code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Agrupamento Isolation Forest - Experimento
#
# Este é um componente que treina um modelo Isolation Forest usando [Scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.IsolationForest.html). <br>
# Scikit-learn é uma biblioteca open source de machine learning que suporta apredizado supervisionado e não supervisionado. Também provê várias ferramentas para ajustes de modelos, pré-processamento de dados, seleção e avaliação de modelos, além de outras funcionalidades.
#
# Este notebook apresenta:
# - como usar o [SDK](https://platiagro.github.io/sdk/) para carregar datasets, salvar modelos e outros artefatos.
# - como declarar parâmetros e usá-los para criar componentes reutilizáveis.
# ## Declaração de parâmetros e hiperparâmetros
#
# Declare parâmetros com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKBqlpGMRcVMdkUMvKID3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMOEYuFFpZbmBUNlXiKOKyoGuULGZcVzluc1VKFNe7JXxjMacsprtMcRAyLiCMBETIq2EAJFiK0aqSYSNJ+1MM/4PgT5JLJtQFGjnmUoUJy/OB/8LtbMz854SYFo0D7i21/DAOBXaBete3vY9uunwD+Z+BKa/rLNWDmk/RqUwsfAb3bwMV1U5P3gMsdoP9JlwzJkfw0hXweeD+jb8oCfbdA16rbW2Mfpw9AmrpaugEODoGRAmWveby7s7W3f880+vsBocZyukMJsmwAAAAGYktHRAD/AP8A/6C9p5MAAAAJcEhZcwAADdcAAA3XAUIom3gAAAAHdElNRQfkBgsMIwnXL7c0AAACDUlEQVQ4y92UP4gTQRTGf29zJxhJZ2NxbMBKziYWlmJ/ile44Nlkd+dIYWFzItiNgoIEtFaTzF5Ac/inE/urtLWxsMqmUOwCEpt1Zmw2xxKi53XitPO9H9978+aDf/3IUQvSNG0450Yi0jXG7C/eB0cFeu9viciGiDyNoqh2KFBrHSilWstgnU7nFLBTgl+ur6/7PwK11kGe5z3n3Hul1MaiuCgKDZwALHA7z/Oe1jpYCtRaB+PxuA8kQM1aW68Kt7e3zwBp6a5b1ibj8bhfhQYVZwMRiQHrvW9nWfaqCrTWPgRWvPdvsiy7IyLXgEJE4slk8nw+T5nDgDbwE9gyxryuwpRSF5xz+0BhrT07HA4/AyRJchUYASvAbhiGaRVWLIMBYq3tAojIszkMoNRulbXtPM8HwV/sXSQi54HvQRDcO0wfhGGYArvAKjAq2wAgiqJj3vsHpbtur9f7Vi2utLx60LLW2hljEuBJOYu9OI6vAzQajRvAaeBLURSPlsBelA+VhWGYaq3dwaZvbm6+m06noYicE5ErrVbrK3AXqHvvd4bD4Ye5No7jSERGwKr3Pms2m0pr7Rb30DWbTQWYcnFvAieBT7PZbFB1V6vVfpQaU4UtDQetdTCZTC557/eA48BlY8zbRZ1SqrW2tvaxCvtt2iRJ0i9/xb4x5uJRwmNlaaaJ3AfqIvKY/+78Av++6uiSZhYMAAAAAElFTkSuQmCC" /> na barra de ferramentas.<br>
# O parâmetro `dataset` identifica os conjuntos de dados. Você pode importar arquivos de dataset com o botão <img src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAABQAAAAUCAYAAACNiR0NAAABhWlDQ1BJQ0MgcHJvZmlsZQAAKJF9kT1Iw0AcxV9TtaIVBzuIOASpThb8QhylikWwUNoKrTqYXPohNGlIUlwcBdeCgx+LVQcXZ10dXAVB8APEydFJ0UVK/F9SaBHjwXE/3t173L0DhFqJqWbbGKB<KEY>3QhiCOMSM/V4aiENz/F1Dx9f7yI8y/vcn6NHyZkM8InEs0w3LOJ14ulNS+e8TxxiRUkhPiceNeiCxI9cl11+41xwWOCZISOdnCMO<KEY> /> na barra de ferramentas.
# + tags=["parameters"]
# parâmetros
dataset = "iris.csv" #@param {type:"string"}
max_samples = "auto" #@param {type:"float"}
contamination = 0.1 #@param {type:"float"}
max_features = 1.0 #@param {type:"float"}
# -
# ## Acesso ao conjunto de dados
#
# O conjunto de dados utilizado nesta etapa será o mesmo carregado através da plataforma.<br>
# O tipo da variável retornada depende do arquivo de origem:
# - [pandas.DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html) para CSV e compressed CSV: .csv .csv.zip .csv.gz .csv.bz2 .csv.xz
# - [Binary IO stream](https://docs.python.org/3/library/io.html#binary-i-o) para outros tipos de arquivo: .jpg .wav .zip .h5 .parquet etc
# +
import pandas as pd
df = pd.read_csv(f'/tmp/data/{dataset}')
# -
# ## Acesso aos metadados do conjunto de dados
#
# Utiliza a função `stat_dataset` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para carregar metadados. <br>
# Por exemplo, arquivos CSV possuem `metadata['featuretypes']` para cada coluna no conjunto de dados (ex: categorical, numerical, or datetime).
# +
import numpy as np
from platiagro import stat_dataset
metadata = stat_dataset(name=dataset)
featuretypes = metadata["featuretypes"]
featuretypes = np.array(featuretypes)
columns = df.columns.to_numpy()
# -
# ## Configuração dos atributos
# +
from platiagro.featuretypes import NUMERICAL
# Selects the indexes of numerical and non-numerical features
numerical_indexes = np.where(featuretypes == NUMERICAL)[0]
non_numerical_indexes = np.where(~(featuretypes == NUMERICAL))[0]
# After the step handle_missing_values,
# numerical features are grouped in the beggining of the array
numerical_indexes_after_handle_missing_values = \
np.arange(len(numerical_indexes))
non_numerical_indexes_after_handle_missing_values = \
np.arange(len(numerical_indexes), len(featuretypes))
# -
# ## Dividindo o conjunto de dados em treino e teste
#
# Conjunto de treino: usado para ajustar o modelo.
# Conjunto de teste: usada para fornecer uma avaliação imparcial de um ajuste de modelo no conjunto de dados de treinamento.
# +
from sklearn.model_selection import train_test_split
X_train, X_test= train_test_split(df, train_size=0.7)
# -
# ## Treina modelo usando sklearn.ensemble.IsolationForest
# +
from sklearn.compose import ColumnTransformer
from sklearn.impute import SimpleImputer
from sklearn.ensemble import IsolationForest
from sklearn.pipeline import Pipeline
from category_encoders.ordinal import OrdinalEncoder
pipeline = Pipeline(steps=[
("handle_missing_values",
ColumnTransformer(
[("imputer_mean", SimpleImputer(strategy="mean"), numerical_indexes),
("imputer_mode", SimpleImputer(strategy="most_frequent"), non_numerical_indexes)],
remainder="drop")),
("handle_categorical_features",
ColumnTransformer(
[("feature_encoder", OrdinalEncoder(), non_numerical_indexes_after_handle_missing_values)],
remainder="passthrough")),
("estimator", IsolationForest(max_samples=max_samples,
contamination=contamination,
max_features=max_features))
])
pipeline.fit(X_train)
features_after_pipeline = \
np.concatenate((columns[numerical_indexes],
columns[non_numerical_indexes]))
# -
# ## Avalia desempenho
#
# No caso do Isolation Florest, podemos medir o desempenho obtendo a anomalia média.
# +
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
# Run all except the last step
df_encoded = Pipeline(steps=pipeline.steps[:-1]).transform(X_test)
# Normalization and dimension reduction
df_encoded_std = StandardScaler().fit_transform(df_encoded)
pca = PCA(n_components=2)
reduced = pca.fit_transform(df_encoded_std)
X_pca = pd.DataFrame(reduced, columns=["X", "Y"])
y_pred_test = pipeline.predict(X_test)
X_pca["Anomaly"] = y_pred_test
# +
import matplotlib.pyplot as plt
import seaborn as sns
from platiagro import save_figure
ax = sns.scatterplot(x="X", y="Y", hue="Anomaly", data=X_pca)
ax.set_title("PCA Graph", {"fontweight": 'bold'})
save_figure(figure=plt.gcf())
# -
# ## Salva métricas
#
# Utiliza a função `save_metrics` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para salvar métricas. Por exemplo: `accuracy`, `precision`, `r2_score`, `custom_score` etc.<br>
# +
from platiagro import save_metrics
save_metrics(anomaly_score=y_pred_test)
# -
# ## Salva alterações no conjunto de dados
#
# O conjunto de dados será salvo (e sobrescrito com as respectivas mudanças) localmente, no container da experimentação, utilizando a função `pandas.DataFrame.to_csv`.<br>
# +
new_columns = list("Anomaly")
score = pipeline.predict(df)
df_anomaly = df.copy()
df_anomaly["Anomaly"] = score
# save dataset changes
df.to_csv(f'/tmp/data/{dataset}', index=False)
# -
# ## Salva modelo e outros artefatos
#
# Utiliza a função `save_model` do [SDK da PlatIAgro](https://platiagro.github.io/sdk/) para salvar modelos e outros artefatos.<br>
# Essa função torna estes artefatos disponíveis para o notebook de implantação.
# +
from platiagro import save_model
save_model(pipeline=pipeline,
columns=columns,
new_columns=new_columns,
features_after_pipeline=features_after_pipeline)
|
samples/isolation-forest-clustering/Experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Loading in the libraries.
# +
# Old libraries that we know and love.
import numpy as np
import matplotlib.pylab as py
import pandas as pa
# %matplotlib inline
# Our new libraries.
from sklearn import datasets
from mpl_toolkits.mplot3d import Axes3D
import mayavi.mlab as mlab
iris = datasets.load_iris()
# -
# # Looking at the data
print iris['DESCR']
iris
X = iris['data']
y = iris['target']
py.plot(X[y==0,0],X[y==0,1],'r.')
py.plot(X[y==1,0],X[y==1,1],'g.')
py.plot(X[y==2,0],X[y==2,1],'b.')
py.plot(X[y==0,2],X[y==0,3],'r.')
py.plot(X[y==1,2],X[y==1,3],'g.')
py.plot(X[y==2,2],X[y==2,3],'b.')
fig = py.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=110)
ax.scatter(X[y==0, 0], X[y==0, 1], X[y==0, 2], c='r')
ax.scatter(X[y==1, 0], X[y==1, 1], X[y==1, 2], c='g')
ax.scatter(X[y==2, 0], X[y==2, 1], X[y==2, 2], c='b')
py.show()
mlab.clf()
mlab.points3d(X[y==0, 0], X[y==0, 1], X[y==0, 2],color=(1,0,0))
mlab.points3d(X[y==1, 0], X[y==1, 1], X[y==1, 2],color=(0,1,0))
mlab.points3d(X[y==2, 0], X[y==2, 1], X[y==2, 2],color=(0,0,1))
mlab.axes()
mlab.show()
# Is there a more principled way to look at the data? Yes! <b>Let's go back to the notes.</b>
# # More principled ways to look at the data, Principle Component Analysis (PCA)!
# Some sample data to demonstrate PCA on.
mu1 = np.array([0,0,0])
mu2 = np.array([6,0,0])
np.random.seed(123)
Sigma = np.matrix(np.random.normal(size=[3,3]))
# U,E,VT = np.linalg.svd(Sigma)
# E[0] = 1
# E[1] = 1
# E[2] = 1
# Sigma = U*np.diag(E)*VT
Xrandom1 = np.random.multivariate_normal(mu1,np.array(Sigma*Sigma.T),size=500)
Xrandom2 = np.random.multivariate_normal(mu2,np.array(Sigma*Sigma.T),size=500)
# Plot the data so that it is "spread out" as much as possible.
mlab.clf()
mlab.points3d(Xrandom1[:,0], Xrandom1[:,1], Xrandom1[:,2],color=(1,0,0))
mlab.points3d(Xrandom2[:,0], Xrandom2[:,1], Xrandom2[:,2],color=(0,1,0))
mlab.axes()
mlab.show()
# Can do the same thing with our classification data.
# +
from sklearn.decomposition import PCA
X2D = PCA(n_components=3).fit_transform(X)
py.plot(X2D[y==0,0],X2D[y==0,1],'r.')
py.plot(X2D[y==1,0],X2D[y==1,1],'g.')
py.plot(X2D[y==2,0],X2D[y==2,1],'b.')
# -
# Just as one can project from a high dimensional space to a two-dimensional space, one can also do the same thing to project to a three-dimensional space.
fig = py.figure(1, figsize=(8, 6))
ax = Axes3D(fig, elev=-150, azim=110)
X3D = PCA(n_components=3).fit_transform(X)
ax.scatter(X3D[y==0, 0], X3D[y==0, 1], X3D[y==0, 2], c='r')
ax.scatter(X3D[y==1, 0], X3D[y==1, 1], X3D[y==1, 2], c='g')
ax.scatter(X3D[y==2, 0], X3D[y==2, 1], X3D[y==2, 2], c='b')
py.show()
# And do the same with Mayavi.
mlab.clf()
mlab.points3d(X3D[y==0, 0], X3D[y==0, 1], X3D[y==0, 2],color=(1,0,0))
mlab.points3d(X3D[y==1, 0], X3D[y==1, 1], X3D[y==1, 2],color=(0,1,0))
mlab.points3d(X3D[y==2, 0], X3D[y==2, 1], X3D[y==2, 2],color=(0,0,1))
mlab.axes()
mlab.show()
# <b>Let's go back to the notes for our first algorithm.</b>
# # Our first classification tool, Linear Support Vector Machines.
# Load in the support vector machine (SVM) library
from sklearn import svm
# +
# If there is one thing that I want to harp on, it is the difference
# between testing and training errors! So, here we create a training
# set on which we computer the parameters of our algorithm, and a
# testing set for seeing how well we generalize (and work on real
# world problems).
np.random.seed(1236)
perm = np.random.permutation(len(y))
trainSize = 100
Xtrain = X[perm[:trainSize],0:2]
Xtest = X[perm[trainSize:],0:2]
yHat = np.zeros([len(y)])
# Exists a separator
#yHat[np.logical_or(y==1,y==2)] = 1
# No perfect separator
#yHat[np.logical_or(y==1,y==0)] = 1
# All the data
yHat = y
yHattrain = yHat[perm[:trainSize]]
yHattest = yHat[perm[trainSize:]]
# -
# ## ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^ <p><font color="red">But why do you do this? See the notes.</font>
# Some parameters we can get to play with
# If there is no perfect separator then how much do you penalize points
# that lay on the wrong side?
C = 100.
# The shape of the loss function for points that lay on the wrong side.
loss = 'l2'
# Run the calculation!
clf = svm.LinearSVC(loss=loss,C=C)
clf.fit(Xtrain, yHattrain)
# +
# Make some plots, inspired by scikit-learn tutorial
from matplotlib.colors import ListedColormap
# step size in the mesh for plotting the decision boundary.
h = .02
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, m_max]x[y_min, y_max].
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
py.figure(1, figsize=(8, 6))
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
py.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot also the training points
py.scatter(Xtrain[:, 0], Xtrain[:, 1], c=yHattrain, cmap=cmap_bold,marker='o')
py.scatter(Xtest[:, 0], Xtest[:, 1], c=yHattest, cmap=cmap_bold,marker='+')
py.xlim(xx.min(), xx.max())
py.ylim(yy.min(), yy.max())
py.show()
# -
# Print out some metrics
print 'training score',clf.score(Xtrain,yHattrain)
print 'testing score',clf.score(Xtest,yHattest)
# <b>Back to the notes to define our next method.</b>
# # Our second classification tool, K-nearest neighbors.
# Import the K-NN solver
from sklearn import neighbors
# +
# If there is one thing that I want to harp on, it is the difference
# between testing and training errors! So, here we create a training
# set on which we computer the parameters of our algorithm, and a
# testing set for seeing how well we generalize (and work on real
# world problems).
np.random.seed(123)
perm = np.random.permutation(len(y))
trainSize = 50
Xtrain = X[perm[:trainSize],0:2]
Xtest = X[perm[trainSize:],0:2]
ytrain = y[perm[:trainSize]]
ytest = y[perm[trainSize:]]
# +
# Some parameters to play around with
# The number of neighbors to use.
n_neighbors = 7
#weights = 'distance'
weights = 'uniform'
# -
# Run the calculation
clf = neighbors.KNeighborsClassifier(n_neighbors, weights=weights)
clf.fit(Xtrain, ytrain)
# +
# Make some plots inspired by sci-kit learn tutorial
# step size in the mesh for plotting the decision boundary.
h = .02
# Create color maps
cmap_light = ListedColormap(['#FFAAAA', '#AAFFAA', '#AAAAFF'])
cmap_bold = ListedColormap(['#FF0000', '#00FF00', '#0000FF'])
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, m_max]x[y_min, y_max].
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
py.figure(1, figsize=(8, 6))
py.pcolormesh(xx, yy, Z, cmap=cmap_light)
# Plot also the training points
py.scatter(Xtrain[:, 0], Xtrain[:, 1], c=ytrain, cmap=cmap_bold,marker='o')
py.scatter(Xtest[:, 0], Xtest[:, 1], c=ytest, cmap=cmap_bold,marker='+')
py.xlim(xx.min(), xx.max())
py.ylim(yy.min(), yy.max())
py.show()
# -
# Print out some scores.
print 'training score',clf.score(Xtrain,ytrain)
print 'testing score',clf.score(Xtest,ytest)
# <b>Back to the notes.</b>
# ## Loading in the libraries for regression.
# +
# Old libraries that we know and love.
import numpy as np
import matplotlib.pylab as py
import pandas as pa
# Our new libraries.
from sklearn import cross_validation, linear_model, feature_selection, metrics
import mayavi.mlab as mlab
# -
# # Supervised Regression
# ## Linear Regression
# Read in the data using
Xy = pa.read_csv('Advertising.csv')
# Take a look at the contents.
Xy
# Normalize data
# We do this to make plotting and processing easier. Many Sklearn functions do this
# for you behind the scenes, but we do it explicitly.
# Note, that this is a cousing of the physics idea of nondimensionalization. Think
# about the case where TV was measured in millions, while Radio was measured in
# thousands. One could imagine TV totally washing out the effect of Radio.
# In effect, after normalization, each predictor now stands on an "even footing".
#
# Is this always a good idea?
Xy = (Xy-Xy.min())/(Xy.max()-Xy.min())
Xy
# Select out our predictor columns and our response columns
X = Xy.ix[:,['TV']]
y = Xy.ix[:,['Sales']]
# Last time we did this by hand, now we are smarter and use the sklearn
# routine. This routine splits data into training and testing subsets.
cross_validation.train_test_split([1,2,3,4,5],
[6,7,8,9,10],
test_size=0.4,
random_state=5)
# Now we do it for the real data.
X_train,X_test,y_train,y_test = cross_validation.train_test_split(X,
y,
test_size=0.8)
# Let's take a quick look at the data.
X_train
# Run the solver
reg = linear_model.LinearRegression(fit_intercept=True)
reg.fit(X_train,y_train)
# There are the slope and intercept of the line we computed.
# Beta_0
print reg.intercept_
# Beta_1
print reg.coef_
# Do a plot
plotX = np.linspace(0,1,100)
plotY = reg.predict(np.matrix(plotX).T)
py.plot(X_train,y_train,'ro')
py.plot(X_test,y_test,'go')
py.plot(plotX,plotY,'b-')
# Use the metrics package to print our errors. See discussion on slides.
print 'training error'
print metrics.mean_squared_error(y_train,reg.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,reg.predict(X_test))
# <b>Back to slides.</b>
# ## Multi-dimensional regression
# +
# Select out our predictor columns and our response columns
X = Xy.ix[:,['TV','Radio']]
y = Xy.ix[:,['Sales']]
# Select subsets for training and testing
X_train,X_test,y_train,y_test = cross_validation.train_test_split(X,
y,
test_size=0.8,
random_state=123)
# -
# Plot the data to get a feel for it.
mlab.clf()
mlab.points3d(X_train.ix[:,0]/X.ix[:,0].std(),
X_train.ix[:,1]/X.ix[:,1].std(),
y_train.ix[:,0]/y.ix[:,0].std(),
color=(1,0,0), scale_factor=0.2)
mlab.points3d(X_test.ix[:,0]/X.ix[:,0].std(),
X_test.ix[:,1]/X.ix[:,1].std(),
y_test.ix[:,0]/y.ix[:,0].std(),
color=(0,1,0), scale_factor=0.2)
mlab.axes()
mlab.show()
# Run the solver
reg = linear_model.LinearRegression(fit_intercept=True)
reg.fit(X_train,y_train)
# Create data for plotting
size=10
xPlot,yPlot = np.meshgrid(np.linspace(0,1,size),
np.linspace(0,1,size))
np.array([xPlot.flatten(),yPlot.flatten()])
zPlot = reg.predict(np.transpose(np.array([xPlot.flatten(),
yPlot.flatten()])))
zPlot = zPlot.reshape([size,size])
# +
# Since we will be plotting many times, we
def myPlot(reg,X_train,y_train,X_test,y_test,xPlot,yPlot,zPlot,size=10,scale_factor=0.05):
mlab.clf()
mlab.points3d(X_train.ix[:,0],
X_train.ix[:,1],
y_train.ix[:,0],
color=(1,0,0), scale_factor=scale_factor)
mlab.points3d(X_test.ix[:,0],
X_test.ix[:,1],
y_test.ix[:,0],
color=(0,1,0), scale_factor=scale_factor)
mlab.mesh(xPlot,yPlot,zPlot,color=(0,0,1))
mlab.axes()
mlab.show()
myPlot(reg,X_train,y_train,X_test,y_test,xPlot,yPlot,zPlot)
# -
# Use the metrics package to print our errors
print 'training error'
print metrics.mean_squared_error(y_train,reg.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,reg.predict(X_test))
# <b>Back to the notes.</b>
# ## Non-linear fitting
# +
# Now we try non-linear fittng. See notes for details.
# Note that we add a new column which is a *non-linear* function
# of the original data!
XyNonlinear = Xy.copy()
XyNonlinear['TV*Radio'] = Xy['TV']*Xy['Radio']
# Select out our predictor columns and our response columns
X = XyNonlinear.ix[:,['TV','Radio','TV*Radio']]
y = XyNonlinear.ix[:,['Sales']]
# Select subsets for training and testing
X_train,X_test,y_train,y_test = cross_validation.train_test_split(X,
y,
test_size=0.8,
random_state=123)
# -
# Run the solver
reg = linear_model.LinearRegression(fit_intercept=True)
reg.fit(X_train,y_train)
# +
# Create data for plotting
size = 10
xPlot,yPlot = np.meshgrid(np.linspace(0,1,size),
np.linspace(0,1,size))
zPlot = reg.predict(np.transpose(np.array([xPlot.flatten(),
yPlot.flatten(),
(xPlot*yPlot).flatten()])))
zPlot = zPlot.reshape([size,size])
# -
myPlot(reg,X_train,y_train,X_test,y_test,xPlot,yPlot,zPlot)
# Use the metrics package to print our errors
print 'training error'
print metrics.mean_squared_error(y_train,reg.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,reg.predict(X_test))
#
# <b>Back to the notes.</b>
# ## Too much of a good thing...
# +
# What about adding many non-linear combinations! See notes for details.
degree=5
XCrazy = np.zeros([Xy.shape[0],degree**2])
for i in range(degree):
for j in range(degree):
XCrazy[:,i*degree + j] = (Xy['TV']**i)*(Xy['Radio']**j)
# Select subsets for training and testing
X_train,X_test,y_train,y_test = cross_validation.train_test_split(XCrazy,
y,
test_size=0.8,
random_state=123)
# -
# Run the solver
regOver = linear_model.LinearRegression(fit_intercept=True)
regOver.fit(X_train,y_train)
print regOver.intercept_
print regOver.coef_
# +
# Create data for plotting
size = 10
xPlot,yPlot = np.meshgrid(np.linspace(0,1,size),
np.linspace(0,1,size))
tmp = []
for i in range(degree):
for j in range(degree):
tmp.append( ( (xPlot**i)*(yPlot**j) ).flatten() )
zPlot = regOver.predict(np.transpose(np.array(tmp)))
zPlot = zPlot.reshape([size,size])
# +
# Plot the data
# Select subsets for training and testing
X_train_plot,X_test_plot = cross_validation.train_test_split(Xy.ix[:,['TV','Radio']],
test_size=0.8,
random_state=123)
myPlot(reg,X_train_plot,y_train,X_test_plot,y_test,xPlot,yPlot,zPlot)
# -
# Use the metrics package to print our errors
print 'training error'
print metrics.mean_squared_error(y_train,regOver.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,regOver.predict(X_test))
# <b>Back to notes.</b>
# ## Model Selection
# +
# Fortunately, there is a *lot* that one can do to help. It is possible to have
# many predictors but still get good answers. See notes for details...
degree=5
XCrazy = np.zeros([Xy.shape[0],degree**2])
names = []
for i in range(degree):
for j in range(degree):
XCrazy[:,i*degree + j] = (Xy['TV']**i)*(Xy['Radio']**j)
names.append('TV**%d*Radio**%d'%(i,j))
# Select subsets for training and testing
X_train,X_test,y_train,y_test = cross_validation.train_test_split(XCrazy,
y,
test_size=0.8,
random_state=123)
# -
# We can try None and 3 to see what we get.
selector = feature_selection.RFE(regOver,n_features_to_select=3)
selector.fit(X_train,y_train)
# Print out the predictors we use. These are the predictors selection by the RFE algorithm
# as the most important.
for i in range(len(names)):
print names[i],
print selector.get_support()[i]
# +
# Create data for plotting
size = 10
xPlot,yPlot = np.meshgrid(np.linspace(0,1,size),
np.linspace(0,1,size))
tmp = []
for i in range(degree):
for j in range(degree):
tmp.append( ( (xPlot**i)*(yPlot**j) ).flatten() )
zPlot = selector.predict(np.transpose(np.array(tmp)))
zPlot = zPlot.reshape([size,size])
# +
# Plot the data
# Select subsets for training and testing
X_train_plot,X_test_plot = cross_validation.train_test_split(Xy.ix[:,['TV','Radio']],
test_size=0.8,
random_state=123)
myPlot(reg,X_train_plot,y_train,X_test_plot,y_test,xPlot,yPlot,zPlot)
# -
# Use the metrics package to print our errors
print 'training error'
print metrics.mean_squared_error(y_train,selector.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,selector.predict(X_test))
# <b>Back to notes.</b>
# ## Lasso!
# +
# Lasso regression is another method for doing feature selection.
# It is, by far, by favorite it is a close cousin of my personal
# research topic. See notes for details...
degree=5
XCrazy = np.zeros([Xy.shape[0],degree**2])
names = []
for i in range(degree):
for j in range(degree):
XCrazy[:,i*degree + j] = (Xy['TV']**i)*(Xy['Radio']**j)
names.append('TV**%d*Radio**%d'%(i,j))
# Select subsets for training and testing
X_train,X_test,y_train,y_test = cross_validation.train_test_split(XCrazy,
y,
test_size=0.8,
random_state=123)
# -
# Run the solver
regLasso = linear_model.Lasso(alpha=0.002,fit_intercept=True,normalize=True)
regLasso.fit(X_train,y_train)
# Print out the predictors we use. These betas with non-zero weights are those
# selected by the Lasso algorithm as being the most important. What do you notice?
print regLasso.intercept_
for i in range(len(regLasso.coef_)):
print names[i],regLasso.coef_[i]
# +
# Create data for plotting
size = 10
xPlot,yPlot = np.meshgrid(np.linspace(0,1,size),
np.linspace(0,1,size))
tmp = []
for i in range(degree):
for j in range(degree):
tmp.append( ( (xPlot**i)*(yPlot**j) ).flatten() )
zPlot = regLasso.predict(np.transpose(np.array(tmp)))
zPlot = zPlot.reshape([size,size])
# +
# Plot the data
# Select subsets for training and testing
X_train_plot,X_test_plot = cross_validation.train_test_split(Xy.ix[:,['TV','Radio']],
test_size=0.8,
random_state=123)
myPlot(reg,X_train_plot,y_train,X_test_plot,y_test,xPlot,yPlot,zPlot)
# -
# Use the metrics package to print our errors
print 'training error'
print metrics.mean_squared_error(y_train,regLasso.predict(X_train))
print 'testing error'
print metrics.mean_squared_error(y_test,regLasso.predict(X_test))
|
lectures/06 Machine Learning Part 1 and Midterm Review/2_ML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="qrM11qy46His"
# # TASK #1: UNDERSTAND THE PROBLEM STATEMENT AND BUSINESS CASE
# + [markdown] id="KzAol5aeBOfp"
# 
# + [markdown] id="jj6vbDkaO7gk"
# 
# + [markdown] id="_aSS2gGhPMW7"
# 
# + [markdown] id="lOnlTCNgPWei"
# 
# + [markdown] id="LfpAhVZC9YHj"
# Any publications based on this dataset should acknowledge the following:
#
# <NAME>. (2013). UCI Machine Learning Repository [http://archive.ics.uci.edu/ml]. Irvine, CA: University of California, School of Information and Computer Science.
#
# The original dataset can be found here at the UCI Machine Learning Repository.
#
#
# + [markdown] id="IJwsNqB76MBy"
# # TASK #2: IMPORT LIBRARIES AND DATASETS
# + id="ETf0YAwI6NDN"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="5oDTMhZP6QNx" outputId="c61a6d7b-8e01-47e9-f92c-c499e2f2f2d0"
# You will need to mount your drive using the following commands:
# For more information regarding mounting, please check this out: https://stackoverflow.com/questions/46986398/import-data-into-google-colaboratory
from google.colab import drive
drive.mount('/content/drive')
# + id="ZhDnHNiN6R7A"
# You have to include the full link to the csv file containing your dataset
creditcard_df = pd.read_csv('/content/drive/My Drive/Colab Notebooks/Modern AI Portfolio Builder/Business AI/UCI_Credit_Card.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="TQf9G6QF6pVV" outputId="e61c8efe-a867-44f7-99a5-06dd69aa8abd"
creditcard_df
# + colab={"base_uri": "https://localhost:8080/", "height": 554} id="UtamBsKS8fNR" outputId="21817acd-6c51-4694-8647-7eaf0e6c6976"
creditcard_df.info()
# 24 features in total, each contains 30000 data points
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="ENzZ7eo28lu2" outputId="f5e98e8b-7299-40cc-9f8c-b4213637f094"
creditcard_df.describe()
# the mean for LIMIT_BAL = 1500, min =1, and max = 30000
# the mean for AGE = 25 years old, min = 21, and max = 79
# PAY_AMT average is around 5k
# + [markdown] id="rhxz_5cAB-Jk"
# # TASK #3: VISUALIZE DATASET
# + colab={"base_uri": "https://localhost:8080/", "height": 416} id="uk1G6de2B-rs" outputId="397f761f-8087-4a26-92fa-0fabd154b3b1"
# Let's see if we have any missing data, luckily we don't!
sns.heatmap(creditcard_df.isnull(), yticklabels = False, cbar = False, cmap="Blues")
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="cmSjFMYyCC_E" outputId="2123824b-faeb-45f2-9bae-95e3c58bb839"
creditcard_df.hist(bins = 30, figsize = (20,20), color = 'r')
# + id="qhWC2_9iCLf7"
# Let's drop the ID column
creditcard_df.drop(['ID'], axis=1, inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="kEydThfXDTiD" outputId="cd0f54e6-43f1-4f92-adce-cb66901a7782"
creditcard_df
# + id="F08VjkV1DWMl"
# Let's see how many customers could potentially default on their credit card payment!
cc_default_df = creditcard_df[creditcard_df['default.payment.next.month'] == 1]
cc_nodefault_df = creditcard_df[creditcard_df['default.payment.next.month'] == 0]
# + colab={"base_uri": "https://localhost:8080/", "height": 101} id="0vPf2N3FDt8B" outputId="58c0d88b-cf56-4c53-a636-0fb373c48f29"
# Count the number of employees who stayed and left
# It seems that we are dealing with an imbalanced dataset
print("Total =", len(creditcard_df))
print("Number of customers who defaulted on their credit card payments =", len(cc_default_df))
print("Percentage of customers who defaulted on their credit card payments =", 1.*len(cc_default_df)/len(creditcard_df)*100.0, "%")
print("Number of customers who did not default on their credit card payments (paid their balance)=", len(cc_nodefault_df))
print("Percentage of customers who did not default on their credit card payments (paid their balance)=", 1.*len(cc_nodefault_df)/len(creditcard_df)*100.0, "%")
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="8nyqOjAfEMlP" outputId="2d34979c-50a0-4fe9-cddd-20125e41a106"
# Let's compare the mean and std of the customers who stayed and left
cc_default_df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 304} id="wnHJpRmfEs_O" outputId="df3b3d0e-f1b4-4d38-c638-36c0284a1695"
# Let's compare the mean and std of the customers who stayed and left
cc_nodefault_df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2N1hbr3iExE-" outputId="814646db-b183-49a0-81c3-5b9048cea564"
correlations = creditcard_df.corr()
f, ax = plt.subplots(figsize = (20, 20))
sns.heatmap(correlations, annot = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 711} id="k0Nnl5pxHoXz" outputId="60c4b841-8473-43cd-976a-a27a8fe0cdf4"
plt.figure(figsize=[25, 12])
sns.countplot(x = 'AGE', hue = 'default.payment.next.month', data = creditcard_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="8p_rit_pIEmw" outputId="3d175cf3-dfce-4a4c-d763-87beb9883803"
plt.figure(figsize=[20,20])
plt.subplot(311)
sns.countplot(x = 'EDUCATION', hue = 'default.payment.next.month', data = creditcard_df)
plt.subplot(312)
sns.countplot(x = 'SEX', hue = 'default.payment.next.month', data = creditcard_df)
plt.subplot(313)
sns.countplot(x = 'MARRIAGE', hue = 'default.payment.next.month', data = creditcard_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 470} id="cXTCL9dGTgda" outputId="d36c1fc0-ff17-4826-a102-259d9bb58659"
# KDE (Kernel Density Estimate) is used for visualizing the Probability Density of a continuous variable.
# KDE describes the probability density at different values in a continuous variable.
plt.figure(figsize=(12,7))
sns.distplot(cc_nodefault_df['LIMIT_BAL'], bins = 250, color = 'r')
sns.distplot(cc_default_df['LIMIT_BAL'], bins = 250, color = 'b')
plt.xlabel('Amount of bill statement in September, 2005 (NT dollar)')
#plt.xlim(0, 200000)
# + colab={"base_uri": "https://localhost:8080/", "height": 470} id="DDXsc1toIuHw" outputId="2e8a1356-ed95-4dd0-97a1-7c87e2dc55c4"
# KDE (Kernel Density Estimate) is used for visualizing the Probability Density of a continuous variable.
# KDE describes the probability density at different values in a continuous variable.
plt.figure(figsize=(12,7))
sns.kdeplot(cc_nodefault_df['BILL_AMT1'], label = 'Customers who did not default (paid balance)', shade = True, color = 'r')
sns.kdeplot(cc_default_df['BILL_AMT1'], label = 'Customers who defaulted (did not pay balance)', shade = True, color = 'b')
plt.xlabel('Amount of bill statement in September, 2005 (NT dollar)')
#plt.xlim(0, 200000)
# + colab={"base_uri": "https://localhost:8080/", "height": 471} id="id_S89-eK-m3" outputId="60a16c78-c0f9-4745-ae85-4921d51a4235"
# KDE (Kernel Density Estimate) is used for visualizing the Probability Density of a continuous variable.
# KDE describes the probability density at different values in a continuous variable.
plt.figure(figsize=(12,7))
sns.kdeplot(cc_nodefault_df['PAY_AMT1'], label = 'Customers who did not default (paid balance)', shade = True, color = 'r')
sns.kdeplot(cc_default_df['PAY_AMT1'], label = 'Customers who defaulted (did not pay balance)', shade = True, color = 'b')
plt.xlabel('PAY_AMT1: Amount of previous payment in September, 2005 (NT dollar)')
plt.xlim(0, 200000)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Iq4eAkJbLWgz" outputId="abf181ef-5cfb-4539-f76f-42bcd140b0ad"
# Let's see the impact of sex on the limit balance
plt.figure(figsize=[10,20])
plt.subplot(211)
sns.boxplot(x = 'SEX', y = 'LIMIT_BAL', data = creditcard_df, showfliers = False)
plt.subplot(212)
sns.boxplot(x = 'SEX', y = 'LIMIT_BAL', data = creditcard_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="MM_Vd6WAWDyS" outputId="bc7f6db1-7c79-44ac-d970-0561b06ce54f"
plt.figure(figsize=[10,20])
plt.subplot(211)
sns.boxplot(x = 'MARRIAGE', y = 'LIMIT_BAL', data = creditcard_df, showfliers = False)
plt.subplot(212)
sns.boxplot(x = 'MARRIAGE', y = 'LIMIT_BAL', data = creditcard_df)
# + [markdown] id="gs1wi5-CkPaS"
# # TASK #4: CREATE TESTING AND TRAINING DATASET & PERFORM DATA CLEANING
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="VghsrG44tH3f" outputId="2a19d79a-3427-412b-d8ce-bd92c382b11b"
creditcard_df
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="Zz5Z0H-YkVYx" outputId="d0a16e5c-ae9c-435b-bc14-08dd631a4d40"
X_cat = creditcard_df[['SEX', 'EDUCATION', 'MARRIAGE']]
X_cat
# + id="nsEXKNl_kVa4"
from sklearn.preprocessing import OneHotEncoder
onehotencoder = OneHotEncoder()
X_cat = onehotencoder.fit_transform(X_cat).toarray()
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="0o3k406tkVdL" outputId="17286c22-e8a9-4719-b612-de6442d1630f"
X_cat.shape
# + id="PGbaiijVkeso"
X_cat = pd.DataFrame(X_cat)
# + colab={"base_uri": "https://localhost:8080/", "height": 402} id="PoJMSzTkkevg" outputId="74d30e31-1578-46e7-cb5a-00ebcdc5d5b6"
X_cat
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="G5fStu_EkeyB" outputId="69f15694-eb47-4de1-acde-988f3bd454ff"
# note that we dropped the target 'default.payment.next.month'
X_numerical = creditcard_df[['LIMIT_BAL', 'AGE', 'PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6',
'BILL_AMT1','BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6',
'PAY_AMT1', 'PAY_AMT2', 'PAY_AMT3', 'PAY_AMT4', 'PAY_AMT5', 'PAY_AMT6']]
X_numerical
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="pOjFqEmCke2S" outputId="802eb6d3-c819-4a1a-fd5b-b648f954887a"
X_all = pd.concat([X_cat, X_numerical], axis = 1)
X_all
# + id="9RnDq2T0ke0o"
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X = scaler.fit_transform(X_all)
# + colab={"base_uri": "https://localhost:8080/", "height": 218} id="Sfsu_l3Xko5d" outputId="84f1ed63-0059-4e6a-896f-c41de045d1bb"
y = creditcard_df['default.payment.next.month']
y
# + [markdown] id="gEtIwsk8JtK5"
# # TASK #5: UNDERSTAND THE THEORY AND INTUITION BEHIND XGBOOST ALGORITHM
# + [markdown] id="UhqcckCwPWsg"
# 
# + [markdown] id="CHG_8stZPWz9"
# 
# + [markdown] id="m4D_1sQ2PW9J"
# 
# + [markdown] id="z6mFqaj1PW6f"
# 
# + [markdown] id="E2jRi15pPW4X"
# 
# + [markdown] id="5U92jIVhQEOK"
# 
# + [markdown] id="-Pcs_NUvQNcB"
# 
# + [markdown] id="OrkKSieRQTgz"
# 
# + [markdown] id="VwuHHxS6QaAm"
# 
# + [markdown] id="H2923IEdQiTv"
# 
# + [markdown] id="bSaiMocIQqiK"
# 
# + [markdown] id="L06ueqFBYS-J"
# # TASK #6: UNDERSTAND XGBOOST ALGORITHM KEY STEPS
# + [markdown] id="jbWwT3jWQw2c"
# 
# + [markdown] id="uCufQqU7QxEd"
# 
# + [markdown] id="6vh5lX0tQxRR"
# 
# + [markdown] id="b9JvNDonSt4S"
# 
# + [markdown] id="pzU8uMEhSuE3"
# 
# + [markdown] id="KCk6QpJ9SuCQ"
# 
# + [markdown] id="ozKbAVu6St_j"
# 
# + [markdown] id="y125yGORSt9l"
# 
# + [markdown] id="Zzolh-bkSt1M"
# 
# + [markdown] id="AI-PVfZjQxaH"
# 
# + [markdown] id="9vrsZhYEQxha"
# 
# + [markdown] id="_oabhKbdlTrJ"
# # TASK #7: TRAIN AND EVALUATE AN XGBOOST CLASSIFIER (LOCALLY)
#
# + id="1Ql5MIMMlWq3"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="KWtxnGyElZ1l" outputId="a571c65d-44c9-4709-c0ef-8e9d259955f5"
X_train.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="6aEFOxqdlZ4M" outputId="2c5980a8-4f2c-47ab-81a5-b38081dab119"
X_test.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 67} id="LeEJoWKzzGGC" outputId="297331c7-f086-4667-97ec-728196aa227f"
# !pip install xgboost
# + colab={"base_uri": "https://localhost:8080/", "height": 134} id="_vnKdYjry4pr" outputId="7c74effe-697d-4883-dcff-9b1c3a5fa4f8"
# Train an XGBoost regressor model
import xgboost as xgb
model = xgb.XGBClassifier(objective ='reg:squarederror', learning_rate = 0.1, max_depth = 5, n_estimators = 100)
model.fit(X_train, y_train)
# + id="rp7L11rclZ75"
from sklearn.metrics import accuracy_score
y_pred = model.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="oZz4g97tlZ6N" outputId="0819b9dd-9dad-404c-d4d7-100dc46650a5"
y_pred
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="a0a526t8ldm2" outputId="67fd771a-3197-4323-96a9-397dc90dc3b9"
from sklearn.metrics import confusion_matrix, classification_report
print("Accuracy {} %".format( 100 * accuracy_score(y_pred, y_test)))
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="Od4hj5_EldrJ" outputId="7cd564b3-7925-4e20-a725-04f0438f7852"
# Testing Set Performance
cm = confusion_matrix(y_pred, y_test)
sns.heatmap(cm, annot=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 168} id="bupPWKEOldtV" outputId="7a5aeef7-0648-4fa0-b1bf-2b2c0712ba44"
print(classification_report(y_test, y_pred))
# + [markdown] id="bxcE1bLGz5hZ"
# # TASK #8: OPTIMIZE XGBOOST HYPERPARAMETERS BY PERFORMING GRID SEARCH
# + id="SE7szVIY0M0d"
param_grid = {
'gamma': [0.5, 1, 5], # regularization parameter
'subsample': [0.6, 0.8, 1.0], # % of rows taken to build each tree
'colsample_bytree': [0.6, 0.8, 1.0], # number of columns used by each tree
'max_depth': [3, 4, 5] # depth of each tree
}
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="eHgd0IfW0D0U" outputId="1544a058-6774-41d9-d08d-a0ad0b103b47"
from xgboost import XGBClassifier
xgb_model = XGBClassifier(learning_rate=0.01, n_estimators=100, objective='binary:logistic')
from sklearn.model_selection import GridSearchCV
grid = GridSearchCV(xgb_model, param_grid, refit = True, verbose = 4)
grid.fit(X_train, y_train)
# + id="OdTsOhAJ0Nz_"
y_predict_optim = grid.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} id="2ukPNDEo0Sx3" outputId="47b387b7-73ed-447b-93a0-ab45a23bc590"
y_predict_optim
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="be4wECDBJAlw" outputId="75084414-e26d-4fea-aa67-4f2a139adef3"
# Testing Set Performance
cm = confusion_matrix(y_predict_optim, y_test)
sns.heatmap(cm, annot=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 168} id="QrPx8TlLJIrF" outputId="66e9369a-f881-47f1-8a19-d5fb9d7355bf"
print(classification_report(y_test, y_predict_optim))
# + [markdown] id="NxoqK3wSVIAW"
# # TASK #9: XG-BOOST ALGORITHM IN AWS SAGEMAKER
# + [markdown] id="6c50JCGnToQr"
# 
# + [markdown] id="Zm188bZUTogy"
# 
# + [markdown] id="XxJRtidpToqE"
# 
# + [markdown] id="Vi66pXGQToeR"
# 
# + [markdown] id="DI9BWVNuTobA"
# 
# + [markdown] id="ZRVviq1nVqSu"
# # TASK #10: TRAIN XG-BOOST USING SAGEMAKER
# + id="h7KAdxi6JJgG"
# Convert the array into dataframe in a way that target variable is set as the first column and followed by feature columns
# This is because sagemaker built-in algorithm expects the data in this format.
train_data = pd.DataFrame({'Target': y_train[:,0]})
for i in range(X_train.shape[1]):
train_data[i] = X_train[:,i]
# + id="mt-itpqwVxQk"
train_data.head()
# + id="h5cI9ZZGVxVD"
val_data = pd.DataFrame({'Target':y_val[:,0]})
for i in range(X_val.shape[1]):
val_data[i] = X_val[:,i]
# + id="a2-Hc-FsVxgN"
val_data.head()
# + id="41_J7DykVxk9"
val_data.shape
# + id="ifZToPLsVxqG"
# save train_data and validation_data as csv files.
train_data.to_csv('train.csv', header = False, index = False)
val_data.to_csv('validation.csv', header = False, index = False)
# + id="5N_vpfK_VxvF"
# Boto3 is the Amazon Web Services (AWS) Software Development Kit (SDK) for Python
# Boto3 allows Python developer to write software that makes use of services like Amazon S3 and Amazon EC2
import sagemaker
import boto3
# Create a sagemaker session
sagemaker_session = sagemaker.Session()
#S 3 bucket and prefix that we want to use
# default_bucket - creates a Amazon S3 bucket to be used in this session
bucket = 'sagemaker-practical-3'
prefix = 'XGBoost-Regressor'
key = 'XGBoost-Regressor'
#Roles give learning and hosting access to the data
#This is specified while opening the sagemakers instance in "Create an IAM role"
role = sagemaker.get_execution_role()
# + id="DbdoRZ1GVxtD"
print(role)
# + id="qfrO5GPaVxok"
# read the data from csv file and then upload the data to s3 bucket
import os
with open('train.csv','rb') as f:
# The following code uploads the data into S3 bucket to be accessed later for training
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', key)).upload_fileobj(f)
# Let's print out the training data location in s3
s3_train_data = 's3://{}/{}/train/{}'.format(bucket, prefix, key)
print('uploaded training data location: {}'.format(s3_train_data))
# + id="QAm6uKHzVxjf"
# read the data from csv file and then upload the data to s3 bucket
with open('validation.csv','rb') as f:
# The following code uploads the data into S3 bucket to be accessed later for training
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation', key)).upload_fileobj(f)
# Let's print out the validation data location in s3
s3_validation_data = 's3://{}/{}/validation/{}'.format(bucket, prefix, key)
print('uploaded validation data location: {}'.format(s3_validation_data))
# + id="zQn_ylRsVxeC"
# creates output placeholder in S3 bucket to store the output
output_location = 's3://{}/{}/output'.format(bucket, prefix)
print('training artifacts will be uploaded to: {}'.format(output_location))
# + id="_FniO4vjVxcA"
# This code is used to get the training container of sagemaker built-in algorithms
# all we have to do is to specify the name of the algorithm, that we want to use
# Let's obtain a reference to the XGBoost container image
# Note that all regression models are named estimators
# You don't have to specify (hardcode) the region, get_image_uri will get the current region name using boto3.Session
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'xgboost','0.90-2') # Latest version of XGboost
# + id="givvDwooVxat"
# Specify the type of instance that we would like to use for training
# output path and sagemaker session into the Estimator.
# We can also specify how many instances we would like to use for training
# Recall that XGBoost works by combining an ensemble of weak models to generate accurate/robust results.
# The weak models are randomized to avoid overfitting
# num_round: The number of rounds to run the training.
# Alpha: L1 regularization term on weights. Increasing this value makes models more conservative.
# colsample_by_tree: fraction of features that will be used to train each tree.
# eta: Step size shrinkage used in updates to prevent overfitting.
# After each boosting step, eta parameter shrinks the feature weights to make the boosting process more conservative.
Xgboost_regressor1 = sagemaker.estimator.Estimator(container,
role,
train_instance_count = 1,
train_instance_type = 'ml.m5.2xlarge',
output_path = output_location,
sagemaker_session = sagemaker_session)
#We can tune the hyper-parameters to improve the performance of the model
Xgboost_regressor1.set_hyperparameters(max_depth = 10,
objective = 'reg:linear',
colsample_bytree = 0.3,
alpha = 10,
eta = 0.1,
num_round = 100
)
# + id="ExKC0xVpVxXf"
# Creating "train", "validation" channels to feed in the model
# Source: https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html
train_input = sagemaker.session.s3_input(s3_data = s3_train_data, content_type='csv',s3_data_type = 'S3Prefix')
valid_input = sagemaker.session.s3_input(s3_data = s3_validation_data, content_type='csv',s3_data_type = 'S3Prefix')
data_channels = {'train': train_input,'validation': valid_input}
Xgboost_regressor1.fit(data_channels)
# + [markdown] id="w0Kw6K67WJAm"
# # TASK #11: DEPLOY MODEL TO PERFORM INFERENCE
# + id="nwEvEyVvVxTj"
# Deploy the model to perform inference
Xgboost_regressor = Xgboost_regressor1.deploy(initial_instance_count = 1, instance_type = 'ml.m5.2xlarge')
# + id="C6aMpTPwWUZE"
'''
Content type over-rides the data that will be passed to the deployed model, since the deployed model expects data
in text/csv format, we specify this as content -type.
Serializer accepts a single argument, the input data, and returns a sequence of bytes in the specified content
type
Reference: https://sagemaker.readthedocs.io/en/stable/predictors.html
'''
from sagemaker.predictor import csv_serializer, json_deserializer
Xgboost_regressor.content_type = 'text/csv'
Xgboost_regressor.serializer = csv_serializer
Xgboost_regressor.deserializer = None
# + id="wqWx9AuIWUdh"
X_test.shape
# + id="Wg-w_2jGWUgU"
# making prediction
predictions1 = Xgboost_regressor.predict(X_test[0:10000])
# + id="iXM7-8eVWUol"
predictions2 = Xgboost_regressor.predict(X_test[10000:20000])
# + id="04TwH3H7WcuS"
predictions3 = Xgboost_regressor.predict(X_test[20000:30000])
# + id="skW2KXkSWc09"
predictions4 = Xgboost_regressor.predict(X_test[30000:31618])
# + id="jN0Th7fdWc5X"
# custom code to convert the values in bytes format to array
def bytes_2_array(x):
# makes entire prediction as string and splits based on ','
l = str(x).split(',')
# Since the first element contains unwanted characters like (b,',') we remove them
l[0] = l[0][2:]
#same-thing as above remove the unwanted last character (')
l[-1] = l[-1][:-1]
# iterating through the list of strings and converting them into float type
for i in range(len(l)):
l[i] = float(l[i])
# converting the list into array
l = np.array(l).astype('float32')
# reshape one-dimensional array to two-dimensional array
return l.reshape(-1,1)
# + id="-IO-hnu_Wc34"
predicted_values_1 = bytes_2_array(predictions1)
# + id="o5l6F7JLWcy4"
predicted_values_1.shape
# + id="vRPi4YAiWcxN"
predicted_values_2 = bytes_2_array(predictions2)
predicted_values_2.shape
# + id="00a3zWcrWcsE"
predicted_values_3 = bytes_2_array(predictions3)
predicted_values_3.shape
# + id="sSaOxKWZWUb_"
predicted_values_4 = bytes_2_array(predictions4)
predicted_values_4.shape
# + id="ls7kOoCqWnKN"
predicted_values = np.concatenate((predicted_values_1, predicted_values_2, predicted_values_3, predicted_values_4))
# + id="DZ61FPBAWnQ3"
predicted_values.shape
# + id="7FGdRyoHWnf5"
from sklearn.metrics import r2_score, mean_squared_error, mean_absolute_error
from math import sqrt
k = X_test.shape[1]
n = len(X_test)
RMSE = float(format(np.sqrt(mean_squared_error(y_test, predicted_values)),'.3f'))
MSE = mean_squared_error(y_test, predicted_values)
MAE = mean_absolute_error(y_test, predicted_values)
r2 = r2_score(y_test, predicted_values)
adj_r2 = 1-(1-r2)*(n-1)/(n-k-1)
print('RMSE =',RMSE, '\nMSE =',MSE, '\nMAE =',MAE, '\nR2 =', r2, '\nAdjusted R2 =', adj_r2)
# + id="yhGR_kDpWni4"
# Delete the end-point
Xgboost_regressor.delete_endpoint()
# + [markdown] id="fqRKoCr7Yylg"
# # EXCELLENT JOB! YOU SHOULD BE PROUD OF YOUR NEWLY ACQUIRED SKILLS
|
AI_in_Business_&_AutoML.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Applicazione del transfer learning con MobileNet_V2
# A high-quality, dataset of images containing fruits. The following fruits are included: Apples - (different varieties: Golden, Golden-Red, Granny Smith, Red, Red Delicious), Apricot, Avocado, Avocado ripe, Banana (Yellow, Red), Cactus fruit, Carambula, Cherry, Clementine, Cocos, Dates, Granadilla, Grape (Pink, White, White2), Grapefruit (Pink, White), Guava, Huckleberry, Kiwi, Kaki, Kumsquats, Lemon (normal, Meyer), Lime, Litchi, Mandarine, Mango, Maracuja, Nectarine, Orange, Papaya, Passion fruit, Peach, Pepino, Pear (different varieties, Abate, Monster, Williams), Pineapple, Pitahaya Red, Plum, Pomegranate, Quince, Raspberry, Salak, Strawberry, Tamarillo, Tangelo.
#
# Training set size: 28736 images.
#
# Validation set size: 9673 images.
#
# Number of classes: 60 (fruits).
#
# Image size: 100x100 pixels.
import numpy as np
from tensorflow.python.keras.utils import to_categorical
from tensorflow.python.keras.preprocessing.image import ImageDataGenerator
from tensorflow.python.keras.preprocessing import image
from skimage import transform
import matplotlib.pyplot as plt
# %matplotlib inline
from tensorflow.python.keras.applications.mobilenet_v2 import MobileNetV2
from tensorflow.python.keras.models import Model
from tensorflow.python.keras.layers import Dense, GlobalAveragePooling2D
from tensorflow.python.keras import backend as K
# ### Change the path of directories!!!
# Setting path location for validation, traing and testing images
validationPath = 'E:/Validation'
trainPath = 'E:/Training'
# ### Plot an image, for example E:/Training/Cocos/15_100.jpg
# ### Now you define the functions ables to read mini-batch of data
# Making an image data generator object with augmentation for training
train_datagen = ImageDataGenerator(rescale=1./255,
rotation_range=30,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
fill_mode='nearest')
# Making an image data generator object with no augmentation for validation
test_datagen = ImageDataGenerator(rescale=1./255)
# ### why train_datagen and test_datagen are different? answer . . .
#
# ...........................................
#
# ...........................................
# Using the generator with batch size 32 for training directory
train_generator = train_datagen.flow_from_directory(trainPath,
target_size=(224, 224),
batch_size=32,
class_mode='categorical')
# Using the generator with batch size 17 for validation directory
validation_generator = test_datagen.flow_from_directory(validationPath,
target_size=(224, 224),
batch_size=17,
class_mode='categorical')
# ### you can control the dimensions of the generator outputs
validation_generator[0][0].shape
validation_generator[0][1].shape
# ### Now you need to define your model . . .
# the default definition of MobileNet_V2 is:
#
# MobileNetV2(input_shape=None, alpha=1.0, depth_multiplier=1, include_top=True, weights='imagenet', input_tensor=None, pooling=None, classes=1000)
#
# but you have a different number of classes . . . . .
# ### Define what layers you want to train . . .
# ### Compile the model . . .
# ## to fit the model you can write an expression as:
# history = model.fit_generator(train_generator,
# epochs=20,validation_data=validation_generator,)
# ### Fine tuning?
# ### once you have obtained the final estimate of the model you must evaluate it with more details . . .
# ### take an image of a papaya from internet and try to apply your model . . .
|
Eserc_Frutta.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''cv-base'': conda)'
# name: python3
# ---
# + tags=["outputPrepend"]
import pandas as pd
def Room(house):
i = house.find('室')
j = house.find('厅')
room = house[0:i]
if room != '--':
return int(room)
else:
return -1
def LivingRoom(house):
i = house.find('室')
j = house.find('厅')
living = house[i + 1:j]
if living != '--':
return int(living)
else:
return -1
def square(mianji):
s = mianji[0:mianji.find('平')]
if s == '0':
return -1
elif s != '--':
return int(s)
else:
return -1
def CJdanjia(cjdanjia):
s = cjdanjia[0:cjdanjia.find('元')]
if int(s) <= 9999:
return -1
else:
return int(s)
def Comp(line, xiaoqu, mendian,attribute):
room = line['room']
living = line['living']
mianji = line['mianji']
danjia = line['danjia']
xiaoqu_name = line['xiaoqu']
mendian_name = line['mendian']
xiaoqu_set = xiaoqu.get_group(xiaoqu_name).reset_index(drop=True)
mendian_set = mendian.get_group(mendian_name).reset_index(drop=True)
xiaoqu_mean = round(xiaoqu_set[attribute].mean())
mendian_mean = round(mendian_set[attribute].mean())
xiaoqu_set = xiaoqu_set.drop(xiaoqu_set[xiaoqu_set[attribute] == -1].index).reset_index(drop=True)
mendian_set = mendian_set.drop(mendian_set[mendian_set[attribute] == -1].index).reset_index(drop=True)
distance_xiaoqu = abs((xiaoqu_set['room'] - room) * 20) + abs((xiaoqu_set['living'] - living) * 20) + \
abs(xiaoqu_set['mianji'] - mianji) + abs(xiaoqu_set['danjia'] - danjia)
distance_mendian = abs((mendian_set['room'] - room) * 20) + abs((mendian_set['living'] - living) * 20) + \
abs(mendian_set['mianji'] - mianji) + abs(mendian_set['danjia'] - danjia)
if distance_xiaoqu.empty and distance_mendian.empty:
if xiaoqu_set.__len__() < 2:
return mendian_mean
else:
return xiaoqu_mean
elif ~distance_xiaoqu.empty and ~distance_mendian.empty:
if distance_xiaoqu.min()<distance_mendian.min():
index = distance_xiaoqu.idxmin()
return xiaoqu_set.iloc[index][attribute]
else:
index = distance_mendian.idxmin()
return mendian_set.iloc[index][attribute]
elif ~distance_xiaoqu.empty and distance_mendian.empty:
index = distance_mendian.idxmin()
return mendian_set.iloc[index][attribute]
else:
index = distance_xiaoqu.idxmin()
return xiaoqu_set.iloc[index][attribute]
data = pd.read_csv('data/lianjia1.csv',encoding='gbk')
data['xiaoqu'],data['huxing'],data['mianji']=data['cjxiaoqu'].str.split(' ').str
data['room'] = data['huxing'].map(Room).fillna(-1)
data['living'] = data['huxing'].map(LivingRoom).fillna(-1)
data['mianji'] = data['mianji'].map(square).fillna(-1)
data['danjia'] = data['cjdanjia'].map(CJdanjia).fillna(-1)
xiaoqu_mean = data.groupby('xiaoqu')['mianji'].mean().astype(int)
mendian_mean = data.groupby('mendian')['mianji'].mean().astype(int)
mendian = data.groupby('mendian')
xiaoqu = data.groupby('xiaoqu')
for i in range(0,data.__len__()):
#i = 243
line = data.iloc[i]
if line['room'] == -1:
line['room'] = Comp(line,xiaoqu,mendian,'room')
if line['living'] == -1:
line['living'] = Comp(line, xiaoqu, mendian,'living')
if line['mianji'] <= 0:
line['mianji'] = Comp(line, xiaoqu, mendian,'mianji')
if line['danjia'] <= 0:
line['danjia'] = Comp(line, xiaoqu, mendian,'danjia')
line['cjzongjia'] = line['mianji'] * line['danjia'] / 10000
data.iloc[i] = line
data['room'] = data['room'].astype(str)
data['living'] = data['living'].astype(str)
data['mianji'] = data['mianji'].astype(str)
data['danjia'] = data['danjia'].astype(str)
data['danjia'] = data['danjia']+'元/平'
data['new'] = data['xiaoqu']+' '+data['room']+'室'+data['living']+'厅'+' '+data['mianji']+'平'
data.drop(['huxing','room','mianji','cjdanjia','living','cjxiaoqu'], axis=1)
data.to_csv('./generated_data/lianjia.csv',encoding='gbk',columns=['cjtaoshu','mendian','cjzongjia','zhiwei','haoping','danjia',
'new','xingming','cjzhouqi','biaoqian','cjlouceng','cjshijian',
'congyenianxian','bankuai'])
# -
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
import pandas as pd
import tqdm
import numpy as np
from sklearn.linear_model import LinearRegression
import scipy.stats as stats
import warnings
warnings.filterwarnings('ignore')
plt.rcParams['font.sans-serif'] = ['SimHei']
plt.rcParams['axes.unicode_minus'] = False
print('加载数据集')
data = pd.read_csv('lianjia.csv', index_col=0,encoding='gbk')
print('属性类别数:', len(data.columns))
print('总行数:', len(data))
print('示例数据:')
data.head(5)
print('提取每一列属性以及名称')
num_fields = data.select_dtypes(include=np.number).columns.values
nom_fields = data.select_dtypes(exclude=np.number).columns.values
print('标称属性:', nom_fields)
print('数值属性:', num_fields)
print(data.shape," ",nom_fields.shape," ",num_fields.shape)
# +
print("统计每一个属性的个数")
print('\nmendian(门店):')
a=data['mendian'].value_counts()
print(a)
print('\nzhiwei(职位):')
b=data['zhiwei'].value_counts()
print(b)
print('\nhaoping(好评):')
c=data['haoping'].value_counts()
print(c)
print('\ndanjia(成交单价):')
d=data['danjia'].value_counts()
print(d)
print('\nnew(成交单价):')
d1=data['new'].value_counts()
print(d1)
print('\nxingming(姓名):')
e=data['xingming'].value_counts()
print(e)
print('\nbiaoqian(标签):')
f=data['biaoqian'].value_counts()
print(f)
print('\ncjlouceng(成交楼层):')
g=data['cjlouceng'].value_counts()
print(g)
print('\ncjshijian(成交时间):')
h=data['cjshijian'].value_counts()
print(h)
print('\ncongyenianxian(从月年限):')
i=data['congyenianxian'].value_counts()
print(i)
print('\nbankuai(板块):')
j=data['bankuai'].value_counts()
print(j)
# -
data.describe()
# +
print('数据可视化:\n')
field = ['cjzongjia','cjzhouqi','danjia','cjtaoshu']
print('数据太多导致显示数据出现重叠看不清具体值,所以取前20个为例:')
for fie in field:
print("{}前20个样例:".format(fie))
data[fie].value_counts(sort=False).head(20).plot.barh()
plt.show()
# +
print("画出不同店家的成交数量饼图")
label=[]
for key in data['mendian'].value_counts().index:
label.append(key)
data['mendian'].value_counts().plot.pie(labels=label,
autopct='%.2f', fontsize=10,figsize=(12, 12))
# +
print("画出不同从月年限的饼图")
label=[]
for key in data['congyenianxian'].value_counts().index:
label.append(key)
data['congyenianxian'].value_counts().plot.pie(labels=label,
autopct='%.2f', fontsize=15,figsize=(12, 12))
# +
print("画出不同板块(地区)的饼图")
label=[]
for key in data['bankuai'].value_counts().index:
label.append(key)
data['bankuai'].value_counts().plot.pie(labels=label,
autopct='%.2f', fontsize=15,figsize=(12, 12))
# +
print("画出不同标签的饼图")
label=[]
for key in data['biaoqian'].value_counts().index:
label.append(key)
data['biaoqian'].value_counts().plot.pie(labels=label,
autopct='%.2f', fontsize=15,figsize=(12, 12))
# -
|
utils/visualization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using pyapetnet to predict anatomy-guided MAP PET reconstructions in image space
#
# In this notebook, we wil learn how to use pre-trained models included in the pyapetnet package to predict anatomy-guided MAP PET reconstructions from (simulated) PET OSEM and T1 MR images.
#
# In this tutorial, we will have a closer look at:
# - loading pre-trained models
# - loading nifti data
# - pre-processing nifti data
# - feeding the pre-processed data into the pre-trained model
# - saving visualizing the results
#
# **If you install pyapetnet from pypi using ```pip install pyapetnet```**, it will create a command line tool that does all those steps in one go. Moreover, it allows allows to load and write dicom data.
#
# For more details on pyapetnet is available here:
# - https://doi.org/10.1016/j.neuroimage.2020.117399 (NeuroImage publication on pyapetnet)
# - https://github.com/gschramm/pyapetnet/ (github repository of pyapetnet)
# + [markdown] id="bwNWY7nD3x4T"
# ## (1) Preparation: Install the pyapetnet package
#
# Before running this notebook, make sure that the pyapetnet package is installed.
# This can by done via <br>
# ```pip install pyapetnet``` <br>
# which will install the package and all its dependencies (e.g. tensorflow). We recommend to use a separate virtual environment.
#
# + [markdown] id="iNVA9mz9cCet"
# ## (2) Data used in this demo
#
# In this tutorial, we wil use simulated PET and MR data that are based on the brainweb phantom.
# The nifti files used in this tutorial, are available at <br>
# https://github.com/gschramm/pyapetnet/tree/master/demo_data <br>
# By changing ```pet_fname``` or ```mr_fname``` other input data sets can be used.
#
# -
# ## (3) Loading modules
# In the next cell, we will load all required python modules. E.g. tensorflow, to load the pre-trained model and pyapetnet for data preprocessing
# +
import nibabel as nib
import json
import os
import tensorflow as tf
import numpy as np
import os
import matplotlib.pyplot as plt
import pyapetnet
from pyapetnet.preprocessing import preprocess_volumes
from pyapetnet.utils import load_nii_in_ras
# -
# ## (4) Specification of input pameters
# In the next cell, we specify the required input parameters:
# - ```model_name``` (name of the pre-trained model shipped with the pyapernet package)
# - ```pet_fname / mr_fname``` (absolute path of the PET and MR input nifti files)
# - ```coreg_inputs``` whether to apply rigid coregistration between PET and MR volumes using mutual information
# - ```crop_mr``` whether to crop both volumes to the bounding box of the MR (usefule to limit memory usage)
# - ```output_fname``` absolute path of the nifti file for the output
#
# + id="10BENAxbbgkH"
# inputs (adapt to your needs)
# the name of the trained CNN
model_name = '200824_mae_osem_psf_bet_10'
# we use a simulated demo data included in pyapetnet (based on the brainweb phantom)
mydata_dir = '.'
pet_fname = os.path.join(mydata_dir, 'brainweb_06_osem.nii')
mr_fname = os.path.join(mydata_dir, 'brainweb_06_t1.nii')
# preprocessing parameters
coreg_inputs = True # rigidly coregister PET and MR using mutual information
crop_mr = True # crop the input to the support of the MR (saves memory + speeds up the computation)
# the name of the ouput file
output_fname = f'prediction_{model_name}.nii'
# + [markdown] id="k5jI9bMRNNEV"
# ## (5) Load the pre-trained CNN (model)
# Now we can load the pretrained model. pyapetnet includes a few preprained models that are installed all installed
# at <br>
# ```os.path.join(os.path.dirname(pyapetnet.__file__),'trained_models')```<br>
# where ```pyapetnet.__file__``` points to the install path of pyapetnet.
#
# A more detailed description of all models can be found at <br>
# https://github.com/gschramm/pyapetnet/blob/master/pyapetnet/trained_models/model_description.md
#
# The dummy dictionary ```custom_objects``` is needed since the model definition depends on 2 custom loss functions (related to SSIM). For inference the loss fucntions are not needed with is why we pass a dummy dictionary.
#
# Last but not least, we read the internal voxel size used to train the model. This is necessary to correctly pre-process the input data (which comes usually in a different voxel size).
# + id="0nGzaQw1Eju_"
# load the trained CNN and its internal voxel size used for training
model_abs_path = os.path.join(os.path.dirname(pyapetnet.__file__),'trained_models',model_name)
model = tf.keras.models.load_model(model_abs_path, custom_objects = {'ssim_3d_loss': None,'mix_ssim_3d_mae_loss': None})
# load the voxel size used for training
with open(os.path.join(model_abs_path,'config.json')) as f:
cfg = json.load(f)
training_voxsize = cfg['internal_voxsize']*np.ones(3)
# + [markdown] id="NTi_LWFmNF8J"
# ## (6) Load and preprocess the input PET and MR volumes
#
# Finally, ee load the data from the input nifti files. The preprocessing function rigidly coregisters the inputs,
# interpolates the volumes to the internal voxel size of the CNN, crops the volumes to the MR support, and does an intensity normalization (division by 99.9% percentile). We use the 99.99% percentile since it is more robust for noisy (PET) volumes.
#
# **The voxelsize of the input volumes is deduced from the affine transforamtion stored in the nifti header. Make sure that the affine stored there is correct.**
# + colab={"base_uri": "https://localhost:8080/", "height": 435} id="SumoESLqL6iL" outputId="1c24b4ac-1879-4ea8-db38-d7f61d9e10d4"
# load and preprocess the input PET and MR volumes
pet, pet_affine = load_nii_in_ras(pet_fname)
mr, mr_affine = load_nii_in_ras(mr_fname)
# preprocess the input volumes (coregistration, interpolation and intensity normalization)
pet_preproc, mr_preproc, o_aff, pet_max, mr_max = preprocess_volumes(pet, mr,
pet_affine, mr_affine, training_voxsize, perc = 99.99, coreg = coreg_inputs, crop_mr = crop_mr)
# -
# ## (7) Show and check pre-processed Input data
#
# Before passing the PET and MR input volumes to the loaded CNN, it is a good idea to check whether both volumes were correctly pre-processed. If the pre-processing was successfull, the volumes should be well aligned, should be interpolated to the internal voxelsize of the CNN, and their 99.99% percentile should be 1.
# +
print(f'PET 99.99% percentile {np.percentile(pet_preproc,99.99):.3f}')
print(f'PET 99.99% percentile {np.percentile(mr_preproc,99.99):.3f}')
fig, ax = plt.subplots(2,3, figsize = (9,6))
ax[0,0].imshow(pet_preproc[:,::-1,pet_preproc.shape[2]//2].T, cmap = plt.cm.Greys, vmax = 1)
ax[0,1].imshow(pet_preproc[:,pet_preproc.shape[1]//2,::-1].T, cmap = plt.cm.Greys, vmax = 1)
ax[0,2].imshow(pet_preproc[pet_preproc.shape[0]//2,:,::-1].T, cmap = plt.cm.Greys, vmax = 1)
ax[1,0].imshow(mr_preproc[:,::-1,pet_preproc.shape[2]//2].T, cmap = plt.cm.Greys_r, vmax = 1)
ax[1,1].imshow(mr_preproc[:,pet_preproc.shape[1]//2,::-1].T, cmap = plt.cm.Greys_r, vmax = 1)
ax[1,2].imshow(mr_preproc[pet_preproc.shape[2]//2,:,::-1].T, cmap = plt.cm.Greys_r, vmax = 1)
for axx in ax.flatten(): axx.set_axis_off()
ax[0,1].set_title('preprocessed input PET')
ax[1,1].set_title('preprocessed input MR')
fig.tight_layout()
# + [markdown] id="3imUqw2qPHE-"
# ## (8) Running the actual CNN prediction
#
# Once the data is read and preprocesed we can run the actual prediction.
# The input to the pyapetnet models is a python list containing two "tensors" (the preprocessed PET and MR volumes). The dimensions of both tensors are (1,n0,n1,n2,1) where n0,n1,n2 are the spatial dimensions of the pre-processed volumes. The left most dimension is the batch size (1 in our case) and the right most dimension is the number of input channels / features (1 in our case).
#
# We decided to input two (1,n0,n1,n2,1) tensors instead of one (1,n0,n1,n2,2) tensor since in the first layer, since in the first layers we decided to learn separte PET and MR features.
#
# Based on the design of the model, there is no restiction on the spatial input shape (n0,n1,n2) provided that enough GPU/CPU memory is available.
#
# Using a recent Nvidia GPU, this step should take roughly 1s.
# + id="xvloGXYiPKRu"
# the actual CNN prediction
x = [np.expand_dims(np.expand_dims(pet_preproc,0),-1), np.expand_dims(np.expand_dims(mr_preproc,0),-1)]
pred = model.predict(x).squeeze()
# + [markdown] id="P2XH9MdOatTH"
# ## (7) Undo the intensity normalization
#
# We undo the intensity normalization that was applied during pre-processing.
# + id="0PFKRt_4awLu"
pred *= pet_max
# + [markdown] id="ClQSF-bFbmyO"
# ## (8) Save the volumes
#
# We save the pre-processed volumes and the prediction to nifti files.
# + id="95s5HaqGbqd8"
nib.save(nib.Nifti1Image(pet_preproc, o_aff), 'pet_preproc.nii')
nib.save(nib.Nifti1Image(mr_preproc, o_aff), 'mr_preproc.nii')
nib.save(nib.Nifti1Image(pred, o_aff), f'prediction_{model_name}.nii')
# + [markdown] id="06mi6lgROvxj"
# ## (9) Display the input and the prediction
#
# Finally we display the results.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 658} id="IElO4PdzMKS6" outputId="c56a5636-b00e-4de8-ea86-90dd1e3190d8"
fig, ax = plt.subplots(3,3, figsize = (9,9))
ax[0,0].imshow(pet_preproc[:,::-1,pet_preproc.shape[2]//2].T, cmap = plt.cm.Greys, vmax = 1)
ax[0,1].imshow(pet_preproc[:,pet_preproc.shape[1]//2,::-1].T, cmap = plt.cm.Greys, vmax = 1)
ax[0,2].imshow(pet_preproc[pet_preproc.shape[0]//2,:,::-1].T, cmap = plt.cm.Greys, vmax = 1)
ax[1,0].imshow(mr_preproc[:,::-1,pet_preproc.shape[2]//2].T, cmap = plt.cm.Greys_r, vmax = 1)
ax[1,1].imshow(mr_preproc[:,pet_preproc.shape[1]//2,::-1].T, cmap = plt.cm.Greys_r, vmax = 1)
ax[1,2].imshow(mr_preproc[pet_preproc.shape[2]//2,:,::-1].T, cmap = plt.cm.Greys_r, vmax = 1)
ax[2,0].imshow(pred[:,::-1,pet_preproc.shape[2]//2].T, cmap = plt.cm.Greys, vmax = pet_max)
ax[2,1].imshow(pred[:,pet_preproc.shape[1]//2,::-1].T, cmap = plt.cm.Greys, vmax = pet_max)
ax[2,2].imshow(pred[pet_preproc.shape[0]//2,:,::-1].T, cmap = plt.cm.Greys, vmax = pet_max)
for axx in ax.flatten(): axx.set_axis_off()
ax[0,1].set_title('pre-processed input PET')
ax[1,1].set_title('pre-processed input MR')
ax[2,1].set_title('predicted MAP Bowsher')
fig.tight_layout()
|
notebooks/04_inference_demo_brainweb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collaboration and Competition
#
# ---
#
# In this notebook, you will learn how to use the Unity ML-Agents environment for the third project of the [Deep Reinforcement Learning Nanodegree](https://www.udacity.com/course/deep-reinforcement-learning-nanodegree--nd893) program.
#
# ### 1. Start the Environment
#
# We begin by importing the necessary packages. If the code cell below returns an error, please revisit the project instructions to double-check that you have installed [Unity ML-Agents](https://github.com/Unity-Technologies/ml-agents/blob/master/docs/Installation.md) and [NumPy](http://www.numpy.org/).
# +
from unityagents import UnityEnvironment
import numpy as np
import random
from collections import namedtuple, deque
import copy
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Next, we will start the environment! **_Before running the code cell below_**, change the `file_name` parameter to match the location of the Unity environment that you downloaded.
#
# - **Mac**: `"path/to/Tennis.app"`
# - **Windows** (x86): `"path/to/Tennis_Windows_x86/Tennis.exe"`
# - **Windows** (x86_64): `"path/to/Tennis_Windows_x86_64/Tennis.exe"`
# - **Linux** (x86): `"path/to/Tennis_Linux/Tennis.x86"`
# - **Linux** (x86_64): `"path/to/Tennis_Linux/Tennis.x86_64"`
# - **Linux** (x86, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86"`
# - **Linux** (x86_64, headless): `"path/to/Tennis_Linux_NoVis/Tennis.x86_64"`
#
# For instance, if you are using a Mac, then you downloaded `Tennis.app`. If this file is in the same folder as the notebook, then the line below should appear as follows:
# ```
# env = UnityEnvironment(file_name="Tennis.app")
# ```
env = UnityEnvironment(file_name="/home/nuno/workspaces/drlnd/deep-reinforcement-learning/p3_collab-compet/Tennis_Linux/Tennis.x86_64")
# Environments contain **_brains_** which are responsible for deciding the actions of their associated agents. Here we check for the first brain available, and set it as the default brain we will be controlling from Python.
# get the default brain
brain_name = env.brain_names[0]
brain = env.brains[brain_name]
# ### 2. Examine the State and Action Spaces
#
# In this environment, two agents control rackets to bounce a ball over a net. If an agent hits the ball over the net, it receives a reward of +0.1. If an agent lets a ball hit the ground or hits the ball out of bounds, it receives a reward of -0.01. Thus, the goal of each agent is to keep the ball in play.
#
# The observation space consists of 8 variables corresponding to the position and velocity of the ball and racket. Two continuous actions are available, corresponding to movement toward (or away from) the net, and jumping.
#
# Run the code cell below to print some information about the environment.
# +
# reset the environment
env_info = env.reset(train_mode=True)[brain_name]
# number of agents
num_agents = len(env_info.agents)
print('Number of agents:', num_agents)
# size of each action
action_size = brain.vector_action_space_size
print('Size of each action:', action_size)
# examine the state space
states = env_info.vector_observations
state_size = states.shape[1]
print('There are {} agents. Each observes a state with length: {}'.format(states.shape[0], state_size))
print('The state for the first agent looks like:', states[0])
# + active=""
# for i in range(1, 6): # play game for 5 episodes
# env_info = env.reset(train_mode=False)[brain_name] # reset the environment
# states = env_info.vector_observations # get the current state (for each agent)
# scores = np.zeros(num_agents) # initialize the score (for each agent)
# while True:
# actions = np.random.randn(num_agents, action_size) # select an action (for each agent)
# actions = np.clip(actions, -1, 1) # all actions between -1 and 1
# env_info = env.step(actions)[brain_name] # send all actions to tne environment
# next_states = env_info.vector_observations # get next state (for each agent)
# rewards = env_info.rewards # get reward (for each agent)
# dones = env_info.local_done # see if episode finished
# scores += env_info.rewards # update the score (for each agent)
# states = next_states # roll over states to next time step
# if np.any(dones): # exit loop if episode finished
# break
# print('Score (max over agents) from episode {}: {}'.format(i, np.max(scores)))
# -
# ### 3. Environment and helper methods
# Some utility functions to be used on the other objects and modules.
# +
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
def soft_update(local_model, target_model, tau):
"""Soft update model parameters.
θ_target = τ*θ_local + (1 - τ)*θ_target
Params
======
local_model: PyTorch model (weights will be copied from)
target_model: PyTorch model (weights will be copied to)
tau (float): interpolation parameter
"""
for target_param, local_param in zip(target_model.parameters(), local_model.parameters()):
target_param.data.copy_(tau*local_param.data + (1.0-tau)*target_param.data)
pass
def hard_update(target, source):
"""Hard update model parameters.
"""
for target_param, source_param in zip(target.parameters(), source.parameters()):
target_param.data.copy_(source_param.data)
pass
def hidden_init(layer):
fan_in = layer.weight.data.size()[0]
lim = 1. / np.sqrt(fan_in)
return (-lim, lim)
def seeding(seed=10):
random.seed(seed)
np.random.seed(seed)
torch.manual_seed(seed)
# -
# ### 4. DDPG Agent
# We are going to implement the MADDPG algorithm. In the following classes we define a simple Agent that implements the DDPG algorithm, except that it uses the the status and actions of all the agents in this multi agent environment.
#
# This code is derived from the ddpg agent for the scnenario of ddpg-bipedal code provided in the [github repository](https://github.com/udacity/deep-reinforcement-learning/tree/master/ddpg-bipedal) using also as a reference the code from the [MADDPG paper](https://github.com/openai/maddpg), and also code from other Udacity students ([gtg162y](https://github.com/gtg162y/DRLND/blob/master/P3_Collab_Compete/Tennis_Udacity_Workspace.ipynb) and [abhismatrix1](https://github.com/abhismatrix1/Tennis-MultiAgent)).
#
# In this implementation instead of using the OUNoise class, it just adds some random normal noise as exploratory noise. The amount of noise is decreased on the parent agent the MADDPG.
# +
class Actor(nn.Module):
"""Actor (Policy) Model."""
def __init__(self, state_size, action_size, fc1_units=512, fc2_units=256):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fc1_units (int): Number of nodes in first hidden layer
fc2_units (int): Number of nodes in second hidden layer
"""
super(Actor, self).__init__()
self.fc1 = nn.Linear(state_size, fc1_units)
self.do1 = nn.Dropout(0.2)
self.fc2 = nn.Linear(fc1_units, fc2_units)
self.do2 = nn.Dropout(0.1)
self.fc3 = nn.Linear(fc2_units, action_size)
self.reset_parameters()
def reset_parameters(self):
self.fc1.weight.data.uniform_(*hidden_init(self.fc1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state):
"""Build an actor (policy) network that maps states -> actions."""
x = F.elu(self.do1(self.fc1(state)))
x = F.elu(self.do2(self.fc2(x)))
return F.tanh(self.fc3(x))
class Critic(nn.Module):
"""Critic (Value) Model."""
def __init__(self, state_size, action_size, fcs1_units=512, fc2_units=256):
"""Initialize parameters and build model.
Params
======
state_size (int): Dimension of each state
action_size (int): Dimension of each action
seed (int): Random seed
fcs1_units (int): Number of nodes in the first hidden layer
fc2_units (int): Number of nodes in the second hidden layer
"""
super(Critic, self).__init__()
self.fcs1 = nn.Linear(state_size, fcs1_units)
self.do1 = nn.Dropout(0.2)
self.fc2 = nn.Linear(fcs1_units+action_size, fc2_units)
self.do2 = nn.Dropout(0.1)
self.fc3 = nn.Linear(fc2_units, 1)
self.reset_parameters()
def reset_parameters(self):
self.fcs1.weight.data.uniform_(*hidden_init(self.fcs1))
self.fc2.weight.data.uniform_(*hidden_init(self.fc2))
self.fc3.weight.data.uniform_(-3e-3, 3e-3)
def forward(self, state, action):
"""Build a critic (value) network that maps (state, action) pairs -> Q-values."""
xs = F.leaky_relu(self.do1(self.fcs1(state)))
x = torch.cat((xs, action), dim=1)
x = F.leaky_relu(self.do2(self.fc2(x)))
return self.fc3(x)
class DDPG(object):
"""Interacts with and learns from the environment.
There are two agents and the observations of each agent has 24 dimensions. Each agent's action has 2 dimensions.
Will use two separate actor networks (one for each agent using each agent's observations only and output that agent's action).
The critic for each agents gets to see the actions and observations of all agents. """
def __init__(self, state_size, action_size, num_agents):
"""Initialize an Agent object.
Params
======
state_size (int): dimension of each state for each agent
action_size (int): dimension of each action for each agent
"""
self.state_size = state_size
self.action_size = action_size
# Actor Network (w/ Target Network)
self.actor_local = Actor(state_size, action_size).to(device)
self.actor_target = Actor(state_size, action_size).to(device)
self.actor_optimizer = optim.Adam(self.actor_local.parameters(), lr=LR_ACTOR, weight_decay=WEIGHT_DECAY_ACTOR)
# Critic Network (w/ Target Network)
self.critic_local = Critic(num_agents*state_size, num_agents*action_size).to(device)
self.critic_target = Critic(num_agents*state_size, num_agents*action_size).to(device)
self.critic_optimizer = optim.Adam(self.critic_local.parameters(), lr=LR_CRITIC, weight_decay=WEIGHT_DECAY_CRITIC)
# Noise process
self.noise_scale = NOISE_START
# Make sure target is initialized with the same weight as the source (makes a big difference)
hard_update(self.actor_target, self.actor_local)
hard_update(self.critic_target, self.critic_local)
def act(self, states):
"""Returns actions for given state as per current policy."""
states = torch.from_numpy(states).float().to(device)
self.actor_local.eval()
with torch.no_grad():
actions = self.actor_local(states).cpu().data.numpy()
self.actor_local.train()
#add noise
actions += self.noise_scale * (2 * np.random.randn(1, self.action_size))
return np.clip(actions, -1, 1)
def reset(self):
pass
def learn(self, experiences, gamma):
#for MADDPG
"""Update policy and value parameters using given batch of experience tuples.
Q_targets = r + γ * critic_target(next_state, actor_target(next_state))
where:
actor_target(state) -> action
critic_target(state, action) -> Q-value
Params
======
experiences (Tuple[torch.Tensor]): tuple of (s, a, r, s', done) tuples
gamma (float): discount factor
"""
full_states, actor_full_actions, full_actions, agent_rewards, agent_dones, full_next_states, critic_full_next_actions = experiences
# ---------------------------- update critic ---------------------------- #
# Get Q values from target models
Q_target_next = self.critic_target(full_next_states, critic_full_next_actions)
# Compute Q targets for current states (y_i)
Q_target = agent_rewards + gamma * Q_target_next * (1 - agent_dones)
# Compute critic loss
Q_expected = self.critic_local(full_states, full_actions)
critic_loss = F.mse_loss(input=Q_expected, target=Q_target) #target=Q_targets.detach() #not necessary to detach
# Minimize the loss
self.critic_optimizer.zero_grad()
critic_loss.backward()
#torch.nn.utils.clip_grad_norm(self.critic_local.parameters(), 1.0) #clip the gradient for the critic network (Udacity hint)
self.critic_optimizer.step()
# ---------------------------- update actor ---------------------------- #
# Compute actor loss
actor_loss = -self.critic_local.forward(full_states, actor_full_actions).mean() #-ve b'cse we want to do gradient ascent
# Minimize the loss
self.actor_optimizer.zero_grad()
actor_loss.backward()
self.actor_optimizer.step()
def soft_update(self):
# ----------------------- update target networks ----------------------- #
soft_update(self.critic_local, self.critic_target, TAU)
soft_update(self.actor_local, self.actor_target, TAU)
# -
# ### 5. MADDPG
# This class implements the rest of the MADDPG agent, and interacts with the DDPG agents. This class keeps the replay buffer that is shared by all the agents, this way they share the experiences acumulated on each episode.
# +
class MADDPG(object):
'''The main class that defines and trains all the agents'''
def __init__(self, state_size, action_size, num_agents):
self.state_size = state_size
self.action_size = action_size
self.num_agents = num_agents
self.whole_action_dim = self.action_size*self.num_agents
# Replay memory
self.memory = ReplayBuffer(BUFFER_SIZE, BATCH_SIZE)
# DDPG agents
self.agents = [DDPG(state_size, action_size, num_agents),\
DDPG(state_size, action_size, num_agents)]
#Noise decay
self.noise_scale = NOISE_START
def reset(self):
for agent in self.agents:
agent.reset()
def step_episode(self, i_episode):
self.noise_scale *= NOISE_REDUCTION
if self.noise_scale < NOISE_END:
self.noise_scale = NOISE_END
for agent in self.agents:
agent.noise_scale = self.noise_scale
def step(self, states, actions, rewards, next_states, dones):
#for stepping maddpg
"""Save experience in replay memory, and use random sample from buffer to learn."""
# index 0 is for agent 0 and index 1 is for agent 1
full_states = np.reshape(states, newshape=(-1))
full_next_states = np.reshape(next_states, newshape=(-1))
# Save experience / reward
self.memory.add(full_states, states, actions, rewards, full_next_states, next_states, dones)
# Learn, if enough samples are available in memory
if len(self.memory) > BATCH_SIZE:
for agent_no in range(self.num_agents):
samples = self.memory.sample()
self.learn(samples, agent_no, GAMMA)
#soft update all the agents
for agent in self.agents:
agent.soft_update()
def learn(self, samples, agent_no, gamma):
#for learning MADDPG
full_states, states, actions, rewards, full_next_states, next_states, dones = samples
critic_full_next_actions = torch.zeros(states.shape[:2] + (self.action_size,), dtype=torch.float, device=device)
for agent_id, agent in enumerate(self.agents):
agent_next_state = next_states[:,agent_id,:]
critic_full_next_actions[:,agent_id,:] = agent.actor_target.forward(agent_next_state)
critic_full_next_actions = critic_full_next_actions.view(-1, self.whole_action_dim)
agent = self.agents[agent_no]
agent_state = states[:,agent_no,:]
actor_full_actions = actions.clone() #create a deep copy
actor_full_actions[:,agent_no,:] = agent.actor_local.forward(agent_state)
actor_full_actions = actor_full_actions.view(-1, self.whole_action_dim)
full_actions = actions.view(-1,self.whole_action_dim)
agent_rewards = rewards[:,agent_no].view(-1,1) #gives wrong result without doing this
agent_dones = dones[:,agent_no].view(-1,1) #gives wrong result without doing this
experiences = (full_states, actor_full_actions, full_actions, agent_rewards, \
agent_dones, full_next_states, critic_full_next_actions)
agent.learn(experiences, gamma)
def act(self, full_states):
# all actions between -1 and 1
actions = []
for agent_id, agent in enumerate(self.agents):
action = agent.act(np.reshape(full_states[agent_id,:], newshape=(1,-1)))
action = np.reshape(action, newshape=(1,-1))
actions.append(action)
actions = np.concatenate(actions, axis=0)
return actions
def save_maddpg(self):
for agent_id, agent in enumerate(self.agents):
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor_' + str(agent_id) + '.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic_' + str(agent_id) + '.pth')
# -
class ReplayBuffer(object):
"""Fixed-size buffer to store experience tuples."""
def __init__(self, buffer_size, batch_size):
"""Initialize a ReplayBuffer object.
Params
======
buffer_size (int): maximum size of buffer
batch_size (int): size of each training batch
"""
self.memory = deque(maxlen=buffer_size) # internal memory (deque)
self.batch_size = batch_size
self.experience = namedtuple("Experience", field_names=["full_state", "state", "action", "reward", \
"full_next_state", "next_state", "done"])
def add(self, full_state, state, action, reward, full_next_state, next_state, done):
"""Add a new experience to memory."""
e = self.experience(full_state, state, action, reward, full_next_state, next_state, done)
self.memory.append(e)
def sample(self):
"""Randomly sample a batch of experiences from memory."""
experiences = random.sample(self.memory, k=self.batch_size)
full_states = torch.from_numpy(np.array([e.full_state for e in experiences if e is not None])).float().to(device)
states = torch.from_numpy(np.array([e.state for e in experiences if e is not None])).float().to(device)
actions = torch.from_numpy(np.array([e.action for e in experiences if e is not None])).float().to(device)
rewards = torch.from_numpy(np.array([e.reward for e in experiences if e is not None])).float().to(device)
full_next_states = torch.from_numpy(np.array([e.full_next_state for e in experiences if e is not None])).float().to(device)
next_states = torch.from_numpy(np.array([e.next_state for e in experiences if e is not None])).float().to(device)
dones = torch.from_numpy(np.array([e.done for e in experiences if e is not None]).astype(np.uint8)).float().to(device)
return (full_states, states, actions, rewards, full_next_states, next_states, dones)
def __len__(self):
"""Return the current size of internal memory."""
return len(self.memory)
# ### 6. Train
# To train this agent, here we define some hyperparameters, that have been tuned. In this case, the agent takes arround 400 episodes to start producing a good enough output. During this episodes, the exploratory noise has to be kept high enough to generate outcomes that produce some random positive rewards, otherwise the agent by itself will not found positive outcomes and will not converge.
# +
#Train hyperparameters
BUFFER_SIZE = int(1e5) # replay buffer size
BATCH_SIZE = 512 # minibatch size
GAMMA = 0.99 # discount factor
TAU = 1e-3 # for soft update of target parameters
LR_ACTOR = 1e-4 # learning rate of the actor
LR_CRITIC = 3e-4 # learning rate of the critic
WEIGHT_DECAY_ACTOR = 0.0 # L2 weight decay
WEIGHT_DECAY_CRITIC = 0.0 # L2 weight decay
#to decay exploration as it learns
NOISE_START=2.0
NOISE_END=0.01
NOISE_REDUCTION=0.995
seeding(seed=3)
state_size=env_info.vector_observations.shape[1]
action_size=brain.vector_action_space_size
num_agents=env_info.vector_observations.shape[0]
maddpg_agent = MADDPG(state_size=state_size, action_size=action_size, num_agents=num_agents)
#Training
def train_MADDPG(n_episodes=2500, t_max=1000):
scores_deque = deque(maxlen=100)
scores_list = []
scores_list_100_avg = []
for i_episode in range(1, n_episodes+1):
env_info = env.reset(train_mode=True)[brain_name] # reset the environment
states = env_info.vector_observations # get the current states (for all agents)
maddpg_agent.reset() #reset the maddpg_agent
scores = np.zeros(num_agents) # initialize the score (for each agent in MADDPG)
num_steps = 0
maddpg_agent.step_episode(i_episode)
for _ in range(t_max):
actions = maddpg_agent.act(states)
env_info = env.step(actions)[brain_name] # send all actions to the environment
next_states = env_info.vector_observations # get next state (for each agent in MADDPG)
rewards = env_info.rewards # get rewards (for each agent in MADDPG)
dones = env_info.local_done # see if episode finished
scores += rewards # update the score (for each agent in MADDPG)
maddpg_agent.step(states, actions, rewards, next_states, dones) #train the maddpg_agent
states = next_states # roll over states to next time step
num_steps += 1
if np.any(dones): # exit loop if episode finished
break
scores_deque.append(np.max(scores))
scores_list.append(np.max(scores))
scores_list_100_avg.append(np.mean(scores_deque))
print('\rEpisode {}, Average Score: {:.2f}, Current Score: {:.2f}, Noise Scaling: {:.2f}, Memory size: {}, Num Steps: {}'.format(i_episode, np.mean(scores_deque), np.max(scores), maddpg_agent.noise_scale, len(maddpg_agent.memory), num_steps), end="")
if i_episode % 100 == 0:
maddpg_agent.save_maddpg()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores_list)+1), scores_list)
plt.plot(np.arange(1, len(scores_list)+1), scores_list_100_avg)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
if np.mean(scores_deque) > 0.5 and len(scores_deque) >= 100:
maddpg_agent.save_maddpg()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores_list)+1), scores_list)
plt.plot(np.arange(1, len(scores_list)+1), scores_list_100_avg)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
break
return scores_list, scores_list_100_avg
# -
# do long-running work here
scores, scores_avg = train_MADDPG()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.plot(np.arange(1, len(scores)+1), scores_avg)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# ### 7. Conclusions
# Compared to other agents that we have used during the course, this algorithms takes a longer time without producing an outcome that is close to the expected average value. But at a certain point it starts converging very fast.
# I think that the nature of the problem (repeat the ball bouncing) produces this behaveiour: once the agents learns how to hit the ball, it has just to repeat the same behaviour over and over.
# The difficulty comes from learning with another agent, that makes the environment hard to predict and where the DADDPG algorithm helps.
|
p3_collab-compet/Tennis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] cell_id="00001-f9f152cc-d6ad-4741-b838-4c6fab6abbba" deepnote_cell_type="markdown" tags=[]
# # 02-Advanced: Data analysis
# For these exercises we are going to do some text and more advanced analysis.
# + [markdown] cell_id="00001-70d86720-3538-4f91-8276-9a4e7e3c5da2" deepnote_cell_type="markdown" tags=[]
# ## Text data
# Often we will need to work with unstructured data like text, images, or audio. To use unstructured data in analyis, we will often need to convert it into something more useable, something more "quantified".
#
# Let's start easy by loading a `.csv` file containing some text data we plan to analyse. Run the code below to load `reviews_sample.csv`.
# + cell_id="00002-1634df66-c1d5-45d5-ab5d-9000f2403efd" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=516 execution_start=1633510549583 source_hash="9416d4fc" tags=[]
import pandas as pd
df_reviews = pd.read_csv('../data/reviews_sample.csv')
df_reviews.head()
# + [markdown] cell_id="00003-288e7094-680c-4292-bf1e-b730185e7cba" deepnote_cell_type="markdown" tags=[]
# ### Exercise-01: Cleaning text
# We want to analyse the `comments` column, but we first need to remove rows with `nan` values in the `comments` column. Use what you learned in `01-Basic.ipynb` to do this below.
# + cell_id="00004-4673738a-93e7-4134-ae6e-22998be22e2d" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=8 execution_start=1633510550110 source_hash="ceb90d56" tags=[]
# (SOLUTION)
# + [markdown] cell_id="00005-93c7f454-b595-40a0-a6c6-c189b129d9bc" deepnote_cell_type="markdown" tags=[]
# ### Exercise-02: Analysing text
# Now the text data is cleaned, we are going to convert the text to quantities of interest. A common way to do this is to estimate the *sentiment* of the text. There are many ways to analyse sentiment, and here we are going to use the [VADER](https://www.nltk.org/_modules/nltk/sentiment/vader.html) sentiment analytics tool which is included in the [NLTK](https://www.nltk.org/) (Natural Language Toolkit) package.
#
# Run the code below to import `nltk`, import the `SentimentIntensityAnalyzer` class from `nltk`, download the `vader_lexicon`, and create an instance of `SentimentIntensityAnalyzer`.
# + cell_id="00006-88f0f313-da50-4de2-8c59-b91ceda052e9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=1831 execution_start=1633510550168 source_hash="307a976d" tags=[]
import nltk
from nltk.sentiment.vader import SentimentIntensityAnalyzer
nltk.download('vader_lexicon')
scorer = SentimentIntensityAnalyzer()
# + [markdown] cell_id="00006-2e37548c-c4e9-48bb-9088-4f3a793e0499" deepnote_cell_type="markdown" tags=[]
# Now we have loaded our sentiment analysis tool, we can calculate the sentiment of the comments in the Airbnb reviews. To do this, we are going to create a `calculate_sentiment` function and then apply it to a small sample (`N=5`) of rows from `df_reviews`. To focus our attention on the columns `['comments','sentiment']` we are going to show only those in the `head()` of `df_reviews_sample`. Please run the code below and watch the magic happen!
# + cell_id="00008-3d2d2f75-8993-4673-ae50-ccd7bef43b8a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=60 execution_start=1633510552007 source_hash="5d36530c" tags=[]
def calculate_sentiment(comment):
return(scorer.polarity_scores(comment)['compound'])
N = 5
df_reviews_sample = df_reviews.sample(N)
df_reviews_sample.loc[:,'sentiment'] = df_reviews_sample['comments'].apply(calculate_sentiment)
df_reviews_sample[['comments','sentiment']].head()
# + [markdown] cell_id="00009-9ff1b9ac-aee6-482e-82e5-dfa41f358959" deepnote_cell_type="markdown" tags=[]
# Run the above code a few times, to get a feel for how the sentiment analysis tool is working. Is there anything unexpected/interesting?
# + [markdown] cell_id="00001-d2b6a883-b571-449b-83d3-2f0379123a4e" deepnote_cell_type="markdown" tags=[]
# ## Segmented data
# Sometimes we want to perform analysis on only a segment on the data. For example, someone might ask *what is the most expensive listing in London?*, and to solve this we would only need to analysis data for listings based in London.
#
# To explore this idea, let's analyse some segments of the `listings_sample.csv` data. Run the code below to load `listings_sample.csv`.
# + cell_id="00002-38d1fa1d-4b4c-47a0-9455-637abde65d4a" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=832 execution_start=1633510552063 source_hash="f5e2616a" tags=[]
import pandas as pd
df_listings = pd.read_csv('../data/listings_sample.csv')
df_listings.head()
# + [markdown] cell_id="00013-7d435d32-23a9-4455-b13f-cd971bae9808" deepnote_cell_type="markdown" tags=[]
# Let's assume we want to segment the listings by `room_type`. First, let's take a quick look at how many listings of each `room_type` there are in `df_listings`. To do this, we can use the `value_counts()` method. Please run the code below.
# + cell_id="00003-589ec3dd-36ab-40c1-90ad-860a97e56620" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=31 execution_start=1633510552905 source_hash="97e6acc6" tags=[]
df_listings['room_type'].value_counts().head()
# + [markdown] cell_id="00002-b862573b-05d6-4559-94cf-493bfabfadf4" deepnote_cell_type="markdown" tags=[]
# ### Exercise-03: Identify luxury homes
# You are a business analyst at a company which a business development manager who asks you to identify the 5 most expensive listings which are entire homes/apartments.
#
# To do this, you first need to create a new `price_$` column in `df_listings` like we did in `01-Basic.ipynb`. Please do that using the `format_price` function written below.
# + cell_id="00015-e16a4998-d0e0-4f43-9f4a-62c9c48ee484" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=58 execution_start=1633510552964 source_hash="1c897ce6" tags=[]
def format_price(price):
return(float(price.replace('$','').replace(',','')))
# (SOLUTION)
# + [markdown] cell_id="00016-05dd5e08-bffb-4942-9ca2-b8d8f0e55f9c" deepnote_cell_type="markdown" tags=[]
# Now we have price values in a format we can use, we can segment the data and run our analysis to meet the manager's request. To do this, we will:
#
# 1. Segment `df_listings` to create a new `DataFrame` named `df_entire_home_apt` which contains only listings with `room_type` equal to `Entire home/apt`.
# 2. Sort the rows of `df_entire_home_apt` by the values in `price_$` from largest (most expensive) to smallest (cheapest).
# 3. Display the top 5 rows to show columns we think most relevant for the 5 most expensive listings which are entire homes/apartments.
#
# We will do all this by running only the 3 lines of code shown below. Please run the code.
# + cell_id="00005-c178ce40-626a-4f1e-a5c5-bae461aca1c3" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=75 execution_start=1633510553008 source_hash="129ca33c" tags=[]
df_entire_home_apt = df_listings[df_listings['room_type']=='Entire home/apt']
df_entire_home_apt = df_entire_home_apt.sort_values(by=['price_$'],ascending=False)
df_entire_home_apt[['id','name','description','neighbourhood','price_$']].head(5)
# + [markdown] cell_id="00004-4fa624fe-6cd9-4e58-93eb-48dd59579c08" deepnote_cell_type="markdown" tags=[]
# ### Exercise 04: Identify budget rooms
# the same business development manager now asks you to identify the 5 cheapest listings which are private rooms. Using the above exercise as a guide, run an analysis to meet the manager's request.
# + cell_id="00004-9098e09e-69e2-414a-b7ac-4fa3391d5686" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=49 execution_start=1633510553086 source_hash="f2cc5168" tags=[]
# (SOLUTION)
# + [markdown] cell_id="00002-0c75d8a2-e584-4da0-98c2-0f13d8152dce" deepnote_cell_type="markdown" tags=[]
# ## Advanced request
# Well done so far! Let's now take it too the next level! Let's used what we have learned so far to meet the following request:
#
# *Identify the 5 listings with highest positive mean sentiment of their reviews, such that the listings have at least 5 reviews less than 3 years old*
# + [markdown] cell_id="00002-7d28c2af-73d4-42d7-b2fb-dcbbb4652939" deepnote_cell_type="markdown" tags=[]
# ### Exercise-05: Filter review data
# First we need to load `reviews_sample.csv` and filter it to only keep reviews with:
# 1. values of not `nan` in the `comments` column,
# 2. values less than 3 years old in the `date` column, and
# 3. values in the `listing_id` column corresponding to listings with at least 5 reviews.
#
# This can all be done by running the following code.
# + cell_id="00000-6f9e41dc-39f8-462f-888c-92d2cc0ff357" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=508 execution_start=1633510553130 source_hash="f353607d" tags=[]
import pandas as pd
df_reviews = pd.read_csv('../data/reviews_sample.csv')
df_reviews = df_reviews[~df_reviews['comments'].isna()]
df_reviews = df_reviews[df_reviews['date']>'2018-10-05']
listing_counts = df_reviews['listing_id'].value_counts()
valid_listings = listing_counts[listing_counts>=5].index
df_reviews = df_reviews[df_reviews['listing_id'].isin(valid_listings)]
# + [markdown] cell_id="00023-60af850b-2d2e-421a-9247-4aeaa401d6ea" deepnote_cell_type="markdown" tags=[]
# We can do a "sanity check" by running the code below, which should all produce value `0`. Consider how you might use such test to check your code as you develop it.
# + cell_id="00024-511e6b18-e2f2-4406-9244-fb00370ff859" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=86 execution_start=1633510650116 source_hash="e03d4b06" tags=[]
print(sum(df_reviews['comments'].isna()))
print(sum(df_reviews['date']<='2018-10-05'))
print(sum(df_reviews['listing_id'].value_counts()<5))
# + [markdown] cell_id="00023-f33b4ce0-222e-47d4-af7b-19dfd829b4f0" deepnote_cell_type="markdown" tags=[]
# ### Exercise-06: Analyse review data
# We already have most of the pieces we need. Simply `apply` the `calculate_sentiment` function (in the way we did before) to the `comments` column of the (now filtered) `df_reviews` to create a new column named `sentiment` containing the a sentiment value for each reviews comment. Please write the code below and run.
#
# (Note that we are no running the `calculate_sentiment` over every row in the filtered `df_reviews` so it might take a while longer...)
# + cell_id="00003-9d0b5a27-934f-4498-a97c-1dc8a75b26f0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=12428 execution_start=1633510553704 source_hash="9a409e5f" tags=[]
# SOLUTION
# + [markdown] cell_id="00005-2e327721-cef6-445c-bd4d-73907ceecc87" deepnote_cell_type="markdown" tags=[]
# Next we need to calculate the mean sentiment for each listing. To do this, we use the `groupby` and `agg` methods on the following way to create a new `DataFrame` named `listing_scored` containing each `listings_id` and the `mean` of its `sentiment` scores. Please run the code below.
# + cell_id="00007-1560c73a-737a-46e3-a62b-b38601b3ddda" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=47 execution_start=1633510566127 source_hash="76c3be3f" tags=[]
listings_scored = df_reviews.groupby('listing_id')['sentiment'].agg(['mean']).reset_index()
listings_scored.head()
# + [markdown] cell_id="00011-4c49117b-bd28-4c19-946a-dcaa92e4838b" deepnote_cell_type="markdown" tags=[]
# Finally, we just need to sort the listings in `listings_scored` by their `mean` value (from high to low) and print the top 5. Easy, you've got this! Please use what you've learned before to do this below.
# + cell_id="00030-f0ac8274-db95-4bb4-aac8-53714e767555" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=39 execution_start=1633510566163 source_hash="79003250" tags=[]
# SOLUTION
# + [markdown] cell_id="00031-07f63de9-ebc3-4dcf-953e-f795dc43d59e" deepnote_cell_type="markdown" tags=[]
# ## Inspect results
# Have a play around with the above code. Maybe try to fetch and inspect the reviews of the listings with high and low sentiment socres, and see what types of listings these are.
#
# Or, if you've found this notebook too easy, have a go at `03-Expert.ipynb`.
# + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
# <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=2c6f047c-21a6-4149-814c-b3f60a9bf973' target="_blank">
# <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPSJ0cmFuc2xhdGUoLTEyMzUuMDAwMDAwLCAtNzkuMDAwMDAwKSI+CiAgICAgICAgICAgIDxnIGlkPSJHcm91cC0zIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgxMjM1LjAwMDAwMCwgNzkuMDAwMDAwKSI+CiAgICAgICAgICAgICAgICA8cG9seWdvbiBpZD0iUGF0aC0yMCIgZmlsbD0iIzAyNjVCNCIgcG9pbnRzPSIyLjM3NjIzNzYyIDgwIDM4LjA0NzY2NjcgODAgNTcuODIxNzgyMiA3My44MDU3NTkyIDU3LjgyMTc4MjIgMzIuNzU5MjczOSAzOS4xNDAyMjc4IDMxLjY4MzE2ODMiPjwvcG9seWdvbj4KICAgICAgICAgICAgICAgIDxwYXRoIGQ9Ik0zNS4wMDc3MTgsODAgQzQyLjkwNjIwMDcsNzYuNDU0OTM1OCA0Ny41NjQ5MTY3LDcxLjU0MjI2NzEgNDguOTgzODY2LDY1LjI2MTk5MzkgQzUxLjExMjI4OTksNTUuODQxNTg0MiA0MS42NzcxNzk1LDQ5LjIxMjIyODQgMjUuNjIzOTg0Niw0OS4yMTIyMjg0IEMyNS40ODQ5Mjg5LDQ5LjEyNjg0NDggMjkuODI2MTI5Niw0My4yODM4MjQ4IDM4LjY0NzU4NjksMzEuNjgzMTY4MyBMNzIuODcxMjg3MSwzMi41NTQ0MjUgTDY1LjI4MDk3Myw2Ny42NzYzNDIxIEw1MS4xMTIyODk5LDc3LjM3NjE0NCBMMzUuMDA3NzE4LDgwIFoiIGlkPSJQYXRoLTIyIiBmaWxsPSIjMDAyODY4Ij48L3BhdGg+CiAgICAgICAgICAgICAgICA8cGF0aCBkPSJNMCwzNy43MzA0NDA1IEwyNy4xMTQ1MzcsMC4yNTcxMTE0MzYgQzYyLjM3MTUxMjMsLTEuOTkwNzE3MDEgODAsMTAuNTAwMzkyNyA4MCwzNy43MzA0NDA1IEM4MCw2NC45NjA0ODgyIDY0Ljc3NjUwMzgsNzkuMDUwMzQxNCAzNC4zMjk1MTEzLDgwIEM0Ny4wNTUzNDg5LDc3LjU2NzA4MDggNTMuNDE4MjY3Nyw3MC4zMTM2MTAzIDUzLjQxODI2NzcsNTguMjM5NTg4NSBDNTMuNDE4MjY3Nyw0MC4xMjg1NTU3IDM2LjMwMzk1NDQsMzcuNzMwNDQwNSAyNS4yMjc0MTcsMzcuNzMwNDQwNSBDMTcuODQzMDU4NiwzNy43MzA0NDA1IDkuNDMzOTE5NjYsMzcuNzMwNDQwNSAwLDM3LjczMDQ0MDUgWiIgaWQ9IlBhdGgtMTkiIGZpbGw9IiMzNzkzRUYiPjwvcGF0aD4KICAgICAgICAgICAgPC9nPgogICAgICAgIDwvZz4KICAgIDwvZz4KPC9zdmc+' > </img>
# Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
|
exercises/02-Advanced.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import re
import string
from nltk import word_tokenize
from nltk.stem import WordNetLemmatizer
from sklearn.feature_extraction.text import TfidfVectorizer
from collections import Counter
from pylab import *
import nltk
import warnings
warnings.filterwarnings('ignore')
data_patio_lawn_garden = pd.read_json('data/reviews_Patio_Lawn_and_Garden_5.json', lines = True)
data_patio_lawn_garden[['reviewText', 'overall']].head()
lemmatizer = WordNetLemmatizer()
data_patio_lawn_garden['cleaned_review_text'] = data_patio_lawn_garden['reviewText'].apply(\
lambda x : ' '.join([lemmatizer.lemmatize(word.lower()) \
for word in word_tokenize(re.sub(r'([^\s\w]|_)+', ' ', str(x)))]))
data_patio_lawn_garden[['cleaned_review_text', 'reviewText', 'overall']].head()
tfidf_model = TfidfVectorizer(max_features=500)
tfidf_df = pd.DataFrame(tfidf_model.fit_transform(data_patio_lawn_garden['cleaned_review_text']).todense())
tfidf_df.columns = sorted(tfidf_model.vocabulary_)
tfidf_df.head()
data_patio_lawn_garden['target'] = data_patio_lawn_garden['overall'].apply(lambda x : 0 if x<=4 else 1)
data_patio_lawn_garden['target'].value_counts()
from sklearn import tree
dtc = tree.DecisionTreeClassifier()
dtc = dtc.fit(tfidf_df, data_patio_lawn_garden['target'])
data_patio_lawn_garden['predicted_labels_dtc'] = dtc.predict(tfidf_df)
pd.crosstab(data_patio_lawn_garden['target'], data_patio_lawn_garden['predicted_labels_dtc'])
from sklearn import tree
dtr = tree.DecisionTreeRegressor()
dtr = dtr.fit(tfidf_df, data_patio_lawn_garden['overall'])
data_patio_lawn_garden['predicted_values_dtr'] = dtr.predict(tfidf_df)
data_patio_lawn_garden[['predicted_values_dtr', 'overall']].head(10)
|
Chapter03/Exercise 3.07.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.1 64-bit (''motorcycles-prices'': conda)'
# language: python
# name: python3
# ---
# ## Data Preprocessing and EDA
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import os
# %matplotlib inline
# -
df = pd.read_csv('../data/iterim/bikes-02-new_features.csv')
df
# +
# Search for the most sales by brand
plt.figure(figsize=(16, 8))
df['brand_name'].value_counts().plot(kind='bar')
# +
# Search for the most sales by model
plt.figure(figsize=(18,9))
df['model_name'].value_counts()[:50].plot(kind='bar')
# -
# It's clear Bajaj and Royal Enfield brands are the most popular sales
# and Pulsar model is the most popular sales
# +
# take a look for the histogram of each feature
df.hist(figsize=(15, 10))
plt.show()
# -
# Look a weird behaviour in "kms_driven" feature
sns.boxplot(x=df['kms_driven'])
# In effect have rows with more than 1M in the column
df = df[df['kms_driven'] < df['kms_driven'].quantile(0.975)]
sns.boxplot(x=df['kms_driven'])
# New distribution shows another outliers, but are realistic values
# Now teke a look in the "price column"
sns.boxplot(x=df['price'])
# Clearly have a outlier with 3M value, i'll fix it
df = df[df['price'] < 3000000]
sns.boxplot(x=df['price'])
df.hist(figsize=(15, 10))
plt.show()
sns.pairplot(df)
plt.figure(figsize=(10, 5))
sns.heatmap(df.corr(), annot=True)
# Can see that the *power* and the *motor size* is bigger influenced in **price**, and have negative but strong correlation with the *mileage*.
DATA_DIR_iterim = '../data/iterim/'
df.to_csv(os.path.join(DATA_DIR_iterim, 'bikes-03-no_outliers.csv'), index=False)
|
notebooks/2.0-motorcycles-visuzlizations-fix-outliers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="acbetMKBt825"
# <div align="center">
# <h1><img width="30" src="https://madewithml.com/static/images/rounded_logo.png"> <a href="https://madewithml.com/">Made With ML</a></h1>
# Applied ML · MLOps · Production
# <br>
# Join 30K+ developers in learning how to responsibly <a href="https://madewithml.com/about/">deliver value</a> with ML.
# <br>
# </div>
#
# <br>
#
# <div align="center">
# <a target="_blank" href="https://newsletter.madewithml.com"><img src="https://img.shields.io/badge/Subscribe-30K-brightgreen"></a>
# <a target="_blank" href="https://github.com/GokuMohandas/MadeWithML"><img src="https://img.shields.io/github/stars/GokuMohandas/MadeWithML.svg?style=social&label=Star"></a>
# <a target="_blank" href="https://www.linkedin.com/in/goku"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
# <a target="_blank" href="https://twitter.com/GokuMohandas"><img src="https://img.shields.io/twitter/follow/GokuMohandas.svg?label=Follow&style=social"></a>
# <br>
# 🔥 Among the <a href="https://github.com/topics/mlops" target="_blank">top MLOps</a> repositories on GitHub
# </div>
#
# <br>
# <hr>
# + [markdown] id="ZbHqEEy3NkSU"
# # MLOps - Tagifai
# + [markdown] id="XTNsIiUrqoJW"
# <div align="left">
# <a target="_blank" href="https://madewithml.com/#mlops"><img src="https://img.shields.io/badge/📖 Read-lessons-9cf"></a>
# <a href="https://github.com/GokuMohandas/MLOps/blob/main/notebooks/tagifai.ipynb" role="button"><img src="https://img.shields.io/static/v1?label=&message=View%20On%20GitHub&color=586069&logo=github&labelColor=2f363d"></a>
# <a href="https://colab.research.google.com/github/GokuMohandas/MLOps/blob/main/notebooks/tagifai.ipynb"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
# </div>
# + [markdown] id="oh-HuNfDrPg0"
# This notebooks contains the code for our `Tagifai` feature including 🔢 Data and 📈 Modeling. After this, we'll be moving all of this code to Python scripts with proper styling, testing, etc.
#
# > Be sure to checkout the accompanying [lessons](https://madewithml.com/courses/mlops) as opposed to just running the code here. The lessons will help us develop an intuition before jumping into the application.
# + [markdown] id="lbqAqitENkSU"
# # 🔢 Data
# + [markdown] id="2ROgHAeQNkSU"
# ## Labeling
# + id="VdrrsquiNkSU"
from collections import Counter, OrderedDict
import ipywidgets as widgets
import itertools
import json
import pandas as pd
from urllib.request import urlopen
# + id="WB-2nm6NNkSU" colab={"base_uri": "https://localhost:8080/"} outputId="3c568eeb-8658-4370-e423-efd084fcc5dc"
# Load projects
url = "https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/datasets/projects.json"
projects = json.loads(urlopen(url).read())
print (json.dumps(projects[-305], indent=2))
# + id="_pJcQOR7NkSU" colab={"base_uri": "https://localhost:8080/"} outputId="a0ffda5f-4b50-460d-a49c-39fbea03244d"
# Load tags
url = "https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/datasets/tags.json"
tags = json.loads(urlopen(url).read())
tags_dict = {}
for item in tags:
key = item.pop("tag")
tags_dict[key] = item
print (f"{len(tags_dict)} tags")
# + id="l8GWm_unNkSU" colab={"base_uri": "https://localhost:8080/", "height": 185, "referenced_widgets": ["bdcf0a4e930047d4bf0233ec50c48de5", "f084e9794c0342c3ba4b7497472f3c86", "c3489da1f1094829a61f7a8a0d3d8de6", "63ca8ab59f9e4eb9a514524b7e3890c2", "3ff1259660ac48b0956b3819fa95b5c0", "02166c9af03d4476bb8b94f6424ed40e", "15fe95599879419280726a9295b169a3"]} outputId="7bed2260-cffd-4095-eb03-f7f5db2133f5"
@widgets.interact(tag=list(tags_dict.keys()))
def display_tag_details(tag='question-answering'):
print (json.dumps(tags_dict[tag], indent=2))
# + id="ZAYXvZ_TNkSU" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="f419a816-0e00-4f2b-8e1b-abbf05071f94"
# Create dataframe
df = pd.DataFrame(projects)
print (f"{len(df)} projects")
df.head(5)
# + [markdown] id="HFifXKl_eKsN"
# ## Preprocessing
# + [markdown] id="RxAZ1AmteRaD"
# Preprocessing the data via feature engineering, filtering and cleaning. Certain preprocessing steps are global (don't depend on our dataset, ex. lower casing text, removing stop words, etc.) and others are local (constructs are learned only from the training split, ex. vocabulary, standardization, etc.). For the local, dataset-dependent preprocessing steps, we want to ensure that we [split](https://madewithml.com/courses/mlops/splitting) the data first before preprocessing to avoid data leaks.
# + [markdown] id="U_001GPyMZsC"
# We can combine existing input features to create new meaningful signal (helping the model learn).
# + id="3x1ldAFQNkSU"
# Feature engineering
df["text"] = df.title + " " + df.description
# + [markdown] id="spTOm3UvMdtH"
# Filter tags above a certain frequency threshold because those with fewer samples won't be adequate for training.
# + id="Lt9j3gz1NkSU"
def filter(l, include=[], exclude=[]):
"""Filter a list using inclusion and exclusion lists of items."""
filtered = [item for item in l if item in include and item not in exclude]
return filtered
# + id="Q1H1lnKXNkSU"
# Inclusion/exclusion criteria for tags
include = list(tags_dict.keys())
exclude = ["machine-learning", "deep-learning", "data-science",
"neural-networks", "python", "r", "visualization", "wandb"]
# + id="FF9cfyfaNkSU"
# Filter tags for each project
df.tags = df.tags.apply(filter, include=include, exclude=exclude)
tags = Counter(itertools.chain.from_iterable(df.tags.values))
# + [markdown] id="Av4fPxjDMjAM"
# We're also going to restrict the mapping to only tags that are above a certain frequency threshold. The tags that don't have enough projects will not have enough samples to model their relationships.
# + id="k1GcLzL6NkSU" tags=["hide-input"] colab={"base_uri": "https://localhost:8080/", "height": 185, "referenced_widgets": ["8e4860a2a7e94b26960e65c928f4850f", "674a18b7ecfc484eb7fd3ef912810c91", "ecafbdf41693451a98e72e5a8b8c9656", "9a10f66838db4dc8b4b3939fc04beb70", "<KEY>", "<KEY>", "ae2f241082ef4c57aef0f63b0766ea32"]} outputId="88d1e2fc-b47b-4fb2-fd30-bee2c573a578"
@widgets.interact(min_tag_freq=(0, tags.most_common()[0][1]))
def separate_tags_by_freq(min_tag_freq=30):
tags_above_freq = Counter(tag for tag in tags.elements()
if tags[tag] >= min_tag_freq)
tags_below_freq = Counter(tag for tag in tags.elements()
if tags[tag] < min_tag_freq)
print ("Most popular tags:\n", tags_above_freq.most_common(5))
print ("\nTags that just made the cut:\n", tags_above_freq.most_common()[-5:])
print ("\nTags that just missed the cut:\n", tags_below_freq.most_common(5))
# + id="JjaEbjzONkSV"
# Filter tags that have fewer than <min_tag_freq> occurrences
min_tag_freq = 30
tags_above_freq = Counter(tag for tag in tags.elements()
if tags[tag] >= min_tag_freq)
df.tags = df.tags.apply(filter, include=list(tags_above_freq.keys()))
# + [markdown] id="-KI1xupRMtgw"
# Remove inputs that have no remaining (not enough frequency) tags.
# + id="PTdbpY73LdP9"
import nltk
from nltk.corpus import stopwords
from nltk.stem import PorterStemmer
import re
# + id="xRK_c_CFNkSV" colab={"base_uri": "https://localhost:8080/"} outputId="daef6e0c-c07a-4373-dec6-e7b62617dc72"
# Remove projects with no more remaining relevant tags
df = df[df.tags.map(len) > 0]
print (f"{len(df)} projects")
# + [markdown] id="8_f1LpKpMwzO"
# Since we're dealing with text data, we can apply some of the common preparation processes:
# + id="VDXLH6QeLd0F" colab={"base_uri": "https://localhost:8080/"} outputId="1a52e80b-110c-49a2-c85f-88d144fef7e1"
nltk.download('stopwords')
STOPWORDS = stopwords.words('english')
porter = PorterStemmer()
# + id="VfdWkkV8LlNR"
def preprocess(text, lower=True, stem=False,
filters="[!\"'#$%&()*\+,-./:;<=>?@\\\[\]^_`{|}~]",
stopwords=STOPWORDS):
"""Conditional preprocessing on our text unique to our task."""
# Lower
if lower:
text = text.lower()
# Remove stopwords
pattern = re.compile(r'\b(' + r'|'.join(stopwords) + r')\b\s*')
text = pattern.sub('', text)
# Spacing and filters
text = re.sub(r"([-;;.,!?<=>])", r" \1 ", text)
text = re.sub(filters, r"", text)
text = re.sub('[^A-Za-z0-9]+', ' ', text) # remove non alphanumeric chars
text = re.sub(' +', ' ', text) # remove multiple spaces
text = text.strip()
# Remove links
text = re.sub(r'http\S+', '', text)
# Stemming
if stem:
text = " ".join([porter.stem(word) for word in text.split(' ')])
return text
# + id="L6aKH-g0LlQt" colab={"base_uri": "https://localhost:8080/", "height": 98, "referenced_widgets": ["3cf807aca9ba4d708ff2576684028ad4", "91a3841aa87a40c8bf9718361b667421", "0125d686888e444891002c469c9900b5", "305981f69def4456a6e9cc587bb551a0", "319c832b5aa14b87bfadddbada2b2da1", "d704e961a2c24e9d8d4d1c9211fed9ad", "d304af7635b741b0842223d6ae61b084", "02b70c2aa0f34e50973e1acee2192a25", "16371fcd16644a3299dc0a3a8dab171b", "d433fbc1e3744ed79d55aa03e4fafe1a"]} outputId="efa84dd2-77e6-4860-9a19-c45a6226edb4"
@widgets.interact(lower=True, stem=False)
def display_preprocessed_text(lower, stem):
text = "Conditional image generation using Variational Autoencoders and GANs."
preprocessed_text = preprocess(text=text, lower=lower, stem=stem)
print (preprocessed_text)
# + id="3LRaq0_5LpE4" colab={"base_uri": "https://localhost:8080/"} outputId="145d600f-30e0-4bdf-e13a-db91d7ad1c4b"
# Apply to dataframe
original_df = df.copy()
df.text = df.text.apply(preprocess, lower=True, stem=False)
print (f"{original_df.text.values[0]}\n{df.text.values[0]}")
# + [markdown] id="WuCrsbxbNkSV"
# ## Exploratory Data Analysis (EDA)
# + id="tHdQmqTBNkSV"
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
from wordcloud import WordCloud, STOPWORDS
sns.set_theme()
warnings.filterwarnings("ignore")
# + id="2hGuYu5ONkSV" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="83f412a3-5c99-49b1-f515-0aa6ff990f77"
# Number of tags per project
num_tags_per_project = [len(tags) for tags in df.tags]
num_tags, num_projects = zip(*Counter(num_tags_per_project).items())
plt.figure(figsize=(10, 3))
ax = sns.barplot(list(num_tags), list(num_projects))
plt.title("Tags per project", fontsize=20)
plt.xlabel("Number of tags", fontsize=16)
ax.set_xticklabels(range(1, len(num_tags)+1), rotation=0, fontsize=16)
plt.ylabel("Number of projects", fontsize=16)
plt.show()
# + id="5JuX5Ju-NkSV" colab={"base_uri": "https://localhost:8080/", "height": 580} outputId="4bdf9935-09b6-4ee9-ffe2-737b6d595f48"
# Distribution of tags
all_tags = list(itertools.chain.from_iterable(df.tags.values))
tags, tag_counts = zip(*Counter(all_tags).most_common())
plt.figure(figsize=(25, 5))
ax = sns.barplot(list(tags), list(tag_counts))
plt.title("Tag distribution", fontsize=20)
plt.xlabel("Tag", fontsize=16)
ax.set_xticklabels(tags, rotation=90, fontsize=14)
plt.ylabel("Number of projects", fontsize=16)
plt.show()
# + id="NgMGuIQrNkSV" colab={"base_uri": "https://localhost:8080/", "height": 335, "referenced_widgets": ["865b1767dc1841069d80d437617851ea", "50a4d6d73187402ea93ce1dd55b93e6f", "8491dca93c1344be88a0c5308865762d", "7aaf0551653340868c77d1b07767b630", "be045013601748f593d89de3c447c251", "919c78de6ac048c5bf5ac451201c0d61", "f047d8dbe7f5433d998388446bfc09bb"]} outputId="b8dd38a8-bf06-49b2-dd53-e59bb28fd8ec"
@widgets.interact(tag=list(tags))
def display_word_cloud(tag="pytorch"):
# Plot word clouds top top tags
plt.figure(figsize=(15, 5))
subset = df[df.tags.apply(lambda tags: tag in tags)]
text = subset.text.values
cloud = WordCloud(
stopwords=STOPWORDS, background_color="black", collocations=False,
width=500, height=300).generate(" ".join(text))
plt.axis("off")
plt.imshow(cloud)
# + [markdown] id="7sfzg9CMNkSV"
# ## Label encoding
# + id="DYKtkRjlNkSV"
import numpy as np
import random
# + id="U-khYwAVNkSV"
# Set seeds for reproducibility
seed = 42
np.random.seed(seed)
random.seed(seed)
# + id="ytQ6_8v7NkSV"
# Shuffle
df = df.sample(frac=1).reset_index(drop=True)
# + id="j4lC_E14NkSV"
# Get data
X = df.text.to_numpy()
y = df.tags
# + [markdown] id="zT6FHkqyNkSV"
# We'll be writing our own LabelEncoder which is based on scikit-learn's [implementation](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html).
# + id="YHPmj1XANkSV"
class LabelEncoder(object):
"""Label encoder for tag labels."""
def __init__(self, class_to_index={}):
self.class_to_index = class_to_index
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
def __len__(self):
return len(self.class_to_index)
def __str__(self):
return f"<LabelEncoder(num_classes={len(self)})>"
def fit(self, y):
classes = np.unique(list(itertools.chain.from_iterable(y)))
for i, class_ in enumerate(classes):
self.class_to_index[class_] = i
self.index_to_class = {v: k for k, v in self.class_to_index.items()}
self.classes = list(self.class_to_index.keys())
return self
def encode(self, y):
y_one_hot = np.zeros((len(y), len(self.class_to_index)), dtype=int)
for i, item in enumerate(y):
for class_ in item:
y_one_hot[i][self.class_to_index[class_]] = 1
return y_one_hot
def decode(self, y):
classes = []
for i, item in enumerate(y):
indices = np.where(item == 1)[0]
classes.append([self.index_to_class[index] for index in indices])
return classes
def save(self, fp):
with open(fp, 'w') as fp:
contents = {'class_to_index': self.class_to_index}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, 'r') as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + id="fIcGw6MNNkSV"
# Encode
label_encoder = LabelEncoder()
label_encoder.fit(y)
num_classes = len(label_encoder)
# + id="pLzecFwWNkSV" colab={"base_uri": "https://localhost:8080/"} outputId="d7fb1bf6-92e8-46e0-da04-f5f6546b0733"
label_encoder.class_to_index
# + id="l8na7YazNkSW" colab={"base_uri": "https://localhost:8080/"} outputId="280c6d43-7972-4561-e90c-53edb6607942"
# Sample
label_encoder.encode([["attention", "data-augmentation"]])
# + id="iMyIbacNNkSW" colab={"base_uri": "https://localhost:8080/"} outputId="67cfb772-ead4-4520-b9cd-7c9f9bc4f270"
# Encode all our labels
y = label_encoder.encode(y)
print (y.shape)
# + [markdown] id="ufCNlDjQNkSW"
# ## Splitting
# + id="XOfLfpMKNxy3" colab={"base_uri": "https://localhost:8080/"} outputId="c5fe36fa-4dad-43d3-9b36-90e6a9781ce0"
# !pip install scikit-multilearn==0.2.0 -q
# + [markdown] id="BBRwUKtfNkSW"
# You need to [clean](https://madewithml.com/courses/applied-ml/preprocessing/) your data first before splitting, at least for the features that splitting depends on. So the process is more like: preprocessing (global, cleaning) → splitting → preprocessing (local, transformations). We're splitting using the tag labels which have already been inspected and cleaned during EDA.
# + [markdown] id="2XMXpueENkSW"
# **Naive split**
# + id="WEhp9SMFNkSW"
from sklearn.model_selection import train_test_split
from skmultilearn.model_selection.measures import get_combination_wise_output_matrix
# + id="TTUju11uNkSW"
# Split sizes
train_size = 0.7
val_size = 0.15
test_size = 0.15
# + [markdown] id="DTRQWabKNkSW"
# For simple multiclass classification, you can specify how to stratify the split by adding the [`stratify`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) keyword argument. But our task is multilabel classification, so we'll need to use other techniques to create even splits.
# + id="fw83_zitNkSW"
# Split (train)
X_train, X_, y_train, y_ = train_test_split(X, y, train_size=train_size)
# + id="WPVutu_dNkSW" colab={"base_uri": "https://localhost:8080/"} outputId="f673c17a-8df8-4643-9c84-381b786ac753"
print (f"train: {len(X_train)} ({(len(X_train) / len(X)):.2f})\n"
f"remaining: {len(X_)} ({(len(X_) / len(X)):.2f})")
# + id="u-CFrR7pNkSW"
# Split (test)
X_val, X_test, y_val, y_test = train_test_split(
X_, y_, train_size=0.5)
# + id="-evG0mc1NkSW" colab={"base_uri": "https://localhost:8080/"} outputId="577f4898-6be9-409e-f0f3-87edf33bf5ae"
print(f"train: {len(X_train)} ({len(X_train)/len(X):.2f})\n"
f"val: {len(X_val)} ({len(X_val)/len(X):.2f})\n"
f"test: {len(X_test)} ({len(X_test)/len(X):.2f})")
# + id="Jw7KBZxsNkSW"
# Get counts for each class
counts = {}
counts['train_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_train, order=1) for combination in row)
counts['val_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_val, order=1) for combination in row)
counts['test_counts'] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_test, order=1) for combination in row)
# + id="XsqdznxBNkSW" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="75bb126a-e9d5-4576-d655-713aa99d58be"
# View distributions
pd.DataFrame({
"train": counts["train_counts"],
"val": counts["val_counts"],
"test": counts["test_counts"]
}).T.fillna(0)
# + [markdown] id="_UIGPYZCNkSW"
# It's hard to compare these because our train and test proportions are different. Let's see what the distribution looks like once we balance it out. What do we need to multiply our test ratio by so that we have the same amount as our train ratio?
#
# $$ \alpha * N_{test} = N_{train} $$
#
# $$ \alpha = \frac{N_{train}}{N_{test}} $$
# + id="Nu1CRlQRNkSW"
# Adjust counts across splits
for k in counts["val_counts"].keys():
counts["val_counts"][k] = int(counts["val_counts"][k] * \
(train_size/val_size))
for k in counts["test_counts"].keys():
counts["test_counts"][k] = int(counts["test_counts"][k] * \
(train_size/test_size))
# + id="TRZcPnAkNkSW" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="864bb271-5db1-4dcf-f602-0f772b4a1920"
dist_df = pd.DataFrame({
"train": counts["train_counts"],
"val": counts["val_counts"],
"test": counts["test_counts"]
}).T.fillna(0)
dist_df
# + [markdown] id="o-XWskGRNkSW"
# We can see how much deviance there is in our naive data splits by computing the standard deviation of each split's class counts from the mean (ideal split).
#
# $ \sigma = \sqrt{\frac{(x - \bar{x})^2}{N}} $
# + id="jYL9nvKvNkSW" colab={"base_uri": "https://localhost:8080/"} outputId="51240f9c-c3af-413a-ac40-36e8178bc1af"
# Standard deviation
np.mean(np.std(dist_df.to_numpy(), axis=0))
# + [markdown] id="vorJjssYNkSW"
# Some of these distributions are not great. Let's try and balance this out a bit better.
# + [markdown] id="etOP2EetNkSW"
# **Stratified split**
# + [markdown] id="whQp6BPxvQaz"
# Now we'll apply [iterative stratification](http://lpis.csd.auth.gr/publications/sechidis-ecmlpkdd-2011.pdf) via the [skmultilearn](http://scikit.ml/index.html) library, which essentially splits each input into subsets (where each label is considered individually) and then it distributes the samples starting with fewest "positive" samples and working up to the inputs that have the most labels.
# + id="31BlT7xPNkSW"
from skmultilearn.model_selection import IterativeStratification
# + id="m1WZAQJGNkSW"
def iterative_train_test_split(X, y, train_size):
"""Custom iterative train test split which
'maintains balanced representation with respect
to order-th label combinations.'
"""
stratifier = IterativeStratification(
n_splits=2, order=1, sample_distribution_per_fold=[1.0-train_size, train_size, ])
train_indices, test_indices = next(stratifier.split(X, y))
X_train, y_train = X[train_indices], y[train_indices]
X_test, y_test = X[test_indices], y[test_indices]
return X_train, X_test, y_train, y_test
# + id="jghaS1edNkSW"
# Get data
X = df.text.to_numpy()
y = df.tags
# + id="Rv0baiseNkSW"
# Binarize y
label_encoder = LabelEncoder()
label_encoder.fit(y)
y = label_encoder.encode(y)
# + id="YMaDgwX7NkSW"
# Split
X_train, X_, y_train, y_ = iterative_train_test_split(
X, y, train_size=train_size)
X_val, X_test, y_val, y_test = iterative_train_test_split(
X_, y_, train_size=0.5)
# + id="qV1HlPP_NkSW" colab={"base_uri": "https://localhost:8080/"} outputId="086238e6-7491-4fa1-8125-f5ffd2ac99ae"
print(f"train: {len(X_train)} ({len(X_train)/len(X):.2f})\n"
f"val: {len(X_val)} ({len(X_val)/len(X):.2f})\n"
f"test: {len(X_test)} ({len(X_test)/len(X):.2f})")
# + id="QqjXL-lRNkSX"
# Get counts for each class
counts = {}
counts["train_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_train, order=1) for combination in row)
counts["val_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_val, order=1) for combination in row)
counts["test_counts"] = Counter(str(combination) for row in get_combination_wise_output_matrix(
y_test, order=1) for combination in row)
# + id="AKFGUgfkNkSX"
# Adjust counts across splits
for k in counts["val_counts"].keys():
counts["val_counts"][k] = int(counts["val_counts"][k] * \
(train_size/val_size))
for k in counts["test_counts"].keys():
counts["test_counts"][k] = int(counts["test_counts"][k] * \
(train_size/test_size))
# + id="Pn6kIt0HNkSX" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="3ee147bf-2a16-4780-cf97-ec3db677b3ca"
# View distributions
pd.DataFrame({
"train": counts["train_counts"],
"val": counts["val_counts"],
"test": counts["test_counts"]
}).T.fillna(0)
# + id="ftgmBEVdNkSX"
dist_df = pd.DataFrame({
'train': counts['train_counts'],
'val': counts['val_counts'],
'test': counts['test_counts']
}).T.fillna(0)
# + id="c02UhVufNkSX" colab={"base_uri": "https://localhost:8080/"} outputId="db142599-6ba4-4932-90d8-26c2cab26d03"
# Standard deviation
np.mean(np.std(dist_df.to_numpy(), axis=0))
# + [markdown] id="1GvMA5m0vWVr"
# > [Iterative stratification](http://scikit.ml/_modules/skmultilearn/model_selection/iterative_stratification.html#IterativeStratification) essentially creates splits while "trying to maintain balanced representation with respect to order-th label combinations". We used to an `order=1` for our iterative split which means we cared about providing representative distribution of each tag across the splits. But we can account for [higher-order](https://arxiv.org/abs/1704.08756) label relationships as well where we may care about the distribution of label combinations.
# + id="rf2b7aGVK_nb" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="8524eb26-86dd-4129-c6ae-7b93d37cd691"
# Split DataFrames
train_df = pd.DataFrame({"text": X_train, "tags": label_encoder.decode(y_train)})
val_df = pd.DataFrame({"text": X_val, "tags": label_encoder.decode(y_val)})
test_df = pd.DataFrame({"text": X_test, "tags": label_encoder.decode(y_test)})
train_df.head()
# + [markdown] id="ZsR6To1zNkSX"
# ## Augmentation
# + [markdown] id="BZQNks8TNkSX"
# We'll often want to increase the size and diversity of our training data split through data augmentation. It involves using the existing samples to generate synthetic, yet realistic, examples.
# + id="PR9WLz0jc3GO" colab={"base_uri": "https://localhost:8080/"} outputId="e19d65a5-7524-4454-f86b-a86d23220688"
# !pip install nlpaug==1.1.0 transformers==3.0.2 -q
# !pip install snorkel==0.9.7 -q
# + id="C6MX5Gm8NkSX"
import nlpaug.augmenter.word as naw
# + id="A9kUIKpsNkSX" colab={"base_uri": "https://localhost:8080/", "height": 113, "referenced_widgets": ["e886090b074c466eb8e0f14f5ef954ca", "747c2da3eb1947fda304f47049f4474b", "73d2d77130b24f3d81034d5f7b127c68", "55e5adaa7a4f4a0ab3da028ac952382d", "7aa6fb7eca57477895777072bd120515", "03ab4f108cab486e88fe8369aa8f0175", "c0d68d668f6747e1acc4fc50462eb7e9", "8e9c83977ac64258a9e5e263f1a6bc45", "cc05303ebebf4b65a46f53cec8db30b3", "<KEY>", "<KEY>", "<KEY>", "580299e4813a485ca6e203af3a6852fb", "<KEY>", "0e314fcf971c4aff995a2f3027db1526", "<KEY>", "<KEY>", "<KEY>", "dad5ee231ca442a69668edd577ffc7ab", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "3892e35b326c4c9782229a628f1a6381", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "25e3abe146a14a18ad5eacd19f4d79b3", "ba07fa7580784b0780c477249e345996", "<KEY>", "c79a85d893004d33823a2189a301e037", "47806adb2a27451b8dcaa398c33a46b3"]} outputId="2e2e620d-dcdb-4b82-aeda-929a4d89054e"
# Load tokenizers and transformers
substitution = naw.ContextualWordEmbsAug(model_path="distilbert-base-uncased", action="substitute")
insertion = naw.ContextualWordEmbsAug(model_path="distilbert-base-uncased", action="insert")
# + id="3YWn96hxNkSX"
text = "Conditional image generation using Variational Autoencoders and GANs."
# + id="NLt1kYXkg3L8" colab={"base_uri": "https://localhost:8080/"} outputId="22c442d4-7a51-4f98-a19a-93392e95d1f3"
# Substitutions
augmented_text = substitution.augment(text)
print (augmented_text)
# + [markdown] id="36DzpSqbNkSX"
# Substitution doesn't seem like a great idea for us because there are certain keywords that provide strong signal for our tags so we don't want to alter those. Also, note that these augmentations are NOT deterministic and will vary every time we run them. Let's try insertion...
# + id="bK8PV0FWb7pE" colab={"base_uri": "https://localhost:8080/"} outputId="72b09ea4-0583-4ccc-ceed-cff82e8f9847"
# Insertions
augmented_text = insertion.augment(text)
print (augmented_text)
# + [markdown] id="x5K29gM5NkSX"
# A little better but still quite fragile and now it can potentially insert key words that can influence false positive tags to appear. Maybe instead of substituting or inserting new tokens, let's try simply swapping machine learning related keywords with their aliases from our [auxiliary data](https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/datasets/tags.json). We'll use Snorkel's [transformation functions](https://www.snorkel.org/use-cases/02-spam-data-augmentation-tutorial) to easily achieve this.
# + id="-8EchR_16aIn"
import inflect
from snorkel.augmentation import transformation_function
inflect = inflect.engine()
# + id="LJoGgiKD_wog" colab={"base_uri": "https://localhost:8080/"} outputId="956cf939-9594-45de-eba1-4697848a07cb"
# Inflect
print (inflect.singular_noun("graphs"))
print (inflect.singular_noun("graph"))
print (inflect.plural_noun("graph"))
print (inflect.plural_noun("graphs"))
# + id="J-WP46Zs0rko"
def replace_dash(x):
return x.replace("-", " ")
# + id="YLComLRexHTN"
flat_tags_dict = {}
for tag, info in tags_dict.items():
tag = tag.replace("-", " ")
aliases = list(map(replace_dash, info["aliases"]))
if len(aliases):
flat_tags_dict[tag] = aliases
for alias in aliases:
_aliases = aliases + [tag]
_aliases.remove(alias)
flat_tags_dict[alias] = _aliases
# + id="8TkafOfS8Iqa"
# Tags that could be singular or plural
can_be_singular = [
'animations',
'cartoons',
'autoencoders',
'conditional random fields',
'convolutional neural networks',
'databases',
'deep q networks',
'gated recurrent units',
'gaussian processes',
'generative adversarial networks',
'graph convolutional networks',
'graph neural networks',
'k nearest neighbors',
'learning rates',
'multilayer perceptrons',
'outliers',
'pos',
'quasi recurrent neural networks',
'recommendation systems',
'recurrent neural networks',
'streaming data',
'data streams',
'support vector machines',
'variational autoencoders']
can_be_plural = [
'annotation',
'data annotation',
'continuous integration',
'continuous deployment',
'crf',
'conversational ai',
'chatbot',
'cnn',
'db',
'dqn',
'expectation maximization',
'fine tuning',
'finetuning',
'finetune',
'gru',
'gan',
'gcn',
'gnn',
'hyperparameter optimization',
'hyperparameter tuning',
'image generation',
'inference',
'prediction',
'knn',
'knowledge base',
'language modeling',
'latent dirichlet allocation',
'lstm',
'machine translation',
'model compression',
'compression',
'perceptron',
'mlp',
'optical character recognition',
'outlier detection',
'pos tagging',
'pca',
'qrnn',
'rnn',
'segmentation',
'image segmentation',
'spatial temporal cnn',
'data streaming',
'svm',
'tabular',
'temporal cnn',
'tcnn',
'vae',
'vqa',
'visualization',
'data visualization']
# + id="hKUwzfZc96gO"
# Add to flattened dict
for tag in can_be_singular:
flat_tags_dict[inflect.singular_noun(tag)] = flat_tags_dict[tag]
for tag in can_be_plural:
flat_tags_dict[inflect.plural_noun(tag)] = flat_tags_dict[tag]
# + id="KU6DGLk26RNR" colab={"base_uri": "https://localhost:8080/"} outputId="822d05be-1e98-40f9-9b2a-e0989873578e"
# Doesn't perfectly match (ex. singlar tag to singlar alias)
# But good enough for data augmentation for char-level tokenization
# Could've also used stemming before swapping aliases
print (flat_tags_dict["gan"])
print (flat_tags_dict["gans"])
print (flat_tags_dict["generative adversarial network"])
print (flat_tags_dict["generative adversarial networks"])
# + id="E2QdYOXSY55L" colab={"base_uri": "https://localhost:8080/"} outputId="ade27f2a-11d5-48d3-f8c5-06918bb99923"
# We want to match with the whole word only
print ("gan" in "This is a gan.")
print ("gan" in "This is gandalf.")
# + id="MHtHER09W8ew"
def find_word(word, text):
word = word.replace("+", "\+")
pattern = re.compile(fr"\b({word})\b", flags=re.IGNORECASE)
return pattern.search(text)
# + id="4zTuKPuRXTRO" colab={"base_uri": "https://localhost:8080/"} outputId="29580797-5b1d-4e46-ea54-f3eef1365cd8"
# Correct behavior (single instance)
print (find_word("gan", "This is a gan."))
print (find_word("gan", "This is gandalf."))
# + id="2r0f6iqeoNSh"
@transformation_function()
def swap_aliases(x):
"""Swap ML keywords with their aliases."""
# Find all matches
matches = []
for i, tag in enumerate(flat_tags_dict):
match = find_word(tag, x.text)
if match:
matches.append(match)
# Swap a random match with a random alias
if len(matches):
match = random.choice(matches)
tag = x.text[match.start():match.end()]
x.text = f"{x.text[:match.start()]}{random.choice(flat_tags_dict[tag])}{x.text[match.end():]}"
return x
# + id="4Hnuu4LZqgxS" colab={"base_uri": "https://localhost:8080/"} outputId="fc71fba3-de8a-4ca3-de46-9a8c6bffd660"
# Swap
for i in range(3):
sample_df = pd.DataFrame([{"text": "a survey of reinforcement learning for nlp tasks."}])
sample_df.text = sample_df.text.apply(preprocess, lower=True, stem=False)
print (swap_aliases(sample_df.iloc[0]).text)
# + id="Zb1PkpxE3jrP" colab={"base_uri": "https://localhost:8080/"} outputId="48234010-b77e-4bd7-bb83-39ade5aa32c0"
# Undesired behavior (needs contextual insight)
for i in range(3):
sample_df = pd.DataFrame([{"text": "Autogenerate your CV to apply for jobs using NLP."}])
sample_df.text = sample_df.text.apply(preprocess, lower=True, stem=False)
print (swap_aliases(sample_df.iloc[0]).text)
# + [markdown] id="7C0z19mFBswx"
# Now we'll define a [augmentation policy](https://snorkel.readthedocs.io/en/v0.9.1/packages/augmentation.html) to apply our transformation functions with certain rules (how many samples to generate, whether to keep the original data point, etc.)
# + id="8Cwju3KrnH7O"
from snorkel.augmentation import ApplyOnePolicy, PandasTFApplier
# + id="NitY8I8jlg-f" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="29593b93-e032-43f7-d16d-f6cd0bb75820"
# Transformation function (TF) policy
policy = ApplyOnePolicy(n_per_original=5, keep_original=True)
tf_applier = PandasTFApplier([swap_aliases], policy)
train_df_augmented = tf_applier.apply(train_df)
train_df_augmented.drop_duplicates(subset=["text"], inplace=True)
train_df_augmented.head()
# + id="UhCHE2eCnXf9" colab={"base_uri": "https://localhost:8080/"} outputId="54d3088b-72f4-4ff3-860f-ca857c7dde0d"
len(train_df), len(train_df_augmented)
# + [markdown] id="XAgm9USuoQUF"
# For now, we'll skip the data augmentation because it's quite fickle and empirically it doesn't improvement performance much. But we can see how this can be very effective once we can control what type of vocabulary to augment on and what exactly to augment with.
#
# > Regardless of what method we use, it's important to validate that we're not just augmenting for the sake of augmentation. We can do this by executing any existing [data validation tests](https://madewithml.com/courses/mlops/testing#data) and even creating specific tests to apply on augmented data.
# + [markdown] id="lGvI2YuuNkSX"
# # 📈 Modeling
# + [markdown] id="IdQjMSQgNkSX"
# We'll begin modeling by starting with the simplest baseline and slowly adding complexity.
# + id="C4O57X2dNkSY"
from sklearn.metrics import precision_recall_fscore_support
import torch
# + id="NXd8flJuNkSY"
def set_seeds(seed=1234):
"""Set seeds for reproducibility."""
np.random.seed(seed)
random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
torch.cuda.manual_seed_all(seed) # multi-GPU
# + id="jf_XCcSguF9o"
def get_data_splits(df, train_size=0.7):
""""""
# Get data
X = df.text.to_numpy()
y = df.tags
# Binarize y
label_encoder = LabelEncoder()
label_encoder.fit(y)
y = label_encoder.encode(y)
# Split
X_train, X_, y_train, y_ = iterative_train_test_split(
X, y, train_size=train_size)
X_val, X_test, y_val, y_test = iterative_train_test_split(
X_, y_, train_size=0.5)
return X_train, X_val, X_test, y_train, y_val, y_test, label_encoder
# + id="6Q9C95ZAhJHv"
class Trainer(object):
def __init__(self, model, device, loss_fn=None, optimizer=None, scheduler=None):
# Set params
self.model = model
self.device = device
self.loss_fn = loss_fn
self.optimizer = optimizer
self.scheduler = scheduler
def train_step(self, dataloader):
"""Train step."""
# Set model to train mode
self.model.train()
loss = 0.0
# Iterate over train batches
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, targets = batch[:-1], batch[-1]
self.optimizer.zero_grad() # Reset gradients
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, targets) # Define loss
J.backward() # Backward pass
self.optimizer.step() # Update weights
# Cumulative Metrics
loss += (J.detach().item() - loss) / (i + 1)
return loss
def eval_step(self, dataloader):
"""Validation or test step."""
# Set model to eval mode
self.model.eval()
loss = 0.0
y_trues, y_probs = [], []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, y_true = batch[:-1], batch[-1]
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, y_true).item()
# Cumulative Metrics
loss += (J - loss) / (i + 1)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
y_trues.extend(y_true.cpu().numpy())
return loss, np.vstack(y_trues), np.vstack(y_probs)
def predict_step(self, dataloader):
"""Prediction step."""
# Set model to eval mode
self.model.eval()
y_probs = []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Forward pass w/ inputs
inputs, targets = batch[:-1], batch[-1]
z = self.model(inputs)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
return np.vstack(y_probs)
def train(self, num_epochs, patience, train_dataloader, val_dataloader,
tolerance=1e-5):
best_val_loss = np.inf
for epoch in range(num_epochs):
# Steps
train_loss = self.train_step(dataloader=train_dataloader)
val_loss, _, _ = self.eval_step(dataloader=val_dataloader)
self.scheduler.step(val_loss)
# Early stopping
if val_loss < best_val_loss - tolerance:
best_val_loss = val_loss
best_model = self.model
_patience = patience # reset _patience
else:
_patience -= 1
if not _patience: # 0
print("Stopping early!")
break
# Logging
print(
f"Epoch: {epoch+1} | "
f"train_loss: {train_loss:.5f}, "
f"val_loss: {val_loss:.5f}, "
f"lr: {self.optimizer.param_groups[0]['lr']:.2E}, "
f"_patience: {_patience}"
)
return best_model
# + [markdown] id="fAQjRWHCNkSY"
# ## Random
# + [markdown] id="StAbvxEGkSmS"
# <u><i>motivation</i></u>: We want to know what random (chance) performance looks like. All of our efforts should be well above this.
# + id="Y6R-g-5INkSY"
# Set seeds
set_seeds()
# + id="SIYPLUMnNkSY"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
# + id="_UWsFo8cNkSY" colab={"base_uri": "https://localhost:8080/"} outputId="fc28f108-0b66-477c-8297-0c009c728a6b"
# Label encoder
print (label_encoder)
print (label_encoder.classes)
# + id="YXTrB3ToNkSY" colab={"base_uri": "https://localhost:8080/"} outputId="23fd053f-244b-493c-a7cf-fd1a0df4cb2b"
# Generate random predictions
y_pred = np.random.randint(low=0, high=2, size=(len(y_test), len(label_encoder.classes)))
print (y_pred.shape)
print (y_pred[0:5])
# + id="tAAoXyNANkSY" colab={"base_uri": "https://localhost:8080/"} outputId="79e80731-1f46-4531-bad5-2eaa54b78b96"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + [markdown] id="BLzOP5fIkYWl"
# We made the assumption that there is an equal probability for whether an input has a tag or not but this isn't true. Let's use the **train split** to figure out what the true probability is.
# + id="pN6tq9z6iW7a" colab={"base_uri": "https://localhost:8080/"} outputId="53435e58-d7f8-4451-8e6d-6421e0aee6ac"
# Percentage of 1s (tag presence)
tag_p = np.sum(np.sum(y_train)) / (len(y_train) * len(label_encoder.classes))
print (tag_p)
# + id="vInlz4R1iA4i"
# Generate weighted random predictions
y_pred = np.random.choice(
np.arange(0, 2), size=(len(y_test), len(label_encoder.classes)),
p=[1-tag_p, tag_p])
# + id="_LnLd-Arj5t0" colab={"base_uri": "https://localhost:8080/"} outputId="1468f0b7-2bf6-4620-9c8e-742b6368c973"
# Validate percentage
np.sum(np.sum(y_pred)) / (len(y_pred) * len(label_encoder.classes))
# + id="hpfBiruhkJME" colab={"base_uri": "https://localhost:8080/"} outputId="b1e0d301-79ce-432c-8883-347a859c1d31"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + [markdown] id="YC9ConMQlPgg"
# <u><i>limitations</i></u>: we didn't use the tokens in our input to affect our predictions so nothing was learned.
# + [markdown] id="OVzBKdt3NkSY"
# ## Rule-based
# + [markdown] id="u_XWLBgQlSn-"
# <u><i>motivation</i></u>: we want to use signals in our inputs (along with domain expertise and auxiliary data) to determine the labels.
# + id="nXlgWoyFNkSY"
# Set seeds
set_seeds()
# + [markdown] id="2RzG44f6mx_4"
# ### Unstemmed
# + id="ZhXtWZcgmxWW"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
# + id="3N961xapm1wS" colab={"base_uri": "https://localhost:8080/"} outputId="60a796d6-d70e-4802-fd8f-1fca46bd54c0"
# Restrict to relevant tags
print (len(tags_dict))
tags_dict = {tag: tags_dict[tag] for tag in label_encoder.classes}
print (len(tags_dict))
# + id="8sN9-ZfEm10i" colab={"base_uri": "https://localhost:8080/"} outputId="a78ea7c7-316d-4a70-99b3-d643f108fcd1"
# Map aliases
aliases = {}
for tag, values in tags_dict.items():
aliases[preprocess(tag)] = tag
for alias in values["aliases"]:
aliases[preprocess(alias)] = tag
aliases
# + id="ntVJvdVBm14U"
def get_classes(text, aliases, tags_dict):
"""If a token matches an alias,
then add the corresponding tag
class (and parent tags if any)."""
classes = []
for alias, tag in aliases.items():
if alias in text:
classes.append(tag)
for parent in tags_dict[tag]["parents"]:
classes.append(parent)
return list(set(classes))
# + id="cKHKKwNCnBzX" colab={"base_uri": "https://localhost:8080/"} outputId="181f0946-1153-4e6d-8c82-6cbd3f2c1d6c"
# Sample
text = "This project extends gans for data augmentation specifically for object detection tasks."
get_classes(text=preprocess(text), aliases=aliases, tags_dict=tags_dict)
# + id="LD-w5gcjnB29"
# Prediction
y_pred = []
for text in X_test:
classes = get_classes(text, aliases, tags_dict)
y_pred.append(classes)
# + id="YEcszZaGnZye"
# Encode labels
y_pred = label_encoder.encode(y_pred)
# + id="erYMwIffnB66" colab={"base_uri": "https://localhost:8080/"} outputId="f1a5b30b-f57a-436c-e568-2d3871e60a32"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + [markdown] id="bKbvhmgam2yG"
# ### Stemmed
# + [markdown] id="id2HJmqXnxPr"
# We're looking for exact matches with the aliases which isn't always perfect, for example:
# + id="dcr2o2HjoI_O" colab={"base_uri": "https://localhost:8080/"} outputId="03803b8c-b8ae-4ac3-b823-73effa1e1a29"
print (aliases[preprocess("gan")])
# print (aliases[preprocess("gans")]) # this won't find any match
print (aliases[preprocess("generative adversarial networks")])
# print (aliases[preprocess("generative adversarial network")]) # this won't find any match
# + [markdown] id="Gjv6GFJZoV6x"
# We don't want to keep adding explicit rules but we can use [stemming](https://nlp.stanford.edu/IR-book/html/htmledition/stemming-and-lemmatization-1.html) to represent different forms of a word uniformly, for example:
# + id="w8YcWdTqpagt" colab={"base_uri": "https://localhost:8080/"} outputId="1e00eeb1-024c-4da2-867b-570f186e55a3"
print (porter.stem("democracy"))
print (porter.stem("democracies"))
# + [markdown] id="EKCMUotL9wLr"
# So let's now stem our aliases as well as the tokens in our input text and then look for matches:
# + id="8BnfNEsf3YQB"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True, stem=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
# + id="3J4YQRZ_NkSY" colab={"base_uri": "https://localhost:8080/"} outputId="3c529340-dc4e-4d41-fc0a-52e4699c214a"
# Map aliases
aliases = {}
for tag, values in tags_dict.items():
aliases[preprocess(tag, stem=True)] = tag
for alias in values["aliases"]:
aliases[preprocess(alias, stem=True)] = tag
aliases
# + id="Qx-tmDpbNkSY" colab={"base_uri": "https://localhost:8080/"} outputId="dc5577c3-900f-45b1-8138-fadc1c9249dc"
# Checks (we will write proper tests soon)
print (aliases[preprocess("gan", stem=True)])
print (aliases[preprocess("gans", stem=True)])
print (aliases[preprocess("generative adversarial network", stem=True)])
print (aliases[preprocess("generative adversarial networks", stem=True)])
# + id="oXoeXojANkSY" colab={"base_uri": "https://localhost:8080/"} outputId="4dc71f1e-7972-491e-b919-c08252cfb51e"
# Sample
text = "This project extends gans for data augmentation specifically for object detection tasks."
get_classes(text=preprocess(text, stem=True), aliases=aliases, tags_dict=tags_dict)
# + id="5mw2GDgLNkSY"
# Prediction
y_pred = []
for text in X_test:
classes = get_classes(text, aliases, tags_dict)
y_pred.append(classes)
# + id="YcOkiIEXNkSY"
# Encode labels
y_pred = label_encoder.encode(y_pred)
# + id="S0UobHBcNkSY" colab={"base_uri": "https://localhost:8080/"} outputId="4f9f6e46-95db-484f-a1e9-3c1dc833aaa5"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + [markdown] id="8qdSlJCMfyBT"
# A nice improvement from the unstemmed approach! This is because rule-based approaches can yield labels with high certainty when there is an absolute condition match but it fails to generalize or learn implicit patterns.
# + [markdown] id="xjoBs0D8VI23"
# ### Inference
# + id="8j4iNdkfVLRX" colab={"base_uri": "https://localhost:8080/"} outputId="1217503e-8010-43af-8e94-f54543bdf01e"
# Infer
text = "Transfer learning with transformers for self-supervised learning"
print (preprocess(text, stem=True))
get_classes(text=preprocess(text, stem=True), aliases=aliases, tags_dict=tags_dict)
# + [markdown] id="VSvxZVVcFFMO"
# Now let's see what happens when we replace the word *transformers* with *BERT*. Sure we can add this as an alias but we can't keep doing this. This is where it makes sense to learn from the data as opposed to creating explicit rules.
# + id="k7cWn6Fgfxp2" colab={"base_uri": "https://localhost:8080/"} outputId="ae0668b0-fac4-498b-c452-dee864b63a37"
# Infer
text = "Transfer learning with BERT for self-supervised learning"
print (preprocess(text, stem=True))
get_classes(text=preprocess(text, stem=True), aliases=aliases, tags_dict=tags_dict)
# + [markdown] id="qdzlfoIzloqc"
# <u><i>limitations</i></u>: we failed to generalize or learn any implicit patterns to predict the labels because we treat the tokens in our input as isolated entities.
# + [markdown] id="q6rPM9Q8fX3-"
# > We would ideally spend more time tuning our model because it's so simple and quick to train. This approach also applies to all the other models we'll look at as well.
# + [markdown] id="eoa5oVgaNkSZ"
# ## Simple ML
# + [markdown] id="Yqg_kc3VmHb9"
# <u><i>motivation</i></u>:
# - *representation*: use term frequency-inverse document frequency [(TF-IDF)](https://en.wikipedia.org/wiki/Tf%E2%80%93idf) to capture the significance of a token to a particular input with respect to all the inputs, as opposed to treating the words in our input text as isolated tokens.
# - *architecture*: we want our model to meaningfully extract the encoded signal to predict the output labels.
# + [markdown] id="KOJW4AONNkSZ"
# So far we've treated the words in our input text as isolated tokens and we haven't really captured any meaning between tokens. Let's use term frequency–inverse document frequency (**TF-IDF**) to capture the significance of a token to a particular input with respect to all the inputs.
#
# $$ w_{i, j} = \text{tf}_{i, j} * log(\frac{N}{\text{df}_i}) $$
#
# $$ w_{i, j}: \text{tf-idf weight for term i in document j} $$
# $$ \text{tf}_{i, j}: \text{# of times term i appear in document j} $$
# $$ N: \text{total # of documents} $$
# $$ {\text{df}_i}: \text{# of documents with token i} $$
# + id="BiRzEi9iNkSZ"
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.multiclass import OneVsRestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import LinearSVC
# + id="J2P0kMvKNkSZ"
from sklearn import metrics
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import accuracy_score, precision_score, recall_score
from sklearn.metrics import precision_recall_curve
from sklearn.preprocessing import MultiLabelBinarizer
# + id="SrJ6fqmlNkSZ"
# Set seeds
set_seeds()
# + id="tIaiaykFNkSZ"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True, stem=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
# + id="ipggqVfzNkSZ" colab={"base_uri": "https://localhost:8080/"} outputId="8146ba87-bf4c-4428-844b-38027c0a9609"
# Tf-idf
vectorizer = TfidfVectorizer()
print (X_train[0])
X_train = vectorizer.fit_transform(X_train)
X_val = vectorizer.transform(X_val)
X_test = vectorizer.transform(X_test)
print (X_train.shape)
print (X_train[0]) # scipy.sparse.csr_matrix
# + id="fEShazTUNkSZ"
def fit_and_evaluate(model):
"""Fit and evaluate each model."""
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
return {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
# + id="3YN1FqKrNkSZ" colab={"base_uri": "https://localhost:8080/"} outputId="c4374ca5-f10a-4de2-b5cb-b2e5f3417968"
# Models
performance = {}
performance["logistic-regression"] = fit_and_evaluate(OneVsRestClassifier(
LogisticRegression(), n_jobs=1))
performance["k-nearest-neighbors"] = fit_and_evaluate(
KNeighborsClassifier())
performance["random-forest"] = fit_and_evaluate(
RandomForestClassifier(n_jobs=-1))
performance["gradient-boosting-machine"] = fit_and_evaluate(OneVsRestClassifier(
GradientBoostingClassifier()))
performance["support-vector-machine"] = fit_and_evaluate(OneVsRestClassifier(
LinearSVC(), n_jobs=-1))
print (json.dumps(performance, indent=2))
# + [markdown] id="gyv7n9ZKpXRu"
# <u><i>limitations</i></u>:
# - *representation*: TF-IDF representations don't encapsulate much signal beyond frequency but we require more fine-grained token representations.
# - *architecture*: we want to develop models that can use better represented encodings in a more contextual manner.
# + [markdown] id="DW_93sOnNkSZ"
# ## CNN w/ Embeddings
# + [markdown] id="8jERhoo07l0b"
# <u><i>motivation</i></u>:
# - *representation*: we want to have more robust (split tokens to characters) and meaningful ([embeddings](https://madewithml.com/courses/basics/embeddings/) representations for our input tokens.
# - *architecture*: we want to process our encoded inputs using [convolution (CNN)](https://madewithml.com/courses/basics/convolutional-neural-networks/) filters that can learn to analyze windows of embedded tokens to extract meaningful signal (n-gram feature extractors).
# + [markdown] id="0XD7VH1MNkSZ"
# ### Set up
# + id="-8X3qlEbNkSZ"
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
# + id="tLn8Z4AINkSZ"
# Set seeds
set_seeds()
# + id="DbSoGGmPNkSZ"
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
X_test_raw = X_test # use for later
# + id="OREUyt-kvMWR"
# Split DataFrames
train_df = pd.DataFrame({"text": X_train, "tags": label_encoder.decode(y_train)})
val_df = pd.DataFrame({"text": X_val, "tags": label_encoder.decode(y_val)})
test_df = pd.DataFrame({"text": X_test, "tags": label_encoder.decode(y_test)})
# + id="oT0xmkCfNuy5" colab={"base_uri": "https://localhost:8080/"} outputId="edc4477d-732d-4158-8516-0b12216e551f"
# Set device
cuda = True
device = torch.device("cuda" if (
torch.cuda.is_available() and cuda) else "cpu")
torch.set_default_tensor_type("torch.FloatTensor")
if device.type == "cuda":
torch.set_default_tensor_type("torch.cuda.FloatTensor")
print (device)
# + [markdown] id="Q6r8kfAkNkSZ"
# ### Tokenizer
# + [markdown] id="Rg0IROLfJY_3"
# We're going to tokenize our input text as character tokens so we can be robust to spelling errors and learn to generalize across tags. (ex. learning that RoBERTa, or any other future BERT based archiecture, warrants same tag as BERT).
# + [markdown] id="krnCpNE6qJvq"
# <img width="500px" src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/foundations/cnn/inputs.png">
# + id="huCZn1stNkSZ"
class Tokenizer(object):
def __init__(self, char_level, num_tokens=None,
pad_token="<PAD>", oov_token="<UNK>",
token_to_index=None):
self.char_level = char_level
self.separator = '' if self.char_level else ' '
if num_tokens: num_tokens -= 2 # pad + unk tokens
self.num_tokens = num_tokens
self.pad_token = pad_token
self.oov_token = oov_token
if not token_to_index:
token_to_index = {pad_token: 0, oov_token: 1}
self.token_to_index = token_to_index
self.index_to_token = {v: k for k, v in self.token_to_index.items()}
def __len__(self):
return len(self.token_to_index)
def __str__(self):
return f"<Tokenizer(num_tokens={len(self)})>"
def fit_on_texts(self, texts):
if not self.char_level:
texts = [text.split(" ") for text in texts]
all_tokens = [token for text in texts for token in text]
counts = Counter(all_tokens).most_common(self.num_tokens)
self.min_token_freq = counts[-1][1]
for token, count in counts:
index = len(self)
self.token_to_index[token] = index
self.index_to_token[index] = token
return self
def texts_to_sequences(self, texts):
sequences = []
for text in texts:
if not self.char_level:
text = text.split(' ')
sequence = []
for token in text:
sequence.append(self.token_to_index.get(
token, self.token_to_index[self.oov_token]))
sequences.append(np.asarray(sequence))
return sequences
def sequences_to_texts(self, sequences):
texts = []
for sequence in sequences:
text = []
for index in sequence:
text.append(self.index_to_token.get(index, self.oov_token))
texts.append(self.separator.join([token for token in text]))
return texts
def save(self, fp):
with open(fp, "w") as fp:
contents = {
"char_level": self.char_level,
"oov_token": self.oov_token,
"token_to_index": self.token_to_index
}
json.dump(contents, fp, indent=4, sort_keys=False)
@classmethod
def load(cls, fp):
with open(fp, "r") as fp:
kwargs = json.load(fp=fp)
return cls(**kwargs)
# + id="ibPSo_PBNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="c0605cde-e360-40ce-efbb-7b6caf1f75aa"
# Tokenize
char_level = True
tokenizer = Tokenizer(char_level=char_level)
tokenizer.fit_on_texts(texts=X_train)
vocab_size = len(tokenizer)
print (tokenizer)
# + id="84JQQqJrNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="f4ca2512-1f39-4ede-e0d1-b3ef11376d25"
tokenizer.token_to_index
# + id="tqFL_J_UNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="093db965-8d78-4998-9062-5d90f544902c"
# Convert texts to sequences of indices
X_train = np.array(tokenizer.texts_to_sequences(X_train))
X_val = np.array(tokenizer.texts_to_sequences(X_val))
X_test = np.array(tokenizer.texts_to_sequences(X_test))
preprocessed_text = tokenizer.sequences_to_texts([X_train[0]])[0]
print ("Text to indices:\n"
f" (preprocessed) → {preprocessed_text}\n"
f" (tokenized) → {X_train[0]}")
# + [markdown] id="bUrzrb06NkSa"
# ### Data imbalance
# + [markdown] id="WUiU_-XVNkSa"
# We'll factor class weights in our objective function ([binary cross entropy with logits](https://pytorch.org/docs/stable/generated/torch.nn.BCEWithLogitsLoss.html)) to help with class imbalance. There are many other techniques such as over sampling from underrepresented classes, undersampling, etc. but we'll cover these in a separate unit lesson on data imbalance.
# + id="ETiBnH_JNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="c181ff94-efbb-4ba2-e87c-4012600c2363"
# Class weights
train_tags = list(itertools.chain.from_iterable(train_df.tags.values))
counts = np.bincount([label_encoder.class_to_index[class_] for class_ in train_tags])
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
print (f"class counts: {counts},\nclass weights: {class_weights}")
# + [markdown] id="UscX0dcrNkSa"
# ### Datasets
# + [markdown] id="g02qQyaoJ9YT"
# We're going to place our data into a [`Dataset`](https://pytorch.org/docs/stable/data.html#torch.utils.data.Dataset) and use a [`DataLoader`](https://pytorch.org/docs/stable/data.html#torch.utils.data.DataLoader) to efficiently create batches for training and evaluation.
# + id="BhSPgWsvNkSa"
def pad_sequences(sequences, max_seq_len=0):
"""Pad sequences to max length in sequence."""
max_seq_len = max(max_seq_len, max(len(sequence) for sequence in sequences))
padded_sequences = np.zeros((len(sequences), max_seq_len))
for i, sequence in enumerate(sequences):
padded_sequences[i][:len(sequence)] = sequence
return padded_sequences
# + id="yGoNL0oaNkSa"
class CNNTextDataset(torch.utils.data.Dataset):
def __init__(self, X, y, max_filter_size):
self.X = X
self.y = y
self.max_filter_size = max_filter_size
def __len__(self):
return len(self.y)
def __str__(self):
return f"<Dataset(N={len(self)})>"
def __getitem__(self, index):
X = self.X[index]
y = self.y[index]
return [X, y]
def collate_fn(self, batch):
"""Processing on a batch."""
# Get inputs
batch = np.array(batch, dtype=object)
X = batch[:, 0]
y = np.stack(batch[:, 1], axis=0)
# Pad inputs
X = pad_sequences(sequences=X, max_seq_len=self.max_filter_size)
# Cast
X = torch.LongTensor(X.astype(np.int32))
y = torch.FloatTensor(y.astype(np.int32))
return X, y
def create_dataloader(self, batch_size, shuffle=False, drop_last=False):
return torch.utils.data.DataLoader(
dataset=self,
batch_size=batch_size,
collate_fn=self.collate_fn,
shuffle=shuffle,
drop_last=drop_last,
pin_memory=True)
# + id="AcsLQ-xcNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="ef9502f0-6f0a-4767-93e5-e0e2b161a4df"
# Create datasets
filter_sizes = list(range(1, 11))
train_dataset = CNNTextDataset(
X=X_train, y=y_train, max_filter_size=max(filter_sizes))
val_dataset = CNNTextDataset(
X=X_val, y=y_val, max_filter_size=max(filter_sizes))
test_dataset = CNNTextDataset(
X=X_test, y=y_test, max_filter_size=max(filter_sizes))
print ("Data splits:\n"
f" Train dataset:{train_dataset.__str__()}\n"
f" Val dataset: {val_dataset.__str__()}\n"
f" Test dataset: {test_dataset.__str__()}\n"
"Sample point:\n"
f" X: {train_dataset[0][0]}\n"
f" y: {train_dataset[0][1]}")
# + id="CUkm-47FNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="3739e916-df5d-44fa-c0eb-d76fd6447b00"
# Create dataloaders
batch_size = 128
train_dataloader = train_dataset.create_dataloader(
batch_size=batch_size)
val_dataloader = val_dataset.create_dataloader(
batch_size=batch_size)
test_dataloader = test_dataset.create_dataloader(
batch_size=batch_size)
batch_X, batch_y = next(iter(train_dataloader))
print ("Sample batch:\n"
f" X: {list(batch_X.size())}\n"
f" y: {list(batch_y.size())}")
# + [markdown] id="O8UyFr9xNkSa"
# ### Model
# + [markdown] id="9HSHRnFmKg5g"
# We'll be using a convolutional neural network on top of our embedded tokens to extract meaningful spatial signal. This time, we'll be using many filter widths to act as n-gram feature extractors. If you're not familiar with CNNs be sure to check out the [CNN lesson](https://madewithml.com/courses/basics/convolutional-neural-networks/) where we walkthrough every component of the architecture.
# + [markdown] id="HJdHiBArqaaH"
# <img width="500px" src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/foundations/cnn/convolution.gif">
# + [markdown] id="fs2AeywNL3yt"
# Let's visualize the model's forward pass.
#
# 1. We'll first tokenize our inputs (`batch_size`, `max_seq_len`).
# 2. Then we'll embed our tokenized inputs (`batch_size`, `max_seq_len`, `embedding_dim`).
# 3. We'll apply convolution via filters (`filter_size`, `vocab_size`, `num_filters`) followed by batch normalization. Our filters act as character level n-gram detecors. We have three different filter sizes (2, 3 and 4) and they will act as bi-gram, tri-gram and 4-gram feature extractors, respectivelyy.
# 4. We'll apply 1D global max pooling which will extract the most relevant information from the feature maps for making the decision.
# 5. We feed the pool outputs to a fully-connected (FC) layer (with dropout).
# 6. We use one more FC layer with softmax to derive class probabilities.
# + [markdown] id="xHRtownxL6X1"
# <img width="5000px" src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/foundations/cnn/model.png">
# + id="ZhteynQANkSa"
# Arguments
embedding_dim = 128
num_filters = 128
hidden_dim = 128
dropout_p = 0.5
# + id="UWe7ktNMNkSa"
class CNN(nn.Module):
def __init__(self, embedding_dim, vocab_size, num_filters, filter_sizes,
hidden_dim, dropout_p, num_classes, padding_idx=0):
super(CNN, self).__init__()
# Initialize embeddings
self.embeddings = nn.Embedding(
embedding_dim=embedding_dim, num_embeddings=vocab_size,
padding_idx=padding_idx)
# Conv weights
self.filter_sizes = filter_sizes
self.conv = nn.ModuleList(
[nn.Conv1d(in_channels=embedding_dim,
out_channels=num_filters,
kernel_size=f) for f in filter_sizes])
# FC weights
self.dropout = nn.Dropout(dropout_p)
self.fc1 = nn.Linear(num_filters*len(filter_sizes), hidden_dim)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, inputs, channel_first=False):
# Embed
x_in, = inputs
x_in = self.embeddings(x_in)
if not channel_first:
x_in = x_in.transpose(1, 2) # (N, channels, sequence length)
z = []
max_seq_len = x_in.shape[2]
for i, f in enumerate(self.filter_sizes):
# `SAME` padding
padding_left = int(
(self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2)
padding_right = int(math.ceil(
(self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2))
# Conv
_z = self.conv[i](F.pad(x_in, (padding_left, padding_right)))
# Pool
_z = F.max_pool1d(_z, _z.size(2)).squeeze(2)
z.append(_z)
# Concat outputs
z = torch.cat(z, 1)
# FC
z = self.fc1(z)
z = self.dropout(z)
z = self.fc2(z)
return z
# + [markdown] id="ugjRPCGClmCc"
# Padding types:
# * **VALID**: no padding, the filters only use the "valid" values in the input. If the filter cannot reach all the input values (filters go left to right), the extra values on the right are dropped.
# * **SAME**: adds padding evenly to the right (preferred) and left sides of the input so that all values in the input are processed.
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/foundations/cnn/padding.png" width="500">
# </div>
# + [markdown] id="oXZ6ZD6AKbfr"
# We're add padding so that the convolutional outputs are the same shape as our inputs. The amount of padding for the `SAME` padding can be determined using the same equation. We want out output to have the same width as our input, so we solve for P:
#
# $ \frac{W-F+2P}{S} + 1 = W $
#
# $ P = \frac{S(W-1) - W + F}{2} $
#
# If $P$ is not a whole number, we round up (using `math.ceil`) and place the extra padding on the right side.
# + id="suLpR5raNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="23ce48c4-42dd-42a2-d216-445900387b99"
# Initialize model
model = CNN(
embedding_dim=embedding_dim, vocab_size=vocab_size,
num_filters=num_filters, filter_sizes=filter_sizes,
hidden_dim=hidden_dim, dropout_p=dropout_p,
num_classes=num_classes)
model = model.to(device)
print (model.named_parameters)
# + [markdown] id="AfcCItXQNkSa"
# ### Training
# + id="6XYetoGvNkSa"
# Arguments
lr = 2e-4
num_epochs = 100
patience = 10
# + id="gmPoArUeNkSa"
# Define loss
class_weights_tensor = torch.Tensor(np.array(list(class_weights.values())))
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
# + id="YfP6jpAsNkSa"
# Define optimizer & scheduler
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode="min", factor=0.1, patience=5)
# + id="36OkuM21NkSa"
# Trainer module
trainer = Trainer(
model=model, device=device, loss_fn=loss_fn,
optimizer=optimizer, scheduler=scheduler)
# + id="XvmUpxZRNkSa" colab={"base_uri": "https://localhost:8080/"} outputId="525b6f30-d827-4ccc-a2d0-913d4212e7e4"
# Train
best_model = trainer.train(
num_epochs, patience, train_dataloader, val_dataloader)
# + [markdown] id="nggIvJFoNkSa"
# ### Evaluation
# + id="0igBVFnYNkSb"
from pathlib import Path
from sklearn.metrics import precision_recall_curve
# + id="WPA5ojyUNkSb" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="35fa0f97-cce1-479a-e2d2-468898ae86ff"
# Threshold-PR curve
train_loss, y_true, y_prob = trainer.eval_step(dataloader=train_dataloader)
precisions, recalls, thresholds = precision_recall_curve(y_true.ravel(), y_prob.ravel())
plt.plot(thresholds, precisions[:-1], "r--", label="Precision")
plt.plot(thresholds, recalls[:-1], "b-", label="Recall")
plt.ylabel("Performance")
plt.xlabel("Threshold")
plt.legend(loc="best")
# + id="7bF16I1WNkSb"
# Determining the best threshold
def find_best_threshold(y_true, y_prob):
"""Find the best threshold for maximum F1."""
precisions, recalls, thresholds = precision_recall_curve(y_true, y_prob)
f1s = (2 * precisions * recalls) / (precisions + recalls)
return thresholds[np.argmax(f1s)]
# + [markdown] id="VlqdPlplaNCG"
# > Even better to determine per-class thresholds but this is fine for now.
# + id="Pv9rdHebNkSb" colab={"base_uri": "https://localhost:8080/"} outputId="933ad945-2682-4a66-a1d3-107fb875d094"
# Best threshold for f1
threshold = find_best_threshold(y_true.ravel(), y_prob.ravel())
threshold
# + id="N0i2EYMYNkSb"
# Determine predictions using threshold
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# + id="jP9rWb-9NkSb" colab={"base_uri": "https://localhost:8080/"} outputId="3a75e533-0819-4d4a-e19c-9f48f12fb209"
# Evaluate
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
print (json.dumps(performance, indent=2))
# + id="ewsYvUnxNkSb"
# Save artifacts
dir = Path("cnn")
dir.mkdir(parents=True, exist_ok=True)
tokenizer.save(fp=Path(dir, "tokenzier.json"))
label_encoder.save(fp=Path(dir, "label_encoder.json"))
torch.save(best_model.state_dict(), Path(dir, "model.pt"))
with open(Path(dir, "performance.json"), "w") as fp:
json.dump(performance, indent=2, sort_keys=False, fp=fp)
# + [markdown] id="C4tJvSbaNkSb"
# ### Inference
# + id="GwI4ft7CNkSb" colab={"base_uri": "https://localhost:8080/"} outputId="c9e059fb-5244-41cd-9a2f-b1de001c3e1d"
# Load artifacts
device = torch.device("cpu")
tokenizer = Tokenizer.load(fp=Path(dir, "tokenzier.json"))
label_encoder = LabelEncoder.load(fp=Path(dir, "label_encoder.json"))
model = CNN(
embedding_dim=embedding_dim, vocab_size=vocab_size,
num_filters=num_filters, filter_sizes=filter_sizes,
hidden_dim=hidden_dim, dropout_p=dropout_p,
num_classes=num_classes)
model.load_state_dict(torch.load(Path(dir, "model.pt"), map_location=device))
model.to(device)
# + id="sLeXU5LRYaal"
# Initialize trainer
trainer = Trainer(model=model, device=device)
# + id="b_SGtypzNkSb"
# Dataloader
text = "Transfer learning with BERT for self-supervised learning"
X = np.array(tokenizer.texts_to_sequences([preprocess(text)]))
y_filler = label_encoder.encode([np.array([label_encoder.classes[0]]*len(X))])
dataset = CNNTextDataset(
X=X, y=y_filler, max_filter_size=max(filter_sizes))
dataloader = dataset.create_dataloader(
batch_size=batch_size)
# + id="YRETV9QyNkSb" colab={"base_uri": "https://localhost:8080/"} outputId="f5d39ea6-23c6-43e6-95ac-f8d0668f2f9e"
# Inference
y_prob = trainer.predict_step(dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
label_encoder.decode(y_pred)
# + [markdown] id="oYBoIl1Q8ZGB"
# <u><i>limitations</i></u>:
# - *representation*: embeddings are not contextual.
# - *architecture*: extracting signal from encoded inputs is limited by filter widths.
# + [markdown] id="pWYOq-WNGaql"
# Since we're dealing with simple architectures and fast training times, it's a good opportunity to explore tuning and experiment with k-fold cross validation to properly reach any conclusions about performance.
# + [markdown] id="BBZ35y4tnB1h"
# ## Tradeoffs
# + [markdown] id="6oCp9vPrTUMx"
# We could experiment with more complex architectures such as [Transformers](https://madewithml.com/cousres/foundations/transformers), but we're going to go with the embeddings via CNN approach and optimize it because it offer decent performance at a reasonable tradeoff values (size, training time, etc.)
# + id="dVzPrFK0Y0b-" colab={"base_uri": "https://localhost:8080/"} outputId="78b91c59-289c-4df0-f519-193ec5d8aabb"
# Performance
with open(Path("cnn", "performance.json"), "r") as fp:
cnn_performance = json.load(fp)
print (f'CNN: f1 = {cnn_performance["f1"]}')
# + [markdown] id="ok2jBSjGsxyL"
# This was just one run on one split so you'll want to experiment with k-fold cross validation to properly reach any conclusions about performance. Also make sure you take the time to tune these baselines since their training periods are quite fast (we can achieve f1 of 0.7 with just a bit of tuning for both CNN / Transformers). We'll cover hyperparameter tuning in a few lessons so you can replicate the process here on your own time. We should also benchmark on other important metrics as we iterate, not just precision and recall.
# + id="oSM_UOkrVV6D" colab={"base_uri": "https://localhost:8080/"} outputId="05f9c814-acfe-4b6d-fae2-2cea780db9b3"
# Size
print (f'CNN: {Path("cnn", "model.pt").stat().st_size/1000000:.1f} MB')
# + [markdown] id="kDcnWxoMup6W"
# > We'll consider other tradeoffs such as maintenance overhead, behavioral test performances, etc. as we develop.
# + id="Xs1g4GW66ITQ"
# Arguments
embedding_dim = 128
num_filters = 128
hidden_dim = 128
dropout_p = 0.5
# + id="VIRktG-n49UI" colab={"base_uri": "https://localhost:8080/"} outputId="faca35de-102c-4d1a-c8a9-4c6979498cdd"
# Load artifacts
dir = Path("cnn")
device = torch.device("cpu")
tokenizer = Tokenizer.load(fp=Path(dir, "tokenzier.json"))
label_encoder = LabelEncoder.load(fp=Path(dir, "label_encoder.json"))
model = CNN(
embedding_dim=embedding_dim, vocab_size=len(tokenizer),
num_filters=num_filters, filter_sizes=filter_sizes,
hidden_dim=hidden_dim, dropout_p=dropout_p,
num_classes=len(label_encoder))
model.load_state_dict(torch.load(Path(dir, "model.pt"), map_location=device))
model.to(device)
# + id="hOBOLE8J6zV5"
# Trainer module
trainer = Trainer(model=model, device=device)
# + [markdown] id="9ofaM94omwgY"
# ## Evaluation
# + [markdown] id="nsj8_EUEmynv"
# So far we've been evaluating our models by determing the overall precision, recall and f1 scores. But since performance is one of the key decision making factors when comparing different models, we should have even more nuanced evaluation strategies.
#
# - Overall metrics
# - Per-class (tag) metrics
# - Confusion matrix sample analysis
# - Slice metrics
# + id="h51AAn1Fu4b5"
# Metrics
metrics = {"overall": {}, "class": {}}
# + id="H8BgzzHBZNMn"
# Data to evaluate
device = torch.device("cuda")
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
trainer = Trainer(model=model.to(device), device=device, loss_fn=loss_fn)
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# + [markdown] id="TiXcls5JoNA8"
# ### Overall metrics
# + id="h2OQtNODrh6c" colab={"base_uri": "https://localhost:8080/"} outputId="467bfbb3-024f-4c51-a8f4-4f85a96f78c6"
# Overall metrics
overall_metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
metrics["overall"]["precision"] = overall_metrics[0]
metrics["overall"]["recall"] = overall_metrics[1]
metrics["overall"]["f1"] = overall_metrics[2]
metrics["overall"]["num_samples"] = np.float64(len(y_true))
print (json.dumps(metrics["overall"], indent=4))
# + [markdown] id="zl3xSuXRutKG"
# ### Per-class metrics
# + id="1zIAI4mwusoX"
# Per-class metrics
class_metrics = precision_recall_fscore_support(y_test, y_pred, average=None)
for i, _class in enumerate(label_encoder.classes):
metrics["class"][_class] = {
"precision": class_metrics[0][i],
"recall": class_metrics[1][i],
"f1": class_metrics[2][i],
"num_samples": np.float64(class_metrics[3][i]),
}
# + id="Rhh-tgpP0dvj" colab={"base_uri": "https://localhost:8080/"} outputId="3e75dad6-8b94-4222-db68-cef968785a78"
# Metrics for a specific class
tag = "transformers"
print (json.dumps(metrics["class"][tag], indent=2))
# + [markdown] id="4RsmVddrmknD"
# As a general rule, the classes with fewer samples will have lower performance so we should always work to identify the class (or fine-grained slices) of data that our model needs to see more samples of to learn from.
# + id="5gXY4M5rcQ4H"
# Number of training samples per class
num_samples = np.sum(y_train, axis=0).tolist()
# + id="r0gIKVbMrkgo"
# Number of samples vs. performance (per class)
f1s = [metrics["class"][_class]["f1"]*100. for _class in label_encoder.classes]
sorted_lists = sorted(zip(*[num_samples, f1s])) # sort
num_samples, f1s = list(zip(*sorted_lists))
# + id="vQVA6G-j__t5" colab={"base_uri": "https://localhost:8080/", "height": 339} outputId="4041b85b-e112-4d0e-aee2-543589cef0ec"
# Plot
n = 7 # num. top classes to label
fig, ax = plt.subplots()
ax.set_xlabel("# of training samples")
ax.set_ylabel("test performance (f1)")
fig.set_size_inches(25, 5)
ax.plot(num_samples, f1s, "bo-")
for x, y, label in zip(num_samples[-n:], f1s[-n:], label_encoder.classes[-n:]):
ax.annotate(label, xy=(x,y), xytext=(-5, 5), ha="right", textcoords="offset points")
# + [markdown] id="G4LFphxErqFa"
# There are, of course, nuances to this general rule such as the complexity of distinguishing between some classes where we may not need as many samples for easier sub-tasks. In our case, classes with over 100 training samples consistently perform better than 0.6 f1 score, whereas the other class' performances are mixed.
# + [markdown] id="f-juex26zvBF"
# ### Confusion matric sample analysis
# + [markdown] id="xPUao0S4k99c"
# - True positives: learn about where our model performs well.
# - False positives: potentially identify samples which may need to be relabeled.
# - False negatives: identify the model's less performant areas to oversample later.
#
# > It's a good to have our FP/FN samples feed back into our annotation pipelines in the event we want to fix their labels and have those changes be reflected everywhere.
# + id="ZG2SgsPAzukL"
# TP, FP, FN samples
index = label_encoder.class_to_index[tag]
tp, fp, fn = [], [], []
for i in range(len(y_test)):
true = y_test[i][index]
pred = y_pred[i][index]
if true and pred:
tp.append(i)
elif not true and pred:
fp.append(i)
elif true and not pred:
fn.append(i)
# + id="ePrxeVkG0mmO" colab={"base_uri": "https://localhost:8080/"} outputId="d123165c-1fdd-4b9d-d75c-f9ddc8381d92"
print (tp)
print (fp)
print (fn)
# + id="-9UCXV7K0ocX" colab={"base_uri": "https://localhost:8080/"} outputId="9818ff73-155b-4b4f-ba1a-f1b165afb340"
index = tp[0]
print (X_test_raw[index])
print (f"true: {label_encoder.decode([y_test[index]])[0]}")
print (f"pred: {label_encoder.decode([y_pred[index]])[0]}\n")
# + id="tVbuLfiH0ofe"
# Sorted tags
sorted_tags_by_f1 = OrderedDict(sorted(
metrics["class"].items(), key=lambda tag: tag[1]["f1"], reverse=True))
# + id="n4jLEJ6f0ojA" colab={"base_uri": "https://localhost:8080/", "height": 831, "referenced_widgets": ["71c8c4424a7d4ac99dd8c198b4804558", "3f0e873fc4a043eabce5603462e9f5a1", "4cbf6da0bdff4f9f874227d2fa9709ee", "f7a3a46903124946a0c54b6c37c3dc1d", "2b82c7e02f994bc196162c462b5dfdbd", "2d557aaa2185415b97e1f00047b2235f", "f95efc06318e4ac8889af520c1ff06c8"]} outputId="68506454-9ed8-409a-ec1d-8a3f0e9c5c3e"
@widgets.interact(tag=list(sorted_tags_by_f1.keys()))
def display_tag_analysis(tag='transformers'):
# Performance
print (json.dumps(metrics["class"][tag], indent=2))
# TP, FP, FN samples
index = label_encoder.class_to_index[tag]
tp, fp, fn = [], [], []
for i in range(len(y_test)):
true = y_test[i][index]
pred = y_pred[i][index]
if true and pred:
tp.append(i)
elif not true and pred:
fp.append(i)
elif true and not pred:
fn.append(i)
# Samples
num_samples = 3
if len(tp):
print ("\n=== True positives ===")
for i in tp[:num_samples]:
print (f" {X_test_raw[i]}")
print (f" true: {label_encoder.decode([y_test[i]])[0]}")
print (f" pred: {label_encoder.decode([y_pred[i]])[0]}\n")
if len(fp):
print ("=== False positives === ")
for i in fp[:num_samples]:
print (f" {X_test_raw[i]}")
print (f" true: {label_encoder.decode([y_test[i]])[0]}")
print (f" pred: {label_encoder.decode([y_pred[i]])[0]}\n")
if len(fn):
print ("=== False negatives ===")
for i in fn[:num_samples]:
print (f" {X_test_raw[i]}")
print (f" true: {label_encoder.decode([y_test[i]])[0]}")
print (f" pred: {label_encoder.decode([y_pred[i]])[0]}\n")
# + [markdown] id="4S9XH4j4wSlS"
# > It's a really good idea to do this kind of analysis using our rule-based approach to catch really obvious labeling errors.
# + [markdown] id="dvS3UpusXP_R"
# ### Slice metrics
# + [markdown] id="eeWWMG38Ny4U"
# Evaluate performance on key slices of data that goes beyond class-level metrics.
# + id="ZyueOtQsXdGm"
from snorkel.slicing import PandasSFApplier
from snorkel.slicing import slice_dataframe
from snorkel.slicing import slicing_function
# + id="coutP2KtXdLG"
@slicing_function()
def pytorch_transformers(x):
"""Projects with the `pytorch` and `transformers` tags."""
return all(tag in x.tags for tag in ["pytorch", "transformers"])
# + id="zNekudM4XfBE"
@slicing_function()
def short_text(x):
"""Projects with short titles and descriptions."""
return len(x.text.split()) < 7 # less than 7 words
# + id="B7jmdmNaXuA2" colab={"base_uri": "https://localhost:8080/", "height": 190} outputId="068f9572-4aa8-41b2-9b23-449a4cfc1de2"
short_text_df = slice_dataframe(test_df, short_text)
short_text_df[["text", "tags"]].head()
# + [markdown] id="kZuDZwTNO93Q"
# We can define even more slicing functions and create a slices record array using the [`PandasSFApplier`](https://snorkel.readthedocs.io/en/v0.9.6/packages/_autosummary/slicing/snorkel.slicing.PandasSFApplier.html). The slices array has N (# of data points) items and each item has S (# of slicing functions) items, indicating whether that data point is part of that slice. Think of this record array as a masking layer for each slicing function on our data.
# + id="mQG8PFovXfEm" colab={"base_uri": "https://localhost:8080/"} outputId="b9ab355a-e90b-4b62-e024-f4eb86681f87"
# Slices
slicing_functions = [pytorch_transformers, short_text]
applier = PandasSFApplier(slicing_functions)
slices = applier.apply(test_df)
slices
# + [markdown] id="tCltCyWwYIvC"
# If our task was multiclass instead of multilabel, we could've used [snorkel.analysis.Scorer](https://snorkel.readthedocs.io/en/v0.9.1/packages/_autosummary/analysis/snorkel.analysis.Scorer.html) to retrieve our slice metrics. But we've implemented a naive version for our multilabel task based on it.
# + id="GqkwQenBXfIa"
# Score slices
metrics["slices"] = {}
for slice_name in slices.dtype.names:
mask = slices[slice_name].astype(bool)
if sum(mask):
slice_metrics = precision_recall_fscore_support(
y_test[mask], y_pred[mask], average="micro"
)
metrics["slices"][slice_name] = {}
metrics["slices"][slice_name]["precision"] = slice_metrics[0]
metrics["slices"][slice_name]["recall"] = slice_metrics[1]
metrics["slices"][slice_name]["f1"] = slice_metrics[2]
metrics["slices"][slice_name]["num_samples"] = len(y_true[mask])
# + id="QapvZ3bgX3J6" colab={"base_uri": "https://localhost:8080/"} outputId="aa298b47-e9e1-4d80-e5c2-e2c2ed7b886a"
print(json.dumps(metrics["slices"], indent=2))
# + [markdown] id="v9yUHIrgYe6s"
# > In our [testing lesson](https://madewithml.com/courses/mlops/testing/), we'll cover another way to evaluate our model known as [behavioral testing](https://madewithml.com/courses/mlops/testing/#behavioral-testing), which we'll also include as part of performance report.
# + [markdown] id="MILv2j74iUMQ"
# ## Experiment tracking
# + [markdown] id="t40bb7o2jyCP"
# So far, we've been training and evaluating our different baselines but haven't really been tracking these experiments. We'll fix this by defining a proper process for experiment tracking which we'll use for all future experiments (including hyperparameter optimization). There are many options for experiment tracking but we're going to use [MLFlow](https://mlflow.org/) (100% free and [open-source](https://github.com/mlflow/mlflow)) because it has all the functionality we'll need (and [growing integration support](https://medium.com/pytorch/mlflow-and-pytorch-where-cutting-edge-ai-meets-mlops-1985cf8aa789)). We can run MLFlow on our own servers and databases so there are no storage cost / limitations, making it one of the most popular options and is used by Microsoft, Facebook, Databricks and others. You can also set up your own Tracking servers to synchronize runs amongst multiple team members collaborating on the same task.
#
# There are also several popular options such as a [Comet ML](https://www.comet.ml/site/) (Used by Google AI, HuggingFace, etc.) and [Weights and Biases](https://www.wandb.com/) (Used by Open AI, Toyota Research, etc.). These are fantastic tools that provide features like dashboards, seamless integration, hyperparameter search, reports and even [debugging](https://wandb.ai/latentspace/published-work/The-Science-of-Debugging-with-W-B-Reports--Vmlldzo4OTI3Ng)!
# + id="GiU-_H58iVvV" colab={"base_uri": "https://localhost:8080/"} outputId="49a0be62-1f0f-4473-e7de-f60de8095741"
# !pip install mlflow==1.13.1 pyngrok -q
# + id="jjBl3l1cl4x6"
from argparse import Namespace
import mlflow
from pathlib import Path
# + id="bDwbBLh4xdFG"
# Specify arguments
args = Namespace(
char_level=True,
filter_sizes=list(range(1, 11)),
batch_size=64,
embedding_dim=128,
num_filters=128,
hidden_dim=128,
dropout_p=0.5,
lr=2e-4,
num_epochs=100,
patience=10,
)
# + [markdown] id="fIHrsmkN7rYi"
# > When we move to Python scripts, we'll use the [Typer](https://typer.tiangolo.com/) package instead of argparse for a better CLI experience.
# + id="Bq5Zoy6YohM3"
# Set tracking URI
MODEL_REGISTRY = Path("experiments")
Path(MODEL_REGISTRY).mkdir(exist_ok=True) # create experiments dir
mlflow.set_tracking_uri("file://" + str(MODEL_REGISTRY.absolute()))
# + id="9DR1CvJeo48U" colab={"base_uri": "https://localhost:8080/"} outputId="7e5fcd24-f355-4863-8c9f-c2f19afe841d"
# !ls
# + [markdown] id="JGvUEKbl9buj"
# ### Training
# + [markdown] id="JZgQlLyHHwEk"
# We're going to log the training epoch metrics within our `Trainer`'s `train` function.
# + id="F16Qt7xSHqph"
# Modified for experiment tracking
class Trainer(object):
def __init__(self, model, device, loss_fn=None, optimizer=None, scheduler=None):
# Set params
self.model = model
self.device = device
self.loss_fn = loss_fn
self.optimizer = optimizer
self.scheduler = scheduler
def train_step(self, dataloader):
"""Train step."""
# Set model to train mode
self.model.train()
loss = 0.0
# Iterate over train batches
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, targets = batch[:-1], batch[-1]
self.optimizer.zero_grad() # Reset gradients
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, targets) # Define loss
J.backward() # Backward pass
self.optimizer.step() # Update weights
# Cumulative Metrics
loss += (J.detach().item() - loss) / (i + 1)
return loss
def eval_step(self, dataloader):
"""Validation or test step."""
# Set model to eval mode
self.model.eval()
loss = 0.0
y_trues, y_probs = [], []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, y_true = batch[:-1], batch[-1]
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, y_true).item()
# Cumulative Metrics
loss += (J - loss) / (i + 1)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
y_trues.extend(y_true.cpu().numpy())
return loss, np.vstack(y_trues), np.vstack(y_probs)
def predict_step(self, dataloader):
"""Prediction step."""
# Set model to eval mode
self.model.eval()
y_probs = []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Forward pass w/ inputs
inputs, targets = batch[:-1], batch[-1]
z = self.model(inputs)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
return np.vstack(y_probs)
def train(self, num_epochs, patience, train_dataloader, val_dataloader,
tolerance=1e-5):
best_val_loss = np.inf
for epoch in range(num_epochs):
# Steps
train_loss = self.train_step(dataloader=train_dataloader)
val_loss, _, _ = self.eval_step(dataloader=val_dataloader)
self.scheduler.step(val_loss)
# Early stopping
if val_loss < best_val_loss - tolerance:
best_val_loss = val_loss
best_model = self.model
_patience = patience # reset _patience
else:
_patience -= 1
if not _patience: # 0
print("Stopping early!")
break
# Tracking
mlflow.log_metrics(
{"train_loss": train_loss, "val_loss": val_loss}, step=epoch
)
# Logging
print(
f"Epoch: {epoch+1} | "
f"train_loss: {train_loss:.5f}, "
f"val_loss: {val_loss:.5f}, "
f"lr: {self.optimizer.param_groups[0]['lr']:.2E}, "
f"_patience: {_patience}"
)
return best_model, best_val_loss
# + [markdown] id="8Mk5Dlcj_aJd"
# And to make things simple, we'll encapsulate all the components for training into one function which returns all the artifacts we want to be able to track from our experiment. The input argument `args`contains all the parameters needed and it's nice to have it all organized under one variable so we can easily log it and tweak it for different experiments (we'll see this when we do hyperparameter optimization).
# + id="OWUtQYLIR9Nn"
def train_cnn(args, df):
"""Train a CNN using specific arguments."""
# Set seeds
set_seeds()
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
num_classes = len(label_encoder)
# Set device
cuda = True
device = torch.device("cuda" if (
torch.cuda.is_available() and cuda) else "cpu")
torch.set_default_tensor_type("torch.FloatTensor")
if device.type == "cuda":
torch.set_default_tensor_type("torch.cuda.FloatTensor")
# Tokenize
tokenizer = Tokenizer(char_level=args.char_level)
tokenizer.fit_on_texts(texts=X_train)
vocab_size = len(tokenizer)
# Convert texts to sequences of indices
X_train = np.array(tokenizer.texts_to_sequences(X_train))
X_val = np.array(tokenizer.texts_to_sequences(X_val))
X_test = np.array(tokenizer.texts_to_sequences(X_test))
# Class weights
train_tags = list(itertools.chain.from_iterable(train_df.tags.values))
counts = np.bincount([label_encoder.class_to_index[class_] for class_ in train_tags])
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
# Create datasets
train_dataset = CNNTextDataset(
X=X_train, y=y_train, max_filter_size=max(args.filter_sizes))
val_dataset = CNNTextDataset(
X=X_val, y=y_val, max_filter_size=max(args.filter_sizes))
test_dataset = CNNTextDataset(
X=X_test, y=y_test, max_filter_size=max(args.filter_sizes))
# Create dataloaders
train_dataloader = train_dataset.create_dataloader(
batch_size=args.batch_size)
val_dataloader = val_dataset.create_dataloader(
batch_size=args.batch_size)
test_dataloader = test_dataset.create_dataloader(
batch_size=args.batch_size)
# Initialize model
model = CNN(
embedding_dim=args.embedding_dim, vocab_size=vocab_size,
num_filters=args.num_filters, filter_sizes=args.filter_sizes,
hidden_dim=args.hidden_dim, dropout_p=args.dropout_p,
num_classes=num_classes)
model = model.to(device)
# Define loss
class_weights_tensor = torch.Tensor(np.array(list(class_weights.values())))
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
# Define optimizer & scheduler
optimizer = torch.optim.Adam(model.parameters(), lr=args.lr)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode="min", factor=0.1, patience=5)
# Trainer module
trainer = Trainer(
model=model, device=device, loss_fn=loss_fn,
optimizer=optimizer, scheduler=scheduler)
# Train
best_model, best_val_loss = trainer.train(
args.num_epochs, args.patience, train_dataloader, val_dataloader)
# Best threshold for f1
train_loss, y_true, y_prob = trainer.eval_step(dataloader=train_dataloader)
precisions, recalls, thresholds = precision_recall_curve(y_true.ravel(), y_prob.ravel())
threshold = find_best_threshold(y_true.ravel(), y_prob.ravel())
# Determine predictions using threshold
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# Evaluate (simple)
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
return {
"args": args,
"tokenizer": tokenizer,
"label_encoder": label_encoder,
"model": best_model,
"performance": performance,
"best_val_loss": best_val_loss,
}
# + [markdown] id="oyqrdKxZ9gFi"
# ### Tracking
# + [markdown] id="7EyOAaaTLlil"
# With MLFlow we need to first initialize an experiment and then you can do runs under that experiment.
# + id="Z5qLhfSJxSkp"
import tempfile
# + id="TJHQ_SwwLjRY" colab={"base_uri": "https://localhost:8080/"} outputId="115cbb5d-99cf-4718-c14c-1ad7041a36ce"
# Set experiment
mlflow.set_experiment(experiment_name="baselines")
# + id="FCuqxD1V2dce"
def save_dict(d, filepath):
"""Save dict to a json file."""
with open(filepath, "w") as fp:
json.dump(d, indent=2, sort_keys=False, fp=fp)
# + id="5rWwqZPWo7Oo" colab={"base_uri": "https://localhost:8080/"} outputId="979c3d56-7b12-45cc-d25d-23c3fcde6609"
# Tracking
with mlflow.start_run(run_name="cnn") as run:
# Train & evaluate
artifacts = train_cnn(args=args, df=df)
# Log key metrics
mlflow.log_metrics({"precision": artifacts["performance"]["precision"]})
mlflow.log_metrics({"recall": artifacts["performance"]["recall"]})
mlflow.log_metrics({"f1": artifacts["performance"]["f1"]})
# Log artifacts
with tempfile.TemporaryDirectory() as dp:
artifacts["tokenizer"].save(Path(dp, "tokenizer.json"))
artifacts["label_encoder"].save(Path(dp, "label_encoder.json"))
torch.save(artifacts["model"].state_dict(), Path(dp, "model.pt"))
save_dict(artifacts["performance"], Path(dp, "performance.json"))
mlflow.log_artifacts(dp)
# Log parameters
mlflow.log_params(vars(artifacts["args"]))
# + [markdown] id="lqWgLPYgCBhi"
# ### Viewing
# + [markdown] id="J9JkJ_7MCC4U"
# Let's view what we've tracked from our experiment. MLFlow serves a dashboard for us to view and explore our experiments on a localhost port but since we're inside a notebook, we're going to use public tunnel ([ngrok](https://ngrok.com/)) to view it.
# + id="nCco6Xa3436x"
from pyngrok import ngrok
# + [markdown] id="gR4xFBQI7Tqj"
# > You may need to rerun the cell below multiple times if the connection times out it is overloaded.
# + id="6gd8i4b941hL" colab={"base_uri": "https://localhost:8080/"} outputId="b160485d-ed4d-4073-9b8d-fcb52b8f34f2"
# https://stackoverflow.com/questions/61615818/setting-up-mlflow-on-google-colab
get_ipython().system_raw("mlflow server -h 0.0.0.0 -p 5000 --backend-store-uri $PWD/experiments/ &")
ngrok.kill()
ngrok.set_auth_token("")
ngrok_tunnel = ngrok.connect(addr="5000", proto="http", bind_tls=True)
print("MLflow Tracking UI:", ngrok_tunnel.public_url)
# + [markdown] id="tfUQpOIDej0z"
# MLFlow creates a main dashboard with all your experiments and their respective runs. We can sort runs by clicking on the column headers.
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/mlops/experiment_tracking/dashboard.png" width="1000" alt="pivot">
# + [markdown] id="bHhjsDJyeeGN"
# We can click on any of our experiments on the main dashboard to further explore it:
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/mlops/experiment_tracking/plots.png" width="1000" alt="pivot">
# + [markdown] id="2RVeCRid9hWs"
# ### Loading
# + [markdown] id="vEbgvyFQfW_e"
# We need to be able to load our saved experiment artifacts for inference, retraining, etc.
# + id="wUdmwNiVy6Dy"
def load_dict(filepath):
"""Load a dict from a json file."""
with open(filepath, "r") as fp:
d = json.load(fp)
return d
# + id="wP_FfHn07943" colab={"base_uri": "https://localhost:8080/"} outputId="42ad027d-0ed1-4e1e-a170-ac7a19e5db1d"
# Load all runs from experiment
experiment_id = mlflow.get_experiment_by_name("baselines").experiment_id
all_runs = mlflow.search_runs(experiment_ids=experiment_id, order_by=["metrics.best_val_loss ASC"])
print (all_runs)
# + id="PHbhpuFq9QPA"
# Best run
device = torch.device("cpu")
best_run_id = all_runs.iloc[0].run_id
best_run = mlflow.get_run(run_id=best_run_id)
client = mlflow.tracking.MlflowClient()
with tempfile.TemporaryDirectory() as dp:
client.download_artifacts(run_id=best_run_id, path="", dst_path=dp)
tokenizer = Tokenizer.load(fp=Path(dp, "tokenizer.json"))
label_encoder = LabelEncoder.load(fp=Path(dp, "label_encoder.json"))
model_state = torch.load(Path(dp, "model.pt"), map_location=device)
performance = load_dict(filepath=Path(dp, "performance.json"))
# + id="uUAGjr5w8oxM" colab={"base_uri": "https://localhost:8080/"} outputId="a1569217-0b30-401d-e621-3b124fbcc343"
print (json.dumps(performance, indent=2))
# + id="HsMvF6K_8LIt" colab={"base_uri": "https://localhost:8080/"} outputId="ac0b7b10-555c-4b76-d440-0e8cf9d8b413"
# Load artifacts
device = torch.device("cpu")
model = CNN(
embedding_dim=args.embedding_dim, vocab_size=len(tokenizer),
num_filters=args.num_filters, filter_sizes=args.filter_sizes,
hidden_dim=args.hidden_dim, dropout_p=args.dropout_p,
num_classes=len(label_encoder))
model.load_state_dict(model_state)
model.to(device)
# + id="B2axMjbk9XgI"
# Initialize trainer
trainer = Trainer(model=model, device=device)
# + id="wcypz1sO9Xk1"
# Dataloader
text = "Transfer learning with BERT for self-supervised learning"
X = np.array(tokenizer.texts_to_sequences([preprocess(text)]))
y_filler = label_encoder.encode([np.array([label_encoder.classes[0]]*len(X))])
dataset = CNNTextDataset(
X=X, y=y_filler, max_filter_size=max(filter_sizes))
dataloader = dataset.create_dataloader(
batch_size=batch_size)
# + id="NiQr5E6L9XrM" colab={"base_uri": "https://localhost:8080/"} outputId="49d794a8-1ff5-46ce-ac36-186e9bb270f6"
# Inference
y_prob = trainer.predict_step(dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
label_encoder.decode(y_pred)
# + [markdown] id="_CzPlupuYPWT"
# ## Optimization
# + [markdown] id="CHM69_fPYXy_"
# Optimization is the process of fine-tuning the hyperparameters in our experiment to optimize towards a particular objective. It can be a computationally involved process depending on the number of parameters, search space and model architectures. Hyperparameters don't just include the model's parameters but they also include parameters (choices) from preprocessing, splitting, etc. When we look at all the different parameters that can be tuned, it quickly becomes a very large search space. However, just because something is a hyperparameter doesn't mean we need to tune it.
#
# - It's absolutely alright to fix some hyperparameters (ex. lower=True during preprocessing) and remove them from the current tuning subset. Just be sure to note which parameters you are fixing and your reasoning for doing so.
# - You can initially just tune a small, yet influential, subset of hyperparameters that you believe will yield best results.
#
# There are many options for hyperparameter tuning ([Optuna](https://github.com/optuna/optuna), [Ray tune](https://github.com/ray-project/ray/tree/master/python/ray/tune), [Hyperopt](https://github.com/hyperopt/hyperopt), etc.) but we'll be using Optuna for it's simplicity and efficiency.
# + id="SFFkEgjSYR9v" colab={"base_uri": "https://localhost:8080/"} outputId="9f069e28-0889-4efa-e010-7ae2d0634dda"
# !pip install optuna==2.4.0 numpyencoder==0.3.0 -q
# + id="VLI2DvYihgH9"
import optuna
# + [markdown] id="1oxHC5gUZw6v"
# There are many factors to consider when performing hyperparameter optimization and luckily Optuna allows us to [implement](https://optuna.readthedocs.io/en/stable/reference/) them with ease. We'll be conducting a small study where we'll tune a set of arguments (we'll do a much more thorough [study](https://optuna.readthedocs.io/en/stable/reference/study.html) of the parameter space when we move our code to Python scripts). Here's the process for the study:
#
# 1. Define an objective (metric) and identifying the [direction](https://optuna.readthedocs.io/en/stable/reference/generated/optuna.study.StudyDirection.html#optuna.study.StudyDirection) to optimize.
# 2. `[OPTIONAL]` Choose a [sampler](https://optuna.readthedocs.io/en/stable/reference/samplers.html) for determining parameters for subsequent trials. (default is a tree based sampler).
# 3. `[OPTIONAL]` Choose a [pruner](https://optuna.readthedocs.io/en/stable/reference/pruners.html) to end unpromising trials early.
# 4. Define the parameters to tune in each [trial](https://optuna.readthedocs.io/en/stable/reference/trial.html) and the [distribution](https://optuna.readthedocs.io/en/stable/reference/generated/optuna.trial.Trial.html#optuna-trial-trial) of values to sample.
#
# > There are many more options (multiple objectives, storage options, etc.) to explore but this basic set up will allow us to optimize quite well.
# + id="M9hsPDs3iXy1"
from argparse import Namespace
# + id="DeGjHoore6Ca"
# Specify arguments
args = Namespace(
char_level=True,
filter_sizes=list(range(1, 11)),
batch_size=64,
embedding_dim=128,
num_filters=128,
hidden_dim=128,
dropout_p=0.5,
lr=2e-4,
num_epochs=100,
patience=10,
)
# + [markdown] id="cSxjDzajfxMF"
# We're going to modify our `Trainer` object to be able to prune unpromising trials based on the trial's validation loss.
# + id="lu9OCdgxfxXx"
# Trainer (modified for experiment tracking)
class Trainer(object):
def __init__(self, model, device, loss_fn=None,
optimizer=None, scheduler=None, trial=None):
# Set params
self.model = model
self.device = device
self.loss_fn = loss_fn
self.optimizer = optimizer
self.scheduler = scheduler
self.trial = trial
def train_step(self, dataloader):
"""Train step."""
# Set model to train mode
self.model.train()
loss = 0.0
# Iterate over train batches
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch]
inputs, targets = batch[:-1], batch[-1]
self.optimizer.zero_grad() # Reset gradients
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, targets) # Define loss
J.backward() # Backward pass
self.optimizer.step() # Update weights
# Cumulative Metrics
loss += (J.detach().item() - loss) / (i + 1)
return loss
def eval_step(self, dataloader):
"""Validation or test step."""
# Set model to eval mode
self.model.eval()
loss = 0.0
y_trues, y_probs = [], []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Step
batch = [item.to(self.device) for item in batch] # Set device
inputs, y_true = batch[:-1], batch[-1]
z = self.model(inputs) # Forward pass
J = self.loss_fn(z, y_true).item()
# Cumulative Metrics
loss += (J - loss) / (i + 1)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
y_trues.extend(y_true.cpu().numpy())
return loss, np.vstack(y_trues), np.vstack(y_probs)
def predict_step(self, dataloader):
"""Prediction step."""
# Set model to eval mode
self.model.eval()
y_probs = []
# Iterate over val batches
with torch.no_grad():
for i, batch in enumerate(dataloader):
# Forward pass w/ inputs
inputs, targets = batch[:-1], batch[-1]
z = self.model(inputs)
# Store outputs
y_prob = torch.sigmoid(z).cpu().numpy()
y_probs.extend(y_prob)
return np.vstack(y_probs)
def train(self, num_epochs, patience, train_dataloader, val_dataloader,
tolerance=1e-5):
best_val_loss = np.inf
for epoch in range(num_epochs):
# Steps
train_loss = self.train_step(dataloader=train_dataloader)
val_loss, _, _ = self.eval_step(dataloader=val_dataloader)
self.scheduler.step(val_loss)
# Early stopping
if val_loss < best_val_loss - tolerance:
best_val_loss = val_loss
best_model = self.model
_patience = patience # reset _patience
else:
_patience -= 1
if not _patience: # 0
print("Stopping early!")
break
# Logging
print(
f"Epoch: {epoch+1} | "
f"train_loss: {train_loss:.5f}, "
f"val_loss: {val_loss:.5f}, "
f"lr: {self.optimizer.param_groups[0]['lr']:.2E}, "
f"_patience: {_patience}"
)
# Pruning based on the intermediate value
self.trial.report(val_loss, epoch)
if self.trial.should_prune():
raise optuna.TrialPruned()
return best_model, best_val_loss
# + [markdown] id="z3dGFhnehUbk"
# We'll also modify our `train_cnn` function to include information about the trial.
# + id="Qm_xEptLhUqm"
def train_cnn(args, df, trial=None):
"""Train a CNN using specific arguments."""
# Set seeds
set_seeds()
# Get data splits
preprocessed_df = df.copy()
preprocessed_df.text = preprocessed_df.text.apply(preprocess, lower=True)
X_train, X_val, X_test, y_train, y_val, y_test, label_encoder = get_data_splits(preprocessed_df)
num_classes = len(label_encoder)
# Set device
cuda = True
device = torch.device("cuda" if (
torch.cuda.is_available() and cuda) else "cpu")
torch.set_default_tensor_type("torch.FloatTensor")
if device.type == "cuda":
torch.set_default_tensor_type("torch.cuda.FloatTensor")
# Tokenize
tokenizer = Tokenizer(char_level=args.char_level)
tokenizer.fit_on_texts(texts=X_train)
vocab_size = len(tokenizer)
# Convert texts to sequences of indices
X_train = np.array(tokenizer.texts_to_sequences(X_train))
X_val = np.array(tokenizer.texts_to_sequences(X_val))
X_test = np.array(tokenizer.texts_to_sequences(X_test))
# Class weights
train_tags = list(itertools.chain.from_iterable(train_df.tags.values))
counts = np.bincount([label_encoder.class_to_index[class_] for class_ in train_tags])
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
# Create datasets
train_dataset = CNNTextDataset(
X=X_train, y=y_train, max_filter_size=max(args.filter_sizes))
val_dataset = CNNTextDataset(
X=X_val, y=y_val, max_filter_size=max(args.filter_sizes))
test_dataset = CNNTextDataset(
X=X_test, y=y_test, max_filter_size=max(args.filter_sizes))
# Create dataloaders
train_dataloader = train_dataset.create_dataloader(
batch_size=args.batch_size)
val_dataloader = val_dataset.create_dataloader(
batch_size=args.batch_size)
test_dataloader = test_dataset.create_dataloader(
batch_size=args.batch_size)
# Initialize model
model = CNN(
embedding_dim=args.embedding_dim, vocab_size=vocab_size,
num_filters=args.num_filters, filter_sizes=args.filter_sizes,
hidden_dim=args.hidden_dim, dropout_p=args.dropout_p,
num_classes=num_classes)
model = model.to(device)
# Define loss
class_weights_tensor = torch.Tensor(np.array(list(class_weights.values())))
loss_fn = nn.BCEWithLogitsLoss(weight=class_weights_tensor)
# Define optimizer & scheduler
optimizer = torch.optim.Adam(model.parameters(), lr=args.lr)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(
optimizer, mode='min', factor=0.1, patience=5)
# Trainer module
trainer = Trainer(
model=model, device=device, loss_fn=loss_fn,
optimizer=optimizer, scheduler=scheduler, trial=trial)
# Train
best_model, best_val_loss = trainer.train(
args.num_epochs, args.patience, train_dataloader, val_dataloader)
# Best threshold for f1
train_loss, y_true, y_prob = trainer.eval_step(dataloader=train_dataloader)
precisions, recalls, thresholds = precision_recall_curve(y_true.ravel(), y_prob.ravel())
threshold = find_best_threshold(y_true.ravel(), y_prob.ravel())
# Determine predictions using threshold
test_loss, y_true, y_prob = trainer.eval_step(dataloader=test_dataloader)
y_pred = np.array([np.where(prob >= threshold, 1, 0) for prob in y_prob])
# Evaluate (simple)
metrics = precision_recall_fscore_support(y_test, y_pred, average="weighted")
performance = {"precision": metrics[0], "recall": metrics[1], "f1": metrics[2]}
return {
"args": args,
"tokenizer": tokenizer,
"label_encoder": label_encoder,
"model": best_model,
"performance": performance,
"best_val_loss": best_val_loss,
"threshold": threshold,
}
# + [markdown] id="ncGdA34Ahod9"
# ### Objective
# + [markdown] id="NzekaGZyeHXb"
# We need to define an `objective` function that will consume a trial and a set of arguments and produce the metric to optimize on (`f1` in our case).
# + id="Jg33IjzSd27e"
def objective(trial, args):
"""Objective function for optimization trials."""
# Paramters (to tune)
args.embedding_dim = trial.suggest_int("embedding_dim", 128, 512)
args.num_filters = trial.suggest_int("num_filters", 128, 512)
args.hidden_dim = trial.suggest_int("hidden_dim", 128, 512)
args.dropout_p = trial.suggest_uniform("dropout_p", 0.3, 0.8)
args.lr = trial.suggest_loguniform("lr", 5e-5, 5e-4)
# Train & evaluate
artifacts = train_cnn(args=args, df=df, trial=trial)
# Set additional attributes
trial.set_user_attr("precision", artifacts["performance"]["precision"])
trial.set_user_attr("recall", artifacts["performance"]["recall"])
trial.set_user_attr("f1", artifacts["performance"]["f1"])
trial.set_user_attr("threshold", artifacts["threshold"])
return artifacts["performance"]["f1"]
# + [markdown] id="K42Yf3OEhp_V"
# ### Study
# + [markdown] id="epa_kTIl9H7b"
# We're ready to kick off our study with our [MLFlowCallback](https://optuna.readthedocs.io/en/stable/reference/generated/optuna.integration.MLflowCallback.html) so we can track all of the different trials.
# + id="WC58iWVjjPN8"
from numpyencoder import NumpyEncoder
from optuna.integration.mlflow import MLflowCallback
# + id="ZbPWc_c8hsA6"
NUM_TRIALS = 50 # small sample for now
# + id="RQByQ8pihjIR" colab={"base_uri": "https://localhost:8080/"} outputId="6f20b411-a44c-4e81-e87a-16dccebc3368"
# Optimize
pruner = optuna.pruners.MedianPruner(n_startup_trials=5, n_warmup_steps=5)
study = optuna.create_study(study_name="optimization", direction="maximize", pruner=pruner)
mlflow_callback = MLflowCallback(
tracking_uri=mlflow.get_tracking_uri(), metric_name="f1")
study.optimize(lambda trial: objective(trial, args),
n_trials=NUM_TRIALS,
callbacks=[mlflow_callback])
# + id="FfRGqQaGkfx8" colab={"base_uri": "https://localhost:8080/"} outputId="8d8762de-760d-4168-d6a4-6ebb9325ebb8"
# MLFlow dashboard
get_ipython().system_raw("mlflow server -h 0.0.0.0 -p 5000 --backend-store-uri $PWD/experiments/ &")
ngrok.kill()
ngrok.set_auth_token("")
ngrok_tunnel = ngrok.connect(addr="5000", proto="http", bind_tls=True)
print("MLflow Tracking UI:", ngrok_tunnel.public_url)
# + [markdown] id="NojJ-Z1X6IEQ"
# You can compare all (or a subset) of the trials in our experiment.
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/mlops/hyperparameter_optimization/compare.png" width="1000" alt="compare">
#
# We can then view the results through various lens (contours, parallel coordinates, etc.)
#
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/mlops/hyperparameter_optimization/contour.png" width="1000" alt="compare">
# <img src="https://raw.githubusercontent.com/GokuMohandas/MadeWithML/main/images/mlops/hyperparameter_optimization/parallel_coordinates.png" width="1000" alt="compare">
# + id="9HtwFRzEikt7" colab={"base_uri": "https://localhost:8080/", "height": 309} outputId="ac488325-f2d7-47eb-e5fc-206975ffca32"
# All trials
trials_df = study.trials_dataframe()
trials_df = trials_df.sort_values(["value"], ascending=False) # sort by metric
trials_df.head()
# + id="6mP99RFjiENR" colab={"base_uri": "https://localhost:8080/"} outputId="04fe04d8-3343-4a09-fc40-d476c9d1efdd"
# Best trial
print (f"Best value (f1): {study.best_trial.value}")
print (f"Best hyperparameters: {study.best_trial.params}")
# + [markdown] id="Pggo1cnaix85"
# > Don't forget to save learned parameters (ex. decision threshold) during training which you'll need later for inference.
# + id="JZIS8RtfiuDc" colab={"base_uri": "https://localhost:8080/"} outputId="667241fc-111a-44ac-e302-5d78bb43c33c"
# Save best parameters
params = {**args.__dict__, **study.best_trial.params}
params["threshold"] = study.best_trial.user_attrs["threshold"]
print (json.dumps(params, indent=2, cls=NumpyEncoder))
# + [markdown] id="KbxtdyENi78d"
# ... and now we're finally ready to move from working in Jupyter notebooks to Python scripts. We'll be revisiting everything we did so far, but this time with proper software engineering prinicples such as object oriented programming (OOPs), styling, testing, etc. → [https://madewithml.com/#mlops](https://madewithml.com/#mlops)
|
notebooks/tagifai.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get data from a spreadsheet
# In this exercise, you'll create a data frame from a "base case" Excel file: one with a single sheet of tabular data. The fcc_survey.xlsx file here has a sample of responses from FreeCodeCamp's annual New Developer Survey. This survey asks participants about their demographics, education, work and home life, plus questions about how they're learning to code. Let's load all of it.
#
# pandas has not been pre-loaded in this exercise, so you'll need to import it yourself before using read_excel() to load the spreadsheet.
# +
# Load pandas as pd
import pandas as pd
# Read spreadsheet and assign it to survey_responses
survey_responses = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl')
# View the head of the data frame
(survey_responses.head())
# -
# # Load a portion of a spreadsheet
# Spreadsheets meant to be read by people often have multiple tables, e.g., a small business might keep an inventory workbook with tables for different product types on a single sheet. Even tabular data may have header rows of metadata, like the New Developer Survey data here. While the metadata is useful, we don't want it in a data frame. You'll use read_excel()'s skiprows keyword to get just the data. You'll also create a string to pass to usecols to get only columns AD and AW through BA, about future job goals.
#
# pandas has been imported as pd.
# +
# Create string of lettered columns to load
col_string = "AD, AW:BA"
# Load data with skiprows and usecols set
survey_responses = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl',
skiprows=2,
usecols=col_string)
# View the names of the columns selected
print(survey_responses.columns)
# -
# # Select a single sheet
# An Excel workbook may contain multiple sheets of related data. The New Developer Survey response workbook has sheets for different years. Because read_excel() loads only the first sheet by default, you've already gotten survey responses for 2016. Now, you'll create a data frame of 2017 responses using read_excel()'s sheet_name argument in a couple different ways.
#
# pandas has been imported as pd.
# +
import matplotlib.pyplot as plt
import pandas as pd
# Create df from second worksheet by referencing its position
responses_2017 = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2,
sheet_name=1)
# Graph where people would like to get a developer job
job_prefs = responses_2017.groupby("JobPref").JobPref.count()
job_prefs.plot.barh()
plt.show()
# +
# Create df from second worksheet by referencing its name
responses_2017 = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2,
sheet_name='2017')
# Graph where people would like to get a developer job
job_prefs = responses_2017.groupby("JobPref").JobPref.count()
job_prefs.plot.barh()
plt.show()
# -
# # Select multiple sheets
# So far, you've read Excel files one sheet at a time, which lets you customize import arguments for each sheet. But if an Excel file has some sheets that you want loaded with the same parameters, you can get them in one go by passing a list of their names or indices to read_excel()'s sheet_name keyword. To get them all, pass None. You'll practice both methods to get data from fcc_survey.xlsx, which has multiple sheets of similarly-formatted data.
#
# pandas has been loaded as pd.
# +
# Load both the 2016 and 2017 sheets by name
all_survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl',
sheet_name=['2016', '2017'])
# View the data type of all_survey_data
print(type(all_survey_data))
# +
# Load all sheets in the Excel file
all_survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl',
sheet_name=[0, '2017'])
# View the sheet names in all_survey_data
print(all_survey_data.keys())
# +
# Load all sheets in the Excel file
all_survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl',
sheet_name=None)
# View the sheet names in all_survey_data
print(all_survey_data.keys())
# -
# # Work with multiple spreadsheets
# Workbooks meant primarily for human readers, not machines, may store data about a single subject across multiple sheets. For example, a file may have a different sheet of transactions for each region or year in which a business operated.
#
# The FreeCodeCamp New Developer Survey file is set up similarly, with samples of responses from different years in different sheets. Your task here is to compile them in one data frame for analysis.
#
# pandas has been imported as pd. All sheets have been read into the ordered dictionary responses, where sheet names are keys and data frames are values, so you can get data frames with the values() method.
# +
# Create an empty data frame
all_responses = pd.DataFrame()
responses = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2,
sheet_name=['2016', '2017'])
# Set up for loop to iterate through values in responses
for df in responses.values():
# Print the number of rows being added
print("Adding {} rows".format(df.shape[0]))
# Append df to all_responses, assign result
all_responses = all_responses.append(df)
# Graph employment statuses in sample
counts = all_responses.groupby("EmploymentStatus").EmploymentStatus.count()
counts.plot.barh()
plt.show()
# -
# # Set Boolean columns
# Datasets may have columns that are most accurately modeled as Boolean values. However, pandas usually loads these as floats by default, since defaulting to Booleans may have undesired effects like turning NA values into Trues.
#
# fcc_survey_subset.xlsx contains a string ID column and several True/False columns indicating financial stressors. You'll evaluate which non-ID columns have no NA values and therefore can be set as Boolean, then tell read_excel() to load them as such with the dtype argument.
#
# pandas is loaded as pd.
# +
# Load the data
survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2)
# Count NA values in each column
survey_data.isna().sum()
# +
# Set dtype to load appropriate column(s) as Boolean data
survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2)
survey_data.dropna()
survey_data['HasDebt'] = survey_data['HasDebt'].astype('bool')
# View financial burdens by Boolean group
print(survey_data.groupby('HasDebt').sum())
# -
# # Set custom true/false values
# In Boolean columns, pandas automatically recognizes certain values, like "TRUE" and 1, as True, and others, like "FALSE" and 0, as False. Some datasets, like survey data, can use unrecognized values, such as "Yes" and "No".
#
# For practice purposes, some Boolean columns in the New Developer Survey have been coded this way. You'll make sure they're properly interpreted with the help of the true_values and false_values arguments.
#
# pandas is loaded as pd. You can assume the columns you are working with have no missing values.
# Load file with Yes as a True value and No as a False value
survey_subset = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2)
survey_data['HasDebt'] = survey_data['HasDebt'].astype('bool').replace({
True: 'Yes',
False: 'No'
})
survey_data['AttendedBootcamp'] = survey_data['AttendedBootcamp'].astype('bool').replace({
True: 'Yes',
False: 'No'
})
# View the data
print(survey_subset.head())
# # Parse simple dates
# pandas does not infer that columns contain datetime data; it interprets them as object or string data unless told otherwise. Correctly modeling datetimes is easy when they are in a standard format -- we can use the parse_dates argument to tell read_excel() to read columns as datetime data.
#
# The New Developer Survey responses contain some columns with easy-to-parse timestamps. In this exercise, you'll make sure they're the right data type.
#
# pandas has been loaded as pd.
# +
# Load file, with Part1StartTime parsed as datetime data
survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2,
parse_dates=['Part1StartTime'])
# Print first few values of Part1StartTime
print(survey_data.Part1StartTime.head())
# -
# # Get datetimes from multiple columns
# Sometimes, datetime data is split across columns. A dataset might have a date and a time column, or a date may be split into year, month, and day columns.
#
# A column in this version of the survey data has been split so that dates are in one column, Part2StartDate, and times are in another, Part2StartTime. Your task is to use read_excel()'s parse_dates argument to combine them into one datetime column with a new name.
#
# pandas has been imported as pd.
# +
# Create dict of columns to combine into new datetime column
datetime_cols = {"Part2Start": ["Part2StartTime"]}
# Load file, supplying the dict to parse_dates
survey_data = pd.read_excel('../datasets/fcc-new-coder-survey.xlsx', engine='openpyxl', header=2,
parse_dates=datetime_cols)
# View summary statistics about Part2Start
print(survey_data.Part2Start.describe())
# -
# # Parse non-standard date formats
# So far, you've parsed dates that pandas could interpret automatically. But if a date is in a non-standard format, like 19991231 for December 31, 1999, it can't be parsed at the import stage. Instead, use pd.to_datetime() to convert strings to dates after import.
#
# The New Developer Survey data has been loaded as survey_data but contains an unparsed datetime field. We'll use to_datetime() to convert it, passing in the column to convert and a string representing the date format used.
#
# For more on date format codes, see this reference. Some common codes are year (%Y), month (%m), day (%d), hour (%H), minute (%M), and second (%S).
#
# pandas has been imported as pd.
# +
# Parse datetimes and assign result back to Part2EndTime
survey_data["Part2EndTime"] = pd.to_datetime(survey_data["Part2EndTime"],
format="%Y%m%d %H:%M:%S")
# Print first few values of Part2EndTime
print(survey_data['Part2EndTime'].head())
# -
|
streamlined-data-ingestion-with-pandas/2. Importing Data From Excel Files/notebook_section_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# Copyright 2021 Google LLC
# Use of this source code is governed by an MIT-style
# license that can be found in the LICENSE file or at
# https://opensource.org/licenses/MIT.
# Author(s): <NAME> (<EMAIL>) and <NAME> (<EMAIL>)
# -
# <a href="https://opensource.org/licenses/MIT" target="_parent"><img src="https://img.shields.io/github/license/probml/pyprobml"/></a>
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/notebooks/figures//chapter12_figures.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Cloning the pyprobml repo
# !git clone https://github.com/probml/pyprobml
# %cd pyprobml/scripts
# # Installing required software (This may take few minutes)
# !apt install octave -qq > /dev/null
# !apt-get install liboctave-dev -qq > /dev/null
# ## Figure 12.1:
# The logistic (sigmoid) function $\sigma (x)$ in solid red, with the Gaussian cdf function $\Phi (\lambda x)$ in dotted blue superimposed. Here $\lambda =\sqrt \pi /8 $, which was chosen so that the derivatives of the two curves match at $x=0$. Adapted from Figure 4.9 of \citep BishopBook .
# Figure(s) generated by [probit_plot.py](https://github.com/probml/pyprobml/blob/master/scripts/probit_plot.py)
# %run ./probit_plot.py
# ## Figure 12.2:
# Fitting a probit regression model in 2d using a quasi-Newton method or EM.
# Figure(s) generated by [probitRegDemo.m](https://github.com/probml/pmtk3/blob/master/demos/probitRegDemo.m)
# !octave -W probitRegDemo.m >> _
|
notebooks/figures/chapter12_figures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: keras1
# language: python
# name: keras1
# ---
# + jupyter={"outputs_hidden": true}
#INITIAL TRAIN RPN FIRST
BACKBONE = "thundernetv1"
DATASET = "/home/henri_tomas/projects/faster-rcnn/frcnn-from-scratch-with-keras/2007_2012_trainval.txt"
# !python train_rpn.py --network $BACKBONE -o simple -p $DATASET -n 4 --num_epochs 50
# + jupyter={"outputs_hidden": true}
BACKBONE = "thundernetv1"
DATASET = "/home/henri_tomas/projects/faster-rcnn/frcnn-from-scratch-with-keras/2007_2012_trainval.txt"
MODEL = "./models/thundernetv1/voc_person_cem_3.hdf5"
# !python measure_map.py --network $BACKBONE -p $DATASET --load $MODEL --write -n 4
# + jupyter={"outputs_hidden": true}
#INITIAL TRAINING FRCNN, NOT CONTINUE TRAINING
BACKBONE = "thundernetv1"
DATASET = "/home/henri_tomas/projects/faster-rcnn/frcnn-from-scratch-with-keras/2007_2012_trainval.txt"
RPN = "./models/rpn/rpn.thundernetv1.weights.28-1.45.hdf5"
SAVE_TO = "voc_person_tf_3"
# !python train_frcnn.py --network $BACKBONE -o simple -p $DATASET --rpn $RPN --num_epochs 30 --dataset $SAVE_TO -n 4
# +
# CONTINUE TRAINING
BACKBONE = "thundernetv1"
DATASET = "/home/henri_tomas/projects/faster-rcnn/frcnn-from-scratch-with-keras/2007_2012_trainval.txt"
SAVE_TO = "voc_person_tf_3"
MODEL = "./models/thundernetv1/voc_person_tf_3.hdf5"
# !python train_frcnn.py --network $BACKBONE -o simple -p $DATASET --num_epochs 10 --dataset $SAVE_TO -n 4 --load $MODEL
# +
#TEST ON TEST_IMAGES
BACKBONE = "thundernetv1"
TEST_IMAGES = "./test_images/"
MODEL = "./models/thundernetv1/voc_person_tf_3.hdf5"
# !python test_frcnn.py --network $BACKBONE -p $TEST_IMAGES --load $MODEL --write -n 4
# + jupyter={"outputs_hidden": true}
#SAVE MODELS TO H5 WITH CORRECT, CONSTANT INPUT_SHAPES !!!
BACKBONE = "thundernetv1"
TEST_IMAGES = "./test_images/"
MODEL = "./models/thundernetv1/voc_person_cem_3.hdf5"
# !python tflite_convert.py --network $BACKBONE -p $TEST_IMAGES --load $MODEL --write -n 4
# + jupyter={"outputs_hidden": true}
#CONVERT RPN
import tensorflow as tf
converter = tf.lite.TFLiteConverter.from_keras_model_file("model_rpn.h5")
tflite_model = converter.convert()
open("model_rpn.tflite", "wb").write(tflite_model)
# + jupyter={"outputs_hidden": true}
import tensorflow as tf
from keras_frcnn.RoiPoolingConv import RoiPoolingConv
custom_objects = {
"RoiPoolingConv" : RoiPoolingConv,
}
converter = tf.lite.TFLiteConverter.from_keras_model_file("model_cls.h5",
custom_objects=custom_objects)
tflite_model = converter.convert()
open("model_cls.tflite", "wb").write(tflite_model)
# + jupyter={"outputs_hidden": true}
#CONVERT CLS
import tensorflow as tf
import keras
from keras_frcnn.RoiPoolingConv import RoiPoolingConv
keras_file = "model_cls.h5"
# Convert the model.
custom_objects = {
"RoiPoolingConv" : RoiPoolingConv,
}
model = tf.keras.models.load_model(keras_file, custom_objects=custom_objects)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
tflite_model = converter.convert()
open("model_cls.tflite", "wb").write(tflite_model)
# + jupyter={"outputs_hidden": true}
#Sanity Test (does tf_RPN and tflite_RPN have same output?)
import tensorflow as tf
import numpy as np
import time
tf_model = tf.keras.models.load_model("model_rpn.h5")
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path="model_rpn_320.tflite")
interpreter.allocate_tensors()
# Get input and output tensors.
input_details = interpreter.get_input_details()
output_details = interpreter.get_output_details()
# Test model on random input data.
input_shape = input_details[0]['shape']
input_data = np.array(np.random.random_sample(input_shape), dtype=np.float32)
# +
#Inference
interpreter.set_tensor(input_details[0]['index'], input_data)
interpreter.invoke()
#0=cls, 1=reg, 2=base_layers
tflite_results = interpreter.get_tensor(output_details[0]['index'])
# Test the original TensorFlow model on random input data.
#tf_results = tf_model(tf.constant(input_data)) #produces a tf tensor which doesnt show the FUCKING values
tf_results = tf_model.predict(input_data) #shows values because numpy array yey
#print(tflite_results.shape)
# Compare the result.
# NOTE: only 0 (the cls layer) is tested here, pray that is enough proof
for tf_result, tflite_result in zip(tf_results[0], tflite_results):
decimal = 5
error = np.testing.assert_almost_equal(tf_result, tflite_result, decimal=decimal)
#IF no assertion error raised...
if error==None:
print("TF and TFLITE models are equal up to {} decimal places".format(decimal))
# +
#Sanity Test (get_data)
from keras_frcnn.simple_parser import get_data
voc12_imgs, classes_count, class_mapping = get_data('/home/henri_tomas/projects/voc12/2012_trainval.txt')
voc07_imgs, _, _ = get_data('/home/henri_tomas/projects/voc07/2007_trainval.txt')
#for i in range(len(all_imgs)):
#print(all_imgs[i])
all_imgs = voc12_imgs + voc07_imgs
#Should print 4087 + 2008 = 6095
print(len(all_imgs))
# +
#Sanity Test (get_data)
from keras_frcnn.simple_parser import get_data
all_imgs, classes_count, class_mapping = get_data('/home/henri_tomas/projects/faster-rcnn/frcnn-from-scratch-with-keras/2007_2012_trainval.txt')
#Should print 4087 + 2008 = 6095
print(len(all_imgs))
# -
import tensorflow as tf
print(tf.__version__)
# + jupyter={"outputs_hidden": true}
from keras.applications.mobilenet_v2 import MobileNetV2
feature_extractor = MobileNetV2(input_shape=(224,224,3),
include_top=False,
weights='imagenet',
pooling=None)
feature_extractor.summary()
# -
import numpy as np
print(np.random.randint(0,2))
from tensorflow.keras.backend import is_keras_tensor
|
working_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/robrother/EspacioLatente/blob/main/VGG16EspacioLatente.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="feFRu_qfcPSy"
# # IMPORTAMOS LAS LIBRERIAS QUE SE VAN A UTILIZAR
# + id="9772e998" outputId="cc60ef41-aef3-478b-f042-e594b32c8b46" colab={"base_uri": "https://localhost:8080/"}
import tensorflow as tf
from tensorflow import keras
import tensorflow.keras
import time
import numpy as np
import os
import sys
from keras.applications.vgg16 import VGG16
from PIL import Image
from tensorflow.keras import layers, Input
from tensorflow.keras.models import Sequential, Model
from keras.models import Model
from keras.models import Sequential
# %matplotlib inline
print(f"Tensor Flow Version: {tf.__version__}")
print(f"Keras Version: {tensorflow.keras.__version__}")
print()
print(f"Python {sys.version}")
print("GPU is", "available" if tf.test.is_gpu_available() else "NOT AVAILABLE")
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
print(tf.test.is_built_with_cuda())
# + [markdown] id="JHsgaN-WcXZF"
# # Importamos el modelo completo de VGG16
# + id="e3ebd666" outputId="7c9d21ef-397b-42d5-d323-c6ea042acb12" colab={"base_uri": "https://localhost:8080/"}
model = VGG16()
model.summary()
# + [markdown] id="8cHn9n7SciP9"
# # De nuestro modelo completo creamos un submodelo que será de nuestro encoder, para ello usamos las capas de nuestra arquitectura VGG16 que son solo de extraccion de caracteristicas
# + id="c12c720f" outputId="4fda7681-0c69-4f21-832a-688615f40979" colab={"base_uri": "https://localhost:8080/"}
encoder = Sequential()
for i, layer in enumerate(model.layers):
if i<19:
encoder.add(layer)
encoder.summary()
# + [markdown] id="bs2HhTEPc3zS"
# # Congelamos los pesos de esas capas, en este caso hacemos uso de la técnica de Transfer Learning
# + id="e05a9a37"
for layer in encoder.layers:
layer.trainable = False
# + [markdown] id="dDXkfGmfdCEd"
# # Cargamos nuestras imagenes desde un directorio, en este caso todo nuestro conjunto de datos se encuentra en la carpeta con nombre "CenterDronet" la dirección será reemplazada por el directorio donde se encuentren las imágenes de las manzanas. En automático normaliza y recorta las imágnes al tamaño utilizado por VGG, así que no importa que las imágenes sean de distintos tamaños-
# + id="b1ca8c35" outputId="e6a97617-80d3-4f49-b548-8d86bc154c74"
directorio = 'D:/CenterDronet/'
train_datagen = ImageDataGenerator(rescale=1./255)
Imagenes = train_datagen.flow_from_directory(directorio, target_size=(224,224), batch_size=5614, class_mode=None)
# + [markdown] id="V6EkgSTJdxF6"
# # Hacemos pasar nuestro dataset por la red y obtenemos la representación en el espacio latente.
# + id="abc18422" outputId="9974c466-2d35-4edc-a161-902f1d9b9657"
start_time = time.time()
embedings=[]
ex=[]
cuenta = 0
for i in (Imagenes [0]):
im = np.expand_dims(i, axis=0)
pred = encoder.predict(im)
ex.append(pred)
print(cuenta) #<---------------------Puedes comentar este print, solo sirve para indicar el numero de imagen en el que va
cuenta+=1
print("--- %s seconds ---" % (time.time() - start_time))
print("Se ha obtenido la representación en el Espacio Latente de las imágenes")
# + [markdown] id="y08Vpm4VeLvo"
# # Guardamos el resultado en un archivo de numpy para que ya pueda ser cargado directamente el espacio Latente de las imágenes y no se tenga que ejecutar todo lo anterior.
# + id="bc4b7f43" outputId="16a41244-05e6-4708-9a9f-e600b2ed8fea"
print(len(ex))
print(ex[0].shape)
print(type(ex[0]))
# + id="65f06ccf"
np.save('EspacioLatente.npy', ex)
|
VGG16EspacioLatente.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Combining ReacLib Rates with Electron Capture Tables
#
# Here's an example of using tabulated weak rates from Suzuki et a. (2016) together with rates from the ReacLib database.
#
# We'll build a network suitable for e-capture supernovae.
import pynucastro as pyna
# First create a library from ReacLib using a set of nuclei.
# +
reaclib_library = pyna.ReacLibLibrary()
all_nuclei = ["p","he4","ne20","o20","f20","mg24","al27","o16","si28","s32","p31"]
ecsn_library = reaclib_library.linking_nuclei(all_nuclei,with_reverse=True)
# -
# Here are the rates it chose
print(ecsn_library)
# Now let's specify the weak rates we want from Suzuki et al. -- these tables are included with pynucastro:
ecsn_tabular = ["f20--o20-toki","ne20--f20-toki","o20--f20-toki","f20--ne20-toki"]
# These tables are in terms of $T$ and $\rho Y_e$. We can easily plot the tabulated electron capture rates just like ReacLib rates:
a = pyna.Rate("o20--f20-toki")
fig = a.plot()
# Let's create a rate collection from the ReacLib rates and look to see which are actually important
rc = pyna.RateCollection(libraries=[ecsn_library])
# Here we'll pick thermodynamic conditions appropriate to the oxygen burning shell in a white dwarf
comp = pyna.Composition(rc.get_nuclei())
comp.set_nuc("o16", 0.5)
comp.set_nuc("ne20", 0.3)
comp.set_nuc("mg24", 0.1)
comp.set_nuc("o20", 1.e-5)
comp.set_nuc("f20", 1.e-5)
comp.set_nuc("p", 1.e-5)
comp.set_nuc("he4", 1.e-2)
comp.set_nuc("al27", 1.e-2)
comp.set_nuc("si28", 1.e-2)
comp.set_nuc("s32", 1.e-2)
comp.set_nuc("p31", 1.e-2)
comp.normalize()
rc.plot(rho=7.e9, T=1.e9, comp=comp, ydot_cutoff_value=1.e-20)
# Now, this rate collection already includes some weak rates from ReacLib -- we want to remove those. We'll also take the opportunity to remove any rates that are not important, by looking for a $|\dot{Y}| > 10^{-20}~s^{-1}$.
new_rate_list = []
ydots = rc.evaluate_rates(rho=7.e9, T=1.e9, composition=comp)
for rate in rc.rates:
if abs(ydots[rate]) >= 1.e-20 and not rate.weak:
new_rate_list.append(rate)
# Finally, we'll create a new rate collection by combining this new list of rates with the list of tables:
rc_new = pyna.RateCollection(rates=new_rate_list, rate_files=ecsn_tabular)
rc_new.plot(rho=7.e9, T=1.e9, comp=comp)
# We can see the values of the rates at our thermodynamic conditions easily:
rc_new.evaluate_rates(rho=7.e9, T=1.e9, composition=comp)
# Additionally, we can see the specific energy generation rate for these conditions
rc_new.evaluate_energy_generation(rho=7.e9, T=1.e9, composition=comp)
|
docs/source/electron-captures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"} tags=["Header"]
# <img src="https://drive.google.com/uc?id=1v7YY_rNBU2OMaPnbUGmzaBj3PUeddxrw" alt="ITI MCIT EPITA" style="width: 750px;"/>
#
# ---
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://drive.google.com/uc?id=1R0-FYpJQW5YFy6Yv-RZ1rpyBslay0251" alt="Python Logo" style="width: 400px;"/>
# + [markdown] slideshow={"slide_type": "subslide"} tags=["Header"]
# # Introduction To Python
# ## Session 01 : The Basics
#
# By: **<NAME>**, <EMAIL>
# ___
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ### Intro
#
# * the language is named after the BBC show “Monty Python’s Flying Circus”
# * Implementation started by *<NAME> Rossum* in December 1989
# * first released in 1991
# * Python 3.0, released in 2008
# * only Python 3.6 and later are supported, other versions reached EOL support
# * ranked third on [TIOBE index Jan 2021](https://www.tiobe.com/tiobe-index/python/)
# * Python is [TIOBE's Programming Language of 2020!](https://www.tiobe.com/tiobe-index/)
# * Nowadays it is the favorite language in fields such as data science and machine learning,
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="https://drive.google.com/uc?id=1z3Rertrt2480lecPTMuKDW5XOsfuTE_o" alt="Python Logo" style="width: 75%;"/>
#
#
#
# **further reading:**
# * [[https://python-history.blogspot.com/2009/01/brief-timeline-of-python.html]](https://python-history.blogspot.com/2009/01/brief-timeline-of-python.html)
#
# ----
# + [markdown] slideshow={"slide_type": "slide"}
# #### Why to Learn Python?
# + [markdown] slideshow={"slide_type": "subslide"}
# * Simple language which means that It's easy to learn and implement.
# * Python is a dynamic language
# * reduces complexity so you can implement functionality with less code.
# * it is faster for development when compared to other static type language
# * allowing developers to test machine learning algorithms faster.
# * Python is Platform independent, available on Windows, Mac OS X, and Unix operating systems
# * It comes with a large collection of standard modules that you can use as the basis of your programs
# * Some of these modules provide things like file I/O, system calls, sockets, and even interfaces to graphical user interface toolkits like Tk
# + [markdown] slideshow={"slide_type": "subslide"}
# * the high-level data types allow you to express complex operations in a single statement;
# * statement grouping is done by indentation instead of beginning and ending brackets;
# * no variable or argument declarations are necessary.
# + [markdown] slideshow={"slide_type": "subslide"}
# * A great library ecosystem [PyPi.org/](https://pypi.org/)
# * [Scikit-learn](http://scikit-learn.org/stable/user_guide.html) for basic ML algorithms like clustering, linear and logistic regressions, regression, classification.
# * [Pandas](https://pandas.pydata.org/) for high-level data structures and analysis.
# * [Keras](https://keras.io/) for deep learning.
# * [TensorFlow](https://www.tensorflow.org/) for working with deep learning by setting up, training, and utilizing artificial neural networks with massive datasets.
# * [Matplotlib](https://matplotlib.org/tutorials/index.html) for creating 2D plots, histograms, charts, and other forms of visualization
# * .... and more, a lot more.
# + [markdown] slideshow={"slide_type": "subslide"}
# * Python is an open source language with strong community support built around the programming language.
# * [docs.python.org](https://docs.python.org)
# * [discuss.python.org](https://discuss.python.org)
# * [stackoverflow](https://stackoverflow.com/questions/tagged/python)
# * [medium.com](https://medium.com)
# * [Dev.to](https://dev.to)
#
# **further reading:**
# * [https://docs.python.org/3/tutorial/appetite.html](https://docs.python.org/3/tutorial/appetite.html)
#
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ### First Block of Code
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Shebang/HashBang
# + [markdown] slideshow={"slide_type": "subslide"}
# **scripts are not compiled**
#
# The first line in this file is the "shebang" line. When you execute a file from the shell, the shell tries to run the file using the command specified on the shebang line.
#
# ```{python}
# # # #! /usr/bin/python
# # # #! /usr/bin/env python
# # # #! /usr/local/bin/python
# # # #! python
# ```
#
# we need to tell the shell three things:
#
# 1. That the file is a script
# 2. Which interpreter we want to execute the script
# 3. The path of said interpreter
#
# ```#!/usr/bin/env python``` will use the first version of Python it finds in the user's Path, we can moderate this behaviour by specifying a version number such as ```#!/usr/bin/env pythonX.x```
#
#
# #### 1 Comments & Always commenting
#
# do good for yourself and others and always comment whenever you add something new to code.
# in Python, comment starts with # hash mark
#
# ``` # This is a comment. ```
#
# and
#
# ```python
# # This is
# # a multi-line
# # comment.
# ```
# you can also wrap your multi-line comment inside a set of triple quotes
#
# ```python
# """
# This is
# a multi-line
# comment.
# """
# ```
# but this isn’t technically a comment. It’s a string that’s not assigned to any variable!
# and remember where you place them as they could turn into *docstrings*, which are pieces of documentation that are associated with a function or method.
#
# let's test it
# -
# This is Hello, World!.
print("Hello, World!") # even here it won't be shown
# print("This sentence won't be shown up")
# ### run your first python script
#
# + active=""
# mohamed@fou4d-ubuntu:~$ which python3
# /usr/bin/python3
# -
# hello.py file content
# + active=""
# #!/usr/bin/env python3
#
# print("Hello World")
# + active=""
# mohamed@fou4d-ubuntu:~$ python3 hello.py
# Hello World
# mohamed@fou4d-ubuntu:~$
# -
# #### 2 Arithmetic Operators
#
# simple arithmetic operations can be done directly
# Addition
5 + 4
# Subtraction
2 - 12
# Multiplication
3 * 27
# power / Exponentiation
2 ** 4
# Division
5 / 2
# round down division
5 // 2
# Modulus
18 % 7
8 - 6 * 2 + (8/4) + 7
#13 <NAME>
#-13 fakhry
# 5 everyone
# order of Arithmetic operations in python follows PEMDAS role (Parentheses, Exponentiation, Multiplication and Division, Addition and Subtraction)
# +
# Order
10 / 5 * 6 - (1 + 2)
7.5%5
# -
# #### 3 Comparison Operators
#
# Comparison operators are used to compare two values:
# ```==, !=, <, >, <=, >=```
# and it returns ```True``` or ```False```
#
#
#
# Greater Than
10 > 50
# Lesser Than
11 < 9
# Equal
7 == 7
# Not Equal
10 != 8
# Greater than or equal to
9 >= 9
# Less than or equal to
10 <= 10
# Python has the following operators groups:
#
# * Arithmetic operators
# * Assignment operators
# * Bitwise operators
# * Comparison operators
# * Identity operators
# * Logical operators
# * Membership operators
#
# we will discuss each on time
# #### 4 Basic Statement
#
# **print statement**: outputs the contents inside the parenthesis whatever the data type ```print()```
print('Hello World \'ITI')
print('Hello World')
print(5+7)
print("welco\tme python")
print(12/7)
# **assignment statement**: to assign a value (on the right) to a variable (on the left): ```=```
x = 10
print(x)
# ### our first function
#
# during this section we will create our own function ```hello_world()``` that will return/print "Hello world" when it's called in the code
#
# it starts with ```def```
# Python function block begins with the defining keyword ```def``` followed by the function name and parentheses ```()``` and ends with a colon ```:```.
#
# *actually the code of block inside the function starts with colon*
#
# then the function code block is indented.
#
#
# ```python
# # comment to know what's this function about
# def functionname( parameters ): # parameters are optional
# # code
# # code
# # code outside the function
# ```
#
#
#
# now let's start by defining our fist function
#
# +
# function returning Hello World
def hello_world():
print("Hello, World!")
# + active=""
# running the code above will not return anythin cause we didn't call the function,
# now let's call it
# -
hello_world()
# ##### importance of indentation
#
# each block of code is indented by 1 tab to separate it for example:
#
# ```python
# def newfunctionname(): # first block
# # code here
# # second block of code
# # continue 2nd block of code
# # back to first block of code
# ```
#
# failing to indent the code block correctly will result in errors running your code.
# +
# function returning Hello python
def hello_python():
print("Hello, python!")
# -
# as you can see above it raises error message.
# let's dive more into types of errors
#
# always use 4 spaces for indentation.
#
# Using tabs exclusively is possible but PEP 8, the style guide for Python code, states that spaces are preferred.
#
#
# ##### Errors
#
#
# errors are common thing in your life as a programmer and Python programs will terminates once it encounters an error showing detailed error message for the error it stopped by showing a small arrow below where it detected the error.
#
# **syntax errors**
# they are errors shown when the language parser detects an incorrect statement
#
# **indentation error**: shown above
#
print('hello
world)
print(7))
# **Type Error**
a = "Hi"
b = 3.14
#a + b
print(a + 394)
## correctness
# print( a + ", Pi Equals: " + str(b))
# **Zero Division Error**
0 / 0
# ##### Python Keywords
#
# Function names should be lowercase, with words separated by underscores as necessary to improve readability. [PEP 8 - Style Guide for Python Code](https://www.python.org/dev/peps/pep-0008/#function-and-variable-names)
#
# but Python reserve Keywords that we cannot use as function name or any other identifier
#
# ```
# False await else import pass
# None break except in raise
# True class finally is return
# and continue for lambda try
# as def from nonlocal while
# assert del global not with
# async elif if or yield
# ```
# [Source: Python Docs](https://docs.python.org/3/reference/lexical_analysis.html#keywords)
await = 7
except = 75
# let's try another trick in Python Code Blocks
def hello_function():
print("Hello, from the function!")
print("This Code is outside the function")
hello_function()
help('print')
# -----
#
# ### Assignments
#
# #### 1 arithmetic
#
# in Statistics, the Mean value is calculated by dividing the sum of all values by the count of these values.
# calculate the mean of 5 numbers ``` 11, 7, 4, 18, 37 ```
#
( 11 + 7 + 4 + 18 + 37 ) / 5
# #### 2 what are the results
#
# ```python
# 52 + 41
# 12 - 4
# 4 + 10 - 5
# 42 / 2 + 7
# 12 + (45 - 3) - 2 / 4
# ```
# #### 3 comparison operators
#
# ```python
# 47 >= 27
# 84 == 61
# 1 != 8
#
# ```
# #### 4 what is the output
#
# ```python
# def lambda():
# print(0/0)
# ```
|
Session 01 - Basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 5
# ## Due Date: Tuesday, October 3rd at 11:59 PM
# # Problem 1
# We discussed documentation and testing in lecture and also briefly touched on code coverage. You must write tests for your code for your final project (and in life). There is a nice way to automate the testing process called continuous integration (CI).
#
# This problem will walk you through the basics of CI and show you how to get up and running with some CI software.
# ### Continuous Integration
# The idea behind continuous integration is to automate away the testing of your code.
#
# We will be using it for our projects.
#
# The basic workflow goes something like this:
#
# 1. You work on your part of the code in your own branch or fork
# 2. On every commit you make and push to GitHub, your code is automatically tested on a fresh machine on Travis CI. This ensures that there are no specific dependencies on the structure of your machine that your code needs to run and also ensures that your changes are sane
# 3. Now you submit a pull request to `master` in the main repo (the one you're hoping to contribute to). The repo manager creates a branch off `master`.
# 4. This branch is also set to run tests on Travis. If all tests pass, then the pull request is accepted and your code becomes part of master.
#
# We use GitHub to integrate our roots library with Travis CI and Coveralls. Note that this is not the only workflow people use. Google git..github..workflow and feel free to choose another one for your group.
# ### Part 1: Create a repo
# Create a public GitHub repo called `cs207test` and clone it to your local machine.
#
# **Note:** No need to do this in Jupyter.
# ### Part 2: Create a roots library
# Use the example from lecture 7 to create a file called `roots.py`, which contains the `quad_roots` and `linear_roots` functions (along with their documentation).
#
# Also create a file called `test_roots.py`, which contains the tests from lecture.
#
# All of these files should be in your newly created `cs207test` repo. **Don't push yet!!!**
# + language="bash"
# cd ../../../cs207test
# file roots.py
#
# def linear_roots(a=1.0, b=0.0):
# if a == 0:
# raise ValueError("The linear coefficient is zero. This is not a linear equation.")
# else:
# return ((-b / a))
#
# def quad_roots(a=1.0, b=2.0, c=0.0):
# import cmath # Can return complex numbers from square roots
# if a == 0:
# raise ValueError("The quadratic coefficient is zero. This is not a quadratic equation.")
# else:
# sqrtdisc = cmath.sqrt(b * b - 4.0 * a * c)
# r1 = -b + sqrtdisc
# r2 = -b - sqrtdisc
# return (r1 / 2.0 / a, r2 / 2.0 / a)
#
# -
# ### Part 3: Create an account on Travis CI and Start Building
#
# #### Part A:
# Create an account on Travis CI and set your `cs207test` repo up for continuous integration once this repo can be seen on Travis.
#
# #### Part B:
# Create an instruction to Travis to make sure that
#
# 1. python is installed
# 2. its python 3.5
# 3. pytest is installed
#
# The file should be called `.travis.yml` and should have the contents:
# ```yml
# language: python
# python:
# - "3.5"
# before_install:
# - pip install pytest pytest-cov
# script:
# - pytest
# ```
#
# You should also create a configuration file called `setup.cfg`:
# ```cfg
# [tool:pytest]
# addopts = --doctest-modules --cov-report term-missing --cov roots
# ```
#
# #### Part C:
# Push the new changes to your `cs207test` repo.
#
# At this point you should be able to see your build on Travis and if and how your tests pass.
# ### Part 4: Coveralls Integration
# In class, we also discussed code coverage. Just like Travis CI runs tests automatically for you, Coveralls automatically checks your code coverage. One minor drawback of Coveralls is that it can only work with public GitHub accounts. However, this isn't too big of a problem since your projects will be public.
#
# #### Part A:
# Create an account on [`Coveralls`](https://coveralls.zendesk.com/hc/en-us), connect your GitHub, and turn Coveralls integration on.
#
# #### Part B:
# Update your the `.travis.yml` file as follows:
# ```yml
# language: python
# python:
# - "3.5"
# before_install:
# - pip install pytest pytest-cov
# - pip install coveralls
# script:
# - py.test
# after_success:
# - coveralls
# ```
#
# Be sure to push the latest changes to your new repo.
# ### Part 5: Update README.md in repo
# You can have your GitHub repo reflect the build status on Travis CI and the code coverage status from Coveralls. To do this, you should modify the `README.md` file in your repo to include some badges. Put the following at the top of your `README.md` file:
#
# ```
# [](https://travis-ci.org/dsondak/cs207testing.svg?branch=master)
#
# [](https://coveralls.io/github/dsondak/cs207testing?branch=master)
# ```
#
# Of course, you need to make sure that the links are to your repo and not mine. You can find embed code on the Coveralls and Travis CI sites.
# ---
#
# # Problem 2
# Write a Python module for reaction rate coefficients. Your module should include functions for constant reaction rate coefficients, Arrhenius reaction rate coefficients, and modified Arrhenius reaction rate coefficients. Here are their mathematical forms:
# \begin{align}
# &k_{\textrm{const}} = k \tag{constant} \\
# &k_{\textrm{arr}} = A \exp\left(-\frac{E}{RT}\right) \tag{Arrhenius} \\
# &k_{\textrm{mod arr}} = A T^{b} \exp\left(-\frac{E}{RT}\right) \tag{Modified Arrhenius}
# \end{align}
#
# Test your functions with the following paramters: $A = 10^7$, $b=0.5$, $E=10^3$. Use $T=10^2$.
#
# A few additional comments / suggestions:
# * The Arrhenius prefactor $A$ is strictly positive
# * The modified Arrhenius parameter $b$ must be real
# * $R = 8.314$ is the ideal gas constant. It should never be changed (except to convert units)
# * The temperature $T$ must be positive (assuming a Kelvin scale)
# * You may assume that units are consistent
# * Document each function!
# * You might want to check for overflows and underflows
#
# **Recall:** A Python module is a `.py` file which is not part of the main execution script. The module contains several functions which may be related to each other (like in this problem). Your module will be importable via the execution script. For example, suppose you have called your module `reaction_coeffs.py` and your execution script `kinetics.py`. Inside of `kinetics.py` you will write something like:
# ```python
# import reaction_coeffs
# # Some code to do some things
# # :
# # :
# # :
# # Time to use a reaction rate coefficient:
# reaction_coeffs.const() # Need appropriate arguments, etc
# # Continue on...
# # :
# # :
# # :
# ```
# Be sure to include your module in the same directory as your execution script.
# ---
#
# # Problem 3
# Write a function that returns the **progress rate** for a reaction of the following form:
# \begin{align}
# \nu_{A} A + \nu_{B} B \longrightarrow \nu_{C} C.
# \end{align}
# Order your concentration vector so that
# \begin{align}
# \mathbf{x} =
# \begin{bmatrix}
# \left[A\right] \\
# \left[B\right] \\
# \left[C\right]
# \end{bmatrix}
# \end{align}
#
# Test your function with
# \begin{align}
# \nu_{i}^{\prime} =
# \begin{bmatrix}
# 2.0 \\
# 1.0 \\
# 0.0
# \end{bmatrix}
# \qquad
# \mathbf{x} =
# \begin{bmatrix}
# 1.0 \\
# 2.0 \\
# 3.0
# \end{bmatrix}
# \qquad
# k = 10.
# \end{align}
#
# You must document your function and write some tests in addition to the one suggested. You choose the additional tests, but you must have at least one doctest in addition to a suite of unit tests.
# ---
# # Problem 4
# Write a function that returns the **progress rate** for a system of reactions of the following form:
# \begin{align}
# \nu_{11}^{\prime} A + \nu_{21}^{\prime} B \longrightarrow \nu_{31}^{\prime\prime} C \\
# \nu_{12}^{\prime} A + \nu_{32}^{\prime} C \longrightarrow \nu_{22}^{\prime\prime} B + \nu_{32}^{\prime\prime} C
# \end{align}
# Note that $\nu_{ij}^{\prime}$ represents the stoichiometric coefficient of reactant $i$ in reaction $j$ and $\nu_{ij}^{\prime\prime}$ represents the stoichiometric coefficient of product $i$ in reaction $j$. Therefore, in this convention, I have ordered my vector of concentrations as
# \begin{align}
# \mathbf{x} =
# \begin{bmatrix}
# \left[A\right] \\
# \left[B\right] \\
# \left[C\right]
# \end{bmatrix}.
# \end{align}
#
# Test your function with
# \begin{align}
# \nu_{ij}^{\prime} =
# \begin{bmatrix}
# 1.0 & 2.0 \\
# 2.0 & 0.0 \\
# 0.0 & 2.0
# \end{bmatrix}
# \qquad
# \nu_{ij}^{\prime\prime} =
# \begin{bmatrix}
# 0.0 & 0.0 \\
# 0.0 & 1.0 \\
# 2.0 & 1.0
# \end{bmatrix}
# \qquad
# \mathbf{x} =
# \begin{bmatrix}
# 1.0 \\
# 2.0 \\
# 1.0
# \end{bmatrix}
# \qquad
# k_{j} = 10, \quad j=1,2.
# \end{align}
#
# You must document your function and write some tests in addition to the one suggested. You choose the additional tests, but you must have at least one doctest in addition to a suite of unit tests.
# ---
# # Problem 5
# Write a function that returns the **reaction rate** of a system of irreversible reactions of the form:
# \begin{align}
# \nu_{11}^{\prime} A + \nu_{21}^{\prime} B &\longrightarrow \nu_{31}^{\prime\prime} C \\
# \nu_{32}^{\prime} C &\longrightarrow \nu_{12}^{\prime\prime} A + \nu_{22}^{\prime\prime} B
# \end{align}
#
# Once again $\nu_{ij}^{\prime}$ represents the stoichiometric coefficient of reactant $i$ in reaction $j$ and $\nu_{ij}^{\prime\prime}$ represents the stoichiometric coefficient of product $i$ in reaction $j$. In this convention, I have ordered my vector of concentrations as
# \begin{align}
# \mathbf{x} =
# \begin{bmatrix}
# \left[A\right] \\
# \left[B\right] \\
# \left[C\right]
# \end{bmatrix}
# \end{align}
#
# Test your function with
# \begin{align}
# \nu_{ij}^{\prime} =
# \begin{bmatrix}
# 1.0 & 0.0 \\
# 2.0 & 0.0 \\
# 0.0 & 2.0
# \end{bmatrix}
# \qquad
# \nu_{ij}^{\prime\prime} =
# \begin{bmatrix}
# 0.0 & 1.0 \\
# 0.0 & 2.0 \\
# 1.0 & 0.0
# \end{bmatrix}
# \qquad
# \mathbf{x} =
# \begin{bmatrix}
# 1.0 \\
# 2.0 \\
# 1.0
# \end{bmatrix}
# \qquad
# k_{j} = 10, \quad j = 1,2.
# \end{align}
#
# You must document your function and write some tests in addition to the one suggested. You choose the additional tests, but you must have at least one doctest in addition to a suite of unit tests.
# +
# %%file reaction_coeffs.py
from math import exp
def k_constant(k):
return k
def k_arr(a,e,t):
r=8.314
if a<= 0:
raise ValueError("a<0!")
if t<= 0:
raise ValueError("t<0!")
return (a*exp(-e/(r*t)))
def k_mod_arr(a,e,t,b):
r=8.314
if a<= 0:
raise ValueError("a<0!")
if t<= 0:
raise ValueError("t<0!")
return (a*(t**b)*exp(-e/(r*t)))
# +
# %%file reaction1_rate.py
import numpy as np
import copy
def reaction_rate1(x,v,vv,k):
"""Returns reaction rate of chemical reactoins
INPUTS
=======
x: array of float
concentration of molecule species
v: array of integers
Coefficient of molecule species on the left side of equation
vv: array of integers
Coefficient of melecule species on the right side of equation
kk: array of float
reaction rate coefficient
RETURNS
========
roots: array of floats
EXAMPLES
=========
>>> reaction_rate1(np.array([[1.0],[2.0],[1]]),np.array([[1,0],[2,0],[0,2]]),np.array([[0,1],[0,2],[1,0]]),[10,10])
array([[-40.],
[ 10.],
[-80.],
[ 20.],
[ 40.],
[-20.]])
"""
if (any([kk<0 for kk in k])):
raise ValueError("k<0")
flag=True
for j in range(v.shape[1]):
if all([v[i][j]<=0 for i in range(v.shape[0])]):
flag=False
break
if (flag==False):
raise ValueError("no reactants")
flag=True
for j in range(vv.shape[1]):
if all([vv[i][j]<=0 for i in range(vv.shape[0])]):
flag=False
break
if (flag==False):
raise ValueError("no products")
if x.shape[0]!=v.shape[0] or v.shape!=vv.shape or v.shape[1]!=len(k):
raise ValueError("dimensions not match")
tmp=np.array([x[i][0]**v[i][j] for i in range(v.shape[0]) for j in range(v.shape[1])]).reshape(v.shape)
w=copy.copy(k)
for i in range(len(x)):
for j in range(v.shape[1]):
a=tmp[i][j]
w[j]*=a
return (np.transpose([np.array([w[j]*(vv[i][j]-v[i][j]) for i in range(len(x)) for j in range(v.shape[1])])]))
# +
# %%file reaction1_test.py
import reaction1_rate as rr1
import numpy as np
def test_normal1():
x=np.array([[1.0],[2.0],[1]])
v_i_prime=np.array([[1,0],[2,0],[0,2]])
v_i_prime_prime=np.array([[0,1],[0,2],[1,0]])
k=[10,10]
assert all(rr1.reaction_rate1(x,v_i_prime,v_i_prime_prime,k)==[[-40],[10],[-80],[20],[40],[-20]])
def test_nagative_k1():
try:
x=np.array([[1.0],[2.0],[1]])
v_i_prime=np.array([[1,2],[2,0],[0,2]])
v_i_prime_prime=np.array([[0,0],[0,1],[2,1]])
k=[-10,10]
rr1.reaction_rate1(x,v_i_prime,v_i_prime_prime,k)
except ValueError as err:
assert(True)
def test_reactants1():
try:
x=np.array([[1.0],[2.0],[1]])
v_i_prime=np.array([[0,2],[0,0],[0,2]])
v_i_prime_prime=np.array([[0,0],[0,1],[2,1]])
k=[10,10]
rr1.reaction_rate1(x,v_i_prime,v_i_prime_prime,k)
except ValueError as err:
assert(True)
def test_products1():
try:
x=np.array([[1.0],[2.0],[1]])
v_i_prime=np.array([[1,2],[2,0],[0,2]])
v_i_prime_prime=np.array([[0,0],[0,0],[2,0]])
k=[10,10]
rr1.reaction_rate1(x,v_i_prime,v_i_prime_prime,k)
except ValueError as err:
assert(True)
def test_dimensions1():
try:
x=np.array([[1.0],[2.0]])
v_i_prime=np.array([[1,2],[2,0],[0,2]])
v_i_prime_prime=np.array([[0,0],[0,0],[2,0]])
k=[10,10]
rr1.reaction_rate1(x,v_i_prime,v_i_prime_prime,k)
except ValueError as err:
assert(True)
# -
import doctest
doctest.testmod(verbose=True)
# !pytest
# + language="bash"
# pwd
# -
# !pytest --cov
# ---
# # Problem 6
# Put parts 3, 4, and 5 in a module called `chemkin`.
#
# Next, pretend you're a client who needs to compute the reaction rates at three different temperatures ($T = \left\{750, 1500, 2500\right\}$) of the following system of irreversible reactions:
# \begin{align}
# 2H_{2} + O_{2} \longrightarrow 2OH + H_{2} \\
# OH + HO_{2} \longrightarrow H_{2}O + O_{2} \\
# H_{2}O + O_{2} \longrightarrow HO_{2} + OH
# \end{align}
#
# The client also happens to know that reaction 1 is a modified Arrhenius reaction with $A_{1} = 10^{8}$, $b_{1} = 0.5$, $E_{1} = 5\times 10^{4}$, reaction 2 has a constant reaction rate parameter $k = 10^{4}$, and reaction 3 is an Arrhenius reaction with $A_{3} = 10^{7}$ and $E_{3} = 10^{4}$.
#
# You should write a script that imports your `chemkin` module and returns the reaction rates of the species at each temperature of interest given the following species concentrations:
#
# \begin{align}
# \mathbf{x} =
# \begin{bmatrix}
# H_{2} \\
# O_{2} \\
# OH \\
# HO_{2} \\
# H_{2}O
# \end{bmatrix} =
# \begin{bmatrix}
# 2.0 \\
# 1.0 \\
# 0.5 \\
# 1.0 \\
# 1.0
# \end{bmatrix}
# \end{align}
#
# You may assume that these are elementary reactions.
# ---
# # Problem 7
# Get together with your project team, form a GitHub organization (with a descriptive team name), and give the teaching staff access. You can have has many repositories as you like within your organization. However, we will grade the repository called **`cs207-FinalProject`**.
#
# Within the `cs207-FinalProject` repo, you must set up Travis CI and Coveralls. Make sure your `README.md` file includes badges indicating how many tests are passing and the coverage of your code.
|
homeworks/HW5/HW5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Caculate F1_Score
# +
import numpy as np
import sklearn.metrics as metrics
# +
y = np.array([1,1,1,1,0,0]) #0은 정상(실력자), 1은 음치
p = np.array([1,1,0,0,0,0]) #나의 예측
# -
y
y.shape
# accuracy = np.mean(np.equal(y,p))
# right = np.sum(y * p == 1)
# precision = right / np.sum(p)
# recall = right / np.sum(y)
# f1 = 2 * precision*recall/(precision+recall)
#
# print('accuracy',accuracy)
# print('precision', precision)
# print('recall', recall)
# print('f1_Score', f1)
# ### Method
# +
print('accuracy', metrics.accuracy_score(y,p) )
print('precision', metrics.precision_score(y,p) )
print('recall', metrics.recall_score(y,p) )
print('f1', metrics.f1_score(y,p) )
print(metrics.classification_report(y,p))
print(metrics.confusion_matrix(y,p))
# -
|
Python/numpy/1-F1_score.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Plot with Floors
# ================
#
# Add a floor/wall at the boundary of the rendering scene.
#
# +
import pyvista as pv
from pyvista import examples
mesh = examples.download_dragon()
plotter = pv.Plotter()
plotter.add_mesh(mesh)
plotter.add_floor('-y')
plotter.add_floor('-z')
plotter.show()
|
examples/02-plot/floors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Download** (right-click, save target as ...) this page as a jupyterlab notebook from: [Lab18](http://192.168.127.12/engr-1330-webroot/8-Labs/Lab18/Lab18.ipynb)
#
# ___
# # <font color=darkgreen>Laboratory 18: Causality, Simulation, and Probability </font>
#
# LAST NAME, FIRST NAME
#
# R00000000
#
# ENGR 1330 Laboratory 14 - In-Lab
#
# ---
#
#
# ## Exercise 0 *Profile your computer*
#
# Execute the cell below
# Preamble script block to identify host, user, and kernel
import sys
# ! hostname
# ! whoami
print(sys.executable)
print(sys.version)
print(sys.version_info)
# ---
#
# # <font color=purple>Python for Simulation</font>
# ## What is Russian roulette?
# >Russian roulette (Russian: русская рулетка, russkaya ruletka) is a lethal game of chance in which a player places a single round in a revolver, spins the cylinder, places the muzzle against their head, and pulls the trigger in hopes that the loaded chamber does not align with the primer percussion mechanism and the barrel, causing the weapon to discharge. Russian refers to the supposed country of origin, and roulette to the element of risk-taking and the spinning of the revolver's cylinder, which is reminiscent of a spinning roulette wheel. <br>
# - Wikipedia @ https://en.wikipedia.org/wiki/Russian_roulette
# 
# >A game of dafts, a game of chance <br>
# One where revolver's the one to dance <br>
# Rounds and rounds, it goes and spins <br>
# Makes you regret all those sins <br> \
# A game of fools, one of lethality <br>
# With a one to six probability <br>
# There were two guys and a gun <br>
# With six chambers but only one... <br> \
# CLICK, one pushed the gun <br>
# CLICK, one missed the fun <br>
# CLICK, "that awful sound" ... <br>
# BANG!, one had his brains all around! <br>
# ___
# ### Example: Simulate a game of Russian Roulette:
# - For 2 rounds
# - For 5 rounds
# - For 10 rounds
# + jupyter={"outputs_hidden": false}
import numpy as np #import numpy
revolver = np.array([1,0,0,0,0,0]) #create a numpy array with 1 bullet and 5 empty chambers
print(np.random.choice(revolver,2)) #randomly select a value from revolver - simulation
# -
print(np.random.choice(revolver,2))
print(np.random.choice(revolver,2))
print(np.random.choice(revolver,2))
# + jupyter={"outputs_hidden": false}
print(np.random.choice(revolver,5))
# + jupyter={"outputs_hidden": false}
print(np.random.choice(revolver,10))
# -
# 
# ___
# ### Example: Simulate the results of throwing a D6 (regular dice) for 10 times.
# + jupyter={"outputs_hidden": false}
import numpy as np #import numpy
dice = np.array([1,2,3,4,5,6]) #create a numpy array with values of a D6
np.random.choice(dice,10) #randomly selecting a value from dice for 10 times- simulation
# -
# ___
# ### Example: Assume the following rules:
#
# - If the dice shows 1 or 2 spots, my net gain is -1 dollar.
#
# - If the dice shows 3 or 4 spots, my net gain is 0 dollars.
#
# - If the dice shows 5 or 6 spots, my net gain is 1 dollar.
#
# __Define a function to simulate a game with the above rules, assuming a D6, and compute the net gain of the player over any given number of rolls. <br>
# Compute the net gain for 5, 50, and 500 rolls__
# + jupyter={"outputs_hidden": false}
def D6game(nrolls):
import numpy as np #import numpy
dice = np.array([1,2,3,4,5,6]) #create a numpy array with values of a D6
rolls = np.random.choice(dice,nrolls) #randomly selecting a value from dice for nrolls times- simulation
gainlist =[] #create an empty list for gains|losses
for i in np.arange(len(rolls)): #Apply the rules
if rolls[i]<=2:
gainlist.append(-1)
elif rolls[i]<=4:
gainlist.append(0)
elif rolls[i]<=6:
gainlist.append(+1)
return (np.sum(gainlist)) #sum up all gains|losses
# return (gainlist,"The net gain is equal to:",np.sum(gainlist))
# + jupyter={"outputs_hidden": false}
D6game(2)
# + jupyter={"outputs_hidden": false}
D6game(50)
# + jupyter={"outputs_hidden": false}
D6game(500)
# -
# 
# ### Let's Make A Deal Game Show and Monty Hall Problem
# __The Monty Hall problem is a brain teaser, in the form of a probability puzzle, loosely based on the American television game show Let's Make a Deal and named after its original host, <NAME>. The problem was originally posed (and solved) in a letter by <NAME> to the American Statistician in 1975 (Selvin 1975a), (Selvin 1975b).__
#
# >"Suppose you're on a game show, and you're given the choice of three doors: Behind one door is a car; behind the others, goats. You pick a door, say No. 1, and the host, who knows what's behind the doors, opens another door, say No. 3, which has a goat. He then says to you, "Do you want to pick door No. 2?" Is it to your advantage to switch your choice?"
#
# __*From Wikipedia: https://en.wikipedia.org/wiki/Monty_Hall_problem*__
# 
# /data/img1.png)
# 
# 
# ___
# ### Example: Simulate Monty Hall Game for 1000 times. Use a barplot and discuss whether players are better off sticking to their initial choice, or switching doors?
# + jupyter={"outputs_hidden": false}
def othergoat(x): #Define a function to return "the other goat"!
if x == "Goat 1":
return "Goat 2"
elif x == "Goat 2":
return "Goat 1"
# + jupyter={"outputs_hidden": false}
Doors = np.array(["Car","Goat 1","Goat 2"]) #Define a list for objects behind the doors
goats = np.array(["Goat 1" , "Goat 2"]) #Define a list for goats!
def MHgame():
#Function to simulate the Monty Hall Game
#For each guess, return ["the guess","the revealed", "the remaining"]
userguess=np.random.choice(Doors) #randomly selects a door as userguess
if userguess == "Goat 1":
return [userguess, "Goat 2","Car"]
if userguess == "Goat 2":
return [userguess, "Goat 1","Car"]
if userguess == "Car":
revealed = np.random.choice(goats)
return [userguess, revealed,othergoat(revealed)]
# + jupyter={"outputs_hidden": false}
# Check and see if the MHgame function is doing what it is supposed to do:
for i in np.arange(1):
a =MHgame()
print(a)
print(a[0])
print(a[1])
print(a[2])
# + jupyter={"outputs_hidden": false}
c1 = [] #Create an empty list for the userguess
c2 = [] #Create an empty list for the revealed
c3 = [] #Create an empty list for the remaining
for i in np.arange(1000): #Simulate the game for 1000 rounds - or any other number of rounds you desire
game = MHgame()
c1.append(game[0]) #In each round, add the first element to the userguess list
c2.append(game[1]) #In each round, add the second element to the revealed list
c3.append(game[2]) #In each round, add the third element to the remaining list
# + jupyter={"outputs_hidden": false}
import pandas as pd
#Create a data frame (gamedf) with 3 columns ("Guess","Revealed", "Remaining") and 1000 (or how many number of rounds) rows
gamedf = pd.DataFrame({'Guess':c1,
'Revealed':c2,
'Remaining':c3})
gamedf
# + jupyter={"outputs_hidden": false}
# Get the count of each item in the first and 3rd column
original_car =gamedf[gamedf.Guess == 'Car'].shape[0]
remaining_car =gamedf[gamedf.Remaining == 'Car'].shape[0]
original_g1 =gamedf[gamedf.Guess == 'Goat 1'].shape[0]
remaining_g1 =gamedf[gamedf.Remaining == 'Goat 1'].shape[0]
original_g2 =gamedf[gamedf.Guess == 'Goat 2'].shape[0]
remaining_g2 =gamedf[gamedf.Remaining == 'Goat 2'].shape[0]
# + jupyter={"outputs_hidden": false}
# Let's plot a grouped barplot
import matplotlib.pyplot as plt
# set width of bar
barWidth = 0.25
# set height of bar
bars1 = [original_car,original_g1,original_g2]
bars2 = [remaining_car,remaining_g1,remaining_g2]
# Set position of bar on X axis
r1 = np.arange(len(bars1))
r2 = [x + barWidth for x in r1]
# Make the plot
plt.bar(r1, bars1, color='darkorange', width=barWidth, edgecolor='white', label='Original Guess')
plt.bar(r2, bars2, color='midnightblue', width=barWidth, edgecolor='white', label='Remaining Door')
# Add xticks on the middle of the group bars
plt.xlabel('Item', fontweight='bold')
plt.xticks([r + barWidth/2 for r in range(len(bars1))], ['Car', 'Goat 1', 'Goat 2'])
# Create legend & Show graphic
plt.legend()
plt.show()
# -
# <font color=crimson>__According to the plot, it is statitically beneficial for the players to switch doors because the initial chance for being correct is only 1/3__</font>
# 
# # <font color=purple>Python for Probability</font>
# 
# ### <font color=purple>Important Terminology:</font>
#
# __Experiment:__ An occurrence with an uncertain outcome that we can observe. <br>
# *For example, rolling a die.*<br>
# __Outcome:__ The result of an experiment; one particular state of the world. What Laplace calls a "case."<br>
# *For example: 4.*<br>
# __Sample Space:__ The set of all possible outcomes for the experiment.<br>
# *For example, {1, 2, 3, 4, 5, 6}.*<br>
# __Event:__ A subset of possible outcomes that together have some property we are interested in.<br>
# *For example, the event "even die roll" is the set of outcomes {2, 4, 6}.*<br>
# __Probability:__ As Laplace said, the probability of an event with respect to a sample space is the number of favorable cases (outcomes from the sample space that are in the event) divided by the total number of cases in the sample space. (This assumes that all outcomes in the sample space are equally likely.) Since it is a ratio, probability will always be a number between 0 (representing an impossible event) and 1 (representing a certain event).<br>
# *For example, the probability of an even die roll is 3/6 = 1/2.*<br>
#
# __*From https://people.math.ethz.ch/~jteichma/probability.html*__
# + jupyter={"outputs_hidden": false}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
# ___
# ### Example: In a game of Russian Roulette, the chance of surviving each round is 5/6 which is almost 83%. Using a for loop, compute probability of surviving
# - For 2 rounds
# - For 5 rounds
# - For 10 rounds
# + jupyter={"outputs_hidden": false}
nrounds =[]
probs =[]
for i in range(3):
nrounds.append(i)
probs.append((5/6)**i) #probability of surviving- not getting the bullet!
RRDF = pd.DataFrame({"# of Rounds": nrounds, "Probability of Surviving": probs})
RRDF
# + jupyter={"outputs_hidden": false}
nrounds =[]
probs =[]
for i in range(6):
nrounds.append(i)
probs.append((5/6)**i) #probability of surviving- not getting the bullet!
RRDF = pd.DataFrame({"# of Rounds": nrounds, "Probability of Surviving": probs})
RRDF
# + jupyter={"outputs_hidden": false}
nrounds =[]
probs =[]
for i in range(11):
nrounds.append(i)
probs.append((5/6)**i) #probability of surviving- not getting the bullet!
RRDF = pd.DataFrame({"# of Rounds": nrounds, "Probability of Surviving": probs})
RRDF
# + jupyter={"outputs_hidden": false}
RRDF.plot.scatter(x="# of Rounds", y="Probability of Surviving",color="red")
# -
# ___
# ### Example: What will be the probability of constantly throwing an even number with a D20 in
# - For 2 rolls
# - For 5 rolls
# - For 10 rolls
# - For 15 rolls
# + jupyter={"outputs_hidden": false}
nrolls =[]
probs =[]
for i in range(1,16,1):
nrolls.append(i)
probs.append((1/2)**i) #probability of throwing an even number-10/20 or 1/2
DRDF = pd.DataFrame({"# of Rolls": nrolls, "Probability of constantly throwing an even number": probs})
DRDF
# + jupyter={"outputs_hidden": false}
DRDF.plot.scatter(x="# of Rolls", y="Probability of constantly throwing an even number",color="crimson")
# -
# ___
# ### Example: What will be the probability of throwing at least one 6 with a D6:
# - For 2 rolls
# - For 5 rolls
# - For 10 rolls
# - For 50 rolls - Make a scatter plot for this one!
# + jupyter={"outputs_hidden": false}
nRolls =[]
probs =[]
for i in range(1,51,1):
nRolls.append(i)
probs.append(1-(5/6)**i) #probability of at least one 6: 1-(5/6)
rollsDF = pd.DataFrame({"# of Rolls": nRolls, "Probability of rolling at least one 6": probs})
rollsDF
# + jupyter={"outputs_hidden": false}
nRolls =[]
probs =[]
for i in range(1,6,1):
nRolls.append(i)
probs.append(1-(5/6)**i) #probability of at least one 6: 1-(5/6)
rollsDF = pd.DataFrame({"# of Rolls": nRolls, "Probability of rolling at least one 6": probs})
rollsDF
# + jupyter={"outputs_hidden": false}
nRolls =[]
probs =[]
for i in range(1,11,1):
nRolls.append(i)
probs.append(1-(5/6)**i) #probability of at least one 6: 1-(5/6)
rollsDF = pd.DataFrame({"# of Rolls": nRolls, "Probability of rolling at least one 6": probs})
rollsDF
# + jupyter={"outputs_hidden": false}
nRolls =[]
probs =[]
for i in range(1,51,1):
nRolls.append(i)
probs.append(1-(5/6)**i) #probability of at least one 6: 1-(5/6)
rollsDF = pd.DataFrame({"# of Rolls": nRolls, "Probability of rolling at least one 6": probs})
# + jupyter={"outputs_hidden": false}
rollsDF.plot.scatter(x="# of Rolls", y="Probability of rolling at least one 6")
# -
# ___
# ### Example: What is the probability of drawing an ace at least once (with replacement):
# - in 2 tries
# - in 5 tries
# - in 10 tries
# - in 20 tries - make a scatter plot.
#
# + jupyter={"outputs_hidden": false}
nDraws =[]
probs =[]
for i in range(1,100,1):
nDraws.append(i)
probs.append(1-(48/52)**i) #probability of drawing an ace least once : 1-(48/52)
DrawsDF = pd.DataFrame({"# of Draws": nDraws, "Probability of drawing an ace at least once": probs})
DrawsDF
# + jupyter={"outputs_hidden": false}
nDraws =[]
probs =[]
for i in range(1,6,1):
nDraws.append(i)
probs.append(1-(48/52)**i) #probability of drawing an ace least once : 1-(48/52)
DrawsDF = pd.DataFrame({"# of Draws": nDraws, "Probability of drawing an ace at least once": probs})
DrawsDF
# + jupyter={"outputs_hidden": false}
nDraws =[]
probs =[]
for i in range(1,11,1):
nDraws.append(i)
probs.append(1-(48/52)**i) #probability of drawing an ace least once : 1-(48/52)
DrawsDF = pd.DataFrame({"# of Draws": nDraws, "Probability of drawing an ace at least once": probs})
DrawsDF
# + jupyter={"outputs_hidden": false}
nDraws =[]
probs =[]
for i in range(1,21,1):
nDraws.append(i)
probs.append(1-(48/52)**i) #probability of drawing an ace at least once : 1-(48/52)
DrawsDF = pd.DataFrame({"# of Draws": nDraws, "Probability of drawing an ace at least once": probs})
DrawsDF
# + jupyter={"outputs_hidden": false}
DrawsDF.plot.scatter(x="# of Draws", y="Probability of drawing an ace at least once")
# -
# ___
# ### Example:
# - A) Write a function to find the probability of an event in percentage form based on given outcomes and sample space
# - B) Use the function and compute the probability of rolling a 4 with a D6
# - C) Use the function and compute the probability of drawing a King from a standard deck of cards
# - D) Use the function and compute the probability of drawing the King of Hearts from a standard deck of cards
# - E) Use the function and compute the probability of drawing an ace after drawing a king
# - F) Use the function and compute the probability of drawing an ace after drawing an ace
# - G) Use the function and compute the probability of drawing a heart OR a club
# - F) Use the function and compute the probability of drawing a Royal Flush <br>
# *hint: (in poker) a straight flush including ace, king, queen, jack, and ten all in the same suit, which is the hand of the highest possible value
#
# __This problem is designed based on an example by *<NAME>* from DataCamp, accessible @ *https://www.datacamp.com/community/tutorials/statistics-python-tutorial-probability-1*__
# + jupyter={"outputs_hidden": false}
# A
# Create function that returns probability percent rounded to one decimal place
def Prob(outcome, sampspace):
probability = (outcome / sampspace) * 100
return round(probability, 1)
# + jupyter={"outputs_hidden": false}
# B
outcome = 1 #Rolling a 4 is only one of the possible outcomes
space = 6 #Rolling a D6 can have 6 different outcomes
Prob(outcome, space)
# + jupyter={"outputs_hidden": false}
# C
outcome = 4 #Drawing a king is four of the possible outcomes
space = 52 #Drawing from a standard deck of cards can have 52 different outcomes
Prob(outcome, space)
# + jupyter={"outputs_hidden": false}
# D
outcome = 1 #Drawing the king of hearts is only 1 of the possible outcomes
space = 52 #Drawing from a standard deck of cards can have 52 different outcomes
Prob(outcome, space)
# + jupyter={"outputs_hidden": false}
# E
outcome = 4 #Drawing an ace is 4 of the possible outcomes
space = 51 #One card has been drawn
Prob(outcome, space)
# + jupyter={"outputs_hidden": false}
# F
outcome = 3 #Once Ace is already drawn
space = 51 #One card has been drawn
Prob(outcome, space)
# + jupyter={"outputs_hidden": false}
# G
hearts = 13 #13 cards of hearts in a deck
space = 52 #total number of cards in a deck
clubs = 13 #13 cards of clubs in a deck
Prob_heartsORclubs= Prob(hearts, space) + Prob(clubs, space)
print("Probability of drawing a heart or a club is",Prob_heartsORclubs,"%")
# + jupyter={"outputs_hidden": false}
# F
draw1 = 5 #5 cards are needed
space1 = 52 #out of the possible 52 cards
draw2 = 4 #4 cards are needed
space2 = 51 #out of the possible 51 cards
draw3 = 3 #3 cards are needed
space3 = 50 #out of the possible 50 cards
draw4 = 2 #2 cards are needed
space4 = 49 #out of the possible 49 cards
draw5 = 1 #1 cards is needed
space5 = 48 #out of the possible 48 cards
#Probability of a getting a Royal Flush
Prob_RF= 4*(Prob(draw1, space1)/100) * (Prob(draw2, space2)/100) * (Prob(draw3, space3)/100) * (Prob(draw4, space4)/100) * (Prob(draw5, space5)/100)
print("Probability of drawing a royal flush is",Prob_RF,"%")
# -
# ___
# ### Example: Two unbiased dice are thrown once and the total score is observed. Define an appropriate function and use a simulation to find the estimated probability that :
# - the total score is greater than 10?
# - the total score is even and greater than 7?
#
#
# __This problem is designed based on an example by *<NAME>*
# from Medium.com, accessible @ *https://medium.com/future-vision/simulating-probability-events-in-python-5dd29e34e381*__
# + jupyter={"outputs_hidden": false}
import numpy as np
def DiceRoll1(nSimulation):
count =0
dice = np.array([1,2,3,4,5,6]) #create a numpy array with values of a D6
for i in range(nSimulation):
die1 = np.random.choice(dice,1) #randomly selecting a value from dice - throw the D6 once
die2 = np.random.choice(dice,1) #randomly selecting a value from dice - throw the D6 once again!
score = die1 + die2 #summing them up
if score > 10: #if it meets our desired condition:
count +=1 #add one to the "count"
return count/nSimulation #compute the probability of the desired event by dividing count by the total number of trials
nSimulation = 10000
print("The probability of rolling a number greater than 10 after",nSimulation,"rolld is:",DiceRoll1(nSimulation)*100,"%")
# + jupyter={"outputs_hidden": false}
import numpy as np
def DiceRoll2(nSimulation):
count =0
dice = np.array([1,2,3,4,5,6]) #create a numpy array with values of a D6
for i in range(nSimulation):
die1 = np.random.choice(dice,1) #randomly selecting a value from dice - throw the D6 once
die2 = np.random.choice(dice,1) #randomly selecting a value from dice - throw the D6 once again!
score = die1 + die2
if score %2 ==0 and score > 7: #the total score is even and greater than 7
count +=1
return count/nSimulation
nSimulation = 10000
print("The probability of rolling an even number and greater than 7 after",nSimulation," rolls is:",DiceRoll2(nSimulation)*100,"%")
# -
# ___
# ### Example: An urn contains 10 white balls, 20 reds and 30 greens. We want to draw 5 balls with replacement. Use a simulation (10000 trials) to find the estimated probability that:
# - we draw 3 white and 2 red balls
# - we draw 5 balls of the same color
#
#
# __This problem is designed based on an example by *<NAME>*
# from Medium.com, accessible @ *https://medium.com/future-vision/simulating-probability-events-in-python-5dd29e34e381*__
# + jupyter={"outputs_hidden": false}
# A
import numpy as np
import random
d = {} #Create an empty dictionary to associate numbers and colors
for i in range(0,60,1): #total of 60 balls
if i <10: #10 white balls
d[i]="White"
elif i>9 and i<30: #20 red balls
d[i]="Red"
else: #60-30=30 green balls
d[i]="Green"
#
nSimulation= 100000 #How many trials?
outcome1= 0 #initial value on the desired outcome counter
for i in range(nSimulation):
draw=[] #an empty list for the draws
for i in range(5): #how many balls we want to draw?
draw.append(d[random.randint(0,59)]) #randomly choose a number from 0 to 59- simulation of drawing balls
drawarray = np.array(draw) #convert the list into a numpy array
white = sum(drawarray== "White") #count the white balls
red = sum(drawarray== "Red") #count the red balls
green = sum(drawarray== "Green") #count the green balls
if white ==3 and red==2: #If the desired condition is met, add one to the counter
outcome1 +=1
print("The probability of drawing 3 white and 2 red balls is",(outcome1/nSimulation)*100,"%")
# + jupyter={"outputs_hidden": false}
# B
import numpy as np
import random
d = {}
for i in range(0,60,1):
if i <10:
d[i]="White"
elif i>9 and i<30:
d[i]="Red"
else:
d[i]="Green"
#
nSimulation= 10000
outcome1= 0
outcome2= 0 #we can consider multiple desired outcomes
for i in range(nSimulation):
draw=[]
for i in range(5):
draw.append(d[random.randint(0,59)])
drawarray = np.array(draw)
white = sum(drawarray== "White")
red = sum(drawarray== "Red")
green = sum(drawarray== "Green")
if white ==3 and red==2:
outcome1 +=1
if white ==5 or red==5 or green==5:
outcome2 +=1
print("The probability of drawing 3 white and 2 red balls is",(outcome1/nSimulation)*100,"%")
print("The probability of drawing 5 balls of the same color is",(outcome2/nSimulation)*100,"%")
# -
# ___
#  <br>
#
# *Here are some of the resources used for creating this notebook:*
#
#
# - __"Poker Probability and Statistics with Python"__ by __<NAME>__ available at *https://www.datacamp.com/community/tutorials/statistics-python-tutorial-probability-1*<br>
# - __"Simulating probability events in Python"__ by __<NAME>__ available at *https://medium.com/future-vision/simulating-probability-events-in-python-5dd29e34e381*<br>
#
#
# *Here are some great reads on this topic:*
# - __"Simulate the Monty Hall Problem Using Python"__ by __randerson112358__ available at *https://medium.com/swlh/simulate-the-monty-hall-problem-using-python-7b76b943640e* <br>
# - __"The Monty Hall problem"__ available at *https://scipython.com/book/chapter-4-the-core-python-language-ii/examples/the-monty-hall-problem/*<br>
# - __"Introduction to Probability Using Python"__ by __<NAME>__ available at *https://medium.com/future-vision/simulating-probability-events-in-python-5dd29e34e381* <br>
# - __"Introduction to probability and statistics for Data Scientists and machine learning using python : Part-1"__ by __<NAME>__ available at *https://medium.com/@anayan/introduction-to-probability-and-statistics-for-data-scientists-and-machine-learning-using-python-377a9b082487*<br>
#
# *Here are some great videos on these topics:*
# - __"Monty Hall Problem - Numberphile"__ by __Numberphile__ available at *https://www.youtube.com/watch?v=4Lb-6rxZxx0* <br>
# - __"The Monty Hall Problem"__ by __D!NG__ available at *https://www.youtube.com/watch?v=TVq2ivVpZgQ* <br>
# - __"21 - Monty Hall - PROPENSITY BASED THEORETICAL MODEL PROBABILITY - MATHEMATICS in the MOVIES"__ by __Motivating Mathematical Education and STEM__ available at *https://www.youtube.com/watch?v=iBdjqtR2iK4* <br>
# - __"The Monty Hall Problem"__ by __niansenx__ available at *https://www.youtube.com/watch?v=mhlc7peGlGg* <br>
# - __"The Monty Hall Problem - Explained"__ by __AsapSCIENCE__ available at *https://www.youtube.com/watch?v=9vRUxbzJZ9Y* <br>
# - __"Introduction to Probability | 365 Data Science Online Course"__ by __365 Data Science__ available at *https://www.youtube.com/watch?v=soZRfdnkUQg* <br>
# - __"Probability explained | Independent and dependent events | Probability and Statistics | Khan Academy"__ by __Khan Academy__ available at *https://www.youtube.com/watch?v=uzkc-qNVoOk* <br>
# - __"Math Antics - Basic Probability"__ by __mathantics__ available at *https://www.youtube.com/watch?v=KzfWUEJjG18* <br>
# ___
#  <br>
#
# ## Exercise 1: Risk or Probability <br>
#
# ### Are they the same? Are they different? Discuss your opinion.
#
# #### _Make sure to cite any resources that you may use._
# 
|
8-Labs/Lab18/Lab18.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from utils import load_caption, load_vocab, decode_caption, load_annotations, print_image
from nltk.translate import bleu_score
import pickle
import numpy as np
from tqdm import tqdm
from matplotlib import pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
plt.style.use('ggplot')
# -
vocab = load_vocab('/home/spb61/coco2014_vocab.json')
image_id_to_index, index_to_image_id, annotations_dict = load_annotations(annotations_dir='/home/spb61/annotations',
annotations_file='captions_val2014.json',
map_file = '/home/spb61/val_image_id_to_idx.csv')
def get_vote_caption(image_id, voted_pickle_file='../outputs/vote_captions_100_unigram_overlap.pickle'):
with open(voted_pickle_file, 'rb') as handle:
voted_captions = pickle.load(handle)
caption_object = voted_captions[image_id]
return caption_object[0][0]
def find_vote_index(beam_captions, image_id, voted_pickle_file='../outputs/vote_captions_100_unigram_overlap.pickle'):
vote_caption = get_vote_caption(image_id, voted_pickle_file)
for i, cap in enumerate(beam_captions):
if np.array_equal(cap['sentence'], vote_caption):
return i
raise ValueError("Could not find corresponding beam")
def get_annotations(image_id):
return [' '.join(cap) for cap in annotations_dict[image_id]]
fig = plt.figure(figsize=(15, 5))
# image_ids = np.random.choice(40000, size=10, replace=False)
image_ids = [36290, 10622, 6660, 33297, 33626, 20514, 38585, 33347, 38586, 28086]
print(image_ids)
for i, image_id in enumerate(image_ids):
axes = fig.add_subplot(2, 5, i+1)
image_id = int(image_id)
beam_size = 100
image_dir = "/datadrive/val_beam_{}_states/".format(beam_size)
caption_object = load_caption(image_id, image_dir=image_dir)['captions']
probabilities = np.array([c['score'] for c in caption_object])
# print(probabilities * 100)
annotations = get_annotations(image_id)
bleu_scores = []
for j, cap in enumerate(caption_object):
sentence = ' '.join(decode_caption(cap['sentence'], vocab))
# print(sentence)
bleu_scores.append(bleu_score.sentence_bleu(annotations, sentence))
axes.scatter(probabilities, bleu_scores, s=2)
axes.scatter(probabilities[0], bleu_scores[0], marker='*', color='blue')
vote_index = find_vote_index(caption_object, image_id)
print(vote_index)
axes.scatter(probabilities[vote_index], bleu_scores[vote_index], marker='o', color='green')
axes.set_xscale('log')
axes.set_xlim([1e-6, 0.1])
axes.set_xlabel("Probability")
axes.set_ylabel("Bleu score")
plt.tight_layout()
# plt.savefig('../outputs/figs/sentence_beam_bleu.png', bbox_inches='tight')
# Beam bleus
beams = pickle.load(open('/datadrive/beams_{}.pickle'.format(10), 'rb'))
print(len(beams))
for i in range(100):
beam_cap = ' '.join(decode_caption(beams[i][0]['sentence'], vocab))
print(beam_cap)
# +
beams = {}
for k in [1,2,10,100]:
beams[k] = pickle.load(open('/datadrive/beams_{}.pickle'.format(k), 'rb'))
beam_bleus = {}
image_ids = sorted(beams[1].keys())
print(len(image_ids))
for k in [1,2,10,100]:
annotations = []
generated_caps = []
bleus = []
for image_id in tqdm(image_ids):
beam_captions = beams[k][image_id]
beam_cap = decode_caption(beam_captions[0]['sentence'], vocab)
annotations_caps = annotations_dict[image_id]
bleu = bleu_score.sentence_bleu(annotations_caps, beam_cap, weights=[0.25, 0.25, 0.25, 0.25])
bleus.append(bleu)
beam_bleus[k] = bleus
# +
image_id = image_ids[np.argmax(np.array(beam_bleus[100]) - np.array(beam_bleus[1]))]
print_image(image_id, image_dir='/home/spb61/val_2014/')
plt.xticks([])
plt.yticks([])
for i, k in enumerate([1,2,10,100]):
plt.text(0, 400 + i * 45, "{:3d}: {}".format(k, ' '.join(decode_caption(beams[k][image_id][0]['sentence'], vocab))), fontsize=14)
plt.savefig("../outputs/figs/example_image_large_beam1.png", bbox_inches='tight')
# +
image_id = image_ids[np.argsort(np.array(beam_bleus[100]) - np.array(beam_bleus[2]))[-5]]
print_image(image_id, image_dir='/home/spb61/val_2014/')
plt.xticks([])
plt.yticks([])
for i, k in enumerate([1,2,10,100]):
plt.text(0, 540 + i * 45, "{:3d}: {}".format(k, ' '.join(decode_caption(beams[k][image_id][0]['sentence'], vocab)), fontsize=0), fontsize=14)
plt.savefig("../outputs/figs/example_image_large_beam2.png", bbox_inches='tight')
# +
image_id = image_ids[np.argsort(np.array(beam_bleus[100]) - np.array(beam_bleus[10]))[-3]]
print_image(image_id, image_dir='/home/spb61/val_2014/')
plt.xticks([])
plt.yticks([])
for i, k in enumerate([1,2,10,100]):
plt.text(-20, 350 + i * 45, "{:3d}: {}".format(k, ' '.join(decode_caption(beams[k][image_id][0]['sentence'], vocab))), fontsize=14)
plt.savefig("../outputs/figs/example_image_large_beam3.png", bbox_inches='tight')
# -
# # Position ranks analysis
beams_10 = pickle.load(open('/datadrive/beams_10.pickle', 'rb'))
positions = []
for image_id, caption_object in tqdm(beams_10.items()):
positions.append(find_vote_index(caption_object, image_id, voted_pickle_file='../outputs/vote_captions_10_unigram_overlap.pickle'))
beams_10[0]
np.mean(positions)
|
evaluation/Beam bleu scores per image.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/kreshuklab/vem-primer-examples/blob/main/mitochondria/3_instance_segmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # vEM-Mitochondria: Instance Segmentation
#
# Lorem ipsum
# ## Google Colab
#
# IMPORTANT: Run the next cells until `Instance Segmentation` only if you execute this notebook on Google Colab. If you run this notebook locally, you need to set up a python environment with the correct dependencies beforehand, check out [these instructions](https://github.com/kreshuklab/vem-primer-examples#setting-up-conda-environments-advanced).
# mount google drive
from google.colab import drive
drive.mount("/content/gdrive")
# ## Instance Segmentation
#
# Lorem ipsum
# import all the required dependencies
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import zarr
from skimage.measure import label
from skimage.segmentation import watershed
from numcodecs import GZip
# RUN THIS IF YOU WORK LOCALLY
data_path = "vem-primer-data/MitoEM-H-predictions.ome.zarr"
# RUN THIS ON GOOGLE COLAB
data_path = "/content/gdrive/MyDrive/vem-primer-data/MitoEM-H-cropped.ome.zarr"
# apply connected components labeling with foreground and boundary pmaps
def connected_components(foreground, boundaries, threshold=0.5, min_size=250):
seeds = label(np.clip(foreground - boundaries, 0, 1) > threshold)
mask = foreground > threshold
seg = watershed(boundaries, markers=seeds, mask=mask)
# apply size filter
seg_ids, sizes = np.unique(seg, return_counts=True)
bg_ids = seg_ids[sizes < min_size]
seg[np.isin(seg, bg_ids)] == 0
return seg
# TODO adapt everything so that it works out of core
# load the (probabilistic) semantic segmentation
store = zarr.DirectoryStore(data_path)
with zarr.open(store, "a") as f:
foreground = f["predictions/foreground"][:]
boundaries = f["predictions/boundaries"][:]
chunks = f["predictions/boundaries"].chunks
instance_seg = connected_components(foreground, boundaries)
f.create_dataset("segmentation/instance_segmentation", data=instance_seg, chunks=chunks, compression=GZip())
fig, ax = plt.subplots(1, 3)
ax[0].imshow(foreground[10])
ax[1].imshow(boundaries[10])
ax[2].imshow(instance_seg[10])
plt.show()
|
mitochondria/3_instance_segmentation.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# +
using Distributions
using StatsPlots
using Turing
using Logging
using LaTeXStrings
default(labels=false)
# -
# Code 2.1
ways = [0, 3, 8, 9, 0]
ways = ways ./ sum(ways)
println(ways)
# Code 2.2
b = Binomial(9, 0.5)
pdf(b, 6)
# Code 2.3
# +
# size of the grid
size = 20
# grid and prior
p_grid = range(0, 1; length=size)
prior = repeat([1.0], size)
# compute likelihood at each value in grid
likelyhood = [pdf(Binomial(9, p), 6) for p in p_grid]
# compute product of likelihood and prior
unstd_posterior = likelyhood .* prior
# standardize the posterior, so it sums to 1
posterior = unstd_posterior / sum(unstd_posterior);
# -
# Code 2.4
plot(p_grid, posterior;
xlabel="probability of water",
ylabel="posterior probability",
title="$size points",
markershape=:circle
)
# Code 2.5
# +
size = 20
p_grid = range(0, 1; length=size)
# prior is different - 0 if p < 0.5, 1 if >= 0.5
prior = convert(Vector{AbstractFloat}, p_grid .>= 0.5)
# another prior to try (uncomment the line below)
# prior = exp.(-5*abs.(p_grid .- 0.5))
# the rest is the same
likelyhood = [pdf(Binomial(9, p), 6) for p in p_grid]
unstd_posterior = likelyhood .* prior
posterior = unstd_posterior / sum(unstd_posterior);
plot(p_grid, posterior;
xlabel="probability of water",
ylabel="posterior probability",
title="$size points",
markershape=:circle
)
# -
# Code 2.6
# +
@model function water_land(W, L)
p ~ Uniform(0, 1)
W ~ Binomial(W + L, p)
end
Logging.disable_logging(Logging.Warn)
chain = sample(water_land(6, 3), NUTS(0.65), 1000)
display(chain)
# -
# Code 2.7
# +
# analytical calculation
W = 6
L = 3
x = range(0, 1; length=101)
b = Beta(W+1, L+1)
plot(x, pdf(b, x); label = L"\mathcal{Beta}")
# quadratic approximation
b = Normal(0.6374, 0.1414)
plot!(x, pdf(b, x); style=:dash, label=L"\mathcal{N}(0.64, 0.14)")
# -
# Code 2.8
# +
n_samples = 1000
p = Vector{Float64}(undef, n_samples)
p[1] = 0.5
W, L = 6, 3
for i ∈ 2:n_samples
p_old = p[i-1]
p_new = rand(Normal(p_old, 0.1))
if p_new < 0
p_new = abs(p_new)
elseif p_new > 1
p_new = 2-p_new
end
q0 = pdf(Binomial(W+L, p_old), W)
q1 = pdf(Binomial(W+L, p_new), W)
u = rand(Uniform())
p[i] = (u < q1 / q0) ? p_new : p_old
end
# -
# Code 2.9
density(p; label = "MCMC")
b = Beta(W+1, L+1)
plot!(x, pdf(b, x); label = L"\mathcal{Beta}", style=:dash)
|
02-Chapter 2. Small Worlds and Large World.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
import numpy as np
env = gym.make('Pong-v0')
env.env.frameskip = 3
# +
def iteration(env, name):
env.reset()
done = False
frames = []
rewards = []
i=0
while not done:
action = np.random.randint(0, 6)
state, reward, done, _ = env.step(action)
rewards.append(reward)
frames.append(state)
i+=1
frames = np.stack(frames, 0)
rewards = np.array(rewards)
np.savez_compressed("./traces/{}_trace".format(name), frames=frames, rewards=rewards)
return i
def generate_frames(env, num_frames):
frames = 0
i = 0
while frames < num_frames:
frames += iteration(env, i)
i+=1
print("At iteration {}, {} frames generated".format(i, frames))
# -
generate_frames(env, 100000)
data = np.load("./traces/5_trace.npz")
for i in range(81):
data = np.load("./traces/{}_trace.npz".format(i))
|
extras/Pong Data Generation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3.8
# ---
# # Train with tensorflow
#
# description: train tensorflow NN model on mnist data
# +
from azureml.core import Workspace
ws = Workspace.from_config()
ws
# +
import git
from pathlib import Path
# get root of git repo
prefix = Path(git.Repo(".", search_parent_directories=True).working_tree_dir)
# training script
script_dir = prefix.joinpath("code", "models", "tensorflow", "mnist-nn")
script_name = "train.py"
# environment file
environment_file = prefix.joinpath("environments", "tf-gpu.yml")
# azure ml settings
environment_name = "tf-gpu"
experiment_name = "tf-mnist-example"
compute_target = "gpu-cluster"
# + tags=[]
print(open(script_dir.joinpath(script_name)).read())
# +
from azureml.core import ScriptRunConfig, Experiment, Environment
env = Environment.from_conda_specification(environment_name, environment_file)
env.docker.enabled = True
src = ScriptRunConfig(
source_directory=script_dir,
script=script_name,
environment=env,
compute_target=compute_target,
)
run = Experiment(ws, experiment_name).submit(src)
run
# +
from azureml.widgets import RunDetails
RunDetails(run).show()
# -
run.wait_for_completion(show_output=True)
|
notebooks/tensorflow/train-mnist-nn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # data preprocessing
# # step1 : import libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# +
# step2: import dataset
dataset=pd.read_csv('Social_Ads.csv')
# -
dataset
# +
# step 3: to create feature vector (x) and variable vector(Y)
x=dataset.iloc[:,:-1].values
y=dataset.iloc[:,-1].values
# -
x
y
# step4: replacing missing data
from sklearn.impute import SimpleImputer
imputer=SimpleImputer(missing_values=np.nan,strategy='mean' )
imputer.fit(x)
x=imputer.transform(x)
# +
# step5: encoding (not required)
# -
#step6: splitting to data set into training data set and testing data set
from sklearn.model_selection import train_test_split
xtrain,xtest,ytrain,ytest=train_test_split(x,y,test_size=0.2,random_state=1)
# step 7: feature scaling
from sklearn.preprocessing import StandardScaler
sc=StandardScaler()
xtrain=sc.fit_transform(xtrain)
xtest=sc.fit_transform(xtest)
# # step B build KNN classification model
#
from sklearn.neighbors import KNeighborsClassifier
KC=KNeighborsClassifier(n_neighbors=5,weights='uniform',p=2)
KC.fit(xtrain,ytrain)
yestimated=KC.predict(xtest)
# performance matrix
from sklearn.metrics import confusion_matrix
CM=confusion_matrix(ytest,yestimated)
print(CM)
from sklearn.metrics import accuracy_score
CM=accuracy_score(ytest,yestimated)
print(CM)
# +
from sklearn.metrics import precision_score
CM=precision_score(ytest,yestimated)
print(CM)
# -
np.mean([True,True,False])
error_rate=[]
for i in range(1,30):
KC=KNeighborsClassifier(n_neighbors=i)
KC.fit(xtrain,ytrain)
ypred_i=KC.predict(xtest)
error_rate.append(np.mean(ypred_i!=ytest))
plt.plot(range(1,30),error_rate,marker='o',markerfacecolor='red',markersize=10)
|
Python-Week 5/31 august 2021 Day 16 K nearest neighbors algo.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ROOT C++
# language: c++
# name: root
# ---
# ## 计算沉积能量Q
#
# ### 准备
#
# +
TFile *ipf = new TFile("AddVeto.root");
TTree *fChain = (TTree*)ipf->Get("tree");
Int_t iqu, iqd;
fChain->SetBranchAddress("iqu", &iqu);
fChain->SetBranchAddress("iqd", &iqd);
// new file
TFile *opf = new TFile("QProcessor.root", "recreate");
TTree *opt = new TTree("tree", "q process step 1");
// new data
Double_t q0;
// new branch
opt->Branch("q0", &q0, "q0/D");
fChain->AddFriend(opt);
TCanvas *c1 = new TCanvas;
# -
# ### 处理qu
fChain->Draw("iqu >> hqu(420, 0, 4200)", "", "goff");
TH1D *hqu = (TH1D*)gDirectory->Get("hqu");
TF1 *f1 = new TF1("f1", "gaus", 0, 250);
hqu->Fit(f1, "R");
Double_t pedUSigma = f1->GetParameter(2);
Double_t pedU = f1->GetParameter(1);
TString sPedU(Form("%lf", pedU));
TString squLimit(Form("iqu > %lf+3.0*%lf && iqu < 4095", pedU, pedUSigma));
opt->SetAlias("qua", ("iqu-"+sPedU).Data());
opt->SetAlias("quLimit", squLimit.Data());
delete f1;
c1->Draw();
# ### 计算qd
fChain->Draw("iqd >> hqd(420, 0, 4200)", "", "goff");
TH1D *hqd = (TH1D*)gDirectory->Get("hqd");
f1 = new TF1("f1", "gaus", 0, 250);
hqd->Fit(f1, "R");
Double_t pedDSigma = f1->GetParameter(2);
Double_t pedD = f1->GetParameter(1);
TString sPedD{Form("%lf", pedD)};
TString sqdLimit(Form("iqd > %lf+3.0*%lf && iqd < 4095", pedD, pedDSigma));
opt->SetAlias("qda", ("iqd-"+sPedD).Data());
opt->SetAlias("qdLimit", sqdLimit.Data());
opt->SetAlias("qLimit", "qdLimit && quLimit");
delete f1;
c1->Draw();
# ### 存储
Long64_t nentries = fChain->GetEntries();
for (Long64_t jentry = 0; jentry != nentries; ++jentry) {
fChain->GetEntry(jentry);
q0 = sqrt((iqu-pedUSigma) * (iqd-pedDSigma));
opt->Fill();
}
fChain->Draw("q0:(itd+itu)/2.0 >> (450, 0, 4500, 450, 0, 4500) ", "qLimit", "colz");
c1->Draw();
opt->Write();
opf->Close();
# <div>
# <span style="float:left"><a href="hw1_2_3.ipynb">prev</a></span>
# <span style="float:right"><a href="hw1_2_5.ipynb">next</a></span>
# </div>
|
hw1_2/hw1_2_4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Continuación técnicas de reducción de varianza
#
# ## b). Números complementarios
#
# - Se tiene un cierto número de observaciones o valores aleatorios
# - Estos valores en ocasiones puede tener algún sesgo no deseado, además de no haber sido obligado o intencionado, como ejemplo supongamos que se hacen 10 observaciones donde los valores posibles son valores entre [0,1] y en todos los casos se obtuvieron datos menores a 0.5, lo cual sería inusual.
# - Obtener más observaciones puede ser costoso, no posible bajo las mismas condiciones o simplemente tiene un elevado costo computacional, así que lo que sugiere la técnica de “números complementarios” es obtener otro tanto de valores utilizando la fórmula
#
# > Aleatorio nuevo = Límite superior aleatorio generado - Aleatorio generado + Límite inferior aleatorio generado.
# 
#
# > **Ejemplo:** si $x\sim U[a,b]$ el número complementario para este número aleatorio es
# >$$x_{comp}=b-x+a$$
# > *Caso particular a=0,b=1* $$x_{comp}=1-x$$
#
#
# - Estos valores le dan equilibrio forzado a las observaciones o números aleatorios y permiten hacer una evaluación del proceso con valores que presentarán menor varianza.
# - Además que se logra obtener el doble de números respecto a los observados para simular el proceso.
#
# ## Ejemplo de ilustración
#
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle # Librería para hacer ciclos
import matplotlib.pyplot as plt
import scipy.stats as st # Librería estadística
import pandas as pd
cycol = cycle('bgrcmk') # variable que contiene los colores a graficar
a = 2; b = 8
x = np.random.uniform(a,b,4)
xc = b - x + a
# print(x,xc)
for i in range(len(x)):
c = next(cycol)
plt.plot(x[i],0,'>', c=c)
plt.plot(xc[i],0,'o',c=c)
plt.hlines(0,a,b)
plt.xlim(a,b)
plt.show()
# ## Ejemplo de aplicación
# Tomando como base el ejemplo de generación de números aleatorios exponenciales visto la clase pasada, ilustraremos este método.
#
# 
# +
np.random.seed(95555)
# Función para generar variables alestorias exponenciales
xi = lambda ri: -np.log(ri)
# Generación de Números aleatorios
ri =
# Media de observaciones aleatorias (Montecarlo estándar)
m_rand =
print('Media de observaciones aleatorias = ', m_rand)
# Números aleatorios complementarios
ri_c =
xi_c =
# Media de observaciones complementarias
m_comple =
print('Media de observaciones complementarias = ', m_comple)
m_estimada = (m_rand+m_comple)/2
print('La media estimada con el M.N.C es = ',m_estimada)
# -
# ## Análisis: ¿Por qué el método funciona?
#
# ### Recordar
# Ahora analicemos matemáticamente el efecto que esta sucediendo con este método.
# Recordemos la expresión de la varianza del estimador de media (promedio):
# 
# donde
# $$\rho _{X,Y}={\sigma _{XY} \over \sigma _{X}\sigma _{Y}}={E[(X-\mu _{X})(Y-\mu _{Y})] \over \sigma _{X}\sigma _{Y}}= {cov(X,Y) \over \sigma_X \sigma_Y}$$
#
# es el coeficiente de correlación de Pearson, y su valor varía en el intervalo [-1,1], indicando el signo el sentido de la relación. En la siguiente imagen se ilustra, varios grupos de puntos (x, y), con el coeficiente de correlación para cada grupo
# 
# - la **covarianza** es un valor que indica el grado de variación conjunta de dos variables aleatorias respecto a sus medias. Es el dato básico para determinar si existe una dependencia entre ambas variables.
# $$Cov(X,Y)=E[XY]-E[X]E[Y]$$
# - **coeficiente de correlación de Pearson** es una medida de la relación lineal entre dos variables aleatorias cuantitativas. A diferencia de la covarianza, la correlación de Pearson es independiente de la escala de medida de las variables
# Ahora recordemos el promedio de dos observaciones viene dado por la siguiente expresión:
# $$X^{(i)} = {X_1+X_2 \over 2}$$
# Ahora consideremos la media de una muestra $\bar X(n)$ basado en las muestras promediadas $X^{(i)}$, donde su varianza estará dada por:
# $$\begin{aligned}Var[\bar X(n)]&={Var(X^{(i)})\over n}\\
# &= {Var(X_1)+Var(X_2)+2Cov(X_1,X_2)\over 4n}\\
# &= {Var(X)\over 2n}(1+\rho(X_1,X_2))\end{aligned}$$
#
# > Concluimos que para poder obtener una reducción de varianza, con respecto al método de monte carlo tradicional, el coeficiente de correlación $\rho(X_1,X_2)<0$. Pero la pregunta entonces es, ¿Cómo podemos inducir una correlación negativa?.
#
# **Dibujar en el tablero la relación entre las variables {$U\sim U(0,1)$} y {$1-U$}**
#
# Relación entre las variables x1 = U y x2 = 1 - U
# x2 = np.random.rand(5)
x2 = np.array([1,.5,0])
x1 = 1-x2
plt.plot(x1,x2,'o-')
plt.show()
# El ejemplo de aplicación anterior mostró como el método es bastante sencillo y trabaja bastante bien. Pero, ¿Podemos esperar siempre un patrón similar? Desafortunadamente, la respuesta es no. La razón por la cual el enfoque funciona bastante bien en el ejemplo son dos:
# 1. Existe una fuerte correlación (positiva) entre $U$ y $e^u$ en el intervalo [0,1], porque la función es casi lineal allí. Esto significa que se conserva una fuerte correlación en la entrada de simulación y se convierte en una fuerte correlación en la salida de la simulación. No deberíamos esperar resultados impresionantes con funciones más complicadas y no lineales.
U1 = np.random.rand(10)
U2 = np.exp(U1)
plt.plot(U1,U2,'o')
plt.title(r'$u_1 = e^{u_2}$')
plt.show()
# En el caso de nuestro ejemplo de generación variables aleatorias exponenciales el método de la transformada inversa nos arrojó los siguientes resultados:
# $$
# x_i = -\ln u_i \ \rightarrow e^{-x_i} = u_i
# $$
# Lo cuál graficándolo nos arroja el siguiente resultado:
xi = np.random.rand(10)
ui = np.exp(-xi)
plt.plot(xi,ui,'o')
plt.title(r'$u_i = e^{-x_i}$')
plt.show()
# 2.Otra razón es que la función exponencial es creciente monótona. Como veremos en breve, la monotonicidad es una condición importante para el método de muestreo estratificado.
#
# >**Característica de las funciones monótonas:** Es decir una función es monótona cuando es creciente o decreciente en todo su dominio.
#
# ## Ejemplo donde el método de números complemantarios puede fallar
#
# Considere la función $h(x)$ definida como:
# $$h(x)=\begin{cases}0,& x<0,\\ 2x,& 0 \leq x \leq 0.5,\\2-2x,& 0.5\leq x\leq 1,\\0, & x>1,\end{cases}$$
#
# y supongamos que queremos aproximar la integrar $\int_0^1h(x)dx$ usando monte carlo.
#
# Como se puede observar, la función $h(x)$ es un triangulo, y el area encerrada bajo su curva es:
# $$\int_0^1h(x)dx \equiv E[h(U)]=\int_0^1h(u)\cdot 1 du = {1\over 2}$$
#
# Ahora entonces estimemos el valor de esta integral con el método tradicional de monte carlo y con el método de números complementarios
#
# $$\textbf{Monte carlo tradicional}\rightarrow X_I=\frac{h(U_1)+h(U_2)}{ 2}$$
#
# $$\textbf{Método números complemantarios}\rightarrow X_c={h(U)+h(1-U) \over 2}$$
#
# Ahora comparemos las dos varianzas de los dos estimadores:
# $$Var(X_I)={Var[h(U)]\over 2}\\
# Var(X_c)={Var[h(U)]\over 2}+{Cov[h(U),h(1-U)]\over 2}$$
#
# > **Recordar la expresión para calcular la esperanza:**
# > $$ \mathbb {E} [X]=\int_{-\infty }^{\infty }x f(x)dx $$
#
# Para saber que varianza es mayor, encontramos la diferencia de estas dos varianzas, en donde se obtiene:
# $$\begin{aligned}\Delta &= Var(X_c)-Var(X_I)={Cov[h(U),h(1-U)]\over 2} \\ &={1\over2}\{ E[h(U)h(1-U)]-E[h(U)]E[h(1-U)]\}\end{aligned}
# $$
#
# En este caso, debido a la forma de $h(x)$, tenemos que:
# $$E[h(U)]=E[h(1-U)]={1\over 2} \rightarrow \text{expresión de la media para $U\sim U[0,1]$}$$
# $$
# \begin{aligned}
# E[h(U)h(1-U)]&= \int_0^{1/2} h(U)h(1-U)\underbrace{f(x)}_{U\sim [0,1] = 1} du + \int_{1/2}^1 h(U)h(1-U)\underbrace{f(x)}_{U\sim [0,1] = 1} du \\
# E[h(u)h(1-u)] & = \int_0^{1/2} 2u\cdot(2-2(1-u))du + \int_{1/2}^1 2(1-u)\cdot(2-2u)du \\
# &= \int_0^{1/2} 4u^2du + \int_{1/2}^1 (2-2u)^2du = \frac{1}{3}
# \end{aligned}
# $$
#
# Por lo tanto, $Cov[h(U),h(1-U)]={1\over 3}-{1\over 4}={1\over 12}$ y de esta manera $\Delta ={1\over 24}>0$ y se concluye entonces que la varianza del método de números complementarios es mayor que la varianza del método de monte carlo común
# # Validación del resultado anterior
# +
np.random.seed(514)
# Programar función h(x)
# Gráfica de la función h(x)
x = np.arange(-.5,1.5,0.01)
plt.plot(x,list(map(f,x)),label='f(x)')
plt.legend()
plt.show()
# aproximar el valor de la integral usando montecarlo típico
# Aproximación usando método de los números complementarios
# Nota: Para ser justos tome la misma cantidad de términos para efectos de comparación
print('Media usando montecarlo estándar =',media_montecarlo)
print('Media usando números complementarios =',media_complementario)
print('Media teórica =',(0 + .5 + 1)/3 )
# Distrubución triangular (media)
# https://en.wikipedia.org/wiki/Triangular_distribution
# -
# ## ¿Por qué fallo el método en este ejemplo?
#
# Se demostró que las variables $U$ y $1-U$ están negativamente correlacionadas, pero en general no se puede garantizar que $X_1$ y $X_2$ cumplan esta propiedad en general.
#
# Para estar seguros de que la correlación negativa en los números aleatorios de entrada produce una correlación negativa en la salida observada, debemos exigir una relación monótona entre ellos. La función exponencial es una función monótona, pero la función triángulo del segundo ejemplo no lo es.
# ## Ejemplo de aplicación:
# Ejercicio tomado de: Introduction to Operations Research, 9ª ed. pag, 1148.
# 
# +
# Cantidad de términos
N = 10
# Función inversa
f_inv = lambda u:
# MÉTODOS PARA APROXIMAR LA MEDIA DE LA DISTRIBUCIÓN
# 1. Montecarlo crudo
# 2. Método estratificado
# 3. Método números complementarios
# -
# # <font color = red> Tarea
# **Esta tarea incluye el ejercicio dejado en la clase anterior**
# 
# Además use el método de estratificación donde se divide en B estratos, visto la clase pasada y compare el resultado con los métodos anteriores cuando se toman $2,4,6,10$ estratos respectivamente, concluya.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
|
TEMA-2/Clase14_ContinuacionReduccionVarianza.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Crop input image into seperate letters
# ### Import Modules
import cv2
import numpy as np
import imutils
# +
# Load image data
img = cv2.imread('data/2.jpg', -1)
rgb_planes = cv2.split(img)
result_planes = []
result_norm_planes = []
# Remove shadows
for plane in rgb_planes:
dilated_img = cv2.dilate(plane, np.ones((7,7), np.uint8))
bg_img = cv2.medianBlur(dilated_img, 21)
diff_img = 255 - cv2.absdiff(plane, bg_img)
norm_img = cv2.normalize(diff_img,None, alpha=0, beta=255, norm_type=cv2.NORM_MINMAX, dtype=cv2.CV_8UC1)
result_norm_planes.append(norm_img)
result_norm_planes_image = cv2.merge(result_norm_planes)
image = imutils.resize(result_norm_planes_image, height=800)#width=800,
# make a copy
original = image.copy()
#convert to grayscale
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
#parameters for threshold
thresholdValue = 0
maxVal = 255
thresholdingTechnique = cv2.THRESH_OTSU + cv2.THRESH_BINARY
thresh = cv2.threshold(gray,thresholdValue,maxVal,thresholdingTechnique)[1]
# Selecting ROIs
ROIs = cv2.selectROIs("Select Rois",original)
cv2.destroyAllWindows()
# +
crop_number=0
#loop over every bounding box save in array "ROIs"
for rect in ROIs:
x1=rect[0]
y1=rect[1]
x2=rect[2]
y2=rect[3]
#crop roi from original image
img_crop=thresh[y1:y1+y2,x1:x1+x2]
#save cropped image
cv2.imwrite("crop"+str(crop_number)+".jpeg",img_crop)
crop_number+=1
cv2.waitKey(0)
cv2.destroyAllWindows()
# -
|
script/1_crop_image2letters.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data loading/preparation
#
# Recommend you use [Pandas](https://pandas.pydata.org/)
#
# Other options:
# * [NumPy](https://www.numpy.org/)
#
# Scikit learn includes a number of publicly available datasets that can be used for learning ML. From the documentation:
#
# ***A dataset is a dictionary-like object that holds all the data and some metadata about the data. This data is stored in the `.data` member, which is a `n_samples, n_features` array. In the case of supervised problem, one or more response variables are stored in the `.target` member. More details on the different datasets can be found in the dedicated section.***
#
# Some of the steps involved:
# * Removing erroneous data
# * Correcting errors
# * Extracting parts of a corpus of data with automated tools.
# * Integrating data from various sources
# * Feature engineering/data enrichment
# * Semantic mapping
#
# **NOTE:** Most machine learning models/functions in Scikit expect data to be normalized (mean centered and scaled by the standard deviation times n_samples). Tree based methods do not usually require this.
#
# These steps are often repeated multiple times as a project progresses - data visualization and modeling often result in more data preparation.
#
# Data Cleaning takes 50 - 90% of a data scientists time:
# * https://thumbor.forbes.com/thumbor/960x0/https%3A%2F%2Fblogs-images.forbes.com%2Fgilpress%2Ffiles%2F2016%2F03%2FTime-1200x511.jpg
# * https://dataconomy.com/2016/03/why-your-datascientist-isnt-being-more-inventive/
#
# For more instruction, seee this excellent tutorial showing some examples of data loading, preparation, and cleaning: https://pythonprogramming.net/machine-learning-tutorial-python-introduction/
# +
# This is an addition
# -
# import some of the libraries that we'll need
from sklearn import datasets
import numpy as np
import pandas as pd
# Documentation for the Diabetes dataset is available at: https://scikit-learn.org/stable/datasets/index.html#diabetes-dataset
#
# Columns in the dataset:
# * Age
# * Sex
# * Body mass index
# * Average blood pressure
# * S1
# * S2
# * S3
# * S4
# * S5
# * S6
#
# **Each of these 10 feature variables have been mean centered and scaled by the standard deviation times n_samples (i.e. the sum of squares of each column totals 1).**
#
# Target:
# * A quantitative measure of disease progression one year after baseline
#
# ### Load dataset
# +
diabetes = datasets.load_diabetes()
with np.printoptions(linewidth=130):
print('Data - first 5\n', diabetes.data[0:5,:])
print('Target - first 5\n', diabetes.target[0:5])
# -
diabetes.target.shape
diabetes.data.shape
df = pd.DataFrame(data=diabetes.data, columns=['age', 'sex', 'bmi', 'abp', 's1', 's2', 's3', 's4', 's5', 's6'])
df['target'] = diabetes.target
df.head()
# ### Load human readable version of dataset
# compare original data set to see what data looks like in native format
url="https://www4.stat.ncsu.edu/~boos/var.select/diabetes.tab.txt"
df=pd.read_csv(url, sep='\t')
# change column names to lowercase for easier reference
df.columns = [x.lower() for x in df.columns]
df.head()
df.describe()
# # Data visualization/exploration
#
# Recommend you start with [Seaborn](http://seaborn.pydata.org/) - Makes matplotlib easier; can access any part of matplotlib if necessary. Other recommendations include:
#
# * [matplotlib](https://matplotlib.org/) One of the older and more widespread in use
# * [Altair](https://altair-viz.github.io/)
# * [Bokeh](https://bokeh.pydata.org/en/latest/)
# * [Plot.ly](https://plot.ly/python/)
import seaborn as sns
sns.set()
sns.set_style("ticks", {
'axes.grid': True,
'grid.color': '.9',
'grid.linestyle': u'-',
'figure.facecolor': 'white', # axes
})
sns.set_context("notebook")
sns.scatterplot(x=df.age, y=df.y, hue=df.sex, palette='Set1')
sns.scatterplot(x=df.age, y=df.bmi, hue=df.sex, palette='Set1')
sns.jointplot(x=df.age, y=df.bmi, kind='hex')
tdf = df[df.sex == 1]
sns.jointplot(x=tdf.age, y=tdf.bmi, kind='hex')
tdf = df[df.sex == 2]
sns.jointplot(x=tdf.age, y=tdf.bmi, kind='hex')
sns.distplot(df.y, rug=True)
sns.pairplot(df, hue="sex", palette='Set1')
# ### Load the matplotlib extension for interactivity
#
# This will affect all subsequent plots, regardless of cell location.
#
# Best to run this before any plotting in notebook
# +
# # %matplotlib widget
# -
sns.scatterplot(x=df.age, y=df.bmi, hue=df.sex, palette='Set1')
# # Machine learning
#
# Recommend you use [scikit-learn](https://scikit-learn.org/stable/)
#
# Deep Learning options:
#
# * [Caffe](http://caffe.berkeleyvision.org/)
# * [Fastai](https://docs.fast.ai/) - Simplifies deep learning similar to scikit-learn; based on PyTorch
# * [Keras](https://keras.io/)
# * [PyTorch](https://pytorch.org/)
# * [TensorFlow](https://www.tensorflow.org/overview/)
#
# Natural Language Processing options:
#
# * [nltk](http://www.nltk.org/)
# * [spaCy](https://spacy.io/)
# * [Stanford NLP Libraries](https://nlp.stanford.edu/software/)
#
# Computer Vision:
# * [OpenCV](https://opencv.org/)
#
# Forecasting/Time Series:
#
# * [Prophet](https://facebook.github.io/prophet/)
# * [statsmodels](https://www.statsmodels.org/stable/index.html) - Also does other statistical techniques and machine learning
# ## Regression
#
# ### Linear Regression
# +
from sklearn import preprocessing, model_selection, svm
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
# Create linear regression object
regr = LinearRegression()
# by convention, X is features and y is target
# random_state: Set a number here to allow for same results each time
X_train, X_test, y_train, y_test = model_selection.train_test_split(diabetes.data, diabetes.target, test_size=0.2, random_state=42)
# Train the model using the training sets
regr.fit(X_train, y_train)
# -
# To see documentation on `train_test_split()`
??model_selection.train_test_split
# Make predictions using the testing set
y_pred = regr.predict(X_test)
# The coefficients
print('Coefficients: \n', regr.coef_)
# The mean squared error
print("Mean squared error: %.2f"
% mean_squared_error(y_test, y_pred))
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % r2_score(y_test, y_pred))
# +
# from https://stackoverflow.com/questions/26319259/sci-kit-and-regression-summary
import sklearn.metrics as metrics
def regression_results(y_true, y_pred):
# Regression metrics
explained_variance=metrics.explained_variance_score(y_true, y_pred)
mean_absolute_error=metrics.mean_absolute_error(y_true, y_pred)
mse=metrics.mean_squared_error(y_true, y_pred)
mean_squared_log_error=metrics.mean_squared_log_error(y_true, y_pred)
median_absolute_error=metrics.median_absolute_error(y_true, y_pred)
r2=metrics.r2_score(y_true, y_pred)
print('explained_variance: ', round(explained_variance,4))
print('mean_squared_log_error: ', round(mean_squared_log_error,4))
print('r2: ', round(r2,4))
print('MAE: ', round(mean_absolute_error,4))
print('MSE: ', round(mse,4))
print('RMSE: ', round(np.sqrt(mse),4))
regression_results(y_test, y_pred)
# -
# An `explained_variance` of `0.455` means that approximately 45% of the variance in the Target variable is explained by the linear regression formula
#
# ### Support Vector Machine Regression
#
# The objective of this algorithm is to maximize the distance between the decision boundary and the samples that are closest to the decision boundary. Decision boundary is called the “Maximum Margin Hyperplane.” Samples that are closest to the decision boundary are the support vectors. Through mapping of the various dimensions of data (n) into higher dimensional space via a kernel function e.g. k(x,y) each individual maybe separated from its neighbor to better identify those classified into each category.
#
# +
# Create Support Vector Machine regression object
svm_regr = svm.SVR(gamma='auto')
# Train the model using the training sets
svm_regr.fit(X_train, y_train)
# Make predictions using the testing set
y_pred = svm_regr.predict(X_test)
regression_results(y_test, y_pred)
# -
# ### XGBoost Regression
#
# XGBoost (eXtreme Gradient Boosting) is an algorithm that a few years ago was considered state of the art for applied machine learning and Kaggle competitions when dealing with structured data.
#
# XGBoost is an implementation of gradient boosted decision trees designed for speed and performance.
# +
from xgboost.sklearn import XGBRegressor
from sklearn.model_selection import RandomizedSearchCV
import scipy.stats as st
one_to_left = st.beta(10, 1)
from_zero_positive = st.expon(0, 50)
params = {
'n_estimators': st.randint(3, 40),
'max_depth': st.randint(3, 40),
'learning_rate': st.uniform(0.05, 0.4),
'colsample_bytree': one_to_left,
'subsample': one_to_left,
'gamma': st.uniform(0, 10),
'reg_alpha': from_zero_positive,
'min_child_weight': from_zero_positive,
'objective': ['reg:squarederror']
}
xgbreg = XGBRegressor(nthreads=-1)
# -
gs = RandomizedSearchCV(xgbreg, params, n_jobs=1, cv=5, iid=False)
gs.fit(X_train, y_train)
gs_pred = gs.predict(X_test)
gs
regression_results(y_test, gs_pred)
# ## Classification
#
# As we want to demonstrate classification (Target values are part of a class, not continuous numbers) we will switch to a different dataset. See https://scikit-learn.org/stable/datasets/index.html#breast-cancer-wisconsin-diagnostic-dataset for details.
#
# Attribute Information:
#
# * radius (mean of distances from center to points on the perimeter)
# * texture (standard deviation of gray-scale values)
# * perimeter
# * area
# * smoothness (local variation in radius lengths)
# * compactness (perimeter^2 / area - 1.0)
# * concavity (severity of concave portions of the contour)
# * concave points (number of concave portions of the contour)
# * symmetry
# * fractal dimension (“coastline approximation” - 1)
#
# Class/Target:
# * WDBC-Malignant
# * WDBC-Benign
# ### Support Vector Machine Classification
# +
bc = datasets.load_breast_cancer()
with np.printoptions(linewidth=160):
print('Data - first 5\n', bc.data[0:5,:])
print('Target - first 5\n', bc.target[0:5])
# +
# by convention, X is features and y is target
# random_state: Set a number here to allow for same results each time
X_train, X_test, y_train, y_test = model_selection.train_test_split(bc.data, bc.target, test_size=0.2, random_state=42)
# Create Support Vector Machine Classifier object
svmc = svm.SVC(kernel='linear', gamma='auto')
# Train the model using the training sets
svmc.fit(X_train, y_train)
# Make predictions using the testing set
y_pred = svmc.predict(X_test)
svmc
# -
print("Classification report for classifier %s:\n%s\n"
% (svmc, metrics.classification_report(y_test, y_pred)))
print("Confusion matrix:\n%s" % metrics.confusion_matrix(y_test, y_pred))
# +
data = {'y_pred': y_pred,
'y_test': y_test
}
df = pd.DataFrame(data, columns=['y_test','y_pred'])
confusion_matrix = pd.crosstab(df['y_test'], df['y_pred'], rownames=['Actual'], colnames=['Predicted'])
sns.heatmap(confusion_matrix, annot=True)
# -
# ### XGBoost Classifier
# +
from xgboost.sklearn import XGBClassifier
from sklearn.model_selection import cross_val_score
xclas = XGBClassifier()
xclas.fit(X_train, y_train)
xg_y_pred = xclas.predict(X_test)
cross_val_score(xclas, X_train, y_train)
# -
print("Classification report for classifier %s:\n%s\n"
% (xclas, metrics.classification_report(y_test, xg_y_pred)))
print("Confusion matrix:\n%s" % metrics.confusion_matrix(y_test, xg_y_pred))
# ## Clustering (unlabeled data)
#
# Principle Component Analysis (PCA) is a technique used to emphasize variation and bring out strong patterns in a dataset. It's often used to make data easy to explore and visualize as you can use it to find those variables that are most unique and just keep 2 or 3 which can then be easily visualized.
# +
from sklearn.decomposition import IncrementalPCA
X = bc.data
y = bc.target
n_components = 2
ipca = IncrementalPCA(n_components=n_components, batch_size=10)
X_ipca = ipca.fit_transform(X)
# -
# if plot data in 2 dimensions, are there any obvious clusters?
sns.scatterplot(x=X_ipca[:, 0], y=X_ipca[:, 1], palette='Set1')
# what if we label data by Target variable?
sns.scatterplot(x=X_ipca[y == 0, 0], y=X_ipca[y == 0, 1], palette='Set1')
sns.scatterplot(x=X_ipca[y == 1, 0], y=X_ipca[y == 1, 1], palette='Set1')
# ### K-Means clustering
#
# This technique requires you to know the number of clusters when you start. Since you may not know the number of clusters, you can visually determine the number based on distortion. See https://towardsdatascience.com/k-means-clustering-with-scikit-learn-6b47a369a83c
# +
from sklearn.cluster import KMeans
import matplotlib.pyplot as plt
# calculate distortion for a range of number of cluster
distortions = []
for i in range(1, 11):
km = KMeans(
n_clusters=i, init='random',
n_init=10, max_iter=300,
tol=1e-04, random_state=0
)
km.fit(X)
distortions.append(km.inertia_)
# plot
plt.plot(range(1, 11), distortions, marker='o')
plt.xlabel('Number of clusters')
plt.ylabel('Distortion')
plt.show()
# +
from sklearn.cluster import KMeans
km = KMeans(
n_clusters=2,
init='random',
n_init=10,
max_iter=300,
tol=1e-04,
random_state=0
)
y_km = km.fit_predict(bc.data)
# +
# plot the 3 clusters
plt.scatter(
bc.data[y_km == 0, 0], bc.data[y_km == 0, 1],
s=50, c='lightgreen',
marker='s', edgecolor='black',
label='cluster 1'
)
plt.scatter(
bc.data[y_km == 1, 0], bc.data[y_km == 1, 1],
s=50, c='orange',
marker='o', edgecolor='black',
label='cluster 2'
)
# plot the centroids
plt.scatter(
km.cluster_centers_[:, 0], km.cluster_centers_[:, 1],
s=250, marker='*',
c='red', edgecolor='black',
label='centroids'
)
plt.legend(scatterpoints=1)
plt.grid()
plt.show()
# -
# ## Understanding/Explaining the model
#
# See:
#
# * LIME (Local Interpretable Model-agnostic Explanations)
# * Github: https://github.com/marcotcr/lime
# * Paper: https://arxiv.org/abs/1602.04938
# * SHAP (SHapley Additive exPlanations)
# * Github: https://github.com/slundberg/shap
# * Paper: http://papers.nips.cc/paper/7062-a-unified-approach-to-interpreting-model-predictions
# ## Bonus: Deep Learning with structured data
#
# Using Fastai Library and the Diabetes data set used for regression examples.
#
# https://www.kaggle.com/magiclantern/deep-learning-structured-data
|
code/hands-on-machine-learning-with-python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="-K0TVsmwJmun"
# # 1. Text Cleaning
# + id="SKWHSGnB1GqW"
#########
# Import
#########
# %%capture
import os
import sys
import pandas as pd
import numpy as np
# + id="VJHMMzyk2K9M"
########################
# Set directories paths
########################
YOUR_PATH = "..."
proj_dir = os.path.join(YOUR_PATH, "net2020-main")
base_dir = os.path.join(YOUR_PATH, "net2020-main/Semantic_Group/text_preprocessing")
# + id="OX14Jxj_2Voh"
####################################
# Import text-cleaning utils folder
####################################
sys.path.append(os.path.join(base_dir, "utils"))
# + id="dAhJvDBy1i4m"
##############
# Import data
##############
os.chdir(proj_dir)
# import the csv file with all the comments and post togheter
comDB = pd.read_csv(r"database/com_liwc.csv", sep='\t', engine='python', encoding='utf-8')
# import the csv file with JUST the politicians post
postDB = pd.read_csv(r"database/postDB.csv", engine='python')
# + id="Nbvnpcqo2K9Q"
# create the Data Frame
df = pd.DataFrame(data=comDB)
df_post = pd.DataFrame(data=postDB)
# add a new colum with sequence numbers
df['Count']=1
df_post['Count']=1
# print all the DF
pd.set_option('display.max_columns', None)
pd.set_option('display.max_row', 5)
# + id="5xvKnScreGkL"
########################
# Set working directory
########################
os.chdir(base_dir)
# + [markdown] id="O0YzWUSw4Oya"
# # Data Analysis
# + [markdown] id="Sl1UHq6F9hyJ"
# ## NaN values
# + colab={"base_uri": "https://localhost:8080/"} id="pfUiM6Du9lF9" outputId="bf5370b2-183e-46da-8ffa-8f970ec0dade"
print('Columns with Nan in df:\n', [(col, df[col].isna().sum()) for col in df.columns if df[col].isna().sum()>0], '\n')
print('Columns with Nan in df_post:\n', [(col, df_post[col].isna().sum()) for col in df_post.columns if df_post[col].isna().sum()>0])
# + [markdown] id="heENbub--lic"
# For the moment we are concerned about the NaN in the columns related to posts and comments text.
# + [markdown] id="DWr3GVGPDIV2"
# ### NaN in comments dataframe
# + colab={"base_uri": "https://localhost:8080/"} id="o0dhq8de_Xd-" outputId="a2fd4976-91d0-40fb-abf3-120e86c37aa4"
# Identify rows with NaN in post text in df (comments dataframe)
df[df['p_text'].isna()][['Origin_file_order']]
# + colab={"base_uri": "https://localhost:8080/"} id="saM7NH-p_6TP" outputId="4900d4e1-b972-4da4-8592-bbe4d554e210"
# Identify rows with NaN in comment text in df (comments dataframe)
df[df['c_text'].isna()][['Origin_file_order']]
# + [markdown] id="RgsdMJFLACJf"
# Row 45804 in comments dataframe can be removed since we have neither the text of the post nor the text of the comment associated with it.
# + colab={"base_uri": "https://localhost:8080/"} id="Teou5YsZE5cw" outputId="8c071e2a-cd7f-4b34-d55c-27a4e21be354"
print('df shape before dropping row: \t', df.shape)
df = df[df['c_text'].notna()]
print('df shape after dropping row: \t', df.shape)
print('Number of Nan in comments text: ', df['c_text'].isna().sum())
# + colab={"base_uri": "https://localhost:8080/"} id="QDtr2m-ZFZz6" outputId="0cec14a9-964b-4aac-8d93-395c122f7694"
df.shape
# + [markdown] id="Du_FS0UyDOlW"
# ### NaN in posts dataframe
# + colab={"base_uri": "https://localhost:8080/"} id="1Ep9V-AWB7hh" outputId="b003da56-ab0e-41cc-f70b-cbdc93c3e2e9"
# Identify rows with NaN in post text in df_post (posts dataframe)
df_post[df_post['p_text'].isna()][['Origin_file_order']]
# + [markdown] id="-_PCClP-MkXx"
# # Comments Text Preprocessing
# + [markdown] id="cC4q2Sp3H7eO"
# Let us create a dataframe containing only the comments' text
# + id="e1ZkrN7v2g2x"
# comments = df[['c_text']].sample(n=1000, random_state=1).copy() # work with a sample
comments = df[['c_text']].copy()
comments.rename(columns={'c_text':'text'}, inplace=True)
# + id="AvtNyx1ieNdE"
###############################
# Set seed for reproducibility
###############################
import random
random.seed(0)
np.random.seed(0)
# + colab={"base_uri": "https://localhost:8080/"} id="hb-kYXKieZY6" outputId="1904d1fd-0682-4d4b-9e19-3ce27c5cb815"
for i in list(np.random.choice(list(comments.index), 5)):
print(f'Comment {i}')
print(comments.loc[i]['text'], '\n')
# + [markdown] id="NCzIR5gs0dH5"
# ## Word cloud with raw data
# + [markdown] id="9fuXQsjtJ5r2"
# What if we generate a word cloud with no-preprocessed text?
# + id="83g71J8pKYjC"
from PIL import Image
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
import matplotlib.pyplot as plt
% matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="uGpJRJh4KdlH" outputId="d583ec1c-764a-4465-8f29-09134a4dff4c"
full_text = " ".join(comm for comm in comments['text'])
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(full_text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + [markdown] id="LUsSEXK5Kyqr"
# The word cloud we get is full of so-called stop words, the only significant words we can recognize are names of parties or politicians. A bit of text pre-processing is therefore mandatory.
#
#
# + [markdown] id="8V7UUObN1xVx"
# ## Text pre-processing
# + [markdown] id="YmvoR7G13q3F"
# There are differnt types of text preprocessing steps which can be applied and the choice of these steps depends on the tasks to be performed.
#
#
# For this initial step, our goal is to identify the most used words in the comments and the main topics of discussion.
# + id="IwEQRL4VTxpr"
comments["text_clean"] = comments["text"].copy()
# + [markdown] id="9WmiGhvH20co"
# ### Lower casing
# + id="X3TquD_T20cq"
def lower_casing(text):
return(text.lower())
# + id="9U3zalqT20cq"
comments["text_clean"] = comments["text_clean"].apply(lambda text: lower_casing(text))
# + [markdown] id="fw6LGqSO3ZpE"
# ### Removal of redundant spaces
# + id="XtgxEdLi3ZpG"
def remove_spaces(text):
return ' '.join([token for token in text.split()])
# + id="nlmfOV4y3ZpH"
comments["text_clean"] = comments["text_clean"].apply(lambda text: remove_spaces(text))
# + [markdown] id="5ogs-gPr2osT"
# ### Removal of patterns
# + id="cuR_nf-5Kle6"
import re
import regex
from collections import Counter
def remove_patterns(text, patterns):
for pattern in patterns:
try: r = re.findall(pattern, text)
except: r = regex.findall(pattern, text)
for i in r:
try: text = re.sub(re.escape(i), ' ', text)
except: text = regex.sub(regex.escape(i), ' ', text)
return text
def pattern_freq(docs, pattern):
p_freq = Counter()
for text in docs:
p_found= re.findall(pattern, text)
for p in p_found:
p_freq[p] += 1
return p_freq
# + id="SPj-m9H6zG5z"
# %%capture
# !pip install emoji
import emoji
# + id="JZ_1-IXF2znf"
PATTERNS = {'urls': re.compile(r'https?://\S+|www\.\S+'),
'users': re.compile(r'@[\w]*'),
#'hashtags': re.compile(r'#[\w]*'),
'emojis': emoji.get_emoji_regexp(),
'laughs': re.compile(r'\b(?:(?:hah+|ah+|hah+a|ah+a|lo+l) ?)+\b'),
'blabla': re.compile(r'\b(?:(?:bla+) ?)+\b'),
'dates': re.compile(r'\b(?:(?:[0-9]+[:\/,\.])+[0-9]+)+\b'),
'digits': r'#\S+(*SKIP)(*FAIL)|\b\d+\s|\s\d+\s|\s\d+|\b\d+\W|\W\d+\W|\W\d+$'
}
# + [markdown] id="76c2iPpd9XIA"
# Before removing patterns we can answer dollowing questions:
# * Which are the most used hashtags?
# * Which are most tagged users?
# * Are there frequent URLs?
# * Which are most frequent emojis/emoticons?
# + colab={"base_uri": "https://localhost:8080/"} id="au91pHLa9bVj" outputId="85e9c632-4d93-41a6-cf0c-02b521972a05"
hashtags_patt = re.compile(r'#[\w]*')
hashtags_freq = pattern_freq(comments['text'].values, hashtags_patt)
hashtags_freq.most_common(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="nF04TBt__V-Q" outputId="b738b5f4-e475-4a95-ff21-f2d866209e98"
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate_from_frequencies(hashtags_freq)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="9Q3owLSg-l7d" outputId="de065f9b-bda9-48dc-f1e0-b0e07f3ae08a"
users_freq = pattern_freq(comments['text'].values, PATTERNS['users'])
users_freq.most_common(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="E9LnE_oY_OZs" outputId="7c051208-a663-4573-dc37-3484f483d95a"
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate_from_frequencies(users_freq)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="w7q7e5ig_8m1" outputId="39392563-9307-4758-99db-5a09d4baee95"
urls_freq = pattern_freq(comments['text'].values, PATTERNS['urls'])
urls_freq.most_common(10)
# + colab={"base_uri": "https://localhost:8080/"} id="Hg86byYT5LaH" outputId="3a9da2b4-dd82-4b7c-90a8-5f7793dc632a"
emojis_freq = pattern_freq(comments['text'].values, PATTERNS['emojis'])
emojis_freq.most_common(10)
# + id="NRXchEB14Ufa"
comments["text_clean"] = comments["text_clean"].apply(lambda text: remove_patterns(text, PATTERNS.values()))
# + [markdown] id="k_fM6XO1BWO2"
# ### Unicode replacer
# + id="6347g6BifLib"
from unicode_replacer import *
# + id="UCTwcPYh2eyL"
comments['text_clean'] = comments['text_clean'].apply(lambda text: UnicodeReplacer().replace(text))
# + [markdown] id="7w0FiawCRZM7"
# ### Removal of punctuation
# + id="1f3l6hUZRZM8" colab={"base_uri": "https://localhost:8080/"} outputId="499b9653-50d7-4193-b131-f48396c0f5b8"
import string
punct = string.punctuation
print(f"Punctuation symbols: {punct}")
# + id="haD557CSRZM8"
def remove_punctuation(text, keep=[]):
punctuation = string.punctuation
for p in keep:
punctuation = punctuation.replace(p, "")
return(''.join([char if not char in punctuation else ' ' for char in text]))
# + id="JIK-CALtRkV4"
comments['text_clean'] = comments['text_clean'].apply(lambda text: remove_punctuation(text, keep="#',.")).apply(lambda text: remove_spaces(text))
# + id="_k3Pm4M4JIyE" colab={"base_uri": "https://localhost:8080/"} outputId="ad2e9000-77a0-47cf-ad76-8b9d83f5a982"
for i in list(np.random.choice(list(comments.index), 5)):
print(f'Comment {i}')
print(comments.loc[i]['text'])
print(comments.loc[i]['text_clean'], '\n')
print()
# + [markdown] id="aBw6RmERIAPv"
# ### NLP with Spacy
# + id="ZIwHw8oC4IE1"
##########
# Install
##########
# %%capture
# !pip install --upgrade spacy
# !python -m spacy download it_core_news_lg
# #!python -m spacy download it_core_news_sm
# + id="fhZVgNGdILr3"
import it_core_news_lg
nlp = it_core_news_lg.load()
# + [markdown] id="IcKRTGF14ST_"
# #### **Stop words collection**
# + id="pB2_14DM4SUB"
#################################################
# Import list of stopwords from it_stop_words.py
#################################################
from it_stop_words import get_italian_stop_words
my_it_stop_words = get_italian_stop_words()
# + id="GX93AqZX4SUB"
#######################################
# Import stopwords from spacy and nltk
#######################################
from spacy.lang.it.stop_words import STOP_WORDS as it_spacy_stopwords
import nltk
nltk.download('stopwords')
it_nltk_stopwords = nltk.corpus.stopwords.words('italian')
# Gather stopwords
it_stopwords = set(it_spacy_stopwords) | set(it_nltk_stopwords) | my_it_stop_words
# + [markdown] id="Iw4IfZhNEd7t"
# #### **Stop words to keep**
# + id="a9o6EH82hDd6"
keep_words = set(['casa', 'citta', 'città', 'governo', 'lontano', 'milione',
'persona', 'piedi', 'torino', 'vicino'])
it_stopwords = it_stopwords - keep_words
# + id="XlUiMnAaEl3m"
for stopword in keep_words:
nlp_vocab = nlp.vocab[stopword]
nlp_vocab.is_stop = False
for stopword in it_stopwords:
nlp_vocab = nlp.vocab[stopword]
nlp_vocab.is_stop = True
# + [markdown] id="3bEjxmp-f0PY"
# #### **Lemmatization errors**
# + id="2nP0aQjGsL8E"
###########################################################
# Import lemmatization corrections from it_lemma_errors.py
###########################################################
from it_lemma_errors import get_it_lemma_errors
lemma_errors = get_it_lemma_errors()
# + id="WadszXl-9jNS"
def lemma(token):
if token.text in lemma_errors:
if token.pos_ in lemma_errors[token.text]:
return lemma_errors[token.text][token.pos_]
return token.lemma_
# + [markdown] id="YH-26bsvZITw"
# #### **Hashtags as tokens**
#
# + id="MyxpF4DBaUr4"
from spacy.tokenizer import _get_regex_pattern
# get default pattern for tokens that don't get split
re_token_match = _get_regex_pattern(nlp.Defaults.token_match)
# add your patterns (here: hashtags and in-word hyphens)
re_token_match = f"({re_token_match}|#\w+|\w+-\w+)"
# + id="6BL8S2FTbtbk"
# overwrite token_match function of the tokenizer
nlp.tokenizer.token_match = re.compile(re_token_match).match
# + [markdown] id="WBoa2jtIEoNy"
# #### **Text processing**
# + id="BsOtGqjAI5f_"
text_nlp = comments["text_clean"].apply(lambda text: nlp(text))
comments['text_nlp'] = text_nlp
# + id="zOVsht2n8dPr"
# Run this cell to save the nlp-comments
# comments['text_nlp'].to_pickle(os.path.join(base_dir, 'comments_nlp'))
# + id="Dea_ICkL8dPs"
# Run this cell to load the nlp-comments
# comments['text_nlp'] = pd.read_pickle(os.path.join(base_dir, 'comments_nlp'))
# + colab={"base_uri": "https://localhost:8080/"} id="EltPz8mHhe2U" outputId="5a0ef282-2ad3-4384-96dd-fded936fa6c8"
print(f"{'Token':<20}\t{'Lemma':<20}\t{'POS':<20}\t{'is-stop':<8}\t{'is-punct':<8}")
for token in comments['text_nlp'].loc[0]:
print(f"{token.text:<20}\t{lemma(token):<20}\t{token.pos_:<20}\t{token.is_stop or lemma(token) in it_stopwords:^8}\t{token.is_punct:^8}")
# + [markdown] id="wtepnsm27VrG"
# ### Removal of Stop-Words
# + id="VyWmkWx-7VrH"
def remove_stop(tokens):
return(' '.join([lemma(token) for token in tokens if not (token.is_stop or lemma(token) in it_stopwords)]))
# + id="DQoOTHpZ7VrH"
comments['text_clean'] = comments["text_nlp"].apply(lambda tokens: remove_stop(tokens))
# + [markdown] id="qezRUmhYgD-Z"
# ### Further removal of punctuation
# + id="mRxrsOy5gbsG"
comments['text_clean'] = comments['text_clean'].apply(lambda text: remove_punctuation(text, keep=['#']))
# + id="OC0t1MUa9EKO"
comments["text_clean"] = comments['text_clean'].apply(lambda text: remove_spaces(text))
# + id="UPLN7aAjg7Vy" colab={"base_uri": "https://localhost:8080/"} outputId="1c82bfab-edda-4cb7-b440-82b64f9d5882"
import random
for i in list(np.random.choice(list(comments.index), 20)):
print(f'Comment {i}')
print(comments.loc[i]['text'])
print(comments.loc[i]['text_clean'])
print()
# + [markdown] id="3Hfozl0OmYZw"
# ## Save cleaned text
# + id="BaGLgIvZmX4_"
# comments['text_clean'].to_pickle(os.path.join(base_dir, 'comments_cleaned'))
# + id="1TN7jbGxyshE"
# comments['text_clean'] = pd.read_pickle(os.path.join(base_dir, 'comments_cleaned'))
# + [markdown] id="oxOrrNpT423N"
# ## Resulting word-cloud
# + colab={"base_uri": "https://localhost:8080/", "height": 198} id="u9eYYFnGjDDr" outputId="a30440ad-7783-4743-8c8a-a6f71b43d318"
full_cleaned_text = ' '.join([doc for doc in comments['text_clean']])
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(full_cleaned_text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# + [markdown] id="Tic5y-eB2p37"
# # Text Cleaning from function
# + [markdown] id="n0aSyRql3fYR"
# See text_preprocessing folder
# + colab={"base_uri": "https://localhost:8080/"} id="KwerO6uo2p3-" outputId="def7d643-7740-447c-c880-812828f02062"
# Note: text_preprocessing folder has to be updasted. We'll do it after fixing text-preprocessing problems
'''
# Import list of stopwords from it_stop_words.py
import sys
sys.path.append(os.path.join(base_dir, "Semantic_Group/text_preprocessing"))
from text_cleaning import *
cleaned_text = clean_content(comments['text'])
'''
# + id="CfZSx9UX2p4C"
'''
full_cleaned_text = ' '.join(cleaned_text)
wordcloud = WordCloud(max_font_size=50, max_words=100, background_color="white").generate(full_cleaned_text)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
'''
# + [markdown] id="9YfmT5N3uMOH"
# # Cleaned Data Analysis
# + [markdown] id="k5fWIz7akGYV"
# ## Most popular words
# + id="wQ4r1ZbNkGYW"
def words_freq(docs):
words_count = Counter()
for text in docs:
for word in text.split():
words_count[word] += 1
return words_count
# + id="mzbc9iT0kGYX"
words_count = words_freq(comments['text_clean'])
# + colab={"base_uri": "https://localhost:8080/"} id="7VKVjnrXkGYY" outputId="6738ca0b-0c8f-49c7-de34-f52c99d8b333"
print('Total words\n')
len(words_count.most_common())
# + id="P0wrP87WK2rF" colab={"base_uri": "https://localhost:8080/"} outputId="a92913d6-c585-43f8-90fe-c8fa05910533"
print('Most common words\n')
words_count.most_common(50)
# + colab={"base_uri": "https://localhost:8080/"} id="lqFcBeY6kGYz" outputId="e750be8d-88ed-4f39-d682-e16ba2a9c580"
less_common = [(w, wc) for (w, wc) in words_count.most_common()[:-51:-1]]
for (w, wc) in less_common: print((w, wc), end='\n')
# + [markdown] id="lMgBJoPN9hBP"
# **How many words occur 1 time? How many 2 times? ...**
# + id="bvrdC_jxkGY0"
import seaborn as sns
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="3nu0pOHtkGY1" outputId="c9e8c45e-a760-4762-d959-b189c34578bd"
counts = [wc for (w, wc) in words_count.most_common()]
sns.histplot(counts, bins=[1,2,3,4,5,6,7,8,9,10], discrete=True, binrange=[1,10])
|
Semantic_Group/text_preprocessing/text_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating Labeled Data from a Planet Mosaic with Label Maker
#
# In this notebook, we create labeled data for training a machine learning algorithm. As inputs, we use [OpenStreetMap](https://www.openstreetmap.org/#map=4/38.01/-95.84) as the ground truth source and a Planet mosaic as the image source. Development Seed's [Label Maker](https://developmentseed.org/blog/2018/01/11/label-maker/) tool is used to download and prepare the ground truth data, chip the Planet imagery, and package the two to feed into the training process.
#
# The primary interface for Label Maker is through the command-line interface (cli). It is configured through the creation of a configuration file. More information about that configuration file and command line usage can be found in the Label Maker repo [README](https://github.com/developmentseed/label-maker/blob/master/README.md).
#
# **RUNNING NOTE**
#
# This notebook is meant to be run in a docker image specific to this folder. The docker image must be built from the custom [Dockerfile](Dockerfile) according to the directions below.
#
# In label-data directory:
# ```
# docker build -t planet-notebooks:label .
# ```
#
# Then start up the docker container as you usually would, specifying `planet-notebooks:label` as the image.
#
# ## Install Dependencies
#
# In addition to the python packages imported below, the label-maker python package is also a dependency. However, it's primary usage is through the command-line interface (cli), so we use juypter notebook bash magic to run label-maker via the cli instead of importing the python package.
# +
import json
import os
import ipyleaflet as ipyl
import ipywidgets as ipyw
from IPython.display import Image
import numpy as np
# -
# ## Define Mosaic Parameters
#
# In this tutorial, we use the Planet mosaic [tile service](https://developers.planet.com/docs/api/tile-services/). There are many mosaics to choose from. For a list of mosaics available, visit https://api.planet.com/basemaps/v1/mosaics.
#
# We first build the url for the xyz basemap tile service, then we add authorization in the form of the Planet API key.
# Planet tile server base URL (Planet Explorer Mosaics Tiles)
mosaic = 'global_monthly_2018_02_mosaic'
mosaicsTilesURL_base = 'https://tiles.planet.com/basemaps/v1/planet-tiles/{}/gmap/{{z}}/{{x}}/{{y}}.png'.format(mosaic)
mosaicsTilesURL_base
# Planet tile server url with auth
planet_api_key = os.environ['PL_API_KEY']
planet_mosaic = mosaicsTilesURL_base + '?api_key=' + planet_api_key
# url is not printed because it will show private api key
# ## Prepare label maker config file
#
# This config file is pulled from the label-maker repo [README.md](https://github.com/developmentseed/label-maker/blob/master/README.md) example and then customized to utilize the Planet mosaic. The imagery url is set to the Planet mosaic url and the zoom is changed to 15, the maximum zoom supported by the [Planet tile services](https://developers.planet.com/docs/api/tile-services/).
#
# See the label-maker README.md file for a description of the config entries.
# create data directory
data_dir = os.path.join('data', 'label-maker-mosaic')
if not os.path.isdir(data_dir):
os.makedirs(data_dir)
# label-maker doesn't clean up, so start with a clean slate
# !cd $data_dir && rm -R *
# +
# create config file
bounding_box = [1.09725, 6.05520, 1.34582, 6.30915]
config = {
"country": "togo",
"bounding_box": bounding_box,
"zoom": 15,
"classes": [
{ "name": "Roads", "filter": ["has", "highway"] },
{ "name": "Buildings", "filter": ["has", "building"] }
],
"imagery": planet_mosaic,
"background_ratio": 1,
"ml_type": "classification"
}
# define project files and folders
config_filename = os.path.join(data_dir, 'config.json')
# write config file
with open(config_filename, 'w') as cfile:
cfile.write(json.dumps(config))
print('wrote config to {}'.format(config_filename))
# -
# ### Visualize Mosaic at config area of interest
# +
# calculate center of map
bounds_lat = [bounding_box[1], bounding_box[3]]
bounds_lon = [bounding_box[0], bounding_box[2]]
def calc_center(bounds):
return bounds[0] + (bounds[1] - bounds[0])/2
map_center = [calc_center(bounds_lat), calc_center(bounds_lon)] # lat/lon
print(bounding_box)
print(map_center)
# +
# create and visualize mosaic at approximately the same bounds as defined in the config file
map_zoom = 12
layout=ipyw.Layout(width='800px', height='800px') # set map layout
mosaic_map = ipyl.Map(center=map_center, zoom=map_zoom, layout=layout)
mosaic_map.add_layer(ipyl.TileLayer(url=planet_mosaic))
mosaic_map
# -
mosaic_map.bounds
# ## Download OSM tiles
#
# In this step, label-maker downloads the OSM vector tiles for the country specified in the config file.
#
# According to Label Maker documentation, these can be visualized with [mbview](https://github.com/mapbox/mbview). So far I have not been successful getting mbview to work. I will keep on trying and would love to hear how you got this to work!
# !cd $data_dir && label-maker download
# ## Create ground-truth labels from OSM tiles
#
# In this step, the OSM tiles are chipped into label tiles at the zoom level specified in the config file. Also, a geojson file is created for visual inspection.
# !cd $data_dir && label-maker labels
# Visualizing `classification.geojson` in QGIS gives:
#
# 
#
# Although Label Maker doesn't tell us which classes line up with the labels (see the legend in the visualization for labels), it looks like the following relationships hold:
# - (1,0,0) - no roads or buildings
# - (0,1,1) - both roads and buildings
# - (0,0,1) - only buildings
# - (0,1,0) - only roads
#
# Most of the large region with no roads or buildings at the bottom portion of the image is the water off the coast.
# ## Preview image chips
#
# Create a subset of the image chips for preview before creating them all. Preview chips are placed in subdirectories named after each class specified in the config file.
#
# **NOTE** This section is commented out because preview fails due to imagery-offset arg. See more:
# https://github.com/developmentseed/label-maker/issues/79
# +
# # !cd $data_dir && label-maker preview -n 3
# +
# # !ls $data_dir/data/examples
# +
# for fclass in ('Roads', 'Buildings'):
# example_dir = os.path.join(data_dir, 'data', 'examples', fclass)
# print(example_dir)
# for img in os.listdir(example_dir):
# print(img)
# display(Image(os.path.join(example_dir, img)))
# -
# Other than the fact that 4 tiles were created instead of the specified 3, the results look pretty good! All Road examples have roads, and all Building examples have buildings.
# ## Create image tiles
#
# In this step, we invoke `label-maker images`, which downloads and chips the mosaic into tiles that match the label tiles.
#
# Interestingly, only 372 image tiles are downloaded, while 576 label tiles were generated. Looking at the label tile generation output (370 Road tiles, 270 Building tiles) along with the `classification.geojson` visualization (only two tiles that are Building and not Road), we find that there are only 372 label tiles that represent at least one of the Road/Building classes. This is why only 372 image tiles were generated.
# !cd $data_dir && label-maker images
# look at three tiles that were generated
tiles_dir = os.path.join(data_dir, 'data', 'tiles')
print(tiles_dir)
for img in os.listdir(tiles_dir)[:3]:
print(img)
display(Image(os.path.join(tiles_dir, img)))
# ## Package tiles and labels
#
# Convert the image and label tiles into train and test datasets.
# will not be able to open image tiles that weren't generated because the label tiles contained no classes
# !cd $data_dir && label-maker package
# ## Check Package
#
# Let's load the packaged data and look at the train and test datasets.
data_file = os.path.join(data_dir, 'data', 'data.npz')
data = np.load(data_file)
for k in data.keys():
print('data[\'{}\'] shape: {}'.format(k, data[k].shape))
# 297 x (image) and y (label) datasets were created in the train set, and 75 x and y datasets were created in the test set, adding up to 372 sets total, equal to the number of image tiles downloaded.
# ## Next Steps
#
# The next step after creating labeled data is to train the machine learning algorithm.
#
# This Development Seed [walkthrough](https://github.com/developmentseed/label-maker/blob/master/examples/walkthrough-classification-aws.md) demonstrates how to train a neural network classifier.
|
jupyter-notebooks/label-data/label_maker_pl_mosaic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="j0a0a2Djcm9a" colab_type="code" colab={}
# install plotly (chart-studio):
# !pip install chart-studio
# for data manipulation:
import numpy as np
import pandas as pd
import datetime as dt
# for textual analysis:
import nltk
import re
nltk.download('punkt')
# silence warnings:
import warnings
warnings.filterwarnings("ignore")
# for plotting:
import plotly.graph_objects as go
# !pip install FedTools
from FedTools import MonetaryPolicyCommittee
dataset = MonetaryPolicyCommittee().find_statements()
# + id="acBlr4C2c9UG" colab_type="code" colab={}
for i in range(len(dataset)):
dataset.iloc[i,0] = dataset.iloc[i,0].replace('\\n','. ')
dataset.iloc[i,0] = dataset.iloc[i,0].replace('\n',' ')
dataset.iloc[i,0] = dataset.iloc[i,0].replace('\r',' ')
dataset.iloc[i,0] = dataset.iloc[i,0].replace('\xa0',' ')
# + id="wbF9_gnfdLyU" colab_type="code" colab={}
# Loughran and McDonald Sentiment Word Lists (https://sraf.nd.edu/textual-analysis/resources/)
lmdict = {'Negative': ['abandon', 'abandoned', 'abandoning', 'abandonment', 'abandonments', 'abandons', 'abdicated',
'abdicates', 'abdicating', 'abdication', 'abdications', 'aberrant', 'aberration', 'aberrational',
'aberrations', 'abetting', 'abnormal', 'abnormalities', 'abnormality', 'abnormally', 'abolish',
'abolished', 'abolishes', 'abolishing', 'abrogate', 'abrogated', 'abrogates', 'abrogating',
'abrogation', 'abrogations', 'abrupt', 'abruptly', 'abruptness', 'absence', 'absences',
'absenteeism', 'abuse', 'abused', 'abuses', 'abusing', 'abusive', 'abusively', 'abusiveness',
'accident', 'accidental', 'accidentally', 'accidents', 'accusation', 'accusations', 'accuse',
'accused', 'accuses', 'accusing', 'acquiesce', 'acquiesced', 'acquiesces', 'acquiescing',
'acquit', 'acquits', 'acquittal', 'acquittals', 'acquitted', 'acquitting', 'adulterate',
'adulterated', 'adulterating', 'adulteration', 'adulterations', 'adversarial', 'adversaries',
'adversary', 'adverse', 'adversely', 'adversities', 'adversity', 'aftermath', 'aftermaths',
'against', 'aggravate', 'aggravated', 'aggravates', 'aggravating', 'aggravation', 'aggravations',
'alerted', 'alerting', 'alienate', 'alienated', 'alienates', 'alienating', 'alienation',
'alienations', 'allegation', 'allegations', 'allege', 'alleged', 'allegedly', 'alleges',
'alleging', 'annoy', 'annoyance', 'annoyances', 'annoyed', 'annoying', 'annoys', 'annul',
'annulled', 'annulling', 'annulment', 'annulments', 'annuls', 'anomalies', 'anomalous',
'anomalously', 'anomaly', 'anticompetitive', 'antitrust', 'argue', 'argued', 'arguing',
'argument', 'argumentative', 'arguments', 'arrearage', 'arrearages', 'arrears', 'arrest',
'arrested', 'arrests', 'artificially', 'assault', 'assaulted', 'assaulting', 'assaults',
'assertions', 'attrition', 'aversely', 'backdating', 'bad', 'bail', 'bailout', 'balk', 'balked',
'bankrupt', 'bankruptcies', 'bankruptcy', 'bankrupted', 'bankrupting', 'bankrupts', 'bans',
'barred', 'barrier', 'barriers', 'bottleneck', 'bottlenecks', 'boycott', 'boycotted',
'boycotting', 'boycotts', 'breach', 'breached', 'breaches', 'breaching', 'break', 'breakage',
'breakages', 'breakdown', 'breakdowns', 'breaking', 'breaks', 'bribe', 'bribed', 'briberies',
'bribery', 'bribes', 'bribing', 'bridge', 'broken', 'burden', 'burdened', 'burdening', 'burdens',
'burdensome', 'burned', 'calamities', 'calamitous', 'calamity', 'cancel', 'canceled',
'canceling', 'cancellation', 'cancellations', 'cancelled', 'cancelling', 'cancels', 'careless',
'carelessly', 'carelessness', 'catastrophe', 'catastrophes', 'catastrophic', 'catastrophically',
'caution', 'cautionary', 'cautioned', 'cautioning', 'cautions', 'cease', 'ceased', 'ceases',
'ceasing', 'censure', 'censured', 'censures', 'censuring', 'challenge', 'challenged',
'challenges', 'challenging', 'chargeoffs', 'circumvent', 'circumvented', 'circumventing',
'circumvention', 'circumventions', 'circumvents', 'claiming', 'claims', 'clawback', 'closed',
'closeout', 'closeouts', 'closing', 'closings', 'closure', 'closures', 'coerce', 'coerced',
'coerces', 'coercing', 'coercion', 'coercive', 'collapse', 'collapsed', 'collapses',
'collapsing', 'collision', 'collisions', 'collude', 'colluded', 'colludes', 'colluding',
'collusion', 'collusions', 'collusive', 'complain', 'complained', 'complaining', 'complains',
'complaint', 'complaints', 'complicate', 'complicated', 'complicates', 'complicating',
'complication', 'complications', 'compulsion', 'concealed', 'concealing', 'concede', 'conceded',
'concedes', 'conceding', 'concern', 'concerned', 'concerns', 'conciliating', 'conciliation',
'conciliations', 'condemn', 'condemnation', 'condemnations', 'condemned', 'condemning',
'condemns', 'condone', 'condoned', 'confess', 'confessed', 'confesses', 'confessing',
'confession', 'confine', 'confined', 'confinement', 'confinements', 'confines', 'confining',
'confiscate', 'confiscated', 'confiscates', 'confiscating', 'confiscation', 'confiscations',
'conflict', 'conflicted', 'conflicting', 'conflicts', 'confront', 'confrontation',
'confrontational', 'confrontations', 'confronted', 'confronting', 'confronts', 'confuse',
'confused', 'confuses', 'confusing', 'confusingly', 'confusion', 'conspiracies', 'conspiracy',
'conspirator', 'conspiratorial', 'conspirators', 'conspire', 'conspired', 'conspires',
'conspiring', 'contempt', 'contend', 'contended', 'contending', 'contends', 'contention',
'contentions', 'contentious', 'contentiously', 'contested', 'contesting', 'contraction',
'contractions', 'contradict', 'contradicted', 'contradicting', 'contradiction', 'contradictions',
'contradictory', 'contradicts', 'contrary', 'controversial', 'controversies', 'controversy',
'convict', 'convicted', 'convicting', 'conviction', 'convictions', 'corrected', 'correcting',
'correction', 'corrections', 'corrects', 'corrupt', 'corrupted', 'corrupting', 'corruption',
'corruptions', 'corruptly', 'corruptness', 'costly', 'counterclaim', 'counterclaimed',
'counterclaiming', 'counterclaims', 'counterfeit', 'counterfeited', 'counterfeiter',
'counterfeiters', 'counterfeiting', 'counterfeits', 'countermeasure', 'countermeasures', 'crime',
'crimes', 'criminal', 'criminally', 'criminals', 'crises', 'crisis', 'critical', 'critically',
'criticism', 'criticisms', 'criticize', 'criticized', 'criticizes', 'criticizing', 'crucial',
'crucially', 'culpability', 'culpable', 'culpably', 'cumbersome', 'curtail', 'curtailed',
'curtailing', 'curtailment', 'curtailments', 'curtails', 'cut', 'cutback', 'cutbacks',
'cyberattack', 'cyberattacks', 'cyberbullying', 'cybercrime', 'cybercrimes', 'cybercriminal',
'cybercriminals', 'damage', 'damaged', 'damages', 'damaging', 'dampen', 'dampened', 'danger',
'dangerous', 'dangerously', 'dangers', 'deadlock', 'deadlocked', 'deadlocking', 'deadlocks',
'deadweight', 'deadweights', 'debarment', 'debarments', 'debarred', 'deceased', 'deceit',
'deceitful', 'deceitfulness', 'deceive', 'deceived', 'deceives', 'deceiving', 'deception',
'deceptions', 'deceptive', 'deceptively', 'decline', 'declined', 'declines', 'declining',
'deface', 'defaced', 'defacement', 'defamation', 'defamations', 'defamatory', 'defame',
'defamed', 'defames', 'defaming', 'default', 'defaulted', 'defaulting', 'defaults', 'defeat',
'defeated', 'defeating', 'defeats', 'defect', 'defective', 'defects', 'defend', 'defendant',
'defendants', 'defended', 'defending', 'defends', 'defensive', 'defer', 'deficiencies',
'deficiency', 'deficient', 'deficit', 'deficits', 'defraud', 'defrauded', 'defrauding',
'defrauds', 'defunct', 'degradation', 'degradations', 'degrade', 'degraded', 'degrades',
'degrading', 'delay', 'delayed', 'delaying', 'delays', 'deleterious', 'deliberate',
'deliberated', 'deliberately', 'delinquencies', 'delinquency', 'delinquent', 'delinquently',
'delinquents', 'delist', 'delisted', 'delisting', 'delists', 'demise', 'demised', 'demises',
'demising', 'demolish', 'demolished', 'demolishes', 'demolishing', 'demolition', 'demolitions',
'demote', 'demoted', 'demotes', 'demoting', 'demotion', 'demotions', 'denial', 'denials',
'denied', 'denies', 'denigrate', 'denigrated', 'denigrates', 'denigrating', 'denigration',
'deny', 'denying', 'deplete', 'depleted', 'depletes', 'depleting', 'depletion', 'depletions',
'deprecation', 'depress', 'depressed', 'depresses', 'depressing', 'deprivation', 'deprive',
'deprived', 'deprives', 'depriving', 'derelict', 'dereliction', 'derogatory', 'destabilization',
'destabilize', 'destabilized', 'destabilizing', 'destroy', 'destroyed', 'destroying', 'destroys',
'destruction', 'destructive', 'detain', 'detained', 'detention', 'detentions', 'deter',
'deteriorate', 'deteriorated', 'deteriorates', 'deteriorating', 'deterioration',
'deteriorations', 'deterred', 'deterrence', 'deterrences', 'deterrent', 'deterrents',
'deterring', 'deters', 'detract', 'detracted', 'detracting', 'detriment', 'detrimental',
'detrimentally', 'detriments', 'devalue', 'devalued', 'devalues', 'devaluing', 'devastate',
'devastated', 'devastating', 'devastation', 'deviate', 'deviated', 'deviates', 'deviating',
'deviation', 'deviations', 'devolve', 'devolved', 'devolves', 'devolving', 'difficult',
'difficulties', 'difficultly', 'difficulty', 'diminish', 'diminished', 'diminishes',
'diminishing', 'diminution', 'disadvantage', 'disadvantaged', 'disadvantageous', 'disadvantages',
'disaffiliation', 'disagree', 'disagreeable', 'disagreed', 'disagreeing', 'disagreement',
'disagreements', 'disagrees', 'disallow', 'disallowance', 'disallowances', 'disallowed',
'disallowing', 'disallows', 'disappear', 'disappearance', 'disappearances', 'disappeared',
'disappearing', 'disappears', 'disappoint', 'disappointed', 'disappointing', 'disappointingly',
'disappointment', 'disappointments', 'disappoints', 'disapproval', 'disapprovals', 'disapprove',
'disapproved', 'disapproves', 'disapproving', 'disassociates', 'disassociating',
'disassociation', 'disassociations', 'disaster', 'disasters', 'disastrous', 'disastrously',
'disavow', 'disavowal', 'disavowed', 'disavowing', 'disavows', 'disciplinary', 'disclaim',
'disclaimed', 'disclaimer', 'disclaimers', 'disclaiming', 'disclaims', 'disclose', 'disclosed',
'discloses', 'disclosing', 'discontinuance', 'discontinuances', 'discontinuation',
'discontinuations', 'discontinue', 'discontinued', 'discontinues', 'discontinuing', 'discourage',
'discouraged', 'discourages', 'discouraging', 'discredit', 'discredited', 'discrediting',
'discredits', 'discrepancies', 'discrepancy', 'disfavor', 'disfavored', 'disfavoring',
'disfavors', 'disgorge', 'disgorged', 'disgorgement', 'disgorgements', 'disgorges', 'disgorging',
'disgrace', 'disgraceful', 'disgracefully', 'dishonest', 'dishonestly', 'dishonesty', 'dishonor',
'dishonorable', 'dishonorably', 'dishonored', 'dishonoring', 'dishonors', 'disincentives',
'disinterested', 'disinterestedly', 'disinterestedness', 'disloyal', 'disloyally', 'disloyalty',
'dismal', 'dismally', 'dismiss', 'dismissal', 'dismissals', 'dismissed', 'dismisses',
'dismissing', 'disorderly', 'disparage', 'disparaged', 'disparagement', 'disparagements',
'disparages', 'disparaging', 'disparagingly', 'disparities', 'disparity', 'displace',
'displaced', 'displacement', 'displacements', 'displaces', 'displacing', 'dispose', 'dispossess',
'dispossessed', 'dispossesses', 'dispossessing', 'disproportion', 'disproportional',
'disproportionate', 'disproportionately', 'dispute', 'disputed', 'disputes', 'disputing',
'disqualification', 'disqualifications', 'disqualified', 'disqualifies', 'disqualify',
'disqualifying', 'disregard', 'disregarded', 'disregarding', 'disregards', 'disreputable',
'disrepute', 'disrupt', 'disrupted', 'disrupting', 'disruption', 'disruptions', 'disruptive',
'disrupts', 'dissatisfaction', 'dissatisfied', 'dissent', 'dissented', 'dissenter', 'dissenters',
'dissenting', 'dissents', 'dissident', 'dissidents', 'dissolution', 'dissolutions', 'distort',
'distorted', 'distorting', 'distortion', 'distortions', 'distorts', 'distract', 'distracted',
'distracting', 'distraction', 'distractions', 'distracts', 'distress', 'distressed', 'disturb',
'disturbance', 'disturbances', 'disturbed', 'disturbing', 'disturbs', 'diversion', 'divert',
'diverted', 'diverting', 'diverts', 'divest', 'divested', 'divesting', 'divestiture',
'divestitures', 'divestment', 'divestments', 'divests', 'divorce', 'divorced', 'divulge',
'divulged', 'divulges', 'divulging', 'doubt', 'doubted', 'doubtful', 'doubts', 'downgrade',
'downgraded', 'downgrades', 'downgrading', 'downsize', 'downsized', 'downsizes', 'downsizing',
'downsizings', 'downtime', 'downtimes', 'downturn', 'downturns', 'downward', 'downwards', 'drag',
'drastic', 'drastically', 'drawback', 'drawbacks', 'drop', 'dropped', 'drought', 'droughts', 'duress',
'dysfunction', 'dysfunctional', 'dysfunctions', 'easing', 'egregious', 'egregiously', 'embargo',
'embargoed', 'embargoes', 'embargoing', 'embarrass', 'embarrassed', 'embarrasses',
'embarrassing', 'embarrassment', 'embarrassments', 'embezzle', 'embezzled', 'embezzlement',
'embezzlements', 'embezzler', 'embezzles', 'embezzling', 'encroach', 'encroached', 'encroaches',
'encroaching', 'encroachment', 'encroachments', 'encumber', 'encumbered', 'encumbering',
'encumbers', 'encumbrance', 'encumbrances', 'endanger', 'endangered', 'endangering',
'endangerment', 'endangers', 'enjoin', 'enjoined', 'enjoining', 'enjoins', 'erode', 'eroded',
'erodes', 'eroding', 'erosion', 'erratic', 'erratically', 'erred', 'erring', 'erroneous',
'erroneously', 'error', 'errors', 'errs', 'escalate', 'escalated', 'escalates', 'escalating',
'evade', 'evaded', 'evades', 'evading', 'evasion', 'evasions', 'evasive', 'evict', 'evicted',
'evicting', 'eviction', 'evictions', 'evicts', 'exacerbate', 'exacerbated', 'exacerbates',
'exacerbating', 'exacerbation', 'exacerbations', 'exaggerate', 'exaggerated', 'exaggerates',
'exaggerating', 'exaggeration', 'excessive', 'excessively', 'exculpate', 'exculpated',
'exculpates', 'exculpating', 'exculpation', 'exculpations', 'exculpatory', 'exonerate',
'exonerated', 'exonerates', 'exonerating', 'exoneration', 'exonerations', 'exploit',
'exploitation', 'exploitations', 'exploitative', 'exploited', 'exploiting', 'exploits', 'expose',
'exposed', 'exposes', 'exposing', 'expropriate', 'expropriated', 'expropriates', 'expropriating',
'expropriation', 'expropriations', 'expulsion', 'expulsions', 'extenuating', 'fail', 'failed',
'failing', 'failings', 'fails', 'failure', 'failures', 'fallout', 'false', 'falsely',
'falsification', 'falsifications', 'falsified', 'falsifies', 'falsify', 'falsifying', 'falsity',
'fatalities', 'fatality', 'fatally', 'fault', 'faulted', 'faults', 'faulty', 'fear', 'fears',
'felonies', 'felonious', 'felony', 'fictitious', 'fined', 'fines', 'fired', 'firing', 'flaw',
'flawed', 'flaws', 'forbid', 'forbidden', 'forbidding', 'forbids', 'force', 'forced', 'forcing',
'foreclose', 'foreclosed', 'forecloses', 'foreclosing', 'foreclosure', 'foreclosures', 'forego',
'foregoes', 'foregone', 'forestall', 'forestalled', 'forestalling', 'forestalls', 'forfeit',
'forfeited', 'forfeiting', 'forfeits', 'forfeiture', 'forfeitures', 'forgers', 'forgery',
'fraud', 'frauds', 'fraudulence', 'fraudulent', 'fraudulently', 'frivolous', 'frivolously',
'frustrate', 'frustrated', 'frustrates', 'frustrating', 'frustratingly', 'frustration',
'frustrations', 'fugitive', 'fugitives', 'gratuitous', 'gratuitously', 'grievance', 'grievances',
'grossly', 'groundless', 'guilty', 'halt', 'halted', 'hamper', 'hampered', 'hampering',
'hampers', 'harass', 'harassed', 'harassing', 'harassment', 'hardship', 'hardships', 'harm',
'harmed', 'harmful', 'harmfully', 'harming', 'harms', 'harsh', 'harsher', 'harshest', 'harshly',
'harshness', 'hazard', 'hazardous', 'hazards', 'hinder', 'hindered', 'hindering', 'hinders',
'hindrance', 'hindrances', 'hostile', 'hostility', 'hurt', 'hurting', 'idle', 'idled', 'idling',
'ignore', 'ignored', 'ignores', 'ignoring', 'ill', 'illegal', 'illegalities', 'illegality',
'illegally', 'illegible', 'illicit', 'illicitly', 'illiquid', 'illiquidity', 'imbalance',
'imbalances', 'immature', 'immoral', 'impair', 'impaired', 'impairing', 'impairment',
'impairments', 'impairs', 'impasse', 'impasses', 'impede', 'impeded', 'impedes', 'impediment',
'impediments', 'impeding', 'impending', 'imperative', 'imperfection', 'imperfections', 'imperil',
'impermissible', 'implicate', 'implicated', 'implicates', 'implicating', 'impossibility',
'impossible', 'impound', 'impounded', 'impounding', 'impounds', 'impracticable', 'impractical',
'impracticalities', 'impracticality', 'imprisonment', 'improper', 'improperly', 'improprieties',
'impropriety', 'imprudent', 'imprudently', 'inability', 'inaccessible', 'inaccuracies',
'inaccuracy', 'inaccurate', 'inaccurately', 'inaction', 'inactions', 'inactivate', 'inactivated',
'inactivates', 'inactivating', 'inactivation', 'inactivations', 'inactivity', 'inadequacies',
'inadequacy', 'inadequate', 'inadequately', 'inadvertent', 'inadvertently', 'inadvisability',
'inadvisable', 'inappropriate', 'inappropriately', 'inattention', 'incapable', 'incapacitated',
'incapacity', 'incarcerate', 'incarcerated', 'incarcerates', 'incarcerating', 'incarceration',
'incarcerations', 'incidence', 'incidences', 'incident', 'incidents', 'incompatibilities',
'incompatibility', 'incompatible', 'incompetence', 'incompetency', 'incompetent',
'incompetently', 'incompetents', 'incomplete', 'incompletely', 'incompleteness', 'inconclusive',
'inconsistencies', 'inconsistency', 'inconsistent', 'inconsistently', 'inconvenience',
'inconveniences', 'inconvenient', 'incorrect', 'incorrectly', 'incorrectness', 'indecency',
'indecent', 'indefeasible', 'indefeasibly', 'indict', 'indictable', 'indicted', 'indicting',
'indictment', 'indictments', 'ineffective', 'ineffectively', 'ineffectiveness', 'inefficiencies',
'inefficiency', 'inefficient', 'inefficiently', 'ineligibility', 'ineligible', 'inequitable',
'inequitably', 'inequities', 'inequity', 'inevitable', 'inexperience', 'inexperienced',
'inferior', 'inflicted', 'infraction', 'infractions', 'infringe', 'infringed', 'infringement',
'infringements', 'infringes', 'infringing', 'inhibited', 'inimical', 'injunction', 'injunctions',
'injure', 'injured', 'injures', 'injuries', 'injuring', 'injurious', 'injury', 'inordinate',
'inordinately', 'inquiry', 'insecure', 'insensitive', 'insolvencies', 'insolvency', 'insolvent',
'instability', 'insubordination', 'insufficiency', 'insufficient', 'insufficiently',
'insurrection', 'insurrections', 'intentional', 'interfere', 'interfered', 'interference',
'interferences', 'interferes', 'interfering', 'intermittent', 'intermittently', 'interrupt',
'interrupted', 'interrupting', 'interruption', 'interruptions', 'interrupts', 'intimidation',
'intrusion', 'invalid', 'invalidate', 'invalidated', 'invalidates', 'invalidating',
'invalidation', 'invalidity', 'investigate', 'investigated', 'investigates', 'investigating',
'investigation', 'investigations', 'involuntarily', 'involuntary', 'irreconcilable',
'irreconcilably', 'irrecoverable', 'irrecoverably', 'irregular', 'irregularities',
'irregularity', 'irregularly', 'irreparable', 'irreparably', 'irreversible', 'jeopardize',
'jeopardized', 'justifiable', 'kickback', 'kickbacks', 'knowingly', 'lack', 'lacked', 'lacking',
'lackluster', 'lacks', 'lag', 'lagged', 'lagging', 'lags', 'lapse', 'lapsed', 'lapses',
'lapsing', 'late', 'laundering', 'layoff', 'layoffs', 'lie', 'limitation', 'limitations',
'lingering', 'liquidate', 'liquidated', 'liquidates', 'liquidating', 'liquidation',
'liquidations', 'liquidator', 'liquidators', 'litigant', 'litigants', 'litigate', 'litigated',
'litigates', 'litigating', 'litigation', 'litigations', 'lockout', 'lockouts', 'lose', 'loses',
'losing', 'loss', 'losses', 'lost', 'lying', 'malfeasance', 'malfunction', 'malfunctioned',
'malfunctioning', 'malfunctions', 'malice', 'malicious', 'maliciously', 'malpractice',
'manipulate', 'manipulated', 'manipulates', 'manipulating', 'manipulation', 'manipulations',
'manipulative', 'markdown', 'markdowns', 'misapplication', 'misapplications', 'misapplied',
'misapplies', 'misapply', 'misapplying', 'misappropriate', 'misappropriated', 'misappropriates',
'misappropriating', 'misappropriation', 'misappropriations', 'misbranded', 'miscalculate',
'miscalculated', 'miscalculates', 'miscalculating', 'miscalculation', 'miscalculations',
'mischaracterization', 'mischief', 'misclassification', 'misclassifications', 'misclassified',
'misclassify', 'miscommunication', 'misconduct', 'misdated', 'misdemeanor', 'misdemeanors',
'misdirected', 'mishandle', 'mishandled', 'mishandles', 'mishandling', 'misinform',
'misinformation', 'misinformed', 'misinforming', 'misinforms', 'misinterpret',
'misinterpretation', 'misinterpretations', 'misinterpreted', 'misinterpreting', 'misinterprets',
'misjudge', 'misjudged', 'misjudges', 'misjudging', 'misjudgment', 'misjudgments', 'mislabel',
'mislabeled', 'mislabeling', 'mislabelled', 'mislabels', 'mislead', 'misleading', 'misleadingly',
'misleads', 'misled', 'mismanage', 'mismanaged', 'mismanagement', 'mismanages', 'mismanaging',
'mismatch', 'mismatched', 'mismatches', 'mismatching', 'misplaced', 'misprice', 'mispricing',
'mispricings', 'misrepresent', 'misrepresentation', 'misrepresentations', 'misrepresented',
'misrepresenting', 'misrepresents', 'miss', 'missed', 'misses', 'misstate', 'misstated',
'misstatement', 'misstatements', 'misstates', 'misstating', 'misstep', 'missteps', 'mistake',
'mistaken', 'mistakenly', 'mistakes', 'mistaking', 'mistrial', 'mistrials', 'misunderstand',
'misunderstanding', 'misunderstandings', 'misunderstood', 'misuse', 'misused', 'misuses',
'misusing', 'monopolistic', 'monopolists', 'monopolization', 'monopolize', 'monopolized',
'monopolizes', 'monopolizing', 'monopoly', 'moratoria', 'moratorium', 'moratoriums',
'mothballed', 'mothballing', 'negative', 'negatively', 'negatives', 'neglect', 'neglected',
'neglectful', 'neglecting', 'neglects', 'negligence', 'negligences', 'negligent', 'negligently',
'nonattainment', 'noncompetitive', 'noncompliance', 'noncompliances', 'noncompliant',
'noncomplying', 'nonconforming', 'nonconformities', 'nonconformity', 'nondisclosure',
'nonfunctional', 'nonpayment', 'nonpayments', 'nonperformance', 'nonperformances',
'nonperforming', 'nonproducing', 'nonproductive', 'nonrecoverable', 'nonrenewal', 'nuisance',
'nuisances', 'nullification', 'nullifications', 'nullified', 'nullifies', 'nullify',
'nullifying', 'objected', 'objecting', 'objection', 'objectionable', 'objectionably',
'objections', 'obscene', 'obscenity', 'obsolescence', 'obsolete', 'obstacle', 'obstacles',
'obstruct', 'obstructed', 'obstructing', 'obstruction', 'obstructions', 'offence', 'offences',
'offend', 'offended', 'offender', 'offenders', 'offending', 'offends', 'omission', 'omissions',
'omit', 'omits', 'omitted', 'omitting', 'onerous', 'opportunistic', 'opportunistically',
'oppose', 'opposed', 'opposes', 'opposing', 'opposition', 'oppositions', 'outage', 'outages',
'outdated', 'outmoded', 'overage', 'overages', 'overbuild', 'overbuilding', 'overbuilds',
'overbuilt', 'overburden', 'overburdened', 'overburdening', 'overcapacities', 'overcapacity',
'overcharge', 'overcharged', 'overcharges', 'overcharging', 'overcome', 'overcomes',
'overcoming', 'overdue', 'overestimate', 'overestimated', 'overestimates', 'overestimating',
'overestimation', 'overestimations', 'overload', 'overloaded', 'overloading', 'overloads',
'overlook', 'overlooked', 'overlooking', 'overlooks', 'overpaid', 'overpayment', 'overpayments',
'overproduced', 'overproduces', 'overproducing', 'overproduction', 'overrun', 'overrunning',
'overruns', 'overshadow', 'overshadowed', 'overshadowing', 'overshadows', 'overstate',
'overstated', 'overstatement', 'overstatements', 'overstates', 'overstating', 'oversupplied',
'oversupplies', 'oversupply', 'oversupplying', 'overtly', 'overturn', 'overturned',
'overturning', 'overturns', 'overvalue', 'overvalued', 'overvaluing', 'panic', 'panics',
'penalize', 'penalized', 'penalizes', 'penalizing', 'penalties', 'penalty', 'peril', 'perils',
'perjury', 'perpetrate', 'perpetrated', 'perpetrates', 'perpetrating', 'perpetration', 'persist',
'persisted', 'persistence', 'persistent', 'persistently', 'persisting', 'persists', 'pervasive',
'pervasively', 'pervasiveness', 'petty', 'picket', 'picketed', 'picketing', 'plaintiff',
'plaintiffs', 'plea', 'plead', 'pleaded', 'pleading', 'pleadings', 'pleads', 'pleas', 'pled',
'poor', 'poorly', 'poses', 'posing', 'postpone', 'postponed', 'postponement', 'postponements',
'postpones', 'postponing', 'precipitated', 'precipitous', 'precipitously', 'preclude',
'precluded', 'precludes', 'precluding', 'predatory', 'prejudice', 'prejudiced', 'prejudices',
'prejudicial', 'prejudicing', 'premature', 'prematurely', 'pressing', 'pretrial', 'preventing',
'prevention', 'prevents', 'problem', 'problematic', 'problematical', 'problems', 'prolong',
'prolongation', 'prolongations', 'prolonged', 'prolonging', 'prolongs', 'prone', 'prosecute',
'prosecuted', 'prosecutes', 'prosecuting', 'prosecution', 'prosecutions', 'protest', 'protested',
'protester', 'protesters', 'protesting', 'protestor', 'protestors', 'protests', 'protracted',
'protraction', 'provoke', 'provoked', 'provokes', 'provoking', 'punished', 'punishes',
'punishing', 'punishment', 'punishments', 'punitive', 'purport', 'purported', 'purportedly',
'purporting', 'purports', 'question', 'questionable', 'questionably', 'questioned',
'questioning', 'questions', 'quit', 'quitting', 'racketeer', 'racketeering', 'rationalization',
'rationalizations', 'rationalize', 'rationalized', 'rationalizes', 'rationalizing',
'reassessment', 'reassessments', 'reassign', 'reassigned', 'reassigning', 'reassignment',
'reassignments', 'reassigns', 'recall', 'recalled', 'recalling', 'recalls', 'recession',
'recessionary', 'recessions', 'reckless', 'recklessly', 'recklessness', 'redact', 'redacted',
'redacting', 'redaction', 'redactions', 'redefault', 'redefaulted', 'redefaults', 'redress',
'redressed', 'redresses', 'redressing', 'refusal', 'refusals', 'refuse', 'refused', 'refuses',
'refusing', 'reject', 'rejected', 'rejecting', 'rejection', 'rejections', 'rejects',
'relinquish', 'relinquished', 'relinquishes', 'relinquishing', 'relinquishment',
'relinquishments', 'reluctance', 'reluctant', 'renegotiate', 'renegotiated', 'renegotiates',
'renegotiating', 'renegotiation', 'renegotiations', 'renounce', 'renounced', 'renouncement',
'renouncements', 'renounces', 'renouncing', 'reparation', 'reparations', 'repossessed',
'repossesses', 'repossessing', 'repossession', 'repossessions', 'repudiate', 'repudiated',
'repudiates', 'repudiating', 'repudiation', 'repudiations', 'resign', 'resignation',
'resignations', 'resigned', 'resigning', 'resigns', 'restate', 'restated', 'restatement',
'restatements', 'restates', 'restating', 'restructure', 'restructured', 'restructures',
'restructuring', 'restructurings', 'retaliate', 'retaliated', 'retaliates', 'retaliating',
'retaliation', 'retaliations', 'retaliatory', 'retribution', 'retributions', 'revocation',
'revocations', 'revoke', 'revoked', 'revokes', 'revoking', 'ridicule', 'ridiculed', 'ridicules',
'ridiculing', 'riskier', 'riskiest', 'risky', 'sabotage', 'sacrifice', 'sacrificed',
'sacrifices', 'sacrificial', 'sacrificing', 'scandalous', 'scandals', 'scrutinize',
'scrutinized', 'scrutinizes', 'scrutinizing', 'scrutiny', 'secrecy', 'seize', 'seized', 'seizes',
'seizing', 'sentenced', 'sentencing', 'serious', 'seriously', 'seriousness', 'setback',
'setbacks', 'sever', 'severe', 'severed', 'severely', 'severities', 'severity', 'sharply',
'shocked', 'shortage', 'shortages', 'shortfall', 'shortfalls', 'shrinkage', 'shrinkages', 'shut',
'shutdown', 'shutdowns', 'shuts', 'shutting', 'slander', 'slandered', 'slanderous', 'slanders',
'slippage', 'slippages', 'slow', 'slowdown', 'slowdowns', 'slowed', 'slower', 'slowest',
'slowing', 'slowly', 'slowness', 'sluggish', 'sluggishly', 'sluggishness', 'solvencies',
'solvency', 'spam', 'spammers', 'spamming', 'staggering', 'stagnant', 'stagnate', 'stagnated',
'stagnates', 'stagnating', 'stagnation', 'standstill', 'standstills', 'stolen', 'stoppage',
'stoppages', 'stopped', 'stopping', 'stops', 'strain', 'strained', 'straining', 'strains',
'stress', 'stressed', 'stresses', 'stressful', 'stressing', 'stringent', 'strong', 'subjected',
'subjecting', 'subjection', 'subpoena', 'subpoenaed', 'subpoenas', 'substandard', 'sue', 'sued',
'sues', 'suffer', 'suffered', 'suffering', 'suffers', 'suing', 'summoned', 'summoning',
'summons', 'summonses', 'susceptibility', 'susceptible', 'suspect', 'suspected', 'suspects',
'suspend', 'suspended', 'suspending', 'suspends', 'suspension', 'suspensions', 'suspicion',
'suspicions', 'suspicious', 'suspiciously', 'taint', 'tainted', 'tainting', 'taints', 'tampered',
'tense', 'terminate', 'terminated', 'terminates', 'terminating', 'termination', 'terminations',
'testify', 'testifying', 'threat', 'threaten', 'threatened', 'threatening', 'threatens',
'threats', 'tightening', 'tolerate', 'tolerated', 'tolerates', 'tolerating', 'toleration',
'tortuous', 'tortuously', 'tragedies', 'tragedy', 'tragic', 'tragically', 'traumatic', 'trouble',
'troubled', 'troubles', 'turbulence', 'turmoil', 'unable', 'unacceptable', 'unacceptably',
'unaccounted', 'unannounced', 'unanticipated', 'unapproved', 'unattractive', 'unauthorized',
'unavailability', 'unavailable', 'unavoidable', 'unavoidably', 'unaware', 'uncollectable',
'uncollected', 'uncollectibility', 'uncollectible', 'uncollectibles', 'uncompetitive',
'uncompleted', 'unconscionable', 'unconscionably', 'uncontrollable', 'uncontrollably',
'uncontrolled', 'uncorrected', 'uncover', 'uncovered', 'uncovering', 'uncovers', 'undeliverable',
'undelivered', 'undercapitalized', 'undercut', 'undercuts', 'undercutting', 'underestimate',
'underestimated', 'underestimates', 'underestimating', 'underestimation', 'underfunded',
'underinsured', 'undermine', 'undermined', 'undermines', 'undermining', 'underpaid',
'underpayment', 'underpayments', 'underpays', 'underperform', 'underperformance',
'underperformed', 'underperforming', 'underperforms', 'underproduced', 'underproduction',
'underreporting', 'understate', 'understated', 'understatement', 'understatements',
'understates', 'understating', 'underutilization', 'underutilized', 'undesirable', 'undesired',
'undetected', 'undetermined', 'undisclosed', 'undocumented', 'undue', 'unduly', 'uneconomic',
'uneconomical', 'uneconomically', 'unemployed', 'unemployment', 'unethical', 'unethically',
'unexcused', 'unexpected', 'unexpectedly', 'unfair', 'unfairly', 'unfavorability', 'unfavorable',
'unfavorably', 'unfavourable', 'unfeasible', 'unfit', 'unfitness', 'unforeseeable', 'unforeseen',
'unforseen', 'unfortunate', 'unfortunately', 'unfounded', 'unfriendly', 'unfulfilled',
'unfunded', 'uninsured', 'unintended', 'unintentional', 'unintentionally', 'unjust',
'unjustifiable', 'unjustifiably', 'unjustified', 'unjustly', 'unknowing', 'unknowingly',
'unlawful', 'unlawfully', 'unlicensed', 'unliquidated', 'unmarketable', 'unmerchantable',
'unmeritorious', 'unnecessarily', 'unnecessary', 'unneeded', 'unobtainable', 'unoccupied',
'unpaid', 'unperformed', 'unplanned', 'unpopular', 'unpredictability', 'unpredictable',
'unpredictably', 'unpredicted', 'unproductive', 'unprofitability', 'unprofitable', 'unqualified',
'unrealistic', 'unreasonable', 'unreasonableness', 'unreasonably', 'unreceptive',
'unrecoverable', 'unrecovered', 'unreimbursed', 'unreliable', 'unremedied', 'unreported',
'unresolved', 'unrest', 'unsafe', 'unsalable', 'unsaleable', 'unsatisfactory', 'unsatisfied',
'unsavory', 'unscheduled', 'unsellable', 'unsold', 'unsound', 'unstabilized', 'unstable',
'unsubstantiated', 'unsuccessful', 'unsuccessfully', 'unsuitability', 'unsuitable', 'unsuitably',
'unsuited', 'unsure', 'unsuspected', 'unsuspecting', 'unsustainable', 'untenable', 'untimely',
'untrusted', 'untruth', 'untruthful', 'untruthfully', 'untruthfulness', 'untruths', 'unusable',
'unwanted', 'unwarranted', 'unwelcome', 'unwilling', 'unwillingness', 'upset', 'urgency',
'urgent', 'usurious', 'usurp', 'usurped', 'usurping', 'usurps', 'usury', 'vandalism', 'verdict',
'verdicts', 'vetoed', 'victims', 'violate', 'violated', 'violates', 'violating', 'violation',
'violations', 'violative', 'violator', 'violators', 'violence', 'violent', 'violently',
'vitiate', 'vitiated', 'vitiates', 'vitiating', 'vitiation', 'voided', 'voiding', 'volatile',
'volatility', 'vulnerabilities', 'vulnerability', 'vulnerable', 'vulnerably', 'warn', 'warned',
'warning', 'warnings', 'warns', 'wasted', 'wasteful', 'wasting', 'weak', 'weaken', 'weakened',
'weakening', 'weakens', 'weaker', 'weakest', 'weakly', 'weakness', 'weaknesses', 'willfully',
'worries', 'worry', 'worrying', 'worse', 'worsen', 'worsened', 'worsening', 'worsens', 'worst',
'worthless', 'writedown', 'writedowns', 'writeoff', 'writeoffs', 'wrong', 'wrongdoing',
'wrongdoings', 'wrongful', 'wrongfully', 'wrongly',
'negative', 'negatives', 'fail', 'fails', 'failing', 'failure', 'weak', 'weakness', 'weaknesses',
'difficult', 'difficulty', 'hurdle', 'hurdles', 'obstacle', 'obstacles', 'slump', 'slumps',
'slumping', 'slumped', 'uncertain', 'uncertainty', 'unsettled', 'unfavorable', 'downturn',
'depressed', 'disappoint', 'disappoints', 'disappointing', 'disappointed', 'disappointment',
'risk', 'risks', 'risky', 'threat', 'threats', 'penalty', 'penalties', 'down', 'decrease',
'decreases', 'decreasing', 'decreased', 'decline', 'declines', 'declining', 'declined', 'fall',
'falls', 'falling', 'fell', 'fallen', 'drop', 'drops', 'dropping', 'dropped', 'deteriorate',
'deteriorates', 'deteriorating', 'deteriorated', 'worsen', 'worsens', 'worsening', 'weaken',
'weakens', 'weakening', 'weakened', 'worse', 'worst', 'low', 'lower', 'lowest', 'less', 'least',
'smaller', 'smallest', 'shrink', 'shrinks', 'shrinking', 'shrunk', 'below', 'under', 'challenge',
'challenges', 'challenging', 'challenged'
],
'Positive': ['able', 'abundance', 'abundant', 'acclaimed', 'accomplish', 'accomplished', 'accomplishes',
'accomplishing', 'accomplishment', 'accomplishments', 'achieve', 'achieved', 'achievement',
'achievements', 'achieves', 'achieving', 'adequately', 'advancement', 'advancements', 'advances',
'advancing', 'advantage', 'advantaged', 'advantageous', 'advantageously', 'advantages',
'alliance', 'alliances', 'assure', 'assured', 'assures', 'assuring', 'attain', 'attained',
'attaining', 'attainment', 'attainments', 'attains', 'attractive', 'attractiveness', 'beautiful',
'beautifully', 'beneficial', 'beneficially', 'benefit', 'benefited', 'benefiting', 'benefitted',
'benefitting', 'best', 'better', 'bolstered', 'bolstering', 'bolsters', 'boom', 'booming',
'boost', 'boosted', 'breakthrough', 'breakthroughs', 'brilliant', 'charitable', 'collaborate',
'collaborated', 'collaborates', 'collaborating', 'collaboration', 'collaborations',
'collaborative', 'collaborator', 'collaborators', 'compliment', 'complimentary', 'complimented',
'complimenting', 'compliments', 'conclusive', 'conclusively', 'conducive', 'confident',
'constructive', 'constructively', 'courteous', 'creative', 'creatively', 'creativeness',
'creativity', 'delight', 'delighted', 'delightful', 'delightfully', 'delighting', 'delights',
'dependability', 'dependable', 'desirable', 'desired', 'despite', 'destined', 'diligent',
'diligently', 'distinction', 'distinctions', 'distinctive', 'distinctively', 'distinctiveness',
'dream', 'easier', 'easily', 'easy', 'effective', 'efficiencies', 'efficiency', 'efficient',
'efficiently', 'empower', 'empowered', 'empowering', 'empowers', 'enable', 'enabled', 'enables',
'enabling', 'encouraged', 'encouragement', 'encourages', 'encouraging', 'enhance', 'enhanced',
'enhancement', 'enhancements', 'enhances', 'enhancing', 'enjoy', 'enjoyable', 'enjoyably',
'enjoyed', 'enjoying', 'enjoyment', 'enjoys', 'enthusiasm', 'enthusiastic', 'enthusiastically',
'excellence', 'excellent', 'excelling', 'excels', 'exceptional', 'exceptionally', 'excited',
'excitement', 'exciting', 'exclusive', 'exclusively', 'exclusiveness', 'exclusives',
'exclusivity', 'exemplary', 'fantastic', 'favorable', 'favorably', 'favored', 'favoring',
'favorite', 'favorites', 'friendly', 'gain', 'gained', 'gaining', 'gains', 'good', 'great',
'greater', 'greatest', 'greatly', 'greatness', 'happiest', 'happily', 'happiness', 'happy',
'highest', 'honor', 'honorable', 'honored', 'honoring', 'honors', 'ideal', 'impress',
'impressed', 'impresses', 'impressing', 'impressive', 'impressively', 'improve', 'improved',
'improvement', 'improvements', 'improves', 'improving', 'incredible', 'incredibly',
'influential', 'informative', 'ingenuity', 'innovate', 'innovated', 'innovates', 'innovating',
'innovation', 'innovations', 'innovative', 'innovativeness', 'innovator', 'innovators',
'insightful', 'inspiration', 'inspirational', 'integrity', 'invent', 'invented', 'inventing',
'invention', 'inventions', 'inventive', 'inventiveness', 'inventor', 'inventors', 'leadership',
'leading', 'loyal', 'lucrative', 'meritorious', 'opportunities', 'opportunity', 'optimistic',
'outperform', 'outperformed', 'outperforming', 'outperforms', 'perfect', 'perfected',
'perfectly', 'perfects', 'pleasant', 'pleasantly', 'pleased', 'pleasure', 'plentiful', 'popular',
'popularity', 'positive', 'positively', 'preeminence', 'preeminent', 'premier', 'premiere',
'prestige', 'prestigious', 'proactive', 'proactively', 'proficiency', 'proficient',
'proficiently', 'profitability', 'profitable', 'profitably', 'progress', 'progressed',
'progresses', 'progressing', 'prospered', 'prospering', 'prosperity', 'prosperous', 'prospers',
'rebound', 'rebounded', 'rebounding', 'receptive', 'regain', 'regained', 'regaining', 'resolve',
'revolutionize', 'revolutionized', 'revolutionizes', 'revolutionizing', 'reward', 'rewarded',
'rewarding', 'rewards', 'satisfaction', 'satisfactorily', 'satisfactory', 'satisfied',
'satisfies', 'satisfy', 'satisfying', 'smooth', 'smoothing', 'smoothly', 'smooths', 'solves',
'solving', 'spectacular', 'spectacularly', 'stability', 'stabilization', 'stabilizations',
'stabilize', 'stabilized', 'stabilizes', 'stabilizing', 'stable', 'strength', 'strengthen',
'strengthened', 'strengthening', 'strengthens', 'strengths', 'strong', 'stronger', 'strongest',
'succeed', 'succeeded', 'succeeding', 'succeeds', 'success', 'successes', 'successful',
'successfully', 'superior', 'surpass', 'surpassed', 'surpasses', 'surpassing', "sustainable", 'transparency',
'tremendous', 'tremendously', 'unmatched', 'unparalleled', 'unsurpassed', 'upturn', 'upturns',
'valuable', 'versatile', 'versatility', 'vibrancy', 'vibrant', 'win', 'winner', 'winners', 'winning', 'worthy',
'positive', 'positives', 'success', 'successes', 'successful', 'succeed', 'succeeds',
'succeeding', 'succeeded', 'accomplish', 'accomplishes', 'accomplishing', 'accomplished',
'accomplishment', 'accomplishments', 'strong', 'strength', 'strengths', 'certain', 'certainty',
'definite', 'solid', 'excellent', 'good', 'leading', 'achieve', 'achieves', 'achieved',
'achieving', 'achievement', 'achievements', 'progress', 'progressing', 'deliver', 'delivers',
'delivered', 'delivering', 'leader', 'leading', 'pleased', 'reward', 'rewards', 'rewarding',
'rewarded', 'opportunity', 'opportunities', 'enjoy', 'enjoys', 'enjoying', 'enjoyed',
'encouraged', 'encouraging', 'up', 'increase', 'increases', 'increasing', 'increased', 'rise',
'rises', 'rising', 'rose', 'risen', 'improve', 'improves', 'improving', 'improved', 'improvement',
'improvements', 'strengthen', 'strengthens', 'strengthening', 'strengthened', 'stronger',
'strongest', 'better', 'best', 'more', 'most', 'above', 'record', 'high', 'higher', 'highest',
'greater', 'greatest', 'larger', 'largest', 'grow', 'grows', 'growing', 'grew', 'grown', 'growth',
'expand', 'expands', 'expanding', 'expanded', 'expansion', 'exceed', 'exceeds', 'exceeded',
'exceeding', 'beat', 'beats', 'beating']
}
negate = ["aint", "arent", "cannot", "cant", "couldnt", "darent", "didnt", "doesnt", "ain't", "aren't", "can't",
"couldn't", "daren't", "didn't", "doesn't", "dont", "hadnt", "hasnt", "havent", "isnt", "mightnt", "mustnt",
"neither", "don't", "hadn't", "hasn't", "haven't", "isn't", "mightn't", "mustn't", "neednt", "needn't",
"never", "none", "nope", "nor", "not", "nothing", "nowhere", "oughtnt", "shant", "shouldnt", "wasnt",
"werent", "oughtn't", "shan't", "shouldn't", "wasn't", "weren't", "without", "wont", "wouldnt", "won't",
"wouldn't", "rarely", "seldom", "despite", "no", "nobody"]
# + id="xi-NyGxvgFSG" colab_type="code" colab={}
def negator(word):
'''
Determine if the prior word is a negator.
'''
if word.lower() in negate:
return True
else:
return False
# + id="CaXCb7jWjLBQ" colab_type="code" colab={}
def bag_of_words_using_negator(word_dictionary, article):
'''
Count the number of positive and negative words, whilst considering negation for positive words.
Negation is considered as a negator preceeding a positive word by three words or less.
'''
# initialise word counts at 0:
positive_word_count = 0
negative_word_count = 0
# initialise empty lists:
positive_words = []
negative_words = []
# find all words:
input_words = re.findall(r'\b([a-zA-Z]+n\'t|[a-zA-Z]+\'s|[a-zA-Z]+)\b', article.lower())
# determine word_count:
word_count = len(input_words)
# for each word in the article:
for i in range(0, word_count):
# determine if negative. if so, incease negative count and append:
if input_words[i] in word_dictionary['Negative']:
negative_word_count += 1
negative_words.append(input_words[i])
# if the input word is positive, check the three prior words for negators:
if input_words[i] in word_dictionary['Positive']:
# if a negator exists 3 or less words prior, assign negative, otherwise positive:
if i >= 3:
if negator(input_words[i - 1]) or negator(input_words[i - 2]) or negator(input_words[i - 3]):
negative_word_count += 1
negative_words.append(input_words[i] + ' (with negation)')
else:
positive_word_count += 1
positive_words.append(input_words[i])
# if a negator exists 2 or less words prior, assign negative, otherwise positive:
elif i == 2:
if negator(input_words[i - 1]) or negator(input_words[i - 2]):
negative_word_count += 1
negative_words.append(input_words[i] + ' (with negation)')
else:
positive_word_count += 1
positive_words.append(input_words[i])
# if a negator exists 1 word prior, assign negative, otherwise positive:
elif i == 1:
if negator(input_words[i - 1]):
negative_word_count += 1
negative_words.append(input_words[i] + ' (with negation)')
else:
positive_word_count += 1
positive_words.append(input_words[i])
# otherwise assign positive:
elif i == 0:
positive_word_count += 1
positive_words.append(input_words[i])
# collect the findings as a list:
results = [word_count, positive_word_count, negative_word_count, positive_words, negative_words]
return results
# + id="v8Gd9B0bmljZ" colab_type="code" colab={}
def build_dataset(dataset):
'''
This function constructs the dataset by interatively calling the bag_of_words_using_negator
function, using the input argument of 'lmdict'.
'''
# call the bag_of_words_using_negator function for each policy statement, place results into DataFrame:
temporary = [bag_of_words_using_negator(lmdict,x) for x in dataset['FOMC_Statements']]
temporary = pd.DataFrame(temporary)
# Transpose the various columns to the initial 'dataset' DataFrame:
dataset['Total Word Count'] = temporary.iloc[:,0].values
dataset['Number of Positive Words'] = temporary.iloc[:,1].values
dataset['Number of Negative Words'] = temporary.iloc[:,2].values
dataset['Positive Words'] = temporary.iloc[:,3].values
dataset['Negative Words'] = temporary.iloc[:,4].values
# Calculate additional useful metrics:
dataset['Net Sentiment'] = (dataset['Number of Positive Words'] - dataset['Number of Negative Words'])
dataset['2 Year Sentiment MA'] = dataset['Net Sentiment'].rolling(window=16).mean()
dataset['Sentiment Change'] = (dataset['Net Sentiment'].shift(1) / dataset['Net Sentiment'])
dataset['Wordcount Normalized Net Sentiment'] = (dataset['Net Sentiment'] / dataset['Total Word Count'])
return dataset
# + id="dbWoTk_DoWKf" colab_type="code" colab={}
def plot_figure():
'''
This function constructs a Plotly chart by calling the build_dataset function
and subsequently plotting the relevant data.
'''
# call the build_dataset function, using the input argument of the pre-defined dataset:
data = build_dataset(dataset)
# initialise figure:
fig = go.Figure()
# add figure traces:
fig.add_trace(go.Scatter(x = data.index, y = data['Total Word Count'],
mode = 'lines',
name = 'Total Word Count',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['Number of Positive Words'],
mode = 'lines',
name = 'Number of Positive Words',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['Number of Negative Words'],
mode = 'lines',
name = 'Number of Negative Words',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['Net Sentiment'],
mode = 'lines',
name = 'Net Sentiment',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['2 Year Sentiment MA'],
mode = 'lines',
name = '2 Year Sentiment MA',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['Sentiment Change'],
mode = 'lines',
name = 'Sentiment Change',
connectgaps=True))
fig.add_trace(go.Scatter(x = data.index, y = data['Wordcount Normalized Net Sentiment'],
mode = 'lines',
name = 'Wordcount Normalized Net Sentiment',
connectgaps=True))
# add a rangeslider and buttons:
fig.update_xaxes(
rangeslider_visible=True,
rangeselector=dict(
buttons=list([
dict(count=1, label="YTD", step="year", stepmode="todate"),
dict(count=5, label="5 Years", step="year", stepmode="backward"),
dict(count=10, label="10 Years", step="year", stepmode="backward"),
dict(count=15, label="15 Years", step="year", stepmode="backward"),
dict(label="All", step="all")
])))
# add a chart title and axis title:
fig.update_layout(
title="Federal Reserve Bag of Words",
xaxis_title="Date",
yaxis_title="",
font=dict(
family="Arial",
size=11,
color="#7f7f7f"
))
# add toggle buttons for data display:
fig.update_layout(
updatemenus=[
dict(
buttons=list([
dict(
label = 'All',
method = 'update',
args = [{'visible': [True, True, True, True, True, True, True]}]
),
dict(
label = 'Word Count',
method = 'update',
args = [{'visible': [True, False, False, False, False, False, False]}]
),
dict(
label = 'Positive Words',
method = 'update',
args = [{'visible': [False, True, False, False, False, False, False,]}]
),
dict(
label = 'Negative Words',
method = 'update',
args = [{'visible': [False, False, True, False, False, False, False,]}]
),
dict(
label = 'Net Sentiment',
method = 'update',
args = [{'visible': [False, False, False, True, False, False, False,]}]
),
dict(
label = '2 Year Sentiment MA',
method = 'update',
args = [{'visible': [False, False, False, False, True, False, False,]}]
),
dict(
label = 'Sentiment Change',
method = 'update',
args = [{'visible': [False, False, False, False, False, True, False,]}]
),
dict(
label = 'Wordcount Normalized Net Sentiment',
method = 'update',
args = [{'visible': [False, False, False, False, False, False, True]}]
),
]),
direction="down",
pad={"r": 10, "t": 10},
showactive=True,
x=1.0,
xanchor="right",
y=1.2,
yanchor="top"
),])
return fig.show()
# + id="bgnFexSQoXEP" colab_type="code" colab={}
plot_figure()
|
Federal_Reserve_Bag_of_Words.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: models_eval_py2
# language: python
# name: models_eval_py2
# ---
# ### Imports
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from IPython.display import display
import json
import numpy as np
import pandas as pd
import os
import random
import re
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn.metrics as metrics
import tensorflow as tf
# -
# ### Read scored test data
# +
standard_data_path = 'gs://conversationai-models/biosbias/scored_data/test_standard_0409.csv'
scrubbed_data_path = 'gs://conversationai-models/biosbias/scored_data/test_scrubbed_0409.csv'
very_scrubbed_data_path = 'gs://conversationai-models/biosbias/scored_data/test_very_scrubbed_0409.csv'
gender_data_path = 'gs://conversationai-models/biosbias/scored_data/test_data_gender.csv'
perf_df = pd.read_csv(tf.gfile.Open(standard_data_path)).drop_duplicates(subset=['tokens'])
scrubbed_df = pd.read_csv(tf.gfile.Open(scrubbed_data_path)).drop_duplicates(subset=['tokens'])
very_scrubbed_df = pd.read_csv(tf.gfile.Open(very_scrubbed_data_path)).drop_duplicates(subset=['tokens'])
gender_df = pd.read_csv(tf.gfile.Open(gender_data_path)).drop_duplicates(subset=['tokens'])
# -
print(perf_df.shape)
print(scrubbed_df.shape)
df = perf_df.join(scrubbed_df, rsuffix = '_scrubbed')
df = df.join(very_scrubbed_df, rsuffix = '_very_scrubbed')
df.head()
df.shape
df = df.dropna()
print(df.shape)
# ### Preprocessing
def get_class_from_col_name(col_name):
#print(col_name)
pattern = r'^.*_(\d+)$'
return int(re.search(pattern, col_name).group(1))
def find_best_class(df, model_name, class_names):
model_class_names = ['{}_{}'.format(model_name, class_name) for class_name in class_names]
sub_df = df[model_class_names]
df['{}_class'.format(model_name)] = sub_df.idxmax(axis=1).apply(get_class_from_col_name)
# +
# Can check model names here
# df.columns.values
# -
# May have to change.
# Can look them up in experiment tracker.
MODEL_NAMES = {
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_174837': 'debiased_tolga',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_174941': 'debiased_biosbias',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175003': 'strong_debiased_1',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175019': 'strong_debiased_2',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175034': 'strong_debiased_3',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175055': 'strong_debiased_4',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190328_103117': 'glove',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175113': 'strong_no_equalize',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175131': 'strong_no_projection',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190315_112954': 'scrubbed',
'tf_trainer_tf_gru_attention_multiclass_biosbias_glove:v_20190410_175254': 'very_scrubbed'
}
CLASS_NAMES = range(33)
for _model in MODEL_NAMES:
find_best_class(df, _model, CLASS_NAMES)
# Labels with either gender having too few examples
bad_labels = df.groupby('label').gender.value_counts().reset_index(name = 'count').query('count < 5').label.values
assert len(bad_labels) == 0
# ### Accuracy Calculation
accuracy_list = []
for _model in MODEL_NAMES:
is_correct = (df['{}_class'.format(_model)] == df['label'])
_acc = sum(is_correct)/len(is_correct)
accuracy_list.append(_acc)
print ('Accuracy for model {}: {}'.format(MODEL_NAMES[_model], _acc))
# ### Fairness Metrics
for _class in CLASS_NAMES:
df['label_{}'.format(_class)] = (df['label'] == _class)
# Gender ratios of classes
gender_counts = df.groupby('label').gender.value_counts().reset_index(name = 'count')
def frac_female(df):
m_count = df[df['gender'] == "M"]['count'].values[0]
f_count = df[df['gender'] == "F"]['count'].values[0]
return {'label': df['label'].values[0], 'frac_female': f_count/(m_count+f_count)}
frac_female_df = pd.DataFrame(list(gender_counts.groupby('label', as_index = False).apply(frac_female)))
# +
def compute_tpr(df, _class, _model, threshold = 0.5):
tpr = metrics.recall_score(df['label_{}'.format(_class)],
df['{}_{}'.format(_model,_class)] > threshold)
return tpr
def compute_tpr_by_gender(df, _class, _model, threshold = 0.5):
tpr_m = compute_tpr(df.query('gender == "M"'), _class, _model, threshold)
tpr_f = compute_tpr(df.query('gender == "F"'), _class, _model, threshold)
return {'M': tpr_m, 'F': tpr_f}
# +
def compute_tpr_tnr(df, _class, _model, threshold = 0.5):
#cm = metrics.confusion_matrix(df['label_{}'.format(_class)],
# df['{}_{}'.format(_model,_class)] > threshold)
cm = pd.crosstab(df['label_{}'.format(_class)], df['{}_{}'.format(_model,_class)] > threshold)
#display(cm)
if cm.shape[0] > 1:
tn = cm.iloc[0,0]
fp = cm.iloc[0,1]
fn = cm.iloc[1,0]
tp = cm.iloc[1,1]
tpr = tp/(tp+fn)
tnr = tn/(tn+fp)
else:
tpr = 0
tnr = 1
return tpr, tnr
def compute_tr_by_gender(df, _class, _model, threshold = 0.5):
tpr_m, tnr_m = compute_tpr_tnr(df.query('gender == "M"'), _class, _model, threshold)
tpr_f, tnr_f = compute_tpr_tnr(df.query('gender == "F"'), _class, _model, threshold)
return {'TPR_m': tpr_m, 'TPR_f': tpr_f, 'TNR_m': tnr_m, 'TNR_f': tnr_f}
# -
for _class in CLASS_NAMES:
for _model in MODEL_NAMES:
tpr_1 = compute_tpr(df, _class, _model)
tpr_2, _ = compute_tpr_tnr(df, _class, _model)
assert tpr_1 == tpr_2, '{} != {}'.format(tpr_1, tpr_2)
#print('{} == {}'.format(tpr_1, tpr_2))
tpr_df = pd.DataFrame()
for _class in frac_female_df.label:
row = {}
row['label'] = _class
for _model, _model_type in MODEL_NAMES.items():
tpr, tnr = compute_tpr_tnr(df, _class, _model)
row['{}_tpr'.format(_model_type)] = tpr
row['{}_tnr'.format(_model_type)] = tnr
gender_trs = compute_tr_by_gender(df, _class, _model)
row['{}_tpr_F'.format(_model_type)] = gender_trs['TPR_f']
row['{}_tpr_M'.format(_model_type)] = gender_trs['TPR_m']
row['{}_tpr_gender_gap'.format(_model_type)] = gender_trs['TPR_f'] - gender_trs['TPR_m']
row['{}_tnr_F'.format(_model_type)] = gender_trs['TNR_f']
row['{}_tnr_M'.format(_model_type)] = gender_trs['TNR_m']
row['{}_tnr_gender_gap'.format(_model_type)] = gender_trs['TNR_f'] - gender_trs['TNR_m']
tpr_df = tpr_df.append(row, ignore_index = True)
results_df = pd.merge(tpr_df, frac_female_df, on = 'label')
TITLE_LABELS = [
'accountant', 'acupuncturist', 'architect', 'attorney', 'chiropractor', 'comedian', 'composer', 'dentist',
'dietitian', 'dj', 'filmmaker', 'interior_designer', 'journalist', 'landscape_architect', 'magician',
'massage_therapist', 'model', 'nurse', 'painter', 'paralegal', 'pastor', 'personal_trainer',
'photographer', 'physician', 'poet', 'professor', 'psychologist', 'rapper',
'real_estate_broker', 'software_engineer', 'surgeon', 'teacher', 'yoga_teacher']
results_df['label_profession'] = results_df['label'].apply(lambda x: TITLE_LABELS[int(x)])
results_df[['frac_female']+['{}_tpr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]].corr()[['frac_female']]
tpr_gender_gap_cols = ['{}_tpr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]
tnr_gender_gap_cols = ['{}_tnr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]
gender_gap_df = results_df[['label_profession', 'frac_female']+tpr_gender_gap_cols+tnr_gender_gap_cols]
#gender_gap_df.columns = ['label_profession', 'frac_female']+['{}'.format(_model) for _model in MODEL_NAMES.values()]
gender_gap_df.sort_values('frac_female', ascending = False)
# +
# Fraction of comments where new model has lower
# TPR gap than the baseline
def compute_fraction_improved(df, baseline_model, improved_model):
is_improved = np.abs(df[baseline_model]) >= np.abs(df[improved_model])
return np.mean(is_improved)
# -
for _model in MODEL_NAMES.values():
print(_model)
print(compute_fraction_improved(gender_gap_df, 'glove_tpr_gender_gap', '{}_tpr_gender_gap'.format(_model)))
tpr_cols = ['{}_tpr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]
tnr_cols = ['{}_tnr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]
gender_gap_cols = tpr_cols + tnr_cols
gender_gap_df[gender_gap_cols].apply(lambda x: np.mean(x**2))
gender_gap_df[gender_gap_cols].apply(lambda x: np.mean(np.abs(x)))
def plot_tpr_gap(df, _model):
fig, ax = plt.subplots(figsize=(15, 6))
x = 'frac_female'
y = '{}_tpr_gender_gap'.format(_model)
p1 = sns.regplot(x = x, y = y, data = df)
p1.set(xlabel = "% Female", ylabel = "TPR Gender Gap", title = _model)
for line in range(0,df.shape[0]):
p1.text(results_df[x][line]+0.01, df[y][line], df['label_profession'][line], horizontalalignment='left', size='medium', color='black')
plt.show()
for _model in MODEL_NAMES.values():
if 'untuned' in _model:
plot_tpr_gap(results_df, _model)
results_df[['frac_female']+['{}_tpr_gender_gap'.format(_model) for _model in MODEL_NAMES.values()]].corr()[['frac_female']]
# ### Gender Prediction Analysis
# Which model does this correspond to?
model_name = 'tf_gru_attention_multiclass_gender_biosbias_glove:v_20190405_142640'
gender_df['correct'] = ((gender_df['gender'] == 'M') == gender_df[model_name])
acc = gender_df.correct.sum()/gender_df.correct.count()
print('Accuracy: {:.4f}'.format(acc))
|
model_evaluation/BiosBias Evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 7 Fungsi Tanggal Pada Pandas Yang Wajib Diketahui
#
# Artikel berikut merupakan tutorial Pandas yang akan membahasa mengenai fungsi-fungsi Pandas yang wajib diketahui jika berurusan dengan tipe data **DATE**. Seringkali dalam analisa data kita menggunakan DATE atau tanggal sebagai sebuah dimensi atau variable atau feature, karena banyak hal yang dihitung berdasarkan tanggal.
#
# Artikel tutorial Pandas ini dibuat karena ada seorang teman yang bertanya mengenai salah satu fungsi yang berhubungan dengan tanggal, setelah tutorial Pandas sebelumnya 1 dan 2 dipublikasikan. Yuk langsung saja kita **KODING !**
# ## Dataset
# Dataset yang digunakan adalah data sintetis yang terdiri dari tanggal dengan **3 format penulisan tanggal** dan field **value**.
# +
import pandas as pd
df_sample = pd.DataFrame(
{'date_fmt1' :['4/1/20', '4/2/20', '4/3/20', '4/4/20', '4/5/20', '4/6/20', '4/7/20'],
'date_fmt2' :['20200401', '20200402', '20200403', '20200404', '20200405', '20200406', '20200407'],
'date_fmt3' :['Apr.01.2020', 'Apr.02.2020', 'Apr.03.2020', 'Apr.04.2020', 'Apr.05.2020', 'Apr.06.2020', 'Apr.07.2020'],
'value' :[103, 112, 134, 150, 164, 192, 204]
})
df_sample
# -
df_sample.dtypes
# ## 1. Merubah format string ke tipe date
# Panda memiliki fungsi **to_datetime** yang bisa digunakan untuk mengkonversi STRING menjadi tipe data DATETIME. Format pembacaan tanggal dapat disesuaikan dengan menggunakan parameter **format**.
# Berikut ini kita akan mengkonversi field date_fmt1 yang memiliki format **'%m/%d/%y'** ke dalam field baru yaitu **date01**
df_sample['date01'] = pd.to_datetime(df_sample['date_fmt1'], format='%m/%d/%y')
df_sample
# Sebagai contoh lain adalah format tanggal seperti 20200401, jika dibaca maka menggunakan format **'%Y%m%d'**
df_sample['date02'] = pd.to_datetime(df_sample.date_fmt2, format='%Y%m%d')
df_sample
# Jika penulisan bulan dengan menggunakan format singkatan 3 karakter nama bulan, seperti Apr untuk April, maka parameter format untuk pembacaannya dapat menggunakan **%b**
df_sample['date03'] = pd.to_datetime(df_sample['date_fmt3'], format='%b.%d.%Y')
df_sample
# Terlihat dibawah, bahwa field **date01**, **date02** dan **date03** memiliki tipe data **datetime64**
df_sample.dtypes
# ## 2. Mengekstrak Feature Pada Date
# Sebelum meneruskan, kita melakukan persiapan untuk dataframe yang akan digunakan, yaitu membentuknya menjadi 2 field yaitu **date_id** dan **value**
df = df_sample[['date01', 'value']].copy()
df.rename(columns={'date01': 'date_id'}, inplace=True)
df.dtypes
# Dengan tipe data **datetime64**, kita akan dengan mudah mengekstrak fitur, seperti tahun, bulan, tanggal dan fitur lain dengan mudah. Panda menyediakan beberapa atribut yang dapat digunakan seperti dt.year, dt.month, dt.day dan dt.quarter untuk memperoleh nilai tahun, bulan, tanggal dan quarter.
df['year'] = df['date_id'].dt.year
df['month'] = df['date_id'].dt.month
df['day'] = df['date_id'].dt.day
df['querter-of-year'] = df['date_id'].dt.quarter
# Untuk memperoleh nilai minggu, dapat menggunakan **dt.isocalendar().week**. Atrubut **dt.weekofyear** and **dt.week** disarankan untuk tidak digunakan lagi.
df['week-of-year'] = df['date_id'].dt.isocalendar().week
# Hasil yang didapat adalah
df
# ## 3. Menghitung Selisih Antar Tanggal
# Menghitung selisih hari biasanya digunakan ketika menghitung durasi dalam satuan hari. Untuk melakukan perhitungan selisih hari antara 2 tanggal, dapat dilakukan dengan mudah jika telah memiliki tipe data **DATETIME64**
df['diff'] = df.date_id - pd.to_datetime('20200330', format='%Y%m%d')
df
# Untuk mengkonversikan menjadi numerik, dapat menngunakan atribut **dt.days**
df['diff-in-days'] = df['diff'].dt.days
df
df.dtypes
# ## 4. Operasi Penambahan n Hari
# Jika ingin melakukan penambahan hari, maka dapat dilakukan seperti kode dibawah ini
df['seven-days-later'] = df.date_id + pd.Timedelta(days=7)
df
# ## 5. Mengkonversi DATE ke format STRING
# Salah satu fungsi yang dapat digunakan untuk mengkonversi **DATE** ke tipe **STRING** adalah **dt.strftime**. Fungsi ini memiliki paramater untuk format yang diinginkan, seperti halnya fungsi to_datetime.
df['weekday-name'] = df.date_id.dt.strftime('%a')
df['month-name'] = df.date_id.dt.strftime('%b')
df['day-of-week'] = df.date_id.dt.strftime('%w')
df
# https://docs.python.org/3/library/datetime.html#strftime-and-strptime-behavior
# ## 6. Filtering
# Secara umum fungsi-fungsi yang telah dijelaskan diatas dapat digunakan untuk memfilter data. Misalkan kita akan memfilter untuk menampilkan data-data antara tanggal 3 April hingga 7 April
df[['date_id', 'value']] [(df.date_id > '04-03-2020') & (df.date_id < '04-07-2020')]
# Perintah diatas memiliki 3 bagian:
# 1. df adalah data frame yang digunakan
# 2. **[['date_id', 'value']]** adalah nama field yang akan ditampilkan
# 3. **[(df.date_id > '04-03-2020') & (df.date_id < '04-09-2020')]** adalah filter yang digunakan, dalam hal ini adalah untuk menampilkan data-data antara tanggal 3 April hingga 9 April
# Contoh lain adalah menampilkan data-data pada hari Rabu
df[['date_id', 'value']] [(df.date_id.dt.strftime('%a') == 'Wed')]
# ## 7. Filtering menggunakan index
# Jika akan melakukan banyak filtering menggunakan field **date_id**, akan lebih cepat jika field tanggal digunakan sebagai index, sehingga dapat memanfaatkan optimasi yang disediakan oleh Pandas.
df = df.set_index(['date_id'])
df
# Untuk memfilter berdasarkan urutan tanggal, misal data antara tanggal **5 April hingga 7 April**, dapat dilakukan dengan menggunakan index seperti berikut
df.loc['2020-04-05' : '2020-04-07']
# Untuk menampilkan data bulan **April** dengan menggunakan index, dapat dilakukan seperti berikut
df.loc['2020-4']
# ## Penutup
# Demikian tutorial Pandas, dengan 7 fungsi DATE yang bakalan sering kamu gunakan jika melakukan pengolahan data. Semoga kamu bisa memperoleh ide bagaimana (sebetulnya) mudah melakukan pengolahan data dengan menggunakan Pandas.
# Tutorial ini juga dipublikasikan di (idBigData
#
# Untuk yang belum pernah menggunakan Python dapat membaca Berkenalan dengan Python
#
# Enjoy learning and have fun with data !
|
7 Fungsi Tanggal Pada Pandas Yang Wajib Diketahui Oleh Data Engineer dan Data Scientist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
pd.set_option('display.max_colwidth', -1)
sinergise_list = 'data/sinergise-tiles.list.gz'
element84_list = 'data/element84-tiles.list.gz'
# + tags=[]
s = pd.read_csv(sinergise_list, header=None, names=['sinergise'])
e = pd.read_csv(element84_list, header=None, names=['element84'])
print(f"Found {len(s)} Sinergise and {len(e)} Element84 scenes.")
# -
s[['tile_id']] = s.sinergise.apply(
lambda x: pd.Series("{0[4]}{0[5]}{0[6]}_{0[7]}{0[8]:0>2}{0[9]:0>2}_{0[10]}".format(str(x).split("/"))))
e[['tile_id']] = e.element84.apply(
lambda x: pd.Series("{0}".format(str(x).split("/")[5][4:20])))
all_tiles = s.merge(e, on=['tile_id','tile_id'], how='outer', indicator=True)
# + tags=[]
sinergise = all_tiles[all_tiles._merge == 'left_only'].drop(columns=['element84', '_merge'])
element84 = all_tiles[all_tiles._merge == 'right_only'].drop(columns=['sinergise', '_merge'])
print(f"Found {len(sinergise)} scenes that are only in Sinergise and {len(element84)} that are only in Element84.")
# -
sinergise.to_csv('data/sinergise-only.txt', columns=['sinergise', 'tile_id'], index=False)
element84.to_csv('data/element84-only.txt', columns=['element84', 'tile_id'], index=False)
|
1-sinergise-element84-differences.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kW0dyx1VWf2Z"
# <a href="https://colab.research.google.com/drive/1n4O-rRmKNn7jDJO30kRRyVPDn6-HICkb?usp=sharing" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="XGKobxm4T4-b"
# <i>Mutual Funds Prediction</i><br>
# --
# Coded by : sammyon7<br>
# Dataset <a href="https://drive.google.com/drive/folders/1jjufCJRUcaPOkbjvG7Xl0kfkvuCsAV85?usp=sharing">link</a>
# + id="XDw_-lowTWSj"
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn import preprocessing
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier,GradientBoostingClassifier
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="IPLF8UPnTheD" outputId="aea5d439-c4f2-4dd4-e71b-3c3f64cb3a56"
fund_specs=pd.read_csv('Datasets/fund_specs.csv',thousands=',')
fund_specs.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="rApnDZKqTyVS" outputId="ed252fb0-a4cd-4229-a8db-5e3ae72687eb"
fund_specs.nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="HdKJZw4GUIlz" outputId="12cb998d-d283-4722-bd42-1f6ff6fe3069"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="CZyW67BGUJIa" outputId="42a87813-229b-49b1-94e3-259fa1235b46"
fund_specs.drop(labels=['currency'],axis=1,inplace=True)
fund_specs.info()
# + colab={"base_uri": "https://localhost:8080/"} id="znHvq7FIUjFq" outputId="fa48ef41-e574-447d-c421-873dd3dc7e32"
date_array=fund_specs['inception_date'].str.split('-')
print(date_array)
# + colab={"base_uri": "https://localhost:8080/"} id="lV_fk0Q2UmET" outputId="2d3ec043-f289-42dd-d9a2-c2413ea18476"
for i in range(0,fund_specs.inception_date.size):
fund_specs['inception_date'][i]=date_array[i][0]
# + id="Zly37krsUqGa"
fund_specs.inception_date=fund_specs.inception_date.astype('int32')
# + colab={"base_uri": "https://localhost:8080/"} id="o9jBunqGUtNh" outputId="ffc6e355-939c-4f52-8217-b2a139b9d0d9"
fund_specs.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 359} id="s-M-PztzUtwJ" outputId="4d7cdf59-7ec2-43d7-aaa8-fcb77d1cebd1"
fund_specs[['investment_class','fund_size']]=fund_specs[['investment_class','fund_size']].fillna(method='bfill')
fund_specs.investment_class.replace({'Growth':0,'Blend':1,'Value':2},inplace=True)
fund_specs.fund_size.replace({'Small':0,'Medium':1,'Large':2},inplace=True)
fund_specs.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="SJgOPi_hUxp5" outputId="4f5383eb-3512-4312-a9ac-46f47b3dd726"
fund_specs.nunique()
# + colab={"base_uri": "https://localhost:8080/"} id="B7eBAri2U8RB" outputId="824d5b0c-d94f-4c00-9bd3-25954b338d57"
fund_specs.total_assets.fillna(fund_specs.total_assets.median(),inplace=True)
fund_specs.corrwith(fund_specs.greatstone_rating)
# + id="GDh0v0wsVD9w"
fund_specs.return_ytd.fillna(fund_specs.return_ytd.median(),inplace=True)
fund_specs['yield'].fillna(fund_specs['yield'].median(),inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="USn_Omw2VI-p" outputId="499e2557-5aa8-4671-a9af-5b95e77cc480"
corr=fund_specs.corr()
fund_specs_corr=sns.heatmap(corr,annot=True, fmt='.2f')
b,t=fund_specs_corr.get_ylim()
fund_specs_corr.set_ylim(b+.5,t-.5)
plt.tight_layout()
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="ypjg6fJBVMuJ" outputId="83146241-2186-444e-88d9-1238748753ec"
corr
# + id="6MwIATKKVNu5"
fund_ratios=pd.read_csv('Datasets/fund_ratios.csv',thousands=',')
# + colab={"base_uri": "https://localhost:8080/", "height": 529} id="oXGR0xj_Vecx" outputId="e1b2d377-9e26-4350-f5e8-bb1fc54ed413"
fund_ratios.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="5eyoVNcrVhSp" outputId="b745c375-b7d8-4286-e10d-c4dc11ce08b8"
fund_ratios.describe().T
# + id="nNEIiL7xVjLh"
fund_ratios['pb_ratio'].fillna(fund_ratios['pb_ratio'].mean(),inplace=True)
fund_ratios['ps_ratio'].fillna(fund_ratios['ps_ratio'].mean(),inplace=True)
fund_ratios['mmc'].fillna(fund_ratios['mmc'].median(),inplace=True)
fund_ratios['pc_ratio'].fillna(fund_ratios['pc_ratio'].mean(),inplace=True)
fund_ratios['pe_ratio'].fillna(fund_ratios['pe_ratio'].mean(),inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="DdMN8Q6LVlXg" outputId="bf3e7c50-5294-4a18-c555-a58b352fcc53"
fund_ratios.info()
# + id="GDhA6_-6Vn-5"
bond_ratings=pd.read_csv('Datasets/bond_ratings.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="4w4-nPwBV2KQ" outputId="767b19c2-42e4-4d12-a18c-9298b75ed488"
bond_ratings.head(10)
# + id="V8S32dEyV_3B"
bond_ratings.maturity_bond.fillna(bond_ratings.maturity_bond.mean(),inplace=True)
bond_ratings.duration_bond.fillna(bond_ratings.duration_bond.mean(),inplace=True)
bond_ratings.drop(labels='us_govt_bond_rating',axis=1,inplace=True)
bond_ratings.fillna(0,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="YWtZnChdWEXo" outputId="262fb634-8b0d-4bfd-8756-320341f180b8"
bond_ratings
# + colab={"base_uri": "https://localhost:8080/"} id="hLgxzGNnWHHY" outputId="fbb3775a-993e-44ae-ece1-87f700eb1550"
print(bond_ratings.info())
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="XhEIQgk1WJQP" outputId="32a6035a-ccda-4990-ba9e-134a63aeb64a"
fund_allocations=pd.read_csv('Datasets/fund_allocations.csv')
fund_allocations.head(10)
# + id="sLKsqIe7W2RZ"
fund_allocations['portfolio_tech_allocation'].fillna(fund_allocations['portfolio_tech_allocation'].median(),inplace=True)
# + id="bbdpOt15W6kY"
fund_allocations['portfolio_utils_allocation'].fillna(fund_allocations['portfolio_utils_allocation'].median(),inplace=True)
# + id="XRiqRAhRXFpo"
fund_allocations.fillna(fund_allocations.mean(),inplace=True)
fund_allocations.rename(columns={'id':'tag'}, inplace=True)
# + id="GmDxaOs6XYbH"
other_specs=pd.read_csv('Datasets/other_specs.csv',thousands=',')
# + id="fsdaTWCYXe8P"
other_specs.drop(labels=['pc_ratio','pb_ratio','pe_ratio','mmc','ps_ratio'],axis=1,inplace=True)
other_specs[['cash_percent_of_portfolio',
'stock_percent_of_portfolio',
'portfolio_others',
'bond_percentage_of_porfolio',
'portfolio_preferred',
'portfolio_convertable']]=other_specs[['cash_percent_of_portfolio',
'stock_percent_of_portfolio',
'portfolio_others',
'bond_percentage_of_porfolio',
'portfolio_preferred',
'portfolio_convertable']].fillna(16.67)
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="W23ViZvjXjSf" outputId="7ed521cf-6732-4eb6-e442-b2cbf0b217d5"
other_specs.head(10)
# + id="xVKaCNVKXpjv"
nul_cat_tag_list=fund_specs[['tag','inception_date']]
# + id="1zeRmzHeXsMf"
# Define the empty lists
dateEqual2018=[]
dateEqual2017=[]
dateEqual2016=[]
dateEqual2015=[]
dateEqual2014=[]
dateEqual2013=[]
dateEqual2012=[]
dateEqual2011=[]
dateEqual2010=[]
# + id="IVrfqbsvXx9f"
for tagVal,date in nul_cat_tag_list.itertuples(index=False):
if date==2018:
dateEqual2018.append(tagVal)
elif date==2017:
dateEqual2017.append(tagVal)
elif date==2016:
dateEqual2016.append(tagVal)
elif date==2015:
dateEqual2015.append(tagVal)
elif date==2014:
dateEqual2014.append(tagVal)
elif date==2013:
dateEqual2013.append(tagVal)
elif date==2012:
dateEqual2012.append(tagVal)
elif date==2011:
dateEqual2011.append(tagVal)
elif date>2011:
dateEqual2010.append(tagVal)
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="NTEEMAlzXz1f" outputId="caadaada-c538-40df-e301-d606cc074e8e"
other_specs.head(10)
# + id="fZ9fdGQxX1Se"
for tagVal in dateEqual2011:
if other_specs[other_specs.tag==tagVal]['2010_return_category'].isna().values[0]:
other_specs['2010_return_fund']=other_specs['2010_return_fund'].fillna(0)
other_specs['2010_return_category']=other_specs['2010_return_category'].fillna(0)
for tagVal in dateEqual2012:
if other_specs[other_specs.tag==tagVal]['2011_return_category'].isna().values[0]:
other_specs['2011_return_fund']=other_specs['2011_return_fund'].fillna(0)
other_specs['2011_return_category']=other_specs['2011_return_category'].fillna(0)
for tagVal in dateEqual2013:
if other_specs[other_specs.tag==tagVal]['2012_return_category'].isna().values[0]:
other_specs['2012_fund_return']=other_specs['2012_fund_return'].fillna(0)
other_specs['2012_return_category']=other_specs['2012_return_category'].fillna(0)
for tagVal in dateEqual2014:
if other_specs[other_specs.tag==tagVal]['2013_category_return'].isna().values[0]:
other_specs['2013_return_fund']=other_specs['2013_return_fund'].fillna(0)
other_specs['2013_category_return']=other_specs['2013_category_return'].fillna(0)
for tagVal in dateEqual2015:
if other_specs[other_specs.tag==tagVal]['2014_category_return'].isna().values[0]:
other_specs['2014_return_fund']=other_specs['2014_return_fund'].fillna(0)
other_specs['2014_category_return']=other_specs['2014_category_return'].fillna(0)
for tagVal in dateEqual2016:
if other_specs[other_specs.tag==tagVal]['category_return_2015'].isna().values[0]:
other_specs['2015_return_fund']=other_specs['2015_return_fund'].fillna(0)
other_specs['category_return_2015']=other_specs['category_return_2015'].fillna(0)
for tagVal in dateEqual2017:
if other_specs[other_specs.tag==tagVal]['2016_return_category'].isna().values[0]:
other_specs['2016_return_fund']=other_specs['2016_return_fund'].fillna(0)
other_specs['2016_return_category']=other_specs['2016_return_category'].fillna(0)
for tagVal in dateEqual2018:
if other_specs[other_specs.tag==tagVal]['2017_category_return'].isna().values[0]:
other_specs['2017_return_fund']=other_specs['2017_return_fund'].fillna(0)
other_specs['2017_category_return']=other_specs['2017_category_return'].fillna(0)
# + colab={"base_uri": "https://localhost:8080/"} id="EnRZo438X9aO" outputId="44fced92-ea38-4d3f-a7b9-9c049b2209e5"
other_specs.info()
# + id="h0c4Lr8RYUU-"
other_specs=other_specs.drop(labels=['greatstone_rating'],axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="gtv6cZCLYWcf" outputId="3df4e30d-501e-4da1-cdae-9c3ecdd3184f"
other_specs.fillna(other_specs.mean(),inplace=True)
other_specs.nunique()
# + id="zcOTHwEIYZNL"
return_3year=pd.read_csv('Datasets/return_3year.csv',thousands=',')
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="Tx-xgbDUYfZW" outputId="fa80f144-ac38-4cd9-eb8d-6f5b129e8351"
return_3year.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="xPscrA-SYhK-" outputId="c3814639-1fdc-4534-efa8-6acd3d7e1064"
return_3year.fillna(return_3year.mean(),inplace=True)
return_3year.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 781} id="rA118cRRYqQ-" outputId="6fc048da-fd47-4b71-ea87-f0273858310d"
plt.figure(figsize=(20,15))
return_3year_corr=sns.heatmap(return_3year.corr(), annot=True)
b,t=return_3year_corr.get_ylim()
return_3year_corr.set_ylim(b+.5,t-.5)
plt.tight_layout()
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="fM64OA9fYtYW" outputId="b3c1c0a9-cb0d-4607-ae90-3e86ba70d6d8"
return_5year=pd.read_csv('Datasets/return_5year.csv',thousands=',')
return_5year.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="sgL6FHAcY4ql" outputId="7c10b517-bef0-46f8-9b41-00cf3a1a53bc"
return_5year.fillna(return_5year.mean(),inplace=True)
return_5year.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 781} id="3KfT62sKZDTm" outputId="3783335b-4b37-4151-8fb3-4fed3f7f508b"
plt.figure(figsize=(20,15))
return_5year_corr=sns.heatmap(return_5year.corr(), annot=True)
b,t=return_5year_corr.get_ylim()
return_5year_corr.set_ylim(b+.5,t-.5)
plt.tight_layout()
# + id="tYcRPNdxZFuO"
return_10year=pd.read_csv('Datasets/return_10year.csv',thousands=',')
# + colab={"base_uri": "https://localhost:8080/", "height": 889} id="OSrXq0ZrZL5W" outputId="b00ae248-1ce3-4074-faeb-6d622b91f7b1"
return_10year.head(10)
# + id="vdAlSVMNZNSf"
return_10year.fillna(0,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 781} id="wbuYO_vYZSz-" outputId="4a2959b5-d218-455f-c137-add8d8d92a6c"
plt.figure(figsize=(20,15))
return_10year_corr=sns.heatmap(return_10year.corr(), annot=True)
b,t=return_10year_corr.get_ylim()
return_10year_corr.set_ylim(b+.5,t-.5)
plt.tight_layout()
# + id="j7DKicqfZWA2"
# Now we will clean the data!
dataset=fund_specs.merge(fund_ratios)
dataset=dataset.merge(bond_ratings)
dataset=dataset.merge(fund_allocations)
dataset=dataset.merge(other_specs)
dataset=dataset.merge(return_3year)
dataset=dataset.merge(return_5year)
dataset=dataset.merge(return_10year)
# + id="zvkjDEGtZZ4e"
# Test and train!
train_data=dataset[dataset.greatstone_rating.notna()]
# + id="NGhOfyseZpi2"
train_target=train_data['greatstone_rating'].copy(deep=True)
train_data=train_data.drop(labels=['fund_id','tag','greatstone_rating'],axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="QYT7hHOhZvPd" outputId="95f37c03-9339-416a-c5d2-2b09d620391e"
rfc=RandomForestClassifier(n_estimators=500,random_state=11,criterion='entropy',class_weight='balanced')
rfc.fit(train_data,train_target)
# + colab={"base_uri": "https://localhost:8080/"} id="bQhDehNgZxMF" outputId="dc233b78-d836-4ddc-87e1-f5f10e1cb527"
test_data=dataset[dataset.greatstone_rating.isna()]
# + colab={"base_uri": "https://localhost:8080/", "height": 949} id="f7wKYINdaPNe" outputId="4f231f6c-ef88-4b1d-804d-5f5dda357ccc"
test_data
# + id="xXRKGQk8aXIF"
test_fund_id=test_data.fund_id.copy()
test_data=test_data.drop(labels=['fund_id','tag','greatstone_rating'],axis=1)
# + id="Ipw745yGacV0"
output=rfc.predict(test_data)
# + colab={"base_uri": "https://localhost:8080/"} id="dqzmABEVae09" outputId="bcc9830b-27cb-4280-e13c-4ba911dab61e"
output
# + id="8dr7Rkriafst"
result=pd.DataFrame()
result['fund_id']=test_fund_id
result['greatstone_rating']=output
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="dhdiMzB6amLV" outputId="817cc77e-b6eb-461c-9913-b5e3724c7d4d"
result
# + colab={"base_uri": "https://localhost:8080/"} id="y7NVD3n4am0t" outputId="a17a2ed6-e94a-4bd4-83ce-e5d718f61372"
result.greatstone_rating.value_counts()
# + id="Ecc_w2ggap1N"
result.to_csv("result.csv",index=False)
|
Py_NUB/MutualFundsPrediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extending Auto-Sklearn with LGB Component
# +
from ConfigSpace.configuration_space import ConfigurationSpace
from ConfigSpace.hyperparameters import UniformFloatHyperparameter, \
UniformIntegerHyperparameter, CategoricalHyperparameter
import sklearn.metrics
import autosklearn.regression
import autosklearn.pipeline.components.regression
from autosklearn.pipeline.components.base import AutoSklearnRegressionAlgorithm
from autosklearn.pipeline.constants import SPARSE, DENSE, \
SIGNED_DATA, UNSIGNED_DATA, PREDICTIONS
from sklearn.datasets import load_diabetes
from sklearn.model_selection import train_test_split
import xgboost as xgb
print("xgb", xgb.__version__)
print("autosklearn", autosklearn.__version__)
# -
# ## Generate data
X, y = load_diabetes(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y)
# ## Test LGB Regressor
# +
# lgb_model = lgb_model = lgb.LGBMRegressor(
# boosting_type='gbdt', class_weight=None, colsample_bytree=1.0,
# importance_type='split', learning_rate=0.1, max_depth=-1,
# min_child_samples=20, min_child_weight=0.001, min_split_gain=0.0,
# n_estimators=100, n_jobs=-1, num_leaves=31, objective=None,
# random_state=None, reg_alpha=0.0, reg_lambda=0.0, silent=True,
# subsample=1.0, subsample_for_bin=200000, subsample_freq=0
# )
# lgb_model.fit(X_train, y_train, verbose=1)
# lgb_model.predict(X_test)
# -
# ## Implement LGBcomponent
# +
class LGBReg(AutoSklearnRegressionAlgorithm):
def __init__(self, boosting_type, num_leaves, max_depth, learning_rate, n_estimators, reg_alpha, reg_lambda
,random_state=None):
self.boosting_type=boosting_type
self.num_leaves=num_leaves
self.max_depth=max_depth
self.learning_rate=learning_rate
self.n_estimators=n_estimators
self.reg_alpha=reg_alpha
self.reg_lambda=reg_lambda
self.random_state = random_state
self.estimator = None
def fit(self, X, y):
import lightgbm as lgb
self.estimator = lgb.LGBMRegressor(boosting_type=self.boosting_type
,num_leaves=self.num_leaves
,max_depth=self.max_depth
,learning_rate=self.learning_rate
,n_estimators=self.n_estimators
,reg_alpha=self.reg_alpha
,reg_lambda=self.reg_lambda
)
self.estimator.fit(X, y)
return self
def predict(self, X):
if self.estimator is None:
raise NotImplementedError
return self.estimator.predict(X)
@staticmethod
def get_properties(dataset_properties=None):
return {'shortname': 'LGBReg',
'name': 'LGB Regression',
'handles_regression': True,
'handles_classification': False,
'handles_multiclass': False,
'handles_multilabel': False,
'handles_multioutput': True,
'is_deterministic': True,
'input': (SPARSE, DENSE, UNSIGNED_DATA, SIGNED_DATA),
'output': (PREDICTIONS,)}
@staticmethod
def get_hyperparameter_search_space(dataset_properties=None):
cs = ConfigurationSpace()
boosting_type = CategoricalHyperparameter(
name='boosting_type',choices=['gbdt','dart','goss','rf'],default_value='gbdt')
num_leaves=UniformIntegerHyperparameter(
name='num_leaves', lower=1, upper=1000, default_value=31)
max_depth=UniformIntegerHyperparameter(
name='max_depth', lower=-1, upper=1000, default_value=-1)
learning_rate = UniformFloatHyperparameter(
name='learning_rate', lower=0.000000001, upper=1,default_value=0.1) #log=True
n_estimators=UniformIntegerHyperparameter(
name='n_estimators', lower=1, upper=2000, default_value=100)
reg_alpha = UniformFloatHyperparameter(
name='reg_alpha', lower=0.0000000, upper=1, default_value=0.0) #log=True
reg_lambda = UniformFloatHyperparameter(
name='reg_lambda', lower=0.0000000, upper=1, default_value=1) #log=True
cs.add_hyperparameters([boosting_type,num_leaves, max_depth, learning_rate,n_estimators, reg_alpha,reg_lambda])
return cs
# Add XGB component to auto-sklearn.
autosklearn.pipeline.components.regression.add_regressor(LGBReg)
cs = LGBReg.get_hyperparameter_search_space()
print(cs)
# -
#
#
#
# ## Build and Fit the model using the created LGB component
automl = autosklearn.regression.AutoSklearnRegressor(
time_left_for_this_task=30
,per_run_time_limit=10
,ensemble_size=1
,include_estimators=['LGBReg']
,initial_configurations_via_metalearning=0,
)
automl.fit(X_train, y_train)
# ## Print prediction
#
#
y_pred = automl.predict(X_test)
y_pred
# ## Print search results
print(automl.sprint_statistics())
# ## Print model parameters
print("r2 score: ", sklearn.metrics.r2_score(y_pred, y_test))
print(automl.show_models())
|
autosklearn/pipeline/components/regression/extending_regression_LGB .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import h5py
from matplotlib.colors import LinearSegmentedColormap
from matplotlib.patches import Rectangle
import netCDF4 as nc
from cartopy import config
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import shapefile as shp
from datetime import date, timedelta
from descartes import PolygonPatch
with h5py.File("data/mcmc_save_data.hdf5", "r") as fp:
samples = fp['samples'][:, :]
objectives = fp['objectives'][:]
plt.plot(objectives)
true_particle_lon = -88.365997
true_particle_lat = 28.736628
# Starting particle release location
particle_lon = -87.8
particle_lat = 29.2
# +
cmap_nodes = [0.0, 1.0]
# cmap_colors = [(232 / 255, 117 / 255, 0, 0.0), (232 / 255, 117 / 255, 0, 1.0)]
cmap_colors = [(1.00, 0.325, 0.286, 0.0), (1.00, 0.325, 0.286, 1.0)]
my_cmap = LinearSegmentedColormap.from_list("mycmap", cmap_colors)
# +
fig, ax = plt.subplots(1, 1, figsize=(10, 7))
im = ax.hexbin(samples[500:, 1], samples[500:, 0], zorder=0, gridsize=30, cmap=my_cmap, edgecolor=None);
ax.scatter(true_particle_lon, true_particle_lat, color='red', marker='x', zorder=10, label='True Location')
ax.scatter(particle_lon, particle_lat, color='red', zorder=10, label='Starting Location')
ax.set_aspect('equal')
ax.legend()
ax.set_xlabel('Degrees Longitude')
ax.set_ylabel('Degrees Latitude')
# fig.colorbar(im);
fig.savefig('source_probability.png', dpi=300)
# +
with nc.Dataset('data/hycom_gomu_501_2010042000_t000.nc') as ds:
shape = (ds['lat'].shape[0], ds['lon'].shape[0])
lon = np.array(ds['lon'][:])
lat = np.array(ds['lat'][:])
lon[1] - lon[0]
# +
shp_start_date = date(2010, 6, 13)
shapes = []
for i in range(1):
current_date = shp_start_date + timedelta(days=i)
with shp.Reader("data/shapefiles/S_{}_{}.shp".format(current_date.month, current_date.day)) as sf:
shapes.append({
'date': current_date,
'shape_records': sf.shapeRecords()
})
# +
fig = plt.figure(figsize=(15, 10))
ax = fig.add_axes([0.1, 0.1, 0.7, 0.85], projection=ccrs.PlateCarree())
im = ax.hexbin(samples[400:, 1], samples[400:, 0], zorder=0, gridsize=20, cmap=my_cmap, edgecolor=None);
ax.scatter(true_particle_lon, true_particle_lat, color='orange', marker='x', zorder=10, label='True Location')
ax.scatter(particle_lon, particle_lat, color='orange', ec='white', zorder=10, label='Starting Location')
ax.set_aspect('equal')
ax.set_xlabel('Degrees Longitude')
ax.set_ylabel('Degrees Latitude')
ax.coastlines(resolution='10m', color='black', linewidth=1)
ax.add_feature(cfeature.OCEAN)
ax.add_feature(cfeature.LAND)
ax.add_feature(cfeature.LAKES)
ax.add_feature(cfeature.RIVERS)
ax.set_xlim(-90.5, -86.8)
ax.set_ylim(28, 30.5)
ax.scatter(-90.0715, 29.9511, color='red')
ax.text(-90.07, 30, 'New Orleans, LA', color='red')
for i, shape in enumerate(shapes[0]['shape_records']):
poly = shape.__geo_interface__['geometry']
if i == 0:
ax.add_patch(PolygonPatch(poly, ec='black', fc='gray', alpha=0.25, label='Deepwater Horizon Oil Spill'))
else:
ax.add_patch(PolygonPatch(poly, ec='black', fc='gray', alpha=0.25))
# fig.colorbar(im);
ax2 = fig.add_axes([0.55, 0.05, 0.45, 0.45], projection=ccrs.PlateCarree())
ax2.hexbin(samples[500:, 1], samples[500:, 0], zorder=0, gridsize=30, cmap=my_cmap, edgecolor=None);
ax2.scatter(true_particle_lon, true_particle_lat, color='white', marker='x', zorder=10, label='True Location')
# ax2.add_feature(cfeature.OCEAN)
xlim = ax2.get_xlim()
ylim = ax2.get_ylim()
ax.add_patch(Rectangle((xlim[0], ylim[0]), xlim[1] - xlim[0], ylim[1] - ylim[0], fc=(0,0,0,0), ec='black', label='Enlarged Region'))
ax.legend()
# ax2.add_feature(cfeature.OCEAN)
fig.savefig('source_probability_without_burnin_with_map_sw.png', dpi=300)
# -
|
examples/HYCOM_deepwater_horizon/HYCOM_inversion_pictures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Traffic Sign Recognition**
#
# ## Writeup
#
# ### You can use this file as a template for your writeup if you want to submit it as a markdown file, but feel free to use some other method and submit a pdf if you prefer.
#
# ---
#
#
# The goals / steps of this project are the following:
# * Load the data set (see below for links to the project data set)
# * Explore, summarize and visualize the data set
# * Design, train and test a model architecture
# * Use the model to make predictions on new images
# * Analyze the softmax probabilities of the new images
# * Summarize the results with a written report
#
#
# [//]: # (Image References)
#
# [image1]: ./visual/Train_Dataset.png "Train Data Set"
# [image2]: ./visual/Valid_Dataset.png "Validation Data Set"
# [image3]: ./visual/Test_Dataset.png "Test Data Set"
# [image4]: ./visual/signs_gray_color.jpg "Colored and gray signs"
# [image5]: ./visual/jittered.jpg "Jittered Data sets"
# [image6]: ./visual/test1.jpg "German Sign 1"
# [image7]: ./visual/test2.jpg "German Sign 2"
# [image8]: ./visual/test3.jpg "German Sign 3"
# [image9]: ./visual/test4.jpg "German Sign 4"
# [image10]: ./visual/test5.jpg "German Sign 5"
#
#
#
# ## Rubric Points
# ### Here I will consider the [rubric points](https://review.udacity.com/#!/rubrics/481/view) individually and describe how I addressed each point in my implementation.
#
# ---
# ### Writeup / README
#
# #### 1. Provide a Writeup / README that includes all the rubric points and how you addressed each one. You can submit your writeup as markdown or pdf. You can use this template as a guide for writing the report. The submission includes the project code.
#
# You're reading it! and here is a link to my [project code](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/Traffic_Sign_Classifier.ipynb)
#
# ### Data Set Summary & Exploration
#
# #### 1. Provide a basic summary of the data set. In the code, the analysis should be done using python, numpy and/or pandas methods rather than hardcoding results manually.
#
# I used the pandas library to calculate summary statistics of the traffic
# signs data set:
#
# * The size of training set is 34799
# * The size of the validation set is 4410
# * The size of test set is 12630
# * The shape of a traffic sign image is 32x32x3 pixels. RGB color space.
# * The number of unique classes/labels in the data set is 43
#
# #### 2. Include an exploratory visualization of the dataset.
# #### Statistics about the data sets
# Following figures show the amount of different classes (traffic signs) and how many samples each class contains.
#
# ![alt text][image1] ![alt text][image2]
# ![alt text][image3]
#
# As one can see, not all classes are represented eaqually. It may happen, that classes will a small number of examples are trained less good.
#
# ### Design and Test a Model Architecture
#
# #### 1. Describe how you preprocessed the image data. What techniques were chosen and why did you choose these techniques? Consider including images showing the output of each preprocessing technique. Pre-processing refers to techniques such as converting to grayscale, normalization, etc. (OPTIONAL: As described in the "Stand Out Suggestions" part of the rubric, if you generated additional data for training, describe why you decided to generate additional data, how you generated the data, and provide example images of the additional data. Then describe the characteristics of the augmented training set like number of images in the set, number of images for each class, etc.)
#
# As a first step, I decided to convert the images to grayscale because it decreases the amount of input channels respectivly data without loosing too much information. The recognition of traffic signs doesn't depend on the color information. Each sign is recognisable by only its shape and imprinted graphics..
#
# Here are random examples of a traffic sign images before and after grayscaling.
#
# ![alt text][image4]
#
# Additionally, I normalized the image data because data should have mean zero and equal variance for better performance.
#
# #### Generating jittered data
# As suggested in LeCun's paper "Traffic Sign Recognition with Multi-Scale Convolutional Networks" one way to increase validation accuracy of a CNN is to generate more training data. As shown above, some sign classes are underrepresented. By adding jittered images (Applying warp function and perspective transformation) the training set is increased significantly from 34799 images to 89860 images. The validation set is increased from 4410 images to 22465 images.
#
# ![alt text][image5]
#
# #### 2. Describe what your final model architecture looks like including model type, layers, layer sizes, connectivity, etc.) Consider including a diagram and/or table describing the final model.
#
# My final model consisted of the following layers:
#
# | Layer | Description |
# |:---------------------:|:---------------------------------------------:|
# | Input | 32x32x1 Gray scale image |
# | Convolution 5x5 | 1x1 stride, valid padding, outputs 28x28x6 |
# | RELU | |
# | Max pooling | 2x2 stride, outputs 14x14x6 |
# | Convolution 5x5 | 1x1 stride, valid padding, outputs 10x10x16 |
# | RELU | |
# | Max pooling | 2x2 stride, outputs 5x5x16 |
# | Convolution 5x5 | 1x1 stride, valid padding, outputs 1x1x400 |
# | Fully connected | input 400, output 200 |
# | RELU | |
# | Droput | 50 % |
# | Fully connected | input 200, output 84 |
# | RELU | |
# | Droput | 50 % |
# | Fully connected | input 84, output 43 |
#
# #### 3. Describe how you trained your model. The discussion can include the type of optimizer, the batch size, number of epochs and any hyperparameters such as learning rate.
#
# To train the model, I used the ADAM Optimizer. Batchs size was set to 150 and 20 epochs. The learning rate was set to 0.0009.
#
# #### 4. Describe the approach taken for finding a solution and getting the validation set accuracy to be at least 0.93. Include in the discussion the results on the training, validation and test sets and where in the code these were calculated. Your approach may have been an iterative process, in which case, outline the steps you took to get to the final solution and why you chose those steps. Perhaps your solution involved an already well known implementation or architecture. In this case, discuss why you think the architecture is suitable for the current problem.
#
# My final model results were:
# * Training set accuracy of 99.9%
# * Validation set accuracy of 99.6%
# * Test set accuracy of 94%
#
# An iterative approach was chosen:
# At first I implemented the LeNet architecture which was shown in the LeNet Lab. I used the RBG images as data sets for training. With this approach the model realized ~85% validation accuracy.
#
# To improve it, RGB to Gray function and the normalisation step were added. With this improvements the accuracy increased to about 92%. I also adjusted the drop out hyperparameter and tried the softmax function instead of Relu function for the activation step. This didn't have a big impact on the result.
# Adding one more convolutional layer to the LeNet architecture and raising the epochs to 20 was increasing the validation accuracy up to 99%.
#
# ### Test a Model on New Images
#
# #### 1. Choose five German traffic signs found on the web and provide them in the report. For each image, discuss what quality or qualities might be difficult to classify.
#
# Following 5 german traffic signs were choosen to test real data:
#
# ![alt text][image6] ![alt text][image7] ![alt text][image8]
# ![alt text][image9] ![alt text][image10]
#
# The second image (Priority Road) may yield a bad prediction since it is very dark foreground with low contrast and a bright background. This could lead to misclassification. The third image (No entry) is quite blur which could yield to wrong predictions. The 4th image (Right-of-way at next intersection) could be difficult to classify because there is half of another sign shown under the actual traffic sign.
#
# #### 2. Discuss the model's predictions on these new traffic signs and compare the results to predicting on the test set. At a minimum, discuss what the predictions were, the accuracy on these new predictions, and compare the accuracy to the accuracy on the test set (OPTIONAL: Discuss the results in more detail as described in the "Stand Out Suggestions" part of the rubric).
#
# Here are the results of the prediction:
#
# | Image | Prediction |
# |:---------------------:|:---------------------------------------------:|
# | 30 km/h | No Passing |
# | Priority Road | Priority Road |
# | No entry | No entry |
# | Right-of-way at next intersection| Right-of-way at next intersection |
# | Stop | Stop |
#
#
# The model was able to correctly guess 5 of the 5 traffic signs, which gives an accuracy of 100%. This fits to the results of the Validation set (99.6%) and Test set accuracy (99.9%).
#
# #### 3. Describe how certain the model is when predicting on each of the five new images by looking at the softmax probabilities for each prediction. Provide the top 5 softmax probabilities for each image along with the sign type of each probability. (OPTIONAL: as described in the "Stand Out Suggestions" part of the rubric, visualizations can also be provided such as bar charts)
#
# The code for making predictions on my final model is located in the 14th cell of the Ipython notebook.
#
# The top five soft max probabilities of all 5 images are 100 %, 0% ,0% ,0% ,0%. The model is very sure about it's predictions.
#
|
writeup.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
#import matplotlib.pyplot as plt
from matplotlib import pyplot, pylab
# +
data1 = pd.read_csv('progress10-128.csv')
data2 = pd.read_csv('progress10k-128.csv')
data3 = pd.read_csv('progress10k-512.csv')
data4 = pd.read_csv('progress10k-1024.csv')
data5 = pd.read_csv('progress100-128.csv')
data4.head()
# +
pyplot.figure(figsize=(15,5))
for data in [data1, data2, data3, data4, data5]:
pyplot.plot(data['epoch'], data['test/mean_Q'], label='sample')
pylab.xlabel('Epoch')
pylab.ylabel('Mean Q value')
#pyplot.fill_between(color=color)
#pyplot.xlim(0,6)
pyplot.title('Training results comparison for Fetch-PickAndPlace')
L=pyplot.legend()
L.get_texts()[0].set_text('10-128')
L.get_texts()[1].set_text('10k-128')
L.get_texts()[2].set_text('10k-512')
L.get_texts()[3].set_text('10k-1024')
L.get_texts()[4].set_text('100-128')
# L.get_texts()[4].set_text('6 Layers')
# pyplot.savefig('FetchPickPlaceNeuronMean.jpg')
# +
pyplot.figure(figsize=(15,5))
for data in [data1, data2, data3, data4, data5]:
pyplot.plot(data['epoch'], data['test/success_rate'], label='sample')
pylab.xlabel('Epoch')
pylab.ylabel('Success Rate')
#pyplot.fill_between(color=color)
#pyplot.xlim(0,6)
pyplot.title('Training results comparison for Fetch-PickAndPlace')
L=pyplot.legend()
L.get_texts()[0].set_text('10-128')
L.get_texts()[1].set_text('10k-128')
L.get_texts()[2].set_text('10k-512')
L.get_texts()[3].set_text('10k-1024')
L.get_texts()[4].set_text('100-128')
# -
data1.mean()
data2.mean()
data3.mean()
data4.mean()
data5.mean()
|
Results/Jupyter/LinePlot.ipynb
|
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 10
# > NLP
#
# - toc: true
# - badges: true
# - comments: true
#
#hide
from utils import *
from IPython.display import display,HTML
# + active=""
# [[chapter_nlp]]
# -
# # NLP Deep Dive: RNNs
# In <<chapter_intro>> we saw that deep learning can be used to get great results with natural language datasets. Our example relied on using a pretrained language model and fine-tuning it to classify reviews. That example highlighted a difference between transfer learning in NLP and computer vision: in general in NLP the pretrained model is trained on a different task.
#
# What we call a language model is a model that has been trained to guess what the next word in a text is (having read the ones before). This kind of task is called *self-supervised learning*: we do not need to give labels to our model, just feed it lots and lots of texts. It has a process to automatically get labels from the data, and this task isn't trivial: to properly guess the next word in a sentence, the model will have to develop an understanding of the English (or other) language. Self-supervised learning can also be used in other domains; for instance, see ["Self-Supervised Learning and Computer Vision"](https://www.fast.ai/2020/01/13/self_supervised/) for an introduction to vision applications. Self-supervised learning is not usually used for the model that is trained directly, but instead is used for pretraining a model used for transfer learning.
# > jargon: Self-supervised learning: Training a model using labels that are embedded in the independent variable, rather than requiring external labels. For instance, training a model to predict the next word in a text.
# The language model we used in <<chapter_intro>> to classify IMDb reviews was pretrained on Wikipedia. We got great results by directly fine-tuning this language model to a movie review classifier, but with one extra step, we can do even better. The Wikipedia English is slightly different from the IMDb English, so instead of jumping directly to the classifier, we could fine-tune our pretrained language model to the IMDb corpus and then use *that* as the base for our classifier.
#
# Even if our language model knows the basics of the language we are using in the task (e.g., our pretrained model is in English), it helps to get used to the style of the corpus we are targeting. It may be more informal language, or more technical, with new words to learn or different ways of composing sentences. In the case of the IMDb datset, there will be lots of names of movie directors and actors, and often a less formal style of language than that seen in Wikipedia.
#
# We already saw that with fastai, we can download a pretrained English language model and use it to get state-of-the-art results for NLP classification. (We expect pretrained models in many more languages to be available soon—they might well be available by the time you are reading this book, in fact.) So, why are we learning how to train a language model in detail?
#
# One reason, of course, is that it is helpful to understand the foundations of the models that you are using. But there is another very practical reason, which is that you get even better results if you fine-tune the (sequence-based) language model prior to fine-tuning the classification model. For instance, for the IMDb sentiment analysis task, the dataset includes 50,000 additional movie reviews that do not have any positive or negative labels attached. Since there are 25,000 labeled reviews in the training set and 25,000 in the validation set, that makes 100,000 movie reviews altogether. We can use all of these reviews to fine-tune the pretrained language model, which was trained only on Wikipedia articles; this will result in a language model that is particularly good at predicting the next word of a movie review.
#
# This is known as the Universal Language Model Fine-tuning (ULMFit) approqch. The [paper](https://arxiv.org/abs/1801.06146) showed that this extra stage of fine-tuning of the language model, prior to transfer learning to a classification task, resulted in significantly better predictions. Using this approach, we have three stages for transfer learning in NLP, as summarized in <<ulmfit_process>>.
# <img alt="Diagram of the ULMFiT process" width="700" caption="The ULMFiT process" id="ulmfit_process" src="images/att_00027.png">
# We'll now explore how to apply a neural network to this language modeling problem, using the concepts introduced in the last two chapters. But before reading further, pause and think about how *you* would approach this.
# ## Text Preprocessing
# It's not at all obvious how we're going to use what we've learned so far to build a language model. Sentences can be different lengths, and documents can be very long. So, how can we predict the next word of a sentence using a neural network? Let's find out!
#
# We've already seen how categorical variables can be used as independent variables for a neural network. The approach we took for a single categorical variable was to:
#
# 1. Make a list of all possible levels of that categorical variable (we'll call this list the *vocab*).
# 1. Replace each level with its index in the vocab.
# 1. Create an embedding matrix for this containing a row for each level (i.e., for each item of the vocab).
# 1. Use this embedding matrix as the first layer of a neural network. (A dedicated embedding matrix can take as inputs the raw vocab indexes created in step 2; this is equivalent to but faster and more efficient than a matrix that takes as input one-hot-encoded vectors representing the indexes.)
#
# We can do nearly the same thing with text! What is new is the idea of a sequence. First we concatenate all of the documents in our dataset into one big long string and split it into words, giving us a very long list of words (or "tokens"). Our independent variable will be the sequence of words starting with the first word in our very long list and ending with the second to last, and our dependent variable will be the sequence of words starting with the second word and ending with the last word.
#
# Our vocab will consist of a mix of common words that are already in the vocabulary of our pretrained model and new words specific to our corpus (cinematographic terms or actors names, for instance). Our embedding matrix will be built accordingly: for words that are in the vocabulary of our pretrained model, we will take the corresponding row in the embedding matrix of the pretrained model; but for new words we won't have anything, so we will just initialize the corresponding row with a random vector.
# Each of the steps necessary to create a language model has jargon associated with it from the world of natural language processing, and fastai and PyTorch classes available to help. The steps are:
#
# - Tokenization:: Convert the text into a list of words (or characters, or substrings, depending on the granularity of your model)
# - Numericalization:: Make a list of all of the unique words that appear (the vocab), and convert each word into a number, by looking up its index in the vocab
# - Language model data loader creation:: fastai provides an `LMDataLoader` class which automatically handles creating a dependent variable that is offset from the independent variable by one token. It also handles some important details, such as how to shuffle the training data in such a way that the dependent and independent variables maintain their structure as required
# - Language model creation:: We need a special kind of model that does something we haven't seen before: handles input lists which could be arbitrarily big or small. There are a number of ways to do this; in this chapter we will be using a *recurrent neural network* (RNN). We will get to the details of these RNNs in the <<chapter_nlp_dive>>, but for now, you can think of it as just another deep neural network.
#
# Let's take a look at how each step works in detail.
# ### Tokenization
# When we said "convert the text into a list of words," we left out a lot of details. For instance, what do we do with punctuation? How do we deal with a word like "don't"? Is it one word, or two? What about long medical or chemical words? Should they be split into their separate pieces of meaning? How about hyphenated words? What about languages like German and Polish where we can create really long words from many, many pieces? What about languages like Japanese and Chinese that don't use bases at all, and don't really have a well-defined idea of *word*?
#
# Because there is no one correct answer to these questions, there is no one approach to tokenization. There are three main approaches:
#
# - Word-based:: Split a sentence on spaces, as well as applying language-specific rules to try to separate parts of meaning even when there are no spaces (such as turning "don't" into "do n't"). Generally, punctuation marks are also split into separate tokens.
# - Subword based:: Split words into smaller parts, based on the most commonly occurring substrings. For instance, "occasion" might be tokenized as "o c ca sion."
# - Character-based:: Split a sentence into its individual characters.
#
# We'll be looking at word and subword tokenization here, and we'll leave character-based tokenization for you to implement in the questionnaire at the end of this chapter.
# > jargon: token: One element of a list created by the tokenization process. It could be a word, part of a word (a _subword_), or a single character.
# ### Word Tokenization with fastai
# Rather than providing its own tokenizers, fastai instead provides a consistent interface to a range of tokenizers in external libraries. Tokenization is an active field of research, and new and improved tokenizers are coming out all the time, so the defaults that fastai uses change too. However, the API and options shouldn't change too much, since fastai tries to maintain a consistent API even as the underlying technology changes.
#
# Let's try it out with the IMDb dataset that we used in <<chapter_intro>>:
from fastai2.text.all import *
path = untar_data(URLs.IMDB)
# We'll need to grab the text files in order to try out a tokenizer. Just like `get_image_files`, which we've used many times already, gets all the image files in a path, `get_text_files` gets all the text files in a path. We can also optionally pass `folders` to restrict the search to a particular list of subfolders:
files = get_text_files(path, folders = ['train', 'test', 'unsup'])
# Here's a review that we'll tokenize (we'll just print the start of it here to save space):
txt = files[0].open().read(); txt[:75]
# As we write this book, the default English word tokenizer for fastai uses a library called *spaCy*. It has a sophisticated rules engine with special rules for URLs, individual special English words, and much more. Rather than directly using `SpacyTokenizer`, however, we'll use `WordTokenizer`, since that will always point to fastai's current default word tokenizer (which may not necessarily be spaCy, depending when you're reading this).
#
# Let's try it out. We'll use fastai's `coll_repr(collection, n)` function to display the results. This displays the first *`n`* items of *`collection`*, along with the full size—it's what `L` uses by default. Note that fastai's tokenizers take a collection of documents to tokenize, so we have to wrap `txt` in a list:
spacy = WordTokenizer()
toks = first(spacy([txt]))
print(coll_repr(toks, 30))
# As you see, spaCy has mainly just separated out the words and punctuation. But it does something else here too: it has split "it's" into "it" and "'s". That makes intuitive sense; these are separate words, really. Tokenization is a surprisingly subtle task, when you think about all the little details that have to be handled. Fortunately, spaCy handles these pretty well for us—for instance, here we see that "." is separated when it terminates a sentence, but not in an acronym or number:
first(spacy(['The U.S. dollar $1 is $1.00.']))
# fastai then adds some additional functionality to the tokenization process with the `Tokenizer` class:
tkn = Tokenizer(spacy)
print(coll_repr(tkn(txt), 31))
# Notice that there are now some tokens that start with the characters "xx", which is not a common word prefix in English. These are *special tokens*.
#
# For example, the first item in the list, `xxbos`, is a special token that indicates the start of a new text ("BOS" is a standard NLP acronym that means "beginning of stream"). By recognizing this start token, the model will be able to learn it needs to "forget" what was said previously and focus on upcoming words.
#
# These special tokens don't come from spaCy directly. They are there because fastai adds them by default, by applying a number of rules when processing text. These rules are designed to make it easier for a model to recognize the important parts of a sentence. In a sense, we are translating the original English language sequence into a simplified tokenized language—a language that is designed to be easy for a model to learn.
#
# For instance, the rules will replace a sequence of four exclamation points with a single exclamation point, followed by a special *repeated character* token, and then the number four. In this way, the model's embedding matrix can encode information about general concepts such as repeated punctuation rather than requiring a separate token for every number of repetitions of every punctuation mark. Similarly, a capitalized word will be replaced with a special capitalization token, followed by the lowercase version of the word. This way, the embedding matrix only needs the lowercase versions of the words, saving compute and memory resources, but can still learn the concept of capitalization.
#
# Here are some of the main special tokens you'll see:
#
# - `xxbos`:: Indicates the beginning of a text (here, a review)
# - `xxmaj`:: Indicates the next word begins with a capital (since we lowercased everything)
# - `xxunk`:: Indicates the next word is unknown
#
# To see the rules that were used, you can check the default rules:
defaults.text_proc_rules
# As always, you can look at the source code of each of them in a notebook by typing:
#
# ```
# ??replace_rep
# ```
#
# Here is a brief summary of what each does:
#
# - `fix_html`:: Replaces special HTML characters with a readable version (IMDb reviews have quite a few of these)
# - `replace_rep`:: Replaces any character repeated three times or more with a special token for repetition (`xxrep`), the number of times it's repeated, then the character
# - `replace_wrep`:: Replaces any word repeated three times or more with a special token for word repetition (`xxwrep`), the number of times it's repeated, then the word
# - `spec_add_spaces`:: Adds spaces around / and #
# - `rm_useless_spaces`:: Removes all repetitions of the space character
# - `replace_all_caps`:: Lowercases a word written in all caps and adds a special token for all caps (`xxcap`) in front of it
# - `replace_maj`:: Lowercases a capitalized word and adds a special token for capitalized (`xxmaj`) in front of it
# - `lowercase`:: Lowercases all text and adds a special token at the beginning (`xxbos`) and/or the end (`xxeos`)
# Let's take a look at a few of them in action:
coll_repr(tkn('© Fast.ai www.fast.ai/INDEX'), 31)
# Now let's take a look at how subword tokenization would work.
# ### Subword Tokenization
# In addition to the *word tokenization* approach seen in the last section, another popular tokenization method is *subword tokenization*. Word tokenization relies on an assumption that spaces provide a useful separation of components of meaning in a sentence. However, this assumption is not always appropriate. For instance, consider this sentence: 我的名字是郝杰瑞 ("My name is <NAME>" in Chinese). That's not going to work very well with a word tokenizer, because there are no spaces in it! Languages like Chinese and Japanese don't use spaces, and in fact they don't even have a well-defined concept of a "word." There are also languages, like Turkish and Hungarian, that can add many subwords together without spaces, creating very long words that include a lot of separate pieces of information.
#
# To handle these cases, it's generally best to use subword tokenization. This proceeds in two steps:
#
# 1. Analyze a corpus of documents to find the most commonly occurring groups of letters. These become the vocab.
# 2. Tokenize the corpus using this vocab of *subword units*.
#
# Let's look at an example. For our corpus, we'll use the first 2,000 movie reviews:
txts = L(o.open().read() for o in files[:2000])
# We instantiate our tokenizer, passing in the size of the vocab we want to create, and then we need to "train" it. That is, we need to have it read our documents and find the common sequences of characters to create the vocab. This is done with `setup`. As we'll see shortly, `setup` is a special fastai method that is called automatically in our usual data processing pipelines. Since we're doing everything manually at the moment, however, we have to call it ourselves. Here's a function that does these steps for a given vocab size, and shows an example output:
def subword(sz):
sp = SubwordTokenizer(vocab_sz=sz)
sp.setup(txts)
return ' '.join(first(sp([txt]))[:40])
# Let's try it out:
subword(1000)
# When using fastai's subword tokenizer, the special character `▁` represents a space character in the original text.
#
# If we use a smaller vocab, then each token will represent fewer characters, and it will take more tokens to represent a sentence:
subword(200)
# On the other hand, if we use a larger vocab, then most common English words will end up in the vocab themselves, and we will not need as many to represent a sentence:
subword(10000)
# Picking a subword vocab size represents a compromise: a larger vocab means fewer tokens per sentence, which means faster training, less memory, and less state for the model to remember; but on the downside, it means larger embedding matrices, which require more data to learn.
#
# Overall, subword tokenization provides a way to easily scale between character tokenization (i.e., using a small subword vocab) and word tokenization (i.e., using a large subword vocab), and handles every human language without needing language-specific algorithms to be developed. It can even handle other "languages" such as genomic sequences or MIDI music notation! For this reason, in the last year its popularity has soared, and it seems likely to become the most common tokenization approach (it may well already be, by the time you read this!).
# Once our texts have been split into tokens, we need to convert them to numbers. We'll look at that next.
# ### Numericalization with fastai
# *Numericalization* is the process of mapping tokens to integers. The steps are basically identical to those necessary to create a `Category` variable, such as the dependent variable of digits in MNIST:
#
# 1. Make a list of all possible levels of that categorical variable (the vocab).
# 1. Replace each level with its index in the vocab.
#
# Let's take a look at this in action on the word-tokenized text we saw earlier:
toks = tkn(txt)
print(coll_repr(tkn(txt), 31))
# Just like with `SubwordTokenizer`, we need to call `setup` on `Numericalize`; this is how we create the vocab. That means we'll need our tokenized corpus first. Since tokenization takes a while, it's done in parallel by fastai; but for this manual walkthrough, we'll use a small subset:
toks200 = txts[:200].map(tkn)
toks200[0]
# We can pass this to `setup` to create our vocab:
num = Numericalize()
num.setup(toks200)
coll_repr(num.vocab,20)
# Our special rules tokens appear first, and then every word appears once, in frequency order. The defaults to `Numericalize` are `min_freq=3,max_vocab=60000`. `max_vocab=60000` results in fastai replacing all words other than the most common 60,000 with a special *unknown word* token, `xxunk`. This is useful to avoid having an overly large embedding matrix, since that can slow down training and use up too much memory, and can also mean that there isn't enough data to train useful representations for rare words. However, this last issue is better handled by setting `min_freq`; the default `min_freq=3` means that any word appearing less than three times is replaced with `xxunk`.
#
# fastai can also numericalize your dataset using a vocab that you provide, by passing a list of words as the `vocab` parameter.
#
# Once we've created our `Numericalize` object, we can use it as if it were a function:
nums = num(toks)[:20]; nums
# This time, our tokens have been converted to a tensor of integers that our model can receive. We can check that they map back to the original text:
' '.join(num.vocab[o] for o in nums)
# Now that we have numbers, we need to put them in batches for our model.
# ### Putting Our Texts into Batches for a Language Model
# When dealing with images, we needed to resize them all to the same height and width before grouping them together in a mini-batch so they could stack together efficiently in a single tensor. Here it's going to be a little different, because one cannot simply resize text to a desired length. Also, we want our language model to read text in order, so that it can efficiently predict what the next word is. this means that each new batch should begin precisely where the previous one left off.
#
# Suppose we have the following text:
#
# > : In this chapter, we will go back over the example of classifying movie reviews we studied in chapter 1 and dig deeper under the surface. First we will look at the processing steps necessary to convert text into numbers and how to customize it. By doing this, we'll have another example of the PreProcessor used in the data block API.\nThen we will study how we build a language model and train it for a while.
#
# The tokenization process will add special tokens and deal with punctuation to return this text:
#
# > : xxbos xxmaj in this chapter , we will go back over the example of classifying movie reviews we studied in chapter 1 and dig deeper under the surface . xxmaj first we will look at the processing steps necessary to convert text into numbers and how to customize it . xxmaj by doing this , we 'll have another example of the preprocessor used in the data block xxup api . \n xxmaj then we will study how we build a language model and train it for a while .
#
# We now have 90 tokens, separated by spaces. Let's say we want a batch size of 6. We need to break this text into 6 contiguous parts of length 15:
# + hide_input=false
#hide_input
stream = "In this chapter, we will go back over the example of classifying movie reviews we studied in chapter 1 and dig deeper under the surface. First we will look at the processing steps necessary to convert text into numbers and how to customize it. By doing this, we'll have another example of the PreProcessor used in the data block API.\nThen we will study how we build a language model and train it for a while."
tokens = tkn(stream)
bs,seq_len = 6,15
d_tokens = np.array([tokens[i*seq_len:(i+1)*seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# -
# In a perfect world, we could then give this one batch to our model. But that approach doesn't scale, because outside of this toy example it's unlikely that a signle batch containing all the texts would fit in our GPU memory (here we have 90 tokens, but all the IMDb reviews together give several million).
#
# So, we need to divide this array more finely into subarrays of a fixed sequence length. It is important to maintain order within and across these subarrays, because we will use a model that maintains a state so that it remembers what it read previously when predicting what comes next.
#
# Going back to our previous example with 6 batches of length 15, if we chose a sequence length of 5, that would mean we first feed the following array:
# + hide_input=true
#hide_input
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15:i*15+seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# -
# Then this one:
# + hide_input=true
#hide_input
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15+seq_len:i*15+2*seq_len] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# -
# And finally:
# + hide_input=true
#hide_input
bs,seq_len = 6,5
d_tokens = np.array([tokens[i*15+10:i*15+15] for i in range(bs)])
df = pd.DataFrame(d_tokens)
display(HTML(df.to_html(index=False,header=None)))
# -
# Going back to our movie reviews dataset, the first step is to transform the individual texts into a stream by concatenating them together. As with images, it's best to randomize the order of the inputs, so at the beginning of each epoch we will shuffle the entries to make a new stream (we shuffle the order of the documents, not the order of the words inside them, or the texts would not make sense anymore!).
#
# We then cut this stream into a certain number of batches (which is our *batch size*). For instance, if the stream has 50,000 tokens and we set a batch size of 10, this will give us 10 mini-streams of 5,000 tokens. What is important is that we preserve the order of the tokens (so from 1 to 5,000 for the first mini-stream, then from 5,001 to 10,000...), because we want the model to read continuous rows of text (as in the preceding example). An `xxbos` token is added at the start of each during preprocessing, so that the model knows when it reads the stream when a new entry is beginning.
#
# So to recap, at every epoch we shuffle our collection of documents and concatenate them into a stream of tokens. We then cut that stream into a batch of fixed-size consecutive mini-streams. Our model will then read the mini-streams in order, and thanks to an inner state, it will produce the same activation whatever sequence length we picked.
#
# This is all done behind the scenes by the fastai library when we create an `LMDataLoader`. We do this by first applying our `Numericalize` object to the tokenized texts:
nums200 = toks200.map(num)
# and then passing that to `LMDataLoader`:
dl = LMDataLoader(nums200)
# Let's confirm that this gives the expected results, by grabbing the first batch:
x,y = first(dl)
x.shape,y.shape
# and then looking at the first row of the independent variable, which should be the start of the first text:
' '.join(num.vocab[o] for o in x[0][:20])
# The dependent variable is the same thing offset by one token:
' '.join(num.vocab[o] for o in y[0][:20])
# This concludes all the preprocessing steps we need to apply to our data. We are now ready to train our text classifier.
# ## Training a Text Classifier
# As we saw at the beginning of this chapter, thare are two steps to training a state-of-the-art text classifier using transfer learning: first we need to fine-tune our language model pretrained on Wikipedia to the corpus of IMDb reviews, and then we can use that model to train a classifier.
#
# As usual, let's start with assembling our data.
# ### Language Model Using DataBlock
# fastai handles tokenization and numericalization automatically when `TextBlock` is passed to `DataBlock`. All of the arguments that can be passed to `Tokenize` and `Numericalize` can also be passed to `TextBlock`. In the next chapter we'll discuss the easiest ways to run each of these steps separately, to ease debugging—but you can always just debug by running them manually on a subset of your data as shown in the previous sections. And don't forget about `DataBlock`'s handy `summary` method, which is very useful for debugging data issues.
#
# Here's how we use `TextBlock` to create a language model, using fastai's defaults:
# +
get_imdb = partial(get_text_files, folders=['train', 'test', 'unsup'])
dls_lm = DataBlock(
blocks=TextBlock.from_folder(path, is_lm=True),
get_items=get_imdb, splitter=RandomSplitter(0.1)
).dataloaders(path, path=path, bs=128, seq_len=80)
# -
# One thing that's different to previous types we've used in `DataBlock` is that we're not just using the class directly (i.e., `TextBlock(...)`, but instead are calling a *class method*. A class method is a Python method that, as the name suggests, belongs to a *class* rather than an *object*. (Be sure to search online for more information about class methods if you're not familiar with them, since they're commonly used in many Python libraries and applications; we've used them a few times previously in the book, but haven't called attention to them.) The reason that `TextBlock` is special is that setting up the numericalizer's vocab can take a long time (we have to read and tokenize every document to get the vocab). To be as efficient as possible preforms a few optimizations:
#
# - It save the tokenized documents in a temporary folder, so it doesn't have to tokenize them more than once
# - It runs multiple tokenization processes in parallel, to take advantage of your computer's CPUs
#
# We need to tell `TextBlock` how to access the texts, so that it can do this initial preprocessing—that's what `from_folder` does.
#
# `show_batch` then works in the usual way:
dls_lm.show_batch(max_n=2)
# Now that our data is ready, we can fine-tune the pretrained language model.
# ### Fine-Tuning the Language Model
# To convert the integer word indices into activations that we can use for our neural network, we will use embeddings, just like we did for collaborative filtering and tabular modeling. Then we'll feed those embeddings into a *recurrent neural network* (RNN), using an architecture called *AWD-LSTM* (we will show you how to write such a model from scratch in <<chapter_nlp_dive>>). As we discussed earlier, the embeddings in the pretrained model are merged with random embeddings added for words that weren't in the pretraining vocabulary. This is handled automatically inside `language_model_learner`:
learn = language_model_learner(
dls_lm, AWD_LSTM, drop_mult=0.3,
metrics=[accuracy, Perplexity()]).to_fp16()
# The loss function used by default is cross-entropy loss, since we essentially have a classification problem (the different categories being the words in our vocab). The *perplexity* metric used here is often used in NLP for language models: it is the exponential of the loss (i.e., `torch.exp(cross_entropy)`). We also include the accuracy metric, to see how many times our model is right when trying to predict the next word, since cross-entropy (as we've seen) is both hard to interpret, and tells us more about the model's confidence than its accuracy.
#
# Let's go back to the process diagram from the beginning of this chapter. The first arrow has been completed for us and made available as a pretrained model in fastai, and we've just built the `DataLoaders` and `Learner` for the second stage. Now we're ready to fine-tune our language model!
# <img alt="Diagram of the ULMFiT process" width="450" src="images/att_00027.png">
# It takes quite a while to train each epoch, so we'll be saving the intermediate model results during the training process. Since `fine_tune` doesn't do that for us, we'll use `fit_one_cycle`. Just like `cnn_learner`, `language_model_learner` automatically calls `freeze` when using a pretrained model (which is the default), so this will only train the embeddings (the only part of the model that contains randomly initialized weights—i.e., embeddings for words that are in our IMDb vocab, but aren't in the pretrained model vocab):
learn.fit_one_cycle(1, 2e-2)
# This model takes a while to train, so it's a good opportunity to talk about saving intermediary results.
# ### Saving and Loading Models
# You can easily save the state of your model like so:
learn.save('1epoch')
# This will create a file in `learn.path/models/` named *1epoch.pth*. If you want to load your model in another machine after creating your `Learner` the same way, or resume training later, you can load the content of this file with:
learn = learn.load('1epoch')
# Once the initial training has completed, we can continue fine-tuning the model after unfreezing:
learn.unfreeze()
learn.fit_one_cycle(10, 2e-3)
# Once this is done, we save all of our model except the final layer that converts activations to probabilities of picking each token in our vocabulary. The model not including the final layer is called the *encoder*. We can save it with `save_encoder`:
learn.save_encoder('finetuned')
# > jargon: Encoder: The model not including the task-specific final layer(s). This term means much the same thing as _body_ when applied to vision CNNs, but "encoder" tends to be more used for NLP and generative models.
# This completes the second stage of the text classification process: fine-tuning the language model. We can now use it to fine-tune a clasifier using the IMDb sentiment labels.
# ### Text Generation
# Before we move on to fine-tuning the classifier, let's quickly try something different: using our model to generate random reviews. Since it's trained to guess what the next word of the sentence is, we can use the model to write new reviews:
TEXT = "I liked this movie because"
N_WORDS = 40
N_SENTENCES = 2
preds = [learn.predict(TEXT, N_WORDS, temperature=0.75)
for _ in range(N_SENTENCES)]
print("\n".join(preds))
# As you can see, we add some randomness (we pick a random word based on the probabilities returned by the model) so we don't get exactly the same review twice. Our model doesn't have any programmed knowledge of the structure of a sentence or grammar rules, yet it has clearly learned a lot about English sentences: we can see it capitalizes properly (*I* is just transformed to *i* because our rules require two characters or more to consider a word as capitalized, so it's normal to see it lowercased) and is using consistent tense. The general review makes sense at first glance, and it's only if you read carefully that you can notice something is a bit off. Not bad for a model trained in a couple of hours!
#
# But our end goal wasn't to train a model to generate reviews, but to classify them... so let's use this model to do just that.
# ### Creating the Classifier DataLoaders
# We're now moving from language model fine-tuning to classifier fine-tuning. To recap, a language model predicts the next word of a document, so it doesn't need any external labels. A classifier, however, predicts some external label—in the case of IMDb, it's the sentiment of a document.
#
# This means that the structure of our `DataBlock` for NLP classification will look very familiar. It's actually nearly the same as we've seen for the many image classification datasets we've worked with:
dls_clas = DataBlock(
blocks=(TextBlock.from_folder(path, vocab=dls_lm.vocab),CategoryBlock),
get_y = parent_label,
get_items=partial(get_text_files, folders=['train', 'test']),
splitter=GrandparentSplitter(valid_name='test')
).dataloaders(path, path=path, bs=128, seq_len=72)
# Just like with image classification, `show_batch` shows the dependent variable (sentiment, in this case) with each independent variable (movie review text):
dls_clas.show_batch(max_n=3)
# Looking at the `DataBlock` definition, every piece is familiar from previous data blocks we've built, with two important exceptions:
#
# - `TextBlock.from_folder` no longer has the `is_lm=True` parameter.
# - We pass the `vocab` we created for the language model fine-tuning.
#
# The reason that we pass the `vocab` of the language model is to make sure we use the same correspondence of token to index. Otherwise the embeddings we learned in our fine-tuned language model won't make any sense to this model, and the fine-tuning step won't be of any use.
#
# By passing `is_lm=False` (or not passing `is_lm` at all, since it defaults to `False`) we tell `TextBlock` that we have regular labeled data, rather than using the next tokens as labels. There is one challenge we have to deal with, however, which is to do with collating multiple documents into a mini-batch. Let's see with an example, by trying to create a mini-batch containing the first 10 documents. First we'll numericalize them:
nums_samp = toks200[:10].map(num)
# Let's now look at how many tokens each of these 10 movie reviews have:
nums_samp.map(len)
# Remember, PyTorch `DataLoader`s need to collate all the items in a batch into a single tensor, and a single tensor has a fixed shape (i.e., it has some particular length on every axis, and all items must be consistent). This should sound familiar: we had the same issue with images. In that case, we used cropping, padding, and/or squishing to make all the inputs the same size. Cropping might not be a good idea for documents, because it seems likely we'd remove some key information (having said that, the same issue is true for images, and we use cropping there; data augmentation hasn't been well explored for NLP yet, so perhaps there are actually opportunities to use cropping in NLP too!). You can't really "squish" a document. So that leaves padding!
#
# We will expand the shortest texts to make them all the same size. To do this, we use a special padding token that will be ignored by our model. Additionally, to avoid memory issues and improve performance, we will batch together texts that are roughly the same lengths (with some shuffling for the training set). We do this by (approximately, for the training set) sorting the documents by length prior to each epoch. The result of this is that the documents collated into a single batch will tend of be of similar lengths. We won't pad every batch to the same size, but will instead use the size of the largest document in each batch as the target size. (It is possible to do something similar with images, which is especially useful for irregularly sized rectangular images, but at the time of writing no library provides good support for this yet, and there aren't any papers covering it. It's something we're planning to add to fastai soon, however, so keep an eye on the book's website; we'll add information about this as soon as we have it working well.)
#
# The sorting and padding are automatically done by the data block API for us when using a `TextBlock`, with `is_lm=False`. (We don't have this same issue for language model data, since we concatenate all the documents together first, and then split them into equally sized sections.)
#
# We can now create a model to classify our texts:
learn = text_classifier_learner(dls_clas, AWD_LSTM, drop_mult=0.5,
metrics=accuracy).to_fp16()
# The final step prior to training the classifier is to load the encoder from our fine-tuned language model. We use `load_encoder` instead of `load` because we only have pretrained weights available for the encoder; `load` by default raises an exception if an incomplete model is loaded:
learn = learn.load_encoder('finetuned')
# ### Fine-Tuning the Classifier
# The last step is to train with discriminative learning rates and *gradual unfreezing*. In computer vision we often unfreeze the model all at once, but for NLP classifiers, we find that unfreezing a few layers at a time makes a real difference:
learn.fit_one_cycle(1, 2e-2)
# In just one epoch we get the same result as our training in <<chapter_intro>>: not too bad! We can pass `-2` to `freeze_to` to freeze all except the last two parameter groups:
learn.freeze_to(-2)
learn.fit_one_cycle(1, slice(1e-2/(2.6**4),1e-2))
# Then we can unfreeze a bit more, and continue training:
learn.freeze_to(-3)
learn.fit_one_cycle(1, slice(5e-3/(2.6**4),5e-3))
# And finally, the whole model!
learn.unfreeze()
learn.fit_one_cycle(2, slice(1e-3/(2.6**4),1e-3))
# We reached 94.3% accuracy, which was state-of-the-art performance just three years ago. By training another model on all the texts read backwards and averaging the predictions of those two models, we can even get to 95.1% accuracy, which was the state of the art introduced by the ULMFiT paper. It was only beaten a few months ago, by fine-tuning a much bigger model and using expensive data augmentation techniques (translating sentences in another language and back, using another model for translation).
#
# Using a pretrained model let us build a fine-tuned language model that was pretty powerful, to either generate fake reviews or help classify them. This is exciting stuff, but it's good to remember that this technology can also be used for malign purposes.
# ## Disinformation and Language Models
# Even simple algorithms based on rules, before the days of widely available deep learning language models, could be used to create fraudulent accounts and try to influence policymakers. <NAME>, now a computational journalist at ProPublica, analyzed the comments that were sent to the US Federal Communications Commission (FCC) regarding a 2017 proposal to repeal net neutrality. In his article ["More than a Million Pro-Repeal Net Neutrality Comments Were Likely Faked"](https://hackernoon.com/more-than-a-million-pro-repeal-net-neutrality-comments-were-likely-faked-e9f0e3ed36a6), he reports how he discovered a large cluster of comments opposing net neutrality that seemed to have been generated by some sort of Mad Libs-style mail merge. In <<disinformation>>, the fake comments have been helpfully color-coded by Kao to highlight their formulaic nature.
# <img src="images/ethics/image16.png" width="700" id="disinformation" caption="Comments received by the FCC during the net neutrality debate">
# Kao estimated that "less than 800,000 of the 22M+ comments… could be considered truly unique" and that "more than 99% of the truly unique comments were in favor of keeping net neutrality."
#
# Given advances in language modeling that have occurred since 2017, such fraudulent campaigns could be nearly impossible to catch now. You now have all the necessary tools at your disposal to create a compelling language model—that is, something that can generate context-appropriate, believable text. It won't necessarily be perfectly accurate or correct, but it will be plausible. Think about what this technology would mean when put together with the kinds of disinformation campaigns we have learned about in recent years. Take a look at the Reddit dilaogue shown in <<ethics_reddit>>, where a language model based on OpenAI's GPT-2 algorithm is having a conversation with itself about whether the US government should cut defense spending.
# <img src="images/ethics/image14.png" id="ethics_reddit" caption="An algorithm talking to itself on Reddit" alt="An algorithm talking to itself on Reddit" width="600">
# In this case, it was explicitly said that an algorithm was used, but imagine what would happen if a bad actor decided to release such an algorithm across social networks. They could do it slowly and carefully, allowing the algorithm to gradually develop followers and trust over time. It would not take many resources to have literally millions of accounts doing this. In such a situation we could easily imagine getting to a point where the vast majority of discourse online was from bots, and nobody would have any idea that it was happening.
#
# We are already starting to see examples of machine learning being used to generate identities. For example, <<katie_jones>> shows a LinkedIn profile for <NAME>.
# <img src="images/ethics/image15.jpeg" width="400" id="katie_jones" caption="<NAME>'s LinkedIn profile">
# <NAME> was connected on LinkedIn to several members of mainstream Washington think tanks. But she didn't exist. That image you see was auto-generated by a generative adversarial network, and somebody named <NAME> has not, in fact, graduated from the Center for Strategic and International Studies.
#
# Many people assume or hope that algorithms will come to our defense here—that we will develop classification algorithms that can automatically recognise autogenerated content. The problem, however, is that this will always be an arms race, in which better classification (or discriminator) algorithms can be used to create better generation algorithms.
# ## Conclusion
# In this chapter we exlored the last application covered out of the box by the fastai library: text. We saw two types of models: language models that can generate texts, and a classifier that determines if a review is positive or negative. To build a state-of-the art classifier, we used a pretrained language model, fine-tuned it to the corpus of our task, then used its body (the encoder) with a new head to do the classification.
#
# Before we end this section, we'll take a look at how the fastai library can help you assemble your data for your specific problems.
# ## Questionnaire
# 1. What is "self-supervised learning"?
# 1. What is a "language model"?
# 1. Why is a language model considered self-supervised?
# 1. What are self-supervised models usually used for?
# 1. Why do we fine-tune language models?
# 1. What are the three steps to create a state-of-the-art text classifier?
# 1. How do the 50,000 unlabeled movie reviews help us create a better text classifier for the IMDb dataset?
# 1. What are the three steps to prepare your data for a language model?
# 1. What is "tokenization"? Why do we need it?
# 1. Name three different approaches to tokenization.
# 1. What is `xxbos`?
# 1. List four rules that fastai applies to text during tokenization.
# 1. Why are repeated characters replaced with a token showing the number of repetitions and the character that's repeated?
# 1. What is "numericalization"?
# 1. Why might there be words that are replaced with the "unknown word" token?
# 1. With a batch size of 64, the first row of the tensor representing the first batch contains the first 64 tokens for the dataset. What does the second row of that tensor contain? What does the first row of the second batch contain? (Careful—students often get this one wrong! Be sure to check your answer on the book's website.)
# 1. Why do we need padding for text classification? Why don't we need it for language modeling?
# 1. What does an embedding matrix for NLP contain? What is its shape?
# 1. What is "perplexity"?
# 1. Why do we have to pass the vocabulary of the language model to the classifier data block?
# 1. What is "gradual unfreezing"?
# 1. Why is text generation always likely to be ahead of automatic identification of machine-generated texts?
# ### Further Research
# 1. See what you can learn about language models and disinformation. What are the best language models today? Take a look at some of their outputs. Do you find them convincing? How could a bad actor best use such a model to create conflict and uncertainty?
# 1. Given the limitation that models are unlikely to be able to consistently recognize machine-generated texts, what other approaches may be needed to handle large-scale disinformation campaigns that leverage deep learning?
|
_notebooks/10_nlp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Zwecc6kFZCMz"
# # **Connect to Google Drive**
# + colab={"base_uri": "https://localhost:8080/"} id="I7cmjYtcqtHf" executionInfo={"status": "ok", "timestamp": 1619136948132, "user_tz": -180, "elapsed": 19913, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="803b95ed-72f1-4872-ea82-c7ada89d64e7"
from google.colab import drive
drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="2DbK7LxsvmpW" executionInfo={"status": "ok", "timestamp": 1619136950372, "user_tz": -180, "elapsed": 806, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="d403c924-2ae7-4f2f-a469-c013b85834fb"
# cd gdrive/My\ Drive/Colab\ Notebooks/
# + id="7C6t60rGSRwf" executionInfo={"status": "ok", "timestamp": 1619136950373, "user_tz": -180, "elapsed": 524, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + id="RiV7e8P5SUiQ" executionInfo={"status": "ok", "timestamp": 1619136950845, "user_tz": -180, "elapsed": 770, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="FhHTjk0pZZB4"
# # **Preprocessing the dataset**
#
# - Load the data from Google Drive.
# - In Google Drive, have 3 directories: train_data - valid_data - test_data.
# - In each folder, data is stored in h5py format.
# - Standardization (z-score).
# + id="hg58Jdv_wnQ7" executionInfo={"status": "ok", "timestamp": 1619136951222, "user_tz": -180, "elapsed": 655, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
import numpy as np
import h5py
# + id="ISqT2YpNHn0t" executionInfo={"status": "ok", "timestamp": 1619136951518, "user_tz": -180, "elapsed": 687, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
def preprocessing_dataset(num_file, filename_prefix):
X = []
y = []
# Load the data from h5py
for i in range(1, num_file):
filename = filename_prefix + str(i) + '.hdf5'
print('Loading batch ' + str(i) + ' ...')
with h5py.File(filename, 'r') as f:
for j in f['X'][:]:
X.append(j)
for j in f['y'][:]:
y.append(j)
# Convert to numpy array
X = np.array(X)
y = np.array(y)
# Calculate the mean of each channel
mean_0 = X[:, 0, :, :].mean()
mean_1 = X[:, 1, :, :].mean()
mean_2 = X[:, 2, :, :].mean()
# Calculate the standard deviation of each channel
std_0 = X[:, 0, :, :].std()
std_1 = X[:, 1, :, :].std()
std_2 = X[:, 2, :, :].std()
# Standardization
X[:, 0, :, :] = (X[:, 0, :, :] - mean_0) / std_0
X[:, 1, :, :] = (X[:, 1, :, :] - mean_1) / std_1
X[:, 2, :, :] = (X[:, 2, :, :] - mean_2) / std_2
return X, y
# + id="9ShtHx4ETpfp" executionInfo={"status": "ok", "timestamp": 1619136951847, "user_tz": -180, "elapsed": 740, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="iihRe7PLJtWa"
# **Configure the dataset**
# + id="PJvsDiR5JrFa" executionInfo={"status": "ok", "timestamp": 1619136952963, "user_tz": -180, "elapsed": 677, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
train_num_file = 10
train_filename_prefix = 'train_data/X_train_'
valid_num_file = 4
valid_filename_prefix = 'valid_data/X_valid_'
test_num_file = 4
test_filename_prefix = 'test_data/X_test_'
# + id="dt9M5zOKSYeZ" executionInfo={"status": "ok", "timestamp": 1619136953475, "user_tz": -180, "elapsed": 832, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="fK1yHloTqq0M"
# ### **Load the training set**
# + colab={"base_uri": "https://localhost:8080/"} id="6Tr-B93DyS2g" executionInfo={"status": "ok", "timestamp": 1619137040924, "user_tz": -180, "elapsed": 87798, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}} outputId="7b7e4022-bfb4-4701-9c7b-395000b3f9f5"
X_train, y_train = preprocessing_dataset(train_num_file, train_filename_prefix)
# + colab={"base_uri": "https://localhost:8080/"} id="NbGvwTXfzuIE" executionInfo={"status": "ok", "timestamp": 1619137040925, "user_tz": -180, "elapsed": 87543, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="09003707-a254-4952-c59b-969103dc8de6"
print(X_train.shape)
print(y_train.shape)
# + id="r-e26mJtSZH1" executionInfo={"status": "ok", "timestamp": 1619137040927, "user_tz": -180, "elapsed": 87357, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="dv0w0DlUrEpX"
# ### **Load the validation set**
# + colab={"base_uri": "https://localhost:8080/"} id="54FTpMpF7P3k" executionInfo={"status": "ok", "timestamp": 1619137066042, "user_tz": -180, "elapsed": 111980, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}} outputId="c23e5160-8d41-46e6-dd89-b88805f6fd17"
X_valid, y_valid = preprocessing_dataset(valid_num_file, valid_filename_prefix)
# + colab={"base_uri": "https://localhost:8080/"} id="SN9_Tb7NQjww" executionInfo={"status": "ok", "timestamp": 1619137066044, "user_tz": -180, "elapsed": 111693, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="6d5f7eb0-5e43-4606-e218-49f4b5882db1"
print(X_valid.shape)
print(y_valid.shape)
# + id="EJY7KP0pSZqM" executionInfo={"status": "ok", "timestamp": 1619137066045, "user_tz": -180, "elapsed": 111444, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="8sh-TSIjnIhU"
# ### **Load the test set**
# + colab={"base_uri": "https://localhost:8080/"} id="3el2jGb-nOQs" executionInfo={"status": "ok", "timestamp": 1619137096669, "user_tz": -180, "elapsed": 141409, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="99b80c7d-4386-4b58-b6e5-2ef555adfcae"
X_test, y_test = preprocessing_dataset(test_num_file, test_filename_prefix)
# + colab={"base_uri": "https://localhost:8080/"} id="zgJ4jkibnOqj" executionInfo={"status": "ok", "timestamp": 1619137096671, "user_tz": -180, "elapsed": 141075, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="62ef5efc-d103-479b-c583-e6df27b109d0"
print(X_test.shape)
print(y_test.shape)
# + id="ZqVc1Y3YSaQw" executionInfo={"status": "ok", "timestamp": 1619137096672, "user_tz": -180, "elapsed": 140706, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + id="3WeOXNGYSbIB" executionInfo={"status": "ok", "timestamp": 1619137096673, "user_tz": -180, "elapsed": 140343, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="46SO5TPuHCqT"
# # **Building Model Convolution Neural Network**
# + id="xZT0mpOOQ-q2" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619137100343, "user_tz": -180, "elapsed": 143308, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="5ffcab9b-ff59-4758-8b36-9d057fa7a9cb"
import math
import time
import random
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sn
import torch
import torch.nn as nn
from torch import optim
from imblearn.over_sampling import SMOTE
import collections
from warnings import simplefilter, filterwarnings
# ignore all future warnings
simplefilter(action='ignore', category=FutureWarning)
filterwarnings("ignore")
# + [markdown] id="9n2NO9hbxwbB"
# ### **Parameters of the model**
# + id="J8DbgXlox9DK" executionInfo={"status": "ok", "timestamp": 1619137100344, "user_tz": -180, "elapsed": 142662, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
EPOCHS = 80
BATCH_SIZE = 128
LR = 1e-6
# + id="FCfrIllISdGu" executionInfo={"status": "ok", "timestamp": 1619137100345, "user_tz": -180, "elapsed": 142302, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="IKQeUCpDVokL"
# ### **Defining the Model**
# + [markdown] id="F9oKDoeJXV8d"
# **Model description:**
#
# - Input model is RGB image data (3 color channels, 50 pixels height, 50 pixels width)
# - Transform input image to multi-dimensional matrix with size 3 - 50 - 50.
#
#
# **Activation function**
#
# - The ReLU is the most used activation function in the world right now. Since, it is used in almost all the convolutional neural networks or deep learning.
# - Computationally efficient—allows the network to converge very quickly.
# - Non-linear—although it looks like a linear function, ReLU has a derivative function and allows for backpropagation.
#
# 
#
# **Avoid overfitting**
#
# To avoid overfitting in neural networks, use two methods of Dropout and Batch normalization.
# - **Dropout**: is one of the most commonly used and the most powerful regularization techniques used in deep learning. Dropout is applied to intermediate layers of the model during the training time. Let's look at an example of how dropout is applied on a linear layer's output that generates 10 values:
#
# + The figure shows what happens when dropout is applied to the linear layer output with a threshold value of 0.2. It randomly masks or zeros 20% of data, so that the model will not be dependent on a particular set of weights or patterns, thus overfitting.
#
#  is a method used to make artificial neural networks faster and more stable through normalization of the input layer by re-centering and re-scaling. To increase the stability of a neural network, batch normalization normalizes the output of a previous activation layer by subtracting the batch mean and dividing by the batch standard deviation.
#
# + id="f-PHjODbRNb1" executionInfo={"status": "ok", "timestamp": 1619137100346, "user_tz": -180, "elapsed": 140485, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=2)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=2)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=2)
self.conv4 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, padding=2)
self.conv5 = nn.Conv2d(in_channels=256, out_channels=512, kernel_size=3, padding=2)
self.bn1 = nn.BatchNorm2d(32)
self.bn2 = nn.BatchNorm2d(64)
self.bn3 = nn.BatchNorm2d(128)
self.bn4 = nn.BatchNorm2d(256)
self.bn5 = nn.BatchNorm2d(512)
self.mp = nn.MaxPool2d(kernel_size=2)
self.relu = nn.ReLU()
self.do = nn.Dropout()
self.fc1 = nn.Linear(4608, 1024)
self.fc2 = nn.Linear(1024, 256)
self.fc3 = nn.Linear(256, 64)
self.fc4 = nn.Linear(64, 16)
self.fc5 = nn.Linear(16, 2)
def forward(self, x):
x = self.mp(self.relu(self.bn1(self.conv1(x))))
x = self.mp(self.relu(self.bn2(self.conv2(x))))
x = self.mp(self.relu(self.bn3(self.conv3(x))))
x = self.mp(self.relu(self.bn4(self.conv4(x))))
x = self.do(self.mp(self.relu(self.bn5(self.conv5(x)))))
x = x.view(x.shape[0], -1)
x = self.relu(self.fc1(x))
x = self.relu(self.fc2(x))
x = self.relu(self.fc3(x))
x = self.relu(self.fc4(x))
out = self.fc5(x)
return out
# + colab={"base_uri": "https://localhost:8080/"} id="ABX8AhN1RRpQ" executionInfo={"status": "ok", "timestamp": 1619137100347, "user_tz": -180, "elapsed": 140095, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="7285a0bf-87e0-4819-ed5c-61e8fbc0a551"
model = CNN()
print(model)
# + id="gJAFxg_eSusn" executionInfo={"status": "ok", "timestamp": 1619137100349, "user_tz": -180, "elapsed": 139595, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="j298Jstjk3Hf"
# **Calculate the number of trainable parameters (weights and biases)**
# + id="-53ysjW9RUhL" executionInfo={"status": "ok", "timestamp": 1619137100351, "user_tz": -180, "elapsed": 138794, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
# + colab={"base_uri": "https://localhost:8080/"} id="Uwng3UiXRW-5" executionInfo={"status": "ok", "timestamp": 1619137100353, "user_tz": -180, "elapsed": 138264, "user": {"displayName": "Trung L\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="11b3ca85-a8a5-4b05-afcb-bae8286f9162"
print(f'The model has {count_parameters(model):,} trainable parameters')
# + id="_EMFt1yKSxFQ" executionInfo={"status": "ok", "timestamp": 1619137100354, "user_tz": -180, "elapsed": 137435, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="PkoWZdg1lk1E"
# **Optimization Algorithm**
#
# - Steps to optimization
#
# + Pass a batch of data through your model.
# + Calculate the loss of your batch by comparing your model's predictions against the actual labels.
# + Calculate the gradient of each of your parameters with respect to the loss.
# + Update each of your parameters by subtracting their gradient multiplied by a small learning rate parameter.
# + id="Vy7JDLV3RZJY" executionInfo={"status": "ok", "timestamp": 1619137100355, "user_tz": -180, "elapsed": 136519, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
optimizer = optim.Adam(model.parameters(), lr=LR)
# + id="uwqIpOEsSy_4" executionInfo={"status": "ok", "timestamp": 1619137100356, "user_tz": -180, "elapsed": 136024, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="lL6ZwLSFna9u"
# **Loss Fuction**
#
# - **Cross-entropy** can be used to define a loss function in machine learning and optimization.
# + id="Ellkdw4mRbIR" executionInfo={"status": "ok", "timestamp": 1619137100357, "user_tz": -180, "elapsed": 135100, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
loss_fn = nn.CrossEntropyLoss()
# + id="0LxlbzOzSzss" executionInfo={"status": "ok", "timestamp": 1619137100358, "user_tz": -180, "elapsed": 134713, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="HnAa7POyomp4"
# **Calculate the accuracy**
# - This takes the index of the highest value for your prediction and compares it against the actual class label. Then divide how many our model got correct by the amount in the batch to calculate accuracy across the batch.
# + id="Q2C1ydL2RjDm" executionInfo={"status": "ok", "timestamp": 1619137100359, "user_tz": -180, "elapsed": 133929, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def calculate_accuracy(y_pred, y):
top_pred = y_pred.argmax(1, keepdim = True)
correct = top_pred.eq(y.view_as(top_pred)).sum()
acc = correct.float() / y.shape[0]
return acc
# + id="uCnsmr96S0d4" executionInfo={"status": "ok", "timestamp": 1619137100361, "user_tz": -180, "elapsed": 133358, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="OJN5Z-YYo0KM"
# **Create mini-batch**
# + id="PtW_l5uGRlPL" executionInfo={"status": "ok", "timestamp": 1619137100362, "user_tz": -180, "elapsed": 132484, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
def next_batch(X, y, batch_size):
# Mini-batch
for i in np.arange(0, X.shape[0], batch_size):
yield X[i:i + batch_size], y[i:i + batch_size]
# + id="Qw-P-Qt0S1Mg" executionInfo={"status": "ok", "timestamp": 1619137100723, "user_tz": -180, "elapsed": 132274, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="ohXGSlXgoJAc"
# **Transform the model and data to GPU**
# + id="5aA6REEXRdIe" executionInfo={"status": "ok", "timestamp": 1619137100725, "user_tz": -180, "elapsed": 131316, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + id="bYkiWamORfNN" executionInfo={"status": "ok", "timestamp": 1619137105739, "user_tz": -180, "elapsed": 135708, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
model = model.to(device)
loss_fn = loss_fn.to(device)
# + id="rsSTDNtxS1uH" executionInfo={"status": "ok", "timestamp": 1619137105740, "user_tz": -180, "elapsed": 135125, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + id="YKabQCgtS10v" executionInfo={"status": "ok", "timestamp": 1619137105740, "user_tz": -180, "elapsed": 134621, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="VWN6jKWulMmz"
# ### **Training the Model**
# + [markdown] id="dJjXwVq3OV8_"
# **Define Traning step**
# - Put the model into train mode: **model.train()**
# - Iterate over batches of data.
# - Transform the batch to GPU mode.
# - Clear the gradients calculated from the last batch.
# - Impliment model for batch of images: x, get predictions, y_pred.
# - Calculate the loss between prediction labels and the actual labels.
# - Calculate the accuracy between prediction labels and the actual labels.
# - Calculate the gradients of each parameter.
# - Update the parameters by taking optimizer step.
# - Return the loss and the accuracy of epoch.
# + id="uQag77GvRoIo" executionInfo={"status": "ok", "timestamp": 1619137105741, "user_tz": -180, "elapsed": 132837, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def train(model, X, y, optimizer, loss_fn, device, batch_size):
epoch_loss = 0
epoch_acc = 0
# Impliment train mode
model.train()
for (batchX, batchY) in next_batch(X, y, batch_size):
# SMOTE
x_tmp = batchX.reshape(batchX.shape[0], 3 * 50 * 50)
y_tmp = batchY
count = collections.Counter(y_tmp)
x_smote, y_smote = SMOTE(sampling_strategy={0: count[0], 1: count[0]}, k_neighbors=3).fit_resample(x_tmp, y_tmp)
x_smote = x_smote.reshape(x_smote.shape[0], 3, 50, 50)
# Create tensor
x = torch.from_numpy(x_smote).to(device)
y = torch.from_numpy(y_smote).to(device)
# Clear the gradients calculated from the last batch
optimizer.zero_grad()
y_pred = model(x)
loss = loss_fn(y_pred, y)
acc = calculate_accuracy(y_pred, y)
# Calculate the gradients of each parameter
loss.backward()
# Update the parameters
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
num_of_batchs = math.ceil(len(X) / batch_size)
return epoch_loss / num_of_batchs, epoch_acc / num_of_batchs
# + id="DzOgNwIYS3_h" executionInfo={"status": "ok", "timestamp": 1619137105741, "user_tz": -180, "elapsed": 130779, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="slsqH7YBQZrU"
# **Define Evaluation step**
#
# - Similar the training step. The differences are:
# + Put the model into evalution mode: **model.eval()**
# + Don't calculate the gradients of each parameter.
# + Don't update the parameters by taking optimizer step.
#
# - **torch.no_grad()** ensures that gradients are not calculated for whatever is inside the with block. As our model will not have to calculate gradients it will be faster and use less memory.
# + id="DASp2ZTYSVzh" executionInfo={"status": "ok", "timestamp": 1619137105742, "user_tz": -180, "elapsed": 129377, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def evaluate(model, X, y, loss_fn, device, batch_size):
epoch_loss = 0
epoch_acc = 0
# Impliment evaluation mode
model.eval()
with torch.no_grad():
for (batchX, batchY) in next_batch(X, y, batch_size):
x = torch.from_numpy(batchX).to(device)
y = torch.from_numpy(batchY).to(device)
y_pred = model(x)
loss = loss_fn(y_pred, y)
acc = calculate_accuracy(y_pred, y)
epoch_loss += loss.item()
epoch_acc += acc.item()
num_of_batchs = math.ceil(len(X) / batch_size)
return epoch_loss / num_of_batchs, epoch_acc / num_of_batchs
# + id="o9o2ZxaQS42-" executionInfo={"status": "ok", "timestamp": 1619137105742, "user_tz": -180, "elapsed": 128879, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="3D-urAR5RrK3"
# **Calculate the time need to run each epoch**
# + id="gPKz1vGhSiNH" executionInfo={"status": "ok", "timestamp": 1619137105743, "user_tz": -180, "elapsed": 127951, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + id="qJatZ8upS5ZD" executionInfo={"status": "ok", "timestamp": 1619137105744, "user_tz": -180, "elapsed": 127478, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="zuP4bbb8R4ki"
# **Training the Model**
# - Iterate over epochs.
# - In each epochs:
# - Training the Model with training set.
# - Evaluate the Model with validation set.
# - Check the best validation loss and save the model parameters
# - Print the result metrics: Loss and Accuracy.
# + colab={"base_uri": "https://localhost:8080/"} id="_-pyaes-SlCC" executionInfo={"status": "ok", "timestamp": 1619143064287, "user_tz": -180, "elapsed": 6084911, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}} outputId="8c8c27e2-69fe-4e8f-9e1b-bdcfdda93ec7"
best_valid_loss = float('inf')
train_loss_list = list()
valid_loss_list = list()
for epoch in range(EPOCHS):
start_time = time.monotonic()
train_loss, train_acc = train(model, X_train, y_train, optimizer, loss_fn, device, batch_size=BATCH_SIZE)
valid_loss, valid_acc = evaluate(model, X_valid, y_valid, loss_fn, device, batch_size=BATCH_SIZE)
train_loss_list.append(train_loss)
valid_loss_list.append(valid_loss)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'smote_v1.pt')
end_time = time.monotonic()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
print(f'Epoch: {epoch+1:02} | Epoch Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
# + id="AO_4rnTlS6qM" executionInfo={"status": "ok", "timestamp": 1619143064288, "user_tz": -180, "elapsed": 6084133, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="rclO3ZcaT3I_"
# **Load the parameters of the best model**
# + colab={"base_uri": "https://localhost:8080/"} id="W3pRCywIld27" executionInfo={"status": "ok", "timestamp": 1619143064289, "user_tz": -180, "elapsed": 6083386, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="042638eb-3bde-4fd2-c12f-98d9989e4102"
model.load_state_dict(torch.load('smote_v1.pt'))
# + id="s7jDC-emS7SI" executionInfo={"status": "ok", "timestamp": 1619143064290, "user_tz": -180, "elapsed": 6083156, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + id="Lxo29pkTS7bU" executionInfo={"status": "ok", "timestamp": 1619143064290, "user_tz": -180, "elapsed": 6082857, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="rP-QoefNUOJL"
# ### **Examining the Model**
# + id="vsuC-nhhTIIy" executionInfo={"status": "ok", "timestamp": 1619143064291, "user_tz": -180, "elapsed": 6082364, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
from sklearn.metrics import confusion_matrix, accuracy_score, balanced_accuracy_score, f1_score, precision_score, recall_score
# + [markdown] id="C8eXQ8b7CQRe"
# **Visualizing loss of train set and valid set**
# + id="8EnfxhZ2Cffe" executionInfo={"status": "ok", "timestamp": 1619143064291, "user_tz": -180, "elapsed": 6081874, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
epoch_list = np.arange(1, EPOCHS + 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="nHpukHOwC85C" executionInfo={"status": "ok", "timestamp": 1619143064292, "user_tz": -180, "elapsed": 6081638, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="6bfe79d3-3aa3-42dc-e9a4-e037770b3020"
fig, ax = plt.subplots(figsize=(14,8))
ax.plot(epoch_list, train_loss_list, label='train loss')
ax.plot(epoch_list, valid_loss_list, label='validate loss')
ax.set_xlabel('Epoch')
ax.set_ylabel('Loss')
ax.legend()
plt.show()
# + id="9eC3_r9dS82t" executionInfo={"status": "ok", "timestamp": 1619143064293, "user_tz": -180, "elapsed": 6081383, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="xcSAID4QUZEH"
# **Get the predictions**
# + id="LcWx6hIj8Ucv" executionInfo={"status": "ok", "timestamp": 1619143064294, "user_tz": -180, "elapsed": 6080954, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def get_predictions(model, X, y, device, batch_size):
model.eval()
y_predict = []
with torch.no_grad():
for (batchX, batchY) in next_batch(X, y, batch_size):
X = torch.from_numpy(batchX).to(device)
y_tmp = model(X)
top_pred = y_tmp.argmax(1, keepdim = True)
for i in top_pred:
y_predict.append(i.item())
return np.array(y_predict)
# + id="OYDZ8I7E_B1m" executionInfo={"status": "ok", "timestamp": 1619143068233, "user_tz": -180, "elapsed": 6084629, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
y_predict = get_predictions(model, X_test, y_test, device, batch_size=BATCH_SIZE)
# + id="UKGucRbAS9on" executionInfo={"status": "ok", "timestamp": 1619143068234, "user_tz": -180, "elapsed": 6084439, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="D6cEYYL2Uir5"
# **Confusion matrix**
# + colab={"base_uri": "https://localhost:8080/"} id="I3199D2tCKGK" executionInfo={"status": "ok", "timestamp": 1619143068234, "user_tz": -180, "elapsed": 6083943, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="ce80ef26-d010-4a3b-d092-4e77308c7761"
cm = confusion_matrix(y_test, y_predict)
print(cm)
# + id="2zOKEZraEiZp" executionInfo={"status": "ok", "timestamp": 1619143068235, "user_tz": -180, "elapsed": 6083704, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
pd_cm = pd.DataFrame(cm, index = ['negative', 'positive'], columns = ['negative', 'positive'])
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="48rztXHME2Uu" executionInfo={"status": "ok", "timestamp": 1619143068992, "user_tz": -180, "elapsed": 6084199, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}} outputId="f79a44e5-6eb0-4c78-d60f-44ccf319b2b1"
plt.figure(figsize=(14,10))
ax = plt.axes()
sn.heatmap(pd_cm, annot=True, fmt='d', cmap='Blues')
ax.set_xlabel('Predict label')
ax.set_ylabel('True label')
plt.show()
# + id="4A0px8nbTSY_" executionInfo={"status": "ok", "timestamp": 1619143068993, "user_tz": -180, "elapsed": 6084021, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
# + [markdown] id="F-OYpxThG_n5"
# **Quantifying the quality of predictions**
# + id="OhAg9GrbHPSX" executionInfo={"status": "ok", "timestamp": 1619143068994, "user_tz": -180, "elapsed": 6083540, "user": {"displayName": "<NAME>\u01b0\u01a1ng", "photoUrl": "", "userId": "02044496887486952450"}}
def show_metrics_scores(y_test, y_predict):
print(f'Accuracy score\t\t: {accuracy_score(y_test, y_predict):.4f}\n')
print(f'Balanced accuracy score\t: {balanced_accuracy_score(y_test, y_predict):.4f}\n')
print(f'F1 score\t\t: {f1_score(y_test, y_predict):.4f}\n')
print(f'Recall score\t\t: {recall_score(y_test, y_predict):.4f}')
# + colab={"base_uri": "https://localhost:8080/"} id="YEtRmFG7lx5l" executionInfo={"status": "ok", "timestamp": 1619143068994, "user_tz": -180, "elapsed": 6083297, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}} outputId="ae4c7312-ee80-4bce-fc58-a67f7ae148b8"
show_metrics_scores(y_test, y_predict)
# + id="2OuGt43Tap63" executionInfo={"status": "ok", "timestamp": 1619143068995, "user_tz": -180, "elapsed": 6083193, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02044496887486952450"}}
|
CNN_SMOTE_100.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: gfm
# language: python
# name: gfm
# ---
# +
import sys
sys.path.append('../src/')
import data_io
import pandas as pd
import numpy as np
import re
# -
unique_locations = pd.read_csv(data_io.input_cleaned/'geolocations'/
'unique_locations_w_fips_scraped.csv',
encoding='utf-8',
dtype={'state_county_fips_str':'str'})
unique_locations.dropna(subset=["state_county_fips_str"],inplace=True)
print(len(unique_locations))
unique_locations.head()
df = pd.read_csv(data_io.input_cleaned/'gfm'/'US_cancer_campaigns_2018_2021.csv',index_col=[0],
sep='|',encoding='utf-8')
df.head()
# +
# exclusion_df = pd.read_csv(data_io.input_cleaned/'gfm'/'exclusion_tracker_rd_2.csv',
# index_col = 0)
# +
#drop locations that didn't geocode
unique_locations = unique_locations.replace('nan',np.nan).replace('none',np.nan)
unique_locations.dropna(subset=['county_name'], inplace=True)
county_dict = dict(
zip(unique_locations['location_city'].to_list(), unique_locations['county_name'].to_list()))
long_fips_dict = dict(zip(unique_locations['location_city'].to_list(),
unique_locations['state_county_fips_str'].to_list()))
cleaned_location_city = df['location_city']
df['location_county'] = cleaned_location_city.map(county_dict)
df['location_state_county_fip'] = cleaned_location_city.map(long_fips_dict)
# -
unique_locs_to_scrape = df.loc[df.location_state_county_fip.isna(),
['location_city','location_city_only','location_stateprefix']].drop_duplicates()
print(unique_locs_to_scrape.shape)
unique_locs_to_scrape.to_csv(data_io.input_cleaned/'geolocations'/'unique_locations_to_scrape_again.csv')
# +
#exclusion_df.loc['deleted', 'failed_geocode'] = df['county'].isnull().sum()
df = df.dropna(subset = ['location_county'])
# exclusion_df.loc['total', 'failed_geocode'] = len(df)
# exclusion_df.to_csv(data_io.input_cleaned/'gfm'/'final_exclusion_tracker.csv')
geo_fail = df[pd.isnull(df['location_county'])]
#save failed geocodes to make sure nothing in the US failed
save = False
if save:
geo_fail.to_csv(data_io.input_cleaned/'gfm'/'master_failed_geocode.csv', encoding='utf-8-sig')
df.dropna(subset=['location_county'], inplace=True)
# -
# #### Define text mining functions
# +
#functions for additional text mining
SEARCH_OPTIONS = pd.read_csv(data_io.gfm/'gfm'/'free_text_search_terms_w_covid.csv')
SEARCH_DICT = {k:SEARCH_OPTIONS[k].dropna().to_list() for k in SEARCH_OPTIONS.columns}
def create_dict(search_type):
key_col = 'collapsed_'+search_type
new = SEARCH_OPTIONS.dropna(subset=[search_type])
this_dict = pd.Series(new[key_col].values,index=new[search_type].values).to_dict()
return this_dict
INSURE_DICT = create_dict('insurance_type')
#OOP_DICT = create_dict('oop_type')
import string
#import regex as re
def extract_search_term_regex(x, search_type = 'cancer_type', return_context = False,
find_uninsured = False, collapse_dict = 'none'):
if type(x) == str:
x = x.lower()
else:
return np.nan
search_terms = SEARCH_DICT[search_type]
#match only if char after match is a space or punctuation
if 'cancer' in search_type:
for s in search_terms:
smatch = re.search(s+'\W', x)
if smatch:
if return_context == True:
end_smatch = smatch.span()[1]
new = x[smatch.span()[0]:]
new = new[0: new.find('.')]
return new
return(x[smatch.span()[0]:smatch.span()[1]])
return np.nan
else:
return_val = False
uninsure = False
mention = []
collapsed_mention = []
for s in search_terms:
smatch = re.search(s, x)
if smatch:
if return_context == True:
new = x[smatch.span()[0]:smatch.span()[1]]
#print(new)
mention.append(new)
if find_uninsured == True:
if INSURE_DICT[s] == 'uninsured' or INSURE_DICT[s] == 'underunisured':
return True
else:
return_val = True
if type(collapse_dict) != str:
collapsed_mention.append(collapse_dict[s])
return_val = True
if len(mention) >= 1:
mention = ','.join(mention)
else:
mention = None
if len(collapsed_mention) >= 1:
collapsed_mention = np.unique(np.asarray(collapsed_mention))
collapsed_mention = list(collapsed_mention)
collapsed_mention = ','.join
else:
collapsed_mention = None
if type(collapse_dict)!= str:
return collapsed_mention
return mention if return_context == True else return_val
def search_story_and_title(story, title, search_type):
story_truth = extract_search_term_regex(story, search_type = search_type)
title_truth = extract_search_term_regex(title, search_type = search_type)
if title_truth == True or story_truth == True:
return True
elif title_truth == False and story_truth == False:
return False
def get_all_mentions(story, title, search_type):
story_truth = extract_search_term_regex(story,
search_type = search_type,
return_context = True)
title_truth = extract_search_term_regex(title,
search_type = search_type,
return_context = True)
if type(story_truth) == str:
if type(title_truth) == str:
story_truth += title_truth
return story_truth
elif type(title_truth) == str:
return title_truth
else:
return None
#Returns a comma separated string of the features that match the search in question
def extract_feature(story, feature = 'tx_type_search', title = None):
features = SEARCH_DICT[feature]
if pd.isnull(title):
searches = [story]
else:
searches = [story, title]
return_str = ''
for x in searches:
if type(x) == str:
x = x.lower()
for f in features:
if f in x:
if len(return_str) == 0:
return_str += f
else:
return_str += ', '
return_str += f
if len(return_str) == 0:
return np.nan
else:
return return_str
def collapse_feature(mentions, feature_dict):
if type(mentions) == str:
temp_mentions = mentions.split(', ')
new_mentions = []
for t in temp_mentions:
new_mentions.append(feature_dict[t])
new_mentions = np.unique(new_mentions)
new_mentions = ', '.join(new_mentions)
return new_mentions
def assign_num_occurrences(mentions):
if type(mentions) == str:
if ',' in mentions:
new = mentions.split(',')
return len(new)
else:
if mentions != '':
return 1
else:
return 0
# -
# #### Mine each text feature
# +
#Look for clinical/financial details
recode_feats_to_search = ['oop_type', 'insurance_type', 'tx_type', 'cancer_type']
df['story_and_title'] = df['title'] + ' ' + df['story']
for r in recode_feats_to_search:
new_col = r + '_is_mentioned'
print(f"searching for {r}")
df[new_col] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = r),
axis = 1)
print(f"extracting {r}")
df[r] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = r,
return_context = True),
axis = 1)
recode = 'collapsed_' + r
feat_dict = create_dict(r)
print(f"collapsing {r}")
df[recode] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = r,
collapse_dict = feat_dict),
axis = 1)
df['num_tx'] = df['collapsed_tx_type'].apply(assign_num_occurrences)
df['num_oop'] = df['collapsed_oop_type'].apply(assign_num_occurrences)
df['uninsured'] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = 'insurance_type',
find_uninsured = True),
axis = 1)
#Look for mention of Covid
df["covid_is_mentioned"] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = "covid"),
axis = 1)
#Look for worth indicators
worth_indicators = ['brave', 'nice', 'thank', 'self_reliance', 'battle']
for w in worth_indicators:
new_col = w + '_is_mentioned'
print(f"searching for {w}")
df[new_col] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = w),
axis = 1)
print(f"extracting {w}")
df[w] = df.apply(lambda x: extract_search_term_regex(x['story_and_title'],
search_type = w,
return_context = True),
axis = 1)
save = True
if save:
df.drop(columns=['story_and_title'],inplace=True)
df.to_csv(data_io.input_cleaned/'gfm'/'US_cancer_campaigns_2018_2021_with_fips_and_text_features.csv',
encoding='utf-8',sep='|')
# -
df.columns
# Number of campaigns that mentioned COVID vs all campaigns
df.covid.sum(),df.shape
|
notebooks/Assign location FIPs and text indicators.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os import path
# Third-party
import astropy.coordinates as coord
from astropy.table import Table, vstack
from astropy.io import fits, ascii
import astropy.units as u
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
from scipy.interpolate import InterpolatedUnivariateSpline
from scipy.optimize import minimize
from pyia import GaiaData
import gala.coordinates as gc
import gala.dynamics as gd
from gala.dynamics import mockstream
import gala.integrate as gi
import gala.potential as gp
from gala.units import galactic
from gala.mpl_style import center_emph
# -
plt.style.use('notebook')
track = Table.read('../data/stream_track.fits')
# +
# Koposov et al.
kop_pm = ascii.read("""phi1 pm_phi1 pm_phi2 err
-55.00 -13.60 -5.70 1.30
-45.00 -13.10 -3.30 0.70
-35.00 -12.20 -3.10 1.00
-25.00 -12.60 -2.70 1.40
-15.00 -10.80 -2.80 1.00""")
kop_phi2 = ascii.read("""phi1 phi2 err
-60.00 -0.64 0.15
-56.00 -0.89 0.27
-54.00 -0.45 0.15
-48.00 -0.08 0.13
-44.00 0.01 0.14
-40.00 -0.00 0.09
-36.00 0.04 0.10
-34.00 0.06 0.13
-32.00 0.04 0.06
-30.00 0.08 0.10
-28.00 0.03 0.12
-24.00 0.06 0.05
-22.00 0.06 0.13
-18.00 -0.05 0.11
-12.00 -0.29 0.16
-2.00 -0.87 0.07""")
kop_dist = ascii.read("""phi1 dist err
-55.00 7.20 0.30
-45.00 7.59 0.40
-35.00 7.83 0.30
-25.00 8.69 0.40
-15.00 8.91 0.40
0.00 9.86 0.50""")
kop_vr = ascii.read("""phi1 phi2 vr err
-45.23 -0.04 28.8 6.9
-43.17 -0.09 29.3 10.2
-39.54 -0.07 2.9 8.7
-39.25 -0.22 -5.2 6.5
-37.95 0.00 1.1 5.6
-37.96 -0.00 -11.7 11.2
-35.49 -0.05 -50.4 5.2
-35.27 -0.02 -30.9 12.8
-34.92 -0.15 -35.3 7.5
-34.74 -0.08 -30.9 9.2
-33.74 -0.18 -74.3 9.8
-32.90 -0.15 -71.5 9.6
-32.25 -0.17 -71.5 9.2
-29.95 -0.00 -92.7 8.7
-26.61 -0.11 -114.2 7.3
-25.45 -0.14 -67.8 7.1
-24.86 0.01 -111.2 17.8
-21.21 -0.02 -144.4 10.5
-14.47 -0.15 -179.0 10.0
-13.73 -0.28 -191.4 7.5
-13.02 -0.21 -162.9 9.6
-12.68 -0.26 -217.2 10.7
-12.55 -0.23 -172.2 6.6""")
# +
dt = 0.5 * u.Myr
n_steps = 250
_phi2_sigma = 0.2 # deg
_dist_sigma = 0.1 # kpc
_vr_sigma = 1 # km/s
def ln_likelihood(p, phi1_0, data, ham, gc_frame):
# initial conditions at phi1_0
phi2, d, pm1, pm2, vr = p
c = gc.GD1(phi1=phi1_0, phi2=phi2*u.deg, distance=d*u.kpc,
pm_phi1_cosphi2=pm1*u.mas/u.yr,
pm_phi2=pm2*u.mas/u.yr,
radial_velocity=vr*u.km/u.s)
w0 = gd.PhaseSpacePosition(c.transform_to(gc_frame).cartesian)
orbit = ham.integrate_orbit(w0, dt=dt, n_steps=n_steps)
model_gd1 = orbit.to_coord_frame(gc.GD1, galactocentric_frame=gc_frame)
model_x = model_gd1.phi1.wrap_at(180*u.deg).degree
if model_x[-1] < -180:
return -np.inf
model_phi2 = model_gd1.phi2.degree
model_dist = model_gd1.distance.to(u.kpc).value
model_pm1 = model_gd1.pm_phi1_cosphi2.to(u.mas/u.yr).value
model_pm2 = model_gd1.pm_phi2.to(u.mas/u.yr).value
model_vr = model_gd1.radial_velocity.to(u.km/u.s).value
# plt.errorbar(data['phi2'][0], data['phi2'][1], marker='o', linestyle='none')
# plt.errorbar(data['pm2'][0], data['pm2'][1], marker='o', linestyle='none')
# plt.plot(model_x, model_pm2)
# return
ix = np.argsort(model_x)
model_x = model_x[ix]
# define interpolating functions
order = 3
bbox = [-180, 180]
chi2 = 0
phi2_interp = InterpolatedUnivariateSpline(model_x, model_phi2[ix],
k=order, bbox=bbox)
dist_interp = InterpolatedUnivariateSpline(model_x, model_dist[ix],
k=order, bbox=bbox)
pm1_interp = InterpolatedUnivariateSpline(model_x, model_pm1[ix],
k=order, bbox=bbox)
pm2_interp = InterpolatedUnivariateSpline(model_x, model_pm2[ix],
k=order, bbox=bbox)
vr_interp = InterpolatedUnivariateSpline(model_x, model_vr[ix],
k=order, bbox=bbox)
phi2_sigma = np.sqrt(_phi2_sigma**2 + data['phi2'][2]**2)
chi2 += np.sum(-(phi2_interp(data['phi2'][0]) - data['phi2'][1])**2 / phi2_sigma**2 - 2*np.log(phi2_sigma))
dist_sigma = np.sqrt(_dist_sigma**2 + data['dist'][2]**2)
chi2 += np.sum(-(dist_interp(data['dist'][0]) - data['dist'][1])**2 / dist_sigma**2 - 2*np.log(dist_sigma))
pm1_sigma = data['pm1'][2]
chi2 += np.sum(-(pm1_interp(data['pm1'][0]) - data['pm1'][1])**2 / pm1_sigma**2 - 2*np.log(pm1_sigma))
pm2_sigma = data['pm2'][2]
chi2 += np.sum(-(pm2_interp(data['pm2'][0]) - data['pm2'][1])**2 / pm2_sigma**2 - 2*np.log(pm2_sigma))
vr_sigma = np.sqrt(data['vr'][2]**2 + _vr_sigma**2)
chi2 += np.sum(-(vr_interp(data['vr'][0]) - data['vr'][1])**2 / vr_sigma**2 - 2*np.log(vr_sigma))
return chi2
# +
data = dict()
# Koposov data:
# data['phi2'] = (kop_phi2['phi1'], kop_phi2['phi2'], kop_phi2['err'])
# data['dist'] = (kop_dist['phi1'], kop_dist['dist'], kop_dist['err'])
# data['pm1'] = (kop_pm['phi1'], kop_pm['pm_phi1'], kop_pm['err'])
# data['pm2'] = (kop_pm['phi1'], kop_pm['pm_phi2'], kop_pm['err'])
# data['vr'] = (kop_vr['phi1'], kop_vr['vr'], kop_vr['err'])
# Ana's track:
data['phi2'] = (track['phi1'], track['phi2'], track['w'])
data['dist'] = (kop_dist['phi1'], kop_dist['dist'], kop_dist['err'])
data['pm1'] = (track['phi1'], track['pm_phi1_cosphi2'], track['pm_phi1_cosphi2_error'])
data['pm2'] = (track['phi1'], track['pm_phi2'], track['pm_phi2_error'])
data['vr'] = (kop_vr['phi1'], kop_vr['vr'], kop_vr['err'])
# +
ham = gp.Hamiltonian(gp.LogarithmicPotential(v_c=225*u.km/u.s, r_h=0*u.kpc, q1=1, q2=1, q3=1,
units=galactic))
# ham = gp.Hamiltonian(gp.load('../output/mwpot.yml'))
print(ham.potential.parameters)
xyz = np.zeros((3, 128))
xyz[0] = np.linspace(1, 25, 128)
plt.plot(xyz[0], ham.potential.circular_velocity(xyz))
plt.ylim(200, 240)
plt.axvline(8)
# +
gc_frame = coord.Galactocentric(galcen_distance=8*u.kpc, z_sun=0*u.pc)
phi1_0 = 10. * u.deg
p0 = (-3., 9., -5.5, -0, -270.)
# -
res = minimize(lambda *x: -ln_likelihood(*x), x0=p0, args=(phi1_0, data, ham, gc_frame))
res_out = np.hstack([np.array([phi1_0.to(u.deg).value]), res.x])
np.save('../data/log_orbit', res_out)
pos = np.load('../data/log_orbit.npy')
# +
phi1, phi2, d, pm1, pm2, vr = pos
c = gc.GD1(phi1=phi1*u.deg, phi2=phi2*u.deg, distance=d*u.kpc,
pm_phi1_cosphi2=pm1*u.mas/u.yr,
pm_phi2=pm2*u.mas/u.yr,
radial_velocity=vr*u.km/u.s)
w0 = gd.PhaseSpacePosition(c.transform_to(gc_frame).cartesian)
# +
# dt = 0.5*u.Myr
t = 56*u.Myr
n_steps = 1000
dt = t/n_steps
fit_orbit = ham.integrate_orbit(w0, dt=dt, n_steps=n_steps)
model_gd1 = fit_orbit.to_coord_frame(gc.GD1, galactocentric_frame=gc_frame)
model_x = model_gd1.phi1.wrap_at(180*u.deg).degree
# +
fig, axes = plt.subplots(5, 1, figsize=(12, 15), sharex=True)
axes[0].errorbar(data['phi2'][0], data['phi2'][1], data['phi2'][2], marker='o', linestyle='none', color='k')
axes[1].errorbar(data['dist'][0], data['dist'][1], data['dist'][2], marker='o', linestyle='none', color='k')
axes[2].errorbar(data['pm1'][0], data['pm1'][1], data['pm1'][2], marker='o', linestyle='none', color='k')
axes[3].errorbar(data['pm2'][0], data['pm2'][1], data['pm2'][2], marker='o', linestyle='none', color='k')
axes[4].errorbar(data['vr'][0], data['vr'][1], data['vr'][2], marker='o', linestyle='none', color='k')
axes[0].plot(model_x, model_gd1.phi2.degree, 'r-', zorder=100)
axes[1].plot(model_x, model_gd1.distance, 'r-', zorder=100)
axes[2].plot(model_x, model_gd1.pm_phi1_cosphi2.to(u.mas/u.yr).value, 'r-', zorder=100)
axes[3].plot(model_x, model_gd1.pm_phi2.to(u.mas/u.yr).value, 'r-', zorder=100)
axes[4].plot(model_x, model_gd1.radial_velocity.to(u.km/u.s).value, 'r-', zorder=100)
ylabels = ['$\phi_2$ [deg]', 'distance [kpc]', '$\mu_{\phi_1}$ [mas yr$^{-1}$]',
'$\mu_{\phi_2}$ [mas yr$^{-1}$]', '$V_r$ [km s$^{-1}$]']
for i in range(5):
plt.sca(axes[i])
plt.ylabel(ylabels[i])
plt.xlabel('$\phi_1$ [deg]')
# axes[0].set_xlim(-100, 20)
plt.tight_layout()
plt.savefig('../plots/log_orbit_fit.png', dpi=100)
# -
fit_orbit.pos.get_xyz()
fit_orbit.vel.get_d_xyz().to(u.km/u.s)
|
notebooks/log_orbit.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.5
# language: julia
# name: julia-1.0
# ---
# # Julia for Data Science
# Prepared by [@nassarhuda](https://github.com/nassarhuda)! 😃
#
# `Last updated on 03/Jan/2020`
#
# In this tutorial, we will discuss why *Julia* is the tool you want to use for your data science applications.
#
# We will cover the following:
# * **Data**
# * Data processing
# * Visualization
#
# ### Data: Build a strong relationship with your data.
# Every data science task has one main ingredient, the _data_! Most likely, you want to use your data to learn something new. But before the _new_ part, what about the data you already have? Let's make sure you can **read** it, **store** it, and **understand** it before you start using it.
#
# Julia makes this step really easy with data structures and packages to process the data, as well as, existing functions that are readily usable on your data.
#
# The goal of this first part is get you acquainted with some Julia's tools to manage your data.
# First, let's download a csv file from github that we can work with.
#
# Note: `download` depends on external tools such as curl, wget or fetch. So you must have one of these.
P = download("https://raw.githubusercontent.com/nassarhuda/easy_data/master/programming_languages.csv","programminglanguages.csv")
# We can use shell commands like `ls` in Julia by preceding them with a semicolon.
;ls
# And there's the *.csv file we downloaded!
#
# Now let's load it into Julia
# We'll use the `DelimitedFiles` standard library package and its `readdlm()` function as shown
# below.
#
# (Today, the [CSV.jl](https://juliadata.github.io/CSV.jl/stable/) package is the recommended way to load CSVs in Julia. We can install it via `Pkg.add()`, and load .csv files using `CSV.read()`. This tutorial hasn't been updated to use CSV.jl yet.)
# +
# using Pkg
# Pkg.add("CSV") # for CSV.read()
# Pkg.add("DelimitedFiles") # for readdlm
# using CSV
# P = CSV.read("programminglanguages.csv",header=true)
# or
using DelimitedFiles # Standard library in Base
# -
# By default, `readdlm` will fill an array with the data stored in the input .csv file. If we set the keyword argument `header` to `true`, we'll get a second output array for just the headers.
P,H = readdlm("programminglanguages.csv", ',', header=true)
P # stores the dataset
H # stores the header names
# Here we write our first small function. <br>
# Now you can answer questions such as, "when was language X created?"
function language_created_year(P,language::String)
loc = findfirst(P[:,2].==language)
return P[loc,1]
end
language_created_year(P,"Julia")
language_created_year(P,"julia")
# As expected, this will not return what you want, but thankfully, string manipulation is really easy in Julia!
function language_created_year_v2(P,language::String)
loc = findfirst(lowercase.(P[:,2]).==lowercase.(language))
return P[loc,1]
end
language_created_year_v2(P,"julia")
# **Reading and writing to files is really easy in Julia.** <br>
#
# You can use different delimiters with the function `readdlm`, from the `DelimitedFiles` package. <br>
#
# To write to files, we can use `writedlm`. <br>
#
# Let's write this same data to a file with a different delimiter.
writedlm("programming_languages_data.txt", P, '-')
# We can now check that this worked using a shell command to glance at the file,
;head -10 programming_languages_data.txt
# and also check that we can use `readdlm` to read our new text file correctly.
P_new_delim = readdlm("programming_languages_data.txt", '-');
P == P_new_delim
# ### Dictionaries
# Let's try to store the above data in a dictionary format!
#
# First, let's initialize an empty dictionary
dict = Dict{Integer,Vector{String}}()
# Here we told Julia that we want `dict` to only accept integers as keys and vectors of strings as values.
#
# However, we could have initialized an empty dictionary without providing this information (depending on our application).
dict2 = Dict()
# This dictionary takes keys and values of any type!
#
# Now, let's populate the dictionary with years as keys and vectors that hold all the programming languages created in each year as their values.
for i = 1:size(P,1)
year,lang = P[i,:]
if year in keys(dict)
dict[year] = push!(dict[year],lang)
else
dict[year] = [lang]
end
end
# Now you can pick whichever year you want and find what programming languages were invented in that year
dict[2003]
# ### DataFrames!
# *Shout out to R fans!*
# One other way to play around with data in Julia is to use a DataFrame.
#
# This requires loading the `DataFrames` package. Thankfully, this tutorial came with a Project.toml file that specifies exactly which version of DataFrames to install...
# #### Project.toml files
#
# For this tutorial (Julia for Data Science), you may have noticed that there are files in this folder called [`Project.toml`](/edit/introductory-tutorials/broader-topics-and-ecosystem/intro-to-julia-for-data-science/Project.toml) and [`Manifest.toml`](/edit/introductory-tutorials/broader-topics-and-ecosystem/intro-to-julia-for-data-science/Manifest.toml). These are files autogenerated by Julia's package manager, `Pkg`, that describe the _exact set of packages_ installed for a julia project, allowing you to share your work in a perfectly reproducible way.
#
# Jupyter was able to detect those `.toml` files, and so this notebook was automatically started with _this project activated!_ Note that this means any packages you add or remove inside this notebook will only affect this "Julia for Data Science" _project_.
#
# To install all of the package dependencies used in the rest of the tutorial, you only need to run this next cell (commands that start with `]` are package repl commands):
] instantiate
# You can read more about package manager commands, here: https://docs.julialang.org/en/v1/stdlib/Pkg/index.html
#
# **Now back to DataFrames!**
using DataFrames
df = DataFrame(year = P[:,1], language = P[:,2])
# You can access columns by header name, or column index.
#
# In this case, `df[1]` is equivalent to `df.year` or `df[!, :year]`.
#
# Note that if we want to index columns by header name, we precede the header name with a colon. In Julia, this means that the header names are treated as *symbols*.
df.year
# **`DataFrames` provides some handy features when dealing with data**
#
# First, it uses julia's "missing" type.
a = missing
typeof(a)
# Let's see what happens when we try to add a "missing" type to a number.
a + 1
# `DataFrames` provides the `describe` function, which can give you quick statistics about each column in your dataframe
describe(df)
# ### RDatasets
#
# We can use RDatasets to play around with pre-existing datasets
using RDatasets
iris = dataset("datasets", "iris")
# Note that data loaded with `dataset` is stored as a DataFrame. 😃
typeof(iris)
# The summary we get from `describe` on `iris` gives us a lot more information than the summary on `df`!
describe(iris)
# ### More on Missing Values
#
# Julia 1.0 and beyond has native support for `missing` values. (Before Julia 1.0, this was done via the DataArrays.jl package.)
# More information on using arrays with missing values can be found
# [in the Julia documentation](https://docs.julialang.org/en/v1/manual/missing/#Arrays-With-Missing-Values-1).
foods = ["apple", "cucumber", "tomato", "banana"]
calories = [missing,47,22,105]
typeof(calories)
using Statistics # julia's standard library for stats
mean(calories)
# Missing values ruin everything! 😑
#
# Luckily we can ignore them with `skipmissing`!
mean(skipmissing(calories))
# Oh WAIT! Detour. How did I get the emoji there?
#
# Try this out:
#
# ```
# \:expressionless: + <TAB>
# ```
😑 = 0 # expressionless
😀 = 1
😞 = -1
# *Back to missing values*
prices = [0.85,1.6,0.8,0.6,]
dataframe_calories = DataFrame(item=foods,calories=calories)
dataframe_prices = DataFrame(item=foods,price=prices)
# We can also `join` two dataframes together
DF = join(dataframe_calories,dataframe_prices,on=:item)
# ### FileIO
using FileIO
julialogo = download("https://avatars0.githubusercontent.com/u/743164?s=200&v=4","julialogo.png")
# Again, let's check that this download worked!
;ls
#
# Next, let's load the Julia logo, stored as a .png file
#
# **NOTE: You may see errors below, that certain Image packages could not be found. If so:**
# - This is because these packages are specific to your OS, so aren't installed by default.
# - Simply run the suggested commands to install the packages, then try again.
# +
# These commands may vary depending on your operating system.
#import Pkg; Pkg.add("QuartzImageIO")
#import Pkg; Pkg.add("ImageMagick")
# -
X1 = load("julialogo.png")
# We see here that Julia stores this logo as an array of colors.
@show typeof(X1);
@show size(X1);
# And if we load the Images package, it will display in Jupyter as an image:
using Images
X1
# ### File types
# In Julia, many file types are supported so you do not have to transfer a file you from another language to a text file before you read it.
#
# *Some packages that achieve this:*
# MAT CSV NPZ JLD FASTAIO
#
#
# Let's try using MAT to write a file that stores a matrix.
using MAT
A = rand(5,5)
matfile = matopen("densematrix.mat", "w")
write(matfile, "A", A)
close(matfile)
# Now try opening densematrix.mat with MATLAB!
newfile = matopen("densematrix.mat")
read(newfile,"A")
names(newfile)
close(newfile)
|
introductory-tutorials/broader-topics-and-ecosystem/intro-to-julia-for-data-science/1. Julia for Data Science - Data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bias in Data
#
# ## Purpose
#
# This project explores the concept of *bias* by examinging how the number and quality of Wikipedia articles about political figures vary among countries.
#
# Several specific questions are addressed:
# - Which countries have the greatest and the least coverage of politicians on Wikipedia compared to their population?
# - which countries have the highest and lowest proportion of high quality articles about politicians?
# - Which regions have the most articles about politicians, relative to their populations?
# - Which regions have teh highest proprtion of high-quality articles about politicians?
#
# Article quality is estimated using a machine learning service called ORES. NOTES ABOUT PACKAGES AND THINGS THAT ARE USED TO CONDUCT ANALYSIS.
# ## Data Ingestion and Cleaning
#
# ### Data Sources
# The data used in this analysis is drawn from two sources:
# - The Wikipedia politicians by country dataset, found on Figshare: https://figshare.com/articles/Untitled_Item/5513449
# - A subset of the world population datasheet published by the Population Reference Bureau
#
# ### Data Cleaning
#
# The Wikipedia *Politicians by Country* dataset contains some pages which are not Wikipedia articles. These pages are filtered out before we conduct our analysis by removing all page names that begin with the string "Template:".
#
# The Population Reference Bureau *World Population Datasheet* contains some rows relating to regional population counts. These are filtered out prior to country-level analyses performed below, but utilized in the final two tables in the Analysis section and in the Reflection section to address coverage and quality by region.
# +
# import needed packages
import pandas as pd
# read the csv files in to Pandas data frames
politicos_full = pd.read_csv("page_data.csv")
pops_regions = pd.read_csv("WPDS_2018_data.csv")
# check that the imports have worked correctly
#print(politicos_full.head())
# remove the no-Wikipedia articles by filtering the politicos data frame to remove instances of the string "Template:"
politicos = politicos_full[~politicos_full.page.str.contains("Template:")]
# check that the filtering step has worked correctly
#print(politicos.head())
# remove the regions from the population data frame by removing rows where the geography col is all caps
# first we make a deep copy of the dataframe because we want a dataframe free of regions, but we also want the region data
pops_countries = pops_regions.copy(deep=True)
# drop regions from the new countries dataframe for the upcoming analysis
pops_countries.drop(pops_countries[pops_countries['Geography'].str.isupper()].index, inplace = True)
# drop countries from the regions dataframe so the two will be completely distinct
#pops_regions = pops_regions[pops_regions['Geography'].str.isupper()]
# check that both dataframes are correct
#print(pops_regions.head())
#print(pops_countries.head())
# -
# ### Quality Predictions
#
# In the following code we use the ORES API to get json files which contain predictions about the quality of individual articles.
#
# ORES documentation: https://ores.wikimedia.org/v3/#!/scoring/get_v3_scores_context
#
# There are six total quality categories. The first two categories (FA and GA) are considered high quality.
#
# FA - Featured article
# GA - Good article
# B - B-class article
# C - C-class article
# Start - Start-class article
# Stub - Stub-class article
#
# The first fuction in the following code get_ores_data is taken from https://github.com/Ironholds/data-512-a2/blob/master/hcds-a2-bias_demo.ipynb and modified only so that it returns the result (rather than simply printing it).
# +
# import needed packages
import requests
import json
# this block of code is taken from https://github.com/Ironholds/data-512-a2/blob/master/hcds-a2-bias_demo.ipynb
# it is modified only so that get_ores_data returns the result response
headers = {'User-Agent' : 'https://github.com/chisquareatops', 'From' : '<EMAIL>'}
def get_ores_data(revision_ids, headers):
# Define the endpoint
endpoint = 'https://ores.wikimedia.org/v3/scores/{project}/?models={model}&revids={revids}'
# Specify the parameters - smushing all the revision IDs together separated by | marks.
# Yes, 'smush' is a technical term, trust me I'm a scientist.
# What do you mean "but people trusting scientists regularly goes horribly wrong" who taught you tha- oh.
params = {'project' : 'enwiki',
'model' : 'wp10',
'revids' : '|'.join(str(x) for x in revision_ids)
}
api_call = requests.get(endpoint.format(**params))
response = api_call.json()
#print(json.dumps(response, indent=4, sort_keys=True
return response
# +
# we need to extract the overall prediction from the above function, which also returns sores for all page types
# make a list of the ids
revids = list(politicos['rev_id'])
# loop through the list of ids in chunks of 100
def get_pred(df, block_size):
start = 0
end = block_size
output_final = list()
while start < len(revids):
revids_temp = revids[start:end]
output_temp = get_ores_data(revids_temp, headers)
for key, item in output_temp['enwiki']['scores'].items():
dict_temp = dict()
dict_temp['rev_id'] = key
if 'error' in item['wp10']:
dict_temp['prediction'] = 'no score'
else:
dict_temp['prediction'] = item['wp10']['score']['prediction']
output_final.append(dict_temp)
start += 100
end += 100
scores = pd.DataFrame(output_final)
return scores
# +
# call the above functions to get the predictions for our data frame; divide the articles into blocks of 100
politicos_preds = get_pred(politicos, 100)
# check that the above step worked correctly
#print(politicos_preds.head())
# save the articles with no score to a csv and then remove them from the data frame
pred[pred.prediction == 'no score'][['rev_id']].to_csv('wp_wpds_articles-no_score.csv')
politicos_preds = politicos_preds[~politicos_preds.prediction.str.contains("no score")]
# -
# ### Merge and Output Data
#
# In the following code we merge our data so that the predictions we are interested in are associated with the individual articles in our data set. We then export a csv of this combined data.
# +
# make copies just in case before merging
politicos_final = politicos.copy(deep=True)
politicos_preds_final = politicos_preds.copy(deep=True)
pops_countries_final = pops_countries.copy(deep=True)
# merge the politcal article data and the quality predictions on the rev_id/revision_id cols
politicos_preds_final = politicos_preds_final.astype({'rev_id': 'int64'})
combined_final = politicos_final.merge(politicos_preds_final, how='right', left_on='rev_id', right_on='rev_id')
# merge the new data frame with the population data on the country/Geography cols
combined_final = combined_final.merge(pops_countries_final, how='right', left_on='country', right_on='Geography')
# check that the above step worked
#print(combined_final.head())
# rename the cols to comply with assignment
combined_final.rename(columns={'page':'article_name','Population mid-2018 (millions)':'population','rev_id':'revision_id','prediction':'article_quality'}, inplace=True)
# save the rows that have no match on the country field to a csv, then drop from the final data frame
combined_final[combined_final.Geography.isnull()].to_csv('wp_wpds_countries-no_match.csv')
combined_final.dropna(inplace=True)
# remove Geography col to comply with assigment (now that rows with no country match are gone)
combined_final.drop('Geography', axis=1)
# check that the above step worked
print(combined_final.head())
# -
# change some data types so the following analysis will work
combined_final['population'] = combined_final['population'].str.replace(',', '')
combined_final = combined_final.astype({'population':'float'})
# # Analysis
#
# In this section we create the following six individual tables:
#
# - Top 10 countries by coverage: 10 highest-ranked countries in terms of number of politician articles as a proportion of country population
# - Bottom 10 countries by coverage: 10 lowest-ranked countries in terms of number of politician articles as a proportion of country population
# - Top 10 countries by relative quality: 10 highest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
# - Bottom 10 countries by relative quality: 10 lowest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
# - Geographic regions by coverage: Ranking of geographic regions (in descending order) in terms of the total count of politician articles from countries in each region as a proportion of total regional population
# - Geographic regions by coverage: Ranking of geographic regions (in descending order) in terms of the relative proportion of politician articles from countries in each region that are of GA and FA-quality
# +
# select for high quality articles by keeping only the FA and GA designations in the article_quality field
combined_final_2 = combined_final.copy(deep=True)
hq_articles = combined_final_2.loc[combined_final_2['article_quality'].isin(['FA','GA'])]
# count total number of high quality articles in each country using group by
hq_articles_country = hq_articles.groupby('country').count()['article_name']
# make this result into a dataframe with appropriate cols so we can bring back population data and report the proportion
hq_articles_country_df = hq_articles_country.to_frame()
hq_articles_country_df['country'] = hq_articles_country_df.index
hq_articles_country_df.reset_index(drop=True, inplace=True)
hq_articles_country_df = hq_articles_country_df.merge(pops_countries_final, how='inner', left_on='country', right_on='Geography')
# find the actual proprtion: divide number of high quality articles by total population
hq_articles_country_df = hq_articles_country_df.astype({'article_name': 'float'})
hq_articles_country_df['Population mid-2018 (millions)'] = hq_articles_country_df['Population mid-2018 (millions)'].str.replace(',', '')
hq_articles_country_df = hq_articles_country_df.astype({'Population mid-2018 (millions)': 'float'})
hq_articles_country_df['article_proportion'] = hq_articles_country_df['article_name'] / (hq_articles_country_df['Population mid-2018 (millions)'] * 1000000)
# -
# #### Top 10 countries by coverage: 10 highest-ranked countries in terms of number of politician articles as a proportion of country population
# sort by proportion and display a table of the top 10
articles_over_pop = hq_articles_country_df[['country','article_proportion']]
articles_over_pop = articles_over_pop.sort_values('article_proportion', ascending=False)
print(articles_over_pop.head(10))
# #### Bottom 10 countries by coverage: 10 highest-ranked countries in terms of number of politician articles as a proportion of country population
#Bottom 10 countries by coverage
articles_over_pop = articles_over_pop.sort_values('article_proportion', ascending=True)
print(articles_over_pop.head(10))
# +
#Top 10 countries by relative quality
# group the same way we did in previous steps, but this time using all articles
all_articles = combined_final.loc[combined_final['article_quality'].isin(['FA','GA','B','C','Start','Stub'])]
# count total number of high quality articles in each country using group by
all_articles_country = all_articles.groupby('country').count()['article_name']
# make a dataframe with this total number of articles per country so it can be merged with dataframe from prev step
all_articles_country_df = all_articles_country.to_frame()
all_articles_country_df['country'] = all_articles_country_df.index
all_articles_country_df.reset_index(drop=True, inplace=True)
all_articles_country_df = all_articles_country_df.astype({'article_name': 'float'})
all_articles_country_df.rename(columns = {'article_name':'total_articles'}, inplace = True)
all_articles_country_df = all_articles_country_df.merge(hq_articles_country_df, how='right', left_on='country', right_on='country')
# -
# #### Top 10 countries by relative quality: 10 highest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
# find the proprtion: divide number of high quality articles by total articles
all_articles_country_df['quality_to_total'] = all_articles_country_df['article_name'] / all_articles_country_df['total_articles']
hqarticles_over_total = all_articles_country_df[['country','quality_to_total']]
hqarticles_over_total = hqarticles_over_total.sort_values('quality_to_total', ascending=False)
print(hqarticles_over_total.head(10))
# #### Bottom 10 countries by relative quality: 10 highest-ranked countries in terms of the relative proportion of politician articles that are of GA and FA-quality
hqarticles_over_total = hqarticles_over_total.sort_values('quality_to_total', ascending=True)
print(hqarticles_over_total.head(10))
# +
# regions by coverage
# The only data source we have that connects countries to regions is the original WPDS_2018_data.csv data (now pops_regions)
# countries in this data belong to the region that precedes them in the file, so we need to loop through it.
# create an empty dict to hold country/region pairs as we find them
region_dict = {}
# loop through the original data we preserved (as a list) to identify countries vs. regions, then store pairs
for value in pops_regions['Geography'].tolist():
# if the current row is a region, make it the current region (the first row is a region)
if value.isupper():
region = value
# if the current row is a country, add a new country/region pair to the dict
else:
region_dict.update({value:region})
# use a lambda to make a new col in the most recent dataframe and use the dict to insert a region value
all_articles_country_df['region'] = all_articles_country_df['country'].apply(lambda x: region_dict[x])
# test that the above step worked correctly
#print(all_articles_country_df.head())
# -
# #### Geographic regions by coverage: Ranking of geographic regions (in descending order) in terms of the total count of politician articles from countries in each region as a proportion of total regional population
# +
# add up the total number of articles in each region using group by
all_articles_region = all_articles_country_df.groupby('region').sum()['total_articles']
# turn that result back into a data frame
all_articles_region_df = all_articles_region.to_frame()
all_articles_country_df.reset_index(drop=True, inplace=True)
# add up the total number of articles in each region using group by
pop_region = all_articles_country_df.groupby('region').sum()['Population mid-2018 (millions)']
# turn that result back into a data frame
pop_region_df = pop_region.to_frame()
pop_region_df.reset_index(inplace=True)
#all_articles_region_df = all_articles_region_df.sort_values('article_proportion', ascending=False)
all_articles_over_pop = all_articles_region_df.merge(pop_region_df, how='right', left_on='region', right_on='region')
all_articles_over_pop['total_articles_over_pop'] = all_articles_over_pop['total_articles']/all_articles_over_pop['Population mid-2018 (millions)']
all_articles_over_pop = all_articles_over_pop[['region', 'total_articles_over_pop']]
all_articles_over_pop = all_articles_over_pop.sort_values('total_articles_over_pop', ascending=False)
print(all_articles_over_pop)
# -
# #### Geographic regions by coverage: Ranking of geographic regions (in descending order) in terms of the relative proportion of politician articles from countries in each region that are of GA and FA-quality
# +
# add up the total number of articles in each region using group by
all_articles_region = all_articles_country_df.groupby('region').sum()['total_articles']
# turn that result back into a data frame
all_articles_region_df = all_articles_region.to_frame()
all_articles_country_df.reset_index(drop=True, inplace=True)
# add up the total number of high quality articles in each region using group by
hq_region = all_articles_country_df.groupby('region').sum()['article_name']
# turn that result back into a data frame
hq_region_df = hq_region.to_frame()
hq_region_df.reset_index(inplace=True)
hq_over_all_articles_df = all_articles_region_df.merge(hq_region_df, how='right', left_on='region', right_on='region')
hq_over_all_articles_df['hq_over_all_articles'] = hq_over_all_articles_df['article_name']/hq_over_all_articles_df['total_articles']
hq_over_all_articles_df = hq_over_all_articles_df[['region', 'hq_over_all_articles']]
hq_over_all_articles_df = hq_over_all_articles_df.sort_values('hq_over_all_articles', ascending=False)
print(hq_over_all_articles_df)
# -
# # Reflection
#
# Potentially the most significant source of potential bias is this analysis is the ORES scores themselves and the way in which they are generated. Unfortunately I can't speak to this as I don't know how the detailed algorithms behind this scores. I can only state that I cannot be sure whether these scores thoroughly account for possible cultural differences when evaluating the 'quality' of an article.
#
# Another potential problem with the data is different political structures in different countries. Governments can vary widely in size and different branches of government vary wiely in how much power they weild, how long they are in office, and other important measures both within and between countries. Therefore, it is possible that one country may have proportionaly more political offices which warrant (and require) lengthy or high quality explanation than another country.
#
# The tables generated here suggest possible hypothesis about internet access: it's possible that countries where a higher percentage of the population has internet access will tend to have more total articles and proportionally more high-quality articles. This would have to be tested by brining in additional data. It would also be interesting to combine the above data set with national GDP data to investigate any possible correlations with wealth at the national or regional level.
|
hcds-a2-bias.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 11장 - 심층 신경망 훈련
# ## 11.1 그래디언트 소실과 폭주 문제
# #### 11.1.1 세이비어 초기화와 He 초기화
# #### 11.1.2 수렴하지 않는 활성화 함수
# #### 11.1.3 배치 정규화
# #### 11.1.4 그래디언트 클리핑
#
# ## 11.2 미리 훈련된층 재사용하기
# #### 11.2.1 텐서플로 모델 재사용하기
# #### 11.2.2 다른 프레임워크의 모델 재사용하기
# #### 11.2.3 신경망의 하위층 학습에서 제외하기
# #### 11.2.4 동결된 층 캐싱하기
# #### 11.2.5 상위층을 변경, 삭제, 대체하기
# #### 11.2.6 모델 저장소
# #### 11.2.7 비지도 사전훈련
# #### 11.2.8 보조 작업으로 사전훈련
# ## 11.3 고속 옵티마이저
# #### 11.3.1 모멘텀 최적화
# #### 11.3.2 네스테로프 가속 경사
# #### 11.3.3 AdaGrad
# #### 11.3.4 RMSProp
# #### 11.3.5 Adam 최적화
# #### 11.3.6 학습률 스케줄링
# ## 11.4 과대적합을 피하기 위한 규제 방법
# #### 11.4.1 조기종료
# #### 11.4.2 l1과 l2 규제
# #### 11.4.3 드롭아웃
# #### 11.4.4 맥스-노름 규제
# #### 11.4.5 데이터 증식
# ## 11.5 실용적 가이드라인
# ## 11.6 연습문제
# ----
# +
# # %load_ext watermark
# # %watermark -v -p numpy,sklearn,scipy,matplotlib,tensorflow
# +
# 파이썬 2와 파이썬 3 지원
from __future__ import division, print_function, unicode_literals
# 공통
import numpy as np
import os
# 일관된 출력을 위해 유사난수 초기화
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# 맷플롯립 설정
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# 한글출력
plt.rcParams['font.family'] = 'NanumBarunGothic'
plt.rcParams['axes.unicode_minus'] = False
# 그림을 저장할 폴더
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
# -
import tensorflow as tf
#
# ----
# ## 11.1 그래디언트 소실과 폭주 문제
#
# * 그래디언트 소실 : 출력층에서 입력층으로 역전파 알고리즘이 진행됨에 따라 그래디언트가 점점 작아져서 하위층(입력층)에 가까운 층들은 연결가중치가 거의 갱신되지 않는 것
# 
# * 그래디언트 폭주 : 반대로 그래디언트가 점점 커져서 여러개의 층이 비정상적으로 큰 가중치로 갱신(발산)되는 것
# * 2010년 <NAME>과 <NAME>의 "Understandimg the Difficulty of Training Deep Feedforward Neural Networks" 논문
# - 로지스틱 시그모이드 활성화 함수 + 정규분포 가중치 초기화 사용 -> 각 층의 출력에서의 분산이 입력의 분산보다 더 커짐
# - 신경망의 위쪽(출력층)으로 갈수록 층을 지날때마다 분산이 계속 커져 높은층(출력층)에서는 활성화 함수가 0 또는 1로 수렴하게 됨
# - 시그모이드의 도함수는 f(1-f)이므로 함수의 값이 0 또는 1에 가까우면 도함수의 결과가 매우작아지고 층이 거듭될 수록 더 작아짐
def logit(z):
return 1 / (1 + np.exp(-z))
# +
z = np.linspace(-5, 5, 200)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [1, 1], 'k--')
plt.plot([0, 0], [-0.2, 1.2], 'k-')
plt.plot([-5, 5], [-3/4, 7/4], 'g--')
plt.plot(z, logit(z), "b-", linewidth=2)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('수렴', xytext=(3.5, 0.7), xy=(5, 1), arrowprops=props, fontsize=14, ha="center")
plt.annotate('수렴', xytext=(-3.5, 0.3), xy=(-5, 0), arrowprops=props, fontsize=14, ha="center")
plt.annotate('선형', xytext=(2, 0.2), xy=(0, 0.5), arrowprops=props, fontsize=14, ha="center")
plt.grid(True)
plt.title("로지스틱 활성화 함수", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("sigmoid_saturation_plot")
plt.show()
# -
# ----
# ### 11.1.1 세이비어 초기화와 He 초기화
# * 그래디언트 소실/발산을 완화하려면
# - 적절한 신호가 흐르기 위해서는 각 층의 출력에 대한 분산이 입력에 대한 분산과 같아야 한다
# - 역방향에서 층을 통과하기 전과 후의 그래디언트 분산이 동일해야 한다
# 
# * <NAME>과 <NAME>의 가중치 초기화 방법
# - 출력의 분산은 입력 연결 수에 비례하여 늘어나므로 가중치의 분산을 1/n_inputs로 하여 입출력의 분산을 맞춘다
# - 역방향일 경우에는 입출력이 반대로 바뀌어 분산이 1/n_outputs가 되어야 한다
# - 두 방향에 대해 절충안으로 2/(n_inputs + n_outputs)를 가중치의 분산으로 한다
# - tf.contrib.layers에 구현됨
# - (세이비어 초기화 전략을 사용하면 훈련 속도를 상당히 높일 수 있다!)
# 
# tf의 xavier_initializer 모듈을 이용한 예시
W = tf.get_variable("W", shape=[784, 256], initializer=tf.contrib.layers.xavier_initializer())
# * He 초기화
# - ReLU 활성화 함수 및 ELU 등의 변종들을 위한 초기화 전략
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
# -
# tf.layers.dense는 기본적으로 세이비어 초기화함수를 사용하므로 아래와 같이 variance_scaling_initializer를 지정하여 He 초기화를 사용할 수 있다
he_init = tf.variance_scaling_initializer()
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, kernel_initializer=he_init, name="hidden1")
# ----
# ### 11.1.2 수렴하지 않는 활성화 함수
# * 시그모이드 활성화 함수에 대한 오해
# - 생물학적 뉴런과 비슷하므로 최선의 선택일 것이라는 오해
# - 실제로는 ReLU가 더 잘 작동함
#
# * 죽은 ReLU (dying ReLU) 문제
# - 훈련하는 동안 일부 뉴런이 0 이외의 값을 출력하지 않음
# - 훈련 도중 가중치의 합이 음수가 되면 그 다음부턴 계속 0만 출력하게 됨
#
# * Leaky ReLU
# - 하이퍼파라미터 alpha는 함수가 새는(leaky) 정도를 의미
# - 새는 정도는 x < 0 일 때의 기울기이며 일반적으로 0.01로 설정함
# - 작은 기울기가 leakyrelu를 절대 죽지 않게 만든다
# - 아래 기술할 다른 ReLU 변현들에 비해 속도가 빠르다
# - tf.nn.leaky_relu, tf.keras.layers.LeakyReLU 로 구현됨
# 
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
# +
plt.plot(z, leaky_relu(z, 0.05), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([0, 0], [-0.5, 4.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('통과', xytext=(-3.5, 0.5), xy=(-5, -0.2), arrowprops=props, fontsize=14, ha="center")
plt.title("Leaky ReLU 활성화 함수", fontsize=14)
plt.axis([-5, 5, -0.5, 4.2])
save_fig("leaky_relu_plot")
plt.show()
# -
# * RReLU (randomized leaky ReLU)
# - 훈련 동안 주어진 범위 내에서 alpha를 무작위로 선택하고, 테스트 시에는 평균을 사용하는 방법
# - 신경망이 과대적합시 사용하기 좋음
#
# * PReLU(parametric leaky ReLU)
# - alpha가 훈련 동안 학습되는 ReLU
# - alpha가 하이퍼파라미터가 아니라 역전파에 의해 갱신되므로, 훈련세트가 클 경우에 사용하기 좋음
# * ELU (exponential linear unit)
# - 다른 모든 ReLU의 성능을 앞지르는 활성화 함수
# - alpha값은 x가 음수일 때 수렴할 값을 설정
# - x < 0일 떄, 음숫값이 들어오므로 평균 출력이 0에 더 가까워짐 -> 그래디언트 소실 완화
# - x < 0이어도 그래디언트가 0이 아니므로 죽은 ReLU 문제가 생기지 않음
# - alpha가 1이면 x = 0에서 연속적이므로 모든면에서 매끄러워서 경사하강법의 속도를 높여줌
# - 단, ReLU나 ReLU의 변종 함수들에 비해 계산이 느리다
# - tf.nn.elu, tf.keras.layers.ELU로 구현됨
# 
# 
def elu(z, alpha=1):
return np.where(z < 0, alpha * (np.exp(z) - 1), z)
# +
plt.plot(z, elu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1, -1], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"ELU 활성화 함수 ($\alpha=1$)", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("elu_plot")
plt.show()
# -
# 텐서플로에서 ELU를 구현하는 것은 간단합니다. 층을 구성할 때 활성화 함수에 지정하기만 하면 됩니다:
# +
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
# -
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.elu, name="hidden1")
# ### SELU
# 이 활성화 함수는 <NAME>, <NAME>, <NAME>가 2017년에 쓴 [논문](https://arxiv.org/pdf/1706.02515.pdf)에서 소개되었습니다(나중에 책에 추가하겠습니다). 훈련할 때 SELU 활성화 함수를 사용한 완전 연결 신경망은 스스로 정규화를 합니다. 각 층의 출력은 훈련하는 동안 같은 평균과 분산을 유지하려는 경향이 있어 그래디언트 소실과 폭주 문제를 해결합니다. 이 활성화 함수는 심층 신경망에서 다른 활성화 함수보다 뛰어난 성능을 내므로 꼭 이 함수를 시도해봐야 합니다.
def selu(z,
scale=1.0507009873554804934193349852946,
alpha=1.6732632423543772848170429916717):
return scale * elu(z, alpha)
# +
plt.plot(z, selu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1.758, -1.758], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"SELU 활성화 함수", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("selu_plot")
plt.show()
# -
# 기본적으로 SELU 하이퍼파라미터(`scale`과 `alpha`)는 평균이 0, 표준 편차가 1에 가깝게 유지되도록 조정합니다(입력도 평균이 0, 표준 편차가 1로 표준화되었다고 가정합니다). 이 활성화 함수를 사용하면 100층으로 된 심층 신경망도 그래디언트 소실/폭주 문제없이 모든 층에서 대략 평균이 0이고 표준 편차가 1을 유지합니다:
np.random.seed(42)
Z = np.random.normal(size=(500, 100))
for layer in range(100):
W = np.random.normal(size=(100, 100), scale=np.sqrt(1/100))
Z = selu(np.dot(Z, W))
means = np.mean(Z, axis=1)
stds = np.std(Z, axis=1)
if layer % 10 == 0:
print("층 {}: {:.2f} < 평균 < {:.2f}, {:.2f} < 표준 편차 < {:.2f}".format(
layer, means.min(), means.max(), stds.min(), stds.max()))
# 텐서플로 1.4 버전에 `tf.nn.selu()` 함수가 추가되었습니다. 이전 버전을 사용할 때는 다음 구현을 사용합니다:
def selu(z,
scale=1.0507009873554804934193349852946,
alpha=1.6732632423543772848170429916717):
return scale * tf.where(z >= 0.0, z, alpha * tf.nn.elu(z))
# 하지만 SELU 활성화 함수는 일반적인 드롭아웃과 함께 사용할 수 없습니다(드롭아웃은 SELU 활성화 함수의 자동 정규화 기능을 없애버립니다). 다행히 같은 논문에 실린 알파 드롭아웃(Alpha Dropout)을 사용할 수 있습니다. 텐서플로 1.4에 `tf.contrib.nn.alpha_dropout()`이 추가되었습니다(Linz 대학교 생물정보학 연구소(Institute of Bioinformatics)의 <NAME>가 만든 [구현](https://github.com/bioinf-jku/SNNs/blob/master/selu.py)을 확인해 보세요).
# SELU 활성화 함수를 사용한 신경망을 만들어 MNIST 문제를 풀어 보겠습니다:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=selu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=selu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 40
batch_size = 50
# -
# 이제 훈련할 차례입니다. 입력을 평균 0, 표준 편차 1로 스케일 조정해야 합니다:
# +
means = X_train.mean(axis=0, keepdims=True)
stds = X_train.std(axis=0, keepdims=True) + 1e-10
X_val_scaled = (X_valid - means) / stds
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
X_batch_scaled = (X_batch - means) / stds
sess.run(training_op, feed_dict={X: X_batch_scaled, y: y_batch})
if epoch % 5 == 0:
acc_batch = accuracy.eval(feed_dict={X: X_batch_scaled, y: y_batch})
acc_valid = accuracy.eval(feed_dict={X: X_val_scaled, y: y_valid})
print(epoch, "배치 데이터 정확도:", acc_batch, "검증 세트 정확도:", acc_valid)
save_path = saver.save(sess, "./my_model_final_selu.ckpt")
# -
# ----
# ### 11.1.3 배치 정규화
#
# * ELU 등 ReLU 응용 함수와 He 초기화를 사용하면 훈련 초기단계에서 그래디언트 소실/폭주를 크게 감소시킬 수 있지만, 훈련 중 다시 발생할 수 있다.
#
# * 내부 공변량 변화 문제
# - 훈련하는 동안 이전 층의 파라미터가 변함에 따라 각 층에 들어오는 입력의 분포가 변하는 문제
# * 2015년 <NAME>, <NAME>가 배치 정규화 (Batch Normalization, BN) 기법을 제안함
# * 배치 정규화
# - 각 층에서 데이터의 분포를 정규화하는 기법
# - 입력 데이터의 평균이 0이 되도록 정규화 및 스케일링하고 이동시킴
# - 테스트 시에는 전체 훈련 세트의 평균과 표준편차를 대신 사용함
# - 배치 정규화가 적용된 모든 층은 gamma(스케일), beta(이동), mu(평균), sigma(표준편차) 총 4개의 파라미터를 학습하게 된다
# 
# * 배치 정규화의 성능
# - 로지스틱 등 수렴되는 활성화 함수를 사용해도 소실 문제가 크게 감소됨
# - 네트워크가 가중치 초기화에 덜 민감해짐
# - 학습률을 크게 줄 수 있어 학습 속도가 개선됨
# - 배치 정규화가 규제 역할을 하므로 다른 규제 기법의 필요성이 줄어듦
# - "사람의 판단 능력을 뛰어 넘는 4.9%의 top-5 검증 에러를 달성함"
# * 배치 정규화의 단점
# - 모델의 복잡도 증가
# - 실행 시간이 오래 걸림. 층마다 추가되는 계산으로 인해 신경망의 예측이 느려짐 -> 예측이 빨라야 하는 경우에 사용하기 어려움
# * 텐서플로우 모듈
# - tf.nn.batch_normalization : 평균, 표준편차를 직접 계산해 입력해야하며, 스케일 조정과 이동을 위한 파라미터를 생성해야 함
# - tf.layers.batch_normalization : 모든 일을 대신 해주는 layer
# - tf.keras.layers.BatchNormalization : tf.keras 버전
# 각 은닉층의 활성화 함수 전에 배치 정규화를 추가하기 위해 ELU 활성화 함수를 배치 정규화 층 이후에 수동으로 적용하겠습니다.
#
# 노트: `tf.layers.dense()` 함수가 (책에서 사용하는) `tf.contrib.layers.arg_scope()`와 호환되지 않기 때문에 대신 파이썬의 `functools.partial()` 함수를 사용합니다. 이를 사용해 `tf.layers.dense()`에 필요한 매개변수가 자동으로 설정되도록 `my_dense_layer()`를 만듭니다(그렇지 않으면 `my_dense_layer()`를 호출할 때마다 덮어씌여질 것입니다). 다른 코드는 이전과 비슷합니다.
# +
reset_graph()
import tensorflow as tf
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
training = tf.placeholder_with_default(False, shape=(), name='training')
hidden1 = tf.layers.dense(X, n_hidden1, name="hidden1")
bn1 = tf.layers.batch_normalization(hidden1, training=training, momentum=0.9) # momentum 값은 이동평균 계산에 사용됨
bn1_act = tf.nn.elu(bn1)
hidden2 = tf.layers.dense(bn1_act, n_hidden2, name="hidden2")
bn2 = tf.layers.batch_normalization(hidden2, training=training, momentum=0.9)
bn2_act = tf.nn.elu(bn2)
logits_before_bn = tf.layers.dense(bn2_act, n_outputs, name="outputs")
logits = tf.layers.batch_normalization(logits_before_bn, training=training,
momentum=0.9)
# -
# 같은 매개변수를 계속 반복해서 쓰지 않도록 파이썬의 `partial()` 함수를 사용합니다:
# +
reset_graph()
learning_rate = 0.01
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
training = tf.placeholder_with_default(False, shape=(), name='training')
# +
from functools import partial
my_batch_norm_layer = partial(tf.layers.batch_normalization,
training=training, momentum=0.9)
hidden1 = tf.layers.dense(X, n_hidden1, name="hidden1")
bn1 = my_batch_norm_layer(hidden1)
bn1_act = tf.nn.elu(bn1)
hidden2 = tf.layers.dense(bn1_act, n_hidden2, name="hidden2")
bn2 = my_batch_norm_layer(hidden2)
bn2_act = tf.nn.elu(bn2)
logits_before_bn = tf.layers.dense(bn2_act, n_outputs, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
# -
# 각 층에 ELU 활성화 함수와 배치 정규화를 사용하여 MNIST를 위한 신경망을 만듭니다:
# +
reset_graph()
batch_norm_momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
training = tf.placeholder_with_default(False, shape=(), name='training')
with tf.name_scope("dnn"):
he_init = tf.variance_scaling_initializer()
my_batch_norm_layer = partial(
tf.layers.batch_normalization,
training=training,
momentum=batch_norm_momentum)
my_dense_layer = partial(
tf.layers.dense,
kernel_initializer=he_init)
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
bn1 = tf.nn.elu(my_batch_norm_layer(hidden1))
hidden2 = my_dense_layer(bn1, n_hidden2, name="hidden2")
bn2 = tf.nn.elu(my_batch_norm_layer(hidden2))
logits_before_bn = my_dense_layer(bn2, n_outputs, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
# 노트: 배치 정규화를 위해 별도의 업데이트 연산을 실행해 주어야 합니다(`sess.run([training_op, extra_update_ops],...`).
n_epochs = 20
batch_size = 200
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train = X_train.astype(np.float32).reshape(-1, 28*28) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, 28*28) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_valid, X_train = X_train[:5000], X_train[5000:]
y_valid, y_train = y_train[:5000], y_train[5000:]
def shuffle_batch(X, y, batch_size):
rnd_idx = np.random.permutation(len(X))
n_batches = len(X) // batch_size
for batch_idx in np.array_split(rnd_idx, n_batches):
X_batch, y_batch = X[batch_idx], y[batch_idx]
yield X_batch, y_batch
# +
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run([training_op, extra_update_ops],
feed_dict={training: True, X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# 어!? MNIST 정확도가 좋지 않네요. 물론 훈련을 더 오래하면 정확도가 높아지겠지만 이런 얕은 신경망에서는 배치 정규화와 ELU가 큰 효과를 내지 못합니다. 대부분 심층 신경망에서 빛을 발합니다.
# 업데이트 연산에 의존하는 훈련 연산을 만들 수도 있습니다:
#
# ```python
# with tf.name_scope("train"):
# optimizer = tf.train.GradientDescentOptimizer(learning_rate)
# extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
# with tf.control_dependencies(extra_update_ops):
# training_op = optimizer.minimize(loss)
# ```
#
# 이렇게 하면 훈련할 때 `training_op`만 평가하면 텐서플로가 업데이트 연산도 자동으로 실행할 것입니다:
#
# ```python
# sess.run(training_op, feed_dict={training: True, X: X_batch, y: y_batch})
# ```
# 한가지 더, 훈련될 변수 개수가 전체 전역 변수 개수보다 적습니다. 이동 평균을 위한 변수는 훈련되는 변수가 아니기 때문입니다. 미리 학습한 신경망을 재사용할 경우(아래 참조) 이런 훈련되지 않는 변수를 놓쳐서는 안됩니다.
[v.name for v in tf.trainable_variables()]
[v.name for v in tf.global_variables()]
# ----
# #### 11.1.4 그래디언트 클리핑
#
# * 그래디언트 클리핑 (Gradient Clipping)
# - 역전파도리 때 일정 임계값을 넘어서지 못하게 그래디언트를 단순히 잘라내는 방법
# - 순환신경망(14장)에서 널리 사용되며 보통은 BN이 우선시된다
# - Optimizer의 minimize() 함수는 그래디언트 계산과 적용을 모두 수행하므로 compute_gradients(), clip_by_value(), apply_gradients() 세 개 함수로 분할하여 적용해야 한다
# MNIST를 위한 간단한 신경망을 만들고 그래디언트 클리핑을 적용해 보겠습니다. 시작 부분은 이전과 동일합니다(학습한 모델을 재사용하는 예를 만들기 위해 몇 개의 층을 더 추가했습니다. 아래 참조):
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_hidden3 = 50
n_hidden4 = 50
n_hidden5 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3")
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4")
hidden5 = tf.layers.dense(hidden4, n_hidden5, activation=tf.nn.relu, name="hidden5")
logits = tf.layers.dense(hidden5, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
# -
learning_rate = 0.01
# 이제 그래디언트 클리핑을 적용합니다. 먼저 그래디언트를 구한 다음 `clip_by_value()` 함수를 사용해 클리핑하고 적용합니다:
# +
threshold = 1.0
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var) # 그래디언트를 -1.0 ~ 1.0 사이로 클리핑해서 적용한다
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
# -
grads_and_vars[0][0]
grads_and_vars[0][1]
# 나머지는 이전과 동일합니다:
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# ----
# ## 11.2 미리 훈련된 층 재사용하기
# * 전이 학습 (transfer learning)
# - 큰 규모의 DNN을 처음부터 새로 훈련시키는 것은 좋지 못함
# - 비슷한 유형의 문제를 처리한 신경망을 찾아 하위층을 재사용하는 것이 훈련속도도 빠르고 필요한 학습 데이터도 적음
# 
# ----
# ### 11.2.1 텐서플로 모델 재사용하기
# * 저장된 모델 로드하여 새로운 데이터셋으로 학습하기
#
# 먼저 그래프 구조를 로드해야 합니다. `import_meta_graph()` 함수가 그래프 연산들을 로드하여 기본 그래프에 적재하고 모델의 상태를 복원할 수 있도록 `Saver` 객체를 반환합니다. 기본적으로 `Saver` 객체는 `.meta` 확장자를 가진 파일에 그래프 구조를 저장하므로 이 파일을 로드해야 합니다:
reset_graph()
saver = tf.train.import_meta_graph("./my_model_final.ckpt.meta")
# 다음으로 훈련해야 할 모든 연산을 가져와야 합니다. 그래프 구조를 모를 때는 모든 연산을 출력해 볼 수 있습니다:
for op in tf.get_default_graph().get_operations():
print(op.name)
# 웁스, 연산이 엄청 많네요! 텐서보드로 그래프를 시각화해보는 것이 더 좋을 것 같습니다. 다음 코드는 주피터에서 그래프를 그려줍니다(만약 브라우저에서 보이지 않는다면 `FileWriter`로 그래프를 저장한 다음 텐서보드에서 열어 보세요):
from tensorflow_graph_in_jupyter import show_graph
show_graph(tf.get_default_graph())
# 필요한 연산을 찾았다면 그래프의 `get_operation_by_name()`이나 `get_tensor_by_name()` 메서드를 사용하여 추출할 수 있습니다:
# +
X = tf.get_default_graph().get_tensor_by_name("X:0")
y = tf.get_default_graph().get_tensor_by_name("y:0")
accuracy = tf.get_default_graph().get_tensor_by_name("eval/accuracy:0")
training_op = tf.get_default_graph().get_operation_by_name("GradientDescent")
# -
# 원본 모델을 만들 때 다른 사람이 재사용하기 쉽게 연산에 명확한 이름을 부여하고 문서화를 하는 것이 좋습니다. 또 다른 방법은 처리해야 할 중요한 연산들을 모두 모아 놓은 컬렉션을 만드는 것입니다:
for op in (X, y, accuracy, training_op):
tf.add_to_collection("my_important_ops", op)
# 이렇게 하면 모델을 재사용할 때 다음과 같이 간단하게 쓸 수 있습니다:
X, y, accuracy, training_op = tf.get_collection("my_important_ops")
# 이제 세션을 시작하고 모델을 복원하여 준비된 훈련 데이터로 훈련을 계속할 수 있습니다:
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
# 모델 훈련 계속하기...
# 실제로 테스트를 해보죠!
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# * import_meta_graph() 없이 직접 코드로 그래프 구현
#
# 또 다른 방법으로 원본 그래프를 만든 파이썬 코드에 접근할 수 있다면 `import_meta_graph()` 대신 사용할 수 있습니다:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_hidden3 = 50
n_hidden4 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3")
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4")
hidden5 = tf.layers.dense(hidden4, n_hidden5, activation=tf.nn.relu, name="hidden5")
logits = tf.layers.dense(hidden5, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
threshold = 1.0
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var)
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
# 그 다음 훈련을 계속할 수 있습니다:
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# * import한 그래프에 새로운 layer 추가하기
#
# 일반적으로 하위층만 재사용할 것입니다. `import_meta_graph()`를 사용하면 전체 그래프를 로드하지만 필요하지 않은 부분은 무시하면 됩니다. 이 예에서는 학습된 3번째 층 위에 4번째 은닉층을 새로 추가합니다(원래 4번째 층은 무시됩니다). 새로운 출력층도 추가하고 이 출력으로 손실을 계산하고 이를 최소화하기 위한 새로운 옵티마이저를 만듭니다. 전체 그래프(원본 그래프 전체와 새로운 연산)를 저장할 새로운 `Saver` 객체와 새로운 모든 변수를 초기화할 초기화 연산도 필요합니다:
# +
reset_graph()
n_hidden4 = 20 # 새 층
n_outputs = 10 # 새 층
saver = tf.train.import_meta_graph("./my_model_final.ckpt.meta")
X = tf.get_default_graph().get_tensor_by_name("X:0")
y = tf.get_default_graph().get_tensor_by_name("y:0")
hidden3 = tf.get_default_graph().get_tensor_by_name("dnn/hidden3/Relu:0")
new_hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="new_hidden4")
new_logits = tf.layers.dense(new_hidden4, n_outputs, name="new_outputs")
with tf.name_scope("new_loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=new_logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("new_eval"):
correct = tf.nn.in_top_k(new_logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("new_train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
new_saver = tf.train.Saver()
# -
# 새로운 모델을 훈련시킵니다:
with tf.Session() as sess:
init.run()
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = new_saver.save(sess, "./my_new_model_final.ckpt")
# 원본 모델을 만든 파이썬 코드에 접근할 수 있다면 필요한 부분만 재사용하고 나머지는 버릴 수 있습니다:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1") # 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2") # 재사용
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3") # 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# -
# 그러나 이전에 학습된 모델을 복원하기 위해 (복원할 변수 리스트를 전달합니다. 그렇지 않으면 그래프와 맞지 않는다고 에러를 낼 것입니다) `Saver` 객체를 하나 만들고 훈련이 끝난 후 새로운 모델을 저장하기 위해 또 다른 `Saver` 객체를 만들어야 합니다:
# +
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs): # 책에는 없음
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size): # 책에는 없음
sess.run(training_op, feed_dict={X: X_batch, y: y_batch}) # 책에는 없음
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# -
# ### 11.2.2 다른 프레임워크의 모델 재사용하기
# 이 예에서는 재사용하려는 각 변수에 대해 변수 초기화 할당 연산을 찾고, 초기화 될 값에 해당하는 두 번째 입력 핸들을 구합니다. 초기화가 실행될 때 여기에 `feed_dict` 매개변수를 사용하여 초깃값 대신 원하는 값을 주입합니다:
# +
reset_graph()
n_inputs = 2
n_hidden1 = 3
# +
original_w = [[1., 2., 3.], [4., 5., 6.]] # 다른 프레임워크로부터 가중치를 로드
original_b = [7., 8., 9.] # 다른 프레임워크로부터 편향을 로드
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
# [...] 모델의 나머지 부분을 구성
# hidden1 변수의 할당 노드에 대한 핸들을 구합니다
graph = tf.get_default_graph()
assign_kernel = graph.get_operation_by_name("hidden1/kernel/Assign")
assign_bias = graph.get_operation_by_name("hidden1/bias/Assign")
init_kernel = assign_kernel.inputs[1]
init_bias = assign_bias.inputs[1]
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init, feed_dict={init_kernel: original_w, init_bias: original_b})
# [...] 새 작업에 모델을 훈련시킵니다
print(hidden1.eval(feed_dict={X: [[10.0, 11.0]]})) # 책에는 없음
# -
# 또 다른 방법은 전용 할당 노드와 플레이스홀더를 만든는 것입니다. 이 방법은 더 번거롭고 효율적이지 않지만 하려는 방식이 잘 드러나는 방법입니다:
# +
reset_graph()
n_inputs = 2
n_hidden1 = 3
original_w = [[1., 2., 3.], [4., 5., 6.]] # 다른 프레임워크로부터 가중치를 로드
original_b = [7., 8., 9.] # 다른 프레임워크로부터 편향을 로드
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
# [...] 모델의 나머지를 구성
# hidden1 변수의 할당 노드에 대한 핸들을 구합니다
with tf.variable_scope("", default_name="", reuse=True): # 루트 범위
hidden1_weights = tf.get_variable("hidden1/kernel")
hidden1_biases = tf.get_variable("hidden1/bias")
# 전용 플레이스홀더와 할당 노드를 만듭니다
original_weights = tf.placeholder(tf.float32, shape=(n_inputs, n_hidden1))
original_biases = tf.placeholder(tf.float32, shape=n_hidden1)
assign_hidden1_weights = tf.assign(hidden1_weights, original_weights)
assign_hidden1_biases = tf.assign(hidden1_biases, original_biases)
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
sess.run(assign_hidden1_weights, feed_dict={original_weights: original_w})
sess.run(assign_hidden1_biases, feed_dict={original_biases: original_b})
# [...] 새 작업에 모델을 훈련시킵니다
print(hidden1.eval(feed_dict={X: [[10.0, 11.0]]}))
# -
# `get_collection()`에 `scope`를 지정하여 변수의 핸들을 가져올 수도 있습니다:
tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES, scope="hidden1")
# 또는 그래프의 `get_tensor_by_name()` 메서드를 사용할 수 있습니다:
tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
tf.get_default_graph().get_tensor_by_name("hidden1/bias:0")
# ### 11.2.3 하위층 동결하기
# * Optimizer 변수목록에서 제외하기
#
# Optimizer의 minimize() 함수를 실행할 때, var_list 파라미터에 동결할 층들을 제외하고 전달하는 방법
#
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1") # 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2") # 재사용
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3") # 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
# -
with tf.name_scope("train"): # 책에는 없음
optimizer = tf.train.GradientDescentOptimizer(learning_rate) # 책에는 없음
train_vars = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope="hidden[34]|outputs")
training_op = optimizer.minimize(loss, var_list=train_vars)
init = tf.global_variables_initializer()
new_saver = tf.train.Saver()
# +
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# -
# * tf.stop_gradient() 사용하기
#
# tf.stop_gradient()가 적용된 아래의 모든 층이 고정됨
#
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
# -
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
name="hidden1") # 동결층 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
name="hidden2") # 동결층 재사용
hidden2_stop = tf.stop_gradient(hidden2)
hidden3 = tf.layers.dense(hidden2_stop, n_hidden3, activation=tf.nn.relu,
name="hidden3") # 동결하지 않고 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu,
name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
# +
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# -
# 훈련하는 코드는 이전과 완전히 동일합니다:
# +
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# -
# ### 11.2.4 동결된 층 캐싱하기
#
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
name="hidden1") # 동결층 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
name="hidden2") # 동결층 재사용 & 캐싱
hidden2_stop = tf.stop_gradient(hidden2)
hidden3 = tf.layers.dense(hidden2_stop, n_hidden3, activation=tf.nn.relu,
name="hidden3") # 동결하지 않고 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu,
name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
# +
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
import numpy as np
n_batches = len(X_train) // batch_size
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
h2_cache = sess.run(hidden2, feed_dict={X: X_train})
h2_cache_valid = sess.run(hidden2, feed_dict={X: X_valid}) # 책에는 없음
for epoch in range(n_epochs):
shuffled_idx = np.random.permutation(len(X_train))
hidden2_batches = np.array_split(h2_cache[shuffled_idx], n_batches)
y_batches = np.array_split(y_train[shuffled_idx], n_batches)
for hidden2_batch, y_batch in zip(hidden2_batches, y_batches):
sess.run(training_op, feed_dict={hidden2:hidden2_batch, y:y_batch})
accuracy_val = accuracy.eval(feed_dict={hidden2: h2_cache_valid, # 책에는 없음
y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_new_model_final.ckpt")
# -
# # 고속 옵티마이저
# ## 모멘텀 옵티마이저
optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate,
momentum=0.9)
# ## 네스테로프 가속 경사
optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate,
momentum=0.9, use_nesterov=True)
# ## AdaGrad
optimizer = tf.train.AdagradOptimizer(learning_rate=learning_rate)
# ## RMSProp
optimizer = tf.train.RMSPropOptimizer(learning_rate=learning_rate,
momentum=0.9, decay=0.9, epsilon=1e-10)
# ## Adam 최적화
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
# ## 학습률 스케줄링
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
# -
with tf.name_scope("train"): # 책에는 없음
initial_learning_rate = 0.1
decay_steps = 10000
decay_rate = 1/10
global_step = tf.Variable(0, trainable=False, name="global_step")
learning_rate = tf.train.exponential_decay(initial_learning_rate, global_step,
decay_steps, decay_rate)
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum=0.9)
training_op = optimizer.minimize(loss, global_step=global_step)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 5
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# # 규제로 과대적합 피하기
# ## $\ell_1$과 $\ell_2$ 규제
# $\ell_1$ 규제를 직접 구현해 보죠. 먼저 평상시처럼 모델을 만듭니다(간단하게 하기 위해 은닉층을 하나만 두겠습니다):
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
logits = tf.layers.dense(hidden1, n_outputs, name="outputs")
# -
# 그다음, 층의 가중치에 대한 핸들을 얻어 크로스 엔트로피 손실에 $\ell_1$ 손실(즉, 가중치의 절댓값)을 더해 전체 손실을 계산합니다:
# +
W1 = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
W2 = tf.get_default_graph().get_tensor_by_name("outputs/kernel:0")
scale = 0.001 # l1 규제 하이퍼파라미터
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y,
logits=logits)
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy")
reg_losses = tf.reduce_sum(tf.abs(W1)) + tf.reduce_sum(tf.abs(W2))
loss = tf.add(base_loss, scale * reg_losses, name="loss")
# -
# 나머지는 이전과 동일합니다:
# +
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# 다른 방법으로는 `tf.layers.dense()` 함수에 규제 함수를 전달할 수 있습니다. 이 함수는 규제 손실을 계산하기 위한 연산을 만들고 규제 손실 컬렉션에 이 연산을 추가합니다. 모델 선언부는 이전과 동일합니다:
# +
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
# -
# 그다음, 동일한 매개변수를 매번 반복하지 않으려고 파이썬의 `partial()` 함수를 사용합니다. `kernel_regularizer` 매개변수를 지정해야 합니다:
scale = 0.001
# +
my_dense_layer = partial(
tf.layers.dense, activation=tf.nn.relu,
kernel_regularizer=tf.contrib.layers.l1_regularizer(scale))
with tf.name_scope("dnn"):
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
hidden2 = my_dense_layer(hidden1, n_hidden2, name="hidden2")
logits = my_dense_layer(hidden2, n_outputs, activation=None,
name="outputs")
# -
# 기본 손실에 규제 손실을 추가합니다:
with tf.name_scope("loss"): # 책에는 없음
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits( # 책에는 없음
labels=y, logits=logits) # 책에는 없음
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy") # 책에는 없음
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
loss = tf.add_n([base_loss] + reg_losses, name="loss")
# 나머지는 평상시와 동일합니다:
# +
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# ## 드롭아웃
# +
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
# +
training = tf.placeholder_with_default(False, shape=(), name='training')
dropout_rate = 0.5 # == 1 - keep_prob
X_drop = tf.layers.dropout(X, dropout_rate, training=training)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X_drop, n_hidden1, activation=tf.nn.relu,
name="hidden1")
hidden1_drop = tf.layers.dropout(hidden1, dropout_rate, training=training)
hidden2 = tf.layers.dense(hidden1_drop, n_hidden2, activation=tf.nn.relu,
name="hidden2")
hidden2_drop = tf.layers.dropout(hidden2, dropout_rate, training=training)
logits = tf.layers.dense(hidden2_drop, n_outputs, name="outputs")
# +
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum=0.9)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# +
n_epochs = 20
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={training: True, X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
# -
# ## 맥스 노름
# 2개의 은닉층을 가진 간단한 MNIST 신경망을 만들어 보겠습니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
# -
# 다음으로 첫 번째 은닉층의 가중치에 대한 핸들을 얻고 `clip_by_norm()` 함수를 사용해 가중치를 클리핑하는 연산을 만듭니다. 그런 다음 클리핑된 가중치를 가중치 변수에 할당하는 연산을 만듭니다:
threshold = 1.0
weights = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
clipped_weights = tf.clip_by_norm(weights, clip_norm=threshold, axes=1)
clip_weights = tf.assign(weights, clipped_weights)
# 두 번째 층에 대해서도 동일하게 할 수 있습니다:
weights2 = tf.get_default_graph().get_tensor_by_name("hidden2/kernel:0")
clipped_weights2 = tf.clip_by_norm(weights2, clip_norm=threshold, axes=1)
clip_weights2 = tf.assign(weights2, clipped_weights2)
# 초기와 연산과 `Saver` 객체를 만듭니다:
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# 이제 모델을 훈련시킵니다. 이전과 매우 동일한데 `training_op`을 실행한 후에 `clip_weights`와 `clip_weights2` 연산을 실행하는 것만 다릅니다:
n_epochs = 20
batch_size = 50
with tf.Session() as sess: # 책에는 없음
init.run() # 책에는 없음
for epoch in range(n_epochs): # 책에는 없음
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size): # 책에는 없음
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
clip_weights.eval()
clip_weights2.eval() # 책에는 없음
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_model_final.ckpt") # 책에는 없음
# 위 구현은 이해하기 쉽고 잘 작동하지만 조금 번거롭습니다. 더 나은 방법은 `max_norm_regularizer()` 함수를 만드는 것입니다:
def max_norm_regularizer(threshold, axes=1, name="max_norm",
collection="max_norm"):
def max_norm(weights):
clipped = tf.clip_by_norm(weights, clip_norm=threshold, axes=axes)
clip_weights = tf.assign(weights, clipped, name=name)
tf.add_to_collection(collection, clip_weights)
return None # 규제 손실을 위한 항이 없습니다
return max_norm
# 그런 다음 (필요한 임계값을 지정해서) 맥스 노름 규제 매개변수에 넘길 함수를 만들기 위해 이 함수를 호출합니다. 은닉층을 만들 때 이 규제 함수를 `kernel_regularizer` 매개변수를 통해 전달할 수 있습니다:
# +
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
# +
max_norm_reg = max_norm_regularizer(threshold=1.0)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
# +
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
# -
# 훈련 연산이 실행된 후에 가중치 클리핑 연산을 실행하는 것을 제외하면 이전과 동일합니다:
n_epochs = 20
batch_size = 50
# +
clip_all_weights = tf.get_collection("max_norm")
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
sess.run(clip_all_weights)
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_model_final.ckpt") # 책에는 없음
# -
# # 연습문제 해답
# 11장의 연습문제는 [11_deep_learning_exercise](11_deep_learning_exercises.ipynb) 노트북에 있습니다.
|
11_deep_learning_reading.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
# import seaborn as sns
# import matplotlib.pyplot as plt
# import matplotlib
# matplotlib.pyplot.style.use('seaborn')
# matplotlib.rcParams['figure.figsize'] = (15, 5)
# # %matplotlib inline
# # %pylab inline
# -
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from IPython.core.display import SVG
import math
# $\lim\limits_{x \to 0^+} {1 \over 0} \ne \infty$
# # $\lim\limits_{x \to a} {\left( f(x) + g(x) \right)} =
# \lim\limits_{x \to a} {f(x)} +
# \lim\limits_{x \to a} {g(x)}$
#
# # $\lim\limits_{x \to a} {\left( f(x) - g(x) \right)} =
# \lim\limits_{x \to a} {f(x)} -
# \lim\limits_{x \to a} {g(x)}$
#
# # $\lim\limits_{x \to a} {\left( f(x) \cdot g(x) \right)} =
# \lim\limits_{x \to a} {f(x)} \cdot
# \lim\limits_{x \to a} {g(x)}$
#
# # $\lim\limits_{x \to a} {\frac{f(x)}{g(x)}} = \frac{
# \lim\limits_{x \to a} {f(x)}
# }{
# \lim\limits_{x \to a} {g(x)}
# }$
#
# # $\lim\limits_{x \to a} {\left( c \cdot f(x) \right)} =
# c \cdot \lim\limits_{x \to a} {f(x)}$
#
# # $\lim\limits_{x \to a} {\left( f(x) \right)^n} =
# \left( \lim\limits_{x \to a} {f(x)} \right)^n$
# # Strategy in finding limits
#
# 1) Direct substitution
#
# - $f(a) = \frac{b}{0}$ probably asymtote (DNE)
# - $f(a) = b$ lim eq $b$
# - $f(a) = \frac{0}{0}$ Indeterminate form
#
# 2) Indeterminate form
#
# - factoring
# - conjucates
# - trig identities
# - approximation
#
# 3) Direct substitution (again) and repeat
# $\lim\limits_{x \to 3} \frac{x^2 - 9x + 18}{x - 3}$
#
# 1) Direct substitution
#
# $\lim\limits_{x \to 3} \frac{9 - 27 + 18}{3 - 3} = \frac{0}{0}$
#
# 2) Indeterminate form -> factoring
#
# $\lim\limits_{x \to 3} \frac{x^2 - 9x + 18}{x - 3} =
# \lim\limits_{x \to 3} \frac{(x-6)(x-3)}{x - 3} =
# \lim\limits_{x \to 3} (x-6)
# $
#
# $a + b = -9$
#
# $ab = 18$
#
# $(-6, -3)$
#
# 3) Direct substitution (again)
#
# $\lim\limits_{x \to 3} (x-6) = \lim\limits_{x \to 3} (3-6) = -3$
#
# 4) Limit found
# $\lim\limits_{x \to -\frac{\pi}{4}} \frac{cos(2x)}{cos(x) + sin(x)}$
#
# 1) Direct substitution
#
# $\frac{cos(2(-\frac{\pi}{4}))}{cos(-\frac{\pi}{4}) + sin(-\frac{\pi}{4})} =
# \frac{cos(-\frac{\pi}{2})}{cos(-\frac{\pi}{4}) + sin(-\frac{\pi}{4})} =
# \frac{0}{\frac{\sqrt2}{2} + -\frac{\sqrt2}{2}} = \frac{0}{0}
# $
#
# 2) Indeterminate form -> trig identities
#
# $\frac{cos^2(x) - sin^2(x)}{cos(x) + sin(x)} =
# \frac{(cos(x) + sin(x))(cos(x) - sin(x))}{cos(x) + sin(x)} =
# cos(x) - sin(x)$
#
# 3) Direct substitution (again)
#
# $cos(-\frac{\pi}{4}) - sin(-\frac{\pi}{4}) =
# \frac{\sqrt2}{2} - -\frac{\sqrt2}{2} =
# \frac{\sqrt2}{2} + \frac{\sqrt2}{2} =
# 2\frac{\sqrt2}{2} = \sqrt2$
#
# 4) Limit found
# $\lim\limits_{x \to 4} \frac{x-4}{\sqrt{3x-3} - 3}$
#
# 1) Direct substitution
#
# ${0 \over \sqrt{9} - 3} = \frac{0}{0}$
#
# 2) Indeterminate form -> conjucates
#
# $\lim\limits_{x \to 3} \frac{(x-4)(\sqrt{3x-3} + 3)}{(\sqrt{3x-3} - 3)(\sqrt{3x-3} + 3)} =
# \lim\limits_{x \to 3} \frac{(x-4)(\sqrt{3x-3} + 3)}{3x-3 - 9} =
# \lim\limits_{x \to 3} \frac{(x-4)(\sqrt{3x-3} + 3)}{3x-12} =
# \lim\limits_{x \to 3} \frac{(x-4)(\sqrt{3x-3} + 3)}{3(x-4)} =
# \lim\limits_{x \to 3} \frac{\sqrt{3x-3} + 3}{3}$
#
# 3) Direct substitution (again)
#
# $\frac{\sqrt{9} + 3}{3} = 2$
#
# 4) Limit found
# $\lim\limits_{x \to-1} \frac{1 + \sqrt{5x + 30} }{x^2 - 1}$
#
# 1) Direct substitution - probably asymtote (DNE)
#
# $\frac{1 + \sqrt{5(-1) + 30} }{(-1)^2 - 1} = \frac{1 + \sqrt{25} }{0}$
#
# 4) Done
# $\lim\limits_{x \to 5} \frac{x^2 - 25}{x^2 -10x +25}$
#
# 1) Direct substitution
#
# $\frac{x^2 - 25}{x^2 -10x +25} =
# \frac{x^2 - 25}{(x - 5)^2} =
# \frac{(5)^2 - 25}{((5) - 5)^2}= \frac{0}{0}$
#
# 2) Indeterminate form -> factoring
#
# $\frac{x^2 - 25}{x^2 -10x +25} =
# \frac{x^2 - 25}{(x - 5)^2} =
# \frac{(x - 5)(x + 5)}{(x - 5)^2} =
# \frac{x+5}{x-5}
# $
#
# 3) Direct substitution (again) - probably asymtote (DNE)
#
# $\frac{x+5}{x-5} = \frac{10}{0}$
#
# 4) Done
# # Limits by rationalizing
#
# ### $\lim\limits_{x \to 1}{ \frac{\sqrt{5x+4}-3}{x-1} }$
#
# $\lim\limits_{x \to 1}{ \frac{\sqrt{5x+4}-3}{x-1} } =
# \frac{
# \lim\limits_{x \to 1}{\sqrt{5x+4}-3}
# }{
# \lim\limits_{x \to 1}{x-1}
# }
# =
# \frac{0}{0}
# $
#
# So refac:
#
# $ \frac{\sqrt{5x+4}-3}{x-1}$
#
# $a^2 - b^2 = (a-b)(a+b)$
#
# $ \frac{\sqrt{5x+4}-3}{x-1} \cdot \frac{\sqrt{5x+4}+3}{\sqrt{5x+4}+3}$
#
# $ \frac{5x + 5}{(x-1)(\sqrt{5x+4}+3)}$
#
# $ \frac{5(x + 1)}{(x-1)(\sqrt{5x+4}+3)}$
#
# $\lim\limits_{x \to 1}{ \frac{5(x+1)}{(x-1)(\sqrt{5x+4}+3)} }$
#
# $\lim\limits_{x \to 1}{ \frac{5}{\sqrt{5(1)+4}+3} } = \frac{5}{6}$
# ### $\lim\limits_{x \to \frac{\pi}{2}}{sin(2x) \over cos(x)}$
#
# $\frac{sin(\pi)}{cos({\pi \over 2})} = \frac{0}{0}$
#
# Use $sin(2a) = 2sin(a)cos(a)$
#
# $\frac{sin(2x)}{cos(x)} = \frac{2sin(x)cos(x)}{cos(x)} = 2sin(x)$, for $x \ne \{..., \frac{\pi}{2}, ...\}$
#
# $2sin(\frac{\pi}{2}) = (2)(1) = 2$
#
# $\lim\limits_{x \to \frac{\pi}{2}}{sin(2x) \over cos(x)} = 2$
# ### $\lim\limits_{x \to \frac{\pi}{2}}{cot^2(x) \over 1 - sin(x)}$
#
# ---
#
# $cot^2 x = \frac{1 + cos2x}{1 - cos2x} = \frac{1 + (1 - 2sin^2x)}{1 - (1 - 2sin^2x)} =
# \frac{2 - 2sin^2x}{2sin^2x} =
# \frac{1 - sin^2x}{sin^2x}$
#
# or:
#
# $cot^2 x = \frac{1}{tan^2 x} = \frac{1}{\frac{sin^2x}{1 - sin^2x}} = \frac{1 - sin^2 x}{sin^2x}$
#
# ---
#
# $\frac{1 - sin^2 x}{sin^2x} = \frac{(1-sinx)(1+sinx)}{sin^2x}$
#
# ---
#
# ${1 \over 1 - sin(x)} \cdot \frac{(1-sinx)(1+sinx)}{sin^2x} =
# \frac{(1-sinx)(1+sinx)}{(sin^2x)(1 - sin(x))} = \frac{1+sinx}{sin^2x}$
#
# $\frac{1+sin(\frac{\pi}{2})}{sin^2(\frac{\pi}{2})} = 2$
# ### $\lim\limits_{\theta \to -\frac{\pi}{4}} \frac{
# 1 + \sqrt{2} sin(\theta)
# }{
# cos(2\theta)
# }$
#
# $\lim\limits_{\theta \to -\frac{\pi}{4}} \frac{
# 1 + \sqrt{2} sin(\theta)
# }{
# 1 - 2sin^2\theta
# }$
#
# $\lim\limits_{\theta \to -\frac{\pi}{4}} \frac{
# 1 + \sqrt{2} sin(\theta)
# }{
# (1 - \sqrt{2}sin\theta)(1 + \sqrt{2}sin\theta)
# }$
#
# $\lim\limits_{\theta \to -\frac{\pi}{4}} \frac{
# 1
# }{
# 1 - \sqrt{2}sin\theta
# }$
#
# $\lim\limits_{\theta \to -\frac{\pi}{4}} \frac{
# 1
# }{
# 1 - \sqrt{2} (-\frac{\sqrt{2}}{2})
# } = \frac{1}{2}$
# ### $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 3cos^2(x)
# }{
# 2 - 2sin(x)
# }$
#
# Use:
#
# $sin^2(x) + cos^2(x) = 1$
#
# $cos^2(x) = 1 - sin^2(x)$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 3(1 - sin^2(x))
# }{
# 2(1 - sin(x))
# }$
#
# Use: $a^2 - b^2 = (a-b)(a+b)$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 3(1 - sin(x))(1 + sin(x))
# }{
# 2(1 - sin(x))
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 3(1 + sin(x))
# }{
# 2
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 3(1 + 1)
# }{
# 2
# } = \frac{6}{2} = 3$
# ### $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# sin^2(2x)
# }{
# 1 - sin^2(x)
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# sin^2(2x)
# }{
# 1 - (1 - cos^2(x))
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# sin^2(2x)
# }{
# cos^2(x)
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# sin(2x)sin(2x)
# }{
# cos(x)cos(x)
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} \frac{
# 2sin(x)cos(x)2sin(x)cos(x)
# }{
# cos(x)cos(x)
# }$
#
# $\lim\limits_{x \to \frac{\pi}{2}} 4sin^2(x) = 4$
# ### $\lim\limits_{x \to -2} \frac{
# \sqrt{3x + 10} - 2
# }{
# x+2
# }$
#
# $\lim\limits_{x \to -2} \frac{
# \sqrt{3x + 10} - 2
# }{
# x+2
# }$
#
# $\lim\limits_{x \to -2} \frac{
# (\sqrt{3x + 10} - 2)(\sqrt{3x + 10} + 2)
# }{
# (x+2)(\sqrt{3x + 10} + 2)
# }$
#
# $\lim\limits_{x \to -2} \frac{
# 3x + 10 - 4
# }{
# (x+2)(\sqrt{3x + 10} + 2)
# }$
#
# $\lim\limits_{x \to -2} \frac{
# 3(x + 2)
# }{
# (x+2)(\sqrt{3x + 10} + 2)
# }$
#
# $\lim\limits_{x \to -2} \frac{
# 3
# }{
# \sqrt{3x + 10} + 2
# } =
# \frac{3}{\sqrt{3(-2) + 10} + 2} = \frac{3}{4}
# $
# ### $\lim\limits_{x \to 2} \frac{x^3 -2x^2}{x^2 - 4}$
#
# $\lim\limits_{x \to 2} \frac{x^2(x - 2)}{(x - 2)(x + 2)}$
#
# $\lim\limits_{x \to 2} \frac{x^2}{(x + 2)} = \frac{4}{4} = 1$
# $\lim_\limits{x \to \frac{\pi}{4}} \frac{cos(2x)}{\sqrt{2}cos(x) - 1}$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \frac{
# cos(2x)(\sqrt{2}cos(x) + 1)
# }{
# (\sqrt{2}cos(x) - 1)(\sqrt{2}cos(x) + 1)
# }$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \frac{
# cos(2x)(\sqrt{2}cos(x) + 1)
# }{
# 2cos^2(x) - 1
# }$
#
# Use:
# * $cos^2 x = 1 - sin^2 x$
# * $cos(2x) = 1 - 2sin^2(x)$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \frac{
# (1 - 2sin^2(x))(\sqrt{2}cos(x) + 1)
# }{
# 1 - 2sin^2(x)
# }$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \sqrt{2}cos(x) + 1$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \sqrt{2}cos(\frac{\pi}{4}) + 1$
#
# $\lim_\limits{x \to \frac{\pi}{4}} \sqrt{2} \cdot \frac{\sqrt{2}}{2} + 1 = 2$
# $\lim_\limits{x\to 2} \frac{3 - \sqrt{5x - 1}}{x - 2}$
#
# $\lim_\limits{x\to 2} \frac{
# (3 - \sqrt{5x - 1})(3 + \sqrt{5x - 1})
# }{
# (x - 2)(3 + \sqrt{5x - 1})
# }$
#
# $\lim_\limits{x\to 2} \frac{
# 9 - (5x - 1)
# }{
# (x - 2)(3 + \sqrt{5x - 1})
# }$
#
# $\lim_\limits{x\to 2} \frac{
# 9 - 5x + 1
# }{
# (x - 2)(3 + \sqrt{5x - 1})
# }$
#
# $\lim_\limits{x\to 2} \frac{
# -5(-2 + x)
# }{
# (x - 2)(3 + \sqrt{5x - 1})
# }$
#
# $\lim_\limits{x\to 2} \frac{
# -5
# }{
# 3 + \sqrt{5x - 1}
# }
# =
# -\frac{5}{6}
# $
# # Limits at infinity
#
# two approaches:
#
# # $\lim\limits_{x\to\infty} \frac{-4x^3 + 5x}{2x^3 + 7} = \frac{-4x^3}{2x^3} = \frac{-4}{2} = -2$
#
# # $\lim\limits_{x\to\infty} \frac{-4x^3 + 5x}{2x^3 + 7} =
# \frac{\frac{-4x^3}{x^3} + \frac{5x}{x^3}}{\frac{2x^3}{x^3} + \frac{7}{x^3}} =
# \frac{-4 + \frac{5}{x^2}}{2 + \frac{7}{x^3}} =
# \frac{-4 + 0}{2 + 0} = -2$
#
# ---
#
# # $\lim\limits_{x\to\infty} \frac{-5x^4 + 2x}{x^3} = \frac{-5x^4}{x^3} = -5x = -\infty = $ 𝑢𝑛𝑏𝑜𝑢𝑛𝑑𝑒𝑑
#
# # $\lim\limits_{x\to\infty} \frac{-5x^4 + 2x}{x^3} = \lim\limits_{x\to\infty} \frac{\frac{-5x^4 + 2x}{x^4}}{\frac{x^3}{x^4}} = \frac{-5 + 0}{0} = -\infty = unbounded$
#
# ---
#
# # $\lim\limits_{x\to\infty} \frac{-5x^3 + 2x^2 -7}{x^4 + 3x} = \frac{-5x^3}{x^4} = \frac{-5}{x} = 0$
#
# # $\lim\limits_{x\to\infty} \frac{-5x^3 + 2x^2 -7}{x^4 + 3x} =
# \frac{\frac{-5x^3}{x^4} + \frac{2x^2}{x^4} - \frac{7}{x^4}}{\frac{x^4}{x^4} + \frac{3x}{x^4}} =
# \frac{\frac{-5}{x} + \frac{2}{x^2} - \frac{7}{x^4}}{1 + \frac{3}{x^3}} =
# \frac{0 + 0 - 0}{1 + 0} =
# \frac{0}{1} = 0
# $
#
# ---
# # $\lim\limits_{x\to-\infty} \frac{\sqrt{16x^6 -x^2}}{6x^3 + x^2} =
# \frac{\sqrt{(4x^3)^2 -x^2}}{6x^3 + x^2} =
# \frac{\sqrt{(4x^3)^2}}{6x^3} =
# \frac{|4x^3|}{6x^3} =
# \frac{4}{-6} =
# -\frac{2}{3}
# $
#
# hint: $\sqrt{a^2} = |a|$
#
# ## $\lim\limits_{x\to-\infty} \frac{\sqrt{16x^6 -x^2}}{6x^3 + x^2} =
# \frac{\frac{\sqrt{16x^6 -x^2}}{x^3}}{\frac{6x^3}{x^3} + \frac{x^2}{x^3}} =
# \frac{\frac{\sqrt{16x^6 -x^2}}{-\sqrt{x^6}}}{\frac{6x^3}{x^3} + \frac{x^2}{x^3}} =
# -\frac{\sqrt{\frac{16x^6}{x^6} - \frac{x^2}{x^6}}}{\frac{6x^3}{x^3} + \frac{x^2}{x^3}} =
# -\frac{\sqrt{16 - \frac{1}{x^4}}}{6 + \frac{1}{x}} =
# -\frac{\sqrt{16 - 0}}{6 + 0} =
# -\frac{4}{6} =
# -\frac{2}{3}
# $
#
# ---
#
# # $\lim\limits_{x\to-\infty} \frac{\sqrt{4x^4 - x}}{2x^2 +3} =
# \frac{\sqrt{4x^4}}{2x^2} =
# \frac{\sqrt{(2x^2)^2}}{2x^2} =
# \frac{|2x^2|}{2x^2} =
# \frac{2}{+2} = 1
# $
#
# # $\lim\limits_{x\to-\infty} \frac{\sqrt{4x^4 - x}}{2x^2 +3} =
# \frac{ \frac{\sqrt{4x^4 - x}}{x^2} }{ \frac{2x^2 +3}{x^2} } =
# \frac{ \frac{\sqrt{4x^4 - x}}{+\sqrt{x^4}} }{ \frac{2x^2 +3}{x^2} } =
# \frac{ \sqrt{\frac{4x^4 - x}{x^4}} }{ \frac{2x^2}{x^2} + \frac{3}{x^2} } =
# \frac{ \sqrt{4 - \frac{1}{x^3}} }{ 2 + \frac{3}{x^2} } =
# \frac{ \sqrt{4 - 0} }{ 2 + 0 } =
# \frac{ 2 }{ 2 } = 1
# $
#
# ---
#
# # $\lim\limits_{x\to\infty} \frac{\sqrt{9x^6 + 4x^2}}{x^3 - 1} =
# \frac{\sqrt{9x^6}}{x^3} =
# \frac{\sqrt{(3x^3)^2}}{x^3} =
# \frac{|3x^3|}{x^3} =
# \frac{3}{+1} = 3
# $
#
# ## $\lim\limits_{x\to\infty} \frac{\sqrt{9x^6 + 4x^2}}{x^3 - 1} =
# \frac{\sqrt{9x^6 + 4x^2}}{x^3 - 1} =
# \frac{ \frac{\sqrt{9x^6 + 4x^2}}{x^3} }{ \frac{x^3 - 1}{x^3} } =
# \frac{ \frac{\sqrt{9x^6 + 4x^2}}{+\sqrt{x^6}} }{ \frac{x^3 - 1}{x^3} } =
# \frac{ \sqrt{\frac{9x^6}{x^6} + \frac{4x^2}{x^6}} }{ \frac{x^3}{x^3} - \frac{1}{x^3} } =
# \frac{ \sqrt{9 + \frac{4}{x^4}} }{ 1 - \frac{1}{x^3} } =
# \frac{ \sqrt{9 + 0} }{ 1 - 0 } =
# \frac{3}{1} = 3
# $
#
# ---
# # lim-approaching
# # $\frac{3x}{(x+2)^2}, @x = -2$
#
# # $\frac{3x}{(x+2)^2}, x \to -2^+ \to-\infty$
#
# $\frac{3(-1.9)}{(-1.9 + 2)^2} =
# \frac{-3(1.9)}{(0.1)^2} =
# \frac{-3(1.9)}{0.01}
# $
#
# # $\frac{3x}{(x+2)^2}, x \to -2^- \to-\infty$
#
# $\frac{3(-2.1)}{(-2.1 + 2)^2} =
# \frac{-3(2.1)}{(-0.1)^2} =
# \frac{-3(2.1)}{0.01}
# $
# # $\frac{x}{ln(x - 2)}, @x = 3$
#
# # $\frac{x}{ln(x - 2)}, x \to 3^+ \to \infty$
#
# $\frac{3.1}{ln(3.1 - 2)} = \frac{3.1}{ln(1.1)} = \frac{3}{0.01}$
#
# # $\frac{x}{ln(x - 2)}, x \to 3^- \to -\infty$
#
# $\frac{2.9}{ln(2.9 - 2)} = \frac{3.1}{ln(0.9)} = \frac{3}{-0.1}$
|
005_calc/lim.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandas compatible data sources
# - text, - csv, josn, html tables etc
# - binary - for io performance or other software like excel
# - relational dbs (all major dbs)
#
# ----
# ### Tate Collection
import pandas as pd
import os
path = os.path.join('.', 'data', 'artwork_data.csv')
df = pd.read_csv(path, nrows=5)
df
# Pandas assigns incremental indices to the data frame, but we already have an id column...
df = pd.read_csv(path, index_col='id', nrows=5)
df
# for our needs, we only need the artist column...
df = pd.read_csv(path, index_col='id', usecols=['id', 'artist'], nrows=5)
# we could also pass column indices 0 and 2 or the names
df
# Good to define columns in constant cariable
COLS_TO_USE = ['id', 'artist', 'title', 'medium', 'year', 'acquisitionYear', 'height', 'width', 'units']
df = pd.read_csv(path, index_col='id', usecols=COLS_TO_USE)
df.head()
# +
# We have a warning, pandas cannot predict data type. We can deal with this later
# As this is the data we want to use for forthcoming work, we should save this subset
# The pickle format is python-native format, we will only be using this with python code.
df.to_pickle(os.path.join('.','data_frame.pickle'))
# -
# ----
# ## Artists
#
# Artist data is found in different json files, with similar schema. This is not the ideal format for querying, so we'll need to convert it first. Best to use the method family:
#
# ```
# pd.DataFrame.from_*()
# ```
# Example use of from_records
records = [('Espresso', '5$'), ('Falt White', '10$')]
pd.DataFrame.from_records(records, columns=['Coffee', 'Price'])
# Read the artist json files...
KEYS_TO_USE = ['id', 'all_artists', 'title', 'medium', 'dateText', 'acquisitionYear', 'height', 'width', 'units']
import json
def get_record_from_file(file_path , keys_to_use):
""" Process single json file and return a
tuple containing specific fields."""
with open(file_path) as artwork_file:
content = json.load(artwork_file)
record = []
for field in keys_to_use:
record.append(content[field])
return tuple(record)
# +
# Use get_record_from_file to process the first files
import os
SAMPLE_JSON = os.path.join('.', 'data','artworks','a','000','a00001-1035.json')
sample_record = get_record_from_file(SAMPLE_JSON, KEYS_TO_USE)
sample_record
# -
# Iterate over multiple files...
# +
import os
def read_artworks_from_json(keys_to_use):
""" Traverse the directories with JSON files.
For first file in each directory call function
for processing single file and go to the next
directory.
"""
JSON_ROOT = os.path.join('.', 'data','artworks')
artworks = []
for root, _, files in os.walk(JSON_ROOT):
for f in files:
if f.endswith('json'):
record = get_record_from_file( os.path.join(root,f), keys_to_use)
artworks.append(record)
break # only do first file
df = pd.DataFrame.from_records(artworks, columns=keys_to_use, index="id")
return df
# -
df = read_artworks_from_json(KEYS_TO_USE)
df.head()
|
course-notes/pluralsight/pandas fundamentals/3-exploring pandas data input capabilities.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp utils
# -
# # Utils
#
# > This file holds function to load the data and create folds for cross-validation.
#hide
# %load_ext autoreload
# %autoreload 2
#hide
from nbdev.showdoc import *
from plant_pathology.config import TEST_DATA_PATH
import numpy as np
from fastcore.all import *
# +
#export
from typing import List, Tuple
import pandas as pd
from fastcore.all import *
# -
# For some of our tests, we need access to the competition's data, which is stored in our `config` module.
# ## Load Data
# This reads the training CSV into a pandas DataFrame. You can choose to load the CSV that includes the cross-validation (CV) folds already added to it (if you've created it). You can also choose to load the training data with pseudo-labeled examples added as well (if you've created a CSV of pseudo-labels).
#export
def load_data(
data_path: Path, with_folds: bool = False, pseudo_labels_path: str = None
) -> Tuple[Path, pd.DataFrame]:
"""Load data (with/without cross-validation folds) into DataFrame."""
train_df = pd.read_csv(
data_path / ("train_folds.csv" if with_folds else "train.csv")
)
# Add pseudo-labels to DataFrame
if pseudo_labels_path is not None:
train_df = pd.concat(
[train_df, pd.read_csv(pseudo_labels_path)], ignore_index=True
)
return data_path, train_df
path, df = load_data(TEST_DATA_PATH, with_folds=False)
df.head()
# ## Average Predictions
#export
def average_preds(dfs: List[pd.DataFrame]) -> pd.DataFrame:
"""Average predictions on test examples across prediction DataFrames in `dfs`."""
all_preds_df = pd.concat(dfs)
avg_preds_df = all_preds_df.groupby(all_preds_df.image_id).mean()
return avg_preds_df
#hide
# Create fake test set prediction dataframes
NUM_EXAMPLES = 5
all_zeros_prediction_dfs = []
for _ in range(5): # One dataframe for each fold
# Make dataframe with fake predictions
fake_preds = np.zeros((NUM_EXAMPLES, len(df.columns)))
preds_df = pd.DataFrame(fake_preds, columns=df.columns)
# Fix test filenames
test_fns = [f"Test_{i}" for i in range(NUM_EXAMPLES)]
preds_df["image_id"] = test_fns
all_zeros_prediction_dfs.append(preds_df)
# Let's test this by confirming that if the predictions for everything are `0.0`, the average of all the predictions should also be `0.0`.
len(all_zeros_prediction_dfs)
all_zeros_prediction_dfs[0]
averaged_preds_df = average_preds(all_zeros_prediction_dfs); averaged_preds_df
assert np.all(averaged_preds_df == 0.) # Average of a bunch of 0's is 0
test_eq(averaged_preds_df.shape, (5, 4)) # 5 examples, 4 classes
# ### Save Averaged Preds
# Utility function to load and average all test set prediction CSVs matching naming pattern `"predictions_fold_[0-4].csv"`, which is the default naming scheme when running the training script using 5-fold cross-validation.
#export
def get_averaged_preds(path: Path, verbose: bool = False) -> Path:
"""Returns DataFrame of averaged of averaged predictions of prediction CSVs in `path` dir."""
# Load test set prediction CSVs for each of 5 CV folds
prediction_files = list(path.glob("predictions_fold_[0-4].csv"))
if verbose:
print(prediction_files)
return average_preds([pd.read_csv(fn) for fn in prediction_files])
#hide
from nbdev.export import notebook2script; notebook2script()
|
nbks/00_utils.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # deepctr
# +
# %load_ext autoreload
# %autoreload 2
import os
import warnings
warnings.filterwarnings('ignore')
import sys
sys.path.append(os.path.abspath('..'))
# ---------------------------------
from time import sleep
import numpy as np
import pandas as pd
import scipy
import tqdm
from copy import deepcopy
import tensorflow as tf
from tensorflow.keras.layers import Activation
import matplotlib.pyplot as plt
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
from sklearn.preprocessing import LabelEncoder
from hyperopt import hp
from deepctr.models import xDeepFM
from deepctr.inputs import SparseFeat, DenseFeat, get_feature_names
# ---------------------------------
from tools import CV, Tuning, CVGetScore, IdxValEncoder, LE, CyclicLR, MaxLrFinder
# ---------------------------------
from tools import focal_loss, gelu, mish
from tensorflow.keras.utils import get_custom_objects
get_custom_objects().update({'focal_loss': focal_loss()})
get_custom_objects().update({'mish': mish})
get_custom_objects().update({'gelu': gelu})
# +
train_df = pd.read_csv('../data/train.csv', index_col='id')
test_df = pd.read_csv('../data/test.csv', index_col='id')
# ord_5
for i in range(2):
train_df[f'ord_5_{i}'] = train_df['ord_5'].str[i]
test_df[f'ord_5_{i}'] = test_df['ord_5'].str[i]
# null
train_df['null'] = train_df.isna().sum(axis=1)
test_df['null'] = test_df.isna().sum(axis=1)
for col in test_df.columns:
train_df[col].fillna('isnull', inplace=True)
test_df[col].fillna('isnull', inplace=True)
# target
target = train_df['target']
y_train = target.values
# drop
train_df.drop(['target', 'ord_5'], axis=1, inplace=True)
test_df.drop(['ord_5'], axis=1, inplace=True)
# +
feature_col = train_df.columns
bin_col = ['null']
class_col = ['bin_0', 'bin_1', 'bin_2', 'bin_3', 'bin_4',
'nom_0', 'nom_1', 'nom_2', 'nom_3', 'nom_4',
'nom_5', 'nom_6', 'nom_7', 'nom_8', 'nom_9',
'ord_0', 'ord_1', 'ord_2', 'ord_3', 'ord_4',
'day', 'month', 'ord_5_0', 'ord_5_1']
# +
ecd = LE(feature_col, bin_col=bin_col, class_col=class_col)
ecd.fit(train_df, verbose=1)
ecd.fit(test_df, verbose=1)
x_train_arr = ecd.transform(train_df, verbose=1)
x_test_arr = ecd.transform(test_df, verbose=1)
del train_df, test_df
# +
# x_train_df = pd.DataFrame(data=x_train_arr, columns=feature_col)
# x_test_df = pd.DataFrame(data=x_test_arr, columns=feature_col)
# -
def col_func(vocabulary, sparse_features, dense_features, k=5):
# sparse
feature_col = list()
for f in sparse_features:
feature_col.append(SparseFeat(f, vocabulary_size=vocabulary[f], embedding_dim=k))
for f in dense_features:
feature_col.append(DenseFeat(f, 1))
dnn_f = feature_col
linear_f= feature_col
fn = get_feature_names(linear_f + dnn_f)
return dnn_f, linear_f, fn
def xdeepfm(vocabulary, k, loss, metrics, optimizer,
num_deep_layer=2, num_neuron=256,
num_cin_layer=2, num_cin=128,**kwargs):
dnn_f, linear_f, _ = col_func(vocabulary, sparse_features=class_col, dense_features=bin_col, k=k)
tf.random.set_seed(1024)
model = xDeepFM(linear_feature_columns=linear_f,
dnn_feature_columns=dnn_f,
cin_layer_size=tuple(num_cin for _ in range(num_cin_layer)),
dnn_hidden_units=tuple(num_neuron for _ in range(num_deep_layer)),
**kwargs)
model.compile(loss=loss, metrics=metrics, optimizer=optimizer)
return model
def mkinput(input_arr, feature_col):
return dict(zip(feature_col, input_arr.T))
# # Search Max LR
# see - ./main/main_8_xdeepfm_relu.ipynb
# # fit one
def cv_score(batch_size, epochs, nflod, base_lr, max_lr, model_params, model_func, verbose=1):
clr = CyclicLR(base_lr=0.1**(base_lr),
max_lr = 0.1**(max_lr),
step_size= int(4.0*(x_train_arr.shape[0]*((nflod-1)/nflod)) / batch_size),
mode='triangular2',
gamma=1.0)
es = tf.keras.callbacks.EarlyStopping(monitor='val_AUC', patience=2, mode='max', restore_best_weights=True)
fit_param = {'batch_size': batch_size, 'epochs':epochs, 'verbose': verbose, 'callbacks':[es, clr]}
model = model_func(**model_params)
cv = CV(model, nflod, random_state=2333)
score = cv.fit(x=mkinput(x_train_arr, feature_col),
y=y_train,
metrics_func=roc_auc_score,
split_method=StratifiedKFold,
fit_params=fit_param,
eval_param={'batch_size':batch_size},
use_proba=False,
verbose=verbose,
fit_use_valid=True)
tf.keras.backend.clear_session()
print(score)
# # tunning
# +
batch_size = 8192
epochs = 100
nflod = 5
seed = 2333
# fit param
clr = CyclicLR(base_lr=0.1**(4.5),
max_lr = 0.1**(3.5),
step_size= int(4.0*(x_train_arr.shape[0]*((nflod-1)/nflod)) / batch_size),
mode='triangular2',
gamma=1.0)
es = tf.keras.callbacks.EarlyStopping(monitor='val_AUC',
patience=2,
mode='max',
restore_best_weights=True)
fit_param = {
'batch_size': batch_size,
'epochs':epochs,
'verbose': 0,
'callbacks':[es, clr]
}
cv_fit_param = {'fit_params': fit_param,
'eval_param': {'batch_size':batch_size},
'use_proba':False,
'fit_use_valid': True}
# model_fix_param & model_search_space
model_fix_param = {'vocabulary': ecd.get_vocabulary(),
'loss': 'binary_crossentropy',
'metrics': ['AUC'],
'optimizer': 'Adam',
'dnn_activation': 'mish',
'cin_activation': 'linear',
'dnn_use_bn': False,
'num_deep_layer': 2,
'num_neuron': 256,
'num_cin_layer': 2}
ss = {
'num_cin': (hp.choice, (64, 100)),
'k': (hp.choice, (5, 6, 7, 8, 9, 10)),
'l2_reg_linear': (hp.loguniform, (-20, 0)),
'l2_reg_embedding': (hp.loguniform, (-7.5, -2.5)),
'l2_reg_dnn': (hp.loguniform, (-20, -7.5)),
'l2_reg_cin': (hp.loguniform, (-15, 0)),
'dnn_dropout': (hp.loguniform, (-20, -1))
}
# cv get score
def neg_auc(y_true, y_pred):
return - roc_auc_score(y_true, y_pred)
gs = CVGetScore(x=mkinput(x_train_arr, feature_col),
y=y_train,
metrics_func=neg_auc,
split_method=StratifiedKFold,
nfolds=nflod,
random_state=seed,
model=xdeepfm,
cv_fit_params=cv_fit_param,
model_fix_params=model_fix_param,
model_search_space=ss)
tuning = Tuning(gs, verbose=1)
tuning.fmin(gs.GET_SEARCH_SPACE(), max_evals=100)
# -
log = tuning.log.get_log()
log.sort_values('score').head()
log.sort_values('score').tail()
tuning.log.plot(score_interval=[-0.789, -0.788])
seed = np.random.randint(2**32)
print(seed)
# ## 3355867947
######
log.to_csv(f'/data/{seed}.csv', index=False)
#####
######
seed = 4293006264
log = pd.read_csv(f'/data/{seed}.csv')
#####
log.sort_values('score').head()
# # stacking
# +
batch_size = 8192
epochs = 400
nflod = 40
nmodel = 5
# model params
model_tuning_param = log.sort_values('score').head(nmodel).reset_index(drop=True).to_dict()
model_param = {'vocabulary': ecd.get_vocabulary(),
'loss': 'binary_crossentropy',
'metrics': ['AUC'],
'optimizer': 'Adam',
'dnn_activation': 'mish',
'cin_activation': 'linear',
'dnn_use_bn': False,
'num_deep_layer': 2,
'num_neuron': 256,
'num_cin_layer': 2}
# callbacks
clr = CyclicLR(
base_lr=0.1**(5),
max_lr = 0.1**(3.5),
step_size= int(4.0*(x_train_arr.shape[0]*((nflod-1)/nflod)) / batch_size),
mode='triangular2',
gamma=1.0)
es = tf.keras.callbacks.EarlyStopping(monitor='val_AUC',
patience=5,
mode='max',
restore_best_weights=True)
# fit
fit_param = {
'batch_size': batch_size,
'epochs':epochs,
'verbose': 0,
'callbacks':[es, clr]
}
pred_lst = []
score_lst = []
pred_arr_lst = []
for i in range(nmodel):
model_params = deepcopy(model_param)
for param_name in model_tuning_param.keys():
if param_name not in ['score', 'update', 'usetime', 'index']:
model_params[param_name] = model_tuning_param[param_name][i]
# cv
model = xdeepfm(**model_params)
cv = CV(model, nflod)
score, pred_arr = cv.fit(x=mkinput(x_train_arr, feature_col),
y=y_train,
metrics_func=roc_auc_score,
split_method=StratifiedKFold,
fit_params=fit_param,
eval_param={'batch_size':batch_size},
use_proba=False,
verbose=True,
fit_use_valid=True,
output_oof_pred=True)
pred = cv.predict(x=mkinput(x_test_arr, feature_col), pred_param={'batch_size': batch_size})
pred_lst.append(pred)
score_lst.append(score)
pred_arr_lst.append(pred_arr)
print('score: ', score)
del model, cv
tf.keras.backend.clear_session()
# -
(0.7895477195367968 + 0.7895494333002201 + 0.7895496775194 + 0.7895507896600742 + 0.7895647491197464)/5
pred_arr = np.array(pred_arr_lst).squeeze().T
np.save(f'/data/{seed}stacking1.npy', pred_arr)
pred_arr.shape
pred = np.array(pred_lst).squeeze().T
np.save(f'/data/{seed}predict.npy', pred)
pred.shape
submission = pd.read_csv('../data/sample_submission.csv', index_col='id')
submission['target'] = np.mean(pred_lst, axis=0)
submission.to_csv(f'/data/main_8_xdeepfm_mish_{seed}.csv')
|
main/main_8_xdeepfm_mish.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HarmanBhutani/Data-Science-DTI5125/blob/main/DTI5125_chatbot.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="NIZ2uCQnpO99" outputId="a09b5096-94d7-4b6f-93e4-267bbb234ae9"
# !pip install rasa==1.10.3
# + colab={"base_uri": "https://localhost:8080/"} id="CVlR5YawpSD7" outputId="225fb44f-c4be-44f3-caf0-984c031489eb"
# !python -m spacy download en
# + colab={"base_uri": "https://localhost:8080/"} id="M_Xm_MEZpcyk" outputId="9b5c8531-8231-4f85-9ae7-3a8b4ec21fca"
# !pip install nest_asyncio==1.3.3
# + colab={"base_uri": "https://localhost:8080/"} id="gdyMMNyzpe95" outputId="ee11edb7-c329-4d0e-ad6c-762daf438f88"
import os
import rasa
import nest_asyncio
nest_asyncio.apply()
print("Event loop ready.")
# + id="jJXOoTJUphFx"
from rasa.cli.scaffold import create_initial_project
# + id="mdlW5ZSEqc6C"
project = "DTI5125-Chatbot"
create_initial_project(project)
# + colab={"base_uri": "https://localhost:8080/"} id="X4sFZLErqhza" outputId="72440a58-4310-4ae9-a430-008b293facda"
# move into project directory and show files
os.chdir(project)
print(os.listdir("."))
# + colab={"base_uri": "https://localhost:8080/"} id="2vO-ynF6qkZ_" outputId="37b7af61-0771-49a1-e96e-580f10ab22f4"
config = "config.yml"
training_files = "data/"
domain = "domain.yml"
output = "models/"
print(config, training_files, domain, output)
# + id="ZryVb0c3qrXs"
import os
# + colab={"base_uri": "https://localhost:8080/"} id="VBRzsPJwqnUZ" outputId="ef648c92-72f6-45ef-f5f0-53dcdd739b21"
model_path = rasa.train(domain, config, [training_files], output)
print(model_path)
# + colab={"base_uri": "https://localhost:8080/"} id="K-sJp9fbqpiY" outputId="0d764400-e685-4452-8cf1-082af010f9fa"
from rasa.jupyter import chat
endpoints = 'endpoints.yml'
chat(model_path, endpoints)
# + colab={"base_uri": "https://localhost:8080/"} id="mtec-PaArBWf" outputId="0ff95205-5203-4a85-8be2-aaa03988e985"
chat(model_path, endpoints)
# + colab={"base_uri": "https://localhost:8080/"} id="54CztHMRtsTq" outputId="d8dfec68-7a88-4f2a-dc5f-c8397c286688"
# %%writefile data/nlu.md
nlu_md = """
## intent:greet
- hey
- hello there
- hi
- hello
- good morning
- good evening
- hey there
- let's go
- hey dude
- goodmorning
- goodevening
- good afternoon
- start
## intent:affirm
- great
- OK
- okay
- thank you
- thanks
- yes, thanks
- cheers
- thanks a lot
- thank you very much
## intent: health_about
- Please tell me about [health department](HealthInsurance),
- Please tell me the [health insurance](HealthInsurance) Plans for family
- What is [health insurance](HealthInsurance)
- For how many year's [health insurance](HealthInsurance) is covered
- Please tell me about [abc health insurance](HealthInsurance)
- Please tell me the [abc health insurance](HealthInsurance) Plans for family
- What is [abc health insurance](HealthInsurance)
- For how many year's [abc health insurance](HealthInsurance) is covered
- How to renew my [health insurance](HealthInsurance)
- What is [health insurance](HealthInsurance)
- I would like to know more about [health insurance](HealthInsurance)
- Please tell something about [health insurance](HealthInsurance)
- What is meant by [health insurance](HealthInsurance)
- Explain about [health insurance](HealthInsurance)
- Give me the details of [health insurance](HealthInsurance)
- Tell me about [health insurance](HealthInsurance)
- Please tell about [health insurance](HealthInsurance)
- How to renew my [abc health insurance](HealthInsurance)
- What is [abc health insurance](HealthInsurance)
- I would like to know more about [abc health insurance](HealthInsurance)
- Please tell something about [abc health insurance](HealthInsurance)
- What is meant by [abc health insurance](HealthInsurance)
- Explain about [abc health insurance](HealthInsurance)
- Give me the details of [abc health insurance](HealthInsurance)
- Tell me about [abc health insurance](HealthInsurance)
- Please tell about [abc health insurance](HealthInsurance)
## intent: vehicle_about
- How to renew my [abc vehicle insurance](VehicleInsurance)
- What is [abc vehicle insurance](VehicleInsurance)
- I would like to know more about [abc vehicle insurance](VehicleInsurance)
- Please tell something about [abc vehicle insurance](VehicleInsurance)
- What is meant by [abc vehicle insurance](VehicleInsurance)
- Explain about [abc vehicle insurance](VehicleInsurance)
- Give me the details of [abc vehicle insurance](VehicleInsurance)
- Tell me about [abc vehicle insurance](VehicleInsurance)
- Please tell about [abc vehicle insurance](VehicleInsurance)
- How to renew my [vehicle insurance](VehicleInsurance)
- What is [vehicle insurance](VehicleInsurance)
- I would like to know more about [vehicle insurance](VehicleInsurance)
- Please tell something about [vehicle insurance](VehicleInsurance)
- What is meant by [vehicle insurance](VehicleInsurance)
- Explain about [vehicle insurance](VehicleInsurance)
- Give me the details of [vehicle insurance](VehicleInsurance)
- Tell me about [vehicle insurance](VehicleInsurance)
- Please tell about [vehicle insurance](VehicleInsurance)
## intent: vehicle_benefits
- What are the [vehicle insurance](VehicleInsurance) [benefits](Benefits)?
- Are there any [vehicle insurance](VehicleInsurance) benefits?
- Tell me about the [benefits](Benefits) of [vehicle insurance](VehicleInsurance)?
- What do you know about [vehicle insurance](VehicleInsurance) [benefits](Benefits)?
- What are the [benefits](Benefits) of taking ABC [vehicle insurance](VehicleInsurance)?
- What are the [benefits](Benefits) over other [vehicle insurance](VehicleInsurance) companies?
- What are the [abc vehicle insurance](VehicleInsurance) [benefits](Benefits)?
- Are there any [abc vehicle insurance](VehicleInsurance) [benefits](Benefits)?
- Tell me about the benifits of [abc vehicle insurance](VehicleInsurance)?
- What do you know about [abc vehicle insurance](VehicleInsurance) [benefits](Benefits)?
- What are the [benefits](Benefits) of taking ABC [abc vehicle insurance](VehicleInsurance)?
- What are the [benefits](Benefits) over other [abc vehicle insurance](VehicleInsurance) companies?
- What are the [vehicle insurance](VehicleInsurance) [tax benefits](Benefits)?
- Are there any [vehicle insurance](VehicleInsurance) tax benefits?
- Tell me about the [tax benefits](Benefits) of [vehicle insurance](VehicleInsurance)?
- What do you know about [vehicle insurance](VehicleInsurance) [tax benefits](Benefits)?
- What are the [tax benefits](Benefits) of taking ABC [vehicle insurance](VehicleInsurance)?
- What are the [tax benefits](Benefits) over other [vehicle insurance](VehicleInsurance) companies?
- What are the [abc vehicle insurance](VehicleInsurance) [tax benefits](Benefits)?
- Are there any [abc vehicle insurance](VehicleInsurance) [tax benefits](Benefits)?
- Tell me about the benifits of [abc vehicle insurance](VehicleInsurance)?
- What do you know about [abc vehicle insurance](VehicleInsurance) [tax benefits](Benefits)?
- What are the [tax benefits](Benefits) of taking ABC [abc vehicle insurance](VehicleInsurance)?
- What are the [tax benefits](Benefits) over other [abc vehicle insurance](VehicleInsurance) companies?
- What are the [vehicle insurance](VehicleInsurance) [regular benefits benefits](Benefits)?
- Are there any [vehicle insurance](VehicleInsurance) regular benefits benefits?
- Tell me about the [regular benefits benefits](Benefits) of [vehicle insurance](VehicleInsurance)?
- What do you know about [vehicle insurance](VehicleInsurance) [regular benefits benefits](Benefits)?
- What are the [regular benefits benefits](Benefits) of taking ABC [vehicle insurance](VehicleInsurance)?
- What are the [regular benefits benefits](Benefits) over other [vehicle insurance](VehicleInsurance) companies?
- What are the [abc vehicle insurance](VehicleInsurance) [regular benefits benefits](Benefits)?
- Are there any [abc vehicle insurance](VehicleInsurance) [regular benefits benefits](Benefits)?
- Tell me about the benifits of [abc vehicle insurance](VehicleInsurance)?
- What do you know about [abc vehicle insurance](VehicleInsurance) [regular benefits benefits](Benefits)?
- What are the [regular benefits benefits](Benefits) of taking ABC [abc vehicle insurance](VehicleInsurance)?
- What are the [regular benefits benefits](Benefits) over other [abc vehicle insurance](VehicleInsurance) companies?
## intent: health_benefits
- What are the [health insurance](HealthInsurance) [benefits](Benefits)?
- Are there any [health insurance](HealthInsurance) [benefits](Benefits)?
- Tell me about the [benefits](Benefits) of [health insurance](HealthInsurance)?
- What do you know about [health insurance](HealthInsurance) [benefits](Benefits)?
- What are the [benefits](Benefits) of taking ABC [health insurance](HealthInsurance)?
- What are the [benefits](Benefits) over other [health insurance](HealthInsurance) companies?
- What are the [abc health insurance](HealthInsurance) [benefits](Benefits)?
- Are there any [abc health insurance](HealthInsurance) [benefits](Benefits)?
- Tell me about the [benefits](Benefits) of [abc health insurance](HealthInsurance)?
- What do you know about [abc health insurance](HealthInsurance) [benefits](Benefits)?
- What are the [benefits](Benefits) of taking ABC [abc health insurance](HealthInsurance)?
- What are the [benefits](Benefits) over other [abc health insurance](HealthInsurance) companies?
- What are the [health insurance](HealthInsurance) [regular benefits](Benefits)?
- Are there any [health insurance](HealthInsurance) [regular benefits](Benefits)?
- Tell me about the [regular benefits](Benefits) of [health insurance](HealthInsurance)?
- What do you know about [health insurance](HealthInsurance) [regular benefits](Benefits)?
- What are the [regular benefits](Benefits) of taking ABC [health insurance](HealthInsurance)?
- What are the [regular benefits](Benefits) over other [health insurance](HealthInsurance) companies?
- What are the [abc health insurance](HealthInsurance) [regular benefits](Benefits)?
- Are there any [abc health insurance](HealthInsurance) [regular benefits](Benefits)?
- Tell me about the [regular benefits](Benefits) of [abc health insurance](HealthInsurance)?
- What do you know about [abc health insurance](HealthInsurance) [regular benefits](Benefits)?
- What are the [regular benefits](Benefits) of taking ABC [abc health insurance](HealthInsurance)?
- What are the [regular benefits](Benefits) over other [abc health insurance](HealthInsurance) companies?
- What are the [health insurance](HealthInsurance) [complementary benefits](Benefits)?
- Are there any [health insurance](HealthInsurance) [complementary benefits](Benefits)?
- Tell me about the [complementary benefits](Benefits) of [health insurance](HealthInsurance)?
- What do you know about [health insurance](HealthInsurance) [complementary benefits](Benefits)?
- What are the [complementary benefits](Benefits) of taking ABC [health insurance](HealthInsurance)?
- What are the [complementary benefits](Benefits) over other [health insurance](HealthInsurance) companies?
- What are the [abc health insurance](HealthInsurance) [complementary benefits](Benefits)?
- Are there any [abc health insurance](HealthInsurance) [complementary benefits](Benefits)?
- Tell me about the [complementary benefits](Benefits) of [abc health insurance](HealthInsurance)?
- What do you know about [abc health insurance](HealthInsurance) [complementary benefits](Benefits)?
- What are the [complementary benefits](Benefits) of taking ABC [abc health insurance](HealthInsurance)?
- What are the [complementary benefits](Benefits) over other [abc health insurance](HealthInsurance) companies?
## intent: health_timeperiod
- What is the [validity](TimePeriod) of [abc health insurance](HealthInsurance)?
- What is the [duration](TimePeriod) of [abc health insurance](HealthInsurance)?
- How [long](TimePeriod) is the [abc health insurance](HealthInsurance) valid?
- What is the [validity](TimePeriod) of [health insurance](HealthInsurance)?
- What is the [duration](TimePeriod) of [health insurance](HealthInsurance)?
- How [long](TimePeriod) is the [health insurance](HealthInsurance) valid?
- What is the [validity](TimePeriod) of [health insurance](HealthInsurance)?
- What is the [duration](TimePeriod) of [health insurance](HealthInsurance)?
- How [long](TimePeriod) is the [health insurance](HealthInsurance) valid?
- What is the [validity](TimePeriod) of [abc health insurance](HealthInsurance)?
- What is the [duration](TimePeriod) of [abc health insurance](HealthInsurance)?
- How [long](TimePeriod) is the [abc health insurance](HealthInsurance) valid?
## intent: vehicle_timeperiod
- What is the [validity](TimePeriod) of [vehicle insurance](VehicleInsurance)?
- What is the [duration](TimePeriod) of [vehicle insurance](VehicleInsurance)?
- How [long](TimePeriod) is the [vehicle insurance](VehicleInsurance) valid?
- What is the [validity](TimePeriod) of [abc vehicle insurance](VehicleInsurance)?
- What is the [duration](TimePeriod) of [abc vehicle insurance](VehicleInsurance)?
- How [long](TimePeriod) is the [abc vehicle insurance](VehicleInsurance) valid?
## intent: vehicle_insuranceclaim
- What are the [documents](Documents) required for [vehicle insurance](VehicleInsurance)?
- What are the [documents](Documents) that i need to apply for [vehicle insurance](VehicleInsurance)?
- What all [documents](Documents) do i need for [vehicle insurance](VehicleInsurance)?
- WHAT [factors](Documents) DO I NEED TO CONSIDER BEFORE APPLYING FOR [vehicle insurance](VehicleInsurance)?
- What are the [documents](Documents) required for [abc vehicle insurance](VehicleInsurance)?
- What are the [documents](Documents) that i need to apply for [abc vehicle insurance](VehicleInsurance)?
- What all [documents](Documents) do i need for [abc vehicle insurance](VehicleInsurance)?
- WHAT [factors](Documents) DO I NEED TO CONSIDER BEFORE APPLYING FOR [abc vehicle insurance](VehicleInsurance)?
## intent: health_insuranceclaim
- What are the [documents](Documents) required for [health insurance](HealthInsurance)?
- What are the [documents](Documents) that i need to apply for [health insurance](HealthInsurance)?
- What all [documents](Documents) do i need for [health insurance](HealthInsurance)?
- WHAT [factors](Documents) DO I NEED TO CONSIDER BEFORE APPLYING FOR [health insurance](HealthInsurance)?
- What are the [documents](Documents) required for [abc health insurance](HealthInsurance)?
- What are the [documents](Documents) that i need to apply for [abc health insurance](HealthInsurance)?
- What all [documents](Documents) do i need for [abc health insurance](HealthInsurance)?
- WHAT [factors](Documents) DO I NEED TO CONSIDER BEFORE APPLYING FOR [abc health insurance](HealthInsurance)?
## intent: health_linking
- How to [link](Link) my aadhar?
- How to [link](Link) my pan card?
- How to [link](Link) my GST?
- Is it possible to [link](Link) my aadhar with my [health insurance](HealthInsurance)?
- Is it possible to [link](Link) my GST with my [health insurance](HealthInsurance)?
- Is it possible to [link](Link) my pan card with my [health insurance](HealthInsurance)?
- Is it necessasary to [link](Link) my aadhar with my [health insurance](HealthInsurance)?
- Is it necessasary to [link](Link) my GST with my [health insurance](HealthInsurance)?
- Is it necessasary to [link](Link) my pan card with my [health insurance](HealthInsurance)?
- How can I [link](Link) my aadhar with my [health insurance](HealthInsurance)?
- How can I [link](Link) my GST with my [health insurance](HealthInsurance)?
- How can I [link](Link) my pan card with my [health insurance](HealthInsurance)?
- How do I [link](Link) my aadhar?
- How do I [link](Link) my GST?
- How do I [link](Link) my pan card?
## intent: vehicle_linking
- How to [link](Link) my aadhar?
- How to [link](Link) my pan card?
- How to [link](Link) my GST?
- Is it possible to [link](Link) my aadhar with my [vehicle insurance](VehicleInsurance)?
- Is it possible to [link](Link) my GST with my [vehicle insurance](VehicleInsurance)?
- Is it possible to [link](Link) my pan card with my [vehicle insurance](VehicleInsurance)?
- Is it necessasary to [link](Link) my aadhar with my [vehicle insurance](VehicleInsurance)?
- Is it necessasary to [link](Link) my GST with my [vehicle insurance](VehicleInsurance)?
- Is it necessasary to [link](Link) my pan card with my [vehicle insurance](VehicleInsurance)?
- How can I [link](Link) my aadhar with my [vehicle insurance](VehicleInsurance)?
- How can I [link](Link) my GST with my [vehicle insurance](VehicleInsurance)?
- How can I [link](Link) my pan card with my [vehicle insurance](VehicleInsurance)?
- How do I [link](Link) my aadhar?
- How do I [link](Link) my GST?
- How do I [link](Link) my pan card?
## intent: generalinfo
- I forgot my [password](Credentials)
- How do I reset my [password](Credentials)?
- I forgot my [username](Credentials)
- I am not able to [login](Credentials) in
- How to update [password](Credentials)?
- Retrieve [username](Credentials)
- Retrieve [password](Credentials)
- How to update [username](Credentials)?
- How to change [username](Credentials)?
- How to change [password](Credentials)?
- Failed to remember [username](Credentials)?
- Failed to remember [password](Credentials)?
## intent: vehicle_sum_assured
- How much [amount](InsuranceSum) is guaranteed for [vehicle insurance](VehicleInsurance)?
- How much [money](InsuranceSum) is guaranteed for [vehicle insurance](VehicleInsurance)?
- What is the minimum [amount](InsuranceSum) of [money](InsuranceSum) assured for [vehicle insurance](VehicleInsurance)?
- How much [insurance sum](InsuranceSum) is guaranteed for [vehicle insurance](VehicleInsurance)?
- How much [insurance sum](InsuranceSum) is guaranteed for [vehicle insurance](VehicleInsurance)?
- What is the minimum [amount](InsuranceSum) of [insurance sum](InsuranceSum) assured for [vehicle insurance](VehicleInsurance)?
## intent: health_sum_assured
- How much [amount](InsuranceSum) is guaranteed for [health insurance](HealthInsurance)?
- How much [money](InsuranceSum) is guaranteed for [health insurance](HealthInsurance)?
- What is the minimum [amount](InsuranceSum) of [money](InsuranceSum) assured for [health insurance](HealthInsurance)?
- How much [insurance sum](InsuranceSum) is guaranteed for [health insurance](HealthInsurance)?
- How much [insurance sum](InsuranceSum) is guaranteed for [health insurance](HealthInsurance)?
- What is the minimum [amount](InsuranceSum) of [insurance sum](InsuranceSum) assured for [health insurance](HealthInsurance)?
## intent: health_sublimit
- What is [health insurance](HealthInsurance) [sub limit](SubLimit)?
- What is the [sub limit](SubLimit) of [health insurance](HealthInsurance)?
- What is [abc health insurance](HealthInsurance) [sub limit](SubLimit)?
- What is the [sub limit](SubLimit) of [abc health insurance](HealthInsurance)?
## intent: vehicle_sublimit
- What is [vehicle insurance](VehicleInsurance) [sub limit](SubLimit)?
- What is the [sub limit](SubLimit) of [vehicle insurance](VehicleInsurance)?
- What is [abc vehicle insurance](VehicleInsurance) [sub limit](SubLimit)?
- What is the [sub limit](SubLimit) of [abc vehicle insurance](VehicleInsurance)?
#intent: insurance_renewal
- When do I [renew](Renew) my [policy](Policy)?
- What are the procedures that is required to [renew](Renew) my [policy](Policy)?
- What are the documents required to [renew](Renew) my [policy](Policy)?
- What are the documents needed for [renewing](Renew) my [policy](Policy)?
- How to [renew](Renew) my [policy](Policy)?
- How can I [renew](Renew) my [policy](Policy)?
- When should I [renew](Renew) my [policy](Policy)?
- Can I [renew](Renew) my insurance [policy](Policy)?
- When do I [renew](Renew) my [isurance policy](Policy)?
- What are the procedures that is required to [renew](Renew) my [isurance policy]?
- What are the documents required to [renew](Renew) my [isurance policy]?
- What are the documents needed for [renewing](Renew) my [isurance policy]?
- How to [renew](Renew) my [isurance policy]?
- How can I [renew](Renew) my [isurance policy]?
- When should I [renew](Renew) my [isurance policy]?
- Can I [renew](Renew) my insurance [isurance policy]?
#intent: vehicle_coverage
- What is the basic [vehicle insurance](VehicleInsurance) [coverage](Coverage)?
- What are the different [vehicle insurance](VehicleInsurance) [coverage](Coverage)?
- What is the [coverage](Coverage) for [vehicle insurance](VehicleInsurance)?
- What is the basic [abc vehicle insurance](VehicleInsurance) [coverage](Coverage)?
- What are the different [abc vehicle insurance](VehicleInsurance) [coverage](Coverage)?
- What is the [coverage](Coverage) for [abc vehicle insurance](VehicleInsurance)?
- What is the basic [vehicle insurance](VehicleInsurance) [insurance coverage](Coverage)?
- What are the different [vehicle insurance](VehicleInsurance) [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for [vehicle insurance](VehicleInsurance)?
- What is the basic [abc vehicle insurance](VehicleInsurance) [insurance coverage](Coverage)?
- What are the different [abc vehicle insurance](VehicleInsurance) [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for [abc vehicle insurance](VehicleInsurance)?
- What is the basic vehicle insurance [coverage](Coverage)?
- What are the different vehicle insurance [coverage](Coverage)?
- What is the [coverage](Coverage) for vehicle insurance?
- What is the basic abc vehicle insurance [coverage](Coverage)?
- What are the different abc vehicle insurance [coverage](Coverage)?
- What is the [coverage](Coverage) for abc vehicle insurance?
- What is the basic vehicle insurance [insurance coverage](Coverage)?
- What are the different vehicle insurance [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for vehicle insurance?
- What is the basic abc vehicle insurance [insurance coverage](Coverage)?
- What are the different abc vehicle insurance [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for abc vehicle insurance?
#intent: health_coverage
- What is the basic [health insurance](HealthInsurance) [coverage](Coverage)?
- What are the different [health insurance](HealthInsurance) [coverage](Coverage)?
- What is the [coverage](Coverage) for [health insurance](HealthInsurance)?
- What is the basic [abc health insurance](HealthInsurance) [coverage](Coverage)?
- What are the different [abc health insurance](HealthInsurance) [coverage](Coverage)?
- What is the [coverage](Coverage) for [abc health insurance](HealthInsurance)?
- What is the basic [health insurance](HealthInsurance) [insurance coverage](Coverage)?
- What are the different [health insurance](HealthInsurance) [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for [health insurance](HealthInsurance)?
- What is the basic [abc health insurance](HealthInsurance) [insurance coverage](Coverage)?
- What are the different [abc health insurance](HealthInsurance) [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for [abc health insurance](HealthInsurance)?
- What is the basic health insurance [coverage](Coverage)?
- What are the different health insurance [coverage](Coverage)?
- What is the [coverage](Coverage) for health insurance?
- What is the basic abc health insurance [coverage](Coverage)?
- What are the different abc health insurance [coverage](Coverage)?
- What is the [coverage](Coverage) for abc health insurance?
- What is the basic health insurance [insurance coverage](Coverage)?
- What are the different [health insurance] [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for health insurance?
- What is the basic abc health insurance [insurance coverage](Coverage)?
- What are the different abc health insurance [insurance coverage](Coverage)?
- What is the [insurance coverage](Coverage) for abc health insurance?
"""
# + colab={"base_uri": "https://localhost:8080/"} id="CV1tjpq4uPDW" outputId="51b193ee-feb1-4c18-9d61-880b4a1a26b6"
# %%writefile domain.yml
intents:
- greet
- health_about
- vehicle_about
- vehicle_timeperiod
- health_timeperiod
- vehicle_benefits
- health_benefits
- affirm
- health_insuranceclaim
- vehicle_insuranceclaim
- generalinfo
- health_sum_assured
- vehicle_sum_assured
- health_sublimit
- vehicle_sublimit
- insurance_renewal
- health_coverage
- vehicle_coverage
slots:
HealthInsurance:
type: categorical
values:
- health insurance
- abc health insurance
VehicleInsurance:
type: categorical
values:
- vehicle insurance
- abc vehicle insurance
Benefits:
type: categorical
values:
- benefits
- tax benefits
- regular benefits
- complementary benefits
TimePeriod:
type: categorical
values:
- timeperiod
- duration
- long
- validity
Documents:
type: categorical
values:
- documents
- factors
Link:
type: categorical
values:
- link
Credentials:
type: categorical
values:
- password
- username
- login
InsuranceSum:
type: categorical
values:
- money
- amount
- insurance sum
SubLimit:
type: categorical
values:
- sub limit
Renew:
type: categorical
values:
- renew
- renewing
Policy:
type: categorical
values:
- policy
- insurance policy
Coverage:
type: categorical
values:
- coverage
- insurance coverage
entities:
- HealthInsurance
- VehicleInsurance
- Benefits
- TimePeriod
- Documents
- Link
- Credentials
- InsuranceSum
- SubLimit
- Renew
- Policy
- Coverage
actions:
- utter_greet
- utter_affirm
- utter_action_health_about
- utter_action_vehicle_about
- utter_action_vehicle_benefits
- utter_action_health_benefits
- utter_action_vehicle_timeperiod
- utter_action_health_timeperiod
- utter_action_health_document
- utter_action_vehicle_document
- utter_action_health_link
- utter_action_vehicle_link
- utter_action_credentials
- utter_action_health_insurancesum
- utter_action_vehicle_insurancesum
- utter_action_health_sublimit
- utter_action_vehicle_sublimit
- utter_action_insurance_renew
- utter_action_health_policy
- utter_action_vehicle_policy
- utter_action_health_coverage
- utter_action_vehicle_coverage
- action_default
- __main__.ApiAction
responses:
utter_greet:
- text: "Welcome"
utter_action_health_about:
- text: "It is health insurance, which can be covered for all medical day-care treatments such as skin treatment and Lasik surgery. The policy covered for 5 yrs. The Free look period is 15 days, the grace period and waiting period is 30 days."
utter_action_vehicle_about:
- text: "It is vehicle insurance, which can be covered for all own damage treatments such as fire, theft, accident. The policy covered for 5 yrs. The Free look period is 15 days, the grace period and waiting period is 30 days."
utter_action_vehicle_benefits:
- text: "a. Renewal Benefits :rn i. Cumulative Bonus (Additional Sum Insured) - An Additional Sum Insured of 10% of Annual Sum Insured provided on each renewal for every claim-free year up to a maximum of 50%.rnii. Complimentary vehicle service Coupons: One coupon per individual policy and two coupons per Floater policy will be offered.rn b. Tax Benefits: Income tax exemption for 1 lakh rupees"
utter_action_health_benefits:
- text: "a. Renewal Benefits :i. a) Cumulative Bonus (Additional Sum Insured) - An Additional Sum Insured of 10% of Annual Sum Insured provided on each renewal for every claim-free year up to a maximum of 50% ii. b) Complementary Health Check-Up Coupons: One coupon per individual policy and two coupons per Floater policy will be offered. b. Tax Benefits: Income tax exemption for 1 lakh rupees "
utter_action_vehicle_timeperiod:
- text: "The policy covered for 5 yrs. The Free look period is 15 days, the grace period and waiting period is 30 days."
utter_action_health_timeperiod:
- text: "The policy covered for 5 yrs. The Free look period is 15 days, the grace period and waiting period is 30 days."
utter_action_health_document:
- text: "a. Medical bills b. Policy copy"
utter_action_vehicle_document:
- text: "a. Driving License copy. b. Original FIR copy c. RC copy of the vehicle with all original keys."
action_default:
- text: "Default"
utter_action_health_link:
- text: "https://www.abc.com/health"
utter_action_vehicle_link:
- text: "https://www.abc.com/vehicle"
utter_action_credentials:
- text: "https://www.abc.com/vehicle/passwordreset"
utter_action_health_insurancesum:
- text: ""
utter_action_vehicle_insurancesum:
- text: ""
utter_action_health_sublimit:
- text: ""
utter_action_vehicle_sublimit:
- text: ""
utter_action_insurance_renew:
- text: ""
utter_action_health_coverage:
- text: ""
utter_action_vehicle_coverage:
- text: ""
utter_action_health_policy:
- text: ""
utter_action_vehicle_policy:
- text: ""
utter_affirm:
- text: "You're welcome"
"""
# + colab={"base_uri": "https://localhost:8080/"} id="reU8ZOWmvGht" outputId="ff1a653b-fb20-46f8-927f-b5873bd642d6"
# %%writefile data/stories.md
## path 0
* greet
- utter_greet
* health_about{"HealthInsurance":"health insurance"}
- utter_action_health_about
* affirm
- utter_affirm
## path 1
* greet
- utter_greet
* health_about{"HealthInsurance":"abc health insurance"}
- utter_action_health_about
* affirm
- utter_affirm
## path 2
* greet
- utter_greet
* vehicle_about{"VehicleInsurance":"vehicle insurance"}
- utter_action_vehicle_about
* affirm
- utter_affirm
## path 3
* greet
- utter_greet
* vehicle_benefits{"Benefits":"benefits"}
- utter_action_vehicle_benefits
* affirm
- utter_affirm
## path 4
* greet
- utter_greet
* vehicle_benefits{ "Benefits":"tax benefits"}
- utter_action_vehicle_benefits
* affirm
- utter_affirm
## path 5
* greet
- utter_greet
* vehicle_benefits{ "Benefits":"regular benefits"}
- utter_action_vehicle_benefits
* affirm
- utter_affirm
## path 6
* greet
- utter_greet
* health_benefits{"Benefits":"benefits"}
- utter_action_health_benefits
* affirm
- utter_affirm
## path 7
* greet
- utter_greet
* health_benefits{ "Benefits":"regular benefits"}
- utter_action_health_benefits
* affirm
- utter_affirm
## path 8
* greet
- utter_greet
* health_benefits{ "Benefits":"complementary benefits"}
- utter_action_health_benefits
* affirm
- utter_affirm
## path 9
* greet
- utter_greet
* vehicle_timeperiod{ "TimePeriod":"timeperiod"}
- utter_action_vehicle_timeperiod
* affirm
- utter_affirm
## path 10
* greet
- utter_greet
* vehicle_timeperiod{ "TimePeriod":"long"}
- utter_action_vehicle_timeperiod
* affirm
- utter_affirm
## path 11
* greet
- utter_greet
* vehicle_timeperiod{ "TimePeriod":"duration"}
- utter_action_vehicle_timeperiod
* affirm
- utter_affirm
## path 12
* greet
- utter_greet
* vehicle_timeperiod{ "TimePeriod":"validity"}
- utter_action_vehicle_timeperiod
* affirm
- utter_affirm
## path 13
* greet
- utter_greet
* health_timeperiod{ "TimePeriod":"timeperiod"}
- utter_action_health_timeperiod
* affirm
- utter_affirm
## path 14
* greet
- utter_greet
* health_timeperiod{ "TimePeriod":"long"}
- utter_action_health_timeperiod
* affirm
- utter_affirm
## path 15
* greet
- utter_greet
* health_timeperiod{ "TimePeriod":"duration"}
- utter_action_health_timeperiod
* affirm
- utter_affirm
## path 16
* greet
- utter_greet
* health_timeperiod{ "TimePeriod":"validity"}
- utter_action_health_timeperiod
* affirm
- utter_affirm
## path 17
* greet
- utter_greet
* health_insuranceclaim{ "Documents":"documents"}
- utter_action_health_document
* affirm
- utter_affirm
## path 18
* greet
- utter_greet
* health_insuranceclaim{ "Documents":"factors"}
- utter_action_health_document
* affirm
- utter_affirm
## path 19
* greet
- utter_greet
* vehicle_insuranceclaim{ "Documents":"documents"}
- utter_action_health_document
* affirm
- utter_affirm
## path 20
* greet
- utter_greet
* vehicle_insuranceclaim{ "Documents":"factors"}
- utter_action_health_document
* affirm
- utter_affirm
## path 21
* greet
- utter_greet
* health_timeperiod{ "Link":"link"}
- utter_action_health_link
* affirm
- utter_affirm
## path 22
* greet
- utter_greet
* health_timeperiod{ "Link":"link"}
- utter_action_vehicle_link
* affirm
- utter_affirm
## path 23
* greet
- utter_greet
* generalinfo{ "Credentials":"password"}
- utter_action_credentials
* affirm
- utter_affirm
## path 24
* greet
- utter_greet
* generalinfo{ "Credentials":"username"}
- utter_action_credentials
* affirm
- utter_affirm
## path 25
* greet
- utter_greet
* generalinfo{ "Credentials":"login"}
- utter_action_credentials
* affirm
- utter_affirm
## path 26
* greet
- utter_greet
* health_sum_assured{ "InsuranceSum":"amount"}
- utter_action_health_insurancesum
* affirm
- utter_affirm
## path 27
* greet
- utter_greet
* health_sum_assured{ "InsuranceSum":"insurance sum"}
- utter_action_health_insurancesum
* affirm
- utter_affirm
## path 28
* greet
- utter_greet
* health_sum_assured{ "InsuranceSum":"money"}
- utter_action_health_insurancesum
* affirm
- utter_affirm
## path 29
* greet
- utter_greet
* vehicle_sum_assured{ "InsuranceSum":"amount"}
- utter_action_vehicle_insurancesum
* affirm
- utter_affirm
## path 30
* greet
- utter_greet
* vehicle_sum_assured{ "InsuranceSum":"insurance sum"}
- utter_action_vehicle_insurancesum
* affirm
- utter_affirm
## path 31
* greet
- utter_greet
* vehicle_sum_assured{ "InsuranceSum":"money"}
- utter_action_vehicle_insurancesum
* affirm
- utter_affirm
## path 32
* greet
- utter_greet
* health_sublimit{ "SubLimit":"sub limit"}
- utter_action_health_sublimit
* affirm
- utter_affirm
## path 33
* greet
- utter_greet
* vehicle_sublimit{ "SubLimit":"sub limit"}
- utter_action_vehicle_sublimit
* affirm
- utter_affirm
## path 34
* greet
- utter_greet
* insurance_renewal{ "Renew":"renew"}
- utter_action_insurance_renew
* affirm
- utter_affirm
## path 35
* greet
- utter_greet
* insurance_renewal{ "Renew":"renewing"}
- utter_action_insurance_renew
* affirm
- utter_affirm
## path 36
* greet
- utter_greet
* insurance_renewal{ "Policy":"policy"}
- utter_action_health_policy
* affirm
- utter_affirm
## path 37
* greet
- utter_greet
* insurance_renewal{ "Policy":"insurance policy"}
- utter_action_health_policy
* affirm
- utter_affirm
## path 38
* greet
- utter_greet
* insurance_renewal{ "Policy":"policy"}
- utter_action_vehicle_policy
* affirm
- utter_affirm
## path 39
* greet
- utter_greet
* insurance_renewal{ "Policy":"insurance policy"}
- utter_action_vehicle_policy
* affirm
- utter_affirm
## path 40
* greet
- utter_greet
* health_coverage{ "Coverage":"coverage"}
- utter_action_health_coverage
* affirm
- utter_affirm
## path 41
* greet
- utter_greet
* health_coverage{ "Coverage":"insurance coverage"}
- utter_action_health_coverage
* affirm
- utter_affirm
## path 42
* greet
- utter_greet
* vehicle_coverage{ "Coverage":"coverage"}
- utter_action_vehicle_coverage
* affirm
- utter_affirm
## path 43
* greet
- utter_greet
* vehicle_coverage{ "Coverage":"insurance coverage"}
- utter_action_vehicle_coverage
* affirm
- utter_affirm
## path 1000
* affirm
- utter_affirm
# + colab={"base_uri": "https://localhost:8080/"} id="Twp9eGVaregY" outputId="a3723046-8a96-4ee4-aa2a-d1b7a49533fd"
# + colab={"base_uri": "https://localhost:8080/"} id="jMaPFn6zrkuF" outputId="810d8404-0c74-4fc6-b475-5ff8c6b0c65b"
model_path = rasa.train(domain, config, [training_files], output)
print(model_path)
# + id="U8BkV8wProLw"
endpoints = "endpoints.yml"
chat(model_path, endpoints)
# + id="_djKlxmksQ0N"
chat(model_path, endpoints)
# + colab={"base_uri": "https://localhost:8080/"} id="236vSfSdsfdY" outputId="faea8af4-5031-4db9-d70e-28812a8acb5e"
import rasa.data as data
stories_directory, nlu_data_directory = data.get_core_nlu_directories(training_files)
print(stories_directory, nlu_data_directory)
# + id="Vkf48LYAskce"
rasa.test(model_path, stories_directory, nlu_data_directory)
print("Done testing...")
# + id="cB43vU97soGw"
|
DTI5125_chatbot.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from tools import *
from models import *
import plotly.graph_objects as go
import plotly.figure_factory as ff
from Bio.SeqUtils import GC
from Bio import SeqIO
import os
from random import sample
from plotly.subplots import make_subplots
import pickle
from scipy import stats
from collections import Counter
plt.ioff()
import warnings
warnings.filterwarnings('ignore')
# +
#RECORDING THE PERFORMANCE
TFs = ["JUND", "HNF4A", "MAX", "SP1", "SPI1"]
results = {}
real_bm_include_target = {}
real_bm_no_target = {}
fake_bm_include_target = {}
fake_bm_no_target = {}
for TF in TFs:
real_bm_include_target[TF] = []
real_bm_no_target[TF] = []
fake_bm_include_target[TF] = []
fake_bm_no_target[TF] = []
for i in range(1,6):
pkl_file = open("../RESULTS_BM_SUBSAMPLE_R_DANQ_True_I_True/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_true = pickle.load(pkl_file)
pkl_file.close()
real_bm_include_target[TF].append(list(mccoef_true_true.values())[0])
pkl_file = open("../RESULTS_BM_SUBSAMPLE_R_DANQ_True_I_False/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_false = pickle.load(pkl_file)
pkl_file.close()
real_bm_no_target[TF].append(list(mccoef_true_false.values())[0])
pkl_file = open("../RESULTS_BM_SUBSAMPLE_R_DANQ_False_I_True/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_false_true = pickle.load(pkl_file)
pkl_file.close()
fake_bm_include_target[TF].append(list(mccoef_false_true.values())[0])
pkl_file = open("../RESULTS_BM_SUBSAMPLE_R_DANQ_False_I_False/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_false_false = pickle.load(pkl_file)
pkl_file.close()
fake_bm_no_target[TF].append(list(mccoef_false_false.values())[0])
real_bm_include_target = pd.Series(real_bm_include_target)
real_bm_no_target = pd.Series(real_bm_no_target)
fake_bm_include_target = pd.Series(fake_bm_include_target)
fake_bm_no_target = pd.Series(fake_bm_no_target)
# +
results = {}
real_cofactor_include_target = {}
real_cofactor_no_target = {}
for TF in TFs:
real_cofactor_include_target[TF] = []
real_cofactor_no_target[TF] = []
for i in range(1,6):
pkl_file = open("../RESULTS_COFACTOR_DANQ_SUBSAMPLE_I_True/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_true = pickle.load(pkl_file)
pkl_file.close()
real_cofactor_include_target[TF].append(list(mccoef_true_true.values())[0])
pkl_file = open("../RESULTS_COFACTOR_DANQ_SUBSAMPLE_I_False/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_false = pickle.load(pkl_file)
pkl_file.close()
real_cofactor_no_target[TF].append(list(mccoef_true_false.values())[0])
real_cofactor_include_target = pd.Series(real_cofactor_include_target)
real_cofactor_no_target = pd.Series(real_cofactor_no_target)
# +
results = {}
real_string_include_target = {}
real_string_no_target = {}
for TF in TFs:
real_string_include_target[TF] = []
real_string_no_target[TF] = []
for i in range(1,6):
pkl_file = open("../RESULTS_STRING_DANQ_SUBSAMPLE_I_True/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_true = pickle.load(pkl_file)
pkl_file.close()
real_string_include_target[TF].append(list(mccoef_true_true.values())[0])
pkl_file = open("../RESULTS_STRING_DANQ_SUBSAMPLE_I_False/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_false = pickle.load(pkl_file)
pkl_file.close()
real_string_no_target[TF].append(list(mccoef_true_false.values())[0])
real_string_include_target = pd.Series(real_string_include_target)
real_string_no_target = pd.Series(real_string_no_target)
# +
real_lowcorbm_include_target = {}
real_lowcorbm_no_target = {}
for TF in TFs:
real_lowcorbm_include_target[TF] = []
real_lowcorbm_no_target[TF] = []
for i in range(1,6):
pkl_file = open("../RESULTS_LOWCORBM_DANQ_SUBSAMPLE_I_True/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_true = pickle.load(pkl_file)
pkl_file.close()
real_lowcorbm_include_target[TF].append(list(mccoef_true_true.values())[0])
pkl_file = open("../RESULTS_LOWCORBM_DANQ_SUBSAMPLE_I_False/"+
TF+"_"+str(i)+"/mccoef.pkl", 'rb')
mccoef_true_false = pickle.load(pkl_file)
pkl_file.close()
real_lowcorbm_no_target[TF].append(list(mccoef_true_false.values())[0])
real_lowcorbm_include_target = pd.Series(real_lowcorbm_include_target)
real_lowcorbm_no_target = pd.Series(real_lowcorbm_no_target)
# +
fig = go.Figure()
TF = "SPI1"
fig.add_trace(go.Box(
y=real_bm_include_target[TF],
x=[TF]*5,
name='Same binding mode',
marker_color='blue',
showlegend=True
))
fig.add_trace(go.Box(
y=real_cofactor_include_target[TF],
x=[TF]*5,
name='Co-factors',
marker_color='darkblue',
showlegend=True
))
fig.add_trace(go.Box(
y=real_lowcorbm_include_target[TF],
x=[TF]*5,
name='Same binding mode (low correlation)',
marker_color='magenta',
showlegend=True
))
fig.add_trace(go.Box(
y=real_string_include_target[TF],
x=[TF]*5,
name='STRING partners',
marker_color='coral',
showlegend=True
))
fig.add_trace(go.Box(
y=fake_bm_include_target[TF],
x=[TF]*5,
name='Random',
marker_color='seagreen',
showlegend=True
))
###########################################
fig.add_trace(go.Box(
y=real_bm_no_target[TF],
x=[TF]*5,
name='Same binding mode',
marker_color='blue',
showlegend=False
))
fig.add_trace(go.Box(
y=real_cofactor_no_target[TF],
x=[TF]*5,
name='Co-factors',
marker_color='darkblue',
showlegend=False
))
fig.add_trace(go.Box(
y=real_lowcorbm_no_target[TF],
x=[TF]*5,
name='Same binding mode (low correlation)',
marker_color='magenta',
showlegend=False
))
fig.add_trace(go.Box(
y=real_string_no_target[TF],
x=[TF]*5,
name='STRING partners',
marker_color='coral',
showlegend=False
))
fig.add_trace(go.Box(
y=fake_bm_no_target[TF],
x=[TF]*5,
name='Random',
marker_color='seagreen',
showlegend=False
))
fig.update_layout(title='',
plot_bgcolor='rgba(0,0,0,0)', paper_bgcolor='rgba(0,0,0,0)',
boxmode='group',
font=dict(
family="Courier New, monospace",
size=18,
color="black"
))
fig.update_layout(legend=dict(
yanchor="top",
y=0.99,
xanchor="right",
x=1.4,
font=dict(
size=10,
color="black"
)
))
#fig.update_layout(autosize=False,width=500,height=333)
fig.update_yaxes(range=[0, 1], title= 'MCC')
fig.update_xaxes(showline=True, linewidth=2, linecolor='black',
tickfont=dict(size=18))
fig.update_yaxes(showline=True, linewidth=2, linecolor='black')
fig.show()
|
notebooks/TL_exploring_pentad_TFs_DanQ.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setting up Python on your own computer
#
# We're going to walk through the steps needed to install all of the components used during this boot camp -- mainly, Python 3, `virtualenv` and `virtualenvwrapper`. If you get stuck, [please contact me](mailto:<EMAIL>) and I'll help you get things sorted out.
#
# For Mac/Linux, we're going to mostly follow [<NAME>'s excellent Python installation guides](http://docs.python-guide.org/en/latest/starting/installation/). For PCs, we'll be working through [Anthony DeBarros's guide to installing Python on Windows](http://www.anthonydebarros.com/2015/08/16/setting-up-python-in-windows-10/).
# ## Other resources:
#
# - [The Hitchhiker's Guide to Python](http://docs.python-guide.org/en/latest/)
# - [Think Python](http://www.greenteapress.com/thinkpython/html/index.html)
# - [Useful pandas snippets](http://www.swegler.com/becky/blog/2014/08/06/useful-pandas-snippets/)
# - [First Python Notebook tutorial](http://www.firstpythonnotebook.org/)
# - [Codecademy's Python lessons](https://www.codecademy.com/learn/learn-python)
# - [The Python style guide](https://www.python.org/dev/peps/pep-0008/)
|
exercises/18. Setting up Python on your own computer-working.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # ODE RLC circuit
# This notebook shows how to solve numerically a second order ODE of a RLC series circuit.
# <img src="images/RLC_series_circuit_v1.png" alt="Drawing" style="width: 300px;"/>
# We define
#
# - $u(t)$ input voltage
# - $x(t)$ voltage over the capacitor to the ground
from scipy.integrate import solve_ivp
import numpy as np
import matplotlib.pyplot as plt
# We have a second order ODE:
#
# \begin{align}
# LC \ddot{x} + RC \dot{x} + x = u(t)
# \end{align}
# In oder to calculate a numerical solution we have to rearrange the second order equation into a system of first order ODEs. A system of first order ODEs can be solved with Euler method.
# We rewrite the ODE so that the highest order is the first - we substitute $v=\dot{x}$:
#
# \begin{align}
# LC \dot{v} + RC v + x = u(t)
# \end{align}
# The final ODE system is:
# \begin{align}
# \dot{v} = \frac{u(t) - RC v - x}{LC} \\
# \dot{x} = v
# \end{align}
# imitates European line voltage
def input_signal(t):
return 230*np.sin(2*np.pi*100*t)
R = 100; C = 1e-3; L = 100e-3
def ode_right_side(t, vars):
v, x = vars[0], vars[1]
vdot = (input_signal(t) - R*C*v - x)/(L*C)
xdot = v
return np.r_[vdot, xdot]
initial_state = np.r_[0, 0]
integral_range = [0, 1]
eval_times = np.linspace(0, 1, 10000)
result = solve_ivp(ode_right_side, integral_range, y0=initial_state, t_eval=eval_times)
plt.figure(figsize=(15,5))
plt.plot(result.t, result.y[1, :], label="x(t)")
plt.xlabel("time [s]")
plt.ylabel("voltage [V]")
plt.legend()
plt.xlim([0, 0.5])
# imitates step function from t > 0
def input_signal(t):
return 230
result = solve_ivp(ode_right_side, integral_range, y0=initial_state, t_eval=eval_times)
plt.figure(figsize=(15,5))
plt.plot(result.t, result.y[1, :], label="x(t)")
plt.xlabel("time [s]")
plt.ylabel("voltage [V]")
plt.legend()
plt.xlim([0, 0.5])
|
ODE simulation RLC circuit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# +
# Import standard libraries
import csv
import OmicsIntegrator as oi
import pandas as pd
import numpy as np
import scipy.stats as ss
import networkx as nx
import matplotlib.pyplot as plt
from matplotlib_venn import venn2, venn3
import pickle
import tqdm
import time
# Import custom libraries
import drugs2
import prizes
import sensitivity_analysis as sensitivity
import sensitivity_sarspartners as svp
import neighborhoods as nbh
# -
# # Create prized list
# ## Select IAV differentially expressed genes
# Load IAV DE genes data
iav_de_genes_file_name = '../Data/iav_genes.csv'
iav_genes_df = pd.read_csv(iav_de_genes_file_name)
# Minor reformatting
iav_genes_df.columns = ['name','log2FC_blanco','log2FC_ageing']
iav_genes_df = iav_genes_df.replace([np.inf, -np.inf], np.nan)
iav_genes_df = iav_genes_df.dropna(subset = ['log2FC_blanco','log2FC_ageing'], how='any')
iav_genes_df['name'] = iav_genes_df['name'].str.upper().str.strip()
# Keep only protein coding genes
protein_coding_genes_file_name = '../Data/protein_coding_ensembl_gene_id_hgnc_hg19.txt'
coding_genes = prizes.load_protein_coding_genes(protein_coding_genes_file_name)
iav_genes_df = iav_genes_df.merge(coding_genes, on = 'name', how = 'inner')
iav_genes_df.head()
# ## Create final prized list
terminal_df = iav_genes_df.copy()
terminal_df.insert(1,'prize', np.abs(terminal_df['log2FC_blanco']))
terminal_df.sort_values(by='prize', ascending=False, inplace=True)
terminal_df.head()
# Plot histogram of prizes
plt.figure()
plt.hist(terminal_df['prize'], bins=50)
plt.plot()
# Reduce number of terminals to top 150 differentially expressed genes (see red vertical line in histogram above)
terminal_df = terminal_df.iloc[:150]
terminal_df.head()
# Save terminal df to tsv
terminal_df.to_csv(r'../Save_iav_noage/terminals_ppi_analysis.tsv', header=True, index=None, sep='\t', quoting = csv.QUOTE_NONE, escapechar = '\t')
# # Prepare the sentivity analysis for Stein tree parameters
# ## W range
# Load prizes data
prizes_data = terminal_df
terminals = list(prizes_data['name'])
n_terminals = len(terminals)
# Load IREF interactome
interactome_file = "../Data/iRefIndex_v14_MIScore_interactome_C9.costs.allcaps.txt"
graph = oi.Graph(interactome_file)
# Distribution of cheapest path between any two terminals without penalty (g=-\infty)
network = graph.interactome_graph
shortest_dist_mat = np.zeros(shape = (n_terminals,n_terminals))
for ix_prot1 in tqdm.tqdm(np.arange(n_terminals)):
time.sleep(0.01)
for ix_prot2 in np.arange(ix_prot1+1, n_terminals, 1):
shortest_dist_mat[ix_prot1,ix_prot2] = nx.dijkstra_path_length(network,
source = terminals[ix_prot1],
target = terminals[ix_prot2],
weight = 'cost')
shortest_dist_mat[ix_prot2,ix_prot2] = shortest_dist_mat[ix_prot1,ix_prot2]
# Plot heatmap of cheapest path
plt.matshow(np.transpose(shortest_dist_mat))
plt.xlabel('terminals')
plt.ylabel('terminals')
plt.colorbar()
# Plot histogram of cheapest path
plt.subplot(1,2,1)
flat_mat = shortest_dist_mat[np.triu_indices(n_terminals, k=1)]
plt.hist(flat_mat,40)
plt.xlabel('Cost of shortest path between two terminals')
plt.ylabel('Number of pairs of terminals')
# ## G range
# Create a dictionary containing edge costs under several choices of g
g_range = [0,1,2,3,4,5]
edge_costs_dict={'g = -Inf': graph.edge_costs}
for g in g_range:
params = {"w": 0, "b": 1, "g": g, "edge_noise": 0, "dummy_mode": "terminals", "seed": 0, "skip_checks": False}
graph._reset_hyperparameters(params)
edge_costs_dict['g='+str(g)] = graph.costs
edge_costs_df = pd.DataFrame(data = edge_costs_dict)
# Boxplot of edge costs under several choices of g
plt.figure()
edge_costs_df.boxplot()
plt.yscale('log')
plt.ylabel('penalized edge cost')
plt.title('Boxplots of penalized edge costs in IREF for different values of g')
plt.show()
# ## B range
# For a range of g, compute the maximum penalized edge cost
g_range = [0,1,2,3,4,5]
max_penalized_edge_cost = pd.DataFrame(edge_costs_df.max())
max_penalized_edge_cost.columns=['max_penalized_edge_cost']
max_penalized_edge_cost
# # Sensitivity analysis for Steiner tree parameters
# ## Run sensitivity analysis
# Load set of virus interacting genes from Gordon et al.
virus_partners_file_name = "../Data/SARSCov_targets_df.tsv"
virus_interacting_genes = sensitivity.import_virus_partners(virus_partners_file_name)
# Parameters for sensitivity analysis
interactome_file_name = "../Data/iRefIndex_v14_MIScore_interactome_C9.costs.allcaps.txt"
prize_file_name = "../Save_iav_noage/terminals_ppi_analysis.tsv"
# Graph hyperparameters
graph_params = {
"noise": 0.0,
"dummy_mode": "terminals",
"exclude_terminals": False,
"seed": 1,
"pruning": 'strong',
"verbosity_level": 0
}
# Set sweeping parameters, i.e. configurations (w,b,g)
W_list = np.linspace(start = 0.2, stop = 2, num = 10)
B_list = np.array([5., 10., 15., 20., 25., 30., 35., 40., 45., 50.])
# Run sensitivity analysis
networks_dict = sensitivity.run_sensitivity_analysis(interactome_file_name,
prize_file_name,
graph_params,
W_list,
B_list,
G=0)
# Save as pickle
with open("../Save_iav_noage/networks_dict.pkl", "wb") as f:
pickle.dump(networks_dict, f)
# Add metatadata
networks_dict = pickle.load(open("../Save_iav_noage/networks_dict.pkl", "rb"))
networks_dict = sensitivity.add_metadata(networks_dict, virus_interacting_genes)
# Make summary
networks_summary_df = sensitivity.make_summary(networks_dict, n_terminals, g=0)
networks_summary_df.head()
# ## Plot node stability heatmaps
# Stability of selected nodes
mat_allnodes = sensitivity.create_matrix_gene_overlap_between_networks(networks_summary_df, networks_dict)
plt.figure()
plt.matshow(mat_allnodes)
plt.xlabel('networks')
plt.ylabel('networks')
plt.title('Nodes stability')
plt.colorbar()
plt.show()
# Stability of selected terminals
mat_terminals = sensitivity.create_matrix_terminal_overlap_between_networks(networks_summary_df, networks_dict)
plt.figure()
plt.matshow(mat_terminals)
plt.xlabel('networks')
plt.ylabel('networks')
plt.title('Terminals stability')
plt.colorbar()
plt.show()
# ## Select robust parameters
# Select network corresponding to g=0, w=1.4 and b=40
index_selected = 67
paramstring_selected = networks_summary_df[networks_summary_df['index']==index_selected].index[0]
network_selected = networks_dict[paramstring_selected]
# Save selected network to file
oi.output_networkx_graph_as_interactive_html(network_selected, filename="../Save_iav_noage/network_selected.html")
oi.output_networkx_graph_as_pickle(network_selected, filename= '../Save_iav_noage/network_selected.pickle')
#oi.output_networkx_graph_as_graphml_for_cytoscape(robust_network, filename= '../Save_iav_noage/network_selected.graphml')
# # Add drug targets to selected network
# ## Construct drug/target data
# Compute degree centrality for all nodes in the interactome
graph = oi.Graph(interactome_file_name)
centrality_dic = nx.degree_centrality(graph.interactome_graph)
# Load drug/target data from DrugCentral
drug_target_file_name = '../Data/drug.target.interaction.tsv'
drugcentral_df = drugs2.load_drug_target_data(drug_target_file_name, aff_cst_thresh=5)
drugcentral_df['degree_centrality'] = [centrality_dic[gene] if (gene in list(centrality_dic.keys())) else None for gene in drugcentral_df['gene']]
drugcentral_df.head()
# Construct table that gives the number of targets per drug
num_targets_df0 = drugcentral_df.groupby('drug', as_index=False)[['gene']].agg({
('num_targets','count'),
('num_terminal_targets',lambda gs: len(set(gs).intersection(set(terminals))))
})
max_centrality_df0 = drugcentral_df.groupby('drug', as_index=False)[['degree_centrality']].agg({
('max_degree_centrality',max)
})
num_targets_df = pd.DataFrame({'drug': num_targets_df0.index,
'num_targets': num_targets_df0['gene']['num_targets'],
'num_terminal_targets': num_targets_df0['gene']['num_terminal_targets'],
'max_target_centrality': max_centrality_df0['degree_centrality']['max_degree_centrality']}).reset_index(drop=True)
num_targets_df = num_targets_df.sort_values(by='num_targets', ascending=False, inplace=False)
num_targets_df.head(10)
# Histogram of number of targets per drug
plt.figure()
plt.hist(num_targets_df['num_targets'],50)
plt.yscale('log')
plt.xlabel('Number of targets')
plt.ylabel('Count')
plt.plot()
# +
# Load L1000 drugs with correlations
embedded_drugs_file_name = '../Data/iav_correlations_autoencoder_space.txt'
bestdrugs_df = pd.read_csv(embedded_drugs_file_name, header=None)
bestdrugs_df.columns = ['drug','corr']
bestdrugs_df['drug'] = bestdrugs_df['drug'].str.strip("()' ").str.lower()
bestdrugs_df['corr'] = pd.to_numeric(bestdrugs_df['corr'].str.strip("() "))
bestdrugs_df.sort_values(by='corr', axis=0, ascending=False, inplace=True)
# Plot histogram of correlations
plt.figure()
plt.hist(bestdrugs_df['corr'],50)
plt.xlabel('anticorrelation')
plt.ylabel('count')
plt.show()
# Select top drugs (most anticorrelated with IAV signature)
bestdrugs_df = bestdrugs_df.iloc[:142] # so that we have the same number (142) of selected drugs as in the A549-SARS-Cov-2 analysis
bestdrugs_df.head()
# -
# Merge L1000 drugs with DrugCentral drug/target dataset
targets_and_drugs_df = drugcentral_df.merge(bestdrugs_df, on = 'drug', how = 'inner')
targets_and_drugs_df.head()
# ## Add drug target information to selected network
network_selected = pickle.load(open('../Save_iav_noage/network_selected.pickle', "rb"))
network_selected = drugs2.add_drug_info_to_selected_network(network_selected, targets_and_drugs_df)
# Save enriched network as pickle
oi.output_networkx_graph_as_pickle(network_selected, filename= '../Save_iav_noage/network_selected_with_drug_info.pickle')
# Construct table of drug targets in the network
drug_targets_df = drugs2.drug_targets_in_selected_network(network_selected)
drug_targets_df.to_csv(r'../Save_iav_noage/drug_targets_in_network.tsv', header=True, index=None, sep='\t', quoting = csv.QUOTE_NONE, escapechar = '\t')
drug_targets_df['affinity'] = pd.to_numeric(drug_targets_df['affinity'], errors='coerce')
drug_targets_df.dropna(subset=['affinity'], inplace=True)
drug_targets_df['terminal'] = drug_targets_df['name'].isin(terminals)
drug_targets_df
# Save drug/target dataframe to csv
drug_targets_df.to_csv(r'../Save_iav_noage/final_drug_target_table.tsv', header=True, index=None, sep='\t', quoting = csv.QUOTE_NONE, escapechar = '\t')
# # Compare to A549 cell type
# Drug/target dataframe for A549
drug_targets_df_a549_file_name = '../Save/final_drug_target_table.tsv'
drug_targets_df_a549 = pd.read_csv(drug_targets_df_a549_file_name, sep = '\t')
drug_targets_df_a549.head()
# Common gene targets
targets_iav = set(drug_targets_df['name'])
targets_a549 = set(drug_targets_df_a549['name'])
venn2(subsets = [targets_iav,targets_a549],set_labels = ('Gene targets (IAV, no age)','Gene targets (A549)'))
print(targets_iav.intersection(targets_a549))
# +
# Common drugs
drugs_iav = set(drug_targets_df['drug'])
drugs_a549 = set(drug_targets_df_a549['drug'])
venn2(subsets = [drugs_iav,drugs_a549],set_labels = ('IAV drugs','A549 drugs'))
a549_minus_iav = num_targets_df.loc[num_targets_df['drug'].isin(drugs_a549.difference(drugs_iav))]
iav_minus_a549 = num_targets_df.loc[num_targets_df['drug'].isin(drugs_iav.difference(drugs_a549))]
a549_inter_iav = num_targets_df.loc[num_targets_df['drug'].isin(drugs_a549.intersection(drugs_iav))]
print(a549_minus_iav)
print(iav_minus_a549)
print(a549_inter_iav)
# -
|
Code_ppi/Code/.ipynb_checkpoints/SteinerTree_notebook-IAVNoAge-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mowhamadrexa/EECS3481Assignments/blob/main/Q10_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="q5C_tBB0MciF"
# q=71, alpha = 7
# + [markdown] id="U1zC0OIaMrhR"
# # a)
# + [markdown] id="wtKlXZ4UN6dN"
# $X_{a} = 15$
# + colab={"base_uri": "https://localhost:8080/"} id="PbyLfY1bMtBZ" outputId="fa3b09cb-5b70-463a-ac32-0a7d33bdd8f1"
q = 71
a = 7
Xa = 15
Ya = a**Xa % q
Ya
# + [markdown] id="eqZWCXt_NH2p"
# # b)
# + [markdown] id="TwLlIyBNN9n5"
# $X_{b} = 27$
# + colab={"base_uri": "https://localhost:8080/"} id="MTo-qY3TNJ67" outputId="3218f65f-5afb-419e-a633-835a5a5c1db6"
q = 71
a = 7
Xb = 27
Yb = a**Xb % q
Yb
# + [markdown] id="AwYfFb4mN2nj"
# # c)
# + colab={"base_uri": "https://localhost:8080/"} id="mIFg1jPON4DK" outputId="46f90b2b-9c33-498d-95d6-cc3286cc0707"
AliceSharedKey = Yb**Xa % q
AliceSharedKey
# + colab={"base_uri": "https://localhost:8080/"} id="ACOVGFZ5Oj9h" outputId="c37fe923-fa6f-4c23-c2c5-61f99d5209f4"
BobSharedKey = Ya**Xb % q
BobSharedKey
# + [markdown] id="zfmdzQkYOzLS"
# The shared key is 34
|
Q10_1.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# # Redfish schemas: Why should I care
#
# Powered by [HPE DEV Team](hpedev.io)
#
# ### Speakers : <NAME>
#
#
# Version 0.1
#
# <img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 125px;"/>
#
# ## Introduction
#
# This Jupyter notebook defines environment variables that will be used through the rest of the notebook. Then, it explains how Redfish schemas can help you in your development. For didactic reasons, commands presented in this notebook are not optimized.
#
# ## Create environment variables
#
# The following `bash` code defines environment variables (i.e. IP address, username, password....) depending on your student ID number stored in variable `$Stud`. It creates as well a `.json` file containing the credentials of your OpenBMC appliance required to open a Redfish session.
#
# Click in the cell below and then click on the
# <img src="Pictures/RunButton.png" style="display:inline;width=55px;height=25px"/>
# icon to create the environment variables and the json file.
# +
# Set Student ID number
Stud="00"
echo "You are Student $Stud"
# Create BMC location/ports variables
BmcIP=openbmcs:443${Stud} # OpenBMC simulator
BmcURI="https://${BmcIP}"
# BMC Administrator credentials
BmcUser="student"
BmcPassword='<PASSWORD>!'
# Minimum required Redfish headers
HeaderODataVersion="OData-Version: 4.0"
HeaderContentType="Content-Type: application/json"
# Data files
ResponseHeaders="ResponseHeaders.txt" # Used to hold HTTP response headers
SessionData="./CreateSession-data.json" # Body/Workload used to create the Redfish session
cat > ${SessionData} << __EOF__
{
"UserName": "$BmcUser",
"Password": <PASSWORD>"
}
__EOF__
# Verify we can reach the remote Bmc
curl --insecure --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--request GET ${BmcURI}/redfish | jq &>>/dev/null || echo "WARNING: Problem reaching the remote BMC"
# -
# ## Retrieve the Redfish Service Entry point content (Root)
#
# The Redfish Service Entry point is **`/redfish/v{RedfishVersion}/`**.
#
# Run the next cell to retrieve Redfish version(s) available today in your OpenBMC.
#
# This request does not require any authentication.
#
# If you are not familiar with cURL, get help from its [manual page](https://curl.haxx.se/docs/manpage.html).
# ## Create a Redfish session using cURL
#
#
# All the URIs below the Root entry point requires authentication. In this section you'll go through the session authentication method as it may differ from other Rest APIs (i.e. OneView).
#
# The following `curl` command sends a `POST` request toward the standard `/redfish/v1/SessionService/Sessions` URI of your BMC. The body/workload of this request is in the `@${SessionData}` json file populated in the very first `bash` cell of this notebook (Environment variables). You can view its content by clicking on it from the left pane of your Jupyter environment.
#
# Select and run the following cell.
#
# If this `POST` request is successful, the BMC sends back a `Token` and a `Session Location` in the **headers of the response**. Response headers are saved in the `$ResponseHeaders` text file now present in the left pane of your Jupyter environment.
#
# +
echo 'Create Session and print body response:'
curl --dump-header $ResponseHeaders \
--insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--request POST --data "@$SessionData" \
${BmcURI}/redfish/v1/SessionService/Sessions | jq
BmcToken=$(awk '/X-Auth-Token/ {print $NF}' $ResponseHeaders | tr -d '\r')
BmcSessionLocation="$BmcURI"$(awk '/^Loca.*Se/ {gsub("https://.*/red", "/red", $NF);print $NF}' $ResponseHeaders | tr -d '\r')
echo "Bmc Token: $BmcToken"
echo -e "Bmc Session Location: $BmcSessionLocation\n"
# -
# The following cell extracts the name of the BMCs present in your system and then, for each BMC it extracts its properties.
#
# Run it and review the properties returned by your OpenBMC. Among them you should notice the `Actions` and the `Oem` resources which need some explanation.
#
# The `Actions` collection contains all the possible actions that can be performed on your BMC; With this version of OpenBMC, you can perform a reset of the BMC by posting the value `GracefulRestart` at `/redfish/v1/Managers/bmc/Actions/Manager.Reset`. You'll do this later.
#
# The `Oem` collection contains resources specific to `OpenBmc` and not part of the Redfish standard. This is a way to allow computer makers to expose their specific and added value resources to the Rest API.
# +
BmcList=$(curl --insecure --silent --noproxy "localhost, 127.0.0.1" \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET ${BmcURI}/redfish/v1/Managers | jq '.Members[]' | \
awk -F/ '/@odata.id/ {print $NF}' | tr -d '"' )
echo "List of BMC(s) present in this system:"
echo -e "$BmcList\n"
for bmc in $BmcList ; do
echo "Properties of BMC: $bmc"
curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET ${BmcURI}/redfish/v1/Managers/${bmc} | jq
done
# -
# If you want to view the network protocols supported by your BMC, you can retrieve them with a `GET` of the `NetworkProtocol` URI mentionned in the output of the above `GET` request.
#
# Run the next `curl`command. Its output should show an empty array of `NTPServers` (if not, contact your instructor). It contains as well the **Type** of the resources in this sub-tree: `@odata.type = #ManagerNetworkProtocol.v1_4_0.ManagerNetworkProtocol`.
#
# Said differently, the `NetworkProtocol` resources falls in the **`ManagerNetworkProtocol`**. You will need this info later.
#
echo "Network Protocol configuration:"
curl --insecure --silent --noproxy "localhost, 127.0.0.1" \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET ${BmcURI}/redfish/v1/Managers/${bmc}/NetworkProtocol | jq
# As it is always good to have the correct date and time in a BMC, you may want to supply at least one server IP in the `NTPServers` array of your BMC. To reach that goal you have first to verify in the Redfish Schema whether the `NTPServers` array can be modified.
#
# Generally speaking the location of the Redfish Schemas of a particular OData type is under the `/redfish/v1/JsonSchemas/{type}` endpoint.
#
# Run the following command listing the location(s) of the `ManagerNetworkProtocol` schema used by your BMC and study its output.
#
# The **`PublicationUri`** URI requires an Internet connection to reach `http://redfish.dmtf.org`.
#
# However, the `URI` pointer does not require any Internet access to view its content as it is embedded in the BMC at `/redfish/v1/JsonSchemas/ManagerNetworkProtocol/ManagerNetworkProtocol.json`
# +
echo "Manager Network Protocol schema locations:"
curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET \
${BmcURI}/redfish/v1/JsonSchemas/ManagerNetworkProtocol | jq
# -
# Using the embedded `URI` You can retrieve the definition of the `NTPServers` object and verify you will be able to modify it.
#
# Run the following `curl` command which extracts the `NTPServers` schema definition and note the **`readonly = false`** property.
# +
echo "NTPServers schema definition:"
curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET \
${BmcURI}/redfish/v1/JsonSchemas/ManagerNetworkProtocol/ManagerNetworkProtocol.json | \
jq '.definitions | .NTPProtocol | .properties | .NTPServers'
# -
# You are now sure that it is possible to alter/populate the list of `NTPServers` in your BMC.
#
# The following commands performs a `PATCH` of the `NetworkProtocol` endpoint with a single NTP server IP address.
#
# This `PATCH` request does not return any response data. Other Redfish implementation (i.e. HPE iLO) are more verbose. However, by checking the response header file `$ResponseHeaders`, you should see an `HTTP/1.1: 204` return code stating that the request was successful.
echo "Patching NTP Servers"
curl --dump-header $ResponseHeaders \
--insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request PATCH --data '{ "NTP": {"NTPServers": ["192.168.0.99", ""]} }' \
${BmcURI}/redfish/v1/Managers/bmc/NetworkProtocol | jq
# Verify with the following command that the `NTPServers` list contains the IP address you supplied.
echo "NTP Server list:"
curl --insecure --silent --noproxy "localhost, 127.0.0.1" \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET ${BmcURI}/redfish/v1/Managers/${bmc}/NetworkProtocol | jq '.NTP'
# ## Perform an action: Reset OpenBMC
#
# In the previous section you modified a resource that is not requiring a reset of the BMC to be taken into account. However other parameters may require a restart when changed.
#
# In this paragraph you will perform the `GracefulRestart` action seen previously in your OpenBMC using a `POST` request toward the corresponding target.
#
# After you run this reset command, run the next `bash` cell in order to wait until the BMC is back online.
# +
echo "Starting a reset of the BMC at" ; date
echo
curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request POST --data '{ "ResetType": "GracefulRestart"}' \
${BmcURI}/redfish/v1/Managers/bmc/Actions/Manager.Reset | jq
# -
# ## Wait until OpenBMC is back online
#
# The following cell loops until the BMC returns a valid output to a `GET` request. Run it and wait until the BMC is back on line. This should take about two minutes.
# +
ret=""
while [ "X${ret}" != "X0" ] ; do
timeout 3 curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" \
--header "X-Auth-Token: $BmcToken" \
--request GET ${BmcURI}/redfish/v1/Managers/$bmc > /dev/null
ret=$?
done
echo "BMC is back online at " ; date
echo
# -
# ## Delete sessions
#
# It is extremely important to delete Redfish sessions to avoid reaching the maximum number of opened sessions in a BMC, preventing any access to it. Read this [article](https://developer.hpe.com/blog/managing-ilo-sessions-with-redfish) for more detail.
# +
echo "Body response of a session deletion:"
curl --insecure --noproxy "localhost, 127.0.0.1" --silent \
--header "$HeaderContentType" --header "$HeaderODataVersion" --header "X-Auth-Token: $BmcToken" \
--request DELETE $BmcSessionLocation | jq
# -
# ### What next ?
#
# If you want to re-run this notebook against an **HPE iLO 5**, from your Jupyter Home page, you can duplicate it and then modify the **`BmcIP`** variable with **`16.31.87.207`**.
#
# Then, you will be able to compare the output of OpenBMC and HPE iLO 5 Redfish implementations.
# It is time now to go through the **[Lab 3 notebook](3-Aspire-RedfishPython.ipynb)** to study a Python code suitable for several Redfish implementation
# <img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/><img src="https://redfish.dmtf.org/sites/default/files/DMTF_Redfish_logo_R.jpg" alt="Redfish Logo" style="width: 50px;"/>
|
Redfish/Current/RedfishSchemas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Alih87/Cats-and-Dogs-Classification/blob/main/YOLO_Detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="HHTPSmkYQXYR"
import tensorflow as tf
from keras import backend as K
import numpy as np
import pandas as pd
import PIL
from keras.layers import Input, Lambda, Conv2D
from keras.models import load_model, Model
import matplotlib.pyplot as plt
import os
from scipy import io
from scipy import misc
import argparse
# + id="cIZiL-a0dVgz"
def yolo_filter_boxes(box_confidence, boxes, box_class_probs, threshold = 0.6):
box_scores = box_confidence*box_class_probs
box_classes = K.argmax(box_scores, axis = -1)
box_class_scores = K.max(box_scores, axis = -1)
filtering_mask = box_class_scores >= threshold
scores = tf.boolean_mask(box_class_scores, filtering_mask)
boxes = tf.boolean_mask(boxes, fitering_mask)
classes= tf.boolean_mask(box_classes, filtering_mask)
return scores, boxes, classes
# + id="BvQm0ODXFuXN"
def iou(box1, box2):
xi1 = np.max(box1[0], box2[0])
xi2 = np.max(box1[2], box2[2])
yi1 = np.max(box1[1], box2[1])
yi2 = np.max(box1[3], box2[3])
inter_area = max((yi2-yi1), 0)*max((xi2-xi1), 0)
box1_area = (box1[3] - box1[0]) * (box1[2] - box1[0])
box2_area = (box2[3] - box2[1]) * (box2[2] - box2[0])
union_area = box1_area + box2_area - inter_area
iou = inter_area/union_area
return iou
# + id="ZkkgjtjuTg4N"
def yolo_non_max_suppression(scores, boxes, classes, max_boxes = 10, iou_threshold = 0.5):
max_boxes_tensor = K.variable(max_boxes, dtype='int32')
K.get_session().run(tf.variables_initializeer([max_boxes_tensor]))
|
YOLO_Detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decision Tree Classification
# It is out-of-date model, so just have a look what it's like.
# ## Decision Tree Classifier
import pandas as pd
df = pd.read_excel('./files/wholesale.xls', index_col=0)
df.head()
df.info()
# Set x, y
x = df.iloc[:, 0:8]
y = df['label']
# Import module
from sklearn.tree import DecisionTreeClassifier
# Create and fit model
tr = DecisionTreeClassifier()
tr.fit(x, y)
# Get features & class labels - already known
tr.max_features_, tr.classes_
# Predict data
tr.predict(x)[10:21]
# Compare with actual data
y[10:21].to_numpy()
# Get score - Mean accuracy between self.predict(x) & y
tr.score(x, y)
# Predict unseen data
tr.predict([[0, 4, 13265, 1208, 3821, 6400, 468, 3200]])
# ## Plot the Tree
# * Root node
# * Split node = Internal node = Decision node
# * Leaf node = External node = Terminal node = Class(Label)
#
# Tree node number is counted by BFS method. **If data returns TRUE at a split node, it takes on the left branch** and FALSE vice versa. **'Threshold'** is the value used in the condition of features in split node. **'Value'** in the node represents the sample count in each label.
# Import module
from sklearn.tree import plot_tree
import matplotlib.pyplot as plt
# Turn fitted model into tree visual
# Feature and threshold only apply to split nodes
plt.figure(figsize=(50, 60))
plot_tree(tr, feature_names=['Channel', 'Region', 'Fresh', 'Milk', 'Grocery', 'Frozen','Detergents_Paper', 'Delicassen'])
plt.show()
|
Day32_02_decision_tree_final.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p><font size="6"><b> 02 - Pandas: Basic operations on Series and DataFrames</b></font></p>
#
#
# > *DS Data manipulation, analysis and visualisation in Python*
# > *December, 2017*
#
# > *© 2016, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# + run_control={"frozen": false, "read_only": false}
# %matplotlib inline
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# As you play around with DataFrames, you'll notice that many operations which work on NumPy arrays will also work on dataframes.
#
# + run_control={"frozen": false, "read_only": false}
# redefining the example objects
population = pd.Series({'Germany': 81.3, 'Belgium': 11.3, 'France': 64.3,
'United Kingdom': 64.9, 'Netherlands': 16.9})
countries = pd.DataFrame({'country': ['Belgium', 'France', 'Germany', 'Netherlands', 'United Kingdom'],
'population': [11.3, 64.3, 81.3, 16.9, 64.9],
'area': [30510, 671308, 357050, 41526, 244820],
'capital': ['Brussels', 'Paris', 'Berlin', 'Amsterdam', 'London']})
# + run_control={"frozen": false, "read_only": false}
countries.head()
# -
# # The 'new' concepts
# ## Elementwise-operations
# Just like with numpy arrays, many operations are element-wise:
# + run_control={"frozen": false, "read_only": false}
population / 100
# + run_control={"frozen": false, "read_only": false}
countries['population'] / countries['area']
# + run_control={"frozen": false, "read_only": false}
np.log(countries['population'])
# -
# which can be added as a new column, as follows:
# + run_control={"frozen": false, "read_only": false}
countries["log_population"] = np.log(countries['population'])
# + run_control={"frozen": false, "read_only": false}
countries.columns
# + run_control={"frozen": false, "read_only": false}
countries['population'] > 40
# -
# <div class="alert alert-info">
#
# <b>REMEMBER</b>:
#
# <ul>
# <li>When you have an operation which does NOT work element-wise or you have no idea how to do it directly in Pandas, use the **apply()** function</li>
# <li>A typical use case is with a custom written or a **lambda** function</li>
# </ul>
# </div>
# + run_control={"frozen": false, "read_only": false}
countries["population"].apply(np.log) # but this works as well element-wise...
# + run_control={"frozen": false, "read_only": false}
countries["capital"].apply(lambda x: len(x)) # in case you forgot the functionality: countries["capital"].str.len()
# + run_control={"frozen": false, "read_only": false}
def population_annotater(population):
"""annotate as large or small"""
if population > 50:
return 'large'
else:
return 'small'
# + run_control={"frozen": false, "read_only": false}
countries["population"].apply(population_annotater) # a custom user function
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Calculate the population numbers relative to Belgium</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
population / population['Belgium']
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Calculate the population density for each country and add this as a new column to the dataframe.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
countries['population']*1000000 / countries['area']
# + clear_cell=true run_control={"frozen": false, "read_only": false}
countries['density'] = countries['population']*1000000 / countries['area']
countries
# -
# <div class="alert alert-danger">
#
# **WARNING**: **Alignment!** (unlike numpy)
#
# <ul>
# <li>Pay attention to **alignment**: operations between series will align on the index: </li>
# </ul>
#
# </div>
# + run_control={"frozen": false, "read_only": false}
s1 = population[['Belgium', 'France']]
s2 = population[['France', 'Germany']]
# + run_control={"frozen": false, "read_only": false}
s1
# + run_control={"frozen": false, "read_only": false}
s2
# + run_control={"frozen": false, "read_only": false}
s1 + s2
# -
# ## Aggregations (reductions)
# Pandas provides a large set of **summary** functions that operate on different kinds of pandas objects (DataFrames, Series, Index) and produce single value. When applied to a DataFrame, the result is returned as a pandas Series (one value for each column).
# The average population number:
# + run_control={"frozen": false, "read_only": false}
population.mean()
# -
# The minimum area:
# + run_control={"frozen": false, "read_only": false}
countries['area'].min()
# -
# For dataframes, often only the numeric columns are included in the result:
# + run_control={"frozen": false, "read_only": false}
countries.median()
# -
# # Application on a real dataset
# Reading in the titanic data set...
# + run_control={"frozen": false, "read_only": false}
df = pd.read_csv("../data/titanic.csv")
# -
# Quick exploration first...
# + run_control={"frozen": false, "read_only": false}
df.head()
# + run_control={"frozen": false, "read_only": false}
len(df)
# -
# The available metadata of the titanic data set provides the following information:
#
# VARIABLE | DESCRIPTION
# ------ | --------
# survival | Survival (0 = No; 1 = Yes)
# pclass | Passenger Class (1 = 1st; 2 = 2nd; 3 = 3rd)
# name | Name
# sex | Sex
# age | Age
# sibsp | Number of Siblings/Spouses Aboard
# parch | Number of Parents/Children Aboard
# ticket | Ticket Number
# fare | Passenger Fare
# cabin | Cabin
# embarked | Port of Embarkation (C = Cherbourg; Q = Queenstown; S = Southampton)
#
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>What is the average age of the passengers?</li>
# </ul>
#
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Age'].mean()
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>Plot the age distribution of the titanic passengers</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Age'].hist() #bins=30, log=True
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>What is the survival rate? (the relative number of people that survived)</li>
# </ul>
#
# Note: the 'Survived' column indicates whether someone survived (1) or not (0).
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Survived'].sum() / len(df['Survived'])
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Survived'].mean()
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>What is the maximum Fare? And the median?</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Fare'].max()
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Fare'].median()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Calculate the 75th percentile (`quantile`) of the Fare price (Tip: look in the docstring how to specify the percentile)</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Fare'].quantile(0.75)
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>Calculate the normalized Fares (relative to its mean)</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
df['Fare'] / df['Fare'].mean()
# -
# # Acknowledgement
#
#
# > This notebook is partly based on material of <NAME> (https://github.com/jakevdp/OsloWorkshop2014).
#
# ---
|
_solved/pandas_02_basic_operations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
from sklearn import preprocessing, cross_validation, neighbors, naive_bayes, neural_network, svm, tree
from sklearn.metrics import confusion_matrix, roc_curve, auc
from sklearn.model_selection import KFold, StratifiedKFold
from sklearn.ensemble import AdaBoostClassifier, BaggingClassifier, VotingClassifier
from sklearn.decomposition import PCA
from scipy import interp
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import HTML, display
import tabulate
# # Reading Data
df = pd.read_csv('data/pd_speech_features.txt')
df.drop(['id'], 1, inplace=True)
X = np.array(df.drop(['class'], 1))
y = np.array(df['class'])
# # Normalizing Data
Z = np.divide((X - X.mean(0)), X.std(0))
# # PCA
pca = PCA(n_components = 168)
Z_PCA = pca.fit_transform(Z)
print(Z_PCA.shape)
# # Building the models
# +
# KNN
# knn1 = neighbors.KNeighborsClassifier(n_neighbors=7)
knn = neighbors.KNeighborsClassifier(n_neighbors=5)
# Naive Bayes
gnb = naive_bayes.GaussianNB()
gnb_bagging = BaggingClassifier(naive_bayes.GaussianNB(), max_samples = 0.5, max_features = 1.0, n_estimators = 20)
# Support Vector Machine
svmc = svm.SVC(kernel='linear', probability=True,
random_state=1)
# Neural Network
nn = neural_network.MLPClassifier(
# activation = 'logistic',
# solver = 'sgd',
# max_iter=3000,
# learning_rate_init = 0.001,
# momentum = 0.9,
# epsilon = 1e-04,
hidden_layer_sizes = (200),
random_state = 42)
# Decision Tree
dt = tree.DecisionTreeClassifier(random_state = 42)
dt_boost = AdaBoostClassifier(random_state = 42, base_estimator=tree.DecisionTreeClassifier(random_state = 42, max_depth=1), n_estimators= 100, learning_rate = 1)
boost = AdaBoostClassifier(n_estimators= 100)
vote = VotingClassifier(estimators=[('dt', dt_boost), ('knn', knn), ('nn', nn), ('gnb', gnb)],
voting='soft', weights=[1, 1, 1, 1])
# -
# # Testing using cross validation
# +
# model = knn
# scores = cross_validation.cross_val_score(model, Z, y, cv=5, scoring='accuracy')
# y_pred = cross_validation.cross_val_predict(model, Z, y, cv=5)
# acc = scores.mean()
# conf_mat = confusion_matrix(y, y_pred)
# +
# print(acc)
# print(conf_mat)
# -
# ## Accuracy, ROC/AUC, and confuson matrix
# +
folds = 5
cv = StratifiedKFold(n_splits=folds)
# Classifiers:
# Decision Tree: dt
# Decision Tree (with boosting): dt_boost
# MLP: nn
# Gaussian Naive Bayes: gnb
# KNN: knn
classifier = nn
inp = Z
acc = np.zeros(folds)
confm = np.zeros((2, 2))
tprs = []
aucs = []
mean_fpr = np.linspace(0, 1, 100)
i = 0
for train, test in cv.split(inp, y):
probas_ = classifier.fit(inp[train], y[train]).predict_proba(inp[test])
# Compute accuracy
y_pred = classifier.predict(inp[test])
acc[i] = (y_pred == y[test]).mean()
# Confusion matrix
confm = confm + confusion_matrix(y[test], y_pred)
# Compute ROC curve and area the curve
fpr, tpr, thresholds = roc_curve(y[test], probas_[:, 1])
tprs.append(interp(mean_fpr, fpr, tpr))
tprs[-1][0] = 0.0
roc_auc = auc(fpr, tpr)
aucs.append(roc_auc)
plt.plot(fpr, tpr, lw=1, alpha=0.3,
label='ROC fold %d (AUC = %0.2f)' % (i, roc_auc))
i += 1
plt.plot([0, 1], [0, 1], linestyle='--', lw=2, color='r',
label='Chance', alpha=.8)
mean_tpr = np.mean(tprs, axis=0)
mean_tpr[-1] = 1.0
mean_auc = auc(mean_fpr, mean_tpr)
std_auc = np.std(aucs)
plt.plot(mean_fpr, mean_tpr, color='b',
label=r'Mean ROC (AUC = %0.2f $\pm$ %0.2f)' % (mean_auc, std_auc),
lw=2, alpha=.8)
std_tpr = np.std(tprs, axis=0)
tprs_upper = np.minimum(mean_tpr + std_tpr, 1)
tprs_lower = np.maximum(mean_tpr - std_tpr, 0)
plt.fill_between(mean_fpr, tprs_lower, tprs_upper, color='grey', alpha=.2,
label=r'$\pm$ 1 std. dev.')
plt.xlim([-0.05, 1.05])
plt.ylim([-0.05, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic')
plt.legend(loc="lower right")
plt.show()
# -
'{:.2f}% +- {:.2f}%'.format(acc.mean() * 100, acc.std() * 100)
cm = np.zeros((3,3))
cm[0:2, 0:2] = confm
cm[0,2] = (cm[0,0] / cm[0,0:2].sum())* 100
cm[1,2] = (cm[1,1] / cm[1,0:2].sum())* 100
cm[2,0] = (cm[0,0] / cm[0:2,0].sum())* 100
cm[2,1] = (cm[1,1] / cm[0:2,1].sum())* 100
display(HTML(tabulate.tabulate(cm, tablefmt='html')))
|
MLP 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Testing PASTIS in imaging mode </h1>
#
# ## -- JWST aperture --
# Here we're testing the module "image_pastis.py" which is the version of PASTIS that still generates images. Since this is a module, it's a tiny bit harder to test, so I'm basically just going through the code step by step.
#
# Since this is just copy-pasted code from that module, this will probably be out of date. This is an older version for the JWST. Maybe I can update this at some point - I will when I come back to working on the analytical model.
# +
import os
import numpy as np
from astropy.io import fits
import astropy.units as u
import poppy.zernike as zern
import poppy.matrixDFT as mft
import poppy
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
os.chdir('../../pastis/')
from config import CONFIG_PASTIS
import util as util
# Change to output folder for this script
#dir = '/Users/ilaginja/Documents/data_from_repos/pastis_data/active/calibration'
#os.chdir(dir)
# -
# ## Setup and single segment aberration
#
# Since this is a module, its function will be called with input parameters, which I will define separetaly here to be able to use them.
#
# Taken from `calibration.py`.
# +
# Define the aberration coeffitients "coef"
which_tel = CONFIG_PASTIS.get('telescope', 'name')
nb_seg = CONFIG_PASTIS.getint(which_tel, 'nb_subapertures')
zern_max = CONFIG_PASTIS.getint('zernikes', 'max_zern')
nm_aber = CONFIG_PASTIS.getfloat('calibration', 'single_aberration') * u.nm # [nm] amplitude of aberration
zern_number = CONFIG_PASTIS.getint('calibration', 'zernike') # Which (Noll) Zernike we are calibrating for
wss_zern_nb = util.noll_to_wss(zern_number) # Convert from Noll to WSS framework
### What segmend are we aberrating? ###
i = 0 # segment 1 --> i=0, seg 2 --> i=1, etc.
### ------------------------------- ###
# Create arrays to hold Zernike aberration coefficients
Aber_WSS = np.zeros([nb_seg, zern_max]) # The Zernikes here will be filled in the WSS order!!!
# Because it goes into _apply_hexikes_to_seg().
Aber_Noll = np.copy(Aber_WSS) # This is the Noll version for later.
# Feed the aberration nm_aber into the array position
# that corresponds to the correct Zernike, but only on segment i
Aber_WSS[i, wss_zern_nb-1] = nm_aber.to(u.m).value # Aberration on the segment we're currenlty working on;
# convert to meters; -1 on the Zernike because Python starts
# numbering at 0.
Aber_Noll[i, zern_number-1] = nm_aber.value # Noll version - in input units directly!
# Vector of aberration coefficients takes all segments, but only for the Zernike we currently work with
coef = Aber_Noll[:,zern_number-1]
# Make sure the aberration coefficients have correct units
coef *= u.nm
# Define the (Noll) zernike number
zernike_pol = zern_number
# We're not calibrating
cali=False
print('coef: {}'.format(coef))
print('Aberration: {}'.format(nm_aber))
print('Zernike (Noll): {}'.format(util.zernike_name(zern_number, framework='Noll')))
print('Zernike (WSS): {}'.format(util.zernike_name(wss_zern_nb, framework='WSS')))
print('Zernike number (Noll): {}'.format(zernike_pol))
# -
# Now we'll start with the actual code in the module `image_pastis.py`.
# +
#-# Parameters
dataDir = os.path.join(CONFIG_PASTIS.get('local', 'local_data_path'), 'active')
nb_seg = CONFIG_PASTIS.getint(which_tel, 'nb_subapertures')
tel_size_m = CONFIG_PASTIS.getfloat(which_tel, 'diameter') * u.m
real_size_seg = CONFIG_PASTIS.getfloat(which_tel, 'flat_to_flat') # size in meters of an individual segment flatl to flat
size_seg = CONFIG_PASTIS.getint('numerical', 'size_seg') # pixel size of an individual segment tip to tip
wvln = CONFIG_PASTIS.getint(which_tel, 'lambda') * u.nm
inner_wa = CONFIG_PASTIS.getint(which_tel, 'IWA')
outer_wa = CONFIG_PASTIS.getint(which_tel, 'OWA')
tel_size_px = CONFIG_PASTIS.getint('numerical', 'tel_size_px') # pupil diameter of telescope in pixels
im_size_pastis = CONFIG_PASTIS.getint('numerical', 'im_size_px_pastis') # image array size in px
im_size_e2e = CONFIG_PASTIS.getint('numerical', 'im_size_px_webbpsf')
sampling = CONFIG_PASTIS.getfloat(which_tel, 'sampling') # sampling
largeur = tel_size_px * sampling # size of pupil (?) with taking the sampling into account
size_px_tel = tel_size_m / tel_size_px # size of one pixel in pupil plane in m
px_sq_to_rad = (size_px_tel * np.pi / tel_size_m) * u.rad
zern_max = CONFIG_PASTIS.getint('zernikes', 'max_zern')
# Create Zernike mode object for easier handling
zern_mode = util.ZernikeMode(zernike_pol)
#-# Mean subtraction for piston
if zernike_pol == 1:
coef -= np.mean(coef)
#-# Generic segment shapes
# Load pupil from file
pupil = fits.getdata(os.path.join(dataDir, 'segmentation', 'pupil.fits'))
pup_im = np.copy(pupil)
print('Pupil shape:', pupil.shape)
plt.imshow(pupil, origin='lower')
plt.show()
# -
# ### Creating a mini segment
#
# At this point, you have to make sure the pixel size **size_seg** of your individual segment is correct, and this will be different depending on the pixel size of your total pupil.
#
# With that, we create a mini segment.
# +
# Creat a mini-segment (one individual segment from the segmented aperture)
mini_seg_real = poppy.NgonAperture(name='mini', radius=real_size_seg) # creating real mini segment shape with poppy
#test = mini_seg_real.sample(wavelength=wvln, grid_size=flat_diam, return_scale=True) # fix its sampling with wavelength
mini_hdu = mini_seg_real.to_fits(wavelength=wvln, npix=size_seg) # make it a fits file
mini_seg = mini_hdu[0].data
plt.imshow(mini_seg, origin='lower')
plt.title('One mini segment')
plt.show()
print('Mini-segment array size:', mini_seg.shape)
# -
# We managed to cut the array to a square where the mini-segment is just about touching the array edges. The size of this array is the size of our mini-segment and that's a number that we have to enter into the configfile. Enter the pixel size of the mini-segment array into the configfile section: **[numerical] --> size_seg**
# ### Generating the dark hole
# +
#-# Generate a dark hole
dh_area = util.create_dark_hole(pup_im, inner_wa, outer_wa, sampling)
print('DH array shape:', dh_area.shape)
boxsize = 50
plt.figure(figsize=(16, 8))
plt.subplot(1, 2, 1)
plt.imshow(dh_area, origin='lower')
plt.title('Dark hole')
plt.subplot(1, 2, 2)
plt.imshow(util.zoom_cen(dh_area, boxsize), origin='lower')
plt.title('Dark hole zoomed in')
plt.show()
# -
# ### Importing the matrix
# +
#-# Import information form aperture generation script
Projection_Matrix = fits.getdata(os.path.join(dataDir, 'segmentation', 'Projection_Matrix.fits'))
vec_list = fits.getdata(os.path.join(dataDir, 'segmentation', 'vec_list.fits'))
NR_pairs_list = fits.getdata(os.path.join(dataDir, 'segmentation', 'NR_pairs_list_int.fits'))
NR_pairs_nb = NR_pairs_list.shape[0]
plt.imshow(Projection_Matrix[:,:,0], origin='lower')
plt.title('Projection matrix displaying NRPs')
plt.colorbar()
plt.show()
print('Non-redundant pairs (' + str(NR_pairs_nb) + '):')
print(NR_pairs_list)
# -
# ### Calculating the (uncalibrated) analytical image
#
# We don't have calibration coefficients yet, so we skip the "if cali:" part.
#
# Move on to the calculation of eq. 13 in the paper Leboulleux et al. 2018 that calculates the image intensity of the analytical images:
#
# $$I(u) = ||\hat{Z_l}(u)||^2 \Bigg[ \sum_{k=1}^{n_{seg}} a^2_{k,l} + 2 \sum_{q=1}^{n_{NRP}} A_q cos(b_q \cdot u) \Bigg] $$
#
# #### Generic coefficients
#
# $A_q$... *generic coefficients*
#
# $$A_q = \sum_{(i,j)} a_{i,l} a_{j,l}$$
# +
#-# Generic coefficients
generic_coef = np.zeros(NR_pairs_nb) * u.nm * u.nm # coefficients in front of the non redundant pairs,
# the A_q in eq. 13 in Leboulleux et al. 2018
for q in range(NR_pairs_nb):
for i in range(nb_seg):
for j in range(i+1, nb_seg):
if Projection_Matrix[i, j, 0] == q+1:
print('q:', q, 'i:', i, 'j:', j)
generic_coef[q] += coef[i] * coef[j]
print('ci:', coef[i], 'cj:', coef[j])
# -
print('Generic coefficients:')
print(generic_coef)
# #### Sum over the aberration coefficients and sum over the cosines
#
# $cos(b_q \cdot u)$... cos_u_mat
#
# Sum over $a^2_{k,l}$... sum1
# +
#-# Constant sum and cosine sum - calculating eq. 13 from Leboulleux et al. 2018
i_line = np.linspace(-im_size_pastis/2., im_size_pastis/2., im_size_pastis)
tab_i, tab_j = np.meshgrid(i_line, i_line)
cos_u_mat = np.zeros((int(im_size_pastis), int(im_size_pastis), NR_pairs_nb))
# The -1 with each NR_pairs_list is because the segment names are saved starting from 1, but Python starts
# its indexing at zero, so we have to make it start at zero here too.
for q in range(NR_pairs_nb):
cos_u_mat[:,:,q] = np.cos(px_sq_to_rad * (vec_list[NR_pairs_list[q,0]-1, NR_pairs_list[q,1]-1, 0] * tab_i) +
px_sq_to_rad * (vec_list[NR_pairs_list[q,0]-1, NR_pairs_list[q,1]-1, 1] * tab_j)) * u.dimensionless_unscaled
sum1 = np.sum(coef**2) # sum of all a_{k,l} in eq. 13 - this works only for single Zernikes (l fixed), because np.sum would sum over l too, which would be wrong.
print('cos:', cos_u_mat)
print('sum1:', sum1)
# -
# $\Bigg[ \sum_{k=1}^{n_{seg}} a^2_{k,l} + 2 \sum_{q=1}^{n_{NRP}} A_q cos(b_q \cdot u) \Bigg]$ = sum2 + generic_coef[q] * cos_u_mat[:,:,q]
# +
sum2 = np.zeros((int(im_size_pastis), int(im_size_pastis))) * u.nm * u.nm
for q in range(NR_pairs_nb):
sum2 = sum2 + generic_coef[q] * cos_u_mat[:,:,q]
print('sum2:', sum2)
# -
# #### The global envelope from the mini segment Zernike
# +
#-# Local Zernike
# Generate a basis of Zernikes with the mini segment being the support
isolated_zerns = zern.hexike_basis(nterms=zern_max, npix=size_seg, rho=None, theta=None, vertical=False, outside=0.0)
# Calculate the Zernike that is currently being used and put it on one single subaperture, the result is Zer
# Apply the currently used Zernike to the mini-segment.
if zernike_pol == 1:
Zer = np.copy(mini_seg)
elif zernike_pol in range(2, zern_max-2):
Zer = np.copy(mini_seg)
Zer = Zer * isolated_zerns[zernike_pol-1]
plt.imshow(Zer, origin='lower')
plt.title('Zernike mode on mini segment')
plt.show()
# -
# Simply to show what it would look like for other Zernikes:
# +
# Calculate them
zern_iter = np.arange(8) + 1 # +1 so that they start at 1
print('Zernike numbers (Noll):', zern_iter)
minizern_stack = []
for i in range(zern_iter.shape[0]):
if i+1 == 1: # for piston
minizern_stack.append(mini_seg)
elif i+1 in range(2, zern_max-2):
minizern_stack.append(mini_seg * isolated_zerns[i])
minizern_stack = np.array(minizern_stack)
# -
# Display them
plt.figure(figsize=(16, 8))
plt.suptitle('Different Zernikes on one mini-segment')
for i in range(minizern_stack.shape[0]):
plt.subplot(2, 4, i+1)
plt.imshow(minizern_stack[i], origin='lower')
plt.title('Noll Zernike: ' + str(i+1))
plt.show()
# +
# Fourier Transform of the Zernike - the global envelope
mf = mft.MatrixFourierTransform()
ft_zern = mf.perform(Zer, im_size_pastis/sampling, im_size_pastis)
box_e2e = int(im_size_e2e/2) # We set the image zoom to the same like WebbPSF
plt.figure(figsize=(15, 15))
plt.subplot(2, 2, 1)
plt.imshow(np.abs(ft_zern), norm=LogNorm(), origin='lower')
plt.title('FT of mini Zernike')
plt.subplot(2, 2, 2)
plt.imshow(util.zoom_cen(np.abs(ft_zern), 50), norm=LogNorm(), origin='lower')
plt.title('FT of mini Zernike - zoom')
plt.subplot(2, 2, 3)
plt.imshow(util.zoom_cen(np.abs(ft_zern), box_e2e), norm=LogNorm(), origin='lower')
plt.title('FT of mini Zernike - zoom like WebbPSF')
plt.show()
# -
# #### What do those envelopes look like for other Zernikes?
#
# Also check the global envelopes from the other Zernikes on the mini-segment.
# +
mini_ft = []
for i in range(minizern_stack.shape[0]):
ft_klein = mf.perform(minizern_stack[i], im_size_pastis/sampling, im_size_pastis)
mini_ft.append(ft_klein)
mini_ft = np.array(mini_ft)
# +
zoomin = 50
plt.figure(figsize=(18, 18))
plt.suptitle('Different Zernikes envelopes')
for i in range(mini_ft.shape[0]):
plt.subplot(3, 3, i+1)
plt.imshow(util.zoom_cen(np.abs(mini_ft[i]), zoomin), norm=LogNorm(), origin='lower')
plt.title('Noll Zernike: ' + str(i+1))
plt.show()
# -
# Now in the image size of the WebbPSF images from 3. notebook
plt.figure(figsize=(18, 18))
plt.suptitle('Different Zernikes envelopes')
for i in range(mini_ft.shape[0]):
plt.subplot(3, 3, i+1)
plt.imshow(util.zoom_cen(np.abs(mini_ft[i]), box_e2e), norm=LogNorm(), origin='lower')
plt.title('<NAME>: ' + str(i+1))
plt.show()
# They correspond to what Lucie has in her paper, which is pretty cool. The sampling is very different though, as I tried to match the sampling to the WebbPSF default.
#
# #### Putting things together and calculating the full image
#
# Moving on. Calculating the full $I(u)$ now.
#
# $$I(u) = ||\hat{Z_l}(u)||^2 \Bigg[ \sum_{k=1}^{n_{seg}} a^2_{k,l} + 2 \sum_{q=1}^{n_{NRP}} A_q cos(b_q \cdot u) \Bigg] $$
# +
#-# Final image - old and WRONG version!!
# Generating the final image that will get passed on to the outer scope, I(u) in eq. 13
intensity_wrong = np.abs(ft_zern**2 * (sum1.value + 2. * sum2.value)) # old and WRONG!!!
#intensity = np.abs(ft_zern)**2 * (sum1.value + 2. * sum2.value)
plt.figure(figsize=(18,18))
plt.subplot(2, 2, 1)
plt.imshow(intensity_wrong, norm=LogNorm(), origin='lower')
plt.title('Final image')
plt.subplot(2, 2, 2)
plt.imshow(util.zoom_cen(intensity_wrong, 50), norm=LogNorm(), origin='lower') #[450:575, 450:575]
plt.title('Zoomed final image')
plt.subplot(2, 2, 3)
plt.imshow(util.zoom_cen(intensity_wrong, box_e2e), norm=LogNorm(), origin='lower')
plt.title('Final image - zoom like WebbPSF')
plt.show()
# +
#-# Final image
# Generating the final image that will get passed on to the outer scope, I(u) in eq. 13
#intensity = np.abs(ft_zern**2 * (sum1.value + 2. * sum2.value)) # old and WRONG!!!
intensity = np.abs(ft_zern)**2 * (sum1.value + 2. * sum2.value)
plt.figure(figsize=(18,18))
plt.subplot(2, 2, 1)
plt.imshow(intensity, norm=LogNorm(), origin='lower')
plt.title('Final image')
plt.subplot(2, 2, 2)
plt.imshow(util.zoom_cen(intensity, 50), norm=LogNorm(), origin='lower') #[450:575, 450:575]
plt.title('Zoomed final image')
plt.subplot(2, 2, 3)
plt.imshow(util.zoom_cen(np.abs(ft_zern), box_e2e), norm=LogNorm(), origin='lower')
plt.title('Final image - zoom like WebbPSF')
plt.show()
# -
#
#
# #### Extracting the dark hole
# +
# PASTIS is only valid inside the dark hole.
tot_dh_im_size = sampling*(outer_wa+3) # zoom box must be big enough to capture entire DH
intensity_zoom = util.zoom_cen(intensity, tot_dh_im_size)
dh_area_zoom = util.zoom_cen(dh_area, tot_dh_im_size)
dh_psf = dh_area_zoom * intensity_zoom
# Display dark hole and inner part of image next ot each other, on the same scale
plt.figure(figsize=(19,10))
plt.subplot(1, 2, 1)
plt.imshow(dh_psf, norm=LogNorm(), origin='lower')
plt.title('Final image - dark hole only')
plt.subplot(1, 2, 2)
plt.imshow(util.zoom_cen(intensity, tot_dh_im_size), norm=LogNorm(), origin='lower')
plt.title('Without mask for comparison, same image size')
plt.show()
# -
# Crop out the DH for the Zernike envelopes.
# +
mini_im = np.abs(mini_ft) # don't forget that mini_ft is the E-field
mini_dh_stack = []
plt.figure(figsize=(18, 60))
plt.suptitle('DH area of individual Zernike envelopes')
for i in range(mini_ft.shape[0]):
mini_zoom = util.zoom_cen(mini_im[i], tot_dh_im_size)
mini_dh = dh_area_zoom * mini_zoom
mini_dh_stack.append(mini_dh)
plt.subplot(8, 2, i*2+1)
plt.imshow(np.abs(mini_dh), norm=LogNorm(), origin='lower')
plt.title('Noll Zernike: ' + str(i+1))
plt.subplot(8, 2, i*2+2)
plt.imshow(np.abs(mini_zoom), norm=LogNorm(), origin='lower')
plt.title('Noll Zernike: ' + str(i+1))
plt.show()
# -
# ### Aberrating pairs of segments
#
# We now want to explore what the final analytical image looks like when we aberrate two segments at a time, with the same aberration.
# +
# Decide which two segments you want to aberrate
segnum1 = 9 # Which segments are we aberrating - I number them starting with 1
segnum2 = 15
segnum_array = np.array([segnum1, segnum2])
zern_pair = 1 # Which Noll Zernike are we putting on the segments.
print('Aberrated segments:', segnum_array)
print('Noll Zernike used:', zern_pair)
# Create aberration vector
Aber_Noll = np.zeros([nb_seg, zern_max])
print('nm_aber: {}'.format(nm_aber))
# Fill aberration array
for i, nseg in enumerate(segnum_array):
Aber_Noll[nseg-1, zern_pair-1] = nm_aber.value # fill only the index for current Zernike, on segment i - in input units
# Define the aberration coefficient vector
coef = Aber_Noll[:, zern_pair-1]
coef *= u.nm
print('coef:', coef)
# +
#-# Generic coefficients
generic_coef = np.zeros(NR_pairs_nb) * u.nm * u.nm # coefficients in front of the non redundant pairs,
# the A_q in eq. 13 in Leboulleux et al. 2018
for q in range(NR_pairs_nb):
for i in range(nb_seg):
for j in range(i+1, nb_seg):
if Projection_Matrix[i, j, 0] == q+1:
generic_coef[q] += coef[i] * coef[j]
print('Generic coefficients:')
print(generic_coef)
#-# Constant sum and cosine sum - calculating eq. 13 from Leboulleux et al. 2018
i_line = np.linspace(-im_size_pastis/2., im_size_pastis/2., im_size_pastis)
tab_i, tab_j = np.meshgrid(i_line, i_line) # these are arrys for the image plane coordinate u
cos_u_mat = np.zeros((int(im_size_pastis), int(im_size_pastis), NR_pairs_nb))
# The -1 with each NR_pairs_list is because the segment names are saved starting from 1, but Python starts
# its indexing at zero, so we have to make it start at zero here too.
for q in range(NR_pairs_nb):
cos_u_mat[:,:,q] = np.cos(px_sq_to_rad * (vec_list[NR_pairs_list[q,0]-1, NR_pairs_list[q,1]-1, 0] * tab_i) +
px_sq_to_rad * (vec_list[NR_pairs_list[q,0]-1, NR_pairs_list[q,1]-1, 1] * tab_j)) * u.dimensionless_unscaled
sum1 = np.sum(coef**2) # sum of all a_{k,l} in eq. 13 - this works only for single Zernikes (l fixed), because np.sum would sum over l too, which would be wrong.
#print('cos:', cos_u_mat)
#print('sum1:', sum1)
sum2 = np.zeros((int(im_size_pastis), int(im_size_pastis))) * u.nm * u.nm
for q in range(NR_pairs_nb):
sum2 = sum2 + generic_coef[q] * cos_u_mat[:,:,q]
#print('sum2:', sum2)
# +
# Calculate the Zernike that is currently being used and put it on one single subaperture, the result is Zer
# Apply the currently used Zernike to the mini-segment.
if zern_pair == 1:
Zer = np.copy(mini_seg)
elif zern_pair in range(2, zern_max-2):
Zer = np.copy(mini_seg)
Zer = Zer * isolated_zerns[zern_pair-1]
plt.imshow(Zer, origin='lower')
plt.title('Zernike mode on mini segment')
plt.show()
# +
# Fourier Transform of the Zernike - the global envelope
mf = mft.MatrixFourierTransform()
ft_zern = mf.perform(Zer, im_size_pastis/sampling, im_size_pastis)
#xcen = int(ft_zern.shape[1]/2.)
#ycen = int(ft_zern.shape[0]/2.)
#boxw = int(161/2) # We can see in the 3. notebook that WebbPSF produces 161 x 161 px images by default.
#
#plt.figure(figsize=(15, 15))
#plt.subplot(2, 2, 1)
#plt.imshow(np.abs(ft_zern), norm=LogNorm())
#plt.title('FT of mini Zernike')
#plt.subplot(2, 2, 2)
#plt.imshow(np.abs(ft_zern)[485:540, 485:540], norm=LogNorm())
#plt.title('FT of mini Zernike - zoom')
#plt.subplot(2, 2, 3)
#plt.imshow(np.abs(ft_zern)[ycen-boxw:ycen+boxw, xcen-boxw:xcen+boxw], norm=LogNorm())
#plt.title('FT of mini Zernike - zoom like WebbPSF')
#plt.show()
# +
#-# Final image
# Generating the final image that will get passed on to the outer scope, I(u) in eq. 13
intensity = np.abs(ft_zern)**2 * (sum1.value + 2. * sum2.value)
boxw2 = box_e2e/2
plt.figure(figsize=(10,10))
plt.imshow(util.zoom_cen(intensity, boxw2), norm=LogNorm(), origin='lower')
plt.title('Zoomed final image')
plt.show()
# -
# #### Saving some of the images
#
# I will save a couple of images down here to be able to display them next to each other:
# +
#segs_3_11_noll_1 = np.copy(intensity)
#segs_11_17_noll_1 = np.copy(intensity)
#segs_6_11_noll_1 = np.copy(intensity)
#segs_9_2_noll_1 = np.copy(intensity)
#segs_9_5_noll_1 = np.copy(intensity)
#segs_9_15_noll_1 = np.copy(intensity)
#segs_8_2_noll_1 = np.copy(intensity)
#segs_8_3_noll_1 = np.copy(intensity)
#segs_8_12_noll_1 = np.copy(intensity)
#segs_8_18_noll_1 = np.copy(intensity)
#segs_2_6_noll_1 = np.copy(intensity)
#segs_10_16_noll_1 = np.copy(intensity)
#segs_8_1_noll_1 = np.copy(intensity)
#segs_8_6_noll_1 = np.copy(intensity)
#segs_8_16_noll_1 = np.copy(intensity)
#segs_10_16_noll_2 = np.copy(intensity)
#segs_2_6_noll_2 = np.copy(intensity)
#segs_8_18_noll_2 = np.copy(intensity)
#segs_8_12_noll_2 = np.copy(intensity)
#segs_8_3_noll_2 = np.copy(intensity)
#segs_8_2_noll_2 = np.copy(intensity)
#segs_9_15_noll_2 = np.copy(intensity)
#segs_9_2_noll_2 = np.copy(intensity)
# +
# Save them all to fits files
save_dir = '/astro/opticslab1/PASTIS/jwst_data/uncalibrated_analytical_images/2019-01-25-16h-00min'
#util.write_fits(segs_3_11_noll_1, os.path.join(save_dir, 'segs_3_11_noll_1.fits'))
#util.write_fits(segs_11_17_noll_1, os.path.join(save_dir, 'segs_11_17_noll_1.fits'))
#util.write_fits(segs_6_11_noll_1, os.path.join(save_dir, 'segs_6_11_noll_1.fits'))
#util.write_fits(segs_9_2_noll_1, os.path.join(save_dir, 'segs_9_2_noll_1.fits'))
#util.write_fits(segs_9_5_noll_1, os.path.join(save_dir, 'segs_9_5_noll_1.fits'))
#util.write_fits(segs_9_15_noll_1, os.path.join(save_dir, 'segs_9_15_noll_1.fits'))
#util.write_fits(segs_8_2_noll_1, os.path.join(save_dir, 'segs_8_2_noll_1.fits'))
#util.write_fits(segs_8_3_noll_1, os.path.join(save_dir, 'segs_8_3_noll_1.fits'))
#util.write_fits(segs_8_12_noll_1, os.path.join(save_dir, 'segs_8_12_noll_1.fits'))
#util.write_fits(segs_8_18_noll_1, os.path.join(save_dir, 'segs_8_18_noll_1.fits'))
#util.write_fits(segs_2_6_noll_1, os.path.join(save_dir, 'segs_2_6_noll_1.fits'))
#util.write_fits(segs_10_16_noll_1, os.path.join(save_dir, 'segs_10_16_noll_1.fits'))
#util.write_fits(segs_8_1_noll_1, os.path.join(save_dir, 'segs_8_1_noll_1.fits'))
#util.write_fits(segs_8_6_noll_1, os.path.join(save_dir, 'segs_8_6_noll_1.fits'))
#util.write_fits(segs_8_16_noll_1, os.path.join(save_dir, 'segs_8_16_noll_1.fits'))
#util.write_fits(segs_10_16_noll_2, os.path.join(save_dir, 'segs_10_16_noll_2.fits'))
#util.write_fits(segs_2_6_noll_2, os.path.join(save_dir, 'segs_2_6_noll_2.fits'))
#util.write_fits(segs_8_18_noll_2, os.path.join(save_dir, 'segs_8_18_noll_2.fits'))
#util.write_fits(segs_8_12_noll_2, os.path.join(save_dir, 'segs_8_12_noll_2.fits'))
#util.write_fits(segs_8_3_noll_2, os.path.join(save_dir, 'segs_8_3_noll_2.fits'))
#util.write_fits(segs_8_2_noll_2, os.path.join(save_dir, 'segs_8_2_noll_2.fits'))
#util.write_fits(segs_9_15_noll_2, os.path.join(save_dir, 'segs_9_15_noll_2.fits'))
#util.write_fits(segs_9_2_noll_2, os.path.join(save_dir, 'segs_9_2_noll_2.fits'))
# -
# #### Display and compare the images
#
# I started making PASTIS images from the direct entrance pupil of JWST, which I needed to change to be using the exit pupil instead, because I needed to make it the same like in WebbPSF. That's what's happening now in the code, but below here, I am just showing the general properties of PASTIS images. I have made a lot more different data with the entrance pupil, which is why I am using those images for the demo below here. This data is stored in: '/astro/opticslab1/PASTIS/jwst_data/uncalibrated_analytical_images/2018-01-17-17h-35min_piston_pairs'.
#
# #### *Fringe orientation and spacing*
#
# Let's start with the image in which we put **piston** on **segments 3 and 11**. When we check in what relation those two segments lie to each other on the (exit!) pupil (open the pupil image file for help), we can see:
#
# 1. They like on a diagonal offset to each other, about 40 degrees tilted from the horizontal.
# 2. They are very close together, the closest two segments on the JWST can be.
#
# This means for the image:
# 1. The fringes are are tilted by the same amount like the connection vector between the two segments, but flipped by 90 degrees.
# 2. Since the segments are **close together**, the fringes in the Fourier plane, here the final image, will be **wide**.
# +
# I need to read the images in now
read_dir1 = '/astro/opticslab1/PASTIS/jwst_data/uncalibrated_analytical_images/2019-01-17-17h-35min_piston_1000nm_pairs'
segs_3_11_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_3_11_noll_1.fits'))
segs_11_17_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_11_17_noll_1.fits'))
segs_6_11_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_6_11_noll_1.fits'))
segs_9_2_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_9_2_noll_1.fits'))
segs_9_5_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_9_5_noll_1.fits'))
segs_9_15_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_9_15_noll_1.fits'))
segs_8_2_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_8_2_noll_1.fits'))
segs_8_3_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_8_3_noll_1.fits'))
segs_8_12_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_8_12_noll_1.fits'))
segs_8_18_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_8_18_noll_1.fits'))
segs_2_6_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_2_6_noll_1.fits'))
segs_10_16_noll_1 = fits.getdata(os.path.join(read_dir1, 'segs_10_16_noll_1.fits'))
read_dir2 = '/astro/opticslab1/PASTIS/jwst_data/uncalibrated_analytical_images/2019-01-17-17h-45min_tip_1000nm_pairs'
segs_10_16_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_10_16_noll_2.fits'))
segs_2_6_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_2_6_noll_2.fits'))
segs_8_18_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_8_18_noll_2.fits'))
segs_8_12_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_8_12_noll_2.fits'))
segs_8_3_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_8_3_noll_2.fits'))
segs_8_2_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_8_2_noll_2.fits'))
segs_9_15_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_9_15_noll_2.fits'))
segs_9_2_noll_2 = fits.getdata(os.path.join(read_dir2, 'segs_9_2_noll_2.fits'))
boxw2 = box_e2e/2
# -
plt.figure(figsize=(10, 10))
plt.imshow(util.zoom_cen(segs_3_11_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 3 and 11')
plt.show()
# Moving on to the image iwhere we put **piston** on **segments 11 and 17**. As opposed to the previous image of segments 3 and 11 being aberrated:
#
# 1. They are connected by the same diagonal.
# 2. They are very far apart, they have the largest possible segment distance on the JWST.
#
# This means for the image:
# 1. The fringes have the same orientation in the image like the previous one, because the two segment pairs have the same orientation.
# 2. Since the segments are **far apart**, the fringes in the Fourier plane, here the final image, will be **narrow**. In fact the fringes are so narrow, that we cannot clearly see them when the sampling is low (aliasing).
plt.figure(figsize=(10, 10))
plt.imshow(util.zoom_cen(segs_11_17_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 11 and 17')
plt.show()
# For **piston** on **segments 6 and 11**, we have the same fringe orientation again, because the segment pair has the same orientation in the pupil like the two examples before, but because their distance is in between the two previous cases, the fringe spacing will also be somewhere in between.
plt.figure(figsize=(10, 10))
plt.imshow(util.zoom_cen(segs_6_11_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 6 and 11')
plt.show()
# For **piston** on the segment pairs **9-2**, **9-5** and **9-15**, we can see that they have a pair orientation that is rotated by 90 degrees with respect to the cases we have looked at before, and since these three pairs have different segment separations, you can see how their fringe spacing differs.
# +
plt.figure(figsize=(18, 6))
plt.subplot(1, 3, 1)
plt.imshow(util.zoom_cen(segs_9_2_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 9 and 2')
plt.subplot(1, 3, 2)
plt.imshow(util.zoom_cen(segs_9_5_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 9 and 5')
plt.subplot(1, 3, 3)
plt.imshow(util.zoom_cen(segs_9_15_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 9 and 15')
plt.show()
# -
# For **piston** on the segment pairs **8-2**, **8-3** and **8-12**, we can see that they have a pair orientation that is vertical in the pupil plane, so our fringes will be horizontal this time. And since these three pairs have different segment separations, you can see how their fringe spacing differs - like in the previous examples.
# +
plt.figure(figsize=(18, 6))
plt.subplot(1, 3, 1)
plt.imshow(util.zoom_cen(segs_8_2_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 8 and 2')
plt.subplot(1, 3, 2)
plt.imshow(util.zoom_cen(segs_8_3_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 8 and 3')
plt.subplot(1, 3, 3)
plt.imshow(util.zoom_cen(segs_8_12_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 8 and 12')
plt.show()
# -
# #### *Pair redundancy*
#
# Below, I will display the images for **piston** on pair **8-18** and **2-6**.
#
# As you can see - there is absolutely no difference. Except that we know that the two images come from a practically different setup, but effectively, they show the same result.
# This is explained in Fig. 2 and Sec. 2.2.2 of Leboulleux et al. (2018). The only defining parameters for the influence a segment pair has on the image plane are its orientation and separation. Both of these things are exactly the same for the two pairs displayed here, hence the images resulting from these two seemingly different setups are the same.
# +
plt.figure(figsize=(18, 9))
plt.subplot(1, 2, 1)
plt.imshow(util.zoom_cen(segs_8_18_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 8 and 18')
plt.subplot(1, 2, 2)
plt.imshow(util.zoom_cen(segs_2_6_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 2 and 6')
plt.show()
# -
# And as one last example with **piston** only: pair **10-16**. The fringe orientation will be same like in the previous example, but the spacing will be different, because this pair is further apart.
plt.figure(figsize=(10, 10))
plt.imshow(util.zoom_cen(segs_10_16_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 10 and 16')
plt.show()
# #### *A different Zernike than piston*
#
# The principles described above will hold true for whatever Zernike we use to aberrate a segment pair. the morphology of the image will change with the chosen Zernike though, as we have see further above when we displayed the different envelopes coming from local Zernikes on the mini segment.
# +
im_list2 = np.array([segs_10_16_noll_2, segs_2_6_noll_2, segs_8_18_noll_2, segs_8_12_noll_2, \
segs_8_3_noll_2, segs_8_2_noll_2, segs_9_15_noll_2, segs_9_2_noll_2])
pair_list2 = np.array(['10-16', '2-6', '2-18', '8-12', '8-3', '8-2', '9-15', '9-2'])
plt.figure(figsize=(20, 50))
for i in range(im_list2.shape[0]):
plt.subplot(4, 2, i+1)
plt.imshow(util.zoom_cen(im_list2[i], boxw2), norm=LogNorm(), origin='lower')
plt.title('Segment pair ' + pair_list2[i])
plt.show()
# -
# If we compare the pair **9-2** between the **piston** and the **tip** version, we can see that especially the core loos different. And there is that extra dark vertical line in the tip image, although it would be much easier to spot with increased image sampling.
# +
plt.figure(figsize=(18, 9))
plt.subplot(1, 2, 1)
plt.imshow(util.zoom_cen(segs_9_2_noll_1, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 9 and 2 - Piston')
plt.subplot(1, 2, 2)
plt.imshow(util.zoom_cen(segs_9_2_noll_2, boxw2), norm=LogNorm(), origin='lower')
plt.title('Piston on segments 9 and 2 - Tip')
plt.show()
# -
|
notebooks/JWST and WebbPSF/2_Testing PASTIS in imaging mode before calibration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # datetime
#
# Python has the datetime module to help deal with timestamps in your code. Time values are represented with the time class. Times have attributes for hour, minute, second, and microsecond. They can also include time zone information. The arguments to initialize a time instance are optional, but the default of 0 is unlikely to be what you want.
#
# ## time
# Let's take a look at how we can extract time information from the datetime module. We can create a timestamp by specifying datetime.time(hour,minute,second,microsecond)
# +
import datetime
t = datetime.time(4, 20, 1)
# Let's show the different components
print(t)
print('hour :', t.hour)
print('minute:', t.minute)
print('second:', t.second)
print('microsecond:', t.microsecond)
print('tzinfo:', t.tzinfo)
# -
# Note: A time instance only holds values of time, and not a date associated with the time.
#
# We can also check the min and max values a time of day can have in the module:
print('Earliest :', datetime.time.min)
print('Latest :', datetime.time.max)
print('Resolution:', datetime.time.resolution)
# The min and max class attributes reflect the valid range of times in a single day.
# ## Dates
# datetime (as you might suspect) also allows us to work with date timestamps. Calendar date values are represented with the date class. Instances have attributes for year, month, and day. It is easy to create a date representing today’s date using the today() class method.
#
# Let's see some examples:
today = datetime.date.today()
print(today)
print('ctime:', today.ctime())
print('tuple:', today.timetuple())
print('ordinal:', today.toordinal())
print('Year :', today.year)
print('Month:', today.month)
print('Day :', today.day)
# As with time, the range of date values supported can be determined using the min and max attributes.
print('Earliest :', datetime.date.min)
print('Latest :', datetime.date.max)
print('Resolution:', datetime.date.resolution)
# Another way to create new date instances uses the replace() method of an existing date. For example, you can change the year, leaving the day and month alone.
# +
d1 = datetime.date(2015, 3, 11)
print('d1:', d1)
d2 = d1.replace(year=1990)
print('d2:', d2)
# -
# # Arithmetic
# We can perform arithmetic on date objects to check for time differences. For example:
d1
d2
d1-d2
# This gives us the difference in days between the two dates. You can use the timedelta method to specify various units of times (days, minutes, hours, etc.)
#
# Great! You should now have a basic understanding of how to use datetime with Python to work with timestamps in your code!
|
13-Advanced Python Modules/02-Datetime.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Sursa: Proiect Diana
#LUCRUL CU LISTE -> [" "]
#1. Creare Lista Actori
actoriLoveRain = ["Adriana"]
print("Lista de actori ai serialului Love Rain este:")
print(actoriLoveRain)
# -
#2. Lungime lista actori
print("Lungime lista:")
print(len(actoriLoveRain))
#3.Adaugare actrita Hwang BoRa la sfarsitul listei:
actoriLoveRain.append("Hwang BoRa")
print("Dupa adaugarea actritei Hwang BoRa:")
print(actoriLoveRain)
#4. Adaugarea actorului Seo In Guk pe pozitia 3 (numerotarea incepe de la 0)
actoriLoveRain.insert(3,"Seo In Guk")
print("Dupa adaugarea actorului Seo In Guk pe pozitia 3:")
print(actoriLoveRain)
#5. Eliminarea actritei Im Yoon Ah din lista
actoriLoveRain.remove("Im Yoon Ah")
print("Dupa eliminarea actritei Im Yoon Ah:")
print(actoriLoveRain)
#6. Inlocuirea primului actor cu actorul Yoo Seung Ho
actoriLoveRain[0]="Yoo Seung Ho"
print("Dupa inlocuirea primului actor:")
print(actoriLoveRain)
#7. Inversarea elementelor listei de actori
actoriLoveRain.reverse()
print("Dupa inversarea listei de actori:")
print(actoriLoveRain)
#8. Extragerea actorului de pe pozitia 5 (locul 6 - numerotarea se incepe de la 0)
print("Actorul de pe pozitia 5 este:")
actoriLoveRain.pop(5)
#9. Golire lista
actoriLoveRain.clear()
print("Lista de actori se va sterge:")
print(actoriLoveRain)
|
Proiect/2. Python/Code/01.Liste.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/Machine_Learning_in_Business/blob/main/BlackScholesReplicationExamplePython.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="YAfO2gP6xZss"
# Expect this program to run for 30 minutes or more
# Load package
import numpy as np
from scipy.stats import norm
import random
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
import tensorflow as tf
from tensorflow.keras.layers import Dense
from tensorflow import keras
from numpy.random import seed
seed(100)
import matplotlib.pyplot as plt
from IPython.display import clear_output
n = norm.pdf
N = norm.cdf
# + colab={"base_uri": "https://localhost:8080/", "height": 322} id="YAwzhEWwxZsv" outputId="db2c967a-8cb4-480f-a12a-c7612448d9e1"
# Load option data
option_dataset = pd.read_csv('Option_Data.csv')
option_dataset.head()
# + [markdown] id="Ewaa9r1sxZsw"
# ## Divide data into Training, Validation and Test set
# + id="AmvwaPeYxZsy"
# Include option price with and without noise in data set splitting for later BS mean error calculation on test set
y = option_dataset[['Option Price with Noise','Option Price']]
X = option_dataset[['Spot price', 'Strike Price', 'Risk Free Rate','Volatility','Maturity','Dividend']]
# Divide data into training set and test set(note that random seed is set)
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.2,random_state=100)
# Divide training set into training and validation set
X_train,X_val,y_train,y_val=train_test_split(X_train,y_train,test_size=0.25,random_state=100)
# + [markdown] id="DFY2eL7NxZsy"
# ## Feature Scaling
# + id="VUUyDpZcxZsz"
# Scale features based on Z-Score
scaler = StandardScaler()
scaler.fit(X_train)
X_scaled_train = scaler.transform(X_train)
X_scaled_vals = scaler.transform(X_val)
X_scaled_test = scaler.transform(X_test)
y_train = np.asarray(y_train)
y_val = np.asarray(y_val)
y_test = np.asarray(y_test)
# + [markdown] id="ZmeyBNfwxZs0"
# ## Define Neural Network
# + colab={"base_uri": "https://localhost:8080/"} id="Obkq4SdMxZs1" outputId="490eb98b-42af-45d1-a8ca-66e332febb8e"
# Create ML Model
# Sequential function allows you to define your Neural Network in sequential order
# Within Sequential, use Dense function to define number of nodes, activation function and other related parameters
# For more information regrading to activation functoin, please refer to https://keras.io/activations/
model = keras.models.Sequential([Dense(20,activation = "sigmoid",input_shape = (6,)),
Dense(20,activation = "sigmoid"),Dense(20,activation = "sigmoid"),
Dense(1)])
# Model summary function shows what you created in the model
model.summary()
# + id="uV4Q4yXWxZs2"
# Complie function allows you to choose your measure of loss and optimzer
# For other optimizer, please refer to https://keras.io/optimizers/
model.compile(loss = "mae",optimizer = "Adam")
# + id="9383j2TxxZs3"
# Checkpoint function is used here to periodically save a copy of the model.
# Currently it is set to save the best performing model
checkpoint_cb = keras.callbacks.ModelCheckpoint("bs_pricing_model_vFinal.h5",save_best_only = True)
# Early stopping allows you to stop your training early if no improvment is shown after cerain period
# Currently it is set at if no improvement occured in 5000 epochs, at the stop the model will also revert back to the best weight
early_stopping_cb = keras.callbacks.EarlyStopping(patience = 5000,restore_best_weights = True)
# Remark: checkpoint could be redundant here as early stopping function can also help restoring to the best weight
# We put both here just to illustrate different ways to keep the best model
# + id="xxVlQjLDxZs4"
# train your model
# The fit function allows you to train a NN model. Here we have training data, number of epochs, validation data,
# and callbacks as input
# Callback is an optional parameters that allow you to enable tricks for training such as early stopping and checkpoint
# Remarks: Altough we put 50000 epochs here, the model will stop its training once our early stopping criterion is triggered
# Also, select the first column of y_train data array, which is the option price with noise column
history=model.fit(X_scaled_train,y_train[:,0],epochs= 50000,verbose = 0, validation_data=(X_scaled_vals,y_val[:,0]),
callbacks=[checkpoint_cb,early_stopping_cb])
# + [markdown] id="vfTYxPXKxZs4"
# ## Calculate prediction error for both NN and BS analytical formula
# + colab={"base_uri": "https://localhost:8080/"} id="mLh-mICuxZs5" outputId="6964fd37-93fb-47c2-a3c3-47edcf830988"
# Load the best model you saved and calcuate MAE for testing set
model = keras.models.load_model("bs_pricing_model_vFinal.h5")
mae_test = model.evaluate(X_scaled_test,y_test[:,0],verbose=0)
print('Nerual network mean absoluste error on test set:', mae_test)
# + colab={"base_uri": "https://localhost:8080/"} id="wpK_hOmCxZs5" outputId="52509fc3-f83b-49ab-eb2c-6f22ca2f3c15"
model_prediction = model.predict(X_scaled_test)
mean_error = np.average(model_prediction.T - y_test[:,0])
std_error = np.std(model_prediction.T - y_test[:,0])
mean_error_vs_BS_price = np.average(model_prediction.T - y_test[:,1])
std_error_vs_BS_price = np.std(model_prediction.T - y_test[:,1])
BS_mean_error = np.average(y_test[:,0] - y_test[:,1])
BS_std_error = np.std(y_test[:,0] - y_test[:,1])
print('Black-Scholes Statistics:')
print('Mean error on test set:',BS_mean_error)
print('Standard deviation of error on test set:',BS_std_error)
print(" ")
print('Neural Network Statistics:')
print('Mean error on test set vs. option price with noise:',mean_error)
print('Standard deviation of error on test set vs. option price with noise:',std_error)
print('Mean error on test set vs. BS analytical formula price:',mean_error_vs_BS_price)
print('Standard deviation of error on test set vs. BS analytical formula price:',std_error_vs_BS_price)
# + [markdown] id="X0AK3C55xZs6"
# ## Review your results and export training history
# + colab={"base_uri": "https://localhost:8080/", "height": 324} id="cdBeCS9XxZs6" outputId="2baf40c5-21d9-448b-a307-a93bf084694c"
# Plot training history
pd.DataFrame(history.history).plot(figsize=(8,5))
plt.grid(True)
plt.gca().set_ylim(0.1,0.2)
plt.show()
#Export your training history for MSE
output = pd.DataFrame(history.history)
output.to_csv("mae_history.csv")
# + id="tCuQ6d8PxZs6"
|
BlackScholesReplicationExamplePython.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting 1-D data
#
# `scipp` offers a number of different ways to plot data from a `DataArray` or a `Dataset`. It uses the `matplotlib` graphing library to do so.
import numpy as np
import scipp as sc
# ## Basic plot
#
# Plotting is done using the `plot` function.
# Generally the information in a dataset is sufficient to produce a useful plot out of the box.
#
# For example, a simple plot from a 1D dataset is produced as follows:
d = sc.Dataset()
N = 50
d['Signal'] = sc.Variable(dims=['tof'], values=5.0 + 5.0*np.sin(np.arange(N, dtype=float)/5.0),
unit=sc.units.counts)
d.coords['tof'] = sc.Variable(dims=['tof'], values=np.arange(N).astype(float),
unit=sc.units.us)
sc.plot(d)
# ## With error bars
#
# Error bars are shown automatically if variances are present in the data:
d['Signal'].variances = np.square(np.random.random(N))
sc.plot(d)
# Note that the length of the errors bars is the standard-deviation, i.e., the square root of the variances stored in the data.
#
# ## Multiple variables on the same axes
#
# ### Plotting a Dataset with multiple entries
#
# If a dataset contains more than one 1D variable with the same coordinates, they are plotted on the same axes:
d['Background'] = sc.Variable(dims=['tof'], values=3.0*np.random.random(N),
unit=sc.units.counts)
sc.plot(d)
# It is possible to hide the error bars with
sc.plot(d, errorbars=False)
# We can always plot just a single item of the dataset.
sc.plot(d['Background'])
# ### Overplotting using a dict of DataArrays
#
# One can also supply the `plot` function with a `dict` of data arrays.
# Compatible data arrays will be overplotted on the same axes:
sc.plot({"My sample": d['Signal'], "My background": d["Background"]})
# Note that the newly supplied names (keys) have also been adopted as the graph names.
#
# We can also overplot sections of interest with the help of slicing.
sc.plot({"My sample": d['Signal']['tof', 10:40], "My background": d["Background"]})
# This overplotting is useful for plotting slices of a 2D data array:
M = 100
L = 5
xx = np.arange(M, dtype=float)
yy = np.arange(L, dtype=float)
x, y = np.meshgrid(xx, yy)
b = M/20.0
c = M/5.0
e = L/10.0
r = np.sqrt(((x-c)/b)**2 + (y/e)**2)
a = np.sin(r)
d2d = sc.DataArray(data=sc.Variable(dims=['y', 'x'], values=a, unit=sc.units.counts))
d2d.coords['x'] = sc.Variable(dims=['x'], values=xx, unit=sc.units.m)
d2d.coords['y'] = sc.Variable(dims=['y'], values=yy, unit=sc.units.m)
sc.plot({f'slice-{i}': d2d['y', i] for i in range(L)})
# Or using the `collapse` helper function, which returns a `dict` of data arrays:
sc.plot(sc.collapse(d2d, keep='x'))
# ## Customizing linestyles, markers and colors
#
# Linestyles can be customized following the Matplotlib syntax.
# For instance, it is possible to connect the dots by setting `linestyle='solid'`:
sc.plot(d, linestyle='solid')
# Marker colors and symbols can be changed via the `color` and `marker` keyword arguments:
sc.plot(d, color=['red', '#30D5F9'], marker=['s', 'x'])
# The supplied `color` and `marker` arguments can also be a list of integers, which correspond to one of the pre-defined [colors](https://matplotlib.org/3.1.1/users/dflt_style_changes.html) or [markers](https://matplotlib.org/3.1.1/api/markers_api.html) (which were taken from matplotlib). In addition, the grid can also be displayed:
sc.plot(d, color=[6, 8], grid=True)
# ## Logarithmic scales
#
# Use the keyword arguments `scale` and `norm` to apply logarithmic scales to the horizontal and vertical axes, respectively.
sc.plot(d, norm='log', scale={'tof': 'log'})
# ## Histograms
# Histograms are automatically generated if the coordinate is bin edges:
histogram = sc.DataArray(data=sc.Variable(dims=['tof'],
values=20.0 + 20.0*np.cos(np.arange(N-1, dtype=float) / 3.0),
unit=sc.units.counts),
coords={'tof':d.coords['tof']})
sc.plot(histogram)
# and with error bars
histogram.variances = 5.0*np.random.random(N-1)
sc.plot(histogram)
# The histogram color can be customized:
sc.plot(histogram, color='#000000')
# ## Multiple 1D variables with different dimensions
#
# `scipp.plot` also supports multiple 1-D variables with different dimensions (note that the data entries are grouped onto the same graph if they have the same dimension and unit):
M = 60
d['OtherSample'] = sc.Variable(dims=['x'], values=10.0*np.random.rand(M),
unit=sc.units.s)
d['OtherNoise'] = sc.Variable(dims=['x'], values=7.0*np.random.rand(M),
variances=3.0*np.random.rand(M),
unit=sc.units.s)
d['SomeKgs'] = sc.Variable(dims=['x'], values=20.0*np.random.rand(M),
unit=sc.units.kg)
d.coords['x'] = sc.Variable(dims=['x'],
values=np.arange(M).astype(float),
unit=sc.units.m)
sc.plot(d)
# ## Custom labels along x axis
#
# Sometimes one wishes to have labels along the `x` axis instead of the dimension-coordinate.
# This can be achieved via the `labels` keyword argument by specifying which dimension should run along the `x` axis:
d1 = sc.Dataset()
N = 100
x = np.arange(N, dtype=float)
d1['Sample'] = sc.Variable(dims=['tof'],
values=np.sqrt(x),
unit=sc.units.counts)
d1.coords['tof'] = sc.Variable(dims=['tof'],
values=x,
unit=sc.units.us)
d1.coords['somelabels'] = sc.Variable(dims=['tof'],
values=np.linspace(101., 105., N),
unit=sc.units.s)
sc.plot(d1, labels={'tof': 'somelabels'})
# ## Plotting masks
#
# If a dataset item contains masks, the symbols of masks data points will have a thick black contour in a 1D plot:
d4 = sc.Dataset()
N = 50
x = np.arange(N).astype(float)
d4['Sample'] = sc.Variable(dims=['tof'], values=3*np.sin(x/5)+3,
unit=sc.units.counts)
d4['Background'] = sc.Variable(dims=['tof'], values=1.0*np.random.rand(N),
unit=sc.units.counts)
d4.coords['tof'] = sc.Variable(dims=['tof'], values=x,
unit=sc.units.us)
d4['Sample'].masks['mask1'] = sc.Variable(dims=['tof'],
values=np.where(np.abs(x-40) < 10, True, False))
d4['Background'].masks['mask1'] = ~d4['Sample'].masks['mask1']
d4['Background'].masks['mask1']['tof', 0:20].values = np.zeros(20, dtype=bool)
sc.plot(d4)
# Checkboxes below the plot can be used to hide/show the individual masks.
# A global toggle button is also available to hide/show all masks in one go.
#
# The color of the masks can be changed as follows:
sc.plot(d4, masks={'color': 'red'})
# ### Masks on histograms
#
# Masks on a histogram show up as a thick black line:
N = 50
x = np.arange(N+1).astype(float)
d4 = sc.DataArray(data=sc.Variable(dims=['tof'], values=3*np.sin(x[:-1]/5)+3, unit=sc.units.counts))
d4.coords['tof'] = sc.Variable(dims=['tof'], values=x, unit=sc.units.us)
d4.masks['mask1'] = sc.Variable(dims=['tof'], values=np.where(np.abs(x[:-1]-40) < 10, True, False))
sc.plot(d4)
# ## Plotting time series
#
# When plotting data with time-series (`dtype=datetime64`) coordinates,
# a special axis tick label formatting, which dynamically adapts to the zoom level, is used.
time = sc.array(dims=['time'], values=np.arange(np.datetime64('2017-01-01T12:00:00'),
np.datetime64('2017-01-01T13:00:00'), 20))
N = time.sizes['time']
data = sc.DataArray(data=sc.array(dims=['time'],
values=np.arange(N) + 50.*np.random.random(N),
unit="K"),
coords={'time':time})
data.plot(title="Temperature as a function of time")
# ## Saving figures
# Static `pdf` or `png` copies of the figures can be saved to file (note that any buttons displayed under a figure are not saved to file). This is achieved as follows:
sc.plot(d4, filename='my_1d_figure.pdf')
|
docs/visualization/plotting/plotting-1d-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating a QComponent - Basic
# Now that you have become familiar with Qiskit Metal and feel comfortable using the available aspects and functionality, the next step is learning how to make your own circuit component in Metal.
#
# We will start off by going over the sample `my_qcomponent` in `qiskit_metal>qlibrary>user_components` as a basis, which we will walk through below.
# ## Reviewing my_qcomponent
# +
# -*- coding: utf-8 -*-
# This code is part of Qiskit.
#
# (C) Copyright IBM 2017, 2021.
#
# This code is licensed under the Apache License, Version 2.0. You may
# obtain a copy of this license in the LICENSE.txt file in the root directory
# of this source tree or at http://www.apache.org/licenses/LICENSE-2.0.
#
# Any modifications or derivative works of this code must retain this
# copyright notice, and modified files need to carry a notice indicating
# that they have been altered from the originals.
# -
# Always be sure to include the proper copyright and license information, and give yourself credit for any components you create!
from qiskit_metal import draw, Dict
from qiskit_metal.toolbox_metal import math_and_overrides
from qiskit_metal.qlibrary.core import QComponent
# Import any classes or functions you will be wanting to use when creating your component. The geometries in Metal are shapely objects, which can be readily generated and manipulated through functions in `draw`. Mathematical functions can be accessed via `math_and_overrides`. Any imports that are part of the Metal requirement list can also be used.
#
# The key import is what the parent class of your new component will be. For this example, we are using `QComponent` as the parent for `MyQComponent`, which is the base component class in Metal and contains a great deal of automated functionality. All component hierarchy must have QComponent as the top base class.
dir(QComponent)
# `MyQComponent` creates a simple rectangle of a variable width, height, position and rotation.
class MyQComponent(QComponent):
"""
Use this class as a template for your components - have fun
Description:
Options:
"""
# Edit these to define your own tempate options for creation
# Default drawing options
default_options = Dict(width='500um',
height='300um',
pos_x='0um',
pos_y='0um',
orientation='0',
layer='1')
"""Default drawing options"""
# Name prefix of component, if user doesn't provide name
component_metadata = Dict(short_name='component',
_qgeometry_table_poly='True')
"""Component metadata"""
def make(self):
"""Convert self.options into QGeometry."""
p = self.parse_options() # Parse the string options into numbers
# EDIT HERE - Replace the following with your code
# Create some raw geometry
# Use autocompletion for the `draw.` module (use tab key)
rect = draw.rectangle(p.width, p.height, p.pos_x, p.pos_y)
rect = draw.rotate(rect, p.orientation)
rect = draw.translate(rect,p.pos_x,p.pos_y)
geom = {'my_polygon': rect}
self.add_qgeometry('poly', geom, layer=p.layer, subtract=False)
# The docstring at the start of the class should clearly explain what the component is, what the parameterized values of the component refer to (in a sense the 'inputs'), and any other information you believe would be relevant for a user making use of your component.
#
# `default_options` is a dictionary to be included in the class of all components. The keywords, of type string, in the default dictionary are the parameters the front-end user is allowed to modify. The keywords in the above indicate that the width and height can be modified via the components options, but have a default value of 500um and 300 um respectively. Further, the position and rotation can also be changed. The `layer` is an expected keyword in a default dictionary, as it is used by renderers to help determine further properties of the `qgeometry` of the component when rendered, eg. GDS QRenderer uses the layer # to define which layer the qgeometry is on.
#
# `component_metadata` is a dictionary which contains some important pieces of information, such as the default/shorthand name of the component (`short_name`), or indicating what types of qgeometry tables are included in this component, eg. `_qgeometry_table_poly='True'`.
# The `component_metadata` must contain the flags for each type of qgeometry table being used via `add_qgeometry` methods at the end of the `make()` function, in order for renderer options to be updated correctly. Currently the options are:
#
# `_qgeometry_table_path='True'`
#
# `_qgeometry_table_poly='True'`
#
# `_qgeometry_table_junction='True'`
#
#
# The `make()` method is where the actual generation of the component's layout is written.
# ### The make() method
# Although not required, a good first line is `p = self.parse_options()` to cut down on your amount of typing. The `parse_options()` translates the keywords in `self.options` from strings into their appropriate value with respect to the prefix included, e.g.,`p.width`=> "500um" -> 0.0005
#
# Following this, all code generating the shapely geometries that are to represent your circuit layout should be written, via the `draw` module or even written in directly. It is a good practice to play around with the geometries in a jupyter notebook first for quick visual feedback, such as:
draw.rectangle(1,2,0,0)
draw.rotate(draw.rectangle(1,2,0,0), 45)
# Or for a little more complexity:
# +
face = draw.shapely.geometry.Point(0, 0).buffer(1)
eye = draw.shapely.geometry.Point(0, 0).buffer(0.2)
eye_l = draw.translate(eye, -0.4, 0.4)
eye_r = draw.translate(eye, 0.4, 0.4)
smile = draw.shapely.geometry.Point(0, 0).buffer(0.8)
cut_sq = draw.shapely.geometry.box(-1, -0.3, 1, 1)
smile = draw.subtract(smile, cut_sq)
face = draw.subtract(face, smile)
face = draw.subtract(face, eye_r)
face = draw.subtract(face, eye_l)
face
# -
# Once you are happy with your geometries, and have them properly parameterized to allow the Front End User as much customization of your component as you wish, it is time to convert the geometries into Metal `qgeometries` via `add_qgeometry`
# #### add_qgeometry
import qiskit_metal as metal
# ?metal.qlibrary.core.QComponent.add_qgeometry
# `add_qgeometry` is the method by which the shapely geometries you have drawn are converted into Metal qgeometries, the format which allows for the easy translatability between different renderers and the variable representation of quantum elements such as Josephson junctions.
#
# Currently there are three kinds of qgeometries, `path`, `poly` and `junction`.
# `path` -> shapely LineString
# `poly` -> any other shapely geometry (currently)
# `junction` -> shapely LineString
#
# Both `path` and `junction` also take and input of `width`, with is added to the qgeometry table to inform renderers of, as an example, how much to buffer the LineString of a cpw transmission line to turn it into a proper 2D sheet.
#
# `subtract` indicates this qgeometry is to be subtracted from the ground plane of that layer#. A ground plane is automatically included for that layer at the dimension of the chip size if any qgeometry has `subtract = True`. As an example, a cpw transmission line's dielectric gap could be drawn by using the same LineString as previously, having the `width = cpw_width + 2*cpw_gap` and setting `subtract = True`.
#
# `fillet` is an option that informs a renderer that the vertices of this qgeometry are to be filleted by that value (eg. `fillet = "100um"`).
# #### add_pin
# ?metal.qlibrary.core.QComponent.add_pin
# The final step for creating your QComponent is adding of pins. This is not necessarily a requirement for your component, but to have full functionality with Metal and be able to make use of auto-routing components with it, you will want to indicate where the "ports" of your component are.
#
# Following from the above docstring, pins can be added from two coordinates indicating either an orthogonal vector to the port plane of your component, or a tangent to the port plane of your component. For the former, you want the vector to be pointing to the middle point of your intended plane, with the latter being across the length of your intended plane (as indicated in the above figure). The `width` should be the size of the plane (say, in the case of a CPW transmission line, the trace width), with the `gap` following the same logic (though this value can be ignored for any non-coplanar structure).
# ## Example
#
#
# Below is a simple QComponent that implements everything we have reviewed.
# +
from qiskit_metal import draw, Dict
from qiskit_metal.toolbox_metal import math_and_overrides
from qiskit_metal.qlibrary.core import QComponent
class MySimpleGapCapacitor(QComponent):
"""
Inherits 'QComponent' class.
Description:
A simple CPW style gap capacitor, with endcap islands each coupled to their own
cpw transmission line that ends in a pin.
Options:
* cpw_width: width of the cpw trace of the transmission line
* cpw_gap: dielectric gap of the cpw transmission line
* cap_width: width of the gap capacitor (size of the charge islands)
* cap_gap: dielectric space between the two islands
* pos_x/_y: position of the capacitor on chip
* orientation: 0-> is parallel to x-axis, with rotation (in degrees) counterclockwise.
* layer: the layer number for the layout
"""
# Edit these to define your own tempate options for creation
# Default drawing options
default_options = Dict(cpw_width='15um',
cpw_gap='9um',
cap_width='35um',
cap_gap='3um',
pos_x='0um',
pos_y='0um',
orientation='0',
layer='1')
"""Default drawing options"""
# Name prefix of component, if user doesn't provide name
component_metadata = Dict(short_name='component',
_qgeometry_table_poly='True',
_qgeometry_table_path='True')
"""Component metadata"""
def make(self):
"""Convert self.options into QGeometry."""
p = self.parse_options() # Parse the string options into numbers
pad = draw.rectangle(p.cpw_width, p.cap_width, 0, 0)
pad_left = draw.translate(pad,-(p.cpw_width+p.cap_gap)/2,0)
pad_right = draw.translate(pad,(p.cpw_width+p.cap_gap)/2,0)
pad_etch = draw.rectangle(2*p.cpw_gap+2*p.cpw_width+p.cap_gap,2*p.cpw_gap+p.cap_width)
cpw_left = draw.shapely.geometry.LineString([[-(p.cpw_width+p.cap_gap/2),0],[-(p.cpw_width*3 +p.cap_gap/2),0]])
cpw_right = draw.shapely.geometry.LineString([[(p.cpw_width+p.cap_gap/2),0],[(p.cpw_width*3 +p.cap_gap/2),0]])
geom_list = [pad_left,pad_right,cpw_left,cpw_right,pad_etch]
geom_list = draw.rotate(geom_list,p.orientation)
geom_list = draw.translate(geom_list,p.pos_x,p.pos_y)
[pad_left,pad_right,cpw_left,cpw_right,pad_etch] = geom_list
self.add_qgeometry('path', {'cpw_left':cpw_left, 'cpw_right':cpw_right}, layer=p.layer, width = p.cpw_width)
self.add_qgeometry('path', {'cpw_left_etch':cpw_left, 'cpw_right_etch':cpw_right}, layer=p.layer, width = p.cpw_width+2*p.cpw_gap, subtract=True)
self.add_qgeometry('poly', {'pad_left':pad_left, 'pad_right':pad_right}, layer=p.layer)
self.add_qgeometry('poly', {'pad_etch':pad_etch}, layer=p.layer, subtract=True)
self.add_pin('cap_left', cpw_left.coords, width = p.cpw_width, gap = p.cpw_gap, input_as_norm=True)
self.add_pin('cap_right', cpw_right.coords, width = p.cpw_width, gap = p.cpw_gap, input_as_norm=True)
# +
design = metal.designs.DesignPlanar()
gui = metal.MetalGUI(design)
# -
my_cap = MySimpleGapCapacitor(design,'my_cap')
gui.rebuild()
gui.autoscale()
# You should now see *my_cap* in the Metal gui. One can work on the layout of the component through these cells by changing the above class, such as how the parameterized values are used. By enabling `overwrite_enabled` (which should normally be kept to False), the code can quickly be iterated through until you, the component designer, is happy with the qcomponent you have just created.
design.overwrite_enabled = True
# We will delve into more complex QComponent topics in the next notebook, `Creating a QComponent - Advanced` </br>
#
# Use the command below to close Metal GUI.
# + tags=["nbsphinx-thumbnail"]
gui.screenshot()
# -
gui.main_window.close()
|
docs/tut/components/3.2 Creating a QComponent - Basic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sbhattac/ai-workshop/blob/master/ANN/AI_Workshop_ANN.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="o7DsY44Bvla4" colab_type="text"
# # Artificial Neural Networks (ANNs)
#
# ### A Team Project to:
# * Introduce basic concepts of ANNs
# * Demonstrate set of computational techniques inspired by natural neural systems
# * To simulate or model natural systems with ANNs
#
# <p><em><strong>Attribution: This jupyter notebook builds upon material found at <a href="https://cspogil.org/Home">https://cspogil.org/Home</a> </strong></em></p>
# <p> </p>
# <p><a href="http://creativecommons.org/licenses/by-nc-sa/4.0/" rel="license"><img style="border-width: 0;" src="https://i.creativecommons.org/l/by-nc-sa/4.0/88x31.png" alt="Creative Commons License" /></a><br />This work is licensed under a <a href="http://creativecommons.org/licenses/by-nc-sa/4.0/" rel="license">Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License</a>.</p>
#
# + id="3RYUC50U_O15" colab_type="code" cellView="form" colab={}
#@title # Assigning Roles and Responsibilities within each group
#@markdown ---
#@markdown ### Enter Instructor Name:
Instructor_Name = "" #@param {type:"string"}
#@markdown 1. Introduces activities. Assigns roles to participants.
#@markdown 2. Responds for help or clarification request.
#@markdown 3. Collects the Jupiter notebooks from Recorder and Evaluator
#@markdown ---
#@markdown ### Enter Facilitator Name:
Facilitator_Name = "" #@param {type:"string"}
#@markdown ### Enter Backup Facilitator Name:
Backup_Facilitator_Name = "" #@param {type:"string"}
#@markdown 1. Reads aloud each question and ask for volunteers to answer. If there is no volunteer then he/she starts the discussion and asks one participant after another for comments, solutions, answers, or clarifications. When majority participants agree then she/he asks Recorder to record the answer. Also coordinates discussion about the code execution and the output like any other question.
#@markdown 2. Involves each participant equally in the discussions.
#@markdown 3. Turn the coordinating role to Evaluator after finishing each activity.
#@markdown ---
#@markdown ### Enter Recorder Name:
Recorder_Name = "" #@param {type:"string"}
#@markdown ### Enter Backup Recorder Name:
Backup_Recorder_Name = "" #@param {type:"string"}
#@markdown 1. Coordinates Zoom screen access. Displays his/her screen when asking questions. Gives access to screen sharing as requested
#@markdown 2. Records all answers for each question inside the Jupiter Notebook
#@markdown 3. Use "Run all" in menu "Runtime" and then "Save" Jupiter Workbook with all answers and results.
#@markdown 4. Submit Jupiter notebook with all answers and results of the running code.
#@markdown ---
#@markdown ### Enter Evaluator Name:
Evaluator_Name = "" #@param {type:"string"}
#@markdown ### Enter Backup Evaluator Name:
Backup_Evaluator_Name = "" #@param {type:"string"}
#@markdown 1. Keeps track of time for each designated Activity.
#@markdown 2. After each activity leads the discussion about material and collects feedback in the form of the table below.
#@markdown 3. Submit Jupiter notebook with all comments and results of discussion at the end of each activity.
#@markdown ---
#@markdown ### Enter Participant names
Participant_4_Name = "" #@param {type:"string"}
Participant_5_Name = "" #@param {type:"string"}
Participant_6_Name = "" #@param {type:"string"}
Participant_7_Name = "" #@param {type:"string"}
Participant_8_Name = "" #@param {type:"string"}
#@markdown 1. Participates actively in team work to answer all questions.
#@markdown 2. Executes the code and shares the comments.
#@markdown ---
# + id="Y8eS79mDTxxl" colab_type="code" cellView="form" colab={}
#@title (RUN CELL) Download files to work on and install necessary libraries
# !pip install palettable
# + id="wZKeKvW7Vnq-" colab_type="code" cellView="form" colab={}
#@title (RUN CELL) Library Imports
from matplotlib import pyplot
import matplotlib.patches as patches
from math import cos, sin, atan
from palettable.tableau import Tableau_10
from time import localtime, strftime
import torch
from torchvision import datasets, transforms
import torch.nn.functional as F
from torch import nn
import ipywidgets as widgets
import numpy as np
import matplotlib.pyplot as plt
# + id="6C7vlhwtVl9F" colab_type="code" cellView="form" colab={}
#@title (RUN CELL) Helper Classes and Functions for ANN visualization
# This code is a modified version of the libray written by <NAME> found at https://github.com/jzliu-100/visualize-neural-network.git
from matplotlib import pyplot
import matplotlib.patches as patches
from math import cos, sin, atan
from palettable.tableau import Tableau_10
from time import localtime, strftime
import numpy as np
class Neuron():
def __init__(self, x, y):
self.x = x
self.y = y
def draw(self, neuron_radius, id=-1):
circle = pyplot.Circle((self.x, self.y), radius=neuron_radius, fill=False)
pyplot.gca().add_patch(circle)
pyplot.gca().text(self.x, self.y-0.15, str(id), size=10, ha='center')
class Layer():
def __init__(self, network, number_of_neurons, number_of_neurons_in_widest_layer):
self.vertical_distance_between_layers = 6
self.horizontal_distance_between_neurons = 2
self.neuron_radius = 0.5
self.number_of_neurons_in_widest_layer = number_of_neurons_in_widest_layer
self.previous_layer = self.__get_previous_layer(network)
self.y = self.__calculate_layer_y_position()
self.neurons = self.__intialise_neurons(number_of_neurons)
def __intialise_neurons(self, number_of_neurons):
neurons = []
x = self.__calculate_left_margin_so_layer_is_centered(number_of_neurons)
for iteration in range(number_of_neurons):
neuron = Neuron(x, self.y)
neurons.append(neuron)
x += self.horizontal_distance_between_neurons
return neurons
def __calculate_left_margin_so_layer_is_centered(self, number_of_neurons):
return self.horizontal_distance_between_neurons * (self.number_of_neurons_in_widest_layer - number_of_neurons) / 2
def __calculate_layer_y_position(self):
if self.previous_layer:
return self.previous_layer.y + self.vertical_distance_between_layers
else:
return 0
def __get_previous_layer(self, network):
if len(network.layers) > 0:
return network.layers[-1]
else:
return None
def __line_between_two_neurons(self, neuron1, neuron2, i,j, weight=0.4, textoverlaphandler=None, show_weights=True):
angle = atan((neuron2.x - neuron1.x) / float(neuron2.y - neuron1.y))
x_adjustment = self.neuron_radius * sin(angle)
y_adjustment = self.neuron_radius * cos(angle)
# assign colors to lines depending on the sign of the weight
color=Tableau_10.mpl_colors[0]
if weight > 0: color=Tableau_10.mpl_colors[1]
# assign different linewidths to lines depending on the size of the weight
abs_weight = abs(weight)
if abs_weight > 0.5:
linewidth = 10*abs_weight
elif abs_weight > 0.8:
linewidth = 100*abs_weight
else:
linewidth = abs_weight
# draw the weights and adjust the labels of weights to avoid overlapping
if True: #abs_weight > 0.5:
# while loop to determine the optimal locaton for text lables to avoid overlapping
index_step = 2
num_segments = 10
txt_x_pos = neuron1.x - x_adjustment+index_step*(neuron2.x-neuron1.x+2*x_adjustment)/num_segments
txt_y_pos = neuron1.y - y_adjustment+index_step*(neuron2.y-neuron1.y+2*y_adjustment)/num_segments
while ((not textoverlaphandler.getspace([txt_x_pos-0.5, txt_y_pos-0.5, txt_x_pos+0.5, txt_y_pos+0.5])) and index_step < num_segments):
index_step = index_step + 1
txt_x_pos = neuron1.x - x_adjustment+index_step*(neuron2.x-neuron1.x+2*x_adjustment)/num_segments
txt_y_pos = neuron1.y - y_adjustment+index_step*(neuron2.y-neuron1.y+2*y_adjustment)/num_segments
# print("Label positions: ", "{:.2f}".format(txt_x_pos), "{:.2f}".format(txt_y_pos), "{:3.2f}".format(weight))
# a=pyplot.gca().text(txt_x_pos, txt_y_pos, "{:3.2f}".format(weight), size=8, ha='center')
if show_weights:
a=pyplot.gca().text(txt_x_pos, txt_y_pos, "w"+str(i+1)+str(j+1), size=8, ha='left',fontsize=12, color='red')
a.set_bbox(dict(facecolor='white', alpha=0))
# print(a.get_bbox_patch().get_height())
line = pyplot.Line2D((neuron1.x - x_adjustment, neuron2.x + x_adjustment), (neuron1.y - y_adjustment, neuron2.y + y_adjustment), linewidth=linewidth, color=color)
x1,x2 = neuron1.x - x_adjustment, neuron2.x + x_adjustment
y1,y2 = neuron1.y - y_adjustment, neuron2.y + y_adjustment
pyplot.annotate("",
xy=(x1, y1), xycoords='data',
xytext=(x2, y2), textcoords='data',
arrowprops=dict(arrowstyle="->", color="0",
shrinkA=5, shrinkB=5,
patchA=None, patchB=None,
connectionstyle="arc3,rad=0.",
),
)
#pyplot.gca().add_line(line)
def draw(self, layerType=0, weights=None, textoverlaphandler=None, show_weights=True):
j=0 # index for neurons in this layer
for neuron in self.neurons:
i=0 # index for neurons in previous layer
neuron.draw( self.neuron_radius, id=j+1 )
if self.previous_layer:
for previous_layer_neuron in self.previous_layer.neurons:
self.__line_between_two_neurons(neuron, previous_layer_neuron, i,j, weights[i,j], textoverlaphandler, show_weights=show_weights)
i=i+1
j=j+1
# write Text
x_text = self.number_of_neurons_in_widest_layer * self.horizontal_distance_between_neurons
if layerType == 0:
pyplot.text(x_text, self.y, 'Input Layer', fontsize = 12)
elif layerType == -1:
pyplot.text(x_text, self.y, 'Output Layer', fontsize = 12)
else:
pyplot.text(x_text, self.y, 'Hidden Layer '+str(layerType), fontsize = 12)
# A class to handle Text Overlapping
# The idea is to first create a grid space, if a grid is already occupied, then
# the grid is not available for text labels.
class TextOverlappingHandler():
# initialize the class with the width and height of the plot area
def __init__(self, width, height, grid_size=0.2):
self.grid_size = grid_size
self.cells = np.ones((int(np.ceil(width / grid_size)), int(np.ceil(height / grid_size))), dtype=bool)
# input test_coordinates(bottom left and top right),
# getspace will tell you whether a text label can be put in the test coordinates
def getspace(self, test_coordinates):
x_left_pos = int(np.floor(test_coordinates[0]/self.grid_size))
y_botttom_pos = int(np.floor(test_coordinates[1]/self.grid_size))
x_right_pos = int(np.floor(test_coordinates[2]/self.grid_size))
y_top_pos = int(np.floor(test_coordinates[3]/self.grid_size))
if self.cells[x_left_pos, y_botttom_pos] and self.cells[x_left_pos, y_top_pos] \
and self.cells[x_right_pos, y_top_pos] and self.cells[x_right_pos, y_botttom_pos]:
for i in range(x_left_pos, x_right_pos):
for j in range(y_botttom_pos, y_top_pos):
self.cells[i, j] = False
return True
else:
return False
class NeuralNetwork():
def __init__(self, number_of_neurons_in_widest_layer):
self.number_of_neurons_in_widest_layer = number_of_neurons_in_widest_layer
self.layers = []
self.layertype = 0
def add_layer(self, number_of_neurons ):
layer = Layer(self, number_of_neurons, self.number_of_neurons_in_widest_layer)
self.layers.append(layer)
def draw(self, weights_list=None, show_weights=True):
# vertical_distance_between_layers and horizontal_distance_between_neurons are the same with the variables of the same name in layer class
vertical_distance_between_layers = 6
horizontal_distance_between_neurons = 2
overlaphandler = TextOverlappingHandler(\
self.number_of_neurons_in_widest_layer*horizontal_distance_between_neurons,\
len(self.layers)*vertical_distance_between_layers, grid_size=0.2 )
pyplot.figure(figsize=(12, 9))
for i in range( len(self.layers) ):
layer = self.layers[i]
if i == 0:
layer.draw( layerType=0, show_weights=show_weights)
elif i == len(self.layers)-1:
layer.draw( layerType=-1, weights=weights_list[i-1], textoverlaphandler=overlaphandler, show_weights=show_weights)
else:
layer.draw( layerType=i, weights=weights_list[i-1], textoverlaphandler=overlaphandler, show_weights=show_weights)
pyplot.axis('scaled')
pyplot.axis('off')
pyplot.title( 'Neural Network architecture', fontsize=15 )
#figureName='ANN_'+strftime("%Y%m%d_%H%M%S", localtime())+'.png'
#pyplot.savefig(figureName, dpi=300, bbox_inches="tight")
pyplot.show()
class ANN_Arch_View():
# para: neural_network is an array of the number of neurons
# from input layer to output layer, e.g., a neural network of 5 nerons in the input layer,
# 10 neurons in the hidden layer 1 and 1 neuron in the output layer is [5, 10, 1]
# para: weights_list (optional) is the output weights list of a neural network which can be obtained via classifier.coefs_
def __init__( self, neural_network, weights_list=None ):
self.neural_network = neural_network
self.weights_list = weights_list
# if weights_list is none, then create a uniform list to fill the weights_list
if weights_list is None:
weights_list=[]
for first, second in zip(neural_network, neural_network[1:]):
tempArr = np.ones((first, second))*0.4
weights_list.append(tempArr)
self.weights_list = weights_list
def draw( self, show_weights=True):
widest_layer = max( self.neural_network )
network = NeuralNetwork( widest_layer )
for l in self.neural_network:
if l>0:
network.add_layer(l)
network.draw(self.weights_list,show_weights=show_weights)
# + [markdown] id="FnjeuzFszg8g" colab_type="text"
# # Activity 1. Biological Nervous Systems
# + [markdown] id="4tkqF7CXwb_R" colab_type="text"
# ## Activity 1 Scientific Basis of ANNs:
#
# In (most) natural systems, information processing is done by the nervous system, which consists of the brain, spinal cord, and peripheral nerves. Each of these parts contains many nerve cells, also called neurons. The table above shows the typical number of neurons in various animal species. Describe the relationship between number of neurons and the animal’s:
# + [markdown] id="KOmRci6FzvaC" colab_type="text"
# # Fig 1
#
# Consider the following diagram in Fig. 1, and answer questions 1-2:
#
# 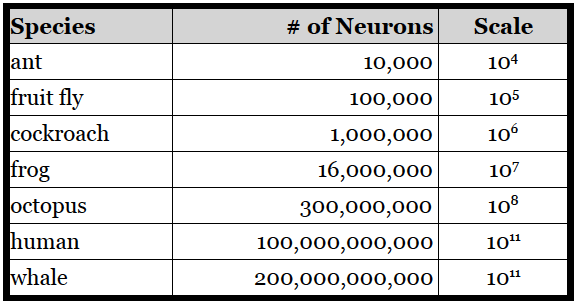
#
# + id="auqHE4eOG2Mt" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 1 Questions 1 - 2
#@markdown 1. Describe the relationship between number of neurons and the animal’s size
activity1_answer1 = "" #@param {type:"string"}
#@markdown 2. Describe the relationship between number of neurons and the animal’s intelligence
activity1_answer2 = "" #@param {type:"string"}
# + [markdown] id="89uZGcc80EgY" colab_type="text"
# # Fig 2
# Neurons can be grouped into 3 broad categories:
#
# ● <b>afferent neurons</b> send signals toward the brain
#
# ● <b>efferent neurons</b> send signals away from the brain
#
# ● <b>interneurons connect</b> other neurons.
#
# (Remember that <b>afferents approach</b> the brain,
# and <b>efferents exit</b> the brain.)
#
# Answer questions 3-11 based on these terms.
#
# 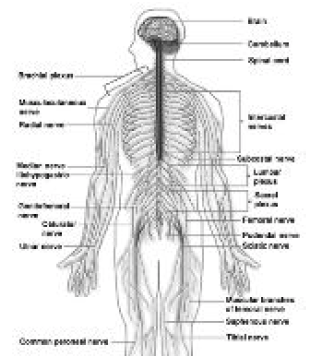
#
# + id="KJ9CvEiBKqSZ" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 1 Questions 3 - 11
#@markdown ###Label each of the following as A (afferent) or E (efferent) :
#@markdown 3. Photoreceptor (light-sensitive) cells in the retina of the eye
activity1_answer3 = "" #@param {type:"string"}
#@markdown 4. Hair cells that react to sound vibrations in the cochlea of the ear
activity1_answer4 = "" #@param {type:"string"}
#@markdown 5. Cells in muscles that cause the muscles to move
activity1_answer5 = "" #@param {type:"string"}
#@markdown 6. Cells in muscles that sense the relative position of body parts (used for proprioception)
activity1_answer6 = "" #@param {type:"string"}
#@markdown 7. Cells that cause the mouth to produce saliva
activity1_answer7 = "" #@param {type:"string"}
#@markdown ###Sensory neurons sense information, and motor neurons control muscles and glands.
#@markdown 8. Which examples above (3-7) are sensory neurons?
activity1_answer8 = "" #@param {type:"string"}
#@markdown 9. Which examples above (3-7) are motor neurons?
activity1_answer9 = "" #@param {type:"string"}
#@markdown 10. Are sensory neurons afferent or efferent?
activity1_answer10 = "" #@param {type:"string"}
#@markdown 11. Are motor neurons afferent or efferent?
activity1_answer11 = "" #@param {type:"string"}
# + [markdown] id="QTChWW2DiPQy" colab_type="text"
# # Fig 3
# Different types of neurons have different shapes and structures, but most contain a similar set of components, shown below.
#
# When a neuron <b>fires</b>, a electrical signal travels from the soma, through the axon, and to terminals that connect to the dendrites of other neurons, so that some of these other neurons may also fire as a result of this signal. This signal is called an <b>action potential</b> and is how the system <b>reacts</b>.
#
# The dendritic connections between neurons change over time (more slowly). These changing connections are how the system <b>learns</b>.
#
# To create artificial neural systems (either as simulations or to solve other problems), we need to consider both processes - <b>reacting</b> and <b>learning</b>.
#
# Consider the following diagram in Fig. 3, and answer questions 12-15:
#
# 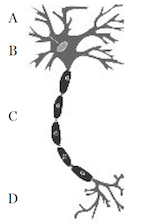
#
# + id="k9gX3vs4jEno" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 1 Questions 12 - 15
#@markdown ###Match the labels (A-D) to the descriptions below:
#@markdown 12. The soma is the main cell body with the nucleus .
activity1_answer3 = "" #@param {type:"string"}
#@markdown 13. The soma has branching dendrites that receive signals from other cells.
activity1_answer4 = "" #@param {type:"string"}
#@markdown 14. The axon is the long arm that sends signals from the soma.
activity1_answer5 = "" #@param {type:"string"}
#@markdown 15. Terminals connect the axon to the dendrites of other neurons.
activity1_answer6 = "" #@param {type:"string"}
# + [markdown] id="_ZdciI4UPSJY" colab_type="text"
# # Activity 2. Perceptrons
#
#
#
# + [markdown] id="oOpFg2hLQQQm" colab_type="text"
# ## A) Bridging the gap between biological and artificial NNs: Perceptron
#
# In a typical Artificial Neural Network (ANN), each neuron has a set of <b>inputs</b> and one <b>output</b>. Each input value is multipled by a <b>weight</b> (positive or negative) to determine its effect.
#
# The weighted inputs are added together, and then an <b>activation function</b> (also called a <b>transfer function</b>) is applied to the sum to determine the output value.
#
# 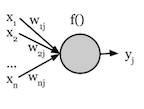
#
# Thus the neuron’s operation can be written as:
# $$y_j=f(\sum_{i=1}^n x_iw_{ij})$$
#
# This model of a neuron is called a <b>perceptron</b>.
# + colab_type="code" cellView="form" id="ROeK6A14-ndA" colab={}
#@title #(RUN upon completion) Activity 2 (A) Questions 1 - 7
#@markdown ###Refer to the diagram and equation above and identify:
#@markdown 1. the letter used for the <b>input(s)</b> ?
activity2_answer1 = "" #@param {type:"string"}
#@markdown 2. the letter used for the <b>output(s)</b> ?
activity2_answer2 = "" #@param {type:"string"}
#@markdown 3. the letter used for the <b>transfer function</b> ?
activity2_answer3 = "" #@param {type:"string"}
#@markdown 4. the letter & subscripts used for the <b>weight</b> between input #2 and output #1.
activity2_answer4 = "" #@param {type:"string"}
#@markdown 5. the letter that best fits a neuron’s <b>dendrites</b> ?
activity2_answer5 = "" #@param {type:"string"}
#@markdown 6. the letter that best fits a neuron’s <b>terminals</b> ?
activity2_answer6 = "" #@param {type:"string"}
#@markdown 7. the letter that best fits a neuron’s <b>soma</b> ?
activity2_answer7 = "" #@param {type:"string"}
# + [markdown] id="N4R7cnJR-qlV" colab_type="text"
# ## B) Modifying code to visualize different ANN architectures:
# The code cell below when executed will show you the architecture of an ANN with some small differences with the picture shown above. The inputs also have their own nodes (circles). Note that they <b> do not </b> apply an activation function. All other nodes apply activation function on their weighted inputs. Also, note that the inputs are just labeled with numbers and not with 'x' and a subscript. The output y is also understood and not shown. Only weights are shown.
#
# <b>This code cell below needs to be "shown" by choosing Edit -> Show/hide code. You are asked to change some values in it and re-execute it </b> in questions 8 - 12 below. After this work you can hide it again by choosing Edit -> Show/hide code.
#
# In Python comments begin with # and those lines with comments that have the words <b>TRY CHANGING</b> are usually the ones you will modify in the exercises.
# + id="L43xlDq-6_qw" colab_type="code" cellView="form" colab={}
#@title # (MODIFY AND RUN CELL) Coding for Activity 2 (B) to answer Questions 8 - 10
network=ANN_Arch_View([2,1]) # TRY CHANGING, the first number in this list is number of input nodes, next number is number of output nodes
network.draw(show_weights=True)
# + id="Zg7fZ04QRSM9" colab_type="code" cellView="form" colab={}
#@title # Activity 2 Questions 8 - 10
#@markdown 8. Change the code cell above and run it to visualize an ANN with 5 input nodes and 1 output node. What is the name of the weight connecting input node 4 to the output node?
activity2_answer2 = "" #@param {type:"string"}
#@markdown 9. Change the code cell above and run it to visualize an ANN with 5 input nodes and 3 output nodes. What is the name of the weight connecting input node 2 to the output node 3?
activity2_answer3 = "" #@param {type:"string"}
#@markdown 10. How many weight values will an ANN of 7 input nodes and 5 output nodes have? You may change the code and run it to visualize this ANN.
activity2_answer3 = "" #@param {type:"string"}
# + colab_type="code" id="vYFxXxF7NVLI" cellView="form" colab={}
#@title # (MODIFY AND RUN CELL) Coding for Activity 2 (B) to answer Questions 11 - 13
network=ANN_Arch_View([4, 2, 2, 1]) # TRY CHANGING, list of number of nodes in all layers starting from input through hidden layers and ending with number of nodes in output layer
network.draw(show_weights=True)
# + id="atUXsiSlSfzY" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 2 (B) Questions 11 - 13
#@markdown 11. Why are some layers called hidden layers as seen in the ANN architecture above? Hint: think of an approximate analogy to biological nervous systems.
activity2_answer11 = "" #@param {type:"string"}
#@markdown 12. An ANN of architecture <b>[4, 2, 2, 1]</b> has two weights named <b>w22</b> - are they the same?
activity2_answer12 = "" #@param {type:"string"}
#@markdown 13. How many weights would an ANN of architecture <b>[5, 4, 3, 2, 1]</b> have? You may change and re-run the code cell above to visualize it. <b>Extra fun</b>: to unclutter the picture you can hide the weight names by setting the show_weights input to False, with show_weights=False (just change True to False)
activity2_answer13 = "" #@param {type:"string"}
# + [markdown] id="vFeAVzqhfqiS" colab_type="text"
# # Activity 3. Perceptrons and Activation Functions
#
#
# + id="KJLvnTDRcVew" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Class and Object for a [3,1] ANN creation in PyTorch
class MyANNModel(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(3, 1,bias=False)
def forward(self, x):
x = x.view(x.shape[0], -1)
x = self.fc1(x)
x = my_activation(x)
return x
ANN_model = MyANNModel()
# + colab_type="code" cellView="form" id="6PkA-jNgoEKc" colab={}
#@title #(RUN CELL) Helper Functions to create widgets for controlling the [3,1] PyTorch ANN
def plot_output_node_activation(model, num_input_nodes, out_node_num, **k):
'''Plots the activation output of one node given a model which has no hidden layers'''
plt.figure(figsize=(7,5))
plt.title('Output of activation function from node ' + str(out_node_num))
plt.ylim(-1.2, 1.2)
plt.xlim(-2, 2)
wsums = np.arange(-2,2,0.1)
pl=plt.plot(wsums,my_activation(torch.tensor(wsums)))
x_vals = []
w_vals = []
for i in range(num_input_nodes):
x_vals.append(k['x'+str(i+1)])
w_vals.append(k['w'+str(i+1)+str(out_node_num)])
model.fc1.__dict__['_parameters']['weight'][out_node_num-1][i] = k['w'+str(i+1)+str(out_node_num)]
xv = np.dot(x_vals,w_vals)
yv = model.forward(torch.tensor([x_vals])).item()
plt.scatter([xv], [yv], color='red', s=30)
plt.xlabel('weighted sum of inputs')
plt.ylabel('activation function output')
plt.grid(True)
plt.tight_layout()
def make_output_node_widgets(model, num_input_nodes, out_node_num):
wght_wdgts = [] # weight widgets
inpt_wdgts = [] # input widgets
for i in range(1,(num_input_nodes+1)):
inpt_wdgts.append(widgets.FloatSlider(min=-1.0,max=1.0,description='x'+str(i)))
wght_wdgts.append(widgets.FloatSlider(min=0.0,max=1.0,description='w'+str(i)+str(out_node_num)))
ui = widgets.HBox([widgets.VBox(inpt_wdgts), widgets.VBox(wght_wdgts)])
all_wgts ={}
all_wgts['out_node_num'] = widgets.fixed(out_node_num)
all_wgts['num_input_nodes'] = widgets.fixed(num_input_nodes)
all_wgts['model'] = widgets.fixed(model)
for xws in inpt_wdgts:
all_wgts[xws.description] = xws
for wws in wght_wdgts:
all_wgts[wws.description] = wws
inter = widgets.interactive_output(plot_output_node_activation,all_wgts)
display(ui,inter)
# + [markdown] id="OXjRo2VrYVnU" colab_type="text"
# ## A) Understanding Activation Functions:
#
# An <b>activation</b> (or <b>transfer</b> ) function converts the weighted sum of input values into an output value for a perceptron (you may want to look back at Activity 2 (A) for the perceptron model).
#
# There are several good choices of activation functions for ANNs which are biology inspired and work well with ANNs. Some of them are shown below in the diagram.
# 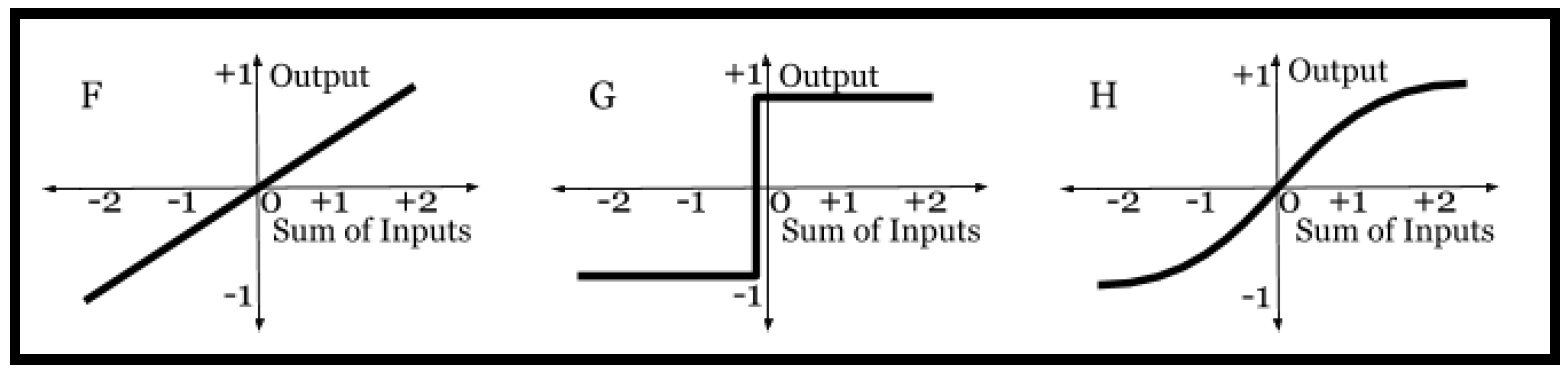
# The MODIFY AND RUN CELL below has code for an activation function of a <b>[3,1]</b> ANN i.e. one with 3 inputs and 1 output. It shows the graphical output of the function as computed by the ANN built in PyTorch in one of the earlier code cells. Through widgets you will be able to directly modify the weights of this ANN (in practice the weights are "learned" which is something we will see later).
#
# For answer questions 1 - 15 you will need to modify and run the code cell below multiple times. Be <b>attentive</b> to the hints in comments.
#
# + id="kjL5MGmwpMhk" colab_type="code" cellView="form" colab={}
#@title #(MODIFY AND RUN CELL) Coding for Activity 3 (A) to answer Questions 1 - 17
#@markdown <b>INSTRUCTION</b>: have ONLY one line uncommented at a time, i.e. no hashtag (#) in front of only one out_value computing line
def my_activation(in_value):
out_value = in_value/2.0 # Function F
#out_value = torch.sign(in_value) # Function G
#out_value = torch.tanh(in_value) # Function H
#out_value = torch.sign(in_value - 1.5) # Mystery Function X
#out_value = torch.nn.functional.relu(in_value) # Mystery Function Y
return out_value
make_output_node_widgets(ANN_model, ANN_model.fc1.in_features, ANN_model.fc1.out_features)
# + id="eKy7htAIfpPL" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 3 Questions 1 - 17
#@markdown <b>For Function F (make sure the correct statement is uncommented in the code cell above to plot function):</b>
#@markdown 1. If the sum of inputs is 2, what is the output?
activity3_answer1 = "" #@param {type:"string"}
#@markdown 2. If the sum of inputs is 1, what is the output?
activity3_answer2 = "" #@param {type:"string"}
#@markdown 3. What is the minimum output value for any input?
activity3_answer3 = "" #@param {type:"string"}
#@markdown 4. Could a small change in one input cause a large change in the output?
activity3_answer4 = "" #@param {type:"string"}
#@markdown <b>For Function G (make sure the correct statement is uncommented in the code cell above to plot function):</b>
#@markdown 5. If the sum of inputs is 2, what is the output?
activity3_answer5 = "" #@param {type:"string"}
#@markdown 6. If the sum of inputs is 1, what is the output?
activity3_answer6 = "" #@param {type:"string"}
#@markdown 7. What is the minimum output value for any input?
activity3_answer7 = "" #@param {type:"string"}
#@markdown 8. Could a small change in one input cause a large change in the output?
activity3_answer8 = "" #@param {type:"string"}
#@markdown <b>For Function H (make sure the correct statement is uncommented in the code cell above to plot function):</b>
#@markdown 9. If the sum of inputs is 2, what is the output?
activity3_answer9 = "" #@param {type:"string"}
#@markdown 10. If the sum of inputs is 1, what is the output?
activity3_answer10 = "" #@param {type:"string"}
#@markdown 11. What is the minimum output value for any input?
activity3_answer11 = "" #@param {type:"string"}
#@markdown 12. Could a small change in one input cause a large change in the output?
activity3_answer12 = "" #@param {type:"string"}
#@markdown <b>Which of the activation functions (F,G,H) is:</b>
#@markdown 13. A step function?
activity3_answer13 = "" #@param {type:"string"}
#@markdown 14. A linear function?
activity3_answer14 = "" #@param {type:"string"}
#@markdown 15. A sigmoid function?
activity3_answer15 = "" #@param {type:"string"}
#@markdown 16. How would you describe mystery function X?
activity3_answer16 = "" #@param {type:"string"}
#@markdown 17. How would you describe mystery function Y?
activity3_answer17 = "" #@param {type:"string"}
# + [markdown] id="INqtQdftlEPc" colab_type="text"
# # Activity 4. Perceptrons for Logic
# + id="f7dLpj0j5YrC" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Class and Object for a [2,1] ANN creation in PyTorch suitable for Logic functions
class MyANNModel(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(2, 1,bias=False)
def forward(self, x):
x = x.view(x.shape[0], -1)
x = self.fc1(x)
x = my_activation(x)
return x
ANN_model = MyANNModel()
# + id="kfeb4Jbb9P_e" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Helper Functions to create widgets for controlling the [2,1] PyTorch ANN
def make_output_node_widgets(model, num_input_nodes, out_node_num):
wght_wdgts = [] # weight widgets
inpt_wdgts = [] # input widgets
for i in range(1,(num_input_nodes+1)):
inpt_wdgts.append(widgets.FloatSlider(min=0,max=1,step=1,value=0,description='x'+str(i)))
wght_wdgts.append(widgets.FloatSlider(min=0,max=2,step=1,value=-1,description='w'+str(i)+str(out_node_num)))
ui = widgets.HBox([widgets.VBox(inpt_wdgts), widgets.VBox(wght_wdgts)])
all_wgts ={}
all_wgts['out_node_num'] = widgets.fixed(out_node_num)
all_wgts['num_input_nodes'] = widgets.fixed(num_input_nodes)
all_wgts['model'] = widgets.fixed(model)
for xws in inpt_wdgts:
all_wgts[xws.description] = xws
for wws in wght_wdgts:
all_wgts[wws.description] = wws
inter = widgets.interactive_output(plot_output_node_activation,all_wgts)
display(ui,inter)
# + [markdown] id="D_b0jAzTYujS" colab_type="text"
# ## A) Thinking Logically with ANNs:
#
# Logical thinking is ingrained in human language and patterns of communication. Can we create a simple [2,1] ANN to create a "logically thinking" ANN to reproduce the input/output functionality of the AND and OR functions shown below?
#
# Note that the AND and OR logic need to be implemented by two separate set of choices for weights. The inputs x1 and x2 are limited to 0 (for F) and 1 (for T). The weights can be 0, 1 or 2. <b>These constraints are set in how the sliders operate to select those values.</b>
# 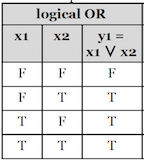
# 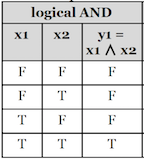
#
# + id="_Ehqq2mw622K" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Coding for Activity 4 (A) to answer Questions 1 - 2
def my_activation(in_value):
out_value = torch.sign(torch.relu(in_value - 1))
return out_value
make_output_node_widgets(ANN_model, ANN_model.fc1.in_features, ANN_model.fc1.out_features)
# + id="9xge1bSBmlvw" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 4 (A) Questions 1 - 2
#@markdown 1. What set of weights did you choose to implement the AND logic?
activity5_answer1 = "" #@param {type:"string"}
#@markdown 2. What set of weights did you choose to implement the OR logic?
activity5_answer2 = "" #@param {type:"string"}
# + [markdown] id="_3f-lt4Kvzqf" colab_type="text"
# # Activity 5. Perceptrons for Images
# + id="eSOi6hrBIK4q" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Class and Object for a [9,4] ANN creation in PyTorch for 3 X 3 image input and 4 classes
class MyANNModel(nn.Module):
def __init__(self):
super().__init__()
self.fc1 = nn.Linear(9, 4,bias=False)
def forward(self, x):
x = x.view(x.shape[0], -1)
x = self.fc1(x)
x = my_activation(x)
return x
ANN_model = MyANNModel()
# + id="HXW6yFw5LMCN" colab_type="code" cellView="form" colab={}
#@title #(RUN CELL) Helper Functions to create widgets for controlling the [9,4] PyTorch ANN
def plot_output_node_activation(model, num_input_nodes, out_node_num, **k):
'''Plots the activation output of one node given a model which has no hidden layers'''
plt.figure(figsize=(7,5))
plt.title('Output of activation function from node ' + str(out_node_num))
plt.ylim(-1.2, 1.2)
plt.xlim(-2, 2)
wsums = np.arange(-2,2,0.1)
pl=plt.plot(wsums,my_activation(torch.tensor(wsums)))
x_vals = []
w_vals = []
for i in range(num_input_nodes):
x_vals.append(k['x'+str(i+1)])
w_vals.append(k['w'+str(i+1)+str(out_node_num)])
model.fc1.__dict__['_parameters']['weight'][out_node_num-1][i] = k['w'+str(i+1)+str(out_node_num)]
xv = np.dot(x_vals,w_vals)
yv = model.forward(torch.tensor([x_vals]))[0][out_node_num].item()
plt.scatter([xv], [yv], color='red', s=30)
plt.xlabel('weighted sum of inputs')
plt.ylabel('activation function output')
plt.grid(True)
plt.tight_layout()
def make_output_node_widgets(model, num_input_nodes, out_node_num):
wght_wdgts = [] # weight widgets
inpt_wdgts = [] # input widgets
for i in range(1,(num_input_nodes+1)):
inpt_wdgts.append(widgets.FloatSlider(min=0,max=1,step=1,value=0,description='x'+str(i)))
wght_wdgts.append(widgets.FloatSlider(min=-1,max=1,value=-1,description='w'+str(i)+str(out_node_num)))
ui = widgets.HBox([widgets.VBox(inpt_wdgts), widgets.VBox(wght_wdgts)])
all_wgts ={}
all_wgts['out_node_num'] = widgets.fixed(out_node_num)
all_wgts['num_input_nodes'] = widgets.fixed(num_input_nodes)
all_wgts['model'] = widgets.fixed(model)
for xws in inpt_wdgts:
all_wgts[xws.description] = xws
for wws in wght_wdgts:
all_wgts[wws.description] = wws
inter = widgets.interactive_output(plot_output_node_activation,all_wgts)
display(ui,inter)
# + id="JerPpXXjI-eR" colab_type="code" cellView="form" colab={}
#@title # (RUN CELL) Visualize the [9,4] ANN
network=ANN_Arch_View([9,4]) # TRY CHANGING, the first number in this list is number of input nodes, next number is number of output nodes
network.draw(show_weights=True)
# + [markdown] id="irvMDlP_wPmx" colab_type="text"
# # A) Simple ANN to classify 3 X 3 optical character
#
# ANNs have many applications, including computer vision and image recognition. You may have used computer software that is able to scan printed pages and convert them to digitized form to make them searchable. In order for the software to do this it has to understand the character represented in digital images which are tables of pixels. These automated tasks have different names like optical character recognition, handwriting recognition. Inside such software you will find ANNs which are more complex than the one you will build here, but those ANNs have similar building blocks. This section explores an example with 3x3 pixel black-and-white images, as show below. These 9 pixels will be the inputs (x1,...,x9) for an ANN, with outputs for each of 4 symbols (X, 十 , T, L).
#
# For each symbol, choose a set of 9 input weights and a step function threshold
# to produce the correct output. All of the weights can be either +1.0 or -1.0 as constrained by the widgets below and pixels are either black (0) or white (1). In other words you have to answer questions like:
#
# What set of 9 weights to choose to make the output node 1 fire for symbol X at the input?
#
# What set of 9 weights to choose to make the output node 2 fire for symbol 十 at the input?
#
# ... and so on ...
#
# Suggestions: start by choosing an output node number from the 1, 2, 3, 4 which corresponding to the 4 symbols (X, 十 , T, L). Then configure the input pixels to be that input symbol and then choose a proper set of 9 weights so that the output node fires.
#
# 
#
# + id="3uNyWtYsLyBz" colab_type="code" cellView="form" colab={}
#@title #(MODIFY AND RUN CELL) Coding for Activity 5 (A) to answer Questions 1 - 5
#@markdown <b>INSTRUCTION:</b> modify value of the output node number as shown in code below to get different output nodes
def my_activation(in_value):
out_value = torch.sign(in_value)
return out_value
output_node_num = 1 # TRY CHANGING, can be 1, 2, 3 or 4 corresponding to the 4 symbols (X, 十 , T, L)
make_output_node_widgets(ANN_model, ANN_model.fc1.in_features, output_node_num)
# + id="lgQVSd_3OnZp" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 5 (A) Questions 1 - 5
#@markdown 1. What set of weights did you choose to make the output node 1 fire for symbol X?
activity5_answer1 = "" #@param {type:"string"}
#@markdown 2. What set of weights did you choose to make the output node 1 fire for symbol +?
activity5_answer2 = "" #@param {type:"string"}
#@markdown 3. What set of weights did you choose to make the output node 1 fire for symbol T?
activity5_answer3 = "" #@param {type:"string"}
#@markdown 4. What set of weights did you choose to make the output node 1 fire for symbol L?
activity5_answer4 = "" #@param {type:"string"}
#@markdown 5. This approach could be used for larger images (more pixels), and with more outputs (for more symbols). Explain why these sorts of ANNs are called classifiers.
activity5_answer5 = "" #@param {type:"string"}
# + [markdown] id="jzR08yJNP5sB" colab_type="text"
# # B) Future Activity: teaching an ANN
#
# In future activities we will explore algorithms to adjust weights so that ANNs can learn.
# Learning (also called training ) usually requires a set of sample inputs.
# When each input should give a known output, it is supervised learning .
# When inputs are provided but the outputs are unknown, it is unsupervised learning.
# + id="AiMPcw2qQXfX" colab_type="code" cellView="form" colab={}
#@title #(RUN upon completion) Activity 5 (A) Questions 6 - 9
#@markdown Categorize each example below as supervised or unsupervised learning.
#@markdown 6. Given images of faces, identify the most common faces.
activity5_answer6 = "" #@param {type:"string"}
#@markdown 7. Given images of specific people, learn to recognize each person.
activity5_answer7 = "" #@param {type:"string"}
#@markdown 8. Given sets of test results for many patients, identify patients with similar symptoms.
activity5_answer8 = "" #@param {type:"string"}
#@markdown 9. Given sets of test results for many patients, provide the right diagnosis for each patient.
activity5_answer9 = "" #@param {type:"string"}
# + id="CWtttduI0mfY" colab_type="code" cellView="form" colab={}
#@title Team Work Evaluation for the ANN notebook based activities.
#@markdown 1. How much time was required for completion of the ANN notebook?
activity1_evaluation1 = "" #@param {type:"string"}
#@markdown 2. Was the contribution from each participant equal?
activity1_evaluation2 = "" #@param {type:"string"}
#@markdown 3. How could the team work and learn more effectively?
activity1_evaluation3 = "" #@param {type:"string"}
#@markdown 4. How many participants thought the problems were too simple (trivial)?
activity1_evaluation4 = "" #@param {type:"string"}
#@markdown 5. How many participants thought the problems were at the proper level of difficulty?
activity1_evaluation5 = "" #@param {type:"string"}
#@markdown 6. How many participants thought the problems were too hard?
activity1_evaluation6 = "" #@param {type:"string"}
#@markdown 7. Was help needed? Where?
activity1_evaluation7 = "" #@param {type:"string"}
#@markdown 8. Does the team have any suggestions about how the ANN notebook could be improved? If so, how?
activity1_evaluation8 = "" #@param {type:"string"}
# + [markdown] id="MapMslmOgdsb" colab_type="text"
# # A Project With Real Images
# # Code cells are ready to run (note that execution may be long ...)
# # Discuss with instructor for ideas on what can be done
#
# This is a project to train an ANN to classify fashion images. See below for samples.
# 
# + id="zdVeQmp_gEv2" colab_type="code" cellView="form" colab={}
#@title # (RUN CELL) LIBRARY INSTALL COMMAND
# !pip3 install --upgrade torchfusion
# + id="nP7FgOpNHjtf" colab_type="code" cellView="form" colab={}
#@title # (RUN CELL) TRAINING CODE
from torchfusion.layers import *
from torchfusion.datasets import *
from torchfusion.metrics import *
import torch.nn as nn
import torch.cuda as cuda
from torch.optim import Adam
from torchfusion.learners import StandardLearner
train_loader = fashionmnist_loader(size=28,batch_size=32)
test_loader = fashionmnist_loader(size=28,train=False,batch_size=32)
model = nn.Sequential(
Flatten(),
Linear(784,100),
Swish(),
Linear(100,100),
Swish(),
Linear(100,100),
Swish(),
Linear(100,10)
)
if cuda.is_available():
model = model.cuda()
optimizer = Adam(model.parameters())
loss_fn = nn.CrossEntropyLoss()
train_metrics = [Accuracy()]
test_metrics = [Accuracy()]
learner = StandardLearner(model)
if __name__ == "__main__":
print(learner.summary((1,28,28)))
learner.train(train_loader,train_metrics=train_metrics,optimizer=optimizer,loss_fn=loss_fn,test_loader=test_loader,test_metrics=test_metrics,num_epochs=40,batch_log=False)
# + id="Y4xHC801ngVn" colab_type="code" cellView="form" colab={}
#@title # (RUN CELL) TRAINING CODE
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/sbhattac/ai-workshop/master/ANN/"
for filename in ("sample-1.jpg", "sample-2.jpg", "sample-3.jpg", "sample-4.jpg"):
print("Downloading", filename)
url = DOWNLOAD_ROOT + filename
urllib.request.urlretrieve(url, filename)
# + id="chtWcy2rfroB" colab_type="code" cellView="form" colab={}
#@title # (RUN CELL) TESTING CODE
import torch
from torchfusion.layers import *
import torch.nn as nn
import torch.cuda as cuda
from torchfusion.learners import StandardLearner
from torchfusion.utils import load_image
model = nn.Sequential(
Flatten(),
Linear(784,100),
Swish(),
Linear(100,100),
Swish(),
Linear(100,100),
Swish(),
Linear(100,10)
)
if cuda.is_available():
model = model.cuda()
#learner = StandardLearner(model)
#learner.load_model("best_models\model_20.pth")
if __name__ == "__main__":
#map class indexes to class names
class_map = {0:"T-Shirt",1:"Trouser",2:"Pullover",3:"Dress",4:"Coat",5:"Sandal",6:"Shirt",7:"Sneaker",8:"Bag",9:"Ankle Boot"}
#Load the image
image = load_image("sample-1.jpg",grayscale=True,target_size=28,mean=0.5,std=0.5)
#add batch dimension
image = image.unsqueeze(0)
#run prediction
pred = learner.predict(image)
#convert prediction to probabilities
pred = torch.softmax(pred,0)
#get the predicted class
pred_class = pred.argmax().item()
#get confidence for the prediction
pred_conf = pred.max().item()
#Map class_index to name
class_name = class_map[pred_class]
print("Predicted Class: {}, Confidence: {}".format(class_name,pred_conf))
|
ANN/AI_Workshop_ANN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.11
# language: julia
# name: julia-0.3
# ---
addprocs(7);
@everywhere begin
using NetworkDiscovery
using POMDPs
import POMDPs: solve
using POMCP
using POMDPToolbox
end
nodes = 100
comms = 5
probes = 10
p_inter = 0.01
p_intra = 0.3
N = 10000
@everywhere solve(solver::DiscoveryHeuristic, pomdp) = solver
@everywhere function tick_hack(x::Float64,y::Float64)
print(".")
return x+y
end
function est_rew(solver, nodes, comms, probes, p_intra, p_inter, N)
sum = @parallel (tick_hack) for i in 1:N
prob_rng = MersenneTwister(i)
sim_rng = MersenneTwister(i)
nw = generate_network(prob_rng, nodes, comms, p_intra, p_inter)
pomdp = generate_problem(prob_rng, nw, probes, 1, 100.0, 10, 10, p_intra, p_inter)
policy = solve(solver, pomdp)
revealed = initial_belief(pomdp)
sim = RolloutSimulator(rng=sim_rng, initial_state=nw, initial_belief=revealed)
simulate(sim, pomdp, policy)
end
# print("\r")
end
guess_rng = MersenneTwister(1)
policy = DiscoveryHeuristic(ProbeHighestDegree(false), GuessBasedOnNeighbors(guess_rng))
@time est_rew(policy, nodes, comms, probes, p_intra, p_inter, N)/N
rollout_rng = MersenneTwister(1)
pomcp_rng = MersenneTwister(1)
rollout_policy = DiscoveryHeuristic(ProbeHighestDegree(false), GuessBasedOnNeighbors(rollout_rng))
solver = POMCPSolver(rollout_policy, 0.0, 100.0, 1000, pomcp_rng, false, FullBeliefConverter(), 0)
@time est_rew(solver, nodes, comms, probes, p_intra, p_inter, N)
|
notebooks/Compare Heuristic vs POMCP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Training Sample: train.csv
# #### Evaluation Sample: validation_under.csv
# #### Method: OOB
# #### Output: Best hyperparameters; Pr-curve; ROC AUC
# # Training Part
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer,classification_report, matthews_corrcoef, accuracy_score, average_precision_score, roc_auc_score
# #### Input data is read and named as the following
transactions = pd.read_csv('../Data/train.csv')
X_train = transactions.drop(labels='Class', axis=1)
y_train = transactions.loc[:,'Class']
# #### Tuning parameters
# +
test = 0
rf = RandomForestClassifier(n_jobs=-1, random_state=1)
if test== 0:
n_estimators = [75,150,800,1000,1200]
min_samples_split = [2, 5]
min_samples_leaf = [1, 5]
else:
n_estimators = [70]
min_samples_split = [2]
min_samples_leaf = [1]
param_grid_rf = {'n_estimators': n_estimators,
'min_samples_split': min_samples_split,
'min_samples_leaf': min_samples_split,
'oob_score': [True]
}
# -
grid_rf = GridSearchCV(estimator=rf, param_grid=param_grid_rf,cv = 2,
n_jobs=-1, pre_dispatch='2*n_jobs', verbose=1, return_train_score=False)
grid_rf.fit(X_train, y_train)
# #### The best score and the estimator
grid_rf.best_score_
grid_rf.best_params_
# # Evaluation Part
evaluation = pd.read_csv('../Data/validation_under.csv')
X_eval = evaluation.drop(labels='Class', axis=1)
y_eval = evaluation.loc[:,'Class']
def Random_Forest_eval(estimator, X_test, y_test):
y_pred = estimator.predict(X_test)
print('Classification Report')
print(classification_report(y_test, y_pred))
if y_test.nunique() <= 2:
try:
y_score = estimator.predict_proba(X_test)[:,1]
except:
y_score = estimator.decision_function(X_test)
print('AUPRC', average_precision_score(y_test, y_score))
print('AUROC', roc_auc_score(y_test, y_score))
Random_Forest_eval(grid_rf, X_eval, y_eval)
# ### Precision Recall Curve
from sklearn.metrics import precision_recall_curve
precision_rf,recall_rf,thresholds_rf = precision_recall_curve(
y_eval, grid_rf.predict_proba(X_eval)[:,1])
import matplotlib.pyplot as plt
close_default_rf = np.argmin(np.abs(thresholds_rf * 0.5))
plt.plot(precision_rf,recall_rf,label="rf")
plt.plot(precision_rf[close_default_rf],recall_rf[close_default_rf],'^',c='k',
label = "threashold 0.5 rf")
plt.xlabel("precision")
plt.ylabel("recall")
plt.legend(loc="best")
plt.title("PR_Curve")
# ### Area Under the Receiver Operating Characteristic Curve
from sklearn.metrics import roc_curve
fpr_rf,tpr_rf,th_rf = roc_curve(
y_eval, grid_rf.predict_proba(X_eval)[:,1])
close_default_rf = np.argmin(np.abs(th_rf * 0.5))
plt.plot(fpr_rf,tpr_rf,label="rf")
plt.plot(fpr_rf[close_default_rf],tpr_rf[close_default_rf],'^',c='k',
label = "threashold 0.5 rf")
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.legend(loc="best")
plt.title("ROC")
|
Random_Forest/.ipynb_checkpoints/RF_new_data-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 64-bit
# name: python374jvsc74a57bd07945e9a82d7512fbf96246d9bbc29cd2f106c1a4a9cf54c9563dadf10f2237d4
# ---
# # Data Science Bootcamp - The Bridge
# ## Precurso
# En este notebook vamos a ver, uno a uno, los conceptos básicos de Python. Constarán de ejercicios prácticos acompañados de una explicación teórica dada por el profesor.
#
# Los siguientes enlaces están recomendados para el alumno para profundizar y reforzar conceptos a partir de ejercicios y ejemplos:
#
# - https://www.kaggle.com/learn/python
#
# - https://facundoq.github.io/courses/aa2018/res/02_python.html
#
# - https://www.w3resource.com/python-exercises/
#
# - https://www.practicepython.org/
#
# - https://es.slideshare.net/egutierrezru/python-paraprincipiantes
#
# - https://www.sololearn.com/Play/Python#
#
# - https://github.com/mhkmcp/Python-Bootcamp-from-Basic-to-Advanced
#
# Ejercicios avanzados:
#
# - https://github.com/darkprinx/100-plus-Python-programming-exercises-extended/tree/master/Status (++)
#
# - https://github.com/mahtab04/Python-Programming-Practice (++)
#
# - https://github.com/whojayantkumar/Python_Programs (+++)
#
# - https://www.w3resource.com/python-exercises/ (++++)
#
# - https://github.com/fupus/notebooks-ejercicios (+++++)
#
# Tutor de ayuda PythonTutor:
#
# - http://pythontutor.com/
#
# ## 1. Variables y tipos
#
# ### Cadenas
# +
# Entero - Integer - int
x = 7
# Cadena - String - Lista de caracteres - str
x = "lorena"
print(x)
x = 7
print(x)
# -
# built-in
type(x)
# built-in
type
# +
x = 5
y = 7
z = x + y
print(z)
# +
x = "'lorena' \""
l = 'silvia----'
g = x + l
# Las cadenas se concatenan
print(g)
# -
# type muestra el tipo de la variable
type(g)
type(5)
print(x)
# +
# print es una función que recibe varios argumentos y cada argumento está diferenciado por la coma. Después de cada coma, la función 'print' añade un espacio.
# mala praxis
print( g,z , 6, "cadena")
# buena praxis - PEP8
print(g, z, 6, "cadena")
# +
x = 5
s = "6"
print(x + s)
# +
# Cambiar de int a str
#built-in
x = 6
print(type(x))
x = str(6)
print(type(x))
# -
x = 5
z = "558" + str(x)
print(z)
# +
# Cambiar de str a int
s = "56"
print(s)
print(type(s))
s = int(s)
print(s)
print(type(s))
z = 7 + s
print(z)
# -
s = "56a"
s = int(s)
print(s)
# +
# Para ver la longitud de una lista de caracteres (lista)
p = "lista caracteres"
print(len(p))
# +
# String, Integer, Float, List, None (NaN), bool
# str, int, float, list, bool
# -
x = 5.875
print(type(x))
o = x + 2
print(o)
x = int(x)
print(x)
print(type(x))
x = float(x)
print(x)
print(type(x))
l = (2 + 5) * 3
print(l)
l = ((2 + 5) * 3)
print(l)
m = ((((65 + int("22")) * 2)))
m
s = "89"
print(type(int(s)))
# +
# los decimales son float. Python permite las operaciones entre int y float
x = 4
y = 4.2
print(x + y)
# -
t = 5
g = 2
print(t / g)
t = 4
g = 2
print(t / g)
print(type(t / g))
d = t / g
print(d)
print(type(d))
d = int(d * 6.1 + 1.1 / 6.2)
print(d)
# La división normal (/) es siempre float
# La división absoluta (//) puede ser:
# - float si uno de los dos números (o los dos) son float
# - int si los dos son int
j = 15
k = 4
division = j // k
print(division)
division = j / k
print(division)
print(type(division))
# +
num1 = 12
num2 = 3
suma = num1 + num2
resta = num1 - num2
multiplicacion = num1 * num2
division = num1 / num2
division_absoluta = num1 // num2
print("suma:", suma)
print("resta:", resta)
print("multiplicación:", multiplicacion)
print("division:", division)
print("division_absoluta:", division_absoluta)
print(type(division))
print(type(division_absoluta))
# +
# Jupyter notebook permite que la última línea se imprima por pantalla ( la variable )
x = 2
k = 4
k
x
k
# -
# Soy un comentario
# print("Hello Python world!")
# Estoy creando una variable que vale 2
"""
Esto es otro comentario
"""
# ## Ejercicio:
# ### Crear una nueva celda.
# ### Declarar tres variables:
# - Una con el nombre "edad" con valor vuestra edad
# - Otra "edad_compañero_der" que contengan la edad de tipo entero de vuestro compañero de la derecha
# - Otra "suma_anterior" que contenga la suma de las dos variables anteriormente declaradas
#
# Mostrar por pantalla la variable "suma_anterior"
#
#
edad = 18
edad_companero_der = 27
suma_anterior = edad + edad_companero_der
print(suma_anterior)
# +
# funciones built-in de string
string1 = "Hola"
print(string1.upper())
# -
# ## 2. Números y operadores
# +
x = 2
print(x**)
# -
x = 4
x = x + 1
print(x)
x = 4
x += 1
# +
### Enteros ###
x = 3
print("- Tipo de x:")
print(type(x)) # Imprime el tipo (o `clase`) de x
print("- Valor de x:")
print(x) # Imprimir un valor
print("- x+1:")
print(x + 1) # Suma: imprime "4"
print("- x-1:")
print(x - 1) # Resta; imprime "2"
print("- x*2:")
print(x * 2) # Multiplicación; imprime "6"
print("- x^2:")
print(x ** 2) # Exponenciación; imprime "9"
# Modificación de x
x += 1
print("- x modificado:")
print(x) # Imprime "4"
x *= 2
print("- x modificado:")
print(x) # Imprime "8"
#print("- El módulo de x con 40")
#print(40 % x)
print("- Varias cosas en una línea:")
print(1, 2, x, 5*2) # imprime varias cosas a la vez
# -
# El módulo muestra el resto de la división entre dos números
x = 4 % 2
print(x)
# +
numero = 99
numero % 2 # Si el resto es 0, el número es par. Sino, impar.
# +
numero = 99
numero % 3 # El 3 es divisor de 'numero' si el resultado del módulo da 0
# -
4%5
# Actualizar variable en cada ejecución
x = 5
x += 2
j = 7 + x
print(j)
# ## Título markdown
#
# #### otro título
# # INPUT
x = input()
print(type(x))
print(x)
x = int(input())
print(type(x))
l = 4 + x
print(x)
x = int(input("Escribe un número:"))
print(type(x))
l = 4 + x
print(l)
x = float(input("Escribe un número:"))
print(type(x))
l = 4 + x
print(l)
x = int(float("57.9"))
x
edad = input("Escribe tu edad")
print(edad)
edad = float(edad)
print(edad)
# ## 3. Tipo None
# +
# str int float none
x = None
p = 5 + x
print(p)
# -
x = None
x = str(x)
print(x)
x = None
print(type(x))
# ### Booleans
cierto = True
falso = False
print(type(cierto))
# ## 4. Listas y colecciones
# +
# Una colección:
# lista, conjunto (set), tupla, array, string, diccionarios
# +
# Lista de elementos:
# Las posiciones se empiezan a contar desde 0
s = "Cadena"
# -
print(s[0])
s = ""
print(type(s))
s[0]
print(len(s))
lista = []
print(type(lista))
# +
# La última posición de una colección coincide con el (número de elementos - 1)
# -
lista = [3, 7.2, "hola"]
print(lista[2]) # último
print(lista[1]) # penúltimo
print(lista[0]) # antepenúltimo
print(str(lista[0]) + "\n" + str(lista[1]) + "\n" + lista[2])
print(str(lista[0]), str(lista[1]), lista[2], sep="\n")
# Salto de línea \n
print("Hol\n me llamo G\nbriel")
lista = [3, 7.2, "hola"]
print("Primer elemento:\n", lista[0], "\nSegundo elemento:\n",
lista[1], "\nÚltimo elemento:\n", lista[-1])
# No pasamos el último elemento a String porque ya es String
print("Primer elemento:\n" + str(lista[0]) + "\nSegundo elemento:\n" + str(lista[1]) + "\nÚltimo elemento:\n" + lista[-1])
lista = [3, 7.2, "hola"]
tamano_lista = len(lista)
elemento_que_esta_en_la_mitad = len(lista) // 2
lista[elemento_que_esta_en_la_mitad]
print(lista[-1]) # último
print(lista[-2]) # penúltimo
print(lista[-3]) # antepenúltimo
print(lista[0:2])
print(lista)
lista2 = ["hola", 2, [7.2, "x"]]
print(len(lista2))
print(lista2[-1])
print(lista2[-1][-1])
lista3 = ["hola", 2, [7.2, "xa"]]
print(lista3[-1][-1][0])
lista4 = ["hola", 2, [[7.2, "xa"],]]
print(type(lista4))
print(type(lista4[-1]))
print(type(lista4[-1][1]))
lista4 = ["hola", 2, [[7.2, "xa"]]]
print(lista4)
print(type(lista4))
print(lista4[-1])
print(type(lista4[-1]))
print(lista4[-1][0])
print(type(lista4[-1][0]))
print(lista4[-1][0][-1])
print(type(lista4[-1][0][-1]))
print(lista4[-1][0][-1][-1])
print(type(lista4[-1][0][-1][-1]))
lista5 = [4,]
print(lista5)
# +
s = "hola"
lenght = 4
s[1:4]
# +
s = "<NAME> Clara"
print(s[::-1])
# +
# Para acceder a varios elementos, se especifica con la nomenclatura "[N:M]". N es el primer elemento a obtener, M es el último elemento a obtener pero no incluido. Ejemplo:
# Queremos mostrar desde las posiciones 3 a la 7. Debemos especificar: [3:8]
# Si M no tiene ningún valor, se obtiene desde N hasta el final.
# Si N no tiene ningún valor, es desde el principio de la colección hasta M
s[3:len(s)]
# -
# Coge desde el elemento en la posición 3 (incluido) hasta el final
s[3:]
# Coge todos los elementos hasta la posición 3 (sin incluir)
s[:3]
# Agrega un valor a la última posición de la lista
s = "hola"
s = s.upper()
print(s)
lista = [4, "x"]
print(lista)
lista.append(7)
print(lista)
print(len(lista))
lista = [4, "x"]
print(lista)
lista.append(7)
print(lista)
print(len(lista))
# remove elimina el primer elemento que se encuentra que coincide con el valor del argumento
lista = [4, 'y', 7]
lista.remove(7)
lista.remove(7)
print(lista)
# remove elimina el primer elemento que se encuentra que coincide con el valor del argumento
lista = [4, 'y', 7, 'y']
lista.remove('y')
print(lista)
lista = [4, 'y', 7, 'y']
lista.append('y') # append siempre añade el elemento al final de la lista
print(lista)
lista.remove('y') # elimina la primera coincidencia
print(lista)
lista = [4, 'x', 'y', ['z']]
lista[-1].append(4) # añade un entero 4 a la lista ['z']
print(lista)
lista = [4, 'x', 'y', ['z']]
lista[-1].append([4]) # añade una lista [4] a la lista ['z']
print(lista)
h = 4
l = [4]
print("tipo de h:", type(h))
print("tipo de l:", type(l))
l.append(3)
print(l)
# +
# Accedemos a la posición 1 del elemento que está en la posición 2 de lista
lista = ['l', 9.4, [8.1, 'xs']]
print(lista[2][1])
# -
lista2 = ['l', 9.4, [8.1, 'xs']]
lista2.reverse()
print(lista2)
lista3 = [2, 4, 6, 8, 'o']
print(len(lista3))
lista_1 = [2, 4]
lista_2 = [8, 9]
lista_3 = lista_1 + lista_2 # concatenamos
print(lista_3)
lista3 = [2, 4, 6, 8, 'o']
lista3.append(lista2) # añadir al final de la lista
print(lista3)
lista3[5][0].append(lista2)
print(lista3)
print(lista3[5][0][2])
lista = ["a", "b"]
lista.reverse()
print(lista)
lista = ["a", "b"]
lista = lista.reverse()
print(lista)
lista = ["a", "b"]
lista = lista[::-1]
print(lista)
# +
# Ejemplo práctico
lista_pacientes = [["Gabriel..", "algo"], ["Lorena", "inflamación"]]
lista_pacientes[0]
# -
# ### Colecciones
#
# 1. Listas
# 2. String (colección de caracteres)
# 3. Tuplas
# 4. Conjuntos (Set)
# 5. Diccionarios
s = "j"
s[0][0]
n = 2
n[0]
# Listas --> Mutables
lista = [2, 5, "caract", [9, "g", ["j"]]]
print(lista[-1][-1][-1])
lista[3][2][0]
x = ((2 + 2))
x
x = 2
print(type(x))
lista = [2, 5]
tupla = (2, 4)
print(len(tupla))
tupla[1]
# +
# Tuplas --> Inmutables
tupla = (2, 5, "caract", [9, "g", ["j"]])
print(type(tupla))
tupla
# -
# Update listas
lista = [2, "6", ["k", "m"]]
lista[1] = 1
lista
tupla = (2, "b", ["k", "m"])
tupla[1] = 1
tupla
tupla[2].append("Otro")
tupla
# +
# La tupla es inmutable pero sí se puede modificar las colecciones mutables dentro de ella
tupla[2].remove("k")
tupla[2].remove("m")
tupla[2].remove("Otro")
tupla
# -
tupla[2] = [2]
tupla = (2, 'b', [])
# Cambiar una tupla a lista (mutable) y volver a tupla una vez modificada
print(tupla)
tupla = list(tupla)
print(tupla)
tupla[1] = "5"
print(tupla)
tupla = tuple(tupla)
tupla
# +
# Conjuntos
conjunto = [2, 4, 6, "a", "z", "h", 2]
conjunto = set(conjunto)
conjunto
# -
conjunto = ["a", "z", "h", 2, 2, 4, 6, True, True, False]
conjunto = set(conjunto)
conjunto
conjunto = ["a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False]
conjunto = set(conjunto)
conjunto
conjunto_tupla = ("a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False)
conjunto = set(conjunto_tupla)
conjunto
conjunto = {"a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False}
conjunto
conjunto = {"a", "z", "h", 2, 2, 4, 6, 2.1, 2.4, 2.3, True, True, False, None, (24, 2)}
conjunto
conjunto = list(conjunto)
conjunto
# Si quiero que mi conjunto no tenga datos repetidos:
# 1. Lo paso a conjunto
# 2. Lo vuelvo a pasar al tipo del conjunto original
lista = [2, 2, "perro", "gato"]
lista = set(lista)
lista = list(lista)
lista
# Si quiero que mi conjunto no tenga datos repetidos:
# 1. Lo paso a conjunto
# 2. Lo vuelvo a pasar al tipo del conjunto original
lista = [2, 2, "perro", "gato"]
lista = list(set(lista))
lista
# Creación de tupla vacía
tupla_vacia = ()
type(tupla_vacia)
# Creación de tupla vacía
tupla_vacia = (2,)
type(tupla_vacia)
# Lista vacía
type([])
# Conjunto vacío necesita un elemento. Si no añades un elemento, es un diccionario
conjunto_vacio = {0,}
type(conjunto_vacio)
c = {0,}
type(c)
# String vacío
s = ""
# Lista vacía
[]
# Tupla vacía
()
# Conjunto vacío
{0,}
y = (2)
y
y = {2,}
print(type(y))
y
s = "String"
lista_s = list(s)
lista_s
s = "String"
conj = set(s)
conj
# Cambiar de tupla a lista, remover elemento y convertir en tupla
tupla = (2, 6, ["x"], True)
tupla = list(tupla)
tupla.remove(["x"])
tupla = tuple(tupla)
tupla
# Eliminar por posición en una lista
lista = [2, 6, ["x"], True]
lista.pop() # pop elimina el último elemento por defecto
lista
lista = [2, 6, ["x"], True]
lista.pop(2)
lista
# pop retorna el valor eliminado
lista = [2, 6, ["x"], True]
valor_eliminado = lista.pop(1)
print(lista)
valor_eliminado
conjunto = {2, 5, "h"}
lista_con_conjunto = [conjunto]
lista_con_conjunto[0]
# No podemos acceder a elementos de un conjunto
lista_con_conjunto[0][0]
# ## 5. Condiciones: if, elif, else
x = 5
if x > 3:
print("Es mayor a 3")
x = 5
if x > 3: # siempre que hay dos puntos, hay indentación
print("Es mayor a 3")
x = 5
if x > 3: # siempre que hay dos puntos, hay indentación
print("Es mayor a 3")
x = 2
print(x)
# +
# dos bloques de condiciones diferentes
x = 5
if x > 3: # siempre que hay dos puntos, hay indentación
print("Es mayor a 3")
if x < 4:
x = 2
print(x)
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x >:
print("2")
else:
print("3")
print("Fuera del bloque")
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 30:
print("2")
elif x == 5:
print("x es 5")
else:
print("3")
print("Fuera del bloque")
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 30:
print("2")
elif x == 6:
print("x es 5")
else:
print("3")
print("Fuera del bloque")
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 30:
print("2")
elif x == 6:
print("x es 5")
print("Fuera del bloque")
# -
#
# - if inicia bloque de condiciones
# - pueden existir todos los elif que se necesiten
# - solo puede existir un else
# - no son necesarios ni los elif ni el else
#
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 30:
print("2")
elif x == 6:
print("x es 5")
else:
print("primer else")
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 3:
print("2")
elif x == 5:
print("x es 5")
else:
print("primer else")
print("Fuera del bloque")
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 3:
print("2")
x = 6
elif x == 6:
print("x es 5")
else:
print("primer else")
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 9:
print("2")
x = 6
elif x == 6:
print("x es 6")
else:
print("primer else")
print("Fuera del bloque")
# +
x = 5
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x > 3:
print("2")
x = 10
if x < 4: # inicio de un bloque de condiciones
print("1")
elif x == 6:
print("x es 5")
else:
print("else interno")
elif x == 5:
print("x es 5")
else:
print("primer else")
print("Fuera del bloque")
# +
"""
== -> Igualdad
!= -> Diferecia
< -> Menor que
> -> Mayor que
<= -> Menor o igual
>= -> Mayor o igual
"""
print(2 == 2)
# -
if True:
print("hola")
else:
print("adios")
# +
if (2 == 2 and 2 == 3): # and necesita que ambas condiciones sean True
print("hola")
else:
print("adios")
if True and False:
print("hola")
else:
print("adios")
# +
# and | or
if 1 == 1 or 2 == 3: # or necesita que solo una condición sea True
print("hola")
else:
print("adios")
# -
if False or False and True:
print("hola")
else:
print("adios")
if ((1 == 2) or (2 == 2)) and (2 == 2):
print("hola")
else:
print("adios")
if 1==2 or 1==2 or (2==2 and 2==2):
print("hola")
else:
print("adios")
if 1==2 or 1==2 or 2==2 and 2==3:
print("hola")
else:
print("adios")
if 1==2 or 1==2 or (2==2 and 2==3):
print("hola")
else:
print("adios")
type(False)
not(False)
if False or True and not(False):
print("hola")
else:
print("adios")
if False or False or True and False:
print("hola")
else:
print("adios")
if 1==2 or 2==2 or 2==1 and 2==1:
print("hola")
else:
print("adios")
if 1==2 or 2==1 or 2==2 and 2==2:
print("hola")
else:
print("adios")
if 1==2 or 2==1 or (True):
print(2)
if 1==2 or (2==1 and 7==7 and 2>1 and 4>4):
print(2)
else:
print(False)
if 1==2 or (2==2 and 7==7 and 2>1) or (2==2 and 4>4):
print(2)
else:
print(False)
if 1==2 or (2==2 and 7==7 and 2>1) or (2==2 and 4>4):
print(2)
else:
print(False)
if 1!=2 or (2>=2 and 7<=7 and 2>1) or (2==2 and 4<5):
print(2)
else:
print(False)
if 2:
print("hola")
""" Los elementos en python que son False:
- 0 (cero)
- Conjuntos vacíos ([], (), {}, "", set())
- False
- None
"""
if 0:
print("hola")
elif [] or ():
print("adios")
elif False:
print("difente")
elif None:
print(None)
if -5815:
print(True)
lista = [0, False, "Hola"]
if lista[0]:
print("1")
elif lista[1]:
print(2)
elif lista[2]:
print(3)
lista = [0, False, "Hola"]
if lista[0]:
print("1")
if lista[1]:
print(2)
if lista[2]:
print(lista[2])
lista = [1, True, "Hola"]
if lista[0]:
print("1")
if lista[1]:
print(2)
if lista[2]:
print(3)
lista = [1, True, "Hola"]
if lista[0]:
print("1")
elif lista[1]:
print(2)
elif lista[2]:
print(3)
# ### Boolean
# and solo devolverá True si TODOS son True
True and False
# +
# or devolverá True si UNO es True
# +
# Sinónimos de False para las condiciones:
# None
# False
# 0 (int or float)
# Cualquier colección vacía --> [], "", (), {}
# El None no actúa como un número al compararlo con otro número
"""
El True lo toma como un numérico 1
"""
# -
if (0 and 1) or 2 or (1 and ()):
print("hola")
if [] or () and None and 1 or 2:
print("adios")
if False and True or True:
print("difente")
if None or False or 0 or 1 and 1:
print(None)
if False or True or False:
print("hola")
if False or False or True:
print("adios")
if False or True:
print("difente")
if False or False or False or True:
print(None)
if True:
print("hola")
if [] or () and None and 1 or 2:
print("adios")
if False and True or True:
print("difente")
if None or False or 0 or 1 and 1:
print(None)
if (0 and 1) or (False) or (1 and ()):
print("hola")
if [] or () and None and 1 or 2:
print("adios")
if False and True or True:
print("difente")
if None or False or 0 or 1 and 1:
print(None)
if 2==2 and not 3==3 or 1==1:
print(1)
if True and False or True:
print(1)
if not 2==2 and 3==3 or not 1==1:
print(1)
if False and True or False:
print(1)
# # Bucles For, While
# Lista alturas
lista = [2, "x", [6]]
print(lista[0])
print(lista[1])
print(lista[2])
# +
lista = [2, "x", [6]]
for x in lista:
print(x)
print("fin")
# +
lista = [2, 1, 7]
# Mostrar todos los elementos de la lista restándoles 1
for elem in lista:
print("--------")
print(elem - 1)
# +
lista = ["x", "y", "z"]
# Mostrar todos los elementos de la lista concatenándoles "F"
for elem in lista:
print(elem + "F")
print(lista)
# +
tupla = ("X", 1, "G")
# Mostrar todos los elementos de la lista concatenándoles "F"
for valor_del_elemento in tupla:
print(valor_del_elemento + "F")
# +
tupla = ("X", 1, "G")
# Mostrar todos los elementos de la lista concatenándoles "F"
for valor_del_elemento in tupla:
print(str(valor_del_elemento) + "F")
# +
tupla = ("X", 1, "G")
tupla_set = set(tupla)
print(tupla_set)
# Mostrar todos los elementos de la lista concatenándoles "F"
for valor_del_elemento in tupla_set:
print(str(valor_del_elemento) + "F")
# +
tupla = ("X", 1, "G")
# Mostrar todos los elementos de la lista concatenándoles "F"
for valor_del_elemento in tupla[1:2]:
if valor_del_elemento == 1:
print(str(valor_del_elemento) + "F")
# -
lista = [2, 4, 5, 8]
x = lista[2] # esto es un escalar Entero
y = lista[2:3] # esto sigue siendo lista
print("x:", x)
print("y:", y)
# +
lista = [[2, 4], [6, "8"]]
for x in lista:
print(x)
# +
lista = [[2, 4], "x", 2, [6, "8"]]
for x in lista:
print(x)
if len(x) >= 3:
print(x)
print("fuera del for")
# +
lista = [[2, 4], "xyz", 2, [6, "8"]]
for x in lista:
print("primer print", x)
if len(x) >= 3: # el error para la ejecución de python
print("segundo print", x)
print("-------------")
print("fuera del for")
# +
x = 2
print(type(x))
# -
x = 2
if type(x) == int or type(x) == float:
print("x es Entero o Float")
x = 2
if isinstance(x, int): # built-in
print("x es Entero")
x = 2
if isinstance(x,(int, float)):
print("x es Entero")
# +
lista = [[2, 4], "xyz", 2, [6, "8"]]
# Mostrar solo los que sean Listas
for x in lista:
if isinstance(x, list): # ¿es x de tipo lista?
print(x)
print("fuera del for")
# +
lista_alturas = [1.66,1.82,1.73,1.75,1.54,1.70,1.70,1.80,1.81,1.87, 1.86, 1.52, 1.70, 1.78, 1.73, 1.78, 1.60, 1.77,1.80, 1.64, 1.79]
# mostrar solo las alturas menores a 1.7
for altura in lista_alturas:
if altura < 1.7:
print(altura)
# +
lista_alturas = [1.66,1.82,1.73,1.75,1.54,1.70,1.70,1.80,1.81,1.87, 1.86, 1.52, 1.70, 1.78, 1.73, 1.78, 1.60, 1.77,1.80, 1.64, 1.79]
# mostrar solo las alturas menores a 1.7
# & guardar en una variable 'alturas_menores' aquellas
alturas_menores = []
for altura in lista_alturas:
if altura < 1.7:
alturas_menores.append(altura)
print(altura)
alturas_menores
# +
lista_alturas = [1.66,1.82,1.73,1.75,1.54,1.70,1.70,1.80,1.81,1.87, 1.86, "x", None, 200, 1.52, 1.70, 1.78, 1.73, 1.78, 1.60, 1.77,1.80, 1.64, 1.79, 164]
# mostrar solo las alturas menores a 1.7
# & guardar en una variable 'alturas_menores' aquellas
alturas_menores = []
# si isinstance se pone en primer lugar, la condición del for solo se va a hacer para aquellos que sea True el isintance
for altura in lista_alturas:
if isinstance(altura, float) and altura < 1.7:
alturas_menores.append(altura)
print(altura)
alturas_menores
# +
lista_alturas = [1.66,1.82,1.73,1.75,1.54,1.70,1.70,1.80,1.81,1.87, 1.86, 1.52, 1.70, 1.78, 1.73, "x", None, 200, 1.78, 1.60, 1.77,1.80, 1.64,1.85, 1.79, 1.63, 164]
# mostrar solo las alturas menores a 1.7
# & guardar en una variable 'alturas_menores' aquellas
# & mostrar los valores que sean enteros
alturas_menores = []
# si isinstance se pone en primer lugar, la condición del for solo se va a hacer para aquellos que sea True el isintance
for altura in lista_alturas:
if isinstance(altura, (float,int)):
if isinstance(altura, int): # Es de tipo int
print("Este valor es Entero:", altura)
elif altura < 1.7: # Es de tipo float
alturas_menores.append(altura)
print(altura)
alturas_menores
# +
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
# Mostrar sólo los nombres:
for x in lista_listas:
print(x[0])
print("END")
# +
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
# Mostrar sólo los nombres : dinero:
for x in lista_listas:
print(x[0], ":", x[1])
print("END")
# -
# range en for
print(list(range(2)))
print(list(range(20)))
lista = ["x", "y", "z"]
rango_de_lista = range(len(lista))
print(list(rango_de_lista))
rango_de_lista
# +
# [start:stop:step]
list(range(0, 4, 1))
# -
list(range(0, 4, 2))
list(range(1, 4, 2))
# +
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
print(list(range(len(lista_listas))))
# +
lista = ["x", "y", "z"]
rango = list(range(len(lista)))
# Mostrar cada uno de los elementos
for i in rango:
print(lista[i])
print("END")
# -
range(3)
# +
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
# Mostrar sólo los nombres:
for i in range(len(lista_listas)):
print(lista_listas[i][0])
print("END")
# -
list(range(4))
# +
nombres = ["Pepe", "Juan", "Lorena", "Rocío"]
dineros = [200, 300, 150, 500]
for i in range(len(nombres)):
print("Iteración:", i + 1)
print("valor de i:", i)
print(nombres[i], ":", dineros[i])
print("----------")
# +
# for anidado
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
for _ in lista_listas:
for elem in _:
print(elem)
# +
# for anidado para mostrar --> nombre : dinero
lista_listas = [["Pepe", 200], ["Juan", 300], ["Lorena", 150], ["Rocío", 500]]
for elemento_lista in lista_listas:
for elem in elemento_lista:
print(elem, ":", elemento_lista[1])
break
# -
lista_nombre_altura = [["<NAME>", 1.75], ]
# +
a = [1, 2]
b = [4, 5]
c = []
contador = 0
for num in a:
c.append(num + b[contador])
contador += 1
c
# -
list(range(len(a)))
# +
a = [1, 2]
b = [4, 5]
c = []
for i in range(len(a)):
c.append(a[i] + b[i])
c
# -
# for normal
lista = [2, 4, 6]
for elem in lista: # elem representa el VALOR del elemento
print(elem)
# +
# for normal con contador
# num representa el VALOR del elemento
# contador representa la POSICIÓN del elemento
c = []
contador = 0
for num in a:
c.append(num + b[contador])
contador += 1
c
# +
# for range
lista = [2, 4, 6]
for i in range(len(lista)): # i representa la POSICIÓN
print(lista[i])
# +
# Enumerate
# i representa la POSICIÓN del elemento y elem es el VALOR del elemento
lista = [2, 4, 6]
for i, elem in enumerate(lista):
print("-------")
print("i:", i)
print("elem:", elem)
# +
a = [1, 2]
b = [4, 5]
c = []
for i, elem in enumerate(a):
c.append(elem + b[i])
c
# +
a = [1, 2, "x", "f", 9]
b = [4, 5, "s", "h", 3]
for i, elem in enumerate(b):
if type(elem) == str: # isinstance(elem, str)
print(elem, a[i])
# -
list(range(5, 10))
# +
# el anterior con un for range
a = [1, 2, "x", "f", 9]
b = [4, 5, "s", "h", 3]
for pos in range(5): # range(len(b))
if type(b[pos]) == str: # isinstance(elem, str)
print(b[pos], a[pos])
# +
b = [4, 5, "s", "h", 3]
# mostrar los elementos de dos en dos si su tipo es int
for pos, v_elem in enumerate(b):
if (pos % 2) == 0 and isinstance(v_elem, int):
print(v_elem)
# +
b = [4, 5, "s", "h", 3]
# mostrar los elementos int que sean pares
for pos, v_elem in enumerate(b):
if isinstance(v_elem, int) and (v_elem % 2) == 0: # el orden al ser el tipo un posible problema se pone primero en la condición
print(v_elem)
# -
list(range(8))
# +
lista = [4, 5, "s2", "h1", 3, ["xs"], "3", []]
# 1. mostrar los elementos en posiciones pares (posiciones: 0, 2, 4)
# 2. si es string mostrar también el segundo caracter si tiene
# 3. si es lista mostrar también el primer elemento si tiene
# 4. mostrar sí o sí el quinto elemento (una vez)
for i, v_elem in enumerate(lista):
if i % 2 == 0: # es una posición par (1.)
print(v_elem)
if isinstance(v_elem, str) and len(v_elem) > 1: # >= 2
print(v_elem[1]) # (2.)
elif isinstance(v_elem, list) and len(v_elem) > 0: # >= 1
print(v_elem[0]) # (3.)
print(lista[4]) # fuera del for para que se ejecute una vez(4.0)
lista
# +
# ** Esto NO muestra un elemento si cumple cualquiera de las 2 condiciones después de comprobar que está en una posición par
lista = [[2, 4], 5, "s2", "h1", 3, ["xs"], "3", []]
# 1. mostrar los elementos en posiciones pares (posiciones: 0, 2, 4)
# 2. si es string mostrar también el segundo caracter si tiene
# 3. si es lista mostrar también el primer elemento si tiene
# 4. mostrar sí o sí el quinto elemento (una vez)
# 5. Sin mostrar el string 's2' pero sí '2'
for i, v_elem in enumerate(lista):
if i % 2 == 0: # es una posición par (1.)
if isinstance(v_elem, str) and len(v_elem) > 1: # >= 2
print(v_elem[1]) # (2.)
elif isinstance(v_elem, list) and len(v_elem) > 0: # >= 1
print(v_elem[0]) # (3.)
else:
print(v_elem)
print(lista[4]) # fuera del for para que se ejecute una vez(4.0)
lista
# +
lista = [[2, 4], 5, "s2", "h1", 3, ["xs"], "3", []]
# 1. mostrar los elementos en posiciones pares (posiciones: 0, 2, 4)
# 2. si es string mostrar también el segundo caracter si tiene
# 3. si es lista mostrar también el primer elemento si tiene
# 4. mostrar sí o sí el quinto elemento (una vez)
# 5. Sin mostrar el string 's2' pero sí '2'
for i, v_elem in enumerate(lista):
if i % 2 == 0: # es una posición par (1.)
if i != 2: # v_elem != "s2"
print(v_elem)
if isinstance(v_elem, str) and len(v_elem) > 1: # >= 2
print(v_elem[1]) # (2.)
elif isinstance(v_elem, list) and len(v_elem) > 0: # >= 1
print(v_elem[0]) # (3.)
print(lista[4]) # fuera del for para que se ejecute una vez(4.0)
lista
# -
# ## Funciones
# +
x = 2
y = 3
suma = x + y
# -
s = "s"
s.upper()
# +
# def nombre_de_la_funcion():
def nombre_de_la_funcion():
print("Me ejecuto desde dentro de una función")
nombre_de_la_funcion() #llamar-invocar-ejecutar la funcion
# -
nombre_de_la_funcion #sin parentesis no se invoca
def suma():
x = 5
y = 7
print(x + y)
suma()
def suma_con_parametro(nombre_parametro):
print(5 + int(nombre_parametro))
suma_con_parametro(2) # no hay que olvidar poener el parametro #suma_con_paramentro(nombre_parametro=2) es lo mismo
x = input("un número:")
suma_con_parametro(nombre_parametro=x)
# +
def multiplicacion_con_parametros(parametro1, parametro2):
print("parametro1:", parametro1)
print("parametro2:", parametro2)
if parametro1 > 0 and parametro2 > 0:
print(parametro1 * parametro2)
else:
print("solo se admiten números mayores a 0")
multiplicacion_con_parametros(parametro1=-2, parametro2=3)
# +
def multiplicacion_con_parametros(parametro1, parametro2):
print("parametro1:", parametro1)
print("parametro2:", parametro2)
if parametro1 > 0 and parametro2 > 0:
print(parametro1 * parametro2)
else:
print("solo se admiten números mayores a 0")
param1 = int(input())
param2 = int(input())
multiplicacion_con_parametros(parametro1=param1, parametro2=param2)
# -
#repaso dia 4
l[1,None,4,0,51,None]
for elem in l:
if elem: # esto es para ver si existe(si es verdadero) no va a saltar el none y el 0 si hay 0 y quieres recogerlo como un numero esto es una mala praxis
#cuidado con ` y otros signos que se cuelan
#repaso dia 4
tupla=("x","1","g")
for elem in tupla [1:2]:
if elem=="1":
print(elem+"F")
#hemos visto 3 tipos de for:
# 1. for normal:
for valor_elemento in nombre_lista:
#for con range
# 2. for con range:
for posicion in range(len(nombre_lista)):
# 3. for con enumerate
for posicion, valor_elemento in enumerate(nombre_lista):
# las funciones pueden utilizar variables preexistentes
param=4
def nf(param1):
print(2)
print(param)
if param==2:
print (param+2)
nf(param1=65) # imprime el 2 y el valor de param, creado fuera. el if no lo cumple,
# aunque tu metas un valor a la param1 en este caso param1=65 dentro de la funcion, da igual el valor que le des porque devuelve lo que le pidas
# +
#RETURN EN LA FUNCION: guarda informacion en una variable
#Print se imprime-muestra en pantalla
#print retorna el valor
# -
#las funciones por defecto tiene un "return None" TODAS las funciones tienen un return none aunque no lo veamos
def f():
print("hola")
return None # no se ve pero existe ahora lo escribimos para verlo
print(f()) # aqui se muestra en la pantalla la resolucion de la funcion y tambien te va a mostra rel retorno
#jupyter Notebooks no muestra None como output por defecto cuando el none esta en la ultima fila, si no hace el print(x) muestra el none xq no es la ultima linea SOLO EN JUPYTER
x =f() # x = asignamos a la variable x la funcion f()
x #asi ejecuta la funcion pero no muestra el retorno
# si hago el f() sin el print no va a mostra r el none xq es la ultima linea de jupyter y en jupyter no funciona
f()
#buenas practicas son para que muestre el none
x=f() #asi me ejecuta la funcion y asigno a x la funcion f()
print(x) # asi me muestra la el retorno de la variable x que le hemos asignado la funcion f()
print(f()) # asi me muestra el retorno de la funcion
# +
#ejemplo de lo anterior
def f_con_return():
print(2)
return 3
x=f_con_return() #x estamos invocando a la funcion como la invocacion es un print de 2 es lo que devuelve ese print y estamos asignando a x el valor de esa funcion y guarda el return 3 en la variable x
print(x) # como le hemos asignado a la variable x el valor de la funcion, con print de x muestra el retorno
# -
#diferencias return sin return
def suma_sin_return (a,b):
print(a+b)
def suma_con_retorno(a,b):
print(a+b)
return a+b
suma_sin_return(a=2,b=2) #ejecuta la fucnion
suma_con_retorno(a=2,b=2) # ejecuta la funcion
lo_que_retorna_la_funcion_sin_retorno=suma_sin_return(a=2,b=2) #asignamos suma_sin_return a la variable lo_que_retorna_la_funcion_sin_retorno no tiene nada retornado en lo_que_retorna_la_funcion_sin_retorno asi que si hacemos print no sale nada. ejecutala funcion tambien
print(lo_que_retorna_la_funcion_sin_retorno)
# +
lo_que_retorna_la_funcion_con_retorno= suma_con_retorno(a=2,b=2) #asignamos la lo_que_retorna_la_funcion_con_retorno a suma_con_retorno y le guarda el valor 4 (return) y la ejecuta
print(lo_que_retorna_la_funcion_con_retorno) #printea lo que esta guardado e lo_que_retorna_la_funcion_con_retorno
# -
def multiplicar_lo_sumado(lo_sumado):
print(lo_sumado*2)
def suma_sin_return(a,b):
print(a+b) #return None
def suma_con_retorno(a,b):
print(a+b)
return a+b # tiene return
def multiplicar_lo_sumado(lo_sumado):
print(lo_sumado*2) # return None
# +
#ejemplo 1: suma con return
x=suma_con_retorno(2,5) # asignamos a la varbialbe x la funcion suma_con_retorno. ademas ejecuta la funcion con esos valores y guarda en x el return a+b
y=multiplicar_lo_sumado(lo_sumado=x) # asignamos a la varbialbe y la funcion multiplicar_lo_sumado. ademas ejecuta la funcion con esos valores que estamos diciendo que la variable de la funcion, llamada lo_sumado sea igual al valor de ejecutar x. no se guarda nada None
print(y) #muestra el retorno de la variable y ya que esta asignada a una variable
print(x) #muestra el retorno de la variable x ya que esta asignada a una variable, valor guardado en x (return)
# -
#ejemplo 3: con suma con returin y sin print al final
#x es lo q retorna la funcion a la derecha de la igualdad
x=suma_con_retorno(a=2,b=4) #asignamos a la varbialbe x la funcion suma_con_retorno. ejecuta la funcion, como dentro de la funcion hay un print lo muestra
y=multiplicar_lo_sumado(lo_sumado=x) # asignamos a la varbialbe y la funcion multiplicar_lo_sumado. ejecuta la funcion, como hay un print lo muestra
y #no retorna el retorno ya que es none habria que hacer un print para que lo mostrase la ultima fila si no hay print no muestra el retorno. no ha guardado nada en y por lo tanto no lo muestra si
print(y) #ves ahora muestra el retorno no retorna el retorno
def f():
print("hola")
c= 2+7
print("c de dentro:", c)
h=(2,4,6)
x=f() #asignamos a x la funcion f() y se ejecuta y no hemos guardado nada (return) en x
print(x) # muestra el retorno de la varabiale x, ya que se ha asignado a una funcion
# +
def f():
return 1 # lo ultimo que lee una funcion durante su ejecucion es el return
print("hola")
c= 2+7
print("c de dentro:", c)
h=(2,4,6)
x=f() #asigna a una variable una funcion y la ejecuta hemos guardado 1 en la funcion
x #como el retorno es diferfente a none lo muestra aunque no se haga con print mejor siempre hacer print guardado en x
# -
def f():
print("hola")
c= 2+7
print("c de dentro:", c)
h=(2,4,6)
return h
return 1
x=f() #asigna a una variable una funcion y la ejecuta guarda al tupla en x
print(x) #esto muestra el retorno de la variable que le hemos asignado la funcion solo lee el primer return muestra la tupla guardad en x
def f():
print("hola")
c= 2+7
print("c de dentro:", c)
h=(2,4,6)
return h
return 1
x=f() #ejecuta la funcion y le asignamos a x la funcion f no muestra el retorno hasta que se ponga x al final o print x
#las funcioens predeterminadas, o bien se ponen directamente lista.append(2) o bien se ponen con el = lista=lista.append, prueba las dos si no hay retorno no hace falta el =. se puede mirar en google para ver si tiene o no return
# #append, remove, del, pop(elimina el ultimom elemento) etc
lista= [2,3,4]
lista.append(2)
print(lista)
lista= [2,3,4]
lista=lista.append(2) #aqui se ve que no tiene return asi que no añade el 2 a la lista si no que devuelve lo guardado en lista
print(lista)
lista=[2,4,6]
p=lista.pop() #elimina y alamacena en p el elemento eliminado, return
print(lista) #muestra la lista con el elemento borrado
p #muestra el print del return de p
#si lo haces sin el =
lista2=[2,4,6]
lista2.pop()
print(lista2) # tambien se elimina aunque tenga return la funcion
print(lista2.pop()) #se alamacena el ultimo elemento de la lista que se va a eleminar
x=print("2") #ejecuta la funcion print
print(x) # no hay nada almacenado en x, nada retornado
# +
#crear documentacion sirve para explicar cosas en proyectos grupales, o para tu yo del futuro. no afecta a funciones si haces help (jk) te enseña la ayuda del a funcion
def jk(a):
"""
#esto es documentacion
esta funcion lo q hace es crear una variable q alberga la suma del paraemtro "a más 2
parametros:
"a": una variable que será sumada con 2
return:variable "suma"
"""
suma=2+a
return suma
help(jk) # esta es la funcion de ayuda de la funcion x y su docuemtnacion, informacion de la funcion
# -
#if
def multiplicacion_con_parametros (a,b):
a=int(a) # o bien se pone el int aqui o fuera en la parte de los param1 y param2
b=int(b)
print("Parametro1:", a)
print("parametro2:",b)
if a>0 and b>0:
print(a*b)
else:
print("solo se admiten mayores a 0")
param1=input()
param2=input()
multiplicacion_con_parametros(a=param1,b=param2)
# +
#pass #este elemento significa no hagas nada para que no te de error la identacion "pasa de lo que estoy haceindo" es para que no te de error una idetnacion (:)
# +
#lista
def elimina_ultimo_elemento (lista):
lista.pop() #elimina el ultimo elemetno, salvo que se indique en que posicion esta el elemento que quieres borrar
elimina_ultimo_elemento(lista= [2,4]) #aqui hay que meter una lista en verdad puedes poner el nombre que quieras (lista pero esto no significa que este correcto)
lista #printea el return
# -
def resta_dos (x): #este es 4
x= x-2 # esta variable es local dentro de la funcion 1º hacemos la derecha primero sustituyo la x de la derecha es 4 y luego ejecuta dentro es 2
return(x) #esta x es la izquierda de la igualdad que es 2
x=4 #variable global
resta_dos(x=x) #llamando a una funcion al que le voy a dar el argumento x= a la x varaible global (4) # en esta funcion es donde se esta haciendo el return ya que no lo hemos asignad oa ninguna variable
x #asi vale 4 porque no estoy modiicando ninguna variable, es decir me devuelve el x=4 como print de x
print(resta_dos(x=x)) #muestra el return que se almacena aqui y no en ninguna variable se almacena en la propia funcion
#para que el x me saliera el return habria que asignarlo a una variable
x=print(resta_dos(x=x)) #lee la ultima linea superior
x #muestra el print del return
# +
#una # una variable x que tenga los nombres concatenados de la lista nombres altura utilizando una funcion para ello.
# x=nombre_funcion(lista=lista_nombres_altura)
lista_nombres_alturas = [("<NAME>", 1.75),("<NAME>", 1.70),("<NAME>", 1.82),("<NAME>", 1.80),('<NAME>', 1.86),('<NAME>', 1.73), ('<NAME>', 1.79), ("<NAME>", 1.52), ('<NAME>', 1.75), ("<NAME>",1.78),("<NAME>",1.70), ("<NAME>", 1.78),("<NAME>", 1.54), ("<NAME>", 1.87), ("<NAME>", 1.66), ("<NAME>", 1.63), ("<NAME>", 1.70), ("<NAME>", 1.64), ("<NAME>",1.80), ("<NAME>", 1.77), ("<NAME>", 1.64), ("<NAME>", 1.85), ("<NAME>", 1.82), ("<NAME>", 1.81), ("<NAME>", 1.60),("<NAME>", 1.84),("<NAME>",1.61)]
def nombre_funcion (lista):
nombres="" #crear una coleccion vacia
for tupla in lista:
#tupla va a representar cada uno de estos eelemtnos las tuplas de la lista ("<NAME>", 1.75) tupla =("<NAME>", 1.75) por iteracion...
nombres=lnombres+tupla[0]) # el primer elemento de cada tupla el elemento primero de cada tupla de la lista
return nombres #porque todos los nombres concatenados estan en esta variable
x=nombre_funcion(lista=lista_nombres_alturas) # al final los nobmres e van a guardar en return de la funcion. cuando se resuelva lo de la derecha, lo que retorna la parte de la derecha se guarda en x. #al realiar esta igualdad dnetro de la funcion se dice que la variable lista_nombres_altura se va a llamar lista
print(x)
#añadido si pongo en nombre =0.0 me suma todas las alturas si el resto de variables estan sumando las alturas
# -
#extra replace algo por algo
g=("aa s d gggh hh")
g=g.replace(" ","") #quitar al menos donde haya un espacio y poner 0 espacios
print(g)
#repaso para calcular la media
lista=[2,5,11,20]
for i, value in enumerate(lista[0:len(lista)//2]):
print ("i:",i)
print("value:",value)
#repaso para calcular la media
lista=[2,5,11,20]
for i, value in enumerate(lista[len(lista)//2:]): #desde la mitad hasta el final
print ("i:",i)
print("value:",value)
A=[1,2,"x","y"]
b=[4,5,"s","h"]
for i,value in enumerate(b): #isinstance(elem,str) es lo mismo, hay que poner elemento y el tipo
if type elem(a)==str:
print(elem,b[i])
# +
#1. lo que se vaya a utilziar en una funcion normalmente se crean dentro de la funcion ya qeue no se utiliza fuera(ejemplo contadores)
#2. si necesito utilizar dentro de una funcion un valor de fuera se lo paso por parametro. los parametros son los que usamos al definir la funcion def funcion1 (a,b,c) -> estos son los parametros
#se pueden retornar varias cosas a la vez
# -
x=2
x,y=2,6 #podemos meter xa varias variables varios valores x sera 2 y y sera 6
t=2,6,5 # si guardo mas de un elemento en una variable se guarda como una tupla
print(x)
print(y)
print(t)
# +
x,y=10,20,30 #nse puede guardar en x e y mas de 2 valores, no se guardan como tupla
# -
def retorna_varios():
print("yo retorno")
return 2,4,6
x=retorna_varios() #ejecuta la funcion
print(x) #retorna el retorno de x como una tupla
y,z,g=retorna_varios() #ejecuta la funcion
print(y)#printea el retorno de y
print(z)#printea el retorno de z
print(g)#printea el retorno de g
lista=["x","y","z"]
for t in enumerate(lista): #con enumerate si solo se pone una variable, se muestra las dos variables juntas en una tupla, posicion y valor
print(type(y))
print(t)
for tt in enumerate(lista):
print("posicion",t[0])#muestra la posicion de la tupla creada
print("valor del elemento:",t[1]) #muestra el valor de la tupla creada
# +
#DICCIONARIOS -> es un tipo de coleccion
#lista, tupla, string, conjuntos y diccionarios
# -
lista=[]
tupla=() #recordar que si usamos un unico elemento en una tupla hay que escribir el elemento y la coma xq si no era un numero entero
string=""
conjunto=set() #asi se inicia un conjunto
diccionario={}#asi se inicia un diccionario
type(diccionario)
#los diccionarios siempre guardan una key y un valor. la key suele ser un string
diccionario={"key_string": "valor", 0:"valor"} #key a la izquierda valor a la derecha el 0: es otra clave con otros valores. no podemos teener dos claves con el mismo nombre pero si dos varaibles con el mismo numero
diccionario
diccionario={"key_string": "valor", 0:"valor",0:"a","key":999}
diccionario #ves sale solo una clave con el nombre. si hay dos claves con el mismo nombre la ultima sustituye a la primera
#acceder a un valor en un diccionario
list(diccionario["key"])
list(diccionario.keys()) #accedemos a las claves con una lista xq hemos puesto lista
list(diccionario.values())#surgen los valores en una lista xq se ha puesto lista
#queremos saber aquellos valores mayores que 0
for value in diccionario.values(): #recorremos los valores del diccionario
if isinstance(value,int) and value>0: #comprobar que el tipo del valor es entero y mayor a 0
print(value)
for key in diccionario.keys(): #recorremos las claves del diccionario
if isinstance(key,str): #comprobar que el tipo de la clave es entero y mayor a 0
print(key) #muestra clave
print(diccionario[key])#muestra valor asociado a la clave
print("-------------------")
#con .items es como el enumnerate de las tablas i y valor
for key,value in diccionario.items():
print("key:",key)
print("value:",value) #print(diccionario[key] hacer esto es sinonimo con lo que se ejecuta, muestra el valor
print("------")
dicc={}
#para añadir clave:valor. no existen valores o claves sueltos
dicc["nombre_de_la_clave"]="valor_de_esa_clave" #asi se añade la key y el valor
dicc
#eliminar clave:valor se eliminan a pares
del dicc["nombre_de_la_clave"] #poniendolo asi se elimina la clave valor
dicc
#crear un diccionario ap artir de dos listaS:
#lista "a" tiene claves
#lista "b" tiene los valores asociados a cada clave
a=["x","y","z"]
b=[0,10,20]
diccionario={}
diccionario[a[0]]=b[0] #seria asi con cda valor muy tedioso, vamos a usar otras opcciones
#utilizando el rango
diccionario={}
for i in range(len(a)):
diccionario[a[i]]=b[i]
diccionario
#con contador
a=["x","y","z"]
b=[0,10,20]
contador=0 #con posiciones creo este contador
diccionario={}
for v_elemento in a:
diccionario[v_elem]=b[contador]
contador+=1 #esto es igual a contador mas 1
print(diccionario)
#con enumerate
a=["x","y","z"]
b=[0,10,20]
for pos,v_element in enumerate(a):
diccionario[v_element]=b[pos]
diccionario
#recorrer lista con contador EJEMPLO
a=[2,4,6,8,10]
contador=0 #si queremos que reresente la posion del elemento que estamos recorriendo es como una posicion
for valor in a:
if contador>=2: #para mostrar los elementos en la posicion mayor e igual a 2
print(valor)
print(contador)
contador+=1
print("-------------------")
a=[2,4,6,8,10] #que nos muesre las posiciones pares
#con rango
for pos in range(len(a)):
if pos%2==0:#para posiciones pareces
print(a[pos])
# +
#añadimos /actualizamos el par de valores
dicc["70423563R"]=["pablo",28,1.81,73]
#asi se cogeria y se actualizaria el valor del elemento
dicc["70423563R"]=["pablo",28,1.81,74] # se ha cambiado
dicc
# -
#la clave tiene que ser un str,float un entero o una tupla lista NO se puede
dicc[("70423563R",1)]={1:("Z","H")}
dicc
#para acceder a la "Z" que esta como parte del valor del diccionario
dicc[("70423563R",1)] [1][0] #accedemos al segundo diccionario y luego a la tupla y luego ya estariamos en el valor que buscamos se accede con la clave
# +
##WHILE
# -
for i in range(20):
print(i) #esta linea se repite 20 veces, sea lo que sea lo que se printea, str, lista etc
for i in range(20):
if i<5:
print(i)
#while es una condicion que mientras sea true se va a hacer todo lo de abajo del while en bucle hasta que se quite el true
while True: #por ejemplo la app esta abierta hasta que de al boton de cerrar (no hacerlo porque salen unos de manera infinita) al lado del while se pone la condicion
print("1")#(no hacerlo porque salen unos de manera infinita)
break #break para que se muestre una vez
# +
#while (conditions):
# lo que se va a ejecutar en bucle
# -
#quiero ejecutar 5 prints con contador
contador=0
while contador <5: #en este caso en cuanto el contador llega a 5 hasta que se satisfaace la condicion
print(contador)
contador+=1
#la difeerencia con for es que recorre algo sobre un valor while no hace falta asi se haria con un for es mas eficiente el while
for i in range(100): #con el for sigue leyendo hasta rango a 100
if i<5:
print(i)
#dentro del while puede haber de todo con while se accede por POSICION POR ELEMENTO, no por valor el while no recorre como el for simplemente tiene en cuenta la condicion
a=["x","y","z","o"]
contador=0
#[0,3]->[] incluido
#[0,4) -> 0 incluido 4 sin incluir
while contador<len(a): # ==contador<=len(a)-1 con esto no puede valer mas de la ultima posocion
print("contador:", contador) #mientras uno exista se va a reproducir de manera infinita
print(a[contador])
contador+=1 # si se deja asi daria error
# +
#para acceder a las posciones pares
a=["x","y","z","o"] # a diferencia del for haciendo este ejercicio no lee las posiciones impares
contador=0
while contador<len(a): # ==contador<=len(a)-1 con esto no puede valer mas de la ultima posocion
print("contador:", contador) #mientras uno exista se va a reproducir de manera infinita
print(a[contador])
contador+=2
# +
#como funciona while
a=["x","y","z","o"]
contador=0
while (contador %2==0) and (contador<len(a)): # ==contador<=len(a)-1 en la posicion 0 se cumple las condiciones asi que lee lo de abajo y lo ejecuta
print("contador:", contador)
print(a[contador])
contador+=1
print("hola") #como vemos en el printeado del while, muestra tanto el printeo de " contador" y el contado (0) luego suma uno al contador (posicion 1) y printea "1" en la siguiente lectura la posicion 1 no es par asi que como no se cumple las 2 condiciones deja de leer y while deja de leer todo lo demas.
# -
#no hace falta para ahora ignorar
import time #importar la libreria time
acumulador = 0
while acumulador<3:
print("acumulador:",acumulador)
time.sleep(1) # con esto se para 1 segundo y luego sigue mostrando el resto
acumulador +=1
#programa que pregunte al usuario que numero quiere que se sume con el "3"
num=int(input("escribe un numero para sumar a 3"))
def suma_con_3(lo_que_el_usuario_escriba):
print(lo_que_el_usuario_escriba+3)
suma_con_3(lo_que_el_usuario_escriba=2)
while True:
num=input("escribe un numero para sumar a 3")
if num=="STOP":
print("has parado")
break #asi se consigue que si se escribe STOP se djee de ejecutar
suma_con_3(lo_que_el_usuario_escriba=int(num))#si escribo un str que no sea "STOP" daria error
# +
#is.digit() es para ver los numeros enteros dentro de cadenas
while True:
num=input("escribe un numero para sumar a 3")
if num=="STOP": #si num es stop
print("has parado")
break #asi se consigue que si se escribe STOP se djee de ejecutar
elif num.isdigit(): #si num no es stop y ademas solo contiene numeros
suma_con_3(lo_que_el_usuario_escriba=int(num))
else:
print("insterta solo numeros o 'STOP'") #si por ejemplo meto una lista, una letra etc sale esto
# +
### continue or break
l=[2,"x",6,[0,2],"y"]
for pos,v_elem in enumerate(l):
if isinstance(v_elem,str): #enseña los elementos string de los valores
print ("------------")
continue #con el continue se dirige a la siguiente interaccion en el for. no lee lo de abajo del continue. el break para toda la interaccion
print ("pos:", pos)
print("v_elem:",v_elem)
#si ahora poniese un break lo recorre un avez y ya esta
# -
l=[2,"x",6,[0,2],"y"]
for pos,v_elem in enumerate(l):
if pos %2==0:#la posicion par esto es igual a if pos %2==1: print (v_elem) y daria lo mismo
continue # si esto es par no leas mas y ve a la siguiente interracion se puede poner cualquier funcion sin el continue es mas facil a ve ces cuando hy muchas condiciones
else:
print(v_elem)
# son sinonimos las dos siguiente entradas
l=[2,"x",6,[0,2],"y"]
for pos,v_elem in enumerate(l):
if v_elem ==6:
continue
else:
print(v_elem)
l=[2,"x",6,[0,2],"y"]
for pos,v_elem in enumerate(l):
if v_elem !=6:
print(v_elem)
#PARA ESTAS COSAS MIRAR ARCHIVO PYTHON_BASIC DENTRO DE LA CARPETA DE PYTHON EN EL INDICE ESTA Todo bastante descrito
lista=[2,34,6]
lista.insert(1,"x") #se agrega en la posicion 1 la "x"
print(lista)
# +
def shout(text):
return text.upper() #pone un texto en mayusculas
def whisper(text):
return text.lower() #pone un texto en minus
def greet(func): #saluda
# storing the function in a variable
greeting = func("Hi, I am created by a function passed as an argument.") #le meto el argumento directamente a func
print(greeting)
greet(shout)
greet(whisper)
# -
def up (text):
return text.upper()
def f_recibe_una_funcion(func):
print(func("Hola")) #etso muestra el retorno de la funcion, en este caso el retorno de up
f_recibe_una_funcion(func=up)
x=up # esto es hacer una referencia de una funcion
x(text="hola")
def up (text):
return text.upper()
def dos():
return "2"
def f_recibe_una_funcion(func):
print(func("Hola")) #este error es xq dos no tiene ningun argumento definido () no tendrias que poner ningun argumento para poder ejecutarlo y yo le estoy metiendo un argumento de manera forzada
f_recibe_una_funcion(func=dos)
def up (text):
return text.upper()
def dos(num_string):
return num_string
def f_recibe_una_funcion(func):
print(func("Hola")) #el "hola" es el nombre del parametro x el que se va a sustituir el num_string dentro de dos y text dentro de up. no paso la referencia si no la llamada en este caso es la llamada
f_recibe_una_funcion(func=dos)
def up (text):
return text.upper()
def dos(num_string):
return num_string
def f_recibe_una_funcion(func):
print(func(text="Hola"))# en este caso no daria error con up xq text esta definido en la funcion up, ya que func se sustituye por up pero si se sustituye por la funcion dos al text no estar definido daria error esto es una referencia, referencias un parametro/argumentos a un valor
f_recibe_una_funcion(func=dos)
# +
#argfumentos / parametros es lo mismo dentro de una funcion (son las cosas en los parentesis cuando defines la funcion)
|
week1_precurse_python_I/day5_python_IV/theory/Python_IV_Precurse.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Chapter 1 – The Machine Learning landscape**
#
# _This is the code used to generate some of the figures in chapter 1._
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/01_the_machine_learning_landscape.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# </td>
# <td>
# <a target="_blank" href="https://kaggle.com/kernels/welcome?src=https://github.com/ageron/handson-ml2/blob/master/01_the_machine_learning_landscape.ipynb"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
# </td>
# </table>
# # Code example 1-1
# Although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead.
# + slideshow={"slide_type": "-"}
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# -
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# This function just merges the OECD's life satisfaction data and the IMF's GDP per capita data. It's a bit too long and boring and it's not specific to Machine Learning, which is why I left it out of the book.
def prepare_country_stats(oecd_bli, gdp_per_capita):
oecd_bli = oecd_bli[oecd_bli["INEQUALITY"]=="TOT"]
oecd_bli = oecd_bli.pivot(index="Country", columns="Indicator", values="Value")
gdp_per_capita.rename(columns={"2015": "GDP per capita"}, inplace=True)
gdp_per_capita.set_index("Country", inplace=True)
full_country_stats = pd.merge(left=oecd_bli, right=gdp_per_capita,
left_index=True, right_index=True)
full_country_stats.sort_values(by="GDP per capita", inplace=True)
remove_indices = [0, 1, 6, 8, 33, 34, 35]
keep_indices = list(set(range(36)) - set(remove_indices))
return full_country_stats[["GDP per capita", 'Life satisfaction']].iloc[keep_indices]
# The code in the book expects the data files to be located in the current directory. I just tweaked it here to fetch the files in `datasets/lifesat`.
import os
datapath = os.path.join("datasets", "lifesat", "")
# To plot pretty figures directly within Jupyter
# %matplotlib inline
import matplotlib as mpl
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Download the data
import urllib.request
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml2/master/"
os.makedirs(datapath, exist_ok=True)
for filename in ("oecd_bli_2015.csv", "gdp_per_capita.csv"):
print("Downloading", filename)
url = DOWNLOAD_ROOT + "datasets/lifesat/" + filename
urllib.request.urlretrieve(url, datapath + filename)
# +
# Code example
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn.linear_model
# Load the data
oecd_bli = pd.read_csv(datapath + "oecd_bli_2015.csv", thousands=',')
gdp_per_capita = pd.read_csv(datapath + "gdp_per_capita.csv",thousands=',',delimiter='\t',
encoding='latin1', na_values="n/a")
# Prepare the data
country_stats = prepare_country_stats(oecd_bli, gdp_per_capita)
X = np.c_[country_stats["GDP per capita"]]
y = np.c_[country_stats["Life satisfaction"]]
# Visualize the data
country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction')
plt.show()
# Select a linear model
model = sklearn.linear_model.LinearRegression()
# Train the model
model.fit(X, y)
# Make a prediction for Cyprus
X_new = [[22587]] # Cyprus' GDP per capita
print(model.predict(X_new)) # outputs [[ 5.96242338]]
# -
# Replacing the Linear Regression model with k-Nearest Neighbors (in this example, k = 3) regression in the previous code is as simple as replacing these two
# lines:
#
# ```python
# import sklearn.linear_model
# model = sklearn.linear_model.LinearRegression()
# ```
#
# with these two:
#
# ```python
# import sklearn.neighbors
# model = sklearn.neighbors.KNeighborsRegressor(n_neighbors=3)
# ```
# +
# Select a 3-Nearest Neighbors regression model
import sklearn.neighbors
model1 = sklearn.neighbors.KNeighborsRegressor(n_neighbors=3)
# Train the model
model1.fit(X,y)
# Make a prediction for Cyprus
print(model1.predict(X_new)) # outputs [[5.76666667]]
# -
# # Note: you can ignore the rest of this notebook, it just generates many of the figures in chapter 1.
# Create a function to save the figures.
# +
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "fundamentals"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# -
# Make this notebook's output stable across runs:
np.random.seed(42)
# # Load and prepare Life satisfaction data
# If you want, you can get fresh data from the OECD's website.
# Download the CSV from http://stats.oecd.org/index.aspx?DataSetCode=BLI
# and save it to `datasets/lifesat/`.
oecd_bli = pd.read_csv(datapath + "oecd_bli_2015.csv", thousands=',')
oecd_bli = oecd_bli[oecd_bli["INEQUALITY"]=="TOT"]
oecd_bli = oecd_bli.pivot(index="Country", columns="Indicator", values="Value")
oecd_bli.head(2)
oecd_bli["Life satisfaction"].head()
# # Load and prepare GDP per capita data
# Just like above, you can update the GDP per capita data if you want. Just download data from http://goo.gl/j1MSKe (=> imf.org) and save it to `datasets/lifesat/`.
gdp_per_capita = pd.read_csv(datapath+"gdp_per_capita.csv", thousands=',', delimiter='\t',
encoding='latin1', na_values="n/a")
gdp_per_capita.rename(columns={"2015": "GDP per capita"}, inplace=True)
gdp_per_capita.set_index("Country", inplace=True)
gdp_per_capita.head(2)
full_country_stats = pd.merge(left=oecd_bli, right=gdp_per_capita, left_index=True, right_index=True)
full_country_stats.sort_values(by="GDP per capita", inplace=True)
full_country_stats
full_country_stats[["GDP per capita", 'Life satisfaction']].loc["United States"]
# +
remove_indices = [0, 1, 6, 8, 33, 34, 35]
keep_indices = list(set(range(36)) - set(remove_indices))
sample_data = full_country_stats[["GDP per capita", 'Life satisfaction']].iloc[keep_indices]
missing_data = full_country_stats[["GDP per capita", 'Life satisfaction']].iloc[remove_indices]
# -
sample_data.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(5,3))
plt.axis([0, 60000, 0, 10])
position_text = {
"Hungary": (5000, 1),
"Korea": (18000, 1.7),
"France": (29000, 2.4),
"Australia": (40000, 3.0),
"United States": (52000, 3.8),
}
for country, pos_text in position_text.items():
pos_data_x, pos_data_y = sample_data.loc[country]
country = "U.S." if country == "United States" else country
plt.annotate(country, xy=(pos_data_x, pos_data_y), xytext=pos_text,
arrowprops=dict(facecolor='black', width=0.5, shrink=0.1, headwidth=5))
plt.plot(pos_data_x, pos_data_y, "ro")
plt.xlabel("GDP per capita (USD)")
save_fig('money_happy_scatterplot')
plt.show()
sample_data.to_csv(os.path.join("datasets", "lifesat", "lifesat.csv"))
sample_data.loc[list(position_text.keys())]
# +
import numpy as np
sample_data.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(5,3))
plt.xlabel("GDP per capita (USD)")
plt.axis([0, 60000, 0, 10])
X=np.linspace(0, 60000, 1000)
plt.plot(X, 2*X/100000, "r")
plt.text(40000, 2.7, r"$\theta_0 = 0$", fontsize=14, color="r")
plt.text(40000, 1.8, r"$\theta_1 = 2 \times 10^{-5}$", fontsize=14, color="r")
plt.plot(X, 8 - 5*X/100000, "g")
plt.text(5000, 9.1, r"$\theta_0 = 8$", fontsize=14, color="g")
plt.text(5000, 8.2, r"$\theta_1 = -5 \times 10^{-5}$", fontsize=14, color="g")
plt.plot(X, 4 + 5*X/100000, "b")
plt.text(5000, 3.5, r"$\theta_0 = 4$", fontsize=14, color="b")
plt.text(5000, 2.6, r"$\theta_1 = 5 \times 10^{-5}$", fontsize=14, color="b")
save_fig('tweaking_model_params_plot')
plt.show()
# -
from sklearn import linear_model
lin1 = linear_model.LinearRegression()
Xsample = np.c_[sample_data["GDP per capita"]]
ysample = np.c_[sample_data["Life satisfaction"]]
lin1.fit(Xsample, ysample)
t0, t1 = lin1.intercept_[0], lin1.coef_[0][0]
t0, t1
sample_data.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(5,3))
plt.xlabel("GDP per capita (USD)")
plt.axis([0, 60000, 0, 10])
X=np.linspace(0, 60000, 1000)
plt.plot(X, t0 + t1*X, "b")
plt.text(5000, 3.1, r"$\theta_0 = 4.85$", fontsize=14, color="b")
plt.text(5000, 2.2, r"$\theta_1 = 4.91 \times 10^{-5}$", fontsize=14, color="b")
save_fig('best_fit_model_plot')
plt.show()
cyprus_gdp_per_capita = gdp_per_capita.loc["Cyprus"]["GDP per capita"]
print(cyprus_gdp_per_capita)
cyprus_predicted_life_satisfaction = lin1.predict([[cyprus_gdp_per_capita]])[0][0]
cyprus_predicted_life_satisfaction
sample_data.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(5,3), s=1)
plt.xlabel("GDP per capita (USD)")
X=np.linspace(0, 60000, 1000)
plt.plot(X, t0 + t1*X, "b")
plt.axis([0, 60000, 0, 10])
plt.text(5000, 7.5, r"$\theta_0 = 4.85$", fontsize=14, color="b")
plt.text(5000, 6.6, r"$\theta_1 = 4.91 \times 10^{-5}$", fontsize=14, color="b")
plt.plot([cyprus_gdp_per_capita, cyprus_gdp_per_capita], [0, cyprus_predicted_life_satisfaction], "r--")
plt.text(25000, 5.0, r"Prediction = 5.96", fontsize=14, color="b")
plt.plot(cyprus_gdp_per_capita, cyprus_predicted_life_satisfaction, "ro")
save_fig('cyprus_prediction_plot')
plt.show()
sample_data[7:10]
(5.1+5.7+6.5)/3
# +
backup = oecd_bli, gdp_per_capita
def prepare_country_stats(oecd_bli, gdp_per_capita):
oecd_bli = oecd_bli[oecd_bli["INEQUALITY"]=="TOT"]
oecd_bli = oecd_bli.pivot(index="Country", columns="Indicator", values="Value")
gdp_per_capita.rename(columns={"2015": "GDP per capita"}, inplace=True)
gdp_per_capita.set_index("Country", inplace=True)
full_country_stats = pd.merge(left=oecd_bli, right=gdp_per_capita,
left_index=True, right_index=True)
full_country_stats.sort_values(by="GDP per capita", inplace=True)
remove_indices = [0, 1, 6, 8, 33, 34, 35]
keep_indices = list(set(range(36)) - set(remove_indices))
return full_country_stats[["GDP per capita", 'Life satisfaction']].iloc[keep_indices]
# +
# Code example
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn.linear_model
# Load the data
oecd_bli = pd.read_csv(datapath + "oecd_bli_2015.csv", thousands=',')
gdp_per_capita = pd.read_csv(datapath + "gdp_per_capita.csv",thousands=',',delimiter='\t',
encoding='latin1', na_values="n/a")
# Prepare the data
country_stats = prepare_country_stats(oecd_bli, gdp_per_capita)
X = np.c_[country_stats["GDP per capita"]]
y = np.c_[country_stats["Life satisfaction"]]
# Visualize the data
country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction')
plt.show()
# Select a linear model
model = sklearn.linear_model.LinearRegression()
# Train the model
model.fit(X, y)
# Make a prediction for Cyprus
X_new = [[22587]] # Cyprus' GDP per capita
print(model.predict(X_new)) # outputs [[ 5.96242338]]
# -
oecd_bli, gdp_per_capita = backup
missing_data
position_text2 = {
"Brazil": (1000, 9.0),
"Mexico": (11000, 9.0),
"Chile": (25000, 9.0),
"Czech Republic": (35000, 9.0),
"Norway": (60000, 3),
"Switzerland": (72000, 3.0),
"Luxembourg": (90000, 3.0),
}
# +
sample_data.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(8,3))
plt.axis([0, 110000, 0, 10])
for country, pos_text in position_text2.items():
pos_data_x, pos_data_y = missing_data.loc[country]
plt.annotate(country, xy=(pos_data_x, pos_data_y), xytext=pos_text,
arrowprops=dict(facecolor='black', width=0.5, shrink=0.1, headwidth=5))
plt.plot(pos_data_x, pos_data_y, "rs")
X=np.linspace(0, 110000, 1000)
plt.plot(X, t0 + t1*X, "b:")
lin_reg_full = linear_model.LinearRegression()
Xfull = np.c_[full_country_stats["GDP per capita"]]
yfull = np.c_[full_country_stats["Life satisfaction"]]
lin_reg_full.fit(Xfull, yfull)
t0full, t1full = lin_reg_full.intercept_[0], lin_reg_full.coef_[0][0]
X = np.linspace(0, 110000, 1000)
plt.plot(X, t0full + t1full * X, "k")
plt.xlabel("GDP per capita (USD)")
save_fig('representative_training_data_scatterplot')
plt.show()
# +
full_country_stats.plot(kind='scatter', x="GDP per capita", y='Life satisfaction', figsize=(8,3))
plt.axis([0, 110000, 0, 10])
from sklearn import preprocessing
from sklearn import pipeline
poly = preprocessing.PolynomialFeatures(degree=30, include_bias=False)
scaler = preprocessing.StandardScaler()
lin_reg2 = linear_model.LinearRegression()
pipeline_reg = pipeline.Pipeline([('poly', poly), ('scal', scaler), ('lin', lin_reg2)])
pipeline_reg.fit(Xfull, yfull)
curve = pipeline_reg.predict(X[:, np.newaxis])
plt.plot(X, curve)
plt.xlabel("GDP per capita (USD)")
save_fig('overfitting_model_plot')
plt.show()
# -
full_country_stats.loc[[c for c in full_country_stats.index if "W" in c.upper()]]["Life satisfaction"]
gdp_per_capita.loc[[c for c in gdp_per_capita.index if "W" in c.upper()]].head()
# +
plt.figure(figsize=(8,3))
plt.xlabel("GDP per capita")
plt.ylabel('Life satisfaction')
plt.plot(list(sample_data["GDP per capita"]), list(sample_data["Life satisfaction"]), "bo")
plt.plot(list(missing_data["GDP per capita"]), list(missing_data["Life satisfaction"]), "rs")
X = np.linspace(0, 110000, 1000)
plt.plot(X, t0full + t1full * X, "r--", label="Linear model on all data")
plt.plot(X, t0 + t1*X, "b:", label="Linear model on partial data")
ridge = linear_model.Ridge(alpha=10**9.5)
Xsample = np.c_[sample_data["GDP per capita"]]
ysample = np.c_[sample_data["Life satisfaction"]]
ridge.fit(Xsample, ysample)
t0ridge, t1ridge = ridge.intercept_[0], ridge.coef_[0][0]
plt.plot(X, t0ridge + t1ridge * X, "b", label="Regularized linear model on partial data")
plt.legend(loc="lower right")
plt.axis([0, 110000, 0, 10])
plt.xlabel("GDP per capita (USD)")
save_fig('ridge_model_plot')
plt.show()
# -
|
01_the_machine_learning_landscape.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
EVCのためのEV-GMMを構築します. そして, 適応学習する.
詳細 : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580aabf534c4dbb8bc.pdf
This program make EV-GMM for EVC. Then, it make adaptation learning.
Check detail : https://pdfs.semanticscholar.org/cbfe/71798ded05fb8bf8674580abf534c4dbb8bc.pdf
"""
# +
from __future__ import division, print_function
import os
from shutil import rmtree
import argparse
import glob
import pickle
import time
import numpy as np
from numpy.linalg import norm
from sklearn.decomposition import PCA
from sklearn.mixture import GMM # sklearn 0.20.0から使えない
from sklearn.preprocessing import StandardScaler
import scipy.signal
import scipy.sparse
# %matplotlib inline
import matplotlib.pyplot as plt
import IPython
from IPython.display import Audio
import soundfile as sf
import wave
import pyworld as pw
import librosa.display
from dtw import dtw
import warnings
warnings.filterwarnings('ignore')
# +
"""
Parameters
__Mixtured : GMM混合数
__versions : 実験セット
__convert_source : 変換元話者のパス
__convert_target : 変換先話者のパス
"""
# parameters
__Mixtured = 40
__versions = 'pre-stored0.1.2'
__convert_source = 'input/EJM10/V01/T01/TIMIT/000/*.wav'
__convert_target = 'adaptation/EJF01/V01/T01/ATR503/A/*.wav'
# settings
__same_path = './utterance/' + __versions + '/'
__output_path = __same_path + 'output/EJF01/' # EJF01, EJF07, EJM04, EJM05
Mixtured = __Mixtured
pre_stored_pickle = __same_path + __versions + '.pickle'
pre_stored_source_list = __same_path + 'pre-source/**/V01/T01/**/*.wav'
pre_stored_list = __same_path + "pre/**/V01/T01/**/*.wav"
#pre_stored_target_list = "" (not yet)
pre_stored_gmm_init_pickle = __same_path + __versions + '_init-gmm.pickle'
pre_stored_sv_npy = __same_path + __versions + '_sv.npy'
save_for_evgmm_covarXX = __output_path + __versions + '_covarXX.npy'
save_for_evgmm_covarYX = __output_path + __versions + '_covarYX.npy'
save_for_evgmm_fitted_source = __output_path + __versions + '_fitted_source.npy'
save_for_evgmm_fitted_target = __output_path + __versions + '_fitted_target.npy'
save_for_evgmm_weights = __output_path + __versions + '_weights.npy'
save_for_evgmm_source_means = __output_path + __versions + '_source_means.npy'
for_convert_source = __same_path + __convert_source
for_convert_target = __same_path + __convert_target
converted_voice_npy = __output_path + 'sp_converted_' + __versions
converted_voice_wav = __output_path + 'sp_converted_' + __versions
mfcc_save_fig_png = __output_path + 'mfcc3dim_' + __versions
f0_save_fig_png = __output_path + 'f0_converted' + __versions
converted_voice_with_f0_wav = __output_path + 'sp_f0_converted' + __versions
# +
EPSILON = 1e-8
class MFCC:
"""
MFCC() : メル周波数ケプストラム係数(MFCC)を求めたり、MFCCからスペクトルに変換したりするクラス.
動的特徴量(delta)が実装途中.
ref : http://aidiary.hatenablog.com/entry/20120225/1330179868
"""
def __init__(self, frequency, nfft=1026, dimension=24, channels=24):
"""
各種パラメータのセット
nfft : FFTのサンプル点数
frequency : サンプリング周波数
dimension : MFCC次元数
channles : メルフィルタバンクのチャンネル数(dimensionに依存)
fscale : 周波数スケール軸
filterbankl, fcenters : フィルタバンク行列, フィルタバンクの頂点(?)
"""
self.nfft = nfft
self.frequency = frequency
self.dimension = dimension
self.channels = channels
self.fscale = np.fft.fftfreq(self.nfft, d = 1.0 / self.frequency)[: int(self.nfft / 2)]
self.filterbank, self.fcenters = self.melFilterBank()
def hz2mel(self, f):
"""
周波数からメル周波数に変換
"""
return 1127.01048 * np.log(f / 700.0 + 1.0)
def mel2hz(self, m):
"""
メル周波数から周波数に変換
"""
return 700.0 * (np.exp(m / 1127.01048) - 1.0)
def melFilterBank(self):
"""
メルフィルタバンクを生成する
"""
fmax = self.frequency / 2
melmax = self.hz2mel(fmax)
nmax = int(self.nfft / 2)
df = self.frequency / self.nfft
dmel = melmax / (self.channels + 1)
melcenters = np.arange(1, self.channels + 1) * dmel
fcenters = self.mel2hz(melcenters)
indexcenter = np.round(fcenters / df)
indexstart = np.hstack(([0], indexcenter[0:self.channels - 1]))
indexstop = np.hstack((indexcenter[1:self.channels], [nmax]))
filterbank = np.zeros((self.channels, nmax))
for c in np.arange(0, self.channels):
increment = 1.0 / (indexcenter[c] - indexstart[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexstart[c], indexcenter[c])):
filterbank[c, i] = (i - indexstart[c]) * increment
decrement = 1.0 / (indexstop[c] - indexcenter[c])
# np,int_ は np.arangeが[0. 1. 2. ..]となるのをintにする
for i in np.int_(np.arange(indexcenter[c], indexstop[c])):
filterbank[c, i] = 1.0 - ((i - indexcenter[c]) * decrement)
return filterbank, fcenters
def mfcc(self, spectrum):
"""
スペクトルからMFCCを求める.
"""
mspec = []
mspec = np.log10(np.dot(spectrum, self.filterbank.T))
mspec = np.array(mspec)
return scipy.fftpack.realtransforms.dct(mspec, type=2, norm="ortho", axis=-1)
def delta(self, mfcc):
"""
MFCCから動的特徴量を求める.
現在は,求める特徴量フレームtをt-1とt+1の平均としている.
"""
mfcc = np.concatenate([
[mfcc[0]],
mfcc,
[mfcc[-1]]
]) # 最初のフレームを最初に、最後のフレームを最後に付け足す
delta = None
for i in range(1, mfcc.shape[0] - 1):
slope = (mfcc[i+1] - mfcc[i-1]) / 2
if delta is None:
delta = slope
else:
delta = np.vstack([delta, slope])
return delta
def imfcc(self, mfcc, spectrogram):
"""
MFCCからスペクトルを求める.
"""
im_sp = np.array([])
for i in range(mfcc.shape[0]):
mfcc_s = np.hstack([mfcc[i], [0] * (self.channels - self.dimension)])
mspectrum = scipy.fftpack.idct(mfcc_s, norm='ortho')
# splrep はスプライン補間のための補間関数を求める
tck = scipy.interpolate.splrep(self.fcenters, np.power(10, mspectrum))
# splev は指定座標での補間値を求める
im_spectrogram = scipy.interpolate.splev(self.fscale, tck)
im_sp = np.concatenate((im_sp, im_spectrogram), axis=0)
return im_sp.reshape(spectrogram.shape)
def trim_zeros_frames(x, eps=1e-7):
"""
無音区間を取り除く.
"""
T, D = x.shape
s = np.sum(np.abs(x), axis=1)
s[s < 1e-7] = 0.
return x[s > eps]
# +
def analyse_by_world_with_harverst(x, fs):
"""
WORLD音声分析合成器で基本周波数F0,スペクトル包絡,非周期成分を求める.
基本周波数F0についてはharvest法により,より精度良く求める.
"""
# 4 Harvest with F0 refinement (using Stonemask)
frame_period = 5
_f0_h, t_h = pw.harvest(x, fs, frame_period=frame_period)
f0_h = pw.stonemask(x, _f0_h, t_h, fs)
sp_h = pw.cheaptrick(x, f0_h, t_h, fs)
ap_h = pw.d4c(x, f0_h, t_h, fs)
return f0_h, sp_h, ap_h
def wavread(file):
"""
wavファイルから音声トラックとサンプリング周波数を抽出する.
"""
wf = wave.open(file, "r")
fs = wf.getframerate()
x = wf.readframes(wf.getnframes())
x = np.frombuffer(x, dtype= "int16") / 32768.0
wf.close()
return x, float(fs)
def preEmphasis(signal, p=0.97):
"""
MFCC抽出のための高域強調フィルタ.
波形を通すことで,高域成分が強調される.
"""
return scipy.signal.lfilter([1.0, -p], 1, signal)
def alignment(source, target, path):
"""
タイムアライメントを取る.
target音声をsource音声の長さに合うように調整する.
"""
# ここでは814に合わせよう(targetに合わせる)
# p_p = 0 if source.shape[0] > target.shape[0] else 1
#shapes = source.shape if source.shape[0] > target.shape[0] else target.shape
shapes = source.shape
align = np.array([])
for (i, p) in enumerate(path[0]):
if i != 0:
if j != p:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
else:
temp = np.array(target[path[1][i]])
align = np.concatenate((align, temp), axis=0)
j = p
return align.reshape(shapes)
# -
"""
pre-stored学習のためのパラレル学習データを作る。
時間がかかるため、利用できるlearn-data.pickleがある場合はそれを利用する。
それがない場合は一から作り直す。
"""
timer_start = time.time()
if os.path.exists(pre_stored_pickle):
print("exist, ", pre_stored_pickle)
with open(pre_stored_pickle, mode='rb') as f:
total_data = pickle.load(f)
print("open, ", pre_stored_pickle)
print("Load pre-stored time = ", time.time() - timer_start , "[sec]")
else:
source_mfcc = []
#source_data_sets = []
for name in sorted(glob.iglob(pre_stored_source_list, recursive=True)):
print(name)
x, fs = sf.read(name)
f0, sp, ap = analyse_by_world_with_harverst(x, fs)
mfcc = MFCC(fs)
source_mfcc_temp = mfcc.mfcc(sp)
#source_data = np.hstack([source_mfcc_temp, mfcc.delta(source_mfcc_temp)]) # static & dynamic featuers
source_mfcc.append(source_mfcc_temp)
#source_data_sets.append(source_data)
total_data = []
i = 0
_s_len = len(source_mfcc)
for name in sorted(glob.iglob(pre_stored_list, recursive=True)):
print(name, len(total_data))
x, fs = sf.read(name)
f0, sp, ap = analyse_by_world_with_harverst(x, fs)
mfcc = MFCC(fs)
target_mfcc = mfcc.mfcc(sp)
dist, cost, acc, path = dtw(source_mfcc[i%_s_len], target_mfcc, dist=lambda x, y: norm(x - y, ord=1))
#print('Normalized distance between the two sounds:' + str(dist))
#print("target_mfcc = {0}".format(target_mfcc.shape))
aligned = alignment(source_mfcc[i%_s_len], target_mfcc, path)
#target_data_sets = np.hstack([aligned, mfcc.delta(aligned)]) # static & dynamic features
#learn_data = np.hstack((source_data_sets[i], target_data_sets))
learn_data = np.hstack([source_mfcc[i%_s_len], aligned])
total_data.append(learn_data)
i += 1
with open(pre_stored_pickle, 'wb') as output:
pickle.dump(total_data, output)
print("Make, ", pre_stored_pickle)
print("Make pre-stored time = ", time.time() - timer_start , "[sec]")
# +
"""
全事前学習出力話者からラムダを推定する.
ラムダは適応学習で変容する.
"""
S = len(total_data)
D = int(total_data[0].shape[1] / 2)
print("total_data[0].shape = ", total_data[0].shape)
print("S = ", S)
print("D = ", D)
timer_start = time.time()
if os.path.exists(pre_stored_gmm_init_pickle):
print("exist, ", pre_stored_gmm_init_pickle)
with open(pre_stored_gmm_init_pickle, mode='rb') as f:
initial_gmm = pickle.load(f)
print("open, ", pre_stored_gmm_init_pickle)
print("Load initial_gmm time = ", time.time() - timer_start , "[sec]")
else:
initial_gmm = GMM(n_components = Mixtured, covariance_type = 'full')
initial_gmm.fit(np.vstack(total_data))
with open(pre_stored_gmm_init_pickle, 'wb') as output:
pickle.dump(initial_gmm, output)
print("Make, ", initial_gmm)
print("Make initial_gmm time = ", time.time() - timer_start , "[sec]")
weights = initial_gmm.weights_
source_means = initial_gmm.means_[:, :D]
target_means = initial_gmm.means_[:, D:]
covarXX = initial_gmm.covars_[:, :D, :D]
covarXY = initial_gmm.covars_[:, :D, D:]
covarYX = initial_gmm.covars_[:, D:, :D]
covarYY = initial_gmm.covars_[:, D:, D:]
fitted_source = source_means
fitted_target = target_means
# +
"""
SVはGMMスーパーベクトルで、各pre-stored学習における出力話者について平均ベクトルを推定する。
GMMの学習を見てみる必要があるか?
"""
timer_start = time.time()
if os.path.exists(pre_stored_sv_npy):
print("exist, ", pre_stored_sv_npy)
sv = np.load(pre_stored_sv_npy)
print("open, ", pre_stored_sv_npy)
print("Load pre_stored_sv time = ", time.time() - timer_start , "[sec]")
else:
sv = []
for i in range(S):
gmm = GMM(n_components = Mixtured, params = 'm', init_params = '', covariance_type = 'full')
gmm.weights_ = initial_gmm.weights_
gmm.means_ = initial_gmm.means_
gmm.covars_ = initial_gmm.covars_
gmm.fit(total_data[i])
sv.append(gmm.means_)
sv = np.array(sv)
np.save(pre_stored_sv_npy, sv)
print("Make pre_stored_sv time = ", time.time() - timer_start , "[sec]")
# +
"""
各事前学習出力話者のGMM平均ベクトルに対して主成分分析(PCA)を行う.
PCAで求めた固有値と固有ベクトルからeigenvectorsとbiasvectorsを作る.
"""
timer_start = time.time()
#source_pca
source_n_component, source_n_features = sv[:, :, :D].reshape(S, Mixtured*D).shape
# 標準化(分散を1、平均を0にする)
source_stdsc = StandardScaler()
# 共分散行列を求める
source_X_std = source_stdsc.fit_transform(sv[:, :, :D].reshape(S, Mixtured*D))
# PCAを行う
source_cov = source_X_std.T @ source_X_std / (source_n_component - 1)
source_W, source_V_pca = np.linalg.eig(source_cov)
print(source_W.shape)
print(source_V_pca.shape)
# データを主成分の空間に変換する
source_X_pca = source_X_std @ source_V_pca
print(source_X_pca.shape)
#target_pca
target_n_component, target_n_features = sv[:, :, D:].reshape(S, Mixtured*D).shape
# 標準化(分散を1、平均を0にする)
target_stdsc = StandardScaler()
#共分散行列を求める
target_X_std = target_stdsc.fit_transform(sv[:, :, D:].reshape(S, Mixtured*D))
#PCAを行う
target_cov = target_X_std.T @ target_X_std / (target_n_component - 1)
target_W, target_V_pca = np.linalg.eig(target_cov)
print(target_W.shape)
print(target_V_pca.shape)
# データを主成分の空間に変換する
target_X_pca = target_X_std @ target_V_pca
print(target_X_pca.shape)
eigenvectors = source_X_pca.reshape((Mixtured, D, S)), target_X_pca.reshape((Mixtured, D, S))
source_bias = np.mean(sv[:, :, :D], axis=0)
target_bias = np.mean(sv[:, :, D:], axis=0)
biasvectors = source_bias.reshape((Mixtured, D)), target_bias.reshape((Mixtured, D))
print("Do PCA time = ", time.time() - timer_start , "[sec]")
# +
"""
声質変換に用いる変換元音声と目標音声を読み込む.
"""
timer_start = time.time()
source_mfcc_for_convert = []
source_sp_for_convert = []
source_f0_for_convert = []
source_ap_for_convert = []
fs_source = None
for name in sorted(glob.iglob(for_convert_source, recursive=True)):
print("source = ", name)
x_source, fs_source = sf.read(name)
f0_source, sp_source, ap_source = analyse_by_world_with_harverst(x_source, fs_source)
mfcc_source = MFCC(fs_source)
#mfcc_s_tmp = mfcc_s.mfcc(sp)
#source_mfcc_for_convert = np.hstack([mfcc_s_tmp, mfcc_s.delta(mfcc_s_tmp)])
source_mfcc_for_convert.append(mfcc_source.mfcc(sp_source))
source_sp_for_convert.append(sp_source)
source_f0_for_convert.append(f0_source)
source_ap_for_convert.append(ap_source)
target_mfcc_for_fit = []
target_f0_for_fit = []
target_ap_for_fit = []
for name in sorted(glob.iglob(for_convert_target, recursive=True)):
print("target = ", name)
x_target, fs_target = sf.read(name)
f0_target, sp_target, ap_target = analyse_by_world_with_harverst(x_target, fs_target)
mfcc_target = MFCC(fs_target)
#mfcc_target_tmp = mfcc_target.mfcc(sp_target)
#target_mfcc_for_fit = np.hstack([mfcc_t_tmp, mfcc_t.delta(mfcc_t_tmp)])
target_mfcc_for_fit.append(mfcc_target.mfcc(sp_target))
target_f0_for_fit.append(f0_target)
target_ap_for_fit.append(ap_target)
# 全部numpy.arrrayにしておく
source_data_mfcc = np.array(source_mfcc_for_convert)
source_data_sp = np.array(source_sp_for_convert)
source_data_f0 = np.array(source_f0_for_convert)
source_data_ap = np.array(source_ap_for_convert)
target_mfcc = np.array(target_mfcc_for_fit)
target_f0 = np.array(target_f0_for_fit)
target_ap = np.array(target_ap_for_fit)
print("Load Input and Target Voice time = ", time.time() - timer_start , "[sec]")
# +
"""
適応話者学習を行う.
つまり,事前学習出力話者から目標話者の空間を作りだす.
適応話者文数ごとにfitted_targetを集めるのは未実装.
"""
timer_start = time.time()
epoch=100
py = GMM(n_components = Mixtured, covariance_type = 'full')
py.weights_ = weights
py.means_ = target_means
py.covars_ = covarYY
fitted_target = None
for i in range(len(target_mfcc)):
print("adaptation = ", i+1, "/", len(target_mfcc))
target = target_mfcc[i]
for x in range(epoch):
print("epoch = ", x)
predict = py.predict_proba(np.atleast_2d(target))
y = np.sum([predict[:, i: i + 1] * (target - biasvectors[1][i])
for i in range(Mixtured)], axis = 1)
gamma = np.sum(predict, axis = 0)
left = np.sum([gamma[i] * np.dot(eigenvectors[1][i].T,
np.linalg.solve(py.covars_, eigenvectors[1])[i])
for i in range(Mixtured)], axis=0)
right = np.sum([np.dot(eigenvectors[1][i].T,
np.linalg.solve(py.covars_, y)[i])
for i in range(Mixtured)], axis = 0)
weight = np.linalg.solve(left, right)
fitted_target = np.dot(eigenvectors[1], weight) + biasvectors[1]
py.means_ = fitted_target
print("Load Input and Target Voice time = ", time.time() - timer_start , "[sec]")
# -
"""
変換に必要なものを残しておく.
"""
np.save(save_for_evgmm_covarXX, covarXX)
np.save(save_for_evgmm_covarYX, covarYX)
np.save(save_for_evgmm_fitted_source, fitted_source)
np.save(save_for_evgmm_fitted_target, fitted_target)
np.save(save_for_evgmm_weights, weights)
np.save(save_for_evgmm_source_means, source_means)
|
old-exp/adapt/adapt1/make-evgmm0.1.2-EJF01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 64-bit
# name: python38264bit903daaff972d42a981642e2de4b1a84b
# ---
# + tags=[]
# Beyond Bitswap Test Bed
# Be able to create a yaml file custom.
import ipywidgets as widgets
from IPython.display import display
print("Use case configuration")
testcase = widgets.Text(description="Testcase")
input_data = widgets.Text(description="Input Data Type")
file_size = widgets.Text(description="File Size")
files_directory = widgets.Text(description="Files Directory")
run_count = widgets.IntSlider(description="Run Count", min=0, max=300)
display(testcase, input_data, file_size, files_directory, run_count)
print("Network configuration")
n_nodes = widgets.IntSlider(description="Number nodes", min=0, max=300)
n_leechers = widgets.IntSlider(description="Number leechers", min=0, max=300)
n_passive = widgets.IntSlider(description="Number passive nodes", min=0, max=300)
max_peer_connections = widgets.FloatSlider(description="Max peer connections (%)", min=0, max=100)
churn_rate = widgets.IntSlider(description="Churn Rate (%)", min=0, max=100)
display(n_nodes, n_leechers, n_passive, max_peer_connections, churn_rate)
# + tags=[]
import ui
l = ui.Layout()
l.show()
#display(l.testcase, l.input_data, l.file_size, l.files_directory, l.run_count, \
# l.n_nodes, l.n_leechers, l.n_passive, l.max_peer_connections, l.churn_rate)
# + tags=[]
# Building config and running testcase
import utils
#testid = utils.runner(utils.process_yaml_config("./config.yaml"))
testid = utils.runner(utils.process_layout_config(l))
print(testid)
# + tags=[]
# Collecting the data.
utils.collect_data(testid)
# + tags=[]
import process
agg, testcases = process.aggregate_results()
byLatency = process.groupBy(agg, "latencyMS")
byNodeType = process.groupBy(agg, "nodeType")
byFileSize = process.groupBy(agg, "fileSize")
byBandwidth = process.groupBy(agg, "bandwidthMB")
byTopology = process.groupBy(agg, "topology")
# -
process.plot_latency(byLatency, byBandwidth, byFileSize)
process.plot_messages(byFileSize, byTopology)
process.plot_bw_overhead(byFileSize, byTopology)
process.plot_througput(byLatency, byBandwidth, byFileSize, byTopology, testcases)
|
beyond-bitswap/scripts/dashboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Importing libraries</h1>
from __future__ import print_function
import keras
from matplotlib import pyplot as plt
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
import numpy as np
from sklearn.model_selection import train_test_split
import pandas as pd
# <h1>importing dataset</h1>
data=pd.read_csv('fmnist.csv')
# <h1>Reshaping the dataset</h1>
# +
# input image dimensions
img_rows, img_cols = 28, 28
n_instances=10000
#number of intances can effect performance and time in training
X = data.iloc[0:n_instances,1:].values
Y = data.iloc[0:n_instances,0].values
test_size=0.20##test size
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=test_size,random_state=3)
xx = np.expand_dims(x_train, axis=0)
xt = np.expand_dims(x_test, axis=0)
x_train = xx.reshape(len(x_train),28,28,1)##reshaping dataset into 4 dimensions
x_test = xt.reshape(len(x_test),28,28,1)##reshaping dataset into 4 dimensions
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255 ##dividing each pixel by 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
num_classes=10
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# -
# <h1>Fitting model</h1>
# +
##activation is the activation function in each layer
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
batch_size = 128
#The batch size is a hyperparameter that defines the number of samples to work through before updating
#the internal model parameters.
epochs = 3 ##a number of epochs means how many times you go through your training set.
##more epochs can get more accuracy but it will need more time and performance
#By setting verbose 0, 1 or 2 you just say how do you want to 'see' the training progress for each epoch.
history=model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
# -
# <h1>Accuracy results</h1>
# +
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# summarize history for accuracy
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
# <h1>Image Results</h1>
pred=model.predict(x_test)
numberof_images_display=5
n= np.random.randint(0,len(x_test),numberof_images_display)
for i in n:
two_d = (np.reshape(x_test[i], (28, 28)) * 255).astype(np.uint8)
print("Predicted value: ",np.argmax(pred[i],axis=None,out=None)) ##from binary to real value
print("Real value: ",np.argmax(y_test[i], axis=None, out=None))
plt.imshow(two_d, interpolation='nearest',cmap='gray')
plt.show()
|
Pattern recognition II/tutorial 6/Code/CNN fmnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import glob
import os
import sys
from deep_utils import dump_pickle, load_pickle
import time
from itertools import chain
from argparse import ArgumentParser
import torch
from pretrainedmodels.utils import ToRange255
from pretrainedmodels.utils import ToSpaceBGR
from scipy.spatial.distance import cdist
from torch.utils.data import DataLoader
from torch.utils.data.dataloader import default_collate
from torchvision import transforms
from data.inshop import InShop
from metric_learning.util import SimpleLogger
from metric_learning.sampler import ClassBalancedBatchSampler
from PIL import Image
import metric_learning.modules.featurizer as featurizer
import metric_learning.modules.losses as losses
import numpy as np
from evaluation.retrieval import evaluate_float_binary_embedding_faiss, _retrieve_knn_faiss_gpu_inner_product
from PIL import Image
import matplotlib.pyplot as plt
def adjust_learning_rate(optimizer, epoch, epochs_per_step, gamma=0.1):
"""Sets the learning rate to the initial LR decayed by 10 every epochs"""
# Skip gamma update on first epoch.
if epoch != 0 and epoch % epochs_per_step == 0:
for param_group in optimizer.param_groups:
param_group['lr'] *= gamma
print("learning rate adjusted: {}".format(param_group['lr']))
# +
dataset = "InShop"
dataset_root = ""
batch_size = 64
model_name = "resnet50"
lr = 0.01
gamma = 0.1
class_balancing = True
images_per_class = 5
lr_mult = 1
dim = 2048
test_every_n_epochs = 2
epochs_per_step = 4
pretrain_epochs = 1
num_steps = 3
output = "data1/output"
create_pkl = False
model_path = '/home/ai/projects/symo/classification_metric_learning/data1/output/InShop/2048/resnet50_75/epoch_30.pth'
# +
def get_most_similar(feature, features_dict, n=10, distance='cosine'):
features = list(features_dict.values())
ids = list(features_dict.keys())
p = cdist(np.array(features),
np.expand_dims(feature, axis=0),
metric=distance)[:, 0]
group = zip(p, ids.copy())
res = sorted(group, key=lambda x: x[0])
r = res[:n]
return r
def extract_feature(model, loader, gpu_device):
"""
Extract embeddings from given `model` for given `loader` dataset on `gpu_device`.
"""
model.eval()
model.to(gpu_device)
db_dict = {}
log_every_n_step = 10
with torch.no_grad():
for i, (im, class_label, instance_label, index) in enumerate(loader):
im = im.to(device=gpu_device)
embedding = model(im)
for i,em in zip(index, embedding):
db_dict[loader.dataset.image_paths[int(i)]] = em.detach().cpu().numpy()
if (i + 1) % log_every_n_step == 0:
print('Process Iteration {} / {}:'.format(i, len(loader)))
dump_pickle('db.pkl', db_dict)
return db_dict
# -
def main(query_img):
torch.cuda.set_device(0)
gpu_device = torch.device('cuda')
output_directory = os.path.join(output, dataset, str(dim),
'_'.join([model_name, str(batch_size)]))
if not os.path.exists(output_directory):
os.makedirs(output_directory)
out_log = os.path.join(output_directory, "train.log")
sys.stdout = SimpleLogger(out_log, sys.stdout)
# Select model
model_factory = getattr(featurizer, model_name)
model = model_factory(dim)
weights = torch.load(model_path)
model.load_state_dict(weights)
eval_transform = transforms.Compose([
transforms.Resize((256, 256)),
transforms.CenterCrop(max(model.input_size)),
transforms.ToTensor(),
ToSpaceBGR(model.input_space == 'BGR'),
ToRange255(max(model.input_range) == 255),
transforms.Normalize(mean=model.mean, std=model.std)
])
# Setup dataset
# train_dataset = InShop('../data1/data/inshop', transform=train_transform)
query_dataset = InShop('data1/data/inshop', train=False, query=True, transform=eval_transform)
index_dataset = InShop('data1/data/inshop', train=False, query=False, transform=eval_transform)
query_loader = DataLoader(query_dataset,
batch_size=batch_size,
drop_last=False,
shuffle=False,
pin_memory=True,
num_workers=0)
model.to(device='cuda')
model.eval()
query_image = Image.open(query_img).convert('RGB')
with torch.no_grad():
query_image = model(eval_transform(query_image).to('cuda').unsqueeze(0))[0].cpu().numpy()
index_dataset = InShop('data1/data/inshop', train=False, query=False, transform=eval_transform)
index_loader = DataLoader(index_dataset,
batch_size=75,
drop_last=False,
shuffle=False,
pin_memory=True,
num_workers=0)
if create_pkl:
db_list = extract_feature(model, index_loader, 'cuda')
else:
db_list = load_pickle('db.pkl')
return get_most_similar(query_image, db_list)
def visualize(query_img, images):
img = Image.open(query_img)
plt.imshow(img)
plt.title('main_image')
plt.show()
for score, img_path in images:
img = Image.open(img_path)
plt.imshow(img)
plt.title(str(score))
plt.show()
query_img = "/home/ai/Pictures/im3.png"
images = main(query_img)
visualize(query_img, images)
|
demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: venv
# ---
# +
# Small scripts to play with data and perform simple tasks
# You can play with various datasets from here: https://archive.ics.uci.edu/ml/index.php
# +
# Import necessary libraries
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as stats
import seaborn as sns
from matplotlib import rcParams
pd.set_option('float_format', '{:f}'.format)
# -
# Read data using pandas
df = pd.read_csv('./data/kc_house_data.csv')
# Print dimensions
print(df.shape)
# Print a few rows
df.head()
# +
# Other way to read file, using numpy
from numpy import genfromtxt
# Read 2 specific columns
data = genfromtxt('./data/kc_house_data.csv',delimiter=',',usecols=(2,5))
# Print dimensions
print(f'Dimensions: {data.shape}\n')
# Print data
print('price\tsqft_living')
for i in range(1,5):
print(np.array2string(data[i], formatter={'float_kind':'{0:.1f}'.format}))
# -
# Continue with the dataframe
# Check to see if there are any null values in the data
df.isnull().any()
# Check out the data types
df.dtypes
# Get a summary of the data
df.describe()
# Describe a specific attribute
df['price'].describe()
# +
# Some comments:
# We are working with a data set that contains 21,613 observations (see count)
# Mean price is approximately $540k
# Median price is approximately $450k
# Average house area is ~ 2080 ft2
# +
# Let's plot some histograms
fig = plt.figure(figsize=(24, 6))
sqft = fig.add_subplot(121)
cost = fig.add_subplot(122)
sqft.hist(df.sqft_living, bins=80)
sqft.set_xlabel('Ft^2')
sqft.set_title("Histogram of House Square Footage")
cost.hist(df.price, bins=80)
cost.set_xlabel('Price ($)')
cost.set_title("Histogram of Housing Prices")
# +
# Observation:
# Both variables have a distribution that is right-skewed.
# +
# Let's do some regression analysis
from sklearn.linear_model import LinearRegression
from numpy import linspace, matrix
linreg = LinearRegression()
prices = np.array(list(df['price']))
prices = prices.reshape(-1,1)
area = np.array(list(df['sqft_living']))
area = area.reshape(-1,1)
linreg.fit(area, prices)
# Plot outputs
plt.scatter(area, prices, color = 'red')
plt.plot(area, linreg.predict(area), color = 'blue')
plt.title('Area vs Prices')
plt.xlabel('Area')
plt.ylabel('Price')
plt.show()
# -
# +
# Generate synthetic data examples
from faker import Faker
fake = Faker()
name = fake.name()
address = fake.address()
print(f'Random name:\n{name}\n')
print(f'Random address:\n{address}')
# +
# continue with synthetic data
job = fake.job()
num = fake.pyint()
phone = fake.phone_number()
name_f = fake.first_name_female()
email = fake.ascii_email()
geo = fake.latlng()
date = fake.date()
company = fake.company()
print(f'Random job: {job}')
print(f'Random number: {num}')
print(f'Random phone: {phone}')
print(f'Random female name: {name_f}')
print(f'Random email: {email}')
print(f'Random geolocation: {geo}')
print(f'Random date: {date}')
print(f'Random company: {company}')
# +
# What about GR synthetic data?
fake = Faker('el_GR')
name = fake.name()
address = fake.address()
print(f'Random name:\n{name}\n')
print(f'Random address:\n{address}\n')
job = fake.job()
num = fake.pyint()
phone = fake.phone_number()
name_f = fake.first_name_female()
email = fake.ascii_email()
geo = fake.latlng()
date = fake.date()
company = fake.company()
print(f'Random job: {job}')
print(f'Random phone: {phone}')
print(f'Random female name: {name_f}')
print(f'Random company: {company}')
# +
# Play with SQL
import sqlite3
# Create a connection
conn = sqlite3.connect('./data/example.db')
# Create a table
conn.execute('''CREATE TABLE EMPLOYEE
(ID INT PRIMARY KEY NOT NULL,
NAME TEXT NOT NULL,
AGE INT NOT NULL,
ADDRESS CHAR(50),
SALARY REAL);''')
# Close connection
conn.close()
# +
# Insert some records
conn = sqlite3.connect('./data/example.db')
conn.execute("INSERT INTO EMPLOYEE (ID,NAME,AGE,ADDRESS,SALARY) \
VALUES (1, 'Bob', 32, 'California', 20000.00 )");
conn.execute("INSERT INTO EMPLOYEE (ID,NAME,AGE,ADDRESS,SALARY) \
VALUES (2, 'Alice', 25, 'Texas', 15000.00 )");
conn.execute("INSERT INTO EMPLOYEE (ID,NAME,AGE,ADDRESS,SALARY) \
VALUES (3, 'Joe', 23, 'Norway', 20000.00 )");
conn.execute("INSERT INTO EMPLOYEE (ID,NAME,AGE,ADDRESS,SALARY) \
VALUES (4, 'Mary', 25, 'Rich-Mond ', 65000.00 )");
conn.commit()
conn.close()
# +
# Perform query
conn = sqlite3.connect('./data/example.db')
cursor = conn.execute("SELECT id, name, address, salary from EMPLOYEE")
for row in cursor:
print(f'ID: {row[0]}')
print(f'NAME: {row[1]}')
print(f'ADDRESS: {row[2]}')
print(f'SALARY: {row[3]}')
print()
conn.close()
# +
# Perform update
conn = sqlite3.connect('./data/example.db')
conn.execute("UPDATE EMPLOYEE set SALARY = 30000.00 where ID = 1")
conn.commit()
print(f'Total number of rows updated: {conn.total_changes}')
# Check the updated result
cursor = conn.execute("SELECT id, name, address, salary from EMPLOYEE")
for row in cursor:
print(f'ID: {row[0]}')
print(f'NAME: {row[1]}')
print(f'ADDRESS: {row[2]}')
print(f'SALARY: {row[3]}')
print()
conn.close()
# +
# Perform delete
conn = sqlite3.connect('./data/example.db')
conn.execute("DELETE from EMPLOYEE where ID = 2;")
conn.commit()
print(f'Total number of rows updated: {conn.total_changes}')
# Check the updated result
cursor = conn.execute("SELECT id, name, address, salary from EMPLOYEE")
for row in cursor:
print(f'ID: {row[0]}')
print(f'NAME: {row[1]}')
print(f'ADDRESS: {row[2]}')
print(f'SALARY: {row[3]}')
print()
conn.close()
# +
# Accessing SQL records with pandas
conn = sqlite3.connect('./data/example.db')
df = pd.read_sql_query("SELECT * from EMPLOYEE", conn)
conn.close()
df
# +
# Then, query the dataframe
# Example: get the persons with salary >= 25000
df[df.SALARY >= 25000]
# -
|
playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create Cases in Batch and Run in Parallel
#
# This notebook demonstrates creating cases in batch and running them in parallel.
# ## Create Cases in Batch
#
# The approach to create cases in batch following this procedure:
#
# - Load the base case from file
# - For each desired case output:
# * Alter parameters to the desired value
# * Save each system to a new case file
# +
import andes
import numpy as np
from andes.utils.paths import get_case
andes.config_logger()
# -
# create directory for output cases
# !rm -rf batch_cases
# !mkdir -p batch_cases
kundur = get_case('kundur/kundur_full.xlsx')
ss = andes.load(kundur)
# We demonstrate running the Kundur's system under different loading conditions.
#
# Cases are created by modifying the `p0` of `PQ` with `idx == PQ_0`.
# As always, input parameters can be inspected in `cache.df_in`.
p0_base = ss.PQ.get('p0', "PQ_0")
# Create 10 cases so that the load increases from `p0_base` to `1.2 * p0_base`.
p0_values = np.linspace(p0_base, 1.2 * p0_base, 10)
for value in p0_values:
ss.PQ.alter('p0', 'PQ_0', value)
file_name = f'batch_cases/kundur_p_{value:.2f}.xlsx'
andes.io.dump(ss, 'xlsx', file_name, overwrite=True)
# ## Parallel Simulation
# Parallel simulation is easy with the command line tool.
#
# Change directory to `batch_cases`:
# +
import os
# change the Python working directory
os.chdir('batch_cases')
# -
# !ls -la
# ### Running from Command line
# !andes run *.xlsx -r tds
# ### Number of CPUs
# In some cases, you don't want the simulatino to use up all resources.
#
# ANDES allows to control the number of processes to run in parallel through `--ncpu NCPU`, where `NCPU` is the maximum number of processes (equivalent to the number of CPU cores) allowed.
# !andes run *.xlsx -r tds --ncpu 4
# ### Running with APIs
# Setting `pool = True` allows returning all system instances in a list.
#
# This comes with a penalty in computation time but can be helpful if you want to extract data directly.
systems = andes.run('*.xlsx', routine='tds', pool=True, verbose=10)
systems[0]
systems
# ### Example plots
#
# Plotting or data analyses can be carried out as usual.
ss = systems[0]
systems[0].TDS.plotter.plot(ss.GENROU.omega, latex=False)
systems[5].TDS.plotter.plot(ss.GENROU.omega, latex=False)
|
examples/7. parallel-simulation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="C_Vi5KyGEA2t"
# -*- coding: utf-8 -*-
Sbox = [
["63", "7C", "77", "7B", "F2", "6B", "6F", "C5", "30", "01", "67", "2B", "FE", "D7", "AB", "76"],
["CA", "82", "C9", "7D", "FA", "59", "47", "F0", "AD", "D4", "A2", "AF", "9C", "A4", "72", "C0"],
["B7", "FD", "93", "26", "36", "3F", "F7", "CC", "34", "A5", "E5", "F1", "71", "D8", "31", "15"],
["04", "C7", "23", "C3", "18", "96", "05", "9A", "07", "12", "80", "E2", "EB", "27", "B2", "75"],
["09", "83", "2C", "1A", "1B", "6E", "5A", "A0", "52", "3B", "D6", "B3", "29", "E3", "2F", "84"],
["53", "D1", "00", "ED", "20", "FC", "B1", "5B", "6A", "CB", "BE", "39", "4A", "4C", "58", "CF"],
["D0", "EF", "AA", "FB", "43", "4D", "33", "85", "45", "F9", "02", "7F", "50", "3C", "9F", "A8"],
["51", "A3", "40", "8F", "92", "9D", "38", "F5", "BC", "B6", "DA", "21", "10", "FF", "F3", "D2"],
["CD", "0C", "13", "EC", "5F", "97", "44", "17", "C4", "A7", "7E", "3D", "64", "5D", "19", "73"],
["60", "81", "4F", "DC", "22", "2A", "90", "88", "46", "EE", "B8", "14", "DE", "5E", "0B", "DB"],
["E0", "32", "3A", "0A", "49", "06", "24", "5C", "C2", "D3", "AC", "62", "91", "95", "E4", "79"],
["E7", "C8", "37", "6D", "8D", "D5", "4E", "A9", "6C", "56", "F4", "EA", "65", "7A", "AE", "08"],
["BA", "78", "25", "2E", "1C", "A6", "B4", "C6", "E8", "DD", "74", "1F", "4B", "BD", "8B", "8A"],
["70", "3E", "B5", "66", "48", "03", "F6", "0E", "61", "35", "57", "B9", "86", "C1", "1D", "9E"],
["E1", "F8", "98", "11", "69", "D9", "8E", "94", "9B", "1E", "87", "E9", "CE", "55", "28", "DF"],
["8C", "A1", "89", "0D", "BF", "E6", "42", "68", "41", "99", "2D", "0F", "B0", "54", "BB", "16"]
]
Sbox_inv = [
["52", "09", "6A", "D5", "30", "36", "A5", "38", "BF", "40", "A3", "9E", "81", "F3", "D7", "FB"],
["7C", "E3", "39", "82", "9B", "2F", "FF", "87", "34", "8E", "43", "44", "C4", "DE", "E9", "CB"],
["54", "7B", "94", "32", "A6", "C2", "23", "3D", "EE", "4C", "95", "0B", "42", "FA", "C3", "4E"],
["08", "2E", "A1", "66", "28", "D9", "24", "B2", "76", "5B", "A2", "49", "6D", "8B", "D1", "25"],
["72", "F8", "F6", "64", "86", "68", "98", "16", "D4", "A4", "5C", "CC", "5D", "65", "B6", "92"],
["6C", "70", "48", "50", "FD", "ED", "B9", "DA", "5E", "15", "46", "57", "A7", "8D", "9D", "84"],
["90", "D8", "AB", "00", "8C", "BC", "D3", "0A", "F7", "E4", "58", "05", "B8", "B3", "45", "06"],
["D0", "2C", "1E", "8F", "CA", "3F", "0F", "02", "C1", "AF", "BD", "03", "01", "13", "8A", "6B"],
["3A", "91", "11", "41", "4F", "67", "DC", "EA", "97", "F2", "CF", "CE", "F0", "B4", "E6", "73"],
["96", "AC", "74", "22", "E7", "AD", "35", "85", "E2", "F9", "37", "E8", "1C", "75", "DF", "6E"],
["47", "F1", "1A", "71", "1D", "29", "C5", "89", "6F", "B7", "62", "0E", "AA", "18", "BE", "1B"],
["FC", "56", "3E", "4B", "C6", "D2", "79", "20", "9A", "DB", "C0", "FE", "78", "CD", "5A", "F4"],
["1F", "DD", "A8", "33", "88", "07", "C7", "31", "B1", "12", "10", "59", "27", "80", "EC", "5F"],
["60", "51", "7F", "A9", "19", "B5", "4A", "0D", "2D", "E5", "7A", "9F", "93", "C9", "9C", "EF"],
["A0", "E0", "3B", "4D", "AE", "2A", "F5", "B0", "C8", "EB", "BB", "3C", "83", "53", "99", "61"],
["17", "2B", "04", "7E", "BA", "77", "D6", "26", "E1", "69", "14", "63", "55", "21", "0C", "7D"]
]
rows, cols = 4, 4
"""# Encryption"""
def text2hex(s):
s = s.encode('utf-8')
return s.hex()
def hex2text(h):
return bytes.fromhex(h).decode('utf-8')
#make a matrix with 00 in the blocks as an initial matrix
def get2dMatrix(rows, cols):
Matrix = [[0 for x in range(cols)] for y in range(rows)]
return Matrix
#hex values to assign in matrix each block 2 hex value
def hexToMatrixAssign(my_matrix, my_hex):
th = my_hex
lenth = len(th)
cnt = 0
for i in range(rows):
for j in range(cols):
if cnt >= lenth:
my_matrix[j][i] = "00"
cnt +=2
else:
my_matrix[j][i] = th[cnt]+th[cnt+1]
# print(f"{j}{i} = {th[cnt]+th[cnt+1]}")
cnt +=2
return my_matrix
def printMatrix(my_matrix, rows, cols):
for i in range(rows):
for j in range(cols):
print(f"{my_matrix[i][j]} ",end = '')
print()
def hex2Binary(hexadecimal):
# hexadecimal = "54776f20"
end_length = len(hexadecimal) * 4
hex_as_int = int(hexadecimal, 16)
hex_as_binary = bin(hex_as_int)
padded_binary = hex_as_binary[2:].zfill(end_length)
return padded_binary
def binary2Hex(binary_no):
return hex(int(binary_no, 2)).replace("0x","")
def _Xor(a, b):
a = hex2Binary(a)
b = hex2Binary(b)
y = int(a,2) ^ int(b,2)
y = '{0:b}'.format(y)
y = binary2Hex(str(y))
if len(y) == 1:
y = "0"+y
return y
def xorIn2matrix(matrix_1, matrix_2, rows, cols):
n_matrix = get2dMatrix(rows, cols)
for i in range(rows):
for j in range(cols):
n_matrix[i][j] = _Xor(matrix_1[i][j], matrix_2[i][j])
return n_matrix
def _1shift(my_matrix, rows, cols):
temp = my_matrix[0][0]
for i in range(cols):
my_matrix[0][i] = my_matrix[0][i+1]
if i+2 == cols:
break
my_matrix[0][cols-1] = temp
return my_matrix
def _2shift(my_matrix, rows, cols):
temp = my_matrix[1][0]
temp1 = my_matrix[1][1]
for i in range(cols):
my_matrix[1][i] = my_matrix[1][i+2]
if i+3 == cols:
break
my_matrix[1][cols-1] = temp1
my_matrix[1][cols-2] = temp
return my_matrix
def getFromSbox(x):
a = 0
b = 0
if ord(x[0]) >= 48 and ord(x[0]) <= 75:
a = ord(x[0]) -48
elif ord(x[0]) >= 97 and ord(x[0]) <= 122:
a = ord(x[0]) -87
else:
a = int(x[0])
if ord(x[1]) >= 48 and ord(x[1]) <= 57:
b = ord(x[1]) -48
elif ord(x[1]) >= 97 and ord(x[1]) <= 122:
b = ord(x[1]) -87
else:
b = int(x[1])
return Sbox[a][b].lower()
# 3rd row s-box implementation
def _3sbox(my_matrix, rows, cols):
for i in range(cols):
my_matrix[2][i] = getFromSbox(my_matrix[2][i])
return my_matrix
#last row execution
def _4addbit(my_matrix, rows, cols):
temp = my_matrix[3][0]
temp = hex2Binary(temp)
temp = bin(int(temp,2) + int("01",2)).replace("0b","")
temp = binary2Hex(temp)
my_matrix[3][0] = temp
return my_matrix
def encryptData(myText, myKey1, no_round):
#plain text
t_matrix = 0
#key
k_matrix = 0
th = text2hex(myText)
kh = text2hex(myKey1)
rows, cols = 4, 4
# cols = getNoOfColumn(len(myText), len(myKey1))
cipherText = get2dMatrix(rows, cols)
cipherText = hexToMatrixAssign(cipherText, th)
# printMatrix(cipherText, rows, cols)
cipherKey = get2dMatrix(rows, cols)
cipherKey = hexToMatrixAssign(cipherKey, kh)
# printMatrix(cipherKey, rows, cols)
printMatrix(cipherText, 4, 4)
print("")
printMatrix(cipherKey, 4, 4)
print("")
roundCount = 0
#total 11 rounds
for i in range(no_round+1):
t_matrix = xorIn2matrix(cipherText, cipherKey, rows, cols)
if i <=9:
k_matrix = _1shift(cipherKey, rows, cols)
k_matrix = _2shift(cipherKey, rows, cols)
k_matrix = _3sbox(cipherKey, rows, cols)
if i < 9:
k_matrix = _4addbit(cipherKey, rows, cols)
#_________________________________________________________
roundCount +=1
print(f"#Round {roundCount}")
print("---------------------------")
printMatrix(t_matrix, 4, 4)
print()
printMatrix(k_matrix, 4, 4)
print("\n\n")
#_________________________________________________________
cipherText = t_matrix
cipherKey = k_matrix
return t_matrix, k_matrix
"""# Decryption"""
def HexMatrixToText(my_matrix, rows, cols):
mystr = ""
for i in range(rows):
for j in range(cols):
mystr += my_matrix[j][i]
return hex2text(mystr)
def decryptData(t_matrix, k_matrix, no_round):
roundCount = no_round+1
kpo = ""
for i in range(no_round):
#key 10 will remain the same
if i > 0:
#changes after that
if i > 1:
kpo = d_4addbit(k_matrix, rows, cols)
kpo = d_3sbox(k_matrix, rows, cols)
kpo = _2shift(k_matrix, rows, cols)
for i in range(3):
kpo = _1shift(k_matrix, rows, cols)
else:
kpo = k_matrix
# tpo = xorIn2matrix(kpo, t_matrix, rows, cols)
tpo = xorIn2matrix(kpo, t_matrix, 4, 4)
# roundCount -=1
# print(f"#Round {roundCount}")
# print("---------------------------")
# printMatrix(tpo, 4, 4)
# print()
# printMatrix(kpo, 4, 4)
# print("\n\n")
k_matrix = kpo
t_matrix = tpo
actualText = HexMatrixToText(t_matrix, rows, cols)
return t_matrix, k_matrix, actualText.replace("\x00", "")
#4th row subtraction
def d_4addbit(my_matrix, rows, cols):
temp = my_matrix[3][0]
temp = hex2Binary(temp)
temp = bin(int(temp,2) - int("01",2)).replace("0b","")
temp = binary2Hex(temp)
my_matrix[3][0] = temp
return my_matrix
#inverse s-box define
def dgetFromSbox(x):
a = 0
b = 0
if ord(x[0]) >= 48 and ord(x[0]) <= 57:
a = ord(x[0]) -48
elif ord(x[0]) >= 97 and ord(x[0]) <= 122:
a = ord(x[0]) -87
else:
a = int(x[0])
if ord(x[1]) >= 48 and ord(x[1]) <= 57:
b = ord(x[1]) -48
elif ord(x[1]) >= 97 and ord(x[1]) <= 122:
b = ord(x[1]) -87
else:
b = int(x[1])
return Sbox_inv[a][b].lower()
#assign inverse s-box
def d_3sbox(my_matrix, rows, cols):
for i in range(cols):
my_matrix[2][i] = dgetFromSbox(my_matrix[2][i])
return my_matrix
# + id="100ykCgXywAs" outputId="2ddfa590-ca52-4dd2-9f74-0ed35802f7ef" colab={"base_uri": "https://localhost:8080/"}
"""# Main Work For Encryption"""
myText = "Two One Nine Two"
myKey1 = "Thats my Kung Fu"
keyLen = len(myKey1)
textLen = len(myText)
if keyLen > 16:
print("Key Length Is Exceed 128 Bits")
else:
#start assigning key in matrix from 0 to 15
listText = []
cn = 0
temp = ""
for i in range(textLen):
temp += myText[i]
cn +=1
if cn == 16 or i == textLen-1:
listText.append(temp)
temp = ""
cn = 0
#print key 10 and main cipher text
for x in listText:
t_matrix, k_matrix = encryptData(x, myKey1, 10)
print(f"Text = [{x}] ---------------------------T/K")
print()
printMatrix(t_matrix, 4, 4)
print()
printMatrix(k_matrix, 4, 4)
print("\n\n")
# + id="HWT8_bWINEVG" outputId="68339e85-52b7-4ca0-88a7-ade610d544a1" colab={"base_uri": "https://localhost:8080/"}
#decryption process output
enText = [
["73", "1c", "09", "13"],
["54", "6e", "4a", "54"],
["c1", "6e", "02", "50"],
["21", "20", "65", "6f"]
]
enKey = [
["20", "67", "54", "73"],
["68", "20", "4b", "20"],
["26", "10", "19", "34"],
["7d", "79", "6e", "75"]
]
#11th round execution
dt_matrix, dk_matrix, actualText = decryptData(t_matrix, k_matrix, 11)
print(actualText)
|
Test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + cell_id="00000-20ec9206-9165-45bb-86ba-5f55ad92a2ef" deepnote_cell_type="code"
with open('input.txt', 'r') as fp:
data = fp.read()
# + cell_id="00001-056d8c61-f272-490a-abb3-ce101dee014d" deepnote_cell_type="code"
data
# + cell_id="00002-5a17fc82-0e5f-4b84-afce-371118f8fddf" deepnote_cell_type="code"
result=0
it = 0
for char in data:
it += 1
if char=="(":
result=result+1
else:
result=result-1
# + cell_id="00003-6eae5865-04e7-4300-ba29-2eeaf3fbaaff" deepnote_cell_type="code"
result
# + cell_id="00004-57df87b6-daa5-41a1-b607-9df9af7e0780" deepnote_cell_type="code"
result=0
it = 0
for char in data:
it += 1
if char=="(":
result=result+1
else:
result=result-1
if result==-1:
print(it)
break
# + cell_id="00005-5f8886f5-6b86-4db4-8cbf-039817b548d1" deepnote_cell_type="code"
sum([1 if char == '(' else -1 for char in data])
# + [markdown] tags=[] created_in_deepnote_cell=true deepnote_cell_type="markdown"
# <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=978e47b7-a961-4dca-a945-499e8b781a34' target="_blank">
# <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODBweCIgaGVpZ2h0PSI4MHB4IiB2aWV3Qm94PSIwIDAgODAgODAiIHZlcnNpb249IjEuMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayI+CiAgICA8IS0tIEdlbmVyYXRvcjogU2tldGNoIDU0LjEgKDc2NDkwKSAtIGh0dHBzOi8vc2tldGNoYXBwLmNvbSAtLT4KICAgIDx0aXRsZT5Hcm91cCAzPC90aXRsZT4KICAgIDxkZXNjPkNyZWF0ZWQgd2l0aCBTa2V0Y2guPC9kZXNjPgogICAgPGcgaWQ9IkxhbmRpbmciIHN0cm9rZT0ibm9uZSIgc3Ryb2tlLXdpZHRoPSIxIiBmaWxsPSJub25lIiBmaWxsLXJ1bGU9ImV2ZW5vZGQiPgogICAgICAgIDxnIGlkPSJBcnRib2FyZCIgdHJhbnNmb3JtPSJ0cmFuc2xhdGUoLTEyMzUuMDAwMDAwLCAtNzkuMDAwMDAwKSI+CiAgICAgICAgICAgIDxnIGlkPSJHcm91cC0zIiB0cmFuc2Zvcm09InRyYW5zbGF0ZSgxMjM1LjAwMDAwMCwgNzkuMDAwMDAwKSI+CiAgICAgICAgICAgICAgICA8cG9seWdvbiBpZD0iUGF0aC0yMCIgZmlsbD0iIzAyNjVCNCIgcG9pbnRzPSIyLjM3NjIzNzYyIDgwIDM4LjA0NzY2NjcgODAgNTcuODIxNzgyMiA3My44MDU3NTkyIDU3LjgyMTc4MjIgMzIuNzU5MjczOSAzOS4xNDAyMjc4IDMxLjY4MzE2ODMiPjwvcG9seWdvbj4KICAgICAgICAgICAgICAgIDxwYXRoIGQ9Ik0zNS4wMDc3MTgsODAgQzQyLjkwNjIwMDcsNzYuNDU0OTM1OCA0Ny41NjQ5MTY3LDcxLjU0MjI2NzEgNDguOTgzODY2LDY1LjI2MTk5MzkgQzUxLjExMjI4OTksNTUuODQxNTg0MiA0MS42NzcxNzk1LDQ5LjIxMjIyODQgMjUuNjIzOTg0Niw0OS4yMTIyMjg0IEMyNS40ODQ5Mjg5LDQ5LjEyNjg0NDggMjkuODI2MTI5Niw0My4yODM4MjQ4IDM4LjY0NzU4NjksMzEuNjgzMTY4MyBMNzIuODcxMjg3MSwzMi41NTQ0MjUgTDY1LjI4MDk3Myw2Ny42NzYzNDIxIEw1MS4xMTIyODk5LDc3LjM3NjE0NCBMMzUuMDA3NzE4LDgwIFoiIGlkPSJQYXRoLTIyIiBmaWxsPSIjMDAyODY4Ij48L3BhdGg+CiAgICAgICAgICAgICAgICA8cGF0aCBkPSJNMCwzNy43MzA0NDA1IEwyNy4xMTQ1MzcsMC4yNTcxMTE0MzYgQzYyLjM3MTUxMjMsLTEuOTkwNzE3MDEgODAsMTAuNTAwMzkyNyA4MCwzNy43MzA0NDA1IEM4MCw2NC45NjA0ODgyIDY0Ljc3NjUwMzgsNzkuMDUwMzQxNCAzNC4zMjk1MTEzLDgwIEM0Ny4wNTUzNDg5LDc3LjU2NzA4MDggNTMuNDE4MjY3Nyw3MC4zMTM2MTAzIDUzLjQxODI2NzcsNTguMjM5NTg4NSBDNTMuNDE4MjY3Nyw0MC4xMjg1NTU3IDM2LjMwMzk1NDQsMzcuNzMwNDQwNSAyNS4yMjc0MTcsMzcuNzMwNDQwNSBDMTcuODQzMDU4NiwzNy43MzA0NDA1IDkuNDMzOTE5NjYsMzcuNzMwNDQwNSAwLDM3LjczMDQ0MDUgWiIgaWQ9IlBhdGgtMTkiIGZpbGw9IiMzNzkzRUYiPjwvcGF0aD4KICAgICAgICAgICAgPC9nPgogICAgICAgIDwvZz4KICAgIDwvZz4KPC9zdmc+' > </img>
# Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
|
members/eszter/Kalo_AoC.1/AoC.1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
india_weather = pd.DataFrame({
"city":["Mumbai","Pune","Banglore"],
"temperature":[7,12,20],
"humidity":[3,5,1],
})
india_weather
usa_weather = pd.DataFrame({
"city":["New York","Chicago","Los Angles"],
"temperature":[17,22,30],
"humidity":[13,10,15],
})
usa_weather
idt_us = pd.concat([india_weather,usa_weather])
idt_us
idt_us = pd.concat([india_weather,usa_weather],ignore_index=True)
idt_us
idt_us = pd.concat([india_weather,usa_weather],keys=["India","USA"])
idt_us
idt_us.loc["USA"]
temperature = pd.DataFrame({
"city":["Mumbai","Pune","Banglore"],
"temperature":[7,12,20]
})
temperature
humidity = pd.DataFrame({
"city":["Mumbai","Pune","Banglore"],
"humidity":[13,10,15]
})
humidity
df = pd.concat([temperature,humidity])
df
df = pd.concat([temperature,humidity],axis=1)
df
temperature = pd.DataFrame({
"city":["Mumbai","Pune","Banglore"],
"temperature":[7,12,20]
})
temperature
humidity = pd.DataFrame({
"city":["Pune","Mumbai"],
"humidity":[13,10]
})
humidity
df = pd.concat([temperature,humidity],axis=1)
df
temperature = pd.DataFrame({
"city":["Mumbai","Pune","Banglore"],
"temperature":[7,12,20]
},index = [0,1,2])
temperature
humidity = pd.DataFrame({
"city":["Pune","Mumbai"],
"humidity":[13,10]
}, index = [1,0])
humidity
df = pd.concat([temperature,humidity],axis=1)
df
s = pd.Series(["Humid","Sunny","Rain"],name="Event")
s
df = pd.concat([temperature,s],axis=1)
df
|
path_of_ML/Pandas/Concat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Elementarströmungen 2
# Wir haben bereits die Translationsströmung, Quellen- und Senkenströmung sowie deren Überlagerung kennengelernt. Im zweiten Teil der Elementarströmungen kommen jetzt noch zwei wichtige hinzu: die Dipolströmung und der Potentialwirbel.
# ### Dipolströmung
#
# Die Dipolströmung ist ein Sonderfall der Überlagerung einer Quellen- und einer Senkenströmung mit gleicher Stärke $Q$, mit verschwindendem Abstand $\Delta x$. Dabei wird beim Grenzübergang $\Delta x \rightarrow 0$ das Produkt $M=Q\cdot\Delta x$, das sog. **Dipolmoment**, konstant gehalten, indem $Q$ umgekehrt proportional erhöht wird. Die resultierende Potentialfunktion wird dann:
#
# $$\phi = \frac{M}{2\pi}\cdot \lim_{\Delta x\rightarrow 0} \left( \frac{\ln\sqrt{(x+\Delta x)^2+y^2}-\ln\sqrt{x^2+y^2} } {\Delta x} \right)$$
#
# $$\phi = \frac{M}{2\pi}\cdot \frac{x}{x^2+y^2}$$
#
# Mit der gleichen Vorgehensweise erhält man die Stromfunktion der Dipolströmung:
#
# $$\psi = \frac{M}{2\pi}\cdot\lim_{\Delta x\rightarrow 0}\left(\frac{\text{arctan} \frac{y}{x} - \text{arctan} \frac{y}{x+\Delta x}}{\Delta x}\right)$$
#
# $$\psi = -\frac{M}{2\pi}\cdot \frac{y}{x^2+y^2}$$
#
# Durch Ableitung der Potential- oder der Stromfunktion erhält man schließlich die Geschwindigkeitskomponenten der Dipolströmung:
#
# $$u = -\frac{M}{2\pi} \cdot \frac{x^2-y^2}{(x^2+y^2)^2}$$
# $$v = -\frac{M}{2\pi} \cdot \frac{2xy}{(x^2+y^2)^2}$$
# Für die Implementierung in Python importieren wir wieder die benötigten Bibliotheken und definieren die benötigten Variablen:
# +
import math
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
nx = 400 # Anzahl der Punkte in x-Richtung
ny = 200 # Anzahl der Punkte in y-Richtung
x = np.linspace(-5, 15, nx) # 1D-Array mit x-Koordinaten
y = np.linspace(-5, 5, ny) # 1D-Array mit y-Koordinaten
X, Y = np.meshgrid(x, y) # erzeugt das Gitter mit nx * ny Punkten
# -
# Die oben gezeigten Gleichungen für die Geschwindigkeitskomponenten sowie die Potential- und Stromfunktionen implementieren wir als Funktionen:
# +
def dipolx_v(x, y, xs, ys, M): # der Geschwindigkeitsvek. der Quellenstr.
s = -M/(2*math.pi) / ((x-xs)**2+(y-ys)**2)**2
return s*((x-xs)**2-(y-ys)**2), s*2*(x-xs)*(y-ys)
def dipolx_psi(x, y, xs, ys, M): # die Stromfunktion der Quellenströmung
return -M/(2*math.pi) * (y-ys) / ((x-xs)**2+(y-ys)**2)
def dipolx_phi(x, y, xs, ys, M): # die Potentialfunktion der Quellenströmung
return M/(2*math.pi) * (x-xs) / ((x-xs)**2+(y-ys)**2)
# -
# Damit ist alles bereit, um die Strom- und Potentiallinien der Dipolströmung zu berechnen und diese zu zeichnen:
# +
M = 20.0 # Dipolmoment in m^3/s
u_dipol, v_dipol = dipolx_v(X, Y, 0, 0, M) # Dipolströmung bei (0,0)
psi_dipol = dipolx_psi(X, Y, 0, 0, M)
phi_dipol = dipolx_phi(X, Y, 0, 0, M)
plt.figure(figsize=(10, 5))
plt.xlabel('x')
plt.ylabel('y')
plt.streamplot(X, Y, u_dipol, v_dipol,
density=2, linewidth=1, arrowsize=2, arrowstyle='->')
plt.contour(X, Y, phi_dipol,
[-1.5, -0.5, -0.3, -0.2, -0.1, 0.1, 0.2, 0.3, 0.5, 1.5],
colors='green', linewidths=1, linestyles='solid')
#plt.contour(X, Y, psi_dipol,
# [-1.5, -0.2, -0.05, -0.01, 0.01, 0.05, 0.2, 1.5],
# colors='#407eb4', linewidths=1, linestyles='solid')
plt.scatter(0, 0, color='red', s=50, marker='o', linewidth=0);
# -
# Die Dipolströmung für sich alleine betrachtet ist nicht besonders sinnvoll. Wenn wir Sie jedoch mit einer Translationsströmung überlagern, erhalten wir die Umströmung eines Zylinders.
#
# Wir müssen hierzu nur die bereits zuvor definierten Funktionen für die Translationsströmung übernehmen:
# +
def trans_v(x, y, u1, v1): # der Geschwindigkeitsvek. der Translationsstr.
return np.full_like(x, u1), np.full_like(y, v1)
def trans_psi(x, y, u1, v1): # die Stromfunktion der Translationsströmung
return -v1*x+u1*y
def trans_phi(x, y, u1, v1): # die Potentialfunktion der Translationsströmung
return u1*x+v1*y
# -
# und diese mit der Dipolströmung überlagern:
# +
u1 = 0.3
u_trans, v_trans = trans_v(X, Y, u1, 0) # Translationsströmung
u_gesamt = u_dipol + u_trans # lineare Überlagerung
v_gesamt = v_dipol + v_trans
psi_trans = trans_psi(X, Y, u1, 0.0)
psi_gesamt = psi_dipol + psi_trans
phi_trans = trans_phi(X, Y, u1, 0.0)
phi_gesamt = phi_dipol + phi_trans
# -
# und schließlich grafisch darstellen:
# +
# Neuen Plot einrichten
plt.figure(figsize=(10, 5))
plt.xlabel('x')
plt.ylabel('y')
# Stromlinien mit Matplotlib-Funktion darstellen
plt.streamplot(X, Y, u_gesamt, v_gesamt,
density=2, linewidth=1, arrowsize=2, arrowstyle='->')
# Staustromlinie in rot eintragen
plt.contour(X, Y, psi_gesamt, levels=[0],
colors='red', linewidths=1, linestyles='solid');
# -
# ### Potentialwirbel (drehungsfreier Wirbel)
#
# Unter einem Potentialwirbel versteht man eine Strömung, auf der die Fluidelemente auf kreisförmigen Bahnen um einen zentralen Punkt umlaufen und sich dabei nicht um die eigene Achse drehen. Man spricht dann von einem drehungsfreien Wirbel.
#
# 
#
# Beim Potentialwirbel ist die Radialkomponente der Geschwindigkeit gleich Null:
#
# $$u_r = 0$$
#
# Die Tangentialkomponente erhalten wir durch die Erfüllung der Drehungsfreiheit
#
# $$\text{rot}\overrightarrow{v} =
# \begin{pmatrix}
# \frac{\partial w}{\partial y}-\frac{\partial v}{\partial z} \\
# \frac{\partial u}{\partial z}-\frac{\partial w}{\partial x} \\
# \frac{\partial v}{\partial x}-\frac{\partial u}{\partial y}
# \end{pmatrix}_{x,y,z}
# =
# \begin{pmatrix}
# \frac{1}{r}\frac{\partial u_z}{\partial \varphi}-\frac{\partial u_\varphi}{\partial z} \\
# \frac{\partial u_r}{\partial z}-\frac{\partial u_z}{\partial r} \\
# \frac{1}{r}\frac{\partial (ru_\varphi)}{\partial r}-\frac{1}{r}\frac{\partial u_r}{\partial \varphi}
# \end{pmatrix}_{r,\varphi,z} = 0,$$
#
# die hier auch in Zylinderkoordinaten gegeben ist, da die Herleitung der tangentialien Geschwindigkeitskomponente des Potentialwirbels in Zylinderkoordinaten einfacher ist. Im Zweidimensionalen sind die $z$-Komponente der Geschwindigkeit und alle Ableitungen nach $z$ Null und es muss nur die folgende Gleichung erfüllt werden:
#
# $$\frac{\partial (ru_\varphi)}{\partial r}-\frac{\partial u_r}{\partial \varphi} = 0$$
#
# Da der Potentialwirbel axialsymmetrisch ist, ändert sich die Geschwindigkeit in Umfangsrichtung nicht, so dass $\frac{\partial u_r}{\partial \varphi} = 0$ ist und sich damit die Forderung der Drehungsfreiheit auf
#
# $$\frac{\partial (ru_\varphi)}{\partial r} = 0$$
#
# reduziert. Die Gleichung ist erfüllt, wenn $ru_\varphi = const$ ist. Wobei es üblich ist, die Konstante durch die [Zirkulation](https://de.wikipedia.org/wiki/Zirkulation_%28Feldtheorie%29) $\Gamma$ auszudrücken und bei positiver Zirkulation eine Drehrichtung des Wirbels im Uhrzeigersinn zu definieren. Damit sind die Geschwindigkeitskomponenten des Potentialwirbels bekannt:
#
# $$u_\varphi = -\frac{\Gamma}{2\pi r}, \qquad u_r = 0$$
#
# bzw. in kartesischen Koordinaten:
#
# $$u_x = \frac{\Gamma}{2\pi}\frac{y}{x^2+y^2}, \qquad u_y = -\frac{\Gamma}{2\pi}\frac{x}{x^2+y^2}$$
#
# Durch Integration der Geschwindigkeitskomponenten finden wir wieder die Potential- und Stromfunktionen (vgl. Herleitung der Quellenströmung):
#
# $$\phi = -\frac{\Gamma}{2\pi}\text{arctan}\left(\frac{y}{x}\right), \qquad \psi = \frac{\Gamma}{2\pi} \text{ln}\sqrt{x^2+y^2}$$
#
#
# Als nächstes sollen die Strom- und Potentiallinien des Potentialwirbels dargestellt werden. Dazu definieren wir wieder entsprechende Python-Funktionen:
# +
def vortex_v(x, y, x1, y1, Gamma): # Geschwindigkeitsvek. des Potentialwirbels
s = Gamma/(2*math.pi*((x-x1)**2+(y-y1)**2))
return s*(y-y1), -s*(x-x1)
def vortex_psi(x, y, x1, y1, Gamma): # Stromfunktion des Potentialwirbels
s = -Gamma/(2*math.pi)
return -s*np.log((x-x1)**2+(y-y1)**2)
def vortex_phi(x, y, x1, y1, Gamma): # Potentialfunktion des Potentialwirbels
s = -Gamma/(2*math.pi)
return np.arctan2((y-y1),(x-x1))
# -
# Und plotten alles wieder mit der Matplotlib:
# +
Gamma = 2
u_vortex, v_vortex = vortex_v(X, Y, 0, 0, Gamma)
phi_vortex = vortex_phi(X, Y, 0, 0, Gamma)
# Neuen Plot einrichten
plt.figure(figsize=(10, 5))
plt.xlabel('x')
plt.ylabel('y')
# Stromlinien mit Matplotlib-Funktion darstellen
plt.streamplot(X, Y, u_vortex, v_vortex,
density=2, linewidth=1, arrowsize=2, arrowstyle='->')
# Potentiallinien in grün einzeichnen
plt.contour(X, Y, phi_vortex,
[-math.pi*7/8, -math.pi*3/4, -math.pi*5/8, -math.pi/2,
-math.pi*3/8, -math.pi/4, -math.pi/8,
0, math.pi/8, math.pi/4, math.pi*3/8, math.pi/2,
math.pi*5/8, math.pi*3/4, math.pi*7/8],
colors='green', linewidths=1, linestyles='solid');
# -
# Der Potentialwirbel ist bis auf sein Zentrum drehungsfrei. D.h. die Zirkulation
#
# $$\Gamma = \oint_C {\overrightarrow{v}(\overrightarrow{r}) \text{d} \overrightarrow{r}}$$
#
# entlang einer beliebigen Kurve $C$ in der Umgebung des Wirbels, die das Zentrum nicht mit einschließt gerade Null ist. Nur wenn das Zentrum innerhalb der geschlossenen Kurve liegt, ist die Zirkulation $\Gamma$ ungleich Null und entspricht dem vorgegebenen Wert.
# ### Übersicht Elementarströmungen
#
# Die folgende Tabelle gibt eine Übersicht, über die zuvor hergeleiteten Elementarströmungen. Zusätzlich ist die Staupunktströmung enthalten, auf die wir im Rahmen der Vorlesung nicht näher eingehen. Die Übersicht enthält die Gleichungen der Geschwindigkeitsvektoren sowie der Potential- und Stromfunktionen in kartesischen sowie Polarkoordinaten.
#
# 
#
# [Hier auch zum Runterladen als PDF](Formelsammlung_Potentialtheorie.pdf)
#
# Im nächsten Kapitel werden wir die Elementarströmungen so überlagern, dass wir auch den Auftrieb von umströmten Körpern berechnen können.
#
# [Hier](2_4-Potentialtheorie_Auftrieb.ipynb) geht's weiter oder [hier](index.ipynb) zurück zur Übersicht.
#
# ---
# ###### Copyright (c) 2018, <NAME> und <NAME>
#
# Der folgende Python-Code darf ignoriert werden. Er dient nur dazu, die richtige Formatvorlage für die Jupyter-Notebooks zu laden.
from IPython.core.display import HTML
def css_styling():
styles = open('TFDStyle.css', 'r').read()
return HTML(styles)
css_styling()
|
4_3-Potentialtheorie_Elementarstroemungen2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importando elementos importantes
# +
import torch
import matplotlib.pyplot as plt
import itertools
from nn.rbf import RBF
from sklearn.datasets import load_boston
from sklearn.metrics import mean_squared_error
# -
# # Carregando a base Boston Housing e Inicializando a RBF
x, y = load_boston(return_X_y=True)
rbf = RBF(x=x, y=y, c=5, gamma=1e-3)
# # Treinamento com 80% das amostras (1a)
thetas = rbf.fit()
y_pred = rbf.predict(thetas=thetas)
# # Erro quadratico medio
rbf.rmse(y_pred=y_pred).item()
# # Definindo variacoes para o numero de centroides e para o gamma
cvalues = [c for c in range(3, 9)]
gvalues = [10**-g for g in range(1, 7)]
count = 0
for c, g in itertools.product(cvalues, gvalues):
separator = '' if (count % 2) == 0 else '|\n'
rbf.set_params(c=c, gamma=g, x=rbf.x_train)
thetas = rbf.fit()
y_pred = rbf.predict(thetas=thetas)
rmse = rbf.rmse(y_pred=y_pred).item()
print(f'| c = {c:02d} | gamma = {g:.6f} | RMSE = {rmse:07.4f} ', end=separator)
count+=1
# # Conclusão
# A definição de centróides para este problema é de suma relevância. Caso os valores escolhidos não sejam valores adequados, o modelo pode convergir para outro problema e caso haja centróides de mais, o modelo pode simplesmente se adaptar de mais (overfitting). Em comparação com o modelo linear feito no exercício anterior, este modelo se sai melhor quando tem seus parâmetros bem ajustados, pois sua conversão é mais rápida e mais precisa. (1b)
#
# Acima podemos ver diversos testes com variações nos valores de C e de GAMMA. O melhor valor obtido foi c=3, gamma=1e-6.
#
# # Autores
# - **<NAME>** @ https://github.com/chrismachado
# - **<NAME>** @ https://github.com/vitorverasm
|
rp-ex4rbf-1-answer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Data Fields Exploration
# ### *This notebook explores the data fields in the USGS Wind Turbine Data Fields*
link_to_metadata = 'https://eerscmap.usgs.gov/uswtdb/assets/data/uswtdb_v1_1_20180710.xml'
link_to_usgs_data = 'https://eerscmap.usgs.gov/uswtdb/assets/data/uswtdbCSV.zip'
import re
import sys
import xml.etree.ElementTree as ET
from urllib.request import urlopen
import pandas as pd
from bs4 import BeautifulSoup as BS
usgs_data = pd.read_csv('./uswtdbCSV/uswtdb_v1_1_20180710.csv')
usgs_data.shape
# * There are 57646 rows in the dataset with 24 columns.
usgs_data.columns
print(usgs_data.head().to_string())
# * To explore the meaning of the columns in the dataset, we will need to traverse the XML Tree of the metadata.
# * I downloaded the raw XML file from the website and saved it in my working directory from [HERE](https://eerscmap.usgs.gov/uswtdb/assets/data/uswtdb_v1_1_20180710.xml).
tree = ET.parse('./uswtdb_v1_1_20180710.xml')
root = tree.getroot()
ean_info = root.find('eainfo')
all_attributes = ean_info.find('detailed')
counter = 0
for elem in all_attributes:
if elem.tag != 'attr':
continue
counter += 1
try:
attr_label = elem.find('attrlabl').text
except:
attr_label = None
try:
attr_def = elem.find('attrdef').text
except:
attr_def = None
try:
attr_defs = elem.find('attrdefs').text
except:
attr_defs = None
try:
attr_domv = elem.find('attrdomv').find('udom').text
except:
try:
attr_domv = elem.find('attrdomv').find('rdom').find('attrunit').text
except:
attr_domv = None
print('#', counter, attr_label)
print('\n')
print('Definition: ')
print(attr_def)
print('\n')
print('Additional Details: ')
print(attr_defs)
print('\n')
print('Attribute Units Explanation: ')
print(attr_domv)
print('- - - - - - - - -')
|
01 - Metadata XML Read and Exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="direction:rtl">با سلام خدمت استاد گرامی این فایل صرفا برای نمایش کد ها و جواب آن ها تهیه شده است در این جا قابلیت اجرای کد ها نیست </div>
# <div style="direction:rtl">این قدم اول یک پروژه است در مرحله اول ما باید یک مدل ساده و ابتدایی طراحی میکردیم سپس نتیجه های به دست آمده را با مدل های گذشته مقایسه کنیم اگر مدل ما ارزش سرمایه گذاری داشته باشد مدل را پیشرفت خواهیم داد</div>
#
#
#
# ## Problem Statement
#
# The max flow problem can be formulated mathematically as a linear programming problem using the following model.
#
# ### Sets
#
# $N$ = nodes in the network
# $A$ = network arcs
#
# ### Parameters
#
# $s$ = source node
# $t$ = sink node
# $c_{ij}$ = arc flow capacity, $\forall (i,j) \in A$
#
# ### Variables
# $f_{i,j}$ = arc flow, $\forall (i,j) \in A$
#
# ### Objective
#
# Maximize the flow into the sink nodes
# $\max \sum_{\{i \mid (i,t) \in A\}} c_{i,t} f_{i,t}$
#
# ### Constraints
#
# Enforce an upper limit on the flow across each arc
# $f_{i,j} \leq c_{i,j}$, $\forall (i,j) \in A$
#
# Enforce flow through each node
# $\sum_{\{i \mid (i,j) \in A\}} f_{i,j} = \sum_{\{i \mid (j,i) \in A\}} f_{j,i}$, $\forall j \in N - \{s,t\}$
#
# Flow lower bound
# $f_{i,j} \geq 0$, $\forall (i,j) \in A$
#
# ## Pyomo Formulation
#
#
# +
from pyomo.environ import *
model = AbstractModel()
# -
# Nodes in the network
model.N = Set()
# Network arcs
model.A = Set(within=model.N*model.N)
# Source node
model.s = Param(within=model.N)
# Sink node
model.t = Param(within=model.N)
# Flow capacity limits
model.c = Param(model.A)
# The flow over each arc
model.f = Var(model.A, within=NonNegativeReals)
# Maximize the flow into the sink nodes
def total_rule(model):
return sum(model.f[i,j] for (i, j) in model.A if j == value(model.t))
model.total = Objective(rule=total_rule, sense=maximize)
# +
# Enforce an upper limit on the flow across each arc
def limit_rule(model, i, j):
return model.f[i,j] <= model.c[i, j]
model.limit = Constraint(model.A, rule=limit_rule)
# Enforce flow through each node
def flow_rule(model, k):
if k == value(model.s) or k == value(model.t):
return Constraint.Skip
inFlow = sum(model.f[i,j] for (i,j) in model.A if j == k)
outFlow = sum(model.f[i,j] for (i,j) in model.A if i == k)
return inFlow == outFlow
model.flow = Constraint(model.N, rule=flow_rule)
# -
# !cat maxflow.py
# ## Model Data
#
# <div style="direction:rtl">یکی از قابلیت های مدل ما این است که داده بعد از تشکیل شدن طرح مدل به آن تزریق میشود و میتوان از شیوه مرسوم برای بقیه نرم افزار ها هم استفاده کرد </div>
# !cat maxflow.dat
#
#
# ## Solution
#
#
# !pyomo solve --solver=glpk maxflow.py maxflow.dat
# By default, the optimization results are stored in the file `results.yml`:
# !cat results.yml
# ## References
#
# * <NAME>, (2002). "On the history of the transportation and maximum flow problems". Mathematical Programming 91 (3): 437–445.
|
maxflow.ipynb
|
# ##### Copyright 2020 The OR-Tools Authors.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # simple_lp_program
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/linear_solver/simple_lp_program.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/ortools/linear_solver/samples/simple_lp_program.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010-2018 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Minimal example to call the GLOP solver."""
# [START program]
# [START import]
from __future__ import print_function
from ortools.linear_solver import pywraplp
# [END import]
# [START solver]
# Create the linear solver with the GLOP backend.
solver = pywraplp.Solver.CreateSolver('GLOP')
# [END solver]
# [START variables]
# Create the variables x and y.
x = solver.NumVar(0, 1, 'x')
y = solver.NumVar(0, 2, 'y')
print('Number of variables =', solver.NumVariables())
# [END variables]
# [START constraints]
# Create a linear constraint, 0 <= x + y <= 2.
ct = solver.Constraint(0, 2, 'ct')
ct.SetCoefficient(x, 1)
ct.SetCoefficient(y, 1)
print('Number of constraints =', solver.NumConstraints())
# [END constraints]
# [START objective]
# Create the objective function, 3 * x + y.
objective = solver.Objective()
objective.SetCoefficient(x, 3)
objective.SetCoefficient(y, 1)
objective.SetMaximization()
# [END objective]
# [START solve]
solver.Solve()
# [END solve]
# [START print_solution]
print('Solution:')
print('Objective value =', objective.Value())
print('x =', x.solution_value())
print('y =', y.solution_value())
# [END print_solution]
|
examples/notebook/linear_solver/simple_lp_program.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Доверительные интервалы для доли
# ## Генерация данных
import numpy as np
# +
np.random.seed(1)
statistical_population = np.random.randint(2, size = 100000)
random_sample = np.random.choice(statistical_population, size = 1000)
# -
#истинное значение доли
statistical_population.mean()
# ## Точечная оценка доли
random_sample.mean()
# ## Доверительный интервал для доли
from statsmodels.stats.proportion import proportion_confint
# ### Доверительный интервал на основе нормального распределения
# $$\hat{p}\pm z_{1-\frac{\alpha}{2}} \sqrt{\frac{\hat{p}\left(1-\hat{p}\right)}{n}}$$
normal_interval = proportion_confint(sum(random_sample), len(random_sample), method = 'normal')
print('normal_interval [%f, %f] with width %f' % (normal_interval[0],
normal_interval[1],
normal_interval[1] - normal_interval[0]))
# ### Доверительный интервал Уилсона
# $$\frac1{ 1 + \frac{z^2}{n} } \left( \hat{p} + \frac{z^2}{2n} \pm z \sqrt{ \frac{ \hat{p}\left(1-\hat{p}\right)}{n} + \frac{
# z^2}{4n^2} } \right), \;\; z \equiv z_{1-\frac{\alpha}{2}}$$
wilson_interval = proportion_confint(sum(random_sample), len(random_sample), method = 'wilson')
print('wilson_interval [%f, %f] with width %f' % (wilson_interval[0],
wilson_interval[1],
wilson_interval[1] - wilson_interval[0]))
# ## Размер выборки для интервала заданной ширины
from statsmodels.stats.proportion import samplesize_confint_proportion
n_samples = int(np.ceil(samplesize_confint_proportion(random_sample.mean(), 0.01)))
n_samples
np.random.seed(1)
random_sample = np.random.choice(statistical_population, size = n_samples)
normal_interval = proportion_confint(sum(random_sample), len(random_sample), method = 'normal')
print('normal_interval [%f, %f] with width %f' % (normal_interval[0],
normal_interval[1],
normal_interval[1] - normal_interval[0]))
|
4. Stats for Data Analysis/Confidence Intervals/proporion_conf_int.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Time: O(n + p)
# Space: O(n + p)
from collections import defaultdict
def min_window(text, ptr):
ascii_text, ascii_ptr = defaultdict(int), defaultdict(int)
for p in ptr: ascii_ptr[p] += 1
n, p = len(text), len(ascii_ptr)
count = start = 0
window = ''
for i in range(n):
ascii_text[text[i]] += 1
if count < p:
if ascii_text[text[i]] == ascii_ptr[text[i]]:
count += 1
if count == p:
while ascii_text[text[start]] > ascii_ptr[text[start]]:
ascii_text[text[start]] -= 1
start += 1
window = min(window, text[start:i + 1], key = len) if window else text[start:i + 1]
return window
if __name__=='__main__':
tc = [["ADOBECODEBANC","ABC"],
["zaaskzaa", "zsk"],
["tutorial","oti"]]
for t, p in tc:
print(min_window(t, p))
# -
|
assignments/array/Minimum Window Substring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 优化算法在机器学习中扮演着至关重要的角色,了解常用的优化算法对于机器学习爱好者和从业者有着重要的意义。
#
# 这系列文章先讲述优化算法和机器学习的关系,然后罗列优化算法分类,尤其是机器学习中常用的几类.接下来明确下数学符号,开始按照历史和逻辑顺序,依次介绍各种机器学习中常用的优化算法.
#
# 这篇先讲其中基于一阶导数的标准梯度下降法和Momentum,其中穿插学习率退火方法和基于二阶导数的优化算法来辅助说明各算法的意义和背后的想法.
#
# # 优化算法和机器学习的关系
#
# 机器学习的过程往往是
#
# - 建模实际问题,定义损失函数
# - 代入训练数据,利用优化算法来优化损失函数并更新参数,直到终止条件(比如迭代数或者更新的收益或者损失函数的大小)
# 可见优化算法和损失函数在机器学习中占有重要的地位.
# # 优化算法分类
#
# 优化算法有很多种,常见的包括
#
# - 基于导数的,比如基于一阶导数的梯度下降法(GD, Grandient Descent)和基于二阶导数的牛顿法等,要求损失函数(运筹学中更多叫做目标函数)可导
# - 群体方法(population method),比如遗传算法(Genetic Algo)和蚁群算法(Ant Colony Optimization),不依赖于问题(problem-independent),不需要对目标函数结构有太多的了解
# - 单体方法(single-state method),比如模拟退火算法(Simulated Annealing),同样,不依赖于问题(problem-independent),不需要对目标函数结构有太多的了解
# 等.
#
# 机器学习中常用的是基于导数,尤其是基于一阶导数的优化算法,包括
#
# - 标准梯度下降法(GD, standard Gradient Descent)
# - 带有momentum的GD
# - RMSProp (Root Mean Square Propagation)
# - AdaM (Adaptive Moment estimates)
# - AdaGrad (Adaptive Gradient Algo)
# - AdaDelta
# # 符号规定
#
# 在具体解释前先规定下符号
# - 损失函数为 $L(x)$ (很多地方也会写作$ J(x)$ )
# - 梯度为 $ g(x) = \frac{\partial L(x)}{\partial x} $
# - $g_t$ 表示第t次迭代的梯度,
# - 第t次迭代时, $x_{t+1} = x_t + \Delta x_t $
# - 学习率为 $\eta $
# - $o(f(x))$ 表示 $f(x)$ 的高阶无穷小,也就是当 $f(x) $无限接近0时, $g(x) = o(f(x)), \lim_{f(x)\to0} \frac{g(x)}{f(x)} = 0 $,比如 $x^2$ 就是 $x$的高阶无穷小
#
# # 标准梯度下降法(GD, standard Gradient Descent)
#
# 每次迭代的更新为
# $\Delta x_t = - \eta* g_t $
#
#
# 标准GD的想法来源于一阶泰勒展开
# $f(x_1) = f(x_0) + f'(x)|_{x=x_0} * (x_1 - x_0) + o(x_1-x_0) $.
#
# 其中 $o(x_1-x_0)$ 叫做皮亚诺(Peano)余项,当 $x_1 - x_0$ 很小时,这个余项可以忽略不计.
# 当 $x_1 - x_0$ 和一阶导数也就是梯度相反方向时, 损失函数下降最快.
#
# 一个经典的解释是:想象我们从山上下来,每步都沿着坡度最陡的方向.这时,水平面是我们的定义域,海拔是值域.
#
# ### GD缺点
# 但GD有两个主要的缺点:
# - 优化过程中,保持一定的学习率,并且这个学习率是人工设定.当学习率过大时,可能在靠近最优点附近震荡(想象一步子太大跨过去了);学习率过小时,优化的速度太慢
# - 学习率对于每个维度都一样,而我们经常会遇到不同维度的曲率(二阶导数)差别比较大的情况,这时GD容易出现zig-zag路径.(参考图2,优化路径呈现zig-zag形状,该图绘制代码放在附录1中)
# 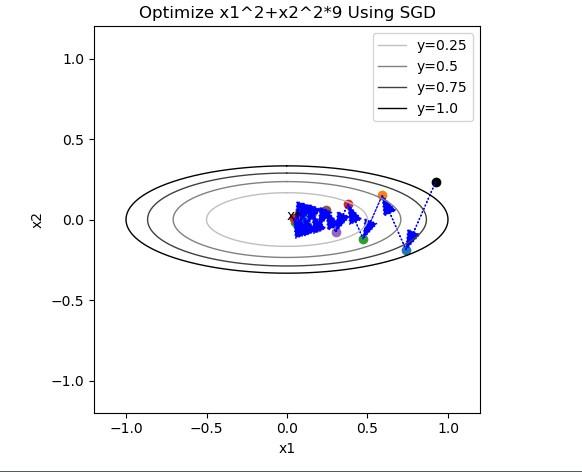
# ### 考虑
# 所以人们考虑
# - 动态选择更好的学习率,比如前期大些来加速优化,靠近低点了小些避免在低点附近来回震荡,甚至
# - 为每个维度选择合适的学习率 $\eta$ .
# # 学习率退火 (Learning Rate Annealing)
#
# 出于考虑1,人们参考了单体优化方法中的模拟退火(Simulated Annealing),学习率随着迭代次数的增加或者损失函数在验证集上的表现变好而衰减(decay).
# 学习率退化可以直接加在GD上.
#
# ### 改进方向
# AdaGrad等算法([我的一篇知乎文章](https://zhuanlan.zhihu.com/p/109521635)有介绍)就借鉴了退火的学习率衰减的思想.不过这个不是这篇的重点.
#
# # 牛顿法 (Newton's Method)
#
# 出于考虑2(为每个维度选择合适的$\eta$),基于二阶导数的牛顿法被提出.它来源于泰勒二阶展开.
# $f(x_1) = f(x_0) + f'(x)|_{x=x_0} * (x_1 - x_0) + \frac{(x_1-x_0)^2}{2!} f''(x)|_{x={x_0}} + o((x_1-x_0)^2) $.
#
# 对于多元函数 x ,
# $f(x_1) = f(x_0) + g(x_0) * (x_1 - x_0) + \frac{1}{2!} (x_1-x_0) H(x_0) (x_1-x_0)^t+ o((x_1-x_0)^2) $
#
# 其中 $H(x)$ 为Hessian矩阵
# 有 $H[i, j] = \frac{\partial^2{L}}{\partial{x^i}\partial{x^j}}$ .
#
# 这样每次迭代都会考虑损失函数的曲率(二阶导数)来选择步长.对比上图中的标准GD,牛顿法可以一步就到达最优点.
# ### 牛顿法缺点
#
# 但是牛顿法的计算复杂度很高,因为Hessian矩阵的维度是参数个数的平方,而参数的个数往往很多.
#
#
# ### 改进方向
# 不同的方法随即被提出,比如
# - Becker和LeCun提出的[用对角线元素来代替Hessian全矩阵](https://nyuscholars.nyu.edu/en/publications/improving-the-convergence-of-back-propagation-learning-with-secon)
#
# - 依靠历史的梯度信息来模拟二阶方法,包括Momentum,RMSProp(用二阶距来模拟二阶导数),AdaM(用一阶矩和二阶矩的比例来模拟二阶导数)等.
#
# 我们先介绍Momentum
# # Momentum
#
# 借鉴了物理中动量(momentum)的概念,让 $\Delta x_t $保留一部分之前的方向,而不是完全用梯度的负方向.
#
# 每次迭代的更新为
# $\Delta x_t = - \eta* g_t + \rho * \Delta x_{t-1} $
# 或
# $\Delta x_t = \eta * (\mu * \Delta x_{t-1} - g_t) $
# 其中$\mu$也可以写成momentum
#
# 这样预期可以达到两个效果:
# - 某个维度在近几次迭代中正负号总是改变时,说明二阶导数可能相对其他维度比较大或者说每次步子迈得太大了,需要改变幅度小些或者迈得小点来避免zig-zag路径
# - 某个维度在近几次迭代中符号几乎不变,说明二阶导数可能相对其他维度比较小或者说大方向是正确的,这个维度改变的幅度可以扩大些,来加速改进.
#
#
# 如下图所示,加入了Momentum,前期的训练加快了,靠近低点时也减小了震荡.
# 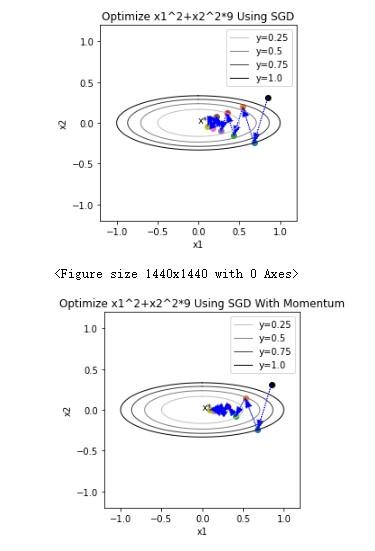
#
|
ipynbs/appendix/optimizer/1. Standard GD and Momentum.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''Py37'': conda)'
# name: python3
# ---
# # Comprehensive Analysis of our Adahybrid strategy
#
import numpy as np
import pandas as pd
import seaborn as sns
import glob
import csv
import traceback
import datetime
import os
pd.options.display.max_columns=50
# ## 1. [Prelim] Comparison between 90%+10% hybrid strategies and Adahybrid (Take a chance model vs Our model)
# - xgb(90%)+random(10%)
# - DATE(90%)+bATE(10%)
# - Ada DATE+bATE
# - Ada xgb+random
results = glob.glob('../results/performances/ada-prelim-result-*') # quick- or www21- or fld-
list1, list2 = zip(*sorted(zip([os.stat(result).st_size for result in results], results)))
# ### Collecting Result Files: Results of Individual Experiments
# +
import matplotlib.pyplot as plt
from collections import defaultdict
# %matplotlib inline
full_results = defaultdict(list)
# Retrieving results
num_logs = len([i for i in list1 if i > 1000])
count= 0
for i in range(1,num_logs+1):
try:
df = pd.read_csv(list2[-i])
var = 'norm-precision'
rolling_mean7 = df[var].rolling(window=7).mean()
rolling_mean14 = df[var].rolling(window=14).mean()
filename = list2[-i][list2[-i].index('16'):list2[-i].index('16')+10]
info = ','.join(list(df[['data', 'sampling', 'subsamplings']].iloc[0]))
full_results[info].append(rolling_mean14)
count += 1
# Draw individual figures
# plt.figure()
# plt.title(info+','+filename)
# plt.plot(df['numWeek'], df[var], color='skyblue', label='Weekly')
# plt.plot(df['numWeek'], rolling_mean7, color='teal', label='MA (7 weeks)')
# plt.plot(df['numWeek'], rolling_mean14, color='blue', label='MA (14 weeks)')
# plt.legend(loc='upper left')
# plt.ylabel(var)
# plt.xlabel('numWeeks')
# plt.show()
except:
print('loading error:', list2[-i])
continue
print(count)
# plt.close()
# -
full_results.keys()
# ### Mdata Simulation Results - Hybrid
full_results.keys()
# +
plt.figure()
# info = ','.join(list(df[['data', 'samplings']].iloc[0]))
result_one_dataset = [key for key in full_results.keys() if 'real-m' in key]
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
# print(pd.concat([*full_results[key]], axis=1)) # Check current running status: debug purpose
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
plt.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Mdata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left')
plt.ylabel(var)
plt.xlabel('numWeeks')
# plt.ylim(0.4, 0.6)
plt.show()
plt.close()
# -
# ### Tdata Simulation Results - Hybrid
# +
plt.figure()
result_one_dataset = [key for key in full_results.keys() if 'real-t' in key]
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
plt.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Tdata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left')
plt.ylabel(var)
plt.xlabel('numWeeks')
plt.show()
plt.close()
# -
# ## 2. [Ada] Comparison between Hybrid Strategies (by changing Exploration part) and Adahybrid
cresults = glob.glob('../results/performances/ada-result-*') # quick- or www21- or fld-
clist1, clist2 = zip(*sorted(zip([os.stat(result).st_size for result in cresults], cresults)))
# +
import matplotlib.pyplot as plt
from collections import defaultdict
# %matplotlib inline
full_results = defaultdict(list)
# Retrieving results
num_logs = len([i for i in clist1 if i > 1000])
count= 0
for i in range(1,num_logs+1):
try:
df = pd.read_csv(clist2[-i])
var = 'norm-precision'
rolling_mean7 = df[var].rolling(window=7).mean()
rolling_mean14 = df[var].rolling(window=14).mean()
filename = clist2[-i][clist2[-i].index('16'):clist2[-i].index('16')+10]
info = ','.join(list(df[['data', 'sampling', 'subsamplings', 'current_weights']].iloc[0]))
full_results[info].append(rolling_mean14)
count += 1
# Draw individual figures
# plt.figure()
# plt.title(info+','+filename)
# plt.plot(df['numWeek'], df[var], color='skyblue', label='Weekly')
# plt.plot(df['numWeek'], rolling_mean7, color='teal', label='MA (7 weeks)')
# plt.plot(df['numWeek'], rolling_mean14, color='blue', label='MA (14 weeks)')
# plt.legend(loc='upper left')
# plt.ylabel(var)
# plt.xlabel('numWeeks')
# plt.show()
except:
print('loading error:', clist2[-i])
continue
print(count)
# plt.close()
# +
fig = plt.figure()
ax = fig.add_subplot(111)
# info = ','.join(list(df[['data', 'samplings']].iloc[0]))
plt.style.use('seaborn-dark')
colors = sns.color_palette("icefire", 12)
ax.set_prop_cycle('color', colors)
result_one_dataset = sorted([key for key in full_results.keys() if 'real-m' in key])
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
# print(pd.concat([*full_results[key]], axis=1)) # Check current running status: debug purpose
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
ax.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Mdata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
plt.ylabel(var)
plt.xlabel('numWeeks')
# plt.ylim(0.4, 0.6)
plt.show()
plt.close()
# +
fig = plt.figure()
ax = fig.add_subplot(111)
# info = ','.join(list(df[['data', 'samplings']].iloc[0]))
plt.style.use('seaborn-dark')
colors = sns.color_palette("icefire", 12)
ax.set_prop_cycle('color', colors)
result_one_dataset = sorted([key for key in full_results.keys() if 'real-t' in key])
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
# print(pd.concat([*full_results[key]], axis=1)) # Check current running status: debug purpose
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
ax.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Tdata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
plt.ylabel(var)
plt.xlabel('numWeeks')
# plt.ylim(0.4, 0.6)
plt.show()
plt.close()
# +
fig = plt.figure()
ax = fig.add_subplot(111)
# info = ','.join(list(df[['data', 'samplings']].iloc[0]))
plt.style.use('seaborn-dark')
colors = sns.color_palette("icefire", 12)
ax.set_prop_cycle('color', colors)
result_one_dataset = sorted([key for key in full_results.keys() if 'real-c' in key])
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
# print(pd.concat([*full_results[key]], axis=1)) # Check current running status: debug purpose
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
ax.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Cdata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
plt.ylabel(var)
plt.xlabel('numWeeks')
# plt.ylim(0.4, 0.6)
plt.show()
plt.close()
# +
fig = plt.figure()
ax = fig.add_subplot(111)
# info = ','.join(list(df[['data', 'samplings']].iloc[0]))
plt.style.use('seaborn-dark')
colors = sns.color_palette("icefire", 12)
ax.set_prop_cycle('color', colors)
result_one_dataset = sorted([key for key in full_results.keys() if 'real-n' in key])
print('The number of trials for each setting (Results are averaged):')
for key in result_one_dataset:
avg_result = pd.concat([*full_results[key]], axis=1).mean(axis=1)
# print(pd.concat([*full_results[key]], axis=1)) # Check current running status: debug purpose
print(key, len(full_results[key]), round(np.mean(avg_result[-13:]), 4))
ax.plot(avg_result.index, avg_result, label=key)
# # printing test_illicit_rate
# tir = pd.read_csv(list2[-1])['test_illicit_rate'].rolling(window=7).mean()
# plt.plot(tir.index, tir, label='Test illicit rate (ref)')
plt.title('<Ndata> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
plt.legend(loc='upper left', bbox_to_anchor=(1.05, 1))
plt.ylabel(var)
plt.xlabel('numWeeks')
# plt.ylim(0.4, 0.6)
plt.show()
plt.close()
# -
# ## 3. Weights distribution visualization
exp_prefix = 'ada-prelim'
wresults = glob.glob(f'../results/ada_ratios/{exp_prefix}-result-*') # quick- or www21- or fld-
wlist1, wlist2 = zip(*sorted(zip([os.stat(result).st_size for result in wresults], wresults)))
# +
import matplotlib.pyplot as plt
from collections import defaultdict
# %matplotlib inline
full_results = defaultdict(list)
weights = defaultdict(dict)
# Retrieving results
num_logs = len([i for i in wlist1 if i > 1000])
count= 0
file_no = 0
for i in range(1,num_logs+1):
try:
cols = [f'{i/20} explore rate' for i in range(20)]
df = pd.read_csv(wlist2[-i])
filename = wlist2[-i][wlist2[-i].index('16'):wlist2[-i].index('16')+10]
dataname = df['data'].iloc[0]
full_results[(filename, dataname)] = df[cols].transpose()
weights[(filename, dataname)] = df['chosen_rate']
count += 1
# Draw individual figures
# plt.figure()
# plt.title(info+','+filename)
# plt.plot(df['numWeek'], df[var], color='skyblue', label='Weekly')
# plt.plot(df['numWeek'], rolling_mean7, color='teal', label='MA (7 weeks)')
# plt.plot(df['numWeek'], rolling_mean14, color='blue', label='MA (14 weeks)')
# plt.legend(loc='upper left')
# plt.ylabel(var)
# plt.xlabel('numWeeks')
# plt.show()
except:
print('loading error:', wlist2[-i])
continue
print(count)
# plt.close()
# -
full_results.keys()
keys = list(full_results.keys())
keys
# +
ind = 1
runid = keys[ind][0]
data = keys[ind][1]
wresults = glob.glob(f'../results/ada_ratios/{exp_prefix}-result-{runid}*')[0] # quick- or www21- or fld-
fig = plt.figure(figsize=(16,8))
n = full_results[(runid, data)].shape[1]
ax = fig.add_subplot(211)
ax = sns.heatmap(full_results[(runid, data)], linewidth=0, cmap = 'YlGnBu')
ax.set_title(f'<{wresults[38:-9]}-{data}> Train: 1 months, Valid: 28 days, Test: 7 days, fast linear decay')
ax.legend(loc='upper left')
ax.set_ylabel('Exploration')
ax.set_xlabel('numWeeks')
ax.set_xticks(np.arange(0, n, 10))
ax.set_xticklabels(np.arange(0, n, 10))
ax2 = fig.add_subplot(212)
ax2.plot(weights[(runid, data)])
ax2.set_ylabel('Exploration')
ax2.set_xlabel('numWeeks')
ax2.set_xticks(np.arange(0, n, 10))
ax2.set_xticklabels(np.arange(0, n, 10))
plt.show()
plt.close()
|
analysis/analysis-ada.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0
# language: julia
# name: julia-1.3
# ---
using ProgressMeter
using Printf
using BSON: @save, @load
using PyCall
pygui(:qt)
using PyPlot
pygui(true)
include("escape_dimer_one_hit.jl")
H=.22
# +
t_end = 1e3
width = 2.25; height =4.5
n_iter_P = 100;n_iter_Q =100;
N = n_iter_P * n_iter_Q;
ArrQ = range(-width, stop = width, length = n_iter_Q)
ArrP = range(0, stop = height, length = n_iter_P)
mesh = [(Q, P) for Q in ArrQ, P in ArrP]
mesh_list = reshape(mesh, 1, :)
t_list = t_end * ones(N);
Energy = H* ones(N);
h = replace(@sprintf("%.13f",H), "." => "_")
nQ = @sprintf("_%d",n_iter_Q)
nP = @sprintf("_%d",n_iter_P)
h_BSON="num_hhits_escape_data"*h*nQ*nP*".bson"
location = "/mnt/bdd38f66-9ece-451a-b915-952523c139d2/Escape/"
println(h_BSON)
one_hit_exit=[]
# -
include("escape_dimer_one_hit.jl")
exit_points= @showprogress map(escape_after_one_hit, mesh_list, t_list, Energy);
cleaned_points=[]
for item in exit_points
if item!=nothing
push!(cleaned_points, item)
end
end
cleaned_points
Q0,P0,Q1,P1=unzipper3(cleaned_points)
Q_0=vcat(Q0,-Q0); P_0=vcat(P0,-P0);
Q_1=vcat(Q1,-Q1); P_1=vcat(P1,-P1);
plot(Q_0,P_0, ".",markersize=1,c=:blue);
plot(Q_1,P_1, ".",markersize=1,c=:red);
xlim(-2.5,2.5)
ylim(-4.5,4.5) ;
exit_points
Q0,P0,Q1,P1=unzipper3(cleaned_points)
Q_0=vcat(Q0,-Q0); P_0=vcat(P0,-P0);
Q_1=vcat(Q1,-Q1); P_1=vcat(P1,-P1);
plot(Q_0,P_0, ".",markersize=1,c=:blue);
plot(Q_1,P_1, ".",markersize=1,c=:red);
xlim(-2.5,2.5)
ylim(-4.5,4.5) ;
cleaned_points
sol[4,1]
function add_eight!(point,radius, one_hit_exit, t_end, energy)
X=point[2]
Y=point[4]
X_plus=X+radius
Y_plus=Y+radius
X_minus=X-radius
Y_minus=Y-radius
point_1=[X_minus, Y_plus]
point_2=[X, Y_plus]
point_3=[X_minus, Y_plus]
point_4=[X_minus, Y]
point_5=[X_plus, Y]
point_6=[X_minus, Y_minus]
point_7=[X, Y_minus]
point_8=[X_minus, Y_minus];
new_points= [point_1, point_2, point_3, point_4, point_5, point_6, point_7, point_8]
for point in new_points
new_point=escape_after_one_hit(point,t_end,energy)
if new_point!=nothing
push!(one_hit_exit, new_point)
end
end
end
function add_four!(point,radius, one_hit_exit, t_end, energy)
X=point[2]
Y=point[4]
X_plus=X+radius
Y_plus=Y+radius
X_minus=X-radius
Y_minus=Y-radius
point_2=[X, Y_plus]
point_4=[X_minus, Y]
point_5=[X_plus, Y]
point_7=[X, Y_minus]
new_points= [point_2, point_4, point_5, point_7]
for point in new_points
new_point=escape_after_one_hit(point,t_end,energy)
if new_point!=nothing
push!(one_hit_exit, new_point)
end
end
end
function addpoint!(list)
push!(list,[-10,10])
end
function unzipper2(array_of_points)
N=length(array_of_points)
Q=zeros(N)
P=zeros(N)
for (index,point) in enumerate(array_of_points)
Q[index]=point[2]
P[index]=point[4]
end
return Q,P
end
function unzipper3(array_of_points)
N=length(array_of_points)
Q0=zeros(N)
P0=zeros(N)
Q1=zeros(N)
P1=zeros(N)
for (index,point) in enumerate(array_of_points)
Q0[index]=point[2]
P0[index]=point[4]
Q1[index]=point[6]
P1[index]=point[8]
end
return Q0,P0,Q1,P1
end
Q0,P0,Q1,P1=unzipper3(new_forward)
plot(Q0,P0, ".",markersize=1,c=:blue);
plot(Q1,P1, ".",markersize=1,c=:red);
xlim(-2.5,2.5)
ylim(-4.5,4.5) ;
# +
old_list=copy(cleaned_points)
energy=.25
radius=.1
@showprogress for (index,point) in enumerate(old_list)
add_eight!(point,radius, cleaned_points, t_end, energy)
end
cleaned_points
length(cleaned_points)
Q0,P0,Q1,P1=unzipper3(cleaned_points)
Q_0=vcat(Q0,-Q0); P_0=vcat(P0,-P0);
Q_1=vcat(Q1,-Q1); P_1=vcat(P1,-P1);
plot(Q_0,P_0, ".",markersize=1,c=:blue);
plot(Q_1,P_1, ".",markersize=1,c=:red);
xlim(-2.5,2.5)
ylim(-4.5,4.5) ;
# -
Q,P=unzipper2(cleaned_points)
Q_one=vcat(Q,-Q); P_one=vcat(P,-P);
plot(Q_one,P_one, ".",markersize=1,c=:blue);
cleaned_points
SAVE_DATA=Dict("one_hit_exit"=>one_hit_exit)
include("forward_one.jl")
# +
new_forward=[]
@showprogress for point in cleaned_points
forward_point=forward_one(point, t_end, H)
if forward_point!=nothing
push!(new_forward,forward_point)
end
end
new_forward
# -
Q,P=unzipper2(new_forward)
Q_escape=vcat(Q,-Q); P_escape=vcat(P,-P);
Q,P=unzipper2(cleaned_points)
Q_one=vcat(Q,-Q); P_one=vcat(P,-P);
# plot(Q_one,P_one, ".",markersize=1,c=:blue);
plot(Q_escape,P_escape, ".",markersize=1,c=:red);
xlim(-2.5,2.5)
ylim(-4.5,4.5) ;
include("negative_time.jl")
# +
forward_back=[]
@showprogress for point in cleaned_points
back_point=back_one(point, t_end, H)
if back_point!=nothing
push!(forward_back,back_point)
end
end
# -
new_forward
cleaned_points
# +
Q0,P0=unzipper2(new_forward)
Q_escape=vcat(Q0,-Q0); P_escape=vcat(P0,-P0);
Q,P=unzipper2(cleaned_points)
Q_one=vcat(Q,-Q); P_one=vcat(P,-P);
Q1,P1=unzipper2(forward_back)
Q_b=vcat(Q1,-Q1); P_b=vcat(P1,-P1);
plot(Q0,P0, ".",markersize=1,c=:blue);
plot(Q,P, ".",markersize=1,c=:red);
plot(Q1,P1, ".",markersize=1,c=:green);
xlim(-2.5,2.5)
ylim(-4.5,4.5)
# -
forward_back
log10(200)
@load "escape_TIME_cdata0_250000001000.bson" SAVE_DATA
time_to_exit=SAVE_DATA["escape_times"]
mesh=SAVE_DATA["mesh"]
ArrP=SAVE_DATA["ArrP"]
ArrQ=SAVE_DATA["ArrQ"]
width = 2.5;
height=5;
logz=log1p.(time_to_exit);
mesh_P=[P for Q in ArrQ, P in ArrP];
mesh_Q=[Q for Q in ArrQ, P in ArrP];
M,N=size(logz)
# for i in 1:M
# for j in 1:N
# if isnan(logz[i,j])
# logz[i,j] = 0
# end
# end
# end
ArrP = range(-width, stop = width, length = N)
pcolormesh(ArrP,ArrQ,logz)
pcolormesh(reverse(ArrP),-ArrQ,logz)
colorbar()
contourf(ArrP,ArrQ,logz)
contourf(reverse(ArrP),-ArrQ,logz)
colorbar()
size(logz)
for i in 1:1000
for j in 1:1000
if isnan(logz[i,j])
logz[i,j] = 0
end
end
end
logz
isnan(NaN)
# + active=""
# reverse(ArrQ)
# -
ArrQ
SAVE_DATA
|
point_brancher.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to get the $\nabla^2$operator in tensorflow
# In tensorflow, the default 2nd derivative operator will contract over the hessian matrix in an undesirable way. For example:
import tensorflow as tf
import tensorflow_probability as tfp
def generate_inputs(nwalkers, n_particles, dimension):
x = tf.random.uniform(shape=[nwalkers, n_particles, dimension])
return x
# This function generates walkers in the same format as metropolis walkers, with random values
inputs = generate_inputs(4,1,2)
# The function below creates a scalar value: f(x, y) = $\alpha x^2 + \beta y^2 + \gamma x y$
def wavefunction(inputs, alpha=0.1, beta=0.2, gamma=0.5):
# return x^2 + y^2 + xy, with constants:
ret = alpha * inputs[:,:,0]**2
ret += beta * inputs[:,:,1]**2
ret += gamma * inputs[:,:,0]*inputs[:,:,1]
return tf.squeeze(ret)
output = wavefunction(inputs)
output.shape
# Here is the forward pass, telling tensorflow to watch the inputs since that's what we want to differentiate with respect to:
# +
inputs = generate_inputs(4,1,2)
with tf.GradientTape(persistent=True) as outer_tape:
outer_tape.watch(inputs)
with tf.GradientTape(persistent=True) as inner_tape:
inner_tape.watch(inputs)
outputs = wavefunction(inputs)
# Compute the first derivative with respect to the inputs
dw_dx = inner_tape.gradient(outputs, inputs)
# -
# Note that we have to compute the first derivative, above, within the block of the outer_tape to compute a second derivative
# This should have the same shape as the inputs, and the values are analytically computable (and thus checkable):
assert dw_dx.shape == inputs.shape
def analytic_derivative(inputs, alpha=0.1, beta=0.2, gamma=0.5):
_x = inputs[:,:,0]
_y = inputs[:,:,1]
x = 2*alpha*_x + gamma*_y
y = 2*beta*_y + gamma*_x
return tf.stack([x,y], axis=2)
dw_dx - analytic_derivative(inputs)
# The challenge, then, is the 2nd derivative. We know what it *should* be if we're computing $\nabla^2$, but this is not what tensorflow computes:
def nabla_squared(inputs, alpha=0.1, beta=0.2, gamma=0.5):
_x = tf.constant(2 * alpha, shape=inputs[:,:,0].shape)
_y = tf.constant(2*beta, shape=inputs[:,:,1].shape)
return tf.stack([_x, _y], axis=2)
nabla_squared(inputs)
d2w_dx2 = outer_tape.gradient(dw_dx, inputs)
print(d2w_dx2)
# If you look closely, each value here is off by $\gamma$ - tensorflow is contracting the hessian to compute this second derivative!
d2wdx2 = outer_tape.jacobian(dw_dx, inputs)
print(d2wdx2)
# The jacobian comes out with a dimension that is much too big. We can contract it with "einsum":
tf.einsum("wpdwpd->wpd",d2wdx2)
# And now, that is the correct value for $\nabla^2$ of this function!
# This is an equivalent implementation in our case where the output (per "batch") depends only on that input "batch":
tf.einsum("wpdpd->wpd",outer_tape.batch_jacobian(dw_dx, inputs))
|
Tensorflow 2nd Derivatives.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="knOigRU1UJ9Y"
# # Estimating Work Tour Scheduling
#
# This notebook illustrates how to re-estimate the mandatory tour scheduling component for ActivitySim. This process
# includes running ActivitySim in estimation mode to read household travel survey files and write out
# the estimation data bundles used in this notebook. To review how to do so, please visit the other
# notebooks in this directory.
# -
# # Load libraries
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="s53VwlPwtNnr" outputId="d1208b7a-c1f2-4b0b-c439-bf312fe12be0"
import os
import larch # !conda install larch -c conda-forge # for estimation
import pandas as pd
# -
# We'll work in our `test` directory, where ActivitySim has saved the estimation data bundles.
os.chdir('test')
# # Load data and prep model for estimation
# +
modelname = "mandatory_tour_scheduling_work"
from activitysim.estimation.larch import component_model
model, data = component_model(modelname, return_data=True)
# -
# # Review data loaded from the EDB
#
# The next (optional) step is to review the EDB, including the coefficients, utilities specification, and chooser and alternative data.
# ## Coefficients
data.coefficients
# ## Utility specification
data.spec
# ## Chooser data
data.chooser_data
# ## Alternatives data
data.alt_values
# # Estimate
#
# With the model setup for estimation, the next step is to estimate the model coefficients. Make sure to use a sufficiently large enough household sample and set of zones to avoid an over-specified model, which does not have a numerically stable likelihood maximizing solution. Larch has a built-in estimation methods including BHHH, and also offers access to more advanced general purpose non-linear optimizers in the `scipy` package, including SLSQP, which allows for bounds and constraints on parameters. BHHH is the default and typically runs faster, but does not follow constraints on parameters.
model.estimate()
# ### Estimated coefficients
model.parameter_summary()
# + [markdown] colab_type="text" id="TojXWivZsx7M"
# # Output Estimation Results
# -
from activitysim.estimation.larch import update_coefficients
result_dir = data.edb_directory/"estimated"
update_coefficients(
model, data, result_dir,
output_file=f"{modelname}_coefficients_revised.csv",
);
# ### Write the model estimation report, including coefficient t-statistic and log likelihood
model.to_xlsx(
result_dir/f"{modelname}_model_estimation.xlsx",
data_statistics=False,
)
# # Next Steps
#
# The final step is to either manually or automatically copy the `*_coefficients_revised.csv` file to the configs folder, rename it to `*_coefficients.csv`, and run ActivitySim in simulation mode.
pd.read_csv(result_dir/f"{modelname}_coefficients_revised.csv")
|
activitysim/examples/example_estimation/notebooks/08_work_tour_scheduling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nterop={"id": "17"}
# %run algorithms.ipynb
# + nterop={"id": "3"}
import ctypes
from copy import deepcopy
from functools import partial
from itertools import product
import multiprocessing
from multiprocessing.managers import BaseManager
import numpy as np
import os
import sys
import time
# + nterop={"id": "11"}
class SharedState(object):
def __init__(self, slot_count=None, level_count_per_slot=None, additive=None):
self.regret_shared_array_base = multiprocessing.Array(ctypes.c_double, simulation_count * horizon)
self.regret = np.ctypeslib.as_array(self.regret_shared_array_base.get_obj())
self.regret = self.regret.reshape((simulation_count, horizon))
self.reward_shared_array_base = multiprocessing.Array(ctypes.c_double, simulation_count * horizon)
self.reward = np.ctypeslib.as_array(self.reward_shared_array_base.get_obj())
self.reward = self.reward.reshape((simulation_count, horizon))
if slot_count is None or level_count_per_slot is None:
# Fixed world.
slot_count = 3
level_count_per_slot = 3
input_marginals = np.array([[0.08, 0.10, 0.09], [0.11, 0.11, 0.06], [0.05, 0.16, 0.07]])
self.worlds = [World(slot_count=slot_count,level_count_per_slot=level_count_per_slot, input_marginals=input_marginals)] \
* simulation_count
else:
self.worlds = []
for s in range(simulation_count):
self.worlds.append(World(slot_count=slot_count,level_count_per_slot=level_count_per_slot,
additive=additive))
def get_world(self, s):
return self.worlds[s]
def update_regret(self, s, regret_per_period):
self.regret[s, :] = regret_per_period[:]
def get_regret(self):
return self.regret
def update_reward(self, s, reward_per_period):
self.reward[s, :] = reward_per_period[:]
def get_reward(self):
return self.reward
BaseManager.register('SharedState', SharedState)
def single_simulation(s, agent_type, shared_state):
t0 = time.time()
world = shared_state.get_world(s)
if (agent_type == "IndependentBernoulliArmsTSAgent"):
agent = IndependentBernoulliArmsTSAgent(world = world, horizon = horizon)
elif (agent_type == "MarginalPosteriorTSAgent"):
agent = MarginalPosteriorTSAgent(world = world, horizon = horizon)
elif (agent_type == "MarginalPosteriorUCBAgent"):
agent = MarginalPosteriorTSAgent(world = world, horizon = horizon)
elif (agent_type == "LogisticRegressionTSAgent"):
agent = LogisticRegressionTSAgent(world = world, horizon = horizon, regularization_parameter = regularization_parameter)
elif (agent_type == "LogisticRegressionUCBAgent"):
agent = LogisticRegressionUCBAgent(world = world, horizon = horizon, regularization_parameter = regularization_parameter)
else:
print("This agent_type is not supported.")
return
agent.run()
shared_state.update_regret(s, agent.regret_per_period)
shared_state.update_reward(s, agent.reward_per_period)
t1 = time.time()
def run(agent_types, output_prefix, slot_count=None, level_count_per_slot=None, additive=None):
if (seed > 0):
np.random.seed(seed)
manager = BaseManager()
manager.start()
pool = multiprocessing.Pool(core_count)
shared_state = manager.SharedState(slot_count=slot_count, level_count_per_slot=level_count_per_slot)
for agent_type in agent_types:
t0 = time.time()
func = partial(single_simulation, agent_type = agent_type, shared_state = shared_state)
pool.map(func, range(simulation_count))
t1 = time.time()
print("{} Elapsed Time: {}".format(agent_type, (t1 - t0)))
parameters = ""
if (agent_type == "LogisticRegressionTSAgent" or agent_type == "LogisticRegressionUCBAgent"):
parameters = "_R{}".format(regularization_parameter)
if not os.path.exists("Results/"):
os.makedirs("Results/")
filename = "Results/{}_{}_H{}_S{}{}".format(output_prefix, agent_type, horizon, simulation_count, parameters)
np.save(filename + "_Regret.npy", np.mean(shared_state.get_regret(), axis = 0))
np.save(filename + "_RegretVar.npy", np.var(shared_state.get_regret(), axis = 0))
np.save(filename + "_ET.npy", (t1-t0) / simulation_count)
# + nterop={"id": "15"}
horizon = 50000
simulation_count = 1000
regularization_parameter = 10
seed = 1704
core_count = multiprocessing.cpu_count()
agent_types = ["IndependentBernoulliArmsTSAgent", "MarginalPosteriorTSAgent", "LogisticRegressionTSAgent"]
slot_count_list = [2, 3, 4]
level_count_per_slot_list = [2, 3, 4, 5]
additive_world_type_list = [True, False]
for slot_count, level_count_per_slot, additive_world_type in product(slot_count_list, level_count_per_slot_list,
additive_world_type_list):
output_prefix = "SLOT{}LEVEL{}_ADD{}".format(slot_count, level_count_per_slot, additive_world_type)
print(output_prefix)
run(agent_types = agent_types, output_prefix=output_prefix,
slot_count=slot_count, level_count_per_slot=level_count_per_slot)
# + nterop={"id": "18"}
|
simulate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../Pierian-Data-Logo.PNG">
# <br>
# <strong><center>Copyright 2019. Created by <NAME>.</center></strong>
# # Linear Regression with PyTorch
# In this section we'll use PyTorch's machine learning model to progressively develop a best-fit line for a given set of data points. Like most linear regression algorithms, we're seeking to minimize the error between our model and the actual data, using a <em>loss function</em> like mean-squared-error.
#
# <img src='../Images/linear-regression-residuals.png' width='400' style="display: inline-block"><br>
#
# Image source: <a href='https://commons.wikimedia.org/wiki/File:Residuals_for_Linear_Regression_Fit.png'>https://commons.wikimedia.org/wiki/File:Residuals_for_Linear_Regression_Fit.png</a>
#
# To start, we'll develop a collection of data points that appear random, but that fit a known linear equation $y = 2x+1$
# ## Perform standard imports
# +
import torch
import torch.nn as nn # we'll use this a lot going forward!
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ## Create a column matrix of X values
# We can create tensors right away rather than convert from NumPy arrays.
# +
X = torch.linspace(1,50,50).reshape(-1,1)
# Equivalent to
# X = torch.unsqueeze(torch.linspace(1,50,50), dim=1)
# -
# ## Create a "random" array of error values
# We want 50 random integer values that collectively cancel each other out.
torch.manual_seed(71) # to obtain reproducible results
e = torch.randint(-8,9,(50,1),dtype=torch.float)
print(e.sum())
# ## Create a column matrix of y values
# Here we'll set our own parameters of $\mathrm {weight} = 2,\; \mathrm {bias} = 1$, plus the error amount.<br><strong><tt>y</tt></strong> will have the same shape as <strong><tt>X</tt></strong> and <strong><tt>e</tt></strong>
y = 2*X + 1 + e
print(y.shape)
# ## Plot the results
# We have to convert tensors to NumPy arrays just for plotting.
plt.scatter(X.numpy(), y.numpy())
plt.ylabel('y')
plt.xlabel('x');
# Note that when we created tensor $X$, we did <em>not</em> pass <tt>requires_grad=True</tt>. This means that $y$ doesn't have a gradient function, and <tt>y.backward()</tt> won't work. Since PyTorch is not tracking operations, it doesn't know the relationship between $X$ and $y$.
# ## Simple linear model
# As a quick demonstration we'll show how the built-in <tt>nn.Linear()</tt> model preselects weight and bias values at random.
# +
torch.manual_seed(59)
model = nn.Linear(in_features=1, out_features=1)
print(model.weight)
print(model.bias)
# -
# Without seeing any data, the model sets a random weight of 0.1060 and a bias of 0.9638.
# ## Model classes
# PyTorch lets us define models as object classes that can store multiple model layers. In upcoming sections we'll set up several neural network layers, and determine how each layer should perform its forward pass to the next layer. For now, though, we only need a single <tt>linear</tt> layer.
class Model(nn.Module):
def __init__(self, in_features, out_features):
super().__init__()
self.linear = nn.Linear(in_features, out_features)
def forward(self, x):
y_pred = self.linear(x)
return y_pred
# <div class="alert alert-info"><strong>NOTE:</strong> The "Linear" model layer used here doesn't really refer to linear regression. Instead, it describes the type of neural network layer employed. Linear layers are also called "fully connected" or "dense" layers. Going forward our models may contain linear layers, convolutional layers, and more.</div>
# When <tt>Model</tt> is instantiated, we need to pass in the size (dimensions) of the incoming and outgoing features. For our purposes we'll use (1,1).<br>As above, we can see the initial hyperparameters.
torch.manual_seed(59)
model = Model(1, 1)
print(model)
print('Weight:', model.linear.weight.item())
print('Bias: ', model.linear.bias.item())
# As models become more complex, it may be better to iterate over all the model parameters:
for name, param in model.named_parameters():
print(name, '\t', param.item())
# <div class="alert alert-info"><strong>NOTE:</strong> In the above example we had our Model class accept arguments for the number of input and output features.<br>For simplicity we can hardcode them into the Model:
#
# <tt><font color=black>
# class Model(torch.nn.Module):<br>
# def \_\_init\_\_(self):<br>
# super().\_\_init\_\_()<br>
# self.linear = Linear(1,1)<br><br>
# model = Model()
# </font></tt><br><br>
#
# Alternatively we can use default arguments:
#
# <tt><font color=black>
# class Model(torch.nn.Module):<br>
# def \_\_init\_\_(self, in_dim=1, out_dim=1):<br>
# super().\_\_init\_\_()<br>
# self.linear = Linear(in_dim,out_dim)<br><br>
# model = Model()<br>
# <em>\# or</em><br>
# model = Model(i,o)</font></tt>
# </div>
# Now let's see the result when we pass a tensor into the model.
x = torch.tensor([2.0])
print(model.forward(x)) # equivalent to print(model(x))
# which is confirmed with $f(x) = (0.1060)(2.0)+(0.9638) = 1.1758$
# ## Plot the initial model
# We can plot the untrained model against our dataset to get an idea of our starting point.
x1 = np.array([X.min(),X.max()])
print(x1)
# +
w1,b1 = model.linear.weight.item(), model.linear.bias.item()
print(f'Initial weight: {w1:.8f}, Initial bias: {b1:.8f}')
print()
y1 = x1*w1 + b1
print(y1)
# -
plt.scatter(X.numpy(), y.numpy())
plt.plot(x1,y1,'r')
plt.title('Initial Model')
plt.ylabel('y')
plt.xlabel('x');
# ## Set the loss function
# We could write our own function to apply a Mean Squared Error (MSE) that follows<br>
#
# $\begin{split}MSE &= \frac {1} {n} \sum_{i=1}^n {(y_i - \hat y_i)}^2 \\
# &= \frac {1} {n} \sum_{i=1}^n {(y_i - (wx_i + b))}^2\end{split}$<br>
#
# Fortunately PyTorch has it built in.<br>
# <em>By convention, you'll see the variable name "criterion" used, but feel free to use something like "linear_loss_func" if that's clearer.</em>
criterion = nn.MSELoss()
# ## Set the optimization
# Here we'll use <a href='https://en.wikipedia.org/wiki/Stochastic_gradient_descent'>Stochastic Gradient Descent</a> (SGD) with an applied <a href='https://en.wikipedia.org/wiki/Learning_rate'>learning rate</a> (lr) of 0.001. Recall that the learning rate tells the optimizer how much to adjust each parameter on the next round of calculations. Too large a step and we run the risk of overshooting the minimum, causing the algorithm to diverge. Too small and it will take a long time to converge.
#
# For more complicated (multivariate) data, you might also consider passing optional <a href='https://en.wikipedia.org/wiki/Stochastic_gradient_descent#Momentum'><tt>momentum</tt></a> and <a href='https://en.wikipedia.org/wiki/Tikhonov_regularization'><tt>weight_decay</tt></a> arguments. Momentum allows the algorithm to "roll over" small bumps to avoid local minima that can cause convergence too soon. Weight decay (also called an L2 penalty) applies to biases.
#
# For more information, see <a href='https://pytorch.org/docs/stable/optim.html'><strong><tt>torch.optim</tt></strong></a>
# +
optimizer = torch.optim.SGD(model.parameters(), lr = 0.001)
# You'll sometimes see this as
# optimizer = torch.optim.SGD(model.parameters(), lr = 1e-3)
# -
# ## Train the model
# An <em>epoch</em> is a single pass through the entire dataset. We want to pick a sufficiently large number of epochs to reach a plateau close to our known parameters of $\mathrm {weight} = 2,\; \mathrm {bias} = 1$
# <div class="alert alert-info"><strong>Let's walk through the steps we're about to take:</strong><br>
#
# 1. Set a reasonably large number of passes<br>
# <tt><font color=black>epochs = 50</font></tt><br>
# 2. Create a list to store loss values. This will let us view our progress afterward.<br>
# <tt><font color=black>losses = []</font></tt><br>
# <tt><font color=black>for i in range(epochs):</font></tt><br>
# 3. Bump "i" so that the printed report starts at 1<br>
# <tt><font color=black> i+=1</font></tt><br>
# 4. Create a prediction set by running "X" through the current model parameters<br>
# <tt><font color=black> y_pred = model.forward(X)</font></tt><br>
# 5. Calculate the loss<br>
# <tt><font color=black> loss = criterion(y_pred, y)</font></tt><br>
# 6. Add the loss value to our tracking list<br>
# <tt><font color=black> losses.append(loss)</font></tt><br>
# 7. Print the current line of results<br>
# <tt><font color=black> print(f'epoch: {i:2} loss: {loss.item():10.8f}')</font></tt><br>
# 8. Gradients accumulate with every backprop. To prevent compounding we need to reset the stored gradient for each new epoch.<br>
# <tt><font color=black> optimizer.zero_grad()</font></tt><br>
# 9. Now we can backprop<br>
# <tt><font color=black> loss.backward()</font></tt><br>
# 10. Finally, we can update the hyperparameters of our model<br>
# <tt><font color=black> optimizer.step()</font></tt>
# </div>
# +
epochs = 50
losses = []
for i in range(epochs):
i+=1
y_pred = model.forward(X)
loss = criterion(y_pred, y)
losses.append(loss)
print(f'epoch: {i:2} loss: {loss.item():10.8f} weight: {model.linear.weight.item():10.8f} \
bias: {model.linear.bias.item():10.8f}')
optimizer.zero_grad()
loss.backward()
optimizer.step()
# -
# ## Plot the loss values
# Let's see how loss changed over time
plt.plot(range(epochs), losses)
plt.ylabel('Loss')
plt.xlabel('epoch');
# ## Plot the result
# Now we'll derive <tt>y1</tt> from the new model to plot the most recent best-fit line.
# +
w1,b1 = model.linear.weight.item(), model.linear.bias.item()
print(f'Current weight: {w1:.8f}, Current bias: {b1:.8f}')
print()
y1 = x1*w1 + b1
print(x1)
print(y1)
# -
plt.scatter(X.numpy(), y.numpy())
plt.plot(x1,y1,'r')
plt.title('Current Model')
plt.ylabel('y')
plt.xlabel('x');
# ## Great job!
|
01-Linear-Regression-with-PyTorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## Series
import pandas as pd
import numpy as np
series1 = pd.Series(['a', 'b', 'c'])
print(series1[1:3])
# +
# Viewing Specific Keys
record = pd.Series({
'firstname': 'Charles',
'lastname': 'Givre',
'middle': 'classfied'
})
record[['firstname', 'lastname']]
# firstname Charles
# lastname Givre
# String manipulations: Show True/False for each record
record.str.contains('Cha')
# firstname True
# lastname False
# middle False
# Filter based on the returned True/False
record[record.str.contains('Cha')]
# Other string manipulations
# Record.str.contains... functions
# => contains, count, extract, find, findall, len
# +
# Display head/tail number of records
randomNumbers = pd.Series(
np.random.randint(1, 100, 50)
)
randomNumbers.head(10)
# randomNumbers.tail(7)
# -
# Filtering Data in a Series
# 1. Generate True/False values for all values
randomNumbers < 10
# 2. Filter based on True/False values and only show records
randomNumbers[randomNumbers < 10]
small_rands = randomNumbers[randomNumbers < 10]
small_rands
def addTwo(n):
return n + 2
# iterates through the entire series and apply addTwo function
small_rands.apply(addTwo)
# Use lambda instead
small_rands.apply(lambda x: x + 1)
# +
# Remove Missing Values:
Series.dropna()
# Replace missing values with "something" value
Series.fillna(value="<something>"
# -
#
# ---
#
# ## DataFrame
# +
data = pd.DataFrame(<data>, <index>, <column_names>)
# 1. Pass two dimentional data -> Series
# 2. Usually, reading from outside sources
data = pd.read_csv(<file>)
data = pd.read_excel('file.xls')
data = pd.read_json(<file>/<url>)
data = pd.read_sql(<query>, <connection_obj>)
data = pd.read_html(<source>)
logdf = pd.read_table('../data/mysql.log', names=['raw'])
# refer to a column:
logdf['raw'].str.extract(
'(?P<data>]d{6}\s\d{2}:\d{2}:\d{2}:\d{2})...',
expand=False
)
# -
# Web Server Logs
# 1. Complicated to parse - use apache_log_parser package
import apache_log_parser
line_parser = apache_log_parser.make_parser("%h %l %u %t \'%r\' %>s %b \'%{Referer}i\' \'%{User-agent}i\' ")
# pandas are moving to arrow data structure
server_log = open("../data/hackers-access.httpd", "r")
parsed_server_data = []
# loop and add to parsed_server_data and move into a dataframe
# +
# Manipulating datafarme
df = data['column'] # returns series
df['ip'].value_counts().head() # counts unique IPs!
# return the following columns
df = data[['column1', 'column2', 'column3']] # returns a DataFrame
# extract columns and filter
# 1. show specified columns
# 2. Filter(any column)
df[['col1', 'col2']][col3 > 5]
# pull out individual rows
data.loc[<index>]
data.loc[<list of indexes>]
data.sample[<n>] # return a random sample of a dataset
# Apply to DataFrame
data.apply(<function>)
# - function will receive Series == each row
# - function will return a new row => allows us to add new columns
# -
# Apply a function to a column and then create a new column
df = pd.read_csv('data/dailybots.csv')
# df['orgs'] + 2 => will add 2 to the entire column
df['orgs2'] = df['orgs'] + 2 # this will create a new column
# +
# Transpose
data.T => Reshaping data
# aggregation
# 1. the sum of columns
data.sum(axis=0)
# 2. the sum of the rows
data.sum(axis=1) # from operating on a column to row
# drop
# 1. inplace => change the current dataframe
# 2. errors => specify an error
data.drop(labels, axis=0, level=None, inplace=False, errors='raise')
# +
# Merging datasets
# Union
Series 1 + Series 2
combinedSeries = pd.concat([series1, series2], ...)
# Join
# 1. Inner Join -> common things between sets
# 2. Outer Join ->
# 3. Left Join -> all the data in set A, A and B, but not B
# 4. Right Join -> same
pd.merge(leftData, rightData,
how="<join type / inner,lerft,right,outer>",
on="list of fields")
# +
# Grouping and Aggregating data
df_grouped = df.groupby(
['Protocol', 'Source', 'Destination']
)
print(df_grouped.size())
stats_packets = df_grouped['Length'].agg({'No Packets': len, 'Volume': sum,
'SD': lambda x: np.std(x, ddoff=1...)})
|
1m_ML_Security/notebooks/day_1/day_1_concepts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3.7 `while` Statement
product = 3
while product <= 50:
product = product * 3
product
##########################################################################
# (C) Copyright 2019 by Deitel & Associates, Inc. and #
# Pearson Education, Inc. All Rights Reserved. #
# #
# DISCLAIMER: The authors and publisher of this book have used their #
# best efforts in preparing the book. These efforts include the #
# development, research, and testing of the theories and programs #
# to determine their effectiveness. The authors and publisher make #
# no warranty of any kind, expressed or implied, with regard to these #
# programs or to the documentation contained in these books. The authors #
# and publisher shall not be liable in any event for incidental or #
# consequential damages in connection with, or arising out of, the #
# furnishing, performance, or use of these programs. #
##########################################################################
|
examples/ch03/snippets_ipynb/03_07.ipynb
|