Boston foram coletados em 1978 e cada uma das 489 entradas representa dados agregados sobre 14 atributos para imóveis de vários subúrbios de Boston. Para o propósito deste projeto, os passos de pré-processamento a seguir foram feitos para esse conjunto de dados:
# - 16 observações de dados possuem um valor `'MEDV'` de 50.0. Essas observações provavelmente contêm **valores ausentes ou censurados** e foram removidas.
# - 1 observação de dados tem um valor `'RM'` de 8.78. Essa observação pode ser considerada **aberrante** e foi removida.
# - Os atributos `'RM'`, `'LSTAT'`, `'PTRATIO'`, and `'MEDV'` são essenciais. O resto dos **atributos irrelevantes** foram excluídos.
# - O atributo `'MEDV'` foi **escalonado multiplicativamente** para considerar 35 anos de inflação de mercado.
#
# Execute a célula de código abaixo para carregar o conjunto dos dados dos imóveis de Boston, além de algumas bibliotecas de Python necessárias para este projeto. Você vai saber que o conjunto de dados carregou com sucesso se o seu tamanho for reportado.
# +
# Importar as bibliotecas necessárias para este projeto
import numpy as np
import pandas as pd
import visuals as vs # Supplementary code
from sklearn.cross_validation import ShuffleSplit
# Formatação mais bonita para os notebooks
# %matplotlib inline
# Executar o conjunto de dados de imóveis de Boston
data = pd.read_csv('housing.csv')
prices = data['MEDV']
features = data.drop('MEDV', axis = 1)
# Êxito
print "O conjunto de dados de imóveis de Boston tem {} pontos com {} variáveis em cada.".format(*data.shape)
# -
# ## Explorando os Dados
# Na primeira seção deste projeto, você fará uma rápida investigação sobre os dados de imóveis de Boston e fornecerá suas observações. Familiarizar-se com os dados durante o processo de exploração é uma prática fundamental que ajuda você a entender melhor e justificar seus resultados.
#
# Dado que o objetivo principal deste projeto é construir um modelo de trabalho que tem a capacidade de estimar valores dos imóveis, vamos precisar separar os conjuntos de dados em **atributos** e **variável alvo**. O **atributos**, `'RM'`, `'LSTAT'` e `'PTRATIO'`, nos dão informações quantitativas sobre cada ponto de dado. A **variável alvo**, `'MEDV'`, será a variável que procuramos estimar. Eles são armazenados em `features` e ` prices`, respectivamente.
# ### Implementação: Calcular Estatísticas
# Para a sua primeira implementação de código, você vai calcular estatísticas descritivas sobre preços dos imóveis de Boston. Dado que o `numpy` já foi importado para você, use essa biblioteca para executar os cálculos necessários. Essas estatísticas serão extremamente importantes depois para analisar várias estimativas resultantes do modelo construído.
#
# Na célula de código abaixo, você precisará implementar o seguinte:
# - Calcular o mínimo, o máximo, a média, a mediana e o desvio padrão do `'MEDV'`, que está armazenado em `prices`.
# - Armazenar cada cálculo em sua respectiva variável.
# +
# TODO: Preço mínimo dos dados
minimum_price = None
# TODO: Preço máximo dos dados
maximum_price = None
# TODO: Preço médio dos dados
mean_price = None
# TODO: Preço mediano dos dados
median_price = None
# TODO: Desvio padrão do preço dos dados
std_price = None
# Mostrar as estatísticas calculadas
print "Estatísticas para os dados dos imóveis de Boston:\n"
print "Preço mínimo: ${:,.2f}".format(minimum_price)
print "Preço máximo: ${:,.2f}".format(maximum_price)
print "Preço médio: ${:,.2f}".format(mean_price)
print "Preço mediano: ${:,.2f}".format(median_price)
print "Desvio padrão dos preços: ${:,.2f}".format(std_price)
# -
# ### Questão 1 - Observação de Atributos
# Para lembrar, estamos utilizando três atributos do conjunto de dados dos imóveis de Boston: 'RM'`, `'LSTAT'` e `'PTRATIO'`. Para cada observação de dados (vizinhança):
# - `'RM'` é o número médio de quartos entre os imóveis na vizinhança.
# - `'LSTAT'` é a porcentagem de proprietários na vizinhança considerados de "classe baixa" (proletariado).
# - `'PTRATIO'` é a razão de estudantes para professores nas escolas de ensino fundamental e médio na vizinhança.
#
# _Utilizando sua intuição, para cada um dos atributos acima, você acha que um aumento no seu valor poderia levar a um **aumento** no valor do `'MEDV'` ou uma **diminuição** do valor do `'MEDV'`? Justifique sua opinião para cada uma das opções._
# **Dica:** Você espera que um imóvel que tem um valor `'RM'` de 6 custe mais ou menos que um imóvel com valor `'RM'` de 7?
# **Resposta: **
# ----
#
# ## Desenvolvendo um Modelo
# Na segunda seção deste projeto, você vai desenvolver ferramentas e técnicas necessárias para um modelo que faz estimativas. Ser capaz de fazer avaliações precisas do desempenho de cada modelo através do uso dessas ferramentas e técnicas ajuda a reforçar a confiança que você tem em suas estimativas.
# ### Implementação: Definir uma Métrica de Desempenho
# É difícil medir a qualidade de um modelo dado sem quantificar seu desempenho durante o treinamento e teste. Isso é geralmente feito utilizando algum tipo de métrica de desempenho, através do cálculo de algum tipo de erro, qualidade de ajuste, ou qualquer outra medida útil. Para este projeto, você irá calcular o [*coeficiente de determinação*](https://pt.wikipedia.org/wiki/R%C2%B2), R<sup>2</sup>, para quantificar o desempenho do seu modelo. O coeficiente da determinação para um modelo é uma estatística útil em análise regressa, como se ele frequentemente descrevesse como "good" a capacidade do modelo de fazer estimativas.
#
# Os valores para R<sup>2</sup> têm um alcance de 0 a 1, que captura a porcentagem da correlação ao quadrado entre a estimativa e o valor atual da **variável alvo**. Um modelo R<sup>2</sup> de valor 0 sempre falha ao estimar a variável alvo, enquanto que um modelo R<sup>2</sup> de valor 1, estima perfeitamente a variável alvo. Qualquer valor entre 0 e 1 indica qual a porcentagem da variável alvo, ao utilizar esse modelo, ele pode ser explicado pelos **atributos**. *Um modelo pode dar também um R<sup>2</sup> negativo, que indica que o modelo não é melhor do que aquele que estima ingenuamente a média da variável alvo.*
#
# Para a função ‘performance_metric’ na célula de código abaixo, você irá precisar implementar o seguinte:
# - Utilizar o `r2_score` do `sklearn.metrics` para executar um cálculo de desempenho entre `y_true` e `y_predict`.
# - Atribuir a pontuação do desempenho para a variável `score`.
# +
# TODO: Importar 'r2_score'
def performance_metric(y_true, y_predict):
""" Calcular e retornar a pontuação de desempenho entre
valores reais e estimados baseado na métrica escolhida. """
# TODO: Calcular a pontuação de desempenho entre 'y_true' e 'y_predict'
score = None
# Devolver a pontuação
return score
# -
# ### Questão 2 - Qualidade do Ajuste
# Admita que um conjunto de dados que contém cinco observações de dados e um modelo fez a seguinte estimativa para a variável alvo:
#
# | Valores Reais | Estimativa |
# | :-------------: | :--------: |
# | 3.0 | 2.5 |
# | -0.5 | 0.0 |
# | 2.0 | 2.1 |
# | 7.0 | 7.8 |
# | 4.2 | 5.3 |
# *Você consideraria que esse modelo foi capaz de capturar a variação da variável alvo com sucesso? Por que ou por que não?*
#
# Executar a célula de código abaixo para usar a função `performance_metric’ e calcular o coeficiente de determinação desse modelo.
# Calcular o desempenho deste modelo
score = performance_metric([3, -0.5, 2, 7, 4.2], [2.5, 0.0, 2.1, 7.8, 5.3])
print "O coeficiente de determinação, R^2, do modelo é {:.3f}.".format(score)
# **Resposta:**
# ### Implementação: Misturar e Separar os Dados
# Sua próxima implementação exige que você pegue o conjunto de dados de imóveis de Boston e divida os dados em subconjuntos de treinamento e de teste. Geralmente os dados são também misturados em uma ordem aleatória ao criar os subconjuntos de treinamento e de teste para remover qualquer viés (ou erro sistemático) na ordenação do conjunto de dados.
#
# Para a célula de código abaixo, você vai precisar implementar o seguinte:
# - Utilize `train_test_split` do `sklearn.cross_validation` para misturar e dividir os dados de `features` e `prices` em conjuntos de treinamento e teste.
# - Dividq os dados em 80% treinamento e 20% teste.
# - Mude o `random_state` do `train_test_split` para um valor de sua escolha. Isso garante resultados consistentes.
# - Atribuir a divisão de treinamento e teste para X_train`, `X_test`, `y_train` e `y_test`.
# +
# TODO: Importar 'train_test_split'
# TODO: Misturar e separar os dados em conjuntos de treinamento e teste
X_train, X_test, y_train, y_test = (None, None, None, None)
# Êxito
print "Separação entre treino e teste feita com êxito."
# -
# ### Questão 3 - Treinamento e Teste
# *Qual o benefício de separar o conjunto de dados em alguma relação de subconjuntos de treinamento e de teste para um algoritmo de aprendizagem?*
# **Dica:** O que pode dar errado se não houver uma maneira de testar seu modelo?
# **Resposta: **
# ----
#
# ## Analisando o Modelo de Desempenho
# Na terceira parte deste projeto, você verá o desempenho em aprendizagem e teste de vários modelos em diversos subconjuntos de dados de treinamento. Além disso, você irá investigar um algoritmo em particular com um parâmetro `'max_depth'` (profundidade máxima) crescente, em todo o conjunto de treinamento, para observar como a complexidade do modelo afeta o desempenho. Plotar o desempenho do seu modelo baseado em critérios diversos pode ser benéfico no processo de análise, por exemplo: para visualizar algum comportamento que pode não ter sido aparente nos resultados sozinhos.
# ### Curvas de Aprendizagem
# A célula de código seguinte produz quatro gráficos para um modelo de árvore de decisão com diferentes níveis de profundidade máxima. Cada gráfico visualiza a curva de aprendizagem do modelo para ambos treinamento e teste, assim que o tamanho do conjunto treinamento aumenta. Note que a região sombreada da curva de aprendizagem denota a incerteza daquela curva (medida como o desvio padrão). O modelo é pontuado em ambos os conjuntos treinamento e teste utilizando R<sup>2</sup>, o coeficiente de determinação.
#
# Execute a célula de código abaixo e utilizar esses gráficos para responder as questões a seguir.
# Criar curvas de aprendizagem para tamanhos de conjunto de treinamento variável e profundidades máximas
vs.ModelLearning(features, prices)
# ### Questão 4 - Compreendendo os Dados
# *Escolha um dos gráficos acima e determine a profundidade máxima para o modelo. O que acontece com a pontuação da curva de treinamento se mais pontos de treinamento são adicionados? E o que acontece com a curva de teste? Ter mais pontos de treinamento beneficia o modelo?*
# **Dica:** As curvas de aprendizagem convergem para uma pontuação em particular?
# **Resposta: **
# ### Curvas de Complexidade
# A célula de código a seguir produz um gráfico para um modelo de árvore de decisão que foi treinada e validada nos dados de treinamento utilizando profundidades máximas diferentes. O gráfico produz duas curvas de complexidade – uma para o treinamento e uma para a validação. Como a **curva de aprendizagem**, a área sombreada de ambas as curvas de complexidade denota uma incerteza nessas curvas, e o modelo pontuou em ambos os conjuntos de treinamento e validação utilizando a função `performance_metric`.
#
# Execute a célula de código abaixo e utilize o gráfico para responder as duas questões a seguir.
vs.ModelComplexity(X_train, y_train)
# ### Questão 5 - Equilíbrio entre viés e variância
# *Quando o modelo é treinado com o profundidade máxima 1, será que o modelo sofre mais de viés (erro sistemático) ou variância (erro aleatório)? E o que acontece quando o modelo é treinado com profundidade máxima 10? Quais pistas visuais existem no gráfico para justificar suas conclusões?*
# **Dica:** Como você sabe que um modelo está experimentando viés alto ou variância alta?
# **Resposta: **
# ### Questão 6 - Modelo Ótimo de Melhor Suposição
# *Qual profundidade máxima (`'max_depth'`) você acredita que resulta em um modelo que melhor generaliza um dado desconhecido? Que intuição te levou a essa resposta?*
# **Resposta: **
# -----
#
# ## Avaliando o Desempenho do Modelo
# Nesta parte final do projeto, você irá construir um modelo e fazer uma estimativa de acordo com o conjunto de atributos do cliente utilizando um modelo otimizado a partir de `fit_model`.
# ### Questão 7 - Busca em Matriz
# *O que é a técnica de busca em matriz (*grid search*) e como ela pode ser aplicada para otimizar um algoritmo de aprendizagem?*
# **Resposta: **
# ### Questão 8 - Validação Cruzada
# *O que é a técnica de treinamento de validação-cruzada k-fold? Quais benefícios essa técnica proporciona para busca em matriz ao otimizar um modelo?*
# **Dica:** Assim como há um raciocínio por trás de utilizar um conjunto de teste, o que poderia dar errado ao utilizar busca em matriz sem um conjunto de validação cruzada?
# **Resposta: **
# ### Implementação: Ajustar um Modelo
# Na sua última implementação, você vai precisar unir tudo o que foi aprendido e treinar um modelo utilizando o **algoritmo de árvore de decisão**. Para garantir que você está produzindo um modelo otimizado, você treinará o modelo utilizando busca em matriz para otimizar o parâmetro de profundidade máxima (`'max_depth'`) para uma árvore de decisão. Esse parâmetro pode ser entendido como o número de perguntas que o algoritmo de árvore de decisão pode fazer sobre os dados antes de fazer uma estimativa. Árvores de decisão são parte de uma classe de algoritmos chamados *algoritmos de aprendizagem supervisionada*.
#
# Para a função `fit_model` na célula de código abaixo, você vai precisar implementar o seguinte:
# - Utilize o [`DecisionTreeRegressor`](http://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeRegressor.html) do `sklearn.tree` para gerar um objeto regressor de árvore de decisão.
# - Atribua esse objeto à variável `'regressor'`.
# - Gere um dicionário para `'max_depth'` com os valores de 1 a 10 e atribua isso para a variável `'params'`.
# - Utilize o [`make_scorer`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.make_scorer.html) do `sklearn.metrics` para gerar um objeto de função de pontuação.
# - Passe a função `performance_metric` como um parâmetro para esse objeto.
# - Atribua a função de pontuação à variável `'scoring_fnc'`.
# - Utilize o [`GridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.grid_search.GridSearchCV.html) do `sklearn.grid_search` para gerar um objeto de busca por matriz.
# - Passe as variáveis `'regressor'`, `'params'`, `'scoring_fnc'` and `'cv_sets'` como parâmetros para o objeto.
# - Atribua o objeto `GridSearchCV` para a variável `'grid'`.
# +
# TODO: Importar 'make_scorer', 'DecisionTreeRegressor' e 'GridSearchCV'
def fit_model(X, y):
""" Desempenhar busca em matriz sobre o parâmetro the 'max_depth' para uma
árvore de decisão de regressão treinada nos dados de entrada [X, y]. """
# Gerar conjuntos de validação-cruzada para o treinamento de dados
cv_sets = ShuffleSplit(X.shape[0], n_iter = 10, test_size = 0.20, random_state = 0)
# TODO: Gerar uma árvore de decisão de regressão de objeto
regressor = None
# TODO: Gerar um dicionário para o parâmetro 'max_depth' com um alcance de 1 a 10
params = {}
# TODO: Transformar 'performance_metric' em uma função de pontuação utilizando 'make_scorer'
scoring_fnc = None
# TODO: Gerar o objeto de busca em matriz
grid = None
# Ajustar o objeto de busca em matriz com os dados para calcular o modelo ótimo
grid = grid.fit(X, y)
# Devolver o modelo ótimo depois de realizar o ajuste dos dados
return grid.best_estimator_
# -
# ### Fazendo Estimativas
# Uma vez que o modelo foi treinado em conjunto de dados atribuído, ele agora pode ser utilizado para fazer estimativas em novos conjuntos de entrada de dados. No caso do *regressor da árvore de decisão*, o modelo aprendeu *quais são as melhores perguntas sobre a entrada de dados*, e pode responder com uma estimativa para a **variável alvo**. Você pode utilizar essas estimativas para conseguir informações sobre os dados dos quais o valor da variável alvo é desconhecida – por exemplo, os dados dos quais o modelo não foi treinado.
# ### Questão 9 - Modelo Ótimo
# _Qual profundidade máxima do modelo ótimo? Como esse resultado se compara com a sua suposição na **Questão 6**?
#
# Executar a célula de código abaixo para ajustar o regressor da árvore de decisão com os dados de treinamento e gerar um modelo ótimo.
# +
# Ajustar os dados de treinamento para o modelo utilizando busca em matriz
reg = fit_model(X_train, y_train)
# Produzir valores para 'max_depth'
print "O parâmetro 'max_depth' é {} para o modelo ótimo.".format(reg.get_params()['max_depth'])
# -
# **Resposta: **
# ### Questão 10 -Estimando Preços de Venda
# Imagine que você era um corretor imobiliário na região de Boston ansioso para utilizar esse modelo que ajuda os imóveis que seus clientes desejam vender. Você coletou as seguintes informações de três dos seus clientes:
#
# | Atributos | Cliente 1 | Cliente 2 | Cliente 3 |
# | :---: | :---: | :---: | :---: |
# | Número total de quartos em um imóvel | 5 quartos | 4 quartos | 8 quartos |
# | Nível de pobreza da vizinhança (em %) | 17% | 32% | 3% |
# | Razão estudante:professor das escolas próximas | 15-to-1 | 22-to-1 | 12-to-1 |
# *Qual valor você sugeriria para cada um dos seus clientes para a venda de suas casas? Esses preços parecem razoáveis dados os valores para cada atributo?*
# **Dica:** Utilize as estatísticas que você calculou na seção **Explorando Dados** para ajudar a justificar sua resposta.
#
# Execute a célula de códigos abaixo para que seu modelo otimizado faça estimativas para o imóvel de cada um dos clientes.
# +
# Gerar uma matriz para os dados do cliente
client_data = [[5, 17, 15], # Cliente 1
[4, 32, 22], # Cliente 2
[8, 3, 12]] # Cliente 3
# Mostrar estimativas
for i, price in enumerate(reg.predict(client_data)):
print "Preço estimado para a casa do cliente {}: ${:,.2f}".format(i+1, price)
# -
# **Resposta: **
# ### Sensibilidade
# Um modelo ótimo não é necessariamente um modelo robusto. Às vezes, um modelo é muito complexo ou muito simples para generalizar os novos dados. Às vezes, o modelo pode utilizar um algoritmo de aprendizagem que não é apropriado para a estrutura de dados especificado. Outras vezes, os próprios dados podem ter informação excessiva ou exemplos insuficientes para permitir que o modelo apreenda a variável alvo – ou seja, o modelo não pode ser ajustado. Execute a célula de código abaixo para rodar a função `fit_model` dez vezes com diferentes conjuntos de treinamento e teste para ver como as estimativas para um cliente específico mudam se os dados foram treinados.
vs.PredictTrials(features, prices, fit_model, client_data)
# ### Questão 11 - Aplicabilidade
# *Em poucas linhas, argumente se o modelo construído deve ou não ser utilizado de acordo com as configurações do mundo real.*
# **Dica:** Algumas questões para responder:
# - *Quão relevante dados coletados em 1978 podem ser nos dias de hoje?*
# - *Os atributos presentes são suficientes para descrever um imóvel?*
# - *Esse modelo é robusto o suficiente para fazer estimativas consistentes?*
# - *Dados coletados em uma cidade urbana como Boston podem ser aplicados para uma cidade rural?*
# **Resposta: **
|
resources/boston_housing/boston_housing_PT.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Intro
#
# EGG provides multiple components to create different types of games. In particular, EGG has strong tooling for creating one-step Sender/Receiver games. In these games, two agents, Sender and Receiver, are trained to perform a common task while following a simple communication protocol:
#
# * Sender receives an input (e.g., an image) and sends a single message to Receiver;
# * Receiver obtains the message and (optionally) its own input, produces an output.
#
# This type of games includes, for instance, signaling games and discrete auto-encoders. In a nutshell, any Sender/Receiver game only differs in terms of (a) input data, (b) agents' architecture, (c) communication type (one-symbol, fixed-lentgh or variable-length multiple-symbol messages), (d) loss. EGG allows us to create new games by only specifying those components.
#
# In most cases, training of agents with discrete channel communication is done either via Gumbel-Softmax relaxation or by means of Reinforce. EGG allows to switch between the two with minimal changes in the logic.
#
#
#
# ## MNIST Auto-encoder game
#
# In this tutorial we will create a conceptually simple, but fully-featured Sender/Receiver game. In this game, Sender and Receiver are trained together such that Sender tries to encode a MNIST digit in a discrete message and Receiver will try to decode it from the message.
#
# While implementing this game, we will walk through several steps, typical for creating a new game:
# * pre-train a _vision_ module for the Sender agent by training it to classify digits;
# * use the pre-trained vision module to implement Sender and Receiver that communicate via a single symbol messages and analyse the trained models;
# * update the agents to allow variable-length multi-symbol messages.
#
# NB: depending on the computational resources you have available, this might be slow to run. To get at least a sense of the full pipeline, consider reducing the number of training epochs, or, alternatively, run a Google Colab version of the tutorial using a GPU.
#
# So let's start!
# if you are running this notebook via Google Colab, you have to install EGG first
# !pip install git+https://github.com/facebookresearch/EGG.git
# also you'll need to change the runtime to GPU (Runtime -> Change runtime type -> Hardware Accelator -> GPU)
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import torch
import torch.nn as nn
import egg.core as core
from torchvision import datasets, transforms
from torch import nn
from torch.nn import functional as F
import matplotlib.pyplot as plt
import random
import numpy as np
from pylab import rcParams
rcParams['figure.figsize'] = 5, 10
# For convenince and reproducibility, we set some EGG-level command line arguments here
opts = core.init(params=['--random_seed=7', # will initialize numpy, torch, and python RNGs
'--lr=1e-3', # sets the learning rate for the selected optimizer
'--batch_size=32',
'--optimizer=adam'])
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# -
# In most cases, we want to start a script by initializing EGG by calling `core.init` Typically, we use command-line arguments for that (see https://github.com/fairinternal/EGG/blob/master/egg/core/CL.md for more details)
# Next, we implement our ```Vision``` module (a part of a standard MNIST [model](https://github.com/pytorch/examples/blob/master/mnist/main.py)) that maps a MNIST image into a 500-dimensional vector.
# To pre-train this ```Vision``` module, we'll use the auxilary task of classifying MNIST digits. For this, we also define a ```PretrainNet``` model that takes the output of ```Vision``` and classifies images into 10 classes (0, 1, ..., 9).
# +
class Vision(nn.Module):
def __init__(self):
super(Vision, self).__init__()
self.conv1 = nn.Conv2d(1, 20, 5, 1)
self.conv2 = nn.Conv2d(20, 50, 5, 1)
self.fc1 = nn.Linear(4*4*50, 500)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2, 2)
x = x.view(-1, 4*4*50)
x = F.relu(self.fc1(x))
return x
class PretrainNet(nn.Module):
def __init__(self, vision_module):
super(PretrainNet, self).__init__()
self.vision_module = vision_module
self.fc = nn.Linear(500, 10)
def forward(self, x):
x = self.vision_module(x)
x = self.fc(F.leaky_relu(x))
return x
# -
# We also need DataLoaders for MNIST....
# +
kwargs = {'num_workers': 1, 'pin_memory': True} if torch.cuda.is_available() else {}
transform = transforms.ToTensor()
batch_size = opts.batch_size # set via the CL arguments above
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('./data', train=True, download=True,
transform=transform),
batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('./data', train=False, transform=transform),
batch_size=batch_size, shuffle=False, **kwargs)
# -
# Now, everything is ready to actually run the pre-training. First, we instantiate the modules and then run 10 epochs of MNIST digit recognition by a PretrainNet instance:
vision = Vision()
class_prediction = PretrainNet(vision) # note that we pass vision - which we want to pretrain
optimizer = core.build_optimizer(class_prediction.parameters()) # uses command-line parameters we passed to core.init
class_prediction = class_prediction.to(device)
# Now we can run very typical Pytorch training loop to pretrain the vision module:
for epoch in range(10):
mean_loss, n_batches = 0, 0
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = class_prediction(data)
loss = F.cross_entropy(output, target)
loss.backward()
optimizer.step()
mean_loss += loss.mean().item()
n_batches += 1
print(f'Train Epoch: {epoch}, mean loss: {mean_loss / n_batches}')
# Awesome! Now ```vision``` is a pre-trained ```Vision``` module and we can proceed to actually implement the communication game.
# ## Single-symbol communication
#
# As said before, Sender/Receiver games only differ in the data used, the architecture of the agents, the type of the channel, and the loss.
# We will start with the simplest communication case, one-symbol communication.
#
#
# Let's define our Sender (that will re-use the pre-trained ```vision``` module) and Receiver.
# For implementing single-symbol games, EGG provides special wrappers, ```GumbelSoftmaxWrapper``` and ```ReinforceWrapper``` that wrap an agent and implement the required Gumbel-Softmax or Reinforce-specific logic. Both wrappers assume that the agent returns log-probabilities over all terms for the vocabulary.
#
# We start by defining the logic of our agents:
# +
class Sender(nn.Module):
def __init__(self, vision, output_size):
super(Sender, self).__init__()
self.fc = nn.Linear(500, output_size)
self.vision = vision
def forward(self, x, aux_input=None):
with torch.no_grad():
x = self.vision(x)
x = self.fc(x)
return x
class Receiver(nn.Module):
def __init__(self, input_size):
super(Receiver, self).__init__()
self.fc = nn.Linear(input_size, 784)
def forward(self, channel_input, receiver_input=None, aux_input=None):
x = self.fc(channel_input)
return torch.sigmoid(x)
sender = Sender(vision, output_size=400)
receiver = Receiver(input_size=400)
# -
# While ```GumbelSoftmaxWrapper``` and ```ReinforceWrapper``` have the same interface to the user's defined Sender and Receiver, they do differ in their output.
#
# Since ```GumbelSoftmaxWrapper``` uses relaxation of the symbols, it doesn't output the index of the used symbol; instead, it outputs a tensor of dimensions (batch size; vocab size) with potentially all elements being non-zero. In evaluation mode, it outputs a tensor of the same dimensionality, but with the one-hot encoded symbol of highest probability. In contrast, ```ReinforceWrapper``` directly outputs the index of the transmitted symbol (plus other data to be discussed below).
#
# To make the change of wrappers simpler, we define a special helper layer, `core.RelaxedEmbedding` which behaves as a standard `torch.nn.Embedding` layer if the inputs are indexes (i.e. are of type torch.long), or provides their relaxation, if they are a tensor of floats. You don't have to use this, but it makes changing the training from GS-based to RF-based and vice-versa very simple, because you do not need to worry about the type of the input that the Receiver will get.
#
# Furthermore, you can also wrap your Receiver agent logic in the `core.SymbolReceiverWrapper`. This wrapper would embed the input symbol via the `core.RelaxedEmbedding` and pass the resulting embedding to the user-implement agent.
#
# (NB: `core.RelaxedEmbedding` has the same initialization as `torch.nn.Embedding` - different from that of `nn.Linear`!)
# Apart from specifying the agents' architecture, we have to define the loss that the agents experience. By default, EGG expects the loss function to take the following arguments: Sender's input, the message transmitted, the input of the receiver, the output of the receiver, and the labels provided from the dataset.
# Any of those can be used or ignored. In the case of our MNIST auto-encoder game, there is no specific input for Receiver, so the game mechanics passes `None` instead.
def loss(sender_input, _message, _receiver_input, receiver_output, _labels, _aux_input=None):
loss = F.binary_cross_entropy(receiver_output, sender_input.view(-1, 784), reduction='none').mean(dim=1)
return loss, {}
# Note that this loss function is differentiable; it doesn't have to be differentiable if the training is done via Reinforce. However, if it is the case, the training of Receiver might be able to exploit that. See the [readme](https://github.com/facebookresearch/EGG/blob/master/README.md#an-important-technical-point) for more discussion.
#
# Another common parameter that would be shared across all the games we'll implement is the size of the vocabulary. We set it to 10 as MNIST has 10 digits; although it might be fun to play with other values.
vocab_size = 10
# ### Training with Gumbel-Softmax
#
# The next step is to actually create the agents. We first discuss how training can be done with Gumbel-Softmax. For that, we instantiate our Sender and Receiver and wrap Sender into `core.GumbelSoftmaxWrapper`.
sender = Sender(vision, vocab_size)
sender = core.GumbelSoftmaxWrapper(sender, temperature=1.0) # wrapping into a GS interface, requires GS temperature
receiver = Receiver(input_size=400)
receiver = core.SymbolReceiverWrapper(receiver, vocab_size, agent_input_size=400)
# Once we have the agents created, we can adopt a pre-defined game logic which juggles the agents, messages,
# data, and the loss. Essentially, the game instance is also a `torch.nn.Module` - hence it can be handled as that, i.e.
# we can extract the trainable parameters of the agents, push it around devices, etc. Each game is expected to output a two-tuple. The first element of the tuple is a loss that is to be minimized. The second is a python dict `{}` of some auxilary values that are averaged over the epoch and printed. Accuracy or entropy of the communication channel are good examples of such auxilary data.
#
#
# Another entity that we will need is `core.Trainer`. Essentially, `Trainer` implements the training loop, handles checkpointing, etc.
#
#
# The Gumbel-Softmax distribution is parameterised by its temperature. In general, the latter has strong impact on training. To anneal the temperature of Sender, we create a callback that `trainer` invokes after each training epoch is over. In this snippet, the callback applies a multiplicative decay rule on the temperature value.
# +
game = core.SymbolGameGS(sender, receiver, loss)
optimizer = torch.optim.Adam(game.parameters())
trainer = core.Trainer(
game=game, optimizer=optimizer, train_data=train_loader,
validation_data=test_loader, callbacks=[core.TemperatureUpdater(agent=sender, decay=0.9, minimum=0.1)]
)
# -
# Yay, everything is ready for actual training!
n_epochs = 15
trainer.train(n_epochs)
# Now when Sender and Receiver are trained, we firstly have a look at the codebook they came up with: what does Receiver output when given a particular word from a vocabulary? For that, we'll take every word in the vocabulary [0..9], wrap it into a tensor and pass it to Receiver:
# +
game.eval()
for z in range(vocab_size):
t = torch.zeros(vocab_size).to(device)
t[z] = 1
with torch.no_grad():
# Receiver outputs a single tensor of predictions
sample = game.receiver(t).float().cpu()
sample = sample.view(28, 28)
plt.title(f"Input: symbol {z}")
plt.imshow(sample, cmap='gray')
plt.show()
# -
# Ok, not too bad: some digits are vague, but others do resemble actual digits. Huh!
#
#
# Next, how good are our two agents at auto-encoding the images? Let's take some images from the validation set, feed them to Sender and look at (a) what messages are sent, (b) how does Receiver's output look like?
#
# First, we define a tiny, single-batch 10-digit test dataset:
# +
test_inputs = []
for z in range(10):
index = (test_loader.dataset.targets[:100] == z).nonzero()[0, 0]
img, _ = test_loader.dataset[index]
test_inputs.append(img.unsqueeze(0))
test_inputs = torch.cat(test_inputs)
test_dataset = [[test_inputs, None]]
# -
# Next, we write a small helper function to dump and plot the input-outputs of the agents along with the communicated messages. We will use it throughout the tutorial.
def plot(game, test_dataset, is_gs, variable_length):
interaction = \
core.dump_interactions(game, test_dataset, is_gs, variable_length)
for z in range(10):
src = interaction.sender_input[z].squeeze(0)
dst = interaction.receiver_output[z].view(28, 28)
# we'll plot two images side-by-side: the original (left) and the reconstruction
image = torch.cat([src, dst], dim=1).cpu().numpy()
plt.title(f"Input: digit {z}, channel message {interaction.message[z]}")
plt.imshow(image, cmap='gray')
plt.show()
# Here the left image is the original and the one on the right is the recovered one, each pair is labelled by the original class and the symbol sent over the channel.
plot(game, test_dataset, is_gs=True, variable_length=False)
# Again, not totally unreasonable!
#
# ### Training with Reinforce
# What's the equivalent code when training by Reinforce?
#
# We create exactly the same Sender instance, but then feed it into ```core.ReinforceWrapper```. This wrapper implements the boilerplate code required for Reinforce-based optimization: samples from a distribution, records the log-probability of what was sampled, and reports the entropy of the distribution (*).
#
# Hence, each agent returns a tuple of three elements: the actual sampled output, the log-probability of the sampled output, and the entropy of the sampling distribution. During the evaluation, the sampling process is replaced by taking the most likely output.
#
# But what if an agent (e.g. Receiver) runs a deterministic function? In that case, we use a simple `core.ReinforceDeterministicWrapper` that appends zero entropy and log-probability tensors. `ReinforceDeterministicWrapper` can only be used if Receiver's parameters can be optimized via a gradient-based optimization, i.e. the loss function has to be differentiable.
#
#
#
# (*) Entropy regularization is used to encourage the agents to do some exploration.
# +
sender = Sender(vision, output_size=vocab_size)
sender = core.ReinforceWrapper(sender) # wrapping into a Reinforce interface
receiver = Receiver(input_size=400)
receiver = core.SymbolReceiverWrapper(receiver, vocab_size, agent_input_size=400)
receiver = core.ReinforceDeterministicWrapper(receiver)
# -
# This time we use the one-symbol game implementation with Reinforce logic, `core.SymbolGameReinforce`. It also consumes two entropy regularization terms, for Sender and Receiver. The entropies of the output distributions of the agents are weighted by these parameters and added to the loss.
#
# Note that we managed to change from Gumbel Softmax to Reinforce training by only (a) changing the agent wrappers, (b) changing the game type.
# +
game = core.SymbolGameReinforce(sender, receiver, loss, sender_entropy_coeff=0.05, receiver_entropy_coeff=0.0)
optimizer = torch.optim.Adam(game.parameters(), lr=1e-2) # we can also use a manually set optimizer
trainer = core.Trainer(game=game, optimizer=optimizer, train_data=train_loader,
validation_data=test_loader)
# -
n_epochs = 15
trainer.train(n_epochs)
# Now let's check what is in the communication code book and the quality of the auto-encoding. The dumping code is essentially the same as before, the only change is that Receiver returns a tuple of three elements, and we need only one.
# +
game.eval()
for z in range(vocab_size):
t = torch.zeros(vocab_size).to(device)
t[z] = 1
with torch.no_grad():
sample, _1, _2 = game.receiver(t)
sample = sample.float().cpu()
sample = sample.view(28, 28)
plt.title(f"Input: symbol {z}")
plt.imshow(sample, cmap='gray')
plt.show()
# -
plot(game, test_dataset, is_gs=False, variable_length=False)
# Not too bad either! Let us move to a more interesting case: variable-length messages.
# ## Communication with variable-length messages
#
# This time, communication is carried out by RNNs. EGG provides pre-implemented code to use vanilla RNNs, GRUs, or LSTMs. There is also an experimental support for communication via Transformer-based blocks.
#
# Note that, when communicating over variable-length sequences, we need to define a special end-of-sequence symbol. The agents are trained such that the loss is calculated wrt to the Receiver output either after receiving the end-of-sequence symbol or after the maximal number of symbols were produced. EGG handles those scenarios, by defining special wrappers for agents, RnnSenderGS and RnnReceiverGS for Gumbel Softmax-based learning, and RnnSenderReinforce and RnnReceiverReinforce for Reinforce-based learning.
#
# By convention, the user-implemented Sender, after processing its input, outputs the initial hidden state vector for its RNN. EGG unrolls this RNN to produce a message. In turn, this message is fed into Receiver's RNN. The only thing that we need to implement for Receiver is how we map the Receiver RNN's hidden state to the Receiver's output.
#
# **Important points concerning end-of-sequence behaviour: (1) EGG assumes that the end-of-sequence symbol is always 0. (2) `vocab_size` always includes the end-of-sequence symbol - hence `vocab_size=2` specifies the unary encoding. (3) Communication stops immediately if the first symbol emitted by Sender is 0.**
#
# This time we'll have to specify some additional parameters:
# we start by defining the RNN parameters
hidden_size = 20
emb_size = 10
# We will use the same pre-trained vision module. Note, this time **Sender does not output a vector which will be casted as logits of probabilities over the vocab - it only outputs the initial hidden state for its RNN cell**!
# The rest is pretty much the same as before: we instantiate the instances and put them into wrappers (note that we set the RNN cell types to 'rnn' and maximal communication length `max_len=2`!)
#
# First, we'll go through Gumbel Softmax-based training.
#
#
# ### Variable-length messages with Gumbel Softmax
# +
sender = Sender(vision, hidden_size)
receiver = Receiver(hidden_size)
sender_rnn = core.RnnSenderGS(sender, vocab_size, emb_size, hidden_size,
cell="gru", max_len=2, temperature=1.0)
receiver_rnn = core.RnnReceiverGS(receiver, vocab_size, emb_size,
hidden_size, cell="gru")
game_rnn = core.SenderReceiverRnnGS(sender_rnn, receiver_rnn, loss)
# -
# Note that sometimes specifying different learning speeds for the two agents aids learning; this can be done as follows
optimizer = torch.optim.Adam([
{'params': game_rnn.sender.parameters(), 'lr': 1e-3},
{'params': game_rnn.receiver.parameters(), 'lr': 1e-2}
])
# Now we can train the agents
trainer = core.Trainer(game=game_rnn, optimizer=optimizer, train_data=train_loader,
validation_data=test_loader)
trainer.train(15)
# Heh, the validation loss is lower than it used to be with one symbol communication. Let's see how it works. Again, the original image is on the left, the transmitted and decoded - on the right. The communicated message is in the title. Note that since default `vocab_size` is 10 and 0 is used for end-of-sequence, Sender has an effective vocabulary of 9 symbols.
#
plot(game_rnn, test_dataset, is_gs=True, variable_length=True)
# Seems to work better than with length-one!
#
# We can also enumerate all possible 10x10 messages of length two {x = first symbol, y = second symbol} and check the Receiver's interpretation of the code, as shown next. NB: When x=0, the second symbol is ignored, as 0 is interpreted as end-of-sequence. For this reason, in this case Receiver generates the same image irrespective of y's value.
# +
f, ax = plt.subplots(10, 10, sharex=True, sharey=True)
for x in range(10):
for y in range(10):
t = torch.zeros((1, 2, vocab_size)).to(device)
t[0, 0, x] = 1
t[0, 1, y] = 1
with torch.no_grad():
sample = game_rnn.receiver(t).float().cpu()
# 0 is the end-of-sequence symbol, hence we stop immediately when x == 0
output_index = (0 if x == 0 else 1)
sample = sample[0, output_index, :].view(28, 28)
ax[x][y].imshow(sample, cmap='gray')
if y == 0:
ax[x][y].set_ylabel(f'x={x}')
if x == 0:
ax[x][y].set_title(f'y={y}')
ax[x][y].set_yticklabels([])
ax[x][y].set_xticklabels([])
plt.show()
# -
# For completeness, we re-iterate by training with Reinforce.
#
# ### Variable-length messages with Reinforce
# +
sender = Sender(vision, hidden_size)
receiver = Receiver(hidden_size)
sender_rnn = core.RnnSenderReinforce(sender, vocab_size, emb_size, hidden_size,
cell="gru", max_len=2)
receiver_rnn = core.RnnReceiverDeterministic(receiver, vocab_size, emb_size,
hidden_size, cell="gru")
game_rnn = core.SenderReceiverRnnReinforce(sender_rnn, receiver_rnn, loss,
sender_entropy_coeff=0.015,
receiver_entropy_coeff=0.0)
# -
optimizer = torch.optim.Adam([
{'params': game_rnn.sender.parameters(), 'lr': 1e-3},
{'params': game_rnn.receiver.parameters(), 'lr': 1e-2}
])
trainer = core.Trainer(game=game_rnn, optimizer=optimizer, train_data=train_loader,
validation_data=test_loader)
trainer.train(15)
# Now we can check how the auto-encoding is done and the emerging protocol.
#
plot(game_rnn, test_dataset, is_gs=False, variable_length=True)
# +
f, ax = plt.subplots(10, 10, sharex=True, sharey=True)
for x in range(10):
for y in range(10):
t = torch.zeros((1, 2)).to(device).long()
t[0, 0] = x
t[0, 1] = y
with torch.no_grad():
sample = game_rnn.receiver(t)[0].float().cpu()
sample = sample[0, :].view(28, 28)
ax[x][y].imshow(sample, cmap='gray')
if y == 0:
ax[x][y].set_ylabel(f'x={x}')
if x == 0:
ax[x][y].set_title(f'y={y}')
ax[x][y].set_yticklabels([])
ax[x][y].set_xticklabels([])
plt.show()
# -
# Not surprisingly, the results are different.
# Sometimes (eg `x=3`, `x=7`) it looks like the first symbol in the message (x) encodes which digit Receiver should reproduce, while the second (y) has more effect on the style of the digit. But is it true? How would we check that? 🤔
#
# ## (advanced under the hood stuff, may be skipped): optimization with stochastic nodes via Reinforce
#
# When we apply the differentiable relaxation of the communication channel by replacing the discrete messages with their Gumbel Softmax approximations, we make the entire Sender->Receiver pipeline differentiable. In contrast, Reinforce-based training doesn't make any relaxations. Instead, it effectively "smoothes" the channel by operating with a randomized distribution of messages, parameterised by Sender's output distribution.
#
# However, things might get more tricky with Reinforce-based training. In particular, it can happen that the loss on the Receiver's side is differentiable (as in our MNIST example above), and, in this case, we can train Receiver using the standard backprop. It might also happen that the loss is not differentiable (imagine 0-1 accuracy loss). In the latter case, Receiver also has to randomise its outputs.
#
# *EGG handles both cases with the same API*, however, it can be useful to know how it operates internally.
#
# Now, let's have a closer look at the optimisation problem. By $\theta_s$ and $\theta_r$ we denote the parameters of Sender and Receiver, respectively; $m$ is the message sent, and it comes from Sender's output distribution $P(m | i, \theta_s)$; $R(m, \theta_r)$ is Receiver's output, and $i$ is Sender's input.
#
# ### Deterministic Receiver
# When Receiver is deterministic, we want to minimize the following aggregate loss for an input $i$:
#
# $$L = \mathbb{E}_{m \sim P(m | i, \theta_s)}~ l(R(m, \theta_r), m, i)$$
#
# The gradient wrt to ${\theta_r}$ would be:
#
# $$\nabla_{\theta_r} L = \nabla_{\theta_r}\mathbb{E}_{m \sim P(m | i, \theta_s)}~ l(R(m, \theta_r), m, i) = \mathbb{E}_{m \sim P(m | i, \theta_s)}~ \nabla_{\theta_r} l(R(m, \theta_r), m, i) = \mathbb{E}_{m \sim P(m | i, \theta_s)} \frac{\partial}{\partial r } l(r, m, i) \cdot \nabla_{\theta_r} R(m, \theta_r)
# $$
#
# We can estimate it by sampling $N$ messages from Sender:
# $$\frac{1}{N} \sum_m \frac{\partial}{\partial r } l(r, m, i) \cdot \nabla_{\theta_r} R(m, \theta_r)
# $$
# Note that this gradient of $\theta_r$ is obtained by standard back-propogation from the loss with the messages sampled from Sender. By using the Reinforce trick, we estimate the second gradient by sampling $N$ messages from $P(m | i, \theta_s)$:
#
# $$\nabla_{\theta_s} L = \frac{1}{N} \sum_m l(R(m, \theta_r), m, i) \nabla_{\theta_s} \log P(m | i, \theta_s) $$
#
# **In this case, in Pytorch the gradients of both agents can be obtained by differentiating the sum**
#
# ```(l.mean() + (l.detach() * log_prob_sender).mean()).backwards()```
#
# (note: EGG also utilizes the mean baseline to reduce variance of the gradient estimate, but we omit that here for simplicity).
#
#
# ### Stochastic Receiver
# When the loss $l$ is not differentiable, and we sample output $o$ from Receiver $o \sim P(o | m, \theta_r)$, the expected loss becomes:
#
# $$L = \mathbb{E}_{m, o \sim P(m, o | i, \theta_s, \theta_r)} ~ l(o, m, i)$$
#
# The gradient wrt $\theta_r$ would be
# $$\nabla_{\theta_r} L = \mathbb{E}_{m, o \sim P(m, o | i, \theta_s, \theta_r)} l(o, m, i) ~ \nabla_{\theta_r} log P(m, o | i, \theta_s, \theta_r) = \mathbb{E}_{m, o \sim P(m, o | i, \theta_s, \theta_r)} l(o, m, i) ~ \nabla_{\theta_r} log P(o | m, \theta_r)$$
#
# By sampling $N$ $(m, o)$ pairs, we get
# $$\nabla_{\theta_r} L \approx \frac{1}{N} \sum_{o, m} l(o, m, i) ~ \nabla_{\theta_r} log P(o | m, \theta_r)$$
#
# and similarly,
# $$\nabla_{\theta_s} L \approx \frac{1}{N} \sum_{o, m} l(o, m, i) ~ \nabla_{\theta_s} log P(m | i, \theta_s)$$
#
#
# **In this case, in Pytorch the gradients of both agents can be obtained by differentiating the sum**
#
# ```(l.detach() * (log_prob_sender + log_prob_receiver).mean()).backwards()```
#
#
# ### Putting it all together
#
# Now, we can notice that if `l` is not differentiable, the gradient of `l.mean()` would be zero. Conversely, if Receiver is deterministic, `log_prob_receiver` would be the zero constant, hence its gradient is zero, too.
#
# Thus, both cases can be covered by minimizing the loss:
#
# ```(l.mean() + (l.detach() * (log_prob_sender + log_prob_receiver).mean())).backwards()```
#
# And that is what EGG does internally. Depending on whether Receiver is wrapped in `ReinforceDeterministicWrapper`/`RnnReceiverDeterministic` or in `ReinforceWrapper`/`RnnReceiverReinforce` defines which part of the loss would be effective for training Receiver.
#
#
# Overall, this makes it hard to implement a general Reinforce interface for all possible losses. Some involved cases might require re-implementing game mechanics or resorting to Gumbel Softmax relaxation, which doesn't have this problem.
|
tutorials/EGG walkthrough with a MNIST autoencoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# You have to change the path in the next code block
# + pycharm={"name": "#%%\n"}
import os
os.chdir(os.path.expanduser('~/programming/python/phyper/example'))
# +
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from sklearn.datasets import load_iris
import h5py
import numpy as np
from config import instances, Instance
from dataset import Iris
from paths import get_cross_validation_scores_path, get_training_metrics_path
from IPython.display import display
from paths import jupyter_plot
# orca has been installed with `brew cask install orca`
import plotly
plotly.io.orca.config.executable = '/usr/local/Caskroom/orca/1.3.1/orca.app/Contents/MacOS/orca'
# + pycharm={"name": "#%%\n"}
data = load_iris()
df = pd.DataFrame(data=data.data, columns=data.feature_names)
df['target'] = data.target
# print(df)
# + pycharm={"name": "#%%\n"}
from score_models import get_last_epoch
all_scores = pd.DataFrame(columns=['instance_hash', 'training_loss', 'training_accuracy', 'validation_loss', 'validation_accuracy'])
rows = []
for instance in instances:
path = get_training_metrics_path(instance)
with h5py.File(path, 'r') as f5:
keys = f5.keys()
last_epoch = get_last_epoch(keys)
metrics = f5[f'epoch{last_epoch}']
training_loss = metrics['training_loss']
training_accuracy = metrics['training_accuracy']
validation_loss = metrics['validation_loss']
validation_accuracy = metrics['validation_accuracy']
rows.append({'instance_hash': instance.get_instance_hash(),
'training_loss': metrics['training_loss'][...].item(),
'training_accuracy': metrics['training_accuracy'][...].item(),
'validation_loss': metrics['validation_loss'][...].item(),
'validation_accuracy': metrics['validation_accuracy'][...].item()})
all_scores = all_scores.append(rows, ignore_index=True)
all_scores.sort_values(by=['instance_hash'], inplace=True)
cv_scores = pd.read_csv(get_cross_validation_scores_path())
models = Instance.get_resources(instances, resource_name='cross_validated_model')
# + pycharm={"name": "#%%\n"}
plt.figure(figsize=(2, 5))
# plt.pcolor(all_scores[['training_loss', 'validation_loss']])
plt.pcolor(all_scores[['training_accuracy', 'validation_accuracy']])
plt.colorbar()
plt.show()
# + pycharm={"name": "#%%\n"}
def f(column_name):
column = all_scores[[column_name]]
v = column.to_numpy()
m = v.reshape((-1, instance.cv_k)).transpose()
plt.figure(figsize=(5, 2))
plt.matshow(m, fignum=0)
plt.colorbar(orientation='horizontal')
plt.title(column_name)
plt.xlabel('instances')
plt.ylabel('cv fold')
plt.show()
f('training_accuracy')
f('validation_accuracy')
f('training_loss')
f('validation_loss')
# + pycharm={"name": "#%%\n"}
def g(column_name):
m = all_scores[[column_name]].to_numpy().reshape((-1, instance.cv_k))
means = np.mean(m, axis=1)
plt.figure()
plt.hist(means)
plt.title('histogram of mean ' + column_name)
plt.show()
g('training_loss')
g('validation_loss')
g('training_accuracy')
g('validation_accuracy')
# + pycharm={"name": "#%%\n"}
parallel_df = all_scores.copy()
parallel_df.set_index(keys=['instance_hash'], inplace=True)
parallel_df['transformation'] = None
parallel_df['centering'] = None
parallel_df['n_hidden_layers'] = None
transformations = {}
for instance in instances:
instance_hash = instance.get_instance_hash()
parallel_df.at[instance_hash, 'transformation'] = instance.transformation
parallel_df.at[instance_hash, 'centering'] = instance.centering
parallel_df.at[instance_hash, 'n_hidden_layers'] = instance.n_hidden_layers
# + pycharm={"name": "#%%\n"}
import plotly
from plotly.offline import init_notebook_mode
init_notebook_mode(connected = True)
import plotly.express as px
fig = px.parallel_coordinates(parallel_df[['training_loss', 'validation_loss', 'training_accuracy', 'validation_accuracy']], color='validation_accuracy', color_continuous_scale=px.colors.diverging.Tealrose, title='Relation between losses and accuracies')
fig.update_layout(
autosize=False,
width=1600,
height=800
)
plotly.offline.iplot(fig)
# + pycharm={"name": "#%%\n"}
parallel_df.sort_values(by=['validation_accuracy', 'n_hidden_layers'], ascending=False, inplace=True)
df = px.data.tips()
fig = px.parallel_categories(parallel_df, dimensions=['transformation', 'centering', 'n_hidden_layers', 'validation_accuracy'],
color='validation_accuracy', color_continuous_scale=px.colors.sequential.Inferno,
labels={'transformation': 'Transformation', 'centering': 'Centering', 'n_hidden_layers': '# hidden layers', 'validation_accuracy': 'Validation accuracy'},
title='Effect of the hyperparameters on validation accuracy')
fig.update_layout(
autosize=False,
width=1600,
height=800
)
fig.write_image(jupyter_plot('parallel_categories.png'))
fig.show()
# + pycharm={"name": "#%%\n"}
import plotly.graph_objects as go
p_df = parallel_df.copy()
transformation_k = list(set(parallel_df['transformation'].tolist()))
transformation_v = list(range(len(transformation_k)))
transformation_d = dict(zip(transformation_k, transformation_v))
p_df['transformation'] = p_df['transformation'].apply(lambda x: transformation_d[x])
p_df['centering'] = p_df['centering'].apply(lambda x: int(x))
# display(p_df.head(10))
# for x in p_df.iloc[0]:
# print(type(x), x)
# + pycharm={"name": "#%%\n"}
fig = go.Figure(data=
go.Parcoords(
line = dict(color = p_df['validation_accuracy'],
colorscale = 'Viridis',
showscale = True,
cmin = p_df['validation_accuracy'].min(),
cmax = p_df['validation_accuracy'].max()),
dimensions = list([
dict(range = [min(transformation_v), max(transformation_v)],
tickvals = transformation_v,
ticktext = transformation_k,
label = 'Transformation', values = p_df['transformation'].tolist()),
dict(range = [0, 1],
tickvals = [0, 1],
ticktext = ['No', 'Yes'],
label = 'Centering', values = p_df['centering'].tolist()),
dict(range = [p_df['n_hidden_layers'].min(), p_df['n_hidden_layers'].max()],
tickvals = p_df['n_hidden_layers'].unique().tolist(),
label = '# hidden layers', values = p_df['n_hidden_layers'].tolist()),
dict(range = [p_df['training_accuracy'].min(), p_df['training_accuracy'].max()],
label = 'Training accuracy', values = p_df['training_accuracy'].tolist()),
dict(range = [p_df['validation_accuracy'].min(), p_df['validation_accuracy'].max()],
label = 'validation accuracy', values = p_df['validation_accuracy'].tolist()),
dict(range = [p_df['training_loss'].min(), p_df['training_loss'].max()],
label = 'Training loss', values = p_df['training_loss'].tolist()),
dict(range = [p_df['validation_loss'].min(), p_df['validation_loss'].max()],
label = 'validation loss', values = p_df['validation_loss'].tolist()),
# dict(range = [1,5],
# tickvals = [1,2,4,5],
# label = 'C', values = [2,4],
# ticktext = ['text 1', 'text 2', 'text 3', 'text 4']),
# dict(range = [1,5],
# label = 'D', values = [4,2])
])
)
)
fig.update_layout(
template='plotly_dark',
title='Effect of the hyperparameters on validation accuracy',
autosize=False,
width=1600,
height=800
)
fig.show()
# + pycharm={"name": "#%%\n"}
|
example/notebooks/results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ''
# name: sagemath
# ---
# + language="html"
# <link href="http://mathbook.pugetsound.edu/beta/mathbook-content.css" rel="stylesheet" type="text/css" />
# <link href="https://aimath.org/mathbook/mathbook-add-on.css" rel="stylesheet" type="text/css" />
# <style>.subtitle {font-size:medium; display:block}</style>
# <link href="https://fonts.googleapis.com/css?family=Open+Sans:400,400italic,600,600italic" rel="stylesheet" type="text/css" />
# <link href="https://fonts.googleapis.com/css?family=Inconsolata:400,700&subset=latin,latin-ext" rel="stylesheet" type="text/css" /><!-- Hide this cell. -->
# <script>
# var cell = $(".container .cell").eq(0), ia = cell.find(".input_area")
# if (cell.find(".toggle-button").length == 0) {
# ia.after(
# $('<button class="toggle-button">Toggle hidden code</button>').click(
# function (){ ia.toggle() }
# )
# )
# ia.hide()
# }
# </script>
#
# -
# **Important:** to view this notebook properly you will need to execute the cell above, which assumes you have an Internet connection. It should already be selected, or place your cursor anywhere above to select. Then press the "Run" button in the menu bar above (the right-pointing arrowhead), or press Shift-Enter on your keyboard.
# $\newcommand{\identity}{\mathrm{id}}
# \newcommand{\notdivide}{\nmid}
# \newcommand{\notsubset}{\not\subset}
# \newcommand{\lcm}{\operatorname{lcm}}
# \newcommand{\gf}{\operatorname{GF}}
# \newcommand{\inn}{\operatorname{Inn}}
# \newcommand{\aut}{\operatorname{Aut}}
# \newcommand{\Hom}{\operatorname{Hom}}
# \newcommand{\cis}{\operatorname{cis}}
# \newcommand{\chr}{\operatorname{char}}
# \newcommand{\Null}{\operatorname{Null}}
# \newcommand{\lt}{<}
# \newcommand{\gt}{>}
# \newcommand{\amp}{&}
# $
# <div class="mathbook-content"></div>
# <div class="mathbook-content"><p id="p-353">We begin our study of algebraic structures by investigating sets associated with single operations that satisfy certain reasonable axioms; that is, we want to define an operation on a set in a way that will generalize such familiar structures as the integers ${\mathbb Z}$ together with the single operation of addition, or invertible $2 \times 2$ matrices together with the single operation of matrix multiplication. The integers and the $2 \times 2$ matrices, together with their respective single operations, are examples of algebraic structures known as groups.</p></div>
# <div class="mathbook-content"><p id="p-354">The theory of groups occupies a central position in mathematics. Modern group theory arose from an attempt to find the roots of a polynomial in terms of its coefficients. Groups now play a central role in such areas as coding theory, counting, and the study of symmetries; many areas of biology, chemistry, and physics have benefited from group theory.</p></div>
# <div class="mathbook-content"><nav class="summary-links"><li><a href="section-mod-n-sym.ipynb"><span class="codenumber">3.1</span><span class="title">Integer Equivalence Classes and Symmetries</span></a></li><li><a href="section-groups-define.ipynb"><span class="codenumber">3.2</span><span class="title">Definitions and Examples</span></a></li><li><a href="section-subgroups.ipynb"><span class="codenumber">3.3</span><span class="title">Subgroups</span></a></li><li><a href="exercises-groups.ipynb"><span class="codenumber">3.4</span><span class="title">Exercises</span></a></li><li><a href="groups-exercises-detecting-errors.ipynb"><span class="codenumber">3.5</span><span class="title">Additional Exercises: Detecting Errors</span></a></li><li><a href="groups-references.ipynb"><span class="codenumber">3.6</span><span class="title">References and Suggested Readings</span></a></li><li><a href="groups-sage.ipynb"><span class="codenumber">3.7</span><span class="title">Sage</span></a></li><li><a href="groups-sage-exercises.ipynb"><span class="codenumber">3.8</span><span class="title">Sage Exercises</span></a></li></nav></div>
|
aata/groups.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OCEA 201 -- Homework 2
# +
# imports
import unyt
import numpy as np
from IPython.display import Image
from IPython.core.display import HTML
from IPython.display import display
# -
# # Running the code
#
# 80 TSTOP (years)
# 4 IPLTM (years)
# 0.05 X_0 (N/(m*m))
# 1e4 A_H m*m/s
# 0. R_1
# # Steady-state solution
# ## As derived in class, and also in Section 3.4 of Pedlosky, the linear form of the layer equations has steady-state solutions for the stream functions of the 3 layers:
#
# ## $\beta \frac{\partial \psi_1}{\partial x} = \frac{f_0}{H_1} w_E$
#
# ## $\beta \frac{\partial \psi_2}{\partial x} = 0$
#
# ## $\beta \frac{\partial \psi_3}{\partial x} = 0$
# ## Equation 2 (and 3), impose a no-flow condition in Layer 2 (and 3) given the boundary condition of no-flow on the Eastern boundary. This is what we expect to see once the model comes to steady-state.
# ----
# # Year 1
Image(filename = 'Outputs/year1_stream.png', width=700)#, height=100)
# ## Here, we see transport in both layers.
#
# ## In layer 1, we see the Interior motions characterstic of Sverdrup transport and we see the (fast) establishment of the Western boundary current.
#
# ## In layer 2, there is also substantial motion, far more complex than that apparent in layer 1. This includes western motion.
# ----
# # Year 20
Image(filename = 'Outputs/year20_stream.png', width=700)#, height=100)
# ## Here we are at year 20 where the motions in Layer 1 now are very similar to the steady state solution from last week's assignment.
#
# ## In Layer 2, the motions in the lower-right portion of the box have been eliminated by westward-moving Rossby waves. As their speed is inversely propotional to $f$, the effects are greatest towards the bottom of the box (where $f$ is smaller).
#
# ## At the top of the Layer 2 box, however, there are still motions as the Rossby waves have yet to cross the box.
#
# ----
# # Year 40
Image(filename = 'Outputs/year40_stream.png', width=700)#, height=100)
# ## In Layer 2, the motions have now almost entirely disappeared. Only the upper left region has lingering transport.
# ----
# # Year 80
Image(filename = 'Outputs/year80_stream.png', width=700)#, height=100)
# ## Now, in Year 80, Layer 2 shows no transport. The Rossby waves and entirely achieved the steady state solution.
|
OCEA-201/hw/2/OCEA201_Homework2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feos-dft
# language: python
# name: feos-dft
# ---
# +
from feos_dft import *
from feos_dft.si import *
import numpy as np
import matplotlib.pyplot as plt
# -
sigma = ANGSTROM
func = FMTFunctional(np.array([sigma/ANGSTROM]), FMTVersion.WhiteBear)
# # Adsorption isotherms
# +
# %%time
potential = ExternalPotential.HardWall(sigma/ANGSTROM)
pore_size = 5*sigma
pores = {'slit_pore': (Pore1D(func, Geometry.Cartesian, 2*pore_size, potential), pore_size-sigma/2),
'cylindrical_pore': (Pore1D(func, Geometry.Cylindrical, pore_size, potential), np.pi*(pore_size-sigma/2)**2),
'spherical_pore': (Pore1D(func, Geometry.Spherical, pore_size, potential), 4/3*np.pi*(pore_size-sigma/2)**3)}
f, ax = plt.subplots(1,2,figsize=(15,5))
for label, (pore, volume) in pores.items():
pmax = State(func, KELVIN, density=0.75/(sigma**3*NAV)).pressure()
isotherm = Adsorption1D.adsorption_isotherm(func, KELVIN, (1e-10*pmax, pmax, 50), pore)
ax[0].plot(isotherm.pressure/(KB*KELVIN)*sigma**3, isotherm.total_adsorption/volume*sigma**3*NAV, label=label)
profile = isotherm.profiles[-1]
ax[1].plot(profile.r/sigma, (profile.density*sigma**3*NAV).T, label=label)
ax[0].axis([0,5,0,0.75])
ax[0].set_xlabel('$\\beta p\sigma^3$')
ax[0].set_ylabel('$\\bar\\rho\sigma^3$')
ax[0].legend()
ax[1].axis([0,5,0,6])
ax[1].set_xlabel('$r~~/~~\sigma$')
ax[1].set_ylabel('$\\rho\sigma^3$')
ax[1].legend();
# -
# # Pair correlation function
# +
for density in [0.1,0.5,0.95]:
bulk = State(func, KELVIN, density=density/(sigma**3*NAV))
pair_correlation = PairCorrelation(bulk, 2048, 30*sigma).solve()
plt.plot(pair_correlation.r/ANGSTROM, pair_correlation.pair_correlation_function.T, label=f'$\\rho\sigma^3={density}$')
plt.axis([0,4,0,6])
plt.xlabel('$r~~/~~\sigma$')
plt.ylabel('$g(r)$')
plt.legend();
# -
# # Density profiles of mixtures
# +
mix = FMTFunctional(np.array([1.0, 2.0, 0.5]), FMTVersion.WhiteBear)
bulk = State(mix, KELVIN, density=0.42/(ANGSTROM**3*NAV), molefracs=np.array([0.3,0.2,0.5]))
profile = Pore1D(mix, Geometry.Cartesian, 10*ANGSTROM, ExternalPotential.HardWall(1.0)).initialize(bulk).solve()
plt.plot(profile.z/ANGSTROM, (profile.density*ANGSTROM**3*NAV).T)
plt.axis([0,5,0,1])
plt.xlabel('$z~~/~~\mathrm{\AA}$')
plt.ylabel('$\\rho~~/~~\mathrm{\AA^{-3}}$');
# -
|
examples/FundamentalMeasureTheory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
css_file = "./notebook_style.css"
HTML(open(css_file, 'r').read())
# + [markdown] slideshow={"slide_type": "slide"}
# # 2. Testing Scientific Codes
#
# - [When should I test?](#When-should-I-test?)
# - [Effective testing](#Effective-testing)
# * [Unit tests](#Unit-tests)
# * [Integration tests](#Integration-tests)
# * [Convergence tests](#Convergence-tests)
# * [Regression tests](#Regression-tests)
# * [Test driven development](#Test-driven-development)
# - [Common problems and how to solve them](#Common-problems-and-how-to-solve-them)
# * [Randomness](#My-code-has-some-randomness-and-so-its-output-changes-every-time-I-run-it---what-can-I-test-for?)
# * [Unknown solutions](#I-don%27t-know-what-the-correct-solution-should-be)
# * [Building on other people's code](#I-didn%27t-write-most-of-the-code---how-do-I-know-that-the-bit-I-wrote-works?)
# * [Numerical error](#I-know-there-is-some-numerical-error-in-my-code---how-can-I-test-my-code-is-correct-up-to-this-error?)
# - [Further reading](#Further-reading)
# -
import numpy
from numpy.random import rand
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams.update({'font.size': 18})
from scipy.integrate import quad
import unittest
# + [markdown] slideshow={"slide_type": "slide"}
# In the experimental sciences, new theories are developed by applying the scientific method. This involves carrying out tests to ensure that results are accurate, reproducible and reliable. The experimental setup will be tested in order to show that it is working as designed and so as to eliminate or quantify any systematic errors. A result will not be trusted unless the experiment itself has been carried out to a suitable standard.
#
# In computational science, we should apply the same principles to our code. A result should only be trusted if the code that has produced it has undergone rigorous testing which demonstrates that it is working as intended and that any limitations of the code (e.g. numerical errors) are understood and quantified.
#
# Unfortunately, testing scientific codes can be quite challenging. By their very nature, they are often built in order to investigate systems where the behaviour is to some extent unknown. They can be very complex, often built over a number of years (or even decades!) with contributions from a vast number of people. However, even for the most complicated of codes there are a number of different types of tests that we can apply in order for us to build robust, reliable code whose results can be trusted.
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# 
# ##### Writing good code is hard - [xkcd](https://xkcd.com/844/)
# + [markdown] slideshow={"slide_type": "slide"}
# ## When should I test?
#
# ### Always and often
#
# The earlier you start testing the better, as it will be possible to catch bugs as they develop and before they become too entrenched in the code. Once written, you should then try and execute tests every time changes are made. Continuous integration (see below) is a useful tool to use to make sure that tests are run frequently - once the tests are written and the CI setup, they can then be forgetten about to a certain extent, safe in the knowledge that if any bugs are introduced in changes to the code, they should be caught.
#
# **However**, it is important to review your tests regularly. In code that is being actively developed, tests must be amended and new tests written so as to make sure that new features are also tested. Regression tests are useful here to test that changes to the code improve its performance rather than making it worse. Code coverage is a useful tool to make sure that all code is being tested. It's all very well having a testing suite, but if only 20% of the code has been tested, you still cannot trust that the other 80% of the code is producing reliable results.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Effective testing
#
# In order to have an effective set of tests, it's necessary to make sure that the entire parameter space is tested, not just one or two nice cases. Of particular importance are edge and corner cases. If the code needs to run over a set of parameters, then edge cases are those which are at the beginning and end of this range. A corner case is then where one or more edge cases are combined. Such cases tend to be where errors most often arise, as often special code is required to deal with boundary values.
#
# In the code below, we demonstrate the importance of testing edge cases. The code takes a vector $\mathbf{v}$ and normalises it $\hat{\mathbf{v}} = \frac{\mathbf{v} }{ |\mathbf{v}|}$. We see that if the code is run for the vector $(0,0)$, a `RuntimeWarning` is raised as the function is attempting to divide by zero.
# -
def normalise(v):
norm = numpy.sqrt(numpy.sum(v**2))
return v / norm
normalise(numpy.array([0,0]))
# We therefore need to amend our function for the case where the norm of the vector is zero. A possible solution is the function below.
def improved_normalise(v):
norm = numpy.sqrt(numpy.sum(v**2))
if norm == 0.:
return v
return v / norm
improved_normalise(numpy.array([0,0]))
# Our improved function now tests to see if the norm is zero - if so, it returns the original vector rather than attempting to divide by zero. There are often multiple ways to deal with invalid data in a program - the choice of which method to use depends on which is most appropriate for your particular problem. An alternative implementation of the normalise function would be to use [*exception handling*](https://docs.python.org/3/tutorial/errors.html#exceptions). Exceptions are errors which are detected during the execution of the program (rather than syntax or parsing errors which are detected by the parser before the program is run). You have most likely come across some of python's built in exceptions before, e.g. `FloatingPointError`, `IOError`, `ZeroDivisionError`. It's also possible for the user to define their own exceptions. The default behaviour when an exception is detected is for the program to exit (the exception is an *unhandled error*), however we can instead choose to *handle* the exception using a `try...except` block.
#
# By default, numpy sets floating-point errors to be raised as *warnings* rather than exceptions. The first line of the code below turns this behaviour off.
# +
numpy.seterr(invalid='raise')
def exception_normalise(v):
try:
norm = numpy.sqrt(numpy.sum(v**2))
vhat = v / norm
except FloatingPointError:
print('Norm of vector is zero, returning original vector')
vhat = v
return vhat
# -
exception_normalise(numpy.array([0,0]))
# It is also important to check that the code breaks as expected. If the code input is garbage but it still manages to run as normal, that is not good behaviour and suggests some data validation of input parameters is needed. For example, let's try to run our improved normalisation function on a string:
exception_normalise("I am a string")
# Python correctly spots that it cannot perform the power operation on a string and raises a `TypeError` exception. However, it would probably be more useful to implement some kind of type checking of the function inputs before this (e.g. using `numpy.isnumeric`), and/or make sure that the code that calls this function is capable of catching such exceptions. For example, we may decide that if data of an invalid type is passed to the function, it should return 0:
def exception_normalise(v):
try:
norm = numpy.sqrt(numpy.sum(v**2))
vhat = v / norm
except FloatingPointError:
print('Norm of vector is zero, returning original vector')
vhat = v
except TypeError:
print('Invalid input data, returning 0')
vhat = 0
return vhat
exception_normalise("I am a string")
# ### Unit tests
#
# For complicated codes made up of many functions, it is useful to write a serious of tests that check small parts of the code - *units* - at a time. This makes it easier to track down the exact location of bugs. These units may be individual functions or groups of shorter functions. Unit tests therefore encourage good coding practice, as they require code to be modular.
#
# In the example below, we have three (very simple) functions: `squared` which returns the square of its input, `add_2` which adds 2 to its input and `square_plus_2` which calls the two previous functions to return $x^2+2$. To test this code, we could therefore write unit tests for the first two functions to check they are working correctly. We've used the `unittest` module here as it allows us to test that functions correctly raise exceptions when given invalid data.
# +
def squared(x):
return x*x
def add_2(x):
return x + 2
def square_plus_2(x):
return add_2(squared(x))
# +
class test_units(unittest.TestCase):
def test_squared(self):
self.assertTrue(squared(-5) == 25)
self.assertTrue(squared(1e5) == 1e10)
self.assertRaises(TypeError, squared, "A string")
def test_add_2(self):
self.assertTrue(add_2(-5) == -3)
self.assertTrue(add_2(1e5) == 100002)
self.assertRaises(TypeError, add_2, "A string")
test_units().test_squared()
test_units().test_add_2()
# -
# ### Integration tests
#
# Once you've written your unit tests and are pretty confident that individual parts of the code work on their own, you than need to verify that these different parts work together. To see why this is needed, imagine you were asked to build a car, despite only having a vague idea of how everything fits together. You've been given all the different parts (the engine, the wheels, the steering wheel...) - these have all previously undergone rigorous testing and you have been assured that they all work fine. You put them all together to the best of your ability, but unfortunately cannot get the car to work. Much as with your code, despite the individual parts working, this is no guarantee that they will work when put together.
#
# In the above example, we can add an integration test by writing a test for `square_plus_2` - this calls the other two functions, so we'll test that it does this properly.
# +
class test_integration(unittest.TestCase):
def test_square_plus_2(self):
self.assertTrue(square_plus_2(-5) == 27)
self.assertTrue(square_plus_2(1e5) == 10000000002)
self.assertRaises(TypeError, square_plus_2, "A string")
test_integration().test_square_plus_2()
# -
# As we'll see below, integration tests can be difficult to design. They can encompass a small section of the code, e.g. to check that one function correctly calls another, all the way up to the entire code. Because they can involve many different functions, they are often a lot more complex than unit tests.
# ### Convergence tests
#
# Often we want to calculate a solution on some kind of grid. The solution we find is a discretised approximation of the exact continuous solution. As the resolution of the grid increases, the solution should approach exact solution. Convergence tests are a way of checking this. The solution is calculated for grids of various resolutions. If the code is working correctly, the error of the solution should decrease with increasing resolution approximately at an order that depends on the accuracy of the algorithm (until the error becomes so small it then becomes dominated by floating point errors).
# In the example below, we will demonstrate this by using the trapezium rule to approximate the integral of $\sin (x)$ with various different step sizes, $h$. By comparing the calculated errors to a line of gradient $h^2$, it can be seen that the numerical approximation is converging as expected at $O(h^2)$.
# +
hs = numpy.array([1. / (4. * 2.**n) for n in range(8)])
errors = numpy.zeros_like(hs)
for i, h in enumerate(hs):
xs = numpy.arange(0., 1.+h, h)
ys = numpy.sin(xs)
# use trapezium rule to approximate integral
integral_approx = sum((xs[1:] - xs[:-1]) * 0.5 * (ys[1:] + ys[:-1]))
errors[i] = -numpy.cos(1) + numpy.cos(0) - integral_approx
plt.loglog(hs, errors, 'x', label='Error')
plt.plot(hs, 0.1*hs**2, label=r'$h^2$')
plt.xlabel(r'$h$')
plt.ylabel('error')
plt.legend(loc='center left', bbox_to_anchor=[1.0, 0.5])
plt.show()
# -
# ### Regression tests
#
# When building your code, generally you'll be aiming for its performance to improve with time. Results should get more accurate or, at the very least, should not deteriorate. Regression tests are a way to check this. Multiple versions of the code are run and the outputs compared. If the output has changed such that it is significantly different from the previous output, the test fails. Such tests can help catch bugs that other types of tests may not, and can help ensure the project remains backwards-compatible for such cases where that is important.
# ### Test driven development
#
# Above we said that it's best to start testing your code early on in the development of your code. Test driven development takes this to the extreme: tests are written *before* you even write a line of the main code. The tests then take the form of a sort of 'product specification', providing a set of tests for things that you aim for your finished code to be able to do. In order to be able to write such tests, you need to have a good idea of the structure your code will have, so test driven development can be useful in that it forces you to plan out your code before you start writing it.
#
# Unfortunately, as we've seen, scientific codes can be difficult to test, often having unpredictable output. They often implement novel techniques, so it can be hard to plan what the final code will look like. This means that test driven development can be harder to successfully implement for scientific codes.
# ## Common problems and how to solve them
#
# ### My code has some randomness and so its output changes every time I run it - what can I test for?
#
# In time evolution problems, it may be that whilst the output at any individual timestep can be somewhat random, the behaviour averaged over a number of timesteps is to some extent known. Tests can therefore be written to check that this is the case. In other problems, it may be more useful to test the average behaviour across the entire domain or sections of the domain. Even if the behaviour is completely random and so it's not possible to take any meaningful averages, the chances are that it should still be within a set of known values - we can therefore write tests that check the data is within these limits. Another strategy is to try to write tests that isolate the random parts so that you can check the non-random parts of the code work. If you are using a random number generator, it can be possible to eliminate the non-determinism by testing using a fixed seed value for the generator.
#
# In the code below, we generate an array of random data and apply some function to it before plotting the results. It can be seen that the output is different every time the code is run.
# +
data = rand(80,80)
def func(a):
return a**2 * numpy.sin(a)
output = func(data)
plt.imshow(output)
plt.colorbar()
plt.show()
# -
# The output of this code changes every time the code is run, however we can still write some tests for it. We know that all values in the output array must be $0\leq x \leq 1$. In some circumstances, such as in this case, we may know the statistical distribution of the random data. We can therefore calculate what the average output value should be and compare this to our code's output. In our case, the data is generated from a uniform distribution of numbers between 0 and 1, so the average value of the output is given by $\int_0^1 f(x) \,dx \simeq 0.22$
# +
def test_limits(a):
if numpy.all(a >= 0.) and numpy.all(a <= 1.):
return True
return False
def test_average(a):
if numpy.isclose(numpy.average(a), 0.223, rtol=5.e-2):
return True
return False
if test_limits(output):
print('Function output within correct limits')
else:
print('Function output is not within correct limits')
if test_average(output):
print('Function output has correct average')
else:
print('Function output does not have correct average')
# -
# ### I don't know what the correct solution should be
#
# In experimental science, the experimental setup will be tested using a control. This is where the experiment is run using a set of input data for which the outcome is known, so that any bugs in the apparatus or systematic errors can be identified. In computational science, there is often a simple system whose behaviour is known which can be used to test the code. E.g. in time evolution problems, a system which is initially static should remain that way. If this is not the case, then this indicates there is something seriously wrong with the code! In physics, we can also check for symmetries of the system (e.g. rotational symmetry, translation symmetry, reflection symmetry). There are also often conserved quantities (e.g. mass, energy, charge) that we can check the code conserves.
#
# In the below example, we look at a black box function - `scipy.integrate.quad`. Here, this function will stand in for a bit of code that we have written and want to test. Say we wish to use `quad` to calculate the integral of some complicated function and we have little idea what the solution will be. Before we use it on the complicated function, we will test that it behaves correctly for a function whose integral we already know: $f(x) = \sin(x)$.
# +
xs = numpy.linspace(0.0, 2.0 * numpy.pi)
integrals = numpy.zeros_like(xs)
for i in range(len(xs)):
integrals[i] = quad(numpy.sin, 0.0, xs[i])[0]
plt.plot(xs, -numpy.cos(xs)+1, '-', label=r'$\int f(x)$')
plt.plot(xs, integrals, 'x', label='quad')
plt.legend(loc='center left', bbox_to_anchor=[1.0, 0.5])
plt.show()
# -
# As hoped, quad gives the correct solution:
#
# $$
# \int^\alpha_0 \sin(x)\, dx = -\cos(\alpha) + 1
# $$
#
# ### I didn't write most of the code - how do I know that the bit I wrote works?
#
# Unit tests! If the original code can run in isolation, make sure that there are suitable tests which make sure that it works correctly. Any failures in subsequent tests that then incorporate your code will therefore only be the result of bugs in your code. Unit tests of individual functions in your code should also be used.
# ### I know there is some numerical error in my code - how can I test my code is correct up to this error?
#
# In numerical calculations, there will always be some computational error that cannot be avoided. This can from the computer's floating point representation of numerical data or from the choice of algorithm used. It is often the case that we don't require our result to be 100% precise, but rather correct up to some tolerance. We can therefore build tests to reflect this.
#
# In python, we can use `numpy.isclose` and `numpy.allclose` to do this. In the example below, we take some data and add a small amount of random noise to it. This random noise is supposed to represent numerical errors that are introduced over the course of a simulation. If we test that the output array is equal to the original array, python correctly tells us that it is not. However, if we test that the output array is close to the original array, we find that this is true.
# +
x = numpy.linspace(0, 2*numpy.pi, num=500)
initial_data = x**2 * numpy.cos(5*x)
# add noise
noisey_data = initial_data + (rand(len(x)) - 0.5) * 4
plt.plot(x, initial_data, label='initial data')
plt.plot(x, noisey_data, label='noisey data')
plt.legend(loc='center left', bbox_to_anchor=[1.0, 0.5])
plt.xlim(x[0], x[-1])
plt.show()
if numpy.array_equal(initial_data, noisey_data):
print('Noisey data exactly equal to initial data')
else:
print('Noisey data is not exactly equal to initial data')
if numpy.allclose(initial_data, noisey_data, atol=2):
print('Noisey data is close to initial data')
else:
print('Noisey data is not close to initial data')
# -
# Another tactic that can be used here is to employ convergence tests to make sure that the numerical error is decreasing with increasing resolution (at least until the point where this error becomes dominated by floating point erorrs). If you know that the algorithm you are using to solve the system is accurate to some order, your solution should converge at a rate close to this. If not, it is likely that you have made a mistake in your implementation of the algorithm.
# ## Further reading
# - Check out [this presentation](https://www.google.co.uk/url?sa=t&rct=j&q=&esrc=s&source=web&cd=4&ved=0ahUKEwjg64jtyM7UAhVkLcAKHftYDiwQFghJMAM&url=http%3A%2F%2Farchive.euroscipy.org%2Ffile%2F8962%2Fraw%2FEuroScipy_tutorial_2012.pdf&usg=AFQjCNG0K9lmRR5wUbn-8C4YToxMdklW0Q&sig2=BhwDp-UodcNOLsSufqKUhg&cad=rja) by <NAME> from EuroScipy 2012 which gives a detailed demonstration of how to use the python `unittest` library and covers some advanced testing topics such as mocks and patches
# - [The design and use of reference data sets for testing scientific software](http://http.icsi.berkeley.edu/ftp/pub/speech/papers/wikipapers/cox_harris_testing_numerical_software.pdf), <NAME> and <NAME> - outlines the problems with testing scientific software and how you can go about creating suitable test data
#
|
02-Testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
from numpy import array
def read_data(file):
pairs = []
f = open(file, mode='r', encoding ='utf8')
for x in f:
sample = json.loads(x)
pair = [sample['source'], sample['target']]
pairs.append(pair)
return array(pairs)
def predict_sequence(model, tokenizer, source):
prediction = model.predict(source, verbose=0)[0]
integers = [argmax(vector) for vector in prediction]
target = list()
for i in integers:
word = word_for_id(i, tokenizer)
if word is None:
break
target.append(word)
return ' '.join(target)
def word_for_id(integer, tokenizer):
for word, index in tokenizer.word_index.items():
if index == integer:
return word
return None
# +
from tensorflow.keras.models import load_model
from numpy import argmax
model = load_model('mtrans.h5')
myData = read_data('tiny.json')[:10]
from ipynb.fs.full.model import encode_sequences
from ipynb.fs.full.model import src_tok
from ipynb.fs.full.model import src_length
from ipynb.fs.full.model import tgt_tok
myTest = encode_sequences(src_tok, src_length, myData[:,0])
for count, source in enumerate(myTest):
source = source.reshape((1, source.shape[0]))
translation = predict_sequence(model, tgt_tok, source)
print(myData[:,0][count], source, translation, sep=':')
# -
|
mtrans/predict.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Augmenation On The PCB Defect Dataset
#
# This notebook contains all the code needed to use TAO augmenation on subsets of the PCB defect dataset to showcase how augmenatation can be used to improve KPIs for small datasets.
#
# This notebook requires the TAO Launcher, Docker and NGC to be setup
#
# The github readme has steps on setting up the prerequisites
#
# This notebook also requires preprocess_pcb.py to be in the same directory to function.
#
# This notebook takes the following steps
# 1) Download and unpack the PCB defect dataset
#
# 2) Convert the dataset to kitti format
#
# 3) Split the dataset into test and train subsets
#
# 4) Map local directories the the TAO launcher
#
# 5) Generate offline augmenation spec file and apply augmentation to the training sets
#
# 6) Generate TF Records for the test and training sets
#
# 7) Downloads pretrained object detection weights needed for the trainings
#
# 8) Launch trainings and evaluation
#
# The last section of this notebook contains all the commands needed to run training and evaluation on all 6 datasets.
# Steps 1-7 only need to run 1 time. The trainings in step 7 can be run in any order once steps 1-6 have successfully run.
# A common test set of 500 images is used for validation on all trainings
#
# Datasets
# 100 subset x1
# 100 subset x10
# 100 subset x20
# 500 subset x1
# 500 subset x10
# 500 subset x20
#
# !python3 -m pip install matplotlib
import os
from preprocess_pcb import convert_annotation, create_subset
# +
#paths relative to local repository
repo_home = os.path.join(os.getcwd(), "../../")
model_home = os.path.join(repo_home, "workspace/models")
dataset_home = os.path.join(repo_home, "datasets/pcb_defect")
exp_home = os.path.join(repo_home, "workspace/pcb_data_aug")
#paths for inside the container
dataset_home_cont = "/datasets/pcb_defect/"
exp_home_cont = "/tlt_exp/pcb_data_aug/"
# -
# ## Download and unpack the PCB defect dataset
# %cd $dataset_home
#download and unzip
# !wget https://www.dropbox.com/s/h0f39nyotddibsb/VOC_PCB.zip
# !unzip VOC_PCB.zip
# ## Convert the dataset to kitti format
#setup folders for dataset images and labels
os.makedirs("original/images", exist_ok=True)
os.makedirs("original/labels", exist_ok=True)
# !cp -r VOC_PCB/JPEGImages/. original/images
# +
#Setup Paths and make label folder
xml_label_path = "VOC_PCB/Annotations"
kitti_label_output = "original/labels"
#Convert labels to kitti and put into output folder
for x in os.listdir(xml_label_path):
current_label_path = os.path.join(xml_label_path, x)
convert_annotation(current_label_path, kitti_label_output)
# -
# ## Split the dataset into test and train subsets
# +
#Setup folders for dataset subset
test_500 = os.path.join(exp_home, "test_500_list.txt")
train_100 = os.path.join(exp_home, "train_100_list.txt")
train_500 = os.path.join(exp_home, "train_500_list.txt")
os.makedirs("500_subset_test_x1", exist_ok=True)
os.makedirs("100_subset_train_x1", exist_ok=True)
os.makedirs("500_subset_train_x1", exist_ok=True)
#Create the subsets based on predefined lists
create_subset("original", test_500, "500_subset_test_x1")
create_subset("original", train_100, "100_subset_train_x1")
create_subset("original", train_500, "500_subset_train_x1")
# +
## Map local directories the the TAO launcher
# +
# Mapping up the local directories to the TAO docker.
import json
mounts_file = os.path.expanduser("~/.tao_mounts.json")
# Define the dictionary with the mapped drives
drive_map = {
"Mounts": [
# Mapping the data directory
{
"source": os.path.join(repo_home, "datasets"),
"destination": "/datasets"
},
# Mapping the specs directory.
{
"source": os.path.join(repo_home, "workspace"),
"destination": "/tlt_exp"
},
]
}
# Writing the mounts file.
with open(mounts_file, "w") as mfile:
json.dump(drive_map, mfile, indent=4)
# -
# ## Generate offline augmenation spec file and apply augmentation to the training sets
from preprocess_pcb import gen_random_aug_spec, combine_kitti, visualize_images
from random import randint
#Input dataset folder to augment, augment output folder and number of augmentations. Requires local paths and container paths
#For each augment a randomized spec file and augmented dataset is produced
#Also outputs a dataset with all combined augmentations
def generate_augments(dataset_folder, dataset_folder_cont, output_folder, output_folder_cont, num_augments):
for i in range(0,num_augments):
spec_out = os.path.join(output_folder, "aug_spec" + str(i) + ".txt")
spec_out_cont = os.path.join(output_folder_cont, "aug_spec" + str(i) + ".txt")
gen_random_aug_spec(600,600,"jpg", spec_out)
# !cat $spec_out
aug_folder = os.path.join(output_folder, "aug" + str(i))
aug_folder_cont = os.path.join(output_folder_cont, "aug" + str(i))
# !tao augment -a $spec_out_cont -o $aug_folder_cont -d $dataset_folder_cont
if i == 0:
d1 = dataset_folder
d2 = aug_folder
d3 = os.path.join(output_folder, "combined_x2")
combine_kitti(d1,d2,d3)
else:
d1 = os.path.join(output_folder, "combined_x" + str(i+1))
d2 = aug_folder
d3 = os.path.join(output_folder, "combined_x" + str(i+2))
combine_kitti(d1,d2,d3)
# +
#generate augmentations for 100 image subset
dataset_folder = "100_subset_train_x1" #folder for the existing dataset to be augmented. This folder will not be modified
dataset_folder_cont = os.path.join(dataset_home_cont, "100_subset_train_x1")
output_folder = "100_subset_train_aug" #folder for the augmented output. Does not need to exist
output_folder_cont = os.path.join(dataset_home_cont, output_folder)
num_augments = 19 #number of augmented datasets to generate
os.makedirs(output_folder, exist_ok=True)
generate_augments(dataset_folder,dataset_folder_cont,output_folder, output_folder_cont, num_augments)
# -
#Display some of the augmented images
#Rerun to see new images each time
aug_choice = str(randint(0,num_augments-1))
visualize_images(os.path.join(output_folder, "aug"+aug_choice+"/images"), num_images=8)
# +
#generate augmentations for 500 image subset
dataset_folder = "500_subset_train_x1" #folder for the existing dataset to be augmented. This folder will not be modified
dataset_folder_cont = os.path.join(dataset_home_cont, "500_subset_train_x1")
output_folder = "500_subset_train_aug" #folder for the augmented output. Does not need to exist
output_folder_cont = os.path.join(dataset_home_cont, "500_subset_train_aug")
num_augments = 19 #number of augmented datasets to generate
os.makedirs(output_folder, exist_ok=True)
generate_augments(dataset_folder, dataset_folder_cont, output_folder, output_folder_cont, num_augments)
# -
#Display some of the augmented images
#Rerun to see new images each time
aug_choice = str(randint(0,num_augments-1))
visualize_images(os.path.join(output_folder, "aug"+aug_choice+"/images"), num_images=8)
# +
#Place important datasets in the dataset folder
# !mv 100_subset_train_aug/combined_x10 100_subset_train_x10
# !mv 100_subset_train_aug/combined_x20 100_subset_train_x20
# !mv 500_subset_train_aug/combined_x10 500_subset_train_x10
# !mv 500_subset_train_aug/combined_x20 500_subset_train_x20
# -
# ## Generate TF Records for the test and training sets
#Returns the tf record config as a string with the given dataset path
#root directory path must be inside the container
def gen_tf_spec(dataset_path):
spec_str = f"""
kitti_config {{
root_directory_path: "/datasets/pcb_defect/{dataset_path}"
image_dir_name: "images"
label_dir_name: "labels"
image_extension: ".jpg"
partition_mode: "random"
num_partitions: 2
val_split: 20
num_shards: 10
}}
"""
return spec_str
#Loop through all datasets to generate tf records
dataset_paths = ["500_subset_test_x1", "500_subset_train_x1", "500_subset_train_x10", "500_subset_train_x20", "100_subset_train_x1", "100_subset_train_x10", "100_subset_train_x20"]
for path in dataset_paths:
record_path = os.path.join(dataset_home, path, "tfrecord_spec.txt")
record_path_cont = os.path.join(dataset_home_cont, path, "tfrecord_spec.txt")
record_output = os.path.join(dataset_home, path, "tfrecords_rcnn/")
record_output_cont = os.path.join(dataset_home_cont, path, "tfrecords_rcnn/")
print("************" + record_path)
with open(record_path, "w+") as spec:
spec.write(gen_tf_spec(path))
# !tao faster_rcnn dataset_convert -d $record_path_cont -o $record_output_cont
# ## Downloads pretrained object detection weights needed for the trainings
#requires NGC to be configured
os.makedirs(os.path.join(model_home, "fasterRCNN"), exist_ok=True)
# %cd $model_home/fasterRCNN
# !ngc registry model download-version "nvidia/tlt_pretrained_object_detection:resnet18"
# ## Launch trainings and evaluation
# Each cell in this section will train and evaluate on 1 dataset in the experiment. The results will be output to the respective experiment folder.
#
# The trainings may take several hours depending on your hardware.
experiments_cont = os.path.join(exp_home_cont, "experiments")
experiments = os.path.join(exp_home, "experiments")
# !tao faster_rcnn train -e $experiments_cont/offline_aug/100_subset_train_x1/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/100_subset_train_x1/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/100_subset_train_x1/eval_log.txt
# !cat $experiments/offline_aug/100_subset_train_x1/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_aug/100_subset_train_x10/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/100_subset_train_x10/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/100_subset_train_x10/eval_log.txt
# !cat $experiments/offline_aug/100_subset_train_x10/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_aug/100_subset_train_x20/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/100_subset_train_x20/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/100_subset_train_x20/eval_log.txt
# !cat $experiments/offline_aug/100_subset_train_x20/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_aug/500_subset_train_x1/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/500_subset_train_x1/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/500_subset_train_x1/eval_log.txt
# !cat $experiments/offline_aug/500_subset_train_x1/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_aug/500_subset_train_x10/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/500_subset_train_x10/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/500_subset_train_x10/eval_log.txt
# !cat $experiments/offline_aug/500_subset_train_x10/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_aug/500_subset_train_x20/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_aug/500_subset_train_x20/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_aug/500_subset_train_x20/eval_log.txt
# !cat $experiments/offline_aug/500_subset_train_x20/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/100_subset_train_x1/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/100_subset_train_x1/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/100_subset_train_x1/eval_log.txt
# !cat $experiments/offline_online_aug/100_subset_train_x1/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/100_subset_train_x10/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/100_subset_train_x10/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/100_subset_train_x10/eval_log.txt
# !cat $experiments/offline_online_aug/100_subset_train_x10/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/100_subset_train_x20/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/100_subset_train_x20/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/100_subset_train_x20/eval_log.txt
# !cat $experiments/offline_online_aug/100_subset_train_x20/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/500_subset_train_x1/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/500_subset_train_x1/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/500_subset_train_x1/eval_log.txt
# !cat $experiments/offline_online_aug/500_subset_train_x1/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/500_subset_train_x10/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/500_subset_train_x10/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/500_subset_train_x10/eval_log.txt
# !cat $experiments/offline_online_aug/500_subset_train_x10/eval_log.txt
# !tao faster_rcnn train -e $experiments_cont/offline_online_aug/500_subset_train_x20/training_spec.txt -k tlt_encode
# !tao faster_rcnn evaluate -e $experiments_cont/offline_online_aug/500_subset_train_x20/training_spec.txt -k tlt_encode --log_file $experiments_cont/offline_online_aug/500_subset_train_x20/eval_log.txt
# !cat $experiments/offline_online_aug/500_subset_train_x20/eval_log.txt
|
workspace/pcb_data_aug/Process&Train_PCB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import numpy as np
import torch
from PIL import Image
from matplotlib.patches import Rectangle
with open("test.jpg", "rb") as f:
img = np.asarray(Image.open(f))
print("image is from ghibli.jp")
plt.imshow(img)
# + pycharm={"name": "#%%\n"}
def add_bbox(image,
bboxes):
fig, ax = plt.subplots()
if torch.is_tensor(image):
image = image.permute(1, 2, 0).numpy()
if torch.is_tensor(bboxes):
bboxes = [bbox.numpy() for bbox in bboxes.unbind(0)]
ax.imshow(image)
for bbox in bboxes:
left, top, right, bottom = bbox
#top, left, bottom, right = bbox
ax.add_patch(Rectangle((left, top), right-left, bottom-top,
linewidth=2, edgecolor=(0, 1, 0, 1), facecolor="none"))
return ax
bbox = torch.tensor([[240, 40, 310, 100]], dtype=torch.float)
# + pycharm={"name": "#%%\n"}
add_bbox(img, [[240, 40, 310, 100]])
# + pycharm={"name": "#%%\n"}
from homura.vision.transforms import RandomRotation
add_bbox(*RandomRotation(30,target_type="bbox")(img, bbox))
# + pycharm={"name": "#%%\n"}
from homura.vision.transforms import RandomHorizontalFlip
add_bbox(*RandomHorizontalFlip(0.9, target_type="bbox")(img, bbox))
# + pycharm={"name": "#%%\n"}
from homura.vision.transforms import RandomCrop
add_bbox(*RandomCrop((200, 300), target_type="bbox")(img, bbox))
# + pycharm={"name": "#%%\n"}
from homura.vision.transforms import RandomResizedCrop
add_bbox(*RandomResizedCrop((200, 300), target_type="bbox")(img, bbox))
# + pycharm={"name": "#%%\n"}
from homura.vision.transforms import CenterCrop
add_bbox(*CenterCrop(250, target_type="bbox")(img, bbox))
# + pycharm={"name": "#%%\n"}
add_bbox(*(CenterCrop(250, target_type="bbox") * RandomHorizontalFlip(p=0.9, target_type="bbox"))(img, bbox))
# + pycharm={"name": "#%%\n"}
add_bbox(*(RandomRotation(30, target_type="bbox") * CenterCrop(250, target_type="bbox") * RandomHorizontalFlip(p=0.9, target_type="bbox"))(img, bbox))
# + pycharm={"name": "#%%\n"}
|
examples/transforms.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Deep Learning
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# In this notebook, a template is provided for you to implement your functionality in stages, which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission if necessary.
#
# > **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
#
# In addition to implementing code, there is a writeup to complete. The writeup should be completed in a separate file, which can be either a markdown file or a pdf document. There is a [write up template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) that can be used to guide the writing process. Completing the code template and writeup template will cover all of the [rubric points](https://review.udacity.com/#!/rubrics/481/view) for this project.
#
# The [rubric](https://review.udacity.com/#!/rubrics/481/view) contains "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. The stand out suggestions are optional. If you decide to pursue the "stand out suggestions", you can include the code in this Ipython notebook and also discuss the results in the writeup file.
#
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ---
# ## Step 0: Import Module And Load The Data
# Import all the relevant modules.
import cv2
import csv
import matplotlib.image as mpimg
import matplotlib.mlab as mlab
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pickle
from random import randint
import seaborn as sns
from sklearn.utils import shuffle
import tensorflow as tf
from tensorflow.contrib.layers import flatten
# +
#Load the data
training_file = "train.p"
validation_file= "valid.p"
testing_file = "test.p"
with open(training_file, mode='rb') as file:
train = pickle.load(file)
with open(validation_file, mode='rb') as file:
valid = pickle.load(file)
with open(testing_file, mode='rb') as file:
test = pickle.load(file)
X_train_ori, y_train_ori = train['features'], train['labels']
#Create a array large enough to hold the new agumented images
#which will be created in the pre processing section
X_train = np.empty((3*X_train_ori.shape[0],X_train_ori.shape[1],X_train_ori.shape[2],X_train_ori.shape[3]))
y_train = np.empty((3*y_train_ori.shape[0]))
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# - `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# - `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
# - `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
# - `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
#
# Complete the basic data summary below. Use python, numpy and/or pandas methods to calculate the data summary rather than hard coding the results. For example, the [pandas shape method](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.shape.html) might be useful for calculating some of the summary results.
# ### Basic Summary of the Data Set
# +
#Number of original training examples
n_train_ori = X_train_ori.shape[0]
print("Number of original training examples =", n_train_ori)
# Number of training examples after image agumentation
n_train = X_train.shape[0]
print("Number of training examples =", n_train)
# Number of validation examples
n_validation = X_valid.shape[0]
print("Number of validation examples =", n_validation)
# Number of testing examples.
n_test = X_test.shape[0]
print("Number of testing examples =", n_test)
# Shape of an traffic sign image
image_shape = X_train.shape[1:]
print("Image data shape =", image_shape)
# Unique classes/labels there are in the dataset.
n_classes = len(set(y_train_ori))
print("Number of classes =", n_classes)
# -
# ### Include an exploratory visualization of the dataset
# +
### Data exploration visualization
# Visualizations will be shown in the notebook.
# %matplotlib inline
def plotTrafficSign(n_rows, n_cols):
"""
This function displays random images from the trainign data set.
"""
fig, axes = plt.subplots(nrows = n_rows, ncols = n_cols, figsize=(60,30))
for row in axes:
for col in row:
index = randint(0,n_train_ori)
col.imshow(X_train_ori[index,:,:,:])
col.set_title(y_train_ori[index])
#Plot traffic signs for visualization
plotTrafficSign(10, 5)
# -
#Plot distribution of data
sns.distplot(y_train_ori, kde=False, bins=n_classes)
sns.distplot(y_valid, kde=False, bins=n_classes)
sns.distplot(y_test, kde=False, bins=n_classes)
# Histogram of the data shows that the trainign data is unevenly distributed. This might affect the training of CNN model.
# Comparing the distribution across the 3 sets (training/validation/test), it seems that the distribution is similar in all the sets.
# ----
#
# ## Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
#
# With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
#
# There are various aspects to consider when thinking about this problem:
#
# - Neural network architecture (is the network over or underfitting?)
# - Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# - Number of examples per label (some have more than others).
# - Generate fake data.
#
# Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
# ### Pre-process the Data Set (normalization, grayscale, etc.)
# Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
#
# Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
# +
### Preprocess the data.
def dataGeneration():
"""
This function auguments the training data by creating new data (via image rotation)
"""
global X_train
global y_train
global y_train_ori
global X_train_ori
global n_train_ori
#Create new data by fliping the images in the vertical and horizontal directions
X_train[0:n_train_ori,:,:,:] = X_train_ori[:,:,:,:]
y_train[0:n_train_ori] = y_train_ori[:]
width = X_train.shape[1]
height = X_train.shape[2]
center = (width/ 2, height/ 2)
for index in range(n_train_ori):
#Rotate by 10 degrees
rotation = cv2.getRotationMatrix2D(center, 10, 1.0)
X_train[n_train_ori+index,:,:,:] = cv2.warpAffine(X_train_ori[index,:,:,:], rotation, (width, height))
y_train[n_train_ori+index] = y_train_ori[index]
#Flip the image horizontally
rotation = cv2.getRotationMatrix2D(center, -10, 1.0)
X_train[2*n_train_ori+index,:,:,:] = cv2.warpAffine(X_train_ori[index,:,:,:], rotation, (width, height))
y_train[2*n_train_ori+index] = y_train_ori[index]
def normalize(X_input):
"""
This function normalizes the data
"""
#Min-Max normalization of data
range_min = 0.1
range_max = 0.9
data_min = 0
data_max = 255
X_input = range_min + (((X_input - data_min)*(range_max - range_min) )/(data_max - data_min))
return X_input
def randomize(X_input, y_input):
"""
This function randomizes the data.
"""
#Randomize the data
X_input, y_input = shuffle(X_input, y_input)
return X_input, y_input
dataGeneration()
X_train = normalize(X_train)
X_valid = normalize(X_valid)
X_test = normalize(X_test)
X_train, y_train = randomize(X_train, y_train)
# -
# ### Model Architecture
def LeNet(x, keep_prob=1.0):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
global n_classes
# Layer 1: Convolutional. Input = 32x32x3. Output = 28x28x6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 3, 6), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(6))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
# Activation.
conv1 = tf.nn.relu(conv1)
# Pooling. Input = 28x28x6. Output = 14x14x6.
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#Dropout
conv1 = tf.nn.dropout(conv1, keep_prob)
# Layer 2: Convolutional. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
# Activation.
conv2 = tf.nn.relu(conv2)
# Pooling. Input = 10x10x16. Output = 5x5x16.
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#Dropout
conv2 = tf.nn.dropout(conv2, keep_prob)
# Flatten. Input = 5x5x16. Output = 400.
fc0 = flatten(conv2)
# Layer 3: Fully Connected. Input = 400. Output = 300.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 300), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(300))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# Activation.
fc1 = tf.nn.relu(fc1)
#Dropout
fc1 = tf.nn.dropout(fc1, keep_prob)
# Layer 4: Fully Connected. Input = 300. Output = 200.
fc2_W = tf.Variable(tf.truncated_normal(shape=(300, 200), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(200))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# Activation.
fc2 = tf.nn.relu(fc2)
#Dropout
fc2 = tf.nn.dropout(fc2, keep_prob)
# Layer 5: Fully Connected. Input = 200. Output = n_classes.
fc3_W = tf.Variable(tf.truncated_normal(shape=(200, n_classes), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(n_classes))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
class CModel:
def __init__(self, input_conv, target, learning_rate = 0.001,
epochs = 10, batch_size = 128, keep_prob=1.0, debug_logging = False):
"""
This is the ctor for the class CModel.
It initializes various hyper parameters required for training.
"""
self.learning_rate = learning_rate
self.epoch = epochs
self.batch_size = batch_size
self.debug_logging = debug_logging
self.input_conv = input_conv
self.target = target
self.logits = None
self.one_hot_out_class = None
self.keep_prob = keep_prob
def __loss(self):
"""
This function calculates the loss.
"""
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=self.one_hot_out_class, logits=self.logits)
loss_operation = tf.reduce_mean(cross_entropy)
return loss_operation
def __optimize(self, loss_operation):
"""
This function runs the optimizer to train the weights.
"""
optimizer = tf.train.AdamOptimizer(learning_rate = self.learning_rate)
minimize_loss = optimizer.minimize(loss_operation)
return minimize_loss
def trainLeNet(self):
"""
This function trains the LeNet network.
"""
print("n_classes ",n_classes)
self.logits = LeNet(self.input_conv,self.keep_prob)
self.one_hot_out_class = tf.one_hot(self.target, n_classes)
loss_operation = self.__loss()
minimize_loss = self.__optimize(loss_operation)
return minimize_loss
def accuracy(self):
"""
This function calculates the accuracy of the model.
"""
prediction, _ = self.prediction()
correct_prediction = tf.equal(prediction, tf.argmax(self.one_hot_out_class, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
return accuracy_operation
def prediction(self):
return tf.argmax(self.logits, 1), tf.nn.top_k(tf.nn.softmax(self.logits), k=5)
# ### Train, Validate and Test the Model
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
# sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
# +
#Model training
class CEvaluate:
def __init__(self, learning_rate=0.001, epoch=10, batch_size=128):
self.input_conv = tf.placeholder(tf.float32, (None, 32, 32, 3))
self.target = tf.placeholder(tf.int32, (None))
self.keep_prob = tf.placeholder(tf.float32)
self.model = CModel(self.input_conv, self.target, learning_rate, epoch, batch_size, self.keep_prob)
self.train = self.model.trainLeNet()
self.accuracy_operation = self.model.accuracy()
self.epoch = epoch
self.batch_size = batch_size
self.saver = tf.train.Saver()
self.prediction = self.model.prediction()
def __evaluate(self, X_data, y_data, keep_prob=1):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, self.batch_size):
batch_x, batch_y = X_data[offset:offset+self.batch_size], y_data[offset:offset+self.batch_size]
accuracy = sess.run(self.accuracy_operation, feed_dict={self.input_conv: batch_x, \
self.target: batch_y, self.keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
def test(self):
global X_test
global y_test
with tf.Session() as sess:
self.saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = self.__evaluate(X_test, y_test)
print("Test Accuracy = ", test_accuracy)
def predictions(self, test_images):
with tf.Session() as sess:
self.saver.restore(sess, './lenet')
predict, top_k_softmax = sess.run(self.prediction, feed_dict={self.input_conv: test_images, self.keep_prob: 1.0})
return predict, top_k_softmax
def run(self):
global X_train
global y_train
global X_valid
global y_valid
validation_accuracy = []
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
for i in range(self.epoch):
print("Epoch == ", i)
for offset in range(0, num_examples, self.batch_size):
end = offset + self.batch_size
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(self.train, feed_dict={self.input_conv: batch_x, self.target: batch_y, self.keep_prob: 0.9})
validation_accuracy.append(self.__evaluate(X_valid, y_valid))
print("Validation Accuracy == ", validation_accuracy[i])
self.saver.save(sess, './lenet')
plt.plot(validation_accuracy)
plt.xlabel("Epoch")
plt.ylabel("Validation Accuracy")
plt.title("Tracking of validation accuracy")
plt.show()
learning_rate = 0.001
epoch = 30
batch_size = 128
eval_model = CEvaluate(learning_rate, epoch, batch_size)
eval_model.run()
eval_model.test()
# -
# ---
#
# ## Step 3: Test a Model on New Images
#
# To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load and Output the Images
### Load the images and plot them.
import os
test_images = os.listdir('test_images')
num_test_images = 5
X_new_test = np.empty((num_test_images, 32, 32, 3))
y_new_test = np.empty(num_test_images)
dic = {"60.jpg":3, "70.jpg":4, "roadwork.jpg":25, "stop.jpg":14, "yield.jpg":13}
for index, image_name in enumerate(test_images):
image_path = os.path.join('test_images', image_name)
original_image = mpimg.imread(image_path)
X_new_test[index,:,:,:] = cv2.resize(original_image,(32,32),interpolation=cv2.INTER_AREA)
y_new_test[index] = dic[image_name]
plt.imshow(X_new_test[index,:,:,:])
plt.show()
# ### Predict the Sign Type for Each Image/Analyze Performance/ Output Soft Max
#
# +
with open('signnames.csv', mode='r') as file:
reader = csv.reader(file)
sign_mapping = {rows[0]:rows[1] for rows in reader}
X_new_test = normalize(X_new_test)
predict, top_k_softmax = eval_model.predictions(X_new_test)
for output,expected in zip(predict,y_new_test):
print("Expected {} ...... Output {}".format(sign_mapping[str(int(expected))], sign_mapping[str(output)]))
# +
### Calculate the accuracy for these 5 new images.
count = 0
for result, expectation in zip(predict, y_new_test):
if result == expectation:
count = count+1
accuracy = count/num_test_images
print("accuracy of the prediction of new test images", accuracy)
# -
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
print("top_k_softmax == ", top_k_softmax)
# ### Project Writeup
#
# Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ---
#
# ## Step 4 (Optional): Visualize the Neural Network's State with Test Images
#
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
#
# <figure>
# <img src="visualize_cnn.png" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> Your output should look something like this (above)</p>
# </figcaption>
# </figure>
# <p></p>
#
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
|
Traffic_Sign_Classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import copy
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(0)
pd.set_option('display.max_rows', None)
# ## Overview
# This notebook simulates the spread of an infection within a community that serves as a collection of private spaces (houses) and public spaces (landmarks). A random number of people belong to each house. Each simulation run represents a day. People leave their house at the start of each simulation run and visit one landmark during the day. Each individual may contract the virus by 1) being at a landmark where another infected person shows up or 2) from unknown "random" sources (for example, while communiting). At the end of each simulation run, all individuals return to their house. At this stage, an individual may be infected or exposed to the virus. If they are infected, they may infect other people in their house.
# ## Functions to execute distinct steps within a simulation run
# This function builds a dictionary of houses in our virtual community. It takes in (x,y) coordinates for each house, minimum and maximum number of people per house, and probability that each individual has the contact tracing app installed.
def build_houses_dict(houses_x, houses_y, min_household, max_household, app_penetration):
# initializing dictionary to store all houses
houses_dict = {}
# starting count for total number of people
total_people = 0
for i, coordinates in enumerate(zip(houses_x, houses_y)):
# initiliazing a dictionary for a specific house
houses_dict[i] = {}
# generating a random number of people in the house from a uniform distribution
num_people = np.random.randint(low=min_household, high=max_household)
total_people += num_people
houses_dict[i]['coordinates'] = coordinates
houses_dict[i]['num_people'] = num_people
# initialize house with no individual infected or exposed to the virus
# (and therefore no virus present in the house)
houses_dict[i]['infected'] = np.zeros(num_people, dtype=bool)
houses_dict[i]['exposed'] = np.zeros(num_people, dtype=bool)
houses_dict[i]['infection_present'] = False
# randomly determine whether each person has the contact tracing app installed
houses_dict[i]['has_app'] = np.random.choice([False, True],
p=[1-app_penetration, app_penetration],
size=num_people)
return houses_dict, total_people
# This function builds a dictionary of landmarks (proxy for public spaces) in our virtual community. It takes in (x,y) coordinates for each landmark.
def build_landmarks_dict(landmarks_x, landmarks_y):
# initializing dictionary to store all landmarks
landmarks_dict = {}
for i, coordinates in enumerate(zip(landmarks_x, landmarks_y)):
# initiliazing a dictionary for a specific landmark
landmarks_dict[i] = {}
landmarks_dict[i]['coordinates'] = coordinates
return landmarks_dict
# This function determines which individuals leave their house at the beginning of each simulation run (proxy for one day).
def build_individuals_in_dict(houses_dict, total_people, ct_compliance, leave_prob):
# initializing dictionary to store all individuals and their status at the beginning of each simulation run
individuals_in_dict = {}
counter = 0
# loop over each house
for i in houses_dict:
# find the number of people in this house
num_people = houses_dict[i]['num_people']
# loop over each individual in this house
for j in range(num_people):
individuals_in_dict[counter] = {}
individuals_in_dict[counter]['house_number'] = i
individuals_in_dict[counter]['index_number'] = j
individuals_in_dict[counter]['infected_in'] = houses_dict[i]['infected'][j]
individuals_in_dict[counter]['exposed_in'] = houses_dict[i]['exposed'][j]
individuals_in_dict[counter]['has_app'] = houses_dict[i]['has_app'][j]
# determine whether this individual leaves their house
leave_house = np.random.choice([False, True], p=[1-leave_prob, leave_prob])
# update leave house probabibility if they are exposed and have the app
if leave_house:
if individuals_in_dict[counter]['exposed_in'] and individuals_in_dict[counter]['has_app']:
leave_house = np.random.choice([False, True], p=[ct_compliance, 1-ct_compliance])
individuals_in_dict[counter]['leave_house'] = leave_house
# if this individual leaves their house, determine which landmark they go to
if leave_house:
individuals_in_dict[counter]['landmark_visit'] = np.random.randint(low=0, high=landmarks_n)
counter += 1
# verify we have looped over all individuals
assert total_people == counter
return individuals_in_dict
# This function simulates which individuals visit a given landmark and whether any infected individual visits the landmark.
def build_landmarks_visitors_dict(individuals_in_dict):
# initializing dictionary to store all visitors to each landmark
landmarks_visitors_dict = {}
# loop over all landmarks
for i in range(landmarks_n):
# initializing a dictionary for the specific landmark
# and assume infection is not present at the beginning of each simulation run
landmarks_visitors_dict[i] = {}
landmarks_visitors_dict[i]['visitors'] = []
landmarks_visitors_dict[i]['infection_present'] = False
# now loop over each individual
for i in individuals_in_dict:
# if this individual leaves their house, update the appropriate dictionary for the landmark they visit
if individuals_in_dict[i]['leave_house']:
landmark_visit = individuals_in_dict[i]['landmark_visit']
landmarks_visitors_dict[landmark_visit]['visitors'].append(i)
# if this individual is infected, recognize that infection is present at the landmark they visit
if individuals_in_dict[i]['infected_in']:
landmarks_visitors_dict[landmark_visit]['infection_present'] = True
return landmarks_visitors_dict
# This function simulates which individuals get infected and which individuals are exposed to the virus based on the landmark they visit.
def build_individuals_out_dict(landmarks_visitors_dict, individuals_in_dict,
houses_dict, public_spread, random_spread):
# initializing dictionary to store all individuals and their status at the end of each simulation run
# by making a copy of the previous dictionary
individuals_out_dict = copy.deepcopy(individuals_in_dict)
# loop over each individual
for i in individuals_out_dict:
if individuals_out_dict[i]['leave_house']:
landmark_visit = individuals_out_dict[i]['landmark_visit']
# if this individual was exposed earlier, they are also exposed now
if individuals_out_dict[i]['exposed_in']:
individuals_out_dict[i]['exposed_out'] = True
# otherwise they get exposed when they visit a landmark where infection was present
else:
individuals_out_dict[i]['exposed_out'] = landmarks_visitors_dict[landmark_visit]['infection_present']
# if this individual was infected earlier, they are also infected now
if individuals_out_dict[i]['infected_in']:
individuals_out_dict[i]['infected_out'] = True
# if this individual got exposed by visiting a landmark where infection was present,
# they have a probability of contracting the infection
elif individuals_out_dict[i]['exposed_out']:
individuals_out_dict[i]['infected_out'] = np.random.choice([False, True],
p=[1-public_spread, public_spread])
# otherwise there is a probability of contracting the virus from unknown sources
# (perhaps while communiting)
else:
individuals_out_dict[i]['infected_out'] = np.random.choice([False, True],
p=[1-random_spread, random_spread])
# if the individual was exposed or infected, also update their status in their house's dictionary
house_number = individuals_out_dict[i]['house_number']
index_number = individuals_out_dict[i]['index_number']
if individuals_out_dict[i]['infected_out']:
individuals_out_dict[i]['exposed_out'] = True
houses_dict[house_number]['infected'][index_number] = True
houses_dict[house_number]['infection_present'] = True
if individuals_out_dict[i]['exposed_out']:
houses_dict[house_number]['exposed'][index_number] = True
return individuals_out_dict
# This function determines whether an infected individual ends up infecting other people in their house.
def update_houses_dict(houses_dict, house_spread):
# loop over each house
for i in houses_dict:
# loop over each individual in the house
for j in range(houses_dict[i]['num_people']):
# if there is infection present in the house, everyone is exposed
if houses_dict[i]['infection_present']:
houses_dict[i]['exposed'][j] = True
# all un-infected individuals have a probability of contracting the virus
if houses_dict[i]['infected'][j] == False:
houses_dict[i]['infected'][j] = np.random.choice([False, True],
p=[1-house_spread, house_spread])
return houses_dict
# ## Function for running the simulation
def run_simulation(num_days, houses_n, houses_x, houses_y, landmarks_n, landmarks_x, landmarks_y,
min_household, max_household, app_penetration, ct_compliance, leave_prob,
public_spread, random_spread, house_spread, display_dicts):
# start by building dictionaries for houses and landmarks
houses_dict, total_people = build_houses_dict(houses_x, houses_y, min_household, max_household, app_penetration)
landmarks_dict = build_landmarks_dict(landmarks_x, landmarks_y)
# initialize list for storing number of people infected after each simulation run
people_infected = []
# loop over each simulation run
for i in range(num_days):
# for debugging
if display_dicts:
print(f"\nhouses_day{i}")
display(pd.DataFrame(houses_dict).transpose())
# determine who leaves their house
individuals_in_dict = build_individuals_in_dict(houses_dict, total_people,
ct_compliance, leave_prob)
# determine which individuals visit each landmark
landmarks_visitors_dict = build_landmarks_visitors_dict(individuals_in_dict)
# determine who is exposed or infected in this simulation run due to the landmarks they visit
individuals_out_dict = build_individuals_out_dict(landmarks_visitors_dict, individuals_in_dict,
houses_dict, public_spread, random_spread)
# determine who gets exposed or infected in their house at the end of the day
houses_dict = update_houses_dict(houses_dict, house_spread)
# update list with number of people infected at the end of this simulation run
people_infected_temp = 0
for j in houses_dict:
people_infected_temp += sum(houses_dict[j]['infected'])
people_infected.append(people_infected_temp)
# for debugging
if display_dicts:
print(f"\nindividuals_in_day{i}")
display(pd.DataFrame(individuals_in_dict).transpose())
print(f"\nlandmarks_visitors_day{i}")
display(pd.DataFrame(landmarks_visitors_dict).transpose())
print(f"\nindividuals_out_day{i}")
display(pd.DataFrame(individuals_out_dict).transpose())
return houses_dict, total_people, people_infected
# ## Simulation runs
# +
houses_n = 1000
houses1_x = np.random.uniform(0, 800, houses_n//4)
houses1_y = np.random.uniform(0, 200, houses_n//4)
houses2_x = np.random.uniform(800, 1000, houses_n//4)
houses2_y = np.random.uniform(0, 800, houses_n//4)
houses3_x = np.random.uniform(200, 1000, houses_n//4)
houses3_y = np.random.uniform(800, 1000, houses_n//4)
houses4_x = np.random.uniform(0, 200, houses_n//4)
houses4_y = np.random.uniform(200, 1000, houses_n//4)
houses_x = np.concatenate([houses1_x, houses2_x, houses3_x, houses4_x])
houses_y = np.concatenate([houses1_y, houses2_y, houses3_y, houses4_y])
landmarks_n = 10
landmarks_x = np.random.uniform(300, 700, landmarks_n)
landmarks_y = np.random.uniform(300, 700, landmarks_n)
num_days = 20
houses_dict, total_people, people_infected = run_simulation(num_days=num_days,
houses_n=houses_n,
houses_x=houses_x,
houses_y=houses_y,
landmarks_n=landmarks_n,
landmarks_x=landmarks_x,
landmarks_y=landmarks_y,
min_household=1,
max_household=5,
app_penetration=0.20,
ct_compliance=0.80,
leave_prob=0.60,
public_spread=0.10,
random_spread=0.02,
house_spread=0.50,
display_dicts=False)
# +
days = list(range(num_days))
infected_share = [n/total_people for n in people_infected]
plt.figure(figsize=(20,10))
plt.plot(days, infected_share)
plt.xlabel('days', fontsize=16)
plt.ylabel('share of people infected', fontsize=16)
plt.show()
# -
# ## Grid search 1
# +
# keep number of houses below 100 and landmarks below 10 for quick execution while testing
houses_n = 10000
houses1_x = np.random.uniform(0, 800, houses_n//4)
houses1_y = np.random.uniform(0, 200, houses_n//4)
houses2_x = np.random.uniform(800, 1000, houses_n//4)
houses2_y = np.random.uniform(0, 800, houses_n//4)
houses3_x = np.random.uniform(200, 1000, houses_n//4)
houses3_y = np.random.uniform(800, 1000, houses_n//4)
houses4_x = np.random.uniform(0, 200, houses_n//4)
houses4_y = np.random.uniform(200, 1000, houses_n//4)
houses_x = np.concatenate([houses1_x, houses2_x, houses3_x, houses4_x])
houses_y = np.concatenate([houses1_y, houses2_y, houses3_y, houses4_y])
landmarks_n = 50
landmarks_x = np.random.uniform(300, 700, landmarks_n)
landmarks_y = np.random.uniform(300, 700, landmarks_n)
# +
# %%time
steps = 11
num_days = 20
app_penetration_grid = np.linspace(start=0, stop=1, num=steps)
ct_compliance_grid = np.linspace(start=0, stop=1, num=steps)
total_people_array = np.zeros((steps, steps))
people_infected_array = np.zeros((steps, steps, num_days))
for i, p_app in enumerate(app_penetration_grid):
for j, p_comp in enumerate(ct_compliance_grid):
houses_dict, total_people, people_infected = run_simulation(num_days=num_days,
houses_n=houses_n,
houses_x=houses_x,
houses_y=houses_y,
landmarks_n=landmarks_n,
landmarks_x=landmarks_x,
landmarks_y=landmarks_y,
min_household=1,
max_household=5,
app_penetration=p_app,
ct_compliance=p_comp,
leave_prob=0.35,
public_spread=0.02,
random_spread=0.005,
house_spread=0.5,
display_dicts=False)
total_people_array[i,j] = total_people
people_infected_array[i,j,:] = people_infected
# +
infected_share = np.divide(people_infected_array[:,:,-1], total_people_array)
plt.figure(figsize=(20,10))
plt.contourf(app_penetration_grid, ct_compliance_grid, np.transpose(infected_share),
cmap='Reds')
plt.title(f'infected share after {num_days} days', fontsize=16)
plt.xlabel('app_penetration', fontsize=16)
plt.ylabel('ct_compliance', fontsize=16)
plt.colorbar()
plt.show()
# -
# ## Grid search 2
# +
# Urban
# New York City, New York
# 116 Historic Landmarks - https://en.wikipedia.org/wiki/List_of_National_Historic_Landmarks_in_New_York_City
# 3,154,103 Households - https://www.census.gov/quickfacts/fact/table/newyorkcitynewyork/HSD410218
print(f"New York {int((3154103/116)/100)}x")
# Sprawl
# Austin, Texas
# 2 Historic Landmarks - https://en.wikipedia.org/wiki/List_of_National_Historic_Landmarks_in_Texas
# 370,043 Households - https://www.census.gov/quickfacts/fact/table/austincitytexas/PST045219
print(f"Austin {int((370043/2)/100)}x")
# Average
# Omaha, Nebraska
# 3 Historic Landmarks - https://en.wikipedia.org/wiki/List_of_National_Historic_Landmarks_in_Nebraska
# 183,160 Households - https://www.census.gov/quickfacts/fact/table/omahacitynebraska,US/PST045219
print(f"Omaha {int((183160/3)/100)}x")
# +
houses_n = 500
houses1_x = np.random.uniform(0, 800, houses_n//4)
houses1_y = np.random.uniform(0, 200, houses_n//4)
houses2_x = np.random.uniform(800, 1000, houses_n//4)
houses2_y = np.random.uniform(0, 800, houses_n//4)
houses3_x = np.random.uniform(200, 1000, houses_n//4)
houses3_y = np.random.uniform(800, 1000, houses_n//4)
houses4_x = np.random.uniform(0, 200, houses_n//4)
houses4_y = np.random.uniform(200, 1000, houses_n//4)
houses_x = np.concatenate([houses1_x, houses2_x, houses3_x, houses4_x])
houses_y = np.concatenate([houses1_y, houses2_y, houses3_y, houses4_y])
landmarks_n = 50
landmarks_x = np.random.uniform(300, 700, landmarks_n)
landmarks_y = np.random.uniform(300, 700, landmarks_n)
# +
# %%time
steps = 11
num_days = 30
carriers = 0.02 # MedicalNewsToday states 1-3% of people aymptomatic carriers
surface = 0.005 # Small chance of extended time surface transmission
proximate_spread = 0.5 # Change you'll get it from housemates in close proximity
leave_home = 0.35 # Probability you'll leave your home on any day (if you don't have the app)
# Quarantine:
# 35% because 64.6% average drop in mobility in France (mandatory travel restrictions) according to Google Mobility
# https://www.gstatic.com/covid19/mobility/2020-04-17_FR_Mobility_Report_en.pdf
# 95% chance you leave your home on normal day
# 75% chance for city/density studies, since more people will stay out of fear
app_penetration_grid = np.linspace(start=0, stop=1, num=steps)
ct_compliance_grid = np.linspace(start=0, stop=1, num=steps)
total_people_array = np.zeros((steps, steps))
people_infected_array = np.zeros((steps, steps, num_days))
for i, p_app in enumerate(app_penetration_grid):
for j, p_comp in enumerate(ct_compliance_grid):
houses_dict, total_people, people_infected = run_simulation(num_days=num_days,
houses_n=houses_n,
houses_x=houses_x,
houses_y=houses_y,
landmarks_n=landmarks_n,
landmarks_x=landmarks_x,
landmarks_y=landmarks_y,
min_household=1,
max_household=5,
app_penetration=p_app,
ct_compliance=p_comp,
leave_prob=leave_home,
public_spread=carriers,
random_spread=surface,
house_spread=proximate_spread,
display_dicts=False)
total_people_array[i,j] = total_people
people_infected_array[i,j,:] = people_infected
# +
infected_share = np.divide(people_infected_array[:,:,-1], total_people_array)
plt.figure(figsize=(20,10))
plt.contourf(app_penetration_grid, ct_compliance_grid, np.transpose(infected_share),
cmap="Reds") # vmin=0.2, vmax=0.5
plt.title(f'City infected share with stay-at-home orders after {num_days} days')
plt.xlabel('App Penetration (pct of population)')
plt.ylabel('Contact Tracing Compliance (pct of population)')
plt.colorbar()
plt.show()
# -
# ## For debugging
# +
houses_n = 10
houses1_x = np.random.uniform(0, 800, houses_n//4)
houses1_y = np.random.uniform(0, 200, houses_n//4)
houses2_x = np.random.uniform(800, 1000, houses_n//4)
houses2_y = np.random.uniform(0, 800, houses_n//4)
houses3_x = np.random.uniform(200, 1000, houses_n//4)
houses3_y = np.random.uniform(800, 1000, houses_n//4)
houses4_x = np.random.uniform(0, 200, houses_n//4)
houses4_y = np.random.uniform(200, 1000, houses_n//4)
houses_x = np.concatenate([houses1_x, houses2_x, houses3_x, houses4_x])
houses_y = np.concatenate([houses1_y, houses2_y, houses3_y, houses4_y])
landmarks_n = 4
landmarks_x = np.random.uniform(300, 700, landmarks_n)
landmarks_y = np.random.uniform(300, 700, landmarks_n)
num_days = 3
houses_dict, _, _ = run_simulation(num_days=num_days,
houses_n=houses_n,
houses_x=houses_x,
houses_y=houses_y,
landmarks_n=landmarks_n,
landmarks_x=landmarks_x,
landmarks_y=landmarks_y,
min_household=1,
max_household=3,
app_penetration=1,
ct_compliance=0,
leave_prob=0.5,
public_spread=0.20,
random_spread=0.10,
house_spread=0.50,
display_dicts=True)
# -
# ## Appendix
# +
# leave_prob_exposed vs. leave_prob_not_exposed mesh plot for different values of public_spread
# leave_prob_exposed vs. leave_prob_not_exposed (false positives) mesh plot for different values of public_spread
# low density, medium density, high density
# +
# Notes:
# input: accuracy of data
#
# output 1: spread and peak of spread
# output 2: false positive / false negative
# output 3: tracking spread of infection across geography
# +
# f = open(f"data/individuals_in_dict_day{i}.txt","w"); f.write(str(individuals_in_dict)); f.close()
# f = open(f"data/landmarks_visitors_dict_day{i}.txt","w"); f.write(str(landmarks_visitors_dict)); f.close()
# f = open(f"data/individuals_out_dict_day{i}.txt","w"); f.write(str(individuals_out_dict)); f.close()
# f = open(f"data/houses_dict_day{i}.txt","w"); f.write(str(houses_dict)); f.close()
# f = open(f"data/houses_dict_day{i}.csv","w"); w = csv.writer(f); w.writerows(houses_dict.items()); f.close()
# f = open(f"data/individuals_in_dict_day{i}.csv","w"); w = csv.writer(f); w.writerows(individuals_in_dict.items()); f.close()
# f = open(f"data/landmarks_visitors_dict_day{i}.csv","w"); w = csv.writer(f); w.writerows(landmarks_visitors_dict.items()); f.close()
# f = open(f"data/individuals_out_dict_day{i}.csv","w"); w = csv.writer(f); w.writerows(individuals_out_dict.items()); f.close()
# +
# plt.figure(figsize=(20,10))
# plt.scatter(houses_x, houses_y, label="houses")
# plt.scatter(landmarks_x, landmarks_y, color="red", label="landmarks")
# plt.legend()
# plt.show()
# +
# days = list(range(num_days))
# plt.figure(figsize=(20,10))
# for i, p1 in enumerate(ct_compliance_grid):
# for j, p2 in enumerate(leave_prob_grid):
# infected_share = [n/total_people for n in people_infected_array[i,j,:]]
# plt.plot(days, infected_share)
# plt.xlabel('days')
# plt.ylabel('share of people infected')
# plt.show()
# +
# min_household = 1
# max_household = 5
# app_penetration = 0.2
# ct_compliance = 0.20
# leave_prob = 0.5
# public_spread = 0.10
# random_spread = 0.02
# house_spread = 0.50
# num_days = 20
# display_dicts = False
# houses_dict, total_people, people_infected = run_simulation(num_days, houses_n, houses_x, houses_y,
# landmarks_n, landmarks_x, landmarks_y,
# min_household, max_household, app_penetration,
# ct_compliance, leave_prob,
# public_spread, random_spread, house_spread,
# display_dicts)
# days = list(range(num_days))
# infected_share = [n/total_people for n in people_infected]
# plt.figure(figsize=(20,10))
# plt.plot(days, infected_share)
# plt.xlabel('days')
# plt.ylabel('share of people infected')
# plt.show()
# +
# min_household = 1
# max_household = 5
# app_penetration = 0.2
# ct_compliance = 0.05
# leave_prob = 0.5
# public_spread = 0.10
# random_spread = 0.02
# house_spread = 0.50
# num_days = 20
# display_dicts = False
# houses_dict, total_people, people_infected = run_simulation(num_days, houses_n, houses_x, houses_y,
# landmarks_n, landmarks_x, landmarks_y,
# min_household, max_household, app_penetration,
# ct_compliance, leave_prob,
# public_spread, random_spread, house_spread,
# display_dicts)
# days = list(range(num_days))
# infected_share = [n/total_people for n in people_infected]
# plt.figure(figsize=(20,10))
# plt.plot(days, infected_share)
# plt.xlabel('days')
# plt.ylabel('share of people infected')
# plt.show()
|
simulation/ContactTracingSimulation_v2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="s_qNSzzyaCbD"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="jmjh290raIky"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="J0Qjg6vuaHNt"
# # Neural machine translation with attention
# + [markdown] colab_type="text" id="AOpGoE2T-YXS"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/text/nmt_with_attention">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/nmt_with_attention.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/nmt_with_attention.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/nmt_with_attention.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="CiwtNgENbx2g"
# This notebook trains a sequence to sequence (seq2seq) model for Spanish to English translation. This is an advanced example that assumes some knowledge of sequence to sequence models.
#
# After training the model in this notebook, you will be able to input a Spanish sentence, such as *"¿todavia estan en casa?"*, and return the English translation: *"are you still at home?"*
#
# The translation quality is reasonable for a toy example, but the generated attention plot is perhaps more interesting. This shows which parts of the input sentence has the model's attention while translating:
#
# <img src="https://tensorflow.org/images/spanish-english.png" alt="spanish-english attention plot">
#
# Note: This example takes approximately 10 mintues to run on a single P100 GPU.
# + colab={} colab_type="code" id="tnxXKDjq3jEL"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from sklearn.model_selection import train_test_split
import unicodedata
import re
import numpy as np
import os
import io
import time
# + [markdown] colab_type="text" id="wfodePkj3jEa"
# ## Download and prepare the dataset
#
# We'll use a language dataset provided by http://www.manythings.org/anki/. This dataset contains language translation pairs in the format:
#
# ```
# May I borrow this book? ¿Puedo tomar prestado este libro?
# ```
#
# There are a variety of languages available, but we'll use the English-Spanish dataset. For convenience, we've hosted a copy of this dataset on Google Cloud, but you can also download your own copy. After downloading the dataset, here are the steps we'll take to prepare the data:
#
# 1. Add a *start* and *end* token to each sentence.
# 2. Clean the sentences by removing special characters.
# 3. Create a word index and reverse word index (dictionaries mapping from word → id and id → word).
# 4. Pad each sentence to a maximum length.
# + colab={} colab_type="code" id="kRVATYOgJs1b"
# Download the file
path_to_zip = tf.keras.utils.get_file(
'spa-eng.zip', origin='http://storage.googleapis.com/download.tensorflow.org/data/spa-eng.zip',
extract=True)
path_to_file = os.path.dirname(path_to_zip)+"/spa-eng/spa.txt"
# + colab={} colab_type="code" id="rd0jw-eC3jEh"
# Converts the unicode file to ascii
def unicode_to_ascii(s):
return ''.join(c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn')
def preprocess_sentence(w):
w = unicode_to_ascii(w.lower().strip())
# creating a space between a word and the punctuation following it
# eg: "he is a boy." => "he is a boy ."
# Reference:- https://stackoverflow.com/questions/3645931/python-padding-punctuation-with-white-spaces-keeping-punctuation
w = re.sub(r"([?.!,¿])", r" \1 ", w)
w = re.sub(r'[" "]+', " ", w)
# replacing everything with space except (a-z, A-Z, ".", "?", "!", ",")
w = re.sub(r"[^a-zA-Z?.!,¿]+", " ", w)
w = w.rstrip().strip()
# adding a start and an end token to the sentence
# so that the model know when to start and stop predicting.
w = '<start> ' + w + ' <end>'
return w
# + colab={} colab_type="code" id="opI2GzOt479E"
en_sentence = u"May I borrow this book?"
sp_sentence = u"¿Puedo tomar prestado este libro?"
print(preprocess_sentence(en_sentence))
print(preprocess_sentence(sp_sentence).encode('utf-8'))
# + colab={} colab_type="code" id="OHn4Dct23jEm"
# 1. Remove the accents
# 2. Clean the sentences
# 3. Return word pairs in the format: [ENGLISH, SPANISH]
def create_dataset(path, num_examples):
lines = io.open(path, encoding='UTF-8').read().strip().split('\n')
word_pairs = [[preprocess_sentence(w) for w in l.split('\t')] for l in lines[:num_examples]]
return zip(*word_pairs)
# + colab={} colab_type="code" id="cTbSbBz55QtF"
en, sp = create_dataset(path_to_file, None)
print(en[-1])
print(sp[-1])
# + colab={} colab_type="code" id="OmMZQpdO60dt"
def max_length(tensor):
return max(len(t) for t in tensor)
# + colab={} colab_type="code" id="bIOn8RCNDJXG"
def tokenize(lang):
lang_tokenizer = tf.keras.preprocessing.text.Tokenizer(
filters='')
lang_tokenizer.fit_on_texts(lang)
tensor = lang_tokenizer.texts_to_sequences(lang)
tensor = tf.keras.preprocessing.sequence.pad_sequences(tensor,
padding='post')
return tensor, lang_tokenizer
# + colab={} colab_type="code" id="eAY9k49G3jE_"
def load_dataset(path, num_examples=None):
# creating cleaned input, output pairs
targ_lang, inp_lang = create_dataset(path, num_examples)
input_tensor, inp_lang_tokenizer = tokenize(inp_lang)
target_tensor, targ_lang_tokenizer = tokenize(targ_lang)
return input_tensor, target_tensor, inp_lang_tokenizer, targ_lang_tokenizer
# + [markdown] colab_type="text" id="GOi42V79Ydlr"
# ### Limit the size of the dataset to experiment faster (optional)
#
# Training on the complete dataset of >100,000 sentences will take a long time. To train faster, we can limit the size of the dataset to 30,000 sentences (of course, translation quality degrades with less data):
# + colab={} colab_type="code" id="cnxC7q-j3jFD"
# Try experimenting with the size of that dataset
num_examples = 30000
input_tensor, target_tensor, inp_lang, targ_lang = load_dataset(path_to_file, num_examples)
# Calculate max_length of the target tensors
max_length_targ, max_length_inp = max_length(target_tensor), max_length(input_tensor)
# + colab={} colab_type="code" id="4QILQkOs3jFG"
# Creating training and validation sets using an 80-20 split
input_tensor_train, input_tensor_val, target_tensor_train, target_tensor_val = train_test_split(input_tensor, target_tensor, test_size=0.2)
# Show length
print(len(input_tensor_train), len(target_tensor_train), len(input_tensor_val), len(target_tensor_val))
# + colab={} colab_type="code" id="lJPmLZGMeD5q"
def convert(lang, tensor):
for t in tensor:
if t!=0:
print ("%d ----> %s" % (t, lang.index_word[t]))
# + colab={} colab_type="code" id="VXukARTDd7MT"
print ("Input Language; index to word mapping")
convert(inp_lang, input_tensor_train[0])
print ()
print ("Target Language; index to word mapping")
convert(targ_lang, target_tensor_train[0])
# + [markdown] colab_type="text" id="rgCLkfv5uO3d"
# ### Create a tf.data dataset
# + colab={} colab_type="code" id="TqHsArVZ3jFS"
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
steps_per_epoch = len(input_tensor_train)//BATCH_SIZE
embedding_dim = 256
units = 1024
vocab_inp_size = len(inp_lang.word_index)+1
vocab_tar_size = len(targ_lang.word_index)+1
dataset = tf.data.Dataset.from_tensor_slices((input_tensor_train, target_tensor_train)).shuffle(BUFFER_SIZE)
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
# + colab={} colab_type="code" id="qc6-NK1GtWQt"
example_input_batch, example_target_batch = next(iter(dataset))
example_input_batch.shape, example_target_batch.shape
# + [markdown] colab_type="text" id="TNfHIF71ulLu"
# ## Write the encoder and decoder model
#
# Implement an encoder-decoder model with attention which you can read about in the TensorFlow [Neural Machine Translation (seq2seq) tutorial](https://github.com/tensorflow/nmt). This example uses a more recent set of APIs. This notebook implements the [attention equations](https://github.com/tensorflow/nmt#background-on-the-attention-mechanism) from the seq2seq tutorial. The following diagram shows that each input words is assigned a weight by the attention mechanism which is then used by the decoder to predict the next word in the sentence. The below picture and formulas are an example of attention mechanism from [Luong's paper](https://arxiv.org/abs/1508.04025v5).
#
# <img src="https://www.tensorflow.org/images/seq2seq/attention_mechanism.jpg" width="500" alt="attention mechanism">
#
# The input is put through an encoder model which gives us the encoder output of shape *(batch_size, max_length, hidden_size)* and the encoder hidden state of shape *(batch_size, hidden_size)*.
#
# Here are the equations that are implemented:
#
# <img src="https://www.tensorflow.org/images/seq2seq/attention_equation_0.jpg" alt="attention equation 0" width="800">
# <img src="https://www.tensorflow.org/images/seq2seq/attention_equation_1.jpg" alt="attention equation 1" width="800">
#
# This tutorial uses [Bahdanau attention](https://arxiv.org/pdf/1409.0473.pdf) for the encoder. Let's decide on notation before writing the simplified form:
#
# * FC = Fully connected (dense) layer
# * EO = Encoder output
# * H = hidden state
# * X = input to the decoder
#
# And the pseudo-code:
#
# * `score = FC(tanh(FC(EO) + FC(H)))`
# * `attention weights = softmax(score, axis = 1)`. Softmax by default is applied on the last axis but here we want to apply it on the *1st axis*, since the shape of score is *(batch_size, max_length, hidden_size)*. `Max_length` is the length of our input. Since we are trying to assign a weight to each input, softmax should be applied on that axis.
# * `context vector = sum(attention weights * EO, axis = 1)`. Same reason as above for choosing axis as 1.
# * `embedding output` = The input to the decoder X is passed through an embedding layer.
# * `merged vector = concat(embedding output, context vector)`
# * This merged vector is then given to the GRU
#
# The shapes of all the vectors at each step have been specified in the comments in the code:
# + colab={} colab_type="code" id="nZ2rI24i3jFg"
class Encoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.enc_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state = hidden)
return output, state
def initialize_hidden_state(self):
return tf.zeros((self.batch_sz, self.enc_units))
# + colab={} colab_type="code" id="60gSVh05Jl6l"
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
# sample input
sample_hidden = encoder.initialize_hidden_state()
sample_output, sample_hidden = encoder(example_input_batch, sample_hidden)
print ('Encoder output shape: (batch size, sequence length, units) {}'.format(sample_output.shape))
print ('Encoder Hidden state shape: (batch size, units) {}'.format(sample_hidden.shape))
# + colab={} colab_type="code" id="umohpBN2OM94"
class BahdanauAttention(tf.keras.layers.Layer):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, query, values):
# hidden shape == (batch_size, hidden size)
# hidden_with_time_axis shape == (batch_size, 1, hidden size)
# we are doing this to perform addition to calculate the score
hidden_with_time_axis = tf.expand_dims(query, 1)
# score shape == (batch_size, max_length, 1)
# we get 1 at the last axis because we are applying score to self.V
# the shape of the tensor before applying self.V is (batch_size, max_length, units)
score = self.V(tf.nn.tanh(
self.W1(values) + self.W2(hidden_with_time_axis)))
# attention_weights shape == (batch_size, max_length, 1)
attention_weights = tf.nn.softmax(score, axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * values
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
# + colab={} colab_type="code" id="k534zTHiDjQU"
attention_layer = BahdanauAttention(10)
attention_result, attention_weights = attention_layer(sample_hidden, sample_output)
print("Attention result shape: (batch size, units) {}".format(attention_result.shape))
print("Attention weights shape: (batch_size, sequence_length, 1) {}".format(attention_weights.shape))
# + colab={} colab_type="code" id="yJ_B3mhW3jFk"
class Decoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.dec_units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc = tf.keras.layers.Dense(vocab_size)
# used for attention
self.attention = BahdanauAttention(self.dec_units)
def call(self, x, hidden, enc_output):
# enc_output shape == (batch_size, max_length, hidden_size)
context_vector, attention_weights = self.attention(hidden, enc_output)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# output shape == (batch_size * 1, hidden_size)
output = tf.reshape(output, (-1, output.shape[2]))
# output shape == (batch_size, vocab)
x = self.fc(output)
return x, state, attention_weights
# + colab={} colab_type="code" id="P5UY8wko3jFp"
decoder = Decoder(vocab_tar_size, embedding_dim, units, BATCH_SIZE)
sample_decoder_output, _, _ = decoder(tf.random.uniform((64, 1)),
sample_hidden, sample_output)
print ('Decoder output shape: (batch_size, vocab size) {}'.format(sample_decoder_output.shape))
# + [markdown] colab_type="text" id="_ch_71VbIRfK"
# ## Define the optimizer and the loss function
# + colab={} colab_type="code" id="WmTHr5iV3jFr"
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
# + [markdown] colab_type="text" id="DMVWzzsfNl4e"
# ## Checkpoints (Object-based saving)
# + colab={} colab_type="code" id="Zj8bXQTgNwrF"
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(optimizer=optimizer,
encoder=encoder,
decoder=decoder)
# + [markdown] colab_type="text" id="hpObfY22IddU"
# ## Training
#
# 1. Pass the *input* through the *encoder* which return *encoder output* and the *encoder hidden state*.
# 2. The encoder output, encoder hidden state and the decoder input (which is the *start token*) is passed to the decoder.
# 3. The decoder returns the *predictions* and the *decoder hidden state*.
# 4. The decoder hidden state is then passed back into the model and the predictions are used to calculate the loss.
# 5. Use *teacher forcing* to decide the next input to the decoder.
# 6. *Teacher forcing* is the technique where the *target word* is passed as the *next input* to the decoder.
# 7. The final step is to calculate the gradients and apply it to the optimizer and backpropagate.
# + colab={} colab_type="code" id="sC9ArXSsVfqn"
@tf.function
def train_step(inp, targ, enc_hidden):
loss = 0
with tf.GradientTape() as tape:
enc_output, enc_hidden = encoder(inp, enc_hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']] * BATCH_SIZE, 1)
# Teacher forcing - feeding the target as the next input
for t in range(1, targ.shape[1]):
# passing enc_output to the decoder
predictions, dec_hidden, _ = decoder(dec_input, dec_hidden, enc_output)
loss += loss_function(targ[:, t], predictions)
# using teacher forcing
dec_input = tf.expand_dims(targ[:, t], 1)
batch_loss = (loss / int(targ.shape[1]))
variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(gradients, variables))
return batch_loss
# + colab={} colab_type="code" id="ddefjBMa3jF0"
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
enc_hidden = encoder.initialize_hidden_state()
total_loss = 0
for (batch, (inp, targ)) in enumerate(dataset.take(steps_per_epoch)):
batch_loss = train_step(inp, targ, enc_hidden)
total_loss += batch_loss
if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch + 1,
batch,
batch_loss.numpy()))
# saving (checkpoint) the model every 2 epochs
if (epoch + 1) % 2 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print('Epoch {} Loss {:.4f}'.format(epoch + 1,
total_loss / steps_per_epoch))
print('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
# + [markdown] colab_type="text" id="mU3Ce8M6I3rz"
# ## Translate
#
# * The evaluate function is similar to the training loop, except we don't use *teacher forcing* here. The input to the decoder at each time step is its previous predictions along with the hidden state and the encoder output.
# * Stop predicting when the model predicts the *end token*.
# * And store the *attention weights for every time step*.
#
# Note: The encoder output is calculated only once for one input.
# + colab={} colab_type="code" id="EbQpyYs13jF_"
def evaluate(sentence):
attention_plot = np.zeros((max_length_targ, max_length_inp))
sentence = preprocess_sentence(sentence)
inputs = [inp_lang.word_index[i] for i in sentence.split(' ')]
inputs = tf.keras.preprocessing.sequence.pad_sequences([inputs],
maxlen=max_length_inp,
padding='post')
inputs = tf.convert_to_tensor(inputs)
result = ''
hidden = [tf.zeros((1, units))]
enc_out, enc_hidden = encoder(inputs, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word_index['<start>']], 0)
for t in range(max_length_targ):
predictions, dec_hidden, attention_weights = decoder(dec_input,
dec_hidden,
enc_out)
# storing the attention weights to plot later on
attention_weights = tf.reshape(attention_weights, (-1, ))
attention_plot[t] = attention_weights.numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result += targ_lang.index_word[predicted_id] + ' '
if targ_lang.index_word[predicted_id] == '<end>':
return result, sentence, attention_plot
# the predicted ID is fed back into the model
dec_input = tf.expand_dims([predicted_id], 0)
return result, sentence, attention_plot
# + colab={} colab_type="code" id="s5hQWlbN3jGF"
# function for plotting the attention weights
def plot_attention(attention, sentence, predicted_sentence):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attention, cmap='viridis')
fontdict = {'fontsize': 14}
ax.set_xticklabels([''] + sentence, fontdict=fontdict, rotation=90)
ax.set_yticklabels([''] + predicted_sentence, fontdict=fontdict)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
# + colab={} colab_type="code" id="sl9zUHzg3jGI"
def translate(sentence):
result, sentence, attention_plot = evaluate(sentence)
print('Input: %s' % (sentence))
print('Predicted translation: {}'.format(result))
attention_plot = attention_plot[:len(result.split(' ')), :len(sentence.split(' '))]
plot_attention(attention_plot, sentence.split(' '), result.split(' '))
# + [markdown] colab_type="text" id="n250XbnjOaqP"
# ## Restore the latest checkpoint and test
# + colab={} colab_type="code" id="UJpT9D5_OgP6"
# restoring the latest checkpoint in checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# + colab={} colab_type="code" id="WrAM0FDomq3E"
translate(u'hace mucho frio aqui.')
# + colab={} colab_type="code" id="zSx2iM36EZQZ"
translate(u'esta es mi vida.')
# + colab={} colab_type="code" id="A3LLCx3ZE0Ls"
translate(u'¿todavia estan en casa?')
# + colab={} colab_type="code" id="DUQVLVqUE1YW"
# wrong translation
translate(u'trata de averiguarlo.')
# + [markdown] colab_type="text" id="RTe5P5ioMJwN"
# ## Next steps
#
# * [Download a different dataset](http://www.manythings.org/anki/) to experiment with translations, for example, English to German, or English to French.
# * Experiment with training on a larger dataset, or using more epochs
#
|
site/en/tutorials/text/nmt_with_attention.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Challenge - Football match prediction
#
# ## Description
#
# For this week's challenge, we will try to build a model for the football match outcome prediction. The data containing the international football match results from 1872 to 2021 can be found in the Kaggle (***see Kaggle section for link***).
#
# On the other hand, this week's challenge has been created in the format of Kaggle in-class competition: we will develop our model on the given ```train``` dataset and will have to submit a file containing predictions from the ```test``` dataset. Then, according to the accuracy of your prediction, you will be ranked in the leaderboard.
#
#
# ## Kaggle
#
# You can access the competition via [this link](https://www.kaggle.com/c/ucl-ai-society-football-match-prediction/data) where you can also find the detailed description of the challenge and provided data. The key points:
#
# - The data section contains 3 files - ```train.csv```, ```test.csv```, ```sample_submission.csv```. You should develop the model using the **```train.csv```** file.
# - As the outcome is binary, we are going to use the logistic regression model.
# - After training the model, use it to predict the outcomes of the ```test.csv``` dataset. With outputs and id, create a submission file (the format can be seen in the ```sample_submission.csv``` file).
# - As the dataset features are in string format, we are going to use the string-to-integer encoder. Since it will only be covered in the latter tutorials, at this point, we will provide the function that takes the feature and provides transformed output.
#
# 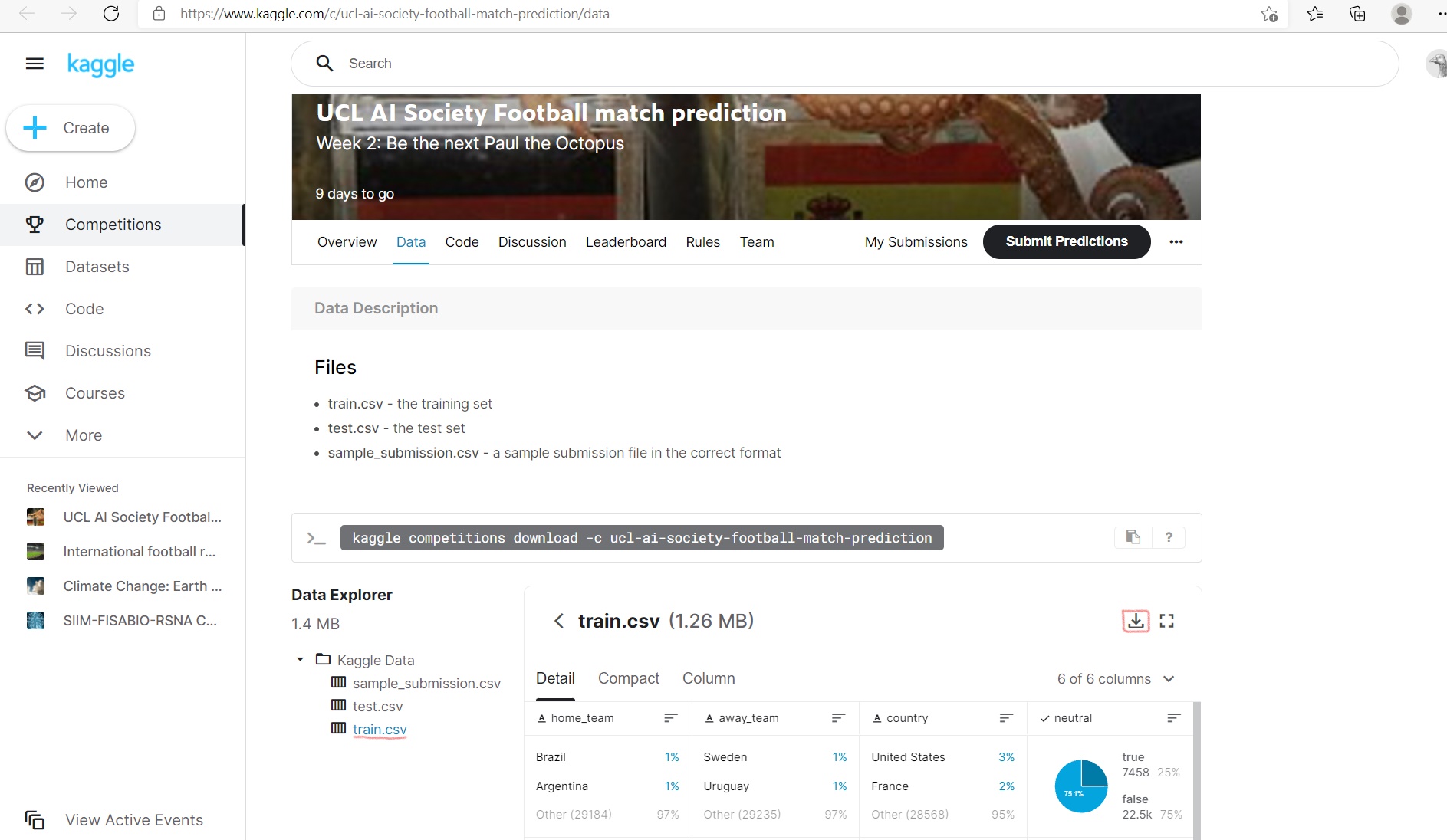
#
# ## Code
#
# You can use the code structure below as a guidance. Some of the steps will be already done for you (string to integer encoding), so **do not change it**.
#Importing libraries
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# After importing the libraries, specify the path for the downloaded ```train.csv``` dataset and read the file.
#Reading the data
PATH =
data =
# As usual, feel free to have a look at data using an appropriate function
#Look at the top dataset values with the Pandas function (.head())
# During the previous time, we learned the importance of checking missing values. Check if there are any missing values and if so, remove them.
#Check missing values and remove them (if there are any)
# Now we came to the data preprocessing step. As you can see, we have already written the ```encode()``` function which takes the data file and outputs ```X``` features. **Do not change this part of the code**.
#
# You will have to write a function that:
# - Takes the data file and passes it through the ```encode()``` function
# - Drops the unwanted columns from the data file
# - Extracts the outcome from the data file and converts it to numpy array.
# +
def encoder(data):
#--------------------------------------
#Extracting teams' names
H_team = np.array(data['home_team'])
A_team = np.array(data['away_team'])
Teams = np.hstack((H_team, A_team))
#--------------------------------------
#Encoding names
team_encoder = LabelEncoder().fit(Teams)
H_encoded = team_encoder.transform(H_team)
A_encoded = team_encoder.transform(A_team)
#--------------------------------------
#Creating X feature encoded values
H = np.expand_dims(H_encoded, axis = -1)
A = np.expand_dims(A_encoded, axis = -1)
X = np.concatenate([H, A], axis = 1)
#--------------------------------------
#Scale values
X = StandardScaler().fit(X).transform(X)
return X
#--------------------------------------------------------------------------------------------------------------------------
def preprocessing(data):
#Removing unwanted columns:
#Extracting encoded features
X =
#Extracting 'Outcome' and converting to the numpy array
y =
return X, y
X, y = preprocessing(data)
# -
# As we have our feature, we can split the data into the train and test sets. Feel free to chose the exact method
#Split X and y into train and test datasets
# Finally, we can build and train out logistic regression model. Feel free to do it from scratch or using Scikit-learn (you should have the code from the previous notebook).
#Build and train logistic regression model
# #### Prepare submission file
# As we now have our trained model, we can pass the ```test.csv``` data file to generate our predictions and convert them to an appropriate submission file.
# +
#Specify your test file path
test_path =
test_data = pd.read_csv(test_path)
#Encoding and preprocessing our test features (do not change this part)
X = encoder(test_data)
#Making prediction
y =
#Extracting id values from the test dataset and convert them to array
idx =
#Converting to DataFrame (do not change this part)
sub_file = pd.DataFrame([idx, y]).T
sub_file.columns = ['id', 'outcome']
#Specify your submission file
saving_path =
#Saving submission file (do not change this part)
sub_file.to_csv(saving_path, index = False)
|
Week-02/4_Challenge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Cleansing of the Data collected for the Best 250 Movies of All Times
# ## Calling the Necessary Libraries
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
# ## Defining the Functions Used in the Data Cleansing
# +
# The following function replaces a string by an integer if the string is purely numerical, and by NaN otherwise.
def Clean_NaN(x):
try:
x=int(float(x))
except:
x=np.nan
return x
# +
# This function removes 'K' and 'M' from a string, and automatically moves the '.' to the right by the
# correct number of places.
def RemoveDot (string):
if string[-1]=='K':
x=string.replace('K','')
t='000'
elif string[-1]=='M':
x=string.replace('M','')
t='000000'
else:
x='Not known!'
t=''
a=0;
b=x.find('.')
if b !=-1:
while x[-1]=='0':
x=x.replace(x[-1],'')
x=x+t
for k in range(len(x)-b):
if x[len(x)-k-1]=='0':
a += 1
else:
break
y=x.replace('.','')[:a+b] + '.' + x.replace('.','')[a+b:]
else:
y=x+t
return y
# +
# The following function takes the runtime in the format of x hours and y mintues, and replace it by total minutes.
def Convert_Runtime(x):
if x=='Not available!':
u=np.nan
else:
b1=x.find('h')
try:
y=int(x[:b1].strip())
except:
y=0
b2=x.find('m')
if b2!=-1:
z=int(x[b1+1:b2].strip())
else:
z=0
u=y*60+z
return u
# +
# The following function, takes the 'Box Office' of a movie in US dollars, and gives the equivalent amount in the
# year 2020. To do the conversion, we have extracted a table (CPI index) from the website
# https://www.usinflationcalculator.com/ and saved it as a txt file. The file is read by the code, and the
# following function uses the content to do the conversion.
def Box_Office_Conv(x,year):
z=int(round(x*CPI[year-1920]))
return z
# +
# The 'Outlier_Elimination' function replaces the outliers of the list x by the bound. UpLow indicates whether
# the given bound is an upper or a lower bound.
def Outlier_Elimination(x,bound,UpLow):
if UpLow=='U':
if x>bound:
return bound
else:
return x
elif UpLow=='L':
if x<bound:
return bound
else:
return x
else:
print('It has not been specified properly whether the bound is a lower bound or an upper bound!')
# -
# ## Calling the Data from Web-scraping
# +
# 'path' indicates where to find the file we obtained from webscraping, and 'path2' is the path for the txt file
# we constructed from the website https://www.usinflationcalculator.com/ for CPI index.
path='/Users/masoud/Dropbox/Private/UMBC-DataScience/DATA-601/Homework-02/250-Best-Movies.csv'
path2='/Users/masoud/Dropbox/Private/UMBC-DataScience/DATA-601/Homework-02/CPI-Conversion.txt'
# +
# Reading the file and storing data in 'CPIIndex'.
with open(path2,'rt') as f:
CPIIndex=f.readlines()
# -
Year=[]
CPI=[]
for k in range(101):
Year.append(int(CPIIndex[k].split(',')[0].strip()))
CPI.append(float(CPIIndex[k].split(',')[1].strip()))
# +
# Reading the data (i.e. the result of webscraping that was stored as a csv file in 'path') in a dataframe.
df=pd.read_csv(path)
# +
# For some reason when the dataframe is read, one new unewanted column is added. In here we just get rid of that
# column.
df.drop(columns=['Unnamed: 0'],inplace=True)
# -
# ### Representing the Dataframe
# The following dataframe has 250 rows (corresponding to the 250 Best Movies), and has 8 columns as follows:
# 1. Movie Name: that represents the name of the movie.
# 2. Movie Year: that represents the year movie was produced and screened.
# 3. Movie url: that represents the electronic address of the movie in the 'rottentomatoes.com' website.
# 4. Genre: that represents the genre of the movie.
# 5. Runtime: that represents the length of the movie.
# 6. Box Office: that represents the amount in US dollars the movie sold tickets in cinemas.
# 7. Critic Ratings: that represents the average ratings by the critics of the 'rottentomatoes.com' website.
# 8. Auidence Ratings: that represents the average ratings by all audience of the 'rottentomatoes.com' website.
df.shape
df.head()
# ## Cleansing the Dataframe
# +
# We define a new dataframe 'New_df'. In the new dataframe, we get rid of the spaces in the name of the columns.
New_df=pd.DataFrame()
New_df['Movie_Name']=df['Movie Name']
New_df['Movie_url']=df['Movie url']
New_df['Genre']=df['Genre']
New_df['Movie_Year']=df['Movie Year'].apply(int)
# +
# All columns of the old dataframe 'df' have string values. For the Box Office, we convert the value to integers
# through 'Clean_NaN' function first, and then we replace all the missing values by the mean of the values of
# the column 'Box Office'.
New_df['Box_Office_(USD)']=df['Box Office'].apply(RemoveDot).apply(Clean_NaN)
New_df['Box_Office_(USD)'].fillna(value=int(round(New_df['Box_Office_(USD)'].mean())),inplace=True)
New_df['Box_Office_(USD)']=New_df['Box_Office_(USD)'].astype(int)
# +
# In order to be able to compare the Box Office values, we need to convert all of values to their equivalent values
# in one year, say 2020. To do this, we take the advantage of the 'function Box_Office_Conv'.
New_df['Box_Office_(USD_2020)']=New_df.apply(
lambda x: Box_Office_Conv(x['Box_Office_(USD)'],x['Movie_Year']), axis=1)
# +
# We replace all the missing values in the column 'Crirtic_Ratings', and then substitute them by the mean of the
# column. The values of the entries of the column are on integer type at the end of the day.
New_df['Critic_Ratings']=df['Critic Ratings'].apply(Clean_NaN)
New_df['Critic_Ratings'].fillna(value=round(New_df['Critic_Ratings'].mean()),inplace=True)
# +
# Similarly, we replace all the missing values in the column 'Audience_Ratings', and then substitute them by
# the mean of the column. The values of the entries of the column are on integer type at the end of the day.
New_df['Audience_Ratings']=df['Audience Ratings'].apply(Clean_NaN)
New_df['Audience_Ratings'].fillna(value=round(New_df['Audience_Ratings'].mean()),inplace=True)
# +
# We convert the values of the column 'Runtime' to the minutes format. After the conversion, the values will have
# integer type.
New_df['Runtime_(min)']=df['Runtime'].apply(Convert_Runtime)
New_df['Runtime_(min)'].fillna(value=round(New_df['Runtime_(min)'].mean()),inplace=True)
# -
# ### Representing the new dataframe New_df
New_df.head()
# ## Checking for Outliers
# ### Outliers of Box Office
# +
# The following is the box plot of the column Box Office. There seems to exist an outlier above the upper limit.
# The outlier belongs to the movie 'Gone With the Wind'. We checked its boxplot against the value presented at
# rottentomatoes.com, and we found that the box office value for this movie is indeed correct. So we will keep this
# in our analysis.
BoxPlot_Box_Office = sns.boxplot(New_df['Box_Office_(USD_2020)'])
# -
New_df.loc[New_df['Box_Office_(USD_2020)']>3500000000]
# ### Outliers of Runtime
# +
# Plotting the box plot of the Runtime column, we particularly see that there are two very lengthy movies. As we
# in below, these two movies are 'The Best of Youth' and 'Satantango'. We checked the exact runtime of the two
# movies, and we realized that we have the correct length of the movies in our dataframe. Therefore, we'll keep
# them as they are.
BoxPlot_Runtime = sns.boxplot(New_df['Runtime_(min)'])
# -
New_df.loc[New_df['Runtime_(min)']>300]
# ### Outliers of Critic Ratings
# +
# It is observed that we have a couple of low outliers for the critic ratings. We do not have an independent way
# of checking the ratings. Therefore, we will replace the lower outliers by the lower fence of the box plot.
BoxPlot_CRatings = sns.boxplot(New_df['Critic_Ratings'])
# -
New_df.loc[New_df['Critic_Ratings']<80]
# +
# We will substitute the lower outliers by the 'Outlier_Elimination' function we defined in HW-1.
LowerBound=84
New_df.Critic_Ratings = New_df.Critic_Ratings.apply(Outlier_Elimination,args=(LowerBound,'L',))
# +
# The new box plot for Critic Ratings shows no outliers, as expected.
BoxPlot_CRatings = sns.boxplot(New_df.Critic_Ratings)
# -
# ### Outliers of Audience Ratings
# +
# It is observed that we have a couple of low outliers for the audience ratings. We do not have an independent way
# of checking the ratings. Therefore, we will replace the lower outliers by the lower fence of the box plot.
BoxPlot_ARatings = sns.boxplot(New_df['Audience_Ratings'])
# -
New_df.loc[New_df['Audience_Ratings']<86]
# +
# We will substitute the lower outliers by the 'Outlier_Elimination' function we defined in HW-1.
LowerBound=88
New_df.Audience_Ratings = New_df.Audience_Ratings.apply(Outlier_Elimination,args=(LowerBound,'L',))
# +
# The new box plot for Audience Ratings shows no outliers, as expected.
BoxPlot_ARatings = sns.boxplot(New_df['Audience_Ratings'])
# -
# ## Saving the Cleansed Dataframe in a csv File
# +
# We save the cleansed dataframe to a newly defined dataframe 'New_df' as a csv file for the next step of the
# process (i.e. Data Analysis)
New_df.to_csv('Cleansed_Data.csv')
# -
|
Assignments/Assignment-02/HW2-Data-Cleansing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import division
import itertools
import warnings
warnings.filterwarnings("ignore")
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import numpy as np
import pandas as pd
import math
from sklearn import metrics
from random import randint
from matplotlib import style
import seaborn as sns
# %matplotlib inline
# # Wrangle Data
# ### Acquire:
# +
colnames=['ip', 'timestamp', 'request_method', 'status', 'size',
'destination', 'request_agent']
df_orig = pd.read_csv('http://python.zach.lol/access.log',
engine='python',
header=None,
index_col=False,
names=colnames,
sep=r'\s(?=(?:[^"]*"[^"]*")*[^"]*$)(?![^\[]*\])',
na_values='"-"',
usecols=[0, 3, 4, 5, 6, 7, 8]
)
new = pd.DataFrame([["172.16.17.32", "[21/Apr/2019:10:02:41+0000]",
"GET /api/v1/items/HTTP/1.1", 200, 1153005, np.nan,
"python-requests/2.21.0"],
["192.168.3.11", "[17/Apr/2019:19:36:41+0000]",
"GET /api/v1/sales?page=79/HTTP/1.1", 301, 1005, np.nan,
"python-requests/2.21.0"],
["192.168.3.11", "[18/Apr/2019:19:42:41+0000]",
"GET /api/v1/sales?page=79/HTTP/1.1", 301, 2560, np.nan,
"python-requests/2.21.0"],
["192.168.127.12", "[19/Apr/2019:19:42:41+0000]",
"GET /api/v1/sales?page=79/HTTP/1.1", 200, 2056327, np.nan,
"python-requests/2.21.0"]], columns=colnames)
df = df_orig.append(new)
# -
df.head()
df.info()
# ### Parse Datetime:
df.timestamp = df.timestamp.str.replace(r'(\[|\+0000\])', '', regex=True)
df.timestamp= pd.to_datetime(df.timestamp.str.replace(':', ' ', 1))
df = df.set_index('timestamp')
# ### Cleanup Text
# +
for col in ['request_method', 'request_agent', 'destination']:
df[col] = df[col].str.replace('"', '')
df['request_method'] = df.request_method.str.replace(r'\?page=[0-9]+', '', regex=True)
df.head()
# -
# ### Add Variable - Converting bytes to mb:
df['size_mb'] = [n/1024/1024 for n in df['size']]
df.describe()
# # Detecting Anomalies in Discrete Variables
# ### Finding anomalies in already existing data:
# #### We can easily see some anomalies around IP addresses.
# +
ip_df = pd.DataFrame(df.ip.value_counts(dropna=False)).reset_index().\
rename(index=str, columns={'index': 'ip', 'ip': 'ip_count'})
ip_df2 = pd.DataFrame(df.ip.value_counts(dropna=False)/df.ip.count()).reset_index().\
rename(index=str, columns={'index': 'ip', 'ip': 'ip_proba'})
ip_df = ip_df.merge(ip_df2)
# see those where rate < 1%
ip_df[ip_df.ip_proba < .01]
# +
print(len(ip_df))
print(ip_df.tail(10))
plt.figure(figsize=(12, 4))
splot = sns.barplot(data=ip_df, x = 'ip', y = 'ip_count', ci = None)
for p in splot.patches:
splot.annotate(format(p.get_height(), '.0f'),
(p.get_x() + p.get_width() / 2., p.get_height()),
ha = 'center', va = 'center', xytext = (0, 10),
textcoords = 'offset points'
)
plt.xticks(rotation='vertical')
# -
# ## Detecting anomalies by establishing a baseline and evaluate as new data arrives.
# # Establish Baseline:
train = df.loc['2019-04-16 19:34:42':'2019-04-17 12:55:14']
train = train[['ip','request_method','status','size','destination','request_agent','size_mb']]
# ## Compute probabilities based on train sample:
ip_df = pd.DataFrame(train.ip.value_counts(dropna=False)/train.ip.count()).reset_index().\
rename(index=str, columns={'index': 'ip', 'ip': 'ip_proba'})
ip_df
# ## Merge probabilities with all data (train + new data):
# - Where the IP address is new, i.e. not seen in the training dataset, fill the probability with a value of 0.
#
df = df.reset_index().merge(ip_df, on=['ip'], how='left').fillna(value=0).set_index('timestamp')
df.ip_proba.value_counts()
# # Conditional Probabilities: Probabilities using 2 discrete variables
# ## Probability of Status given IP Address:
# #### If we are looking for an unexpected status (like authentication failure) from a known/common IP address.
# +
ip_probs = train.groupby('ip').size().div(len(df))
status_given_ip = pd.DataFrame(train.groupby(['ip', 'status']).\
size().div(len(train)).\
div(ip_probs,
axis=0,
level='ip').\
reset_index().\
rename(index=str,
columns={0: 'proba_status_given_ip'})
)
# +
ip_status_count = pd.DataFrame(train.groupby(['ip', 'status'])['request_method'].\
count().reset_index().\
rename(index=str,
columns={'request_method': 'ip_status_count'}))
ip_status = status_given_ip.merge(ip_status_count)
# -
# ## Add these probabilities to original events to detect anomalous events:
df = df.reset_index().merge(ip_status, on=['ip', 'status'], how='left').fillna(value=0).set_index('timestamp')
df.head()
plt.scatter(df.proba_status_given_ip, df.ip_proba)
ip_df
ip_df[ip_df.ip_proba < 0.01]
|
discrete_probabilistic_methods_lesson.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %%checkall
import matplotlib as mpl
try:
import tkinter
except (ImportError, ValueError):
mpl.use('Agg') # Support running in headless mode
import matplotlib.pyplot as plt
import os
import sys
import tempfile
import datetime
import pathlib
import numpy as np
import logging
import pandas as pd
from typing import Any, Sequence, Optional, Tuple, Callable, MutableSequence, MutableSet, Union, List
SEC_PER_DAY = 3600 * 24
_HAS_DISPLAY = None
EPOCH = datetime.datetime.utcfromtimestamp(0)
DATE_FORMAT = '%Y-%m-%d %H:%M:%S'
LOG_FORMAT = '[%(asctime)s.%(msecs)03d %(funcName)s] %(message)s'
def has_display() -> bool:
'''
If we are running in unit test mode or on a server, then don't try to draw graphs, etc.
'''
global _HAS_DISPLAY
if _HAS_DISPLAY is not None: return _HAS_DISPLAY
_HAS_DISPLAY = True
try:
plt.figure()
except tkinter.TclError:
_HAS_DISPLAY = False
return _HAS_DISPLAY
def shift_np(array: np.ndarray, n: int, fill_value: Any = None) -> np.ndarray:
'''
Similar to pandas.Series.shift but works on numpy arrays.
Args:
array: The numpy array to shift
n: Number of places to shift, can be positive or negative
fill_value: After shifting, there will be empty slots left in the array. If set, fill these with fill_value.
If fill_value is set to None (default), we will fill these with False for boolean arrays, np.nan for floats
'''
if array is None: return None
if len(array) == 0: return array
if fill_value is None:
fill_value = False if array.dtype == np.dtype(bool) else np.nan
e = np.empty_like(array)
if n >= 0:
e[:n] = fill_value
e[n:] = array[:-n]
else:
e[n:] = fill_value
e[:n] = array[-n:]
return e
def set_ipython_defaults(jupyter_multiple_display=True) -> None:
from IPython.core.interactiveshell import InteractiveShell
get_ipython().run_line_magic('matplotlib', 'inline') # type: ignore # noqa: 89
if jupyter_multiple_display:
InteractiveShell.ast_node_interactivity = 'all'
# autoreload extension
if 'autoreload' not in get_ipython().extension_manager.loaded: # type: ignore # noqa: 89
get_ipython().magic('load_ext autoreload') # type: ignore # noqa: 89
get_ipython().ipython.run_line_magic('autoreload', '2') # type: ignore # noqa: 89
def set_defaults(df_float_sf: int = 9,
df_display_max_rows: int = 200,
df_display_max_columns: int = 99,
np_seterr: str = 'raise',
plot_style: str = 'ggplot',
mpl_figsize: Tuple[int, int] = (8, 6),
jupyter_multiple_display=True) -> None:
'''
Set some display defaults to make it easier to view dataframes and graphs.
Args:
df_float_sf: Number of significant figures to show in dataframes (default 4). Set to None to use pandas defaults
df_display_max_rows: Number of rows to display for pandas dataframes when you print them (default 200). Set to None to use pandas defaults
df_display_max_columns: Number of columns to display for pandas dataframes when you print them (default 99). Set to None to use pandas defaults
np_seterr: Error mode for numpy warnings. See numpy seterr function for details. Set to None to use numpy defaults
plot_style: Style for matplotlib plots. Set to None to use default plot style.
mpl_figsize: Default figure size to use when displaying matplotlib plots (default 8,6). Set to None to use defaults
jupyter_multiple_display: If set, and you have multiple outputs in a Jupyter cell, output will contain all of them. Default True
'''
if df_float_sf is not None: pd.options.display.float_format = ('{:.' + str(df_float_sf) + 'g}').format
if df_display_max_rows is not None: pd.options.display.max_rows = df_display_max_rows
if df_display_max_columns is not None: pd.options.display.max_columns = df_display_max_columns
if plot_style is not None: plt.style.use(plot_style)
if mpl_figsize is not None: mpl.rcParams['figure.figsize'] = mpl_figsize
if np_seterr is not None: np.seterr(np_seterr) # type: ignore
pd.options.mode.chained_assignment = None # Turn off bogus 'view' warnings from pandas when modifying dataframes
# Display all cell outputs
plt.rcParams.update({'figure.max_open_warning': 100}) # For unit tests, avoid warning when opening more than 20 figures
if in_ipython():
set_ipython_defaults(jupyter_multiple_display)
def str2date(s: Optional[Union[np.datetime64, str]]) -> Optional[np.datetime64]:
'''Converts a string like "2008-01-15 15:00:00" to a numpy datetime64. If s is not a string, return s back'''
if isinstance(s, str): return np.datetime64(s)
return s
def strtup2date(tup: Any) -> Tuple[Optional[np.datetime64], Optional[np.datetime64]]:
'''Converts a string tuple like ("2008-01-15", "2009-01-16") to a numpy datetime64 tuple.
If the tuple does not contain strings, return it back unchanged'''
if tup and type(tup) is tuple and isinstance(tup[0], str): return (str2date(tup[0]), str2date(tup[1]))
return tup
def remove_dups(lst: Sequence[Any], key_func: Callable[[Any], Any] = None) -> MutableSequence[Any]:
'''
Remove duplicates from a list
Args:
lst: list to remove duplicates from
key_func: A function that takes a list element and converts it to a key for detecting dups
Returns (List): A list with duplicates removed. This is stable in the sense that original list elements will retain their order
>>> print(remove_dups(['a', 'd', 'a', 'c']))
['a', 'd', 'c']
>>> print(remove_dups(['a', 'd', 'A']))
['a', 'd', 'A']
>>> print(remove_dups(['a', 'd', 'A'], key_func = lambda e: e.upper()))
['a', 'd']
'''
new_list = []
seen: MutableSet[Any] = set()
for element in lst:
if key_func:
key = key_func(element)
else:
key = element
if key not in seen:
new_list.append(element)
seen.add(key)
return new_list
def np_get_index(array: np.ndarray, value: Any) -> int:
'''Get index of a value in a numpy array. Returns -1 if the value does not exist.'''
x = np.where(array == value)
if len(x[0]): return x[0][0]
return -1
def np_find_closest(a: np.ndarray, v: Any) -> Union[int, np.ndarray]:
'''
From https://stackoverflow.com/questions/8914491/finding-the-nearest-value-and-return-the-index-of-array-in-python
Find index of closest value to array v in array a. Returns an array of the same size as v
a must be sorted
>>> assert(all(np_find_closest(np.array([3, 4, 6]), np.array([4, 2])) == np.array([1, 0])))
'''
idx_ = a.searchsorted(v)
idx = np.clip(idx_, 1, len(a) - 1)
left = a[idx - 1]
right = a[idx]
idx -= v - left < right - v
return idx # type: ignore
def np_rolling_window(a: np.ndarray, window: int) -> np.ndarray:
'''
For applying rolling window functions to a numpy array
See: https://stackoverflow.com/questions/6811183/rolling-window-for-1d-arrays-in-numpy
>>> print(np.std(np_rolling_window(np.array([1, 2, 3, 4]), 2), 1))
[0.5 0.5 0.5]
'''
shape = a.shape[:-1] + (a.shape[-1] - window + 1, window)
strides = a.strides + (a.strides[-1],)
return np.lib.stride_tricks.as_strided(a, shape=shape, strides=strides) # type: ignore
def np_round(a: np.ndarray, clip: float):
'''
Round all elements in an array to the nearest clip
Args:
a: array with elements to round
clip: rounding value
>>> np_round(15.8, 0.25)
15.75
'''
return np.round(np.array(a, dtype=float) / clip) * clip
def np_bucket(a: np.ndarray, buckets: List[Any], default_value=0, side='mid') -> np.ndarray:
'''
Given a numpy array and a sorted list of buckets, assign each element to a bucket.
Args:
a (np.ndarray): The numpy array of values
buckets: (list) List of buckets
default_value: Used when we cannot assign an element to any bucket if side is 'left' or 'right'
side (str): If set to mid, we use the midpoint between buckets to assign elements
'left', assignment <= element
'right', assignment >= element
Default: 'mid'
Return:
np.ndarray of same length as a
>>> a = np.array([1, 5, 18, 3, 6, 10, 4])
>>> buckets = [4, 8, 12]
>>> assert np.alltrue(np_bucket(a, buckets, side='left') == np.array([0, 4, 12, 0, 4, 8, 4]))
>>> assert np.alltrue(np_bucket(a, buckets, default_value=25, side='right') == np.array([4, 8, 25, 4, 8, 12, 4]))
>>> assert(np.alltrue(np_bucket(a, buckets) == np.array([4, 4, 12, 4, 8, 12, 4])))
'''
assert side in ['mid', 'left', 'right'], f'unknown side: {side}'
if side == 'mid':
b = [0.5 * (buckets[i + 1] + buckets[i]) for i in range(len(buckets) - 1)]
conditions = [(a < e) for e in b]
ret = np.select(conditions, buckets[:-1], default=buckets[-1])
else:
conditions = [(a < buckets[i]) for i in range(len(buckets))]
if side == 'left':
buckets = buckets[::-1]
conditions = [(a >= buckets[i]) for i in range(len(buckets))]
else:
conditions = [(a <= buckets[i]) for i in range(len(buckets))]
ret = np.select(conditions, buckets, default=default_value)
return ret
def np_parse_array(s: str, dtype=float) -> np.ndarray:
"""
Create a 1 or 2 d numpy array from a string that looks like:
[[2. 5. 3. 0. 0.]
[3. 5. 0. 4. 3.]]
or
[2. 5. 3. 0. 8.]
>>> x = np_parse_array('[[2. 5. 3. 0. 0.]\\n [3. 5. 0. 4. 3.]]')
>>> assert np.allclose(x, np.array([[2., 5., 3., 0., 0.], [3., 5., 0., 4., 3.]]))
>>> x = np_parse_array('[3 4. 5]')
>>> assert np.allclose(x, np.array([3, 4., 5]))
"""
height = s.count(']') - 1
for char in [']', '[', '\n']:
s = s.replace(char, '')
x = np.fromstring(s, sep=' ', dtype=dtype)
if height > 0:
width = int(len(x) / height)
x = x.reshape(height, width)
return x
def np_inc_dates(dates: np.ndarray, num_days: int = 1) -> np.ndarray:
'''
Increment the given date array so each cell gets the next higher value in this array
>>> dates = np.array(['2021-06-01', '2021-06-01', '2021-08-01', '2021-04-01'], dtype='M8[D]')
>>> check = np.array([dates[2], dates[2], np.datetime64('nat'), dates[0]])
>>> assert np.array_equal(np_inc_dates(dates, 1),
... np.array(['2021-08-01', '2021-08-01', 'NaT', '2021-06-01'], dtype='M8[D]'), equal_nan=True)
>>> assert np.array_equal(np_inc_dates(dates, 2),
... np.array(['NaT', 'NaT', 'NaT', '2021-08-01'], dtype='M8[D]'), equal_nan=True)
>>> assert np.array_equal(np_inc_dates(dates, -1),
... np.array(['2021-04-01', '2021-04-01', '2021-06-01', 'NaT'], dtype='M8[D]'), equal_nan=True)
>>> assert np.array_equal(np_inc_dates(dates, -2),
... np.array(['NaT', 'NaT', '2021-04-01', 'NaT'], dtype='M8[D]'), equal_nan=True)
'''
uniq_dates = np.unique(dates)
date_values = np.concatenate([uniq_dates, [np.datetime64('nat')]])
indices = np.searchsorted(uniq_dates, dates) + num_days
indices = np.where((indices < 0) | (indices > len(uniq_dates)), len(uniq_dates), indices)
return date_values[indices]
def np_uniques(arrays: List[np.ndarray]) -> np.ndarray:
'''
Given a list of numpy arrays that may have different datatype, generate a structured numpy
array with sorted, unique elements from that list
>>> array1 = np.array(['2018-01-02', '2018-01-03', '2018-01-02', '2018-01-03'], dtype='M8[D]')
>>> array2 = np.array(['P', 'P', 'P', 'C'])
>>> x = np_uniques([array1, array2])
>>> assert len(x) == 3
>>> assert x[0][0] == np.datetime64('2018-01-02')
>>> assert x[0][1] == 'P'
'''
size = len(arrays[0])
cols = len(arrays)
data_ = [tuple([arrays[i][j] for i in range(cols)]) for j in range(size)]
data = np.array(data_, dtype=[('', col.dtype) for col in arrays])
return np.unique(data)
def day_of_week_num(a: Union[np.datetime64, np.ndarray]) -> Union[int, np.ndarray]:
'''
From https://stackoverflow.com/questions/52398383/finding-day-of-the-week-for-a-datetime64
Get day of week for a numpy array of datetimes
Monday is 0, Sunday is 6
Args:
a: numpy datetime64 or array of datetime64
Return:
int or numpy ndarray of int: Monday is 0, Sunday is 6
>>> day_of_week_num(np.datetime64('2015-01-04'))
6
'''
int_date: int = a.astype('datetime64[D]').view('int64') # type: ignore
ret = (int_date - 4) % 7
# if np.isscalar(ret): ret = ret.item()
return ret
def percentile_of_score(a: np.ndarray) -> Optional[np.ndarray]:
'''
For each element in a, find the percentile of a its in. From stackoverflow.com/a/29989971/5351549
Like scipy.stats.percentileofscore but runs in O(n log(n)) time.
>>> a = np.array([4, 3, 1, 2, 4.1])
>>> percentiles = percentile_of_score(a)
>>> assert(all(np.isclose(np.array([ 75., 50., 0., 25., 100.]), percentiles)))
'''
assert isinstance(a, np.ndarray), f'expected numpy array, got: {a}'
if not len(a): return None
return np.argsort(np.argsort(a)) * 100. / (len(a) - 1)
def date_2_num(d: Union[np.datetime64, np.ndarray]) -> Union[int, np.ndarray]:
from matplotlib.dates import date2num
return date2num(d)
def resample_vwap(df: pd.DataFrame, sampling_frequency: str) -> Optional[np.ndarray]:
'''
Compute weighted average of vwap given higher frequency vwap and volume
'''
if 'v' not in df.columns: return None
sum_1 = df.vwap * df.v
sum_2 = sum_1.resample(sampling_frequency).agg(np.sum)
volume_sum = df.v.resample(sampling_frequency).agg(np.sum)
vwap = sum_2 / volume_sum
return vwap
def resample_trade_bars(df, sampling_frequency, resample_funcs=None):
'''Downsample trade bars using sampling frequency
Args:
df (pd.DataFrame): Must contain an index of numpy datetime64 type which is monotonically increasing
sampling_frequency (str): See pandas frequency strings
resample_funcs (dict of str: int): a dictionary of column name -> resampling function for any columns that are custom defined. Default None.
If there is no entry for a custom column, defaults to 'last' for that column
Returns:
pd.DataFrame: Resampled dataframe
>>> import math
>>> df = pd.DataFrame({'date': np.array(['2018-01-08 15:00:00', '2018-01-09 13:30:00', '2018-01-09 15:00:00', '2018-01-11 15:00:00'], dtype = 'M8[ns]'),
... 'o': np.array([8.9, 9.1, 9.3, 8.6]),
... 'h': np.array([9.0, 9.3, 9.4, 8.7]),
... 'l': np.array([8.8, 9.0, 9.2, 8.4]),
... 'c': np.array([8.95, 9.2, 9.35, 8.5]),
... 'v': np.array([200, 100, 150, 300]),
... 'x': np.array([300, 200, 100, 400])
... })
>>> df['vwap'] = 0.5 * (df.l + df.h)
>>> df.set_index('date', inplace = True)
>>> df = resample_trade_bars(df, sampling_frequency = 'D', resample_funcs={'x': lambda df,
... sampling_frequency: df.x.resample(sampling_frequency).agg(np.mean)})
>>> assert(len(df) == 4)
>>> assert(math.isclose(df.vwap.iloc[1], 9.24))
>>> assert(np.isnan(df.vwap.iloc[2]))
>>> assert(math.isclose(df.l[3], 8.4))
'''
if sampling_frequency is None: return df
if resample_funcs is None: resample_funcs = {}
if 'vwap' in df.columns: resample_funcs.update({'vwap': resample_vwap})
funcs = {'o': 'first', 'h': 'max', 'l': 'min', 'c': 'last', 'v': 'sum'}
agg_dict = {}
for col in df.columns:
if col in funcs:
agg_dict[col] = funcs[col]
continue
if col not in resample_funcs:
agg_dict[col] = 'last'
resampled = df.resample(sampling_frequency).agg(agg_dict).dropna(how='all')
for k, v in resample_funcs.items():
res = v(df, sampling_frequency)
if res is not None: resampled[k] = res
resampled.reset_index(inplace=True)
return resampled
def resample_ts(dates: np.ndarray, values: np.ndarray, sampling_frequency: str) -> Tuple[np.ndarray, np.ndarray]:
'''Downsample a tuple of datetimes and value arrays using sampling frequency, using the last value if it does not exist at the bin edge.
See pandas.Series.resample
Args:
dates: a numpy datetime64 array
values: a numpy array
sampling_frequency: See pandas frequency strings
Returns:
Resampled tuple of datetime and value arrays
'''
if sampling_frequency is None: return dates, values
s = pd.Series(values, index=dates).resample(sampling_frequency).last()
return s.index.values, s.values
def zero_to_nan(array: np.ndarray) -> np.ndarray:
'''Converts any zeros in a numpy array to nans'''
if array is None: return None
return np.where(array == 0, np.nan, array)
def nan_to_zero(array: np.ndarray) -> np.ndarray:
'''Converts any nans in a numpy float array to 0'''
if array is None: return None
return np.where(np.isnan(array), 0, array)
def monotonically_increasing(array: np.ndarray) -> bool:
'''
Returns True if the array is monotonically_increasing, False otherwise
>>> monotonically_increasing(np.array(['2018-01-02', '2018-01-03'], dtype = 'M8[D]'))
True
>>> monotonically_increasing(np.array(['2018-01-02', '2018-01-02'], dtype = 'M8[D]'))
False
'''
if not len(array): return False
ret: bool = np.all(np.diff(array).astype(float) > 0).astype(bool) # type: ignore
return ret
def infer_frequency(timestamps: np.ndarray) -> float:
'''Returns most common frequency of date differences as a fraction of days
Args:
timestamps: A numpy array of monotonically increasing datetime64
>>> timestamps = np.array(['2018-01-01 11:00:00', '2018-01-01 11:15:00', '2018-01-01 11:30:00', '2018-01-01 11:35:00'], dtype = 'M8[ns]')
>>> print(round(infer_frequency(timestamps), 8))
0.01041667
'''
if isinstance(timestamps, pd.Series): timestamps = timestamps.values
assert(monotonically_increasing(timestamps))
numeric_dates = date_2_num(timestamps)
diff_dates = np.round(np.diff(numeric_dates), 8)
(values, counts) = np.unique(diff_dates, return_counts=True)
return values[np.argmax(counts)]
def series_to_array(series: pd.Series) -> np.ndarray:
'''Convert a pandas series to a numpy array. If the object is not a pandas Series return it back unchanged'''
if type(series) == pd.Series: return series.values
return series
def to_csv(df, file_name: str, index: bool = False, compress: bool = False, *args, **kwargs) -> None:
"""
Creates a temporary file then renames to the permanent file so we don't have half written files.
Also optionally compresses using the xz algorithm
"""
compression = None
suffix = ''
if compress:
compression = 'xz'
suffix = '.xz'
df.to_csv(file_name + '.tmp', index=index, compression=compression, *args, **kwargs)
os.rename(file_name + '.tmp', file_name + suffix)
def millis_since_epoch(dt: datetime.datetime) -> float:
"""
Given a python datetime, return number of milliseconds between the unix epoch and the datetime.
Returns a float since it can contain fractions of milliseconds as well
>>> millis_since_epoch(datetime.datetime(2018, 1, 1))
1514764800000.0
"""
return (dt - EPOCH).total_seconds() * 1000.0
def day_symbol(day_int: Union[int, np.ndarray]) -> Union[str, np.ndarray]:
day_str = np.select([day_int == 0, day_int == 1, day_int == 2, day_int == 3, day_int == 4, day_int == 5, day_int == 6],
['M', 'Tu', 'W', 'Th', 'F', 'Sa', 'Su'], default='')
if day_str.shape == (): day_str = np.asscalar(day_str)
return day_str
def infer_compression(input_filename: str) -> Optional[str]:
"""
Infers compression for a file from its suffix. For example, given "/tmp/hello.gz", this will return "gzip"
>>> infer_compression("/tmp/hello.gz")
'gzip'
>>> infer_compression("/tmp/abc.txt") is None
True
"""
parts = input_filename.split('.')
if len(parts) <= 1: return None
suffix = parts[-1]
if suffix == 'gz': return 'gzip'
if suffix == 'bz2': return 'bz2'
if suffix == 'zip': return 'zip'
if suffix == 'xz': return 'xz'
return None
def touch(fname: str, mode: int = 0o666, dir_fd: Optional[int] = None, **kwargs) -> None:
'''replicate unix touch command, i.e create file if it doesn't exist, otherwise update timestamp'''
flags = os.O_CREAT | os.O_APPEND
with os.fdopen(os.open(fname, flags=flags, mode=mode, dir_fd=dir_fd)) as f:
os.utime(f.fileno() if os.utime in os.supports_fd else fname,
dir_fd=None if os.supports_fd else dir_fd, **kwargs)
def is_newer(filename: str, ref_filename: str) -> bool:
'''whether filename ctime (modfication time) is newer than ref_filename or either file does not exist
>>> import time
>>> import tempfile
>>> temp_dir = tempfile.gettempdir()
>>> touch(f'{temp_dir}/x.txt')
>>> time.sleep(0.1)
>>> touch(f'{temp_dir}/y.txt')
>>> is_newer(f'{temp_dir}/y.txt', f'{temp_dir}/x.txt')
True
>>> touch(f'{temp_dir}/y.txt')
>>> time.sleep(0.1)
>>> touch(f'{temp_dir}/x.txt')
>>> is_newer(f'{temp_dir}/y.txt', f'{temp_dir}/x.txt')
False
'''
if not os.path.isfile(filename) or not os.path.isfile(ref_filename): return True
return os.path.getmtime(filename) > os.path.getmtime(ref_filename)
def get_empty_np_value(np_dtype: np.dtype) -> Any:
'''
Get empty value for a given numpy datatype
>>> a = np.array(['2018-01-01', '2018-01-03'], dtype = 'M8[D]')
>>> get_empty_np_value(a.dtype)
numpy.datetime64('NaT')
'''
kind = np_dtype.kind
if kind == 'f': return np.nan # float
if kind == 'b': return False # bool
if kind == 'i' or kind == 'u': return 0 # signed or unsigned int
if kind == 'M': return np.datetime64('NaT') # datetime
if kind == 'O' or kind == 'S' or kind == 'U': return '' # object or string or unicode
raise Exception(f'unknown dtype: {np_dtype}')
def get_temp_dir() -> str:
if os.access('/tmp', os.W_OK):
return '/tmp'
else:
return tempfile.gettempdir()
def linear_interpolate(a1: Union[np.ndarray, float],
a2: Union[np.ndarray, float],
x1: Union[np.ndarray, float],
x2: Union[np.ndarray, float],
x: Union[np.ndarray, float]) -> Union[np.ndarray, float]:
'''
>>> assert(linear_interpolate(3, 4, 8, 10, 8.9) == 3.45)
>>> assert(linear_interpolate(3, 3, 8, 10, 8.9) == 3)
>>> assert(np.isnan(linear_interpolate(3, 4, 8, 8, 8.9)))
>>> x = linear_interpolate(
... np.array([3., 3.]),
... np.array([4., 3.]),
... np.array([8., 8.]),
... np.array([10, 8.]),
... np.array([8.9, 8.]))
>>> assert(np.allclose(x, np.array([3.45, 3.])))
'''
diff = x2 - x1
diff = np.where(diff == 0, 1, diff)
return np.where((a2 == a1), a1,
np.where(x2 == x1, np.nan, a1 + (a2 - a1) * (x - x1) / diff))
def bootstrap_ci(a: np.ndarray,
ci_level: float = 0.95,
n: int = 1000,
func: Callable[[np.ndarray], np.ndarray] = np.mean) -> Tuple[float, float]: # type: ignore
'''
Non parametric bootstrap for confidence intervals
Args:
a: The data to bootstrap from
ci_level: The confidence interval level, e.g. 0.95 for 95%. Default 0.95
n: Number of boostrap iterations. Default 1000
func: The function to use, e.g np.mean or np.median. Default np.mean
Return:
A tuple containing the lower and upper ci
>>> np.random.seed(0)
>>> x = np.random.uniform(high=10, size=100000)
>>> assert np.allclose(bootstrap_ci(x), (4.9773159, 5.010328))
'''
simulations = np.full(n, np.nan)
sample_size = len(a)
for c in range(n):
itersample = np.random.choice(a, size=sample_size, replace=True)
simulations[c] = func(itersample)
simulations.sort()
u_pval = (1 + ci_level) / 2.
l_pval = (1 - u_pval)
l_indx = int(np.floor(n * l_pval))
u_indx = int(np.floor(n * u_pval))
return(simulations[l_indx], simulations[u_indx])
def _add_stream_handler(logger: logging.Logger, log_level: int = logging.INFO, formatter: logging.Formatter = None) -> None:
if formatter is None: formatter = logging.Formatter(fmt=LOG_FORMAT, datefmt=DATE_FORMAT)
stream_handler = logging.StreamHandler(sys.stdout)
# stream_handler = logging.StreamHandler()
stream_handler.setFormatter(formatter)
stream_handler.setLevel(log_level)
logger.addHandler(stream_handler)
def get_main_logger() -> logging.Logger:
# sys.stderr = sys.stdout
main_logger = logging.getLogger('pq')
if len(main_logger.handlers): return main_logger
_add_stream_handler(main_logger)
main_logger.setLevel(logging.INFO)
main_logger.propagate = False
return main_logger
def get_child_logger(child_name: str) -> logging.Logger:
_ = get_main_logger() # Init handlers if needed
full_name = 'pq.' + child_name if child_name else 'pq'
logger = logging.getLogger(full_name)
return logger
def in_ipython() -> bool:
'''
Whether we are running in an ipython (or Jupyter) environment
'''
import builtins
return '__IPYTHON__' in vars(builtins)
class Paths:
'''
Conventions for where to read / write data and reports
'''
def __init__(self, base_path: str = None) -> None:
if base_path:
self.base_path = pathlib.Path(base_path)
else:
self.base_path = pathlib.Path.cwd()
# Data paths
self.data_path = self.base_path / 'data'
self.raw_data_path = self.data_path / 'raw'
self.interim_data_path = self.data_path / 'interim'
self.processed_data_path = self.data_path / 'processed'
self.external_data_path = self.data_path / 'external'
# Reports paths
self.reports_path = self.base_path / 'reports'
self.figures_path = self.reports_path / 'figures'
def create(self) -> None:
default_mode = 0o755
self.data_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.raw_data_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.interim_data_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.processed_data_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.external_data_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.reports_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
self.figures_path.mkdir(mode=default_mode, parents=False, exist_ok=True)
def get_paths(base_path: str = None) -> Paths:
paths = Paths(base_path)
paths.create()
return paths
if __name__ == "__main__":
import doctest
doctest.testmod(optionflags=doctest.NORMALIZE_WHITESPACE | doctest.ELLIPSIS)
|
pyqstrat/src_nb/pq_utils.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Data Wrangler Processing Job for Claims Dataset
#
# This notebook executes your Data Wrangler Flow `claims.flow` on the entire dataset using a SageMaker
# Processing Job and will save the processed data to S3.
#
# This notebook saves data from the step `Cast Single Data Type` from `Source: Claims.Csv`. To save from a different step, go to Data Wrangler
# to select a new step to export.
#
# ---
#
# ## Contents
#
# 1. [Inputs and Outputs](#Inputs-and-Outputs)
# 1. [Run Processing Job](#Run-Processing-Job)
# 1. [Job Configurations](#Job-Configurations)
# 1. [Create Processing Job](#Create-Processing-Job)
# 1. [Job Status & S3 Output Location](#Job-Status-&-S3-Output-Location)
# ---
#
# 
# ### Loading stored variables
# If you ran this notebook before, you may want to re-use the resources you aready created with AWS. Run the cell below to load any prevously created variables. You should see a print-out of the existing variables. If you don't see anything printed then it's probably the first time you are running the notebook!
import warnings
warnings.filterwarnings('ignore')
# !pip install awswrangler
# +
import boto3
import sagemaker
region = sagemaker.Session().boto_region_name
boto3.setup_default_session(region_name=region)
boto_session = boto3.Session(region_name=region)
s3_client = boto3.client("s3", region_name=region)
sagemaker_client = boto_session.client("sagemaker")
sess = sagemaker.session.Session(
boto_session=boto_session, sagemaker_client=sagemaker_client
)
# -
# %store -r
# %store
# ======> output_paths
processing_dir = "/opt/ml/processing"
# # Inputs and Outputs
#
# The below settings configure the inputs and outputs for the flow export.
#
# <div class="alert alert-info"> 💡 <strong> Configurable Settings </strong>
#
# In <b>Input - Source</b> you can configure the data sources that will be used as input by Data Wrangler
#
# 1. For S3 sources, configure the source attribute that points to the input S3 prefixes
# 2. For all other sources, configure attributes like query_string, database in the source's
# <b>DatasetDefinition</b> object.
#
# If you modify the inputs the provided data must have the same schema and format as the data used in the Flow.
# You should also re-execute the cells in this section if you have modified the settings in any data sources.
# </div>
# +
from sagemaker.processing import ProcessingInput, ProcessingOutput
from sagemaker.dataset_definition.inputs import AthenaDatasetDefinition, DatasetDefinition, RedshiftDatasetDefinition
data_sources = []
# -
# ## Input - S3 Source: claims.csv
data_sources.append(ProcessingInput(
source=f"s3://{bucket}/{prefix}/data/raw/claims.csv", # You can override this to point to other dataset on S3
destination=f"{processing_dir}/claims",
input_name="claims",
s3_data_type="S3Prefix",
s3_input_mode="File",
s3_data_distribution_type="FullyReplicated"
))
print(f"Claims s3 path: s3://{bucket}/{prefix}/data/raw/claims.csv")
# ## Output: S3 settings
#
# <div class="alert alert-info"> 💡 <strong> Configurable Settings </strong>
#
# 1. <b>bucket</b>: you can configure the S3 bucket where Data Wrangler will save the output. The default bucket from
# the SageMaker notebook session is used.
# 2. <b>flow_export_id</b>: A randomly generated export id. The export id must be unique to ensure the results do not
# conflict with other flow exports
# 3. <b>s3_ouput_prefix</b>: you can configure the directory name in your bucket where your data will be saved.
# </div>
# +
import time
import uuid
flow_export_id = f"{time.strftime('%d-%H-%M-%S', time.gmtime())}-{str(uuid.uuid4())[:8]}"
flow_export_name = f"flow-{flow_export_id}"
print(f"Flow export name: {flow_export_name}")
# -
# Below are the inputs required by the SageMaker Python SDK to launch a processing job.
# +
import json
# name of the flow file which should exist in the current notebook working directory
flow_file_name = "flows/claims.flow"
# Load .flow file from current notebook working directory
# !echo "Loading flow file from current notebook working directory: $PWD"
with open(flow_file_name) as f:
flow = json.load(f)
# Output name is auto-generated from the select node's ID + output name from the flow file.
output_name = (f"{flow['nodes'][-1]['node_id']}.{flow['nodes'][-1]['outputs'][0]['name']}")
print(f"Output name: {output_name}")
s3_output_prefix = f"export-{flow_export_name}/output"
s3_output_path = f"s3://{bucket}/{prefix}/flow/output/{s3_output_prefix}"
print(f"Flow S3 export result path: {s3_output_path}")
processing_job_output = ProcessingOutput(
output_name=output_name,
source=f"{processing_dir}/output",
destination=s3_output_path,
s3_upload_mode="EndOfJob"
)
# -
# ## Upload Flow to S3
#
# To use the Data Wrangler as an input to the processing job, first upload your flow file to Amazon S3.
#
# 
# +
# Upload flow to S3
s3_client = boto3.client("s3")
s3_client.upload_file(flow_file_name, bucket, f"{prefix}/data_wrangler_flows/{flow_export_name}.flow", ExtraArgs={"ServerSideEncryption": "aws:kms"})
flow_s3_uri = f"s3://{bucket}/{prefix}/data_wrangler_flows/{flow_export_name}.flow"
print(f"Data Wrangler flow {flow_file_name} uploaded to {flow_s3_uri}")
# -
# The Data Wrangler Flow is also provided to the Processing Job as an input source which we configure below.
## Input - Flow: claims.flow
flow_input = ProcessingInput(
source=flow_s3_uri,
destination=f"{processing_dir}/flow",
input_name="flow",
s3_data_type="S3Prefix",
s3_input_mode="File",
s3_data_distribution_type="FullyReplicated"
)
print(f"Flow s3 path: {flow_s3_uri}")
# # Run Processing Job
# ## Job Configurations
#
# <div class="alert alert-info"> 💡 <strong> Configurable Settings </strong>
#
# You can configure the following settings for Processing Jobs. If you change any configurations you will
# need to re-execute this and all cells below it by selecting the Run menu above and click
# <b>Run Selected Cells and All Below</b>
#
# 1. IAM role for executing the processing job.
# 2. A unique name of the processing job. Give a unique name every time you re-execute processing jobs
# 3. Data Wrangler Container URL.
# 4. Instance count, instance type and storage volume size in GB.
# 5. Content type for each output. Data Wrangler supports CSV as default and Parquet.
# 6. Network Isolation settings
# 7. KMS key to encrypt output data
# </div>
#
# 
# +
from sagemaker import image_uris
# IAM role for executing the processing job.
iam_role = sagemaker.get_execution_role()
# Unique processing job name. Give a unique name every time you re-execute processing jobs
processing_job_name = f"data-wrangler-flow-processing-{flow_export_id}"
print(f"Processing Job Name: {processing_job_name}")
# Data Wrangler Container URL.
container_uri = image_uris.retrieve(framework='data-wrangler',region=region)
print(f"Container uri: {container_uri}")
# Processing Job Instance count and instance type.
instance_count = 2
instance_type = "ml.m5.4xlarge"
# Size in GB of the EBS volume to use for storing data during processing
volume_size_in_gb = 30
# Content type for each output. Data Wrangler supports CSV as default and Parquet.
output_content_type = "CSV"
# Network Isolation mode; default is off
enable_network_isolation = False
# Output configuration used as processing job container arguments
output_config = {
output_name: {
"content_type": output_content_type
}
}
# KMS key for per object encryption; default is None
kms_key = None
# -
# ## Create Processing Job
#
# To launch a Processing Job, you will use the SageMaker Python SDK to create a Processor function.
# +
from sagemaker.processing import Processor
from sagemaker.network import NetworkConfig
processor = Processor(
role=iam_role,
image_uri=container_uri,
instance_count=instance_count,
instance_type=instance_type,
volume_size_in_gb=volume_size_in_gb,
network_config=NetworkConfig(enable_network_isolation=enable_network_isolation),
sagemaker_session=sess,
output_kms_key=kms_key
)
# Start Job
processor.run(
inputs=[flow_input] + data_sources,
outputs=[processing_job_output],
arguments=[f"--output-config '{json.dumps(output_config)}'"],
wait=False,
logs=False,
job_name=processing_job_name
)
# -
# ## Job Status & S3 Output Location
#
# Below you wait for processing job to finish. If it finishes successfully, the raw parameters used by the
# Processing Job will be printed
# +
s3_job_results_path = f"s3://{bucket}/{prefix}/flow/{s3_output_prefix}/{processing_job_name}"
print(f"Job results are saved to S3 path: {s3_job_results_path}")
job_result = sess.wait_for_processing_job(processing_job_name)
job_result
# -
# <a id='aud-datasets'></a>
#
# ## DataSets and Feature Types
# [overview](#all-up-overview)
# ----
#
# 
claims_dtypes = {
"policy_id": int,
"incident_severity": int,
"num_vehicles_involved": int,
"num_injuries": int,
"num_witnesses": int,
"police_report_available": int,
"injury_claim": float,
"vehicle_claim": float,
"total_claim_amount": float,
"incident_month": int,
"incident_day": int,
"incident_dow": int,
"incident_hour": int,
"fraud": int,
"driver_relationship_self": int,
"driver_relationship_na": int,
"driver_relationship_spouse": int,
"driver_relationship_child": int,
"driver_relationship_other": int,
"incident_type_collision": int,
"incident_type_breakin": int,
"incident_type_theft": int,
"collision_type_front": int,
"collision_type_rear": int,
"collision_type_side": int,
"collision_type_na": int,
"authorities_contacted_police": int,
"authorities_contacted_none": int,
"authorities_contacted_fire": int,
"authorities_contacted_ambulance": int,
"event_time": float,
}
# ### Load Processed Data into Pandas
#
# We use the [AWS Data Wrangler library](https://github.com/awslabs/aws-data-wrangler) to load the exported
# dataset into a Pandas dataframe.
import awswrangler as wr
# +
# ======> This is your DataFlow output path if you decide to redo the work in DataFlow on your own
if output_content_type.upper() == "CSV":
claims_preprocessed = wr.s3.read_csv(
path=s3_output_path, dataset=True, dtype=claims_dtypes
)
else:
print(f"Unexpected output content type {output_content_type}")
# %store claims_preprocessed
claims_preprocessed
# -
claims_preprocessed.info()
# We now have a set of Pandas DataFrames that contain the customer and claim data, with the correct data types. When Dat Wrangler encodes a feature as one-hot-encoded feature, it will default to float data types for those resulting features (one feature --> many columns for the one hot encoding).
#
# <font color ='red'> Note: </font> the reason for explicitly converting the data types for categorical features generated by Data Wrangler, is to ensure they are of type integer so that Clarify will treat them as categorical variables.
|
03b-ClaimsProcessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# ## Importing the dataset
dataset = pd.read_csv('Position_Salaries.csv')
X = dataset.iloc[:, 1:2].values
y = dataset.iloc[:, 2].values
print("X is: ",X)
print("y is:",y)
# ## Fitting Linear regression
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X, y)
# ## Fitiing Polynomial regression
from sklearn.preprocessing import PolynomialFeatures
poly_reg = PolynomialFeatures(degree = 5)
X_poly = poly_reg.fit_transform(X)
lin_reg2 = LinearRegression()
lin_reg2.fit(X_poly, y)
# ## visualisation for Linear regression
plt.scatter(X, y, color = 'green')
plt.plot(X, lin_reg.predict(X), color = 'red')
plt.title('Truth or Bluff (Linear regression)')
plt.xlabel('Position')
plt.ylabel('Salary')
plt.show()
# ## visualisation for Polynomial regression
X_grid = np.arange(min(X), max(X), 0.1)
X_grid = X_grid.reshape((len(X_grid), 1))
plt.scatter(X, y, color = 'green')
plt.plot(X, lin_reg2.predict(poly_reg.fit_transform(X)), color = 'red')
plt.title('Truth or Bluff (Polynomial regression)')
plt.xlabel('Position')
plt.ylabel('Salary')
plt.show()
# ## Prediciting a new result using Linear regrssion
a = int(input('Enter Previous Level: '))
print("Estimated salary will be: " , lin_reg.predict(a))
# ## Prediciting a new result using Polynomial regrssion
a = int(input('Enter Previous Level: '))
print("Estimated salary will be: " ,lin_reg2.predict(poly_reg.fit_transform(a)))
|
Polynomial Regression From Scratch/Poly_Reg_Test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing Ford GoBike Data
# ## by <NAME>
#
# ## 1. Preliminary Wrangling
#
# > Ford GoBike is a regional public bicycle sharing system located in San Francisco Bay Area, California. In this project, history data of the whole 2018 year was used.
# +
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime
import glob
import os
# %matplotlib inline
# -
# ### 1.1 Gathering data
# This dataset is of big size, and separated by 12 months, so I need to combine them in the first step.
path = r'/Users/zhenghaoxiao/Documents/udacity/communicate_data_findings/bikedata'
files = glob.glob(os.path.join(path, '*.csv'))
df = pd.concat((pd.read_csv(f) for f in files), ignore_index=True)
df.to_csv('bike.csv', index=False)
# ### 1.2 Assessing data
df = pd.read_csv('bike.csv')
# Now, let's have a glimpse of the dataset:
df.head()
df.sample(10)
# Check data type of columns:
df.info(null_counts=True)
# It is obvious that there are some data type problems: timestamp data are not in time date format, ID should be of string format
df.shape
# There are 1863721 rows and 16 columns in the original dataset.
# Duplicated values check:
df.duplicated().sum()
# Missing values check:
df.isna().sum()
# Missing values exist in start_station_id, start_station_name, end_station_id, end_station_name, member_birth_year, and member_gender columns. Not all the missing values need to be cleaned. I should just cleaned the missing values in member_birth_year column, because there will be an analysis of interest on member birth year (ages).
df.describe()
# By assessing the dataset, there are some data quality issues in this dataset:
#
# * Missing values exist in **start_station_id, start_station_name, end_station_id, end_station_name, member_birth_year**, and **member_gender** columns
#
# * Data type errors:
# - should be object: **start_station_id, end_station_id, bike_id**
# - should be category: **user_type, member_gender, bike_share_for_all_trip** (notice that is not a real problem, it just makes future usage of those data more convenient)
# - should be timestamp: **strat_time, end_time**
# - should be int: **member_birth_year**
#
# * Unrealistic birth year exists in **member_birth_year**
# * Data are not sorted by time order
# ### 1.3 Cleaning data
# Make a copy of the original dataset:
df_cleaned = df.copy()
# Let's first fix the data type problems:
#
# * Data type errors:
# - should be object: **start_station_id, end_station_id, bike_id**
# - should be category: **user_type, member_gender, bike_share_for_all_trip** (notice that is not a real problem, it just makes future usage of those data more convenient)
# - should be timestamp: **strat_time, end_time**
#
# **Define**
#
# Use pandas astype and to_datetime functions to convert the data type of those columns
#
# **Code**
# Convert timestamps:
df_cleaned.start_time = pd.to_datetime(df_cleaned.start_time)
df_cleaned.end_time = pd.to_datetime(df_cleaned.end_time)
# Prepare category data:
df_cleaned.user_type = df_cleaned.user_type.astype('category')
df_cleaned.member_gender = df_cleaned.member_gender.astype('category')
df_cleaned.bike_share_for_all_trip = df_cleaned.bike_share_for_all_trip.astype('category')
# Change IDs to string instead of float:
df_cleaned.bike_id = df_cleaned.bike_id.astype(str)
df_cleaned.start_station_id = df_cleaned.start_station_id.astype(str)
df_cleaned.end_station_id = df_cleaned.end_station_id.astype(str)
# **Test**
df_cleaned.head()
df_cleaned.info(null_counts=True)
# I did not convert the member_birth_year column here because we can do some feature engineering on this column to make it more clear.
# **Define**
#
# Convert the member_birth_year to member_age which can make it more clear for our analysis
# **Code**
df_cleaned['member_age'] = 2019 - df_cleaned['member_birth_year']
# **Test**
df_cleaned.info(null_counts=True)
# **Define**
#
# The problems with the age and birth year column is still existing: data type and outlier values
# **Code**
df_cleaned.member_age.sort_values(ascending=False)
# There are many values higher than 100, which is abnormal, I need to set a threshold and delete those abnormal values higher than the threshold
df_cleaned.member_age.describe(percentiles = [.995])
# Since 70 years old will cover 99.5% of the age distribution, and the smallest age is 19, I decide to use 69 years old as the threshold
df_cleaned = df_cleaned.query('member_age <= 69')
# Now we can convert the data type of those two columns to int
df_cleaned.member_age = df_cleaned.member_age.astype(int)
df_cleaned.member_birth_year = df_cleaned.member_birth_year.astype(int)
# **Test**
df_cleaned.member_age.describe()
df_cleaned.info(null_counts=True)
df_cleaned.member_age.plot.hist();
# **Define**
#
# Extract the month, day of the week, and hour from the start_time column. Calculate the duration in minutes. This is not a real data quality or tidiness problem, it just makes the later analysis easier.
# **Code**
df_cleaned['start_time_month'] = df_cleaned['start_time'].dt.strftime('%B')
df_cleaned['start_time_month_num'] = df_cleaned['start_time'].dt.month.astype(int)
df_cleaned['start_time_weekday'] = df_cleaned['start_time'].dt.strftime('%a')
df_cleaned['start_time_hour'] = df_cleaned['start_time'].dt.hour
df_cleaned['duration_min'] = df_cleaned['duration_sec']/60
df_cleaned['duration_min'] = df_cleaned['duration_min'].astype(int)
# **Test**
df_cleaned.head()
df_cleaned.info(null_counts=True)
# **Define**
# * Data is not sorted by time order, sort the dataset to make it follows the time order: from oldest record to latest record
#
# **Code**
df_cleaned = df_cleaned.sort_values(by='start_time')
# **Test**
df_cleaned.head()
# Now the dataset follows the time order from oldest record to latest record.
# ### 1.4 Storing data
df_cleaned.to_csv('bike_clean.csv', index=False)
# Read the data again for future analysis:
df = pd.read_csv('bike_clean.csv')
#
# ### What is the structure of your dataset?
#
# > The cleaned dataset has 1743806 rows and 22 columns, including ID, name, latitude, longitude of start and end station, start time and data, end time and date, bike ID, user type, member year of birth and member gender.
# In addition to the original 16 columns, I added 6 new columns derived from feature engineering: **member_age, start_time_month, start_time_month_num, start_time_weekday, start_time_hour, duration_min**.
#
# ### What is/are the main feature(s) of interest in your dataset?
#
# > * Time date data can help us check which month of a year, which weekday of a week people tend to use this service, base on that, business decision on promotions can be made to improve the revenue.
# > * User data: user type, gender and age can be used to figure out which kind of people are willing to pay for this sharing bike product, advertisement strategy can be adjusted according to those findings.
#
# ### What features in the dataset do you think will help support your investigation into your feature(s) of interest?
#
# > Trip duration can also be a good help, if we can find the trip duration distribution pattern, we can adjust the pricing strategy, and also predict the average lifetime of a random bicycle.
# ## 2. Univariate Exploration
#
#
# The dataset covered the whole year of 2018, it is ordinary that one would like to check the monthly usage of Ford GoBike:
# **Question**
# * What is the distribution of monthly usage of the Ford GoBike?
# **Code**
# +
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
base_color = sns.color_palette()[0]
# -
month_plt = sns.catplot(data=df, x='start_time_month', kind='count', color=base_color, aspect=1.5);
month_plt.set_axis_labels('Month', 'Number of Rides'),
month_plt.fig.suptitle('Ford GoBike Monthly Usage', fontsize=14, fontweight='bold');
month_plt.set_xticklabels(rotation=50);
# * According to the plot, there is a pattern that people tend to ride more frequently during Summer and mid autumn: May to October, in which October is the most popular month; January, February, and March are months with the lowest rides. This should be attributed to the weather and climate of Bay Area.
# Just like the monthly usage, the weekly usage of the Ford GoBike should also unveil some patterns
#
# **Question**
#
# * What is the distribution of weekly usage of the Ford GoBike?
#
# **Code**
# +
weekday = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
week_plt = sns.catplot(data=df, x='start_time_weekday', kind='count', color=base_color, order=weekday)
week_plt.set_axis_labels('Weekdays', 'Number of Rides')
week_plt.fig.suptitle('Ford GoBike Weekly Usage', fontsize=14, fontweight='bold');
week_plt.set_xticklabels(rotation=50);
# -
# * This plot shows that people ride bike twice the frequency in weekdays than weekends. It indicates that most people ride Ford GoBike for commuting.
# What about the hourly usage distribution?
# **Question**
#
# * What is the distribution of hourly usage of the Ford GoBike?
#
# **Code**
# +
hour_plt = sns.catplot(data=df, x='start_time_hour', kind='count', color=base_color, aspect=1.7);
hour_plt.set_axis_labels('Hours', 'Number of Rides')
hour_plt.fig.suptitle('Ford GoBike Hourly Usage', fontsize=14, fontweight='bold');
# -
# * Most rides are occurring on rush hours. It enhanced the hypothesis we framed on the weekly plot: most usage of Ford GoBike is commuting.
# **Question**
#
# * What is the duration distribution of Ford GoBike data?
#
# **Code**
# Let's first decide the bin_edges we'd like to use in the histogram:
df.duration_min.describe()
# +
bin_edges = np.arange(0, 45, 1)
ticks = [0, 5, 10, 15, 20, 25, 30, 35, 40, 45]
labels = ['{}'.format(val) for val in ticks]
plt.hist(data=df, x='duration_min', bins=bin_edges);
plt.title("Ford GoBike Trip Duration in Minutes", fontsize=14, fontweight='bold')
plt.xlabel('Duration (Min)')
plt.xticks(ticks, labels)
plt.ylabel('Number of Rides');
# -
# * Trip duration data is right skewed with a mean of nearly 10 minutes. It indicates people ride bikes mainly within a short distance.
# **Question**
#
# * What is the distribution of user's age?
#
# **Code**
plt.hist(data=df, x='member_age');
plt.title("Ford GoBike User's Age", fontsize=14, fontweight='bold')
plt.xlabel('Age of Members (Years)')
plt.ylabel('Number of Bike Trips');
# * Majority of the users falls between the age of 25 to 40. The distribution is right skewed.
# **Question**
#
# * What is the users' proportion of different genders?
#
# **Code**
# +
user_gender = df.member_gender.value_counts()
labels=['Male', 'Female', 'Other']
plt.pie(user_gender, labels=labels, startangle=90,
autopct= '%1.1f%%', counterclock = False);
plt.title('Ford GoBike User Proportion by Gender', fontsize=14, fontweight='bold');
# -
# * 73.4% of the users are male, while 25.0% of the users are female and 1.5% of the users are other genders.
# **Question**
#
# * What is the users' proportion by user type (customer or subscriber)?
#
# **Code**
#
# +
user_type = df.user_type.value_counts()
labels=['Subscriber', 'Customer']
plt.pie(user_type, labels=labels, startangle=90,
autopct= '%1.1f%%', counterclock = False);
plt.title('Ford GoBike User Proportion by Type', fontsize=14, fontweight='bold');
# -
# ### Discuss the distribution(s) of your variable(s) of interest. Were there any unusual points? Did you need to perform any transformations?
#
# > There are some general patterns found in the dataset:
# * people tend to ride more from May to October
# * most people use the bike for commuting
# * the average duration per ride is around 10 minutes
# * the major gender of the users is male
# * the majority of user type is subscriber
#
# ### Of the features you investigated, were there any unusual distributions? Did you perform any operations on the data to tidy, adjust, or change the form of the data? If so, why did you do this?
#
# > Yes, as the user ages have some unrealistic outliers that higher than 100 years old, to filter those outliers our, I chose a threshold of 69 years old which already covered 99.5% of the user.
# ## 3. Bivariate Exploration
#
# **Question**
#
# * Is the Ford GoBike Monthly Usage different by Gender?
#
# **Code**
g = sns.catplot(data=df, x='start_time_month', col="member_gender", kind='count',
sharey=False, color=base_color)
g.set_axis_labels("Month", "Bike Rides")
g.set_titles("{col_name}")
g.set_xticklabels(rotation=50)
g.fig.suptitle('Monthly Usage by Gender', y=1.05, fontsize=14, fontweight='bold');
# * Those distributions are nearly the same. People ride more from May to October, no matter which gender they are.
# **Question**
#
# * Is the Ford GoBike Weekly Usage different by gender?
#
# **Code**
weekday = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
g = sns.catplot(data=df, x='start_time_weekday', col="member_gender", kind='count',
sharey=False, color=base_color, order=weekday)
g.set_axis_labels("Weekday", "Bike Rides")
g.set_titles("{col_name}")
g.set_xticklabels(rotation=50)
g.fig.suptitle('Weekly Usage by Gender', y=1.05, fontsize=14, fontweight='bold');
# * Those distributions are nearly the same. People ride more on weekdays, no matter which gender they are.
# **Question**
#
# * Is the Ford GoBike Monthly Usage different by user type(subscriber and customer)?
#
# **Code**
g = sns.catplot(data=df, x='start_time_month', col="user_type", kind='count',
sharey=False, color=base_color)
g.set_axis_labels("Month", "Bike Rides")
g.set_titles("{col_name}")
g.set_xticklabels(rotation=50)
g.fig.suptitle('Monthly Usage by User Type', y=1.05, fontsize=14, fontweight='bold');
# * The distributions remain the same. People ride more from May to October no matter which user type they are. So, there is no need to adjust the advertising strategy for different user types.
# **Question**
#
# * Is the Ford GoBike Weekly Usage different by user type(subscriber and customer)?
#
# **Code**
weekday = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
g = sns.catplot(data=df, x='start_time_weekday', col="user_type", kind='count',
sharey=False, color=base_color, order=weekday)
g.set_axis_labels("Weekday", "Bike Rides")
g.set_titles("{col_name}")
g.set_xticklabels(rotation=50)
g.fig.suptitle('Weekly Usage by User Type', y=1.05, fontsize=14, fontweight='bold');
# * We get some findings here! Compared with subscribers, customers are more likely to ride on weekends. Providing promotions for customers on weekends might be a reasonable way to increasing the revenue.
# **Question**
#
# * Is the Ford GoBike Hourly Usage different by user type(subscriber and customer)?
#
# **Code**
g = sns.catplot(data=df, x='start_time_hour', col="user_type", kind='count',
sharey=False, color=base_color)
g.set_axis_labels("Hours", "Bike Rides")
g.set_titles("{col_name}")
g.fig.suptitle('Hourly Usage by User Type', y=1.05, fontsize=14, fontweight='bold');
# * Yes, seem like customers ride bike often in between 8 am to 7 pm, while subscribers often ride bike on rush hours.
# **Question**
#
# * Is the duration time per ride different between user types?
#
# **Code**
# +
g = sns.catplot(data=df, y='duration_min', col="user_type", kind='violin', color=base_color)
g.set_titles(col_template = '{col_name}')
g.set_axis_labels("", "Duration (Min)")
g.fig.suptitle('Ford GoBike Duration by User Type', y=1.05, fontsize=14, fontweight='bold');
# -
# These distributions is skewed, I should build this visualization again with data subset of duration_min lower than 60 minutes.
# +
low_duration = df.query('duration_min < 60')
g = sns.catplot(data=low_duration, y='duration_min', col="user_type", kind='violin', color=base_color)
g.set_titles(col_template = '{col_name}')
g.set_axis_labels("", "Duration (Min)")
g.fig.suptitle('Ford GoBike Duration by User Type', y=1.05, fontsize=14, fontweight='bold');
# -
# * On average, customers tend to spend more time per ride compared with subscribers.
# ### Talk about some of the relationships you observed in this part of the investigation. How did the feature(s) of interest vary with other features in the dataset?
#
# > Generally, people tend to ride more in summer and autumn, rush hour, and weekdays. But there are some differences lies between customer usage and subscriber usage: customers tend to have higher duration per ride, customers tend to ride on weekends.
#
# ### Did you observe any interesting relationships between the other features (not the main feature(s) of interest)?
#
# > There is some difference of the bike usage between customers and subscribers: customers tend to spend more time per ride, and more likely to ride on weekends instead of weekdays.
# ## 4. Multivariate Exploration
#
# **Question**
#
# * What is the monthly bike usage by user type and then by gender?
#
# **Code**
g = sns.catplot(data=df, x='start_time_month', col="user_type", hue="member_gender",
kind='count', sharey=False, legend=False)
g.set_axis_labels("Month", "Bike Rides")
g.set_xticklabels(rotation=50)
g.set_titles("{col_name}")
plt.legend(loc='upper left')
g.fig.suptitle('Monthly Usage Per User Type and Gender', y=1.05, fontsize=14, fontweight='bold');
# * The plot shows no much difference compared with the bivariate one, we can say that gender and user type have no interactions on monthly usage of Ford GoBike. There is another point we can see from the plot that customer has higher female proportion than subscribers.
# **Question**
#
# * What is the weekly bike usage by user type and then by gender?
#
# **Code**
weekday = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
g = sns.catplot(data=df, x='start_time_weekday', col="user_type", hue="member_gender",
kind='count', sharey=False, legend=False, order=weekday)
g.set_axis_labels("Weekday", "Bike Rides")
g.set_xticklabels(rotation=50)
g.set_titles("{col_name}")
plt.legend(loc='upper left')
g.fig.suptitle('Weekly Usage Per User Type and Gender', y=1.05, fontsize=14, fontweight='bold');
# * There is no much difference compared with the bivariate plot.
# **Question**
#
# * What is the hourly bike usage by user type and then by gender?
#
# **Code**
# +
g = sns.catplot(data=df, x='start_time_hour', col="user_type", hue="member_gender",
kind='count', sharey=False, legend=False)
g.set_axis_labels("Hours", "Bike Rides")
g.set_titles("{col_name}")
plt.legend(loc='upper left')
g.fig.suptitle('Hourly Usage Per User Type and Gender', y=1.05, fontsize=14, fontweight='bold');
# -
# * No new findings compared with the bivariate plot.
# **Question**
#
# * Since the main different between subscriber and customer is on hours and weekdays, we can build a heat map with weekday and hour to illustrate the different trend of riding Ford GoBike between them.
#
# **Code**
# +
df['start_time_weekday'] = pd.Categorical(df['start_time_weekday'],
categories=['Mon','Tue','Wed','Thu','Fri','Sat', 'Sun'],
ordered=True)
plt.figure(figsize=(10, 8))
plt.suptitle('Usage Heatmap for Customers and Subscribers', y=1.05, fontsize=14, fontweight='bold')
# plot for subscriber
plt.subplot(1, 2, 1)
df_subscriber = df.query('user_type == "Subscriber"').groupby(["start_time_hour", "start_time_weekday"])["bike_id"].size().reset_index()
df_subscriber = df_subscriber.pivot("start_time_hour", "start_time_weekday", "bike_id")
sns.heatmap(df_subscriber, cmap="YlGnBu")
plt.title('Subscriber', y=1.01)
plt.xlabel('Weekday')
plt.ylabel('Hour')
# plot for customer
plt.subplot(1, 2, 2)
df_customer = df.query('user_type == "Customer"').groupby(["start_time_hour", "start_time_weekday"])["bike_id"].size().reset_index()
df_customer = df_customer.pivot("start_time_hour", "start_time_weekday", "bike_id")
sns.heatmap(df_customer, cmap="YlGnBu")
plt.title('Customer', y=1.01)
plt.xlabel('Weekday')
plt.ylabel('');
# -
# * This plot an appropriate example of how multi-dimensional visualization can carry more information. According to the heat map, we can conclude those popular time periods for customer to ride Ford GoBike are 5 pm in weekdays and 11 am to 5 pm in weekends.
# ### Talk about some of the relationships you observed in this part of the investigation. Were there features that strengthened each other in terms of looking at your feature(s) of interest?
#
# > The findings in bivariate exploratory part are proven to be right in multivariate exploratory part. In the heat map, I found that is a good example of how multi-dimensional visualization can carry more information if appropriate encoding was used. According to these plot, there is a different trend between subscribers and customers that the most popular time periods for customer to ride Ford GoBike are 5 pm in weekdays and 11 am to 5 pm in weekends while subscriber's is rush hours in weekdays. Based on that, I think there are bunch of promotional actions can be applied to increase the revenue get from customers.
#
# ### Were there any interesting or surprising interactions between features?
#
# > No, there are no interactions.
|
communicate_data_findings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Making Zarr data from NetCDF files
#
# - Funding: Interagency Implementation and Advanced Concepts Team [IMPACT](https://earthdata.nasa.gov/esds/impact) for the Earth Science Data Systems (ESDS) program and AWS Public Dataset Program
# - Software developed during [OceanHackWeek 2020](https://github.com/oceanhackweek)
#
# ### Credits: Tutorial development
# * [Dr. <NAME>](mailto:<EMAIL>) - [Twitter](https://twitter.com/ChelleGentemann) - Farallon Institute
# * [<NAME>](mailto:<EMAIL>) - [Twitter](https://twitter.com/clifgray) - Duke University
# * [<NAME>](mailto:<EMAIL>) - University of Southampton
#
# ## Why data format matters
# - NetCDF sprinkles metadata throughout files, making them slow to access and read data
# - Zarr consolidates the metadata, making them FAST for access and reading
#
# +
# filter some warning messages
import warnings
warnings.filterwarnings("ignore")
#libraries
import datetime as dt
import xarray as xr
import fsspec
import s3fs
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
# make datasets display nicely
xr.set_options(display_style="html")
import os.path
#magic fncts #put static images of your plot embedded in the notebook
# %matplotlib inline
plt.rcParams['figure.figsize'] = 12, 6
# %config InlineBackend.figure_format = 'retina'
# -
def get_geo_data(sat,lyr,idyjl):
# arguments
# sat goes-east,goes-west,himawari
# lyr year
# idyjl day of year
ds,iexist=[],False
d = dt.datetime(lyr,1,1) + dt.timedelta(days=idyjl)
fs = s3fs.S3FileSystem(anon=True) #connect to s3 bucket!
#create strings for the year and julian day
imon,idym=d.month,d.day
syr,sjdy,smon,sdym = str(lyr).zfill(4),str(idyjl).zfill(3),str(imon).zfill(2),str(idym).zfill(2)
#use glob to list all the files in the directory
if sat=='goes-east':
file_location,var = fs.glob('s3://noaa-goes16/ABI-L2-SSTF/'+syr+'/'+sjdy+'/*/*.nc'),'SST'
if sat=='goes-west':
file_location,var = fs.glob('s3://noaa-goes17/ABI-L2-SSTF/'+syr+'/'+sjdy+'/*/*.nc'),'SST'
if sat=='himawari':
file_location,var = fs.glob('s3://noaa-himawari8/AHI-L2-FLDK-SST/'+syr+'/'+smon+'/'+sdym+'/*/*L2P*.nc'),'sea_surface_temperature'
#make a list of links to the file keys
if len(file_location)<1:
return file_ob
file_ob = [fs.open(file) for file in file_location] #open connection to files
#open all the day's data
with xr.open_mfdataset(file_ob,combine='nested',concat_dim='time') as ds:
iexist = True #file exists
#clean up coordinates which are a MESS in GOES
#rename one of the coordinates that doesn't match a dim & should
if not sat=='himawari':
ds = ds.rename({'t':'time'})
ds = ds.reset_coords()
else:
ds = ds.rename({'ni':'x','nj':'y'})
#put in to Celsius
#ds[var] -= 273.15 #nice python shortcut to +- from itself a-=273.15 is the same as a=a-273.15
#ds[var].attrs['units'] = '$^\circ$C'
return ds,iexist
# ## Open GOES-16 (East Coast) Data
# - Careful of what you ask for.... each day is about 3 min to access
# +
# %%time
lyr = 2020
satlist = ['goes-east','goes-west','himawari']
for sat in satlist:
init = 0 #reset new data store
for idyjl in range(180,201): #6/28/2020-7/18/2020
print('starting ', idyjl)
ds,iexist = get_geo_data(sat,lyr,idyjl)
if not iexist:
continue
print('writing zarr store')
if init == 0:
ds.to_zarr(sat)
init = 1
else:
ds.to_zarr(sat,append_dim='time')
# -
# #### Now write this to our shared AWS S3 bucket
#
# Note that in order to do this you need the aws command line tools which can be installed by running from the command line
#
# `pip install awscli`
#
# `aws s3 sync ./goes_east s3://ohw-bucket/goes_east`
#
# `aws s3 sync ./goes_west s3://ohw-bucket/goes_west`
#
# `aws s3 sync ./goes_west s3://ohw-bucket/himawari`
#
# #### note that putting the ! in front of a command in jupyter send it to the terminal so you could run it here with
#
#
# ! pip install awscli
# ! aws s3 sync ./goes_east s3://ohw-bucket/goes_east
# ! aws s3 sync ./goes_west s3://ohw-bucket/goes_west
# ! aws s3 sync ./goes_west s3://ohw-bucket/himawari
# ## Test reading the data
# +
# %%time
file_location = 's3://ohw-bucket/goes_east'
ds = xr.open_zarr(fsspec.get_mapper(file_location,anon=False))
ds
# -
|
notebooks/netcdf_to_zarr_GEO_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import skimage.io
from skimage.transform import resize
from imgaug import augmenters as iaa
from tqdm import tqdm
import PIL
from PIL import Image
import cv2
from sklearn.utils import class_weight, shuffle
from keras.utils import multi_gpu_model, multi_gpu_utils
# create callbacks list
from keras.callbacks import ModelCheckpoint, LearningRateScheduler, EarlyStopping, ReduceLROnPlateau
from sklearn.model_selection import train_test_split
import tensorflow as tf
import warnings
warnings.filterwarnings("ignore")
SIZE = 512
# +
# Load dataset info
path_to_train = '../../Human_Protein_Atlas/input/train/'
data = pd.read_csv('../../Human_Protein_Atlas/input/train.csv')
rare_classes = [8,9,10,15,20,24,27]
train_dataset_info = []
for name, labels in zip(data['Id'], data['Target'].str.split(' ')):
train_dataset_info.append({
'path':os.path.join(path_to_train, name),
'labels':np.array([int(label) for label in labels])})
train_dataset_info = np.array(train_dataset_info)
class data_generator:
@staticmethod
def create_train(dataset_info, batch_size, shape, augument=True, heavy_augment_rares=True, oversample_factor = 0):
assert shape[2] == 3
if oversample_factor > 0:
rare_dataset_info = np.array([item for item in dataset_info if np.isin(item['labels'], rare_classes).any()])
#rare_dataset_info = shuffle(rare_dataset_info)
for i in range(oversample_factor):
#dataset_info
dataset_info = np.append(dataset_info,rare_dataset_info)
while True:
dataset_info = shuffle(dataset_info)
for start in range(0, len(dataset_info), batch_size):
end = min(start + batch_size, len(dataset_info))
batch_images = []
X_train_batch = dataset_info[start:end]
batch_labels = np.zeros((len(X_train_batch), 28))
for i in range(len(X_train_batch)):
image = data_generator.load_image(X_train_batch[i]['path'], shape)
#image = preprocess_input(image)
rare = np.isin(X_train_batch[i]['labels'], rare_classes).any()
if augument:
if heavy_augment_rares and rare:
image = data_generator.heavy_augment(image)
else:
image = data_generator.augment(image)
batch_images.append(image)
batch_labels[i][X_train_batch[i]['labels']] = 1
yield np.array(batch_images, np.float32), batch_labels
@staticmethod
def load_image(path, shape):
image = cv2.imread(path + '.png', cv2.IMREAD_UNCHANGED)
return image
@staticmethod
def augment(image):
augment_img = iaa.Sequential([
iaa.OneOf([
iaa.Affine(rotate=0),
iaa.Affine(rotate=90),
iaa.Affine(rotate=180),
iaa.Affine(rotate=270),
iaa.Fliplr(0.5),
iaa.Flipud(0.5),
])], random_order=True)
image_aug = augment_img.augment_image(image)
return image_aug
@staticmethod
def heavy_augment(image):
augment_img = iaa.Sequential([
iaa.OneOf([
iaa.Affine(scale=(0.5,2.0)),
iaa.Affine(shear=15),
iaa.Affine(rotate=0),
iaa.Affine(rotate=35),
iaa.Affine(rotate=90),
iaa.Affine(rotate=180),
iaa.Affine(rotate=270),
iaa.Affine(translate_percent=0.1),
iaa.Fliplr(0.5),
iaa.Flipud(0.5),
iaa.Noop()
])], random_order=True)
image_aug = augment_img.augment_image(image)
return image_aug
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential, load_model
from keras.layers import Activation, Dropout, Flatten, Dense, GlobalMaxPooling2D, BatchNormalization, Input, Conv2D
from keras.applications.inception_v3 import InceptionV3
from keras.callbacks import ModelCheckpoint
from keras import metrics
from keras.optimizers import Adam
from keras import backend as K
import keras
from keras.models import Model
# -
train_dataset_info
rare_classes = ["Endosomes", "Lysosomes", "Rods & rings"]
def create_model(input_shape, n_out):
input_tensor = Input(shape=input_shape)
base_model = InceptionResNetV2(include_top=False,
weights='imagenet',
input_shape=input_shape)
bn = BatchNormalization()(input_tensor)
x = base_model(bn)
x = Conv2D(32, kernel_size=(1,1), activation='relu')(x)
x = Flatten()(x)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.5)(x)
output = Dense(n_out, activation='sigmoid')(x)
model = Model(input_tensor, output)
return model
epochs = 30; batch_size = 16
checkpoint = ModelCheckpoint('../../Human_Protein_Atlas/working/InceptionV3_512.h5', monitor='val_loss', verbose=1,
save_best_only=True, mode='min', save_weights_only = True)
reduceLROnPlat = ReduceLROnPlateau(monitor='val_loss', factor=0.1, patience=6,
verbose=1, mode='auto', epsilon=0.00001)
early = EarlyStopping(monitor="val_loss",
mode="min",
patience=10)
callbacks_list = [checkpoint, early, reduceLROnPlat]
# +
# split data into train, valid
indexes = np.arange(train_dataset_info.shape[0])
np.random.shuffle(indexes)
train_indexes, valid_indexes = train_test_split(indexes, test_size=0.1, random_state=18)
# create train and valid datagens
train_generator = data_generator.create_train(
train_dataset_info[train_indexes], batch_size, (SIZE,SIZE,3), augument=True)
validation_generator = data_generator.create_train(
train_dataset_info[valid_indexes], 32, (SIZE,SIZE,3), augument=False)
# +
with tf.device('/cpu:0'):
input_shape=(SIZE,SIZE,3)
input_tensor = Input(shape=input_shape)
base_model = InceptionV3(include_top=False,
weights='imagenet',
input_shape=input_shape)
bn = BatchNormalization()(input_tensor)
x = base_model(bn)
x = Conv2D(32, kernel_size=(1,1), activation='relu')(x)
x = Flatten()(x)
x = Dropout(0.5)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.5)(x)
output = Dense(28, activation='sigmoid')(x)
model = Model(input_tensor, output)
# -
model = multi_gpu_model(model, gpus=2)
class data_generator:
def create_train(dataset_info, batch_size, shape, augument=True):
assert shape[2] == 3
while True:
dataset_info = shuffle(dataset_info)
for start in range(0, len(dataset_info), batch_size):
end = min(start + batch_size, len(dataset_info))
batch_images = []
X_train_batch = dataset_info[start:end]
batch_labels = np.zeros((len(X_train_batch), 28))
for i in range(len(X_train_batch)):
image = data_generator.load_image(
X_train_batch[i]['path'], shape)
if augument:
image = data_generator.augment(image)
batch_images.append(image/255.)
batch_labels[i][X_train_batch[i]['labels']] = 1
yield np.array(batch_images, np.float32), batch_labels
def load_image(path, shape):
image_red_ch = Image.open(path+'_red.png')
image_yellow_ch = Image.open(path+'_yellow.png')
image_green_ch = Image.open(path+'_green.png')
image_blue_ch = Image.open(path+'_blue.png')
image = np.stack((
np.array(image_red_ch),
np.array(image_green_ch),
np.array(image_blue_ch)), -1)
image = cv2.resize(image, (shape[0], shape[1]))
return image
def augment(image):
augment_img = iaa.Sequential([
iaa.OneOf([
iaa.Affine(rotate=0),
iaa.Affine(rotate=90),
iaa.Affine(rotate=180),
iaa.Affine(rotate=270),
iaa.Fliplr(0.5),
iaa.Flipud(0.5),
])], random_order=True)
image_aug = augment_img.augment_image(image)
return image_aug
# +
for layer in model.layers:
layer.trainable = False
model.layers[-1].trainable = True
model.layers[-2].trainable = True
model.layers[-3].trainable = True
model.layers[-4].trainable = True
model.layers[-5].trainable = True
model.compile(
loss='binary_crossentropy',
optimizer=Adam(1e-03),
metrics=['acc'])
# model.summary()
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=2,
verbose=1)
# +
epochs = 30
# train all layers
for layer in model.layers:
layer.trainable = True
model.compile(loss='binary_crossentropy',
optimizer=Adam(lr=1e-5),
metrics=['accuracy'])
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=epochs,
verbose=1,
callbacks=callbacks_list)
# -
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=epochs,
verbose=1,
callbacks=callbacks_list)
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=epochs,
verbose=1,
callbacks=callbacks_list)
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=epochs,
verbose=1,
callbacks=callbacks_list)
model.fit_generator(
train_generator,
steps_per_epoch=np.ceil(float(len(train_indexes)) / float(batch_size)),
validation_data=validation_generator,
validation_steps=np.ceil(float(len(valid_indexes)) / float(batch_size)),
epochs=epochs,
verbose=1,
callbacks=callbacks_list)
|
wienerschnitzelgemeinschaft/src/Bojan/InceptionV3_512.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="hjSlbalwC-ro" colab_type="code" colab={}
# shared LSTM / siamese LSTM
from keras import layers
from keras import Input
from keras.models import Model
lstm = layers.LSTM(32)
left_input = Input(shape=(None, 128))
left_output = lstm(left_input)
right_input = Input(shape=(None, 128))
# not! right_output = left_output(right_input)
right_output = lstm(right_output)
merged = layers.concatenate([left_output, right_output], axis=-1)
predictins = layers.Dense(1, activation='sigmoid')(merged)
model = Model([left_input, right_input], predictions)
model.fit([left_data, right_data], targets)
# + id="ql8S9g0kJCtm" colab_type="code" colab={}
from keras import layers
from keras import applications
from keras import Input
xception_base = applications.Xception(weights=None, include_top=False)
left_input = Input(shape=(250, 250, 3))
right_input = Input(shape=(250, 250, 3))
left_features = xception_base(left_input)
right_features = xception_base(right_input)
merged_featrues = layers.concatenate([left_features, right_features], axis=-1)
|
7/7-1-6.ipynb
|