code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning with MNIST Dataset using Tensorflow and Keras
#
# MNIST datatset:
# MNIST datatset is a dataset of hand-written digits, 0 - 9. It contains 28x28 images of these hand-written digits.
#import necessary libraries
import matplotlib.pyplot as plt
import tensorflow as tf
import keras
print(tf.__version__)
print(keras.__version__)
#Mnist dataset
mnist_data = keras.datasets.mnist
#Load the data from dataset
(xtrain, ytrain), (xtest, ytest) = mnist_data.load_data()
#Ananlyse the data
print(xtrain[1])
# +
plt.imshow(xtrain[1])
# -
plt.imshow(xtrain[1], cmap = plt.cm.binary)
plt.show()
#Normalizing values
xtrain = keras.utils.normalize(xtrain, axis=1)
xtest = keras.utils.normalize(xtest, axis=1)
plt.imshow(xtrain[1], cmap = plt.cm.binary)
plt.show()
xtrain[1]
from keras.models import Sequential
#Sequential Model
#Feed forward
model = Sequential()
# +
#We need to flatten array, we don't need 28x28
model.add(keras.layers.Flatten())
model.add(keras.layers.Dense(128, activation = 'relu'))
model.add(keras.layers.Dense(128, activation = 'relu'))
model.add(keras.layers.Dense(10, activation = 'softmax'))
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy',metrics=['accuracy'])
h = model.fit(xtrain,ytrain,epochs=5)
# -
# We are getting an accuracy of 98% after 5 epoch, accuracy increased with epoch and loss went down.
print(h.history.keys())
val_loss, val_accuracy = model.evaluate(xtest,ytest)
print(val_loss)
print(val_accuracy)
# ## Saving the Model and Loading it for making predictions
#Save the model
model.save('number_reader.model')
#Loading the Model
new_model = keras.models.load_model('number_reader.model')
#Predicting
preds = new_model.predict(xtest)
print(preds)
import numpy as np
np.argmax(preds[1])
#Plotting the orginal tes
plt.imshow(xtest[1])
plt.show()
|
DL_mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import glob
import pickle
import re
import numpy as np
import pandas as pd
from random import shuffle
from tensorflow.contrib import learn
# +
def clean_str(string):
"""
Tokenization/string cleaning for all datasets except for SST.
Original taken from https://github.com/yoonkim/CNN_sentence/blob/master/process_data.py
"""
string = re.sub(r"[^A-Za-z0-9(),!?\'\`]", " ", string)
string = re.sub(r"\'s", " \'s", string)
string = re.sub(r"\'ve", " \'ve", string)
string = re.sub(r"n\'t", " n\'t", string)
string = re.sub(r"\'re", " \'re", string)
string = re.sub(r"\'d", " \'d", string)
string = re.sub(r"\'ll", " \'ll", string)
string = re.sub(r",", " , ", string)
string = re.sub(r"!", " ! ", string)
string = re.sub(r"\(", " \( ", string)
string = re.sub(r"\)", " \) ", string)
string = re.sub(r"\?", " \? ", string)
string = re.sub(r"\s{2,}", " ", string)
string = re.sub(r"<br />", " ", string)
return string.strip().lower()
def get_reviews(path, clean = True):
complete_path = path + '/*.txt'
files = glob.glob(complete_path)
reviews = [str(open(rev).readlines()[0]).strip() for rev in files]
if clean:
reviews = [clean_str(rev) for rev in reviews]
return reviews
# +
# Gets all the reviews
train_positive_reviews = get_reviews("data/aclImdb/train/pos")
train_negative_reviews = get_reviews("data/aclImdb/train/neg")
test_positive_reviews = get_reviews("data/aclImdb/test/pos")
test_negative_reviews = get_reviews("data/aclImdb/test/neg")
# Divide The train set into train and validation
# Concat all train reviews and write it on a file
train_reviews = train_positive_reviews + train_negative_reviews
output_train = open('data/all_train.txt', 'w')
for rev in train_reviews:
print>>output_train, rev
output_train.close()
# -
# Saves the Train/Test lists into pickle objects
pickle.dump(train_positive_reviews, open( "data/train_pos.p", "wb" ))
pickle.dump(train_negative_reviews, open( "data/train_neg.p", "wb" ))
pickle.dump(test_positive_reviews, open( "data/test_pos.p", "wb" ))
pickle.dump(test_negative_reviews, open( "data/test_neg.p", "wb" ))
# Loads the Train/Test objects
train_positive_reviews = pickle.load(open("data/train_pos.p","rb"))
train_negative_reviews = pickle.load(open("data/train_neg.p","rb"))
test_positive_reviews = pickle.load(open("data/test_pos.p","rb"))
test_negative_reviews = pickle.load(open("data/test_neg.p","rb"))
# +
def get_train_sets():
train_positive_reviews = pickle.load(open("data/train_pos.p","rb"))
train_negative_reviews = pickle.load(open("data/train_neg.p","rb"))
return train_positive_reviews, train_negative_reviews
def get_test_sets():
test_positive_reviews = pickle.load(open("data/test_pos.p","rb"))
test_negative_reviews = pickle.load(open("data/test_neg.p","rb"))
return test_positive_reviews, test_negative_reviews
def label_data(positive_revs, negative_revs):
# Generate the labels
positive_labels = [[0, 1] for _ in positive_revs]
negative_labels = [[1, 0] for _ in negative_revs]
# Concatenates the positive and negative labels for train and val
y_labels = np.concatenate([positive_labels, negative_labels], 0)
x_train = positive_revs + negative_revs
return [x_train, y_labels]
def __split_train_validation(x_train, y_train, amount_val=.25):
x_train = np.array(x_train)
y_train = np.array(y_train)
# Randomly shuffle data
np.random.seed(10)
shuffle_indices = np.random.permutation(np.arange(len(y_train)))
print (shuffle_indices)
x_shuffled = x_train[shuffle_indices]
y_shuffled = y_train[shuffle_indices]
total_reviews = len(x_shuffled)
training_num = total_reviews - int(total_reviews * amount_val)
x_t = x_shuffled[:training_num]
y_t = y_shuffled[:training_num]
x_dev = x_shuffled[training_num:]
y_dev = y_shuffled[training_num:]
return [x_t, y_t], [x_dev, y_dev]
def get_train_validation(train_pos, train_neg, amount_val=.25):
# Divides the sets
total_reviews = len(train_pos)
print("Num Total Reviews in set:", total_reviews)
training_num = total_reviews - int(total_reviews * amount_val)
print("Num Training Reviews:", training_num)
train_pos_reviews_t = train_pos[:training_num]
train_neg_reviews_t = train_neg[:training_num]
train_pos_reviews_v = train_pos[training_num:]
train_neg_reviews_v = train_neg[training_num:]
# Generate the labels
train_positive_labels = [[0, 1] for _ in train_pos_reviews_t]
val_positive_labels = [[0, 1] for _ in train_pos_reviews_v]
train_negative_labels = [[1, 0] for _ in train_neg_reviews_t]
val_negative_labels = [[1, 0] for _ in train_neg_reviews_v]
# Concatenates the positive and negative labels for train and val
y_train = np.concatenate([train_positive_labels, train_negative_labels], 0)
y_val = np.concatenate([val_positive_labels, val_negative_labels], 0)
# Creates one list for positive and negative reviews
x_train = train_pos_reviews_t + train_neg_reviews_t
x_val = train_pos_reviews_v + train_neg_reviews_v
print("x_train:", len(x_train))
print("y_train:", len(y_train))
print("x_val:", len(x_val))
print("y_val:", len(y_val))
return [x_train, y_train],[x_val, y_val]
def get_test_labeled(test_pos, test_neg):
# Generate the labels
test_positive_labels = [[0, 1] for _ in test_pos]
test_negative_labels = [[1, 0] for _ in test_neg]
y = np.concatenate([test_positive_labels, test_negative_labels], 0)
x_test = test_pos + test_neg
return [x_test, y]
#train, validation = get_train_validation(train_positive_reviews, train_negative_reviews)
x_t, y_t = label_data(train_positive_reviews, train_negative_reviews)
# -
# Label the data
x_train, y_train = label_data(train_positive_reviews, train_negative_reviews)
# Separates in Train and Dev
x_train_list, x_dev_list = split_train_validation(x_train, y_train)
# +
# Shuffle the data
def split_train_validation(x_train, y_train, amount_val=.25):
x_train_shuffled = []
y_train_shuffled = []
np.random.seed(10)
shuffle_indices = np.random.permutation(np.arange(len(y_train)))
for i in shuffle_indices:
x_train_shuffled.append(x_train[i])
y_train_shuffled.append(y_train[i])
total_reviews = len(x_train_shuffled)
training_num = total_reviews - int(total_reviews * amount_val)
x_t = x_train_shuffled[:training_num]
y_t = y_train_shuffled[:training_num]
x_dev = x_train_shuffled[training_num:]
y_dev = y_train_shuffled[training_num:]
return [x_t, y_t], [x_dev, y_dev]
# Separates in Train and Dev
x_train_list, x_dev_list = split_train_validation(x_t, y_t)
print(len(x_train_list[0]))
print(len(x_train_list[1]))
# -
print(x_dev_list[0][1])
print(x_dev_list[1][1])
# Loads the vocabulary
def load_vocabulary(file_path, num_words=10000):
with open(file_path) as vocab:
vocab_list = [next(vocab) for x in range(num_words)]
vocab_list = [str(vocab).strip() for vocab in vocab_list]
return vocab_list
#
#load_vocabulary("data/vocab_unigrams_no_counts/part-00000")
# +
# Spark Unigrams
text_file = sc.textFile('all_train.txt')
counts = text_file.flatMap(lambda line: line.split(" ")).map(lambda word:(word, 1)).reduceByKey(lambda a, b: a+b).sortBy(lambda a: -a[1])
# Comment this line, if you want tuples
just_words = counts.map(lambda tuple: tuple[0])
just_words.saveAsTextFile("vocab_unigrams_no_counts")
# Spark Bi-grams
bigrams = text_file.map(lambda x:x.split()).flatMap(lambda x: [((x[i],x[i+1]),1) for i in range(0,len(x)-1)])
count_bigrams = bigrams.reduceByKey(lambda x, y: x+y).sortBy(lambda a: -a[1])
just_bigrams = count_bigrams.map(lambda tuple: tuple[0][0] + ' ' + tuple[0][1])
just_bigrams.saveAsTextFile("vocab_bigrams_no_counts")
just_bigrams.saveAsTextFile("vocab_oov_bigrams_no_counts")
# +
# This is a test for the vocabulary
vocabulary = load_vocabulary("data/vocab_unigrams_no_counts/part-00000")
vocabulary = [str(vocab).strip() for vocab in vocabulary]
vocabulary[:5]
max_len_vocabulary = len(vocabulary)
print (max_len_vocabulary)
# -
train_reviews = train_positive_reviews + train_negative_reviews
print(len(train_reviews))
# +
def set_oov(reviews, vocabulary):
updated_reviews = []
for review in reviews:
up_review = []
splitted_review = review.split(" ")
for i, word in enumerate(splitted_review):
if word not in vocabulary:
splitted_review[i] = 'oov'
else:
splitted_review[i] = word
new_review = (' ').join(splitted_review)
updated_reviews.append(new_review)
return updated_reviews
def set_oov_tag(reviews, vocabulary):
updated_reviews = []
set_vocabulary = set(vocabulary)
for review in reviews:
set_review = set(review.split(" "))
oov_words = set_review - set_vocabulary
#print(list(oov_words))
dic_oov_words = {k:'oov' for k in oov_words}
#print(dic_oov_words)
if len(dic_oov_words) >= 1:
rep = dict((re.escape(k), v) for k, v in dic_oov_words.items())
pattern = re.compile("|".join(rep.keys()))
oov_review = pattern.sub(lambda m: rep[re.escape(m.group(0))], review)
updated_reviews.append(oov_review)
else:
updated_reviews.append(review)
return updated_reviews
oov_reviews = set_oov(train_reviews, vocabulary)
#print(len(new_reviews))
# -
print(len(oov_reviews))
super_review = ' '.join(oov_reviews)
# +
# Prepares Train/Dev for FaceBook FastText
# Loads the Data
train_positive_reviews = pickle.load(open("data/train_pos.p","rb"))
train_negative_reviews = pickle.load(open("data/train_neg.p","rb"))
# For each review append the label
train_pos_reviews_labeled = [x + ' __label__1' for x in train_positive_reviews]
train_neg_reviews_labeled = [x + ' __label__0' for x in train_negative_reviews]
# +
fb_reviews = train_pos_reviews_labeled + train_neg_reviews_labeled
shuffle(fb_reviews)
print(fb_reviews[0])
with open('fastText/fb_train_shuffled.txt', mode='wt', encoding='utf-8') as output_fb_train:
output_fb_train.write('\n'.join(fb_reviews))
# +
# Prepares Test for Facebook FastText
test_positive_reviews = pickle.load(open("data/test_pos.p","rb"))
test_negative_reviews = pickle.load(open("data/test_neg.p","rb"))
# For each review append the label
test_pos_reviews_labeled = [x + ' __label__1' for x in test_positive_reviews]
test_neg_reviews_labeled = [x + ' __label__0' for x in test_negative_reviews]
fb_test_reviews = test_pos_reviews_labeled + test_neg_reviews_labeled
shuffle(fb_test_reviews)
with open('fastText/fb_test_shuffled.txt', mode='wt', encoding='utf-8') as output_fb_test:
output_fb_test.write('\n'.join(fb_test_reviews))
# +
train_positive_reviews = get_reviews("data/aclImdb/train/pos", clean=False)
train_negative_reviews = get_reviews("data/aclImdb/train/neg", clean=False)
test_positive_reviews = get_reviews("data/aclImdb/test/pos", clean=False)
test_negative_reviews = get_reviews("data/aclImdb/test/neg", clean=False)
from random import shuffle
# For each review append the label
train_pos_reviews_labeled = [x + ' __label__1' for x in train_positive_reviews]
train_neg_reviews_labeled = [x + ' __label__0' for x in train_negative_reviews]
fb_reviews = train_pos_reviews_labeled + train_neg_reviews_labeled
shuffle(fb_reviews)
with open('fastText/fb_train_unclean_shuffled.txt', mode='wt', encoding='utf-8') as output_fb_train:
output_fb_train.write('\n'.join(fb_reviews))
#=============================
# For each review append the label
test_pos_reviews_labeled = [x + ' __label__1' for x in test_positive_reviews]
test_neg_reviews_labeled = [x + ' __label__0' for x in test_negative_reviews]
fb_test_reviews = test_pos_reviews_labeled + test_neg_reviews_labeled
shuffle(fb_test_reviews)
with open('fastText/fb_test_unclean_shuffled.txt', mode='wt', encoding='utf-8') as output_fb_test:
output_fb_test.write('\n'.join(fb_test_reviews))
fb_test_reviews[0]
# -
# BIGRAMS
x_train_reviews_oov = pickle.load(open("data/reviews_oov.p", "rb"))
# Set this to file
with open('data/reviews_oov.txt', mode='wt', encoding='utf-8') as output_reviews_oov:
output_reviews_oov.write('\n'.join(x_train_reviews_oov))
# +
rev_test = x_train_reviews_oov[0]
# Loads vocab
bi_vocabulary = load_vocabulary("data/vocab_oov_bigrams_no_counts/part-00000")
def find_bigrams(review, vocabulary):
split_review = review.split(' ')
zipped = zip(split_review, split_review[1:])
bigrams = [x[0] + '_' + x[1] if x[0] + ' ' + x[1] in vocabulary else 'oov' for x in zipped]
print(len(bigrams))
return ' '.join(bigrams)
#[find_bigrams(rev, bi_vocabulary) for rev in x_train_reviews_oov]
find_bigrams(rev_test, bi_vocabulary)
# -
vp = learn.preprocessing.VocabularyProcessor(10)
list(vp.fit_transform(["a", "dog" , "ran" ,"in" ,"the", "park"]))
# +
x = [[1,2,3], [4,5,6]]
y = [['a', 'b', 'c'], ['d','e','f']]
zipped = zip(x,y)
final_revs = [x[0]+x[1] for x in zipped]
final_revs
|
.ipynb_checkpoints/Prev Data-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from civicpy import civic, version
version()
# ## Exploring CIViC Variant 12
# Get a Variant object by its CIViC Variant ID
variant = civic.get_variant_by_id(12)
variant
# Explore details of the Variant object, and related Gene object
(variant.gene.name, variant.name)
# A list of all evidence associated with this variant
variant.evidence
# A CIViC Assertion tied to this variant, and details corresponding to the Assertion
print(f'There are {len(variant.assertions)} assertions for this variant.')
assertion = variant.assertions[0]
print(f'Description of CIViC Assertion {assertion.id}:\n')
print(assertion.description)
# Collecting all evidence, and refining on evidence status
all_evidence = civic.get_all_evidence()
submitted_and_accepted_evidence = civic.get_all_evidence(include_status=['accepted', 'submitted'])
len(all_evidence) > len(submitted_and_accepted_evidence)
# A list of attributes for any CivicRecord object can be found using the Python dir() built-in.
dir(variant)
# ## Exporting to VCF
from civicpy.exports import VCFWriter
with open("example.vcf", "w") as fh:
writer = VCFWriter(fh)
writer.addrecord(variant)
writer.writerecords()
with open("example.vcf", "r") as fh:
for line in fh:
print(line)
|
examples/basic_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pathfinder Application (Wind and Light)
#
# Author: <NAME> (email: <EMAIL>)
#
# ### Feedback
# Questions, comments, suggestions, or requests for functionality are welcome and can be sent to the email address above. This tool will continue development on an 'as-required' basis (i.e. I will add features when I need them, or when somebody asks me for them)!
#
# ### What is this?
# This is a streamlined version of the Pathfinder application which provides easy configuration for a single light cue and a single wind cue to match the current experimental setup. This version does **not** require any use of configuration files, everything can be done from within the notebook (though configuration is more limited). If you want to create more general/complex simulation scenarios, please see the 'General' notebook.
#
# The software mimics the tests from the sky-compass literature whereby the beetle is placed in the centre of an arena, presented with cues, and allowed to roll to the edge of an arena; the cues may then be changed for the second roll and the absolute change in bearing recorded.
#
# ### Usage
# Start by running the code cell below, this will initialise the software and generate a series of graphical controls. You can use the controls to configure the cues. Once you've configured the software, click 'Run Interact' to generate a plot. If you want to change the scenario, simply modify the configuration using the graphical controls and click 'Run Interact' again. A more detailed control reference can be found below but they should be largely self explanatory.
# Run this cell!
# %matplotlib notebook
from pathfinder.runnable.wind_and_light import generate_controls
from IPython.display import display
controls = generate_controls()
display(controls)
# ### Control reference
# #### Switches/checkboxes
# Three checkboxes are provided:
# * Show individual cues: when enabled this will show the directional reference given by each individual cue (this can be a nice way of visualising the relative weight of each cue).
# * Enable/disable the legend: the legend positioning is not consistent as the plots are resized so they can occassionally obscure the plot. Checking this box will turn the legend on. I find the legends useful for reference but annoying once you know what the plot is showing.
# * Show sensory vectors: sensory vectors are the true geometric representation of the cues which indicate how the simulated beetle perceives the cues. These can be visualised by enabling this setting. For more information please see the General notebook (section "How does the beetle get its bearing?").
#
# #### Generic settings
# * Combination strategy: this defines the method used to combine the cues; currently implemented are:
# * avg: simply take the average; and,
# * wta: winner take all, the strongest cue wins (no decision can be made if there are multiple strongest).
# * proj_wta: projected winner take all. The cue with the strongest sensory vector after projection into the ground plane. This accounts for a perceived strength difference of lights at different elevations.
#
# There are any number of strategies that could be implemented, these are simply the ones that I added during development. Again, see "How does the beetle get its bearing?" in the General notebook for more information.
#
# * Confidence threshold: this represents the minimum magnitude required from the combined cue vector. If the magnitude of the combined cue is less than this threshold we can assume that the beetle would not actually have enough information to orient itself and discard the change in bearing. The default value was chosen based on the contrast experiments from "Stellar performance: Mechanisms underlying milky way orientation in dung beetles" (*Foster et al. 2017*).
#
# #### Cue setings
# Finally we provide individual cue configuration for each roll of the beetle. Each roll has a single light cue and some wind. For the light we can alter the strength, elevation, and azimuth. For the wind we can alter its strength and direction.
|
Wind and Light.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # `GiRaFFE_NRPy`: Main Driver
#
# ## Author: <NAME>
#
# <a id='intro'></a>
#
# **Notebook Status:** <font color=Red><b> Validation in progress </b></font>
#
# **Validation Notes:** This code assembles the various parts needed for GRFFE evolution in order.
#
# ### NRPy+ Source Code for this module:
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Main_Driver.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Main_Driver.py)
#
# ### Other critical files (in alphabetical order):
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Afield_flux_handwritten.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Afield_flux_handwritten.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Afield_flux_handwritten.ipynb) Generates the expressions to find the flux term of the induction equation.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-A2B.ipynb) Generates the driver to compute the magnetic field from the vector potential/
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-BCs.ipynb) Generates the code to apply boundary conditions to the vector potential, scalar potential, and three-velocity.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-C2P_P2C.ipynb) Generates the conservative-to-primitive and primitive-to-conservative solvers.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Metric_Face_Values.ipynb) Generates code to interpolate metric gridfunctions to cell faces.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-PPM.ipynb) Genearates code to reconstruct primitive variables on cell faces.
# * [GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Source_Terms.ipynb) Genearates code to compute the $\tilde{S}_i$ source term.
# * [GiRaFFE_NRPy/Stilde_flux.py](../../edit/in_progress/GiRaFFE_NRPy/Stilde_flux.py) [\[**tutorial**\]](Tutorial-GiRaFFE_NRPy-Stilde-flux.ipynb) Generates the expressions to find the flux term of the Poynting flux evolution equation.
# * [../GRFFE/equations.py](../../edit/GRFFE/equations.py) [\[**tutorial**\]](../Tutorial-GRFFE_Equations-Cartesian.ipynb) Generates code necessary to compute the source terms.
# * [../GRHD/equations.py](../../edit/GRHD/equations.py) [\[**tutorial**\]](../Tutorial-GRHD_Equations-Cartesian.ipynb) Generates code necessary to compute the source terms.
#
# ## Introduction:
# Having written all the various algorithms that will go into evolving the GRFFE equations forward through time, we are ready to write a start-to-finish module to do so. However, to help keep things more organized, we will first create a dedicated module to assemble the various functions we need to run, in order, to perform the evolution. This will reduce the length of the standalone C code, improving that notebook's readability.
#
# <a id='prelim'></a>
# # Table of Contents
# $$\label{prelim}$$
#
# During a given RK substep, we will perform the following steps in this order, based on the order used in the original `GiRaFFE`:
# 0. [Step 0](#prelim): Preliminaries
# 1. [Step 1](#rhs): Calculate the right-hand sides
# 1. [Step 1.a](#operand): Calculate the portion of the gauge terms for $A_k$, $(\alpha \Phi - \beta^j A_j)$ and $\Phi$, $(\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi])$ *inside* the parentheses to be finite-differenced.
# 1. [**GRFFE/equations.py**](../../edit/GRFFE/equations.py), [**GRHD/equations.py**](../../edit/GRHD/equations.py)
# 1. [Step 1.b](#source): Calculate the source terms of $\partial_t A_i$, $\partial_t \tilde{S}_i$, and $\partial_t [\sqrt{\gamma} \Phi]$ right-hand sides
# 1. [**GRFFE/equations.py**](../../edit/GRFFE/equations.py), [**GRHD/equations.py**](../../edit/GRHD/equations.py), [**GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Source_Terms.py)
# 1. [Step 1.c](#flux): Calculate the Flux terms
# 1. In each direction:
# 1. Interpolate the metric gridfunctions to cell faces
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Metric_Face_Values.py)
# 1. Reconstruct primitives $\bar{v}^i$ and $B^i$ on cell faces with the piecewise-parabolic method
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_PPM.py)
# 1. Compute the fluxes of $\tilde{S}_i$ and $A_i$ and add the appropriate combinations to the evolution equation right-hand sides
# 1. [**GiRaFFE_NRPy/Stilde_flux.py**](../../edit/in_progress/GiRaFFE_NRPy/Stilde_flux.py), [**GiRaFFE_NRPy/GiRaFFE_NRPy_Afield_flux_handwritten.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Afield_flux_handwritten.py)
# 1. [Step 2](#poststep): Recover the primitive variables and apply boundary conditions (post-step)
# 1. [Step 2.a](#potential_bc): Apply boundary conditions to $A_i$ and $\sqrt{\gamma} \Phi$
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py)
# 1. [Step 2.b](#a2b): Compute $B^i$ from $A_i$
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_A2B.py)
# 1. [Step 2.c](#c2p): Run the Conservative-to-Primitive solver
# 1. This applies fixes to $\tilde{S}_i$, then computes $\bar{v}^i$. A current sheet prescription is then applied to $\bar{v}^i$, and $\tilde{S}_i$ is recomputed to be consistent.
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_C2P_P2C.py)
# 1. [Step 2.d](#velocity_bc): Apply outflow boundary conditions to $\bar{v}^i$
# 1. [**GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py**](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_BCs.py)
# 1. [Step 3](#write_out): Write out the C code function
# 1. [Step 3](#code_validation): Self-Validation against `GiRaFFE_NRPy_Main_Drive.py`
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
#
# <a id='prelim'></a>
#
# # Step 0: Preliminaries \[Back to [top](#toc)\]
# $$\label{prelim}$$
#
# We begin by importing the NRPy+ core functionality. We also import the GRHD module and the GRFFE module.
# +
# Step 0: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
from outputC import outCfunction, lhrh # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import loop as lp # NRPy+: Generate C code loops
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
thismodule = "GiRaFFE_NRPy_Main_Driver"
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",2)
out_dir = os.path.join("GiRaFFE_standalone_Ccodes")
cmd.mkdir(out_dir)
CoordSystem = "Cartesian"
par.set_parval_from_str("reference_metric::CoordSystem",CoordSystem)
rfm.reference_metric() # Create ReU, ReDD needed for rescaling B-L initial data, generating BSSN RHSs, etc.
# Default Kreiss-Oliger dissipation strength
default_KO_strength = 0.1
diss_strength = par.Cparameters("REAL", thismodule, "diss_strength", default_KO_strength)
outCparams = "outCverbose=False,CSE_sorting=none"
# -
# <a id='rhs'></a>
#
# # Step 1: Calculate the right-hand sides \[Back to [top](#toc)\]
# $$\label{rhs}$$
#
# <a id='operand'></a>
#
# In the method of lines using Runge-Kutta methods, each timestep involves several "RK substeps" during which we will run the same set of function calls. These can be divided into two groups: one in which the RHSs themselves are calculated, and a second in which boundary conditions are applied and auxiliary variables updated (the post-step). Here, we focus on that first group.
#
# ## Step 1.a: Calculate the portion of the gauge terms for $A_k$, $(\alpha \Phi - \beta^j A_j)$ and $\Phi$, $(\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi])$ *inside* the parentheses to be finite-differenced. \[Back to [top](#toc)\]
# $$\label{operand}$$
#
# The gauge terms of our evolution equations consist of two derivative terms: the Lorentz gauge term of $\partial_t A_k$, which is $\partial_k (\alpha \Phi - \beta^j A_j)$ and the non-damping, flux-like term of $\partial_t [\psi^6 \Phi]$, which is $\partial_j (\alpha\sqrt{\gamma}A^j - \beta^j [\sqrt{\gamma} \Phi])$. We can save some effort and execution time (at the cost of memory needed) by computing the derivative operands, storing them, and then finite-differencing that stored variable. For more information, see the notebook for the [implementation](Tutorial-GiRaFFE_NRPy-Source_Terms.ipynb) and the [validation](Tutorial-Start_to_Finish_UnitTest-GiRaFFE_NRPy-Source_Terms.ipynb), as well as [Tutorial-GRFFE_Equations-Cartesian](../Tutorial-GRFFE_Equations-Cartesian.ipynb) and [Tutorial-GRHD_Equations-Cartesian](../Tutorial-GRHD_Equations-Cartesian.ipynb) for the terms themselves.
# +
import GRHD.equations as GRHD # NRPy+: Generate general relativistic hydrodynamics equations
import GRFFE.equations as GRFFE # NRPy+: Generate general relativisitic force-free electrodynamics equations
gammaDD = ixp.register_gridfunctions_for_single_rank2("AUXEVOL","gammaDD","sym01",DIM=3)
betaU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","betaU",DIM=3)
alpha = gri.register_gridfunctions("AUXEVOL","alpha")
AD = ixp.register_gridfunctions_for_single_rank1("EVOL","AD")
BU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","BU")
ValenciavU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","ValenciavU")
psi6Phi = gri.register_gridfunctions("EVOL","psi6Phi")
StildeD = ixp.register_gridfunctions_for_single_rank1("EVOL","StildeD")
ixp.register_gridfunctions_for_single_rank1("AUXEVOL","PhievolParenU",DIM=3)
gri.register_gridfunctions("AUXEVOL","AevolParen")
GRHD.compute_sqrtgammaDET(gammaDD)
GRFFE.compute_AD_source_term_operand_for_FD(GRHD.sqrtgammaDET,betaU,alpha,psi6Phi,AD)
GRFFE.compute_psi6Phi_rhs_flux_term_operand(gammaDD,GRHD.sqrtgammaDET,betaU,alpha,AD,psi6Phi)
parens_to_print = [\
lhrh(lhs=gri.gfaccess("auxevol_gfs","AevolParen"),rhs=GRFFE.AevolParen),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","PhievolParenU0"),rhs=GRFFE.PhievolParenU[0]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","PhievolParenU1"),rhs=GRFFE.PhievolParenU[1]),\
lhrh(lhs=gri.gfaccess("auxevol_gfs","PhievolParenU2"),rhs=GRFFE.PhievolParenU[2]),\
]
subdir = "RHSs"
cmd.mkdir(os.path.join(out_dir, subdir))
desc = "Calculate quantities to be finite-differenced for the GRFFE RHSs"
name = "calculate_AD_gauge_term_psi6Phi_flux_term_for_RHSs"
outCfunction(
outfile = os.path.join(out_dir,subdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *restrict params,const REAL *restrict in_gfs,REAL *restrict auxevol_gfs",
body = fin.FD_outputC("returnstring",parens_to_print,params=outCparams),
loopopts ="AllPoints",
rel_path_to_Cparams=os.path.join("../"))
# -
# <a id='source'></a>
#
# ## Step 1.b: Calculate the source terms of $\partial_t A_i$, $\partial_t \tilde{S}_i$, and $\partial_t [\sqrt{\gamma} \Phi]$ right-hand sides \[Back to [top](#toc)\]
# $$\label{source}$$
#
# With the operands of the gradient of divergence operators stored in memory from the previous step, we can now calculate the terms on the RHS of $A_i$ and $[\sqrt{\gamma} \Phi]$ that involve the derivatives of those terms. We also compute the other term in the RHS of $[\sqrt{\gamma} \Phi]$, which is a straightforward damping term.
# +
xi_damping = par.Cparameters("REAL",thismodule,"xi_damping",0.1)
GRFFE.compute_psi6Phi_rhs_damping_term(alpha,psi6Phi,xi_damping)
AevolParen_dD = ixp.declarerank1("AevolParen_dD",DIM=3)
PhievolParenU_dD = ixp.declarerank2("PhievolParenU_dD","nosym",DIM=3)
A_rhsD = ixp.zerorank1()
psi6Phi_rhs = GRFFE.psi6Phi_damping
for i in range(3):
A_rhsD[i] += -AevolParen_dD[i]
psi6Phi_rhs += -PhievolParenU_dD[i][i]
# Add Kreiss-Oliger dissipation to the GRFFE RHSs:
# psi6Phi_dKOD = ixp.declarerank1("psi6Phi_dKOD")
# AD_dKOD = ixp.declarerank2("AD_dKOD","nosym")
# for i in range(3):
# psi6Phi_rhs += diss_strength*psi6Phi_dKOD[i]*rfm.ReU[i] # ReU[i] = 1/scalefactor_orthog_funcform[i]
# for j in range(3):
# A_rhsD[j] += diss_strength*AD_dKOD[j][i]*rfm.ReU[i] # ReU[i] = 1/scalefactor_orthog_funcform[i]
RHSs_to_print = [\
lhrh(lhs=gri.gfaccess("rhs_gfs","AD0"),rhs=A_rhsD[0]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AD1"),rhs=A_rhsD[1]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","AD2"),rhs=A_rhsD[2]),\
lhrh(lhs=gri.gfaccess("rhs_gfs","psi6Phi"),rhs=psi6Phi_rhs),\
]
desc = "Calculate AD gauge term and psi6Phi RHSs"
name = "calculate_AD_gauge_psi6Phi_RHSs"
source_Ccode = outCfunction(
outfile = "returnstring", desc=desc, name=name,
params ="const paramstruct *params,const REAL *in_gfs,const REAL *auxevol_gfs,REAL *rhs_gfs",
body = fin.FD_outputC("returnstring",RHSs_to_print,params=outCparams),
loopopts ="InteriorPoints",
rel_path_to_Cparams=os.path.join("../")).replace("= NGHOSTS","= NGHOSTS_A2B").replace("NGHOSTS+Nxx0","Nxx_plus_2NGHOSTS0-NGHOSTS_A2B").replace("NGHOSTS+Nxx1","Nxx_plus_2NGHOSTS1-NGHOSTS_A2B").replace("NGHOSTS+Nxx2","Nxx_plus_2NGHOSTS2-NGHOSTS_A2B")
# Note the above .replace() functions. These serve to expand the loop range into the ghostzones, since
# the second-order FD needs fewer than some other algorithms we use do.
with open(os.path.join(out_dir,subdir,name+".h"),"w") as file:
file.write(source_Ccode)
# -
# We also need to compute the source term of the $\tilde{S}_i$ evolution equation. This term involves derivatives of the four metric, so we can save some effort here by taking advantage of the interpolations done of the metric gridfunctions to the cell faces, which will allow us to take a finite-difference derivative with the accuracy of a higher order and the computational cost of a lower order. However, it will require some more complicated coding, detailed in [Tutorial-GiRaFFE_NRPy-Source_Terms](Tutorial-GiRaFFE_NRPy-Source_Terms.ipynb)
# +
import GiRaFFE_NRPy.GiRaFFE_NRPy_Source_Terms as source
# Declare this symbol:
sqrt4pi = par.Cparameters("REAL",thismodule,"sqrt4pi","sqrt(4.0*M_PI)")
source.write_out_functions_for_StildeD_source_term(os.path.join(out_dir,subdir),outCparams,gammaDD,betaU,alpha,
ValenciavU,BU,sqrt4pi)
# -
# <a id='flux'></a>
#
# ## Step 1.c: Calculate the Flux terms \[Back to [top](#toc)\]
# $$\label{flux}$$
#
# Now, we will compute the flux terms of $\partial_t A_i$ and $\partial_t \tilde{S}_i$. To do so, we will first need to interpolate the metric gridfunctions to cell faces and to reconstruct the primitives on the cell faces using the code detailed in [Tutorial-GiRaFFE_NRPy-Metric_Face_Values](Tutorial-GiRaFFE_NRPy-Metric_Face_Values.ipynb) and in [Tutorial-GiRaFFE_NRPy-PPM](Tutorial-GiRaFFE_NRPy-PPM.ipynb).
subdir = "FCVAL"
cmd.mkdir(os.path.join(out_dir, subdir))
import GiRaFFE_NRPy.GiRaFFE_NRPy_Metric_Face_Values as FCVAL
FCVAL.GiRaFFE_NRPy_FCVAL(os.path.join(out_dir,subdir))
subdir = "PPM"
cmd.mkdir(os.path.join(out_dir, subdir))
import GiRaFFE_NRPy.GiRaFFE_NRPy_PPM as PPM
PPM.GiRaFFE_NRPy_PPM(os.path.join(out_dir,subdir))
# Here, we will write the function to compute the electric field contribution to the induction equation RHS. This is coded with documentation in [Tutorial-GiRaFFE_NRPy-Afield_flux_handwritten](Tutorial-GiRaFFE_NRPy-Afield_flux_handwritten.ipynb). The recontructed values in the $i^{\rm th}$ direction will contribute to the $j^{\rm th}$ and $k^{\rm th}$ component of the electric field. That is, in Cartesian coordinates, the component $x$ of the electric field will be the average of the values computed on the cell faces in the $\pm y$- and $\pm z$-directions, and so forth for the other components. However, all of these can be written as only a single function as long as we appropriately pass cyclical permutations of the inputs.
# +
import GiRaFFE_NRPy.GiRaFFE_NRPy_Afield_flux_handwritten as Af
# We will pass values of the gridfunction on the cell faces into the function. This requires us
# to declare them as C parameters in NRPy+. We will denote this with the _face infix/suffix.
alpha_face = gri.register_gridfunctions("AUXEVOL","alpha_face")
gamma_faceDD = ixp.register_gridfunctions_for_single_rank2("AUXEVOL","gamma_faceDD","sym01")
beta_faceU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","beta_faceU")
# We'll need some more gridfunctions, now, to represent the reconstructions of BU and ValenciavU
# on the right and left faces
Valenciav_rU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","Valenciav_rU",DIM=3)
B_rU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","B_rU",DIM=3)
Valenciav_lU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","Valenciav_lU",DIM=3)
B_lU = ixp.register_gridfunctions_for_single_rank1("AUXEVOL","B_lU",DIM=3)
subdir = "RHSs"
Af.GiRaFFE_NRPy_Afield_flux(os.path.join(out_dir,subdir))
# -
# We must do something similar here, albeit a bit simpler. For instance, the $x$ component of $\partial_t \tilde{S}_i$ will be a finite difference of the flux throught the faces in the $\pm x$ direction; for further detail, see [Tutorial-GiRaFFE_NRPy-Stilde_flux](Tutorial-GiRaFFE_NRPy-Stilde_flux.ipynb).
# +
ixp.register_gridfunctions_for_single_rank1("AUXEVOL","Stilde_flux_HLLED")
import GiRaFFE_NRPy.Stilde_flux as Sf
Sf.generate_C_code_for_Stilde_flux(os.path.join(out_dir,subdir), True, alpha_face,gamma_faceDD,beta_faceU,
Valenciav_rU,B_rU,Valenciav_lU,B_lU,sqrt4pi)
# -
# <a id='poststep'></a>
#
# # Step 2: Recover the primitive variables and apply boundary conditions \[Back to [top](#toc)\]
# $$\label{poststep}$$
#
# With the RHSs computed, we can now recover the primitive variables, which are the Valencia three-velocity $\bar{v}^i$ and the magnetic field $B^i$. We can also apply boundary conditions to the vector potential and velocity. By doing this at each RK substep, we can help ensure the accuracy of the following substeps.
#
# <a id='potential_bc'></a>
#
# ## Step 2.a: Apply boundary conditions to $A_i$ and $\sqrt{\gamma} \Phi$ \[Back to [top](#toc)\]
# $$\label{potential_bc}$$
#
# First, we will apply boundary conditions to the vector potential, $A_i$, and the scalar potential $\sqrt{\gamma} \Phi$. The file we generate here contains both functions we need for BCs, as documented in [Tutorial-GiRaFFE_NRPy-BCs](Tutorial-GiRaFFE_NRPy-BCs.ipynb).
subdir = "boundary_conditions"
cmd.mkdir(os.path.join(out_dir,subdir))
import GiRaFFE_NRPy.GiRaFFE_NRPy_BCs as BC
BC.GiRaFFE_NRPy_BCs(os.path.join(out_dir,subdir))
# <a id='a2b'></a>
#
# ## Step 2.b: Compute $B^i$ from $A_i$ \[Back to [top](#toc)\]
# $$\label{a2b}$$
#
# Now, we will calculate the magnetic field as the curl of the vector potential at all points in our domain; this requires care to be taken in the ghost zones, which is detailed in [Tutorial-GiRaFFE_NRPy-A2B](Tutorial-GiRaFFE_NRPy-A2B.ipynb).
subdir = "A2B"
cmd.mkdir(os.path.join(out_dir,subdir))
import GiRaFFE_NRPy.GiRaFFE_NRPy_A2B as A2B
A2B.GiRaFFE_NRPy_A2B(os.path.join(out_dir,subdir),gammaDD,AD,BU)
# <a id='c2p'></a>
#
# ## Step 2.c: Run the Conservative-to-Primitive solver \[Back to [top](#toc)\]
# $$\label{c2p}$$
#
# With these functions, we apply fixes to the Poynting flux, and use that to update the three-velocity. Then, we apply our current sheet prescription to the velocity, and recompute the Poynting flux to agree with the now-fixed velocity. More detail can be found in [Tutorial-GiRaFFE_NRPy-C2P_P2C](Tutorial-GiRaFFE_NRPy-C2P_P2C.ipynb).
# +
import GiRaFFE_NRPy.GiRaFFE_NRPy_C2P_P2C as C2P_P2C
C2P_P2C.GiRaFFE_NRPy_C2P(StildeD,BU,gammaDD,betaU,alpha)
values_to_print = [
lhrh(lhs=gri.gfaccess("in_gfs","StildeD0"),rhs=C2P_P2C.outStildeD[0]),
lhrh(lhs=gri.gfaccess("in_gfs","StildeD1"),rhs=C2P_P2C.outStildeD[1]),
lhrh(lhs=gri.gfaccess("in_gfs","StildeD2"),rhs=C2P_P2C.outStildeD[2]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU0"),rhs=C2P_P2C.ValenciavU[0]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU1"),rhs=C2P_P2C.ValenciavU[1]),
lhrh(lhs=gri.gfaccess("auxevol_gfs","ValenciavU2"),rhs=C2P_P2C.ValenciavU[2])
]
subdir = "C2P"
cmd.mkdir(os.path.join(out_dir,subdir))
desc = "Apply fixes to \tilde{S}_i and recompute the velocity to match with current sheet prescription."
name = "GiRaFFE_NRPy_cons_to_prims"
outCfunction(
outfile = os.path.join(out_dir,subdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *params,REAL *xx[3],REAL *auxevol_gfs,REAL *in_gfs",
body = fin.FD_outputC("returnstring",values_to_print,params=outCparams),
loopopts ="AllPoints,Read_xxs",
rel_path_to_Cparams=os.path.join("../"))
# +
# TINYDOUBLE = par.Cparameters("REAL",thismodule,"TINYDOUBLE",1e-100)
C2P_P2C.GiRaFFE_NRPy_P2C(gammaDD,betaU,alpha, ValenciavU,BU, sqrt4pi)
values_to_print = [
lhrh(lhs=gri.gfaccess("in_gfs","StildeD0"),rhs=C2P_P2C.StildeD[0]),
lhrh(lhs=gri.gfaccess("in_gfs","StildeD1"),rhs=C2P_P2C.StildeD[1]),
lhrh(lhs=gri.gfaccess("in_gfs","StildeD2"),rhs=C2P_P2C.StildeD[2]),
]
desc = "Recompute StildeD after current sheet fix to Valencia 3-velocity to ensure consistency between conservative & primitive variables."
name = "GiRaFFE_NRPy_prims_to_cons"
outCfunction(
outfile = os.path.join(out_dir,subdir,name+".h"), desc=desc, name=name,
params ="const paramstruct *params,REAL *auxevol_gfs,REAL *in_gfs",
body = fin.FD_outputC("returnstring",values_to_print,params=outCparams),
loopopts ="AllPoints",
rel_path_to_Cparams=os.path.join("../"))
# -
# <a id='velocity_bc'></a>
#
# ## Step 2.d: Apply outflow boundary conditions to $\bar{v}^i$ \[Back to [top](#toc)\]
# $$\label{velocity_bc}$$
#
# Now, we can apply outflow boundary conditions to the Valencia three-velocity. This specific type of boundary condition helps avoid numerical error "flowing" into our grid.
#
# This function has already been generated [above](#potential_bc).
# <a id='write_out'></a>
#
# # Step 3: Write out the C code function \[Back to [top](#toc)\]
# $$\label{write_out}$$
#
# Now, we have generated all the functions we will need for the `GiRaFFE` evolution. So, we will now assemble our evolution driver. This file will first `#include` all of the files we just generated for easy access. Then, we will write a function that calls these functions in the correct order, iterating over the flux directions as necessary.
# +
# %%writefile $out_dir/GiRaFFE_NRPy_Main_Driver.h
// Structure to track ghostzones for PPM:
typedef struct __gf_and_gz_struct__ {
REAL *gf;
int gz_lo[4],gz_hi[4];
} gf_and_gz_struct;
// Some additional constants needed for PPM:
const int VX=0,VY=1,VZ=2,BX=3,BY=4,BZ=5;
const int NUM_RECONSTRUCT_GFS = 6;
// Include ALL functions needed for evolution
#include "RHSs/calculate_AD_gauge_term_psi6Phi_flux_term_for_RHSs.h"
#include "RHSs/calculate_AD_gauge_psi6Phi_RHSs.h"
#include "PPM/reconstruct_set_of_prims_PPM_GRFFE_NRPy.c"
#include "FCVAL/interpolate_metric_gfs_to_cell_faces.h"
#include "RHSs/calculate_StildeD0_source_term.h"
#include "RHSs/calculate_StildeD1_source_term.h"
#include "RHSs/calculate_StildeD2_source_term.h"
#include "RHSs/calculate_E_field_flat_all_in_one.h"
#include "RHSs/calculate_Stilde_flux_D0.h"
#include "RHSs/calculate_Stilde_flux_D1.h"
#include "RHSs/calculate_Stilde_flux_D2.h"
#include "RHSs/calculate_Stilde_rhsD.h"
#include "boundary_conditions/GiRaFFE_boundary_conditions.h"
#include "A2B/driver_AtoB.h"
#include "C2P/GiRaFFE_NRPy_cons_to_prims.h"
#include "C2P/GiRaFFE_NRPy_prims_to_cons.h"
void GiRaFFE_NRPy_RHSs(const paramstruct *restrict params,REAL *restrict auxevol_gfs,const REAL *restrict in_gfs,REAL *restrict rhs_gfs) {
#include "set_Cparameters.h"
// First thing's first: initialize the RHSs to zero!
#pragma omp parallel for
for(int ii=0;ii<Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2*NUM_EVOL_GFS;ii++) {
rhs_gfs[ii] = 0.0;
}
// Next calculate the easier source terms that don't require flux directions
// This will also reset the RHSs for each gf at each new timestep.
calculate_AD_gauge_term_psi6Phi_flux_term_for_RHSs(params,in_gfs,auxevol_gfs);
calculate_AD_gauge_psi6Phi_RHSs(params,in_gfs,auxevol_gfs,rhs_gfs);
// Now, we set up a bunch of structs of pointers to properly guide the PPM algorithm.
// They also count the number of ghostzones available.
gf_and_gz_struct in_prims[NUM_RECONSTRUCT_GFS], out_prims_r[NUM_RECONSTRUCT_GFS], out_prims_l[NUM_RECONSTRUCT_GFS];
int which_prims_to_reconstruct[NUM_RECONSTRUCT_GFS],num_prims_to_reconstruct;
const int Nxxp2NG012 = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;
REAL *temporary = auxevol_gfs + Nxxp2NG012*AEVOLPARENGF; //We're not using this anymore
// This sets pointers to the portion of auxevol_gfs containing the relevant gridfunction.
int ww=0;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAVU0GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_RU0GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_LU0GF;
ww++;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAVU1GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_RU1GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_LU1GF;
ww++;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAVU2GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_RU2GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*VALENCIAV_LU2GF;
ww++;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*BU0GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*B_RU0GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*B_LU0GF;
ww++;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*BU1GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*B_RU1GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*B_LU1GF;
ww++;
in_prims[ww].gf = auxevol_gfs + Nxxp2NG012*BU2GF;
out_prims_r[ww].gf = auxevol_gfs + Nxxp2NG012*B_RU2GF;
out_prims_l[ww].gf = auxevol_gfs + Nxxp2NG012*B_LU2GF;
ww++;
// Prims are defined AT ALL GRIDPOINTS, so we set the # of ghostzones to zero:
for(int i=0;i<NUM_RECONSTRUCT_GFS;i++) for(int j=1;j<=3;j++) { in_prims[i].gz_lo[j]=0; in_prims[i].gz_hi[j]=0; }
// Left/right variables are not yet defined, yet we set the # of gz's to zero by default:
for(int i=0;i<NUM_RECONSTRUCT_GFS;i++) for(int j=1;j<=3;j++) { out_prims_r[i].gz_lo[j]=0; out_prims_r[i].gz_hi[j]=0; }
for(int i=0;i<NUM_RECONSTRUCT_GFS;i++) for(int j=1;j<=3;j++) { out_prims_l[i].gz_lo[j]=0; out_prims_l[i].gz_hi[j]=0; }
ww=0;
which_prims_to_reconstruct[ww]=VX; ww++;
which_prims_to_reconstruct[ww]=VY; ww++;
which_prims_to_reconstruct[ww]=VZ; ww++;
which_prims_to_reconstruct[ww]=BX; ww++;
which_prims_to_reconstruct[ww]=BY; ww++;
which_prims_to_reconstruct[ww]=BZ; ww++;
num_prims_to_reconstruct=ww;
// In each direction, perform the PPM reconstruction procedure.
// Then, add the fluxes to the RHS as appropriate.
for(int flux_dirn=0;flux_dirn<3;flux_dirn++) {
// In each direction, interpolate the metric gfs (gamma,beta,alpha) to cell faces.
interpolate_metric_gfs_to_cell_faces(params,auxevol_gfs,flux_dirn+1);
// Then, reconstruct the primitive variables on the cell faces.
// This function is housed in the file: "reconstruct_set_of_prims_PPM_GRFFE_NRPy.c"
reconstruct_set_of_prims_PPM_GRFFE_NRPy(params, auxevol_gfs, flux_dirn+1, num_prims_to_reconstruct,
which_prims_to_reconstruct, in_prims, out_prims_r, out_prims_l, temporary);
// For example, if flux_dirn==0, then at gamma_faceDD00(i,j,k) represents gamma_{xx}
// at (i-1/2,j,k), Valenciav_lU0(i,j,k) is the x-component of the velocity at (i-1/2-epsilon,j,k),
// and Valenciav_rU0(i,j,k) is the x-component of the velocity at (i-1/2+epsilon,j,k).
if(flux_dirn==0) {
// Next, we calculate the source term for StildeD. Again, this also resets the rhs_gfs array at
// each new timestep.
calculate_StildeD0_source_term(params,auxevol_gfs,rhs_gfs);
// Now, compute the electric field on each face of a cell and add it to the RHSs as appropriate
//calculate_E_field_D0_right(params,auxevol_gfs,rhs_gfs);
//calculate_E_field_D0_left(params,auxevol_gfs,rhs_gfs);
// Finally, we calculate the flux of StildeD and add the appropriate finite-differences
// to the RHSs.
calculate_Stilde_flux_D0(params,auxevol_gfs,rhs_gfs);
}
else if(flux_dirn==1) {
calculate_StildeD1_source_term(params,auxevol_gfs,rhs_gfs);
//calculate_E_field_D1_right(params,auxevol_gfs,rhs_gfs);
//calculate_E_field_D1_left(params,auxevol_gfs,rhs_gfs);
calculate_Stilde_flux_D1(params,auxevol_gfs,rhs_gfs);
}
else {
calculate_StildeD2_source_term(params,auxevol_gfs,rhs_gfs);
//calculate_E_field_D2_right(params,auxevol_gfs,rhs_gfs);
//calculate_E_field_D2_left(params,auxevol_gfs,rhs_gfs);
calculate_Stilde_flux_D2(params,auxevol_gfs,rhs_gfs);
}
calculate_Stilde_rhsD(flux_dirn+1,params,auxevol_gfs,rhs_gfs);
for(int count=0;count<=1;count++) {
// This function is written to be general, using notation that matches the forward permutation added to AD2,
// i.e., [F_HLL^x(B^y)]_z corresponding to flux_dirn=0, count=1.
// The SIGN parameter is necessary because
// -E_z(x_i,y_j,z_k) = 0.25 ( [F_HLL^x(B^y)]_z(i+1/2,j,k)+[F_HLL^x(B^y)]_z(i-1/2,j,k)
// -[F_HLL^y(B^x)]_z(i,j+1/2,k)-[F_HLL^y(B^x)]_z(i,j-1/2,k) )
// Note the negative signs on the reversed permutation terms!
// By cyclically permuting with flux_dirn, we
// get contributions to the other components, and by incrementing count, we get the backward permutations:
// Let's suppose flux_dirn = 0. Then we will need to update Ay (count=0) and Az (count=1):
// flux_dirn=count=0 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (0+1+0)%3=AD1GF <- Updating Ay!
// (flux_dirn)%3 = (0)%3 = 0 Vx
// (flux_dirn-count+2)%3 = (0-0+2)%3 = 2 Vz . Inputs Vx, Vz -> SIGN = -1 ; 2.0*((REAL)count)-1.0=-1 check!
// flux_dirn=0,count=1 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (0+1+1)%3=AD2GF <- Updating Az!
// (flux_dirn)%3 = (0)%3 = 0 Vx
// (flux_dirn-count+2)%3 = (0-1+2)%3 = 1 Vy . Inputs Vx, Vy -> SIGN = +1 ; 2.0*((REAL)count)-1.0=2-1=+1 check!
// Let's suppose flux_dirn = 1. Then we will need to update Az (count=0) and Ax (count=1):
// flux_dirn=1,count=0 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (1+1+0)%3=AD2GF <- Updating Az!
// (flux_dirn)%3 = (1)%3 = 1 Vy
// (flux_dirn-count+2)%3 = (1-0+2)%3 = 0 Vx . Inputs Vy, Vx -> SIGN = -1 ; 2.0*((REAL)count)-1.0=-1 check!
// flux_dirn=count=1 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (1+1+1)%3=AD0GF <- Updating Ax!
// (flux_dirn)%3 = (1)%3 = 1 Vy
// (flux_dirn-count+2)%3 = (1-1+2)%3 = 2 Vz . Inputs Vy, Vz -> SIGN = +1 ; 2.0*((REAL)count)-1.0=2-1=+1 check!
// Let's suppose flux_dirn = 2. Then we will need to update Ax (count=0) and Ay (count=1):
// flux_dirn=2,count=0 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (2+1+0)%3=AD0GF <- Updating Ax!
// (flux_dirn)%3 = (2)%3 = 2 Vz
// (flux_dirn-count+2)%3 = (2-0+2)%3 = 1 Vy . Inputs Vz, Vy -> SIGN = -1 ; 2.0*((REAL)count)-1.0=-1 check!
// flux_dirn=2,count=1 -> AD0GF+(flux_dirn+1+count)%3 = AD0GF + (2+1+1)%3=AD1GF <- Updating Ay!
// (flux_dirn)%3 = (2)%3 = 2 Vz
// (flux_dirn-count+2)%3 = (2-1+2)%3 = 0 Vx . Inputs Vz, Vx -> SIGN = +1 ; 2.0*((REAL)count)-1.0=2-1=+1 check!
calculate_E_field_flat_all_in_one(params,
&auxevol_gfs[IDX4ptS(VALENCIAV_RU0GF+(flux_dirn)%3, 0)],&auxevol_gfs[IDX4ptS(VALENCIAV_RU0GF+(flux_dirn-count+2)%3, 0)],
&auxevol_gfs[IDX4ptS(VALENCIAV_LU0GF+(flux_dirn)%3, 0)],&auxevol_gfs[IDX4ptS(VALENCIAV_LU0GF+(flux_dirn-count+2)%3, 0)],
&auxevol_gfs[IDX4ptS(B_RU0GF +(flux_dirn)%3, 0)],&auxevol_gfs[IDX4ptS(B_RU0GF +(flux_dirn-count+2)%3, 0)],
&auxevol_gfs[IDX4ptS(B_LU0GF +(flux_dirn)%3, 0)],&auxevol_gfs[IDX4ptS(B_LU0GF +(flux_dirn-count+2)%3, 0)],
&auxevol_gfs[IDX4ptS(B_RU0GF +(flux_dirn-count+2)%3, 0)],
&auxevol_gfs[IDX4ptS(B_LU0GF +(flux_dirn-count+2)%3, 0)],
&rhs_gfs[IDX4ptS(AD0GF+(flux_dirn+1+count)%3,0)], 2.0*((REAL)count)-1.0, flux_dirn);
}
}
}
void GiRaFFE_NRPy_post_step(const paramstruct *restrict params,REAL *xx[3],REAL *restrict auxevol_gfs,REAL *restrict evol_gfs,const int n) {
// First, apply BCs to AD and psi6Phi. Then calculate BU from AD
apply_bcs_potential(params,evol_gfs);
driver_A_to_B(params,evol_gfs,auxevol_gfs);
//override_BU_with_old_GiRaFFE(params,auxevol_gfs,n);
// Apply fixes to StildeD, then recompute the velocity at the new timestep.
// Apply the current sheet prescription to the velocities
GiRaFFE_NRPy_cons_to_prims(params,xx,auxevol_gfs,evol_gfs);
// Then, recompute StildeD to be consistent with the new velocities
//GiRaFFE_NRPy_prims_to_cons(params,auxevol_gfs,evol_gfs);
// Finally, apply outflow boundary conditions to the velocities.
apply_bcs_velocity(params,auxevol_gfs);
}
# -
# <a id='code_validation'></a>
#
# # Step 4: Self-Validation against `GiRaFFE_NRPy_Main_Drive.py` \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# To validate the code in this tutorial we check for agreement between the files
#
# 1. that were generated in this tutorial and
# 1. those that are generated in the module [`GiRaFFE_NRPy_Main_Driver.py`](../../edit/in_progress/GiRaFFE_NRPy/GiRaFFE_NRPy_Main_Driver.py)
#
# +
gri.glb_gridfcs_list = []
# Define the directory that we wish to validate against:
valdir = os.path.join("GiRaFFE_validation_Ccodes")
cmd.mkdir(valdir)
import GiRaFFE_NRPy.GiRaFFE_NRPy_Main_Driver as md
md.GiRaFFE_NRPy_Main_Driver_generate_all(valdir)
# -
# With both sets of codes generated, we can now compare them against each other.
# +
import difflib
import sys
print("Printing difference between original C code and this code...")
# Open the files to compare
files = ["GiRaFFE_NRPy_Main_Driver.h",
"RHSs/calculate_AD_gauge_term_psi6Phi_flux_term_for_RHSs.h",
"RHSs/calculate_AD_gauge_psi6Phi_RHSs.h",
"PPM/reconstruct_set_of_prims_PPM_GRFFE_NRPy.c",
"PPM/loop_defines_reconstruction_NRPy.h",
"FCVAL/interpolate_metric_gfs_to_cell_faces.h",
"RHSs/calculate_StildeD0_source_term.h",
"RHSs/calculate_StildeD1_source_term.h",
"RHSs/calculate_StildeD2_source_term.h",
"RHSs/calculate_E_field_flat_all_in_one.h",
"RHSs/calculate_Stilde_flux_D0.h",
"RHSs/calculate_Stilde_flux_D1.h",
"RHSs/calculate_Stilde_flux_D2.h",
"boundary_conditions/GiRaFFE_boundary_conditions.h",
"A2B/driver_AtoB.h",
"C2P/GiRaFFE_NRPy_cons_to_prims.h",
"C2P/GiRaFFE_NRPy_prims_to_cons.h"]
for file in files:
print("Checking file " + file)
with open(os.path.join(valdir,file)) as file1, open(os.path.join(out_dir,file)) as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
num_diffs = 0
for line in difflib.unified_diff(file1_lines, file2_lines, fromfile=os.path.join(valdir,file), tofile=os.path.join(out_dir,file)):
sys.stdout.writelines(line)
num_diffs = num_diffs + 1
if num_diffs == 0:
print("No difference. TEST PASSED!")
else:
print("ERROR: Disagreement found with .py file. See differences above.")
sys.exit(1)
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-GiRaFFE_NRPy_Main_Driver](TTutorial-GiRaFFE_NRPy_Main_Driver.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-GiRaFFE_NRPy_Main_Driver",location_of_template_file=os.path.join(".."))
|
in_progress/Tutorial-GiRaFFE_NRPy_Main_Driver.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Scrape MyMovies Web Site
#
# ## Goal
#
# Obtain __two__ web pages:
#
# * Cars
# * Pirates of Carribean 5
#
# Get the description of the movies and make a quick analysis about the content.
#
# * Compare the two movies, do they refer to the same topic?
# * Do they have topic in common?
#
# ## Libraries we need
#
# * `bs4` (`BeautifulSoup module`)
# * `requests` or `urllib2`
#
# ## How to do it?
#
# 1. Save two variables for the `urls`
# 2. Use the library `requests` to get the `html` to parse (? Get the useful information)
# 3. Use `BeautifulSoup` to the the *"soup"* to parse
# 4. Explore a bit the `html` and `BeautifulSoup` to understand how they work
# 5. Produce the Cloud of Word (link in Assignment 3, exercise 3) to get an idea of what's going on
#
# import useful libraries
import requests
from bs4 import BeautifulSoup
# Define url
url1 = 'http://www.mymovies.it/film/2017/cars3/'
url2 = 'http://www.mymovies.it/film/2017/piratideicaraibi5/'
# Perform a `get` request. __What it is?__
#
# Basically, you are asking the server to give you something (so you `get` the information). In this particular case, we want to obtain the `html` of the pages of interest.
#
# We create a function to get both the `html` and the `soup`.
#
def get_page(url):
"""Return the html to analyse.
@url: url of the page"""
# Make the requests and get html
html = requests.get(url).text
# Get the soup
soup = BeautifulSoup(html, 'html.parser')
return soup
soup1 = get_page(url1)
# ### How to use the `soup`?
#
# Reeeeaad the [documentation](https://www.crummy.com/software/BeautifulSoup/bs4/doc/).
#
# We are interest in the body (`corpo`)!
#
# Thus, we proceed like as follows.
#
def get_body_to_process(soup):
"""@soup: outpur get_html"""
# Get all the paragraph
body = []
for p in soup1.find_all('p', {'class':'corpo'}):
print (p.text)
body += [p.text]
# Get body to process: join the strings of the lists
body_to_process = ''.join([str(b).strip() for b in body])
return body_to_process
body = []
for p in soup1.find_all('p', {'class':'corpo'}):
print (p.text)
body += [p.text]
# +
# We do a join on the list of paragraphs
body_to_process = ''.join([str(b).strip() for b in body])
# Let's see it!
body_to_process
# -
# ## We need to pre-process the text, why?
#
# * If I want to compute the frequency of a word, I don't want my program to distinguish between "ciao" and "CiAo". So, it's a good use to tranform all the characters in *lowercase*
# * We don't want to analyse the *stopwords* (e.g. of, on, me, you, they, are, have etc.), common words that have not much impact on the analysis
#
# And many other according to your objectives.
def clean_body(body_to_process):
"""
@body_to_process: list of words that need to be processed"""
# Define the stopwords
stop = stopwords.words('italian')
# Clean each word
cleaned_body = ' '.join([w.strip().lower() for w in body_to_process.split() if len(w) > 3 and w not in stop])
return cleaned_body
# +
# Usually in english we already have the list (which you can always modify)... also in italian!!
# Fantastic library for text analysis N(atural)L(anguage)T(ool)K(it)
from nltk.corpus import stopwords
stop = stopwords.words('italian')
# Let's take a look
stop[:10]
# +
# Clean the body dropping all the words which have
cleaned_body = ' '.join([w.strip().lower() for w in body_to_process.split() if len(w) > 3 and w not in stop])
# What we have now
cleaned_body[:25]
# -
# ## Ready to visualize :-)
# %matplotlib inline
from wordcloud import WordCloud
# +
# Generate a word cloud image
wordcloud = WordCloud().generate(cleaned_body)
# Display the generated image:
# the matplotlib way:
import matplotlib.pyplot as plt
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
# lower max_font_size
wordcloud = WordCloud(max_font_size=40).generate(cleaned_body)
plt.figure()
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
# -
|
05/Scrape MyMovies.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 0. Imports and Settings
# +
import pandas as pd
import requests
from datetime import datetime
from bs4 import BeautifulSoup
import math
pd.options.mode.chained_assignment = None
# +
#The headers for us to look real
headers = {'User-Agent':'Mozilla/5.0 (Macintosh; Intel Mac OS X 10.15; rv:95.0) Gecko/20100101 Firefox/95.0'}
#The home page
home_url = 'https://books.toscrape.com/index.html'
#List of categories you wish to scrap
cats = ['Classics', 'Science Fiction', 'Humor', 'Business']
# -
# # 1. Functions
# ## 1.1 Fetch the Categories Available
# +
#This function is to create a clean data-set of the categories, their URLs and number of books in each.
#This is important to prevent the code from stop working in the case of layout (i.e. order) changes in the webpage.
def categories_pages_urls (home_url, headers):
#Access the page and store it
page = requests.get( home_url, headers=headers )
#Parse the homepage using bs4
soup = BeautifulSoup( page.text, 'html.parser' )
#This is where I found the list including all the categories in the sidebar
categories_list = soup.find( 'ul', class_='nav nav-list' ).find_all('a')
#First, I will create a set with the urls heading to the categories pages.
categories_urls = [p.get('href') for p in categories_list]
#Transform this list in a dataframe, excluding line one which goes for the homepage.
categories_urls_df = pd.DataFrame(categories_urls).iloc[1:]
#Now, I need to get a list of the categories themselves to concatenate with the previous list.
#Using split '\n' because there was some weird spacing back there in the html
categories_titles = [p.get_text().split('\n') for p in categories_list]
#Transforming it in a dataframe and dropping alien columns
categories_titles_df = pd.DataFrame(categories_titles).iloc[1:, [False, False, True, False, False]]
#Move the tables together and drop more alien columns
categories_location = pd.concat([categories_titles_df, categories_urls_df], axis = 1).reset_index().iloc[:, [False, True, True]]
categories_location.columns = ['category', 'category_url']
#remove weird 32 spacing from category names
for i in range(len(categories_location)):
categories_location['category'][i] = categories_location['category'][i][32:]
categories_location['books_in_category'] = 0
#Now, I will use a loop to get the quantity of books in each category (we are going to need it later)
for i in range(len(categories_location)):
url_get_cat_number = 'https://books.toscrape.com/'+categories_location.iloc[i]['category_url']
page = requests.get( url_get_cat_number, headers=headers )
soup = BeautifulSoup( page.text, 'html.parser' )
books_in_cat = soup.find( 'form', class_='form-horizontal' ).find('strong')
qty = int([p.get_text('strong') for p in books_in_cat][0])
categories_location.loc[i, 'books_in_category'] = qty
return categories_location
# -
# ## 1.2 Select only the wished categories
#The input to this function (df_cats_urls_qtys) should be the return of categories_pages_urls.
#cats should be a list of categories to be scrapped
def cats_wish_to_scrap(df_cats_urls_qtys, cats, pagination):
#Filter
boolean = df_categories_urls.category.isin(cats)
wish = df_categories_urls[boolean]
#Add pagination info, we will need it later
wish['pages'] = 0
wish['pages'] = wish['books_in_category']/pagination
wish['pages'] = wish['pages'].apply(lambda x: int(math.ceil(x)))
#===========================================
# Need to add pagination features.
# Suggestion: apply change "../index.html" to "../page-{i}.html" i in range(wish[pages])
# Update the wishlist
#===========================================
return wish
# ## 1.3 Fetch the URLs of every single book in the selected categories
# +
#The input to this function should be the return of cats_wish_to_scrap
def create_worklist_to_scrap(worklist):
df_worklist = pd.DataFrame()
for i in range(len(worklist)):
#let's start with Classics from worklist
url_titles = 'http://books.toscrape.com/'+ worklist.iloc[i, 1]
#Access the page and store it
page = requests.get( url_titles, headers=headers )
#Parse the catalogue page using the html.parser
soup = BeautifulSoup( page.text, 'html.parser' )
titles = soup.find('div', class_='col-sm-8 col-md-9')
category_is = titles.find_all('h1')
c = category_is[0].get_text()
url_is = titles.find_all('a')
titles_url = set([p.get('href') for p in url_is])
titles_url = list(titles_url)
df_titles_url = pd.DataFrame(titles_url)
df_titles_url['category'] = c
df_worklist = df_worklist.append(df_titles_url, ignore_index=True)
df_titles_url = df_titles_url.iloc[0:0]
df_worklist.columns = ['title_url','category']
for i in range(len(df_worklist)):
df_worklist['title_url'][i] = 'https://books.toscrape.com/catalogue'+df_worklist['title_url'][i][8:]
return df_worklist
# -
# ## 1.4 Scrap the books pages and append the data in a single dataframe
def books_scrapping(books_to_scrap):
df_books_scrap = pd.DataFrame(columns=['scrap_time','book_title','book_category', 'book_upc', 'book_price', 'book_availability','book_stars'])
#the date the scrapping was held
scrap_time = datetime.now().strftime( '%Y-%m-%d %H:%M:%S' )
for i in range(len(books_to_scrap)):
book_url = books_to_scrap.iloc[0:]['title_url'][i]
book_category = books_to_scrap.iloc[0:]['category'][i]
#Access the page and store it
page = requests.get( book_url, headers=headers )
#Parse the homepage using bs4
soup = BeautifulSoup( page.text, 'html.parser' )
#the title of the book
book_title = (soup.find('div', class_="col-sm-6 product_main").find_all('h1')[0]).get_text()
#where most the information is:
book_info_table = soup.find('table', class_="table table-striped").find_all('td')
#ua unique identifier for each title
book_upc = (book_info_table[0]).get_text()
#price excl. tax
book_price = (book_info_table[2]).get_text()
#quantity available
book_availability = (book_info_table[5]).get_text()
#rating
book_rate = (soup.find('div', class_="col-sm-6 product_main")).find_all('p')
book_stars = (book_rate[2].get('class'))[1]
df_books_scrap.loc[i,'scrap_time'] = scrap_time
df_books_scrap.loc[i,'book_title'] = book_title
df_books_scrap.loc[i,'book_upc'] = book_upc
df_books_scrap.loc[i,'book_price'] = book_price
df_books_scrap.loc[i,'book_availability'] = book_availability
df_books_scrap.loc[i,'book_category'] = book_category
df_books_scrap.loc[i,'book_stars'] = book_stars
return df_books_scrap
# ## 1.5 Final transformations for a clean delivery
def data_processing(books_scrap):
#Change string stars to numbers
dic = {'One':1, 'Two':2, 'Three':3, 'Four':4, 'Five':5}
books_scrap['book_stars'].replace(dic, inplace=True)
#Change price to number
books_scrap['book_price'] = books_scrap['book_price'].apply(lambda x: x[2:])
#Turn availability data better
books_scrap[['book_in_stock', 'availability']] = books_scrap['book_availability'].str.split('(', 1, expand=True)
books_scrap[['nr_available', 'drop']] = books_scrap['availability'].str.split(' ', 1, expand=True)
books_scrap.drop(columns = ['book_availability', 'availability', 'drop'], axis = 1, inplace=True)
return books_scrap
# # 2. Scrapping
# ## 2.1 Website structure scrapping
# ### 2.1.1 Get the categories in the Sidebar, their URLs and sizes for pagination
#Use the defined function to get the clean dataframe of categories
df_categories_urls = categories_pages_urls(home_url=home_url, headers=headers)
df_categories_urls.sample(5)
#Select only the ones we wish to work on and add pagination info
worklist = cats_wish_to_scrap(cats=cats, df_cats_urls_qtys=df_categories_urls, pagination=20)
worklist.sample(3)
# ### 2.1.2 Get the titles in each category
books_to_scrap = create_worklist_to_scrap(worklist=worklist)
books_to_scrap.sample(5)
# ### 2.1.3 Scrap each book page to retrieve the remaining information and build the dataset
books_scrap = books_scrapping(books_to_scrap=books_to_scrap)
books_scrap.sample(5)
# ### 2.1.4 Transform the data and export the final dataset to .csv
df_deliver = data_processing(books_scrap=books_scrap)
df_deliver.sample(5)
df_deliver.to_csv('CoffeeCookies-dataset.csv')
|
m01_books_scrapping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://pythonista.io)
# # Excepciones personalizadas.
# ## La clase *Exception*.
#
# La clase *Exception* pertenece al módulo *\_\_builtins\_\_* y es la clase base a partir de la cual heredan el resto de las excepciones.
#
# **Ejemplo:**
help(Exception)
# ## Creación de una excepción propia.
# Es posible crear una excepción mediante la definición de una clase que herede de cualquiera de las Excepciones disponibles.
# Cabe hacer notar que una excepción que herede directamente de Exception, capturará todas las excepciones.
#
# **Ejemplo:**
class MiError(Exception):
'''Excepcion propia.'''
def __init__(self):
print("Ya me lo esperaba.")
def pidenumero(numero):
try:
numero = int(numero)
if numero == 20 :
print('Adivinaste.')
else:
raise MiError
except MiError:
print('Ese no era el número.')
except:
print('Hubo un error.')
print('Buen día.')
pidenumero(-1j)
pidenumero(12)
pidenumero(20)
# <p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
# <p style="text-align: center">© <NAME>. 2020.</p>
|
11_excepciones_personalizadas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Seminar: simple question answering
# 
#
# Today we're going to build a retrieval-based question answering model with metric learning models.
#
# _this seminar is based on original notebook by [<NAME>](https://github.com/Omrigan/)_
#
#
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Dataset
#
# Today's data is Stanford Question Answering Dataset (SQuAD). Given a paragraph of text and a question, our model's task is to select a snippet that answers the question.
#
# We are not going to solve the full task today. Instead, we'll train a model to __select the sentence containing answer__ among several options.
#
# As usual, you are given an utility module with data reader and some helper functions
import utils
# !pip install sentencepiece tensorflow_hub
# !wget https://rajpurkar.github.io/SQuAD-explorer/dataset/train-v2.0.json -O squad-v2.0.json 2> log
# backup download link: https://www.dropbox.com/s/q4fuihaerqr0itj/squad.tar.gz?dl=1
train, test = utils.build_dataset('./squad-v2.0.json')
pid, question, options, correct_indices, wrong_indices = train.iloc[40]
print('QUESTION', question, '\n')
for i, cand in enumerate(options):
print(['[ ]', '[v]'][i in correct_indices], cand)
# ### Universal Sentence Encoder
#
# We've already solved quite a few tasks from scratch, training our own embeddings and convolutional/recurrent layers. However, one can often achieve higher quality by using pre-trained models. So today we're gonna use pre-trained Universal Sentence Encoder from [Tensorflow Hub](https://tfhub.dev/google/universal-sentence-encoder/2).
#
#
# [__Universal Sentence Encoder__](https://arxiv.org/abs/1803.11175) is a model that encoders phrases, sentences or short paragraphs into a fixed-size vector. It was trained simultaneosly on a variety of tasks to achieve versatility.
#
# +
import tensorflow as tf
import tensorflow_hub as hub
universal_sentence_encoder = hub.load("https://tfhub.dev/google/universal-sentence-encoder/3")
# consider as well:
# * lite: https://tfhub.dev/google/universal-sentence-encoder-lite/2
# * large: https://tfhub.dev/google/universal-sentence-encoder-large/2
# +
dummy_lines = [
"How old are you?", # 0
"In what mythology do two canines watch over the Chinvat Bridge?", # 1
"I'm sorry, okay, I'm not perfect, but I'm trying.", # 2
"What is your age?", # 3
"Beware, for I am fearless, and therefore powerful.", # 4
]
dummy_vectors_np = universal_sentence_encoder(dummy_lines)["outputs"].numpy()
plt.title('phrase similarity')
plt.imshow(dummy_vectors_np.dot(dummy_vectors_np.T), interpolation='none', cmap='gray')
# -
# As you can see, __the strongest similarity is between lines 0 and 3__. Indeed they correspond to "How old are you?" and "What is your age?"
# ### Model (2 points)
#
# Our goal for today is to build a model that measures similarity between question and answer. In particular, it maps both question and answer into fixed-size vectors such that:
#
# Our model is a pair of $V_q(q)$ and $V_a(a)$ - networks that turn phrases into vectors.
#
# __Objective:__ Question vector $V_q(q)$ should be __closer__ to correct answer vectors $V_a(a^+)$ than to incorrect ones $V_a(a^-)$ .
#
# Both vectorizers can be anything you wish. For starters, let's use a couple of dense layers on top of the Universal Sentence Encoder.
#
#
#
# +
import tensorflow.keras.layers as L
import tensorflow.keras as keras
class Vectorizer(keras.Model):
def __init__(self, output_size=256, hid_size=256, universal_sentence_encoder=universal_sentence_encoder):
""" A small feedforward network on top of universal sentence encoder. 2-3 layers should be enough """
super(Vectorizer, self).__init__()
self.universal_sentence_encoder = universal_sentence_encoder
# define a few layers to be applied on top of u.s.e.
# note: please make sure your final layer comes with _linear_ activation
self.first = L.Dense(256)
def __call__(self, input_phrases, is_train=True):
"""
Apply vectorizer. Use dropout and any other hacks at will.
:param input_phrases: [batch_size] of tf.string
:param is_train: if True, apply dropouts and other ops in train mode,
if False - evaluation mode
:returns: predicted phrase vectors, [batch_size, output_size]
"""
return self.first(self.universal_sentence_encoder(input_phrases)['outputs'])
# -
question_vectorizer = Vectorizer()
answer_vectorizer = Vectorizer()
# +
# dummy_v_q = question_vectorizer(dummy_ph, is_train=True)
# dummy_v_q_det = question_vectorizer(dummy_ph, is_train=False)
# utils.initialize_uninitialized()
# assert sess.run(dummy_v_q, {dummy_ph: dummy_lines}).shape == (5, 256)
assert question_vectorizer(dummy_lines).shape == (5, 256)
assert np.allclose(
question_vectorizer(dummy_lines, is_train=False).numpy(),
question_vectorizer(dummy_lines, is_train=False).numpy(),
atol=1e-7
), "make sure your model doesn't use dropout/noise or non-determinism if is_train=False"
print("Well done!")
# -
# ### Training: minibatches
#
# Our model learns on triples $(q, a^+, a^-)$:
# * q - __q__uestion
# * (a+) - correct __a__nswer
# * (a-) - wrong __a__nswer
#
# Below you will find a generator that samples such triples from data.
# +
import random
def iterate_minibatches(data, batch_size, shuffle=True, cycle=False):
"""
Generates minibatches of triples: {questions, correct answers, wrong answers}
If there are several wrong (or correct) answers, picks one at random.
"""
indices = np.arange(len(data))
while True:
if shuffle:
indices = np.random.permutation(indices)
for batch_start in range(0, len(indices), batch_size):
batch_indices = indices[batch_start: batch_start + batch_size]
batch = data.iloc[batch_indices]
questions = batch['question'].values
correct_answers = np.array([
row['options'][random.choice(row['correct_indices'])]
for i, row in batch.iterrows()
])
wrong_answers = np.array([
row['options'][random.choice(row['wrong_indices'])]
for i, row in batch.iterrows()
])
yield {
'questions' : questions,
'correct_answers': correct_answers,
'wrong_answers': wrong_answers,
}
if not cycle:
break
# -
dummy_batch = next(iterate_minibatches(train.sample(3), 3))
print(dummy_batch)
# ### Training: loss function (2 points)
# We want our vectorizers to put correct answers closer to question vectors and incorrect answers farther away from them. One way to express this is to use is Pairwise Hinge Loss _(aka Triplet Loss)_.
#
# $$ L = \frac 1N \underset {q, a^+, a^-} \sum max(0, \space \delta - sim[V_q(q), V_a(a^+)] + sim[V_q(q), V_a(a^-)] )$$
#
# , where
# * sim[a, b] is some similarity function: dot product, cosine or negative distance
# * δ - loss hyperparameter, e.g. δ=1.0. If sim[a, b] is linear in b, all δ > 0 are equivalent.
#
#
# This reads as __Correct answers must be closer than the wrong ones by at least δ.__
#
# 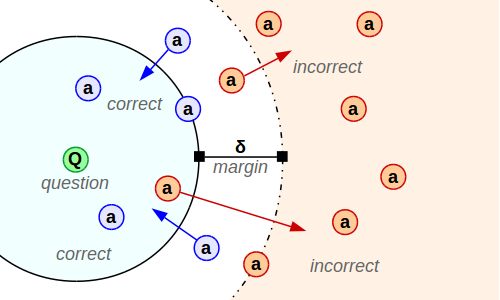
# <center>_image: question vector is green, correct answers are blue, incorrect answers are red_</center>
#
#
# Note: in effect, we train a Deep Semantic Similarity Model [DSSM](https://www.microsoft.com/en-us/research/project/dssm/).
# +
def similarity(a, b):
""" Dot product as a similarity function """
return tf.einsum("ij,ij->i", a, b)
def compute_loss(question_vectors, correct_answer_vectors, wrong_answer_vectors, delta=1.0):
"""
Compute the triplet loss as per formula above.
Use similarity function above for sim[a, b]
:param question_vectors: float32[batch_size, vector_size]
:param correct_answer_vectors: float32[batch_size, vector_size]
:param wrong_answer_vectors: float32[batch_size, vector_size]
:returns: loss for every row in batch, float32[batch_size]
Hint: DO NOT use tf.reduce_max, it's a wrong kind of maximum :)
"""
pos = similarity(question_vectors, correct_answer_vectors)
neg = similarity(question_vectors, wrong_answer_vectors)
return tf.math.maximum(0, delta - pos + neg)
# +
dummy_v1 = tf.constant([[0.1, 0.2, -1], [-1.2, 0.6, 1.0]], dtype=tf.float32)
dummy_v2 = tf.constant([[0.9, 2.1, -6.6], [0.1, 0.8, -2.2]], dtype=tf.float32)
dummy_v3 = tf.constant([[-4.1, 0.1, 1.2], [0.3, -1, -2]], dtype=tf.float32)
assert np.allclose(similarity(dummy_v1, dummy_v2).numpy(), [7.11, -1.84])
assert np.allclose(compute_loss(dummy_v1, dummy_v2, dummy_v3, delta=5.0).numpy(), [0.0, 3.88])
# -
# Once loss is working, let's train our model by our usual means.
opt = keras.optimizers.Adam(1e-3)
# ### Training loop
#
# Just as we always do, we can now train DSSM on minibatches and periodically measure recall on validation data.
#
#
# __Note 1:__ DSSM training may be very sensitive to the choice of batch size. Small batch size may decrease model quality.
#
# __Note 2:__ here we use the same dataset as __"test set"__ and __"validation (dev) set"__.
#
# In any serious scientific experiment, those must be two separate sets. Validation is for hyperparameter tuning and test is for final eval only.
#
# +
import pandas as pd
from IPython.display import clear_output
from tqdm import tqdm
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=256, cycle=True)
loss_history = []
dev_recall_history = []
# -
# infinite training loop. Stop it manually or implement early stopping
for batch in iterate_minibatches(train, batch_size=256, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, v_a_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'])
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'])
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'])
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 100
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
print("Mean recall:", np.mean(dev_recall_history[-10:]))
assert np.mean(dev_recall_history[-10:]) > 0.85, "Please train for at least 85% recall on test set. "\
"You may need to change vectorizer model for that."
print("Well done!")
# # Final evaluation (1 point)
#
# Let's see how well does our model perform on actual question answering.
#
# Given a question and a set of possible answers, pick answer with highest similarity to estimate accuracy.
# +
# optional: build tf graph required for select_best_answer
# <...>
def select_best_answer(question, possible_answers):
"""
Predicts which answer best fits the question
:param question: a single string containing a question
:param possible_answers: a list of strings containing possible answers
:returns: integer - the index of best answer in possible_answer
"""
question_vec = question_vectorizer([question], is_train=False)
answers_vec = answer_vectorizer(possible_answers, is_train=False)
scores = tf.matmul(question_vec, tf.transpose(answers_vec))
return tf.math.argmax(tf.reshape(scores, [-1])).numpy()
# +
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
def draw_results(question, possible_answers, predicted_index, correct_indices):
print("Q:", question, end='\n\n')
for i, answer in enumerate(possible_answers):
print("#%i: %s %s" % (i, '[*]' if i == predicted_index else '[ ]', answer))
print("\nVerdict:", "CORRECT" if predicted_index in correct_indices else "INCORRECT",
"(ref: %s)" % correct_indices, end='\n' * 3)
for i in [1, 100, 1000, 2000, 3000, 4000, 5000]:
draw_results(test.iloc[i].question, test.iloc[i].options,
predicted_answers[i], test.iloc[i].correct_indices)
question = "Why is Putin thief?" # your question here!
possible_answers1 = [
"Because",
"It is lie",
"Because Medvedev's vicious influence"
]
possible_answers2 = [
"Because",
"It is lie",
"Medvedev's vicious influence"
]
predicted_answer1 = select_best_answer(question, possible_answers1)
predicted_answer2 = select_best_answer(question, possible_answers2)
draw_results(question, possible_answers1,
predicted_answer1, [0])
draw_results(question, possible_answers2,
predicted_answer2, [0])
# ### Bonus tasks
#
# There are many ways to improve our question answering model. Here's a bunch of things you can do to increase your understanding and get bonus points.
# At first add early stopping conditionand evaluate default model again
# +
# infinite training loop. Stop it manually or implement early stopping
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=256, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer()
answer_vectorizer = Vectorizer()
for batch in iterate_minibatches(train, batch_size=256, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, v_a_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'])
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'])
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'])
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
# +
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
# ### 1. Hard Negatives (3+ pts)
#
# Not all wrong answers are equally wrong. As the training progresses, _most negative examples $a^-$ will be to easy._ So easy in fact, that loss function and gradients on such negatives is exactly __0.0__. To improve training efficiency, one can __mine hard negative samples__.
#
# Given a list of answers,
# * __Hard negative__ is the wrong answer with highest similarity with question,
#
# $$a^-_{hard} = \underset {a^-} {argmax} \space sim[V_q(q), V_a(a^-)]$$
#
# * __Semi-hard negative__ is the one with highest similarity _among wrong answers that are farther than positive one. This option is more useful if some wrong answers may actually be mislabelled correct answers.
#
# * One can also __sample__ negatives proportionally to $$P(a^-_i) \sim e ^ {sim[V_q(q), V_a(a^-_i)]}$$
#
#
# The task is to implement at least __hard negative__ sampling and apply it for model training.
#
#
#
#
# #### 1.1 Try hard negatives in batch
# +
# infinite training loop. Stop it manually or implement early stopping
BATCH_SIZE = 256
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=BATCH_SIZE, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer()
answer_vectorizer = Vectorizer()
for batch in iterate_minibatches(train, batch_size=BATCH_SIZE, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
all_scores = tf.einsum('ab,cb->ac', v_q, v_a_wrong)
hard_wrong = tf.matmul(tf.one_hot(tf.argmax(all_scores, axis=1), v_q.shape[0]), v_a_wrong)
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, hard_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'])
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'])
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'])
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
# #### Try semi-hard negatives
# +
# infinite training loop. Stop it manually or implement early stopping
BATCH_SIZE = 256
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=BATCH_SIZE, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer()
answer_vectorizer = Vectorizer()
for batch in iterate_minibatches(train, batch_size=BATCH_SIZE, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
all_scores = tf.einsum('ab,cb->ac', v_q, v_a_wrong)
hard_wrong = tf.matmul(tf.one_hot(tf.math.top_k(all_scores, k=2)[1][:, 1], v_q.shape[0]), v_a_wrong)
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, hard_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'])
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'])
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'])
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
# ### 2. Bring Your Own Model (3+ pts)
# In addition to Universal Sentence Encoder, one can also train a new model.
# * You name it: convolutions, RNN, self-attention
# * Use pre-trained ELMO or FastText embeddings
# * Monitor overfitting and use dropout / word dropout to improve performance
#
# __Note:__ if you use ELMO please note that it requires tokenized text while USE can deal with raw strings. You can tokenize data manually or use tokenized=True when reading dataset.
#
#
# * hard negatives (strategies: hardest, hardest farter than current, randomized)
# * train model on the full dataset to see if it can mine answers to new questions over the entire wikipedia. Use approximate nearest neighbor search for fast lookup.
# +
import io
def load_vectors(fname):
fin = io.open(fname, 'r', encoding='utf-8', newline='\n', errors='ignore')
n, d = map(int, fin.readline().split())
data = {}
for line in fin:
tokens = line.rstrip().split(' ')
data[tokens[0]] = list(map(float, tokens[1:]))
return data
# -
fast_text = load_vectors('wiki-news-300d-1M.vec')
class Vectorizer2_0(keras.Model):
def __init__(self, output_size=256, hid_size=256, universal_sentence_encoder=universal_sentence_encoder):
""" A small feedforward network on top of universal sentence encoder. 2-3 layers should be enough """
super(Vectorizer2_0, self).__init__()
self.universal_sentence_encoder = universal_sentence_encoder
self.emb_size = 300
self.out_size = output_size
# define a few layers to be applied on top of u.s.e.
# note: please make sure your final layer comes with _linear_ activation
self.first = L.Dense(self.out_size)
self.gru = L.GRU(self.out_size)
self.bos = self.add_variable("bos", shape=[self.emb_size])
self.eos = self.add_variable("eos", shape=[self.emb_size])
self.unk = self.add_variable("unk", shape=[self.emb_size])
def tokenize(self, phrase):
tokens = [fast_text[tok] if tok in fast_text else self.unk
for tok in phrase.split()]
return [self.bos] + tokens + [self.eos]
def __call__(self, input_phrases, is_train=True):
"""
Apply vectorizer. Use dropout and any other hacks at will.
:param input_phrases: [batch_size] of tf.string
:param is_train: if True, apply dropouts and other ops in train mode,
if False - evaluation mode
:returns: predicted phrase vectors, [batch_size, output_size]
"""
batch_size = len(input_phrases)
splitted_phrases = list(map(self.tokenize, input_phrases))
lens = np.array(list(map(len, splitted_phrases)))
max_len = max(lens)
batch = tf.concat(
[[tf.concat([phrase + [self.eos] * (max_len - len(phrase))], axis=0)] for phrase in splitted_phrases],
axis=0
)
mask = tf.convert_to_tensor(
[[True] * len(phrase) + [False] * (max_len - len(phrase)) for phrase in splitted_phrases]
)
assert batch.shape == (batch_size, max_len, self.emb_size), batch.shape
gru_result = self.gru(inputs=batch, mask=mask)
assert gru_result.shape == (batch_size, self.out_size), gru_result.shape
use_output = self.first(self.universal_sentence_encoder(input_phrases)['outputs'])
result = tf.concat([gru_result, use_output], axis=1)
assert result.shape == (batch_size, self.out_size * 2)
return result
# +
# infinite training loop. Stop it manually or implement early stopping
BATCH_SIZE = 256
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=BATCH_SIZE, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer2_0()
answer_vectorizer = Vectorizer2_0()
for batch in iterate_minibatches(train, batch_size=BATCH_SIZE, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
all_scores = tf.einsum('ab,cb->ac', v_q, v_a_wrong)
hard_wrong = tf.gather(v_a_wrong, tf.argmax(all_scores, axis=1).numpy(), axis=0)
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, hard_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'])
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'])
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'])
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
# Seems, that model overfeet. Try add word_dropout.
class Vectorizer2_0(keras.Model):
def __init__(self, output_size=256, hid_size=256, universal_sentence_encoder=universal_sentence_encoder,
word_dropout=0.0):
""" A small feedforward network on top of universal sentence encoder. 2-3 layers should be enough """
super(Vectorizer2_0, self).__init__()
self.universal_sentence_encoder = universal_sentence_encoder
self.emb_size = 300
self.out_size = output_size
self.word_dropout = word_dropout
# define a few layers to be applied on top of u.s.e.
# note: please make sure your final layer comes with _linear_ activation
self.first = L.Dense(self.out_size)
self.gru = L.GRU(self.out_size)
self.bos = self.add_variable("bos", shape=[self.emb_size])
self.eos = self.add_variable("eos", shape=[self.emb_size])
self.unk = self.add_variable("unk", shape=[self.emb_size])
def tokenize(self, phrases, is_train):
dropout = self.word_dropout if is_train else 0.0
tokens = list(map(lambda phrase:
[fast_text[tok] if tok in fast_text and np.random.rand(1) > dropout else self.unk
for tok in phrase.split()],
phrases
))
return list(map(lambda toks: [self.bos] + toks + [self.eos], tokens))
def __call__(self, input_phrases, is_train=True):
"""
Apply vectorizer. Use dropout and any other hacks at will.
:param input_phrases: [batch_size] of tf.string
:param is_train: if True, apply dropouts and other ops in train mode,
if False - evaluation mode
:returns: predicted phrase vectors, [batch_size, output_size]
"""
batch_size = len(input_phrases)
splitted_phrases = self.tokenize(input_phrases, is_train)
lens = np.array(list(map(len, splitted_phrases)))
max_len = max(lens)
batch = tf.concat(
[[tf.concat([phrase + [self.eos] * (max_len - len(phrase))], axis=0)] for phrase in splitted_phrases],
axis=0
)
mask = tf.convert_to_tensor(
[[True] * len(phrase) + [False] * (max_len - len(phrase)) for phrase in splitted_phrases]
)
assert batch.shape == (batch_size, max_len, self.emb_size), batch.shape
gru_result = self.gru(inputs=batch, mask=mask)
assert gru_result.shape == (batch_size, self.out_size), gru_result.shape
use_output = self.first(self.universal_sentence_encoder(input_phrases)['outputs'])
result = tf.concat([gru_result, use_output], axis=1)
assert result.shape == (batch_size, self.out_size * 2)
return result
# +
# infinite training loop. Stop it manually or implement early stopping
BATCH_SIZE = 256
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=BATCH_SIZE, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer2_0(word_dropout=0.15)
answer_vectorizer = Vectorizer2_0(word_dropout=0.15)
for batch in iterate_minibatches(train, batch_size=BATCH_SIZE, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
all_scores = tf.einsum('ab,cb->ac', v_q, v_a_wrong)
hard_wrong = tf.gather(v_a_wrong, tf.argmax(all_scores, axis=1).numpy(), axis=0)
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, hard_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'], is_train=False)
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'], is_train=False)
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'], is_train=False)
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# #### Try Bidirectional GRU
class Vectorizer2_0(keras.Model):
def __init__(self, output_size=256, hid_size=256, universal_sentence_encoder=universal_sentence_encoder,
word_dropout=0.0):
""" A small feedforward network on top of universal sentence encoder. 2-3 layers should be enough """
super(Vectorizer2_0, self).__init__()
self.universal_sentence_encoder = universal_sentence_encoder
self.emb_size = 300
self.out_size = output_size
self.hid_size = hid_size
self.word_dropout = word_dropout
# define a few layers to be applied on top of u.s.e.
# note: please make sure your final layer comes with _linear_ activation
self.first = L.Dense(self.out_size)
self.gru = L.Bidirectional(L.GRU(self.hid_size))
self.second = L.Dense(self.out_size)
self.bos = self.add_variable("bos", shape=[self.emb_size])
self.eos = self.add_variable("eos", shape=[self.emb_size])
self.unk = self.add_variable("unk", shape=[self.emb_size])
def tokenize(self, phrases, is_train):
dropout = self.word_dropout if is_train else 0.0
tokens = list(map(lambda phrase:
[fast_text[tok] if tok in fast_text and np.random.rand(1) > dropout else self.unk
for tok in phrase.split()],
phrases
))
return list(map(lambda toks: [self.bos] + toks + [self.eos], tokens))
def __call__(self, input_phrases, is_train=True):
"""
Apply vectorizer. Use dropout and any other hacks at will.
:param input_phrases: [batch_size] of tf.string
:param is_train: if True, apply dropouts and other ops in train mode,
if False - evaluation mode
:returns: predicted phrase vectors, [batch_size, output_size]
"""
batch_size = len(input_phrases)
splitted_phrases = self.tokenize(input_phrases, is_train)
lens = np.array(list(map(len, splitted_phrases)))
max_len = max(lens)
batch = tf.concat(
[[tf.concat([phrase + [self.eos] * (max_len - len(phrase))], axis=0)] for phrase in splitted_phrases],
axis=0
)
mask = tf.convert_to_tensor(
[[True] * len(phrase) + [False] * (max_len - len(phrase)) for phrase in splitted_phrases]
)
assert batch.shape == (batch_size, max_len, self.emb_size), batch.shape
gru_result = self.second(self.gru(inputs=batch, mask=mask))
assert gru_result.shape == (batch_size, self.out_size), gru_result.shape
use_output = self.first(self.universal_sentence_encoder(input_phrases)['outputs'])
result = tf.concat([gru_result, use_output], axis=1)
assert result.shape == (batch_size, self.out_size * 2)
return result
# +
# infinite training loop. Stop it manually or implement early stopping
BATCH_SIZE = 256
ewma = lambda x, span: pd.DataFrame({'x': x})['x'].ewm(span=span).mean().values
dev_batches = iterate_minibatches(test, batch_size=BATCH_SIZE, cycle=True)
loss_history = []
dev_recall_history = []
question_vectorizer = Vectorizer2_0(word_dropout=0.15)
answer_vectorizer = Vectorizer2_0(word_dropout=0.15)
for batch in iterate_minibatches(train, batch_size=BATCH_SIZE, cycle=True):
with tf.GradientTape() as tape:
v_q = question_vectorizer(batch['questions'])
v_a_correct = answer_vectorizer(batch['correct_answers'])
v_a_wrong = answer_vectorizer(batch['wrong_answers'])
all_scores = tf.einsum('ab,cb->ac', v_q, v_a_wrong)
hard_wrong = tf.gather(v_a_wrong, tf.argmax(all_scores, axis=1).numpy(), axis=0)
loss = tf.math.reduce_mean(compute_loss(v_q, v_a_correct, hard_wrong))
loss_history.append(loss.numpy())
variables = question_vectorizer.trainable_variables + answer_vectorizer.trainable_variables
grads = tape.gradient(loss, variables)
opt.apply_gradients(zip(grads, variables))
if len(loss_history) % 50 == 0:
# measure dev recall = P(correct_is_closer_than_wrong | q, a+, a-)
dev_batch = next(dev_batches)
test_v_q = question_vectorizer(dev_batch['questions'], is_train=False)
test_v_a_correct = answer_vectorizer(dev_batch['correct_answers'], is_train=False)
test_v_a_wrong = answer_vectorizer(dev_batch['wrong_answers'], is_train=False)
correct_is_closer = tf.greater(similarity(test_v_q, test_v_a_correct),
similarity(test_v_q, test_v_a_wrong))
recall_t = tf.reduce_mean(tf.cast(correct_is_closer, dtype=tf.float32)).numpy()
dev_recall_history.append(recall_t)
if len(loss_history) % 50 == 0:
clear_output(True)
plt.figure(figsize=[12, 6])
plt.subplot(1, 2, 1), plt.title('train loss (hinge)'), plt.grid()
plt.scatter(np.arange(len(loss_history)), loss_history, alpha=0.1)
plt.plot(ewma(loss_history, span=100))
plt.subplot(1, 2, 2), plt.title('dev recall (1 correct vs 1 wrong)'), plt.grid()
dev_time = np.arange(1, len(dev_recall_history) + 1) * 50
plt.scatter(dev_time, dev_recall_history, alpha=0.1)
plt.plot(dev_time, ewma(dev_recall_history, span=10))
plt.show()
if len(loss_history) == 4000:
break
predicted_answers = [
select_best_answer(question, possible_answers)
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
accuracy = np.mean([
answer in correct_ix
for answer, correct_ix in zip(predicted_answers, test['correct_indices'].values)
])
print("Accuracy: %0.5f" % accuracy)
assert accuracy > 0.65, "we need more accuracy!"
print("Great job!")
# -
# ### 3. Search engine (3+ pts)
#
# Our basic model only selects answers from 2-5 available sentences in paragraph. You can extend it to search over __the whole dataset__. All sentences in all other paragraphs are viable answers.
#
# The goal is to train such a model and use it to __quickly find top-10 answers from the whole set__.
#
# * You can ask such model a question of your own making - to see which answers it can find among the entire training dataset or even the entire wikipedia.
# * Searching for top-K neighbors is easier if you use specialized methods: [KD-Tree](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.KDTree.html) or [HNSW](https://github.com/nmslib/hnswlib).
# * This task is much easier to train if you use hard or semi-hard negatives. You can even find hard negatives for one question from correct answers to other questions in batch - do so in-graph for maximum efficiency. See [1.] for more details.
#
#
from sklearn.neighbors import KDTree
# +
questions = [
question
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
]
answers = [
answer
for i, (question, possible_answers) in test[['question', 'options']].iterrows()
for answer in possible_answers
]
answers = list(set(answers))
# +
question_vectors = [
question_vectorizer([question], is_train=False).numpy()
for question in questions
]
answer_vectors = [
answer_vectorizer([answer], is_train=False).numpy()
for answer in answers
]
# -
question_vectors = np.concatenate(question_vectors, axis=0)
answer_vectors = np.concatenate(answer_vectors, axis=0)
# #### Try brute
for i in range(0, len(question_vectors), 5000):
top10_ids = [0] * 10
top10_results = [0] * 10
for j, ans in enumerate(answer_vectors):
res = np.sum(question_vectors[i] * ans)
for k in range(10):
if res > top10_results[k]:
top10_results = top10_results[:k] + [res] + top10_results[k:10]
top10_ids = top10_ids[:k] + [j] + top10_ids[k:10]
break
print(questions[i])
for ans in top10_ids:
print('\t' + answers[ans])
print()
# ### Try KD-Tree
tree = KDTree(answer_vectors)
best_answers = tree.query(question_vectors, k=10)
for i in range(0, len(question_vectors), 5000):
print(questions[i])
for ans in best_answers[1][i]:
print('\t' + answers[ans])
print()
# Result so bad, because KDTree can't optimize similarity metric.
|
week08_conversation/seminar.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bayashi-cl/statistical-learning/blob/main/note/06_LinerModelSelectionAndRegularization.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qlckyD4x6CuU"
# # 6 線形モデル選択と正則化
#
# この章では最小二乗法以外の線形モデルについて考える。
#
# * **部分集合選択** モデルに含まれる予測変数の集合のうち、最も「良い」部分集合を特定し、その部分集合に対して最小二乗法を適用する。
# * **縮小推定** 予測変数をすべて使って推定をするが、何らかの操作により係数の推定値を0やそれに近い値にする。
# * **次元削減** $p$個の予測変数を$M(<p)$次元部分空間に射影して最小二乗法を適用する。
#
# これらのモデルは、最小二乗法に対して次のような利点がある。
#
# * **予測精度** 最小二乗法はデータ数が予測変数の数よりも十分に大きければ十分な性能を発揮するが、そうでない場合は過学習などの問題が発生して分散が大きくなる。縮小推定では回帰係数に制約を課すことで予測精度を向上させる。
# * **モデルの解釈可能性** モデルの中に実際には応答変数と関係のない予測変数が含まれていると、モデルが必要以上に複雑になってしまう。このような無意味な変数を取り除くことでモデルの解釈が容易になる。
# + [markdown] id="stqDCdtXAFG-"
# ## 6.1 部分集合選択
# + [markdown] id="jVrR79WdAcA8"
# ### 補足 計算量の評価について
#
# アルゴリズムのコスト(計算回数)のことを(時間)計算量と呼ぶ。計算量を評価する際に、例えば入力サイズが$n$のアルゴリズムの計算コストが$3n^2 + 2n + 5$である場合、注目するべきは$n^2$の部分であり、それ以外の係数や1次より小さい項は$n$が増加した場合の計算量の増加度合いにはあまり関与しない。
#
# 数列の**オーダー**の記法を計算量でも用いる。
#
# #### $\Theta(O,o,\Omega , \omega)$記法
#
# 2つの計算量の増加度合いが同じ(くらい)であることを言いたい。
#
# $f(n)$が$g(n)$に対して次の条件を満たすとき、$f(n)$のオーダーが$g(n)$であるといい、$f(n) \in \Theta (g(n))$ と表記する。
#
# > ある定数 $n_0$ に対し、ある定数 $c_L, c_U > 0$ が存在し、$n_0$ 以上のすべての $n$ について次の式が成り立つ:
# > $$0 \le c_L \cdot g(n) \le f(n) \le c_U \cdot g(n)$$
#
# また、上の不等式の$c_L$側が成り立つとき、$f(n) \in \Omega (g(n))$、$c_U$側が成り立つとき、$f(n) \in O (g(n))$と表記する。
#
# 更に、大小関係が真に成り立つ($\le$ではなく$\lt$)場合は,
# それぞれ$f(n) \in \omega (g(n))$、$f(n) \in o (g(n))$と表記する。
#
# #### 参考文献
#
# * アルゴリズムイントロダクション 第3版 総合版(p36-)(該当部分は[Amazon](https://www.amazon.co.jp/dp/B078WPYHGN)で試し読みができます。)
#
#
# + [markdown] id="vfDoKe6FSzP2"
# ### 6.1.1 最良部分集合選択
#
# すべての予測変数の組み合わせを評価する手法。以下のアルゴリズムで行われる。
#
# 1. $M_0$を予測変数を持たない**ヌルモデル**とする。ヌルモデルは予測値として標本平均を返す。
# 1. $k = 1, 2, \ldots , p$について以下の手順を行う。
# 1. $k$個の予測変数を持つ$\binom{p}{k}$個のモデルに対してRSSや$R^2$を計算する。
# 1. 最良のモデルを$M_k$とする。
# 1. $M_0, \ldots , M_p$から最良のモデルをBICや修正$R^2$などで選ぶ。
#
# step3でのモデル選択の際にはRSSや$R^2$が特徴の数に対して単調減少/増加することに注意する。RSSが小さい・$R^2$が大きいことは単に訓練データに対して誤差が小さいことを示しているということであり、重要なのはテストデータに対する誤差である。
#
# 最良部分集合選択は単純であり、得られるモデルも厳密に最良であるといえるが、計算量のオーダーがモデル評価の計算量を$\Theta(V)$として$\Theta(2^pV)$であり、$p \ge 40$の場合には現実的な時間で解くことが難しくなる。
# + [markdown] id="v8ye3UCsmPuG"
# ### 6.1.2 ステップワイズ法
#
# モデルを限定することで効率的に探索を行う。
# + [markdown] id="Fat7RhC7zbG0"
# #### 変数増加法
#
# 当てはめを最も良くする予測変数を順にモデルに追加していく手法。以下のアルゴリズムで行われる。
#
# 1. ヌルモデルを$M_0$とする。
# 1. $k = 1, 2, \ldots , p$について以下の手順を行う。
# 1. $M_k$に含まれない予測変数のうち1つを加えた$(p-k)$個のモデルを考える。
# 1. 最良のモデルを$M_{k+1}$とする。
# 1. $M_0, \ldots , M_p$から最良のモデルをBICや修正$R^2$などで選ぶ。
#
# 変数増加法は$p>n$の場合にも適用することができる。
# + [markdown] id="N_iEB6z9zcKN"
# #### 変数減少法
#
# 不要な予測変数を順に削除していく手法。以下のアルゴリズムで行われる。
#
# 1. すべての予測変数を含むモデルをを$M_p$とする。
# 1. $k = p, p-1, \ldots , 1$について以下の手順を行う。
# 1. $M_k$に含まれる予測変数のうち1つを除いた$k$個のモデルを考える。
# 1. 最良のモデルを$M_{k-1}$とする。
# 1. $M_0, \ldots , M_p$から最良のモデルをBICや修正$R^2$などで選ぶ。
#
# + [markdown] id="FwHN1CzH0UQ0"
# #### 変数増減法
#
# 増加法と減少法を混ぜたもの。変数増加法と同様に予測変数を追加していくが、その際にモデルの当てはめに貢献しない予測変数を取り除く。
# + [markdown] id="KdyjrTAJ0v1e"
# これらの手法の計算量は$\Theta(p^2V)$であり最良部分集合選択よりも優れているが、最良部分集合選択と同一のモデルが選択されるとは限らない。
# + [markdown] id="w2T21ZIh18oJ"
# ### 6.1.2 最適モデルの選択
#
# RSSや$R^2$は予測変数の数に依存し、すべての予測変数を含むモデルで最小/最大となるために、予測変数の数の異なるモデルを比較するのに適していない。また、訓練データの誤差の影響を受けているため、テストデータに対する誤差を小さくするためには何らかの修正を加える必要がある。
#
# よく用いられるのは以下の2つの方法である。
#
# 1. 過学習によるバイアスを考慮して訓練誤差を修正することで間接的にテスト誤差を推定する。
# 1. ホールドアウト検証や交差検証法により直接的にテスト誤差を推定する。
# + [markdown] id="VKJmO7L-5PAz"
# #### $C_p$, AIC, BIC
#
# $C_p$, AIC, BICはMSE($:=\textrm{RSS}/n$)に罰則項を追加したものである。それぞれ、
#
# $$C_p = \frac{1}{n}(\textrm{RSS} + 2d\hat{\sigma}^2)$$
# $$\textrm{AIC} = \frac{1}{n\hat{\sigma}^2}(\textrm{RSS} + 2d\hat{\sigma}^2)$$
# $$\textrm{BIC} = \frac{1}{n\hat{\sigma}^2}(\textrm{RSS} + \log{(n)}d\hat{\sigma}^2)$$
#
# となる(定数項は省略されている)。
#
# ここで、$\hat{\sigma}^2$は応答変数の測定値に対する誤差の分散の推定値であり、これが不偏であれば$C_p$はテストデータのMSEとして不偏であることが言える。すなわち、テストデータでの誤差が小さいモデルは$C_p$も小さくなるため、これが最小のモデルを選べば良い。
#
# $C_p$とAICは比例関係にあるが、これらに対してBICは予測変数が多いモデルに対してより重い罰則を与える。
#
# #### 自由度調整済み$R^2$
#
# 自由度調整済み$R^2$は次の式で表される。
#
# $$自由度調整済み R^2 = 1 - \frac{\textrm{RSS}/(n-d-1)}{\textrm{TSS}/(n-1)}$$
#
# モデルに対してRSSをほとんど改善しない予測変数を追加すると、$\textrm{RSS}/(n-d-1)$が増加し、自由度調整済み$R^2$は減少する。
# + [markdown] id="-ilGM-f0F5Xd"
# #### ホールドアウト検証・交差検証
#
# ホールドアウト検証や交差検証を使って直接テスト誤差を推定することもできる。この手法はテスト誤差の直接の推定である点や、真のモデルについての過程がAICなどより少ないという点で優れている。従来は$p$や$n$が大きい場合に交差検証の計算量が大きくなるという問題があったが、コンピュータの計算速度の進歩により改善されている。
# + [markdown] id="XnLOerpBLHsG"
# テスト誤差の推定値は予測変数の個数を変えたとしてもほとんど変化しない場合がある。そのような場合には推定値の標準誤差を計算し、推定値の最小値から1標準誤差以内のモデルのうち最も予測変数の数が少ないものを選択する。これは**1標準誤差ルール**と呼ばれる。
# + [markdown] id="v2I__E8WCdWx"
# ## 考察
#
# ### 最良部分集合選択
#
# * 0-1ナップザック問題に似ている
# * 似たモデルに対して何回も評価をしていて無駄が大きい?
# * 動的計画法で厳密解の計算量を落とせそうな気がする。
# * 時間・空間ともに$\Theta(NB\log{p})$ (Bはバケット数)
# * DPやるには$R^2$に線形性っぽさが必要?
|
note/06_LinerModelSelectionAndRegularization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook can be used to make flow chart of OGGM. I think there is better software out there to do this.
# Eitherway feel free to use/adjust this notebook.
from schemdraw import flow
import schemdraw
from schemdraw import dsp
import matplotlib.pyplot as plt
# +
# Both the schemdraw.Drawing() and the d.add(flow.Start()) commends are needed to start the flow chart.
# From there on you can start drawing line and arrows and add boxes. The .draw() is needed to draw the flow chart.
d = schemdraw.Drawing(fontsize=11, color='black')
d.add(flow.Start(w=2, h=1.5, label='RGI \n & \nDEM'))
d.add(flow.Arrow('down', l=d.unit/2))
topo_pre = d.add(flow.Box(w=5, h=1.5, label="topographical data \n preprocessing"))
d.add(flow.Arrow('down', l=d.unit/2))
flow_pre = d.add(flow.Box(w=5, h=1.5, label="computation of \n the flow lines"))
d.add(flow.Arrow(l=d.unit/2))
width_pre = d.add(flow.Box(w=5, h=1.5, label="geometrical glacier \n width determination"))
d.add(flow.Arrow(l=d.unit/2))
wadjust_pre = d.add(flow.Box(w=5, h=1.5, label='width correction according \n to catchment areas and \n altitude–area distribution'))
d.add(flow.Arrow('right', xy=wadjust_pre.E, l=d.unit/2))
inversion = d.add(flow.Box(w=5, h=1.5, label='ice thinkness inversion', anchor='W'))
d.add(flow.Line('up', xy=inversion.N, l=d.unit/2))
MBcal = d.add(flow.Box(w=5, h=1.5, label='mass balance \n calibration', anchor='S'))
d.add(flow.Arrow(l=d.unit/2))
d.add(flow.Line('up', xy=MBcal.N, l=d.unit/2))
clim_pre = d.add(flow.Box(w=5, h=1.5, label='Climate preprocessing', anchor='S'))
d.add(flow.Arrow(l=d.unit/2))
d.add(flow.Arrow('right', xy=inversion.E, l=d.unit/2))
d7 = d.add(flow.Box(w=5, h=1.5, label='Glacier simulation', anchor='W'))
d.add(flow.Line('right', xy=topo_pre.E, l=d.unit+1))
d.add(flow.Arrow('down', l=d.unit/2 + 0.75))
d.add(flow.Line('right', xy=clim_pre.E, l=d.unit/2))
d8 = d.add(flow.Start(w=3, h=2, label='Baseline \nclimate', anchor='W'))
d.add(flow.Arrow('left', l=d.unit/2, xy=d8.W))
d.draw()
# d.save('your_path/filename');
# +
d = schemdraw.Drawing(fontsize=11, color='black')
d.add(flow.Start(w=3, h=2, label='Glacier outlines \n & DEM'))
d.add(flow.Arrow('down', l=d.unit/4))
topo_pre = d.add(flow.Box(w=5, h=1.5, label="topographical data \n preprocessing"))
d.add(flow.Arrow('down', l=d.unit/2))
flow_pre = d.add(flow.Box(w=5, h=1.5, label="computation of \n the flow lines"))
d.add(flow.Arrow(l=d.unit/2))
width_pre = d.add(flow.Box(w=5, h=1.5, label="geometrical glacier \n width determination"))
d.add(flow.Arrow(l=d.unit/2))
wadjust_pre = d.add(flow.Box(w=5, h=1.5, label='width correction according \n to catchment areas and \n altitude–area distribution'))
d.add(flow.Line('right', xy=wadjust_pre.E, l=d.unit/2))
d.add(flow.Line('right', xy=topo_pre.E, l=d.unit/2))
pre_pro= d.add(flow.Box(w=1.5, h=9, rotation=90, label='preprocessed geometry data'))
d.add(flow.Arrow('right', xy=pre_pro.E, l=d.unit*1.5))
inversion = d.add(flow.Box(w=5, h=1.5, label='ice thinkness inversion', anchor='W'))
d.add(flow.Line('up', l=d.unit/2, xy=inversion.N))
MBcal = d.add(flow.Box(w=5, h=1.5, label='mass balance \nmodel calibration', anchor='S'))
d.add(flow.Line('up', l=d.unit/2, xy=MBcal.N))
clim_prepro = d.add(flow.Box(w=5, h=1.5, label='climate preprocessing', anchor='S'))
d.add(flow.Line('left', l=d.unit/4, xy=MBcal.W))
wgms = d.add(flow.Start(w=3, h=2, label='Mass balance \ndata', anchor='E'))
d.add(flow.Arrow('right', l=d.unit/4, xy=wgms.E))
d.add(flow.Line('left', l=d.unit/4, xy=clim_prepro.W))
clim = d.add(flow.Start(w=3, h=2, label='Baseline \n climate', anchor='E'))
d.add(flow.Arrow('right', l=d.unit/4, xy=clim.E))
d.add(flow.Arrow('down', l=d.unit/2, xy=clim_prepro.S))
d.add(flow.Arrow('down', l=d.unit/2, xy=MBcal.S))
d.add(flow.Arrow('right', l=d.unit/2, xy=inversion.E))
Dynamics = d.add(flow.Box(w=5, h=1.5, label='glacier simulation', anchor='W'))
d.add(flow.Line('right', l=d.unit/2))
d.add(flow.Line('right', l=d.unit*1.3, xy=MBcal.E))
d.add(flow.Line('right', l=d.unit*1.3, xy=clim_prepro.E))
d.add(flow.Arrow('down', l=d.unit*1.75))
d.draw()
# d.save('your_path/filename');
# -
|
OGGM_flowchart.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import matplotlib.pyplot as plt
import json
plt.style.reload_library()
plt.style.use('singlecolumn')
def gsm_fidelity(data):
'''return ground state manifold fidelity'''
if round(data['J']/data['B'], 2) > 1:
return np.sum(data['eigoccs'][:2])
else:
return data['eigoccs'][0]
# -
plt.rcParams.find_all('axes.grid')
# # Loading and checking data
# ## Cooling
# +
data_dir = "../data/TFIM/logsweep/DM/cooling/"
files = sorted(os.listdir(data_dir))
cooling_data = []
for file in files:
if not file.endswith('.json'): continue
cooling_data.append(json.load(open(data_dir+file, 'r')))
# + [markdown] heading_collapsed=true
# ### density matrix norm check
# + hidden=true
print(' L, K, J/B, Log10(Trace of the DM - 1)')
print(
*sorted((d['L'],
d['K'],
round(d['J']/d['B'],2),
round(np.log10(np.abs(np.sum(d['eigoccs'])-1)), 0)
) for d in cooling_data ),
sep='\n'
)
# + [markdown] hidden=true
# **Note:**
# the density matrix simulator accumulates numerical errors, producing a non-normalized final density matrix.
# We cannot get rid of the numerical error, but to get consistent results we nomalize the results (energy, fidelities) during dat analysis.
# -
# ## Reheating
# +
data_dir = "../data/TFIM/logsweep/DM/reheating/"
files = sorted(os.listdir(data_dir))
reheating_data = []
for file in files:
if not file.endswith('.json'): continue
reheating_data.append(json.load(open(data_dir+file, 'r')))
# + [markdown] heading_collapsed=true
# ### density matrix norm check
# + hidden=true
print(' L, K, J/B, Log10(Trace of the DM - 1)')
print(
*sorted((d['L'],
d['K'],
round(d['J']/d['B'],2),
round(np.log10(np.abs(np.sum(d['eigoccs'])-1)), 0),
round(np.sum(d['eigoccs']), 10)
) for d in reheating_data ),
sep='\n'
)
# -
# ## Iterative LogSweep cooling
# +
data_dir = "../data/TFIM/logsweep/DM/iterative/"
files = sorted(os.listdir(data_dir))
iterative_data = []
for file in files:
if not file.endswith('.json'): continue
iterative_data.append(json.load(open(data_dir+file, 'r')))
# + [markdown] heading_collapsed=true
# ### density matrix norm check
# + hidden=true
print(' L, K, J/B, Log10(|Trace of the DM - 1|)')
print(
*sorted((d['L'],
d['K'],
round(d['J']/d['B'],2),
round(np.log10(np.abs(np.sum(d['eigoccs'])-1)), 0),
round(np.sum(d['eigoccs']), 10)
) for d in iterative_data ),
sep='\n'
)
# -
# ## Available data summary
# +
print(' K , L, J/B ')
avail_cooling = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in cooling_data]
avail_reheating = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in reheating_data]
avail_iterative = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in iterative_data]
from itertools import product
for K, L, JvB in np.unique(avail_cooling + avail_reheating + avail_iterative, axis=0):
K = int(K)
L = int(L)
if L!=7: continue
print((K, L, JvB),
'C' if (K, L, JvB) in avail_cooling else ' ',
'R' if (K, L, JvB) in avail_reheating else ' ',
'It' if (K, L, JvB) in avail_iterative else ' '
)
# -
# # Varying energy gradation number K
L = 7
# ## cooling
# ### energy vs K
# + code_folding=[]
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), 'o-', label=f'$J/B={JvB}$')
plt.legend()
# -
# ### GS infidelity vs K
# + code_folding=[]
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'o-', label=f'$J/B={JvB}$')
plt.legend()
plt.xlabel('K')
plt.ylabel('GS manifold infidelity')
plt.xscale('log')
plt.yscale('log')
# + [markdown] heading_collapsed=true
# ## reheating
#
# + [markdown] hidden=true
# ### energy vs K
# + code_folding=[] hidden=true
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), 'x:', label=f'$J/B={JvB}$')
plt.legend()
# + [markdown] hidden=true
# ### GS infidelity vs K
# + code_folding=[] hidden=true
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'x:', label=f'$J/B={JvB}$')
plt.legend()
plt.xscale('log')
plt.yscale('log')
# + [markdown] heading_collapsed=true
# ## iterative
# + [markdown] hidden=true
# ### energy vs K
# + code_folding=[] hidden=true
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), '+--', label=f'$J/B={JvB}$')
plt.legend()
# + [markdown] hidden=true
# ### GS infidelity vs K
# + code_folding=[] hidden=true
# L = 7
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, '+--', label=f'$J/B={JvB}$')
plt.legend()
plt.xscale('log')
plt.yscale('log')
# -
# ## combined
# ### energy vs K
# +
# L = 7
plt.title(f'TFIM chain $L={L}$. Standard LogSweep density matrix sim')
# cooling
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), 'o-', label=f'cooling $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# reheating
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), 'x:', label=f'reheating $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# iterative
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, np.array(E_l)/np.array(norms_l), '+--', label=f'iterative $J/B={JvB}$')
plt.legend()
# -
# #### logarithmic
# +
# L = 7
plt.figure(figsize=(10, 5))
plt.title(f'TFIM chain $L={L}$. Standard LogSweep density matrix sim')
# cooling
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, 1 + np.array(E_l)/np.array(norms_l), 'o-', label=f'cooling $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# reheating
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, 1 + np.array(E_l)/np.array(norms_l), 'x:', label=f'reheating $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# iterative
for JvB in [.2, 1, 5]:
data_iterator = ((d['K'], d['energy'], np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, E_l, norms_l = zip(*sorted(data_iterator))
plt.plot(K_l, 1 + np.array(E_l)/np.array(norms_l), '+--', label=f'iterative $J/B={JvB}$')
plt.yscale('log')
plt.xscale('log')
plt.ylabel('energy relative to GS '
r'$(1 + \mathrm{Tr}[\rho H]/\vert E_\mathrm{GS}\vert)$')
plt.legend(bbox_to_anchor = (1, 0.5), loc = 'center left')
# -
# ### GS infidelity vs K
# + code_folding=[]
L = 7
#plt.title(f'TFIM chain $L={L}$. Standard LogSweep density matrix sim')
JvBlist = [.2, 1, 5]
# cooling
for JvB in JvBlist:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, '+:', label=f'cooling $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# reheating
for JvB in JvBlist:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'x:', label=f'reheating $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
for JvB in JvBlist:
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'o:', label=f'iterative $J/B={JvB}$')
handles, labels = plt.gca().get_legend_handles_labels()
handles = np.array([[[plt.Line2D([],[],marker='',ls='')] + handles[3*i:3*i+3]]
for i in range(3)]).flatten()
ll = ['$JvB = {JvB}$' for JvB in JvBlist]
labels = ['cooling', *ll, 'reheating', *ll, 'iterative', *ll]
plt.legend(handles, labels, ncol=3, bbox_to_anchor = (0.5, 1), loc='lower center')
plt.yscale('log')
plt.xscale('log')
plt.xlabel('K')
plt.xticks([2, 10, 50], [2, 10, 50])
plt.ylabel('ground space infidelity')
# + code_folding=[]
L = 7
#plt.title(f'TFIM chain $L={L}$. Standard LogSweep density matrix sim')
JvBlist = [.2, 1, 5]
def fitfunc(x, a, b):
return a * x**-b
from scipy.optimize import curve_fit
clrs = plt.rcParams['axes.prop_cycle'].by_key()['color']
# reheating
for JvB, c in zip(JvBlist, clrs):
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'x', label=f'reheating $J/B={JvB}$', c=c)
# iterative cooling
annotate_xy_list = [
[4, 0.025],
[10, 0.18],
[6, 0.07]
]
for JvB, c, annotate_xy in zip(JvBlist, clrs, annotate_xy_list):
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, '.', label=f'iterative $J/B={JvB}$', color=c)
# popt, pcov = curve_fit(fitfunc, K_l[10:], infidelity_l[10:])
# plt.plot([2, 50], fitfunc(np.array([2,50]), *popt), '-', c=c, lw=1)
# plt.annotate(r'$\mathbf{\propto K^{-'+f'{popt[1]:.2f}'+r'}}$',
# annotate_xy, fontsize=16, zorder=10,
# va='center', ha='left')
# one-shot cooling
for JvB, c in zip(JvBlist, clrs):
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in cooling_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, '^', label=f'cooling $J/B={JvB}$', c=c)
hndlbl = [[plt.Line2D([],[],marker='',color=clrs[0]), '$J/B=.2$'],
[plt.Line2D([],[],marker='.',ls='',color='grey'), 'iterative\ncooling'],
[plt.Line2D([],[],marker='',color=clrs[1]), '$J/B=1$'],
[plt.Line2D([],[],marker='x',ls='',color='grey'), 'reheating'],
[plt.Line2D([],[],marker='',color=clrs[2]), '$J/B=5$'],
[plt.Line2D([],[],marker='^',ls='',color='grey', lw=1),
'single-shot\ncooling'],
]
plt.legend(*zip(*hndlbl), ncol=3, bbox_to_anchor = (0.5, 1), loc='lower center')
plt.yscale('log')
plt.xscale('log')
plt.xlabel('K')
plt.xticks([2, 10, 50], [2, 10, 50])
plt.ylabel('ground space infidelity')
#plt.savefig('../figures/Ksingleshot.pdf', bbox_inches='tight')
# -
# ### without cooling
# + code_folding=[]
L = 7
#plt.title(f'TFIM chain $L={L}$. Standard LogSweep density matrix sim')
JvBlist = [.2, 1, 5]
def fitfunc(x, a, b):
return a * x**-b
from scipy.optimize import curve_fit
clrs = plt.rcParams['axes.prop_cycle'].by_key()['color']
# reheating
for JvB, c in zip(JvBlist, clrs):
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'x', label=f'reheating $J/B={JvB}$', c=c)
annotate_xy_list = [
[4, 0.025],
[10, 0.18],
[6, 0.07]
]
for JvB, c, annotate_xy in zip(JvBlist, clrs, annotate_xy_list):
data_iterator = ((d['K'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in iterative_data
if d['L'] == L and np.isclose(d['J']/d['B'], JvB))
K_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(K_l, infidelity_l, 'o', label=f'iterative $J/B={JvB}$', color=c)
popt, pcov = curve_fit(fitfunc, K_l[10:], infidelity_l[10:])
plt.plot([2, 50], fitfunc(np.array([2,50]), *popt), '-', c=c, lw=1)
plt.annotate(r'$\mathbf{\propto K^{-'+f'{popt[1]:.2f}'+r'}}$',
annotate_xy, fontsize=16, zorder=10,
va='center', ha='left')
hndlbl = [[plt.Line2D([],[],marker='',color=clrs[0]), '$J/B=.2$'],
[plt.Line2D([],[],marker='o',ls='',color='grey'), 'iterative cooling'],
[plt.Line2D([],[],marker='',color=clrs[1]), '$J/B=1$'],
[plt.Line2D([],[],marker='x',ls='',color='grey'), 'reheating'],
[plt.Line2D([],[],marker='',color=clrs[2]), '$J/B=5$'],
[plt.Line2D([],[],marker='',ls='-',color='grey', lw=1), 'fit'],
]
plt.legend(*zip(*hndlbl), ncol=3, bbox_to_anchor = (0.5, 1), loc='lower center')
plt.yscale('log')
plt.xscale('log')
plt.xlabel('K')
plt.xticks([2, 10, 50], [2, 10, 50])
plt.ylabel('ground space infidelity')
#plt.savefig('../figures/Kinfidelity.pdf', bbox_inches='tight')
# -
# # scaling with system size L
# ## check available data at fixed K
# +
K = 10
print(f'available data for K = {K}:')
print(' K , L, J/B ')
avail_cooling = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in cooling_data if d['K']==K]
avail_reheating = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in reheating_data if d['K']==K]
avail_iterative = [(d['K'], d['L'], round(d['J']/d['B'],1)) for d in iterative_data if d['K']==K]
from itertools import product
for K, L, JvB in np.unique(avail_cooling + avail_reheating + avail_iterative, axis=0):
K = int(K)
L = int(L)
print((K, L, JvB),
'C' if (K, L, JvB) in avail_cooling else ' ',
'R' if (K, L, JvB) in avail_reheating else ' ',
'It' if (K, L, JvB) in avail_iterative else ' '
)
# -
# ### energy vs L
# + code_folding=[]
K=10
plt.figure(figsize=(9, 5))
plt.title(f'TFIM chain. Standard LogSweep(K={K}) density matrix sim')
# cooling
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], d['energy'], np.sum(d['eigoccs']))
for d in cooling_data
if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
L_l, energy_l, norms_l = zip(*sorted(data_iterator))
energy_l /= np.array(norms_l)
plt.plot(L_l, energy_l, 'o-', label=f'cooling $J/B={JvB}$')
# reheating
plt.gca().set_prop_cycle(None)
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], d['energy'], np.sum(d['eigoccs']))
for d in reheating_data
if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
L_l, energy_l, norms_l = zip(*sorted(data_iterator))
energy_l /= np.array(norms_l)
plt.plot(L_l, energy_l, 'x:', label=f'reheating $J/B={JvB}$')
# # iterative
# plt.gca().set_prop_cycle(None)
# for JvB in [.2, 1, 5]:
# data_iterator = ((d['L'], d['energy'], np.sum(d['eigoccs']))
# for d in iterative_data
# if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
# L_l, energy_l, norms_l = zip(*sorted(data_iterator))
# energy_l /= np.array(norms_l)
# plt.plot(L_l, energy_l, '+--', label=f'reheating $J/B={JvB}$')
plt.legend(bbox_to_anchor=(1, .5), loc='center left')
plt.ylim(top=-.95)
plt.xlabel('L')
plt.ylabel('energy')
# -
# ### GS infidelity vs L
# + code_folding=[]
K=10
plt.title(f'TFIM chain. Standard LogSweep(K={K}) density matrix sim')
# cooling
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in cooling_data
if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(L_l, infidelity_l, 'o-', label=f'cooling $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# reheating
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(L_l, infidelity_l, 'x:', label=f'reheating $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# for JvB in [.2, 1, 5]:
# data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
# for d in iterative_data
# if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
# L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
# fidelty_l /= np.array(norms_l)
# infidelity_l = 1 - np.array(fidelty_l)
# plt.plot(L_l, infidelity_l, '+--', label=f'iterative $J/B={JvB}$')
plt.legend()
plt.yscale('log')
plt.xscale('log')
plt.xlabel('L')
plt.ylabel('ground space infidelity')
# -
# ## test: changing K with L
# + code_folding=[]
plt.title(f'TFIM chain. Standard LogSweep(K=L) density matrix sim')
# cooling
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in cooling_data
if d['K'] == d['L'] and np.isclose(d['J']/d['B'], JvB))
L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(L_l, infidelity_l, 'o-', label=f'cooling $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# reheating
for JvB in [.2, 1, 5]:
data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
for d in reheating_data
if d['K'] == d['L'] and np.isclose(d['J']/d['B'], JvB))
L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
fidelty_l /= np.array(norms_l)
infidelity_l = 1 - np.array(fidelty_l)
plt.plot(L_l, infidelity_l, 'x:', label=f'reheating $J/B={JvB}$')
plt.gca().set_prop_cycle(None)
# for JvB in [.2, 1, 5]:
# data_iterator = ((d['L'], gsm_fidelity(d), np.sum(d['eigoccs']))
# for d in iterative_data
# if d['K'] == K and np.isclose(d['J']/d['B'], JvB))
# L_l, fidelty_l, norms_l = zip(*sorted(data_iterator))
# fidelty_l /= np.array(norms_l)
# infidelity_l = 1 - np.array(fidelty_l)
# plt.plot(L_l, infidelity_l, '+--', label=f'iterative $J/B={JvB}$')
plt.legend()
plt.yscale('log')
plt.xscale('log')
plt.xlabel('L')
plt.ylabel('ground space infidelity')
# -
# # Eigenstate occupation plots
# +
L = 7
JvBlist = [0.2, 1, 5]
Klist = [2, 39]
from qdclib import TFIMChain
fig, sbpl = plt.subplots(len(Klist), len(JvBlist),
sharex = True, sharey = True,
gridspec_kw={'hspace': 0, 'wspace': 0})
for i, JvB in enumerate(JvBlist):
system = TFIMChain(L, JvB, 1)
system.normalize()
for j, K in enumerate(Klist):
for d in cooling_data:
if d['L'] == L and np.isclose(JvB, d['J']/d['B']) and d['K'] == K:
break
sbpl[j, i].plot(system.eigvals, d['eigoccs'], '.', label='cooling')
for d in reheating_data:
if d['L'] == L and np.isclose(JvB, d['J']/d['B']) and d['K'] == K:
break
sbpl[j, i].plot(system.eigvals, d['eigoccs'], '_', label='reheating')
sbpl[j, i].set_yscale('log')
sbpl[j, i].text(0.2, 0.005, f'$J/B = {JvB}$\n$K = {K}$')
plt.tight_layout()
sbpl[0,2].legend()
|
data-analysis/TFIM-chain-logsweep-DM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy.optimize import minimize
from numpy.random import rand
# objective function
def objective(x):
return x[0]**2.0 + x[1]**2.0
# derivative of the objective function
def derivative(x):
return [x[0] * 2, x[1] * 2]
# define range for input
r_min, r_max = -5.0, 5.0
# define the starting point as a random sample from the domain
pt = r_min + rand(2) * (r_max - r_min)
# perform the bfgs algorithm search
result = minimize(objective, pt, method='BFGS', jac=derivative)
# summarize the result
print('Status : %s' % result['message'])
print('Total Evaluations: %d' % result['nfev'])
# evaluate solution
solution = result['x']
evaluation = objective(solution)
print('Solution: f(%s) = %.5f' % (solution, evaluation))
|
optimization_algorithms/01 A Gentle Introduction to the BFGS Optimization Algorithm/A Gentle Introduction to the BFGS Optimization Algorithm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> HSV Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt> <dd><a href='./HSV.ipynb'>Matplotlib</a></dd> <dd><a href='../bokeh/HSV.ipynb'>Bokeh</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# HoloViews makes it trivial to work in any color space that can be converted to ``RGB`` by making a simple subclass of ``RGB`` as appropriate. For instance, we also provide the HSV (hue, saturation, value) color space, which is useful for plotting cyclic data (as the Hue) along with two additional dimensions (controlling the saturation and value of the color, respectively).
#
# Like other raster based Element types ``HSV`` accepts gridded data, which may be supplied as a simple array ``NxMx3`` ndarray representing hue, saturation and value channels along with bounds or explicit array coordinates. See the [Gridded Datasets](../../../user_guide/08-Gridded_Datasets.ipynb) user guide to see the accepted data formats.
# +
x,y = np.mgrid[-50:51, -50:51] * 0.1
h = 0.5 + np.sin(0.2*(x**2+y**2)) / 2.0
s = 0.5*np.cos(y*3)+0.5
v = 0.5*np.cos(x*3)+0.5
hsv = hv.HSV(np.dstack([h, s, v]))
hsv
# -
# You can see how this is created from the original channels:
# %opts Image (cmap='gray')
hsv[..., 'H'].relabel('H') + hsv[..., 'S'].relabel('S') + hsv[..., 'V'].relabel('V')
# An ``HSV`` Element can also easily be converted to an ``RGB`` Element using the ``rgb`` property:
print(hsv.rgb)
hsv.rgb[..., 'R'] + hsv.rgb[..., 'G'] + hsv.rgb[..., 'B']
# For full documentation and the available style and plot options, use ``hv.help(hv.HSV).``
|
examples/reference/elements/matplotlib/HSV.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: detectron2
# language: python
# name: detectron2
# ---
# %config IPCompleter.use_jedi = False
import wholeslidedata
from wholeslidedata.iterators import create_batch_iterator
import os
import detectron2
import torch
from detectron2.structures import (
BoxMode,
Instances,
Boxes
)
from detectron2.engine import DefaultTrainer
from detectron2.config import get_cfg
from detectron2 import model_zoo
from wholeslidedata.iterators import BatchIterator
import numpy as np
class Detectron22DataLoader(BatchIterator):
def __next__(self):
x_batch, y_batch = super().__next__()
x_batch = x_batch / 255.0
batch_dicts = []
for idx, x_sample in enumerate(x_batch):
sample_dict = {}
# print(y_batch[idx])
target_gt_boxes = self._get_gt_boxes(y_batch[idx], x_sample.shape[:2])
image = image.transpose(2, 0, 1).astype("float32")
sample_dict['instances'] = target_gt_boxes
sample_dict['image'] = torch.as_tensor(image)
batch_dicts.append(sample_dict)
return batch_dicts
def _get_gt_boxes(self, y_sample, image_size):
y_boxes = y_sample[~np.all(y_sample == 0, axis=-1)]
boxes = [BoxMode.convert(obj[:4], BoxMode.XYXY_ABS, BoxMode.XYXY_ABS) for obj in y_boxes]
target = Instances(image_size)
target.gt_boxes = Boxes(boxes)
classes = [int(obj[-2]) for obj in y_boxes]
classes = torch.tensor(classes, dtype=torch.int64)
target.gt_classes = classes
return target
class WholeSlideDataDetectionTrainer(DefaultTrainer):
@classmethod
def build_train_loader(cls, cfg):
user_config = './configs/detection_config.yml'
cpus = 1
mode = 'training'
training_batch_generator = create_batch_iterator(user_config=user_config,
mode=mode,
cpus=cpus,
iterator_class=Detectron22DataLoader)
return training_batch_generator
# +
user_config = './configs/detection_config.yml'
cpus = 1
mode = 'training'
training_batch_generator = create_batch_iterator(user_config=user_config,
mode=mode,
cpus=cpus,
iterator_class=Detectron22DataLoader)
# +
def train():
# coco_datadict = get_pannuke_coco_datadict(data_folder, fold)
# register(fold, coco_datadict)
cfg = get_cfg()
cfg.merge_from_file(
model_zoo.get_config_file("COCO-Detection/faster_rcnn_X_101_32x8d_FPN_3x.yaml")
)
cfg.DATASETS.TRAIN = ("detection_dataset2",)
cfg.DATASETS.TEST = ()
cfg.DATALOADER.NUM_WORKERS = 1
# cfg.MODEL.WEIGHTS = "detectron2://COCO-Detection/faster_rcnn_X_101_32x8d_FPN_3x/139173657/model_final_68b088.pkl" # Let training initialize from model zoo
# cfg.MODEL.WEIGHTS = None
cfg.SOLVER.IMS_PER_BATCH = 8
cfg.SOLVER.BASE_LR = 0.00001 # pick a good LR
cfg.SOLVER.MAX_ITER = 200000 # 300 iterations seems good enough for this toy dataset; you may need to train longer for a practical dataset
cfg.MODEL.ROI_HEADS.BATCH_SIZE_PER_IMAGE = (
64 # faster, and good enough for this toy dataset (default: 512)
)
cfg.MODEL.ROI_HEADS.NUM_CLASSES = 1
cfg.OUTPUT_DIR = '/home/user/output/'
cfg.SOLVER.STEPS = (1000, 10000, 20000, 50000, 100000)
cfg.SOLVER.WARMUP_ITERS = 100
cfg.SOLVER.GAMMA = 0.5
os.makedirs(cfg.OUTPUT_DIR, exist_ok=True)
trainer = WholeSlideDataDetectionTrainer(cfg)
trainer.resume_or_load(resume=False)
trainer.train()
# -
train()
train()
|
notebooks/detectron2_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [Index](Index.ipynb) - [Back](Widget Events.ipynb) - [Next](Custom Widget - Hello World.ipynb)
# + language="html"
# <style>
# .example-container { background: #999999; padding: 2px; min-height: 100px; }
# .example-container.sm { min-height: 50px; }
# .example-box { background: #9999FF; width: 50px; height: 50px; text-align: center; vertical-align: middle; color: white; font-weight: bold; margin: 2px;}
# .example-box.med { width: 65px; height: 65px; }
# .example-box.lrg { width: 80px; height: 80px; }
# </style>
# -
import ipywidgets as widgets
from IPython.display import display
# + [markdown] slideshow={"slide_type": "slide"}
# # Widget Styling
# -
# ## Basic styling
# The widgets distributed with IPython can be styled by setting the following traits:
#
# - width
# - height
# - fore_color
# - back_color
# - border_color
# - border_width
# - border_style
# - font_style
# - font_weight
# - font_size
# - font_family
#
# The example below shows how a `Button` widget can be styled:
button = widgets.Button(
description='Hello World!',
width=100, # Integers are interpreted as pixel measurements.
height='2em', # em is valid HTML unit of measurement.
color='lime', # Colors can be set by name,
background_color='#0022FF', # and also by color code.
border_color='red')
display(button)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Parent/child relationships
# -
# To display widget A inside widget B, widget A must be a child of widget B. Widgets that can contain other widgets have a **`children` attribute**. This attribute can be **set via a keyword argument** in the widget's constructor **or after construction**. Calling display on an **object with children automatically displays those children**, too.
# +
from IPython.display import display
float_range = widgets.FloatSlider()
string = widgets.Text(value='hi')
container = widgets.Box(children=[float_range, string])
container.border_color = 'red'
container.border_style = 'dotted'
container.border_width = 3
display(container) # Displays the `container` and all of it's children.
# -
# ### After the parent is displayed
# + [markdown] slideshow={"slide_type": "slide"}
# Children **can be added to parents** after the parent has been displayed. The **parent is responsible for rendering its children**.
# +
container = widgets.Box()
container.border_color = 'red'
container.border_style = 'dotted'
container.border_width = 3
display(container)
int_range = widgets.IntSlider()
container.children=[int_range]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Fancy boxes
# -
# If you need to display a more complicated set of widgets, there are **specialized containers** that you can use. To display **multiple sets of widgets**, you can use an **`Accordion` or a `Tab` in combination with one `Box` per set of widgets** (as seen below). The "pages" of these widgets are their children. To set the titles of the pages, one can **call `set_title`**.
# ### Accordion
# +
name1 = widgets.Text(description='Location:')
zip1 = widgets.BoundedIntText(description='Zip:', min=0, max=99999)
page1 = widgets.Box(children=[name1, zip1])
name2 = widgets.Text(description='Location:')
zip2 = widgets.BoundedIntText(description='Zip:', min=0, max=99999)
page2 = widgets.Box(children=[name2, zip2])
accord = widgets.Accordion(children=[page1, page2])
display(accord)
accord.set_title(0, 'From')
accord.set_title(1, 'To')
# + [markdown] slideshow={"slide_type": "slide"}
# ### TabWidget
# +
name = widgets.Text(description='Name:')
color = widgets.Dropdown(description='Color:', options=['red', 'orange', 'yellow', 'green', 'blue', 'indigo', 'violet'])
page1 = widgets.Box(children=[name, color])
age = widgets.IntSlider(description='Age:', min=0, max=120, value=50)
gender = widgets.RadioButtons(description='Gender:', options=['male', 'female'])
page2 = widgets.Box(children=[age, gender])
tabs = widgets.Tab(children=[page1, page2])
display(tabs)
tabs.set_title(0, 'Name')
tabs.set_title(1, 'Details')
# + [markdown] slideshow={"slide_type": "slide"}
# # Alignment
# -
# Most widgets have a **`description` attribute**, which allows a label for the widget to be defined.
# The label of the widget **has a fixed minimum width**.
# The text of the label is **always right aligned and the widget is left aligned**:
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
# + [markdown] slideshow={"slide_type": "slide"}
# If a **label is longer** than the minimum width, the **widget is shifted to the right**:
# -
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
display(widgets.Text(description="aaaaaaaaaaaaaaaaaa:"))
# + [markdown] slideshow={"slide_type": "slide"}
# If a `description` is **not set** for the widget, the **label is not displayed**:
# -
display(widgets.Text(description="a:"))
display(widgets.Text(description="aa:"))
display(widgets.Text(description="aaa:"))
display(widgets.Text())
# + [markdown] slideshow={"slide_type": "slide"}
# ## Flex boxes
# -
# Widgets can be aligned using the `FlexBox`, `HBox`, and `VBox` widgets.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Application to widgets
# -
# Widgets display vertically by default:
buttons = [widgets.Button(description=str(i)) for i in range(3)]
display(*buttons)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Using hbox
# -
# To make widgets display horizontally, you need to **child them to a `HBox` widget**.
container = widgets.HBox(children=buttons)
display(container)
# By setting the width of the container to 100% and its `pack` to `center`, you can center the buttons.
container.width = '100%'
container.pack = 'center'
# + [markdown] slideshow={"slide_type": "slide"}
# ## Visibility
# -
# Sometimes it is necessary to **hide or show widgets** in place, **without having to re-display** the widget.
# The `visible` property of widgets can be used to hide or show **widgets that have already been displayed** (as seen below). The `visible` property can be:
# * `True` - the widget is displayed
# * `False` - the widget is hidden, and the empty space where the widget would be is collapsed
# * `None` - the widget is hidden, and the empty space where the widget would be is shown
w1 = widgets.Latex(value="First line")
w2 = widgets.Latex(value="Second line")
w3 = widgets.Latex(value="Third line")
display(w1, w2, w3)
w2.visible=None
w2.visible=False
w2.visible=True
# + [markdown] slideshow={"slide_type": "slide"}
# ### Another example
# -
# In the example below, a form is rendered, which conditionally displays widgets depending on the state of other widgets. Try toggling the student checkbox.
# +
form = widgets.VBox()
first = widgets.Text(description="First Name:")
last = widgets.Text(description="Last Name:")
student = widgets.Checkbox(description="Student:", value=False)
school_info = widgets.VBox(visible=False, children=[
widgets.Text(description="School:"),
widgets.IntText(description="Grade:", min=0, max=12)
])
pet = widgets.Text(description="Pet's Name:")
form.children = [first, last, student, school_info, pet]
display(form)
def on_student_toggle(name, value):
if value:
school_info.visible = True
else:
school_info.visible = False
student.on_trait_change(on_student_toggle, 'value')
# -
# [Index](Index.ipynb) - [Back](Widget Events.ipynb) - [Next](Custom Widget - Hello World.ipynb)
|
examples/Interactive Widgets/Widget Styling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Invert a binary tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * What does it mean to invert a binary tree?
# * Swap all left and right node pairs
# * Can we assume we already have a Node class?
# * Yes
# * Can we assume the inputs are valid?
# * No
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# <pre>
# Input:
# 5
# / \
# 2 7
# / \ / \
# 1 3 6 9
#
# Output:
# 5
# / \
# 7 2
# / \ / \
# 9 6 3 1
# </pre>
# ## Algorithm
#
# Refer to the [Solution Notebook](). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
# %run ../bst/bst.py
class InverseBst(Bst):
def invert_tree(self):
if self.root is None:
return
self._invertTree(self.root)
return self.root
def _invertTree(self, node):
if node is None:
return
self._invertTree(node.left)
self._invertTree(node.right)
node.left, node.right = node.right, node.left
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_invert_tree.py
from nose.tools import assert_equal
class TestInvertTree(object):
def test_invert_tree(self):
root = Node(5)
bst = InverseBst(root)
node2 = bst.insert(2)
node3 = bst.insert(3)
node1 = bst.insert(1)
node7 = bst.insert(7)
node6 = bst.insert(6)
node9 = bst.insert(9)
result = bst.invert_tree()
assert_equal(result, root)
assert_equal(result.left, node7)
assert_equal(result.right, node2)
assert_equal(result.left.left, node9)
assert_equal(result.left.right, node6)
assert_equal(result.right.left, node3)
assert_equal(result.right.right, node1)
print('Success: test_invert_tree')
def main():
test = TestInvertTree()
test.test_invert_tree()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook]() for a discussion on algorithms and code solutions.
|
graphs_trees/invert_tree/invert_tree_challenge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Лекция 17 "Хеш-таблицы"
#
# ### Финансовый университет при Правительстве РФ, лектор <NAME>
#
# v 0.17
# ### Абстрактный тип данных - ассоциативный массив
# **Словарь (ассоциативного массива (map, dictionary, associative array))** - абстрактная структура данных позволяющая хранить пары вида "ключ - значение" и поддерживающая операции добавления пары, а также поиска и удаления пары по ключу. Предполагается, что ассоциативный массив не может хранить две пары с одинаковыми ключами. Ассоциативный массив с точки зрения интерфейса удобно рассматривать как обычный массив, в котором в качестве индексов можно использовать не только целые числа из определенного диапазона, но и значения других типов — например, строки. В Python словарь реализуется при помощи dict().
# +
# Базовые операции словаря:
d1 = dict() # создание пустого словаря
d1['abc'] = 42 # put(key, val) добавление пары ключ-значение в словарь;
# если по данному ключу уже есть значение - то оно заменяется новым.
v = d1['abc'] # get(key) извлечение значения по ключу;
# в случае отсутсутствия значения по ключу возникает исключительная ситуация KeyError
print(v)
b = 'xyz' in d1 # возвращает True если ключ содержится в словаре, иначе возвращает False
print(b)
it = iter(d1) # итератор по умолчанию генерирует последовательность всех ключей хранящихся в словаре
print(list(it))
l = len(d1) # возвращает количество пар ключ-значение в словаре
print(l)
del d1['abc'] # удаляет из словаря пару ключ-значение с заданным ключом
# -
# Далее мы рассмотрим различные решения задачи реализации словаря.
# ### Таблица с прямой адресацией
# Прямая адресация представляет собой простейшую технологию, которая хорошо работает для небольших множеств ключей. Предположим, что приложению требуется динамическое множество, каждый элемент которого имеет ключ из множества $U = \left \{ 0,1,\ldots, m - 1 \right \} $, где $m$ не слишком велико. Кроме того, предполагается, что никакие два элемента
# не имеют одинаковых ключей.
# Для представления динамического множества мы используем массив, или таблицу с прямой адресацией, каждая ячейка которого соответствует ключу из пространства ключей $U$.
# Возможность прямой индексации элементов обычного массива обеспечивает доступ к произвольной позиции в массиве за время $O(1)$. Прямая индексация применима, если есть возможность выделить массив размера, достаточного для того, чтобы у каждого возможного значения ключа имелась своя ячейка.
# +
# пример: хранение строки строчных латинских символов не более заданной длины
# -
def to_key(s=None):
'''Возвращает неотрицательный ключ по объекту. Если объект не задан, то возвращает максимальное значение ключа.
'''
if s is None:
s = 'z'
return ord(s) - ord('a')
to_key('a')
to_key('c')
to_key()
def seq_to_key(seq):
res = 0
n = to_key() # количество допустимых символов
for i, el in enumerate(seq):
assert 0 <= to_key(el) <= n # проверка корректности рассматриваемого символа
res += to_key(el) * n ** i
return res
seq_to_key('a')
seq_to_key('z')
seq_to_key('bb')
seq_to_key('abc')
seq_to_key('aa') # проблема!
def to_key(s=None):
'''Возвращает ПОЛОЖИТЕЛЬНЫЙ ключ по объекту. Если объект не задан, то возвращает максимальное значение ключа.
'''
if s is None:
s = 'z'
return ord(s) - ord('a') + 1
[seq_to_key(x) for x in ['','a', 'z', 'aa', 'ab', 'zz', 'aaa']]
class DirectStrTable:
def __init__(self, str_len):
self._str_len = str_len # максимальная длина строки в таблице
self._n = self._to_key() # количество допустимых символов
self._none = object() # создаем пустой объект в качестве внутреннего аналога None
total = 0
for i in range(self._str_len + 1):
total *= self._n
total += 1
self._table = [self._none] * total
self._len = 0 # текущее количество элементов, хранящихся в таблице
# internal method
def _to_key(self, s=None):
'''Возвращает ПОЛОЖИТЕЛЬНЫЙ ключ по символу. Если объект не задан, то возвращает максимальное значение ключа.
'''
if s is None:
s = 'z'
return ord(s) - ord('a') + 1
def _from_key(self, k):
return chr(k + ord('a') - 1)
# internal method
def _str_to_key(self, seq):
res = 0
assert len(seq) <= self._str_len
for i, el in enumerate(seq):
assert 0 <= to_key(el) <= self._n # проверка корректности рассматриваемого символа
res += to_key(el) * self._n ** i
return res
def _key_to_str(self, k):
s = ''
while k > 0:
k, m = divmod(k, self._n)
if m == 0:
s += self._from_key(self._n)
k -= 1
else:
s += self._from_key(m)
# s += self._from_key(self._n) if m == 0 else self._from_key(m)
return s
# len(dst)
def __len__(self):
return self._len
# dst[str_key]
def __getitem__(self, str_key):
k = self._str_to_key(str_key)
if self._table[k] is self._none:
raise KeyError(f'Key Error: {repr(s)}') # по данному ключу значения в таблице нет
else:
return self._table[k]
# str_key in dst
def __contains__(self, str_key):
k = self._str_to_key(str_key)
if self._table[k] is self._none:
return False
else:
return True
# dst[str_key] = val
def __setitem__(self, str_key, val):
k = self._str_to_key(str_key)
if self._table[k] is self._none:
# в таблицу добавляется новый элемент
self._table[k] = val
self._len += 1
else:
# в таблице меняется значение существующего элемента
self._table[k] = val
# del dst[str_key]
def __delitem__(self, str_key):
k = self._str_to_key(str_key)
if self._table[k] is self._none:
raise KeyError(f'Key Error: {repr(s)}') # по данному ключу значения в таблице нет
else:
self._table[k] = self._none
self._len -= 1
# for x in dst
def _raw_iter(self):
for rk, val in enumerate(self._table):
if val is not self._none:
yield rk, val # self._key_to_str(rk)
def __iter__(self):
for rk, val in self._raw_iter():
yield self._key_to_str(rk) #, val #
def items(self):
for rk, val in self._raw_iter():
yield self._key_to_str(rk), val
def values(self):
for val in self._table:
if val is not self._none:
yield val
def stat(self):
return f'''Хранится элементов: {self._len};
размер таблицы: {len(self._table)};
доля используемых элементов таблицы: {self._len/len(self._table)}'''
dst1 = DirectStrTable(3)
len(dst1)
dst1['ab'] = 11
len(dst1)
dst1['ab']
'ab' in dst1, 'a' in dst1
dst1['abc'] = 7
len(dst1)
dst1['zz'] = 77
len(dst1)
del dst1['zz']
len(dst1)
dst1['zzz'] = 1
dst1['x'] = 5
dst1['z'] = 6
dst1['aa'] = 7
dst1['zz'] = 8
dst1['aaa'] = 9
len(dst1)
list(dst1)
list(dst1._raw_iter())
list(dst1.items())
list(dst1.values())
d1 = dict([('ab', 11), ('abc', 7)])
list(d1)
list(d1.values())
print(dst1.stat())
# Каждая из приведенных операций очень быстрая: время их работы равно $O(1)$. В некоторых приложениях элементы динамического множества могут храниться непосредственно в таблице с прямой адресацией.
#
# Недостаток прямой адресации очевиден: если пространство ключей $U$ велико, хранение таблицы $Т$ размером $|U|$ непрактично, а то и вовсе невозможно – в зависимости от количества доступной памяти и размера пространства ключей. Кроме того, множество $К$ реально сохраненных ключей может быть мало по сравнению с пространством ключей $U$, а в
# этом случае память, выделенная для таблицы $Т$, в основном расходуется напрасно.
# ### Хеш-таблица
# Хеш-таблица представляет собой эффективную структуру данных для **реализации словарей**. Хотя на поиск элемента в хеш-таблице может в наихудшем случае потребоваться столько же времени, что и в связанном списке, а именно $O(n)$, на практике хеширование исключительно эффективно. При вполне обоснованных допущениях **математическое ожидание времени поиска** элемента в хеш-таблице составляет
# $O(1)$.
# Сравнение разынх методов реализации словарей: https://en.wikipedia.org/wiki/Associative_array
# Хеш-таблица (hash table) представляет собой обобщение обычного массива. Если количество реально хранящихся в массиве ключей мало по сравнению с количеством возможных значений ключей, эффективной альтернативой массива с прямой индексацией становится **хеш-таблица**, которая обычно использует массив с размером, пропорциональным количеству реально хранящихся в нем ключей. Вместо непосредственного использования ключа в качестве индекса массива, **индекс вычисляется по значению ключа**. Идея хеширования состоит в использовании некоторой частичной информации, полученной из ключа, т.е. вычисляется хеш-адрес $h(key)$, который используется для индексации в хеш-таблице.
# Когда множество $К$ хранящихся в словаре ключей гораздо меньше пространства возможных ключей $U$, хеш-таблица требует существенно меньше места, чем таблица с прямой адресацией. Точнее говоря, требования к памяти могут быть снижены до $\Theta(|K|)$, при этом время поиска элемента в хеш-таблице остается равным $O(1)$. Надо только заметить, что это граница среднего времени поиска, в то время как в случае таблицы с прямой адресацией эта граница справедлива для наихудшего случая.
# **Хеш-таблица** это коллекция, хранящая проиндексированные элементы (значения). Каждая позиция в хеш-таблице - слот таблицы (slot, bucket) может содержать элемент адресованный целочисленным не отрицательным индексом внутри таблицы. Т.е. в таблице есть слот с индексом 0, 1 и т.д. При создании все слоты в хеш-таблице пустые. В Python хеш-таблицу мужно реализовать в виде списка заполненного значениями None или их заменителями (для случаев, если пользователю нужно хранить в хеш-таблице значения None).
# В случае прямой адресации элемент с ключом $k$ хранится в ячейке $k$. При хешировании этот элемент хранится в ячейке $h(k)$, т.е. мы используем хеш-функцию $h$ для вычисления ячейки для данного ключа $k$. Функция $h$ отображает пространство ключей $U$ на ячейки хеш-таблицы $Т [0..m - 1]$:
# $$h:U \rightarrow \{ 0,1, \ldots , m-1 \} .$$
# Мы говорим, что элемент с ключом $k$ хешируется в ячейку $h(k)$, величина $h(k)$ называется хеш-значением ключа $k$.
# *Пример*
#
# $T[0, \ldots, 10]$; $h(k) = k \mod 11$
# Пустая хеш-таблица:
# 
def h(k):
return k % 11
keys1 = [54, 26, 93, 17, 77, 31]
[h(k) for k in keys1]
# Хеш-таблица с 6ю элементами:
# 
# При построении хеш-таблиц есть одна проблема: два ключа могут быть хешированы одну и ту же ячейку. Такая ситуация называется **коллизией**.
#
# пример коллизии:
h(12), h(23)
# Т.к. $h$ является детерминистической и для одного и того же значения k всегда дает одно и то же хеш-значение $h(k)$, то поскольку $|U| > m$, должно существовать как минимум два ключа, которые имеют одинаковое хеш-значение. Таким образом,
# полностью **избежать коллизий невозможно** в принципе.
#
# Используются два подхода для борьбы с этой проблемой:
#
# * выбор хеш-функции снижающей вероятность коллизии;
# * использование эффективных алгоритмов разрешения коллизий.
# ### Хеш-функция
# Хеш-функция выполняет преобразование массива входных данных произвольной длины (ключа, сообщения) в (выходную) битовую строку установленной длины (хеш, хеш-код, хеш-сумму).
# Хеш-функции применяются в следующих задачах:
#
# * построение ассоциативных массивов;
# * поиске дубликатов в сериях наборов данных;
# * построение уникальных идентификаторов для наборов данных;
# * вычислении контрольных сумм от данных (сигнала) для последующего обнаружения в них ошибок (возникших случайно или внесённых намеренно), возникающих при хранении и/или передаче данных;
# * сохранении паролей в системах защиты в виде хеш-кода (для восстановления пароля по хеш-коду требуется функция, являющаяся обратной по отношению к использованной хеш-функции);
# * выработке электронной подписи (на практике часто подписывается не само сообщение, а его «хеш-образ»);
# и многих других.
#
# Для решения различных задач требования к хеш-функциям могут очень существенно отличаться.
# "Хорошая" хеш-функция должна удовлетворять двум свойствам:
#
# * быстрое вычисление;
# * минимальное количество коллизий.
# Для обеспечения минимального количества коллизий хеш-функция удовлетворяет (приближенно) предположению простого **равномерного хеширования**: для каждого ключа равновероятно помещение в любую из $m$ ячеек, независимо от хеширования остальных ключей. К сожалению, это условие обычно невозможно проверить, поскольку, как правило, распределение вероятностей, в соответствии с которым поступают вносимые в таблицу ключи, неизвестно; кроме того, вставляемые ключи могут не быть независимыми.
# При построении хеш-функции хорошим подходом является подбор функции таким образом, чтобы она никак **не коррелировала с закономерностями**, которым могут подчиняться существующие данные. Например, мы можем потребовать, чтобы **"близкие" в некотором смысле ключи давали далекие хеш-значения** (например, хеш функция для подряд идущих целых чисел давала далекие хеш-значения). В некоторых приложениях хеш-функций требуется противоположное свойство - непрерывность (близкие ключи длолжны порождать близкие хеш-значения).
# Обычно от хеш-функций ожидается, что значения хеш-функции находиятся в диапазоне от 0 до $m-1$. Причём, часто удобно, есчли $m=2^n$. Таким образом значение хеш-функции может, например, без преобразований хранится в машинном слове.
# #### Метод деления
# Построение хеш-функции методом деления состоит в отображении ключа k в одну из ячеек путем получения остатка от деления $k$ на $m$, т.е. хеш-функция имеет вид $h(k) = k \mod m$.
#
# При использовании данного метода мы обычно стараются избегать некоторых значений $m$. Например, $m$ не должно быть степенью 2, поскольку если $m = 2^n$, то $h(k)$ представляет собой просто $р$ младших битов числа $k$. Если только заранее неизвестно, что все наборы младших $р$ битов ключей равновероятны, лучше строить хеш-функцию таким образом, чтобы ее результат зависел от всех битов ключа. Зачастую хорошие результаты можно получить, выбирая в качестве значения $m$ простое число, достаточно далекое от степени двойки.
# +
# пример хеш-функции для строк, построенной по методу деления:
def str_h(s, m=701):
return sum(ord(symb) for symb in s) % m
# -
s1 = 'Хеш-функция выполняет преобразование массива входных данных произвольной длины (ключа, сообщения) в (выходную) битовую строку установленной длины (хеш, хеш-код, хеш-сумму).'
s1_cod = [(s, str_h(s)) for s in s1.split()]
s1_cod
sorted(s[1] for s in s1_cod)
# #### Метод MAD
# Хеш-функция multiply-add-and-divide (часто именуемая как MAD) преобразует целое число $k$ по следующему алгоритму.
# У хеш-функци имеются следующие параметры: $p$ - большое простое число, $a \in \{1, 2, \ldots, p-1 \}$ и $b \in \{0, 1, \ldots, p-1 \}$, $m$ - количество значений в диапозоне значений хеш-функции.
#
# $$h_{a,b}(k) = ((ak + b)\mod p)\mod m $$
#
#
# Этот класс хеш-функций удобен тем, что размер $m$ выходного диапазона произволен и не обязательно представляет собой простое число. Поскольку число, $а$ можно выбрать $р-1$ способом, и $р$ способами – число $b$, всего в данном семействе будет содержаться $р(р - 1)$ хеш-функций.
# Данную хеш-функцию (семейство хеш-функций) можно использовать для **универсального хеширования**. Универсальным хешированием называется хеширование, при котором используется не одна конкретная хеш-функция, а происходит выбор хеш-функции из заданного семейства по случайному алгоритму. Семейство универсальных хеш-функций называется универсальным, если для любых двух допустимых ключей вероятность коллизии является наименьшей из возможных:
#
# $$\underset{h \in \mathcal{H}}{Pr} \left[ h(x)=h(y) \right] = \frac{1}{m}, x \neq y$$
#
# Нужно отметить, что это утверждение верно дложно выполняться для любых различающихся ключей, при этом случайно выбираются здесь именно функции хеширования $h$ из всего класса универсальных хеш-функций $\mathcal{H}$.
#
# Универсальное хеширование обычно отличается низким числом коллизий и применяется, например, при реализации хеш-таблиц и в криптографии. В частности, можно показать, что хеш-функции MAD для случано выбранных $a$ и $b$ являются классом универсальных хеш-функций (см. https://en.wikipedia.org/wiki/Universal_hashing#Hashing_strings).
import random
2**32 # количество целых чисел в 32 битовом слове
mad_p = 4294967311 # https://www.numberempire.com/primenumbers.php
class MadHash:
def __init__(self, m):
self.m = m
self.p = mad_p
assert self.p > self.m
self.a = random.randint(1, self.p)
self.b = random.randint(0, self.p)
def h(self, k):
return ((self.a * k + self.b) % self.p ) % self.m
mh = MadHash(2**8)
mh.a, mh.b
hr1 = [mh.h(v) for v in range(128)]
hr1
for i, h in enumerate(hr1):
print(i, h)
hr1s = sorted(hr1)
list(zip(hr1s, [x0 == x1 for x0, x1 in zip(hr1s[:-1], hr1s[1:])]))
# Зачастую расчет хеш-функции $h(k)$ можно представить в виде двух операций: рассчета хеш-кода который превращает ключ $k$ в целое число и функции компрессии, которая преобразует хеш-код в целое число в заданном диапазоне $[0, m-1]$.
# Схема построения хеш-функции на базе двух шагов:
# 
# Преимуществом разделения хеш-функции на две компоненты является то, что рассчет хеш-кода проивзодится независимо от того с каким размером хеш-таблицы нужно будет работать. Это **позволяет разрабатывать функции рассчета хеш-кодов для различных типов данных** не ориентируясь на размер хеш-таблицы, который важен только для функции компрессии. Это особенно удобно, т.к. **размер хеш таблицы может быть динамечески изменен в зависимости от количества элементов, хранимых в словаре**.
# Функция MAD может использоваться и в качестве функции построения хеш-кода для целых чисел и в качестве функции компрессии для хеш-кодов, построенных с помощью других функций.
# #### Полиномиальная хеш-функция
# Приведенная ранее функция построения хеш-кода, основанная на суммировании (или опреации xor) хеш-кодов, плохо подходит для работы с символьными строками и другими объектами различной длины, которые могут быть представлены в виде кортежа $(x_0 , x_1 , \ldots , x_{n−1} )$, в котором позиция элемента $x_i$ имеет значение, т.к. такой хеш-код создает коллизии для строк (последовательностей) с одинаковым составом элементов.
# примеры коллизий для хеш-функций строк основанных на суммировании
s2 = ["stop", "tops", "pots", "spot"]
[(s, str_h(s)) for s in s2]
# Такого рода коллизии не будут возникать в хеш-функции, которая учитыват положение элементов в массиве входных данных. Примером такой хеш-функции является функция, использующая константу $a$ ($a \neq 0, a \neq 1$) при построении хеш-функции вида:
#
# $$x_0 a^{n-1} + x_1 a^{n-2} + \ldots + x_{n−2} a + x_{n−1}.$$
#
# Т.е. это полином, использующий элементы массива входных данных $(x_0 , x_1 , \ldots , x_{n−1} )$ в качестве коэффициентов. Такая функция назвается полиниомиальным хеш-кодом. Для использования ее в качестве хеш-функции к ней необходимо только добавить функцию компресии в соответствующий диапозон значений.
# Используя схему Горнера полиномиальный хеш-код можно эффективно вычислить по формуле:
#
# $$x_{n−1} + a (x_{n−2} + a(x_{n−3}+ \ldots + a(x_2 + a(x_1 + ax_0))\ldots))$$
# ### Функция hash в Python
# Стандартным способом для получения хеш-кода в Python является встроенная функция **hash(x)**. Она возвращает целочисленное значение для объекта x. Однако, в Python **только неизменяемые типы** данных могут возвращать значение хеш-кода. Это ограчиние гарантирует, что хеш-код для объека не изменится во время его жизни. Это свойство очень важно для корректной работы при использовании хеш-кодов объектов в хеш-таблицах, например в dict().
hash('Hello world!')
hash(42)
hash(3.141)
hash((1, 2))
hash(None)
hash(frozenset([1, 2]))
# ошибка:
hash([1, 2])
# Важным правилом реализации функции hash для классов явлется необходимость сохранять **консистентность между равенством (x == y) и равенством хеш-функций (hash(x) == hash(y))**. Для любых двух объектов из равенства x == y должно следовать hash(x) == hash(y) (из-за возможности коллизий у хеш-функций следствие в обратную сторону в общем случае не выполняется). Это необходимо для того, чтобы в случае использования объекта в качестве ключа в хеш таблицы для равных объектов (x == y) результат поиска в таблице (который ведется с ипользованием hash(x), hash(y)) был идентичен.
42 == 42.0
hash(42), hash(42.0), hash(42.0000001)
# +
# реализация hash для пользовательского типа данных:
class Color:
def __init__(self, r, g, b):
assert type(r) is int
assert 0 <= r <= 255
self.__red = r
assert type(g) is int
assert 0 <= g <= 255
self.__green = g
assert type(b) is int
assert 0 <= b <= 255
self.__blue = b
@property
def red(self):
return self.__red
@property
def green(self):
return self.__green
@property
def blue(self):
return self.__blue
def __hash__(self):
return hash((self.__red, self.__green, self.__blue))
def __eq__(self, other):
return self.__red == other.red and\
self.__green == other.green and self.__blue == other.blue
# -
c1 = Color(2, 2, 115)
c1.red, c1.green, c1.blue
hash(c1)
c2 = Color(2, 2, 115)
c1 == c2
hash(c1) == hash(c2)
dc = dict()
dc[c1] = 110
dc[c1]
# ### Методы разрешения коллизий
# Разрешение коллизий при помощи цепочек. При использовании данного метода все элементы, хешированные в одну и ту же ячейку, объединяются в связанный список, как показано на рис. Ячейка j или содержит указатель на заголовок списка всех элементов, хеш-значение ключа которых равно j; если таких элементов нет, то ячейка содержит значение None.
# Хеш-таблица использующая цепочки для разрешения коллизий:
# 
def h(k):
return k % 11
keys1 = [54, 26, 93, 17, 77, 31, 44, 20, 55]
[h(k) for k in keys1]
# Время, необходимое для вставки в наихудшем случае, равно $O(1)$. Процедура вставки выполняется очень быстро, поскольку предполагается, что вставляемый элемент отсутствует в таблице. При необходимости это предположение может быть проверено путем выполнения поиска перед вставкой. Время работы поиска в наихудшем случае пропорционально длине списка. Удаление элемента может быть выполнено за время $O(1)$.
# *Открытая адресация*
# При использовании метода открытой адресации все элементы хранятся непосредственно в хеш-таблице, т.е. каждая запись таблицы содержит либо элемент динамического множества, либо специальное значение (None или его заменитель).
#
# При поиске элемента мы систематически проверяем ячейки таблицы до тех пор, пока не найдем искомый элемент или пока
# не убедимся в его отсутствии в таблице. Здесь, в отличие от метода цепочек, нет ни списков, ни элементов, хранящихся вне таблицы. Таким образом, в методе открытой адресации хеш-таблица может оказаться заполненной, делая невозможной вставку новых элементов.
# Коэффициент заполнения таблицы (load factor):
#
# $$\lambda = \frac {number\_of\_items}{table\_size}$$
#
# В хеш-таблице с открытой адресацией коээфициент заполнения таблицы $\lambda$ не может превышать 1.
# Вместо того чтобы следовать по указателям, при открытой адресации мы вычисляем последовательность проверяемых ячеек. Дополнительная память, освобождающаяся в результате отказа от указателей, позволяет использовать хеш-таблицы
# большего размера при том же общем количестве памяти, потенциально
# приводя к меньшему количеству коллизий и более быстрой выборке.
# Для выполнения вставки при открытой адресации мы последовательно проверяем, или исследуем (probe), ячейки хеш-таблицы до тех пор, пока не находим пустую ячейку, в которую помещаем вставляемый ключ.
#
# Для определения исследуемых ячеек хеш-функция расширяется, в нее включается в качестве второго аргумента номер исследования (начинающийся с 0). В методе открытой адресации требуется, чтобы для каждого ключа $k$ последовательность исследований $h(k, 0), h(k, 1),\ldots,h(k, m-1)$ представляла собой перестановку множества $\{0,1,..., m-1\}$, чтобы в
# конечном счете могли быть просмотрены все ячейки хеш-таблицы.
# ```
# % Псевдокод помоещения значения в хеш-таблицу с открытой адресацией:
# HASH_INSERT(T, k)
# 1 i ← 0
# 2 repeat j ← h(k, i)
# 3 if T[j] = NIL
# 4 then T[j] ← k
# 5 return j
# 6 else i ← i + 1
# 7 until i = m
# 8 error "Хеш-таблица переполнена"
# ```
# ```
# % Псевдокод поиска значения в хеш-таблице с открытой адресацией:
# HASH_SEARCH(T, k)
# 1 i ← 0
# 2 repeat j ← h(k,i)
# 3 if T[j] = k
# 4 then return j
# 5 i ← i + 1
# 6 until T[j] = NIL or i = m
# 7 return NIL
# ```
# Процедура **удаления из хеш-таблицы** с открытой адресацией достаточно сложна. При удалении ключа из ячейки i мы не можем просто пометить ее значением NIL. Поступив так, мы можем сделать невозможным выборку ключа k, в процессе вставки которого исследовалась и оказалась занятой ячейка i. Одно из решений состоит в том,
# чтобы помечать такие ячейки специальным значением DELETED вместо NIL. При этом мы должны слегка изменить процедуру
# HASH_INSERT, с тем, чтобы она рассматривала такую ячейку, как пустую и могла вставить в нее новый ключ. В процедуре
# HASH_SEARCH никакие изменения не требуются, поскольку мы просто пропускаем такие ячейки при поиске и исследуем следующие ячейки в последовательности. Однако при использовании специального значения DELETED время поиска перестает зависеть от коэффициента заполнения.
# *Линейное исследование*
#
# Пусть задана обычная хеш-функция $hʹ : U \rightarrow \{0,1, \ldots, m-1\}$, которую мы будем в дальнейшем именовать
# вспомогательной хеш-функцией (auxiliary hash function). Метод линейного исследования для вычисления последовательности исследований использует хеш-функцию
#
# $$h(k, i) = (hʹ(k) + i)\mod m$$
# Линейное исследование легко реализуется, однако с ним связана **проблема первичной кластеризации**, связанной с созданием длинных последовательностей занятых ячеек, что, увеличивает среднее время поиска. Кластеры возникают в связи с тем,
# что вероятность заполнения пустой ячейки, которой предшествуют $i$ заполненных ячеек, равна $(i + 1)/m$. Таким образом, длинные серии заполненных ячеек имеют тенденцию к все большему удлинению, что приводит к увеличению среднего времени поиска.
# Хеш-таблица с открытой адресацией и линейным исследованием:
# 
keys1 = [54, 26, 93, 17, 77, 31, 44, 20, 55]
[h(k) for k in keys1]
# *Квадратичное исследование*
#
# Квадратичное исследование использует хеш-функцию вида:
#
# $$h(k, i) = (hʹ(k) + с_1 i + с_2 i^2) \mod m$$
#
# где $hʹ$ – вспомогательная хеш-функция, $с_1$ и $с_2 \neq 0$ – вспомогательные константы, а $i$ принимает значения от 0 до $m-1$ включительно.
#
# Начальная исследуемая ячейка - $Т[hʹ(k)]$; остальные исследуемые позиции смещены относительно нее на величины, которые описываются квадратичной зависимостью от номера исследования $i$.
#
# Этот метод работает существенно лучше линейного исследования, но для того, чтобы исследование охватывало все ячейки, необходим выбор специальных значений $с_1$, $с_2$ и $m$ . Кроме того, если два ключа имеют одну и то же начальную
# позицию исследования, то одинаковы и последовательности исследования в целом.
# *Двойное хеширование*
#
# Двойное хеширование представляет собой один из наилучших способов использования открытой адресации, поскольку получаемые при этом перестановки обладают многими характеристиками случайно выбираемых перестановок. Двойное хеширование использует хеш-функцию вида
#
# $$h(k, i) = (h_1(k) + i h_2(k)) \mod m$$
#
# где $h_1$ и $h_2$ – вспомогательные хеш-функции. Начальное исследование выполняется в позиции $Т[h_1(k))]$, а смещение каждой из последующих исследуемых ячеек относительно предыдущей равно $h_2(k)$ по модулю $m$.
#
# В отличие от линейного и квадратичного исследования, в данном случае последовательность исследования зависит от клю-
# ча $k$ по двум параметрам – в плане выбора начальной исследуемой ячейки и расстояния между соседними исследуемыми ячейками, так как оба эти параметра зависят от значения ключа. Производительность двойного хеширования достаточно
# близка к производительности "идеальной" схемы равномерного хеширования.
# # Спасибо за внимание!
#
# ----
#
#
# Перехеширование (rehashing)
#
# In the hash table schemes described thus far, it is important that the load factor,
# λ = n/N , be kept below 1. With separate chaining, as λ gets very close to 1, the
# probability of a collision greatly increases, which adds overhead to our operations,
# since we must revert to linear-time list-based methods in buckets that have col-
# lisions.
#
# Experiments and average-case analyses suggest that we should maintain
# λ < 0.9 for hash tables with separate chaining. With open addressing, on the other hand, as the load factor λ grows beyond 0.5 and starts approaching 1, clusters of entries in the bucket array start to grow as well.
#
# If an insertion causes the load factor of a hash table to go above the specified
# threshold, then it is common to resize the table (to regain the specified load factor)
# and to reinsert all objects into this new table. Although we need not define a new
# hash code for each object, we do need to reapply a new compression function that
# takes into consideration the size of the new table. Each rehashing will generally
# scatter the items throughout the new bucket array. When rehashing to a new table, it
# is a good requirement for the new array’s size to be at least double the previous size.
#
|
Course I/Алгоритмы Python/Part2/лекции/lec17_asd4/lec17_asd4_v6.ipynb
|
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ## Lec20 `#23/11`
;; ## REPL Loop in Scheme
;;
;; Source for: [Lec20](../src/Lec20.scm)
;;
;;
;; +
; v1
(define (eval exp)
(cond
((number? exp) exp)
((sum? exp) (eval-sum exp))
(else (error))))
(define (eval-sum exp)
(+ (eval (cadr exp))
(eval (caddr exp))))
(define (sum? exp)
(and (pair? exp) (equal? (car exp) '+)))
;; -
(eval `(+ 2 3))
(eval `(+ 4 (+ 1 4)))
;; ```scheme
;; (define (eval exp)
;; (cond
;; ((number? exp) exp)
;; ((sum? exp) (eval-sum exp))
;; ((symbol? exp) (lookup exp))
;; ((define? exp) (eval-define exp))
;; (else (error))))
;;
;; (define env (make-table))
;; (define (lookup name)
;; (let ((binding (table-get env name)))
;; (if (null? name)
;; (error)
;; (binding-value binding))))
;;
;; (define (eval-define exp)
;; (let ((name (cadr exp))
;; (valexp (caddr exp)))
;; (table-put! env name (eval valexp))))
;; ```
;; Rest is on the source code given in the start!
|
Lectures/.ipynb_checkpoints/Lec20-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Example 2: Convert TFHub Models to OpenVINO IR
# This notebook shows how to convert TFHub models to OpenVINO IR format.
#
# See list of supported models in TFHub-SupportedModelList
#
# Following are the steps
# - Install necessary pip packages.
# - In AWS Sagemaker environment, remove few unused conda envs. This will ensure we have enought space for OpenVINO docker images.
# - Select a model from the supported list
# - Run the conversion
# ### Install necessary pip packages
# !pip3 install --upgrade pip
# !pip3 install -r requirements.txt
# ### Removed few unused conda envs.
# !bash remove-unused-conda-envs.sh
# ## Convert TFHub Models to OpenVINO IR
from ov_utils import create_ir
import csv
# ### Provide below details:
#
# `bucket_name` - S3 bucket name with "sagemaker" as part of the name.
# Example: sagemaker-ir-creation
#
# `tfhub_model_url` - TfHub model URL. See supported list in TFHub-SupportedModelList
#
# `input_shape` - Input shape of the selected model. See TFHub-SupportedModelList
#
# `precision_type` - Use either FP32 or FP16
#
# #### NOTE: If the bucket does not exist then it will be created.
#
# #### Example:
# ```
# bucket_name = "sagemaker-ir-creation"
# tfhub_model_url = "https://tfhub.dev/google/efficientnet/b0/classification/1"
# input_shape = "1,224,224,3"
# precision_type = "FP32"
# ```
bucket_name = "sagemaker-ir-creation" # must have sagemaker as part of the name
tfhub_model_url = "Pick-supported-model-url"
input_shape = "model-input-shape"
precision_type = "FP32" # FP16 also supported
# #### Create IR params.
# +
url_arr = tfhub_model_url.split("/")
url_arr_len = len(url_arr)
model_name = "".join([url_arr[url_arr_len - 4], "-", url_arr[url_arr_len - 3]])
output_dir = "".join(["./", model_name.replace('.','-'), "-tfhub-", precision_type])
if isinstance(input_shape, str):
input_shape = [int(i) for i in input_shape.split(",")]
create_ir_params = {
"tfhub_model_url": tfhub_model_url,
"output_dir": output_dir,
"mo_params": {
"input_shape": input_shape,
"data_type": precision_type,
"model_name": model_name
},
"bucket_name": bucket_name
}
# -
# ### Start the IR creation and upload to S3.
try:
create_ir(create_ir_params)
print (f"IR files created for model:{model_name} and the same is uploaded in S3:{bucket_name}")
except Exception as err:
print(f"FAILED: Please find the error details below:")
print(err)
|
aws/mo-utility/create_ir_for_tfhub.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Belaschich/Projeto_Final_SoulCode/blob/main/ProjetoFinal_(ApacheBeam_Oficial).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hbM8v6YSUM1A"
# # Intalação ApacheBeam
#
# + [markdown] id="QDNI7wZ9XcOm"
# #DEVE-SE REINICIAR O AMBIENTE DE EXECUÇÃO
# + id="CjYhxWBk1ICS" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="a698d0cb-e062-4e68-e3aa-61baeb0d3902"
pip install apache-beam[interactive]
# + [markdown] id="6ZW0gDBFXj4n"
# #DEVE-SE REINICIAR O AMBIENTE DE EXECUÇÃO
# + id="ph74ETYEMk7X"
pip install apache_beam[gcp]
# + [markdown] id="HhMSnmahUa_A"
#
#
# ```
# # Pipeline (DataFlow GCP)
# + id="JeYFj7s0Fxgz" colab={"base_uri": "https://localhost:8080/"} outputId="dc990b49-ee02-4685-d74e-5a1e9d4233e4"
#IMPORT BIBLIOTECAS NECESSÁRIAS PARA A CRIAÇÃO DO PIPELINE
import apache_beam as beam
import os
from apache_beam.options.pipeline_options import PipelineOptions
#OPÇÕES DE CONEXÃO COM O GCP PARA IMPORTAÇÃO/EXPORTAÇÃO
pipeline_options = {
'project': 'testeacesso-332518',
'runner': 'DataflowRunner',
'region': 'southamerica-east1',
'staging_location': 'gs://bucket-projeto-g10/saida/temp',
'temp_location': 'gs://bucket-projeto-g10/saida/temp',
'template_location': 'gs://bucket-projeto-g10/saida/template/template_arq_batch'
}
#CHAVE DE ACESSO AO PROJETO GCP
serviceAccount = r'/content/chaveprogrupo10-332518-4db3212adcd1.json'
os.environ['GOOGLE_APPLICATION_CREDENTIALS'] = serviceAccount
#CRIAÇÃO DA PIPELINE:
pipeline_options = PipelineOptions.from_dictionary(pipeline_options)
p1 = beam.Pipeline(options=pipeline_options)
Pipeline_base_Na = (
p1
#IMPORTAÇÃO DA BASE TRATADA EM NÍVEL SPARK
| "Importar Dados America do Norte" >> beam.io.ReadFromText(r"gs://bucket-projeto-g10/saida_tratado/nivel_spark/Base_ConsumoNA (3.0).csv", skip_header_lines = 1)
| "Separar por Vírgulas" >> beam.Map(lambda record: record.split(','))
#APLICAÇÃO DE FILTROS
| "Filto ano" >> beam.Filter(lambda record: int(record[0]) == 2020 )
| "Filto Categoria" >> beam.Filter(lambda record: str(record[4]) == 'Destilado' )
#EXPORTAÇÃO DA BASE FILTRADA VIA DATAFLOW
| "Saida Para GCP" >> beam.io.WriteToText(r"gs://bucket-projeto-g10/saida_tratado/nivel_apachebeam/Base_ConsumoNA_Destilado2020 (4.0).csv")
)
#EXECUÇÃO DA PIPELINE
p1.run()
|
ProjetoFinal_(ApacheBeam_Oficial).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib import gridspec
df = pd.read_csv('creditcard.csv')
df.head()
df.info()
df.describe()
#Describing the Data
print(df.shape)
print(df.describe())
# Determine number of fraud cases in dataset
fraud = df[df['Class'] == 1]
valid = df[df['Class'] == 0]
outlierFraction = len(fraud)/float(len(valid))
print(outlierFraction)
print('Fraud Cases: {}'.format(len(df[df['Class'] == 1])))
print('Valid Transactions: {}'.format(len(df[df['Class'] == 0])))
print("Amount details of the fraudulent transaction")
fraud.Amount.describe()
# Correlation matrix
corrmat = df.corr()
fig = plt.figure(figsize = (12, 9))
sns.heatmap(corrmat, vmax = .8, square = True)
plt.show()
# dividing the X and the Y from the dataset
X = df.drop(['Class'], axis = 1)
Y = df["Class"]
print(X.shape)
print(Y.shape)
# getting just the values for the sake of processing
# (its a numpy array with no columns)
xData = X.values
yData = Y.values
# Using Skicit-learn to split data into training and testing sets
from sklearn.model_selection import train_test_split
# Split the data into training and testing sets
xTrain, xTest, yTrain, yTest = train_test_split(
xData, yData, test_size = 0.2, random_state = 42)
# Building the Random Forest Classifier (RANDOM FOREST)
from sklearn.ensemble import RandomForestClassifier
# random forest model creation
rfc = RandomForestClassifier()
rfc.fit(xTrain, yTrain)
# predictions
yPred = rfc.predict(xTest)
# +
# Evaluating the classifier
# printing every score of the classifier
# scoring in anything
from sklearn.metrics import classification_report, accuracy_score
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import f1_score, matthews_corrcoef
from sklearn.metrics import confusion_matrix
n_outliers = len(fraud)
n_errors = (yPred != yTest).sum()
print("The model used is Random Forest classifier")
acc = accuracy_score(yTest, yPred)
print("The accuracy is {}".format(acc))
prec = precision_score(yTest, yPred)
print("The precision is {}".format(prec))
rec = recall_score(yTest, yPred)
print("The recall is {}".format(rec))
f1 = f1_score(yTest, yPred)
print("The F1-Score is {}".format(f1))
MCC = matthews_corrcoef(yTest, yPred)
print("The Matthews correlation coefficient is{}".format(MCC))
# -
# printing the confusion matrix
LABELS = ['Normal', 'Fraud']
conf_matrix = confusion_matrix(yTest, yPred)
plt.figure(figsize =(12, 12))
sns.heatmap(conf_matrix, xticklabels = LABELS,
yticklabels = LABELS, annot = True, fmt ="d");
plt.title("Confusion matrix")
plt.ylabel('True class')
plt.xlabel('Predicted class')
plt.show()
|
Credit Card Fraud Detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/araffin/rl-tutorial-jnrr19/blob/sb3/5_custom_gym_env.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="AoxOjIlOImwx" colab_type="text"
# # Stable Baselines3 Tutorial - Creating a custom Gym environment
#
# Github repo: https://github.com/araffin/rl-tutorial-jnrr19/tree/sb3/
#
# Stable-Baselines3: https://github.com/DLR-RM/stable-baselines3
#
# Documentation: https://stable-baselines3.readthedocs.io/en/master/
#
# RL Baselines3 zoo: https://github.com/DLR-RM/rl-baselines3-zoo
#
#
# ## Introduction
#
# In this notebook, you will learn how to use your own environment following the OpenAI Gym interface.
# Once it is done, you can easily use any compatible (depending on the action space) RL algorithm from Stable Baselines on that environment.
#
# ## Install Dependencies and Stable Baselines3 Using Pip
#
#
# + id="Sp8rSS4DIhEV" colab_type="code" colab={}
# !pip install stable-baselines3[extra]
# + [markdown] id="rzevZcgmJmhi" colab_type="text"
# ## First steps with the gym interface
#
# As you have noticed in the previous notebooks, an environment that follows the gym interface is quite simple to use.
# It provides to this user mainly three methods:
# - `reset()` called at the beginning of an episode, it returns an observation
# - `step(action)` called to take an action with the environment, it returns the next observation, the immediate reward, whether the episode is over and additional information
# - (Optional) `render(method='human')` which allow to visualize the agent in action. Note that graphical interface does not work on google colab, so we cannot use it directly (we have to rely on `method='rbg_array'` to retrieve an image of the scene
#
# Under the hood, it also contains two useful properties:
# - `observation_space` which one of the gym spaces (`Discrete`, `Box`, ...) and describe the type and shape of the observation
# - `action_space` which is also a gym space object that describes the action space, so the type of action that can be taken
#
# The best way to learn about gym spaces is to look at the [source code](https://github.com/openai/gym/tree/master/gym/spaces), but you need to know at least the main ones:
# - `gym.spaces.Box`: A (possibly unbounded) box in $R^n$. Specifically, a Box represents the Cartesian product of n closed intervals. Each interval has the form of one of [a, b], (-oo, b], [a, oo), or (-oo, oo). Example: A 1D-Vector or an image observation can be described with the Box space.
# ```python
# # Example for using image as input:
# observation_space = spaces.Box(low=0, high=255, shape=(HEIGHT, WIDTH, N_CHANNELS), dtype=np.uint8)
# ```
#
# - `gym.spaces.Discrete`: A discrete space in $\{ 0, 1, \dots, n-1 \}$
# Example: if you have two actions ("left" and "right") you can represent your action space using `Discrete(2)`, the first action will be 0 and the second 1.
#
#
#
# [Documentation on custom env](https://stable-baselines3.readthedocs.io/en/master/guide/custom_env.html)
# + id="I98IKKyNJl6K" colab_type="code" colab={}
import gym
env = gym.make("CartPole-v1")
# Box(4,) means that it is a Vector with 4 components
print("Observation space:", env.observation_space)
print("Shape:", env.observation_space.shape)
# Discrete(2) means that there is two discrete actions
print("Action space:", env.action_space)
# The reset method is called at the beginning of an episode
obs = env.reset()
# Sample a random action
action = env.action_space.sample()
print("Sampled action:", action)
obs, reward, done, info = env.step(action)
# Note the obs is a numpy array
# info is an empty dict for now but can contain any debugging info
# reward is a scalar
print(obs.shape, reward, done, info)
# + [markdown] id="RqxatIwPOXe_" colab_type="text"
# ## Gym env skeleton
#
# In practice this is how a gym environment looks like.
# Here, we have implemented a simple grid world were the agent must learn to go always left.
# + id="rYzDXA9vJfz1" colab_type="code" colab={}
import numpy as np
import gym
from gym import spaces
class GoLeftEnv(gym.Env):
"""
Custom Environment that follows gym interface.
This is a simple env where the agent must learn to go always left.
"""
# Because of google colab, we cannot implement the GUI ('human' render mode)
metadata = {'render.modes': ['console']}
# Define constants for clearer code
LEFT = 0
RIGHT = 1
def __init__(self, grid_size=10):
super(GoLeftEnv, self).__init__()
# Size of the 1D-grid
self.grid_size = grid_size
# Initialize the agent at the right of the grid
self.agent_pos = grid_size - 1
# Define action and observation space
# They must be gym.spaces objects
# Example when using discrete actions, we have two: left and right
n_actions = 2
self.action_space = spaces.Discrete(n_actions)
# The observation will be the coordinate of the agent
# this can be described both by Discrete and Box space
self.observation_space = spaces.Box(low=0, high=self.grid_size,
shape=(1,), dtype=np.float32)
def reset(self):
"""
Important: the observation must be a numpy array
:return: (np.array)
"""
# Initialize the agent at the right of the grid
self.agent_pos = self.grid_size - 1
# here we convert to float32 to make it more general (in case we want to use continuous actions)
return np.array([self.agent_pos]).astype(np.float32)
def step(self, action):
if action == self.LEFT:
self.agent_pos -= 1
elif action == self.RIGHT:
self.agent_pos += 1
else:
raise ValueError("Received invalid action={} which is not part of the action space".format(action))
# Account for the boundaries of the grid
self.agent_pos = np.clip(self.agent_pos, 0, self.grid_size)
# Are we at the left of the grid?
done = bool(self.agent_pos == 0)
# Null reward everywhere except when reaching the goal (left of the grid)
reward = 1 if self.agent_pos == 0 else 0
# Optionally we can pass additional info, we are not using that for now
info = {}
return np.array([self.agent_pos]).astype(np.float32), reward, done, info
def render(self, mode='console'):
if mode != 'console':
raise NotImplementedError()
# agent is represented as a cross, rest as a dot
print("." * self.agent_pos, end="")
print("x", end="")
print("." * (self.grid_size - self.agent_pos))
def close(self):
pass
# + [markdown] id="Zy5mlho1-Ine" colab_type="text"
# ### Validate the environment
#
# Stable Baselines3 provides a [helper](https://stable-baselines3.readthedocs.io/en/master/common/env_checker.html) to check that your environment follows the Gym interface. It also optionally checks that the environment is compatible with Stable-Baselines (and emits warning if necessary).
# + id="9DOpP_B0-LXm" colab_type="code" colab={}
from stable_baselines3.common.env_checker import check_env
# + id="1CcUVatq-P0l" colab_type="code" colab={}
env = GoLeftEnv()
# If the environment don't follow the interface, an error will be thrown
check_env(env, warn=True)
# + [markdown] id="eJ3khFtkSE0g" colab_type="text"
# ### Testing the environment
# + id="i62yf2LvSAYY" colab_type="code" colab={}
env = GoLeftEnv(grid_size=10)
obs = env.reset()
env.render()
print(env.observation_space)
print(env.action_space)
print(env.action_space.sample())
GO_LEFT = 0
# Hardcoded best agent: always go left!
n_steps = 20
for step in range(n_steps):
print("Step {}".format(step + 1))
obs, reward, done, info = env.step(GO_LEFT)
print('obs=', obs, 'reward=', reward, 'done=', done)
env.render()
if done:
print("Goal reached!", "reward=", reward)
break
# + [markdown] id="Pv1e1qJETfHU" colab_type="text"
# ### Try it with Stable-Baselines
#
# Once your environment follow the gym interface, it is quite easy to plug in any algorithm from stable-baselines
# + id="PQfLBE28SNDr" colab_type="code" colab={}
from stable_baselines3 import PPO, A2C # DQN coming soon
from stable_baselines3.common.cmd_util import make_vec_env
# Instantiate the env
env = GoLeftEnv(grid_size=10)
# wrap it
env = make_vec_env(lambda: env, n_envs=1)
# + id="zRV4Q7FVUKB6" colab_type="code" colab={}
# Train the agent
model = A2C('MlpPolicy', env, verbose=1).learn(5000)
# + id="BJbeiF0RUN-p" colab_type="code" colab={}
# Test the trained agent
obs = env.reset()
n_steps = 20
for step in range(n_steps):
action, _ = model.predict(obs, deterministic=True)
print("Step {}".format(step + 1))
print("Action: ", action)
obs, reward, done, info = env.step(action)
print('obs=', obs, 'reward=', reward, 'done=', done)
env.render(mode='console')
if done:
# Note that the VecEnv resets automatically
# when a done signal is encountered
print("Goal reached!", "reward=", reward)
break
# + [markdown] id="jOggIa9sU--b" colab_type="text"
# ## It is your turn now, be creative!
#
# As an exercise, that's now your turn to build a custom gym environment.
# There is no constrain about what to do, be creative! (but not too creative, there is not enough time for that)
#
# If you don't have any idea, here is is a list of the environment you can implement:
# - Transform the discrete grid world to a continuous one, you will need to change a bit the logic and the action space
# - Create a 2D grid world and add walls
# - Create a tic-tac-toe game
#
# + id="lBDp4Pm-Uh4D" colab_type="code" colab={}
|
5_custom_gym_env.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.2
# language: sage
# name: sagemath
# ---
# +
################################################################################
# THIS IS THE MAIN CODE #
################################################################################
from codebuilder import *
################################################################################
# THE CODE SUPPORTS POLYNOMIALS OF AT MOST 20 VARIABLES. #
# THE VARIABLES ARE x1,x2, ..., x20 #
# FUNCTION "Generator_Matrix" GENERATES THE CODE MATRIX FROM THE POLYNOMIAL #
################################################################################
#example polynomial:
poly = x1+x2*x3
#WARNING: make sure to use consecutive variables, i.e., x2*x3+x1+x4*x5 is good poly, but x1+x4*x3 is not,
# because it is missing x2
#example code generated:
Generator_Matrix(poly,5)
# +
# We use the language of Kasami et.al.
# m is the number of variables.
# r is the degree
# We need: r = m - 4, m - 5, m - 6, ... , 0
# for the code to be triorthogonal.
# We are interested in cases where the weight of the poly is less than 2.5*16 = 40. If r <= m-6, then the weight is
# more than 64. So we only consider the cases where r = m - 4 and r = m - 5
# If r = m - 5, then then only polynomial with weight less than 40 is poly = 1, any other polynomial with will have
# weight more than 32+16=48
# +
################################################################################
# THIS IS THE LIST OF ALL POLYNOMIALS WITH WEIGHT < 40 #
# AND THE CORRESPONDING GENERATOR MATRICES #
# THERE ARE 38 OF THEM #
################################################################################
#Gs is the list of generator matrices
Gs = []
#Polys it the list of all polynomials
Polys = []
m = 4
poly = 1
Polys.append([poly,m])
#ncols = 16
#nrows = 5
m = 6
poly = x1*x2+x3*x4
Polys.append([poly,m])
#ncols = 24
#nrows = 7
m = 6
poly = x1*x2+x3*x4+x5*x6
Polys.append([poly,m])
#ncols = 28
#nrows = 7
m = 7
poly = x1*x2*x3 + x4*x5*x6
Polys.append([poly,m])
#ncols = 28
#nrows = 8
m = 8
poly = x1*x2*x3*x4 + x5*x6*x7*x8
Polys.append([poly,m])
#ncols = 30
#nrows = 9
m = 5
poly = 1
Polys.append([poly,m])
#ncols = 32
#nrows = 6
m = 6
poly = x1*x2 + x3
Polys.append([poly,m])
#ncols = 32
#nrows = 7
m = 6
poly = x2*x3 + x1*x4 + x5
Polys.append([poly,m])
#ncols = 32
#nrows = 7
m = 7
poly = x1*x2*x3 + x1*x4*x5 + x2*x3
Polys.append([poly,m])
#ncols = 32
#nrows = 8
m = 7
poly = x1*x3*x4 + x1*x2*x5 + x2*x3*x6
Polys.append([poly,m])
#ncols = 32
#nrows = 8
m = 7
poly = x1*x2*x4 + x1*x5*x6 + x2*x3*x7
Polys.append([poly,m])
#ncols = 32
#nrows = 8
m = 8
poly = x1*x2*x3 + x1*x2*x3*x4 + x1*x2*x5*x6 + x3*x4*x5*x6
Polys.append([poly,m])
#ncols = 32
#nrows = 9
m = 8
poly = x1*x2*x3*x4 + x2*x3*x4*x5 + x1*x5*x6*x7
Polys.append([poly,m])
#ncols = 32
#nrows = 9
m = 8
poly = x1*x2*x3*x5 + x1*x2*x6 + x1*x2*x4*x6 + x3*x4*x5*x7
Polys.append([poly,m])
#ncols = 32
#nrows = 9
m = 9
poly = x1*x2*x3*x4*x5+(x1+1)*x6*x7*x8*x9
Polys.append([poly,m])
#ncols = 32
#nrows = 10
m = 8
poly = x1*x2*(x3*x4 + x7*x8) + x5*x6*x7*x8
Polys.append([poly,m])
#ncols = 34
#nrows = 9
m = 6
poly = x1*x3 + x4*x5 + x2*x6 + 1
Polys.append([poly,m])
#ncols = 36
#nrows = 7
m = 7
poly = x2*x3*x5 + x1*x4*x6 + x1*x2
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 7
poly = x1*x2*x4 + x1*x3*x4 + x2*x3*x5 + x1*x6*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 7
poly = x2*x3*x4 + x1*x3*x5 + x1*x2*x6 + x1*x4*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 7
poly = x2*x3*x5 + x1*x4*x5 + x3*x4*x6 + x1*x2*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 7
poly = x1*x2*x3 + x2*x3*x4 + x1*x2*x5 + x1*x3*x6 + x4*x5*x6 + x1*x2
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 7
poly = x1*x2*x4 + x1*x3*x4 + x1*x5*x6 + x2*x5*x6 + x2*x3*x7 + x3*x5*x7 + x4*x6*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 8
m = 8
poly = x1*x2*x3*x4 + x1*x2*x5*x6 + x3*x4*x5*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 8
poly = x2*x3*x4*x5 + x1*x2*x4*x7 + x1*x3*x6*x8
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 8
poly = x1*x2*x4*x5 + x1*x2*x5*x6 + x1*x3*x4*x7 + x2*x3*x6*x8
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 8
poly = x1*x2*x3*x4 + x1*x3*x5*x6 + x1*x2*x5*x7 + x2*x4*x6*x7
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 8
poly = x2*x3*x4*x5 + x1*x4*x5*x6 + x2*x4*x5*x6 + x1*x2*x3*x7 + x1*x3*x6*x8
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 8
poly = x1*x2*x4*x6 + x1*x2*x3*x7 + x4*x5*x6*x8 + x3*x5*x7*x8
Polys.append([poly,m])
#ncols = 36
#nrows = 9
m = 9
poly = x1*x2*x3*x4*x5+x6*x7*x8*(x9*(x5+1)+x3*x4)
Polys.append([poly,m])
#ncols = 36
#nrows = 10
m = 8
poly = x1*x2*x3*x4 + x1*x2*x5*x6 + x1*x4*x5*x6 + x2*x3*x7*x8
Polys.append([poly,m])
#ncols = 38
#nrows = 9
m = 8
poly = x1*x2*x3*x5 + x1*x2*x4*x6 + x2*x4*x5*x7 + x1*x3*x6*x8
Polys.append([poly,m])
#ncols = 38
#nrows = 9
m = 8
poly = x1*x2*x4*x5 + x1*x4*x5*x6 + x2*x3*x4*x7 + x1*x5*x6*x7 + x1*x2*x3*x8 + x1*x2*x6*x8 + x2*x3*x7*x8
Polys.append([poly,m])
#ncols = 38
#nrows = 9
m = 8
poly = x1*x2*x3*x4 + x2*x4*x5*x6 + x1*x5*x6*x7 + x1*x3*x7*x8
Polys.append([poly,m])
#ncols = 38
#nrows = 9
m = 8
poly = x1*x2*x3*x4 + x1*x2*x3*x5 + x2*x3*x4*x6 + x1*x4*x5*x6 + x1*x5*x7*x8
Polys.append([poly,m])
#ncols = 38
#nrows = 9
m = 9
poly = x1*x2*x3*x4*x5+x6*x7*x8*(x9*x3+x4*x5)
Polys.append([poly,m])
#ncols = 38
#nrows = 10
m = 9
poly = x1*x2*x3*x4*x5+x6*x7*(x3*x4*x8+x2*(x5+x8)*x9)
Polys.append([poly,m])
#ncols = 38
#nrows = 10
m = 10
poly = x1*x2*x3*x4*x5*x6+x7*x8*x9*x10*(x3*x4+x5*(x6+1))
Polys.append([poly,m])
#ncols = 38
#nrows = 11
for [poly,m] in Polys:
Gs.append(Generator_Matrix(poly,m))
# -
################################################################################
# THIS PART JUST PRINTS THE SUBSPACES AND RELEVANT POLYNOMIALS #
################################################################################
i=0
for G in Gs:
print("Code number",i)
print(G)
print("Number of rows:",G.nrows())
print("Number of columns:",G.ncols())
print("Weight Enumerator function:")
print(Weight_Enumerator(G))
print("-"*100)
i=i+1
# +
#Technical comments:
#Rank of weight 8 vectors in #11 is 7 while in #12 is 8
# #0 is the first Bravyi-Haah code, #1 is the second BH code, #11 is the third BH code
# -
#Check if the subspace Gs[2] is triorthogonal
Is_Tri(Gs[15])
# +
################################################################################
# STUDY OF DESCENDANTS #
# FROM THIS POINT WE COMPUTE THE DESCENDANTS OF #
# TRIORTHOGONAL SUBSPACES AND THEIR DISTANCES. #
################################################################################
#Instructions: There are two main functions here:
# Generate_Even_Desc(G,k,func,arg) and Generate_Odd_Desc(G,k,func,arg)
#These functions take the triorthogonal subspace G, and search through all even
# or odd descendants of G with k logical qubits.
# The other input is a function func, which should have the following form
# func(Desc, arg)
# Any time through the search that a descendant of G with k logical qubits is
# constructed, the func will be called, with the two inputs will be passed:
# Desc: which is the descendant, and
# arg: which is the argument given to Generate_Even_Desc and Generate_odd_Desc
# functions.
# IMPORTANT: If the output of func is True, then the search immediately ends and
# the Generate_Even/Odd_Desc functions return True. If the output of func is
# False then the search continues, and the function return False if func never
# outputs True.
#Example: The following prints one odd and one even descendant of Gs[0] with k=1
# and ends the search
def func(Desc,k):
print(Desc)
print("n + 1 =",Desc.ncols()+1)
return True
print("Odd Descendant:")
Generate_Odd_Desc (Gs[0],1,func,None)
print("Even Descendant:")
Generate_Even_Desc(Gs[0],1,func,None)
# +
################################################################################
# FUNCTIONS FOR CHECKING THE DISTANCE OF A TRIORTHOGONAL MATRIX #
################################################################################
#The function Is_Dist_Larger(Desc,d) takes a descendant Desc and return True if
# its distance is (strictly) larger than d. Otherwise, it returns False.
#Example: The following line check if any even descendant of Gs[0] with k = 2
# has distance larger than 1
k = 2
d = 1
print("Is distance (strictly) larger than",d,"? ",Generate_Even_Desc(Gs[0],k,Is_Dist_Larger,d))
#The output is True
#Example: The following line check if any even descendant of Gs[0] with k = 2
# has distance larger than 2
k = 2
d = 2
print("Is distance (strictly) larger than",d,"? ",Generate_Even_Desc(Gs[0],k,Is_Dist_Larger,d))
#The output is False
#We conclude that the maximum distance of even descendants of Gs[0] with k = 2 is
# equal to 2
# +
################################################################################
# CODE FOR RUNNING THE FAST MULTITHREAD C++ CODE. #
# THE CODE SEARCHES THROUGH THE POLYNOMIALS WITH GIVEN HAMMING WEIGHT #
# AND BASE POLYNOMIALS, AND RETURNS A REPRESENTATIVE FROM EACH AFFINE #
# EQUIVALENCE CLASSES OF THE POLYNOMIALS. SEE THE PAPER FOR DETAILS. #
################################################################################
base_pairs=[]
################################################################################
#Insert the base pairs here.
base1=y1*(y2*y3+y4*y5)
base2=y1*y3*y4
base_pair = [base1,base2]
base_pairs.append(base_pair)
base1=y1*y2*y3
base2=(y1+1)*y3*y4
base_pair = [base1,base2]
base_pairs.append(base_pair)
################################################################################
#The following function writes the instructions and bases in the
# poly_finder_instructions.txt file. number_of_threads should be choses to be
# less than the number of CPU cores. weight is the weight of target Reed-Muller
# polynomials. For other parameters, please see the C++ code.
write_instuctions(base_pairs,weight = 32,number_of_threads=8,trigger_wait=1,trigger_random_jumps=0,max_number_polys=100000)
################################################################################
#Here we compile and run the C++ code.
print("Compiling the C++ code.")
!g++ -O3 classes.cpp -lpthread
print("Running the code:")
print("="*50)
!./a.out
# -
|
Code_Builder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 实战Kaggle比赛:预测房价
# :label:`sec_kaggle_house`
#
# 之前几节我们学习了一些训练深度网络的基本工具和网络正则化的技术(如权重衰减、暂退法等)。
# 本节我们将通过Kaggle比赛,将所学知识付诸实践。
# Kaggle的房价预测比赛是一个很好的起点。
# 此数据集由<NAME>于2011年收集 :cite:`De-Cock.2011`,
# 涵盖了2006-2010年期间亚利桑那州埃姆斯市的房价。
# 这个数据集是相当通用的,不会需要使用复杂模型架构。
# 它比哈里森和鲁宾菲尔德的[波士顿房价](https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.names)
# 数据集要大得多,也有更多的特征。
#
# 本节我们将详细介绍数据预处理、模型设计和超参数选择。
# 通过亲身实践,你将获得一手经验,这些经验将指导你数据科学家职业生涯。
#
# ## 下载和缓存数据集
#
# 在整本书中,我们将下载不同的数据集,并训练和测试模型。
# 这里我们(**实现几个函数来方便下载数据**)。
# 首先,我们建立字典`DATA_HUB`,
# 它可以将数据集名称的字符串映射到数据集相关的二元组上,
# 这个二元组包含数据集的url和验证文件完整性的sha-1密钥。
# 所有类似的数据集都托管在地址为`DATA_URL`的站点上。
#
# + origin_pos=1 tab=["pytorch"]
import hashlib
import os
import tarfile
import zipfile
import requests
#@save
DATA_HUB = dict()
DATA_URL = 'http://d2l-data.s3-accelerate.amazonaws.com/'
# + [markdown] origin_pos=2
# 下面的`download`函数用来下载数据集,
# 将数据集缓存在本地目录(默认情况下为`../data`)中,
# 并返回下载文件的名称。
# 如果缓存目录中已经存在此数据集文件,并且其sha-1与存储在`DATA_HUB`中的相匹配,
# 我们将使用缓存的文件,以避免重复的下载。
#
# + origin_pos=3 tab=["pytorch"]
def download(name, cache_dir=os.path.join('..', 'data')): #@save
"""下载一个DATA_HUB中的文件,返回本地文件名"""
assert name in DATA_HUB, f"{name} 不存在于 {DATA_HUB}"
url, sha1_hash = DATA_HUB[name]
os.makedirs(cache_dir, exist_ok=True)
fname = os.path.join(cache_dir, url.split('/')[-1])
if os.path.exists(fname):
sha1 = hashlib.sha1()
with open(fname, 'rb') as f:
while True:
data = f.read(1048576)
if not data:
break
sha1.update(data)
if sha1.hexdigest() == sha1_hash:
return fname # 命中缓存
print(f'正在从{url}下载{fname}...')
r = requests.get(url, stream=True, verify=True)
with open(fname, 'wb') as f:
f.write(r.content)
return fname
# + [markdown] origin_pos=4
# 我们还需实现两个实用函数:
# 一个将下载并解压缩一个zip或tar文件,
# 另一个是将本书中使用的所有数据集从`DATA_HUB`下载到缓存目录中。
#
# + origin_pos=5 tab=["pytorch"]
def download_extract(name, folder=None): #@save
"""下载并解压zip/tar文件"""
fname = download(name)
base_dir = os.path.dirname(fname)
data_dir, ext = os.path.splitext(fname)
if ext == '.zip':
fp = zipfile.ZipFile(fname, 'r')
elif ext in ('.tar', '.gz'):
fp = tarfile.open(fname, 'r')
else:
assert False, '只有zip/tar文件可以被解压缩'
fp.extractall(base_dir)
return os.path.join(base_dir, folder) if folder else data_dir
def download_all(): #@save
"""下载DATA_HUB中的所有文件"""
for name in DATA_HUB:
download(name)
# + [markdown] origin_pos=6
# ## Kaggle
#
# [Kaggle](https://www.kaggle.com)是一个当今流行举办机器学习比赛的平台,
# 每场比赛都以至少一个数据集为中心。
# 许多比赛有赞助方,他们为获胜的解决方案提供奖金。
# 该平台帮助用户通过论坛和共享代码进行互动,促进协作和竞争。
# 虽然排行榜的追逐往往令人失去理智:
# 有些研究人员短视地专注于预处理步骤,而不是考虑基础性问题。
# 但一个客观的平台有巨大的价值:该平台促进了竞争方法之间的直接定量比较,以及代码共享。
# 这便于每个人都可以学习哪些方法起作用,哪些没有起作用。
# 如果你想参加Kaggle比赛,你首先需要注册一个账户(见 :numref:`fig_kaggle`)。
#
# 
# :width:`400px`
# :label:`fig_kaggle`
#
# 在房价预测比赛页面(如 :numref:`fig_house_pricing` 所示),
# 你在"Data"选项卡下可以找到数据集。
# 你可以通过下面的网址提交预测,并查看排名:
#
# >https://www.kaggle.com/c/house-prices-advanced-regression-techniques
#
# 
# :width:`400px`
# :label:`fig_house_pricing`
#
# ## 访问和读取数据集
#
# 注意,竞赛数据分为训练集和测试集。
# 每条记录都包括房屋的属性值和属性,如街道类型、施工年份、屋顶类型、地下室状况等。
# 这些特征由各种数据类型组成。
# 例如,建筑年份由整数表示,屋顶类型由离散类别表示,其他特征由浮点数表示。
# 这就是现实让事情变得复杂的地方:例如,一些数据完全丢失了,缺失值被简单地标记为“NA”。
# 每套房子的价格只出现在训练集中(毕竟这是一场比赛)。
# 我们将希望划分训练集以创建验证集,但是在将预测结果上传到Kaggle之后,
# 我们只能在官方测试集中评估我们的模型。
# 在 :numref:`fig_house_pricing` 中,"Data"选项卡有下载数据的链接。
#
# 开始之前,我们将[**使用`pandas`读入并处理数据**],
# 这是我们在 :numref:`sec_pandas`中引入的。
# 因此,在继续操作之前,你需要确保已安装`pandas`。
# 幸运的是,如果你正在用Jupyter阅读该书,你可以在不离开笔记本的情况下安装`pandas`。
#
# + origin_pos=8 tab=["pytorch"]
# 如果你没有安装pandas,请取消下一行的注释
# # !pip install pandas
# %matplotlib inline
import numpy as np
import pandas as pd
import torch
from torch import nn
from d2l import torch as d2l
# + [markdown] origin_pos=10
# 为方便起见,我们可以使用上面定义的脚本下载并缓存Kaggle房屋数据集。
#
# + origin_pos=11 tab=["pytorch"]
DATA_HUB['kaggle_house_train'] = ( #@save
DATA_URL + 'kaggle_house_pred_train.csv',
'585e9cc93e70b39160e7921475f9bcd7d31219ce')
DATA_HUB['kaggle_house_test'] = ( #@save
DATA_URL + 'kaggle_house_pred_test.csv',
'fa19780a7b011d9b009e8bff8e99922a8ee2eb90')
# + [markdown] origin_pos=12
# 我们使用`pandas`分别加载包含训练数据和测试数据的两个CSV文件。
#
# + origin_pos=13 tab=["pytorch"]
train_data = pd.read_csv(download('kaggle_house_train'))
test_data = pd.read_csv(download('kaggle_house_test'))
# + [markdown] origin_pos=14
# 训练数据集包括1460个样本,每个样本80个特征和1个标签,
# 而测试数据集包含1459个样本,每个样本80个特征。
#
# + origin_pos=15 tab=["pytorch"]
print(train_data.shape)
print(test_data.shape)
# + [markdown] origin_pos=16
# 让我们看看[**前四个和最后两个特征,以及相应标签**](房价)。
#
# + origin_pos=17 tab=["pytorch"]
print(train_data.iloc[0:4, [0, 1, 2, 3, -3, -2, -1]])
# + [markdown] origin_pos=18
# 我们可以看到,(**在每个样本中,第一个特征是ID,**)
# 这有助于模型识别每个训练样本。
# 虽然这很方便,但它不携带任何用于预测的信息。
# 因此,在将数据提供给模型之前,(**我们将其从数据集中删除**)。
#
# + origin_pos=19 tab=["pytorch"]
all_features = pd.concat((train_data.iloc[:, 1:-1], test_data.iloc[:, 1:]))
# + [markdown] origin_pos=20
# ## 数据预处理
#
# 如上所述,我们有各种各样的数据类型。
# 在开始建模之前,我们需要对数据进行预处理。
# 首先,我们[**将所有缺失的值替换为相应特征的平均值。**]然后,为了将所有特征放在一个共同的尺度上,
# 我们(**通过将特征重新缩放到零均值和单位方差来标准化数据**):
#
# $$x \leftarrow \frac{x - \mu}{\sigma},$$
#
# 其中$\mu$和$\sigma$分别表示均值和标准差。
# 现在,这些特征具有零均值和单位方差,即 $E[\frac{x-\mu}{\sigma}] = \frac{\mu - \mu}{\sigma} = 0$和$E[(x-\mu)^2] = (\sigma^2 + \mu^2) - 2\mu^2+\mu^2 = \sigma^2$。
# 直观地说,我们标准化数据有两个原因:
# 首先,它方便优化。
# 其次,因为我们不知道哪些特征是相关的,
# 所以我们不想让惩罚分配给一个特征的系数比分配给其他任何特征的系数更大。
#
# + origin_pos=21 tab=["pytorch"]
# 若无法获得测试数据,则可根据训练数据计算均值和标准差
numeric_features = all_features.dtypes[all_features.dtypes != 'object'].index
all_features[numeric_features] = all_features[numeric_features].apply(
lambda x: (x - x.mean()) / (x.std()))
# 在标准化数据之后,所有均值消失,因此我们可以将缺失值设置为0
all_features[numeric_features] = all_features[numeric_features].fillna(0)
# + [markdown] origin_pos=22
# 接下来,我们[**处理离散值。**]
# 这包括诸如“MSZoning”之类的特征。
# (**我们用独热编码替换它们**),
# 方法与前面将多类别标签转换为向量的方式相同
# (请参见 :numref:`subsec_classification-problem`)。
# 例如,“MSZoning”包含值“RL”和“Rm”。
# 我们将创建两个新的指示器特征“MSZoning_RL”和“MSZoning_RM”,其值为0或1。
# 根据独热编码,如果“MSZoning”的原始值为“RL”,
# 则:“MSZoning_RL”为1,“MSZoning_RM”为0。
# `pandas`软件包会自动为我们实现这一点。
#
# + origin_pos=23 tab=["pytorch"]
# “Dummy_na=True”将“na”(缺失值)视为有效的特征值,并为其创建指示符特征
all_features = pd.get_dummies(all_features, dummy_na=True)
all_features.shape
# + [markdown] origin_pos=24
# 你可以看到,此转换会将特征的总数量从79个增加到331个。
# 最后,通过`values`属性,我们可以
# [**从`pandas`格式中提取NumPy格式,并将其转换为张量表示**]用于训练。
#
# + origin_pos=25 tab=["pytorch"]
n_train = train_data.shape[0]
train_features = torch.tensor(all_features[:n_train].values, dtype=torch.float32)
test_features = torch.tensor(all_features[n_train:].values, dtype=torch.float32)
train_labels = torch.tensor(
train_data.SalePrice.values.reshape(-1, 1), dtype=torch.float32)
# + [markdown] origin_pos=26
# ## [**训练**]
#
# 首先,我们训练一个带有损失平方的线性模型。
# 显然线性模型很难让我们在竞赛中获胜,但线性模型提供了一种健全性检查,
# 以查看数据中是否存在有意义的信息。
# 如果我们在这里不能做得比随机猜测更好,那么我们很可能存在数据处理错误。
# 如果一切顺利,线性模型将作为*基线*(baseline)模型,
# 让我们直观地知道最好的模型有超出简单的模型多少。
#
# + origin_pos=28 tab=["pytorch"]
loss = nn.MSELoss()
in_features = train_features.shape[1]
def get_net():
net = nn.Sequential(nn.Linear(in_features,1))
return net
# + [markdown] origin_pos=30
# 房价就像股票价格一样,我们关心的是相对数量,而不是绝对数量。
# 因此,[**我们更关心相对误差$\frac{y - \hat{y}}{y}$,**]
# 而不是绝对误差$y - \hat{y}$。
# 例如,如果我们在俄亥俄州农村地区估计一栋房子的价格时,
# 假设我们的预测偏差了10万美元,
# 然而那里一栋典型的房子的价值是12.5万美元,
# 那么模型可能做得很糟糕。
# 另一方面,如果我们在加州豪宅区的预测出现同样的10万美元的偏差,
# (在那里,房价中位数超过400万美元)
# 这可能是一个不错的预测。
#
# (**解决这个问题的一种方法是用价格预测的对数来衡量差异**)。
# 事实上,这也是比赛中官方用来评价提交质量的误差指标。
# 即将$\delta$ for $|\log y - \log \hat{y}| \leq \delta$
# 转换为$e^{-\delta} \leq \frac{\hat{y}}{y} \leq e^\delta$。
# 这使得预测价格的对数与真实标签价格的对数之间出现以下均方根误差:
#
# $$\sqrt{\frac{1}{n}\sum_{i=1}^n\left(\log y_i -\log \hat{y}_i\right)^2}.$$
#
# + origin_pos=32 tab=["pytorch"]
def log_rmse(net, features, labels):
# 为了在取对数时进一步稳定该值,将小于1的值设置为1
clipped_preds = torch.clamp(net(features), 1, float('inf'))
rmse = torch.sqrt(loss(torch.log(clipped_preds),
torch.log(labels)))
return rmse.item()
# + [markdown] origin_pos=34
# 与前面的部分不同,[**我们的训练函数将借助Adam优化器**]
# (我们将在后面章节更详细地描述它)。
# Adam优化器的主要吸引力在于它对初始学习率不那么敏感。
#
# + origin_pos=36 tab=["pytorch"]
def train(net, train_features, train_labels, test_features, test_labels,
num_epochs, learning_rate, weight_decay, batch_size):
train_ls, test_ls = [], []
train_iter = d2l.load_array((train_features, train_labels), batch_size)
# 这里使用的是Adam优化算法
optimizer = torch.optim.Adam(net.parameters(),
lr = learning_rate,
weight_decay = weight_decay)
for epoch in range(num_epochs):
for X, y in train_iter:
optimizer.zero_grad()
l = loss(net(X), y)
l.backward()
optimizer.step()
train_ls.append(log_rmse(net, train_features, train_labels))
if test_labels is not None:
test_ls.append(log_rmse(net, test_features, test_labels))
return train_ls, test_ls
# + [markdown] origin_pos=38
# ## $K$折交叉验证
#
# 你可能还记得,我们在讨论模型选择的部分( :numref:`sec_model_selection`)
# 中介绍了[**K折交叉验证**],
# 它有助于模型选择和超参数调整。
# 我们首先需要定义一个函数,在$K$折交叉验证过程中返回第$i$折的数据。
# 具体地说,它选择第$i$个切片作为验证数据,其余部分作为训练数据。
# 注意,这并不是处理数据的最有效方法,如果我们的数据集大得多,会有其他解决办法。
#
# + origin_pos=39 tab=["pytorch"]
def get_k_fold_data(k, i, X, y):
assert k > 1
fold_size = X.shape[0] // k
X_train, y_train = None, None
for j in range(k):
idx = slice(j * fold_size, (j + 1) * fold_size)
X_part, y_part = X[idx, :], y[idx]
if j == i:
X_valid, y_valid = X_part, y_part
elif X_train is None:
X_train, y_train = X_part, y_part
else:
X_train = torch.cat([X_train, X_part], 0)
y_train = torch.cat([y_train, y_part], 0)
return X_train, y_train, X_valid, y_valid
# + [markdown] origin_pos=40
# 当我们在$K$折交叉验证中训练$K$次后,[**返回训练和验证误差的平均值**]。
#
# + origin_pos=41 tab=["pytorch"]
def k_fold(k, X_train, y_train, num_epochs, learning_rate, weight_decay,
batch_size):
train_l_sum, valid_l_sum = 0, 0
for i in range(k):
data = get_k_fold_data(k, i, X_train, y_train)
net = get_net()
train_ls, valid_ls = train(net, *data, num_epochs, learning_rate,
weight_decay, batch_size)
train_l_sum += train_ls[-1]
valid_l_sum += valid_ls[-1]
if i == 0:
d2l.plot(list(range(1, num_epochs + 1)), [train_ls, valid_ls],
xlabel='epoch', ylabel='rmse', xlim=[1, num_epochs],
legend=['train', 'valid'], yscale='log')
print(f'折{i + 1},训练log rmse{float(train_ls[-1]):f}, '
f'验证log rmse{float(valid_ls[-1]):f}')
return train_l_sum / k, valid_l_sum / k
# + [markdown] origin_pos=42
# ## [**模型选择**]
#
# 在本例中,我们选择了一组未调优的超参数,并将其留给读者来改进模型。
# 找到一组调优的超参数可能需要时间,这取决于一个人优化了多少变量。
# 有了足够大的数据集和合理设置的超参数,$K$折交叉验证往往对多次测试具有相当的稳定性。
# 然而,如果我们尝试了不合理的超参数,我们可能会发现验证效果不再代表真正的误差。
#
# + origin_pos=43 tab=["pytorch"]
k, num_epochs, lr, weight_decay, batch_size = 5, 100, 5, 0, 64
train_l, valid_l = k_fold(k, train_features, train_labels, num_epochs, lr,
weight_decay, batch_size)
print(f'{k}-折验证: 平均训练log rmse: {float(train_l):f}, '
f'平均验证log rmse: {float(valid_l):f}')
# + [markdown] origin_pos=44
# 请注意,有时一组超参数的训练误差可能非常低,但$K$折交叉验证的误差要高得多,
# 这表明模型过拟合了。
# 在整个训练过程中,你将希望监控训练误差和验证误差这两个数字。
# 较少的过拟合可能表明现有数据可以支撑一个更强大的模型,
# 较大的过拟合可能意味着我们可以通过正则化技术来获益。
#
# ## [**提交你的Kaggle预测**]
#
# 既然我们知道应该选择什么样的超参数,
# 我们不妨使用所有数据对其进行训练
# (而不是仅使用交叉验证中使用的$1-1/K$的数据)。
# 然后,我们通过这种方式获得的模型可以应用于测试集。
# 将预测保存在CSV文件中可以简化将结果上传到Kaggle的过程。
#
# + origin_pos=45 tab=["pytorch"]
def train_and_pred(train_features, test_feature, train_labels, test_data,
num_epochs, lr, weight_decay, batch_size):
net = get_net()
train_ls, _ = train(net, train_features, train_labels, None, None,
num_epochs, lr, weight_decay, batch_size)
d2l.plot(np.arange(1, num_epochs + 1), [train_ls], xlabel='epoch',
ylabel='log rmse', xlim=[1, num_epochs], yscale='log')
print(f'训练log rmse:{float(train_ls[-1]):f}')
# 将网络应用于测试集。
preds = net(test_features).detach().numpy()
# 将其重新格式化以导出到Kaggle
test_data['SalePrice'] = pd.Series(preds.reshape(1, -1)[0])
submission = pd.concat([test_data['Id'], test_data['SalePrice']], axis=1)
submission.to_csv('submission.csv', index=False)
# + [markdown] origin_pos=46
# 如果测试集上的预测与$K$倍交叉验证过程中的预测相似,
# 那就是时候把它们上传到Kaggle了。
# 下面的代码将生成一个名为`submission.csv`的文件。
#
# + origin_pos=47 tab=["pytorch"]
train_and_pred(train_features, test_features, train_labels, test_data,
num_epochs, lr, weight_decay, batch_size)
# + [markdown] origin_pos=48
# 接下来,如 :numref:`fig_kaggle_submit2`中所示,
# 我们可以提交预测到Kaggle上,并查看在测试集上的预测与实际房价(标签)的比较情况。
# 步骤非常简单:
#
# * 登录Kaggle网站,访问房价预测竞赛页面。
# * 点击“Submit Predictions”或“Late Submission”按钮(在撰写本文时,该按钮位于右侧)。
# * 点击页面底部虚线框中的“Upload Submission File”按钮,选择你要上传的预测文件。
# * 点击页面底部的“Make Submission”按钮,即可查看你的结果。
#
# 
# :width:`400px`
# :label:`fig_kaggle_submit2`
#
# ## 小结
#
# * 真实数据通常混合了不同的数据类型,需要进行预处理。
# * 常用的预处理方法:将实值数据重新缩放为零均值和单位方法;用均值替换缺失值。
# * 将类别特征转化为指标特征,可以使我们把这个特征当作一个独热向量来对待。
# * 我们可以使用$K$折交叉验证来选择模型并调整超参数。
# * 对数对于相对误差很有用。
#
# ## 练习
#
# 1. 把你的预测提交给Kaggle,它有多好?
# 1. 你能通过直接最小化价格的对数来改进你的模型吗?如果你试图预测价格的对数而不是价格,会发生什么?
# 1. 用平均值替换缺失值总是好主意吗?提示:你能构造一个不随机丢失值的情况吗?
# 1. 通过$K$折交叉验证调整超参数,从而提高Kaggle的得分。
# 1. 通过改进模型(例如,层、权重衰减和dropout)来提高分数。
# 1. 如果我们没有像本节所做的那样标准化连续的数值特征,会发生什么?
#
# + [markdown] origin_pos=50 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1824)
#
|
d2l/pytorch/chapter_multilayer-perceptrons/kaggle-house-price.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
saldo = 500
valor_saque = float(input('Quanto você quer sacar: '))
if saldo >= valor_saque:
saldo -= valor_saque
print('Saque efetuado com sucesso!')
print('Você sacou '+str(valor_saque)+'R$ e agora resta apenas '+str(saldo)+'R$')
else:
print('Saque não efetuado.')
print('Você não tem saldo o suficiente')
#saldo -= valor_saque
# -
valor_saque = 0
saldo -= valor_saque
print('Você sacou '+str(valor_saque)+'R$ e agora resta apenas '+str(saldo)+'R$')
aprendendo = True
if aprendendo:
print('estou aprendendo Python')
print('quero aprender ainda mais')
|
CondicionalSimples.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
df_dep = pd.read_csv('departamentos-nuevo.csv', dtype="string")
df_dep_sc = pd.read_csv('departamentos-nuevo-sc.csv', dtype="string")
df_casa = pd.read_csv('casas-nuevo.csv', dtype="string")
df_casa_sc = pd.read_csv('casas-nuevo-sc.csv', dtype="string")
def convertir(precios):
lista=[]
for i in precios:
if i[0]=='U':
dato=i
i=i.replace("USD ","")
a=float(i)
a=round(a*20.58,1)
a=int(a)
a=str(a)
lista.append(a)
else:
lista.append(i)
return lista
# -
df_dep['nprecio']=df_dep['precio'].str.replace("MN ", "", regex=True).str.replace("$","",regex=False).str.replace(",","",regex=False)
df_dep['nprecio']=convertir(df_dep['nprecio'])
df_dep['nprecio']=pd.to_numeric(df_dep["nprecio"], errors="coerce")
df_dep['nprecio']=df_dep['nprecio'].replace(0,np.NaN)
lista1=df_dep['nprecio']
lista1=list(lista1)
lista1.sort()
print("Departamento más barato = ",df_dep['nprecio'].min())
print("Departamento más caro = ",df_dep['nprecio'].max())
for i in lista1:
print(i)
#moda1=
#mediana1=
#print("Moda = ",moda1)
#print("Mediana = ",mediana1)
df_dep_sc['nprecio']=df_dep_sc['precio'].str.replace("MN ", "", regex=True).str.replace("$","",regex=False).str.replace(",","",regex=False)
df_dep_sc['nprecio']=convertir(df_dep_sc['nprecio'])
df_dep_sc['nprecio']=pd.to_numeric(df_dep_sc["nprecio"], errors="coerce")
df_dep_sc['nprecio']=df_dep_sc['nprecio'].replace(0,np.NaN)
lista2=df_dep_sc['nprecio']
lista2.sort_values()
print('---Precios sin El Campanario---')
print("Departamento más barato = ",df_dep_sc['nprecio'].min())
print("Departamento más caro = ",df_dep_sc['nprecio'].max())
df_casa['nprecio']=df_casa['precio'].str.replace("MN ", "", regex=True).str.replace("$","",regex=False).str.replace(",","",regex=False)
df_casa['nprecio']=pd.to_numeric(df_casa["nprecio"], errors="coerce")
df_casa['nprecio']=df_casa['nprecio'].replace(0,np.NaN)
lista3=df_casa['nprecio']
lista3.sort_values()
print("Casa más barata = ",df_casa['nprecio'].min())
print("Casa más cara = ",df_casa['nprecio'].max())
df_casa_sc['nprecio']=df_casa_sc['precio'].str.replace("MN ", "", regex=True).str.replace("$","",regex=False).str.replace(",","",regex=False)
df_casa_sc['nprecio']=pd.to_numeric(df_casa_sc["nprecio"], errors="coerce")
df_casa_sc['nprecio']=df_casa_sc['nprecio'].replace(0,np.NaN)
lista4=df_casa_sc['nprecio']
lista4.sort_values()
print('Precio sin El Campanario')
print("Casa más barata = ",df_casa_sc['nprecio'].min())
print("Casa más cara = ",df_casa_sc['nprecio'].max())
|
medidas-tendencia-central.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Jug0nljfnzPe"
# # Cで整数を5バイトに押し込む簡単なお仕事
# -
# ビッグデータを処理するような場合、データの読み込みが律速になることが多いです。その場合、データのストレージ上のサイズが小さくなると読み込み時間が減り、全体の処理時間が半分になったりします。そんな時、整数が40bitで十分ならば、40bitで保持したいですね。というわけで、今まで[Go](https://www.soliton-cyber.com/blog/go-uint-40), [Rust](https://www.soliton-cyber.com/blog/rust-uint-40)とやってきましたが、処理効率を考えるとCと比較しないわけにはいきません。そこでCでも同様のことをやってみました。比較のため、clangとgccの両方を試します。
#
# * 以下のコードは全てリトルエンディアンを前提にしております。ビッグエンディアン下の場合は意図しない結果になります。ご注意ください。
# + [markdown] id="E4q2-0YydOYu"
# ## 64bit整数の8バイト配列化
# + [markdown] id="_FafCBqv84Lb"
# まずは愚直な方法
# + colab={"base_uri": "https://localhost:8080/"} id="lE2ajyy1rH5_" outputId="5cec2c2a-5d1f-4c65-e4fe-73fb80436627"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand();
}
uint8_t buf[SIZE][8];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
for (uint8_t j = 0; j < 8; j++) {
buf[idx][j] = v >> (8 * j);
}
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 8; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 8; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="Ae53coZjrHjm" outputId="f02f45e5-1707-43b5-992a-a3067d210779"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="mTyMJY9i8gxw"
# Goでは138秒、Rustでは8秒ですから、Goよりは速いですが、Rustより相当遅い。オプティマイザの性能の違いでしょうか。
# + [markdown] id="Gx6L-1QB9x4D"
# それでは定石通り、内側のfor文を展開しましょう。
# + colab={"base_uri": "https://localhost:8080/"} id="TZT4_VeQ3NT1" outputId="1fd9b1c7-d7df-4345-e362-ef2c03ed03a4"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand();
}
uint8_t buf[SIZE][8];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
uint8_t *p = (uint8_t*)&buf[idx];
p[0] = v;
p[1] = v >> 8;
p[2] = v >> 16;
p[3] = v >> 24;
p[4] = v >> 32;
p[5] = v >> 40;
p[6] = v >> 48;
p[7] = v >> 56;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 8; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 8; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="SvPW8hGCKFRn" outputId="0e31695c-198e-460b-98d3-520fcb3f48e1"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="jpdrzMQw-CTb"
# Goはこれで40秒にまで改善されましたが、Cでは改善されませんでした。
# + [markdown] id="6AzFHFJ4Daov"
# さて、Goではポインターを使うことで8秒、Rustではライブラリを使って4秒まで短縮されました。Cではどうでしょうか。ポインターを使ってみます。
# + colab={"base_uri": "https://localhost:8080/"} id="9S-sHhdwKH0z" outputId="5684a69c-4aeb-44af-a5b7-a0f6015d6605"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand();
}
uint8_t buf[SIZE][8];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
*(uint64_t*)&buf[idx] = vs[idx];
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 8; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 8; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="vyqr5eIPcclg" outputId="4f392ce4-b221-41b2-c532-1eedf1c68923"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="4NWaDNTyrBOm"
# Rustと同程度の速度です。ぎりぎりCの面目を保った感じです。
# + [markdown] id="fGBjk1tFnowv"
# ## 8バイト配列から64bit整数への変換
# + [markdown] id="WPkBvh-oFULO"
# まずは素朴な実装
# + colab={"base_uri": "https://localhost:8080/"} id="HNfasemcv368" outputId="d49d0707-79dd-49ed-aa09-cc81ff497b6a"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][8];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 8; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint8_t *b = (uint8_t*)&buf[idx];
uint64_t v = 0;
for (int j = 0; j < 8; j += 1) {
v += ((uint64_t)b[j] << (8 * j));
}
vs[idx] = v;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="QuM0e9lSv40N" outputId="e0706314-829e-4cf4-94b4-da27ea3904f9"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="WWukRLhZFjW9"
# Goで172秒、Rustで12秒ですから、微妙です。ここでclangとgccで違いが出ました。
# + [markdown] id="IZ92_CLSG1As"
# あまり期待できませんが、念の為、内側のfor文を展開します。
# + colab={"base_uri": "https://localhost:8080/"} id="aChheFBVF35c" outputId="bb12c394-1b02-4bf9-cba3-3830bf36f8e5"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][8];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 8; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint8_t *b = (uint8_t*)&buf[idx];
uint64_t v = (uint64_t)b[0];
v += (uint64_t)b[1] << 8;
v += (uint64_t)b[2] << 16;
v += (uint64_t)b[3] << 24;
v += (uint64_t)b[4] << 32;
v += (uint64_t)b[5] << 40;
v += (uint64_t)b[6] << 48;
v += (uint64_t)b[7] << 56;
vs[idx] = v;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="ylp-aj-dF4bL" outputId="833de4a0-14da-431b-a188-226cfcd794d4"
# !clang -Wall -Ofast main.c
# !./a.out
# !clang -Wall -Ofast main.c
# !./a.out
# + [markdown] id="9dYY06O2G-R1"
# やはり変わりません。ちなみにGoでは39秒、Rustでは19秒でした。Rustでは展開したほうが遅くなりました。と思ったら、gccの方が改善されています。gccではfor文の展開の最適化はなされないのでしょうか。
# + [markdown] id="CB-ZQe5SIGcq"
# それでは本命のポインターによる実装です。
# + colab={"base_uri": "https://localhost:8080/"} id="9vEaDnC0teSH" outputId="93398e4a-fed0-4e80-f85e-3369197d523f"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][8];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 8; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
vs[idx] = *(uint64_t*)&buf[idx];
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="HBfkWq9nvIfn" outputId="363553b4-200e-4d09-e9ef-ff8795be82c9"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="ygFOhkAtx15Y"
# さすがC。Goで13秒、Rustで6秒でしたから、相当速い。
#
# ただ、同様にポインターを使用したRustが遅いのが気になります。Rustの実装は、
# ```
# *(buf[idx].as_ptr() as *mut u64)
# ```
# なのですが、as_ptr()の呼び出しに何らかのコストがかかっていると思われます。なにかもっといい方法があるかもしれません。
#
# + [markdown] id="-fMdlA0ayncR"
# ## 40bit整数の5バイト配列化
# + [markdown] id="G64mg0aaPZj8"
# まずは素朴な実装
# + colab={"base_uri": "https://localhost:8080/"} id="G8eHktAyzcfG" outputId="724fed5a-ea1e-4cc2-dd03-40e48de09ab0"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE][5];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
uint8_t *b = (uint8_t*)buf[idx];
for (uint8_t j = 0; j < 5; j++) {
b[j] = v >> (8 * j);
}
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 5; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="eWNMDxsXzcTp" outputId="1a96944a-5f68-4629-e2cf-e568deda47ea"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="qvSKao-8PfC7"
# 8バイトの時は34秒でしたから、5バイトに減った分だけ順当に短縮されています。ちなみにGoでは90秒、Rustでは22秒でした。
# + [markdown] id="fMIXw7CHJmDr"
# 次に内側のfor文の展開
# + colab={"base_uri": "https://localhost:8080/"} id="YwyJVzqLITyQ" outputId="0992f43b-f395-4cb4-b261-4c06de051333"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE][5];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
uint8_t *b = (uint8_t*)buf[idx];
b[0] = v;
b[1] = v << 8;
b[2] = v << 16;
b[3] = v << 24;
b[4] = v << 32;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 5; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="tlFMkaMPITnT" outputId="c448ea3e-f4fc-4909-cab9-380a2c5878ed"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="VpATg0yNJ2EA"
# こちらはclangもgccも速くなりました。最適化でどのような時にfor文の展開が行われるのか予測が難しい。
# + [markdown] id="_52y57pOSY2g"
# それではmemcpy()を使ってみましょう。
# + colab={"base_uri": "https://localhost:8080/"} id="PR9KZKQy0hux" outputId="884c1900-2d72-49a7-bb72-a6e981fca02d"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#include <string.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE][5];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
memcpy((void*)&buf[idx], (void*)&vs[idx], 5);
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 5; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="WhwRGugm0hpC" outputId="232c69b6-9707-4c19-d5c7-cacad44ef487"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="L1m2lpD6S3J1"
# for文を展開した時や、Rustで関数を利用した時と同程度になりました。Rustでも関数内部でmemcpy()を使っているのかもしれません。
# ちなみにGoの最速も同程度です。
# + [markdown] id="j1iKbNyTUKF8"
# 次に、5バイトを4バイトと1バイトに分けて複製する方法を実装します。4バイトをuint32_tとして一命令で複製できるのが強みです。ただ、このコードはリトルエンディアンが前提なので、ご注意ください。
# + colab={"base_uri": "https://localhost:8080/"} id="OXmggqQU3OaZ" outputId="54fb3bb3-c7fd-443e-eabf-ead928b1a4f6"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#include <string.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE][5];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
uint8_t *b = (uint8_t*)buf[idx];
*(uint32_t*)b = *(uint32_t*)&v;
b[4] = ((uint8_t*)&v)[4];
//b[4] = (uint8_t)(v >> 32);
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 5; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="j8C7zfCt3OTj" outputId="c151b5f5-3b3c-489c-a761-7e7beecd7409"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="KfE9WdJbU3DB"
# for文の展開や同様の処理をRustで実装した場合と同程度です。ただ、64bit整数の8バイト配列化が4秒程度ですから、cpuネイティブのビット幅と異なる整数の取り扱いにはそれなりのコストがかかることが分かります。それについにgccに警告を出されてしまいました。意図的にポイント先の整数をバラしているのですから、当然ですね。この辺りがリトルエンディアン限定である所以でもあります。処理速度的にもこれを採用する意味はないです。
# + [markdown] id="FXMv_7J4yPoh"
# ## 5バイト配列から40bit整数への変換
# + [markdown] id="tQxI4NvdYMsH"
# 素朴な方法
# + colab={"base_uri": "https://localhost:8080/"} id="_yR6If2vOCc-" outputId="7b8d1e71-bb35-43ca-aabf-ee57c6ed62e3"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][5];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 5; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint8_t *b = (uint8_t*)&buf[idx];
uint64_t v = b[0];
for (size_t j = 1; j < 5; j++) {
v += (uint64_t)(b[j]) << (8 * j);
}
vs[idx] = v;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="fZWasSRQOCQ_" outputId="e983fc15-7e07-4666-cfb8-429258f1a80b"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="1dG53yy1Ovcy"
# 内側のfor文の展開
# + colab={"base_uri": "https://localhost:8080/"} id="Rcog0MlR5gQI" outputId="d895b1bf-2c74-46c7-ce39-23b39a1752e6"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][5];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 5; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint8_t *b = (uint8_t*)&buf[idx];
uint64_t v = b[0];
v += (uint64_t)(b[1]) << 8;
v += (uint64_t)(b[2]) << 16;
v += (uint64_t)(b[3]) << 24;
v += (uint64_t)(b[4]) << 32;
vs[idx] = v;
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="dsKtyLcJ5h87" outputId="09f67699-14de-40ac-deab-2f78e8a276fd"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="LkXLiQA8WpAy"
# for文展開しても同じでした。8バイトの時が23秒でしたから、それなりに短縮されています。Goでは24秒、Rustでは10秒でした。
# + [markdown] id="hCKOMJ0bYY2G"
# 次に、uint64_tで取得してから0xFFFFFFFFFFでマスクすることで5バイト化する方法です。
# + colab={"base_uri": "https://localhost:8080/"} id="OgXLDPCp65Yb" outputId="8a9105c8-d2f5-46de-ac8e-54c7ebd5d7fe"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][5];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 5; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
vs[idx] = ((*(uint64_t*)&buf[idx]) & 0xFFFFFFFFFF);
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="jmUM02A765Uy" outputId="ce4fd19e-d493-4022-94a4-3c1a7b674da5"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="GjUGJJXAY1m_"
# 相当速い。ただ、64bitでは4秒でしたので、マスク処理にそれなりのコストがかかっています。ちなみに、Goの最速が14秒、Rustが7秒です。
# + [markdown] id="42SiOnqDZs5j"
# マスクではなく、4バイトと1バイトに分ける方法を試してみます。
# + colab={"base_uri": "https://localhost:8080/"} id="iBr2fZKG7n6p" outputId="f1b8729f-f5d6-4193-8b9c-38a1eef97c72"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][5];
srand(0);
for (int i = 0; i < SIZE; i += 1) {
for (int j = 0; j < 5; j += 1) {
buf[i][j] = (uint8_t)rand();
}
}
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint8_t *b = buf[idx];
vs[idx] = (*(uint32_t*)b) + (((uint64_t)b[4]) << 32);
}
}
end_clock = clock();
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="n39uWojC7n17" outputId="4f081743-920c-46c7-bfb9-b6524cc76275"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="wFB5TMVbZ7CH"
# 遅くなりました。この処理ではマスクの方が有効です。
#
# それとこのコードではgccが警告を出していません。uint8_tへのポインターをuint32_tのポインターに差し替えているので、当然警告が出ると思ったのですが、コンパイラーが検出できる限界を超えたようです。
# + [markdown] id="BloLkQIYyVhG"
# ## 40bit整数の配列の5Nバイト配列化
# + [markdown] id="dpa9W3gebEHD"
# 10万個の整数を5バイトずつのバイト配列にします。まずは素朴な実装。
# + colab={"base_uri": "https://localhost:8080/"} id="oN0rwxX_ctt5" outputId="bce6fbd3-6e4a-4e3d-b857-f1dd487600ca"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE * 5];
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx< SIZE; idx++) {
uint64_t v = vs[idx];
size_t idx5 = idx * 5;
uint8_t *b = (uint8_t*)(buf + idx5);
for (uint8_t j = 0; j < 5; j++) {
b[j] = v >> (8 * j);
}
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE * 5; i += 1) {
total += buf[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="JNbb-0DCc56g" outputId="bbf4f8de-9fb2-43bf-8bbd-2d822f0db276"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="FPQst9zjbN_w"
# Goが104秒、Rustが21秒ですから、Rustと同等程度です。
# + [markdown] id="TQDMoBrFSkHG"
# 次にfor文の展開
# + colab={"base_uri": "https://localhost:8080/"} id="G9Ecuc9dysqd" outputId="b2418530-833d-4126-bc73-f97057062e38"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE * 5];
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx< SIZE; idx++) {
uint64_t v = vs[idx];
size_t idx5 = idx * 5;
uint8_t *b = (uint8_t*)(buf + idx5);
b[0] = v;
b[1] = v >> 8;
b[2] = v >> 16;
b[3] = v >> 24;
b[4] = v >> 32;
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE * 5; i += 1) {
total += buf[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="jHAzeC7yysZL" outputId="a8cdb1ca-4e0f-471f-dff6-16ba5ce9629a"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="FBwDoHm0yz5h"
# Goが16秒、Rustが21秒ですから、Rustと同程度ですが、Goより遅い。CやRustといえど工夫無くしてはGoに負けることがあるようです。
# + [markdown] id="c5rfhgbqcuji"
# 次に下図の戦略での実装。
# + [markdown] id="KpD5B_0cb7hR"
# <image width=500 src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAABAYAAAFeCAYAAAAWvx3kAAAgAElEQVR4Ae29ifMdw/7/HwqpiOJSIglCSGKXUASxE4klsS8XcXEjqyx2EbF85HJFItZCIgjha7nXmpKIoCIEJYK/6Ffzq+e51dGnp2dOn3PendPvmce76l2z9fT069Gv6fPq13S/esAA/iAAAQhAAAIQgAAEIAABCEAAAhCAAAQgAAEIQAACEIAABCAAAQhAAAIQgAAEIAABCEAAAhCAAAQgAAEIQAACEIAABCAAAQhAAAIQgAAEIAABCEAAAhCoFYE///<KEY>AAE<KEY>ABC<KEY>EIAABCEAAAhCAAAQgAAEIQAACEIAABCAAAQhAAAIQgAAEIAABCEAAAhCAAAQgAAEIQAACE<KEY>">
# + colab={"base_uri": "https://localhost:8080/"} id="nVzWZs9mfKYb" outputId="6cdc4ace-d867-469a-c7d7-79b5eed3f64e"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand() & 0xFFFFFFFFFF;
}
uint8_t buf[SIZE * 5 + 3];
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
*(uint64_t*)(buf + idx * 5) = vs[idx];
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 5; i++) printf("%d ", buf[i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE * 5; i += 1) {
total += buf[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="HDNmO65FfKSs" outputId="91a50e21-0da8-478b-9840-df52a684c754"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="-eKt0G3hc9Z8"
# Goが12秒、Rustが8秒ですので、Rustより少し速い程度です。
# + [markdown] id="kWKlHa_dyVcX"
# ### 5Nバイト配列から40bit整数の配列への変換
# + [markdown] id="Z7avANO_ds4G"
# 5バイトずつのバイト配列から10万個の整数を読み出します。素朴な実装は以下になります。
# + colab={"base_uri": "https://localhost:8080/"} id="fkPedFtEUB-D" outputId="a25d9b14-20e3-4937-92fc-17bfcd9a65a4"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE*5];
srand(0);
for (int i = 0; i < SIZE*5; i += 1) {
buf[i] = (uint8_t)rand();
}
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx += 1) {
size_t idx5 = idx * 5;
uint8_t *b = buf + idx5;
uint64_t *v = vs + idx;
*v = 0;
for (size_t j = 0; j < 5; j++) {
*v += (uint64_t)(b[j]) << (8 * j);
}
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="D0_RirCEUBu9" outputId="67ff25d2-7266-4294-dc48-4b09e2589c8d"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="9MGxcCOvUmpv"
# 内側のfor文の展開
# + colab={"base_uri": "https://localhost:8080/"} id="hgYjDdaivK8e" outputId="586d2e9f-3cbc-4ad2-b467-9530b5c647ac"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE*5];
srand(0);
for (int i = 0; i < SIZE*5; i += 1) {
buf[i] = (uint8_t)rand();
}
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx += 1) {
size_t idx5 = idx * 5;
uint8_t *b = buf + idx5;
uint64_t *v = vs + idx;
*v = b[0];
*v += (uint64_t)(b[1]) << 8;
*v += (uint64_t)(b[2]) << 16;
*v += (uint64_t)(b[3]) << 24;
*v += (uint64_t)(b[4]) << 32;
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="JBbMYMx8h1Tk" outputId="b987b2d7-794f-493c-e061-5b82534c6b91"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="iraleSJde9xm"
# Goが23秒、Rustが8秒ですので、間ぐらい。また、このコードでは、最適化によりfor文の展開が行われたようです。
# + [markdown] id="6blzZS9dhI3o"
# 0xFFFFFFFFFFによるマスクの方法。
# + colab={"base_uri": "https://localhost:8080/"} id="gP8Rd3_YkHdE" outputId="318b7f73-c1d2-4d07-fa58-0784a34b4067"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE*5];
srand(0);
for (int i = 0; i < SIZE*5; i += 1) {
buf[i] = (uint8_t)rand();
}
// 測定開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx += 1) {
vs[idx] = *(uint64_t*)(buf + idx * 5) & 0xFFFFFFFFFF;
}
}
end_clock = clock();
// 測定終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
printf("%lu\n", vs[0]);
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="62sER7l5kHUN" outputId="497fca5d-33e5-4d5a-9555-299bf957cb29"
# !clang -Wall -Ofast main.c
# !./a.out
# !gcc -Wall -Ofast main.c
# !./a.out
# + [markdown] id="mQGHjznFhccm"
# Goの最速が12秒、Rustが6.4秒ですから、Cでclangを使った場合が一番速くなりました。
# + [markdown] id="EuHAPQpklySB"
# ## 結論
#
# 整数を5バイトに押し込む簡単なお仕事に関してはCとRustは同程度、Goは約半分の速度。
#
# for文の展開はオプティマイザ任せにせずに自分でやっておいた方が無難。
#
# strict-aliasing rulesの違反は取り扱い注意だが、そこまで踏み込まないとRustに負ける。(といってもRustでもunsafe使ってるけど)
# + [markdown] id="kLXp8UdrjPwj"
# ## 追記
#
# コメントにてコンパイラーの最適化を甘くみると痛い目にあうよというご指摘があったので、確かめてみます。
# + [markdown] id="WlReWIxzkQCh"
# colabに最初から用意されているgccは最新版ではなかったので、まずはgcc-11をインストールします。
# + id="ohitB20obbek"
# !add-apt-repository 'deb http://mirrors.kernel.org/ubuntu hirsute main universe'
# !apt update
# !apt-get install gcc-11
# + [markdown] id="au-phWDBkVvk"
# 次に一番最初の素朴な実装
# + colab={"base_uri": "https://localhost:8080/"} id="-nmuGzx9budg" outputId="a4135e32-ef8f-48f2-9300-eaf7b8b7aa24"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand();
}
uint8_t buf[SIZE][8];
// 計測開始
clock_t start_clock, end_clock;
start_clock = clock();
for (size_t i = 0; i < COUNT/SIZE; i++) {
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
for (uint8_t j = 0; j < 8; j++) {
buf[idx][j] = v >> (8 * j);
}
}
}
end_clock = clock();
// 計測終了
printf("%f sec\n", (double)(end_clock - start_clock) / CLOCKS_PER_SEC);
for (int i = 0; i < 8; i++) printf("%d ", buf[0][i]);
printf("\n");
uint64_t total = 0;
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 8; j += 1) {
total += buf[i][j];
}
}
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="HlVwtf3fjiFT" outputId="9a9cd020-f990-4e8b-9cb8-73d63400fece"
# !gcc-11 -Wall -Ofast main.c
# !./a.out
# + [markdown] id="bjEmelIkkcRN"
# 確かに! 最後のルーブのbufしか使っていないのが最新のコンパイラーに見抜かれてしまったようです。
# + [markdown] id="fZsmXGVwk4Tv"
# より慎重なのはこんなコードでしょうか。内側のループでvsの値が毎回異なるようにし、さらにvsの中身が乱数だということを悟られにくいように関数として切り出しました。そして全てのbufの値にアクセスして、使ってることをアピール。
# + colab={"base_uri": "https://localhost:8080/"} id="dV46fLX8o3qM" outputId="24e975b6-44fb-417a-9b7b-861508db62e5"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
clock_t measure(const uint64_t vs[], uint8_t buf[][8]) {
clock_t start_clock = clock();
for (size_t idx = 0; idx < SIZE; idx++) {
uint64_t v = vs[idx];
for (uint8_t j = 0; j < 8; j++) {
buf[idx][j] = v >> (8 * j);
}
}
return clock() - start_clock;
}
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE][8];
uint64_t total = 0;
srand(0);
clock_t total_clock = 0;
for (size_t n = 0; n < COUNT/SIZE; n++) {
for (int i = 0; i < SIZE*5; i += 1) {
buf[i] = (uint8_t)rand();
}
total_clock += measure(vs, buf);
for (size_t i = 0; i < SIZE; i += 1) {
for (size_t j = 0; j < 8; j += 1) {
total += buf[i][j];
}
}
}
printf("%f sec\n", (double)total_clock / CLOCKS_PER_SEC);
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="lAuy9s4Ajgw7" outputId="c52170db-2b57-44a5-a667-21153a6c04e6"
# %%time
# !gcc-11 -Wall -Ofast main.c
# !./a.out
# + [markdown] id="ARM-SmqMlzFK"
# それらしい計測値が得られました。
#
# 実務で採用する場合は、意図しない最適化の影響を避けるため、関数化して実データを流し込んで比較する必要がありそうです。
# + [markdown] id="yHq6NnciwBNu"
# 全てを関数化すると記事が長くなり過ぎてしまうので、最後のコードだけ書き換えてみました。
# + colab={"base_uri": "https://localhost:8080/"} id="ukA2helxtd_K" outputId="d24a8ac0-84f3-4871-802f-9487f55dfa6e"
# %%writefile main.c
#include <stdio.h>
#include <stdlib.h>
#include <stdint.h>
#include <time.h>
#define COUNT 10000000000
#define SIZE 100000
clock_t measure(uint64_t vs[], const uint8_t buf[]) {
clock_t start_clock = clock();
for (size_t idx = 0; idx < SIZE; idx += 1) {
vs[idx] = *(uint64_t*)(buf + idx * 5) & 0xFFFFFFFFFF;
}
return clock() - start_clock;
}
int main(int argc, char **argv)
{
uint64_t vs[SIZE];
uint8_t buf[SIZE*5];
uint64_t total = 0;
srand(0);
clock_t total_clock = 0;
for (size_t n = 0; n < COUNT/SIZE; n++) {
for (int i = 0; i < SIZE; i += 1) {
vs[i] = rand();
}
total_clock += measure(vs, buf);
for (size_t i = 0; i < SIZE; i += 1) {
total += vs[i];
}
}
printf("%f sec\n", (double)total_clock / CLOCKS_PER_SEC);
printf("%lu\n", total);
return 0;
}
# + colab={"base_uri": "https://localhost:8080/"} id="eofEObNpuLsG" outputId="2ee9002a-c148-4a9f-d114-417bd75d7fe4"
# %%time
# !gcc-11 -Wall -Ofast main.c
# !./a.out
# + [markdown] id="6EDt1mS0wORi"
# 細切れの時間計測を合計するので、誤差が増幅している可能性もあります。処理速度計測は一筋縄ではいきませんね。
|
CUint40.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # King County Real Estate
# ## Designing Data-Driven Price Model
#
# ###### <font color='gray'>By: <NAME>, <NAME>, <NAME></font>
# # Overview
# **This project analyzes King County housing sales data to create both predictive and inferential price model**
#
# The project highlights four key features that most strongly impacts our housing price:
# - Square feet
# - Zip code
# - Water Front
# - View
# # Business Problem
# As a real estate agency firm, their primary role is to connect potential buyers and sellers of real estate properties. It is therefore imperative that they provide an accurate information and services to their clients and ensure both parties consent on a fair property price. Every Door Real Estate Company needs a model that provides statistically significant information on features that impact housing value and predicts a fair price based on those features and will make ssure the client is content on their decision of choosing Every Door Real Estate Company.
#
# Key questions:
# - Which specific features significantly influence the overall value of the house?
# - How significant are the locations of the house within King County?
# - Is it possible to predict the housing price based on its features?
# - How can Every Door Real Estate Company utilize this model to increase their profit?
# ----
# # Data Preparation
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn.preprocessing import OneHotEncoder
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from statsmodels.formula.api import ols
from sklearn.impute import SimpleImputer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import OrdinalEncoder
from sklearn.dummy import DummyRegressor
import matplotlib.pyplot as plt
import statsmodels.api as sm
# %matplotlib inline
# Loading in the given data set into variable housing_df to check the data type for each column
housing_df = pd.read_csv('data/kc_house_data.csv')
housing_df.info()
# Checking to see if any columns in the dataframe have any NaN values
housing_df.isna().sum()
# **Observations**
# <ul>
# <li> We see waterfront, view and yr_renocated columns are the only columns with NaN values.
# <li> We will be using sklearn Simple Imputer to later fill in the missing values in our desired columns.
# <li> Also there are many columns with non-numerical values, so we will convert those values into integers using an appropriate sklearn tools.
# Grabbing the age of the home
housing_df['datetime'] = pd.to_datetime(housing_df['date'])
housing_df['soldyear'] = housing_df['datetime'].dt.year
housing_df['age_when_sold'] = housing_df['soldyear'] - housing_df['yr_built']
# Just getting the numeric values of Graade
housing_df['grade_num'] = housing_df['grade'].str.split()
housing_df['grade_num'] = housing_df['grade_num'].str[0]
housing_df['grade_num'] = housing_df['grade_num'].astype(int)
# Creating a new column 'Basement' that will have a value of 1 or 0 in reference to whther a house has a basement or not.
housing_df['sqft_basement'] = housing_df['sqft_living'] - housing_df['sqft_above']
housing_df['Basement'] = None
housing_df['Basement'] = housing_df['sqft_basement'].map(lambda x: False if x == 0 else True)
housing_df['Basement'] = housing_df['Basement'].astype(int)
housing_df.condition.value_counts()
# Changing all the values in the 'condition' column to integers.
replace_dict2 = {'Poor': 1, 'Fair': 2, 'Average': 3, 'Good': 4, 'Very Good': 5}
housing_df['condition'] = housing_df['condition'].replace(replace_dict2)
wtr_col = housing_df[['waterfront']]
imputer = SimpleImputer(strategy='constant', fill_value = 'NO')
imputer.fit(wtr_col)
waterfront_imputed = imputer.transform(wtr_col)
housing_df.waterfront = waterfront_imputed
wtr_col = housing_df[['waterfront']]
encoder_wtr = OrdinalEncoder()
encoder_wtr.fit(wtr_col)
encoded_wtr = encoder_wtr.transform(wtr_col)
encoded_wtr = encoded_wtr.flatten()
housing_df.waterfront = encoded_wtr
view_col = housing_df[['view']]
imputer = SimpleImputer(strategy='constant', fill_value = 'NONE')
imputer.fit(view_col)
view_imputed = imputer.transform(view_col)
housing_df.view = view_imputed
housing_df.isna().sum()
# We don't see much usage in certain columns or we have already used the column information to its max and will no longer be needing them.
housing_ols = housing_df.drop(['datetime', 'date', 'soldyear', 'yr_built', 'lat', 'long',
'sqft_above', 'sqft_lot15', 'sqft_living15', 'grade', 'sqft_above', 'sqft_basement', 'yr_renovated'], axis = 1)
housing_ols
housing_ols['bathrooms'] = housing_ols['bathrooms'][housing_ols['bathrooms'].between(housing_ols['bathrooms'].quantile(.025), housing_ols['bathrooms'].quantile(.975), inclusive=True)]
housing_ols['floors'] = housing_ols['floors'][housing_ols['floors'].between(housing_ols['floors'].quantile(.025), housing_ols['floors'].quantile(.975), inclusive=True)]
housing_ols['condition'] = housing_ols['condition'][housing_ols['condition'].between(housing_ols['condition'].quantile(.025), housing_ols['condition'].quantile(.975), inclusive=True)]
housing_ols['age_when_sold'] = housing_ols['age_when_sold'][housing_ols['age_when_sold'].between(housing_ols['age_when_sold'].quantile(.025), housing_ols['age_when_sold'].quantile(.975), inclusive=True)]
housing_ols['grade_num'] = housing_ols['grade_num'][housing_ols['grade_num'].between(housing_ols['grade_num'].quantile(.025), housing_ols['grade_num'].quantile(.975), inclusive=True)]
housing_ols['sqft_living'] = housing_ols['sqft_living'][housing_ols['sqft_living'].between(housing_ols['sqft_living'].quantile(.025), housing_ols['sqft_living'].quantile(.975), inclusive=True)]
housing_ols.dropna(inplace=True)
housing_ols.info()
# With new working dataframe we want to see how the remaining columns correlates to our target,price, as well as to other columns to check for collinearity
housing_ols.corr()
# Visually inspecting all columns association with price
fig, (axes1, axes2, axes3, axes4) = plt.subplots(nrows=4, ncols=3, figsize=(15,15))
for xcol, ax in zip(['bedrooms', 'bathrooms', 'sqft_living'], axes1):
housing_ols.plot(kind='scatter', x=xcol, y='price', ax=ax, alpha=0.4, color='b')
for xcols, axs in zip(['sqft_lot', 'floors', 'view'], axes2):
housing_ols.plot(kind='scatter', x=xcols, y='price', ax=axs, alpha=0.4, color='b')
for xcolss, axss in zip(['condition', 'age_when_sold', 'grade_num'], axes3):
housing_ols.plot(kind='scatter', x=xcolss, y='price', ax=axss, alpha=0.4, color='b')
for xcolss, axss in zip(['waterfront', 'Basement'], axes4):
housing_ols.plot(kind='scatter', x=xcolss, y='price', ax=axss, alpha=0.4, color='b')
housing_pred = housing_ols.copy()
# +
# All of these columns are categorical values so we will we will create separate columns with OneHotEncoder.
# Using the new columns found within each categories, we will concat them all into one dataframe.
# Categorical columns include: 'bedrooms', 'grade_num', 'condition', 'bahrooms', 'view', 'floors', 'zipcode'
bedrooms_ohe = housing_pred[['bedrooms']]
ohe_bedrooms = OneHotEncoder(categories ='auto', sparse =False)
ohe_bedrooms.fit(bedrooms_ohe)
ohe_bedrooms_encoded = ohe_bedrooms.transform(bedrooms_ohe)
bedrooms_encoded_ohe = pd.DataFrame(ohe_bedrooms_encoded, columns = ohe_bedrooms.get_feature_names(['bedrooms']), index = housing_pred.index )
housing_pred1 = pd.concat([housing_pred, bedrooms_encoded_ohe ], axis =1)
# -
grade_num_ohe = housing_pred[['grade_num']]
ohe_grade_num = OneHotEncoder(categories ='auto', sparse =False)
ohe_grade_num.fit(grade_num_ohe)
ohe_grade_num_encoded = ohe_grade_num.transform(grade_num_ohe)
grade_num_encoded_ohe = pd.DataFrame(ohe_grade_num_encoded, columns = ohe_grade_num.get_feature_names(['grade_num']), index = housing_pred.index )
housing_pred2 = pd.concat([housing_pred1, grade_num_encoded_ohe], axis =1)
condition_ohe = housing_pred[['condition']]
ohe = OneHotEncoder(categories="auto", sparse=False)
cond_encoded_ohe = pd.DataFrame (ohe.fit_transform(condition_ohe), index = housing_pred.index)
cond_encoded_ohe.columns = ohe.get_feature_names(['condition'])
housing_pred3 = pd.concat([housing_pred2, cond_encoded_ohe], axis = 1)
bathrooms_ohe = housing_pred[['bathrooms']]
ohe = OneHotEncoder(categories='auto', sparse=False, handle_unknown='ignore')
bathrooms_transform = ohe.fit_transform(bathrooms_ohe)
bathrooms_encoded_ohe = pd.DataFrame(bathrooms_transform, columns=ohe.get_feature_names(['bathrooms']), index=housing_pred.index)
housing_pred4 = pd.concat([housing_pred3, bathrooms_encoded_ohe], axis = 1)
view_ohe = housing_pred[['view']]
ohe = OneHotEncoder(categories="auto", sparse=False)
ohe.fit(view_ohe)
view_encoded = ohe.transform(view_ohe)
view_encoded_ohe =pd.DataFrame(view_encoded, columns=ohe.get_feature_names(['view']), index=housing_pred.index)
housing_pred5 = pd.concat([housing_pred4, view_encoded_ohe], axis = 1)
floors_ohe = housing_pred[['floors']]
ohe_floors = OneHotEncoder(categories ='auto', sparse =False)
ohe_floors.fit(floors_ohe)
ohe_floors_encoded = ohe_floors.transform(floors_ohe)
floors_encoded_ohe = pd.DataFrame(ohe_floors_encoded, columns = ohe_floors.get_feature_names(['floors']), index = housing_pred.index )
housing_pred6 = pd.concat([housing_pred5, floors_encoded_ohe ], axis =1)
zipcode_ohe = housing_pred[['zipcode']]
ohe = OneHotEncoder(categories="auto", sparse=False)
ohe.fit(zipcode_ohe)
zipcode_encoded = ohe.transform(zipcode_ohe)
zipcode_encoded_ohe =pd.DataFrame(zipcode_encoded, columns=ohe.get_feature_names(['zipcode']), index=housing_pred.index)
housing_pred_final = pd.concat([housing_pred6, zipcode_encoded_ohe ], axis =1)
# Final dataframe after dealing with the catgoricals and dropping the original columns.
housing_pred_final.drop(['id', 'floors', 'bedrooms', 'bathrooms', 'view', 'condition', 'zipcode', 'grade_num'], axis = 1, inplace=True)
# Finally we want to log transform our target variable, price, in order to account for heteroscesdsitdy in the data
# Creating a new column that takes the log of price
housing_pred_final['log_price'] = np.log(housing_pred_final['price'])
# # Predictive Model Results
# ### Dummy Regressor (Baseline)
# +
# Putting our target in the y_dummy variable and the rest of the dataframe without price in the X_dummy variable
X_dummy = housing_pred_final.drop('price', axis = 1)
y_dummy = housing_pred_final['log_price']
# Splitting up our dummy variables into two subsets: for training data and testing data
X_train, X_test, y_train, y_test = train_test_split(X_dummy, y_dummy, test_size=0.2, random_state=42)
# Setting up our dummy regressor
dummy_regr = DummyRegressor(strategy="mean")
dummy_regr.fit(X_train, y_train)
print(dummy_regr.score(X_train, y_train))
print(dummy_regr.score(X_test, y_test))
# -
# ### Simple Regression
heat_map = housing_pred.corr()
heat_map_price = heat_map[['price']]
sns.heatmap(heat_map_price, annot=True, linewidths =2.5);
# +
heat_map_price = heat_map_price.reset_index().sort_values(by='price', ascending=True)
heat_map_price = heat_map_price[(heat_map_price['index']=='sqft_living')|
(heat_map_price['index']=='grade_num')|
(heat_map_price['index']=='bathrooms')|
(heat_map_price['index']=='bedrooms')|
(heat_map_price['index']=='waterfront')|
(heat_map_price['index']=='floors')]
heat_fig, ax = plt.subplots(figsize=(10,6))
sns.set_style("dark")
x_cor = heat_map_price['index']
y_cor = heat_map_price['price']
clrs=['grey' if (x != 'sqft_living') else 'red' for x in x_cor]
sns.barplot(x=x_cor, y=y_cor, palette=clrs, ax=ax)
ax.tick_params(axis='x', rotation=45)
# -
# **We see from above that sqft_living has the highest correlation from non-catergorical to price therefore it will be used as the feature in the simple model.**
# +
# y variable will always be our target which is price and X variable will contain our housing features
# Here in our simple model, X is only sqft_living
simple_model_df = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living']], axis = 1)
X_simple = simple_model_df.drop('log_price', axis = 1)
y_simple = simple_model_df['log_price']
# Splitting up our dummy variables into two subsets: for training data and testing data
X_train_s, X_test_s, y_train_s, y_test_s = train_test_split(X_simple, y_simple, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_s[cols_scale])
scaled_columns = scaled_fit.transform(X_train_s[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_s.index)
scaled_columns.columns = cols_scale
X_train_s['sqft_living'] = scaled_columns['sqft_living']
# Setting up simple linear regression
simple_reg = LinearRegression()
simple_reg.fit(X_train_s, y_train_s)
print(simple_reg.score(X_train_s, y_train_s))
# Scaling test set
scale_test = X_test_s[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_s.index)
scaled_columns_test.columns = cols_scale
X_test_s['sqft_living'] = scaled_columns_test['sqft_living']
print(simple_reg.score(X_test_s, y_test_s))
# Calculating predicted value of our model
# determine the mean squared error for train and test data
y_hat_train_s = np.exp(simple_reg.predict(X_train_s))
y_hat_test_s = np.exp(simple_reg.predict(X_test_s))
train_mse_s = mean_squared_error(np.exp(y_train_s), y_hat_train_s)
test_mse_s = mean_squared_error(np.exp(y_test_s), y_hat_test_s)
test_rmse_s = np.sqrt(test_mse_s)
print('Train Mean Squarred Error:', train_mse_s)
print('Test Mean Squarred Error:', test_mse_s)
print('Test RMSE: ', test_rmse_s)
# -
# ### Multiple Linear Regression
# #### Model 1
# +
# y variable will always be our target which is price and X variable will contain our housing features
# This is our first muliple linear regression model with just sqft_living and view
multi_model_1 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], view_encoded_ohe], axis = 1)
X_multi1 = multi_model_1.drop('log_price', axis = 1)
y_multi1 = multi_model_1['log_price']
# Splitting up our dummy variables into two subsets: for training data and testing data
X_train_m1, X_test_m1, y_train_m1, y_test_m1 = train_test_split(X_multi1, y_multi1, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m1[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m1[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m1.index)
scaled_columns.columns = cols_scale
X_train_m1['sqft_living'] = scaled_columns['sqft_living']
# Setting up our first multiple linear regression
multi1_reg = LinearRegression()
multi1_reg.fit(X_train_m1, y_train_m1)
print(multi1_reg.score(X_train_m1, y_train_m1))
# Scaling test set
scale_test = X_test_m1[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m1.index)
scaled_columns_test.columns = cols_scale
X_test_m1['sqft_living'] = scaled_columns_test['sqft_living']
print(multi1_reg.score(X_test_m1, y_test_m1))
# Calculating predicted value of our model
# determine the mean squared error for train and test data
y_hat_train_m1 = np.exp(multi1_reg.predict(X_train_m1))
y_hat_test_m1 = np.exp(multi1_reg.predict(X_test_m1))
train_mse_m1 = mean_squared_error(np.exp(y_train_m1), y_hat_train_m1)
test_mse_m1 = mean_squared_error(np.exp(y_test_m1), y_hat_test_m1)
test_rmse_m1 = np.sqrt(test_mse_m1)
print('Train Mean Squarred Error:', train_mse_m1)
print('Test Mean Squarred Error:', test_mse_m1)
print('Test RSME: ', test_rmse_m1)
# -
# #### Model 2
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 2
multi_model_2 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], view_encoded_ohe, cond_encoded_ohe], axis = 1)
X_multi2 = multi_model_2.drop('log_price', axis = 1)
y_multi2 = multi_model_2['log_price']
X_train_m2, X_test_m2, y_train_m2, y_test_m2 = train_test_split(X_multi2, y_multi2, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m2[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m2[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m2.index)
scaled_columns.columns = cols_scale
X_train_m2['sqft_living'] = scaled_columns['sqft_living']
multi2_reg = LinearRegression()
multi2_reg.fit(X_train_m2, y_train_m2)
print(multi2_reg.score(X_train_m2, y_train_m2))
# Scaling test set
scale_test = X_test_m2[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m2.index)
scaled_columns_test.columns = cols_scale
X_test_m2['sqft_living'] = scaled_columns_test['sqft_living']
print(multi2_reg.score(X_test_m2, y_test_m2))
y_hat_train_m2 = multi2_reg.predict(X_train_m2)
y_hat_test_m2 = multi2_reg.predict(X_test_m2)
train_mse_m2 = mean_squared_error(y_train_m2, y_hat_train_m2)
test_mse_m2 = mean_squared_error(y_test_m2, y_hat_test_m2)
print('Train Mean Squarred Error:', train_mse_m2)
print('Test Mean Squarred Error:', test_mse_m2)
# -
# #### Model 3
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 3
multi_model_3 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], zipcode_encoded_ohe], axis = 1)
X_multi3 = multi_model_3.drop('log_price', axis = 1)
y_multi3 = multi_model_3['log_price']
X_train_m3, X_test_m3, y_train_m3, y_test_m3 = train_test_split(X_multi3, y_multi3, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m3[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m3[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m3.index)
scaled_columns.columns = cols_scale
X_train_m3['sqft_living'] = scaled_columns['sqft_living']
multi3_reg = LinearRegression()
multi3_reg.fit(X_train_m3, y_train_m3)
print(multi3_reg.score(X_train_m3, y_train_m3))
# Scaling test set
scale_test = X_test_m3[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m3.index)
scaled_columns_test.columns = cols_scale
X_test_m3['sqft_living'] = scaled_columns_test['sqft_living']
print(multi3_reg.score(X_test_m3, y_test_m3))
y_hat_train_m3 = multi3_reg.predict(X_train_m3)
y_hat_test_m3 = multi3_reg.predict(X_test_m3)
train_mse_m3 = mean_squared_error(y_train_m3, y_hat_train_m3)
test_mse_m3 = mean_squared_error(y_test_m3, y_hat_test_m3)
print('Train Mean Squarred Error:', train_mse_m3)
print('Test Mean Squarred Error:', test_mse_m3)
# -
# #### Model 4
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 4
multi_model_4 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], zipcode_encoded_ohe, bathrooms_encoded_ohe], axis = 1)
X_multi4 = multi_model_4.drop('log_price', axis = 1)
y_multi4 = multi_model_4['log_price']
X_train_m4, X_test_m4, y_train_m4, y_test_m4 = train_test_split(X_multi4, y_multi4, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m4[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m4[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m4.index)
scaled_columns.columns = cols_scale
X_train_m4['sqft_living'] = scaled_columns['sqft_living']
multi4_reg = LinearRegression()
multi4_reg.fit(X_train_m4, y_train_m4)
print(multi4_reg.score(X_train_m4, y_train_m4))
# Scaling test set
scale_test = X_test_m4[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m4.index)
scaled_columns_test.columns = cols_scale
X_test_m4['sqft_living'] = scaled_columns_test['sqft_living']
print(multi4_reg.score(X_test_m4, y_test_m4))
# Bathrooms lead to overfit so omitted the y_hat value and mean square error calculation
# -
# #### Model 5
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 5
multi_model_5 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], zipcode_encoded_ohe, bedrooms_encoded_ohe], axis = 1)
X_multi5 = multi_model_5.drop('log_price', axis = 1)
y_multi5 = multi_model_5['log_price']
X_train_m5, X_test_m5, y_train_m5, y_test_m5 = train_test_split(X_multi5, y_multi5, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m5[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m5[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m5.index)
scaled_columns.columns = cols_scale
X_train_m5['sqft_living'] = scaled_columns['sqft_living']
multi5_reg = LinearRegression()
multi5_reg.fit(X_train_m5, y_train_m5)
print(multi5_reg.score(X_train_m5, y_train_m5))
# Scaling test set
scale_test = X_test_m5[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m5.index)
scaled_columns_test.columns = cols_scale
X_test_m5['sqft_living'] = scaled_columns_test['sqft_living']
print(multi5_reg.score(X_test_m5, y_test_m5))
y_hat_train_m5 = multi5_reg.predict(X_train_m5)
y_hat_test_m5 = multi5_reg.predict(X_test_m5)
train_mse_m5 = mean_squared_error(y_train_m5, y_hat_train_m5)
test_mse_m5 = mean_squared_error(y_test_m5, y_hat_test_m5)
print('Train Mean Squarred Error:', train_mse_m5)
print('Test Mean Squarred Error:', test_mse_m5)
# -
# #### Model 6
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 6
multi_model_6 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], zipcode_encoded_ohe, view_encoded_ohe], axis = 1)
X_multi6 = multi_model_6.drop('log_price', axis = 1)
y_multi6 = multi_model_6['log_price']
X_train_m6, X_test_m6, y_train_m6, y_test_m6 = train_test_split(X_multi6, y_multi6, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m6[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m6[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m6.index)
scaled_columns.columns = cols_scale
X_train_m6['sqft_living'] = scaled_columns['sqft_living']
multi6_reg = LinearRegression()
multi6_reg.fit(X_train_m6, y_train_m6)
print(multi6_reg.score(X_train_m6, y_train_m6))
# Scaling test set
scale_test = X_test_m6[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m6.index)
scaled_columns_test.columns = cols_scale
X_test_m6['sqft_living'] = scaled_columns_test['sqft_living']
print(multi6_reg.score(X_test_m6, y_test_m6))
y_hat_train_m6 = multi6_reg.predict(X_train_m6)
y_hat_test_m6 = multi6_reg.predict(X_test_m6)
train_mse_m6 = mean_squared_error(y_train_m6, y_hat_train_m6)
test_mse_m6 = mean_squared_error(y_test_m6, y_hat_test_m6)
print('Train Mean Squarred Error:', train_mse_m6)
print('Test Mean Squarred Error:', test_mse_m6)
# -
# #### Model 7
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 7
multi_model_7 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], housing_pred_final['waterfront'], zipcode_encoded_ohe, view_encoded_ohe], axis = 1)
X_multi7 = multi_model_7.drop('log_price', axis = 1)
y_multi7 = multi_model_7['log_price']
X_train_m7, X_test_m7, y_train_m7, y_test_m7 = train_test_split(X_multi7, y_multi7, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m7[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m7[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m7.index)
scaled_columns.columns = cols_scale
X_train_m7['sqft_living'] = scaled_columns['sqft_living']
multi7_reg = LinearRegression()
multi7_reg.fit(X_train_m7, y_train_m7)
print(multi7_reg.score(X_train_m7, y_train_m7))
# Scaling test set
scale_test = X_test_m7[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m7.index)
scaled_columns_test.columns = cols_scale
X_test_m7['sqft_living'] = scaled_columns_test['sqft_living']
print(multi7_reg.score(X_test_m7, y_test_m7))
y_hat_train_m7 = multi7_reg.predict(X_train_m7)
y_hat_test_m7 = multi7_reg.predict(X_test_m7)
train_mse_m7 = mean_squared_error(y_train_m7, y_hat_train_m7)
test_mse_m7 = mean_squared_error(y_test_m7, y_hat_test_m7)
print('Train Mean Squarred Error:', train_mse_m7)
print('Test Mean Squarred Error:', test_mse_m7)
# -
# #### Model 8
# +
# Above Steps repeated with different combinations of housing features to see if it improves our model
# Multiple Linear Regression Model number 8
multi_model_8 = pd.concat([housing_pred_final['log_price'], housing_pred_final['sqft_living'], housing_pred_final['waterfront'],
housing_pred_final['Basement'], zipcode_encoded_ohe, view_encoded_ohe,
cond_encoded_ohe, floors_encoded_ohe, bathrooms_encoded_ohe, bedrooms_encoded_ohe, grade_num_encoded_ohe], axis = 1)
X_multi8 = multi_model_8.drop('log_price', axis = 1)
y_multi8 = multi_model_8['log_price']
X_train_m8, X_test_m8, y_train_m8, y_test_m8 = train_test_split(X_multi8, y_multi8, test_size=0.2, random_state=42)
# Scaling sqft_living(standardscaler)
cols_scale = ['sqft_living']
scaler = StandardScaler()
scaled_fit = scaler.fit(X_train_m8[cols_scale])
scaled_columns = scaled_fit.transform(X_train_m8[cols_scale])
scaled_columns = pd.DataFrame(scaled_columns, index = X_train_m8.index)
scaled_columns.columns = cols_scale
X_train_m8['sqft_living'] = scaled_columns['sqft_living']
multi8_reg = LinearRegression()
multi8_reg.fit(X_train_m8, y_train_m8)
print(multi8_reg.score(X_train_m8, y_train_m8))
# Scaling test set
scale_test = X_test_m8[cols_scale]
scaled_columns_test = scaled_fit.transform(scale_test)
scaled_columns_test = pd.DataFrame(scaled_columns_test, index = X_test_m8.index)
scaled_columns_test.columns = cols_scale
X_test_m8['sqft_living'] = scaled_columns_test['sqft_living']
print(multi8_reg.score(X_test_m8, y_test_m8))
y_hat_train_m8 = np.exp(multi8_reg.predict(X_train_m8))
y_hat_test_m8 = np.exp(multi8_reg.predict(X_test_m8))
train_mse_m8 = mean_squared_error(np.exp(y_train_m8), y_hat_train_m8)
test_mse_m8 = mean_squared_error(np.exp(y_test_m8), y_hat_test_m8)
test_rmse_m8 = np.sqrt(test_mse_m8)
print('Train Mean Squarred Error:', train_mse_m8)
print('Test Mean Squarred Error:', test_mse_m8)
print('Test RMSE: ', test_rmse_m8)
# -
y_pred = np.exp(multi8_reg.predict(X_test_m8))
y_act = np.exp(y_test_m8)
fig, ax = plt.subplots()
sns.set_style("dark")
ax.scatter(y_pred, y_act, s = 2)
ax.plot([y_act.min(), y_act.max()], [y_act.min(), y_act.max()], 'k--', lw=1)
ax.set_xlabel('Predicted')
ax.set_ylabel('Actual')
ax.ticklabel_format(style='plain');
# **Final Observations & Insights:**
# <ul>
# </ul>
# <li>
regression_score = pd.DataFrame(index=['Simple', 'M-1', 'M-3', 'M-6', 'M-8'], columns=['test R^2'])
regression_score['test R^2'] = [simple_reg.score(X_test_s, y_test_s),
multi1_reg.score(X_test_m1, y_test_m1),
multi3_reg.score(X_test_m3, y_test_m3),
multi6_reg.score(X_test_m6, y_test_m6),
multi8_reg.score(X_test_m8, y_test_m8)]
# +
model_score_fig, ax = plt.subplots(figsize=(10,6))
sns.set_style("dark")
ax.bar(regression_score.index, regression_score['test R^2'], color=["red", "grey", "grey", "grey", "darkblue"])
plt.title('Model Progression')
plt.xlabel('Model Iteration')
plt.ylabel('Test R^2');
# -
# ----
# # Inferential Model Results
# +
formula = 'price ~ sqft_living + waterfront + C(zipcode) + C(view)'
model_ols = ols(formula=formula, data=housing_pred).fit()
# -
model_summary = model_ols.summary()
model_summary
sm.ProbPlot(model_ols.resid).qqplot(line='s');
plt.title('Q-Q plot');
# +
fitted_vals = model_ols.predict()
resids = model_ols.resid
resids_standardized = model_ols.get_influence().resid_studentized_internal
fig, ax = plt.subplots(1,2)
sns.regplot(x=fitted_vals, y=resids, lowess=True, ax=ax[0], line_kws={'color': 'red'})
ax[0].set_title('Residuals vs Fitted', fontsize=16)
ax[0].set(xlabel='Fitted Values', ylabel='Residuals')
sns.regplot(x=fitted_vals, y=np.sqrt(np.abs(resids_standardized)), lowess=True, ax=ax[1], line_kws={'color': 'red'})
ax[1].set_title('Scale-Location', fontsize=16)
ax[1].set(xlabel='Fitted Values', ylabel='sqrt(abs(Residuals))')
# -
wtr_df = housing_ols.groupby(by = 'waterfront').mean().reset_index()
# +
wtr_fig, ax = plt.subplots(figsize=(10,6))
sns.set_style("dark")
plt.bar(wtr_df['waterfront'], wtr_df['price'], color = 'darkblue')
plt.xlabel('Waterfront(No/Yes)')
plt.ylabel('Average Price')
plt.title('Presence of Waterfront')
ax.ticklabel_format(style='plain', axis = 'y');
# -
view_df = housing_ols.groupby(by = 'view').mean().reset_index()
view_df.sort_values(by = 'price', ascending = True, inplace = True)
view_fig, ax = plt.subplots(figsize=(10,6))
sns.set_style("dark")
color = ['darkblue', 'red', 'grey', 'darkblue', 'darkblue']
plt.bar(view_df['view'], view_df['price'], color = color)
plt.ylabel('Average Price')
plt.xlabel('Quality of View')
plt.title('Avg Price Based on View')
ax.ticklabel_format(style='plain', axis = 'y')
plt.annotate('Blue: Sign. Grey: Not Sign. Red: Reference', (0.02, .95), backgroundcolor='w', xycoords='axes fraction');
# # Conclusions
|
code/Final-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exercise 7 : This covers three types of inheritance.
# Dated : April 15, 2020.
# # - Single inheritance -
# # Question 1
# - Create a class Person, having attributes
# - Name
# - Age
# - Height
# - Weight
#
# - Create another class Worker, having attributes
# - Company
# - Position (Rank)
# - working hours
# - print_info() # method
#
# - Create a child object and assign values to the class members by passing values in the contructor.
# - Print all the informtion using print_info()
#
# Note: Worker class is child class and Person is the parent
# +
# Your code goes here
class Person:
def __init__(self, Name, Age, Height, Weight):
self.Name = Name
self.Age = Age
self.Height = Height
self.Weight = Weight
def about_myself(self):
return "My name is {}, and I am {} years old. I am {} cm tall and weight {} kg".format(self.Name, self.Age, self.Height, self.Weight)
class Worker(Person):
def __init__(self, Name, Age, Height, Weight, Company, Position, Working_Hours):
super(). __init__(Name, Age, Height, Weight)
self.Company = Company
self.Position = Position
self.Working_Hours = Working_Hours
def print_info(self):
return "I work for {}, as an {}, and I work {}".format(self.Company, self.Position, self.Working_Hours)
A = Worker('Johnny', 21, 180, 70, 'Python Corp', 'Employee of the Month', 'Every Day')
print(A.about_myself())
print(A.print_info())
# -
# # - Multiple inheritance -
# # Question 2
#
# - Create a class Person, having attributes
# - Name
# - Age
#
# - Create a class Dancer, having attributes
# - Style
#
# - Create another class Worker, having attributes
# - Company
# - Position (Rank)
# - working hours
# - print_info() # method
#
# - Here we suppose that a Worker is a Dancer and obviously a Person
# - Parent classes : Person and Dancer
# - Child class : Worker
# - Create a child object and assign values to the class members by passing values in the contructor.
# - Print all the informtion
#
# +
# Your code goes here
class Person:
def __init__(self, Name, Age, Height, Weight):
self.Name = Name
self.Age = Age
self.Height = Height
self.Weight = Weight
def about_myself(self):
return "My name is {}, and I am {} years old. I am {} cm tall and weight {} kg".format(self.Name, self.Age, self.Height, self.Weight)
class Dancer:
def __init__(self, style):
self.style = style
class Worker(Person):
def __init__(self, Name, Age, Height, Weight, Company, Position, Working_Hours, style):
self.Company = Company
self.Position = Position
self.Working_Hours = Working_Hours
Person. __init__(self, Name, Age, Height, Weight)
Dancer. __init__(self, style)
def print_info(self):
return (self.about_myself()
+ '\n' + "Company: {}, Position: {}, Working Hours:{}".format(self.Company, self.Position, self.Working_Hours)
+ '\n' + "Dance Style: {}".format(self.style))
A = Worker('Johnny', 21, 180, 70, 'Python Corp', 'Employee of the Month', 'Every Day', 'Disco')
print(A.print_info())
# -
# # - Multi-level inheritance -
# # Question 3
#
# - Create a class GrandFather, having attribute
# - grand_father_name
# - grand_father_age
# - print_grand_info() # method
#
# - Create a class Father(GrandFather), having attribute
# - father_name
# - father_age
# - print_parent_info() # method
#
# - Create a class Son(Father), having attribute
# - son_name
# - son_age
# - print_child_info() # method
#
# - Create an object of Son class, and initialize all the values of parent and grand parent classes.
# - Print all the information using print_info() methods of all three classes
# +
# your code goes here
class GrandFather:
def __init__(self, name, age):
self.name = name
self.age = age
def Grand_info(self):
return ("My name is {} and I am {} years old".format(self.name, self.age))
class Father(GrandFather):
def __init__(self, father_name, father_age):
self.father_name = father_name
self.father_age = father_age
def Father_info(self):
return ("My name is {} and I am {} years old".format(self.father_name, self.father_age))
class Son(Father):
def __init__(self, name, age, father_name, father_age, son_name, son_age):
self.son_name = son_name
self.son_age = son_age
GrandFather. __init__(self, name, age)
Father. __init__(self, father_name, father_age)
def Son_info(self):
return (self.Grand_info() + '\n' + self.Father_info() + '\n'
+ "My name is {} and I am {} years old".format(self.son_name, self.son_age))
S = Son('<NAME>', 75, '<NAME>', 42, 'Vasyl', 18)
print(S.Son_info())
# -
# # Question 4
# - What is a diamond problem?
# - When does diamond problem occur?
# - What is isinstance() method used for? Code an example.
# +
# Your answer goes here
Dimaond problem
- Diamond problem occurs in the context of multiple inheritance
- Example: D inherits from B and C who both inherits from A
- the issue is that python will not be sure which method prevail and which attributes/methods from A are inherited by D
- Python deals with this ambiguity by enforcing order of priority in tuple of vase class
- in the exmaple above, the method resolution order would be D,B,C,A
isinstance()
- syntax: isinstance(object, type)
- object: Required. An object.
- type: A type or a class, or a tuple of types and/or classes
- purpose: rturns TRue if the specified object is of the specified type, otherwise false
# -
isinstance("hello",tuple)
isinstance(12,int)
isinstance(('shoe', 'shirt'), tuple)
# # Happy Coding!
|
Exercises - Qasim/Python. Pandas, Viz/Questions/Exercise 7 (Single, Multiple, Multilevel Inheritance).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
"""
Created on Tue Jun 5 16:32:01 2018
@author: buck06191
"""
import json
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.gridspec as gridspec
import string
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
from PIL import Image
import io
import os
plt.rcParams['figure.figsize'] = (12, 8)
plt.rcParams['figure.dpi'] = 300
def get_axis_limits(ax, scale=.9):
return ax.get_xlim()[1]*scale, ax.get_ylim()[1]*scale
def setAxLinesBW(ax):
"""
Take each Line2D in the axes, ax, and convert the line style to be
suitable for black and white viewing.
"""
MARKERSIZE = 3
COLORMAP = {
'b': {'marker': None, 'dash': (None,None)},
'g': {'marker': None, 'dash': [5,5]},
'r': {'marker': None, 'dash': [5,3,1,3]},
'c': {'marker': None, 'dash': [1,3]},
'm': {'marker': None, 'dash': [5,2,5,2,5,10]},
'y': {'marker': None, 'dash': [5,3,1,2,1,10]},
'k': {'marker': 'o', 'dash': (None,None)} #[1,2,1,10]}
}
lines_to_adjust = ax.get_lines()
try:
lines_to_adjust += ax.get_legend().get_lines()
except AttributeError:
pass
for line in lines_to_adjust:
# if line.get_label()[:3] != "Q10":
# continue
origColor = line.get_color()
line.set_color('black')
line.set_dashes(COLORMAP[origColor]['dash'])
line.set_marker(COLORMAP[origColor]['marker'])
line.set_markersize(MARKERSIZE)
return lines_to_adjust
def setFigLinesBW(fig):
"""
Take each axes in the figure, and for each line in the axes, make the
line viewable in black and white.
"""
fig_label = ["({})".format(s) for s in list(string.ascii_lowercase)]
for jj, ax in enumerate(fig.get_axes()):
lines = setAxLinesBW(ax)
ax.set_title(fig_label[jj], loc='left')
labels = [l.get_label() for l in lines]
fig.legend(lines, labels, 'upper left', prop={'size': 14}, bbox_to_anchor=(0.91, 0.91))
def save_as_tiff(fig, fname):
# Save the image in memory in PNG format
png1 = io.BytesIO()
fig.savefig(png1, format="png", bbox_inches='tight')
# Load this image into PIL
png2 = Image.open(png1)
# Save as TIFF
png2.save("../Figures/{}.tiff".format(fname))
print("saved Tiff")
png1.close()
def get_autoregulation_json_data(model, output, direction):
data_dir = "../data/steady_state/{}/autoregulation/".format(model)
with open(os.path.join(data_dir, '{}_{}.json'.format(output, direction)), 'r') as f:
jsondata = json.load(f)
return jsondata
def get_outputs_json_data(model, direction):
data_dir = "../data/steady_state/{}/model_output/".format(model)
with open(os.path.join(data_dir, 'q_range_runs_{}.json'.format(direction)), 'r') as f:
jsondata = json.load(f)
return jsondata
# -
# ## Blood Pressure
def generate_SS_figure_combo(model):
xlabels = {"P_a": "Arterial Blood\nPressure (mmHg)",
"SaO2sup": 'Arterial Oxygen\nSaturation (%)',
"Pa_CO2": 'Partial Pressure\nof CO$_2$ (mmHg)'}
outputs = list(xlabels.keys())
directions=['down', 'up']
fig, axes = plt.subplots(nrows=2, ncols=len(outputs), sharey=True, sharex='col', figsize=(8,6))
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
n = len(outputs)
for ix in [(i,j) for i in range(2) for j in range(3)]:
direction = directions[ix[0]]
output = outputs[ix[1]]
ax = axes[ix]
jsondata = get_autoregulation_json_data(model, output, direction)
keys = list(jsondata.keys())
keys.sort()
for idx, k in enumerate(keys):
if output == 'SaO2sup':
jsondata[k][output] = [i*100 for i in jsondata[k][output]]
axes[ix].plot(jsondata[k][output], jsondata[k]['CBF'], label='%s$^\circ$C'%k, c=colors[idx], alpha=0.6)
axes[ix].set_title("{})".format(string.ascii_lowercase[3*ix[0]+ix[1]]), loc="left")
if ix[0]==1:
axes[ix].set_xlabel(xlabels[output], size=12)
else:
axes[ix].set_xlabel('')
for item in (axes[ix].get_xticklabels() + axes[ix].get_yticklabels()):
item.set_fontsize(12)
axes[0,0].set_ylabel('Cerebral Blood Flow\n($ml_{blood}ml_{brain}^{-1}s^{-1}$)', size=12)
axes[1,0].set_ylabel('Cerebral Blood Flow\n($ml_{blood}ml_{brain}^{-1}s^{-1}$)', size=12)
axes.flatten()[2].legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0., fontsize=10)
plt.tight_layout()
fig.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/{}/combo_{}_autoreg.png".format(
model,model), transparent=True)
return fig
bph1_ar_fig = generate_SS_figure_combo("bp_hypothermia_1")
bph2_ar_fig = generate_SS_figure_combo("bp_hypothermia_2")
bph3_ar_fig = generate_SS_figure_combo("bp_hypothermia_4")
# +
def generate_SS_figure(output, model, xlabel="", legend_loc="upper left"):
title={'down': 'Decreasing', 'up': 'Increasing'}
fig, axes = plt.subplots(ncols=2, sharey=True, sharex=True)
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
for ix, direction in enumerate(["down", "up"]):
jsondata = get_autoregulation_json_data(model, output, direction)
keys = list(jsondata.keys())
keys.sort()
data = jsondata[keys[0]][output]
for idx, k in enumerate(keys):
axes[ix].plot(jsondata[k][output], jsondata[k]['CBF'], label='Temp: %sC'%k, c=colors[idx])
axes[ix].set_title("{}) {}".format(string.ascii_lowercase[ix], title[direction]), loc="left")
axes[ix].set_xlabel(xlabel, size=12)
for item in (axes[ix].get_xticklabels() + axes[ix].get_yticklabels()):
item.set_fontsize(12)
axes[0].set_ylabel('Cerebral Blood Flow\n($ml_{blood}ml_{brain}^{-1}s^{-1}$)', size=12)
axes[1].legend(loc=legend_loc, fontsize=10)
plt.tight_layout()
fig.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/{}/{}_{}_autoreg.png".format(
model,output,model), transparent=True)
return fig
pa_fig = generate_SS_figure("P_a", "bp_hypothermia_1", "Arterial Blood\nPressure (mmHg)")
sao2_fig = generate_SS_figure("SaO2sup", "bp_hypothermia_1", 'Arterial Oxygen\nSaturation (%)', legend_loc="lower left")
paco2_fig = generate_SS_figure("Pa_CO2", "bp_hypothermia_1", 'Partial Pressure\nof CO2 (mmHg)', legend_loc="lower right")
# -
# +
pa_fig_2 = generate_SS_figure("P_a", "bp_hypothermia_2", "Arterial Blood\nPressure (mmHg)")
sao2_fig_2 = generate_SS_figure("SaO2sup", "bp_hypothermia_2", 'Arterial Oxygen\nSaturation (%)', legend_loc="lower left")
paco2_fig_2 = generate_SS_figure("Pa_CO2", "bp_hypothermia_2", 'Partial Pressure\nof CO2 (mmHg)', legend_loc="lower right")
# +
pa_fig_3 = generate_SS_figure("P_a", "bp_hypothermia_4", "Arterial Blood\nPressure (mmHg)")
sao2_fig_3 = generate_SS_figure("SaO2sup", "bp_hypothermia_4", 'Arterial Oxygen\nSaturation (%)', legend_loc="lower left")
paco2_fig_3 = generate_SS_figure("Pa_CO2", "bp_hypothermia_4", 'Partial Pressure\nof CO2 (mmHg)', legend_loc="lower right")
# -
# ## Model output visualisation ##
bp2_data = get_outputs_json_data(model="bp_hypothermia_2", direction="down")
bp3_data = get_outputs_json_data(model="bp_hypothermia_4", direction="down")
bp1_data = get_outputs_json_data(model="bp_hypothermia_1", direction="down")
# +
title = {"CBF": "CBF", "CCO": "$\Delta$ oxCCO","CMRO2": "CMRO$_{2}$", "TOI": "TOI", "Hbdiff": "$\Delta$ HbD"}
units = {'Hbdiff': "$\mu M$", 'CCO': "$\mu M$", "CBF" : "% of baseline", "CMRO2": "% of baseline", "TOI": "%" }
def parse_json_data(json_data):
parsed_data={}
for k, v in json_data.items():
params = tuple(map(lambda x: round(float(x),2), k.split('_')))
parsed_data[params] = {output: data[-1] for output, data in v.items()}
return parsed_data
def get_cols_and_rows(parsed_json_data):
"""
Get set of col and rows from json keys.
Return
======
cols, rows : sets of the param values
"""
rows = []
cols = []
for k in parsed_json_data.keys():
rows.append(k[0])
cols.append(k[1])
return sorted(set(cols)), sorted(set(rows))
def create_dataframe(parsed_json_data):
cols, rows = get_cols_and_rows(parsed_json_data)
df = pd.DataFrame(index=rows, columns=cols)
return df
def get_baseline_cmro2(json_data):
parsed_data={}
for k, v in json_data.items():
params = tuple(map(lambda x: round(float(x),2), k.split('_')))
parsed_data[params] = {output: data[-1]/data[0] * 100 for output, data in v.items()}
return parsed_data
def get_baseline_NIRS(json_data):
parsed_data={}
for k, v in json_data.items():
params = tuple(map(lambda x: round(float(x),2), k.split('_')))
parsed_data[params] = {output: data[-1]-data[0] for output, data in v.items()}
return parsed_data
def fill_dataframe(json_data, output):
if output == "CMRO2" or output == "CBF":
parsed_json_data = get_baseline_cmro2(json_data)
elif output == "Hbdiff" or output == "CCO":
parsed_json_data = get_baseline_NIRS(json_data)
else:
parsed_json_data = parse_json_data(json_data)
df = create_dataframe(parsed_json_data)
for k, v in parsed_json_data.items():
df.loc[k] = v[output]
return df
def get_output_heatmap(json_data, output, xlabel=None, ylabel=None):
fig, ax = plt.subplots(1, figsize = (4,3.5))
df = fill_dataframe(json_data, output)
if output == "CCO" or output == "Hbdiff":
sns.heatmap(df.apply(pd.to_numeric), ax=ax, cmap="RdBu_r", center=0, vmin= df.values.min(), vmax= df.values.max(),
cbar_kws={'label': '{} ({})'.format(title[output], units[output])})
else:
sns.heatmap(df.apply(pd.to_numeric), ax=ax, cmap="BuPu", vmin= df.values.min(), vmax= df.values.max(),
cbar_kws={'label': '{} ({})'.format(title[output], units[output])})
ax.set_title(title[output])
if xlabel:
ax.set_xlabel(xlabel, size=12)
if ylabel:
ax.set_ylabel(ylabel, size=12)
plt.tight_layout()
return df, fig
def truncate_colormap(cmap, minval=0.0, maxval=1.0, n=100):
new_cmap = mpl.colors.LinearSegmentedColormap.from_list(
'trunc({n},{a:.2f},{b:.2f})'.format(n=cmap.name, a=minval, b=maxval),
cmap(np.linspace(minval, maxval, n)))
return new_cmap
def get_output_timeseries(json_data, output, xlabel=None, ylabel=None):
fig = plt.figure(figsize=(4,3.5))
ax1 = plt.subplot2grid((4, 4), (0, 0), rowspan=4, colspan=3, fig=fig)
ax2 = plt.subplot2grid((4, 4), (0, 3), rowspan=4, fig=fig)
n = len(json_data.keys())
Q10 = np.arange(0.1, 5.1, 0.1)
colors = plt.cm.get_cmap('nipy_spectral')(np.linspace(0.,1,n))
cmap = truncate_colormap(plt.cm.get_cmap('nipy_spectral'), 0., 1)
norm = mpl.colors.Normalize(vmin=0.1, vmax=5.)
# ColorbarBase derives from ScalarMappable and puts a colorbar
# in a specified axes, so it has everything needed for a
# standalone colorbar. There are many more kwargs, but the
# following gives a basic continuous colorbar with ticks
# and labels.
cb1 = mpl.colorbar.ColorbarBase(ax2, cmap=cmap,
norm=norm,
orientation='vertical')
cb1.set_label('$Q_{10}$')
i=0
for k,v in json_data.items():
if output == "CMRO2" or output == "CBF":
v[output] = [i/v[output][0] * 100 for i in v[output]]
ax1.plot(v['temp'], v[output], label=k, color=colors[i])
i+=1
ax1.set_title(title[output])
if xlabel:
ax1.set_xlabel(xlabel, size=12)
if ylabel:
ax1.set_ylabel(ylabel, size=12)
# fig.colorbar(plt.cm.ScalarMappable(norm = pltcol.Normalize(vmin=0.1, vmax=5), cmap=plt.cm.get_cmap('BuPu')))
plt.tight_layout()
return fig
# -
for output in title.keys():
g = get_output_timeseries(bp1_data, output, xlabel="Temp. ($^{\circ}$C)", ylabel="{} ({})".format(title[output], units[output]) )
g.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_1/{}_bph1_outputs.png".format(output), transparent=True)
for output in title.keys():
df, g = get_output_heatmap(bp2_data, output, xlabel="$Q_{10, haemo}$", ylabel="$Q_{10,met}$" )
g# .savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_2/{}_bph2_outputs.png".format(output), transparent=True)
# ## Initial Notes ##
#
# BPH4 with $q_{diff} > 1$ produces no change - odel becomes insensitive to further changes.
for output in title.keys():
df, g = get_output_heatmap(bp3_data, output, xlabel="$q_{diff}$", ylabel="$Q_{10}$" )
g.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_4/{}_bph3_outputs.png".format(output), transparent=True)
df_cco, _ = get_output_heatmap(bp2_data, 'CCO', xlabel="$Q_{10,haemo}$", ylabel="$Q_{10,met}$" )
fig, ax = plt.subplots(1)
for ix, d in enumerate(df_cco.values):
if (ix % 5 == 0) & (ix<25):
ax.plot(df_cco.columns, d, label=df_cco.index[ix])
fig.legend(title = "$Q_{10,met}$", loc=2, bbox_to_anchor=(0.135,0.52))
ax.set_xlabel("$Q_{10,haemo}$", size=12)
ax.set_ylabel("{} ({})".format(title['CCO'], units['CCO']), size=12)
fig.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_2/CCO_bph2_lines.png", transparent=True)
df_cco, _ = get_output_heatmap(bp3_data, 'CCO', xlabel="$q_{diff}$", ylabel="$Q_{10}$" )
fig, ax = plt.subplots(1)
for ix, d in enumerate(df_cco.values):
if ix % 10 == 0:
ax.plot(df_cco.columns, d, label=df_cco.index[ix])
fig.legend(title = "$Q_{10}$", loc=2, bbox_to_anchor=(0.135,0.52))
ax.set_xlabel("$q_{diff}$", size=12)
ax.set_ylabel("{} ({})".format(title['CCO'], units['CCO']), size=12)
fig.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_4/CCO_bph3_lines.png", transparent=True)
df_cco, _ = get_output_heatmap(bp3_data, 'Hbdiff', xlabel="$q_{diff}$", ylabel="$Q_{10}$" )
fig, ax = plt.subplots(1)
for ix, d in enumerate(df_cco.values):
if (ix % 3 == 0) & (ix<15):
ax.plot(df_cco.columns, d, label=df_cco.index[ix])
ax.axhline(0, color='k', alpha=0.5)
fig.legend(title = "$Q_{10}$", loc=2, bbox_to_anchor=(0.75,0.85))
ax.set_xlabel("$q_{diff}$", size=12)
ax.set_ylabel("{} ({})".format(title['Hbdiff'], units['Hbdiff']), size=12)
fig.savefig("/home/buck06191/Dropbox/phd/hypothermia/Figures/steady_state/model_comparisons/bp_hypothermia_4/HbD_bph3_lines.png", transparent=True)
get_output_heatmap(bp2_data, 'HbO2')
# +
## Q10 figures
# with open('q10/220618T1154.json', 'r') as f:
jsondata_q10 = {}
jsondata_q10['down'] = get_outputs_json_data("bp_hypothermia_1", "down")
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
outputs = ['TOI', 'CBF', 'Hbdiff', 'CCO', 'CMRO2']
units = ["(%)", "(% of baseline)", "($\mu M$)", "($\mu M$)", "(% of baseline)"]
keys = list(jsondata_q10['down'].keys())[:6]
ylims = {"up": {}, "down": {}}
# jsondata_q10['up'] = {k:{} for k in keys}
# jsondata_q10['down'] = {k: {} for k in keys}
# for k in keys:
# jsondata_q10['up'][k] = {inner_k: jsondata_q10_all[k][inner_k][:len(jsondata_q10_all[k]['t']) // 2 + 1] for inner_k in jsondata_q10_all[k].keys()}
# jsondata_q10['down'][k] = {inner_k: jsondata_q10_all[k][inner_k][len(jsondata_q10_all[k]['t']) // 2:] for inner_k in jsondata_q10_all[k].keys()}
print(keys)
keys.sort()
# fig, axes = plt.subplots(nrows=2, ncols=3)
for direction in ['down']:
for ii, o in enumerate(outputs):
fig, ax = plt.subplots(1)
print("\n{}\n".format(o))
# ax = axes.flatten()[ii]
for idx, k in enumerate(keys):
if o == 'CBF' or o == 'CMRO2':
print(k, ": ", jsondata_q10[direction][k][o][0])
jsondata_q10[direction][k][o] = [100*(i/jsondata_q10[direction][k][o][0]) for i in jsondata_q10[direction][k][o]]
# l = len(jsondata_q10[k]['t']) // 2
ax.plot(jsondata_q10[direction][k]['temp'], jsondata_q10[direction][k][o], label='Q10 = %s'%k, c=colors[idx])
max_idx = jsondata_q10[direction][k]['temp'].index(37)
# for j, t in enumerate(jsondata_q10[direction][k]['temp']):
# if 33.4<t<33.6:
# min_idx = j
# break
# if direction=="down":
# perc_change = jsondata_q10[direction][k][o][min_idx] / jsondata_q10[direction][k][o][max_idx] * 100
# print("From 37C to 33.5C {} changes to {:.1f}% of its starting value for Q10={}".format(o,perc_change,k))
# elif direction=="up":
# perc_change = jsondata_q10[direction][k][o][max_idx] / jsondata_q10[direction][k][o][min_idx] * 100
# print("From 33.5C to 37C {} changes to {:.1f}% of its starting value for Q10={}".format(o,perc_change,k))
ax.set_xlabel('Temperature ($^\circ$C)', size=16)
if o=='CCO':
ax.set_ylabel("$\Delta$oxCCO {}".format(units[ii]), size=16)
elif o=='DHbdiff':
ax.set_ylabel("$\Delta$HbD {}".format(units[ii]), size=16)
elif o == "TOI":
ax.set_ylabel("StO2 {}".format(units[ii]), size=16)
else:
ax.set_ylabel("{} {}".format(o, units[ii]), size=16)
ax.set_title("{d}".format(d="Rewarming" if direction == "up" else "Cooling"), size=16)
ax.tick_params(axis='both', which='major', labelsize=16)
ax.legend(prop={'size': 14})
ylims[direction][o] = ax.get_ylim()
plt.tight_layout()
fname = "varying_Q10_{}_{}.png".format(o, direction)
plt.show()
print(ylims)
# +
## Q10 figures
# with open('q10/220618T1154.json', 'r') as f:
jsondata_q10 = {}
jsondata_q10['down'] = get_outputs_json_data("bp_hypothermia_1", "down")
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
outputs = ['TOI', 'CBF', 'Hbdiff', 'CCO', 'CMRO2']
units = ["(%)", "(% of baseline)", "($\mu M$)", "($\mu M$)", "(% of baseline)"]
keys = list(jsondata_q10['down'].keys())[:6]
# jsondata_q10['up'] = {k:{} for k in keys}
# jsondata_q10['down'] = {k: {} for k in keys}
# for k in keys:
# jsondata_q10['up'][k] = {inner_k: jsondata_q10_all[k][inner_k][:len(jsondata_q10_all[k]['t']) // 2 + 1] for inner_k in jsondata_q10_all[k].keys()}
# jsondata_q10['down'][k] = {inner_k: jsondata_q10_all[k][inner_k][len(jsondata_q10_all[k]['t']) // 2:] for inner_k in jsondata_q10_all[k].keys()}
print(keys)
keys.sort()
# fig, axes = plt.subplots(nrows=2, ncols=3)
for direction in ['down']:
fig, axes = plt.subplots(nrows=2, ncols=3)
for ii, o in enumerate(outputs):
print("\n{}\n".format(o))
ax = axes.flatten()[ii]
for idx, k in enumerate(keys):
if o == 'CBF' or o == 'CMRO2':
print(k, ": ", jsondata_q10[direction][k][o][0])
jsondata_q10[direction][k][o] = [100*(i/jsondata_q10[direction][k][o][0]) for i in jsondata_q10[direction][k][o]]
# l = len(jsondata_q10[k]['t']) // 2
# l = len(jsondata_q10[k]['t']) // 2
ax.plot(jsondata_q10[direction][k]['temp'], jsondata_q10[direction][k][o], label='Q10 = %s'%k, c=colors[idx])
max_idx = jsondata_q10[direction][k]['temp'].index(37)
for j, t in enumerate(jsondata_q10[direction][k]['temp']):
if 33.4<t<33.6:
min_idx = j
break
if direction=="down":
perc_change = jsondata_q10[direction][k][o][min_idx] / jsondata_q10[direction][k][o][max_idx] * 100
print("From 37C to 33.5C {} changes to {:.1f}% of its starting value for Q10={}".format(o,perc_change,k))
elif direction=="up":
perc_change = jsondata_q10[direction][k][o][max_idx] / jsondata_q10[direction][k][o][min_idx] * 100
print("From 33.5C to 37C {} changes to {:.1f}% of its starting value for Q10={}".format(o,perc_change,k))
ax.set_xlabel('Temperature ($^\circ$C)', size=16)
if o=='CCO':
ax.set_ylabel("$\Delta$oxCCO {}".format(units[ii]), size=16)
elif o=='DHbdiff':
ax.set_ylabel("$\Delta$HbD {}".format(units[ii]), size=16)
elif o == "TOI":
ax.set_ylabel("StO2 {}".format(units[ii]), size=16)
else:
ax.set_ylabel("{} {}".format(o, units[ii]), size=16)
ax.tick_params(axis='both', which='major', labelsize=14)
ax.locator_params(tight=True, nbins=4)
ax.set_ylim(bottom = (ax.get_ylim()[0]-0.054)*0.9975, top=(ax.get_ylim()[1]+0.01)*1.0015)
#ax.legend(prop={'size': 14})
fig.delaxes(axes[-1,-1])
#ax.axvline(33.5)
# fig.suptitle("{d}".format(d="Rewarming" if direction == "up" else "Cooling"), size=18)
fig.subplots_adjust(top=0.8)
axes.flatten()[len(outputs)-1].legend(prop={'size': 14}, bbox_to_anchor=(0.95, 0.91))
plt.tight_layout()
fname = "varying_Q10_{}_combined".format(direction)
# plt.savefig("E:/Dropbox/phd/Conferences/ISOTT-2018/Figures/{}".format(fname))
plt.show()
# +
## Q10 figures
# with open('q10/220618T1154.json', 'r') as f:
print(ylims)
jsondata_q10 = {}
with open('q10/020718T1013.json', 'r') as f:
jsondata_q10['down'] = json.load(f)
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k']
outputs = ['TOI', 'CBF', 'DHbdiff', 'CCO', 'CMRO2']
units = ["(%)", "(% of baseline)", "($\mu M$)", "($\mu M$)", "(% of baseline)"]
keys = list(jsondata_q10_all.keys())[:6]
# jsondata_q10['up'] = {k:{} for k in keys}
# jsondata_q10['down'] = {k: {} for k in keys}
# for k in keys:
# jsondata_q10['up'][k] = {inner_k: jsondata_q10_all[k][inner_k][:len(jsondata_q10_all[k]['t']) // 2 + 1] for inner_k in jsondata_q10_all[k].keys()}
# jsondata_q10['down'][k] = {inner_k: jsondata_q10_all[k][inner_k][len(jsondata_q10_all[k]['t']) // 2:] for inner_k in jsondata_q10_all[k].keys()}
print(keys)
keys.sort()
# fig, axes = plt.subplots(nrows=2, ncols=3)
for direction in ['down']:
for ii, o in enumerate(outputs):
fig, ax = plt.subplots(1)
print("\n{}\n".format(o))
# ax = axes.flatten()[ii]
for idx, k in enumerate(keys):
if o == 'CBF' or o == 'CMRO2':
jsondata_q10[direction][k][o] = [100*(i/jsondata_q10[direction][k][o][0]) for i in jsondata_q10[direction][k][o]]
# l = len(jsondata_q10[k]['t']) // 2
if k == '2.5':
ax.plot(jsondata_q10[direction][k]['temp'], jsondata_q10[direction][k][o], label='Q10 = %s'%k, c=colors[idx])
ax.set_xlabel('Temperature ($^\circ$C)', size=16)
if o=='CCO':
ax.set_ylabel("$\Delta$oxCCO {}".format(units[ii]), size=16)
elif o=='DHbdiff':
ax.set_ylabel("$\Delta$HbD {}".format(units[ii]), size=16)
elif o == "TOI":
ax.set_ylabel("StO2 {}".format(units[ii]), size=16)
else:
ax.set_ylabel("{} {}".format(o, units[ii]), size=16)
#ax.set_title('Effect of decreasing temperature on {}\nfor varying Q10'.format(o), size=16)
ax.set_title("{d}".format(d="Rewarming" if direction == "up" else "Cooling"), size=16)
ax.tick_params(axis='both', which='major', labelsize=16)
ax.legend(prop={'size': 14})
ax.set_ylim(ylims[direction][o])
#fig.delaxes(axes[-1,-1])
setFigLinesBW(fig)
#ax.axvline(33.5)
plt.tight_layout()
fname = "Q10_2_5_{}_{}.png".format(o, direction)
plt.savefig("/home/buck06191/Dropbox/phd/Conferences/ISOTT-2018/Figures/{}".format(fname))
plt.show()
# +
DHbd = jsondata_q10['up']['2.5']['DHbdiff']
CCO = jsondata_q10['up']['2.5']['CCO']
temp = jsondata_q10['up']['2.5']['temp']
for i in temp:
if i >33.5:
idx = temp.index(i)
break
fig, ax = plt.subplots(1)
ax.plot(CCO[idx:], DHbd[idx:], '.k')
ax.axhline(0, color='k')
# set the x-spine (see below for more info on `set_position`)
ax.spines['right'].set_position('zero')
# turn off the right spine/ticks
ax.spines['left'].set_color('none')
ax.yaxis.tick_right()
ax.yaxis.set_label_position('right')
# set the y-spine
ax.spines['bottom'].set_position('zero')
# turn off the top spine/ticks
ax.spines['top'].set_color('none')
ax.spines['bottom'].set_color('none')
#ax.set_ylim(ax.get_ylim()[0],0.02)
ax.tick_params(axis='both', which='major', labelsize=16)
ax.set_xlabel("$\Delta$oxCCO ($\mu M$)", size=16)
ax.set_ylabel("$\Delta$HbD ($\mu M$)", size=16)
fig, ax = plt.subplots(nrows=2, sharex=True, sharey=True, figsize=(12,12))
ax[0].plot(temp[idx:], CCO[idx:], '.k')
ax[0].axhline(0, color='k')
ax[1].plot(temp[idx:], DHbd[idx:], '.k')
ax[1].axhline(0, color='k')
ax[0].tick_params(axis='both', which='major', labelsize=16)
ax[0].set_ylabel("$\Delta$oxCCO ($\mu M$)", size=16)
ax[1].tick_params(axis='both', which='major', labelsize=16)
ax[1].set_ylabel("$\Delta$HbD ($\mu M$)", size=16)
ax[1].set_xlabel('Temperature ($^\circ$C)', size=16)# # fig, axes = plt.subplots(nrows=2, ncols=3)
#plt.tight_layout()
# for ii, o in enumerate(outputs):
# fig, ax = plt.subplots(1)
# print("\n{}\n".format(o))
# # ax = axes.flatten()[ii]
# #l = len(jsondata_q10[k]['t']) // 2
# ax.plot(jsondata_q10["2.5"]['temp'], jsondata_q10["2.5"][o], label='Q10 = %s'%k, c=colors[idx])
# ax.set_xlabel('Temperature ($^\circ$C)', size=16)
# if o=='CCO':
# ax.set_ylabel("$\Delta$oxCCO {}".format(units[ii]), size=16)
# elif o == "TOI":
# ax.set_ylabel("StO2 {}".format(units[ii]), size=16)
# else:
# ax.set_ylabel("{} {}".format(o, units[ii]), size=16)
# #ax.set_title('Effect of decreasing temperature on {}\nfor varying Q10'.format(o), size=16)
# ax.tick_params(axis='both', which='major', labelsize=16)
# #ax.legend(prop={'size': 12})
# #fig.delaxes(axes[-1,-1])
# setFigLinesBW(fig)
# plt.tight_layout()
# fname = "Q10_2_5_{}.png".format(o)
# plt.savefig("/home/buck06191/Dropbox/phd/Conferences/ISOTT-2018/Figures/{}".format(fname))
plt.show()
# -
|
steadystate_Scripts/BORL_Figure_generate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Group Sparse inverse covariance for multi-subject connectome
# =============================================================
#
# This example shows how to estimate a connectome on a group of subjects
# using the group sparse inverse covariance estimate.
#
#
#
# +
import numpy as np
from nilearn import plotting
n_subjects = 4 # subjects to consider for group-sparse covariance (max: 40)
def plot_matrices(cov, prec, title, labels):
"""Plot covariance and precision matrices, for a given processing. """
prec = prec.copy() # avoid side effects
# Put zeros on the diagonal, for graph clarity.
size = prec.shape[0]
prec[list(range(size)), list(range(size))] = 0
span = max(abs(prec.min()), abs(prec.max()))
# Display covariance matrix
plotting.plot_matrix(cov, cmap=plotting.cm.bwr,
vmin=-1, vmax=1, title="%s / covariance" % title,
labels=labels)
# Display precision matrix
plotting.plot_matrix(prec, cmap=plotting.cm.bwr,
vmin=-span, vmax=span, title="%s / precision" % title,
labels=labels)
# -
# Fetching datasets
# ------------------
#
#
# +
from nilearn import datasets
msdl_atlas_dataset = datasets.fetch_atlas_msdl()
adhd_dataset = datasets.fetch_adhd(n_subjects=n_subjects)
# print basic information on the dataset
print('First subject functional nifti image (4D) is at: %s' %
adhd_dataset.func[0]) # 4D data
# -
# Extracting region signals
# --------------------------
#
#
# +
from nilearn import image
from nilearn import input_data
# A "memory" to avoid recomputation
from sklearn.externals.joblib import Memory
mem = Memory('nilearn_cache')
masker = input_data.NiftiMapsMasker(
msdl_atlas_dataset.maps, resampling_target="maps", detrend=True,
low_pass=None, high_pass=0.01, t_r=2.5, standardize=True,
memory='nilearn_cache', memory_level=1, verbose=2)
masker.fit()
subject_time_series = []
func_filenames = adhd_dataset.func
confound_filenames = adhd_dataset.confounds
for func_filename, confound_filename in zip(func_filenames,
confound_filenames):
print("Processing file %s" % func_filename)
# Computing some confounds
hv_confounds = mem.cache(image.high_variance_confounds)(
func_filename)
region_ts = masker.transform(func_filename,
confounds=[hv_confounds, confound_filename])
subject_time_series.append(region_ts)
# -
# Computing group-sparse precision matrices
# ------------------------------------------
#
#
# +
from nilearn.connectome import GroupSparseCovarianceCV
gsc = GroupSparseCovarianceCV(verbose=2)
gsc.fit(subject_time_series)
from sklearn import covariance
gl = covariance.GraphLassoCV(verbose=2)
gl.fit(np.concatenate(subject_time_series))
# -
# Displaying results
# -------------------
#
#
# +
atlas_img = msdl_atlas_dataset.maps
atlas_region_coords = plotting.find_probabilistic_atlas_cut_coords(atlas_img)
labels = msdl_atlas_dataset.labels
plotting.plot_connectome(gl.covariance_,
atlas_region_coords, edge_threshold='90%',
title="Covariance",
display_mode="lzr")
plotting.plot_connectome(-gl.precision_, atlas_region_coords,
edge_threshold='90%',
title="Sparse inverse covariance (GraphLasso)",
display_mode="lzr",
edge_vmax=.5, edge_vmin=-.5)
plot_matrices(gl.covariance_, gl.precision_, "GraphLasso", labels)
title = "GroupSparseCovariance"
plotting.plot_connectome(-gsc.precisions_[..., 0],
atlas_region_coords, edge_threshold='90%',
title=title,
display_mode="lzr",
edge_vmax=.5, edge_vmin=-.5)
plot_matrices(gsc.covariances_[..., 0],
gsc.precisions_[..., 0], title, labels)
plotting.show()
|
experiments/.ipynb_checkpoints/plot_multi_subject_connectome-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="QWlVMMX7avna"
# ## Big O Notation & Algorithm Analysis
# ### An algorithm can be thought of a procedure or formula to solve a particular problem. The question is, which algorithm to use to solve a specific problem when there exist multiple solutions to the problem?
#
# + [markdown] id="lQLsxN_fbKaK"
# ### Big O Notation
# - a statistical measure, used to describe the complexity of the algorithm
# - signifies the relationship between the input to the algorithm and the steps required to execute the algorithm
# - O(n)
#
# ### Why is algorithm analysis important?
# - there are multiple ways to solve most problems, how do you decide which one to use?
# - time and computational power cost money
# + [markdown] id="_zZ0jU4Zdl4R"
# ### Common Big O functions
#
# 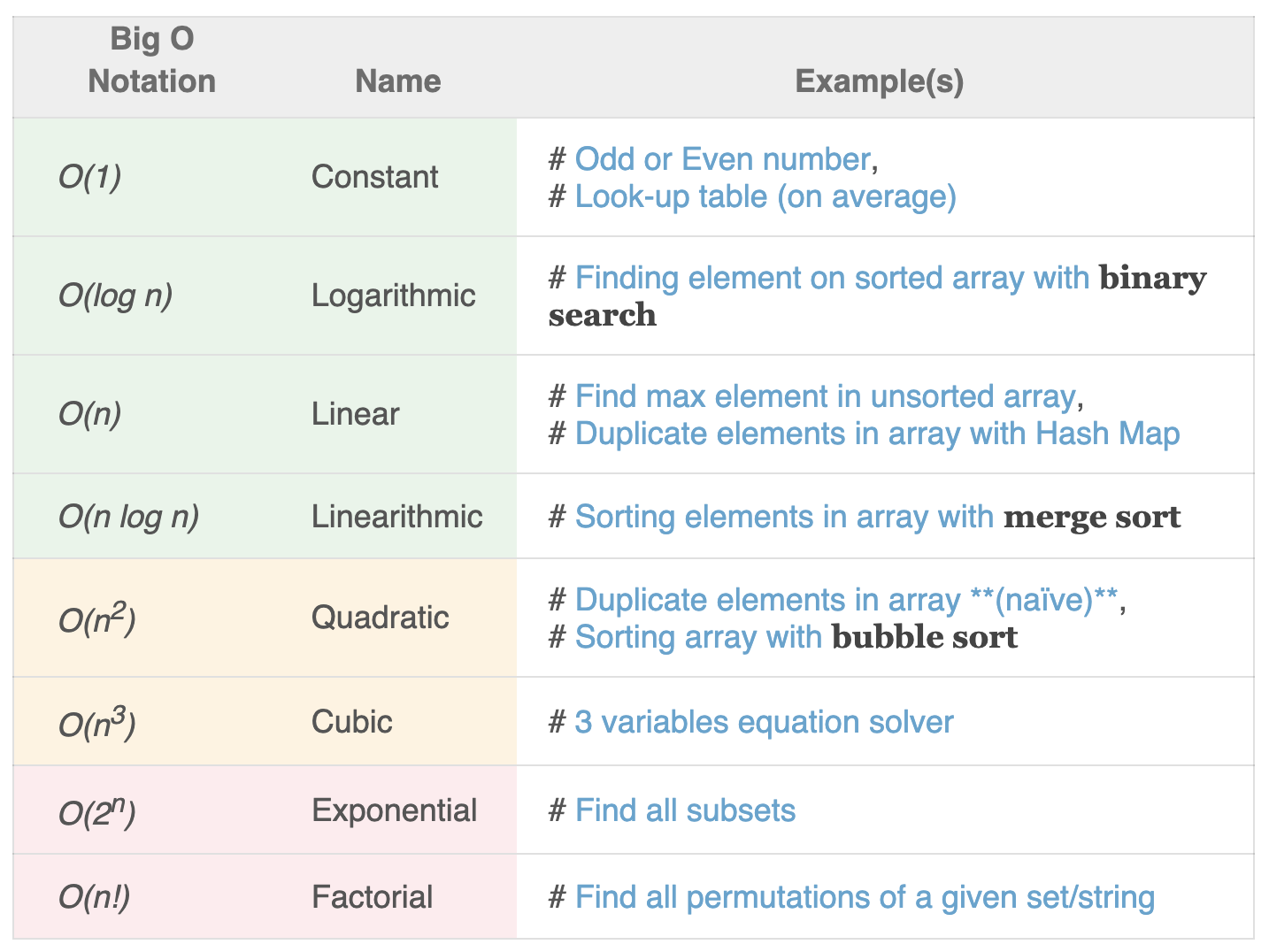
# 
# 
# ### Big 0 Notation steps
# - Break your algorithm/function into individual operations
# - Calculate the Big O of each operation
# - Add up the Big O of each operation together
# - Remove the constants
# - Find the highest order term,this will be what we consider the Big O of our algorithm/function
# + [markdown] id="k7GQEKkgfGU4"
# ## Big O Examples
# + [markdown] id="fhGvApxifMOp"
# ### Constant Complexity ---> (O(C))
# #### is constant if the steps required to complete the execution of an algorithm remain constant, irrespective of the number of inputs
# + id="pwlKoK0gbRFW"
### no matter the size of the 'items' 2 steps are performed, constant ###
def constant_algo(items):
result = items[0] * items[0] # step 1
print(result) # step 2
result = constant_algo([4, 5, 6, 8])
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="Zdp7AnvygojT" outputId="17160286-eebb-4564-ffeb-38a81b437222"
### visualize the model prediction accuracy ###
import matplotlib.pyplot as plt
import numpy as np
x = [2, 4, 6, 8, 10, 12]
y = [2, 2, 2, 2, 2, 2]
### configure the plot ###
print('--- Constant Complexity O(c) --- ')
f, ax = plt.subplots(1, 1, figsize = (10, 7))
ax1 = plt.plot(x, y, color="b", label="input values")
plt.xlabel('Inputs')
plt.ylabel('Steps')
plt.title('Constant Complexity')
plt.legend()
plt.show()
# + [markdown] id="mpVoaMpliZcY"
# ### Linear Complexity (O(n))
# #### is linear if the steps required to complete the execution of an algorithm increase or decrease linearly with the number of inputs
# + id="C_76ff8JggDH"
### number of interations is dependant on the size of the input array ###
def linear_algo(items):
for item in items: # loop through the items
print(item)
result = linear_algo([4, 5, 6, 8])
# + colab={"base_uri": "https://localhost:8080/", "height": 489} id="0iUl_W4FjKtD" outputId="056ce9a0-5318-458f-df02-e3b32c966a97"
import matplotlib.pyplot as plt
import numpy as np
x = [2, 4, 6, 8, 10, 12]
y = [2, 4, 6, 8, 10, 12]
### configure the plot ###
print('--- Linear Complexity O(n) --- ')
f, ax = plt.subplots(1, 1, figsize = (10, 7))
ax1 = plt.plot(x, y, color="b", label="input values")
plt.xlabel('Inputs')
plt.ylabel('Steps')
plt.title('Linear Complexity')
plt.legend()
# + id="_3hAehOmjPIS"
### 2 constant for loops turn to 1, O(n) not O(2n)###
def linear_algo(items):
for item in items: # 1st loop through the items
print(item)
for item in items: # 2nd loop through the items
print(item)
result = linear_algo([4, 5, 6, 8])
# + colab={"base_uri": "https://localhost:8080/", "height": 489} id="WMJlvo8AItLt" outputId="8180f8b7-1a53-431e-f3ad-d0ce8018df48"
import matplotlib.pyplot as plt
import numpy as np
x = [2, 4, 6, 8, 10, 12]
y = [4, 8, 12, 16, 20, 24]
### configure the plot ###
print('--- Linear Complexity O(C) --- ')
f, ax = plt.subplots(1, 1, figsize = (10, 7))
ax1 = plt.plot(x, y, color="b", label="input values")
plt.xlabel('Inputs')
plt.ylabel('Steps')
plt.title('Linear Complexity')
plt.legend()
# + [markdown] id="NfrkZYOpLCli"
#
# + [markdown] id="_r-boaZMJruz"
# ### Quadratic Complexity (O(n^2))
# #### is quadratic when the steps required to execute an algorithm are a quadratic function of the number of items in the input
# + id="YneedK--J7SG"
def quadratic_algo(items):
for item in items: # outter loop
for item2 in items: # nested inner loop
print(item, ' ' ,item)
### number of steps performed is n * n ###
quadratic_algo([4, 5, 6, 8])
# + [markdown] id="L9j0rKwcK_3r"
# 
# + [markdown] id="lxH-eGPaLDyP"
# ### Complexity of Complex Functions
# - always take the worst case complexity
# - if function contains O(n^2) and O(n), choose O(n^2)
# + id="7ePpgIWHM0fS"
def complex_algo(items):
for i in range(5): # 1st loop through set range, O(n)
print ("Python is awesome")
for item in items: # 2nd loop through the items, O(n)
print(item)
for item in items: # 3rd loop through the items, O(n)
print(item)
print("Big O")
print("Big O")
print("Big O")
### O(3n) is O(n) ###
result = complex_algo([4, 5, 6, 8])
# + [markdown] id="QBK-dr-ZOg1O"
# ### Space Complexity
# - refers to the number of spaces you need to allocate in the memory space during the execution of a program
# + id="oO_XZaqFM0ku"
def return_squares(n):
square_list = [] # created list
for num in n: # loop through input
square_list.append(num * num) # append squard to list
### have to allocate memory for the same number of items as in input list O(n) ###
return square_list
nums = [2, 4, 6, 8, 10]
result = return_squares(nums)
|
AlgorithmAnalysis_Notes/BigOAlgorithmAnalysis_Notes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualiser des principes de base de l'IRM
# ## Quelques outils pour observer la composition des images du cerveau
# Ce notebook présente des exemples d'images IRM utilisant différents types de séquences. Ces images ont pour but de visualiser les différents paramètres de l'IRM et leurs effets sur la représention des tissues.
# ### Nilearn
# Nous utiliserons cette [librairie](https://nilearn.github.io/user_guide.html) pour effectuer l'ensemble du travail. Accédez la documentation pour davantage d'info et d'exemples.
#
# * Nilearn 6.2
# Nilearn est gros, nous pouvons donc n'importer que les parties qui nous intéressent. Nilearn contient le module [`plotting`](https://nilearn.github.io/modules/reference.html#module-nilearn.plotting) nous permettant de tirer profit, en (presqu') une seule ligne de code, d'une vaste gamme de techniques de visualisation. Allez voir la liste, mais commençons par la base.
# ## Importer les données
# On commence par récupérer des données. [Ici](https://nilearn.github.io/modules/reference.html#module-nilearn.datasets), vous avez la liste des jeux de données qui se retrouvent à même la librairie Nilearn.
#
# Nous choisissons le *template* (modèle) du Montreal Neurological Institute (MNI). Il s'agit, en quelque sorte, d'un cerveau moyen. Ces données représentent l'espace à partir duquel on repère les différentes régions cérébrales.
from nilearn.datasets import fetch_icbm152_2009
mni = fetch_icbm152_2009() # mni est un dictionnaire contenant plusieurs objets
# 
# ## Images pondérées en T1
# On dit plus couramment qu'on utilise une séquence structurelle lorsqu'on regarde ces images. C'est ce qu'on veut dire lorsqu'on parle tout simplement d'une image par <NAME>.
# `plot_anat` visualise un scan anatomique. Ici, l'image retourne de façon statique. Nous avons une coordonnée représentée sur les trois coupes. Nous restons donc en deux dimensions.
#
# C'est une image typique de l'anatomie. Nous pouvons bien distinguer les tissue. La matière grise est grise, la blanche est blanche, les ventricules sont noirs (tout comme une partie de la boîte crânienne). Comme on l'a vu, c'est à l'aide de cette pondération que nous pouvons mener des analyses structurelles.
from nilearn.plotting import plot_anat
plot_anat(mni.t1, cut_coords=[-17, 0, 17],title='MRI en contraste T1')
# (Pour ceux et celles qui sont curieux.ses, vous avez là un type d'objet qu'on appelle bytes, utilisé pour représenter le spectre noir à blanc sur les trois axes. Les Bytes prennent moins d'espace de stockage)
# +
# mni.t1?
# -
# `view_img`, quant à lui, nous permet d'interagir avec l'image à l'aide de notre curseur. On peut donc modifier la coordonnée d'origine.
from nilearn.plotting import view_img
viewer = view_img(mni.t1, bg_img=None, title='MRI en contraste T1', cmap='gray', symmetric_cmap=False,
black_bg=True, threshold=None,)
viewer.save_as_html('t1.html')
# Nous avons accès à des données HTML qui nous permettent d'explorer les trois coupes.
#
# **[Ouvrez-les](t1.html)** (assurez-vous d'ouvrir ce lien dans un ***nouvel onglet***)
# ## Images pondérées en T2
# Nous utilisons encore la fonction `plot_anat` pour représenter le spectre de noir à blanc (0 à 1) qui nous aide à distinguer les tissues. Le T2 est moins intuitif pour les dissocier; on ne l'utilise pas pour explorer les segmenter.
plot_anat(mni.t2, cut_coords=[-17, 0, 17], title='IRM en contraste T2')
# On peut aussi explorer l'image de façon interactive.
viewer = view_img(mni.t2, bg_img=None, title='IRM en contraste T2', cmap='gray', symmetric_cmap=False,
black_bg=True, threshold=None)
viewer.save_as_html('t2.html')
# **[Ouvrez la visualisation interactive](t2.html)** (assurez-vous d'ouvrir ce lien dans un ***nouvel onglet***)
# ### BOLD ou T2*
# T2* est un autre type de séquence. Elle a plusieurs applications possibles dont l'imagerie fonctionnelle. Cette séquence nous permet d'imager les variations magnétiques du système vasculaire cérébrale, et notamment le signal BOLD (blood-oxygenation level dependent) utilisé en IRMf. Mais, [le contraste T2*](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2799958/) est aussi important pour étudier les traumatismes crâniens, les tumeurs, et les arrêts vasculaires-cérébraux, puisqu'elle nous permet d'imager la **structure** du système vasculaire.
# #### Importer des volumes
# Faisons comme précédemment et importons des données. Cette fois-ci, il s'agit d'un participant de la base de données [ADHD200](https://www.sciencedirect.com/science/article/pii/S105381191630283X). Il s'agit d'une **série de volumes 3D pondérés en T2***.
from nilearn.datasets import fetch_adhd
adhd = fetch_adhd(n_subjects=1)
# ### Volume par volume
# Visualisons le premier volume de la série. `adhd.func` contient les données en 4D. On peut accéder à un volume 3D de la série en spécifiant son index, comme cela : `index_img(adhd.func[0], i)`
from nilearn.plotting import plot_img
from nilearn.image import index_img
plot_img(index_img(adhd.func[0], 0),
bg_img=None,
cut_coords=(36, -27, 66),
black_bg=True,
title="un volume BOLD")
# Le deuxième volume de la série :
from nilearn.plotting import plot_img
from nilearn.image import index_img
plot_img(index_img(adhd.func[0], 1),
bg_img=None,
cut_coords=(36, -27, 66),
black_bg=True,
title="un volume BOLD")
# Les deux volumes sont très similaires! Les changements liés à l'oxygénation du sang sont très petits. On peut regarder le volume de manière interactive.
from nilearn.plotting import view_img
viewer = view_img(index_img(adhd.func[0], 1), bg_img=None, title='un volume BOLD', symmetric_cmap=False,
black_bg=True, threshold=0, cmap='hot')
viewer.save_as_html('bold.html')
# **[Ouvrez la visualisation interactive](bold.html)** (assurez-vous d'ouvrir ce lien dans un ***nouvel onglet***)
# ## Résumé
# ### Contraste T1
# 
# ### Contraste T2
# 
# * T1 - TE et TR courts
# * Contraste rendant le gras clair, comparativement aux fluides (vasculaires, spinaux)
#
# * T2 - TE et TR longs
# * Constraste rendant les fluides clairs, comparativement aux tissues (muscles, gras)
#
# * T2* - TE et TR longs + Angle de bascule bas
# * Contraste rendant les petites variations d'intensité de signal claires (le déphasage)
# * BOLD et Diffusion
#
|
applications_cours2_principes_IRM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: junction
# language: python
# name: junction
# ---
# +
import os
import json
import re
import pandas as pd
import numpy as np
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.model_selection import TimeSeriesSplit, cross_val_score, train_test_split
from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor
import seaborn as sns
from matplotlib import pyplot as plt
from dotenv import load_dotenv, find_dotenv
assert load_dotenv(find_dotenv())
sns.set(font_scale=1.2)
counter_ids = [
'912', #
'1043', #
'1050', #
'1225', #
'922', #
'1246', #
]
import shap
# load JS visualization code to notebook
shap.initjs()
# -
weather_data = pd.read_csv('../data/processed/weather.csv', index_col='Date', parse_dates=['Date'])
weather_data = weather_data.drop(columns=['Unnamed: 0', 'Aikavyöhyke']).fillna(0)
data = pd.read_csv('../data/processed/visitor_counts_6_counters_Nuuksio_with_holidays.csv',
index_col='StartTime',
parse_dates=['StartTime'])
# +
for cid in counter_ids:
thresh = np.quantile(data[cid].dropna(), 0.999)
data[cid][data[cid] > thresh] = np.nan
data.fillna(0, inplace=True)
# -
data = data.join(weather_data)
data['month_day'] = data.index.day
data['weekday'] = data.index.weekday
data['hour'] = data.index.hour
data['month'] = data.index.month
y = data[counter_ids]
X = data.drop(columns=counter_ids)
X_encoded = pd.get_dummies(X)
X_encoded.head()
X_train, X_test, y_train, y_test = train_test_split(X_encoded, y, shuffle=False)
pipe = Pipeline([
('scaler', StandardScaler()),
('reg', RandomForestRegressor(n_estimators=10))
])
scores = -cross_val_score(pipe, X_train, y_train,
cv=TimeSeriesSplit(n_splits=5),
scoring='neg_mean_absolute_error')
scores
model = GradientBoostingRegressor(n_estimators=10).fit(X_train, y_train.values[:, 4])
explainer = shap.TreeExplainer(model)
shap_values = explainer.shap_values(X_test)
y_pred = model.predict(X_test)
y_test.mean()
y_pred.m
ix = 100
shap.force_plot(explainer.expected_value, shap_values[ix,:], X_test.iloc[0, :])
shap.summary_plot(shap_values, X_test)
|
notebooks/Regression model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BERTCRFNER
#
# 可用的中文预训练参数:[`bert-base`](https://storage.googleapis.com/bert_models/2018_11_03/chinese_L-12_H-768_A-12.zip),[`roberta-wwm-ext-base`](https://drive.google.com/uc?export=download&id=1jMAKIJmPn7kADgD3yQZhpsqM-IRM1qZt),[`roberta-wwm-ext-large`](https://drive.google.com/uc?export=download&id=1dtad0FFzG11CBsawu8hvwwzU2R0FDI94),[`macbert-base`](https://drive.google.com/uc?export=download&id=1aV69OhYzIwj_hn-kO1RiBa-m8QAusQ5b),[`macbert-large`](https://drive.google.com/uc?export=download&id=1lWYxnk1EqTA2Q20_IShxBrCPc5VSDCkT)
# +
import uf
print(uf.__version__)
# -
model = uf.BERTCRFNER('../../demo/bert_config.json', '../../demo/vocab.txt')
print(model)
X = ['天亮以前说再见', '笑着泪流满面', '去迎接应该你的', '更好的明天']
y = [['天亮'], [], ['迎接'], ['更好', '明天']]
# # 训练
model.fit(X, y, total_steps=20)
# # 推理
model.predict(X)
# # 评分
model.score(X, y)
|
examples/tutorial/BERTCRFNER.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
# %load_ext autoreload
# %autoreload 2
# -
import warnings
warnings.filterwarnings("ignore")
# +
import tensorflow as tf
import tensorflow_probability as tfp
import mesh_tensorflow as mtf
import flowpm
import flowpm.mesh_ops as mpm
import flowpm.mtfpm as mtfpm
import flowpm.mesh_utils as mesh_utils
from astropy.cosmology import Planck15
from flowpm.tfpm import PerturbationGrowth
from flowpm import linear_field, lpt_init, nbody, cic_paint
from flowpm.utils import r2c3d, c2r3d
sys.path.append('../utils/')
import tools
import diagnostics as dg
import contextlib
import functools
import math, time
from scipy.interpolate import InterpolatedUnivariateSpline as iuspline
import scipy.optimize as sopt
# -
from tensorflow.python.keras import backend as K
# +
from recurrent_inference import rim, ConvRIM3D
from convolutional_recurrent import ConvLSTM3DCell
from tensorflow.python.keras.layers import ConvLSTM2D
from tensorflow.python.keras.layers.convolutional_recurrent import ConvLSTM2DCell
from tensorflow.python.keras.layers import LSTMCell, LSTM, Dense, Conv3D
# +
bs, nc = 100, 32
nsteps = 3
a0, a, nsteps = 0.1, 1.0, nsteps
stages = np.linspace(a0, a, nsteps, endpoint=True)
klin = np.loadtxt('../data/Planck15_a1p00.txt').T[0]
plin = np.loadtxt('../data//Planck15_a1p00.txt').T[1]
ipklin = iuspline(klin, plin)
# Compute necessary Fourier kernels
kvec = tools.fftk((nc, nc, nc), boxsize=nc, symmetric=False)
kmesh = (sum(k**2 for k in kvec)**0.5).astype(np.float32)
priorwt = ipklin(kmesh)
# +
nsims = 200
dpath = '../data/rim-data/L%04d_N%03d_T%02d/'%(bs, nc, nsteps)
alldata = np.array([np.load(dpath + '%04d.npy'%i) for i in range(nsims)]).astype(float32)
# alldata = np.expand_dims(alldata, axis=2)
traindata, testdata = alldata[:int(0.9*nsims)], alldata[int(0.9*nsims):]
ic, fin = traindata[:1,0], traindata[:1,1]
# -
def get_ps(iterand, truth):
ic, fin = truth
ic1, fin1 = iterand
pks = []
if abs(ic1[0].mean()) < 1e-3: ic1[0] += 1
#if abs(ic[0].mean()) < 1e-3: ic[0] += 1
k, p1 = tools.power(ic1[0]+1, boxsize=bs)
k, p2 = tools.power(ic[0]+1, boxsize=bs)
k, p12 = tools.power(ic1[0]+1, f2=ic[0]+1, boxsize=bs)
pks.append([p1, p2, p12])
if fin1[0].mean() < 1e-3: fin1[0] += 1
if fin[0].mean() < 1e-3: fin[0] += 1
k, p1 = tools.power(fin1[0], boxsize=bs)
k, p2 = tools.power(fin[0], boxsize=bs)
k, p12 = tools.power(fin1[0], f2=fin[0], boxsize=bs)
pks.append([p1, p2, p12])
return k, pks
# # GD
# +
@tf.function
def pm(linear):
state = lpt_init(linear, a0=0.1, order=1)
final_state = nbody(state, stages, nc)
tfinal_field = cic_paint(tf.zeros_like(linear), final_state[0])
return tfinal_field
@tf.function
def recon_prototype(linear, data, Rsm, anneal=True):
"""
"""
# linear = tf.reshape(linear, data.shape)
final_field = pm(linear)
residual = final_field - data #.astype(np.float32)
base = residual
if anneal :
print("\nAdd annealing section to graph\n")
Rsmsq = tf.multiply(Rsm*bs/nc, Rsm*bs/nc)
smwts = tf.exp(tf.multiply(-kmesh**2, Rsmsq))
basek = r2c3d(base, norm=nc**3)
basek = tf.multiply(basek, tf.cast(smwts, tf.complex64))
base = c2r3d(basek, norm=nc**3)
chisq = tf.multiply(base, base)
chisq = tf.reduce_sum(chisq)
# chisq = tf.multiply(chisq, 1/nc**3, name='chisq')
#Prior
lineark = r2c3d(linear, norm=nc**3)
priormesh = tf.square(tf.cast(tf.abs(lineark), tf.float32))
prior = tf.reduce_sum(tf.multiply(priormesh, 1/priorwt))
# prior = tf.multiply(prior, 1/nc**3, name='prior')
#
loss = chisq + prior
return loss
@tf.function
def recon_loss_and_grad(x, y, Rsm):
print("val and grad : ", x.shape)
with tf.GradientTape() as tape:
tape.watch(x)
loss = recon_prototype(x, y, Rsm)
grad = tape.gradient(loss, x)
return loss, grad
@tf.function
def recon_grad(x, y, Rsm):
with tf.GradientTape() as tape:
tape.watch(x)
loss = recon_prototype(x, y, Rsm)
grad = tape.gradient(loss, [x])
return grad
@tf.function
def recon_prototype_noanneal(linear, data):
"""
"""
print('new graph')
final_field = pm(linear)
residual = final_field - data #.astype(np.float32)
chisq = tf.multiply(residual, residual)
chisq = tf.reduce_mean(chisq)
# chisq = tf.multiply(chisq, 1/nc**3, name='chisq')
#Prior
lineark = r2c3d(linear, norm=nc**3)
priormesh = tf.square(tf.cast(tf.abs(lineark), tf.float32))
prior = tf.reduce_mean(tf.multiply(priormesh, 1/priorwt))
# prior = tf.multiply(prior, 1/nc**3, name='prior')
#
loss = chisq + prior
return loss, chisq, prior
@tf.function
def recon_noanneal_grad(x, y):
with tf.GradientTape() as tape:
tape.watch(x)
loss = recon_prototype_noanneal(x, y)[0]
grad = tape.gradient(loss, x)
return grad
# +
RRs = [0]
niter = 100
# Create an optimizer for Adam.
opt = tf.keras.optimizers.Adam(learning_rate=0.1)
# opt = tf.keras.optimizers.SGD(learning_rate=100)
#Loop it Reconstruction
##Reconstruction a
x0 = np.random.normal(0, 1, nc**3).reshape(fin.shape).astype(np.float32)
linear = tf.Variable(name='linmesh', shape=(1, nc, nc, nc), dtype=tf.float32,
initial_value=x0, trainable=True)
for iR, RR in enumerate(RRs):
for i in range(niter):
# loss, chisq, prior = recon_prototype_noanneal(linear, tf.constant(fin, dtype=tf.float32))
# print(i, loss.numpy(), chisq.numpy(), prior.numpy())
# grads = recon_grad(linear, tf.constant(fin, dtype=tf.float32), tf.constant(RR, dtype=tf.float32))
# opt.apply_gradients(zip(grads, [linear]))
grads = recon_noanneal_grad(linear, tf.constant(fin, dtype=tf.float32))
opt.apply_gradients(zip([grads], [linear]))
minic = linear.numpy().reshape(fin.shape)
x0 = minic
minfin = pm(tf.constant(minic)).numpy()
# +
# RRs = [0]
# niter = 20
# # Create an optimizer for Adam.
# # opt = tf.keras.optimizers.Adam(learning_rate=0.1)
# opt_sgd = tf.keras.optimizers.SGD(learning_rate=0.1)
# opt_adam = tf.keras.optimizers.Adam(learning_rate=0.1)
# #Loop it Reconstruction
# ##Reconstruction a
# x0 = np.random.normal(0, 1, nc**3).reshape(fin.shape).astype(np.float32)
# linear = tf.Variable(name='linmesh', shape=(1, nc, nc, nc), dtype=tf.float32,
# initial_value=x0, trainable=True)
# linear1 = tf.Variable(name='linmesh2', shape=(1, nc, nc, nc), dtype=tf.float32,
# initial_value=x0, trainable=True)
# steps_adam, steps_sgd = [], []
# for iR, RR in enumerate(RRs):
# for i in range(niter):
# # loss, chisq, prior = recon_prototype_noanneal(linear, tf.constant(fin, dtype=tf.float32))
# # print(i, loss.numpy(), chisq.numpy(), prior.numpy())
# # grads = recon_grad(linear, tf.constant(fin, dtype=tf.float32), tf.constant(RR, dtype=tf.float32))
# # opt.apply_gradients(zip(grads, [linear]))
# l0 = linear.numpy()
# grads = recon_noanneal_grad(linear, tf.constant(fin, dtype=tf.float32))
# opt_adam.apply_gradients(zip([grads], [linear]))
# l1 = linear.numpy()
# steps_adam.append(l1-l0)
# l0 = linear1.numpy()
# opt_sgd.apply_gradients(zip([grads], [linear1]))
# l1 = linear1.numpy()
# steps_sgd.append(l1-l0)
# linear1.assign(linear.numpy())
# minic = linear.numpy().reshape(fin.shape)
# x0 = minic
# minfin = pm(tf.constant(minic)).numpy()
# +
b1, b2 = 0.9, 0.999
m = 0
v = 0
t = 1
eps = 1e-7
lr = 0.1
##Reconstruction a
linear3 = tf.Variable(name='linmesh', shape=(1, nc, nc, nc), dtype=tf.float32,
initial_value=x0, trainable=True)
for i in range(niter):
t = i+1
grads = recon_noanneal_grad(linear3, tf.constant(fin, dtype=tf.float32))
m = b1*m + (1-b1)*grads.numpy()
v = b2*v + (1-b2)*grads.numpy()**2
mc = m/(1-b1**t)
vc = v/(1-b2**t)
delta = lr*mc/(np.sqrt(vc) + 1e-7)
linear3 = linear3 - delta
minic3 = linear3.numpy().reshape(fin.shape)
x3 = minic3
minfin3 = pm(tf.constant(minic3)).numpy()
# -
fig, ax = plt.subplots(1, 3, figsize=(7, 3))
im = ax[0].imshow(ic[0].sum(axis=0))
plt.colorbar(im, ax=ax[0])
im = ax[1].imshow(minic[0].sum(axis=0))
plt.colorbar(im, ax=ax[1])
im = ax[2].imshow(minic3[0].sum(axis=0))
plt.colorbar(im, ax=ax[2])
dg.save2ptfig(0, [minic, minfin], [ic, fin], None, bs, save=False, retfig=True)
dg.save2ptfig(0, [minic3, minfin3], [ic, fin], None, bs, save=False, retfig=True)
# +
# x0 = np.random.normal(0, 1, nc**3).reshape(fin.shape).astype(np.float32)
# linear = tf.Variable(name='linmesh', shape=(1, nc, nc, nc), dtype=tf.float32,
# initial_value=x0, trainable=True)
# RR = 0
# xinitrecons = []
# for j in range(traindata.shape[0]):
# linear.assign(x0)
# for i in range(5):
# grads = recon_noanneal_grad(linear, tf.constant(traindata[j:j+1, 1], dtype=tf.float32))
# opt.apply_gradients(zip([grads], [linear]))
# minic = linear.numpy().reshape(fin.shape)
# xinitrecons.append(linear.numpy().reshape(fin.shape))
# xinitrecons_train = np.array(xinitrecons)
# +
# x0 = np.random.normal(0, 1, nc**3).reshape(fin.shape).astype(np.float32)
# linear = tf.Variable(name='linmesh', shape=(1, nc, nc, nc), dtype=tf.float32,
# initial_value=x0, trainable=True)
# RR = 0
# xinitrecons2 = []
# for j in range(testdata.shape[0]):
# # for j in range(10):
# linear.assign(x0)
# for i in range(5):
# grads = recon_noanneal_grad(linear, tf.constant(testdata[j:j+1, 1], dtype=tf.float32))
# opt.apply_gradients(zip([grads], [linear]))
# minic = linear.numpy().reshape(fin.shape)
# xinitrecons2.append(linear.numpy().reshape(fin.shape))
# xinitrecons_test = np.array(xinitrecons2)
# +
# kvecreal = tools.fftk((nc, nc, nc), bs)
# xinitrecons_train = np.array([tools.gauss(i, kvecreal, 2)
# for i in xinitrecons_train[:,0]]).astype(float32)
# xinitrecons_train = np.expand_dims(xinitrecons_train, 1)
# xinitrecons_test = np.array([tools.gauss(i, kvecreal, 2)
# for i in xinitrecons_test[:, 0]]).astype(float32)
# xinitrecons_test = np.expand_dims(xinitrecons_test, 1)
# +
# jj = 200
# # dg.save2ptfig(0, [xinitrecons_test[jj], minfin], [testdata[jj:jj+1, 0], fin], None, bs, save=False, retfig=True)
# dg.save2ptfig(0, [xinitrecons_train[jj], minfin], [traindata[jj:jj+1, 0], fin], None, bs, save=False, retfig=True)
# # plt.imshow(xinitrecons[1][0].sum(axis=0))
# -
# # Code RIM
# +
# class myRIM3D(tf.keras.Model):
# def __init__(self, cell, output_layer, input_layer, lrgrad, niter):
# super(myRIM3D, self).__init__()
# self.cell = cell
# self.output_layer = output_layer
# self.input_layer = input_layer
# self.lrgrad = lrgrad
# self.niter = niter
# # def call(self, x_init, y, grad_fn, outputs_ta, states_ta, grad_args=[], initstates = None, return_steps=False):
# def call(self, x_init, y, grad_fn, grad_args=[], initstates = None, return_steps=False):
# outputs_ta = tf.TensorArray(size=self.niter+1, dtype=tf.float32)
# states_ta = tf.TensorArray(size=self.niter+1, dtype=tf.float32)
# if initstates is None:
# # stateshape = x_init.shape[:-1] + tuple([self.cell.output_size])
# stateshape = x_init.shape + tuple([self.cell.filters])
# initstates = [tf.zeros(stateshape), tf.zeros(stateshape)]
# i = tf.constant(0, dtype=tf.int32)
# curr_state = initstates
# curr_pos = x_init
# def body(i, pos, states):
# gradient = grad_fn(pos, y, *grad_args)
# gradient = gradient * self.lrgrad
# concat_input = tf.stack([pos, gradient], axis=-1)
# cell_input = self.input_layer(concat_input)
# delta_pos, new_state = self.cell(cell_input, states)
# delta_pos = self.output_layer(delta_pos)[...,0]
# new_pos = pos + delta_pos
# return i +1 , new_pos, new_state
# while tf.less(i, tf.constant(self.niter)):
# outputs_ta = outputs_ta.write(i, curr_pos)
# states_ta = states_ta.write(i, curr_state)
# i, curr_pos, curr_state = body(i, curr_pos, curr_state)
# outputs_ta = outputs_ta.write(i, curr_pos)
# states_ta = states_ta.write(i, curr_state)
# return outputs_ta.stack(), states_ta.stack()
# +
class myAdam(tf.keras.Model):
def __init__(self, niter):
super(myAdam, self).__init__()
self.niter = niter
self.beta_1, self.beta_2 = 0.9, 0.999
self.lr, self.eps = 0.1, 1e-7
def call(self, x_init, y, grad_fn, grad_args=[], ):
outputs_ta = tf.TensorArray(size=self.niter+1, dtype=tf.float32)
i = tf.constant(0, dtype=tf.int32)
curr_pos = x_init
m = tf.zeros_like(x_init)
v = tf.zeros_like(x_init)
def body(i, pos, m, v):
gradient = grad_fn(pos, y, *grad_args)
#get_step = self.optimizer.apply_gradients(zip([gradient],[ pos]))
t = tf.cast(i+1, tf.float32)
m = self.beta_1*m + (1-self.beta_1)*gradient
v = self.beta_2*v + (1-self.beta_2)*gradient**2
mc = m/(1-self.beta_1**t)
vc = v/(1-self.beta_2**t)
delta = -1.*self.lr*mc/(np.sqrt(vc) + self.eps)
new_pos = pos + delta
return i +1 , new_pos, m, v
while tf.less(i, tf.constant(self.niter)):
outputs_ta = outputs_ta.write(i, curr_pos)
i, curr_pos, m, v = body(i, curr_pos, m, v)
outputs_ta = outputs_ta.write(i, curr_pos)
return outputs_ta.stack()
# +
class myRIM3D(tf.keras.Model):
def __init__(self, cell, output_layer, input_layer, niter):
super(myRIM3D, self).__init__()
self.cell = cell
self.output_layer = output_layer
self.input_layer = input_layer
self.niter = niter
self.beta_1, self.beta_2 = 0.9, 0.999
self.lr, self.eps = 0.1, 1e-7
def call(self, x_init, y, grad_fn, grad_args=[], initstates = None, return_steps=False):
outputs_ta = tf.TensorArray(size=self.niter+1, dtype=tf.float32)
states_ta = tf.TensorArray(size=self.niter+1, dtype=tf.float32)
if initstates is None:
stateshape = x_init.shape + tuple([self.cell.filters])
initstates = [tf.zeros(stateshape), tf.zeros(stateshape)]
i = tf.constant(0, dtype=tf.int32)
curr_state = initstates
curr_pos = x_init
m = tf.zeros_like(x_init)
v = tf.zeros_like(x_init)
def body(i, pos, states, m, v):
gradient = grad_fn(pos, y, *grad_args)
t = tf.cast(i+1, tf.float32)
m = self.beta_1*m + (1-self.beta_1)*gradient
v = self.beta_2*v + (1-self.beta_2)*gradient**2
mc = m/(1-self.beta_1**t)
vc = v/(1-self.beta_2**t)
delta = -1.*self.lr*mc/(tf.sqrt(vc) + self.eps)
concat_input = tf.stack([pos, delta], axis=-1)
cell_input = self.input_layer(concat_input)
delta_pos, new_state = self.cell(cell_input, states)
delta_pos = self.output_layer(delta_pos)[...,0]
new_pos = pos + delta_pos
return i +1 , new_pos, new_state, m, v
while tf.less(i, tf.constant(self.niter)):
outputs_ta = outputs_ta.write(i, curr_pos)
states_ta = states_ta.write(i, curr_state)
i, curr_pos, curr_state, m, v = body(i, curr_pos, curr_state, m, v)
outputs_ta = outputs_ta.write(i, curr_pos)
states_ta = states_ta.write(i, curr_state)
return outputs_ta.stack(), states_ta.stack()
# +
rim_iter = 10
input_size = 8
cell_size = 8
input_layer = Conv3D(input_size, kernel_size=5, trainable=True, padding='SAME',
input_shape=(None, nc, nc, nc, 2))
cell = ConvLSTM3DCell(cell_size, kernel_size=5, padding='SAME')
cell.build(input_shape=[None, nc, nc, nc, input_size])
output_layer = Conv3D(1, kernel_size=5, trainable=True, padding='SAME',
input_shape=(None, nc, nc, nc, cell_size))
idx = np.random.randint(0, traindata.shape[0], 1)
xx, yy = traindata[idx, 0].astype(float32), traindata[idx, 1].astype(float32),
x_init = np.random.normal(size=xx.size).reshape(xx.shape).astype(float32)
# stateshape = x_init.shape + tuple([cell.filters])
# initstates = [tf.zeros(stateshape), tf.zeros(stateshape)]
# cell(tf.constant(tf.stack([xx, xx], axis=-1)), initstates);
####
testrim = myRIM3D(cell, output_layer, input_layer, niter=rim_iter)
out, states = testrim(x_init=tf.constant(xx),
y=tf.constant(yy),
grad_fn=recon_noanneal_grad)
# +
# savepath = './rim-models/L%04d_N%03d_T%02d/'%(bs, nc, nsteps)
# print(savepath)
# testrim.load_weights(savepath)
# -
kvecreal = tools.fftk((nc, nc, nc), bs)
train_xinits = np.array([tools.gauss(i/i.mean()-1, kvecreal, bs/nc) for i in traindata[:, 1]]).astype(float32)
test_xinits = np.array([tools.gauss(i/i.mean()-1, kvecreal, bs/nc) for i in testdata[:, 1]]).astype(float32)
def test_callback():
idx = np.random.randint(0, testdata.shape[0], 1)
xx, yy = testdata[idx, 0].astype(float32), testdata[idx, 1].astype(float32),
# x_init = xinitrecons_test[idx, 0]
x_init = np.random.normal(size=xx.size).reshape(xx.shape).astype(float32)
out, states = testrim(x_init=tf.constant(x_init),
y=tf.constant(yy),
grad_fn=recon_noanneal_grad)
out = out.numpy()
fig, ax = plt.subplots(1, 3, figsize = (12, 4))
vmin, vmax = xx[0].sum(axis=0).min(), xx[0].sum(axis=0).max()
ax[0].imshow(xx[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[0].set_title('Truth')
ax[1].imshow(x_init[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[1].set_title('initial point')
ax[2].imshow(out[-1][0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[2].set_title('RIM 10 step')
plt.show()
plt.close()
##
fig, ax = plt.subplots(1, 2, figsize=(9, 4))
k, pks = get_ps([out[0], pm(out[0]).numpy()], [xx, yy])
for i in range(2):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d--'%i)
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d--'%i)
k, pks = get_ps([out[-1], pm(out[-1]).numpy()], [xx, yy])
for i in range(2):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d'%i)
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d'%i)
for axis in ax:
axis.semilogx()
axis.grid(which='both')
plt.show()
plt.close()
# # Train
#
# +
losses = []
# +
piter = 1
testiter = 1
start = time.time()
# Create an optimizer for Adam.
iterweights = np.arange(rim_iter+1)/rim_iter *0. + 1.
iterweights = iterweights.reshape(-1, 1, 1, 1, 1).astype(float32)
# @tf.function
def rim_train(x_true, x_init, y):
with tf.GradientTape() as tape:
x_pred, states = testrim(x_init, y, recon_noanneal_grad)
res = (x_true - x_pred)
loss = tf.reduce_mean(iterweights*tf.square(res))
gradients = tape.gradient(loss, testrim.trainable_variables)
return loss, gradients
lrs = [0.001, 0.0005, 0.0001]
liters = [201, 101, 1001]
trainiter = 0
for il in range(1):
print('Learning rate = %0.3e'%lrs[il])
opt = tf.keras.optimizers.Adam(learning_rate=lrs[il])
for i in range(liters[il]):
idx = np.random.randint(0, traindata.shape[0], 8)
xx, yy = traindata[idx, 0].astype(float32), traindata[idx, 1].astype(float32),
x_init = np.random.normal(size=xx.size).reshape(xx.shape).astype(float32)
#x_init = train_xinits[idx]
#x_init = xinitrecons_train[idx, 0]
loss, gradients = rim_train(x_true=tf.constant(xx),
x_init=tf.constant(x_init),
y=tf.constant(yy))
losses.append(loss.numpy())
opt.apply_gradients(zip(gradients, testrim.trainable_variables))
if i%piter == 0:
print("Time taken for %d iterations : "%piter, time.time() - start)
print("Loss at iteration %d : "%i, losses[-1])
start = time.time()
if i%testiter == 0:
plt.plot(losses)
plt.show()
plt.close()
test_callback()
# savepath = './rim-models/L%04d_N%03d_T%02d-c8/iter-%04d'%(bs, nc, nsteps, trainiter)
# testrim.save_weights(savepath, overwrite=False)
trainiter += 1
# +
# gg = recon_noanneal_grad(tf.constant(np.random.normal(size=yy.size).reshape(yy.shape).astype(float32)), tf.constant(yy)).numpy()
# (gg.max(), gg.min(), gg.mean(), gg.std())
# -
plt.plot(losses)
# plt.semilogx()
plt.loglog()
plt.grid(which='both')
# +
# idx = np.random.randint(0, traindata.shape[0], 1)
# xx, yy = traindata[idx, 0].astype(float32), traindata[idx, 1].astype(float32),
# x_init = xinitrecons_train[idx, 0]
idx = np.random.randint(0, testdata.shape[0], 1)
xx, yy = testdata[idx, 0].astype(float32), testdata[idx, 1].astype(float32),
# x_init = yy.copy() - 1
# x_init = pred2.copy()
# x_init = xinitrecons_test[idx, 0]
x_init = np.random.normal(size=xx.size).reshape(xx.shape).astype(float32)
adam = myAdam(rim_iter)
pred2 = adam(x_init=tf.constant(x_init),
y=tf.constant(yy),
grad_fn=recon_noanneal_grad)[-1].numpy()
adam = myAdam(rim_iter*10)
pred3 = adam(x_init=tf.constant(x_init),
y=tf.constant(yy),
grad_fn=recon_noanneal_grad)[-1].numpy()
out, states = testrim(x_init=tf.constant(x_init),
y=tf.constant(yy),
grad_fn=recon_noanneal_grad)
out = out.numpy()
pred = out[-1]
cstate = out[1][:, 0]
# -
fig, ax = plt.subplots(1, 4, figsize = (12, 4))
vmin, vmax = xx[0].sum(axis=0).min(), xx[0].sum(axis=0).max()
print(vmin, vmax)
# vmin, vmax=None,None
ax[0].imshow(xx[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[0].set_title('Truth')
ax[1].imshow(x_init[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[1].set_title('initial point')
ax[2].imshow(pred[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[2].set_title('RIM %d step'%rim_iter)
ax[3].imshow(pred3[0].sum(axis=0), vmin=vmin, vmax=vmax)
ax[3].set_title('Adam %d step'%(10*rim_iter))
# +
fig, ax = plt.subplots(1, 2, figsize=(9, 4))
k, pks = get_ps([out[0], pm(out[0]).numpy()], [xx, yy])
for i in range(1):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d--'%i, label='Init')
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d--'%i)
k, pks = get_ps([out[-1], pm(out[-1]).numpy()], [xx, yy])
for i in range(1):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d'%i, label='RIM-%d'%rim_iter)
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d'%i)
k, pks = get_ps([pred2, pm(pred2).numpy()], [xx, yy])
for i in range(1):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d:'%i, label='Adam-%d'%rim_iter)
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d:'%i)
k, pks = get_ps([pred3, pm(pred3).numpy()], [xx, yy])
for i in range(1):
ax[0].plot(k, pks[i][2]/(pks[i][0]*pks[i][1])**0.5, 'C%d-.'%i, label='Adam-%d'%(10*rim_iter))
ax[1].plot(k, (pks[i][0]/pks[i][1])**0.5, 'C%d-.'%i)
for axis in ax:
axis.semilogx()
axis.grid(which='both')
axis.legend()
# axis.legend(bbox_to_anchor=(2.6, 1))
ax[0].set_title('Cross Correlation')
ax[1].set_title('transfer Function')
# +
# savepath = './rim-models/L%04d_N%03d_T%02d/'%(bs, nc, nsteps)
# print(savepath)
# testrim.save_weights(savepath, overwrite=False)
# testrim2 = myRIM3D(cell, output_layer, input_layer, lrgrad, niter=rim_iter, optimizer=None)
# testrim2.load_weights(savepath)
# +
# testrim2 = myRIM3D(cell, output_layer, input_layer, lrgrad, niter=rim_iter, optimizer=None)
# testrim2.load_weights(savepath)
# -
# for i in range(out.shape[0]):
# im = plt.imshow(out[i, 0].sum(axis=0))
# plt.colorbar(im)
# plt.title(i)
# plt.show()
# +
# gradlist = []
# gradlist.append(recon_noanneal_grad(tf.constant(xx), tf.constant(yy)))
# for i in range(niter):
# gradlist.append(recon_noanneal_grad(tf.constant(out[i]), tf.constant(yy)))
# +
def summary(x):
print(x.min(), x.max(), x.mean(), x.std())
for j in gradlist:summary(j.numpy())
# -
# plt.hist(gradlist[0].numpy().flatten(), alpha=0.5, density=True)
plt.hist(out[0].flatten(), alpha=0.5, density=True)
plt.hist(out[1].flatten(), alpha=0.5, density=True)
plt.show()
plt.hist(gradlist[1].numpy().flatten(), alpha=0.5, density=True)
plt.hist(gradlist[-1].numpy().flatten(), alpha=0.5, density=True)
for j in out:summary(j)
|
notebooks/Cosmic_Rim_DM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import numpy as np
from random import randint
from tensorflow.examples.tutorials.mnist import input_data
# ### Load MNIST
# Loading mnist data with as one hot encoded
# Please download datasets from http://yann.lecun.com/exdb/mnist/
# Located it under MNIST_data folder of the same directory as this notebook
mnist = input_data.read_data_sets('MNIST_data/', one_hot=True)
# Inspect data
type(mnist)
# Number of examples
mnist.train.num_examples
mnist.test.num_examples
import matplotlib.pyplot as plt
# %matplotlib inline
single_image = mnist.train.images[0].reshape(28, 28)
plt.imshow(single_image, cmap='gray')
single_image.min()
single_image.max()
# ### Parameters
# x is just a placeholder that we want to tell TF to flatten each mnist
# image into a 784-dimensional vector
# 1. Placeholders
x = tf.placeholder(tf.float32, [None, 784])
# Weight is equivalent to 784 images and 10 possible labels
# We are going to learn W and b. It doesn't really matter what they
# are initially
# 2. Variables
W = tf.Variable(tf.zeros([784, 10]))
b = tf.Variable(tf.zeros([10]))
# 3. softmax model
y = tf.nn.softmax(tf.matmul(x, W) + b)
# 4. loss function
# In ML, we typically define what it means for a model to be bad. We call
# this "cost" or "loss". It represents how far off our model is from our
# https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits
y_true = tf.placeholder(tf.float32, [None, 10])
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=y_true, logits=y))
# 5. Optimizer
# Below code automatically performs backpropogation
optimizer = tf.train.GradientDescentOptimizer(0.5)
train_step = optimizer.minimize(cross_entropy)
# 6. Create session
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
# Actual training
for step in range(1000):
batch_xs, batch_ys = mnist.train.next_batch(100)
sess.run(train_step, feed_dict={x: batch_xs, y_true: batch_ys})
# Evaluating model
# We are using softmax: getting highest y probability
# Below basically checks which ones are matching
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(y_true, 1))
# Booleans can be casted to integers T = 1 F = 0
# Get average of them
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# We will get 2 results
# y could be [3, 4] when y_true (testing) can be [3, 9]
print('Checking accuracy x = mnist test images')
print('y_ mnist test labels')
print(sess.run(accuracy,
feed_dict={x: mnist.test.images, y_true: mnist.test.labels}))
# Running some predictions
num = randint(0, mnist.test.images.shape[0])
img1 = mnist.test.images[num]
prediction = sess.run(tf.arg_max(y, 1), feed_dict={x: [img1]})
plt.imshow(img1.reshape(28, 28))
print('NN predicted', prediction[0])
|
009-cnn/MNIST in Tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Calculate the skill of a MJO Index as a function of lead time for Weekly Data
# ### In this example, we demonstrate:
# 1. How to remotely access data from the Subseasonal Experiment (SubX) hindcast database and set it up to be used in `climpred`.
# 2. How to calculate the Anomaly Correlation Coefficient (ACC) using weekly data with `climpred`
# 3. How to calculate and plot historical forecast skill of the real-time multivariate MJO (RMM) indices as function of lead time.
# ### The Subseasonal Experiment (SubX)
#
# Further information on SubX is available from [Pegion et al. 2019](https://journals.ametsoc.org/doi/full/10.1175/BAMS-D-18-0270.1) and the [SubX project website](http://cola.gmu.edu/subx/)
#
# The SubX public database is hosted on the International Research Institute for Climate and Society (IRI) data server http://iridl.ldeo.columbia.edu/SOURCES/.Models/.SubX/
#
# Since the SubX data server is accessed via this notebook, the time for the notebook to run may is several minutes and will vary depending on the speed that data can be downloaded. This is a large dataset, so please be patient. If you prefer to download SubX data locally, scripts are available from https://github.com/kpegion/SubX.
# ### Definitions
#
# RMM
# : Two indices (RMM1 and RMM2) are used to represent the MJO. Together they define the MJO based on 8 phases and can represent both the phase and amplitude of the MJO (Wheeler and Hendon 2004). This example uses the observed RMM1 provided by <NAME> at the Center for Australian Weather and Climate Research. It is the version of the indices in which interannual variability has not been removed.
#
# Skill of RMM
# : Traditionally, the skill of the RMM is calculated as a bivariate correlation encompassing the skill of the two indices together (Rashid et al. 2010; Gottschalck et al 2010). Currently, `climpred` does not have the functionality to calculate the bivariate correlation, thus the anomaly correlation coefficient for RMM1 index is calculated here as a demonstration. The bivariate correlation metric will be added in a future version of `climpred`
# +
import warnings
import matplotlib.pyplot as plt
plt.style.use('ggplot')
plt.style.use('seaborn-talk')
import xarray as xr
import pandas as pd
import numpy as np
from climpred import HindcastEnsemble
import climpred
# -
warnings.filterwarnings("ignore")
# Read the observed RMM Indices
obsds = climpred.tutorial.load_dataset('RMM-INTERANN-OBS')['rmm1'].to_dataset()
obsds
# Read the SubX RMM1 data for the GMAO-GEOS_V2p1 model from the SubX data server. It is important to note that the SubX data contains weekly initialized forecasts where the `init` day varies by model. SubX data may have all NaNs for initial dates in which a model does not make a forecast, thus we apply `dropna` over the `S=init` dimension when `how=all` data for a given `S=init` is missing. This can be slow, but allows the rest of the calculations to go more quickly.
#
# Note that we ran the `dropna` operation offline and then uploaded the post-processed SubX dataset to the `climpred-data` repo for the purposes of this demo. This is how you can do this manually:
#
# ```python
# url = 'http://iridl.ldeo.columbia.edu/SOURCES/.Models/.SubX/.GMAO/.GEOS_V2p1/.hindcast/.RMM/.RMM1/dods/'
# fcstds = xr.open_dataset(url, decode_times=False, chunks={'S': 1, 'L': 45}).dropna(dim='S',how='all')
# ```
fcstds = climpred.tutorial.load_dataset('GMAO-GEOS-RMM1', decode_times=False)
fcstds
# The SubX data dimensions correspond to the following `climpred` dimension definitions: `X=lon`,`L=lead`,`Y=lat`,`M=member`, `S=init`. We will rename the dimensions to their `climpred` names.
fcstds=fcstds.rename({'S': 'init','L': 'lead','M': 'member', 'RMM1' : 'rmm1'})
# Let's make sure that the `lead` dimension is set properly for `climpred`. SubX data stores `leads` as 0.5, 1.5, 2.5, etc, which correspond to 0, 1, 2, ... days since initialization. We will change the `lead` to be integers starting with zero.
fcstds['lead'] = (fcstds['lead'] - 0.5).astype('int')
# Now we need to make sure that the `init` dimension is set properly for `climpred`. We use an `xarray` convenience function to decode it into the proper CFTime calendar. It can detect that this is on a 360 day calendar.
fcstds = xr.decode_cf(fcstds, decode_times=True)
fcstds
# Make Weekly Averages
fcstweekly = fcstds.rolling(lead=7, center=False).mean().dropna(dim='lead')
obsweekly = obsds.rolling(time=7, center=False).mean().dropna(dim='time')
print(fcstweekly)
print(obsweekly)
# Create a new `xr.DataArray` for the weekly fcst data
nleads = fcstweekly['lead'][::7].size
fcstweeklyda = xr.DataArray(fcstweekly['rmm1'][:,:,::7],
coords={'init' : fcstweekly['init'],
'member': fcstweekly['member'],
'lead': np.arange(1,nleads+1),
},
dims=['init', 'member','lead'])
fcstweeklyda = fcstweeklyda.rename('rmm1')
# `climpred` requires that `lead` dimension has an attribute called `units` indicating what time units the `lead` is assocated with. Options are: `years,seasons,months,weeks,pentads,days`. The `lead` `units` are `weeks`.
fcstweeklyda['lead'].attrs = {'units': 'weeks'}
# Create the `climpred HindcastEnsemble` object and add the observations.
hindcast = HindcastEnsemble(fcstweeklyda)
hindcast = hindcast.add_observations(obsweekly)
# Calculate the Anomaly Correlation Coefficient (ACC)
skill = hindcast.verify(metric='acc', comparison='e2o', alignment='maximize', dim='init')
# Plot the skill as a function of lead time
x = np.arange(fcstweeklyda['lead'].size)
plt.bar(x, skill['rmm1'])
plt.title('GMAO_GOES_V2p1 RMM1 Skill')
plt.xlabel('Lead Time (Weeks)')
plt.ylabel('ACC')
plt.ylim(0.0, 1.0)
# ### References
#
# <NAME>., <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME>, 2019: The Subseasonal Experiment (SubX): A Multimodel Subseasonal Prediction Experiment. Bull. Amer. Meteor. Soc., 100, 2043–2060, https://doi.org/10.1175/BAMS-D-18-0270.1
#
# 2. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., … <NAME>. (2017). The Subseasonal Experiment (SubX) [Data set]. IRI Data Library. https://doi.org/10.7916/D8PG249H
#
# 3. <NAME>., & <NAME>. (2004). An all-season real-time multivariate MJO index: Development of an index for monitoring and prediction. Monthly Weather Review, 132(8), 1917–1932. http://doi.org/10.1175/1520-0493(2004)132<1917:AARMMI>2.0.CO;2
#
# 4. <NAME>., <NAME>., <NAME>., & <NAME>. (2010). Prediction of the Madden–Julian oscillation with the POAMA dynamical prediction system. Climate Dynamics, 36(3-4), 649–661. http://doi.org/10.1007/s00382-010-0754-x
#
# 5. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., et al. (2010). A Framework for Assessing Operational Madden–Julian Oscillation Forecasts: A CLIVAR MJO Working Group Project. Bulletin of the American Meteorological Society, 91(9), 1247–1258. http://doi.org/10.1175/2010BAMS2816.1
|
docs/source/examples/subseasonal/weekly-subx-example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item"><li><span><a href="#Video-Numbers" data-toc-modified-id="Video-Numbers-1"><span class="toc-item-num">1 </span>Video Numbers</a></span></li><li><span><a href="#Gender-Balance" data-toc-modified-id="Gender-Balance-2"><span class="toc-item-num">2 </span>Gender Balance</a></span><ul class="toc-item"><li><span><a href="#Face-Size" data-toc-modified-id="Face-Size-2.1"><span class="toc-item-num">2.1 </span>Face Size</a></span></li></ul></li><li><span><a href="#Donald-Trump-and-Hillary-Clinton" data-toc-modified-id="Donald-Trump-and-Hillary-Clinton-3"><span class="toc-item-num">3 </span>Donald Trump and Hillary Clinton</a></span></li><li><span><a href="#James-Comey" data-toc-modified-id="James-Comey-4"><span class="toc-item-num">4 </span>James Comey</a></span></li><li><span><a href="#Shootings" data-toc-modified-id="Shootings-5"><span class="toc-item-num">5 </span>Shootings</a></span></li></ul></div>
# -
from esper.widget import *
from esper.prelude import *
from esper.spark_util import *
from esper.validation import *
from esper.plot_util import *
from esper.spark_identity import get_screen_time_by_canonical_show_spark
from esper.identity_clusters import identity_clustering_workflow
from esper.major_canonical_shows import MAJOR_CANONICAL_SHOWS
from datetime import timedelta, datetime
from collections import OrderedDict
import random
# # Video Numbers
videos = get_videos()
print('Total seconds of 3 years data')
videos.where(videos.threeyears_dataset == True).select('duration').groupBy().sum().collect()
# # Gender Balance
faces = get_faces()
face_genders = get_face_genders()
face_genders = face_genders.where(face_genders.labeler_id != Labeler.objects.get(name='handlabeled-gender').id)
# speakers = get_speakers()
# +
face_genders_nh = face_genders.where(face_genders.host_probability < 0.5)
screen_time_male, screen_time_female = OrderedDict(), OrderedDict()
screen_time_nh_male, screen_time_nh_female = OrderedDict(), OrderedDict()
print('Computing screen time across all channels')
screen_time_male['All Channels'] = sum_over_column(
face_genders, 'duration', probability_column='male_probability'
)
screen_time_female['All Channels'] = sum_over_column(
face_genders, 'duration', probability_column='female_probability'
)
print('Computing screen time across all channels (non-hosts)')
screen_time_nh_male['All Channels'] = sum_over_column(
face_genders_nh, 'duration', probability_column='male_probability'
)
screen_time_nh_female['All Channels'] = sum_over_column(
face_genders_nh, 'duration', probability_column='female_probability'
)
print('Computing screen time by channel')
channel_id_map = {c.id : c.name for c in Channel.objects.all()}
for k, v in sum_over_column(face_genders, 'duration', ['channel_id'],
probability_column='male_probability').items():
screen_time_male[channel_id_map[k[0]]] = v
for k, v in sum_over_column(face_genders, 'duration', ['channel_id'],
probability_column='female_probability').items():
screen_time_female[channel_id_map[k[0]]] = v
print('Computing screen time by channel (non-host)')
for k, v in sum_over_column(face_genders_nh, 'duration', ['channel_id'],
probability_column='male_probability').items():
screen_time_nh_male[channel_id_map[k[0]]] = v
for k, v in sum_over_column(face_genders_nh, 'duration', ['channel_id'],
probability_column='female_probability').items():
screen_time_nh_female[channel_id_map[k[0]]] = v
# +
# speaking_time_male, speaking_time_female = OrderedDict(), OrderedDict()
# gender_id_map = {g.id : g.name for g in Gender.objects.all()}
# for x in speakers.groupBy('gender_id').sum('duration').collect():
# gender_id = x['gender_id']
# value = x['sum(duration)']
# if gender_id_map[gender_id] == 'M':
# speaking_time_male['All Channels'] = (value, 0)
# elif gender_id_map[gender_id] == 'F':
# speaking_time_female['All Channels'] = (value, 0)
# channel_id_map = {c.id : c.name for c in Channel.objects.all()}
# for x in speakers.groupBy('gender_id', 'channel_id').sum('duration').collect():
# channel_id, gender_id = x['channel_id'], x['gender_id']
# value = x['sum(duration)']
# if gender_id_map[gender_id] == 'M':
# speaking_time_male[channel_id_map[channel_id]] = (value, 0)
# elif gender_id_map[gender_id] == 'F':
# speaking_time_female[channel_id_map[channel_id]] = (value, 0)
# -
sort_order = ['All Channels'] + [c.name for c in Channel.objects.all().order_by('name')]
plot_binary_proportion_comparison(
['Male', 'Female'], [screen_time_male, screen_time_female],
'', '', 'Proportion of Screen Time',
raw_data_to_label_fn=None, figsize=(5,5), sort_order=sort_order,
legend_loc=4,
save_path='figures/gender-3y-screen-time.pdf'
)
plot_binary_proportion_comparison(
['Male', 'Female'], [screen_time_nh_male, screen_time_nh_female],
'', '', 'Proportion of Screen Time (Excluding Hosts)',
raw_data_to_label_fn=None, figsize=(5,5), sort_order=sort_order,
legend_loc=4,
save_path='figures/gender-3y-screen-time-no-host.pdf'
)
# plot_binary_proportion_comparison(
# ['Male', 'Female'], [speaking_time_male, speaking_time_female],
# '', '', 'Proportion of Speaking Time',
# raw_data_to_label_fn=None, figsize=(5,5), sort_order=sort_order,
# legend_loc=4,
# save_path='figures/gender-3y-speaking-time.pdf'
# )
canonical_show_map = {c.id : c.name for c in CanonicalShow.objects.all() if c.name in MAJOR_CANONICAL_SHOWS}
screen_time_male_by_show, screen_time_female_by_show = {}, {}
for k, v in sum_over_column(face_genders, 'duration', ['canonical_show_id'],
probability_column='male_probability').items():
if k[0] in canonical_show_map:
screen_time_male_by_show[canonical_show_map[k[0]]] = v
for k, v in sum_over_column(face_genders, 'duration', ['canonical_show_id'],
probability_column='female_probability').items():
if k[0] in canonical_show_map:
screen_time_female_by_show[canonical_show_map[k[0]]] = v
plot_binary_proportion_comparison(
['Male', 'Female'], [screen_time_male_by_show, screen_time_female_by_show],
'Gender Distribution of Screen Time vs. Show', '', 'Proportion of Screen Time',
raw_data_to_label_fn=None, legend_loc=4,
baseline_series_names=[
'Baseline Male (Entire Dataset)',
'Baseline Female (Entire Dataset)'
],
baseline_data=[
screen_time_male['All Channels'][0],
screen_time_female['All Channels'][0]
],
save_path='figures/gender-3y-screen-time-by-show.pdf'
)
show_order = [
(k, screen_time_male_by_show[k][0]/screen_time_female_by_show[k][0])
for k in screen_time_male_by_show
]
show_order.sort(key=lambda x: x[1])
canonical_show_rmap = { v : k for k, v in canonical_show_map.items()}
def face_img(face):
return crop(load_frame(face.person.frame.video, face.person.frame.number, []), face)
def get_face_for_show(canonical_show_name, n_samples=10, dim=(100, 100)):
print('Loading host faces:', canonical_show_name,
[x.name for x in CanonicalShow.objects.filter(name=canonical_show_name)[0].hosts.all()])
host_faces = faces.where(
(faces.canonical_show_id == canonical_show_rmap[canonical_show_name]) &
(faces.host_probability >= 0.9)
).sample(0.1, False).select('id').limit(n_samples)
host_face_ids = [h['id'] for h in host_faces.collect()]
host_face_imgs = par_for(face_img, list(Face.objects.filter(id__in=host_face_ids)), progress=False)
host_face_imgs = [cv2.resize(im, dim) for im in host_face_imgs]
mean_host_im = np.zeros(host_face_imgs[0].shape)
for im in host_face_imgs:
mean_host_im += im
mean_host_im /= len(host_face_imgs)
mean_host_im = mean_host_im.astype(np.uint8)
return mean_host_im
# show_imgs = [get_face_for_show(x[0]) for x in show_order]
show_imgs = par_for(get_face_for_show, [x[0] for x in show_order], workers=4)
assert len(show_order) == len(show_imgs)
host_montage_img = np.hstack(show_imgs)
plt.imsave('figures/show-host-avgs.jpg', cv2.cvtColor(host_montage_img, cv2.COLOR_RGB2BGR))
imshow(host_montage_img)
plt.show()
screen_time_nh_male_by_show, screen_time_nh_female_by_show = {}, {}
for k, v in sum_over_column(face_genders_nh, 'duration', ['canonical_show_id'],
probability_column='male_probability').items():
if k[0] in canonical_show_map:
screen_time_nh_male_by_show[canonical_show_map[k[0]]] = v
for k, v in sum_over_column(face_genders_nh, 'duration', ['canonical_show_id'],
probability_column='female_probability').items():
if k[0] in canonical_show_map:
screen_time_nh_female_by_show[canonical_show_map[k[0]]] = v
plot_binary_proportion_comparison(
['Male (Excl. Hosts)', 'Female (Excl. Hosts)'], [screen_time_nh_male_by_show, screen_time_nh_female_by_show],
'Gender Distribution of Screen Time (Excluding Hosts) vs. Show', '', 'Proportion of Screen Time',
tertiary_series_names=['Male (Incl. Hosts)', 'Female (Incl. Hosts)'],
tertiary_data=[screen_time_male_by_show, screen_time_female_by_show],
# baseline_data=[
# screen_time_male['All Channels'][0],
# screen_time_female['All Channels'][0]
# ],
# baseline_series_names=[
# 'Baseline Male (Entire Dataset)',
# 'Baseline Female (Entire Dataset)'
# ],
raw_data_to_label_fn=None, legend_loc=4,
save_path='figures/gender-3y-screen-time-by-show-no-host.pdf'
)
# +
# speaking_time_male_by_show, speaking_time_female_by_show = {}, {}
# gender_id_map = {g.id : g.name for g in Gender.objects.all()}
# for x in speakers.groupBy('gender_id', 'canonical_show_id').sum('duration').collect():
# canonical_show_id, gender_id = x['canonical_show_id'], x['gender_id']
# value = x['sum(duration)']
# if canonical_show_id in canonical_show_map:
# if gender_id_map[gender_id] == 'M':
# speaking_time_male_by_show[canonical_show_map[canonical_show_id]] = (value, 0)
# elif gender_id_map[gender_id] == 'F':
# speaking_time_female_by_show[canonical_show_map[canonical_show_id]] = (value, 0)
# +
# plot_binary_proportion_comparison(
# ['Male (Speaking Time)', 'Female (Speaking Time)'], [speaking_time_male_by_show, speaking_time_female_by_show],
# 'Gender Distribution of Speaking Time vs. Show', '',
# 'Proportion of Speaking Time',
# tertiary_series_names=['Male (Screen Time)', 'Female (Screen Time)'],
# tertiary_data=[screen_time_male_by_show, screen_time_female_by_show],
# raw_data_to_label_fn=None, legend_loc=4,
# save_path='figures/gender-3y-speaking-time-by-show.pdf'
# )
# + hide_input=false
topic_to_male_screentime, topic_to_female_screentime = ({'abortion': (518294.8301809259, 298815.40602040046),
'affair': (39389.31232893543, 55525.5885530475),
'afghanistan': (1085300.3048727212, 367695.11064195103),
'africa': (295676.98296886904, 95894.06877211915),
'african american': (384330.71684086934, 226270.95589998108),
'agriculture': (6725.373544278547, 3502.248767418472),
'america': (217756.6907928133, 166775.28563761502),
'apple': (180976.48606021717, 66734.37602145101),
'artificial intelligence': (31129.98012259763, 9780.876014142159),
'asia': (1141605.630527936, 372510.1111205174),
'asthma': (339359.0225043069, 83964.431388244),
'autism': (27679.673446264405, 13107.08559609269),
'bankruptcy': (78837.6634167312, 38478.61896356275),
'banks': (96550.74731097101, 66005.33038593784),
'baseball': (274091.54061841004, 151762.4966443527),
'basketball': (180844.9841464684, 92335.38846694138),
'billy bush': (101162.10326908741, 32683.306271344332),
'black lives': (422790.7060048257, 239574.11285279744),
'blockchain': (54516.771888660536, 25852.396131177804),
'book': (317767.23727690626, 81465.36453893088),
'border wall': (790434.1866875102, 349148.14352520194),
'brazil': (87184.90596125394, 54268.0535382443),
'canada': (153524.18500008367, 64193.99272585327),
'cancer': (1180438.1808235142, 316671.52705781977),
'cars': (6670.888493899268, 3320.775273009518),
'catholic': (330303.729550506, 258150.76543767352),
'cats': (28979.570387077274, 9214.089366527223),
'charity': (204798.86084425222, 88472.18198183207),
'charleston church': (111839.93039077295, 59441.16777088708),
'char<NAME>': (664279.8498326603, 316250.8262504142),
'childcare': (21174.53600788235, 12025.711612799776),
'christianity': (79486.77973743765, 75996.69661718622),
'christmas': (146824.5019747962, 74670.01100888346),
'coal': (439630.0139468445, 597247.3653580876),
'college': (406424.2814510059, 229315.1478924913),
'collusion': (1883202.7915484093, 702020.2183103692),
'confederate': (547664.2396951914, 320125.15948140493),
'congress': (86492.74350965022, 28368.94260304476),
'conservatives': (15779.808671273817, 2120.783371604514),
'constitution': (661345.1631899055, 368509.1119868251),
'crime': (511392.745484412, 332953.7455574636),
'cybersecurity': (6676.556084072748, 7972.303337511483),
'dance': (298808.04996236047, 79660.52444297649),
'death penalty': (677261.8199198114, 396053.72422354115),
'democrat': (93644.42535069052, 33436.49167212691),
'disease': (1241021.5409390433, 320389.951661171),
'dogs': (27639.45919906519, 9292.852945616294),
'doping': (46038.52470131569, 32823.89824011052),
'drugs': (22167.191705473677, 18825.741879678426),
'due process': (60834.17494915206, 31431.33060034904),
'earthquake': (95325.66280539987, 71778.51328922086),
'ebola': (27508.322859652657, 9152.448851562669),
'economy': (523535.7784714372, 243115.91057231784),
'education': (523217.5222435232, 531396.0927771898),
'elderly': (13086.991796611876, 10829.243667587816),
'email scandal': (489111.91803879297, 244568.81745981754),
'england': (503772.09215224726, 390284.7233907992),
'europe': (645093.6320127008, 282198.62628515693),
'evangelical': (1119264.1791265535, 421467.65224499255),
'export': (152956.37668341794, 57426.24304025674),
'facebook': (207257.28808802023, 66669.78404600148),
'fashion': (12073.615677556301, 4569.36211837889),
'fiction': (9011.51131970374, 888.9546009303755),
'fifa': (57134.20354719761, 23155.02816213702),
'film': (110180.45290461865, 26896.26335491689),
'flint water crisis': (139064.247302382, 123357.02550061484),
'flood': (725769.3606071043, 690067.0115616816),
'flu': (981581.2873386245, 260143.71532067092),
'food': (597512.7336067666, 186632.5373093995),
'football': (806767.714916063, 368496.9179760397),
'foreign policy': (438097.7061503113, 225208.58995285016),
'france': (799444.860076679, 416621.4583016661),
'gaming': (52664.145851538306, 11925.102502076577),
'gay marriage': (445052.66437707335, 222697.5446355079),
'global warming': (176237.54147089276, 66528.90981610242),
'golf': (197289.95600991137, 67936.83672408364),
'google': (279731.0195125615, 126812.02263627137),
'government shutdown': (467395.1606375679, 151967.62695414774),
'greece': (284450.46758898965, 162047.09708287157),
'guns': (500331.44329546514, 486255.16277422244),
'gym': (171519.5102262545, 58003.7975328875),
'harassment': (951201.1457089252, 398445.83589243225),
'healthcare': (445370.4566424923, 215268.4113285826),
'hispanic': (394998.6033874645, 179724.09385037038),
'homeless': (7703.338247014193, 14164.740674391203),
'hurricane': (1119939.5294325713, 999759.5140730275),
'immigration': (956091.5229206171, 374548.43553251063),
'import': (89074.71619006211, 29196.217885753063),
'india': (105168.25226349467, 45677.08877557278),
'inequality': (265201.71720924275, 125082.30103330349),
'insurance': (2567264.416022875, 1252652.6125356432),
'internet': (357075.60398928344, 117871.43410260581),
'ipo': (912.390000118204, 210.38835366358538),
'iran': (2459518.6824852405, 1003335.196849234),
'iraq': (2212650.0617190553, 874092.7053971062),
'irs': (332584.19550720387, 132787.31971463218),
'isis': (2866859.4547577044, 1051335.1956239496),
'italy': (130406.6039365884, 61941.45894194005),
'jobs': (259050.80016937104, 173634.2135336672),
'july 4th': (83770.649230375, 18840.457850960207),
'lgbt': (148333.08406840145, 110610.45807212126),
'liberals': (1678.2685757006377, 992.5275335888579),
'literature': (2183.8384368099373, 870.4820204732554),
'marijuana': (97301.49001864342, 27619.83956097895),
'media': (174387.86794308136, 58546.37136148416),
'medicine': (2217611.8469176283, 587787.3507599491),
'meme': (1719.4403447382801, 588.5889830953082),
'mexico': (796371.8589637144, 359104.4681115343),
'music': (291002.41952292674, 113761.1324329917),
'nafta': (427332.8673591054, 353616.4583772168),
'nasa': (66151.28684679992, 42291.675013611624),
'national security': (2010769.6860332717, 759818.8567079846),
'native american': (204855.34665983362, 104866.3068458164),
'nato': (642917.8392674389, 244783.97384628997),
'netflix': (134883.26744098682, 36433.52607757323),
'north korea': (3202959.60437709, 1123122.893880117),
'nutrition': (628045.8830887104, 185545.78221704502),
'obamacare': (2413043.659276039, 1945216.4639350078),
'obesity': (16415.15427062277, 3235.8299049281313),
'oil': (631741.4814901062, 424325.1603651201),
'olympics': (127185.8273693312, 48469.825327296705),
'oscars': (35686.94462244695, 9156.613448253622),
'paris attacks': (148227.55932958846, 52373.97777546179),
'paris climate': (268291.2096124075, 102148.83588868032),
'patriot act': (124384.22532170473, 35037.581329157416),
'pets': (7647.105723654945, 5693.893238853053),
'planned parenthood': (353526.37934005586, 204356.7247447404),
'police': (2203366.7164791604, 1646933.0268962663),
'poverty': (271345.64368497813, 214743.08579149973),
'preschool': (153.00593562841277, 63.89683154229696),
'prescription drugs': (1100.4071262470281, 1066.1973296028325),
'president': (428491.59224530857, 203522.45838913354),
'prison': (594942.1010088442, 230060.64546214623),
'public transportation': (1679.6862711204346, 409.727894334501),
'pulse nightclub': (164350.70266352687, 79156.20542516006),
'racism': (204531.93570248978, 115717.95808609245),
'refugee': (626117.5072668382, 299180.83991466195),
'renewable energy': (83209.89136117794, 121375.305567503),
'rifle': (391413.0371698212, 235164.9357345435),
'<NAME>': (1212925.1526060633, 499823.0925165218),
'russia': (1564083.5109740235, 576207.0672263282),
'<NAME>': (550743.7456960728, 301999.6689024732),
'school': (1096258.8891856289, 802791.9080621441),
'shooting': (1944700.2649337302, 1301799.5821853601),
'slavery': (98829.04013686963, 59931.693774785395),
'soccer': (119693.66474707646, 59797.43043575313),
'social security': (368803.8865416375, 164399.36280333457),
'sports': (356053.14269370615, 163246.89031341203),
'spy': (314156.34252784937, 195869.4108524138),
'stock market': (445345.1741998368, 195285.34074265425),
'supreme court': (2171946.8484882284, 1184892.2332038668),
'syria': (2529187.136078459, 953596.1678198774),
'taxes': (1096785.9848850819, 520643.22916893923),
'technology': (862951.168139002, 268157.1648271495),
'tennis': (77323.50280748564, 27151.816212049234),
'terrorism': (315358.44025278196, 235902.7938372964),
'trade': (1176996.2522977474, 736659.0348801608),
'travel': (276649.8170080849, 87448.70633821092),
'travel ban': (1225879.3096513962, 511255.4354447183),
'<NAME>': (118247.72912627066, 47049.936429644426),
'twitter': (88031.15570809695, 29435.38320986443),
'vacation': (130426.90833713741, 31574.75246438502),
'vaccine': (1475874.7212220582, 402000.5869401628),
'vegas shooter': (179437.5703041849, 88900.48992088546),
'veteran': (166842.9228767198, 206926.897767934),
'violence': (284870.12544940674, 184745.6608155794),
'wall street': (809717.0702973759, 357506.1589435399),
'<NAME>': (1177897.8184054664, 701703.5025185598),
'wedding': (96797.98378302893, 31697.39640199462),
'wildfire': (153333.28772372342, 63057.126015616355),
'wildlife': (25859.503427824664, 11018.743415659756),
'wine': (116595.00348645038, 35113.58599143231),
'wiretap': (173900.09471656446, 65505.32814705004),
'yoga': (125075.1809051474, 48030.61142450444)},
{'abortion': (394137.7796622501, 298815.40602040046),
'affair': (33424.83214887562, 55525.5885530475),
'afghanistan': (408656.5841496342, 367695.11064195103),
'africa': (133870.00592001973, 95894.06877211915),
'african american': (212751.3089182232, 226270.95589998108),
'agriculture': (2179.1226185509427, 3502.248767418472),
'america': (106172.85024822761, 166775.28563761502),
'apple': (118582.79328579502, 66734.37602145101),
'artificial intelligence': (11692.225082607576, 9780.876014142159),
'asia': (529472.326462387, 372510.1111205174),
'asthma': (208911.96254740923, 83964.431388244),
'autism': (14950.859419935132, 13107.08559609269),
'bankruptcy': (43363.564811497024, 38478.61896356275),
'banks': (58889.63410377698, 66005.33038593784),
'baseball': (120344.44917137954, 151762.4966443527),
'basketball': (86358.26623724845, 92335.38846694138),
'billy bush': (55367.25108526693, 32683.306271344332),
'black lives': (212890.4334346132, 239574.11285279744),
'blockchain': (29623.71766756244, 25852.396131177804),
'book': (162430.27333370328, 81465.36453893088),
'border wall': (325678.33149734064, 349148.14352520194),
'brazil': (60044.6522636375, 54268.0535382443),
'canada': (63470.74025850829, 64193.99272585327),
'cancer': (709945.7739646005, 316671.52705781977),
'cars': (4488.628256184164, 3320.775273009518),
'catholic': (185085.29754325436, 258150.76543767352),
'cats': (17372.287470780593, 9214.089366527223),
'charity': (103965.00779104963, 88472.18198183207),
'charleston church': (61331.51772400852, 59441.16777088708),
'charlie hebdo': (296120.4230068453, 316250.8262504142),
'childcare': (10596.606801593762, 12025.711612799776),
'christianity': (35634.464173139626, 75996.69661718622),
'christmas': (94193.19165216413, 74670.01100888346),
'coal': (205136.28868912396, 597247.3653580876),
'college': (238206.9075046165, 229315.1478924913),
'collusion': (680603.7657088493, 702020.2183103692),
'confederate': (259854.06530981322, 320125.15948140493),
'congress': (34840.407307833855, 28368.94260304476),
'conservatives': (6885.959430160948, 2120.783371604514),
'constitution': (263571.26256918715, 368509.1119868251),
'crime': (224241.9302246303, 332953.7455574636),
'cybersecurity': (2003.9948001447985, 7972.303337511483),
'dance': (192858.05300727446, 79660.52444297649),
'death penalty': (382071.77924602065, 396053.72422354115),
'democrat': (48765.85960092766, 33436.49167212691),
'disease': (758346.2099785351, 320389.951661171),
'dogs': (21395.465725859696, 9292.852945616294),
'doping': (31182.860350402712, 32823.89824011052),
'drugs': (10500.906726291383, 18825.741879678426),
'due process': (26616.714273403795, 31431.33060034904),
'earthquake': (70062.3091665719, 71778.51328922086),
'ebola': (18590.209672879806, 9152.448851562669),
'economy': (240733.6365102104, 243115.91057231784),
'education': (348314.04235440795, 531396.0927771898),
'elderly': (7112.34654172646, 10829.243667587816),
'email scandal': (320724.8044169953, 244568.81745981754),
'england': (324772.03207197663, 390284.7233907992),
'europe': (285343.3946806591, 282198.62628515693),
'evangelical': (591949.1122648374, 421467.65224499255),
'export': (74515.87727053584, 57426.24304025674),
'facebook': (124714.53346686772, 66669.78404600148),
'fashion': (8531.399337458719, 4569.36211837889),
'fiction': (2433.681106055335, 888.9546009303755),
'fifa': (29420.620276626218, 23155.02816213702),
'film': (60314.37292120689, 26896.26335491689),
'flint water crisis': (94531.20631790478, 123357.02550061484),
'flood': (464055.8468003017, 690067.0115616816),
'flu': (594491.6604091169, 260143.71532067092),
'food': (356636.8293561955, 186632.5373093995),
'football': (349997.87941159715, 368496.9179760397),
'foreign policy': (208749.80402653245, 225208.58995285016),
'france': (377832.28440113127, 416621.4583016661),
'gaming': (26477.17863961938, 11925.102502076577),
'gay marriage': (218912.6439312347, 222697.5446355079),
'global warming': (72682.40917975792, 66528.90981610242),
'golf': (117076.84432375521, 67936.83672408364),
'google': (165326.79600295363, 126812.02263627137),
'government shutdown': (239388.00819793387, 151967.62695414774),
'greece': (160314.12573766994, 162047.09708287157),
'guns': (250362.30715498488, 486255.16277422244),
'gym': (103443.31766830666, 58003.7975328875),
'harassment': (710866.3959327155, 398445.83589243225),
'healthcare': (243335.22283698624, 215268.4113285826),
'hispanic': (218985.80688947934, 179724.09385037038),
'homeless': (4003.3140719714556, 14164.740674391203),
'hurricane': (669115.7790759358, 999759.5140730275),
'immigration': (448896.932901871, 374548.43553251063),
'import': (43752.70626572699, 29196.217885753063),
'india': (45961.12938482034, 45677.08877557278),
'inequality': (133947.23941404698, 125082.30103330349),
'insurance': (1357424.2803067847, 1252652.6125356432),
'internet': (193669.97672476326, 117871.43410260581),
'ipo': (682.2352918404176, 210.38835366358538),
'iran': (907072.4941247015, 1003335.196849234),
'iraq': (857960.7909669633, 874092.7053971062),
'irs': (145823.0120336703, 132787.31971463218),
'isis': (1195396.1967271124, 1051335.1956239496),
'italy': (67009.09676411223, 61941.45894194005),
'jobs': (118379.62495575406, 173634.2135336672),
'july 4th': (44805.61710262437, 18840.457850960207),
'lgbt': (80428.90558792154, 110610.45807212126),
'liberals': (565.1051313397401, 992.5275335888579),
'literature': (983.6473823425413, 870.4820204732554),
'marijuana': (43646.84955426275, 27619.83956097895),
'media': (88308.10970122935, 58546.37136148416),
'medicine': (1302902.9791417532, 587787.3507599491),
'meme': (1056.319081354455, 588.5889830953082),
'mexico': (328525.1350302781, 359104.4681115343),
'music': (196913.9075040999, 113761.1324329917),
'nafta': (176286.41522347747, 353616.4583772168),
'nasa': (32422.0901969104, 42291.675013611624),
'national security': (844179.1071599199, 759818.8567079846),
'native american': (121670.72607957237, 104866.3068458164),
'nato': (236272.8959677966, 244783.97384628997),
'netflix': (78248.1919851057, 36433.52607757323),
'north korea': (1427539.758285573, 1123122.893880117),
'nutrition': (409477.65571675915, 185545.78221704502),
'obamacare': (1045522.9912411453, 1945216.4639350078),
'obesity': (11058.287704685883, 3235.8299049281313),
'oil': (331523.08861333, 424325.1603651201),
'olympics': (91414.80226029831, 48469.825327296705),
'oscars': (21550.380369211314, 9156.613448253622),
'paris attacks': (61736.90880521267, 52373.97777546179),
'paris climate': (109756.92522242744, 102148.83588868032),
'patriot act': (50366.86943972336, 35037.581329157416),
'pets': (5881.198013415431, 5693.893238853053),
'planned parenthood': (282592.4308367877, 204356.7247447404),
'police': (1000404.4475515336, 1646933.0268962663),
'poverty': (118757.24954158161, 214743.08579149973),
'preschool': (119.4682051790624, 63.89683154229696),
'prescription drugs': (470.62523945200456, 1066.1973296028325),
'president': (196400.41342703035, 203522.45838913354),
'prison': (294667.4848770419, 230060.64546214623),
'public transportation': (890.4835653827256, 409.727894334501),
'pulse nightclub': (83570.31802382709, 79156.20542516006),
'racism': (88723.21494816082, 115717.95808609245),
'refugee': (287557.25836546073, 299180.83991466195),
'renewable energy': (40226.51137489136, 121375.305567503),
'rifle': (160485.15532267117, 235164.9357345435),
'<NAME>': (698018.300413622, 499823.0925165218),
'russia': (604499.5506876372, 576207.0672263282),
'<NAME>': (250488.22734363307, 301999.6689024732),
'school': (700394.4628330563, 802791.9080621441),
'shooting': (895033.9286932275, 1301799.5821853601),
'slavery': (39166.31655315375, 59931.693774785395),
'soccer': (68540.34492926665, 59797.43043575313),
'social security': (165746.76337494576, 164399.36280333457),
'sports': (155211.0781271143, 163246.89031341203),
'spy': (116208.85233367849, 195869.4108524138),
'stock market': (274174.66170273215, 195285.34074265425),
'supreme court': (1007601.4350286189, 1184892.2332038668),
'syria': (1023902.3067643837, 953596.1678198774),
'taxes': (445381.8626624654, 520643.22916893923),
'technology': (490162.96609522926, 268157.1648271495),
'tennis': (47792.59322187671, 27151.816212049234),
'terrorism': (112485.19605018759, 235902.7938372964),
'trade': (515312.74993782095, 736659.0348801608),
'travel': (154149.9636392286, 87448.70633821092),
'travel ban': (583350.5388637857, 511255.4354447183),
'<NAME>': (60963.2658353575, 47049.936429644426),
'twitter': (54529.19264025148, 29435.38320986443),
'vacation': (69082.22125912484, 31574.75246438502),
'vaccine': (881866.6849174047, 402000.5869401628),
'vegas shooter': (92242.2866527717, 88900.48992088546),
'veteran': (68943.91833115448, 206926.897767934),
'violence': (138940.0713473897, 184745.6608155794),
'wall street': (450418.72499791963, 357506.1589435399),
'<NAME>': (549474.6485948666, 701703.5025185598),
'wedding': (54230.675876630434, 31697.39640199462),
'wildfire': (98159.61110844158, 63057.126015616355),
'wildlife': (13478.804880483567, 11018.743415659756),
'wine': (73970.4911081436, 35113.58599143231),
'wiretap': (65799.31335817679, 65505.32814705004),
'yoga': (80957.50711620713, 48030.61142450444)})
# +
MIN_TOPIC_SECONDS = 60 * 60
def topic_order(a, b):
keys = set()
keys.update(a.keys())
keys.update(b.keys())
kv = []
for key in keys:
a_val = a.get(key, None)
a_val = 1e-12 if a_val is None else a_val[0]
b_val = b.get(key, None)
b_val = 1e-12 if b_val is None else b_val[0]
if a_val + b_val < MIN_TOPIC_SECONDS:
continue
kv.append((key, a_val / (a_val + b_val)))
kv.sort(key=lambda x: x[1])
return [k for k, _ in kv]
def sum_screentime(a, b, scale):
result = defaultdict(int)
for k, v in a.items():
result[k] += v[0] / scale
for k, v in b.items():
result[k] += v[0] / scale
return result
top_n_topics = {
k for k, v in
sorted(sum_screentime(topic_to_male_screentime, topic_to_female_screentime, 1).items(),
key=lambda x: -x[1])[:50]
}
topic_plot_order = [
x for x in topic_order(topic_to_male_screentime, topic_to_female_screentime) if x in top_n_topics
]
plot_binary_proportion_comparison(
['Male', 'Female'],
[topic_to_male_screentime, topic_to_female_screentime],
'Gender Distribution of Screen Time By Topic',
'', #'Topic',
'Proportion of Screen Time',
baseline_series_names=[
'Baseline Male (All Topics)',
'Baseline Female (All Topics)'
],
baseline_data=[
screen_time_male['All Channels'][0],
screen_time_female['All Channels'][0]
],
sort_order=topic_plot_order,
raw_data_to_label_fn=None,
legend_loc=4,
save_path='figures/gender-3y-screen-time-by-topic.pdf'
)
plot_bar_chart(
['Male + Female'],
[sum_screentime(topic_to_male_screentime, topic_to_female_screentime, 3600 * 24)],
'',
'', 'Coverage (Days)',
figsize=(14, 3), series_colors=['DarkGray'], legend_loc=2,
show_ebars=False,
sort_order=topic_plot_order,
save_path='figures/gender-3y-screen-time-by-topic-bar.pdf'
)
# + hide_input=false
topic_to_male_mentions, topic_to_female_mentions = ({'abortion': (11236.252132713795, 382.40713417580355),
'affordable care act': (9022.325020432472, 178.28504708439905),
'aids': (2994.206518113613, 80.62711396996598),
'american health care act': (548.6910315752029, 9.816415056553373),
'autism': (685.6003779172897, 19.605769053409603),
'back to school': (445.91698437929153, 15.036586701108051),
'birth control': (790.9575905799866, 32.12937705150955),
'bombing': (10231.748476147652, 229.1596507367798),
'breast cancer': (492.06045562028885, 22.139430128109463),
'bullying': (2346.1988904476166, 67.76246679085271),
'business': (91257.73589783907, 2072.886070510939),
'cancer': (9490.374693930149, 318.74710125370245),
'cervical cancer': (70.39318698644638, 2.297435426974729),
'children': (47735.92115211487, 1609.6054159668056),
'cholesterol': (246.24713438749313, 9.821888506982539),
'christianity': (1735.81572920084, 39.62376633400151),
'circumcision': (17.202438831329346, 0.8035193532600359),
'congress': (121732.1553336978, 2253.507099359199),
'contraceptive': (157.66393542289734, 5.82611281824118),
'cooking': (1148.379305779934, 41.961463827020744),
'cryptocurrency': (7.937322318553925, 0.06052572306270672),
'diabetes': (1112.9122722148895, 58.72374729423551),
'divorce': (2735.0869675278664, 109.48168286765573),
'drugs': (13366.578389525414, 326.1088055093154),
'erectile dysfunction': (254.83499819040298, 51.33387254796956),
'evangelicals': (4828.962978422642, 108.15808383762996),
'fashion': (7054.989916741848, 156.77042280390162),
'feminism': (468.977885723114, 18.501466772520207),
'harassment': (7634.083441615105, 189.8308387639827),
'hiv': (734.9643259644508, 22.25738581115057),
'hurricane': (15104.720743238926, 322.8347270487651),
'immigration': (62325.57675892115, 1267.2134431873847),
'influenza': (72.6507505774498, 1.772447656163245),
'isis': (128508.38950520754, 2466.1025610262595),
'islam': (17140.667168080807, 339.810690308258),
'lgbt': (2946.194204747677, 82.94918957705578),
'marijuana': (4614.809868454933, 119.50699899956558),
'marriage': (15073.139418661594, 501.26221638148064),
'massage': (393.5154851078987, 12.879459333881503),
'medicine': (3812.5256739854813, 102.60356735899562),
'millenial': (163.8210887312889, 3.597961624162618),
'mom': (12879.210283339024, 549.0580769178864),
'murder': (28123.603552222252, 803.5589044841867),
'national security': (58577.55921572447, 1074.3868943708096),
'nuclear': (60281.08161979914, 1208.832217647058),
'nutrition': (419.5790290236473, 20.9405775310446),
'obamacare': (58588.064242362976, 1065.369007634213),
'obesity': (342.2370947599411, 6.054233961473762),
'opiods': (30.247527480125427, 0.37525487239480065),
'parenting': (403.7154778242111, 14.166476912983285),
'pregnancy': (760.0743358135223, 26.53600939549398),
'prostate cancer': (135.67578560113907, 3.3653219598482544),
'prostitution': (444.2284245491028, 9.917093827082414),
'racism': (10115.139577507973, 220.70818832073974),
'sex education': (8.993772327899935, 0.3695955099834366),
'sexism': (1769.061091542244, 45.874647775469356),
'social security': (6086.034533202648, 122.3461074949366),
'stem': (1172.9344388246536, 26.100557967605553),
'teens': (1628.375369668007, 47.36954342383142),
'terrorism': (41543.62926822901, 725.19879581253),
'trumpcare': (937.4060715436935, 23.588299493523905),
'violence': (39127.363469302654, 921.3077047006511),
'weight loss': (105.67184317111969, 5.434820085540551),
'women': (97541.35803723335, 3141.858557934175)},
{'abortion': (8481.747867286205, 382.40713417580355),
'affordable care act': (4258.674979567528, 178.28504708439905),
'aids': (1712.7934818863869, 80.62711396996598),
'american health care act': (197.30896842479706, 9.816415056553373),
'autism': (507.39962208271027, 19.605769053409603),
'back to school': (349.0830156207084, 15.036586701108051),
'birth control': (867.0424094200134, 32.12937705150955),
'bombing': (4642.251523852348, 229.1596507367798),
'breast cancer': (694.9395443797112, 22.139430128109463),
'bullying': (1922.8011095523834, 67.76246679085271),
'business': (50323.26410216093, 2072.886070510939),
'cancer': (6103.625306069851, 318.74710125370245),
'cervical cancer': (64.60681301355362, 2.297435426974729),
'children': (35583.07884788513, 1609.6054159668056),
'cholesterol': (166.75286561250687, 9.821888506982539),
'christianity': (744.18427079916, 39.62376633400151),
'circumcision': (36.797561168670654, 0.8035193532600359),
'congress': (53979.844666302204, 2253.507099359199),
'contraceptive': (169.33606457710266, 5.82611281824118),
'cooking': (913.6206942200661, 41.961463827020744),
'cryptocurrency': (1.062677681446075, 0.06052572306270672),
'diabetes': (824.0877277851105, 58.72374729423551),
'divorce': (1853.9130324721336, 109.48168286765573),
'drugs': (6005.4216104745865, 326.1088055093154),
'erectile dysfunction': (700.165001809597, 51.33387254796956),
'evangelicals': (2247.0370215773582, 108.15808383762996),
'fashion': (3500.010083258152, 156.77042280390162),
'feminism': (676.022114276886, 18.501466772520207),
'harassment': (5656.916558384895, 189.8308387639827),
'hiv': (550.0356740355492, 22.25738581115057),
'hurricane': (8933.279256761074, 322.8347270487651),
'immigration': (30598.423241078854, 1267.2134431873847),
'influenza': (34.3492494225502, 1.772447656163245),
'isis': (58204.61049479246, 2466.1025610262595),
'islam': (6615.332831919193, 339.810690308258),
'lgbt': (1935.8057952523232, 82.94918957705578),
'marijuana': (2183.190131545067, 119.50699899956558),
'marriage': (9189.860581338406, 501.26221638148064),
'massage': (320.4845148921013, 12.879459333881503),
'medicine': (2062.4743260145187, 102.60356735899562),
'millenial': (157.1789112687111, 3.597961624162618),
'mom': (12824.789716660976, 549.0580769178864),
'murder': (14401.396447777748, 803.5589044841867),
'national security': (26564.44078427553, 1074.3868943708096),
'nuclear': (24202.918380200863, 1208.832217647058),
'nutrition': (397.4209709763527, 20.9405775310446),
'obamacare': (24388.935757637024, 1065.369007634213),
'obesity': (228.7629052400589, 6.054233961473762),
'opiods': (9.752472519874573, 0.37525487239480065),
'parenting': (412.2845221757889, 14.166476912983285),
'pregnancy': (826.9256641864777, 26.53600939549398),
'prostate cancer': (49.32421439886093, 3.3653219598482544),
'prostitution': (201.77157545089722, 9.917093827082414),
'racism': (4984.860422492027, 220.70818832073974),
'sex education': (36.00622767210007, 0.3695955099834366),
'sexism': (1756.938908457756, 45.874647775469356),
'social security': (2628.965466797352, 122.3461074949366),
'stem': (680.0655611753464, 26.100557967605553),
'teens': (1158.624630331993, 47.36954342383142),
'terrorism': (16324.370731770992, 725.19879581253),
'trumpcare': (376.59392845630646, 23.588299493523905),
'violence': (20052.636530697346, 921.3077047006511),
'weight loss': (178.3281568288803, 5.434820085540551),
'women': (94146.64196276665, 3141.858557934175)})
# +
MIN_TOPIC_MENTIONS = 50
def topic_phrase_order(a, b):
keys = set()
keys.update(a.keys())
keys.update(b.keys())
kv = []
for key in keys:
a_val = a.get(key, None)
a_val = 1e-12 if a_val is None else a_val[0]
b_val = b.get(key, None)
b_val = 1e-12 if b_val is None else b_val[0]
if a_val + b_val < MIN_TOPIC_MENTIONS:
continue
kv.append((key, a_val / (a_val + b_val)))
kv.sort(key=lambda x: x[1])
return [k for k, _ in kv]
def sum_mentions(a, b):
result = defaultdict(int)
for k, v in a.items():
result[k] += v[0]
for k, v in b.items():
result[k] += v[0]
return result
top_n_topic_phrases = {
k for k, v in
sorted(sum_mentions(topic_to_male_mentions, topic_to_female_mentions).items(),
key=lambda x: -x[1])[:50]
}
topic_phrase_plot_order = [
x for x in topic_phrase_order(topic_to_male_mentions, topic_to_female_mentions) if x in top_n_topic_phrases
]
plot_binary_proportion_comparison(
['Male', 'Female'],
[topic_to_male_mentions, topic_to_female_mentions],
'On-Screen Gender Distribution By Keyword/Phrase',
'', #'Keyword',
'Proportion of Faces',
legend_loc=4,
raw_data_to_label_fn=None,
baseline_series_names=[
'Baseline Male (All Topics)',
'Baseline Female (All Topics)'
],
baseline_data=[
screen_time_male['All Channels'][0],
screen_time_female['All Channels'][0]
],
sort_order=topic_phrase_plot_order,
save_path='figures/gender-3y-mentions-by-topic.pdf'
)
plot_bar_chart(
['Male + Female'],
[sum_mentions(topic_to_male_mentions, topic_to_female_mentions)],
'',
'', 'Faces (Count)',
figsize=(14, 3), series_colors=['DarkGray'],
show_ebars=False, legend_loc=2, logy=True,
sort_order=topic_phrase_plot_order,
save_path='figures/gender-3y-mentions-by-topic-bar.pdf'
)
# -
# ## Face Size
gender_map = { g.id : g.name for g in Gender.objects.all() }
def get_gender_face_sizes():
# This touches spark
result = OrderedDict([('M', {}), ('F', {})])
for (gender_id, height), v in count_distinct_over_column(
face_genders.where(face_genders.probability > 0.5),
distinct_columns=[], #['face_id'],
group_by_columns=['gender_id', 'height'],
group_by_key_fn=lambda x: (x[0], math.ceil(x[1] * 100))
).items():
result[gender_map[gender_id]][height] = v[0]
return result
gender_face_size_hists = get_gender_face_sizes()
# +
def get_normalized_gender_face_sizes(hists):
result = defaultdict(dict)
for gender_name, hist in hists.items():
denom = sum(hist.values())
for k, v in hist.items():
result[gender_name][k] = v / denom
return result
norm_gender_face_size_hists = get_normalized_gender_face_sizes(gender_face_size_hists)
def get_cdf_gender_face_sizes(norm_hists):
cdf = defaultdict(dict)
for gender_name, hist in norm_hists.items():
acc = 0.
for k, v in sorted(hist.items(), key=lambda x: x[0]):
cdf[gender_name][k] = acc
acc += v
return cdf
cdf_gender_face_size = get_cdf_gender_face_sizes(norm_gender_face_size_hists)
# +
def plot_curves(curves, title, x_label, y_label,
color_map,
y_lim=None, x_lim=None, figsize=(7, 4),
save_path=None):
fig, ax1 = plt.subplots(figsize=figsize)
for series, values in curves.items():
x = []
y = []
for k in sorted(values):
x.append(k)
y.append(values[k])
ax1.plot(x, y, '-', label=series,
color=color_map[series])
ax1.legend()
ax1.set_title(title)
ax1.set_ylabel(y_label)
ax1.set_xlabel(x_label)
if y_lim is not None:
ax1.set_ylim(y_lim)
if x_lim is not None:
ax1.set_xlim(x_lim)
if save_path is None:
plt.show()
else:
plt.tight_layout()
plt.savefig(save_path)
def full_gender_name(d):
result = {}
gender_names = {'M': 'Male', 'F': 'Female'}
for k, v in d.items():
result[gender_names[k]] = v
return result
gender_color_map = { 'Male': 'Blue', 'Female': 'Red' }
plot_curves(
full_gender_name(norm_gender_face_size_hists),
'Distribution of Face Heights',
'Height (Percentage of Frame)', 'Proportion',
gender_color_map,
save_path='figures/gender-face-height-hist.pdf'
)
plot_curves(
full_gender_name(cdf_gender_face_size),
'Cumulative Distribution of Face Heights',
'Height (Percentage of Frame)', 'CDF',
gender_color_map,
save_path='figures/gender-face-height-cdf.pdf'
)
# -
# # <NAME> and <NAME>
face_identities = get_face_identities()
face_identities = face_identities.where(face_identities.labeler_id.isin(
[l.id for l in Labeler.objects.filter(name__contains='face-identity')]
))
trump_clinton_names = ['<NAME>', '<NAME>']
# +
channel_id_map = {c.id : c.name for c in Channel.objects.all()}
def person_helper(name, date_range):
identity_id = Identity.objects.get(name=name.lower()).id
person_identities = face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5) &
(face_identities.identity_id == identity_id) &
(face_identities.time >= func.to_date(func.lit(date_range[0]))) &
(face_identities.time < func.to_date(func.lit(date_range[1])))
)
screen_time_person = OrderedDict()
screen_time_person['All Channels'] = sum_over_column(
person_identities, 'duration', probability_column='probability'
)
for k, v in sum_over_column(person_identities, 'duration', ['channel_id'],
probability_column='probability').items():
screen_time_person[channel_id_map[k[0]]] = v
return screen_time_person
trump_clinton_3mo = [person_helper(p, ['2016-08-01', '2016-11-09'])
for p in trump_clinton_names]
trump_clinton_1yr = [person_helper(p, ['2015-11-09', '2016-11-09'])
for p in trump_clinton_names]
# -
sort_order = ['All Channels'] + [c.name for c in Channel.objects.all().order_by('name')]
plot_binary_proportion_comparison(
trump_clinton_names,
trump_clinton_3mo,
'Aug 1, 2016 to Nov 9, 2016 (3-Month)',
'',
'Proportion of Screen Time',
series_colors=['LightSalmon', 'LightBlue'],
error_colors=['Red', 'Blue'],
legend_loc=4, figsize=(5, 5),
raw_data_to_label_fn=None,
sort_order=sort_order,
save_path='figures/trump-clinton-3mo-ratio.pdf'
)
plot_binary_proportion_comparison(
trump_clinton_names,
trump_clinton_1yr,
'Nov 9, 2015 to Nov 9, 2016 (1-Year)',
'',
'Proportion of Screen Time',
series_colors=['LightSalmon', 'LightBlue'],
error_colors=['Red', 'Blue'],
legend_loc=4, figsize=(5, 5),
raw_data_to_label_fn=None,
sort_order=sort_order,
save_path='figures/trump-clinton-1yr-ratio.pdf'
)
trump_clinton_3mo_by_show = [
get_screen_time_by_canonical_show_spark(
name.lower(),
face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5)
),
date_range=['2016-08-01', '2016-11-09']
) for name in trump_clinton_names
]
trump_clinton_1yr_by_show = [
get_screen_time_by_canonical_show_spark(
name.lower(),
face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5)
),
date_range=['2015-11-09', '2016-11-09']
) for name in trump_clinton_names
]
# +
def convert_to_hours(d):
return { k : (v[0].total_seconds() / 3600, v[1] / (3600 ** 2)) for k, v in d.items() }
def order_by_ratio(a, b):
keys = set()
keys.update(a.keys())
keys.update(b.keys())
kv = []
for key in keys:
a_val = a.get(key, None)
a_val = 1e-12 if a_val is None else a_val[0].total_seconds()
b_val = b.get(key, None)
b_val = 1e-12 if b_val is None else b_val[0].total_seconds()
kv.append((key, a_val / (a_val + b_val)))
kv.sort(key=lambda x: x[1])
return [k for k, _ in kv]
plot_bar_chart(
trump_clinton_names, [convert_to_hours(x) for x in trump_clinton_3mo_by_show],
'Screen Time Comparison from Aug 1, 2016 to Nov 9, 2016 (3-Month)',
'', 'Hours', figsize=(14, 4), series_colors=['LightSalmon', 'LightBlue'],
sort_order=order_by_ratio(*trump_clinton_3mo_by_show),
save_path='figures/trump-clinton-by-show-3mo-bars.pdf'
)
plot_binary_proportion_comparison(
trump_clinton_names,
trump_clinton_3mo_by_show,
'Distribution of Trump and Clinton Screen Time from Aug 1, 2016 to Nov 9, 2016 (3-Month)',
'',
'Proportion of Screen Time',
series_colors=['LightSalmon', 'LightBlue'],
error_colors=['Red', 'Blue'],
legend_loc=4, figsize=(14, 5),
raw_data_to_label_fn=None,
sort_order=order_by_ratio(*trump_clinton_3mo_by_show),
save_path='figures/trump-clinton-by-show-3mo-ratio.pdf'
)
plot_bar_chart(
trump_clinton_names, [convert_to_hours(x) for x in trump_clinton_1yr_by_show],
'Screen Time Comparison from Nov 9, 2015 to Nov 9, 2016 (1-Year)',
'', 'Hours', figsize=(14, 4), series_colors=['LightSalmon', 'LightBlue'],
sort_order=order_by_ratio(*trump_clinton_1yr_by_show),
save_path='figures/trump-clinton-by-show-1yr-bars.pdf'
)
plot_binary_proportion_comparison(
trump_clinton_names,
trump_clinton_1yr_by_show,
'Distribution of Trump and Clinton Screen Time from Nov 9, 2015 to Nov 9, 2016 (1-Year)',
'',
'Proportion of Screen Time',
series_colors=['LightSalmon', 'LightBlue'],
error_colors=['Red', 'Blue'],
legend_loc=4, figsize=(14, 5),
raw_data_to_label_fn=None,
sort_order=order_by_ratio(*trump_clinton_1yr_by_show),
save_path='figures/trump-clinton-by-show-1yr-ratio.pdf'
)
# -
# # <NAME>
comey_screen_time_by_show = get_screen_time_by_canonical_show_spark(
'<NAME>',
face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5)
),
date_range=['2016-01-01', '2018-01-01']
)
plot_bar_chart(
['<NAME>'], [convert_to_hours(comey_screen_time_by_show)],
'Screen Time from Jan 1, 2016 to Jan 1, 2018',
'', 'Hours', figsize=(14, 6), series_colors=['LightSalmon'],
save_path='figures/comey-by-show-1618.pdf'
)
# +
opinion_shows = [
'The Rachel Maddow Show', 'Hardball',
'Hannity', '<NAME>', 'The OReilly Factor',
]
opinion_show_channels = (['MSNBC'] * 2) + (['FOXNEWS'] * 3)
opinion_canonical_show_ids = { CanonicalShow.objects.get(name=s).id : s for s in opinion_shows }
def comey_show_group_fn(x):
canonical_show_id, timestring = x
year, month, _ = timestring.split(' ')[0].split('-')
return (canonical_show_id, datetime(year=int(year), month=int(month), day=15))
comey_date_range=['2016-01-01', '2018-01-01']
comey_opinion_screen_time_over_time = sum_over_column(
face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5) &
(face_identities.time >= func.to_date(func.lit(comey_date_range[0]))) &
(face_identities.time < func.to_date(func.lit(comey_date_range[1]))) &
(face_identities.canonical_show_id.isin(list(opinion_canonical_show_ids.keys())))
),
'duration',
['canonical_show_id', 'time'],
group_by_key_fn=comey_show_group_fn,
probability_column='probability'
)
comey_opinion_screen_time_series = defaultdict(dict)
for (s, t), v in comey_opinion_screen_time_over_time.items():
comey_opinion_screen_time_series[opinion_canonical_show_ids[s]][t] = v[0] / 3600 # hours
# +
channel_ids = { x.id : x.name for x in Channel.objects.all() }
channels = list(sorted(channel_ids.values()))
def comey_channel_group_fn(x):
channel_id, timestring = x
year, month, _ = timestring.split(' ')[0].split('-')
return (channel_id, datetime(year=int(year), month=int(month), day=15))
comey_channel_screen_time_over_time = sum_over_column(
face_identities.where(
(face_identities.in_commercial == False) &
(face_identities.probability >= 0.3) &
(face_identities.host_probability < 0.5) &
(face_identities.time >= func.to_date(func.lit(comey_date_range[0]))) &
(face_identities.time < func.to_date(func.lit(comey_date_range[1])))
),
'duration',
['channel_id', 'time'],
group_by_key_fn=comey_channel_group_fn,
probability_column='probability'
)
comey_channel_screen_time_series = defaultdict(dict)
for (c, t), v in comey_channel_screen_time_over_time.items():
comey_channel_screen_time_series[channel_ids[c]][t] = v[0] / 3600 # hours
# +
plot_time_series(
['{} ({})'.format(opinion_shows[i], opinion_show_channels[i]) for i in range(len(opinion_shows))] ,
[comey_opinion_screen_time_series[x] for x in opinion_shows],
'<NAME>: Screen Time and Timeline',
'Hours',
# max_time=datetime(year=2017, month=11, day=1),
plotstyle='-', linewidth=2, figsize=(14, 4),
discrete_events=[
('Recommends no charges against Clinton', (datetime(year=2016, month=7, day=5), 18.5)),
('Reopens Clinton email investigation', (datetime(year=2016, month=10, day=28), 5)),
('Trump wins presidential election', (datetime(year=2016, month=11, day=8), 3.5)),
('Informs Trump that he is not under investigation ', (datetime(year=2017, month=1, day=6), 17)),
('Testifies before House Intelligence Committee', (datetime(year=2017, month=3, day=20), 15.5)),
('Testifies before Senate Judiciary Committee', (datetime(year=2017, month=5, day=3), 14)),
('Fired by Trump', (datetime(year=2017, month=5, day=9), 12.5)),
('Mueller appointed as special counsel', (datetime(year=2017, month=5, day=17), 11)),
],
save_path='figures/comey-opinion-time-series.pdf'
)
plot_time_series(
channels,
[comey_channel_screen_time_series[x] for x in channels],
'<NAME>: Screen Time and Timeline',
'Hours',
# max_time=datetime(year=2017, month=11, day=1),
plotstyle='-', linewidth=2, figsize=(14, 4),
# discrete_events=[
# ('Recommends no charges against Clinton', (datetime(year=2016, month=7, day=5), 85)),
# ('Reopens Clinton email investigation', (datetime(year=2016, month=10, day=28), 75)),
# ('Trump wins presidential election', (datetime(year=2016, month=11, day=8), 65)),
# ('Informs Trump that he is not under investigation ', (datetime(year=2017, month=1, day=6), 55)),
# ('Testifies before House Intelligence Committee', (datetime(year=2017, month=3, day=20), 45)),
# ('Testifies before Senate Judiciary Committee', (datetime(year=2017, month=5, day=3), 35)),
# ('Fired by Trump', (datetime(year=2017, month=5, day=9),25)),
# ('Mueller appointed as special counsel', (datetime(year=2017, month=5, day=17), 15)),
# ],
save_path='figures/comey-channel-time-series.pdf'
)
# +
opinion_host_to_shots = {
'<NAME>': [
584292, 584307, 6062987, 22151574, 27039495, 28005192, 28088277, 29310912, 37307674, 41194177,
44419012, 44591079, 44746669, 45164767, 47459051, 49952789, 59278225, 59278241, 60303840
],
'<NAME>': [113168, 113621, 2526491, 10713245, 10714080],
'<NAME>': [157794, 21838447, 23198946, 44426599, 59737649, 52388910, 32578439, 21683169],
'<NAME>': [1678041, 33194816, 44357358, 52691455],
'<NAME>': [
5848914, 8589521, 11292838, 19482668, 22458768, 22458768, 32191124, 41072250, 45188880,
48495239, 50722074, 54541370, 58423117, 58423117
]
}
def montage_shots(shots, cols=4, limit=8):
imgs = [(shot.id, load_frame(shot.video, int((shot.max_frame + shot.min_frame) / 2), []))
for shot in Shot.objects.filter(id__in=shots).order_by('video__time')]
if len(imgs) > limit:
imgs = [img for _, img in sorted(random.sample(list(enumerate(imgs)), limit))]
print('selected:', [i for i, _ in imgs])
shape = (imgs[0][1].shape[1], imgs[0][1].shape[0])
imgs = [cv2.resize(img, shape) for _, img in imgs]
return tile_images(imgs, cols=cols, blank_value=255)
for name, shots in opinion_host_to_shots.items():
print(name)
last_name = name.split(' ')[-1].replace("'", '')
img_out_path = 'figures/comey-{}.jpg'.format(last_name.lower())
img_montage = montage_shots(shots)
plt.imsave(img_out_path, cv2.cvtColor(img_montage, cv2.COLOR_RGB2BGR))
imshow(img_montage)
plt.show()
# -
for year in [2015, 2016, 2017]:
print(year)
for name, shots in opinion_host_to_shots.items():
last_name = name.split(' ')[-1].replace("'", '')
img_out_path = 'figures/comey-{}-{}.jpg'.format(last_name.lower(), year)
shots_in_year = [x.id for x in Shot.objects.filter(id__in=shots, video__time__year=year)]
if len(shots_in_year) > 0:
print(name)
img_montage = montage_shots(shots_in_year)
# plt.imsave(img_out_path, cv2.cvtColor(img_montage, cv2.COLOR_RGB2BGR))
imshow(img_montage)
plt.show()
# # Shootings
raise Exception('This section requires human effort!')
EXAMPLES_PER_CLUSTER = 5
identity_clustering_workflow(
'<NAME>',
examples_per_cluster=EXAMPLES_PER_CLUSTER,
init_clusters=20,
merge_cluster_threshold=0.05,
show_titles=False,
face_probability_threshold=0.5,
duration_label_unit='s',
save_paths=[
'figures/victim-taylor-screen-time.pdf',
'figures/victim-taylor-row-norm.pdf',
'figures/victim-taylor-col-norm.pdf'
]
)
identity_clustering_workflow(
'<NAME>',
examples_per_cluster=EXAMPLES_PER_CLUSTER,
init_clusters=25,
merge_cluster_threshold=0.05,
show_titles=False,
face_probability_threshold=0.5,
save_paths=[
'figures/shooter-farook-screen-time.pdf',
'figures/shooter-farook-row-norm.pdf',
'figures/shooter-farook-col-norm.pdf'
]
)
identity_clustering_workflow(
'<NAME>',
examples_per_cluster=EXAMPLES_PER_CLUSTER,
init_clusters=25,
merge_cluster_threshold=0.05,
show_titles=False,
face_probability_threshold=0.5,
save_paths=[
'figures/shooter-malik-screen-time.pdf',
'figures/shooter-malik-row-norm.pdf',
'figures/shooter-malik-col-norm.pdf'
]
)
|
app/notebooks/document_figures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import os
from pprint import pprint
import requests
from lxml.html import fromstring
import pandas as pd
# -
def random_user_agent(file):
if os.path.exists(file):
with open(file, 'r') as f:
lines = f.readlines()
return str(random.choice(lines)).replace("\n", "")
url_list = [
'https://www.laliga.es/laliga-santander/REPLACE_TEAM/plantilla/clasicas',
'https://www.laliga.es/laliga-santander/REPLACE_TEAM/plantilla/defensivas',
'https://www.laliga.es/laliga-santander/REPLACE_TEAM/plantilla/disciplina',
'https://www.laliga.es/laliga-santander/REPLACE_TEAM/plantilla/ofensivas',
'https://www.laliga.es/laliga-santander/REPLACE_TEAM/plantilla/eficiencia',
]
en_url_list = [
'https://www.laliga.es/en/laliga-santander/REPLACE_TEAM/squad/standings',
'https://www.laliga.es/en/laliga-santander/REPLACE_TEAM/squad/defensives',
'https://www.laliga.es/en/laliga-santander/REPLACE_TEAM/squad/discipline',
'https://www.laliga.es/en/laliga-santander/REPLACE_TEAM/squad/ofensive',
'https://www.laliga.es/en/laliga-santander/REPLACE_TEAM/squad/efficiency',
]
def retrieve_team_list():
head = {
"User-Agent": random_user_agent('resources/user_agent_list.txt'),
"X-Requested-With": "XMLHttpRequest",
"Accept": "text/html,application/xml",
"Accept-Encoding": "gzip, deflate, br",
"Connection": "keep-alive",
}
url = "https://www.laliga.es/laliga-santander"
req = requests.get(url, headers=head)
if req.status_code != 200:
raise ConnectionError("connection error " + str(req.status_code) + ", try again later.")
root = fromstring(req.text)
path = root.xpath(".//div[@id='equipos']/div[contains(@class, 'laliga-santander')]")
strip_from_url = 'https://www.laliga.es/laliga-santander/'
teams = list()
for element in path:
for value in element:
teams.append(value.get('href').replace(strip_from_url, ''))
return teams
def retrieve_team_dict():
head = {
"User-Agent": random_user_agent('resources/user_agent_list.txt'),
"X-Requested-With": "XMLHttpRequest",
"Accept": "text/html,application/xml",
"Accept-Encoding": "gzip, deflate, br",
"Connection": "keep-alive",
}
url = "https://www.laliga.es/laliga-santander"
req = requests.get(url, headers=head)
if req.status_code != 200:
raise ConnectionError("connection error " + str(req.status_code) + ", try again later.")
root = fromstring(req.text)
path = root.xpath(".//div[@id='equipos']/div[contains(@class, 'laliga-santander')]")
strip_from_url = 'https://www.laliga.es/laliga-santander/'
teams = list()
for element in path:
for value in element:
obj = {
'name': value.text_content(),
'tag': value.get('href').replace(strip_from_url, ''),
}
teams.append(obj)
return teams
def retrieve_player_stats(team):
result = list()
for url in url_list:
url = url.replace('REPLACE_TEAM', team['tag'])
head = {
"User-Agent": random_user_agent('resources/user_agent_list.txt'),
"X-Requested-With": "XMLHttpRequest",
"Accept": "text/html,application/xml",
"Accept-Encoding": "gzip, deflate, br",
"Connection": "keep-alive",
}
req = requests.get(url, headers=head)
if req.status_code != 200:
raise ConnectionError("connection error " + str(req.status_code) + ", try again later.")
root = fromstring(req.text)
path = root.xpath(".//div[contains(@class, 'rotar-tabla')]/table[@class='datatable']/thead/tr/th")
index = list()
for element in path:
if 'title' not in element.attrib:
index.append(element.text)
else:
index.append(element.get('title'))
df = pd.DataFrame(columns=index)
path = root.xpath(".//div[contains(@class, 'rotar-tabla')]/table[@class='datatable']/tbody/tr")
row = list()
for element in path:
for value in element:
row.append(value.text_content())
df.loc[len(df)] = row
row = list()
df.insert(loc=0, column='Equipo', value=team['name'])
df = df.drop(columns=['Foto'])
result.append(df)
return result
def retrieve_player_columns_english(team):
result = list()
for url in en_url_list:
url = url.replace('REPLACE_TEAM', team['tag'])
head = {
"User-Agent": random_user_agent('resources/user_agent_list.txt'),
"X-Requested-With": "XMLHttpRequest",
"Accept": "text/html,application/xml",
"Accept-Encoding": "gzip, deflate, br",
"Connection": "keep-alive",
}
req = requests.get(url, headers=head)
if req.status_code != 200:
raise ConnectionError("connection error " + str(req.status_code) + ", try again later.")
root = fromstring(req.text)
path = root.xpath(".//div[contains(@class, 'rotar-tabla')]/table[@class='datatable']/thead/tr/th")
indexes = list()
for element in path:
if 'title' not in element.attrib:
obj = {
'title': element.text,
'col': element.text,
}
indexes.append(obj)
else:
obj = {
'title': element.get('title'),
'col': element.text,
}
indexes.append(obj)
errored_titles = [
'Pictures',
'Goals scored per attempt (outside the box)',
'Goals scored per attempt (inside the box)',
'Goals scored per attempt (left foot)',
'Goals scored per attempt (right foot)',
'Goals scored per attempt (header)',
'Goals scored per attempt (set piece)',
]
for index in indexes:
if index not in result and index['title'] not in errored_titles:
result.append(index)
return result
def concatenate_dataframes(df_list):
result = pd.concat([df for df in df_list], axis=1)
result = result.loc[:,~result.columns.duplicated()]
return result
def __launch__():
team_dict = retrieve_team_dict()
players = pd.DataFrame()
for team in team_dict:
print(team)
df_list = retrieve_player_stats(team)
players = players.append(concatenate_dataframes(df_list), ignore_index=True, sort=False)
# LaLiga Error -> https://www.laliga.es/laliga-santander/barcelona/plantilla/eficiencia
players = players.drop(columns=['Goles marcados por tiros realizados desde fuera del área',
'Goles marcados por tiros realizados desde dentro del área',
'Goles marcados por tiros realizados con el pie izquierdo',
'Goles marcados por tiros realizados con el pie derecho',
'Goles marcados por tiros realizados de cabeza',
'Goles marcados por tiros realizados de jugada a balón parado'])
players.to_csv('../dataset/laliga_player_stats_spanish.csv', encoding='utf-8', index=False)
players.replace(['Portero', 'Defensa', 'Centrocampista', 'Delantero'], ['Goalkeeper', 'Defender', 'Midfielder', 'Forward'], inplace=True)
english_columns = retrieve_player_columns_english({'name': '<NAME>', 'tag': 'barcelona'})
english_columns_list = ['Team']
for english_column in english_columns:
english_columns_list.append(english_column['title'])
players.columns = english_columns_list
players.to_csv('../dataset/laliga_player_stats_english.csv', encoding='utf-8', index=False)
players = __launch__()
|
notebook/laliga_scraper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# ## Translation files
from fastai.text import *
# French/English parallel texts from http://www.statmt.org/wmt15/translation-task.html . It was created by <NAME>, who crawled millions of web pages and then used *a set of simple heuristics to transform French URLs onto English URLs (i.e. replacing "fr" with "en" and about 40 other hand-written rules), and assume that these documents are translations of each other*.
PATH = Path('data/translate')
TMP_PATH = PATH/'tmp'
TMP_PATH.mkdir(exist_ok=True)
fname='giga-fren.release2.fixed'
en_fname = PATH/f'{fname}.en'
fr_fname = PATH/f'{fname}.fr'
# +
re_eq = re.compile('^(Wh[^?.!]+\?)')
re_fq = re.compile('^([^?.!]+\?)')
lines = ((re_eq.search(eq), re_fq.search(fq))
for eq, fq in zip(open(en_fname, encoding='utf-8'), open(fr_fname, encoding='utf-8')))
qs = [(e.group(), f.group()) for e,f in lines if e and f]
# -
pickle.dump(qs, (PATH/'fr-en-qs.pkl').open('wb'))
qs = pickle.load((PATH/'fr-en-qs.pkl').open('rb'))
qs[:5], len(qs)
en_qs,fr_qs = zip(*qs)
en_tok = Tokenizer.proc_all_mp(partition_by_cores(en_qs))
fr_tok = Tokenizer.proc_all_mp(partition_by_cores(fr_qs), 'fr')
en_tok[0], fr_tok[0]
np.percentile([len(o) for o in en_tok], 90), np.percentile([len(o) for o in fr_tok], 90)
keep = np.array([len(o)<30 for o in en_tok])
en_tok = np.array(en_tok)[keep]
fr_tok = np.array(fr_tok)[keep]
pickle.dump(en_tok, (PATH/'en_tok.pkl').open('wb'))
pickle.dump(fr_tok, (PATH/'fr_tok.pkl').open('wb'))
en_tok = pickle.load((PATH/'en_tok.pkl').open('rb'))
fr_tok = pickle.load((PATH/'fr_tok.pkl').open('rb'))
def toks2ids(tok,pre):
freq = Counter(p for o in tok for p in o)
itos = [o for o,c in freq.most_common(40000)]
itos.insert(0, '_bos_')
itos.insert(1, '_pad_')
itos.insert(2, '_eos_')
itos.insert(3, '_unk')
stoi = collections.defaultdict(lambda: 3, {v:k for k,v in enumerate(itos)})
ids = np.array([([stoi[o] for o in p] + [2]) for p in tok])
np.save(TMP_PATH/f'{pre}_ids.npy', ids)
pickle.dump(itos, open(TMP_PATH/f'{pre}_itos.pkl', 'wb'))
return ids,itos,stoi
en_ids,en_itos,en_stoi = toks2ids(en_tok,'en')
fr_ids,fr_itos,fr_stoi = toks2ids(fr_tok,'fr')
def load_ids(pre):
ids = np.load(TMP_PATH/f'{pre}_ids.npy')
itos = pickle.load(open(TMP_PATH/f'{pre}_itos.pkl', 'rb'))
stoi = collections.defaultdict(lambda: 3, {v:k for k,v in enumerate(itos)})
return ids,itos,stoi
en_ids,en_itos,en_stoi = load_ids('en')
fr_ids,fr_itos,fr_stoi = load_ids('fr')
[fr_itos[o] for o in fr_ids[0]], len(en_itos), len(fr_itos)
# ## Word vectors
# fasttext word vectors available from https://fasttext.cc/docs/en/english-vectors.html
# +
# # ! pip install git+https://github.com/facebookresearch/fastText.git
# -
import fastText as ft
# To use the fastText library, you'll need to download [fasttext word vectors](https://github.com/facebookresearch/fastText/blob/master/pretrained-vectors.md) for your language (download the 'bin plus text' ones).
en_vecs = ft.load_model(str((PATH/'wiki.en.bin')))
fr_vecs = ft.load_model(str((PATH/'wiki.fr.bin')))
def get_vecs(lang, ft_vecs):
vecd = {w:ft_vecs.get_word_vector(w) for w in ft_vecs.get_words()}
pickle.dump(vecd, open(PATH/f'wiki.{lang}.pkl','wb'))
return vecd
en_vecd = get_vecs('en', en_vecs)
fr_vecd = get_vecs('fr', fr_vecs)
en_vecd = pickle.load(open(PATH/'wiki.en.pkl','rb'))
fr_vecd = pickle.load(open(PATH/'wiki.fr.pkl','rb'))
# +
ft_words = ft_vecs.get_words(include_freq=True)
ft_word_dict = {k:v for k,v in zip(*ft_words)}
ft_words = sorted(ft_word_dict.keys(), key=lambda x: ft_word_dict[x])
len(ft_words)
# -
dim_en_vec = len(en_vecd[','])
dim_fr_vec = len(fr_vecd[','])
dim_en_vec,dim_fr_vec
en_vecs = np.stack(list(en_vecd.values()))
en_vecs.mean(),en_vecs.std()
# ## Model data
enlen_90 = int(np.percentile([len(o) for o in en_ids], 99))
frlen_90 = int(np.percentile([len(o) for o in fr_ids], 97))
enlen_90,frlen_90
en_ids_tr = np.array([o[:enlen_90] for o in en_ids])
fr_ids_tr = np.array([o[:frlen_90] for o in fr_ids])
class Seq2SeqDataset(Dataset):
def __init__(self, x, y): self.x,self.y = x,y
def __getitem__(self, idx): return A(self.x[idx], self.y[idx])
def __len__(self): return len(self.x)
np.random.seed(42)
trn_keep = np.random.rand(len(en_ids_tr))>0.1
en_trn,fr_trn = en_ids_tr[trn_keep],fr_ids_tr[trn_keep]
en_val,fr_val = en_ids_tr[~trn_keep],fr_ids_tr[~trn_keep]
len(en_trn),len(en_val)
trn_ds = Seq2SeqDataset(fr_trn,en_trn)
val_ds = Seq2SeqDataset(fr_val,en_val)
bs=125
trn_samp = SortishSampler(en_trn, key=lambda x: len(en_trn[x]), bs=bs)
val_samp = SortSampler(en_val, key=lambda x: len(en_val[x]))
trn_dl = DataLoader(trn_ds, bs, transpose=True, transpose_y=True, num_workers=1,
pad_idx=1, pre_pad=False, sampler=trn_samp)
val_dl = DataLoader(val_ds, int(bs*1.6), transpose=True, transpose_y=True, num_workers=1,
pad_idx=1, pre_pad=False, sampler=val_samp)
md = ModelData(PATH, trn_dl, val_dl)
it = iter(trn_dl)
its = [next(it) for i in range(5)]
[(len(x),len(y)) for x,y in its]
# ## Initial model
def create_emb(vecs, itos, em_sz):
emb = nn.Embedding(len(itos), em_sz, padding_idx=1)
wgts = emb.weight.data
miss = []
for i,w in enumerate(itos):
try: wgts[i] = torch.from_numpy(vecs[w]*3)
except: miss.append(w)
print(len(miss),miss[5:10])
return emb
nh,nl = 256,2
class Seq2SeqRNN(nn.Module):
def __init__(self, vecs_enc, itos_enc, em_sz_enc, vecs_dec, itos_dec, em_sz_dec, nh, out_sl, nl=2):
super().__init__()
self.nl,self.nh,self.out_sl = nl,nh,out_sl
self.emb_enc = create_emb(vecs_enc, itos_enc, em_sz_enc)
self.emb_enc_drop = nn.Dropout(0.15)
self.gru_enc = nn.GRU(em_sz_enc, nh, num_layers=nl, dropout=0.25)
self.out_enc = nn.Linear(nh, em_sz_dec, bias=False)
self.emb_dec = create_emb(vecs_dec, itos_dec, em_sz_dec)
self.gru_dec = nn.GRU(em_sz_dec, em_sz_dec, num_layers=nl, dropout=0.1)
self.out_drop = nn.Dropout(0.35)
self.out = nn.Linear(em_sz_dec, len(itos_dec))
self.out.weight.data = self.emb_dec.weight.data
def forward(self, inp):
sl,bs = inp.size()
h = self.initHidden(bs)
emb = self.emb_enc_drop(self.emb_enc(inp))
enc_out, h = self.gru_enc(emb, h)
h = self.out_enc(h)
dec_inp = V(torch.zeros(bs).long())
res = []
for i in range(self.out_sl):
emb = self.emb_dec(dec_inp).unsqueeze(0)
outp, h = self.gru_dec(emb, h)
outp = self.out(self.out_drop(outp[0]))
res.append(outp)
dec_inp = V(outp.data.max(1)[1])
if (dec_inp==1).all(): break
return torch.stack(res)
def initHidden(self, bs): return V(torch.zeros(self.nl, bs, self.nh))
def seq2seq_loss(input, target):
sl,bs = target.size()
sl_in,bs_in,nc = input.size()
if sl>sl_in: input = F.pad(input, (0,0,0,0,0,sl-sl_in))
input = input[:sl]
return F.cross_entropy(input.view(-1,nc), target.view(-1))#, ignore_index=1)
opt_fn = partial(optim.Adam, betas=(0.8, 0.99))
rnn = Seq2SeqRNN(fr_vecd, fr_itos, dim_fr_vec, en_vecd, en_itos, dim_en_vec, nh, enlen_90)
learn = RNN_Learner(md, SingleModel(to_gpu(rnn)), opt_fn=opt_fn)
learn.crit = seq2seq_loss
learn.lr_find()
learn.sched.plot()
lr=3e-3
learn.fit(lr, 1, cycle_len=12, use_clr=(20,10))
learn.save('initial')
learn.load('initial')
# + [markdown] heading_collapsed=true
# ### Test
# + hidden=true
x,y = next(iter(val_dl))
probs = learn.model(V(x))
preds = to_np(probs.max(2)[1])
for i in range(180,190):
print(' '.join([fr_itos[o] for o in x[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in y[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in preds[:,i] if o!=1]))
print()
# + [markdown] heading_collapsed=true
# ## Bidir
# + hidden=true
class Seq2SeqRNN_Bidir(nn.Module):
def __init__(self, vecs_enc, itos_enc, em_sz_enc, vecs_dec, itos_dec, em_sz_dec, nh, out_sl, nl=2):
super().__init__()
self.emb_enc = create_emb(vecs_enc, itos_enc, em_sz_enc)
self.nl,self.nh,self.out_sl = nl,nh,out_sl
self.gru_enc = nn.GRU(em_sz_enc, nh, num_layers=nl, dropout=0.25, bidirectional=True)
self.out_enc = nn.Linear(nh*2, em_sz_dec, bias=False)
self.drop_enc = nn.Dropout(0.05)
self.emb_dec = create_emb(vecs_dec, itos_dec, em_sz_dec)
self.gru_dec = nn.GRU(em_sz_dec, em_sz_dec, num_layers=nl, dropout=0.1)
self.emb_enc_drop = nn.Dropout(0.15)
self.out_drop = nn.Dropout(0.35)
self.out = nn.Linear(em_sz_dec, len(itos_dec))
self.out.weight.data = self.emb_dec.weight.data
def forward(self, inp):
sl,bs = inp.size()
h = self.initHidden(bs)
emb = self.emb_enc_drop(self.emb_enc(inp))
enc_out, h = self.gru_enc(emb, h)
h = h.view(2,2,bs,-1).permute(0,2,1,3).contiguous().view(2,bs,-1)
h = self.out_enc(self.drop_enc(h))
dec_inp = V(torch.zeros(bs).long())
res = []
for i in range(self.out_sl):
emb = self.emb_dec(dec_inp).unsqueeze(0)
outp, h = self.gru_dec(emb, h)
outp = self.out(self.out_drop(outp[0]))
res.append(outp)
dec_inp = V(outp.data.max(1)[1])
if (dec_inp==1).all(): break
return torch.stack(res)
def initHidden(self, bs): return V(torch.zeros(self.nl*2, bs, self.nh))
# + hidden=true
rnn = Seq2SeqRNN_Bidir(fr_vecd, fr_itos, dim_fr_vec, en_vecd, en_itos, dim_en_vec, nh, enlen_90)
learn = RNN_Learner(md, SingleModel(to_gpu(rnn)), opt_fn=opt_fn)
learn.crit = seq2seq_loss
# + hidden=true
learn.fit(lr, 1, cycle_len=12, use_clr=(20,10))
# + hidden=true
learn.save('bidir')
# -
# ## Teacher forcing
class Seq2SeqStepper(Stepper):
def step(self, xs, y, epoch):
self.m.pr_force = (10-epoch)*0.1 if epoch<10 else 0
xtra = []
output = self.m(*xs, y)
if isinstance(output,tuple): output,*xtra = output
self.opt.zero_grad()
loss = raw_loss = self.crit(output, y)
if self.reg_fn: loss = self.reg_fn(output, xtra, raw_loss)
loss.backward()
if self.clip: # Gradient clipping
nn.utils.clip_grad_norm(trainable_params_(self.m), self.clip)
self.opt.step()
return raw_loss.data[0]
class Seq2SeqRNN_TeacherForcing(nn.Module):
def __init__(self, vecs_enc, itos_enc, em_sz_enc, vecs_dec, itos_dec, em_sz_dec, nh, out_sl, nl=2):
super().__init__()
self.emb_enc = create_emb(vecs_enc, itos_enc, em_sz_enc)
self.nl,self.nh,self.out_sl = nl,nh,out_sl
self.gru_enc = nn.GRU(em_sz_enc, nh, num_layers=nl, dropout=0.25)
self.out_enc = nn.Linear(nh, em_sz_dec, bias=False)
self.emb_dec = create_emb(vecs_dec, itos_dec, em_sz_dec)
self.gru_dec = nn.GRU(em_sz_dec, em_sz_dec, num_layers=nl, dropout=0.1)
self.emb_enc_drop = nn.Dropout(0.15)
self.out_drop = nn.Dropout(0.35)
self.out = nn.Linear(em_sz_dec, len(itos_dec))
self.out.weight.data = self.emb_dec.weight.data
self.pr_force = 1.
def forward(self, inp, y=None):
sl,bs = inp.size()
h = self.initHidden(bs)
emb = self.emb_enc_drop(self.emb_enc(inp))
enc_out, h = self.gru_enc(emb, h)
h = self.out_enc(h)
dec_inp = V(torch.zeros(bs).long())
res = []
for i in range(self.out_sl):
emb = self.emb_dec(dec_inp).unsqueeze(0)
outp, h = self.gru_dec(emb, h)
outp = self.out(self.out_drop(outp[0]))
res.append(outp)
dec_inp = V(outp.data.max(1)[1])
if (dec_inp==1).all(): break
if (y is not None) and (random.random()<self.pr_force):
if i>=len(y): break
dec_inp = y[i]
return torch.stack(res)
def initHidden(self, bs): return V(torch.zeros(self.nl, bs, self.nh))
rnn = Seq2SeqRNN_TeacherForcing(fr_vecd, fr_itos, dim_fr_vec, en_vecd, en_itos, dim_en_vec, nh, enlen_90)
learn = RNN_Learner(md, SingleModel(to_gpu(rnn)), opt_fn=opt_fn)
learn.crit = seq2seq_loss
learn.fit(lr, 1, cycle_len=12, use_clr=(20,10), stepper=Seq2SeqStepper)
learn.save('forcing')
# ## Attentional model
def rand_t(*sz): return torch.randn(sz)/math.sqrt(sz[0])
def rand_p(*sz): return nn.Parameter(rand_t(*sz))
class Seq2SeqAttnRNN(nn.Module):
def __init__(self, vecs_enc, itos_enc, em_sz_enc, vecs_dec, itos_dec, em_sz_dec, nh, out_sl, nl=2):
super().__init__()
self.emb_enc = create_emb(vecs_enc, itos_enc, em_sz_enc)
self.nl,self.nh,self.out_sl = nl,nh,out_sl
self.gru_enc = nn.GRU(em_sz_enc, nh, num_layers=nl, dropout=0.25)
self.out_enc = nn.Linear(nh, em_sz_dec, bias=False)
self.emb_dec = create_emb(vecs_dec, itos_dec, em_sz_dec)
self.gru_dec = nn.GRU(em_sz_dec, em_sz_dec, num_layers=nl, dropout=0.1)
self.emb_enc_drop = nn.Dropout(0.15)
self.out_drop = nn.Dropout(0.35)
self.out = nn.Linear(em_sz_dec*2, len(itos_dec))
self.out.weight.data = self.emb_dec.weight.data
self.W1 = rand_p(nh, em_sz_dec)
self.l2 = nn.Linear(em_sz_dec, em_sz_dec)
self.l3 = nn.Linear(em_sz_dec+nh, em_sz_dec)
self.V = rand_p(em_sz_dec)
def forward(self, inp, y=None, ret_attn=False):
sl,bs = inp.size()
h = self.initHidden(bs)
emb = self.emb_enc_drop(self.emb_enc(inp))
enc_out, h = self.gru_enc(emb, h)
h = self.out_enc(h)
dec_inp = V(torch.zeros(bs).long())
res,attns = [],[]
w1e = enc_out @ self.W1
for i in range(self.out_sl):
w2h = self.l2(h[-1])
u = F.tanh(w1e + w2h)
a = F.softmax(u @ self.V, 0)
attns.append(a)
Xa = (a.unsqueeze(2) * enc_out).sum(0)
emb = self.emb_dec(dec_inp)
wgt_enc = self.l3(torch.cat([emb, Xa], 1))
outp, h = self.gru_dec(wgt_enc.unsqueeze(0), h)
outp = self.out(self.out_drop(outp[0]))
res.append(outp)
dec_inp = V(outp.data.max(1)[1])
if (dec_inp==1).all(): break
if (y is not None) and (random.random()<self.pr_force):
if i>=len(y): break
dec_inp = y[i]
res = torch.stack(res)
if ret_attn: res = res,torch.stack(attns)
return res
def initHidden(self, bs): return V(torch.zeros(self.nl, bs, self.nh))
rnn = Seq2SeqAttnRNN(fr_vecd, fr_itos, dim_fr_vec, en_vecd, en_itos, dim_en_vec, nh, enlen_90)
learn = RNN_Learner(md, SingleModel(to_gpu(rnn)), opt_fn=opt_fn)
learn.crit = seq2seq_loss
lr=2e-3
learn.fit(lr, 1, cycle_len=15, use_clr=(20,10), stepper=Seq2SeqStepper)
learn.save('attn')
learn.load('attn')
# + [markdown] heading_collapsed=true
# ### Test
# + hidden=true
x,y = next(iter(val_dl))
probs,attns = learn.model(V(x),ret_attn=True)
preds = to_np(probs.max(2)[1])
# + hidden=true
for i in range(180,190):
print(' '.join([fr_itos[o] for o in x[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in y[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in preds[:,i] if o!=1]))
print()
# + hidden=true
attn = to_np(attns[...,180])
# + hidden=true
fig, axes = plt.subplots(3, 3, figsize=(15, 10))
for i,ax in enumerate(axes.flat):
ax.plot(attn[i])
# -
# ## All
class Seq2SeqRNN_All(nn.Module):
def __init__(self, vecs_enc, itos_enc, em_sz_enc, vecs_dec, itos_dec, em_sz_dec, nh, out_sl, nl=2):
super().__init__()
self.emb_enc = create_emb(vecs_enc, itos_enc, em_sz_enc)
self.nl,self.nh,self.out_sl = nl,nh,out_sl
self.gru_enc = nn.GRU(em_sz_enc, nh, num_layers=nl, dropout=0.25, bidirectional=True)
self.out_enc = nn.Linear(nh*2, em_sz_dec, bias=False)
self.drop_enc = nn.Dropout(0.25)
self.emb_dec = create_emb(vecs_dec, itos_dec, em_sz_dec)
self.gru_dec = nn.GRU(em_sz_dec, em_sz_dec, num_layers=nl, dropout=0.1)
self.emb_enc_drop = nn.Dropout(0.15)
self.out_drop = nn.Dropout(0.35)
self.out = nn.Linear(em_sz_dec, len(itos_dec))
self.out.weight.data = self.emb_dec.weight.data
self.W1 = rand_p(nh*2, em_sz_dec)
self.l2 = nn.Linear(em_sz_dec, em_sz_dec)
self.l3 = nn.Linear(em_sz_dec+nh*2, em_sz_dec)
self.V = rand_p(em_sz_dec)
def forward(self, inp, y=None):
sl,bs = inp.size()
h = self.initHidden(bs)
emb = self.emb_enc_drop(self.emb_enc(inp))
enc_out, h = self.gru_enc(emb, h)
h = h.view(2,2,bs,-1).permute(0,2,1,3).contiguous().view(2,bs,-1)
h = self.out_enc(self.drop_enc(h))
dec_inp = V(torch.zeros(bs).long())
res,attns = [],[]
w1e = enc_out @ self.W1
for i in range(self.out_sl):
w2h = self.l2(h[-1])
u = F.tanh(w1e + w2h)
a = F.softmax(u @ self.V, 0)
attns.append(a)
Xa = (a.unsqueeze(2) * enc_out).sum(0)
emb = self.emb_dec(dec_inp)
wgt_enc = self.l3(torch.cat([emb, Xa], 1))
outp, h = self.gru_dec(wgt_enc.unsqueeze(0), h)
outp = self.out(self.out_drop(outp[0]))
res.append(outp)
dec_inp = V(outp.data.max(1)[1])
if (dec_inp==1).all(): break
if (y is not None) and (random.random()<self.pr_force):
if i>=len(y): break
dec_inp = y[i]
return torch.stack(res)
def initHidden(self, bs): return V(torch.zeros(self.nl*2, bs, self.nh))
rnn = Seq2SeqRNN_All(fr_vecd, fr_itos, dim_fr_vec, en_vecd, en_itos, dim_en_vec, nh, enlen_90)
learn = RNN_Learner(md, SingleModel(to_gpu(rnn)), opt_fn=opt_fn)
learn.crit = seq2seq_loss
learn.fit(lr, 1, cycle_len=15, use_clr=(20,10), stepper=Seq2SeqStepper)
# ### Test
# +
x,y = next(iter(val_dl))
probs = learn.model(V(x))
preds = to_np(probs.max(2)[1])
for i in range(180,190):
print(' '.join([fr_itos[o] for o in x[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in y[:,i] if o != 1]))
print(' '.join([en_itos[o] for o in preds[:,i] if o!=1]))
print()
|
Courses/DeepLearning_2/translate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using Amazon SageMaker Object Detection For Playing Cards
#
# # Table of Contents
# 1. [Setup](#Setup)
# 2. [Data Exploration](#Data-Exploration)
# ## Setup
# Before inspecting and understanding the data, there are some initial steps to prepare the underlying notebook instance with additional Python libraries.
#
# * **jsonlines** is used for easy interaction with JSON records stored as lines in a file. In this workshop, we use a SageMaker object detection [Augmented Manifest](https://docs.aws.amazon.com/sagemaker/latest/dg/object-detection.html#object-detection-augmented-manifest-training) file, allowing SageMaker to stream training data into the training job using Pipe Input mode.
import sys
# !{sys.executable} -m pip install jsonlines
# Set up the S3 bucket where the training data is stored:
bucket_training = 'remars2019-revegas-trainingdata'
# To train the Object Detection algorithm on Amazon SageMaker, we need to setup and authenticate the use of AWS services. To begin with, we need an AWS account role with SageMaker access. Here we will use the execution role the current notebook instance was given when it was created. This role has necessary permissions, including access to your data in S3.
#
# We also import other libraries we need for the rest of the workshop to keep things organized.
# +
# %%time
# %matplotlib inline
import sagemaker
from sagemaker import get_execution_role
import boto3
s3 = boto3.resource('s3')
import json
import jsonlines
import random
import pandas as pd
from pandas.io.json import json_normalize
from collections import Counter
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
import os
import io
role = get_execution_role()
print(role)
sess = sagemaker.Session()
# -
plt.rcParams['figure.figsize'] = [15.5, 6]
# ## Data Exploration
# Let's begin by inspecting the annotated (labelled) data provided for the workshop.
#
# 1. First, we'll download the Augmented Manifest file, which is a text file storing JSON objects on new lines. It stores references to image locations in S3, as well as the corresponding labels of suit and rank.
s3.Bucket(bucket_training).download_file(
'manifests/augmentedManifest.json',
'./full_manifest.json')
# 2. We'll visualize the training data distribution and make sure our classes are balanced. SageMaker uses a key-value map to pass labels as classes to a neural network model. We defined those statically below:
class_map = {"AC": 0, "2C": 1, "3C": 2, "4C": 3, "5C": 4, "6C": 5, "7C": 6, "8C": 7, "9C": 8, "10C": 9, "JC": 10,
"QC": 11,"KC": 12, "AD": 13, "2D": 14, "3D": 15, "4D": 16, "5D": 17, "6D": 18, "7D": 19, "8D": 20,
"9D": 21, "10D": 22, "JD":23, "QD": 24, "KD": 25, "AH": 26, "2H": 27, "3H": 28, "4H": 29, "5H": 30,
"6H": 31, "7H": 32, "8H": 33, "9H": 34, "10H": 35, "JH": 36, "QH": 37, "KH": 38, "AS": 39, "2S": 40,
"3S": 41, "4S": 42, "5S": 43, "6S": 44, "7S": 45, "8S": 46, "9S": 47, "10S": 48, "JS": 49, "QS": 50, "KS": 51}
object_categories = list(class_map.keys())
# +
def get_key(val):
for key, value in class_map.items():
if val == value:
return key
return "key doesn't exist"
labels = []
train_df = pd.read_json('full_manifest.json', lines=True)
# train_df['bounding-box'].values
for x in train_df['bounding-box'].values:
for n in x['annotations']:
# print(get_key(n['class_id']))
labels.append(get_key(n['class_id']))
labels, values = zip(*Counter(labels).items())
indexes = np.arange(len(labels))
width = 1
plt.bar(indexes, values, width)
plt.xticks(indexes + width * 0.5, labels, rotation=45)
plt.show()
# -
# 3. Let's randomly display a training image:
# +
whole_manifest = []
with jsonlines.open('full_manifest.json') as reader:
for obj in reader:
whole_manifest.append(obj)
# +
def display_train_img(whole_manifest):
random_annotation = whole_manifest[random.randint(0,4999)]
s3_uri = random_annotation['source-ref']
s3_key = os.path.basename(s3_uri)
if 'images' not in os.listdir('.'):
os.mkdir('./images')
s3.Bucket(bucket_training).download_file(
s3_key, './images/' + s3_key)
raw_img = mpimg.imread('images/' + s3_key)
plt.imshow(raw_img)
display_train_img(whole_manifest)
# -
# As you can see, we have synthetically generated a training set of 5000 images using data augmentation techniques. By copying a cropped playing card onto various backgrounds and applying image filters such as blur and jpeg compression, the model should be much more robust.
# ## Data Preparation
#
# Now that we have inspected the data, let's perform a few steps to get the data ready to train on Amazon SageMaker.
# ### Create train, validation, and test splits
# +
random.shuffle(whole_manifest)
count_samples = 0
for x in whole_manifest:
count_samples = count_samples+1
print("Total samples: {}".format(count_samples))
train_count = round(count_samples * 0.04)
val_count = round(count_samples * 0.004)
test_count = round(count_samples * 0.002)
print("Train count: " + str(train_count) + '\n' +\
"Validation count: " + str(val_count) + '\n' +\
"Test count: " + str(test_count))
# +
train_manifest = []
for i in range(train_count):
train_manifest.append(whole_manifest.pop())
val_manifest = []
for i in range(val_count):
val_manifest.append(whole_manifest.pop())
test_manifest = []
for i in range(test_count):
test_manifest.append(whole_manifest.pop())
# +
with jsonlines.open('train.manifest', mode='w') as writer:
for i in train_manifest:
writer.write(i)
with jsonlines.open('validate.manifest', mode='w') as writer:
for i in val_manifest:
writer.write(i)
with jsonlines.open('test.manifest', mode='w') as writer:
for i in test_manifest:
writer.write(i)
# -
# ### Upload the manifests to a location in S3 to be used in the training job:
# +
sess.upload_data(path='train.manifest', key_prefix='manifests')
sess.upload_data(path='validate.manifest', key_prefix='manifests')
s3_train_data = 's3://{}/manifests/{}'.format(sess.default_bucket(), 'train.manifest')
s3_validation_data = 's3://{}/manifests/{}'.format(sess.default_bucket(), 'validate.manifest')
# -
# ## Train the model
#
# In the following steps, you will incrementally train a model that we trained in advance over hundreds of thousands of images.
# +
s3_output_path = 's3://{}/card-detection-output/'.format(sess.default_bucket())
# Model URI to our previously trained model:
model_uri = 's3://remars2019-revegas-trainingdata/model.tar.gz'
# Training container image that has the built-in SageMaker algorithm:
from sagemaker.amazon.amazon_estimator import get_image_uri
training_image = sagemaker.amazon.amazon_estimator.get_image_uri(boto3.Session().region_name, 'object-detection', repo_version='latest')
# Create the sagemaker estimator object.
playing_card_model = sagemaker.estimator.Estimator(training_image,
role,
train_instance_count = 1,
train_instance_type = 'ml.p2.xlarge',
input_mode='Pipe',
train_volume_size = 50,
train_max_run = 360000,
output_path = s3_output_path,
base_job_name = 'playingcard-bbox',
sagemaker_session = sess,
model_uri=model_uri)
# -
# Setup hyperparameters
playing_card_model.set_hyperparameters(base_network='resnet-50',
kv_store='dist_sync',
mini_batch_size=16,
use_pretrained_model=1,
num_classes=52, # suit/rank combinations
epochs=30,
image_shape=512,
num_training_samples = train_count,
learning_rate=0.00001,
optimizer='sgd',
early_stopping=False,
lr_scheduler_factor=0.1,
lr_scheduler_step='20,25')
# +
# Create sagemaker s3_input objects
attribute_names = ["source-ref","bounding-box"]
distribution = 'FullyReplicated'
train_data = sagemaker.session.s3_input(s3_train_data, distribution=distribution,
content_type='application/x-recordio',
record_wrapping='RecordIO',
attribute_names=attribute_names,
s3_data_type='AugmentedManifestFile')
validation_data = sagemaker.session.s3_input(s3_validation_data, distribution=distribution,
content_type='application/x-recordio',
record_wrapping='RecordIO',
attribute_names=attribute_names,
s3_data_type='AugmentedManifestFile')
data_channels = {'train': train_data,
'validation': validation_data}
# -
# %%time
playing_card_model.fit(inputs=data_channels, logs=True)
# ## Deploy the model predictor
pcm_predictor = playing_card_model.deploy(initial_instance_count=1, instance_type='ml.m4.xlarge')
pcm_predictor.content_type('image/jpeg')
# +
def display_test_img(test_manifest):
random_annotation = test_manifest[random.randint(0,test_count-1)]
s3_uri = random_annotation['source-ref']
annotations = random_annotation['bounding-box']['annotations']
s3_key = os.path.basename(s3_uri)
s3.Bucket(bucket_training).download_file(
s3_key, 'images/' + s3_key)
raw_img = mpimg.imread('images/' + s3_key)
plt.imshow(raw_img)
return s3_key
s3_key = display_test_img(test_manifest)
print(s3_key)
# +
def generate_predictions(s3_key):
img_bytes = io.BytesIO()
s3.Object(bucket_training, s3_key).download_fileobj(img_bytes)
dets = json.loads(pcm_predictor.predict(img_bytes.getvalue()))
return dets['prediction'], img_bytes
def visualize_detection(img_file, dets, classes=[], thresh=0.4):
"""
visualize detections in one image
Parameters:
----------
img : numpy.array
image, in bgr format
dets : numpy.array
ssd detections, numpy.array([[id, score, x1, y1, x2, y2]...])
each row is one object
classes : tuple or list of str
class names
thresh : float
score threshold
"""
img = mpimg.imread(img_file, "jpg")
plt.imshow(img)
height = img.shape[0]
width = img.shape[1]
colors = dict()
num_detections = 0
for det in dets:
(klass, score, x0, y0, x1, y1) = det
if score < thresh:
continue
num_detections += 1
cls_id = int(klass)
if cls_id not in colors:
colors[cls_id] = (random.random(), random.random(), random.random())
xmin = int(x0 * width)
ymin = int(y0 * height)
xmax = int(x1 * width)
ymax = int(y1 * height)
rect = plt.Rectangle((xmin, ymin), xmax - xmin, ymax - ymin, fill=False,
edgecolor=colors[cls_id], linewidth=3.5)
plt.gca().add_patch(rect)
class_name = str(cls_id)
if classes and len(classes) > cls_id:
class_name = classes[cls_id]
print('{},{}'.format(class_name,score))
plt.gca().text(xmin, ymin - 2,
'{:s} {:.3f}'.format(class_name, score),
bbox=dict(facecolor=colors[cls_id], alpha=0.5),
fontsize=12, color='white')
print('Number of detections: ' + str(num_detections))
plt.show()
# +
detections, img = generate_predictions(s3_key=s3_key)
visualize_detection(img_file=img, dets=detections, classes=object_categories)
# -
# Make the model file accessible so that workshop leads can add your model to the blackjack table system!
# +
from urllib.parse import urlparse
print(playing_card_model.model_data)
o = urlparse(playing_card_model.model_data)
s3object = s3.Object(o.netloc,o.path.lstrip('/'))
print(o.netloc)
print(o.path.lstrip('/'))
s3object.copy_from(
ACL="public-read",
CopySource={"Bucket": o.netloc,
"Key": o.path.lstrip('/')
}
)
# -
pcm_predictor.delete_endpoint()
|
Detecting Playing Cards Using Amazon SageMaker Built-In Object Detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# ### read in csv
# +
#Read in a very simple CSV file
file="F:\\Python_ML\\Course6_Python4DS\\scriptsLecture\\section4\\Resp2.csv"
df1=pd.read_csv(file)
df1.head(6)
# -
#Read in CSV when the seperator is ;
file="F:\\Python_ML\\Course6_Python4DS\\scriptsLecture\\section4\\winequality-red.csv"
#df1=pd.read_csv(file)
df1=pd.read_csv(file, sep = ";")
df1.head()
# ### read in text
#Read in .txt file
#df1=pd.read_csv("bostonTxt.txt") #tab seperated
df1=pd.read_csv("bostonTxt.txt", sep = "\t")
df1.head(6)
|
section2/Lecture9_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pollas_env
# language: python
# name: pollas_env
# ---
# ## import the libraries to use
# +
# This is for webscrapping
from bs4 import BeautifulSoup
import csv
import io
# Pandas
import pandas as pd
# REGEX
import re
# For get the date and time
from datetime import date
from datetime import datetime
# For create the engine and works with db's
import requests
from sqlalchemy.types import Integer, Text, String, DateTime
from sqlalchemy_utils import create_database, database_exists, drop_database
from sqlalchemy import create_engine
#import psycopg2
import sqlite3
# -
# ## functions
# With this function I make the webscrapping I need to extract the data from the tarifaluzahora website
def scrapping (tarifa, day = str(date.today())):
url = 'https://tarifaluzhora.es/?tarifa=' + tarifa
page = requests.get(url)
soup = BeautifulSoup(page.text, "html.parser")
price_ = soup.findAll("span", {"itemprop": "price"})
hours_ = soup.findAll("span", {"itemprop": "description"})
price_hour_ = [price.get_text() for price in price_]
schedule_ = [time.get_text() for time in hours_]
df = pd.DataFrame.from_dict({'precio':price_hour_,'horario':schedule_})
df['hora'] = [int(x[:2]) for x in df['horario']]
df['tarifa'] = tarifa
df['precio'] = [re.sub(r'/[k][W][h]','', str(x)) for x in df['precio']]
df['horario'] = [re.sub(r'[:]','', str(x)) for x in df['horario']]
return df
# ## extract fares from scrapping function
df = scrapping('coche_electrico')
df = df.groupby("precio").min().reset_index()
precio = df.precio[0]
precio
hora = df.hora[0]
hora
tarifa = df.tarifa[0]
tarifa
|
notebooks/Acquisition_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Seaborn - matrix and regression plots
import seaborn as sns
# %matplotlib inline
flights = sns.load_dataset('flights')
flights.head()
flights.shape
# Let us pivot this flights data such that it becomes a 2D matrix. Lets make the Month as row indices
flights_pv = flights.pivot_table(index='month', columns='year', values='passengers')
flights_pv.head()
# Using `pivot_tables` we have also aggregated the data by month and years.
# **ToC**
# - [Heatmap](#Heatmap)
# - [Cluster plot](#Cluster-plot)
# - [Regression Linear model plot](#Regression-linear-model-plot)
# ## Heatmap
# Heatmaps are a great way to represent continually variying data. However, you need to run this on a matrix kind of dataset, one where the row indexes are values themselves instead of serials.
sns.heatmap(flights_pv)
# From the heatmap above, we see there are more passengers in summer (June, July, August) and the number of passengers increases by the year as well.
# ### Heatmap for null data visualization
# SNS Heatmap is great to view how many nulls are in your data.
#from ml chapter, read titanic data
import pandas as pd
titanic = pd.read_csv('../udemy_ml_bootcamp/Machine Learning Sections/Logistic-Regression/titanic_train.csv')
titanic.head()
titanic.shape
titanic.isnull().head()
sns.heatmap(titanic.isnull(), yticklabels=False, cbar=False, cmap='viridis')
# You can see `Age` and `Cabin` columns have lots of null while others have none or very few.
# ## Cluster plot
# Cluster plots are useful to auto group datasets. More of this in machine learning section
sns.clustermap(flights_pv)
# Cluster map rearranges the data to show cells of similar values close by.
# ## Regression linear model plot
# You can do regression plots in two ways. You can decorate a scatter plot to have a fit or you can make a regression plot with scatter on it. We will see the latter.
tips = sns.load_dataset('tips')
tips.head()
#regressin total bill to the tip
sns.lmplot(x='total_bill', y='tip', data=tips)
# You can decorate this by splitting it by sex and assigning a different color for males and females
sns.lmplot(x='total_bill', y='tip', data=tips, hue='sex')
# You can bring in factors like day of week and create a regression for each day
sns.lmplot(x='total_bill', y='tip', data=tips, hue='sex', col='day')
|
python_crash_course/seaborn_cheat_sheet_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Word Prediction
#
# In this notebook, we are going to predict the next word that the writer is going to write. This will help us evaluate that how much the neural network has understood about dependencies between different letters that combine to form a word. We can also get an idea of how much the model has understood about the order of different types of word in a sentence.
#
# Code segments [1] to [5] are same as that in 'train.ipynb' notebook and their detailed explanation can be found over their itself.
import numpy as np
import keras
from keras.models import Sequential
from keras.layers import Dense, LSTM, Dropout, Activation
from keras.optimizers import RMSprop, Adam
from keras.callbacks import ModelCheckpoint
from keras.utils import np_utils
SEQ_LENGTH = 100
def buildmodel(VOCABULARY):
model = Sequential()
model.add(LSTM(256, input_shape = (SEQ_LENGTH, 1), return_sequences = True))
model.add(Dropout(0.2))
model.add(LSTM(256))
model.add(Dropout(0.2))
model.add(Dense(VOCABULARY, activation = 'softmax'))
model.compile(loss = 'categorical_crossentropy', optimizer = 'adam')
return model
file = open('wonderland.txt', encoding = 'utf8')
raw_text = file.read()
raw_text = raw_text.lower()
# +
chars = sorted(list(set(raw_text)))
print(chars)
bad_chars = ['#', '*', '@', '_', '\ufeff']
for i in range(len(bad_chars)):
raw_text = raw_text.replace(bad_chars[i],"")
chars = sorted(list(set(raw_text)))
print(chars)
VOCABULARY = len(chars)
int_to_char = dict((i, c) for i, c in enumerate(chars))
char_to_int = dict((c, i) for i, c in enumerate(chars))
# -
# Now that our model has been defined and we have preprocessed our input file and redefinded our vocabulary, as in train.ipynb file we are ready to proceed. The best model with least loss as we obtained in the last epoch of training is loaded and the model is build and recompiled.
filename = 'saved_models/weights-improvement-49-1.3420.hdf5'
model = buildmodel(VOCABULARY)
model.load_weights(filename)
model.compile(loss = 'categorical_crossentropy', optimizer = 'adam')
from ipywidgets import widgets
from IPython.display import display
# The original model has been defined in a manner to take in 100 character inputs. So when the user initially starts typing the words, the total length of input string will be less than 100 characters. To solve this issue, the input has been padded with series of spaces in the beginning in ordert to make the total length of 100 characters. As the total length exceeds 100 characters, only last 100 characters as the LSTM nodes take care of remembering the context of the document from before.
#
# Succeeding characters are predicted by the model until a space or full stop is encountered. The predicted characters are joined to form the next word, predicted by the model.
# +
original_text = []
predicted_text = []
text = widgets.Text()
display(text)
def handle_submit(sender):
global predicted_text
global original_text
inp = list(text.value)
last_word = inp[len(original_text):]
inp = inp[:len(original_text)]
original_text = text.value
last_word.append(' ')
inp_text = [char_to_int[c] for c in inp]
last_word = [char_to_int[c] for c in last_word]
if len(inp_text) > 100:
inp_text = inp_text[len(inp_text)-100: ]
if len(inp_text) < 100:
pad = []
space = char_to_int[' ']
pad = [space for i in range(100-len(inp_text))]
inp_text = pad + inp_text
while len(last_word)>0:
X = np.reshape(inp_text, (1, SEQ_LENGTH, 1))
next_char = model.predict(X/float(VOCABULARY))
inp_text.append(last_word[0])
inp_text = inp_text[1:]
last_word.pop(0)
next_word = []
next_char = ':'
while next_char != ' ':
X = np.reshape(inp_text, (1, SEQ_LENGTH, 1))
next_char = model.predict(X/float(VOCABULARY))
index = np.argmax(next_char)
next_word.append(int_to_char[index])
inp_text.append(index)
inp_text = inp_text[1:]
next_char = int_to_char[index]
predicted_text = predicted_text + [''.join(next_word)]
print(" " + ''.join(next_word), end='|')
text.on_submit(handle_submit)
# -
# The text box above shows the text as written by the user. The text used over here is the first few characters of the famous children book 'The Cat in the Hat' by Dr. Seuss available [here](http://www.stylist.co.uk/books/100-best-opening-lines-from-childrens-books#gallery-1). As the text is typed over, pressing enter just after the character ends (before the space), gives us the next word suggesstion, followed by a vertical bar to seperate the words, as shown above and in the gif.
#
# Next we summarize the predictions made by the model, in a nice tabular form listing the actual word typed by the user and the word suggessted by the model, before typing it side by side as shown after the code segment below.
# +
from tabulate import tabulate
original_text = original_text.split()
predicted_text.insert(0,"")
predicted_text.pop()
table = []
for i in range(len(original_text)):
table.append([original_text[i], predicted_text[i]])
print(tabulate(table, headers = ['Actual Word', 'Predicted Word']))
# -
# ## Conclusions
# A lot of observations can be made from the table above-
# * Most of the words generated by the model are proper english words, although there are exceptions at many places. This shows that the model has a good understanding of how letters are combined to form different words. Even though it is very obvious to do for a human, but for a computer model to give a reasonable performance at word formation is itself a huge feat.
# * The model has also understood to some extent about grammar of english language. In the above case, we can see that it often suggests verb at place of a verb like 'wet to see' in place of 'wet to play'. Also many a times, words of other part of speech are suggessted but they fit well, for example, 'we sat in the wind' is suggessted in place of 'we sat in the house'. Relationships like this show great hope, although the model has to further learn a lot in this area.
# * There are a few drawbacks as well. One of them is that the model often suggests 'and', both after a comma and a full stop which may be correct in case 1, but is always wrong for case 2.
#
# Overall, this makes up a nice demonstration for word prediction using RNNs with LSTM nodes. Seeing the performance of these models show that how advanced models phone keyboard suggestions use, which are very accurate. Further improvements in this model can be made by further training, tuning the hyperparameters, using a deeper network etc.
|
word_prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from config import Config
import utils
import os
import numpy as np
import json
import cv2
from keras import backend as K
from PIL import Image
from keras.preprocessing.image import *
import model as modellib
import random
import visualize
from visualize import display_images
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from model import log
# %matplotlib
# +
# Root directory of the project
ROOT_DIR = os.getcwd()
MODEL_DIR = os.path.join(ROOT_DIR, "logs")
PLANT = "Sorghum"
MODEL_PATH = os.path.join(ROOT_DIR, "mask_rcnn_%s_trained.h5"% (PLANT))
LOAD_MODEL_PATH = os.path.join(ROOT_DIR, "logs/mask_rcnn_Sorghum_trained.h5")
# -
class GroundTruth():
def __init__(self, polyId, img_height, img_width):
self.next = None
self.mask = np.zeros((img_height,img_width), dtype = "uint8")
self.max_x = 0
self.max_y = 0
self.min_x = 100000
self.min_y = 100000
self.p_Id = polyId
def get_lists(self):
maskList = []
bboxList = []
if(self.next == None):
maskList.append(self.mask)
bboxList.append(np.array([self.min_y, self.min_x, self.max_y, self.max_x]))
return maskList, bboxList
else:
maskList, bboxList = self.next.get_lists()
maskList.append(self.mask)
bboxList.append(np.array([self.min_y, self.min_x, self.max_y, self.max_x]))
return maskList, bboxList
def __str__(self):
temp = "____\nMax_x: " + str(self.max_x) + ", Min_x: " + str(self.min_x)
temp = temp + "\n\nMax_y: " + str(self.max_y) + ", Min_y: " + str(self.min_y)
return temp;
class StalkSpecConfig(Config):
"""Configuration for training on MS COCO.
Derives from the base Config class and overrides values specific
to the COCO dataset.
"""
# Give the configuration a recognizable name
NAME = "StalkSpec"
# We use a GPU with 12GB memory, which can fit two images.
# Adjust down if you use a smaller GPU.
IMAGES_PER_GPU = 1
# Uncomment to train on 8 GPUs (default is 1)
# GPU_COUNT = 8
# Number of classes (including background)
NUM_CLASSES = 2 # Background and sorghum
# Use small images for faster training. Set the limits of the small side
# the large side, and that determines the image shape.
IMAGE_MIN_DIM = 512
IMAGE_MAX_DIM = 512
# Number of training steps per epoch
# This doesn't need to match the size of the training set. Tensorboard
# updates are saved at the end of each epoch, so setting this to a
# smaller number means getting more frequent TensorBoard updates.
# Validation stats are also calculated at each epoch end and they
# might take a while, so don't set this too small to avoid spending
# a lot of time on validation stats.
STEPS_PER_EPOCH = 206
class StalkSpecDataset(utils.Dataset):
"""Load the dataset.
dataset_dir: The root directory of the dataset.
class_ids: If provided, only loads images that have the given classes.
"""
def load_stalks(self, dataset_dir, data_id_file):
self.add_class("stalk", 1, PLANT)
# Path
image_dir = os.path.join(dataset_dir, "images")
train_path = os.path.join(dataset_dir, data_id_file)
image_ids = []
labels = []
paths = []
#fp = filePaths
fp = open(train_path)
lines = fp.readlines()
fp.close()
for line in lines:
line = line.strip('\n')
image_ids.append(int(line))
json_file = open(dataset_dir + "/labels/%s.json"%(line))
json_str = json_file.read()
if(len(json_str) == 0): print(line)
json_data = json.loads(json_str)
labels.append(json_data)
paths.append(line)
pth = os.path.join(image_dir, "0003.jpg")
img = cv2.imread(pth, 0)
h = img.shape[0]
w = img.shape[1]
for ix, i in enumerate(image_ids):
self.add_image("stalk", image_id=i,
path=os.path.join(image_dir, "%s.jpg" % paths[ix]),
data_id_file=data_id_file,
data_dir = dataset_dir,
annotations = labels[ix],
height = h,
width = w)
def load_mask(self, image_id):
"""Load instance masks for the given image.
Different datasets use different ways to store masks. This
function converts the different mask format to one format
in the form of a bitmap [height, width, instances].
Returns:
masks: A bool array of shape [height, width, instance count] with
one mask per instance.
class_ids: a 1D array of class IDs of the instance masks.
bboxes: array [num_instances, (y1, x1, y2, x2)].
"""
image_info = self.image_info[image_id]
json_data = image_info['annotations']
img_scale = 5
img_height = image_info['height']
img_width = image_info['width']
masks = []
bboxes = []
groundTruths = []
for poly in json_data:
polyId = poly['polygon_id']
p = int(polyId)
points = []
currGt = None
#find the ground truth
for gt in groundTruths:
if(gt.p_Id == p):
currGt = gt
break
#if no ground truth was found make a new one and add it to the list
if(currGt == None):
currGt = GroundTruth(p, img_height, img_width)
if(len(groundTruths) > 0):
groundTruths[len(groundTruths)-1].next = currGt
groundTruths.append(currGt)
#update the mask and the min/max x and y
for vertex in poly['vertices']:
x = int((int(vertex['x']) * img_scale)/2)
y = int((int(vertex['y']) * img_scale)/2)
points.append([x,y])
if(x > currGt.max_x): currGt.max_x = x
if(x < currGt.min_x): currGt.min_x = x
if(y > currGt.max_y): currGt.max_y = y
if(y < currGt.min_y): currGt.min_y = y
points = np.array(points)
mask = currGt.mask
cv2.fillConvexPoly(mask, points, 255)
masks, bboxs = groundTruths[0].get_lists()
bboxs = np.asarray(bboxs)
masks = np.asarray(masks)
masks = np.swapaxes(masks, 0, 2)
masks = np.swapaxes(masks, 0, 1)
class_ids = []
for i in range(0, masks.shape[2]):
class_ids.append(1)
class_ids = np.array(class_ids)
return masks, class_ids.astype(np.int32), bboxs
def image_reference(self, image_id):
info = self.image_info[image_id]
if info["source"] == "plants":
return info
def get_ax(rows=1, cols=1, size=16):
"""Return a Matplotlib Axes array to be used in
all visualizations in the notebook. Provide a
central point to control graph sizes.
Adjust the size attribute to control how big to render images
"""
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
# +
inference_config = StalkSpecConfig()
debug_save_dir = "/media/default/ISAACTEGLER/maskrcnn/Mask_RCNN-2.0/debug/"
data_dir = "/media/default/ISAACTEGLER/data_sets/%s/squashed"%(PLANT)
dataset_test = StalkSpecDataset()
dataset_test.load_stalks(data_dir, "test.txt")
dataset_test.prepare()
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir=MODEL_DIR)
model_path = os.path.join(MODEL_DIR, "mask_rcnn_%s_trained.h5"%(PLANT))
# -
model.load_weights(model_path, by_name=True)
image_id = random.choice(dataset_test.image_ids)
print(image_id)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset_test, inference_config,
image_id, use_mini_mask=False)
results = model.detect([original_image], verbose=1)
ax = get_ax(1)
r = results[0]
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset_test.class_names, r['scores'], ax=ax,
title="Predictions")
log("gt_class_id", gt_class_id)
log("gt_bbox", gt_bbox)
log("gt_mask", gt_mask)
|
testResults.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
'''
*****************************************************************************************************************************************
*****************************************************************************************************************************************
* Author : <NAME>
* Author Details : Masters of Science, Computer Science, University of Memphis, Memphis, Tennessee, USA (May 2018)
*****************************************************************************************************************************************
*****************************************************************************************************************************************
* Project Name : Regression Modeling for Housing Price Prediction based on Numerical and Categorical Features
* Description : 1. Built regression model for predicting housing price using 79 numerical and categorical features.
* 2. Built pipelines for machine learning (regression) model training for reading files, creating training
* testing dataset, preprocessing (normalization, label encoding of categorical features), extracting
* features, and training and evaluation in grid search approach for multiple regression models.
* 3. Generated visualization and aggregated report on the performance of various models.
* Procedure : 1. Build pipelines for machine learning model training for reading file, creating training testing
* dataset, preprocessing (normalization, label encoding of categorical features), extracting features,
* and training and evaluation in grid search approach for mutiple regression models.
* 2. Preprocessing unit replaced non standard input features with default value and performed normalization,
* label encoding of categorical features.
* 3. Build feature set using almost all available features.
* 4. Build model training pipeline for regression
* 5. Generate agregated report on performance for various models.
* 6. Visualization for different model performance.
* Input :
* Output :
* Start Date : 09.22.2018
* Last Update :
* Tools Requirement : Anaconda, Python
* Comments : Please use Anaconda editor for visualization and convenience.
* Version History : 1.0.0.0
* Current Version : 1.0.0.0
*****************************************************************************************************************************************
*****************************************************************************************************************************************
'''
import numpy as np
import pandas as pd
import csv
from collections import defaultdict
import string
from string import punctuation
import re
from tokenize import tokenize
import nltk, re, time
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
import tensorflow as tf
from sklearn import metrics
import gc
class File:
def __init__(self, train_X_file, train_Y_file, test_X_file, test_Y_file):
if train_X_file:
self.train_X = pd.read_csv(train_X_file)
if train_Y_file:
self.train_Y = pd.read_csv(train_Y_file)
else:
self.train_Y = None
if test_X_file:
self.test_X = pd.read_csv(test_X_file)
if test_Y_file:
self.test_Y = pd.read_csv(test_Y_file)
else:
self.test_Y = None
def get_content(self):
return self.train_X, self.train_Y, self.test_X, self.test_Y
# +
class Data:
def __init__(self, train_X, train_Y, test_X, test_Y, train_test, train_XY, test_XY, targets):
self.train_X = train_X
self.test_X = test_X
self.train_Y = None
self.test_Y = None
self.feature_cols = None
self.targets = targets
if train_XY:
self.train_Y = self.train_X[self.targets]
self.train_X.drop(columns=self.targets, inplace=True)
else:
self.train_Y = train_Y
if test_XY:
print(targets)
self.test_Y = test_X[self.targets]
self.test_X.drop(columns=targets, inplace=True)
else:
self.test_Y = test_Y
# self.train_X.drop(columns='id', inplace=True)
# self.test_X.drop(columns='id', inplace=True)
def set_feature_cols(self, feature_cols):
if self.feature_cols:
self.feature_cols.append(feature_cols)
else: self.feature_cols = feature_cols
def remove_feature_cols(self, feature_cols):
if self.feature_cols:
self.feature_cols.remove(feature_cols)
else: self.feature_cols = None
def fill_columns_selected(self, columns_names, columns_val, inplace=True):
for cols in columns_names:
self.train_X[cols].fillna(columns_val, inplace=inpl)
self.test_X[cols].fillna(columns_val, inplace=inpl)
def get_data(self):
return self.train_X, self.train_Y, self.test_X, self.test_Y
# +
class Preprocess:
def __init__(self):
pass
def clean_text(self, text, alpha=True, punc=False, case_active=False, remove_stopwords=True):
if alpha:
text = re.sub(r"[^a-z]", " ", text)
if case_active==False:
text = text .lower()
if punc==False:
text = ''.join([c for c in text if c not in punctuation])
if remove_stopwords:
cached_stopwords = stopwords.words("english")
self.text = ' '.join([word for word in text.split() if word not in cached_stopwords])
text = re.sub(r" +", " ", text)
text = text.strip()
text = text .split()
return text
def tokenize(self, text, alpha=True, punc=False, case_active=False, remove_stopwords=True):
if alpha:
text = re.sub(r"[^a-z]", " ", text)
if case_active==False:
text = text.lower()
if punc==False:
text = ''.join([c for c in text if c not in punctuation])
if remove_stopwords:
cached_stopwords = stopwords.words("english")
text = ' '.join([word for word in text.split() if word not in cached_stopwords])
text = re.sub(r" +", " ", text)
text = text.strip()
text = text.split()
return text
def single_char_cnt(self, text, alpha=False, punc=False, remove_stopwords=True):
if alpha:
text = re.sub(r"[^a-z]", " ", text)
if punc==False:
text = ''.join([c for c in text if c not in punctuation])
if remove_stopwords:
cached_stopwords = stopwords.words("english")
text = ' '.join([word for word in text.split() if word not in cached_stopwords])
text = re.sub(r" +", " ", text)
text = text.strip()
text = text.split()
c=0
for tok in text:
if len(tok.strip())==1: c+=1
return c
def find_urls(self, text):
return re.findall('https?://(?:[-\w.]|(?:%[\da-fA-F]{2}))+', text)
def count_modals(self, text):
modals = ['can', 'could', 'may', 'might', 'must', 'will', 'would', 'should']
toks = text.split(' ')
c=0
for tok in toks:
if tok in modals: c+=1
return c
def non_alpha_mid(self, text, alpha=False, punc=False, remove_stopwords=True):
text = re.sub(r" +", " ", text)
if punc==False:
text = ''.join([c for c in text if c not in punctuation])
if remove_stopwords:
cached_stopwords = stopwords.words("english")
text = ''.join([word for word in text if word not in cached_stopwords])
text = text.split()
c=0
for tok in text:
m=0
for ch in tok:
if ch.isalpha()==0: m+=1
if (m>1 and len(tok)>1) or (m>=1): c+=1
return c
# -
class TextFeatures:
def __init__(self, train_X, test_X, columns_text):
self.train_X = train_X
self.test_X = test_X
self.columns_text = columns_text
self.train_X_features = []
self.test_X_features = []
self.train_X_tfidf = None
self.test_X_tfidf = None
def get_features_X(self, X, features):
count = lambda l1, l2: len(list(filter(lambda c: c in l2, l1)))
prep_obj = Preprocess()
features = []
try:
for col in self.columns_text:
length=X[col].size
i=0
features_col = []
for line in range(0, length):
counts = []
text = X[col][line]
counts.append(text.count("!"))
counts.append(text.count("?"))
counts.append(text.count("."))
counts.append(count(text,set(string.punctuation)))
counts.append(len(re.findall('[''""]', text)))
counts.append(prep_obj.single_char_cnt(text, alpha=False, punc=False, remove_stopwords=False))
counts.append(len(prep_obj.find_urls(text)))
counts.append(len(text))
counts.append(len(text.split()))
counts.append(sum(1 for c in text if c.isupper()))
counts.append(prep_obj.count_modals(text))
counts.append(len(re.findall(r'[\U0001f600-\U0001f650]', text)))
counts.append(prep_obj.non_alpha_mid(text, alpha=False, punc=False, remove_stopwords=False))
features_col.append(counts)
i+=1
# if i%10000==0: print(i)
features_col = np.array(features_col, dtype='int64')
features.append(features_col)
except:
print("Error:", line)
return features
def get_tfidf_features(self, tfidf, column):
self.train_X_tfidf = tfidf.fit_transform(train[column])
self.test_X_tfidf = tfidf.transform(test[column])
return self.train_X_tfidf, self.test_X_tfidf
def get_features(self):
self.train_X_features = self.get_features_X(self.train_X, self.columns_text)
self.test_X_features = self.get_features_X(self.test_X, self.columns_text)
return self.train_X_features, self.test_X_features
# +
from sklearn import preprocessing
from sklearn.preprocessing import Normalizer
class PreprocessNumeric:
def __init__(self):
pass
def fill_col(self, train_X, train_Y, test_X, test_Y):
if len(train_X)>0:
train_X.replace(-np.Inf, np.nan)
train_X.replace(np.Inf, np.nan)
train_X.replace(np.nan, 0)
train_X = train_X.fillna(0)
if len(train_Y)>0:
train_Y.replace(-np.Inf, np.nan)
train_Y.replace(np.Inf, np.nan)
train_Y.replace(np.nan, 0)
train_Y = train_Y.fillna(0)
# if len(test_X)>0:
# test_X.replace(-np.Inf, np.nan)
# test_X.replace(np.Inf, np.nan)
# test_X.replace(np.nan, 0)
# test_X = test_X.fillna(0)
# if len(test_Y)>0:
# test_Y.replace(-np.Inf, np.nan)
# test_Y.replace(np.Inf, np.nan)
# test_Y.replace(np.nan, 0)
# test_Y = test_Y.fillna(0)
return train_X, train_Y, test_X, test_Y
def drop_cols(self, train_X, test_X, ignore_cols):
train_X.drop(columns=ignore_cols, inplace=True)
test_X.drop(columns=ignore_cols, inplace=True)
return train_X, test_X
def normalize_target(self, train_Y, test_Y):
normalizer_train_m = {}
normalizer_test_m = {}
for col in train_Y:
normalizer_train = Normalizer(copy=False)
normalizer_test = Normalizer(copy=False)
normalizer_train.fit([train_Y[col]])
train_Y[col] = normalizer_train.transform([train_Y[col]]).flatten().astype('float64')
if test_Y:
normalizer_test.fit([test_Y[col]])
test_Y[col] = normalizer_test.transform([test_Y[col]]).flatten().astype('float64')
normalizer_train_m[col] = normalizer_train
normalizer_test_m[col] = normalizer_test
return train_Y, test_Y, normalizer_train_m, normalizer_test_m
def encoding_type(self, features, rating_score):
dtype_grp = features.columns.to_series().groupby(features.dtypes).groups
# for dtype in dtype_grp:
# print(dtype, len(dtype_grp[dtype]))
# i=0
for dtype in dtype_grp:
if dtype=='int64' :
for col in dtype_grp[dtype]:
features[col] = features[col].fillna(0.0)
features[col]=preprocessing.normalize([features[col]], norm='l2').flatten()
features[col] = features[col].astype('float64')
# i+=1
elif dtype=='float64' :
for col in dtype_grp[dtype]:
features[col] = features[col].fillna(0.0)
features[col]=preprocessing.normalize([features[col]], norm='l2').flatten()
features[col] = features[col].astype('float64')
# i+=1
elif dtype=='object' :
for col in dtype_grp[dtype]:
if col in rating_score:
features[col].replace(rating_score[col], regex=True, inplace=True)
features[col] = features[col].fillna(0.0)
features[col] = features[col].astype('float64')
# i+=1
else:
try:
le = preprocessing.LabelEncoder()
enc = preprocessing.OneHotEncoder()
le.fit(features[col].astype(str))
label_enc_feature_val = le.transform(features[col].astype(str))
features[col] = label_enc_feature_val.astype('float64')
# i+=1
# enc.fit([label_enc_feature_val])
# hot_enc_feature_val = enc.transform([label_enc_feature_val])
# print(hot_enc_feature_val)
# features.join(hot_enc_feature_val)
# features.drop(columns=col)
except ValueError:
print(col)
print("Oops! That was no valid number. Try again...")
# print("i--.", i)
return features
# -
class NumricFeatures:
def __init__(self, train_X, test_X, columns_text):
self.train_X = train_X
self.test_X = test_X
self.columns_text = columns_text
self.train_X_features = []
self.test_X_features = []
self.train_X_tfidf = None
self.test_X_tfidf = None
def get_features(self):
self.train_X_features = self.get_features_X(self.train_X, self.columns_text)
self.test_X_features = self.get_features_X(self.test_X, self.columns_text)
return self.train_X_features, self.test_X_features
# +
from sklearn.model_selection import KFold
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import StratifiedShuffleSplit
from sklearn.model_selection import train_test_split
class Splitter:
def __init__(self, splitter_name, n_splits, test_size, random_state):
self.splitter = None
if splitter_name == 'KFold':
self.splitter = KFold(n_splits=n_splits, shuffle=True, random_state=random_state)
elif splitter_name == 'StratifiedShuffleSplit':
self.splitter = StratifiedShuffleSplit(n_splits=n_splits, test_size=test_size, random_state=random_state)
def get_splitter(self):
return self.splitter
# +
# classifier
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.ensemble import (AdaBoostClassifier, ExtraTreesClassifier, GradientBoostingClassifier, RandomTreesEmbedding, RandomForestClassifier, VotingClassifier)
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.linear_model import LogisticRegression
from sklearn.mixture import GaussianMixture
from sklearn.naive_bayes import BernoulliNB, GaussianNB, MultinomialNB
from sklearn.neighbors import KDTree, KNeighborsClassifier, NearestNeighbors
from sklearn.neural_network import BernoulliRBM, MLPClassifier
from sklearn.svm import LinearSVC, NuSVC
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
# regressor
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.ensemble import (AdaBoostRegressor, BaggingRegressor, ExtraTreesRegressor, GradientBoostingRegressor, RandomTreesEmbedding, RandomForestRegressor, VotingClassifier)
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF
from sklearn.isotonic import IsotonicRegression
from sklearn.kernel_ridge import KernelRidge
from sklearn.linear_model import ARDRegression, LinearRegression, LogisticRegression, LogisticRegressionCV, logistic_regression_path, HuberRegressor, PassiveAggressiveRegressor, RandomizedLogisticRegression, RANSACRegressor, SGDRegressor, TheilSenRegressor
from sklearn.mixture import GaussianMixture
from sklearn.naive_bayes import BernoulliNB, GaussianNB, MultinomialNB
from sklearn.neighbors import KDTree, KNeighborsRegressor, NearestNeighbors, RadiusNeighborsRegressor
from sklearn.neural_network import BernoulliRBM, MLPRegressor
from sklearn.svm import LinearSVR, NuSVR, SVR
from sklearn.tree import DecisionTreeRegressor, ExtraTreeRegressor
class Model:
def __init__(self, model_type):
if model_type == 'Classification':
self.models = {
"AdaBoostClassifier": AdaBoostClassifier(),
"BernoulliNB": BernoulliNB(),
# "BernoulliRBM": BernoulliRBM(),
"DecisionTreeClassifier": DecisionTreeClassifier(),
"ExtraTreesClassifier": ExtraTreesClassifier(),
# "GaussianMixture": GaussianMixture(),
# "GaussianNB": GaussianNB(),
# "GaussianProcessClassifier": GaussianProcessClassifier(),
"GradientBoostingClassifier": GradientBoostingClassifier(),
# "KDTree": KDTree(),
# "KNeighborsClassifier": KNeighborsClassifier(3),
"LogisticRegression": LogisticRegression(),
"LinearSVC": LinearSVC(),
"MLPClassifier": MLPClassifier(),
"MultinomialNB": MultinomialNB(),
# "NearestNeighbors": NearestNeighbors(),
# "NuSVC": NuSVC(),
"QuadraticDiscriminantAnalysis": QuadraticDiscriminantAnalysis(),
"RandomForestClassifier": RandomForestClassifier(),
# "SVC Linear": SVC(kernel="linear", C=0.025),
# "SVC radial": SVC(kernel="radial", C=0.025),
# "SVC": SVC(),
# "SVC Gamma": SVC(gamma=2, C=1)
# VotingClassifier: VotingClassifier(),
}
elif model_type == 'Regression':
self.models = {
"AdaBoostRegressor": AdaBoostRegressor(),
# "ARDRegression": ARDRegression(),
"BaggingRegressor": BaggingRegressor(),
# "BernoulliRBM": BernoulliRBM(),
"DecisionTreeRegressor": DecisionTreeRegressor(),
"ExtraTreesRegressor": ExtraTreesRegressor(),
"ExtraTreeRegressor": ExtraTreeRegressor(),
# "GaussianMixture": GaussianMixture(),
# "GaussianNB": GaussianNB(),
"GaussianProcessRegressor": GaussianProcessRegressor(),
"GradientBoostingRegressor": GradientBoostingRegressor(),
"HuberRegressor": HuberRegressor(),
# "IsotonicRegression": IsotonicRegression(),
# "KernelRidge": KernelRidge(),
# "KDTree": KDTree(),
# "KNeighborsRegressor": KNeighborsRegressor(),
"LinearRegression": LinearRegression(),
# "LogisticRegression": LogisticRegression(),
# "LogisticRegressionCV": LogisticRegressionCV(),
# "logistic_regression_path": logistic_regression_path(),
"LinearSVR": LinearSVR(),
"MLPRegressor": MLPRegressor(),
# "MultinomialNB": MultinomialNB(),
"NuSVR": NuSVR(),
"PassiveAggressiveRegressor": PassiveAggressiveRegressor(),
# "QuadraticDiscriminantAnalysis": QuadraticDiscriminantAnalysis(),
# "RadiusNeighborsRegressor": RadiusNeighborsRegressor(),
"RandomForestRegressor": RandomForestRegressor(),
# "RandomizedLogisticRegression": RandomizedLogisticRegression(),
# "RANSACRegressor": RANSACRegressor(),
# "SGDRegressor": SGDRegressor(),
"SVR": SVR(),
"TheilSenRegressor": TheilSenRegressor(),
}
def get_models(self):
return self.models
# +
from sklearn import metrics
class ModelClassification:
def __init__(self, train_X, train_Y, test_X, test_Y, targets, splitter, models, average, report, details):
self.train_X = train_X
self.train_Y = train_Y
self.test_X = test_X
self.test_Y = test_Y
self.targets = targets
self.splitter = splitter
self.models = models
self.average = average
self.report = report
self.details = details
self.accuracy = 0
self.f1 = 0
self.precision = 0
self.recall = 0
def reset_score(self):
self.accuracy = 0
self.f1 = 0
self.precision = 0
self.recall = 0
def model_evaluation(self, model, target):
self.reset_score()
if self.report:
print("Model Description:")
print(model)
print("-"*100,"\n")
train_Y = np.array(self.train_Y[target], dtype='int64')
test_Y = np.array(self.test_Y[target], dtype='int64')
if self.splitter:
i=0
for train_index, test_index in self.splitter.split(self.train_X, train_Y):
X_train, X_test = self.train_X[train_index], self.train_X[test_index]
y_train, y_test = train_Y[train_index], train_Y[test_index]
model.fit(X_train, y_train)
if len(self.test_X)>0:
predict = model.predict(self.test_X)
accuracy_temp = metrics.accuracy_score(test_Y, predict)
precision_temp = metrics.precision_score(test_Y, predict, average=self.average)
recall_temp = metrics.recall_score(test_Y, predict, average=self.average)
f1_temp = metrics.f1_score(test_Y, predict, average=self.average)
hamming_loss = metrics.hamming_loss(test_Y, predict)
else:
predict = model.predict(X_test)
accuracy_temp = metrics.accuracy_score(y_test, predict)
precision_temp = metrics.precision_score(y_test, predict, average=self.average)
recall_temp = metrics.recall_score(y_test, predict, average=self.average)
f1_temp = metrics.f1_score(y_test, predict, average=self.average)
hamming_loss = metrics.hamming_loss(y_test, predict)
self.accuracy = self.accuracy + accuracy_temp
self.precision = self.precision + precision_temp
self.recall = self.recall+ recall_temp
self.f1= self.f1 + f1_temp
if details:
print("*"*25, " ITERATION - ", i+1, "*"*25)
print("-"*35)
print('%50s%s' % ("Accuracy Score :", accuracy_temp))
print('%50s%s' % ("Precision Score :", precision_temp))
print('%50s%s' % ("Recall Score :", recall_temp))
print('%50s%s' % ("F1 Score :", f1_temp))
print('%50s%s' % ("Hamming Loss :", hamming_loss))
if test_X:
# precision, recall, thresholds = metrics.precision_recall_curve(test_Y, predict)
# print('%50s%s' % ("average_precision_score :", metrics.average_precision_score(test_Y, predict, average=self.average)))
# print('%50s%s' % ("fbeta_score :", metrics.fbeta_score(test_Y, predict)))
# print('%50s%s' % ("roc_auc_score :", metrics.roc_auc_score(test_Y, predict, average=self.average)))
print("-"*35)
print(metrics.classification_report(test_Y, predict))
print("-"*35)
print("Confusion Matrix:\n\n", metrics.confusion_matrix(test_Y, predict))
else:
# precision, recall, thresholds = metrics.precision_recall_curve(y_test, predict)
print('%50s%s' % ("Average Precision Score :", metrics.average_precision_score(y_test, predict, average=self.average)))
print('%50s%s' % ("Fbeta Score :", metrics.fbeta_score(y_test, predict)))
print('%50s%s' % ("Roc Auc Score :", metrics.roc_auc_score(y_test, predict, average=self.average)))
print("-"*35)
print(metrics.classification_report(y_test, predict))
print("-"*35)
print("Confusion Matrix:\n\n", metrics.confusion_matrix(y_test, predict))
print("-"*35)
print("\n")
i+=1
split_num = self.splitter.get_n_splits()
self.accuracy = self.accuracy/split_num
self.precision = self.precision/split_num
self.recall = self.recall/split_num
self.f1 = self.f1/split_num
else:
model.fit(self.train_X, self.train_Y)
predict = model.predict(self.test_X)
self.accuracy = metrics.accuracy_score(test_Y, predict)
self.precision = metrics.precision_score(test_Y, predict, average=self.average)
self.recall = metrics.recall_score(test_Y, predict, average=self.average)
self.f1 = metrics.f1_score(test_Y, predict, average=self.average)
hamming_loss = metrics.hamming_loss(test_Y, predict)
if self.report:
if self.splitter:
print("*"*50, " Average For", i+1, " Folds", "*"*50)
print('%50s%s' % ("Average Accuracy Score: ", self.accuracy))
print('%50s%s' % ("Average Precision Score: ", self.precision))
print('%50s%s' % ("Average Recall Score: ", self.recall))
print('%50s%s' % ("Average F1 Score: ", self.f1))
return self.accuracy, self.precision, self.recall, self.f1
def grid(self, target):
print("Spliter Description:")
print(self.splitter)
print("targets: ", target, "\n\n")
evaluation = {}
if self.models==None:
models_names_obj = Model("Classification")
self.models = models_names_obj.get_models()
for model in self.models:
evaluation_temp = []
accuracy, precision, recall, f1 = self.model_evaluation(self.models[model], target)
evaluation_temp.append(accuracy)
evaluation_temp.append(precision)
evaluation_temp.append(recall)
evaluation_temp.append(f1)
evaluation[model] = evaluation_temp
gc.collect()
rows_list = []
for name in evaluation:
rows_list.append([name]+evaluation[name])
evaluation_pd = pd.DataFrame(rows_list, columns=['model', 'accuracy', 'precision', 'recall', 'f1'])
return evaluation_pd
def multi_target(self):
evaluations_dict = {}
for target in self.targets:
evaluations_dict[target] = self.grid(target)
return evaluations_dict
# +
class ModelRegression:
def __init__(self, train_X, train_Y, test_X, test_Y, targets, splitter, models, average, report, details):
self.train_X = train_X
self.train_Y = train_Y
self.test_X = test_X
self.test_Y = test_Y
self.targets = targets
self.splitter = splitter
self.models = models
self.trained_models = None
self.average = average
self.report = report
self.details = details
self.explained_variance_score_val = 0
self.mean_absolute_error_val = 0
self.mean_squared_error_val = 0
self.mean_squared_log_error_val = 0
self.median_absolute_error_val = 0
self.r2_score_val = 0
def reset_score(self):
self.explained_variance_score_val = 0
self.mean_absolute_error_val = 0
self.mean_squared_error_val = 0
self.mean_squared_log_error_val = 0
self.median_absolute_error_val = 0
self.r2_score_val = 0
def model_evaluation(self, model, target):
self.reset_score()
if report:
print("Model Description:")
print(model)
print("-"*100,"\n")
if splitter:
i=0
train_Y = np.array(self.train_Y[target], dtype='int64')
for train_index, test_index in self.splitter.split(self.train_X, train_Y):
X_train, X_test = self.train_X[train_index], self.train_X[test_index]
y_train, y_test = train_Y[train_index], train_Y[test_index]
model.fit(X_train, y_train)
if test_X:
predict = model.predict(test_X)
explained_variance_score_temp = metrics.explained_variance_score(test_Y, predict)
mean_absolute_error_temp = metrics.mean_absolute_error(test_Y, predict)
mean_squared_error_temp = metrics.mean_squared_error(test_Y, predict)
mean_squared_log_error_temp = metrics.mean_squared_log_error(test_Y, predict)
median_absolute_error_temp = metrics.median_absolute_error(test_Y, predict)
r2_score_temp = metrics.r2_score(test_Y, predict)
else:
predict = model.predict(X_test)
explained_variance_score_temp = metrics.explained_variance_score(y_test, predict)
mean_absolute_error_temp = metrics.mean_absolute_error(y_test, predict)
mean_squared_error_temp = metrics.mean_squared_error(y_test, predict)
mean_squared_log_error_temp = metrics.mean_squared_log_error(y_test, predict)
median_absolute_error_temp = metrics.median_absolute_error(y_test, predict)
r2_score_temp = metrics.r2_score(y_test, predict)
self.explained_variance_score_val = self.explained_variance_score_val + explained_variance_score_temp
self.mean_absolute_error_val = self.mean_absolute_error_val + mean_absolute_error_temp
self.mean_squared_error_val = self.mean_squared_error_val + mean_squared_error_temp
self.mean_squared_log_error_val = self.mean_squared_log_error_val + mean_squared_log_error_temp
self.median_absolute_error_val = self.median_absolute_error_val + median_absolute_error_temp
self.r2_score_val = self.r2_score_val + r2_score_temp
if details:
print("*"*25, " ITERATION - ", i+1, "*"*25)
print("-"*35)
print('%50s%s' % ("explained_variance_score_temp :", explained_variance_score_temp))
print('%50s%s' % ("mean_absolute_error_temp :", mean_absolute_error_temp))
print('%50s%s' % ("mean_squared_error_temp :", mean_squared_error_temp))
print('%50s%s' % ("median_absolute_error_temp :", median_absolute_error_temp))
print('%50s%s' % ("mean_squared_log_error_temp :", mean_squared_log_error_temp))
print('%50s%s' % ("median_absolute_error_temp :", median_absolute_error_temp))
print('%50s%s' % ("r2_score_val :", r2_score_val))
print("-"*35)
print("\n")
i+=1
split_num = splitter.get_n_splits()
self.explained_variance_score_val = self.explained_variance_score_val/split_num
self.mean_absolute_error_val = self.mean_absolute_error_val/split_num
self.mean_squared_error_val = self.mean_squared_error_val/split_num
self.median_absolute_error_val = self.median_absolute_error_val/split_num
self.r2_score_val = self.r2_score_val/split_num
else:
model.fit(train_X, train_Y)
predict = model.predict(test_X)
self.explained_variance_score = metrics.explained_variance_score(test_Y, predict)
self.mean_absolute_error = metrics.mean_absolute_error(test_Y, predict)
self.mean_squared_error = metrics.mean_squared_error(test_Y, predict)
self.mean_squared_log_error = metrics.mean_squared_log_error(test_Y, predict)
self.median_absolute_error = metrics.median_absolute_error(test_Y, predict)
self.r2_score_temp = metrics.r2_score(test_Y, predict)
if report:
if splitter:
print("*"*40, " Average For", i+1, " Folds", "*"*40)
print('%50s%s' % ("Average explained_variance_score: ", self.explained_variance_score_val))
print('%50s%s' % ("Average mean_absolute_error: ", self.mean_absolute_error_val))
print('%50s%s' % ("Average mean_squared_error: ", self.mean_squared_error_val))
print('%50s%s' % ("Average mean_squared_log_error: ", self.mean_squared_log_error_val))
print('%50s%s' % ("Average median_absolute_error: ", self.median_absolute_error_val))
print('%50s%s' % ("Average r2_score: ", self.r2_score_val))
print("\n")
print("*"*100)
return model, self.explained_variance_score_val, self.mean_absolute_error_val, self.mean_squared_error_val, self.mean_squared_log_error_val, self.median_absolute_error_val, self.r2_score_val
def grid(self, target):
print("Spliter Description:")
print(self.splitter)
print("targets: ", target, "\n\n")
trained_models = {}
evaluation = {}
if self.models==None:
models_names_obj = Model("Regression")
self.models = models_names_obj.get_models()
for model in self.models:
evaluation_temp = []
model_new, explained_variance_score_val, mean_absolute_error_val, mean_squared_error_val, mean_squared_log_error_val, median_absolute_error_val, r2_score_val = self.model_evaluation(self.models[model], target)
evaluation_temp.append(explained_variance_score_val)
evaluation_temp.append(mean_absolute_error_val)
evaluation_temp.append(mean_squared_error_val)
evaluation_temp.append(mean_squared_log_error_val)
evaluation_temp.append(median_absolute_error_val)
evaluation_temp.append(r2_score_val)
evaluation[model] = evaluation_temp
trained_models[model] = model_new
# gc.collect()
rows_list = []
for name in evaluation:
rows_list.append([name]+evaluation[name])
evaluation_pd = pd.DataFrame(rows_list, columns=['model', 'explained_variance_score', 'mean_absolute_error', 'mean_squared_error', 'mean_squared_log_error', 'median_absolute_error', 'r2_score'])
return trained_models, evaluation_pd
def multi_target(self):
trained_models_multi_target = {}
evaluations_dict = {}
for target in self.targets:
trained_models, evaluations_dict[target] = self.grid(target)
trained_models_multi_target[target] = trained_models
return trained_models_multi_target, evaluations_dict
# +
train_X_file = r'house-prices-advanced-regression-techniques/train.csv'
test_X_file = r'house-prices-advanced-regression-techniques/test.csv'
file_obj = File(train_X_file, None, test_X_file, None)
train_X, train_Y, test_X, test_Y = file_obj.get_content()
###
targets = ['SalePrice']
data_obj = Data(train_X, None, test_X, None, False, True, False, targets)
train_X, train_Y, test_X, test_Y = data_obj.get_data()
###
ignore_cols = ['BsmtExposure']
val1 = {'Ex':5, 'Gd':4, 'TA':3, 'Fa':2, 'Po':1, 'NA':0}
val2 = {'GLQ':6, 'ALQ':5, 'BLQ':4, 'Rec':3, 'LwQ':2, 'Unf':1, 'NA':0}
rating_score = {'ExterQual':val1, 'ExterCond':val1, 'BsmtQual':val1, 'BsmtCond':val1, 'HeatingQC':val1, 'KitchenQual':val1, 'FireplaceQu':val1, 'GarageQual':val1, 'GarageCond':val1, 'PoolQC':val1, 'BsmtFinType1':val2, 'BsmtFinType2':val2}
#
prep_num = PreprocessNumeric()
train_X, test_X = prep_num.drop_cols(train_X, test_X, ignore_cols)
train_X = prep_num.encoding_type(train_X, rating_score)
test_X = prep_num.encoding_type(test_X, rating_score)
#
# print(train_X)
train_X, train_Y, test_X, test_Y = prep_num.fill_col(train_X, train_Y, None, None)
train_Y, test_Y, normalizer_train, normalizer_test = prep_num.normalize_target(train_Y, None)
# train_Y[targets[0]] = train_Y_temp
#
train_X = train_X.values
norm = train_Y.max()[0].flatten()[0]
train_Y = train_Y/norm
# train_Y[targets[0]] = train_Y[targets[0]]/norm
# train_Y = train_Y.values
# if test_X:
# test_X = test_X.values
# if test_Y:
# test_Y = test_Y.values
# train_Y, test_Y, normalizer_train, normalizer_test = prep_num.normalize(None, train_Y, None, test_Y)
# splitter_name = "StratifiedShuffleSplit"
splitter_name = "KFold"
n_splits = 5
test_size = 0.25
random_state = True
splitter_obj = Splitter(splitter_name, n_splits, test_size, random_state)
splitter = splitter_obj.get_splitter()
#
models = None
average = "weighted"
report = 1
details = 0
m = ModelRegression(train_X, train_Y, None, None, targets, splitter, models, average, report, details)
trained_models, evaluation_pd = m.grid(targets[0])
evaluation = evaluation_pd
# -
evaluation = evaluation_pd
evaluation
# +
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.pyplot import figure
import os
import re
labels= ['explained_variance_score', 'mean_absolute_error', 'mean_squared_error', 'mean_squared_log_error', 'median_absolute_error', 'r2_score']
x = np.array(evaluation_pd['model'])
evaluation = evaluation[labels]
# os.mkdir("output")
# os.mkdir("output/plots")
for n in range(len(labels)):
fig = plt.figure(num=None, figsize=(14, 6), dpi=250)
ax = plt.subplot(111)
y = evaluation[labels[n]]
plt.plot(x, y, label = labels[n])
leg = plt.legend(loc='best', ncol=2, mode="expand", shadow=True, fancybox=True)
leg.get_frame().set_alpha(0.5)
plt.legend()
# plt.xlabel('Model', fontsize=15)
plt.ylabel('Score', fontsize=15)
plt.xticks(rotation=90)
ax.tick_params(labelsize='large', width=5)
ax.grid(True, linestyle='-.')
plt.tight_layout()
plt.title(re.sub('[^a-zA-Z0-9]', ' ', labels[n]).title(), fontsize=20)
plt.show()
fig.savefig("output/plots/"+labels[n])
# +
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.pyplot import figure
labels= ['accuracy', 'precision', 'recall', 'f1']
x = np.array(evaluation['model'])
for item in evaluation_pd:
evaluation = evaluation_pd[item]
figure(num=None, figsize=(14, 6), dpi=250)
ax = plt.subplot(111)
for n in range(len(labels)):
y = evaluation[labels[n]]
plt.plot(x, y, label = labels[n])
plt.title(item)
leg = plt.legend(loc='best', ncol=2, mode="expand", shadow=True, fancybox=True)
leg.get_frame().set_alpha(0.5)
plt.legend()
plt.xlabel('Model')
plt.ylabel('Score')
plt.xticks(rotation=90)
ax.tick_params(labelsize='large', width=5)
ax.grid(True, linestyle='-.')
plt.tight_layout()
plt.show()
|
.ipynb_checkpoints/House Prices Advanced Regression Techniques-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import agama
import galpy
agama.setUnits( mass=1., length=1., velocity=1.) # Msun, kpc, km/s
# # Compare Potentials
# +
from Cautun20 import C20_pot as a_C20
ini_C20 = agama.Potential('Cautun20.ini')
#Load from py or from ini file. Note ini file combines cgm and bulge as single spheroid potential, py gives seperate
a_M17 = agama.Potential("McMillan17.ini")
a_PW19 = agama.Potential("PriceWhelan19.ini")
a_MW14 = agama.Potential("MWPotential2014.ini")
def a_vcirc(pot,R):
vc = np.sqrt(-R*pot.force(np.stack((R,np.zeros_like(R), np.zeros_like(R))).transpose())[:,0])
return vc
def a_pot_R(pot,R):
return pot.potential(np.stack((R,np.zeros_like(R), np.zeros_like(R))).transpose())
# +
nR = 1000
R_space = np.geomspace(0.5, 200, nR)
a_Pots = {'C20':a_C20, 'M17':a_M17, 'PW19':a_PW19, 'MW14':a_MW14, 'ini_C20':ini_C20}
pots = list(a_Pots.keys())
a_vc = {p:a_vcirc(a_Pots[p], R_space) for p in pots}
# +
MWVCircData=np.loadtxt('Eilers19_Vcirc.txt')
vdata_r=MWVCircData[:,0]
vdata_vc=MWVCircData[:,1]
vdata_vc_u=MWVCircData[:,2]
vdata_vc_l=MWVCircData[:,3]
# + tags=[]
plt.figure(figsize=(12,6))
for p in pots:
if p == 'ini_C20':
plt.plot(R_space, a_vc[p], label=p, ls=':')
else:
plt.plot(R_space, a_vc[p], label=p)
plt.ylim([100, 240])
plt.plot(R_space, a_vc[p], label=p)
plt.errorbar(vdata_r, vdata_vc, yerr=[vdata_vc_u, vdata_vc_l], ls = 'none',
label='Eilers 19 Obs', c='black')
plt.scatter(vdata_r, vdata_vc,c='black')
plt.xlim([1, 50])
plt.xlabel('R, kpc')
plt.ylabel('Vcirc km/s')
plt.title('Comparison of Potentials')
plt.legend()
plt.show()
# +
components = ['DM_Halo','disc', 'bulge','cgm']
individual_components = ['thin','thick','HI','H2', 'bulge', 'cgm', 'DM_Halo']
print(a_C20)
a_C20_seperate = {com: a_C20[i] for (i,com) in enumerate(individual_components)}
a_C20_seperate['disc'] = agama.Potential(a_C20_seperate['thin'],a_C20_seperate['thick']
,a_C20_seperate['HI'],a_C20_seperate['H2'])
a_C20_vc_seperate = { com: a_vcirc(a_C20_seperate[com],R_space)
for com in components}
# +
plt.figure(figsize=(12,6))
total_vc = np.zeros_like(a_vc['C20']**2)
for com in components:
plt.plot(R_space, a_C20_vc_seperate[com], label=com)
total_vc = total_vc + (a_C20_vc_seperate[com]**2)
total_vc = np.sqrt(total_vc)
plt.plot(R_space, a_vc['C20'], label='Total', c='k')
plt.plot(R_space, total_vc, label='Resummed Total', c='red', ls = ':', linewidth=3)
plt.ylim([50, 240])
plt.errorbar(vdata_r, vdata_vc, yerr=[vdata_vc_u, vdata_vc_l], ls = 'none', c='black')
plt.scatter(vdata_r, vdata_vc,c='black')
plt.xlim([1, 50])
plt.xlabel('R, kcp')
plt.ylabel('Vcirc km/s')
plt.title('Cautun Components')
plt.legend()
plt.show()
# -
# # Compare with Galpy
# Potential matches, with differences at the very centre (<1 kpc), and 0.1% difference towards R200.
# Implementation, or AGAMA vs Galpy difference?
# A similar difference can be seen in McMillan17 Potential.
# Hopefully not significant!
from galpy.potential.Cautun20 import Cautun20 as g_C20
from galpy.potential.mwpotentials import McMillan17 as g_M17
from galpy.potential import vcirc
g_vc_C20 = vcirc(g_C20,R=R_space/g_C20[0]._ro, use_physical=True)
g_vc_M17 = vcirc(g_M17,R=R_space/g_M17[0]._ro, use_physical=True)
components = ['DM_Halo','disc', 'bulge', 'cgm']
print(g_C20)
g_C20_seperate = { com: g_C20[i] for (i,com) in enumerate(components)}
g_C20_vc_seperate = { com: vcirc(g_C20_seperate[com],R=R_space/g_C20[0]._ro)
for com in components}
# +
plt.figure(figsize=(12,6))
for com in components:
p = plt.plot(R_space, g_C20_vc_seperate[com], label=f'galpy {com}',alpha=0.5)
plt.plot(R_space, a_C20_vc_seperate[com], label=f'agama {com}', ls =':', c = p[0].get_color(),linewidth=4)
plt.plot(R_space, g_vc_C20, label='Total galpy', c='k',alpha=0.4)
plt.plot(R_space, a_vc['C20'], label='Total AGAMA', c='k', ls = ':', linewidth=4)
plt.xlabel('R, kcp')
plt.ylabel('Vcirc km/s')
plt.title('Galpy vs AGAMA Cautun Components')
plt.legend()
plt.show()
# +
plt.figure(figsize=(12,6))
for com in components:
plt.plot(R_space, a_C20_vc_seperate[com]/ g_C20_vc_seperate[com]
, label=f'agama/galpy {com}')
plt.plot(R_space, a_vc['C20']/ g_vc_C20,
label=f'Total agama/galpy ')
plt.xlabel('R, kcp')
plt.ylabel('Vcirc km/s')
plt.title('Galpy vs AGAMA Cautun Components')
plt.yscale('log')
plt.legend()
plt.show()
# +
plt.figure(figsize=(12,6))
plt.plot(R_space, a_vc['M17']/ g_vc_M17,
label=f'M17 agama/galpy ')
plt.xlabel('R, kcp')
plt.ylabel('Vcirc km/s')
plt.title('Galpy vs AGAMA McMillan17')
plt.yscale('log')
plt.legend()
plt.show()
# -
# # Example Contraction
# What would McMillan17 look like contracted?
# +
from contraction_agama import contract_agama_potential
M17_uncontracted_DM_pot = agama.Potential(type="Spheroid",
densityNorm = 8.53702e+06,
axisRatioZ = 1,
gamma = 1,
beta = 3,
scaleRadius = 19.5725)
M17_thin_disc_pot = agama.Potential( type = 'Disk',
surfaceDensity = 8.95679e+08,
scaleRadius = 2.49955,
scaleHeight = 0.3)
M17_thick_disc_pot = agama.Potential( type = "Disk",
surfaceDensity = 1.83444e+08,
scaleRadius = 3.02134,
scaleHeight = 0.9)
M17_HI_disc_pot = agama.Potential( type = "Disk",
surfaceDensity = 5.31319e+07,
scaleRadius = 7,
scaleHeight = -0.085,
innerCutoffRadius = 4)
M17_H2_disc_pot = agama.Potential( type = "Disk",
surfaceDensity = 2.17995e+09,
scaleRadius = 1.5,
scaleHeight = -0.045,
innerCutoffRadius = 12)
M17_bulge_pot = agama.Potential(type="Spheroid",
densityNorm = 9.8351e+10,
axisRatioZ = 0.5,
gamma = 0,
beta = 1.8,
scaleRadius = 0.075,
outerCutoffRadius = 2.1)
M17_Baryon_pot = agama.Potential(M17_thin_disc_pot, M17_thick_disc_pot, M17_HI_disc_pot,
M17_H2_disc_pot, M17_bulge_pot)
M17_contracted_dm_pot = contract_agama_potential(M17_uncontracted_DM_pot, M17_Baryon_pot)
contracted_M17_pot = agama.Potential(M17_contracted_dm_pot, M17_Baryon_pot)
# +
M17_vc_contracted = a_vcirc(contracted_M17_pot,R_space)
plt.figure(figsize=(12,6))
plt.plot(R_space, M17_vc_contracted, label='M17 Contracted')
plt.plot(R_space, a_vc['M17'], label='Original')
plt.xlabel('R, kcp')
plt.ylabel('Vcirc km/s')
plt.title('Contracted McMillan17')
plt.legend()
plt.show()
# -
# # Agama Total Mass vs Scipy Integration
# If agama is not the most recent version, totalMass method fails and required to integrate ourselves. A version of the script is tested below.
# +
from contraction_agama import contract_density_fit, density_from_enclosed_mass, Mcum_from_axi_dens, Mcum_from_sph_dens
def scipy_contract_agama_potential(dm_pot, baryon_pot, fbar=0.157,rmax=500, rmin=0.1, scipy=False):
'''Given spherical DM and axisymmetric baryon agama potentials,
creates a contracted DM agama potential using procedure found in Cautun 20'''
r_space = np.geomspace(rmin,rmax,501)
xyz = np.stack((r_space,np.zeros_like(r_space),np.zeros_like(r_space))).transpose()
dens_dm = dm_pot.density(xyz)
if not scipy:
print('Agama totalMass')
Mcum_dm = np.array([dm_pot.totalMass(r) for r in r_space])
Mcum_bar = np.array([baryon_pot.totalMass(r) for r in r_space])
else:
print('scipy_integration')
Mcum_dm = Mcum_from_sph_dens(r_space, dm_pot.density)
Mcum_bar = Mcum_from_axi_dens(r_space, baryon_pot.density)
dens_bar = density_from_enclosed_mass(r_space,Mcum_bar,r_space) #note spherical average
dens_contracted = contract_density_fit(dens_dm,dens_bar,Mcum_dm,Mcum_bar,fbar)
def contracted_dens_func(xyz):
r = np.linalg.norm(xyz,axis=1)
return np.interp(r, r_space, dens_contracted)
contracted_pot = agama.Potential(type="Multipole", density=contracted_dens_func,
symmetry="spherical", rmin=1e-3, rmax=1e3)
return contracted_pot
# -
M17_contracted_dm_pot = scipy_contract_agama_potential(M17_uncontracted_DM_pot, M17_Baryon_pot)
scipy_M17_contracted_dm_pot = scipy_contract_agama_potential(M17_uncontracted_DM_pot, M17_Baryon_pot, scipy=True)
# +
vc = a_vcirc(M17_contracted_dm_pot, R_space)
scipy_vc = a_vcirc(scipy_M17_contracted_dm_pot, R_space)
plt.figure()
plt.plot(R_space, vc, label='Agama Mtotal')
plt.plot(R_space, scipy_vc, label='Scipy Integration')
plt.legend()
plt.show()
plt.figure()
plt.plot(R_space, vc / scipy_vc, label='Agama Mtotal / Scipy Integration')
plt.legend()
plt.show()
# -
# # Alternate fbar
# +
fb15_M17_contracted_dm_pot = contract_agama_potential(M17_uncontracted_DM_pot, M17_Baryon_pot,fbar=0.15)
contracted_M17_pot = agama.Potential(fb15_M17_contracted_dm_pot, M17_Baryon_pot)
fb12_M17_contracted_dm_pot = contract_agama_potential(M17_uncontracted_DM_pot, M17_Baryon_pot, fbar=0.12)
contracted_M17_pot = agama.Potential(fb12_M17_contracted_dm_pot, M17_Baryon_pot)
# +
fb_vc = a_vcirc(M17_contracted_dm_pot, R_space)
fb15_vc = a_vcirc(fb15_M17_contracted_dm_pot, R_space)
fb12_vc = a_vcirc(fb12_M17_contracted_dm_pot, R_space)
plt.figure()
plt.plot(R_space, fb_vc, label='fbar=0.157')
plt.plot(R_space, fb15_vc, label='fbar=0.15')
plt.plot(R_space, fb12_vc, label='fbar=0.12')
plt.legend()
plt.show()
|
Cautun_Agama-Test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lmcanavals/algorithmic_complexity/blob/main/02_01_nqueens.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1gPMsFPVFEk-"
# # Nqueens (backtracking)
# + id="YIORgYf-zqtK"
import numpy as np
import matplotlib.pyplot as plt
# + id="dJMvPA_9z402"
def draw(board):
n = len(board)
b = np.zeros((n, n, 3), dtype=np.int)
b += [255, 150, 80]
b[::2, ::2] = [255, 220, 150]
b[1::2, 1::2] = [255, 220, 150]
_, ax = plt.subplots()
ax.imshow(b)
for row, col in enumerate(board):
ax.text(col, row, u"\u265b", fontsize=200/n, va="center", ha="center")
ax.set(xticks=[], yticks=[])
ax.axis("image")
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="glUUISPD55Zm" outputId="c9a76f1d-3c71-43d7-ed95-35b25eb00972"
draw([2, 7, 0, 5, 6, 3, 1, 4])
# + id="STu7Yv4S8xq4"
def valid(board, row, col):
n = len(board)
for row_i in range(row):
col_i = board[row_i]
delta = row - row_i
if col_i == col or col_i - delta == col or col_i + delta == col:
return False
return True
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="oMn7k55S6BCt" outputId="db6bdb61-dab1-4cb3-d7a3-3555ce60d249"
board = [0, 0, -1, -1]
draw(board)
assert valid(board, 1, 0) == False
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="S9ujJRYS8kZP" outputId="8b49b136-6119-4c21-c1f1-5b449afcad35"
board = [1, 3, 0, -1]
draw(board)
assert valid(board, 2, 0) == True
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="-mZvhmR79RVJ" outputId="d79d6342-f029-4c22-f62f-e85cf51b26ab"
board = [1, 3, 2, -1]
draw(board)
assert valid(board, 2, 2) == False
# + id="SxKFPD3A9mE-"
def nqueens(board, row):
n = len(board)
if row == n:
draw(board)
else:
for col_i in range(n):
if valid(board, row, col_i):
board[row] = col_i
nqueens(board, row + 1)
# + colab={"base_uri": "https://localhost:8080/"} id="H66zn-HP98F_" outputId="4c570607-c6cf-4f4e-d66d-b696410e0a1c"
[-1]*10
# + colab={"base_uri": "https://localhost:8080/", "height": 487} id="v-2dx6Zn_DE6" outputId="24e7f0fa-19ae-49f7-b69e-640db7b417c9"
n = 4
nqueens([-1]*n, 0)
# + id="nM5By21e_Hzg"
|
02_01_nqueens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Iris plants dataset
# --------------------
#
# **Data Set Characteristics:**
#
# :Number of Instances: 150 (50 in each of three classes)
# :Number of Attributes: 4 numeric, predictive attributes and the class
# :Attribute Information:
# - sepal length in cm
# - sepal width in cm
# - petal length in cm
# - petal width in cm
# - class:
# - Iris-Setosa
# - Iris-Versicolour
# - Iris-Virginica
#
# :Summary Statistics:
#
# ============== ==== ==== ======= ===== ====================
# Min Max Mean SD Class Correlation
# ============== ==== ==== ======= ===== ====================
# sepal length: 4.3 7.9 5.84 0.83 0.7826
# sepal width: 2.0 4.4 3.05 0.43 -0.4194
# petal length: 1.0 6.9 3.76 1.76 0.9490 (high!)
# petal width: 0.1 2.5 1.20 0.76 0.9565 (high!)
# ============== ==== ==== ======= ===== ====================
#
# :Missing Attribute Values: None
# :Class Distribution: 33.3% for each of 3 classes.
# :Creator: <NAME>
# :Donor: <NAME> (<EMAIL>)
# :Date: July, 1988
#
# The famous Iris database, first used by <NAME>. The dataset is taken
# from Fisher's paper. Note that it's the same as in R, but not as in the UCI
# Machine Learning Repository, which has two wrong data points.
#
# This is perhaps the best known database to be found in the
# pattern recognition literature. Fisher's paper is a classic in the field and
# is referenced frequently to this day. (See Duda & Hart, for example.) The
# data set contains 3 classes of 50 instances each, where each class refers to a
# type of iris plant. One class is linearly separable from the other 2; the
# latter are NOT linearly separable from each other.
#
# .. topic:: References
#
# - <NAME>. "The use of multiple measurements in taxonomic problems"
# Annual Eugenics, 7, Part II, 179-188 (1936); also in "Contributions to
# Mathematical Statistics" (John Wiley, NY, 1950).
# - <NAME>., & <NAME>. (1973) Pattern Classification and Scene Analysis.
# (Q327.D83) John Wiley & Sons. ISBN 0-471-22361-1. See page 218.
# - <NAME>. (1980) "Nosing Around the Neighborhood: A New System
# Structure and Classification Rule for Recognition in Partially Exposed
# Environments". IEEE Transactions on Pattern Analysis and Machine
# Intelligence, Vol. PAMI-2, No. 1, 67-71.
# - <NAME>. (1972) "The Reduced Nearest Neighbor Rule". IEEE Transactions
# on Information Theory, May 1972, 431-433.
# - See also: 1988 MLC Proceedings, 54-64. Cheeseman et al"s AUTOCLASS II
# conceptual clustering system finds 3 classes in the data.
# - Many, many more ...
# ### Importing modules
#
# Create the basic setup, load the dataset and separate the variables.
# +
import numpy as np
import seaborn as sns
import pandas as pd
from pandas import DataFrame
from matplotlib import pyplot as plt
from sklearn.datasets import load_iris
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
sns.set(font_scale=.8, rc={'figure.figsize': (12, 12,)})
D = load_iris()
X = D.data
y = D.target
features = D.feature_names
df = pd.DataFrame(data=X, columns=features)
# -
# ### Some facts
#
# We have:
# * 150 measurement
# * Features
# - sepal length in cm
# - sepal width in cm
# - petal length in cm
# - petal width in cm
# * Classes
# - Iris-Setosa
# - Iris-Versicolour
# - Iris-Virginica
# ## Visualization
#
# Now I try to find the best two features where the three classes can be separated clearly with bare eye.
pd.plotting.scatter_matrix(df, c=y, marker='s', cmap='rainbow', grid=False);plt.show()
# Let's see the correlation between the features.
sns.heatmap(df.corr().round(2), annot=True);plt.show()
# So, from these plots, my deduction is that the best two feature is the "Petal Width" and "Petal Length" to distinguish the classes from each other.
plt.scatter(df['petal width (cm)'], df['petal length (cm)'], c=y, cmap='winter');
plt.xlabel('petal width (cm)')
plt.ylabel('petal length (cm)')
plt.show()
# ## Preprocessing
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.model_selection import train_test_split
# First randomly select the train and test datapoints.
X = np.c_[df['petal width (cm)'].values, df['petal length (cm)'].values]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, random_state=0, shuffle=True)
# Let's normalize the datapoints
# +
scaler = MinMaxScaler()
X_train_processed = scaler.fit_transform(X_train)
X_test_processed = scaler.transform(X_test)
X_processed = scaler.transform(X)
# -
# Use one-hot encoding on the target values
# +
encoder = OneHotEncoder()
y_train_processed = encoder.fit_transform(y_train.reshape(-1,1)).todense()
y_test_processed = encoder.transform(y_test.reshape(-1,1)).todense()
y_processed = encoder.transform(y.reshape(-1,1)).todense()
def decode(x):
global encoder
return encoder.inverse_transform(x).reshape(-1,1)
# -
# ## Selecting the right model
#
# As a model I think that the K-Nearest Neighbors classifier is suitable for our classification problem.
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import cross_val_score
# To see what is the difference between a model who got preprocessed variables as input and the other who got raw variables, I make 2 classifier.
# This is model is working with unprocessed data.
# +
_knn = KNeighborsClassifier(n_jobs=-1, n_neighbors=3)
_knn.fit(X_train, y_train)
print('Accuracy on train:', np.mean(cross_val_score(_knn, X_train, y_train, cv=5)))
print('Accuracy on test:', np.mean(cross_val_score(_knn, X_test, y_test, cv=5)))
# -
# This is model is working with preprocessed data.
# +
knn = KNeighborsClassifier(n_jobs=-1, n_neighbors=3)
knn.fit(X_train_processed, y_train_processed)
print('Accuracy on train:', np.mean(cross_val_score(_knn, X_train_processed, y_train_processed, cv=5)))
print('Accuracy on test:', np.mean(cross_val_score(_knn, X_test_processed, y_test_processed, cv=5)))
# -
# ### Visualize the results
# +
# X = np.c_[df['sepal length (cm)'].values, df['petal length (cm)'].values]
xx_min, xx_max = X[:, 0].min() - 1, X[:, 0].max() + 1
yy_min, yy_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(xx_min, xx_max, .02), np.arange(yy_min, yy_max, .02))
plane = knn.predict(scaler.transform(np.c_[xx.ravel(), yy.ravel()]))
plane = decode(plane).reshape(xx.shape)
plt.contourf(xx, yy, plane, cmap='binary', alpha=0.5)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap='inferno', marker='x')
plt.show()
# -
# ### Trying out other classifier models
# +
from sklearn.ensemble import GradientBoostingClassifier
gbc = GradientBoostingClassifier(n_estimators=200, max_depth=10, learning_rate=0.2)
gbc.fit(X_train_processed, y_train)
print('Accuracy on train:', np.mean(cross_val_score(gbc, X_train, y_train, cv=3)))
print('Accuracy on test:', np.mean(cross_val_score(gbc, X_test, y_test, cv=3)))
xx, yy = np.meshgrid(np.arange(X[:, 0].min() - 1, X[:, 0].max() + 1, .02),
np.arange(X[:, 1].min() - 1, X[:, 1].max() + 1, .02))
plane = gbc.predict(scaler.transform(np.c_[xx.ravel(), yy.ravel()])).reshape(xx.shape)
plt.contourf(xx, yy, plane, cmap='binary', alpha=0.5)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap='inferno', marker='x')
plt.title('Classification with Gradient Boosting')
plt.show()
# +
from sklearn.neural_network import MLPClassifier
nn = MLPClassifier(max_iter=1000, hidden_layer_sizes=(500,), learning_rate_init=.002)
nn.fit(X_train_processed, y_train_processed)
print('Accuracy on train:', np.mean(cross_val_score(nn, X_train_processed, y_train_processed, cv=3)))
print('Accuracy on test:', np.mean(cross_val_score(nn, X_test_processed, y_test_processed, cv=3)))
xx, yy = np.meshgrid(np.arange(X[:, 0].min() - 1, X[:, 0].max() + 1, .02),
np.arange(X[:, 1].min() - 1, X[:, 1].max() + 1, .02))
plane = decode(knn.predict(scaler.transform(np.c_[xx.ravel(), yy.ravel()]))).reshape(xx.shape)
plt.contourf(xx, yy, plane, cmap='binary', alpha=0.5)
plt.scatter(X[:, 0], X[:, 1], c=y, cmap='inferno', marker='x')
plt.title('Classification with MLP Neural Network')
plt.show()
|
notebooks/Iris_dataset_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/JBNU-2021/blob/main/Predictive_Analytics/decision_tree/xgboost_pdp_ice.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="KF_Z4KRii4jy"
# ## License
#
# Copyright 2017 - 2020 <NAME> and the H2O.ai team
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="sppVg-qli4j1"
# **DISCLAIMER:** This notebook is not legal compliance advice.
# + [markdown] id="-OkbF5Wni4j3"
# # Engineering Transparency into Your Machine Learning Model with Python and XGBoost
# #### Monotonic XGBoost models, partial dependence, ICE, and Shapley explanations
# + [markdown] id="hyT7K2Hgi4j4"
# A key to building interpretable models is to limit their complexity. The more complex a model is, the harder it is to explain and understand. Overly complex models can also make unstable predictions on new data, which is both difficult to explain and makes models harder to trust. Monotonicity constraints not only simplify models, but do so in a way that is somewhat natural for human reasoning, increasing the transparency of predictive models. Under monotonicity constraints, model predictions can only increase or only decrease as an input variable value increases, and the direction of the constraint is typically specified by the user for logical reasons. For instance, a model might be constrained to produce only increasing probabilities of a certain medical condition as a patient's age increases, or to make only increasing predictions for home prices as a home's square footage increases.
#
# In this notebook a gradient boosting machine (GBM) is trained with monotonicity constraints to predict credit card payment defaults, using the UCI credit card default data, Python, NumPy, Pandas, and XGBoost. First, the credit card default data is loaded and prepared. Then Pearson correlation with the prediction target is used to determine the direction of the monotonicity constraints for each input variable and the model is trained. After the model is trained, partial dependence and individual conditional expectation (ICE) plots are used to analyze and verify the model's monotonic behavior. Finally an example of creating regulator mandated reason codes from high fidelity Shapley explanations for any model prediction is presented. This combination of monotonic XGBoost, partial dependence, ICE, and Shapley explanations is probably the most direct way to create an interpretable machine learning model today.
#
# **Note**: As of the h2o 3.24 "Yates" release, Shapley values are supported in h2o, in addition to GBM monotonicity constraints and partial dependence. To see Shapley values and monotonicity constraints for an h2o GBM in action please see: https://github.com/jphall663/interpretable_machine_learning_with_python/blob/master/dia.ipynb.
# + [markdown] id="3E6oR-SZi4j7"
# #### Python imports
# + [markdown] id="vsXrlpvXi4j8"
# Let's start with Python package imports. NumPy is used for basic arrray, vector, and matrix calculations. Pandas is used for data frame manipulation and plotting, and XGBoost is used to train a GBM with monotonicity constraints.
# + id="XYfC15tyj5G6"
# !pip install shap
# + id="9mE30VYPi4j9"
import numpy as np # array, vector, matrix calculations
import pandas as pd # DataFrame handling
import shap # for consistent, signed variable importance measurements
import xgboost as xgb # gradient boosting machines (GBMs)
import matplotlib.pyplot as plt # plotting
pd.options.display.max_columns = 999 # enable display of all columns in notebook
# enables display of plots in notebook
# %matplotlib inline
np.random.seed(12345) # set random seed for reproducibility
# + [markdown] id="0K4nGpH4i4kA"
# ## 1. Download, explore, and prepare UCI credit card default data
#
# UCI credit card default data: https://archive.ics.uci.edu/ml/datasets/default+of+credit+card+clients
#
# The UCI credit card default data contains demographic and payment information about credit card customers in Taiwan in the year 2005. The data set contains 23 input variables:
#
# * **`LIMIT_BAL`**: Amount of given credit (NT dollar)
# * **`SEX`**: 1 = male; 2 = female
# * **`EDUCATION`**: 1 = graduate school; 2 = university; 3 = high school; 4 = others
# * **`MARRIAGE`**: 1 = married; 2 = single; 3 = others
# * **`AGE`**: Age in years
# * **`PAY_0`, `PAY_2` - `PAY_6`**: History of past payment; `PAY_0` = the repayment status in September, 2005; `PAY_2` = the repayment status in August, 2005; ...; `PAY_6` = the repayment status in April, 2005. The measurement scale for the repayment status is: -1 = pay duly; 1 = payment delay for one month; 2 = payment delay for two months; ...; 8 = payment delay for eight months; 9 = payment delay for nine months and above.
# * **`BILL_AMT1` - `BILL_AMT6`**: Amount of bill statement (NT dollar). `BILL_AMNT1` = amount of bill statement in September, 2005; `BILL_AMT2` = amount of bill statement in August, 2005; ...; `BILL_AMT6` = amount of bill statement in April, 2005.
# * **`PAY_AMT1` - `PAY_AMT6`**: Amount of previous payment (NT dollar). `PAY_AMT1` = amount paid in September, 2005; `PAY_AMT2` = amount paid in August, 2005; ...; `PAY_AMT6` = amount paid in April, 2005.
#
# These 23 input variables are used to predict the target variable, whether or not a customer defaulted on their credit card bill in late 2005. Because XGBoost accepts only numeric inputs, all variables will be treated as numeric.
# + [markdown] id="7wCk_x45i4kE"
# #### Import data and clean
# The credit card default data is available as an `.xls` file. Pandas reads `.xls` files automatically, so it's used to load the credit card default data and give the prediction target a shorter name: `DEFAULT_NEXT_MONTH`.
# + id="9FiSz7-ti4kF"
# import XLS file
path = './credit_cards_dataset.csv'
data = pd.read_csv(path) # skip the first row of the spreadsheet
# remove spaces from target column name
#data = data.rename(columns={'default payment next month': 'DEFAULT_NEXT_MONTH'})
# + [markdown] id="FjQ-_aOji4kH"
# #### Assign modeling roles
# + [markdown] id="xPYGLNs_i4kI"
# The shorthand name `y` is assigned to the prediction target. `X` is assigned to all other input variables in the credit card default data except the row indentifier, `ID`.
# + colab={"base_uri": "https://localhost:8080/"} id="ZDRrPD-ci4kJ" outputId="33252b5b-9a06-4abf-a18d-339dd0dc1dbc"
# assign target and inputs for GBM
y = 'default.payment.next.month'
X = [name for name in data.columns if name not in [y, 'ID']]
print('y =', y)
print('X =', X)
# + [markdown] id="b0di-KBJi4kM"
# #### Display descriptive statistics
# The Pandas `describe()` function displays a brief description of the credit card default data. The input variables `SEX`, `EDUCATION`, `MARRIAGE`, `PAY_0`-`PAY_6`, and the prediction target `DEFAULT_NEXT_MONTH`, are really categorical variables, but they have already been encoded into meaningful numeric, integer values, which is great for XGBoost. Also, there are no missing values in this dataset.
# + id="kGT8URJgi4kN"
data[X + [y]].describe() # display descriptive statistics for all columns
# + [markdown] id="mhKW5NdAi4kP"
# ## 2. Investigate pair-wise Pearson correlations for DEFAULT_NEXT_MONTH
#
# Monotonic relationships are much easier to explain to colleagues, bosses, customers, and regulators than more complex, non-monotonic relationships and monotonic relationships may also prevent overfitting and excess error due to variance for new data.
#
# To train a transparent monotonic classifier, contraints must be supplied to XGBoost that determine whether the learned relationship between an input variable and the prediction target `DEFAULT_NEXT_MONTH` will be increasing for increases in an input variable or decreasing for increases in an input variable. Pearson correlation provides a linear measure of the direction of the relationship between each input variable and the target. If the pair-wise Pearson correlation between an input and `DEFAULT_NEXT_MONTH` is positive, it will be constrained to have an increasing relationship with the predictions for `DEFAULT_NEXT_MONTH`. If the pair-wise Pearson correlation is negative, the input will be constrained to have a decreasing relationship with the predictions for `DEFAULT_NEXT_MONTH`.
#
# Constrainsts are supplied to XGBoost in the form of a Python tuple with length equal to the number of inputs. Each item in the tuple is associated with an input variable based on its index in the tuple. The first constraint in the tuple is associated with the first variable in the training data, the second constraint in the tuple is associated with the second variable in the training data, and so on. The constraints themselves take the form of a 1 for a positive relationship and a -1 for a negative relationship.
# + [markdown] id="Nd3Bmp5Mi4kQ"
# #### Calculate Pearson correlation
# + [markdown] id="QNBUKlufi4kR"
# The Pandas `.corr()` function returns the pair-wise Pearson correlation between variables in a Pandas DataFrame. Because `DEFAULT_NEXT_MONTH` is the last column in the `data` DataFrame, the last column of the Pearson correlation matrix indicates the direction of the linear relationship between each input variable and the prediction target, `DEFAULT_NEXT_MONTH`. According to the calculated values, as a customer's balance limit (`LIMIT_BAL`), bill amounts (`BILL_AMT1`-`BILL_AMT6`), and payment amounts (`PAY_AMT1`-`PAY_AMT6`) increase, their probability of default tends to decrease. However as a customer's number of late payments increase (`PAY_0`, `PAY_2`-`PAY6`), their probability of default usually increases. In general, the Pearson correlation values make sense, and they will be used to ensure that the modeled relationships will make sense as well. (Pearson correlation values between the target variable, DEFAULT_NEXT_MONTH, and each input variable are displayed directly below.)
# + id="ptLYBpGUi4kS"
# displays last column of Pearson correlation matrix as Pandas DataFrame
pd.DataFrame(data[X + [y]].corr()[y]).iloc[:-1]
# + [markdown] id="wi-y4Cw8i4kU"
# #### Create tuple of monotonicity constraints from Pearson correlation values
# + [markdown] id="b2m9rr-pi4kU"
# The last column of the Pearson correlation matrix is transformed from a numeric column in a Pandas DataFrame into a Python tuple of `1`s and `-1`s that will be used to specifiy monotonicity constraints for each input variable in XGBoost. If the Pearson correlation between an input variable and `DEFAULT_NEXT_MONTH` is positive, a positive montonic relationship constraint is specified for that variable using `1`. If the correlation is negative, a negative monotonic constraint is specified using `-1`. (Specifying `0` indicates that no constraints should be used.) The resulting tuple will be passed to XGBoost when the GBM model is trained.
# + id="IXBq09ZVi4kV"
# creates a tuple in which positive correlation values are assigned a 1
# and negative correlation values are assigned a -1
mono_constraints = tuple([int(i) for i in np.sign(data[X + [y]].corr()[y].values[:-1])])
# (-1, -1, 1, -1, 1, 1, 1, 1, 1, 1, 1, -1, -1, -1, -1, -1, -1, -1, -1, -1, -1, -1, -1)
# + [markdown] id="JofPi229i4kW"
# ## 3. Train XGBoost with monotonicity constraints
#
# XGBoost is a very accurate, open source GBM library for regression and classification tasks. XGBoost can learn complex relationships between input variables and a target variable, but here the `monotone_constraints` tuning parameter is used to enforce monotonicity between inputs and the prediction for `DEFAULT_NEXT_MONTH`. XGBoost's early stopping functionality is also used to limit overfitting to the training data
#
# XGBoost is available from: https://github.com/dmlc/xgboost and the implementation of XGBoost is described in detail here: http://www.kdd.org/kdd2016/papers/files/rfp0697-chenAemb.pdf.
#
# After training, GBM variable importance is calculated and displayed. GBM variable importance is a global measure of the overall impact of an input variable on the GBM model predictions. Global variable importance values give an indication of the magnitude of a variable's contribution to model predictions for all observations. To enhance trust in the GBM model, variable importance values should typically conform to human domain knowledge and reasonable expectations.
# + [markdown] id="ZxU260YHi4kX"
# #### Split data into training and test sets for early stopping
# + [markdown] id="Fd8EK9ysi4kY"
# The credit card default data is split into training and test sets to monitor and prevent overtraining. Reproducibility is another important factor in creating trustworthy models, and randomly splitting datasets can introduce randomness in model predictions and other results. A random seed is used here to ensure the data split is reproducible.
# + colab={"base_uri": "https://localhost:8080/"} id="ARUP-rONi4kZ" outputId="25c23b2d-1076-453a-a686-d892b05913f4"
np.random.seed(12345) # set random seed for reproducibility
split_ratio = 0.7 # 70%/30% train/test split
# execute split
split = np.random.rand(len(data)) < split_ratio
train = data[split]
test = data[~split]
# summarize split
print('Train data rows = %d, columns = %d' % (train.shape[0], train.shape[1]))
print('Test data rows = %d, columns = %d' % (test.shape[0], test.shape[1]))
# + [markdown] id="ogh8VzjAi4ka"
# #### Train XGBoost GBM classifier
# To train an XGBoost classifier, the training and test data must be converted from Pandas DataFrames into SVMLight format. The `DMatrix()` function in the XGBoost package is used to convert the data. Many XGBoost tuning parameters must be specified as well. Typically a grid search would be performed to identify the best parameters for a given modeling task. For brevity's sake, a previously-discovered set of good tuning parameters are specified here. Notice that the monotonicity constraints are passed to XGBoost using the `monotone_constraints` parameter. Because gradient boosting methods typically resample training data, an additional random seed is also specified for XGBoost using the `seed` paramter to create reproducible predictions, error rates, and variable importance values. To avoid overfitting, the `early_stopping_rounds` parameter is used to stop the training process after the test area under the curve (AUC) statistic fails to increase for 50 iterations.
# + id="0KcoR4CHi4ka"
# XGBoost uses SVMLight data structure, not Numpy arrays or Pandas DataFrames
dtrain = xgb.DMatrix(train[X], train[y])
dtest = xgb.DMatrix(test[X], test[y])
# used to calibrate predictions to mean of y
base_y = train[y].mean()
# tuning parameters
params = {
'objective': 'binary:logistic', # produces 0-1 probabilities for binary classification
'booster': 'gbtree', # base learner will be decision tree
'eval_metric': 'auc', # stop training based on maximum AUC, AUC always between 0-1
'eta': 0.08, # learning rate
'subsample': 0.9, # use 90% of rows in each decision tree
'colsample_bytree': 0.9, # use 90% of columns in each decision tree
'max_depth': 15, # allow decision trees to grow to depth of 15
'monotone_constraints': 1, # 1 = increasing relationship, -1 = decreasing relationship
'base_score': base_y, # calibrate predictions to mean of y
'seed': 12345 # set random seed for reproducibility
}
# watchlist is used for early stopping
watchlist = [(dtrain, 'train'), (dtest, 'eval')]
# train model
xgb_model = xgb.train(params, # set tuning parameters from above
dtrain, # training data
1000, # maximum of 1000 iterations (trees)
evals=watchlist, # use watchlist for early stopping
early_stopping_rounds=50, # stop after 50 iterations (trees) without increase in AUC
verbose_eval=True) # display iteration progress
# + [markdown] id="fcN3220yi4kc"
# #### Global Shapley variable importance
# By setting `pred_contribs=True`, XGBoost's `predict()` function will return Shapley values for each row of the test set. Instead of relying on traditional single-value variable importance measures, local Shapley values for each input will be ploted below to get a more holistic and consisent measurement for the global importance of each input variable. Shapley values are introduced in greater detail in Section 6 below, but for now notice the monotonicity of the input variable contributions displayed in the Shapley summary plot.
# + id="zEMtwE2ui4kc"
# dtest is DMatrix
# shap_values is Numpy array
shap_values = xgb_model.predict(dtest, pred_contribs=True, ntree_limit=xgb_model.best_ntree_limit)
# + id="YaQ064BHi4kd"
# plot Shapley variable importance summary
shap.summary_plot(shap_values[:, :-1], test[xgb_model.feature_names])
# + [markdown] id="1vT2sv5oi4ke"
# #### Display Shapley variable importance summary
# The variable importance ranking should be parsimonious with human domain knowledge and reasonable expectations. In this case, `PAY_0` is by far the most important variable. As someone's most recent behavior is a very good indicator of future behavior, this checks out.
# + [markdown] id="mvDLxrkVi4kf"
# ## 4. Calculating partial dependence and ICE to validate and explain monotonic behavior
#
# Partial dependence plots are used to view the global, average prediction behavior of a variable under the monotonic model. Partial dependence plots show the average prediction of the monotonic model as a function of specific values of an input variable of interest, indicating how the monotonic GBM predictions change based on the values of the input variable of interest, while taking nonlinearity into consideration and averaging out the effects of all other
# input variables. Partial dependence plots enable increased transparency into the monotonic GBM's mechanisms and enable validation and debugging of the monotonic GBM by comparing a variable's average predictions across its domain to known standards and reasonable expectations. Partial dependence plots are described in greater detail in *The Elements of Statistical Learning*, section 10.13: https://web.stanford.edu/~hastie/ElemStatLearn/printings/ESLII_print12.pdf.
#
# Individual conditional expectation (ICE) plots, a newer and less well-known adaptation of partial dependence plots, can be used to create more localized explanations for a single observation of data using the same basic ideas as partial dependence plots. ICE is also a type of nonlinear sensitivity analysis in which the model predictions for a single observation are measured while a feature of interest is varied over its domain. ICE increases understanding and transparency by displaying the nonlinear behavior of the monotonic GBM. ICE also enhances trust, accountability, and fairness by enabling comparisons of nonlinear behavior to human domain knowledge and reasonable expectations. ICE, as a type of sensitivity analysis, can also engender trust when model behavior on simulated or extreme data points is acceptable. A detailed description of ICE is available in this arXiv preprint: https://arxiv.org/abs/1309.6392.
#
# Because partial dependence and ICE are measured on the same scale, they can be displayed in the same line plot to compare the global, average prediction behavior for the entire model and the local prediction behavior for certain rows of data. Overlaying the two types of curves enables analysis of both global and local behavior simultaneously and provides an indication of the trustworthiness of the average behavior represented by partial dependence. (Partial dependence can be misleading in the presence of strong interactions or correlation. ICE curves diverging from the partial dependence curve can be indicative of such problems.) Histograms are also presented with the partial dependence and ICE curves, to enable a rough measure of epistemic uncertainty for model predictions: predictions based on small amounts of training data are likely less dependable.
# + [markdown] id="jxFgkmzni4kg"
# #### Function for calculating partial dependence
# Since partial dependence and ICE will be calculated for several important variables in the GBM model, it's convenient to have a function doing so. It's probably best to analyze partial dependence and ICE for all variables in a model, but only the top three most important input variables will be investigated here. It's also a good idea to analyze partial dependence and ICE on the test data, or other holdout datasets, to see how the model will perform on new data.
# This simple function is designed to return partial dependence when it is called for an entire dataset and ICE when it is called for a single row. The `bins` argument will be used later to calculate ICE values at the same places in an input variable domain that partial dependence is calculated directly below.
# + id="TgLdZICvi4kg"
def par_dep(xs, frame, model, resolution=20, bins=None):
""" Creates Pandas DataFrame containing partial dependence for a
single variable.
Args:
xs: Variable for which to calculate partial dependence.
frame: Pandas DataFrame for which to calculate partial dependence.
model: XGBoost model for which to calculate partial dependence.
resolution: The number of points across the domain of xs for which
to calculate partial dependence, default 20.
bins: List of values at which to set xs, default 20 equally-spaced
points between column minimum and maximum.
Returns:
Pandas DataFrame containing partial dependence values.
"""
# turn off pesky Pandas copy warning
pd.options.mode.chained_assignment = None
# initialize empty Pandas DataFrame with correct column names
par_dep_frame = pd.DataFrame(columns=[xs, 'partial_dependence'])
# cache original column values
col_cache = frame.loc[:, xs].copy(deep=True)
# determine values at which to calculate partial dependence
if bins == None:
min_ = frame[xs].min()
max_ = frame[xs].max()
by = (max_ - min_)/resolution
bins = np.arange(min_, max_, by)
# calculate partial dependence
# by setting column of interest to constant
# and scoring the altered data and taking the mean of the predictions
for j in bins:
frame.loc[:, xs] = j
dframe = xgb.DMatrix(frame)
par_dep_i = pd.DataFrame(model.predict(dframe, ntree_limit=model.best_ntree_limit))
par_dep_j = par_dep_i.mean()[0]
par_dep_frame = par_dep_frame.append({xs:j,
'partial_dependence': par_dep_j},
ignore_index=True)
# return input frame to original cached state
frame.loc[:, xs] = col_cache
return par_dep_frame
# + [markdown] id="SqEQ2vxBi4kh"
# #### Calculate partial dependence for the most important input variables in the GBM
# The partial dependence for `LIMIT_BAL` can be seen to decrease as credit balance limits increase. This finding is aligned with expectations that the model predictions will be monotonically decreasing with increasing `LIMIT_BAL` and parsimonious with well-known business practices in credit lending. Partial dependence for other important values is displayed in plots further below.
# + colab={"base_uri": "https://localhost:8080/", "height": 672} id="prUazBoqi4ki" outputId="41c2f5d9-4a84-43f1-b247-33ddca0f5044"
par_dep_PAY_0 = par_dep('PAY_0', test[X], xgb_model) # calculate partial dependence for PAY_0
par_dep_LIMIT_BAL = par_dep('LIMIT_BAL', test[X], xgb_model) # calculate partial dependence for LIMIT_BAL
par_dep_BILL_AMT1 = par_dep('BILL_AMT1', test[X], xgb_model) # calculate partial dependence for BILL_AMT1
# display partial dependence for LIMIT_BAL
par_dep_LIMIT_BAL
# + [markdown] id="DxFNJr3zi4kj"
# #### Helper function for finding percentiles of predictions
# + [markdown] id="ZVkhh1F8i4kk"
# ICE can be calculated for any row in the training or test data, but without intimate knowledge of a data source it can be difficult to know where to apply ICE. Calculating and analyzing ICE curves for every row of training and test data set can be overwhelming, even for the example credit card default dataset. One place to start with ICE is to calculate ICE curves at every decile of predicted probabilities in a dataset, giving an indication of local prediction behavior across the dataset. The function below finds and returns the row indices for the maximum, minimum, and deciles of one column in terms of another -- in this case, the model predictions (`p_DEFAULT_NEXT_MONTH`) and the row identifier (`ID`), respectively.
# + id="VV-41wpWi4kk"
def get_percentile_dict(yhat, id_, frame):
""" Returns the percentiles of a column, yhat, as the indices based on
another column id_.
Args:
yhat: Column in which to find percentiles.
id_: Id column that stores indices for percentiles of yhat.
frame: Pandas DataFrame containing yhat and id_.
Returns:
Dictionary of percentile values and index column values.
"""
# create a copy of frame and sort it by yhat
sort_df = frame.copy(deep=True)
sort_df.sort_values(yhat, inplace=True)
sort_df.reset_index(inplace=True)
# find top and bottom percentiles
percentiles_dict = {}
percentiles_dict[0] = sort_df.loc[0, id_]
percentiles_dict[99] = sort_df.loc[sort_df.shape[0]-1, id_]
# find 10th-90th percentiles
inc = sort_df.shape[0]//10
for i in range(1, 10):
percentiles_dict[i * 10] = sort_df.loc[i * inc, id_]
return percentiles_dict
# + [markdown] id="NpqFX1avi4kl"
# #### Find some percentiles of yhat in the test data
# The values for `ID` that correspond to the maximum, minimum, and deciles of `p_DEFAULT_NEXT_MONTH` are displayed below. ICE will be calculated for the rows of the test dataset associated with these `ID` values.
# + id="OqL_aD1Zi4km"
# merge GBM predictions onto test data
yhat_test = pd.concat([test.reset_index(drop=True), pd.DataFrame(xgb_model.predict(dtest,
ntree_limit=xgb_model.best_ntree_limit))],
axis=1)
yhat_test = yhat_test.rename(columns={0:'p_DEFAULT_NEXT_MONTH'})
# find percentiles of predictions
percentile_dict = get_percentile_dict('p_DEFAULT_NEXT_MONTH', 'ID', yhat_test)
# display percentiles dictionary
# ID values for rows
# from lowest prediction
# to highest prediction
percentile_dict
# + [markdown] id="H-meQ656i4kn"
# #### Calculate ICE curve values
# ICE values represent a model's prediction for a row of data while an input variable of interest is varied across its domain. The values of the input variable are chosen to match the values at which partial dependence was calculated earlier, and ICE is calculated for the top three most important variables and for rows at each percentile of the test dataset.
# + id="3quYP24Ai4ko"
# retreive bins from original partial dependence calculation
bins_PAY_0 = list(par_dep_PAY_0['PAY_0'])
bins_LIMIT_BAL = list(par_dep_LIMIT_BAL['LIMIT_BAL'])
bins_BILL_AMT1 = list(par_dep_BILL_AMT1['BILL_AMT1'])
# for each percentile in percentile_dict
# create a new column in the par_dep frame
# representing the ICE curve for that percentile
# and the variables of interest
for i in sorted(percentile_dict.keys()):
col_name = 'Percentile_' + str(i)
# ICE curves for PAY_0 across percentiles at bins_PAY_0 intervals
par_dep_PAY_0[col_name] = par_dep('PAY_0',
test[test['ID'] == int(percentile_dict[i])][X],
xgb_model,
bins=bins_PAY_0)['partial_dependence']
# ICE curves for LIMIT_BAL across percentiles at bins_LIMIT_BAL intervals
par_dep_LIMIT_BAL[col_name] = par_dep('LIMIT_BAL',
test[test['ID'] == int(percentile_dict[i])][X],
xgb_model,
bins=bins_LIMIT_BAL)['partial_dependence']
# ICE curves for BILL_AMT1 across percentiles at bins_BILL_AMT1 intervals
par_dep_BILL_AMT1[col_name] = par_dep('BILL_AMT1',
test[test['ID'] == int(percentile_dict[i])][X],
xgb_model,
bins=bins_BILL_AMT1)['partial_dependence']
# + [markdown] id="puvxfzDli4kp"
# #### Display partial dependence and ICE for `LIMIT_BAL`
# Partial dependence and ICE values for rows at the minimum, maximum and deciles (0%, 10%, 20%, ..., 90%, 99%) of predictions for `DEFAULT_NEXT_MONTH` and at the values of `LIMIT_BAL` used for partial dependence are shown here. Each column of ICE values will be a curve in the plots below. ICE values represent a prediction for a row of test data, at a percentile of interest noted in the column name above, and setting `LIMIT_BAL` to the value of `LIMIT_BAL` at right. Notice that monotonic decreasing prediction behavior for `LIMIT_BAL` holds at each displayed percentile of predicted `DEFAULT_NEXT_MONTH`, helping to validate that the trained GBM predictions are monotonic for `LIMIT_BAL`.
# + id="jdTzHIari4kp"
par_dep_LIMIT_BAL
# + [markdown] id="OFa7YAXii4kq"
# ## 5. Plotting partial dependence and ICE to validate and explain monotonic behavior
#
# Overlaying partial dependence onto ICE in a plot is a convenient way to validate and understand both global and local monotonic behavior. Plots of partial dependence curves overlayed onto ICE curves for several percentiles of predictions for `DEFAULT_NEXT_MONTH` are used to validate monotonic behavior, describe the GBM model mechanisms, and to compare the most extreme GBM behavior with the average GBM behavior in the test data. Partial dependence and ICE plots are displayed for the three most important variables in the GBM: `PAY_0`, `LIMIT_BAL`, and `BILL_AMT1`.
# + id="Sd-YVrPni4kr"
#### Function to plot partial dependence and ICE
def plot_par_dep_ICE(xs, par_dep_frame):
""" Plots ICE overlayed onto partial dependence for a single variable.
Args:
xs: Name of variable for which to plot ICE and partial dependence.
par_dep_frame: Name of Pandas DataFrame containing ICE and partial
dependence values.
"""
# initialize figure and axis
fig, ax = plt.subplots()
# plot ICE curves
par_dep_frame.drop('partial_dependence', axis=1).plot(x=xs,
colormap='gnuplot',
ax=ax)
# overlay partial dependence, annotate plot
par_dep_frame.plot(title='Partial Dependence and ICE for ' + str(xs),
x=xs,
y='partial_dependence',
style='r-',
linewidth=3,
ax=ax)
# add legend
_ = plt.legend(bbox_to_anchor=(1.05, 0),
loc=3,
borderaxespad=0.)
# + [markdown] id="LViEgyspi4ks"
# #### Partial dependence and ICE plot for `LIMIT_BAL`
# The monotonic prediction behavior displayed in the partial dependence, and ICE tables for `LIMIT_BAL` is also visible in this plot. Monotonic decreasing behavior is evident at every percentile of predictions for `DEFAULT_NEXT_MONTH`. Most percentiles of predictions show that sharper decreases in probability of default occur when `LIMIT_BAL` increases just slightly from its lowest values in the test set. However, for the custumers that are most likely to default according to the GBM model, no increase in `LIMIT_BAL` has a strong impact on probabilitiy of default. As mentioned previously, the displayed relationship between credit balance limits and probablility of default is not uncommon in credit lending. As can be seen from the displayed histogram, above ~$NT 500,000 prediction behavior may have been learned from extremely small samples of data.
# + id="mmXyHBJJi4kt"
plot_par_dep_ICE('LIMIT_BAL', par_dep_LIMIT_BAL) # plot partial dependence and ICE for LIMIT_BAL
# + id="QUO4dmHGi4ku"
_ = train['LIMIT_BAL'].plot(kind='hist', bins=20, title='Histogram: LIMIT_BAL')
# + [markdown] id="xuvt7tLIi4kv"
# #### Partial dependence and ICE plot for `PAY_0`
# Monotonic increasing prediction behavior for `PAY_0` is displayed for all percentiles of model predictions. Predition behavior is different at different deciles, but not abnormal or vastly different from the average prediction behavior represented by the red partial dependence curve. The largest jump in predicted probability appears to occur at `PAY_0 = 2`, or when a customer becomes two months late on their most recent payment. Above `PAY_0 = 3` there are few examples from which the model could learn.
# + id="cgYQfRFti4kv"
plot_par_dep_ICE('PAY_0', par_dep_PAY_0) # plot partial dependence and ICE for PAY_0
# + id="RsdvOJw4i4kw"
_ = train['PAY_0'].plot(kind='hist', bins=20, title='Histogram: PAY_0')
# + [markdown] id="aCu0aMbbi4kx"
# #### Partial dependence and ICE plot for `BILL_AMT1`
# Monotonic decreasing prediction behavior for `BILL_AMT1` is also displayed for all percentiles. This mild decrease in probability of default as most recent bill amount increases could be related to wealthier, big-spending customers taking on more debt but also being able to pay it off reliably. Also, customers with negative bills are more likely to default, potentially indicating charge-offs are being recorded as negative bills. In a mission-critical situation, this issue would require more debugging. Also predictions below \$ NT 0 and above \$ NT 400,000 are based on very little training data.
# + id="8DYWIzLIi4kx"
plot_par_dep_ICE('BILL_AMT1', par_dep_BILL_AMT1) # plot partial dependence and ICE for BILL_AMT1
# + id="pxDCu8W0i4ky"
_ = train['BILL_AMT1'].plot(kind='hist', bins=20, title='Histogram: BILL_AMT1')
# + [markdown] id="UfOE0VMNi4kz"
# ## 6. Generate reason codes using the Shapley method
# Now that the monotonic behavior of the GBM has been verified and compared against domain knowledge and reasonable expectations, a method called Shapley explanations will be used to calculate the local variable importance for any one prediction: http://papers.nips.cc/paper/7062-a-unified-approach-to-interpreting-model-predictions. Shapley explanations are the only possible consistent local variable importance values. (Here consistency means that if a variable is more important than another variable in a given prediction, the more important variable's Shapley value will not be smaller in magnitude than the less important variable's Shapley value.) Very crucially Shapley values also *always* sum to the actual prediction of the XGBoost model. When used in a model-specific context for decision tree models, Shapley values are likely the most accurate known local variable importance method available today. In this notebook, XGBoost itself is used to create Shapley values with the `pred_contribs` parameter to `predict()`, but the `shap` package is also available for other types of models: https://github.com/slundberg/shap.
#
# The numeric Shapley values in each column are an estimate of how much each variable contributed to each prediction. Shapley contributions can indicate how a variable and its values were weighted in any given decision by the model. These values are crucially important for machine learning interpretability and are related to "local feature importance", "reason codes", or "turn-down codes." The latter phrases are borrowed from credit scoring. Credit lenders in the U.S. must provide reasons for automatically rejecting a credit application. Reason codes can be easily extracted from Shapley local variable contribution values by ranking the variables that played the largest role in any given decision.
# + [markdown] id="yQPIGV5Oi4kz"
# To find the index corresponding to a particular row of interest later, the index of the `test` DataFrame is reset to begin at 0 and increase sequentially. Without resetting the index, the `test` DataFrame row indices still correspond to the original raw data from which the test set was sampled.
# + id="QzLgKVzZi4k0"
test.reset_index(drop=True, inplace=True)
# + [markdown] id="-45zN_Opi4k0"
# #### Select most risky customer in test data
# One person who might be of immediate interest is the most likely to default customer in the test data. This customer's row will be selected and local variable importance for the corresponding prediction will be analyzed.
# + id="NsENXqZ4i4k1"
decile = 99
row = test[test['ID'] == percentile_dict[decile]]
# + [markdown] id="BvAd0VFCi4k1"
# #### Create a Pandas DataFrame of Shapley values for riskiest customer
# The most interesting Shapley values are probably those that push this customer's probability of default higher, i.e. the highest positive Shapley values. Those values are plotted below.
# + id="RxXvfJVli4k1"
# reset test data index to find riskiest customer in shap_values
# sort to find largest positive contributions
s_df = pd.DataFrame(shap_values[row.index[0], :][:-1].reshape(23, 1), columns=['Reason Codes'], index=X)
s_df.sort_values(by='Reason Codes', inplace=True, ascending=False)
# + id="uMqyGknki4k2"
s_df
# + [markdown] id="v-GVOrDxi4k2"
# #### Plot top local contributions as reason codes
# + id="hbuo89i4i4k3"
_ = s_df[:5].plot(kind='bar',
title='Top Five Reason Codes for a Risky Customer\n',
legend=False)
# + [markdown] id="mkASixIri4k3"
# For the customer in the test dataset that the GBM predicts as most likely to default, the most important input variables in the prediction are, in descending order, `PAY_0`, `PAY_5`, `PAY_6`, `PAY_2`, and `LIMIT_BAL`.
# + [markdown] id="TRSt6h4ki4k4"
# #### Display customer in question
# + [markdown] id="GVom6-1Gi4k4"
# The local contributions for this customer appear reasonable, especially when considering her payment information. Her most recent payment was 3 months late and her payment for 6 months and 5 months previous were 7 months late. Also her credit limit was extremely low, so it's logical that these factors would weigh heavily into the model's prediction for default for this customer.
# + id="AuiqUGUVi4k5"
row # helps understand reason codes
# + [markdown] id="B1OwnNyLi4k5"
# To generate reason codes for the model's decision, the locally important variable and its value are used together. If this customer was denied future credit based on this model and data, the top five Shapley-based reason codes for the automated decision would be:
#
# 1. Most recent payment is 3 months delayed.
# 2. 5th most recent payment is 7 months delayed.
# 3. 6th most recent payment is 7 months delayed.
# 4. 2nd most recent payment is 2 months delayed.
# 5. Credit limit is too low: 10,000 $NT.
#
# (Of course, credit limits are set by the lender and are used to price-in risk to credit decisions, so using credit limits as reason codes or even in a probability of default model is likely questionable. However, in this small, example data set all input columns were used to generate a better model fit. For a slightly more careful treatment of gradient boosting in the context of credit scoring, please see: https://github.com/jphall663/interpretable_machine_learning_with_python/blob/master/dia.ipynb)
# + [markdown] id="4doiJCF3i4k6"
# #### Summary
#
# In this notebook, a highly transparent, nonlinear, monotonic GBM classifier was trained to predict credit card defaults and the monotonic behavior of the classifier was analyzed and validated. To do so, Pearson correlation between each input and the target was used to determine the direction for monotonicity constraints for each input variable in the XGBoost classifier. GBM variable importance, partial dependence, and ICE were calculated, plotted, and compared to one another, domain knowledge, and reasonable expectations. Shapley values were then used to explain the model predictions for the single most risky customer in the test set. These techniques should generalize well for many types of business and research problems, enabling you to train a monotonic GBM model and analyze, validate, and explain it to your colleagues, bosses, and potentially, external regulators.
|
Model/xgboost_pdp_ice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importing libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
dataset=pd.read_csv('Salary_Data.csv')
x=dataset.iloc[:,:-1].values
y=dataset.iloc[:,1].values
# Splitting the dataset into the Training set and Test set
from sklearn.cross_validation import train_test_split
x_train,x_test,y_train,y_test=train_test_split(x,y, test_size=1/3,random_state=0)
# Fitting Linear Regression to the Training Set
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(x_train,y_train)
# Predicting the Test set results
y_pred =regressor.predict(x_test)
# Visualising the Training set results
plt.scatter(x_train,y_train,color='red')
plt.plot(x_train,regressor.predict(x_train),color='blue')
plt.title('Salary vs Experience (Training set)')
plt.xlabel('Years of experience')
plt.ylabel('Salary')
plt.show()
# Visualising the Test set results
plt.scatter(x_test,y_test,color='red')
plt.plot(x_train,regressor.predict(x_train),color='blue')
plt.title('Salary vs Experience (Test set)')
plt.xlabel('Years of experience')
plt.ylabel('Salary')
plt.show()
|
Day 7/Linear Regression/Linear Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,py
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:generic_expression] *
# language: python
# name: conda-env-generic_expression-py
# ---
# # Coverage of MultiPLIER LV
#
# The goal of this notebook is to examine why genes were found to be generic. Specifically, this notebook is trying to answer the question: Are generic genes found in more multiplier latent variables compared to specific genes?
#
# The PLIER model performs a matrix factorization of gene expression data to get two matrices: loadings (Z) and latent matrix (B). The loadings (Z) are constrained to aligned with curated pathways and gene sets specified by prior knowledge [Figure 1B of Taroni et. al.](https://www.cell.com/cell-systems/pdfExtended/S2405-4712(19)30119-X). This ensure that some but not all latent variables capture known biology. The way PLIER does this is by applying a penalty such that the individual latent variables represent a few gene sets in order to make the latent variables more interpretable. Ideally there would be one latent variable associated with one gene set unambiguously.
#
# While the PLIER model was trained on specific datasets, MultiPLIER extended this approach to all of recount2, where the latent variables should correspond to specific pathways or gene sets of interest. Therefore, we will look at the coverage of generic genes versus other genes across these MultiPLIER latent variables, which represent biological patterns.
#
# **Definitions:**
# * Generic genes: Are genes that are consistently differentially expressed across multiple simulated experiments.
#
# * Other genes: These are all other non-generic genes. These genes include those that are not consistently differentially expressed across simulated experiments - i.e. the genes are specifically changed in an experiment. It could also indicate genes that are consistently unchanged (i.e. housekeeping genes)
# +
# %load_ext autoreload
# %autoreload 2
import os
import random
import textwrap
import scipy
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
import rpy2.robjects as ro
from rpy2.robjects import pandas2ri
from rpy2.robjects.conversion import localconverter
from ponyo import utils
from generic_expression_patterns_modules import lv
# +
# Get data directory containing gene summary data
base_dir = os.path.abspath(os.path.join(os.getcwd(), "../"))
data_dir = os.path.join(base_dir, "human_general_analysis")
# Read in config variables
config_filename = os.path.abspath(
os.path.join(base_dir, "configs", "config_human_general.tsv")
)
params = utils.read_config(config_filename)
local_dir = params["local_dir"]
project_id = params["project_id"]
quantile_threshold = 0.98
# -
# Output file
nonzero_figure_filename = "nonzero_LV_coverage.svg"
highweight_figure_filename = "highweight_LV_coverage.svg"
# ## Load data
# Get gene summary file
summary_data_filename = os.path.join(data_dir, f"generic_gene_summary_{project_id}.tsv")
# +
# Load gene summary data
data = pd.read_csv(summary_data_filename, sep="\t", index_col=0, header=0)
# Check that genes are unique since we will be using them as dictionary keys below
assert data.shape[0] == len(data["Gene ID"].unique())
# -
# Load multiplier models
# Converted formatted pickle files (loaded using phenoplier environment) from
# https://github.com/greenelab/phenoplier/blob/master/nbs/01_preprocessing/005-multiplier_recount2_models.ipynb
# into .tsv files
multiplier_model_z = pd.read_csv(
"multiplier_model_z.tsv", sep="\t", index_col=0, header=0
)
# Get a rough sense for how many genes contribute to a given LV
# (i.e. how many genes have a value != 0 for a given LV)
# Notice that multiPLIER is a sparse model
(multiplier_model_z != 0).sum().sort_values(ascending=True)
# ## Get gene data
#
# Define generic genes based on simulated gene ranking. Refer to [figure](https://github.com/greenelab/generic-expression-patterns/blob/master/human_general_analysis/gene_ranking_log2FoldChange.svg) as a guide.
#
# **Definitions:**
# * Generic genes: `Percentile (simulated) >= 60`
#
# (Having a high rank indicates that these genes are consistently changed across simulated experiments.)
#
# * Other genes: `Percentile (simulated) < 60`
#
# (Having a lower rank indicates that these genes are not consistently changed across simulated experiments - i.e. the genes are specifically changed in an experiment. It could also indicate genes that are consistently unchanged.)
generic_threshold = 60
dict_genes = lv.get_generic_specific_genes(data, generic_threshold)
# +
# Check overlap between multiplier genes and our genes
multiplier_genes = list(multiplier_model_z.index)
our_genes = list(data.index)
shared_genes = set(our_genes).intersection(multiplier_genes)
print(len(our_genes))
print(len(shared_genes))
# -
# Drop gene ids not used in multiplier analysis
processed_dict_genes = lv.process_generic_specific_gene_lists(
dict_genes, multiplier_model_z
)
# Check numbers add up
assert len(shared_genes) == len(processed_dict_genes["generic"]) + len(
processed_dict_genes["other"]
)
# ## Get coverage of LVs
#
# For each gene (generic or other) we want to find:
# 1. The number of LVs that gene is present
# 2. The number of LVs that the gene contributes a lot to (i.e. the gene is highly weighted within that LV)
# ### Nonzero LV coverage
dict_nonzero_coverage = lv.get_nonzero_LV_coverage(
processed_dict_genes, multiplier_model_z
)
# Check genes mapped correctly
assert processed_dict_genes["generic"][0] in dict_nonzero_coverage["generic"].index
assert len(dict_nonzero_coverage["generic"]) == len(processed_dict_genes["generic"])
assert len(dict_nonzero_coverage["other"]) == len(processed_dict_genes["other"])
# ### High weight LV coverage
# Quick look at the distribution of gene weights per LV
sns.distplot(multiplier_model_z["LV3"], kde=False)
plt.yscale("log")
dict_highweight_coverage = lv.get_highweight_LV_coverage(
processed_dict_genes, multiplier_model_z, quantile_threshold
)
# Check genes mapped correctly
assert processed_dict_genes["generic"][0] in dict_highweight_coverage["generic"].index
assert len(dict_highweight_coverage["generic"]) == len(processed_dict_genes["generic"])
assert len(dict_highweight_coverage["other"]) == len(processed_dict_genes["other"])
# ### Assemble LV coverage and plot
# +
all_coverage = []
for gene_label in dict_genes.keys():
merged_df = pd.DataFrame(
dict_nonzero_coverage[gene_label], columns=["nonzero LV coverage"]
).merge(
pd.DataFrame(
dict_highweight_coverage[gene_label], columns=["highweight LV coverage"]
),
left_index=True,
right_index=True,
)
merged_df["gene type"] = gene_label
all_coverage.append(merged_df)
all_coverage_df = pd.concat(all_coverage)
# -
all_coverage_df = lv.assemble_coverage_df(
processed_dict_genes, dict_nonzero_coverage, dict_highweight_coverage
)
all_coverage_df.head()
# +
# Plot coverage distribution given list of generic coverage, specific coverage
nonzero_fig = sns.boxplot(
data=all_coverage_df,
x="gene type",
y="nonzero LV coverage",
notch=True,
palette=["#81448e", "lightgrey"],
)
# Manually add statistical annotations based on t-tests below
x1, x2 = 0, 1 # columns 'Sat' and 'Sun' (first column: 0, see plt.xticks())
y, h, col = all_coverage_df["nonzero LV coverage"].max() + 30, 30, "k"
plt.plot([x1, x1, x2, x2], [y, y + h, y + h, y], lw=1.5, c=col)
plt.text(
(x1 + x2) * 0.5, y + h + 10, "p-value = 0.239", ha="center", va="bottom", color=col
)
nonzero_fig.set(ylim=(0, 800))
nonzero_fig.set_xlabel(None)
nonzero_fig.set_xticklabels(
["common DEGs", "other genes"], fontsize=14, fontname="Verdana"
)
nonzero_fig.set_ylabel(
textwrap.fill("Number of LVs", width=30), fontsize=14, fontname="Verdana"
)
nonzero_fig.tick_params(labelsize=14)
nonzero_fig.set_title(
"Number of LVs genes are present in", fontsize=16, fontname="Verdana"
)
# +
# Plot coverage distribution given list of generic coverage, specific coverage
highweight_fig = sns.boxplot(
data=all_coverage_df,
x="gene type",
y="highweight LV coverage",
notch=True,
palette=["#81448e", "lightgrey"],
)
# Manually add statistical annotations based on t-tests below
x1, x2 = 0, 1 # columns 'Sat' and 'Sun' (first column: 0, see plt.xticks())
y, h, col = all_coverage_df["highweight LV coverage"].max() + 10, 10, "k"
plt.plot([x1, x1, x2, x2], [y, y + h, y + h, y], lw=1.5, c=col)
plt.text(
(x1 + x2) * 0.5,
y + h + 5,
"p-value = 6.31e-119",
ha="center",
va="bottom",
color=col,
)
highweight_fig.set(ylim=(0, 150))
highweight_fig.set_xlabel(None)
highweight_fig.set_xticklabels(
["common DEGs", "other genes"], fontsize=14, fontname="Verdana"
)
highweight_fig.set_ylabel(
textwrap.fill("Number of LVs", width=30), fontsize=14, fontname="Verdana"
)
highweight_fig.tick_params(labelsize=14)
highweight_fig.set_title(
"Number of LVs genes contribute highly to", fontsize=16, fontname="Verdana"
)
# -
# ## Calculate statistics
# * Is the reduction in generic coverage significant?
# * Is the difference between generic versus other genes signficant?
# +
# Test: mean number of LVs generic genes present in vs mean number of LVs that generic gene is high weight in
# (compare two blue boxes between plots)
generic_nonzero = all_coverage_df[all_coverage_df["gene type"] == "generic"][
"nonzero LV coverage"
].values
generic_highweight = all_coverage_df[all_coverage_df["gene type"] == "generic"][
"highweight LV coverage"
].values
(stats, pvalue) = scipy.stats.ttest_ind(generic_nonzero, generic_highweight)
print(pvalue)
# +
# Test: mean number of LVs generic genes present in vs mean number of LVs other genes high weight in
# (compare blue and grey boxes in high weight plot)
other_highweight = all_coverage_df[all_coverage_df["gene type"] == "other"][
"highweight LV coverage"
].values
generic_highweight = all_coverage_df[all_coverage_df["gene type"] == "generic"][
"highweight LV coverage"
].values
(stats, pvalue) = scipy.stats.ttest_ind(other_highweight, generic_highweight)
print(pvalue)
# +
# Check that coverage of other and generic genes across all LVs is NOT signficantly different
# (compare blue and grey boxes in nonzero weight plot)
other_nonzero = all_coverage_df[all_coverage_df["gene type"] == "other"][
"nonzero LV coverage"
].values
generic_nonzero = all_coverage_df[all_coverage_df["gene type"] == "generic"][
"nonzero LV coverage"
].values
(stats, pvalue) = scipy.stats.ttest_ind(other_nonzero, generic_nonzero)
print(pvalue)
# -
# ## Get LVs that generic genes are highly weighted in
#
# Since we are using quantiles to get high weight genes per LV, each LV has the same number of high weight genes. For each set of high weight genes, we will get the proportion of generic vs other genes. We will select the LVs that have a high proportion of generic genes to examine.
# Get proportion of generic genes per LV
prop_highweight_generic_dict = lv.get_prop_highweight_generic_genes(
processed_dict_genes, multiplier_model_z, quantile_threshold
)
# Return selected rows from summary matrix
multiplier_model_summary = pd.read_csv(
"multiplier_model_summary.tsv", sep="\t", index_col=0, header=0
)
lv.create_LV_df(
prop_highweight_generic_dict,
multiplier_model_summary,
0.5,
"Generic_LV_summary_table.tsv",
)
# Plot distribution of weights for these nodes
node = "LV61"
lv.plot_dist_weights(
node,
multiplier_model_z,
shared_genes,
20,
all_coverage_df,
f"weight_dist_{node}.svg",
)
# ## Save
# +
# Save plot
nonzero_fig.figure.savefig(
nonzero_figure_filename,
format="svg",
bbox_inches="tight",
transparent=True,
pad_inches=0,
dpi=300,
)
# Save plot
highweight_fig.figure.savefig(
highweight_figure_filename,
format="svg",
bbox_inches="tight",
transparent=True,
pad_inches=0,
dpi=300,
)
# -
# **Takeaway:**
# * In the first nonzero boxplot, generic and other genes are present in a similar number of LVs. This isn't surprising since the number of genes that contribute to each LV is <1000.
# * In the second highweight boxplot, other genes are highly weighted in more LVs compared to generic genes. This would indicate that generic genes contribute alot to few LVs.
#
# This is the opposite trend found using [_P. aeruginosa_ data](1_get_eADAGE_LV_coverage.ipynb). Perhaps this indicates that generic genes have different behavior/roles depending on the organism. In humans, perhaps these generic genes are related to a few hyper-responsive pathways, whereas in _P. aeruginosa_ perhaps generic genes are associated with many pathways, acting as *gene hubs*.
#
# * There are a number of LVs that contain a high proportion of generic genes can be found in [table](Generic_LV_summary_table.tsv). By quick visual inspection, it looks like many LVs are associated with immune response, signaling and metabolism. Which are consistent with the hypothesis that these generic genes are related to hyper-responsive pathways.
|
LV_analysis/1_get_multiplier_LV_coverage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import collections
import os
import string
import sys
import pandas as pd
from nltk import word_tokenize
from nltk.corpus import stopwords
from pprint import pprint
from sklearn.cluster import KMeans
from sklearn.feature_extraction.text import TfidfVectorizer
# +
import nltk
nltk.download('punkt')
# -
data_path = "../data/hopitaux/all/"
# # Choix d'une décennie et du nombre de clusters
DECADE = ''
N_CLUSTERS = 5
# # Chargement des fichiers de la décennie
files = [f for f in sorted(os.listdir(data_path)) if f"_{DECADE[:-1]}" in f]
# Exemple de fichiers
files[:5]
texts = [open(data_path + f, encoding='utf-8').read() for f in files]
# Exemple de textes
texts[0][:400]
# # Vectorisation du texte
def process_text(text, stem=True):
""" Tokenize text and remove punctuation """
text = text.translate(string.punctuation)
tokens = word_tokenize(text)
return tokens
vectorizer = TfidfVectorizer(tokenizer=process_text,
stop_words=stopwords.words('french'),
max_df=0.5,
min_df=0.1,
lowercase=True)
# %time tfidf_vectors = vectorizer.fit_transform(texts)
tfidf_vectors
# Exemple de vecteur TFIDF
pd.Series(
tfidf_vectors[0].toarray()[0],
index=vectorizer.get_feature_names()
).sort_values(ascending=False)
# # Clustering des vecteurs TFIDF
km_model = KMeans(n_clusters=N_CLUSTERS)
km_model.fit(tfidf_vectors)
# +
clustering = collections.defaultdict(list)
for idx, label in enumerate(km_model.labels_):
clustering[label].append(files[idx])
# -
pprint(dict(clustering))
|
module4/s2_clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
from chempy.chemistry import Species, Equilibrium
from chempy.equilibria import EqSystem
# + deletable=true editable=true
Na_p = Species('Na+', 1, composition={11: 1})
Cl_m = Species('Cl-', -1, composition={17: 1})
NaCl = Species('NaCl', phase_idx=1, composition={11: 1, 17: 1})
eq = Equilibrium({'NaCl': 1}, {'Na+': 1, 'Cl-': 1}, 4.0)
eqsys = EqSystem([eq], [Na_p, Cl_m, NaCl])
# + deletable=true editable=true
eqsys.root({'Na+': 5, 'Cl-': 5, 'NaCl': 0})
# + deletable=true editable=true
eqsys.root({'Na+': 1, 'Cl-': 1, 'NaCl': 0})
# + deletable=true editable=true
eqsys.root({'Na+': 2, 'Cl-': 2, 'NaCl': 0})
# + deletable=true editable=true
eqsys.root({'Na+': 0, 'Cl-': 0, 'NaCl': 2})
# + deletable=true editable=true
eqsys.root({'Na+': 0, 'Cl-': 0, 'NaCl': 5.0})
# + deletable=true editable=true
eqsys.root({'Na+': 0, 'Cl-': 0, 'NaCl': 5.0}, rref_preserv=True)
# + deletable=true editable=true
from chempy.equilibria import NumSysLog
eqsys.root({'Na+': 0, 'Cl-': 0, 'NaCl': 5.0}, rref_preserv=True, NumSys=NumSysLog)
# + deletable=true editable=true
from chempy.equilibria import NumSysLin
eqsys.root({'Na+': 0, 'Cl-': 0, 'NaCl': 5.0}, rref_preserv=True, NumSys=(NumSysLog, NumSysLin))
# + deletable=true editable=true
|
examples/NaCl_precipitation.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.1
# language: julia
# name: julia-1.7
# ---
using LinearAlgebra, Plots, Interact
# $$
# u(x) = \int_0^x \cos t dt = \sin x
# $$
n = 10
x = range(0, 1; length=n)
h = step(x) # x[k+1]-x[k] = 1/(n-1)
d = [1; fill(1/h, n-1)]
l = fill(-1/h, n-1)
L = Bidiagonal(d, l, :L)
scatter(x, zero(x); label="grid")
m = (x[1:end-1] + x[2:end])/2
scatter!(m, zero(m); label="mid")
x
@manipulate for n = 2:100, ω=1:100
x = range(0, 1; length=n)
m = (x[1:end-1] + x[2:end])/2
h = step(x) # x[k+1]-x[k] = 1/(n-1)
d = [1; fill(1/h, n-1)]
l = fill(-1/h, n-1)
L = Bidiagonal(d, l, :L)
c = 0
f = x -> cos(ω*x)*ω
u = x -> sin(ω*x)
𝐟ᶠ = f.(x[1:end-1])
𝐟ᵐ = f.(m)
𝐮ᶠ = L \ [c; 𝐟ᶠ]
𝐮ᵐ = L \ [c; 𝐟ᵐ]
g = range(0,1; length=1000)
plot(g, u.(g); label="exact", legend=:bottomright, ylims=(-1,1))
scatter!(x, 𝐮ᶠ; label="forward")
scatter!(x, 𝐮ᵐ; label="mid")
end
# +
function forward_err(u, c, f, n)
x = range(0, 1; length=n)
h = step(x) # x[k+1]-x[k] = 1/(n-1)
d = [1; fill(1/h, n-1)]
l = fill(-1/h, n-1)
L = Bidiagonal(d, l, :L)
𝐟ᶠ = f.(x[1:end-1])
𝐮ᶠ = L \ [c; 𝐟ᶠ]
norm(𝐮ᶠ - u.(x), Inf)
end
function mid_err(u, c, f, n)
x = range(0, 1; length=n)
m = (x[1:end-1] + x[2:end])/2
h = step(x) # x[k+1]-x[k] = 1/(n-1)
d = [1; fill(1/h, n-1)]
l = fill(-1/h, n-1)
L = Bidiagonal(d, l, :L)
𝐟 = f.(m)
𝐮 = L \ [c; 𝐟]
norm(𝐮 - u.(x), Inf)
end
# -
ns = 10 .^(1:8)
scatter(ns, forward_err.(sin, 0, cos, ns); yscale=:log10, xscale=:log10, label="forward")
plot!(ns, ns .^ (-1); label="1/n")
scatter!(ns, mid_err.(sin, 0, cos, ns); yscale=:log10, xscale=:log10, label="mid")
plot!(ns, ns .^ (-2); label="1/n^2")
# We observe forward-diff gives us O(1/n) convergence, mid-diff gives us O(1/n^2).
#
# We observe stability for both methods.
mid_err.(sin, 0, cos, 10^8)
|
lectures/Indefinite Integration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **SVM**
# Resumo
#
#
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.naive_bayes import GaussianNB
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler,MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
from sklearn.compose import make_column_transformer
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
# carregando a base de dados de censo
base = pd.read_csv('../../res/census.csv')
base
base.shape
# +
# separando os dados de previsao e classificacao
previsores = base.iloc[:, 0:14].values
classificadores = base.iloc[:, 14].values
#gerando uma copia dos dados originais para fazer mais testes abaixo
previsores_escalonados=previsores.copy()
#efetuando correcoes nos dados do censo
#transformando dados categorios da base em dados discretos
labelencoder_prev = LabelEncoder()
previsores[:, 1] = labelencoder_prev.fit_transform(previsores[:, 1])
previsores[:, 3] = labelencoder_prev.fit_transform(previsores[:, 3])
previsores[:, 5] = labelencoder_prev.fit_transform(previsores[:, 5])
previsores[:, 6] = labelencoder_prev.fit_transform(previsores[:, 6])
previsores[:, 7] = labelencoder_prev.fit_transform(previsores[:, 7])
previsores[:, 8] = labelencoder_prev.fit_transform(previsores[:, 8])
previsores[:, 9] = labelencoder_prev.fit_transform(previsores[:, 9])
previsores[:, 13] = labelencoder_prev.fit_transform(previsores[:, 13])
# +
# padronizando os valores nao discretos da copia dos previsores ( nao deve ser feito para todos os parametros sob risco de degradar a precisao do algoritimo)
#o min max scaler foi mais interessante para este caso
#scaler = MinMaxScaler(feature_range=(0, 1))
scaler = StandardScaler()
#transformando dados categorios da copia da base em dados discretos
labelencoder_prev = LabelEncoder()
previsores_escalonados[:, 1] = labelencoder_prev.fit_transform(previsores_escalonados[:, 1])
previsores_escalonados[:, 3] = labelencoder_prev.fit_transform(previsores_escalonados[:, 3])
previsores_escalonados[:, 5] = labelencoder_prev.fit_transform(previsores_escalonados[:, 5])
previsores_escalonados[:, 6] = labelencoder_prev.fit_transform(previsores_escalonados[:, 6])
previsores_escalonados[:, 7] = labelencoder_prev.fit_transform(previsores_escalonados[:, 7])
previsores_escalonados[:, 8] = labelencoder_prev.fit_transform(previsores_escalonados[:, 8])
previsores_escalonados[:, 9] = labelencoder_prev.fit_transform(previsores_escalonados[:, 9])
previsores_escalonados[:, 13] = labelencoder_prev.fit_transform(previsores_escalonados[:, 13])
print("\nVisualizando estatisticas dos dados nao discretos antes do escalonamento\n")
for x in range(3):
print('coluna ',x,"\n")
print(previsores_escalonados[:,[4,10,12]][x].min())
print(previsores_escalonados[:,[4,10,12]][x].max())
print(previsores_escalonados[:,[4,10,12]][x].mean())
print(previsores_escalonados[:,[4,10,12]][x].var())
print("\n")
#escalonando os previsores da copia dos dados
previsores_escalonados = scaler.fit_transform(previsores_escalonados)
print("\nVisualizando estatisticas dos dados nao discretos depois do escalonamento\n")
for x in range(3):
print('coluna ',x,"\n")
print(previsores_escalonados[:,[4,10,12]][x].min())
print(previsores_escalonados[:,[4,10,12]][x].max())
print(previsores_escalonados[:,[4,10,12]][x].mean())
print(previsores_escalonados[:,[4,10,12]][x].var())
print("\n")
labelencoder_classe = LabelEncoder()
classificadores = labelencoder_classe.fit_transform(classificadores)
#preprocess = make_column_transformer(( OneHotEncoder(categories='auto'), [1,3,5,6,7,8,9,13] ),remainder="passthrough")
#previsores_escalonados = preprocess.fit_transform(previsores_escalonados).toarray()
#previsores_escalonados = onehotencoder.fit_transform(previsores_escalonados).toarray()
#separando os valores de teste e treinamento para os previsores escalonados e nao escalonados
previsores_treinamento, previsores_teste, classificadores_treinamento1, classificadores_teste1 = train_test_split(previsores, classificadores, test_size=0.33, random_state=0)
previsores_escalonados_treinamento, previsores_escalonados_teste, classificadores_treinamento, classificadores_teste = train_test_split(previsores_escalonados, classificadores, test_size=0.33, random_state=0)
# +
# instanciando o classificador
classificador = classificador = SVC(gamma='scale',kernel='poly',max_iter=10000,degree=3) #kernel = 'linear',random_state = 1 ,max_iter=8000
classificador.fit(previsores_escalonados_treinamento, classificadores_treinamento)
# rodando previsoes com o dados de teste (copia)
previsoes_dados_escalonados = classificador.predict(previsores_escalonados_teste)
# fazendo o fit com os dados normais
classificador.fit(previsores_treinamento, classificadores_treinamento1)
previsoes = classificador.predict(previsores_teste)
# +
#testes dessa instancia algoritimo
# o dado de precisao per se nao quer dizer muita coisa e preciso verificar outras metricas
precisao_escalonados = accuracy_score(classificadores_teste, previsoes_dados_escalonados)
precisao = accuracy_score(classificadores_teste1, previsoes)
# uma dessas metricas eh a matriz de confusao ... ela e capaz de mostrar o desempenho do algoritimo para cada classe
matriz_escalonados = confusion_matrix(classificadores_teste, previsoes_dados_escalonados)
matriz = confusion_matrix(classificadores_teste1, previsoes)
#o scikit tambem possui uma classe utilitaria que prove um report mais detalhado...
report_escalonados = classification_report(classificadores_teste, previsoes_dados_escalonados)
report = classification_report(classificadores_teste1, previsoes)
print("Precisão dados normais / escalonados :\n")
print(precisao,'/',precisao_escalonados)
print("\nMatriz de confusão dados normais / escalonados:\n")
print(matriz)
print("\n")
print(matriz_escalonados)
print("\nReport dados normais / escalonados:\n")
print (report)
print("\n")
print (report_escalonados)
# -
# <br>**TODO :Verificar a base de treinamento para melhorar a distribuicao das classes e verificar se ha alguma melhora**
|
notebooks/estudos_python/machine_learning/svm/classificacao_svm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### First, let's add more features to see if that will improve.
from sklearn import datasets
import numpy as np
iris = datasets.load_iris()
#X = iris["data"][:,3:] # petal width
X = iris["data"][:,:]
y = (iris["target"]==2).astype(np.int)
X.shape, y.shape
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1, random_state=42)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size=0.2, random_state=42)
X_train.shape, X_val.shape, X_test.shape
# -
def sigmoid(z):
return 1 / (1 + np.exp(-z))
def binary_loss(X, Y, weights, alpha=0.00001):
"""
X.shape = (num_instances, num_features)
Y.shape = (num_instances,)
weights.shape = (num_features,)
"""
z = np.dot(X, weights)
h = sigmoid(z)
total_cost = (-Y*np.log(h) - (1-Y)*np.log(1-h)).mean() + alpha * np.sum(np.dot(weights, weights.T))
return total_cost
# +
from sklearn.preprocessing import MinMaxScaler, StandardScaler
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.00001, epochs=5, tolerance=0.0001, print_progress=True):
train_loss_history, val_loss_history = [], []
num_features = X_tr.shape[1]
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
weights = np.random.randn(num_features)
clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
for i in range(len(row)):
gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]
weights[i] = weights[i] - lr*gradient
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
val_loss = binary_loss(X_dev, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
if epoch >=1:
if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
break
return weights, train_loss_history, val_loss_history
# -
n_epochs = 100
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False)
import matplotlib.pyplot as plt
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
# +
#prediction =
# -
# ### Let's try to train with `LogisticRegression`
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression()
log_reg.fit(X_train, y_train)
from sklearn.metrics import confusion_matrix
y_val_pred = log_reg.predict(X_val)
confusion_matrix(y_val, y_val_pred)
y_test_pred = log_reg.predict(X_test)
confusion_matrix(y_test, y_test_pred)
# ### Bias term
# Maybe the problem was that we do not manually add a bias term?
X_train.shape
X_t = np.hstack([X_train, np.ones((X_train.shape[0], 1))])
X_t.shape
X_v = np.hstack([X_val, np.ones((X_val.shape[0], 1))])
X_v.shape
X_te = np.hstack([X_test, np.ones((X_test.shape[0], 1))])
X_te.shape
n_epochs = 100
w, train_hist, val_hist = SGD(X_t, y_train, X_dev=X_v, Y_dev=y_val, epochs=n_epochs, print_progress=False)
import matplotlib.pyplot as plt
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
# No, this is not where the problem lies.
# ### The most suspectible being the function `SGD()`
# Let's look closer into this function.
# ### No `clip()`
# - It is weird to `clip()` the training data values unless we are really sure about the range of our data values
# - I don't know the originally NLP dataset, but `clip()` is not suitable for Iris dataset here.
np.clip(-7, 0, 10), np.clip(38, 0, 10)
X_train[:10, :]
# Problems in the function `SGD()`:
# - The training loss is no longer quite accurate, albeit acceptable. In particular, it is unfair to compare `val_loss` with `train_loss`; only looking the history exclusively within each one makes sense.
# - $\frac{\partial L}{\partial w_i} = (h-y)x_i + 2\alpha w_i\quad$ if $\quad L = -y\log h - (1-y)\log (1-h) + \alpha \lVert w \rVert^2$
row = np.arange(1*3).reshape((1,3))
weights = np.random.randn(3)
row, weights
row @ weights
np.dot(row, weights)
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.00001, epochs=5, tolerance=0.0001, print_progress=True):
train_loss_history, val_loss_history = [], []
num_features = X_tr.shape[1]
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
weights = np.random.randn(num_features)
#clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
for i in range(len(row)):
gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]*2
weights[i] -= lr*gradient
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
val_loss = binary_loss(X_dev, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
#if epoch >=1:
# if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
# break
return weights, train_loss_history, val_loss_history
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False)
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
y_train_pred = (sigmoid(X_train @ w) > 0.5).astype(np.int)
y_train_pred
confusion_matrix(y_train, y_train_pred)
y_val_pred = (sigmoid(X_val @ w) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_test @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
n_epochs = 100
w, train_hist, val_hist = SGD(X_t, y_train, X_dev=X_v, Y_dev=y_val, epochs=n_epochs, print_progress=False)
#w, train_hist, val_hist = SGD(X_t, y_train, X_dev=X_v, Y_dev=y_val, epochs=n_epochs, print_progress=False, lr=0.04)
#w, train_hist, val_hist = SGD(X_t, y_train, X_dev=X_v, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
# ### Add bias term
# **(?)** Why adding bias term gets consistently bad results? Can you explain?
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.legend();
y_val_pred = (sigmoid((X_v @ w)) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_te @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
# ### Curious. I cannot seem to spot the bug.
# Let's try with [`sklearn.linear_model.SGDClassifier`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html) to see how good it can be.
from sklearn.linear_model import SGDClassifier
clf = SGDClassifier(loss="log")
clf.fit(X_train, y_train)
y_train_pred = clf.predict(X_train)
confusion_matrix(y_train, y_train_pred)
y_test_pred = clf.predict(X_test)
confusion_matrix(y_test, y_test_pred)
y_val_pred = clf.predict(X_val)
confusion_matrix(y_val, y_val_pred)
# ### Stopping criterion
# We didn't stop right at where our model's weight give best validation error, let's try that.
np.inf
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.0001, epochs=5, tolerance=0.0001, print_progress=True):
train_loss_history, val_loss_history = [], []
num_features = X_tr.shape[1]
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
weights = np.random.randn(num_features)
#clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
best_val_loss = np.inf
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
for i in range(len(row)):
gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]*2
weights[i] -= lr*gradient
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
val_loss = binary_loss(X_dev, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
#if epoch >=1:
# if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
# break
if val_loss < best_val_loss:
best_val_loss = val_loss
best_weights = weights
print(f"best epoch {epoch}")
return best_weights, train_loss_history, val_loss_history
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False)
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
y_train_pred = (sigmoid(X_train @ w) > 0.5).astype(np.int)
y_train_pred
confusion_matrix(y_train, y_train_pred)
y_val_pred = (sigmoid(X_val @ w) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_test @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
# ### Learning schedule
# +
t0, t1 = 5, 50
def learning_schedule(t):
return t0 / (t + t1)
# -
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.0001, epochs=5, tolerance=0.0001, print_progress=True):
train_loss_history, val_loss_history = [], []
num_features = X_tr.shape[1]
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
weights = np.random.randn(num_features)
#clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
best_val_loss = np.inf
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
for i in range(len(row)):
gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]*2
eta = learning_schedule(epoch*len(X_tr) + idx)
weights[i] -= lr*gradient*eta
#weights[i] -= gradient*eta
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
val_loss = binary_loss(X_dev, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
#if epoch >=1:
# if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
# break
if val_loss < best_val_loss:
best_val_loss = val_loss
best_weights = weights
print(f"best epoch {epoch}")
return best_weights, train_loss_history, val_loss_history
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False)
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
y_train_pred = (sigmoid(X_train @ w) > 0.5).astype(np.int)
y_train_pred
confusion_matrix(y_train, y_train_pred)
y_val_pred = (sigmoid(X_val @ w) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_test @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
# ### <NAME>'s learning schedule
np.zeros(1)
np.zeros(1).dtype
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.0001, epochs=5, tolerance=0.0001, print_progress=True):
def learning_schedule(t):
epsilon = 1e-5
return 1 / (t * alpha + epsilon)
train_loss_history, val_loss_history = [], []
num_features = X_tr.shape[1]
weights = np.zeros(shape=(num_features,))
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
#weights = np.random.randn(num_features)
#clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
best_val_loss = np.inf
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
eta = learning_schedule(epoch*len(X_tr) + idx + 1)
for i in range(len(row)):
#gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]*2
gradient = (sigmoid(row @ weights) - Y_tr[idx]) * row[0, i] + alpha*weights[i]*2
#weights[i] -= lr*gradient
weights[i] -= gradient*eta
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
val_loss = binary_loss(X_dev, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
#if epoch >=1:
# if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
# break
if val_loss < best_val_loss:
best_val_loss = val_loss
best_weights = weights
print(f"best epoch {epoch}")
return best_weights, train_loss_history, val_loss_history
n_epochs = 100
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False)
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=1e-1)
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
y_train_pred = (sigmoid(X_train @ w) > 0.5).astype(np.int)
y_train_pred
confusion_matrix(y_train, y_train_pred)
y_val_pred = (sigmoid(X_val @ w) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_test @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
# ### Did you notice that?
# `SGD()` tends to give all `1`'s or all `0`'s as prediction.<br>
# **(?)** Why was that?
# ### Bias term again
n_epochs = 200
w, train_hist, val_hist = SGD(X_t, y_train, X_dev=X_v, Y_dev=y_val, epochs=n_epochs, print_progress=False)
x = range(1, len(train_hist)+1)
plt.plot(x, train_hist, label="train_loss")
plt.plot(x, val_hist, label="val_loss")
plt.legend();
y_val_pred = (sigmoid((X_v @ w)) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_te @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
# ### `fit_intercept`? Bias term wrong?
def SGD(X_tr, Y_tr, X_dev=[], Y_dev=[], lr=0.1, alpha=0.0001, epochs=5, tolerance=0.0001, print_progress=True):
def learning_schedule(t):
epsilon = 1e-5
return 1 / (t * alpha + epsilon)
train_loss_history, val_loss_history = [], []
mms = MinMaxScaler()
#mms = StandardScaler()
X_tr = mms.fit_transform(X_tr)
X_tr = np.hstack([X_tr, np.ones((X_tr.shape[0], 1))])
num_features = X_tr.shape[1]
weights = np.zeros(shape=(num_features,))
#weights = np.zeros(shape=(num_features,), dtype=np.float64)
#weights = np.random.randn(num_features)
#clip_value = 1e-9
#X_tr = np.clip(X_tr, clip_value, 1-clip_value)
best_val_loss = np.inf
for epoch in range(epochs):
indices = np.arange(X_tr.shape[0])
np.random.shuffle(indices)
X_tr = X_tr[indices]
Y_tr = Y_tr[indices]
# training loop
train_loss = 0
for idx, row in enumerate(X_tr):
row = np.reshape(row, (1, num_features))
train_loss += binary_loss(row, Y_tr[idx], weights, alpha)
for i in range(len(row)):
#gradient = (sigmoid(row @ weights) - float(Y_tr[idx])) * row[0, i] + alpha*weights[i]*2
gradient = (sigmoid(row @ weights) - Y_tr[idx]) * row[0, i] + alpha*weights[i]*2
eta = learning_schedule(epoch*idx + i)
#weights[i] -= lr*gradient
weights[i] -= gradient*eta
train_loss /= X_tr.shape[0]
train_loss_history.append(train_loss)
if len(X_dev) != 0 and len(Y_dev) != 0:
X_dev = mms.transform(X_dev)
X_dex = np.hstack([X_dev, np.ones((X_dev.shape[0], 1))])
#val_loss = binary_loss(X_dev, Y_dev, weights[:-1], alpha)
val_loss = binary_loss(X_dex, Y_dev, weights, alpha)
val_loss_history.append(val_loss)
if print_progress:
print(f"Epoch: {epoch+1}")
print(f"Training loss: {train_loss:.4f}")
print(f"Validation loss: {val_loss:.4f}")
# Wang seems to want the training process to stop when the progress becomes no longer significant.
#if epoch >=1:
# if np.abs(val_loss_history[epoch]-val_loss_history[epoch-1]) < tolerance:
# break
if val_loss < best_val_loss:
best_val_loss = val_loss
best_weights = weights
print(f"best epoch {epoch}")
return best_weights, train_loss_history, val_loss_history
n_epochs = 200
w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=True)
#w, train_hist, val_hist = SGD(X_train, y_train, X_dev=X_val, Y_dev=y_val, epochs=n_epochs, print_progress=False, alpha=0)
y_val_pred = (sigmoid((X_v @ w)) > 0.5).astype(np.int)
y_val_pred
confusion_matrix(y_val, y_val_pred)
y_test_pred = (sigmoid(X_te @ w) > 0.5).astype(np.int)
y_test_pred
confusion_matrix(y_test, y_test_pred)
|
python_taiwan/logistic_regression/yang_wang/02_debug.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Factory
#
# The code does not have to worry about which
# localizer will be instantiated, since the method "localize" will be called
# in the same way independently of the language.
# +
class GreekLocalizer(object):
def __init__(self):
self.translations = {"dog": "σκύλος", "cat": "γάτα"}
def localize(self, msg):
return self.translations.get(msg, msg)
class EnglishLocalizer(object):
def localize(self, msg):
return msg
def get_localizer(lang="English"):
localizers = {
"English": EnglishLocalizer,
"Greek": GreekLocalizer
}
return localizers[lang]()
e, g = get_localizer("English"), get_localizer("Greek")
for msg in "dog parrot cat bear".split():
print(e.localize(msg), g.localize(msg))
|
creational/factory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (learn-env)
# language: python
# name: learn-env
# ---
# # General Overview
# ## Business Understanding
# Microsoft wants to know how to get into the movie businesss. In order for a company to succeed, it requires a sustainable business model. We are here to help substantiate hypotheses of potential business models with data. This particular data set helps us understand the relationship between the production budget of a movie and its gross revenue returns.
# ## Data Understanding
import pandas as pd
df = pd.read_csv('zippedData/tn.movie_budgets.csv.gz')
df.head()
df.info()
# ## Data Cleaning
# Luckily, we have no missing values. However, we need the data columns for production_budget, domestic_gross, and worldwide_gross to be integers, rather than strings, in order to properly analyze them.
#cleaning the production budget column, domestic_gross column, worldwide_gross column to be floats
df['prod_budg_fixed'] = df['production_budget'].str.replace('$', '').str.replace(',','').astype(float)
df['dom_gross_fixed'] = df['domestic_gross'].str.replace('$', '').str.replace(',','').astype(float)
df['world_gross_fixed'] = df['worldwide_gross'].str.replace('$', '').str.replace(',','').astype(float)
df.head()
df.describe()
# ## Data Analysis
# With the cleaning out of the way, we have a column of floats for production budget called prod_budg_fixed, a column of floats for domestic gross called dom_gross_fixed, and a column of floats for worldwide gross called world_gross_fixed. We can now start exploring relationships between production budget and the gross revenue
#Checking the Pearson Correlation Coefficient
df.corr(method='pearson')
# From here, we can see that there is a relatively high correlation between production budget and worldwide gross (0.748306), and an extremely high correlation between the domestic gross and worldwide gross (0.938853), indicating that a movie doing well in domestic theaters will also do well internationally. Logical, but nice to know.
# +
#scatterplot of production budget vs worldwide gross
import matplotlib.pyplot as plt
from matplotlib.axis import Axis
import seaborn as sns
import numpy as np
import pylab as plb
#code for basic scatter plot
x = df['prod_budg_fixed']
y = df['world_gross_fixed']
fig, ax = plt.subplots(figsize=(16, 10))
#going for more of a 'heatmap' style plot for readability, so opacity is low
ax.scatter(x, y, alpha=0.5)
#domestic gross scatter plot
y2 = df['dom_gross_fixed']
ax.scatter(x, y2, alpha=0.5, c='pink')
ax.set_title('Production Budget vs. Gross Revenue')
ax.set_xlabel('Production Budget')
ax.set_ylabel('Worldwide Gross (Blue) and Domestic Gross (Orange)')
ax.set_xlim(xmin=-5000, xmax=(3*10**8))
ax.set_ylim(ymin=-5000, ymax=(2*10**9))
plt.legend(['Worldwide', 'Domestic'])
#finding the trend line
z = np.polyfit(x,y,1)
p = np.poly1d(z)
plb.plot(x, p(x), 'b--')
#Supposedly the equation for the line (need to look into this more to understand it)
print("y=%.6fx+(%.6f)"%(z[0],z[1]))
#second trend line
z2 = np.polyfit(x,y2,1)
p2 = np.poly1d(z2)
plb.plot(x, p2(x), 'r--')
print("y2=%.6fx+(%.6f)"%(z2[0],z2[1]))
# -
# Based on what we see of the scatterplot above, as well as the trend lines, there is a positive correlation for both, with the worldwide gross having a considerably higher slope (meaning higher returns (gross) on investment (production budget)). Releasing a film internationally generally gives better returns than releasing a film purely domestically.
#
# Taking a look at the trend line linear equations:
#
# worldwide gross: y=3.126943x+(-7285667.054592)
#
# domestic gross: y2=1.119087x+(6523888.576660)
#
# Using these equations, we can make extremely general estimates of gross domestic earnings based on our production budget using basic linear algebra.
#
# x = production budget
#
# y-intercept (worldwide gross) = -7285667.05
#
# y2-intercept (domestic gross) = 6523888.57
#
# x-intercept (breakeven point for each):
# y (worldwide
#
#Variance of
df['world_gross_fixed'].var()
df['dom_gross_fixed'].var()
# There is a lower variance on the domestic gross, meaning that there is a higher likelihood that an estimate made using the domestic gross equation will be accurate
# ### Conclusion
#
|
Code/tn_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# Example 1
# =========
#
# Let us test ``cgn`` on a simple unconstrained linear least-squares problem. As an example, we use a regularized version of problem 32 from the article
#
# <NAME>., <NAME>. and <NAME>. "Testing Unconstrained Optimization Software", 1981.
#
# The problem is as follows:
#
# $
# \begin{align}
# \min_{x \in \mathbb R^n} & \frac{1}{2}||F(x)||_2^2 + \frac{\beta}{2}||x||_2^2, \\
# \text{where} \quad & F(x)_j = x_j - \frac{2}{m} \sum_{i=1}^n x_i - 1, \quad \text{for } 1 \leq j \leq n, \\
# & F(x)_j = - \frac{2}{m} \sum_{i=1}^n x_i - 1, \quad \text{for } n < j <= m, \\
# & \beta = \frac{1}{100},
# \end{align}
# $
#
# with $m >= n$. Let us choose $n=200$ and $m=400$.
# -
# First, we implement the affine misfit function $F=F(x)$.
# + pycharm={"name": "#%%\n"}
import numpy as np
m = 400
n = 200
def F(x):
z = np.zeros(m)
for i in range(n):
z[i] = x[i] - 2.0 * sum(x) / m - 1.
for i in range(n,m):
z[i] = - 2.0 * sum(x) / m - 1.
return z
# + [markdown] pycharm={"name": "#%% md\n"}
# We also have to implement its Jacobian:
#
# + pycharm={"name": "#%%\n"}
A = np.zeros((m,n))
# upper half of the Jacobian matrix
for i in range(n):
for j in range(n):
if i == j:
A[i,j] = 1.0 - 2.0 / m
else:
A[i,j] = -2.0 / m
# lower half of the Jacobian matrix
for i in range(n,m):
for j in range(n):
A[i,j] = -2.0 / m
def DF(x):
return A
# + [markdown] pycharm={"name": "#%% md\n"}
# Let us now set up the ``cgn.Parameter`` object. To create a ``cgn.Parameter`` object, the user has to provide a starting guess at initialization. This is necessary so that ``cgn`` can verify the consistency of the user input (e.g. that the dimensions match) before it starts solving the optimization problem. By doing that, it makes it easier to diagnose runtime errors that occur later. For our parameter $x$, we simply use the 1-vector as initial guess.
# + pycharm={"name": "#%%\n"}
import cgn
x = cgn.Parameter(start=np.ones(n), name="x")
# -
# Note that the starting guess can be adapted later via the ``x.start``-property.
# Next, we set up the regularization term by setting $\beta = 1$. The regularization operator defaults to the identity, and the regularizing guess default to the zero vector.
# + pycharm={"name": "#%%\n"}
x.beta = 0.001
# + [markdown] pycharm={"name": "#%% md\n"}
# Next, we can initialize the ``cgn.Problem`` object:
# + pycharm={"name": "#%%\n"}
problem = cgn.Problem(parameters=[x], fun=F, jac=DF)
# + [markdown] pycharm={"name": "#%% md\n"}
# We initialize the solver...
# + pycharm={"name": "#%%\n"}
solver = cgn.CGN()
# -
# and solve the problem:
# + pycharm={"name": "#%%\n"}
x_start = np.ones(n)
solver.options.set_verbosity(2)
solution = solver.solve(problem=problem)
# + [markdown] pycharm={"name": "#%% md\n"}
# Let us view the solution:
# + pycharm={"name": "#%%\n"}
x_min = solution.minimizer("x")
x_min
# + [markdown] pycharm={"name": "#%% md\n"}
# Let us compare this solution to the one obtained with the implementation of ridge regression from ``scikit-learn``:
# + pycharm={"name": "#%%\n"}
from sklearn.linear_model import Ridge
clf = Ridge(alpha=x.beta, fit_intercept=False)
y = np.ones(m)
clf.fit(X=A, y=y)
x_ridge = clf.coef_
difference_to_ridge = np.linalg.norm(x_min - x_ridge)
print(f"Difference to ridge: {difference_to_ridge}")
# + [markdown] pycharm={"name": "#%% md\n"}
# The two solutions agree up to a precision of $10^{-13}$!
#
# Note that the ``solution`` object also provides us access to the minimum of the cost function. Here, it is important to keep in mind that the cost function always has a factor $\frac{1}{2}$ in front.
# + pycharm={"name": "#%%\n"}
cost_at_minimum = solution.cost
print(f"Cost at minimum: {cost_at_minimum}")
# + [markdown] pycharm={"name": "#%% md\n"}
# Let us verify this by manually recomputing the cost at ``x_min``:
# + pycharm={"name": "#%%\n"}
cost_recomputed = 0.5 * np.sum(np.square(F(x_min))) + 0.5 * x.beta * np.sum(np.square(x_min))
print(f"Cost at minimum, recomputed: {cost_at_minimum}")
# + [markdown] pycharm={"name": "#%% md\n"}
# And indeed, both numbers agree.
#
# Finally, the ``solution`` also provides access to the [precision matrix](https://en.wikipedia.org/wiki/Precision_(statistics)) via ``solution.precision``.
#
# + pycharm={"name": "#%%\n"}
precision = solution.precision
precision.shape
# + [markdown] pycharm={"name": "#%% md\n"}
# This is relevant in the case where our optimization problem comes from [maximum likelihood estimation](https://en.wikipedia.org/wiki/Maximum_likelihood_estimation) or
# [Bayesian maximum-a-posteriori estimation](https://en.wikipedia.org/wiki/Maximum_a_posteriori_estimation), and the cost function actually corresponds to a negative log-likelihood or a negative log-posterior density, respectively.
|
examples/example1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pwd
import pickle as pk
import pandas as pd
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.metrics import accuracy_score
# +
##using for generating report later
def classification_report_csv(report, csv_name):
report_data = []
lines = report.split('\n')
for line in lines[2:-3]:
row = {}
row_data = line.split(' ')
row_data = list(filter(None, row_data))
row['class'] = row_data[0]
row['precision'] = float(row_data[1])
row['recall'] = float(row_data[2])
row['f1_score'] = float(row_data[3])
row['support'] = float(row_data[4])
report_data.append(row)
dataframe = pd.DataFrame.from_dict(report_data)
dataframe.to_csv(csv_name, index = False)
# -
# ## get test data from csv file
dfCN = pd.read_csv('./data/df_test_chinese_with_negation.csv',encoding ="ISO-8859-1")
dfFR = pd.read_csv('./data/df_test_french_with_negation.csv',encoding = 'ISO-8859-1')
dfIN = pd.read_csv('./data/df_test_indian_with_negation.csv',encoding = 'ISO-8859-1')
dfJP = pd.read_csv('./data/df_test_japanese_with_negation.csv',encoding = 'ISO-8859-1')
dfCN.head()
dfCN.drop(['Unnamed: 0', 'rating'], inplace=True, axis=1)
dfFR.drop(['Unnamed: 0', 'rating'], inplace=True, axis=1)
dfIN.drop(['Unnamed: 0','rating'], inplace=True, axis=1)
dfJP.drop(['Unnamed: 0','rating'], inplace=True, axis=1)
#dfJP.drop(['category', 'id', 'name', 'rating'], inplace=True, axis=1)
dfJP.head()
testCN = dfCN['review']
testFR = dfFR['review']
testIN = dfIN['review']
testJP = dfJP['review']
testCN_lab = dfCN['sentiment']
testFR_lab = dfFR['sentiment']
testIN_lab = dfIN['sentiment']
testJP_lab = dfJP['sentiment']
print("Chinese Resturant :",len(testCN_lab))
print("French Resturant :",len(testFR_lab))
print("Indian Resturant :",len(testIN_lab))
print("Japanese Resturant :",len(testJP_lab))
# ## testing result using model only trained by scraped data
# ### firstly ,need to change the test data to the same dimention as the training data
with open('./models/preprocessed_negated/vectorise_negation.pk', 'rb') as vectorizer:
vectorizer = pk.load(vectorizer)
test_vectorsCN = vectorizer.transform(testCN)
test_vectorsFR = vectorizer.transform(testFR)
test_vectorsIN = vectorizer.transform(testIN)
test_vectorsJP = vectorizer.transform(testJP)
print(test_vectorsCN.shape)
print(test_vectorsFR.shape)
print(test_vectorsIN.shape)
print(test_vectorsJP.shape)
# ### secondly, check the 4 kinds of resturants test score using Naive Bayes and SVM classifier
# load naivebayes
with open('./models/preprocessed_negated/classifier_negation_naivebayes.pk', 'rb') as naivebayes:
model1 = pk.load(naivebayes)
predCN_na = model1.predict(test_vectorsCN)
predFR_na = model1.predict(test_vectorsFR)
predIN_na = model1.predict(test_vectorsIN)
predJP_na = model1.predict(test_vectorsJP)
predCN_na = list(predCN_na)
predFR_na = list(predFR_na)
predIN_na = list(predIN_na)
predJP_na = list(predJP_na)
cmCN_na = pd.crosstab(pd.Series(testCN_lab), pd.Series(predCN_na), rownames= ['actuals'], colnames=['pred'],margins=True)
cmFR_na = pd.crosstab(pd.Series(testFR_lab), pd.Series(predFR_na), rownames= ['actuals'], colnames=['pred'],margins=True)
cmIN_na = pd.crosstab(pd.Series(testIN_lab), pd.Series(predIN_na), rownames= ['actuals'], colnames=['pred'],margins=True)
cmJP_na = pd.crosstab(pd.Series(testJP_lab), pd.Series(predJP_na), rownames= ['actuals'], colnames=['pred'],margins=True)
#chinese resturant
cmCN_na
print (classification_report(predCN_na, testCN_lab))
print (accuracy_score(predCN_na, testCN_lab))
pd.DataFrame(cmCN_na).to_csv('Chinese_naive_bayes.csv')
report = classification_report(predCN_na, testCN_lab)
classification_report_csv(report,'Chinese_naive_bayes_report.csv')
Chinese_nb_result = pd.Series(predCN_na)
#French resturant
cmFR_na
print (classification_report(predFR_na, testFR_lab))
print (accuracy_score(predFR_na, testFR_lab))
pd.DataFrame(cmFR_na).to_csv('French_naive_bayes.csv')
report = classification_report(predFR_na, testFR_lab)
classification_report_csv(report,'French_naive_bayes_report.csv')
French_nb_result = pd.Series(predFR_na)
#Indian resturant
cmIN_na
print (classification_report(predIN_na, testIN_lab))
print (accuracy_score(predIN_na, testIN_lab))
pd.DataFrame(cmIN_na).to_csv('Indian_naive_bayes.csv')
report = classification_report(predIN_na, testIN_lab)
classification_report_csv(report,'Indian_naive_bayes_report.csv')
Indian_nb_result = pd.Series(predIN_na)
#Japanese resturant
cmJP_na
print (classification_report(predJP_na, testJP_lab))
print (accuracy_score(predJP_na, testJP_lab))
pd.DataFrame(cmJP_na).to_csv('Janpanese_naive_bayes.csv')
report = classification_report(predJP_na, testJP_lab)
classification_report_csv(report,'Japanese_naive_bayes_report.csv')
Japanese_nb_result = pd.Series(predJP_na)
# load SVM
with open('./models/preprocessed_negated/classifier_negation_svm.pk', 'rb') as svm:
model2 = pk.load(svm)
predCN_svm = model2.predict(test_vectorsCN)
predFR_svm = model2.predict(test_vectorsFR)
predIN_svm = model2.predict(test_vectorsIN)
predJP_svm = model2.predict(test_vectorsJP)
predCN_svm = list(predCN_svm)
predFR_svm = list(predFR_svm)
predIN_svm = list(predIN_svm)
predJP_svm = list(predJP_svm)
cmCN_svm = pd.crosstab(pd.Series(testCN_lab), pd.Series(predCN_svm), rownames= ['actuals'], colnames=['pred'],margins=True)
cmFR_svm = pd.crosstab(pd.Series(testFR_lab), pd.Series(predFR_svm), rownames= ['actuals'], colnames=['pred'],margins=True)
cmIN_svm = pd.crosstab(pd.Series(testIN_lab), pd.Series(predIN_svm), rownames= ['actuals'], colnames=['pred'],margins=True)
cmJP_svm = pd.crosstab(pd.Series(testJP_lab), pd.Series(predJP_svm), rownames= ['actuals'], colnames=['pred'],margins=True)
#Chinese resturant
cmCN_svm
print (classification_report(predCN_svm, testCN_lab))
print (accuracy_score(predCN_svm, testCN_lab))
pd.DataFrame(cmCN_svm).to_csv('Chinese_svm.csv')
report = classification_report(predCN_svm, testCN_lab)
classification_report_csv(report,'Chinese_svm_report.csv')
Chinese_svm_result = pd.Series(predCN_svm)
#French resturant
cmFR_svm
print (classification_report(predFR_svm, testFR_lab))
print (accuracy_score(predFR_svm, testFR_lab))
pd.DataFrame(cmFR_svm).to_csv('French_svm.csv')
report = classification_report(predFR_svm, testFR_lab)
classification_report_csv(report,'French_svm_report.csv')
French_svm_result = pd.Series(predFR_svm)
#India resturant
cmIN_svm
print (classification_report(predIN_svm, testIN_lab))
print (accuracy_score(predIN_svm, testIN_lab))
pd.DataFrame(cmIN_svm).to_csv('Indian_svm.csv')
report = classification_report(predIN_svm, testIN_lab)
classification_report_csv(report,'Indian_svm_report.csv')
Indian_svm_result = pd.Series(predIN_svm)
#Japanese resturant
cmJP_svm
print (classification_report(predJP_svm, testJP_lab))
print (accuracy_score(predJP_svm, testJP_lab))
pd.DataFrame(cmJP_svm).to_csv('Japanese_svm.csv')
report = classification_report(predJP_svm, testJP_lab)
classification_report_csv(report,'Japanese_svm_report.csv')
Japanese_svm_result = pd.Series(predJP_svm)
# ### load all the result to csv file
df_result = pd.concat([pd.Series(testCN_lab), Chinese_nb_result, Chinese_svm_result], keys=["actual", "nb", "svm"], axis=1)
df_result.to_csv('Chinese_result.csv')
df_result = pd.concat([pd.Series(testFR_lab), French_nb_result, French_svm_result], keys=["actual", "nb", "svm"], axis=1)
df_result.to_csv('French_result.csv')
df_result = pd.concat([pd.Series(testIN_lab), Indian_nb_result, Indian_svm_result], keys=["actual", "nb", "svm"], axis=1)
df_result.to_csv('Indian_result.csv')
df_result = pd.concat([pd.Series(testJP_lab), Japanese_nb_result, Japanese_svm_result], keys=["actual", "nb", "svm"], axis=1)
df_result.to_csv('Japanese_result.csv')
## concact with the original test csv file and further translate into JSON file
cn = pd.read_csv('./data/df_test_chinese.csv')
cn_score = pd.read_csv('./test_result/chinese_result 3.csv')
cn.head()
cn_score.head()
cn['svm_score']=cn_score['svm']
cn['naive_bayes_score']=cn_score['nb']
cn.to_csv('Chinese_test_final.csv')
cn.head()
# ## Conclusion
# We test the 4 categories of resturants using 3 types of models: model trained with only scraped data; scraped data combined with the standard data; negation combined data.
#
# We measure the overall accuracy and the model trained with negation data performs best among the 3 types (both on SVN and Naive Bayes).Then we save the best one to the csv and generate the JSON format.
# 
# The conclusion of the test result are the followings:
# 1) from the overall accuracy ,we can tell that using the preprocessed data ,the model is robust and thus we can yield much better result on the test data.
# 2) as for the Chinese resturant , because there are some Chinese words can not be indentified ,so the overall accuracy not performs well compared with other 3 types of resturants.
# 3) for the sentiment score , negation plays very improtant role and yile the best result by implementing negation into the preprocess data.
|
TestResults_ 4_Categories_of_Restaurants.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Tag 1. Kapitel 1. Grundlagen
#
# ## Lab 1.
#
# ### Lösungen
#
# In diesem Lab werden folgende Themen geübt:
#
# * Grundlegende Datentypen und Arithmetik
# * Vektoren
# * Vektor Operationen
# * Vergleichsoperatoren
# * Vektor Indexierung und Auswahl
#
# ### Folgenden Aufgaben sind mit R zu lösen
#
# *Tipp: allle Aufgaben können auch in R Studio gelöst werden
#
# ** Was ist zwei hoch zweiunddreißig? **
2 ^ 32
# ** Erstelle einen Vektor namens ** temp.woche1 ** mit folgenden Messwerten: ** 37.8, 38.2, 35.9, 36.2, 36.6
temp.woche1 <- c(37.8, 38.2, 35.9, 36.2, 36.6)
temp.woche1
# ** Weise den Einträgen abgekürzte Namen der Wochentagen von Montag bis Freitag zu (Mo, Di, Mi, usw.) **
names(temp.woche1) <- c('Mo','Di','Mi','Do','Fr')
temp.woche1
# ** Was ist der durchschnittliche Körper-Temperatur beim Patienten (en. mean) für dieser Woche gewesen? **
# (Man kann hier eine eingebaute Funktion verwenden. Siehe Kapitel 8 für Details.)
summary(temp.woche1)
mean(temp.woche1)
# ** Erstelle einen Filter-Vektor ** temp.hoch **, der zeigt, ob beim Patienten gemessene Körper-Temperatur über ** 36.6 ** liegt **
temp.hoch <- temp.woche1 > 36.6
temp.hoch
# ** Wende Filter-Vektor ** temp.hoch ** an, um Fieber-Tage zu auszuwählen **
temp.woche1[temp.hoch]
# ** Erstelle (mit einer eingebauten Funktion) einen Filter-Vektor für maximale Patienten-Temperatur. **
temp.max <- temp.woche1 == max(temp.woche1)
temp.max
# ** Max-Filter für Temperatur-Wochenmessungen anwenden, um herauszuwinden an welchem(n) Tag(en) Patienten-Temperatur am höchsten war **
temp.woche1[temp.max]
# Ende von Lab 1.
|
1.1 R Basics/de-DE/.ipynb_checkpoints/1.1.10 R - Basics. Lab 1. - Solutions-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="k5ZxDky2OZI4"
#default_exp query
# + id="tqmQkGtgOZJA"
#export
# mark rows that contain certain codes in one or more colums
def get_rows(df, codes, cols=None, sep=None, pid='pid', expand=False, codebook=None):
"""
Make a boolean series that is true for all rows that contain the codes
Args
df (dataframe or series): The dataframe with codes
codes (str, list, set, dict): codes to be counted
cols (str or list): list of columns to search in
sep (str): The symbol that separates the codes if there are multiple codes in a cell
pid (str): The name of the column with the personal identifier
"""
# string as input for single codes is allowed
# but then must make it a list
if isinstance(codes, str):
codes = [codes]
# same for cols
# must be a list sine we may loop over it
if not isinstance(cols, list):
cols = [cols]
if any(notation in ''.join(cols) for notation in '*-:']):
cols=expand_cols(cols)
# special case: star notation that does not require expansion
if expand:
# start with special case. for speed: star notation that does not require expansion
# (does not require making a list of unique values)
if any(code.endswith('*') for code in codes):
star_codes=[code for code in codes if code.endswith('*')]
codes=[code for codes if code not in star_codes]
#if codes have both star and hyphen notation
star_codes=expand_hyphen(star_codes)
#get the rows!
endstar_rows=_get_rows_endstar(df=df, codes=codes, cols=cols, codebook=codebook)
# return rows right away if there are no other codes to be checked
if len(codes)==0:
return endstar_rows
# continue with all other codes
# check if any codes need expansion
if any(notation in ''.join(codes) for notation in '*-:']):
codes=expand_codes(codes)
# approach depends on whether we have multi-value cells or not
# if sep exist, then have multi-value cells
if sep:
# have multi-valued cells
codes = [rf'\b{code}\b' for code in codes]
codes_regex = '|'.join(codes)
# starting point: no codes have been found
# needed since otherwise the function might return None if no codes exist
rows = pd.Series(False*len(df),index=df.index)
# loop over all columns and mark when a code exist
for col in cols:
rows=rows | df[col].str.contains(codes_regex, na=False)
# if not multi valued cells
else:
mask = df[cols].isin(codes)
rows = mask.any(axis=1)
if 'endstar_rows' in locals():
rows=rows | endstar_rows
return rows
# + id="CN05UMXsRi6t"
# exporti
def _get_rows_endstar(df, codes, cols):
"""
Returns rows with codes that starts with a given value(s) in one or
more columns, with one or more values in each column
Note: special function for the special (but common) case when the user
wants to pick codes that start with a given string
"""
for col in cols:
if sep:
for code in codes:
code=code.strip('*')
codes_regex=codes_regex + rf'|\b{code}\w+'
#double check, potential problem, hyphen a word boundary in regex. Use b vs w+?
rows = rows | df[col].str.contains(rf'\b{code}\w+', na=False) # single col, multiple cell values
else:
rows = rows | df[col].str.startswith(code) # single col, single cell value
return rows
|
query.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yC6iWdubIxT2"
# # Payment Matching
# At Shopee, bank transfer is a payment method in most countries. When a buyer chooses to place an order using bank transfer, he/she is supposed to make the transfer within 2 days after he/she places the order.
#
# After he/she makes the transfer, Shopee will receive a bank statement from the bank and Shopee needs to compare and match the bank statement with the checkout information in order to confirm that this particular order has been paid. This process is called payment matching.
#
# Two criteria need to be met in order to match a bank statement with a checkout:
# - Amount match: Statement amount equals checkout amount.
# - Name match: Statement description “matches” checkout buyer name (Note: statement description usually contains buyer name)
#
# A proper match occurs when both the amount and the name matches on both bank statement and checkout list.
# + [markdown] id="7r7SKaQISt8_"
# # Preparation of the Dataset
# + id="13otSpPHTQMP" colab={"base_uri": "https://localhost:8080/"} outputId="4678cf77-266e-432a-a062-5fd7d09a7227"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# !pip install fuzzywuzzy
import warnings
warnings.filterwarnings('ignore')
import re
from fuzzywuzzy import fuzz
# + id="XZLlx0L-TzIv"
bank = pd.read_csv('/content/drive/MyDrive/Datasets/NDSC/bank_statement.csv')
check = pd.read_csv('/content/drive/MyDrive/Datasets/NDSC/checkout.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="mP_AXWwRIWgA" outputId="abe4631f-deff-4d36-8640-4c335e3693e0"
bank.head()
# + id="NwtZP0QcyaGY" colab={"base_uri": "https://localhost:8080/"} outputId="af6b3f3d-db5e-40ee-e397-2be90ae198dc"
print(f"This dataset contains {bank.shape[0]} rows and {bank.shape[1]} columns.")
# + id="XosfIvr9qXiG" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="842e7dc6-1aa6-413c-e498-37ab9af79e5e"
check.head()
# + id="Ujz65JIsIeGd" colab={"base_uri": "https://localhost:8080/"} outputId="4c1c58f6-a6ac-4669-d70c-5f434f86ac33"
print(f"This dataset contains {check.shape[0]} rows and {check.shape[1]} columns.")
# + [markdown] id="K-yOg4zfkKNk"
# # Removing Unwanted Characters
# Removing unwanted characters and 'transfer' word on bank_statement.
# + id="UIhU5GbJkI7D"
bank['desc'] = bank['desc'].apply(lambda x: re.sub(r"[^a-zA-Z0-9]+", ' ', x))
check['buyer_name'] = check['buyer_name'].apply(lambda x: re.sub(r"[^a-zA-Z0-9]+", ' ', x))
bank['desc'] = bank['desc'].apply(lambda x: re.sub(r"TRANSFER", ' ', x))
check['buyer_name'] = check['buyer_name'].apply(lambda x: re.sub(r"TRANSFER", ' ', x))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="FalKBuRNkYMS" outputId="93b43d4d-cf71-4c74-e64b-cca5f2ecbb27"
bank.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="7RqCiLhRlChZ" outputId="07c9ab73-cd15-4209-bb78-fda829f11a1e"
check.head()
# + id="HiIjQRoGlDN_"
# Sorting Values for efficiency and convert the strings to sets for better data structure and better time complexity
bank = bank.sort_values(by=['stmt_amount'])
check = check.sort_values(by=['ckt_amount'])
# Converting the 'description' and 'buyer name' to SET Data Type so there are not duplicate words.
bank['desc'] = bank['desc'].apply(lambda x: set(x.split()))
check['buyer_name'] = check['buyer_name'].apply(lambda x: set(x.split()))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="olOUZ7DGlykX" outputId="a87a4356-e01e-4ecb-8633-7fc7e4573115"
bank.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="wTwBPMUWlzfJ" outputId="e63c44d7-5d76-4119-c73c-0cfc04ad405a"
check.head()
# + [markdown] id="WnLmeJ81olqo"
# # 1) Match the Price and Description
# 1. Go through the unique prices one by one
# 2. For each price, query all the bank statements and transactions
# 3. For each transaction, find a bank statement that matches best.
# 4. If the transaction cannot find a fitting bank statement, it will be stored to be processed later using fuzzy search.
# + id="3HQUZVYvl0Gp"
# Sorted unique price
prices = sorted(list(set(check['ckt_amount'])))
# + id="aBbxDSU8otwF"
# Match the price & description
bs = bank.values.tolist()
co = check.values.tolist()
curr_bank, curr_trans, pend_bank, pend_trans = [], [], [], []
answer = []
no_ans_trans, no_ans_bank = [], []
for price in prices:
while bs and bs[0][1] == price:
curr_bank.append(bs.pop(0))
while co and co[0][1] == price:
curr_trans.append(co.pop(0))
for trans in curr_trans:
found = False
curname = trans[2]
bk = list(filter(lambda x: x[2].intersection(curname), curr_bank))
if bk:
b = max(bk, key= lambda x: len(x[2].intersection(curname)))
answer.append((trans, b))
curr_bank.remove(b)
else:
no_ans_trans.append(trans)
no_ans_bank.extend(curr_bank)
curr_bank, curr_trans = [], []
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="zw04LxN9wR-V" outputId="5de66f7d-64de-455e-e2d9-724b7d29a9f8"
# Preview the Transactions
pd.DataFrame(answer)
# + colab={"base_uri": "https://localhost:8080/"} id="i3Zri9c7poXE" outputId="78bf3a4b-1b37-4504-d0e7-5fa2f1aa5b0e"
# Number of transaction that does not match
print(len(no_ans_bank), len(no_ans_trans))
# + colab={"base_uri": "https://localhost:8080/"} id="SPa2FadfrjW2" outputId="eabe2970-fee6-438a-91e2-4704e051f5ad"
no_ans_bank[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="huPzUVbYtcHv" outputId="a6bd9a1d-f95e-4fc7-b040-0989949cc987"
no_ans_trans[:5]
# + [markdown] id="rd4ktUBWtqar"
# # 2) Transactions Which Not Matched (yet)
# For transaction that does not match, using FuzzyWuzzy.
# + id="CWYQ2sbDte0C"
# pick the 'id' only
no_id_bank, no_id_trans = [], []
for x in range(len(no_ans_bank)):
no_id_bank.append(no_ans_bank[x][0])
no_id_trans.append(no_ans_trans[x][0])
# + id="N_hKawfBt49D"
# get the subset of the DataFrame that previously has no match
check_new = check[check['ckt_id'].isin(no_id_trans)]
bank_new = bank[bank['stmt_id'].isin(no_id_bank)]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="_h_Bz9LmuUZJ" outputId="4966caf9-8afd-4f5b-8617-15ec5ff113a7"
# bank DataFrame which has no match transaction
bank_new.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="_THYhZB4ucHs" outputId="af11a26e-1382-4b4c-feaf-80e30d09a7ed"
# checkout DataFrame which has no match transaction
check_new.head()
# + id="47KqLg5Uusiq"
# Convert to string
bank_new['desc'] = bank_new['desc'].apply(lambda x: ' '.join(sorted(list(x))))
check_new['buyer_name'] = check_new['buyer_name'].apply(lambda x: ' '.join(sorted(list(x))))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="8795Muc_vFlX" outputId="6c7de076-9a53-4ad2-bc73-31b2942d42f5"
check_new.head()
# + id="100nzsIEvHjN"
# unique prices for unmatch transaction
prices_new = sorted(list(set(check_new['ckt_amount'])))
# + id="V3K9G2D3vUUc"
bs2, co2 = bank_new.values.tolist(), check_new.values.tolist()
curr_bank2, curr_trans2 = [], []
answer2 = []
for price in prices_new:
while bs2 and bs2[0][1] == price:
curr_bank2.append(bs2.pop(0))
while co2 and co2[0][1] == price:
curr_trans2.append(co2.pop(0))
for trans in curr_trans2:
curname = trans[2]
b = max(curr_bank2, key= lambda x: fuzz.partial_ratio(curname, x[2]))
answer2.append((trans, b))
curr_bank2.remove(b)
curr_bank2, curr_trans2 = [], []
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="iJ1fYGtKv7WT" outputId="93c2df55-04cb-4daf-f4e1-32049ccdb30e"
# Preview the Transactions
pd.DataFrame(answer2)
# + id="J29ne1aFvnRx"
# Construct final dataframe
fs1 = [(x[0][0], x[1][0]) for x in answer]
fs2 = [(x[0][0], x[1][0]) for x in answer2]
fs = pd.DataFrame(fs1+fs2)
fs.columns=['ckt_id', 'stmt_id']
fs=fs[['stmt_id', 'ckt_id']]
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="5qXmLgFzvx3H" outputId="bcfa9c2e-7f25-4f62-f17a-5f148b007552"
fs
# + [markdown] id="dbu3lkm6xKLt"
# # Output the CSV
# + colab={"base_uri": "https://localhost:8080/"} id="ChsJwVrgv5nf" outputId="0fc04935-beae-4624-cb0b-9045403f764a"
# Output CSV
print(fs.shape)
fs.to_csv('submission.csv', index=False)
# + id="ckvQfL6nxL4p"
|
payment_ndsc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inverse Distance Weighting model vs Kriging - tutorial
#
# ## Table of Contents:
#
# 1. Read point data,
# 2. Divide dataset into two sets: modeling and validation set,
# 3. Perform IDW and evaluate it,
# 4. Perform variogram modeling on the modeling set,
# 5. Validate Kriging and compare Kriging and IDW validation results,
# 6. Bonus scenario: only 5% of values are known!
#
# ## Level: Basic
#
# ## Changelog
#
# | Date | Change description | Author |
# |------|--------------------|--------|
# | 2021-05-12 | First version of tutorial | @szymon-datalions |
#
# ## Introduction
#
# In this tutorial we will learn about the one method of validation of our Kriging model. We'll compare it to the Inverse Distance Weighting function where the unknown point value is interpolated as the weighted mean of it's neighbours. Weights are assigned by the inverted distance raised to the n-th power.
#
# (1) **GENERAL FORM OF IDW**
#
# $$z(u) = \frac{\sum_{i}\lambda_{i}*z_{i}}{\sum_{i}\lambda_{i}}$$
#
# where:
#
# - $z(u)$: is the value at unknown location,
# - $i$: is a i-th known location,
# - $z_{i}$: is a value at known location $i$,
# - $\lambda_{i}$: is a weight assigned to the known location $i$.
#
# (2) **WEIGHTING PARAMETER**
#
# $$\lambda_{i} = \frac{1}{d^{p}_{i}}$$
#
# where:
#
# - $d$: is a distance from known point $z_{i}$ to the unknown point $z(u)$,
# - $p$: is a hyperparameter which controls how strong is a relationship between known point and unknown point. You may set large $p$ if you want to show strong relationship between closest point and very weak influence of distant points. On the other hand, you may set small $p$ to emphasize fact that points are influencing each other with the same power irrespectively of their distance.
#
# ---
#
# As you noticed **IDW** is a simple but powerful technique. Unfortunately it has major drawback: **we must set `p` - power - manually** and it isn't derived from the data and variogram. That's why it can be used for other tasks. Example is to use IDW as a baseline for comparison to the other techniques.
# ## Import packages
# +
import numpy as np
import matplotlib.pyplot as plt
from pyinterpolate.idw import inverse_distance_weighting # function for idw
from pyinterpolate.io_ops import read_point_data
from pyinterpolate.semivariance import calculate_semivariance # experimental semivariogram
from pyinterpolate.semivariance import TheoreticalSemivariogram # theoretical models
from pyinterpolate.kriging import Krige # kriging models
# -
# ## 1) Read point data
dem = read_point_data('../sample_data/point_data/poland_dem_gorzow_wielkopolski', data_type='txt')
dem
# ## 2) Divide dataset into two sets: modeling and validation set
#
# In this step we will divide our dataset into two sets:
#
# - modeling set (50%): points used for variogram modeling,
# - validation set (50%): points used for prediction and results validation.
#
# Baseline dataset will be divided randomly.
# +
# Create modeling and validation sets
def create_model_validation_sets(dataset: np.array, frac=0.5):
"""
Function divides base dataset into modeling and validation sets
INPUT:
:param dataset: (numpy array) array with rows of records,
:param frac: (float) number of elements in a validation set
OUTPUT:
return: modeling_set (numpy array), validation_set (numpy array)
"""
removed_idx = np.random.randint(0, len(dem)-1, size=int(frac * len(dem)))
validation_set = dem[removed_idx]
modeling_set = np.delete(dem, removed_idx, 0)
return modeling_set, validation_set
known_points, unknown_points = create_model_validation_sets(dem)
# -
# ## 3) Perform IDW and evaluate it
#
# Inverse Distance Weighting doesn't require variogram modeling or other steps. We pass power to which we want raise distance in weight denominator. Things to remember are:
#
# - Large `power` -> closer neighbours are more important,
# - `power` which is close to the **zero** -> all neighbours are important and we assume that distant process has the same effect on our variable as the closest events.
# +
IDW_POWER = 2
NUMBER_OF_NEIGHBOURS = -1 # Include all points in weighting process (equation 1)
idw_predictions = []
for pt in unknown_points:
idw_result = inverse_distance_weighting(known_points, pt[:-1], NUMBER_OF_NEIGHBOURS, IDW_POWER)
idw_predictions.append(idw_result)
# +
# Evaluation
idw_rmse = np.mean(np.sqrt((unknown_points[:, -1] - np.array(idw_predictions))**2))
print(f'Root Mean Squared Error of prediction with IDW is {idw_rmse}')
# -
# **Clarification:** Obtained Root Mean Squared Error could serve as a baseline for further model development. To build better reference, we create four IDW models of powers:
#
# 1. 0.5,
# 2. 1,
# 3. 2,
# 4. 4.
# +
IDW_POWERS = [0.5, 1, 2, 4]
idw_rmse = {}
for pw in IDW_POWERS:
results = []
for pt in unknown_points:
idw_result = inverse_distance_weighting(known_points, pt[:-1], NUMBER_OF_NEIGHBOURS, pw)
results.append(idw_result)
idw_rmse[pw] = np.mean(np.sqrt((unknown_points[:, -1] - np.array(results))**2))
# -
for pw in IDW_POWERS:
print(f'Root Mean Squared Error of prediction with IDW of power {pw} is {idw_rmse[pw]:.4f}')
# ## 4) Perform variogram modeling on the modeling set
#
# In this step we will go through semivariogram modeling for Kriging interpolation.
# +
search_radius = 0.01
max_range = 0.32
number_of_ranges = 32
exp_semivar = calculate_semivariance(data=known_points, step_size=search_radius, max_range=max_range)
# -
semivar = TheoreticalSemivariogram(points_array=known_points, empirical_semivariance=exp_semivar)
semivar.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
semivar.show_semivariogram()
# ## 5) Validate Kriging and compare Kriging and IDW validation results
#
# In the last we perform Kriging interpolation and compare rersults to the **IDW** models. We use all points to weight values at unknown locations and semivariogram model chosen in the previous step.
# +
# Set Kriging model
model = Krige(semivariogram_model=semivar, known_points=known_points)
# +
kriging_preds = []
for pt in unknown_points:
result = model.ordinary_kriging(pt[:-1], number_of_neighbours=64, test_anomalies=False)
kriging_preds.append(result[0])
# +
# Evaluation
kriging_rmse = np.mean(np.sqrt((unknown_points[:, -1] - np.array(kriging_preds))**2))
print(f'Root Mean Squared Error of prediction with Kriging is {kriging_rmse}')
# -
# **Clarification**: Kriging is better than any from the IDW models and we may assume that our modeling approach gives us more insights into the spatial process which we are observing. But this is not the end... let's consider more complex scenario!
# ## 6) Bonus scenario: only 5% of values are known!
#
# Real world data are rarely as good as sample from the tutorial. It is too expensive to densily sample every location and usually we get only small percent of area covered by data. That's why it is good to compare **IDW vs Kriging** in this scenario! We repeat steps 1-5 with change in division for modeling / validation set. (I encourage you to try it do alone and then compare your code and results with those given in this notebook).
# +
# Data preparation
known_points, unknown_points = create_model_validation_sets(dem, 0.95)
# +
# IDW tests
IDW_POWERS = [0.5, 1, 2, 4]
idw_rmse = {}
for pw in IDW_POWERS:
results = []
for pt in unknown_points:
idw_result = inverse_distance_weighting(known_points, pt[:-1], NUMBER_OF_NEIGHBOURS, pw)
results.append(idw_result)
idw_rmse[pw] = np.mean(np.sqrt((unknown_points[:, -1] - np.array(results))**2))
# +
# Variogram
search_radius = 0.01
max_range = 0.32
number_of_ranges = 32
exp_semivar = calculate_semivariance(data=known_points, step_size=search_radius, max_range=max_range)
# -
semivar = TheoreticalSemivariogram(points_array=known_points, empirical_semivariance=exp_semivar)
semivar.find_optimal_model(weighted=False, number_of_ranges=number_of_ranges)
semivar.show_semivariogram()
# +
# Set Kriging model
model = Krige(semivariogram_model=semivar, known_points=known_points)
# +
kriging_preds = []
for pt in unknown_points:
result = model.ordinary_kriging(pt[:-1], number_of_neighbours=64, test_anomalies=False)
kriging_preds.append(result[0])
kriging_rmse = np.mean(np.sqrt((unknown_points[:, -1] - np.array(kriging_preds))**2))
# +
# Comparison
for pw in IDW_POWERS:
print(f'Root Mean Squared Error of prediction with IDW of power {pw} is {idw_rmse[pw]:.4f}')
print(f'Root Mean Squared Error of prediction with Kriging is {kriging_rmse:.4f}')
# -
# Your results may be different but in the most cases Kriging will be better than IDW. What's even more important is the fact that for the single data source with low number of samples we don't have opportunity to perform validation step and we're not able to guess how big should be the power parameter. With Kriging we model variogram and _voila!_ - model works.
# ---
|
docs/build/html/tutorials/Compare Kriging results to the IDW model (Basic).ipynb
|
# ---
# title: "Random Forest Classifier"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "Training a random forest classifier in scikit-learn."
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ## Preliminaries
# Load libraries
from sklearn.ensemble import RandomForestClassifier
from sklearn import datasets
# ## Load Iris Data
# Load data
iris = datasets.load_iris()
X = iris.data
y = iris.target
# ## Create Random Forest Classifier
# Create random forest classifer object that uses entropy
clf = RandomForestClassifier(criterion='entropy', random_state=0, n_jobs=-1)
# ## Train Random Forest Classifier
# Train model
model = clf.fit(X, y)
# ## Predict Previously Unseen Observation
# +
# Make new observation
observation = [[ 5, 4, 3, 2]]
# Predict observation's class
model.predict(observation)
|
docs/machine_learning/trees_and_forests/random_forest_classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PublicDatasets
# +
import sys
sys.path.append('source/')
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib notebook
# %reload_ext autoreload
# %autoreload 2
from public_data_utils import *
plt.rcParams['figure.figsize'] = 7, 3
matplotlib.rcParams['pdf.fonttype'] = 42
matplotlib.rcParams['ps.fonttype'] = 42
# set the random seed so can reproduce when something didn't work.
# (only when cells are run in order)
np.random.seed(1)
# -
# This notebook contains functionality to evaluate the proposed algorithms on real datasets. To simply reproduce the plots from the paper you can directly run GenerateAllFigures.
#
# # Preparation
#
# ### 1. Download datasets (.mat files)
#
# You can simply run ./bin/download_datasets.sh.
# See datasets/README.md for file descriptions. This notebook was tested with the WiFi and Lawmower datasets.
# ### 2. Choose dataset and range
#
# Note that currently fully functional are only Plaza1 and Plaza2. The others are still kept here for development purposes.
# +
#filename = 'datasets/uah1.mat' # fingers
filename = 'datasets/Plaza1.mat'; # zig zag.
#filename = 'datasets/Plaza2.mat' #
#filename = 'datasets/Gesling1.mat' # not working
#filename = 'datasets/Gesling2.mat' # not working
#filename = 'datasets/Gesling3.mat' #
original_df, anchors_df, traj = read_dataset(filename, verbose=True)
xlim, ylim = get_plotting_params(filename)
print(xlim, ylim)
# +
# %matplotlib inline
from evaluate_dataset import calibrate
calibrate(original_df)
fig, ax = plt.subplots()
for anchor_id in original_df.anchor_id.unique():
if anchor_id == 'GT':
continue
gt_df = original_df[original_df.anchor_id==anchor_id]
fig, axs = plt.subplots(1, 2)
fig.set_size_inches(5, 3)
axs[0].scatter(gt_df.px, gt_df.py, s=1.0)
axs[1].scatter(gt_df.distance_gt, gt_df.distance, s=1.0, label='raw')
axs[1].scatter(gt_df.distance_gt, gt_df.distance_calib, s=1.0, label='calibrated')
axs[1].scatter(gt_df.distance_gt, gt_df.distance_gt, s=1.0, label='ideal')
fig.suptitle(f"anchor id {anchor_id}")
ax.plot(gt_df.timestamp, gt_df.distance, label=anchor_id)
axs[1].legend()
axs[0].set_xlabel('x [m]')
axs[0].set_ylabel('y [m]')
axs[1].set_xlabel('real d [m]')
axs[1].set_ylabel('measured d [m]')
ax.set_xlabel('timestamp')
ax.set_ylabel('distance')
ax.legend()
# -
# ### 3. Prepare dataset
# +
# %matplotlib inline
# #%matplotlib notebook
from public_data_utils import TIME_RANGES
from math import ceil, floor
if 'Plaza1' in filename:
plot_df = original_df[(original_df.timestamp > 300) & (original_df.timestamp < 1400)]
time_ranges = TIME_RANGES
print(original_df.timestamp.min(), original_df.timestamp.max())
print(time_ranges)
fig, ax = plt.subplots()
sns.scatterplot(data=plot_df, x='timestamp', y='px', color='red', linewidth=0.0, ax=ax, s=1.0)
sns.scatterplot(data=plot_df, x='timestamp', y='py', color='green', linewidth=0.0, ax=ax, s=1.0)
side = np.sqrt(len((time_ranges)))
fig, axs2 = plt.subplots(ceil(side), floor(side), sharex=False, sharey=True)
fig.suptitle('piecewise linear coordinates over time', y=0.95)
fig.set_size_inches(1.0*np.array(axs2.shape))
axs2 = axs2.reshape((-1,))
for ax2, time_range in zip(axs2, time_ranges):
plot_df = original_df[(original_df.timestamp > time_range[0]) & (original_df.timestamp < time_range[1])]
ax2.scatter(plot_df.timestamp, plot_df.px, color='red', s=1.0, label='x')
ax2.scatter(plot_df.timestamp, plot_df.py, color='green', s=1.0, label='y')
ax.scatter(plot_df.timestamp, plot_df.px, color='black', s=1.0)
ax.scatter(plot_df.timestamp, plot_df.py, color='black', s=1.0)
ax2.legend(loc='lower left', bbox_to_anchor=[1.0, 0.0])
mask = np.array([False] * len(original_df))
for time_range in time_ranges:
mask = mask | ((original_df.timestamp > time_range[0]) & (original_df.timestamp < time_range[1])).values
full_df = original_df[mask]
else:
full_df = original_df
print('using all measurements for', filename)
# -
# ### 4. (optional) plot distance measurements
# %matplotlib inline
plot_distance_times(full_df)
# ### 5. (optional) plot distance distributions
# +
fig, axs = plt.subplots(5, 1, sharex=False)
fig.set_size_inches(5, 10)
ax = plot_distance_errors(full_df, ax=axs[0])
#savefig(fig, 'results/accuracy.pdf')
axs[1].set_ylabel('(d - d_gt)')
distance_error = full_df.distance - full_df.distance_gt
axs[1].hist(distance_error, bins=30)
axs[2].set_ylabel('1/d(d**2 - d_gt**2)')
distance_error = (full_df.distance.values.astype(np.float32)**2 - full_df.distance_gt.values.astype(np.float32)**2)/(full_df.distance_gt.values.astype(np.float32) + 1e-3)
axs[2].hist(distance_error, bins=30)
axs[3].set_ylabel('(d**2 - d_gt**2)')
distance_error = full_df.distance.values.astype(np.float32)**2 - full_df.distance_gt.values.astype(np.float32)**2
axs[3].hist(distance_error, bins=30)
axs[4].set_ylabel('(d - d_gt)**2')
distance_error = (full_df.distance.values.astype(np.float32) - full_df.distance_gt.values.astype(np.float32))**2
_ = axs[4].hist(distance_error, bins=30)
# -
# ### 6. (optional) plot distance error spacially
# +
range_df = full_df.loc[full_df.system_id=='Range']
anchor_names = sorted(range_df.anchor_name.unique())
print(anchor_names)
fig, axs = plt.subplots(1, len(anchor_names), sharey=True)
fig.set_size_inches(15, 4)
for ax, anchor_name in zip(axs, anchor_names):
plot_df = range_df.loc[range_df.anchor_name==anchor_name].copy()
plot_df.loc[:, 'distance error'] = plot_df.distance.values - plot_df.distance_gt.values
plot_df.loc[:, 'anchor name'] = plot_df.anchor_name.values
anchors_df.loc[:, 'anchor name'] = anchors_df.anchor_name.values
sns.scatterplot(data=plot_df, x='px', y='py', hue='anchor name', size='distance error',
hue_order=anchor_names, linewidth=0.0, alpha=0.8, ax=ax, legend=False)
anchors_df = anchors_df.apply(pd.to_numeric, downcast='float', errors='ignore', axis=0)
sns.scatterplot(data=anchors_df, x='px', y='py', hue='anchor name',
linewidth=0.0, legend=False, ax=ax)
ax.set_title(anchor_name)
g = sns.scatterplot(data=anchors_df, x='px', y='py', hue='anchor name',
linewidth=0.0, legend='full', ax=ax)
g.legend(loc='center left', bbox_to_anchor=(1.25, 0.5), ncol=1)
fig.suptitle('Scatter plots with size proportional to distance error.')
# -
# # Reconstruction Algorithm
# ## bandlimited table
# +
from table_tools import *
def format_here(number):
if number > 10000:
return '{:.2e}'.format(number)
else:
return '{:.1f}'.format(number)
fname = 'results/bandlimited_tuesday.pkl'
outname = 'results/table_bandlimited.tex'
#fname = 'results/bandlimited_tuesday_calib.pkl'
#outname = 'results/table_bandlimited_calib.tex'
result_df = pd.read_pickle(fname)
# convert all numerical columns to float, ignore non-numeric.
result_df = result_df.apply(pd.to_numeric, errors='ignore')
#print_table = result_df[result_df.n_measurements.isin([40, 100, 200, 300, 499])]
print_table = result_df[(result_df.n_complexity >= 5) & (result_df.n_measurements >= 100)]
print_table = print_table[print_table.n_measurements.isin([100, 300, 499])]
methods = ['gt','srls raw', 'srls', 'rls raw', 'rls', 'lm-ellipse',
'lm-ours-weighted', 'ours', 'ours-weighted']
#pretty_print_table(print_table, methods=methods, value='cost_rls')
styler, __ = pretty_print_table(print_table, methods=methods, value='mse')
styler
# -
methods = ['gt','srls raw', 'srls', 'rls raw', 'rls', 'lm-ellipse',
'lm-ours-weighted', 'ours', 'ours-weighted']
__, pt = pretty_print_table(print_table, methods=methods, value='mse')
latex_print(pt, methods, outname, float_format=format_here)
# ## Some bandlimited sanity checks
# plot error vs n measurements and n complexity
plot_df = result_df[result_df.mae < 100]
plot_df = plot_df[plot_df.n_measurements > 100]
fg = sns.FacetGrid(data=plot_df, col='n_complexity', hue='method', legend_out=True)
fg.map(plt.semilogy, 'n_measurements', 'mae', linestyle='', marker='.', alpha=0.5)
legend = plt.gca().get_legend()
plt.legend()
# understand why N=100, K=19 is so bad...
df = result_df.loc[(result_df.n_measurements==100) & (result_df.n_complexity==19), :]
fig, ax = plt.subplots()
for method, df_m in df.groupby('method'):
ax.scatter(df_m.n_it, df_m.mse, label=method)
ax.set_yscale('log')
ax.legend(loc='upper right')
# ## polynomial table
# +
fname = 'results/polynomial_tuesday.pkl'
outname = 'results/table_polynomial.tex'
#fname = 'results/polynomial_tuesday_calib.pkl'
#outname = 'results/table_polynomial_calib.tex'
result_df = pd.read_pickle(fname)
# convert all numerical columns to float, ignore non-numeric.
result_df = result_df.apply(pd.to_numeric, errors='ignore')
print_table = result_df
print_table = print_table[print_table.n_measurements.isin([10, 20, 30, 50])]
methods = ['gt','srls raw', 'srls', 'rls raw', 'rls', 'lm-line',
'lm-ours-weighted', 'ours', 'ours-weighted']
styler, __ = pretty_print_table(print_table, methods=methods, value='mse')
styler
# -
__, pt = pretty_print_table(print_table, methods=methods, value='mse')
latex_print(pt, methods, outname, index_names=False, index=False, float_format=format_here)
# # Sandbox (space to try out stuff)
#
# ### Example reconstructions
# +
from evaluate_dataset import compute_distance_matrix, compute_anchors
chosen_distance = 'distance'
#chosen_distance = 'distance_gt'
anchor_names = None
## Construct anchors.
anchors = compute_anchors(anchors_df, anchor_names)
print(anchors.shape)
## Construct times.
times = full_df[full_df.system_id == range_system_id].timestamp.unique()
## Construct D.
D, times = compute_distance_matrix(full_df, anchors_df, anchor_names, times, chosen_distance)
print(D.shape)
if np.sum(D > 0) > D.shape[0]:
print('Warning: multiple measurements for times:{}/{}!'.format(
np.sum(np.sum(D > 0, axis=1)>1), D.shape[0]))
## Construct ground truth.
points_gt = get_ground_truth(full_df, times)
# +
from other_algorithms import apply_algorithm
from coordinate_fitting import fit_trajectory
print(D.shape)
fig, ax = plt.subplots()
ax.scatter(points_gt.px, points_gt.py, s=10)
traj.set_n_complexity(3)
#method = 'ours-weighted'
method = 'lm-line'
coeffs, __, __ = apply_algorithm(traj, D, times, anchors, method=method)
traj.set_coeffs(coeffs=coeffs)
traj.plot_pretty(times=times, ax=ax, color='red', label='fitted')
traj.print()
coeffs = fit_trajectory(points_gt.T, times, traj)
traj.set_coeffs(coeffs=coeffs)
traj.plot_pretty(times=times, ax=ax, color='green', label='best fit')
ax.set_xlim(*xlim)
ax.set_ylim(*ylim)
ax.legend()
# -
# ### Piecewise linear reconstructions: correlation between cost and reconstruction error
# +
result_df = pd.read_pickle('results/polynomial_tuesday_calib.pkl')
result_df = pd.read_pickle('results/polynomial_tuesday.pkl')
chosen_measure = 'mse'
for N, df_N in result_df.groupby('n_measurements'):
Ks = df_N.n_complexity.unique()
fig, axs = plt.subplots(2, len(Ks), squeeze=False, sharey=True)
fig.suptitle(f'N={N}')
i = 0
for K, df_K in df_N.groupby('n_complexity'):
ax1, ax2 = axs[:, i]
for method, df_method in df_K.groupby('method'):
ax1.plot(df_method[chosen_measure].values, label=method)
ax2.scatter(df_method['cost_rls'].values, df_method[chosen_measure], label=method)
ax2.set_xscale('log')
ax2.set_yscale('log')
ylim_chosen = min(200, df_method[chosen_measure].max())
xlim_rls = min(2000, df_method['cost_rls'].max())
ax1.set_ylim([1, ylim_chosen])
ax2.set_ylabel(str.upper(chosen_measure))
ax2.set_xlim([1, xlim_rls])
ax2.legend(loc='lower left', bbox_to_anchor=[1, 0])
i += 1
# -
|
PublicDatasets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
import math
import geopandas as gpd
import json
from bokeh.io import output_notebook, show, output_file
from bokeh.plotting import figure
from bokeh.models import GeoJSONDataSource, LinearColorMapper, ColorBar, NumeralTickFormatter
from bokeh.palettes import brewer
from bokeh.io.doc import curdoc
from bokeh.models import Slider, HoverTool, Select
from bokeh.layouts import widgetbox, row, column
from bokeh.application import Application
from bokeh.application.handlers import FunctionHandler
output_notebook()
# -
#path = r'C:/Users/ShrekTheOger/Documents/GitHub/final-project-final-project-bowen-and-natasia'
path = r'C:\Users\engel\Documents\GitHub\final-project-final-project-bowen-and-natasia'
#import warnings
#warnings.filterwarnings('ignore')
#Load informational dataframe from refined file folder
df = os.path.join(path+'/refined_data', 'df_income_pop.csv')
df = pd.read_csv(df)
#retrive shape file of us in county level
county_shp = os.path.join(path+'/raw_data', 'cb_2020_us_county_20m', 'cb_2020_us_county_20m.shp')
county = gpd.read_file(county_shp)
#Combining state_id with county_id into same format as in university shape file
l = []
for i in range(len(df['state_id'])):
if len(str(df['state_id'][i])) == 1:
if len(str(df['county_id'][i])) == 1:
a = '0'+ str(df['state_id'][i]) + '00' + str(df['county_id'][i])
elif len(str(df['county_id'][i])) == 2:
a = '0'+ str(df['state_id'][i]) + '0' + str(df['county_id'][i])
else:
a = '0'+ str(df['state_id'][i]) + str(df['county_id'][i])
else:
if len(str(df['county_id'][i])) == 1:
a = str(df['state_id'][i]) + '00' + str(df['county_id'][i])
elif len(str(df['county_id'][i])) == 2:
a = str(df['state_id'][i]) + '0' + str(df['county_id'][i])
else:
a = str(df['state_id'][i]) + str(df['county_id'][i])
l.append(a)
df.insert(3, 'GEOID', l)
df.drop(labels=['state_id', 'county_id'], axis=1, inplace = True)
county.head()
df.head()
df['GEOID2'] = df['COUNTYFIPS'].apply(lambda x: '{0:05}'.format(x))
df.dtypes
df.tail()
#Combine yearly data of county level income and population data with county geometries
df_shape = county.merge(df, how = 'inner', on='GEOID').set_geometry('geometry')
#retrive shape file of universities
uni_df = os.path.join(path+'/refined_data', 'uni_fund_df.csv')
uni_df = pd.read_csv(uni_df)
universities_shp = os.path.join(path+'/raw_data', 'Colleges_and_Universities-shp', 'Colleges_and_Universities.shp')
universities = gpd.read_file(universities_shp)
universities['IPEDSID'] = universities['IPEDSID'].astype(str).astype(int)
uni_shape = universities.merge(uni_df, how = 'inner', on = 'IPEDSID').set_geometry('geometry')
uni_shape = uni_shape.to_crs(df_shape.crs)
df_shape['year']
# # Interactive Geographic Map Using Bokeh
# Here we use Bokeh to display county level population information with hovering and display income discrepensies using density map.
# We also displayed where the top 50 universities at each state, also displaying their fund level using hovering effect.
#
# Our code is heavily inspired by <NAME> from his website https://jimking100.github.io/2019-09-04-Post-3/
def json_data(selectedYear, selectedState):
yr = selectedYear
st = selectedState
# Pull selected year from state data
df_yr = df_shape[(df_shape['year'] == yr) & (df_shape['state'] == st)].copy().reset_index(drop=True)
merged_json = json.loads(df_yr.to_json())
# Convert to json preferred string-like object
json_data = json.dumps(merged_json)
return json_data
def json_data_uni(selectedYear, selectedState):
yr = selectedYear
st = selectedState
# Pull selected year from state data
uni_yr = uni_shape[(uni_shape['year'] == yr) & (uni_shape['state'] == st)].copy().reset_index(drop=True)
merged_json = json.loads(uni_yr.to_json())
# Convert to json preferred string-like object
json_data = json.dumps(merged_json)
return json_data
# +
"""The disadvantage of using a static scale is some county like Los Angelous has a overly dense population which pulled
the scale upper limit really high, so when we have state level inspections, many counties that have less than 100,000 become
insignificant, even those big counties like Cook County become drawfted by the abnormal dense population from city like New
York and Los Angeles."""
# This dictionary contains the formatting for the data in the plots
format_data = [('total_population', 0, 12000000,'0,0', 'Total Population'),
('total_native', 0, 6750000,'0,0', 'Total Native'),
('total_born_in_state', 0, 5200000,'0,0', 'Total Born in State'),
('total_born_out_state', 0, 2000000,'0,0', 'Total Born out State'),
('total_born_outside_us', 0, 130000,'0,0', 'Total Born out State'),
('total_foreign_born', 0, 3500000,'0,0', 'Total Foreigner'),
('income_past12m', 0, 80000,'$0,0', 'Income in Past 12 Months')
]
#Create a DataFrame object from the dictionary
format_df = pd.DataFrame(format_data, columns = ['field' , 'min_range', 'max_range' , 'format', 'verbage'])
# -
# Define the callback function: update_plot
def update_plot(attr, old, new):
# The input yr is the year selected from the slider
yr = slider.value
print(yr)
st = select_st.value
print(st)
new_data = json_data(yr, st)
new_uni = json_data_uni(yr, st)
# The input cr is the criteria selected from the select box
input_field = select.value
input_field = format_df.loc[format_df['verbage'] == input_field, 'field'].iloc[0]
# Update the plot based on the changed inputs
p = make_plot(input_field)
# Update the layout, clear the old document and display the new document
layout = column(p, select, select_st, slider)
curdoc().clear()
curdoc().add_root(layout)
# Update the data
geosource.geojson = new_data
geosource_uni.geojson = new_uni
# Define the callback function: update_plot
def update_plot_uni(attr, old, new):
# The input yr is the year selected from the slider
yr = slider.value
st = select_st.value
new_uni = json_data_uni(yr, st)
# Update the data
geosource_uni.geojson = new_uni
#def update_plot_all(attr, old, new):
#
# update_plot(attr, old, new)
# update_plot_uni(attr, old, new)
# Create a plotting function
def make_plot(field_name):
# Set the format of the colorbar
min_range = format_df.loc[format_df['field'] == field_name, 'min_range'].iloc[0]
max_range = format_df.loc[format_df['field'] == field_name, 'max_range'].iloc[0]
field_format = format_df.loc[format_df['field'] == field_name, 'format'].iloc[0]
# Instantiate LinearColorMapper that linearly maps numbers in a range, into a sequence of colors.
color_mapper = LinearColorMapper(palette = palette, low = min_range, high = max_range)
# Create color bar.
format_tick = NumeralTickFormatter(format=field_format)
color_bar = ColorBar(color_mapper=color_mapper, label_standoff=18, border_line_color=None, location = (0, 0))
# Create figure object.
verbage = format_df.loc[format_df['field'] == field_name, 'verbage'].iloc[0]
p = figure(title = verbage,
plot_height = 650, plot_width = 850)
p.xgrid.grid_line_color = None
p.ygrid.grid_line_color = None
p.axis.visible = False
# Add patch renderer to figure.
r1 = p.patches('xs','ys', source = geosource, fill_color = {'field' : field_name, 'transform' : color_mapper},
line_color = 'black', line_width = 0.25, fill_alpha = 1)
# Specify color bar layout.
p.add_layout(color_bar, 'right')
# Add the hover tool to the graph
p.add_tools(HoverTool(renderers=[r1], tooltips=hover))
r2 = p.circle('x','y', color = 'red', source=geosource_uni, size=5, fill_alpha = 0.7)
p.add_tools(HoverTool(renderers=[r2], tooltips=hover_uni))
return p
# +
# Input geojson source that contains features for plotting for:
# initial year 2010 and initial income per capital for counties -- Income_past12m
geosource = GeoJSONDataSource(geojson = json_data(2015,'Illinois'))
geosource_uni = GeoJSONDataSource(geojson = json_data_uni(2015,'Illinois'))
input_field = 'income_past12m'
# Define a sequential multi-hue color palette.
palette = brewer['Blues'][8]
# Reverse color order so that dark blue is highest obesity.
palette = palette[::-1]
# Add hover tool
hover = [ ('County','@NAME'),('State', '@state'),('Total population', '@total_population{,}'),
('Born instate', '@total_born_in_state{,}'),('Born outstate', '@total_born_out_state{,}'),]
hover_uni = [ ('School','@NAME'),('Fund', '@fund{,}')]
# Call the plotting function
p = make_plot(input_field)
# Make a slider object: slider
slider = Slider(title = 'Year',start = 2010, end = 2019, step = 1, value = 2015)
# Make a selection object: select
select = Select(title='Select Criteria:', value='income_past12m', options=['Income in Past 12 Months', 'Total Population'])
# Make a selection object: select
select_st = Select(title='Select Target State:', value='Illinois', options=uni_shape['state'].unique().tolist())
# Make a column layout of widgetbox(slider) and plot, and add it to the current document
# Display the current document
layout = column(p, select, select_st, slider)
def modify_doc(doc):
doc.add_root(column(layout))
slider.on_change('value', update_plot)
select.on_change('value', update_plot)
select_st.on_change('value', update_plot)
slider.on_change('value', update_plot_uni)
select.on_change('value', update_plot_uni)
select_st.on_change('value', update_plot_uni)
handler = FunctionHandler(modify_doc)
app = Application(handler)
show(app)
# -
|
History/Bokeh with Interact (still working on it)- Natasia Edit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computational Astrophysics
# ## 02. Pyplot Visualization. Example 2
# ---
# ## <NAME>
#
# Observatorio Astronómico Nacional\
# Facultad de Ciencias\
# Universidad Nacional de Colombia
#
# ---
# ### About this notebook
#
# In this worksheet, we use a synthetic dataset to illustrate the use of `%matplotlib inline` and `%matplotlib notebook` in visualization.
#
# ---
# ### Generation of the dataset in astroML
#
# Using the `generate_mu_z` module of the `astroML` package, we simulate a set of 100 supernovas, to obtain the values of the distance modulus ( $\mu$ ), redshift ( $ z $ ) and error ( $d\mu$ ).
# Detailed information about this module can be found at
# https://www.astroml.org/modules/generated/astroML.datasets.generate_mu_z.html
#
import numpy as np
from matplotlib import pyplot as plt
from astroML.datasets import generate_mu_z
z_sample, mu_sample, dmu = generate_mu_z(100, random_state=21)
# ---
#
# ## The `%matplotlib inline` magic
# Plot the dataset using `pyplot` and using `%matplotlib inline`. This allows to specify a default size for all the plots in the notebook.
# %matplotlib inline
plt.rcParams['figure.figsize'] = [9.5, 6]
plt.plot(z_sample, mu_sample, 'o')
plt.plot(z_sample, mu_sample, '.r');
# Note that all plots have the same size.
plt.plot(z_sample, mu_sample, '.')
plt.xlabel(r'$z$')
plt.ylabel(r'$\mu$')
# ---
#
# ## The `%matplotlib notebook` magic
# Plot the dataset using `pyplot` and using `%matplotlib notebook`. This gives a zoomable and resizeable plot. This is the best for quick tests where you need to work interactively.
#
# **Note: To activate this command it is needed to restart the kernel!**
# +
# %matplotlib notebook
import numpy as np
from matplotlib import pyplot as plt
from astroML.datasets import generate_mu_z
z_sample, mu_sample, dmu = generate_mu_z(100, random_state=21)
# -
plt.plot(z_sample, mu_sample, 'o')
plt.plot(z_sample, mu_sample, '.r');
plt.plot(z_sample, mu_sample, '.')
plt.xlabel(r'$z$')
plt.ylabel(r'$\mu$')
|
03._Visualization/presentation/02.pyplotExample02/pyplotExample02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
from scipy.special import expit
import matplotlib.pyplot as plt
from scipy.stats import norm
import warnings
warnings.filterwarnings("ignore")
# # Metroplis Hasting
# +
def affine_z(z, a, b):
return(a*z+b)
def unnormalised_true_dist(z):
a = 10; b = 3
pz = np.exp(-z**2)*expit(affine_z(z,a,b))
return pz
def next_state(current_state, tau):
transition_dist = norm(loc = current_state, scale = tau)
return(transition_dist.rvs(1))
def accept_prob(pos_dist, current_state, next_state):
current_state_p = pos_dist(current_state)
next_state_p = pos_dist(next_state)
return(np.min([1, next_state_p/current_state_p]))
def true_dist(a, b, x_min, x_max, n_samples):
z = np.linspace(x_min, x_max, n_samples)
F = compute_gauss_hermite_approx(z,a,b)
pz = unnormalised_true_dist(z)/F
plt.plot(z, pz, "r", linewidth=2.0)
plt.xlabel("z")
plt.ylabel("probability")
def compute_gauss_hermite_approx(z, a, b):
degree = 100
def only_sigmoid(z):
# returns sigmoid(ax + b) for function exp(-x^2)*sigmoid(10x+3)
return (expit(a*z+b))
points, weights = np.polynomial.hermite.hermgauss( degree) # points.shape = weights.shape = (degree,)
F = np.sum(only_sigmoid(points) * weights)
return (F)
def metropolis_hasting(true_dist, n_iter, burn_in, m, tau):
tau = np.sqrt(tau)
z_p = 0 #initial value of z
rejected = np.array([])
accepted = np.array([])
for i in range(1, burn_in+1):
z_n = next_state(z_p, tau)
prob = accept_prob(true_dist, z_p, z_n)
u = np.random.uniform(0, 1, 1)
if (u <= prob):
z_p = z_n
# print("Burn-in for "+str(burn_in)+" iterations done!")
for i in range(1, n_iter+1):
accept = False
z_n = next_state(z_p, tau)
prob = accept_prob(true_dist, z_p, z_n)
u = np.random.uniform(0,1,1)
if(u<=prob):
accept = True
if(i%m == 0):
if(accept):
accepted = np.hstack((accepted, z_n))
else:
accepted = np.hstack((accepted, z_p))
rejected = np.hstack((rejected, z_n))
if(accept):
z_p = z_n
# print("Sampling for " +str(n_iter) + " iterations done!")
return accepted, rejected
# +
print("\n=======================================================================")
print(" METROPOLIS HASITNG SAMPLING ")
print("=======================================================================")
n_iter = 50000
burn_in = 100000
m = 10 ; a = 10; b = 3
tau_val = [0.01, 0.1, 0.2, 0.5, 1]
accept_rate = np.array([])
plots = []
for tau in tau_val:
# print("\n\ntau = ", tau)
accepted, rejected = metropolis_hasting(unnormalised_true_dist, n_iter, burn_in, m, tau)
accept_rate = np.hstack((accept_rate, rejected.shape[0]/accepted.shape[0]))
plots.append(accepted)
# print("Accept Rate ", 1 - accept_rate[-1])
# plt.hist(accepted, bins=50, density=True)
# true_dist(a, b, -5, 5, 500) # get true distribution
# plt.legend(["True_dist", "tau="+str(tau)])
# plt.savefig("metropolis_hasting_tau"+str(tau).replace('.','_')+".png")
# plt.clf()
# plt.show()
accept_rate = (1 - accept_rate)
#print(accept_rate)
plt.clf()
plt.plot(tau_val, accept_rate, marker='o')
plt.grid()
plt.xlabel("Tau")
plt.ylabel("Acceptance Rate")
plt.show()
# -
fig, axs = plt.subplots(3, 2, subplot_kw=dict(projection="polar"))
# # Hybrid Monte Carlo
# +
def total_energy(state):
z = state[0]
r = state[1]
s = state[2]
u = z**2 - np.log(expit(affine_z(z,a,b)))
k = 0.5*r*r/s
return(1/np.exp(u+k))
def dU_dz(z):
grad = 2*z - (1 - expit(affine_z(z,a,b)))*a
return(grad)
def leapfrog(z, r, s, eps, L):
for i in range(L):
r -= (eps/2)*dU_dz(z)
z += eps*r/s
r -= (eps/2)*dU_dz(z)
return (z, r)
def hybrid_monte_carlo(pos_dist, n_iter, burn_in, m, eps, L):
s = 1
r = norm(loc=0, scale=np.sqrt(s))
z_p = np.array([2], dtype=np.float) # initial value of z
rejected = np.array([])
accepted = np.array([])
for i in range(1, burn_in + 1):
r_p = r.rvs(1) # sampling r from normal distribution
z_n, r_n = leapfrog(np.copy(z_p), np.copy(r_p), s, eps, L)
r_n*=(-1)
prob = accept_prob(pos_dist, [z_p, r_p, s], [z_n, r_n, s])
u = np.random.uniform(0, 1, 1)
if (u <= prob):
z_p = z_n
# print("Burn-in for " + str(burn_in) + " iterations done!")
for i in range(1, n_iter + 1):
accept = False
r_p = r.rvs(1) # sampling r from normal distribution
z_n, r_n = leapfrog(np.copy(z_p), np.copy(r_p), s, eps, L)
r_n *= (-1)
prob = accept_prob(pos_dist, [z_p, r_p, s], [z_n, r_n, s])
u = np.random.uniform(0, 1, 1)
if (u <= prob):
accept = True
if (i % m == 0):
if (accept):
accepted = np.hstack((accepted, z_n))
else:
accepted = np.hstack((accepted, z_p))
rejected = np.hstack((rejected, z_n))
if (accept):
z_p = z_n
# print("Sampling for "+str(n_iter) + " iterations done!")
return accepted, rejected
# +
L = 10
eps_val = [0.005, 0.01, 0.1, 0.2, 0.5]
accept_rate = np.array([])
for eps in eps_val:
# plt.clf()
# print("\n\nepsilon = ", eps, "\t L = ", L)
accepted, rejected = hybrid_monte_carlo(total_energy, n_iter, burn_in, m, eps, L)
accept_rate = np.hstack((accept_rate, rejected.shape[0]/accepted.shape[0]))
# print("Accept Rate ", 1 - accept_rate[-1])
plt.hist(accepted, bins=50, density=True)
true_dist(a, b, -5, 5, 500) # get true distribution
plt.legend(["True_dist", "eps="+str(eps)])
plt.show()
plt.savefig("hybrid_monte_carlo_"+str(eps).replace('.','_')+".png")
accept_rate = (1 - accept_rate)
# print(accept_rate)
plt.clf()
plt.plot(eps_val, accept_rate, marker='o')
plt.xlabel("eps")
plt.ylabel("AcceptanceRate")
plt.grid()
# -
|
assignments/a5/Q1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import os
import torch
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
from torchvision import datasets, models, transforms
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import time
import copy
import cv2
"""
below is for local load only
"""
# im_names = os.listdir('/home/clint/project2/images')
# path = '/home/clint/project2/files/'
# name = 'Xa_test.csv'
# names = os.listdir(path)
# df1 = pd.DataFrame()
# for name in names:
# print(name)
# df2 = pd.read_csv(f'{path}{name}')
# df1 = df1.append(df2, ignore_index=True)
# df_select = df1[df1['Image File'].isin(im_names)]
# df_select2 = df_select.dropna()
# df_val = df_select2.sample(int(len(df_select2)*.2/200))
# df_train = df_select2[~df_select2.index.isin(df_val.index.tolist())]
# df_train = df_train.sample(int(len(df_train)/200))
# data_val = P2DataLoader(df_val, root='/home/clint/project2/images')
# data_train = P2DataLoader(df_train, root='/home/clint/project2/images')
# dataload_val = DataLoader(data_val, batch_size=4, shuffle=True, num_workers=0)
# dataload_train = DataLoader(data_train, batch_size=4, shuffle=True, num_workers=0)
# dataloaders = {'train':dataload_train, 'val': dataload_val}
# dataset_sizes = {'train':len(df_train),'val':len(df_val)}
class P2DataLoader():
def __init__(self, df, root='', train=True, transform=None,):
# path to image data
# self.csv_data = pd.read_csv(csv_file)
self.root = root
self.csv_data = df
self.target = np.array(self.csv_data['Sex (subj)']) # label of image
self.im_file = np.array(self.csv_data['Image File']) # label of image
self.h = np.array(self.csv_data['Image Height'])
self.w = np.array(self.csv_data['Image Width'])
self.x1 = np.array(self.csv_data['X (top left)'])
self.x2 = np.array(self.csv_data['X (bottom right)'])
self.y1 = np.array(self.csv_data['Y (top left)'])
self.y2 = np.array(self.csv_data['Y (bottom right)'])
def __getitem__(self, index):
"""
Args:
index (int): Index
Returns:
dict: {'image': image, 'target': index of target class, 'meta': dict}
"""
#make slicer from bbox
img, target, h, w = io.imread(f'{self.root}/{self.im_file[index]}'), self.target[index], self.h[index], self.w[index]
# slicer from bbox
img = img[self.y1[index]:self.y2[index],self.x1[index]:self.x2[index]]
# resize to a standard size
img = cv2.resize(img , (64, 64))
img = torch.from_numpy(img).float()
img = img.reshape(3, 64, 64)
"""
I have a transform library we can use here
"""
# if self.transform is not None:
# img = self.transform(img)
out = {'image': img,
'target': int(target),
'meta': {'im_size': (h, w), 'index': index, 'class_ID': target}}
return out
def get_image(self, index):
img = index
return img
def __len__(self):
return len(self.csv_data)
def train_model(model, criterion, optimizer, scheduler, num_epochs=25):
start = time.time()
gold_acc, gold_model_wts = 0.0, copy.deepcopy(model.state_dict())
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
scheduler.step()
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
current_corrects, current_loss = 0, 0.0
# Here's where the training happens
print('Iterating through data...')
for data in dataloaders[phase]:
inputs = data['image'].to(device)
labels = data['target'].to(device)
# We need to zero the gradients, don't forget it
optimizer.zero_grad()
"""
forward
"""
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
"""
backward
"""
if phase == 'train':
loss.backward()
optimizer.step()
# loss statistics
current_loss += loss.item() * inputs.size(0)
current_corrects += torch.sum(preds == labels.data)
e_loss = current_loss / dataset_sizes[phase]
epoch_ac = current_corrects.double() / dataset_sizes[phase]
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
phase, e_loss, epoch_ac))
"""
copy only if the model improved
"""
if phase == 'val' and epoch_ac > gold_acc:
gold_acc = epoch_ac
gold_model_wts = copy.deepcopy(model.state_dict())
print()
end = time.time() - start
print('Training complete in {:.0f}m {:.0f}s'.format(
end // 60, end % 60))
print('Best val Acc: {:4f}'.format(gold_acc))
# Now we'll load in the best model weights and return it
model.load_state_dict(gold_model_wts)
return model
def Tain_ResNet(path2trainCSV, val_fract, path2ims, num_epochs=25):
"""
load data
inputs: path2trainCSV, val_fract, path2ims
"""
df = pd.read_csv(path2trainCSV)
df_val = df.sample(int(len(sample)*val_fract))
df_train = df[~df.index.isin(df_val.index)]
data_val = P2DataLoader(df_val, root=path2ims)
data_train = P2DataLoader(df_train, root=path2ims)
dataload_val = DataLoader(data_val, batch_size=4, shuffle=True, num_workers=0)
dataload_train = DataLoader(data_train, batch_size=4, shuffle=True, num_workers=0)
dataloaders = {'train':dataload_train, 'val': dataload_val}
dataset_sizes = {'train':len(df_train),'val':len(df_val)}
"""
build res
"""
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
res_mod = models.resnet34(pretrained=True)
num_ftrs = res_mod.fc.in_features
res_mod.fc = nn.Linear(num_ftrs, 2)
res_mod = res_mod.to(device)
criterion = nn.CrossEntropyLoss()
optimizer_ft = optim.SGD(res_mod.parameters(), lr=0.001, momentum=0.9) # Observe that all parameters are being optimized
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_ft, step_size=7, gamma=0.1)# Decay LR by a factor of 0.1 every 7 epochs
"""
train
"""
base_model = train_model(res_mod, criterion, optimizer_ft, exp_lr_scheduler, num_epochs=num_epochs)
# -
|
resNet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="1HyoZ4SjWtVh"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
dfa=pd.read_csv('/content/Sample_for_AI.csv')
dfb=pd.read_csv('/content/Sample_for_BI.csv')
from google.colab import data_table
# + colab={"base_uri": "https://localhost:8080/", "height": 921} id="Pnggzg_wzAn9" outputId="a263495f-1e7c-4416-8fba-ada47533fe3a"
dfb
# + colab={"base_uri": "https://localhost:8080/", "height": 623} id="T8ljv6JrApVO" outputId="1e3655b0-d83d-48f2-d621-b5f06baadc43"
dfa
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="zxNe8neAWtkW" outputId="a75c5d1b-e844-40db-caca-df58da1bb8d8"
dfa.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="xMuk83QU5IJX" outputId="f241849a-87e6-4a2b-d70a-fabaf2824ea2"
dfa.info
# + colab={"base_uri": "https://localhost:8080/"} id="afbhMeB19hkO" outputId="21e256b5-a1c5-4829-9da8-b3457fd4697b"
dfa.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="NBh2XIBN17tZ" outputId="c7d26387-82d1-4532-f97d-03e475798fde"
dfb.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 830} id="sWu3vewCa6jG" outputId="93e94b21-aa76-4895-812d-7fb84714b824"
import matplotlib.pyplot as plt
dfa.hist(bins=50, figsize=(20,15))
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="65fka-ORoR2W" outputId="f8294bbc-38ec-42af-932a-51180a1f0996"
dfa.columns
# + colab={"base_uri": "https://localhost:8080/"} id="TIG4gBrz1xcH" outputId="d9dcbe64-f448-46a5-f27b-fc588454fdde"
dfb.columns
# + id="yH3Q5XAPg9wb"
dfa.Specialization.fillna('unknown', inplace = True)
dfb['Page Views Per Visit'].fillna(0, inplace = True)
dfb.TotalVisits.fillna(0, inplace = True)
dfa['Page Views Per Visit'].fillna(0, inplace = True)
dfa.TotalVisits.fillna(0, inplace = True)
dfb.Country.fillna('unknown', inplace = True)
dfb.Specialization.fillna('unknown', inplace = True)
dfb['How did you hear about Us'].fillna('unknown', inplace = True)
dfb.Current_Occupation.fillna('unknown', inplace = True)
dfb['What matters most to you in choosing a course'].fillna('unknown', inplace = True)
dfb.Source.fillna(0, inplace = True)
dfb['Last Activity'].fillna('unknown', inplace = True)
# + colab={"base_uri": "https://localhost:8080/"} id="HRirNhbH3E8-" outputId="b60f4522-8c8c-439a-a078-0c5f1d39a3eb"
dfa.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="0ePSSzMFjnOO" outputId="de314162-887c-4b72-c035-d7bb511ca2c9"
dfb.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="CjeRK2HDoBus" outputId="9d08fc15-fbe6-4ab5-ef64-62c399aa9960"
dfa["income_cat"] = pd.cut(dfa["Page Views Per Visit"],
bins=[0., 1.5, 3.0, 4.5, 6., np.inf],
labels=[1, 2, 3, 4, 5])
dfa["income_cat"].hist()
# + colab={"base_uri": "https://localhost:8080/", "height": 800} id="bRYBZ0XQ29mj" outputId="a36d1ae3-36cf-4d27-93b6-8d2ef126eb8f"
from pandas.plotting import scatter_matrix
attributes = ["Purchased", "TotalVisits", "Total Time Spent on Website",
"Page Views Per Visit",'Specialization','Origin']
scatter_matrix(dfa[attributes], figsize=(15, 10))
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="WtXdNzFj5T12" outputId="99521472-f144-464c-a2ec-9c9cf4634226"
dfa.plot(kind="scatter", x="Page Views Per Visit", y="TotalVisits",
alpha=0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 174} id="pE7L1I4W7GV9" outputId="42f008a3-fc5d-4f9a-d8cb-530cdebd01a6"
corr_matrix = dfa.corr()
corr_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="gz11gk5uBq5K" outputId="290c0146-a71d-480d-fa13-91cad80464ad"
dfa.columns
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="tvFwOTcND0wY" outputId="37f26b7b-56b1-477a-b9a2-02362da1a6dc"
Origin_cat = dfa[["Origin"]]
Origin_cat.head(10)
# + id="KOKv-4qX8v8C"
dfa_labels = dfa["Purchased"].copy()
# + id="EsJ1UzovExIc"
from sklearn.preprocessing import OrdinalEncoder
ordinal_encoder = OrdinalEncoder()
Origin = ordinal_encoder.fit_transform(Origin_cat)
# + colab={"base_uri": "https://localhost:8080/"} id="75YfrCMDJ16n" outputId="2cfefc74-fc36-44c7-d89a-b6566e29a371"
Origin[:10]
ordinal_encoder.categories_
# + id="aOlT2J65cAzS"
Y = ordinal_encoder.fit_transform(dfa[["Purchased"]])
Special= ordinal_encoder.fit_transform(dfa[["Specialization"]])
# + colab={"base_uri": "https://localhost:8080/"} id="cwQJbkbdF3QL" outputId="33c1d084-5dcd-4b15-eb4f-ede1613c8d17"
Y
# + colab={"base_uri": "https://localhost:8080/"} id="_wfInIrjrn4s" outputId="7336a630-a351-4f2f-9dfa-9cf1479c107f"
df=[Special,Origin]
df
# + id="XdvXWogbZq9k"
# + colab={"base_uri": "https://localhost:8080/"} id="6eXLIBuyb2KG" outputId="5750c4cc-5cac-4b41-e3d5-aaae3a9f9c23"
Country = ordinal_encoder.fit_transform(dfb[["Country"]])
print(Country[:10])
Specialization = ordinal_encoder.fit_transform(dfb[["Specialization"]])
print(Specialization[:10])
about_Us= ordinal_encoder.fit_transform(dfb[["How did you hear about Us"]])
Current_Occupation = ordinal_encoder.fit_transform(dfb[["Current_Occupation"]])
print(Current_Occupation[:10])
matters_most= ordinal_encoder.fit_transform(dfb[["What matters most to you in choosing a course"]])
Origin = ordinal_encoder.fit_transform(dfb[["Origin"]])
#Source = ordinal_encoder.fit_transform(dfb[["Source"]])
Do_Not_Email = ordinal_encoder.fit_transform(dfb[["Do Not Email"]])
Do_Not_Call = ordinal_encoder.fit_transform(dfb[["Do Not Call"]])
Purchased = ordinal_encoder.fit_transform(dfb[["Purchased"]])
Last_Activity= ordinal_encoder.fit_transform(dfb[["Last Activity"]])
Last_Activity
# + colab={"base_uri": "https://localhost:8080/", "height": 556} id="CBb9Vd-ui7bw" outputId="c3f7c903-db9e-4a45-f7a1-678ea997a415"
df = pd.DataFrame({'Purchased':Purchased.flatten()})
df['Do_Not_Call']=pd.DataFrame({'Do_Not_Call':Do_Not_Call.flatten()})
df['Last_Activity']=pd.DataFrame({'Last_Activity':Last_Activity.flatten()})
df['Do_Not_Email']=pd.DataFrame({'Do_Not_Email':Do_Not_Email.flatten()})
df['Origin']=pd.DataFrame({'Origin':Origin.flatten()})
df['matters_most']=pd.DataFrame({'matters_most':matters_most.flatten()})
df['Current_Occupation ']=pd.DataFrame({'Current_Occupation ':Current_Occupation .flatten()})
df['about_Us']=pd.DataFrame({'about_Us':about_Us.flatten()})
df['Origin']=pd.DataFrame({'Origin':Origin.flatten()})
df['Specialization']=pd.DataFrame({'Specialization':Specialization.flatten()})
df['Country']=pd.DataFrame({'Country':Country.flatten()})
df['TotalVisits']=dfb['TotalVisits']
df['Total Time Spent on Website']=dfb['Total Time Spent on Website']
df['Page Views Per Visit']=dfb['Page Views Per Visit']
x= df.drop("Purchased",axis= 1)
y=df["Purchased"]
x
# + id="QBQ3AjTrKIyP"
from sklearn.preprocessing import OneHotEncoder
cat_encoder = OneHotEncoder()
Email_cat_1hot = cat_encoder.fit_transform(dfa[["Do Not Email"]])
Email=Email_cat_1hot.toarray()
# + id="l6zpxpPfTeG_"
Call_cat_1hot = cat_encoder.fit_transform(dfa[["Do Not Call"]])
Call=Call_cat_1hot.toarray()
# + colab={"base_uri": "https://localhost:8080/"} id="NzfQmzg6Axec" outputId="150dcd33-2aa1-49f0-97f8-6ad985023aca"
#dfa['TotalVisits'].fillna(0)
#dfa['Page Views Per Visit'].fillna(0)
# + colab={"base_uri": "https://localhost:8080/"} id="bT1xbZNbBjZv" outputId="5316a78a-86a1-459a-a132-2fad337212c0"
dfb.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 576} id="ZkxaV5FY_fgL" outputId="e3f21465-0bd2-40ae-94f8-7d0bbbae158a"
corr_matrix = x.corr()
corr_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="T8cGHvnEOKJx" outputId="c5e96078-cc8e-43c6-9693-fb91c5c997e7"
dfa.columns
# + colab={"base_uri": "https://localhost:8080/"} id="4VbGUBL_jqwE" outputId="01d0a006-ae56-4390-fdf4-6a7d09c2fa77"
Yb = pd.get_dummies(y).values
Yb
# + id="heUHU1B2gToo"
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x,Yb,test_size = 0.25, random_state =42)
# + id="ceRlfMrhqpkz"
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense , InputLayer , Dropout
# + colab={"base_uri": "https://localhost:8080/"} id="rb7j-hKjqyIx" outputId="78151b3b-eb82-435a-da06-a9f980ca3ea1"
model1 = Sequential()
model1.add(Dense(6, input_shape = (12,),kernel_initializer='he_uniform',activation='relu'))
model1.add(Dense(2, activation='softmax'))
model1.summary()
# + id="YsacJa-Lq93y"
model1.compile(optimizer="adam",
loss= "categorical_crossentropy",
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="XMWE6bmorC_W" outputId="59db7d06-a897-4c2c-b634-d1da21151191"
model1.get_weights()
# + colab={"base_uri": "https://localhost:8080/"} id="THw-Q0C1rMDj" outputId="83a13e76-f001-4a51-e8d0-cd0a8b29f756"
model1.fit(x_train, y_train, epochs= 100)
# + colab={"base_uri": "https://localhost:8080/"} id="vtcB6NSRrRQF" outputId="7122cafa-39c6-43af-c839-3883928a8275"
model1.evaluate(x_test,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="6bSyJ8JTrjx9" outputId="32b47c3f-7d5b-4630-fe4f-dece534d5932"
y_predict1 = model1.predict(x_test)
y_predict1
# + colab={"base_uri": "https://localhost:8080/"} id="vACgG1iHhYAK" outputId="6d35f251-37c0-4187-a0f8-3eb1d074a254"
pickle.dump(model1, open("AI-model.pkl", "wb"))
# + colab={"base_uri": "https://localhost:8080/"} id="gM_3AYL3uT-J" outputId="e3b42d66-1089-4ed1-e017-8f5ea4e51a78"
print(y_test)
# + id="_ITbB__RvaGq"
y_test_class = np.argmax(y_test, axis=1)
y_predict_class = np.argmax(y_predict1, axis=1)
# + colab={"base_uri": "https://localhost:8080/"} id="ACd5zvqpvhvC" outputId="62251d4a-bec6-49fa-8218-db0e00035741"
y_test_class.shape
# + colab={"base_uri": "https://localhost:8080/"} id="_psG0BxhvoPB" outputId="a9ab15ba-d891-48c5-d93d-4fda56cbf38e"
y_test_class
# + colab={"base_uri": "https://localhost:8080/"} id="MUauooNQvv-c" outputId="04e5da0c-1d98-4d6b-f1d8-4ee57eb2d125"
y_predict_class.shape
# + colab={"base_uri": "https://localhost:8080/"} id="BOXahc5sv35z" outputId="1426b7d6-b829-4bab-d28f-51545692d14b"
y_predict_class
# + colab={"base_uri": "https://localhost:8080/"} id="6vInV6Ecwk2d" outputId="ae4da8c8-f9a0-4840-fab5-544cdadf1ef9"
from sklearn.metrics import confusion_matrix,classification_report
print(classification_report(y_test_class, y_predict_class))
print(confusion_matrix (y_test_class, y_predict_class))
# + colab={"base_uri": "https://localhost:8080/"} id="FIPrrjJ1B8iB" outputId="55cca876-46d0-4a4a-a092-95556424b845"
dfa.columns
# + id="LILYIqLMCEh8"
dft = pd.DataFrame({'Purchased':Y.flatten()})
dft['Email']=pd.DataFrame({'Email':Email.flatten()})
dft['Call']=pd.DataFrame({'Call':Call.flatten()})
dft['Special']=pd.DataFrame({'Special':Special.flatten()})
dft['Origin']=pd.DataFrame({'Origin':Origin.flatten()})
dft['TotalVisits']=dfa['TotalVisits']
dft['Total Time Spent on Website']=dfa['Total Time Spent on Website']
dft['Page Views Per Visit']=dfa['Page Views Per Visit']
X= dft.drop("Purchased",axis= 1)
Y=dft["Purchased"]
# + colab={"base_uri": "https://localhost:8080/"} id="9EV0YJgsP-Ps" outputId="fd7145e1-6c89-4fbc-c088-a97b11ecc687"
headers = list(X)
train_X, test_X, train_Y, test_Y = train_test_split(X, Y, test_size=0.33, stratify=Y, random_state=0)
print(train_X.shape, test_X.shape)
print()
print('Number of rows in Train dataset: {train_df.shape[0]}')
print(train_Y.value_counts())
print()
print('Number of rows in Test dataset: {test_df.shape[0]}')
print(test_Y.value_counts())
# + id="9IpI7fmDRDl8"
import xgboost as xgb
import pickle
# + colab={"base_uri": "https://localhost:8080/"} id="I2oKrifoQfKk" outputId="b938e566-5347-4329-acfe-73a9dbce040e"
model = xgb.XGBClassifier(max_depth=12,
subsample=0.33,
objective='binary:logistic',
n_estimators=300,
learning_rate = 0.01)
eval_set = [(train_X, train_Y), (test_X, test_Y)]
model.fit(train_X, train_Y.values.ravel(), early_stopping_rounds=15, eval_metric=["error", "logloss"], eval_set=eval_set, verbose=True)
# + colab={"base_uri": "https://localhost:8080/"} id="lOnGLwmBRrNo" outputId="5ed1b651-03c6-4bdb-d65f-5d8f28dd9998"
y_pred = model.predict(test_X)
y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="MxaB2lvFR2Dh" outputId="b843a80d-fb86-4320-ab61-f54b4ff71a5f"
from sklearn.metrics import confusion_matrix,classification_report
print(classification_report(test_Y, y_pred))
print(confusion_matrix (test_Y, y_pred))
# + id="_7ZQbEPbhB14"
pickle.dump(model, open("BI-model.pkl", "wb"))
# + id="xLeCFh1Xi1Cj"
from matplotlib import pyplot
# + colab={"base_uri": "https://localhost:8080/", "height": 544} id="81Kj0E1SiVm2" outputId="af4cdf1f-05b5-4518-ec94-d2c1f9313461"
# retrieve performance metrics
results = model.evals_result()
epochs = len(results['validation_0']['error'])
x_axis = range(0, epochs)
# plot log loss
fig, ax = pyplot.subplots()
ax.plot(x_axis, results['validation_0']['logloss'], label='Train')
ax.plot(x_axis, results['validation_1']['logloss'], label='Test')
ax.legend()
pyplot.ylabel('Log Loss')
pyplot.title('XGBoost Log Loss')
pyplot.show()
# plot classification error
fig, ax = pyplot.subplots()
ax.plot(x_axis, results['validation_0']['error'], label='Train')
ax.plot(x_axis, results['validation_1']['error'], label='Test')
ax.legend()
pyplot.ylabel('Classification Error')
pyplot.title('XGBoost Classification Error')
pyplot.show()
|
eda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Student Alcohol Consumption
# ### Introduction:
#
# This time you will download a dataset from the UCI.
#
# ### Step 1. Import the necessary libraries
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/04_Apply/Students_Alcohol_Consumption/student-mat.csv).
# ### Step 3. Assign it to a variable called df.
# ### Step 4. For the purpose of this exercise slice the dataframe from 'school' until the 'guardian' column
# ### Step 5. Create a lambda function that will capitalize strings.
# ### Step 6. Capitalize both Mjob and Fjob
# ### Step 7. Print the last elements of the data set.
# ### Step 8. Did you notice the original dataframe is still lowercase? Why is that? Fix it and capitalize Mjob and Fjob.
# ### Step 9. Create a function called majority that returns a boolean value to a new column called legal_drinker (Consider majority as older than 17 years old)
# ### Step 10. Multiply every number of the dataset by 10.
# ##### I know this makes no sense, don't forget it is just an exercise
|
04_Apply/Students_Alcohol_Consumption/Exercises.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kalz2q/mycolabnotebooks/blob/master/math_calc_ordinarydiff.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="3IRm9LFtUKjN"
# # メモ
#
# pandoc -f mediawiki -t markdown math_calc_ordinarydiff.wiki -o output.md
#
# の実験
#
# https://ja.wikibooks.org/wiki/解析学基礎/常微分方程式
#
# を pandoc してみた。 そこそこ使える。
# + [markdown] id="ctEVSUa4UAX_"
# 常微分方程式とは何ぞや
# ----------------------
#
# 微分方程式とは、独立変数 $x$ と、 $x$ の関数 $y(x)$ 、およびその何階かの導関数を含む方程式である。一般化すれば、微分方程式は
#
# $$f(x, y, y', \cdots, y^{(n)}) = 0$$
#
# の形に書くことのできる方程式である。そして、この方程式に含まれる導関数のうちもっとも高階の導関数が $y^{(n)}$ であるとき、これを $n$ 階微分方程式と呼び、この方程式を満たすような関数を求める操作を、微分方程式を解く、という。
#
# 微分方程式は、大きく分けて常微分方程式と偏微分方程式に分かれる。常微分方程式とは、一変数関数とその導関数からなる方程式である。一方、偏微分方程式とは、多変数関数とその偏導関数との方程式である。ここでは、常微分方程式の解き方について記述することにし、本書では特に断りのない場合「微分方程式」は常微分方程式をさしているものとする。
#
#
# + [markdown] id="dDfM--c7Uj4f"
# ### 解の種類
#
# 微分方程式は微分された関数が含まれた方程式であるから、その解を求めるためには多くの場合積分操作が必要であり、解には積分定数が含まれる。 $n$ 階微分方程式であれば $n$ 個の任意の積分定数が含まれる。このような積分定数を含む形の解を**一般解**と呼ぶ。
#
# 一般解のうち、積分定数にある値を与えた解を**特殊解**と呼ぶ。
#
# さらに、微分方程式の解のなかには、方程式の解であるにもかかわらず、積分定数にどのような値を代入しても表すことのできない解も存在する。このような解を**特異解**と呼ぶ。
#
# * 一般解
# * 特殊解
# * 特異解
#
#
#
#
# + [markdown] id="DXeKaE7XVSbU"
# ### 問題の種類
#
# 微分方程式を解くとき、一般解が重要になる場面はそう多くない。現実にはある独立変数 $x$ における従属変数 $y$ の値が定まっていて、その条件を満たすような特殊解が必要になる場合がほとんどである。
#
# たとえば時刻 $t=0$において、ある関数 $y(t)$ の値が $y_0$ と分かっている時に $y(t)$ に関する微分方程式を解くような場合である。
#
# このような、ある初期条件 $ y(x_0) = y_0 $ を満たすような微分方程式
#
# $$f(x, y, y', \cdots, y^{(n)})$$
#
# の特殊解を求める問題を**初期値問題**といい、これらをみたす特殊解を求めることを初期値問題を解くという。
#
# また、例えば位置 $x=0$ と $x=L$ で常に $y=0$ となるような波(固定端)の変位 $y(x)$ に関する微分方程式を解くという状況もある。
#
# このような、ある境界条件
#
# $$y(x_i) = y_i (i = 0, 1, \cdots, n)$$
#
# を満たすような微分方程式
#
# $$f(x, y, y', \cdots, y^{(n)})$$
#
# の特殊解を求める問題を**境界値問題**といい、これらをみたす特殊解を求めることを境界値問題を解くという。
#
#
#
#
# + [markdown] id="fguKVWTEVoeC"
# 初等解法
# --------
#
# 微分方程式を、有限回の式変形や変数変換や積分によって解く方法を初等解法と呼ぶ。はじめに、微分方程式の初等解法について解説する。なお、どのような微分方程式であっても初等解法によって解くことができるとは限らず、この手法が適用できる場合は限られてくることに注意されたい。
#
# まずは、1 階の微分方程式について考えることにする。
#
#
# + [markdown] id="8ykQ42a7WPID"
# ### 変数分離形
#
# 一般に、 $n$ 階微分方程式が
#
# $$y^{(n)} = f(x, y, y', \cdots, y^{(n-1)})$$
#
# の形で書き表されるとき、これを正規形と呼ぶ。
#
# 1階微分方程式の正規形
#
# $$y' = f(x, y)$$ において、右辺の式が
#
# $$f(x, y) = X(x)Y(y)$$
# のように $x$ のみの関数と $y$ のみの関数との積の形に変形できるとき、これを**変数分離形**の微分方程式と呼ぶ。この場合、微分方程式は
#
# $$y' = X(x)Y(y)$$
#
# の形になっているから、$Y(y) \neq 0$ と仮定して両辺を $Y(y)$ で割ることにより
#
# $$\frac{y'}{Y(y)} = X(x)$$
#
# と変形して、左辺が $y$ とその導関数のみの式、右辺は $x$ のみの式となるように分離することができる。
#
# もし $Y(y) = 0$ を満たす $y$ の値が存在すれば、その値を $y=a$ とすると、もとの微分方程式に代入して
#
# $$y'(x) = 0$$
#
# を得る。一方、いま置いた $y=a$ も $y'=0$ を満たす関数である。すなわち、微分方程式の解は
#
# $$y(x) = a$$
#
# と簡単に求めることができる。これは微分方程式の特殊解である。
#
# では、$Y(y) \neq 0$として変数を分離した式に戻ろう。
# 分離した式の両辺を $x$ で積分して、
#
# $$\int\frac{1}{Y(y(x))}y'(x)dx = \int X(x)dx$$
#
# 左辺は置換積分の公式より、
#
# $$\int\frac{1}{Y(y(x))}y'(x)dx = \int\frac{1}{Y(y)}dy$$
#
# であるので、
#
# $$\int\frac{1}{Y(y)}dy = \int X(x)dx$$
#
# を得る。これで両辺の不定積分が計算できれば、微分方程式の解が求まることになる。これは微分方程式の一般解である。
#
#
#
# + [markdown] id="ytA-WiuZWS4g"
# #### 例題
#
# 微分方程式 $y' = xy$ を解く。
#
# これは変数分離形の1階微分方程式である。 $y=0$ のとき $y'=0$ となって、これは微分方程式を満たす。
#
# $y \neq 0$と仮定して両辺を $y$ で割ると、
#
# $$\frac{y'}{y} = x$$
#
# であるから、両辺を $x$ で積分して、
#
# $$\int\frac{1}{y}dy = \int x dx + C$$
#
# となる。両辺の不定積分を計算すれば、
#
# $$\log |y| = \frac{1}{2}x^2 + C$$ となるから、これより
#
# $$y = e^{\frac{1}{2}x^2 + C} = Ae^{\frac{1}{2}x^2}$$(Aは任意の定数)
#
# とすることができる。これは微分方程式の一般解である。
#
# 先に求めた $y=0$ は、一般解で $A=0$ とした場合であるから、微分方程式の特殊解である。したがって、微分方程式の解は
#
# $$y(x) = Ae^{\frac{1}{2}x^2}$$ である。
#
#
#
# + [markdown] id="klOr4RvaW7cx"
# ### 同次形
#
# 一見変数分離形でないように見える微分方程式であっても、適切な変数変換によって変数分離形へ持ち込むことのできる微分方程式が存在する。
#
# 1 階微分方程式の正規形
#
# $$y' = f(x, y)$$ において、右辺の式が
#
# $$f(x, y) = g\left(\frac{y}{x}\right)$$
#
# のように$\frac{y}{x}$の関数として記述できるとき、これを**同次形**の微分方程式と呼ぶ。このとき微分方程式は
#
# $$y' = g\left(\frac{y}{x}\right)$$ の形をしている。
#
# $z(x) = \frac{y(x)}{x}$とおく。このとき $y = xz$ であるから、
#
# $$y' = (xz)' = z + xz'$$
#
# が成り立つ。これを元の微分方程式に代入すると、
#
# $$z + xz' = g(z)$$ となる。これを$z'$について解くと、
#
# $$z' = \frac{g(z) - z}{x}$$
# となって、変数分離形の微分方程式となる。
#
# 変数分離形の方程式の解き方にしたがってこれを解くと、
#
# $$\begin{align}
# \frac{z'}{g(z)-z} &= \frac{1}{x} \\
# \int\frac{1}{g(z)-z}dz &= \int\frac{1}{x}dx + C \\
# \int\frac{1}{g(z)-z}dz &= \log|x| + C
# \end{align}$$
# となる。これで左辺の不定積分を計算し、$z = \frac{y}{x}$を代入し直せば微分方程式の解が得られる。
#
#
#
# + [markdown] id="WGQxWt-cXQWG"
# #### 例題
#
# 微分方程式$y' = \frac{y}{x} + \frac{x}{y}$を解く。
#
# これは同次形の1階微分方程式である。$z = \frac{y}{x}$とおくと、$y = xz$であるからこの微分方程式は
#
# $$\begin{align}
# z + xz' &= z + \frac{1}{z} \\
# xz' &= \frac{1}{z}
# \end{align}$$
# と書き直すことができる。これは変数分離形の微分方程式である。$z \neq 0$に注意して変数分離を行うと
#
# $$zz' = \frac{1}{x}$$ であるから、両辺を $x$ で積分して式変形を行うと、
#
# $$\begin{align}
# \int z dz &= \int\frac{1}{x} dx \\
# \frac{1}{2}z^2 &= \log |x| + C \\
# \log|x| &= \frac{1}{2}z^2 + C \\
# x &= e^{\frac{1}{2}z^2+C} = Ae^{\frac{1}{2}z^2}
# \end{align}$$ となる。ここで$z = \frac{y}{x}$を代入しなおすと、
#
# $$x = Ae^{\frac{y^2}{2x^2}}$$
#
# となる。これが求める微分方程式の一般解である。
#
#
#
#
#
# + [markdown] id="F1McY4N_XmAW"
# ### 同次形の応用
#
# 正規形の 1 階微分方程式
#
# $$y' = f(x, y)$$
#
# について、右辺が $x$ と $y$ の有理関数になっている場合、すなわち
#
# $$f(x, y) = \frac{h(x, y)}{g(x, y)}$$
#
# の場合を考える。このとき、$g(x,y)$ および $h(x,y)$ が特定の形をしている場合は、上手な式変形や変数変換によって同次形の解法を適用することができることが知られている。ここでは、いくつかの例題を用いてそれらの解法を紹介することにする。
#
#
#
# + [markdown] id="jX-vtVF0hRdS"
# #### 例題 1
#
# 微分方程式
#
# $$y' = \frac{2x^2 + 3xy + y^2}{x^2 - 4xy + 2y^2}$$
#
# を解く。
#
# これは、$g(x,y)$ と $h(x,y)$ がともにすべての項で $x, y$ について同次であるような場合である。例えばこのような場合には、右辺の分子と分母を $x^2$ で割ることで
#
# $$y' = \frac{2 + 3(y/x) + (y/x)^2}{1 - 4(y/x) + 2(y/x)^2}$$
#
# となって、容易に同次形の微分方程式へ持ち込むことができる。あとは同次形の解法に従って解けばよい。
#
#
#
#
#
# + [markdown] id="oXVT0ODWhdhy"
# #### 例題 2
#
# 微分方程式
# $$y' = \frac{2x + 3y -8}{x - y + 1}$$
# を解く。
#
# これは、$g(x,y)$ と $h(x,y)$ がともに $x, y$ の1次式になっている場合である。例えばこのような場合は、次の手順で解くことができることが知られている。
#
# はじめに、連立方程式
#
# $$\begin{cases}
# 2x + 3y - 8 = 0 \\
# x - y + 1 = 0
# \end{cases}$$
#
# を解く。これを解くと、解は$(x,y) = (1, 2)$である。この解を用いて、
#
# $$\begin{cases}
# x = u + 1 \\
# y = v + 2
# \end{cases}$$
#
# とおく。これをもとの微分方程式へ代入すると、
#
# $$\begin{align}
# \frac{dy}{dx} &= \frac{2(u+1) + 3(v+2) - 8}{(u+1) - (v+2) + 1} \\
# \frac{dv}{du} &= \frac{2u + 3v}{u - v}
# \end{align}$$ となる。ここで、
#
# $$\frac{dy}{dx} = \frac{d}{dx}(v+2) = \frac{dv}{dx} = \frac{dv}{du}\frac{du}{dx} = \frac{dv}{du}\frac{d}{dx}(x-1) = \frac{dv}{du}$$
#
# を用いた。
#
# このように $x, y$ から $u, v$ への変数変換を施すと、例題 1で見た形の方程式となり、右辺の分母分子を $u$で割ることによって同次形の微分方程式として扱うことができる。そして $u, v$ の式として同次形の微分方程式を解いた後、変数を $u, v$ から $x, y$ に戻せば、求めるべき微分方程式の解が得られる。
#
#
#
#
# + [markdown] id="agGPVbCuhwGI"
# #### 例題 3
#
# 微分方程式
#
# $$y' = \frac{2x + 3y - 4}{4x + 6y - 3}$$
#
# を解く。
#
# これは、例題 2 のようにして連立方程式を解こうとしても、連立方程式の解が存在しないか、あるいは1つに定まらない場合である。このような場合は、右辺の分母を $z$ と置くことによって一般解を求めることができる。
#
# この問題では、
#
# $$z = 4x + 6y - 3$$ とおくと、分子は
#
# $$2x + 3y - 4 = \frac{1}{2}(4x + 6y - 3) - \frac{5}{2} = \frac{1}{2}z - \frac{5}{2}$$
# である。また、
#
# $$z' = 4 + 6y'$$
#
# であるから、この微分方程式は変数分離形へと変形することができて
#
# $$\begin{align}
# \frac{z' - 4}{6} &= \frac{\frac{1}{2}z - \frac{5}{2}}{z} \\
# \frac{z}{7z-15}z' &= 1 \\
# \int\frac{z}{7z-15}dz &= x + C
# \end{align}$$
#
# と変形できる。この左辺に $z = 4x + 6y - 3$ を代入すれば求めるべき微分方程式の一般解が求まる。
#
#
#
#
# + [markdown] id="MsMN4btyiBej"
# #### 例題 4
#
# 微分方程式
#
# $$y' = \frac{2x + 3y - 4}{3}$$
#
# を解く。
#
# これは、例題 3と同様に連立方程式を解こうとしても解が一意に定まらず、かつ、分母が定数になっている場合である。この場合は右辺を $z$ と置けばよい。
#
# $$z = \frac{2x + 3y - 4}{3}$$
#
# とおくと、
#
# $$z' = \frac{2 + 3y'}{3} = \frac{2 + 3z}{3}$$
#
# となる。これは変数分離形の微分方程式であるから、その方法に従って解いた後で
#
# $$z = \frac{2x + 3y - 4}{3}$$
#
# を代入すれば求めるべき一般解が得られる。
#
#
#
#
# + [markdown] id="_qd795nVXPKi"
# #### 例題 5
#
# 微分方程式
#
# $$y' = \frac{x^2y}{x^3+y}$$
#
# を解く。
#
# これは、同次形をさらに一般化させた微分方程式である。同次形では、正規形の微分方程式
#
# $$y' = f(x,y)$$
#
# の右辺 $f(x,y)$ について、$\lambda$ を定数として
#
# $$f(\lambda x, \lambda y) = f(x, y)$$
#
# が成り立つ。この例題は同次形ではないため、微分方程式の右辺を $f(x,y)$ と置いてもこれは成り立たない。しかし、
#
# $$f(\lambda x, \lambda^3y) = \frac{(\lambda x)^2\lambda^3y}{(\lambda x)^3 + \lambda^3y} = \lambda^2\frac{x^2y}{x^3+y} = \lambda^2f(x,y)$$
#
# が成り立っている。
#
# 一般に、$f(x,y)$ について $n \neq 0$ として
#
# $$f(\lambda x, \lambda^n y) = \lambda^{n-1}f(x, y)$$
#
# が成り立つとき、
#
# $$z = \frac{y}{x^n}$$
#
# とおいて変数変換を施し、式変形を行うことで変数分離形へ持ち込むことができることが知られている。
#
# この例題では、$z = \frac{y}{x^3}$ と置くと、$y = x^3z$ であるから
#
# $$y' = 3x^2z + x^3z'$$
#
# これをもとの微分方程式へ代入すると、
#
# $$\begin{align}
# 3x^2z + x^3z' &= \frac{x^2x^3z}{x^3+x^3z} \\
# &= \frac{x^2z}{1+z} \\
# z' &= -\frac{3z^2 + 2z}{x(1+z)}
# \end{align}$$
#
# となる。これは変数分離形の微分方程式であるから、変数分離形の解法に従って解き、最後に $z = \frac{y}{x^3}$ を代入すればよい。
#
#
#
#
#
# + [markdown] id="hCtenSHUXhOh"
# 1 階線型微分方程式
# -----------------
#
# 1階微分方程式が線型であるとは、与えられた微分方程式が
#
# $$y' + f(x)y = g(x)$$
#
# と書けることである。このように書けない1階微分方程式は1階非線型微分方程式という。
#
#
#
#
# + [markdown] id="xB2VGrg2YEVr"
# ### 斉次 1 階線型微分方程式
#
# 斉次 1 階線型微分方程式とは、1 階線型微分方程式であって、特に $g(x)=0$ であるものをいい、この時この微分方程式は斉次であるという。
#
# $g(x) \neq 0$ の場合は非斉次であるという。また、斉次は「同次」とも呼ばれることがあるが、本書では前者で統一することにする。
#
# まずは斉次1階線型微分方程式を解いてみよう。
#
# 簡単な微分積分法しか知らない我々は、これ程までに限定してやっと解けるようになるのである。
#
# 今解こうとしているのは、次の微分方程式である。
#
# $$y' + f(x)y = 0$$
#
# これは変数分離形の微分方程式である。まず $y \ne 0$ を仮定して、この式を同値変型する。
#
# $${{y'} \over {y}} = -f(x)$$
#
# 両辺を積分して
#
# $${\rm ln}|y| = \int{ - f(x)dx + C_0}$$ 両辺を*e*の肩に掛けて、
#
# $$|y| = e^{\int{ - f(x) dx + C_0}}$$
#
# 右は常に正なので、$e^{C_0}=C$ として、
#
# $$y = Ce^{\int{ - f(x)dx}}$$
#
# この解法を**変数分離法**といい、得られた結果がこの斉次方程式の一般解である。
#
# 一般解はこのようにして求められたが、$y(x_0)=y_0$ となるときの特殊解 $y$ を求めなければならないときもある。斉次1階微分方程式の初期値問題について考えてみよう。
#
# **初期値問題**
#
# $$y' + f(x)y = 0 ; y(x_0)=y_0$$
#
# を解く。
#
# はじめに微分方程式を解くと、先に導いたように一般解
#
# $$y = Ce^{\int{ - f(x)dx}}$$
# を得る。この式の両辺に$(x, y) = (x_0, y_0)$を代入すれば、積分定数*C*の値が求められるため、改めてそれをこの式に代入しなおすことで特殊解が得られる。
#
# あるいは、微分方程式を解く際に不定積分ではなく$x_0$から$x$までの定積分を求めることによって初期値問題を解くこともできる。多少厄介だが、積分記号を外せないときにも解を求めることができる。
#
# 変数分離を施した形
#
# $$\frac{y'}{y} = -f(x)$$ より、両辺を$x_0$から$x$まで定積分する。
#
# $$\begin{align}
# &{\int_{x_0}^{x} {{y'} \over {y}}\ dx'} = {\int_{x_0}^{x} -f(x')\ dx'} \\
# \iff & {\rm ln}y - {\rm ln}y_0= {{\int_{x_0}^{x} -f(x')\ dx'}} \\
# \iff & {{y} \over {y_0}} = e^{\int_{x_0}^{x} -f(x')\ dx'} \\
# \iff & y=y_0e^{\int_{x_0}^{x} -f(x')\ dx' } \\
# \end{align}$$
# 結局、一般解における積分定数*C*が$y_0$に、不定積分が定積分になっただけであった。
#
#
#
# + [markdown] id="2XBMK1aWYLIn"
# #### 例題 1
#
# 微分方程式
#
# $$y' -4xy = 0$$
#
# を解く。
#
# 上の解説の通り、両辺を $y$ で割り変数分離法によって計算する。この微分方程式の一般解は
#
# $$y = Ce^{\int{ - ( -4x )dx }}= Ce^{2x^2}$$
#
# である。
#
#
#
# + [markdown] id="v-WSxgdYcGI1"
# #### 例題 2
#
# 次の微分方程式の初期値問題を解け。
#
# $$y' + y \sin x = 0 ; y(0) = {3 \over 2}$$
#
# この微分方程式の一般解として、変数分離法によって
#
# $$y = Ce^{\cos x}$$
#
# が求められる。この式に$(x,y) = (0, 3/2)$を代入すれば、
#
# $$C = \frac{3/2}{e^{\cos 0}} = \frac{3}{2e}$$
#
# したがって求めるべき特殊解は
#
# $$y = \frac{3}{2e}e^{\cos x} = \frac{3}{2}e^{\cos x -1}$$
#
# あるいは、不定積分の代わりに定積分を行うことにより、
#
# $$y={3 \over 2}e^{-\int_{0}^{x} \sin t\ dt}={3 \over 2}e^{\cos x-1}$$
#
# が導かれる。
#
#
#
#
#
# + [markdown] id="jIb1RRQKcUGE"
# ### 非斉次 1 階線型微分方程式
#
# 次に、非斉次1階線型微分方程式
#
# $$y'+ f(x)y = g(x) \tag {1.1}$$
#
# の解き方を考えてみよう。しかし今、我々にできる事は二つしかない。それは、斉次微分方程式を解くことと、各種式変形を行うことである。これを最大限駆使して解くしかない。具体的には、なんとかして(1.1)を斉次微分方程式
#
# $$z'=\nu(x)$$
#
# の形に式変形して、これを解くのである。
#
# 天下り式であるが、(1.1) にある関数 $h(x)$ をかけて
#
# $$h(x)y' + h(x)f(x)y = h(x)g(x)\tag {1.2}$$
#
# とする。ここで $h(x)$ が
#
# $$\{h(x)y\}'= h(x)y' + h(x)f(x)y \tag {1.3}$$
#
# をみたすような関数であるとすると、$z=h(x)y,\nu(x)=h(x)g(x)$とすれば $z'=\nu(x)$ の形に変形できる。
#
# ではそのような$h(x)$は存在するのだろうか。具体的に求めてみる。
#
# $\{h(x)y\}'=h(x)y'+h'(x)y$ であるから、これを(1.3)に代入すると
#
# $$\begin{align}
# h(x)y' + h'(x)y &= h(x)y' + h(x)f(x)y \\
# h'(x) &= h(x)f(x)
# \end{align}$$
#
# を得る。$h(x)$ についてはこの変数分離形の微分方程式を解けばよい。定数倍は関係ないので、
#
# $$h(x)=e^{\int f(x) dx} \tag{1.4}$$
#
# としてよい。この $h(x)$ は**積分因子**と呼ばれる。
#
# さて、(1.2)と(1.3)より、
#
# $$\{h(x)y\}'=h(x)g(x) \tag{1.5}$$
#
# を得る。これを変形すると、
#
# $$\begin{align}
# h(x)y &= \int h(x)g(x)dx + C \\
# y &= {1 \over {h(x)}}\left(\int h(x)g(x)dx+C\right)
# \end{align}$$
#
# あとはこれに(1.4)を代入すると、一般解
#
# $$y = \frac{1}{e^{\int f(x) dx}}\left(\int e^{\int f(x) dx}g(x)dx + C\right)$$
#
# を得る。
#
# 初期値問題
#
# $$y'+f(x)y=g(x) ; \;\; y(x_0)=y_0$$
#
# を解くには、(1.5) の両辺を積分する際に定積分とすれば、
#
# $$\int_{x_0}^x \{h(x)y\}' dx=\int_{x_0}^x h(x)g(x) dx$$
#
# を得る。あとはこれを $y$ について解けばよい。
#
# 以上、非斉次微分方程式の解法を述べた。手順をまとめると、
#
# 1. 積分因子 $h(x)=e^{\int f(x)dx}$ を求める。
# 2. $g(x)$ に積分因子を掛け積分する。
# 3. それを $h(x)$ で割って一般解とする。
#
# となる。
#
#
#
# + [markdown] id="iD8YKXVPcp10"
# 非斉次1階線型微分方程式の別の解法として、**定数変化法**と呼ばれる方法を紹介する。
#
# 非斉次な微分方程式
#
# $$y' + f(x)y = g(x)$$
#
# を解くのが最終的な目標であるが、ひとまず、右辺を$g(x)=0$とおいて、斉次な微分方程式
#
# $$y' + f(x)y = 0$$
#
# を解くことにする。この形ならば、前々節で見た方法によって、一般解
#
# $$y_h(x) = Ce^{\int{ - f(x)dx}}$$
#
# を得ることができる。ここで、非斉次な場合は積分定数の $C$ が $x$ の関数になると考えて、仮に非斉次微分方程式の解を
#
# $$y = C(x)\; y_h(x)$$
#
# とおく。これを解くべき微分方程式へ代入すると、
#
# $$
# \begin{align}
# \left\{C(x)y_h(x)\right\}' + f(x)C(x)y_h(x) &= g(x) \\
# C'y_h + Cy_h' + f(x)Cy_h &= g(x) \\
# C'y_h + C(y_h' + f(x)y_h) &= g(x)
# \end{align}
# $$
#
# となるが、ここで $y_h$ が斉次微分方程式 $y'+f(x)y = 0$ の解であることから、
#
# $$C'(x)y_h(x) = g(x)$$
#
# が得られる。この中で未知関数は $C'(x)$ のみであるから、両辺を $y_h(x)$ で割って $x$ で積分すると、
#
# $$\begin{align}
# C(x) &= \int \frac{g(x)}{y_h(x)} dx + C \\
# &= \int \frac{g(x)}{e^{\int{ - f(x)dx}}} dx + C \\
# &= \int g(x)e^{\int f(x)dx} dx + C
# \end{align}$$
#
# したがって、求めるべき非斉次微分方程式の一般解は、
#
# $$y = C(x)y_h(x) = e^{\int -f(x)dx} \left\{\int g(x)e^{\int f(x)dx} dx + C\right\}$$
#
# となる。これは積分因子を用いて求めた一般解と等しい。
#
#
#
#
# + [markdown] id="O2PCl2fvnKf4"
# # いまここ
# + [markdown] id="PwTy_5Prcfa9"
# #### 例題 1
#
# 微分方程式
#
# $$y'-2xy=x$$
#
# を解く。
#
# $f(x)=-2x$より、積分因子$h(x)$は、
#
# $$h(x) = e^{\int -2xdx} = e^{-x^2}$$
# である。これを与式右辺($g(x)$)に掛けて積分すると、
#
# $$\int h(x)g(x)dx = \int e^{-x^2}xdx = -\frac{1}{2}e^{-x^2} + C$$ (1.6)
# したがって、微分方程式の一般解は
#
# $$y=\frac{-\frac{1}{2}e^{-x^2} + C}{e^{-x^2}} = -{1\over 2}+Ce^{x^2}$$
# となる。
#
# あるいは、定数変化法によって求めることもできる。仮に斉次な微分方程式
#
# $$y'-2xy=0$$ を解くと、この一般解は
#
# $$y = Ce^{x^2}$$ となる。これより、仮に求めるべき微分方程式の解を
#
# $$y = C(x)e^{x^2}$$ と置いて元の微分方程式に代入すると、
#
# $$C'(x)e^{x^2} = x$$ が得られる。これより、
#
# $$C(x) = \int\frac{x}{e^{x^2}}dx = \int e^{-x^2}xdx = -\frac{1}{2}e^{-x^2} + C$$
# となるから、求める一般解は
#
# $$y = e^{x^2}\left(-\frac{1}{2}e^{-x^2} + C\right) = -\frac{1}{2} + Ce^{x^2}$$
# である。
#
#
# + id="35vYdzF94MpO"
#### 例題 2
初期値問題$y'-2xy=x ; y(1)=2$を解く。
例題1で(1.6)を積分するときに定積分にする。
$$\begin{align}
\int_1^x \{ye^{-t^2}\}'dt &= \int_1^x e^{-t^2}tdt \\
\left[ye^{-t^2}\right]_1^x &= \left[-{1\over 2}e^{-t^2}\right]_1^x \\
ye^{-x^2}-2e^{-1} &= -{1\over 2}e^{-x^2}+{1\over 2}e^{-1} \\
ye^{-x^2} &= -{1\over 2}e^{-x^2}+{5\over 2}e^{-1}
\end{align}$$ したがって求める特殊解は
$$y(x)=-{1\over 2}+{5\over 2}e^{x^2-1}$$
あるいは、例題1で求めた一般解に$(x, y) = (1, 2)$を代入することによって*C*の値を求めてもよい。
# + id="-YVlxxCM4seX"
### ベルヌーイの微分方程式
1 階微分方程式のなかでも、特に
$$y' + f(x)y = g(x)y^n$$
の形の微分方程式をベルヌーイ(Bernoulli)の微分方程式と呼ぶ。$n = 0, 1$であれば上で見た非斉次1階微分方程式あるいは斉次1階微分方程式の形となり、これらの解法が適用できるが、それ以外の場合でも適切な式変形によって線型微分方程式へ帰着できることが知られている。
ベルヌーイの1階微分方程式
$$y' + f(x)y = g(x)y^n, (n \ne 0, 1)$$ の両辺に$(1-n)y^{-n}$をかけると、
$$(1-n)y^{-n}y' + f(x)(1-n)y^{1-n} = g(x)(1-n)$$
となるから、ここで$z = y^{1-n}$とおくと、$z' = (1-n)y^{-n}y'$なので、
$$z' + f(x)(1-n)z = g(x)(1-n)$$
となる。これは $z$ に関する1階線型微分方程式であるから、定数変化法あるいは積分因子を用いる方法によって計算することができて、一般解
$$z = e^{-(1-n)\int f(x)dx} \left\{(1-n)\int g(x)e^{(1-n)\int f(x)dx} dx + C\right\}$$
を得る。これに$z=y^{1-n}$を代入しなおすと、
$$\begin{align}
y^{1-n} &= e^{-(1-n)\int f(x)dx} \left\{(1-n)\int g(x)e^{(1-n)\int f(x)dx} dx + C\right\} \\
y &= e^{-\int f(x)dx} \left\{(1-n)\int g(x)e^{(1-n)\int f(x)dx} dx + C\right\}^\frac{1}{1-n}
\end{align}$$ を得る。
# + id="h1trPCqk4rMx"
### リッカチの微分方程式
1階微分方程式のなかでも、特に
$$y' = f(x)y^2 + g(x)y + h(x)$$
の形に書くことのできる微分法定式をリッカチ(Riccati)の微分方程式と呼ぶ。この形の方程式は初等解法によって一般解を求めることはできない。しかし、なにか1つの特殊解$y_0$が見つかれば、それを元にして一般解を求めることができる。
リッカチの微分方程式
$$y' = f(x)y^2 + g(x)y + h(x)$$
について、ある特殊解$y_0$が与えられているとする。この時、$z = y - y_0$とおいて元の微分方程式へ代入すると、
$$\begin{align}
z' + y_0' &= f(x)(z + y_0)^2 + g(x)(z + y_0) + h(x) \\
z' &= f(x)z^2 + \left\{2f(x)y_0 + g(x)\right\}z + \left\{f(x)y_0^2 + g(x)y_0 + h(x) - y_0'\right\}
\end{align}$$ となる。ここで$y_0$がこの微分方程式の特殊解であることから
$$y_0' = f(x)y_0^2 + g(x)y_0 + g(x)$$ が成り立っているので、
$$z' = f(x)z^2 + \left\{2f(x)y_0 + g(x)\right\}z$$
となる。これはベルヌーイの微分方程式で$n = 2$の場合であるから、[前節で見た方法で解くことができる](#ベルヌーイの微分方程式 "wikilink")。両辺に$-z^{-2}$をかけて
$$-z^{-2}z' = -f(x) - \left\{2f(x)y_0 + g(x)\right\}z^{-1}$$
さらに$u = z^{-1}$とおくと$u' = -z^{-2}z'$であるから
$$u' = -f(x) - \left\{2f(x)y_0 + g(x)\right\}u$$
となって、1階線型微分方程式に帰着する。この一般解は、前節で見た式から
$$z = e^{\int \left\{2f(x)y_0 + g(x)\right\}dx} \left\{-\int f(x)e^{\int \left\{2f(x)y_0 + g(x)\right\}dx} dx + C\right\}^{-1}$$
となり、求めるべき微分方程式の一般解も
$$\begin{align}
y = z + y_0 &= e^{\int \left\{2f(x)y_0 + g(x)\right\}dx} \left\{-\int f(x)e^{\int \left\{2f(x)y_0 + g(x)\right\}dx} dx + C\right\}^{-1} + y_0 \\
&= \frac{e^{\int \left\{2f(x)y_0 + g(x)\right\}dx}}{-\int f(x)e^{\int \left\{2f(x)y_0 + g(x)\right\}dx} dx + C} + y_0
\end{align}$$ と求まる。
# + id="VqZhsy0O4pu4"
### 演習
次の方程式を解け
1. $f'(x)+f(x)\cos x=0$
2. $f'(x)+f(x)\sqrt x \sin x=0$
3. $f'(x)+f(x){{2x}\over {x^2+1}}={1 \over {x^2+1}}$
4. $f'(x)+f(x)=xe^x$
5. $f'(x)+f(x)x^2=1$
6. $f'(x)+f(x){x \over {x^2+1}}=1-{x^3 \over {x^4+1}}f(x)$
7. $f'(x)+f(x)\sqrt{x^2+1}=0,f(0)=\sqrt 5$
8. $f'(x)+f(x)\sqrt{x^2+1}e^-x=0,f(0)=1$
9. $f'(x)+f(x)\sqrt{x^2+1}e^-x=0,f(0)=0$
10. $f'(x)-2xf(x)=x,f(0)=1$
11. $f'(x)+xf(x)=x+1,f({3 \over 2})=0$
12. $f'(x)+2f(x)={1 \over {x^2+1}},f(1)=2$
13. $f'(x)-2xf(x)=x,f(0)=1$
14. $(x^2+1)f'(x)+xf(x)=(x^2+1)^{5 \over 2}$
15. $(x^2+1)f'(x)+3xf(x)=(x^2+1)^{5 \over 2},f(1)={1 \over 3}$
### 原子核の崩壊速度
線型微分方程式のひとつの応用例として、原子核の崩壊に関するものを見てみよう。
物理学者ラザフォードは、放射性元素の原子核は不安定で、一定の割合で崩壊する事を示した。つまり、原子核の数をyという関数で表すことにすれば
: y\'=-λy (5.1)
という関係式が成り立つ。ここで比例定数λは崩壊定数と呼ばれる正数である。
この関係式は、まさに一階線形常微分方程式となっているので、これまでに述べた方法で解くことができる。
y(x~0~)=y~0~とすれば、(5.1)は
$$y=y_0e^{-\lambda(x-x_0)}$$ (5.2)
と解ける。適当に文字を置き換えると、[高等学校理科 物理II
原子と原子核の](高等学校理科_物理II_原子と原子核 "wikilink")1.2.3で述べた式が導かれたことになる。
# + id="BsV5Ti_l4nqg"
一階定数係数連立線型常微分方程式と高階定数係数線型常微分方程式
--------------------------------------------------------------
### 連立線型常微分方程式と行列の指数関数
上の節では一階の線型常微分方程式の解法を述べた。その中でも最もやさしい定数係数の方程式
$$y'=ay$$ の解は、変数分離法により簡単に求まり、
$$y=Ce^{xa}$$ であった。ただし、C=y(0)である。
次に、 $n$ 本の一階定数係数線型常微分方程式を連立させた方程式
$$\begin{cases}
y_1'=a_{11}y_1+a_{12}y_2+\cdots a_{1n}y_n \\
y_2'=a_{21}y_1+a_{22}y_2+\cdots a_{2n}y_n \\
\vdots \\
y_n'=a_{n1}y_1+a_{n2}y_2+\cdots a_{nn}y_n
\end{cases}$$ を考えよう。この方程式は、行列を用いて
$$\mathbf{y}'=A\mathbf{y} \cdots \bigstar$$ と表すことができる。ただし
$\mathbf{y}=\begin{pmatrix}
y_1 \\
y_2 \\
\vdots \\
y_n
\end{pmatrix},A=\begin{pmatrix}
a_{11}&a_{12}&\cdots&a_{1n} \\
a_{21}&a_{22}&\cdots&a_{2n} \\
\vdots&&&\vdots\\
a_{n1}&a_{n2}&\cdots&a_{nn} \\
\end{pmatrix}$である。
方程式が1本のときの例から類推すれば、この連立方程式の解は
$$e^{xA}$$
のようなものが定義できれば、それを用いて表せそうである。しかし、行列の指数関数をどうやって定義すればよいだろうか?そのために、そもそも実数上の関数としての指数関数がどのように定義されるかを考えてみると、次のようにしてTaylor展開で定義できることが思い出される。
$$e^x=\sum_{k=0}^\infty \frac{x^k}{k!}$$
行列であっても、この式に代入することは可能そうである。すなわち、次のように定義する。
**定義** 正方行列*A*に対して、$e^{xA}:=\sum_{k=0}^\infty \frac{(xA)^k}{k!}$
この級数が収束するのか、またどの程度よい収束をするのかが問題だが、結論から言えば一様絶対収束する。詳しい証明は省くが、ゆえにこの級数を項別微分することができ、
$$(e^{xA})'=Ae^{xA}$$ が成り立つ。
このことから、連立線型微分方程式は初期条件を与えると次のように解けることがわかる。
**定理**
$$\mathbf{y}=e^{xA}\mathbf{y}(0)$$
: は、方程式$\bigstar$の初期値$\mathbf{y}(0)=\begin{pmatrix}c_1 \\ c_2 \\ \cdots \\ c_n\end{pmatrix}$における解になっている。
実際に解になっていることは代入によって確かめることができる。
### 高階定数係数線型常微分方程式
次に、 $n$ 階の定数係数線型常微分方程式
$$y^{(n)}=a_{n-1}y^{(n-1)}+\cdots+a_1y'+a_0y$$
を考える。この方程式は、実は次のようにして連立常微分方程式とみなして行列を使って表せる。
$$\frac{d}{dx}\begin{pmatrix}y \\ y' \\ \vdots \\ y^{(n-2)} \\ y^{(n-1)}\end{pmatrix}=
\begin{pmatrix}
0&1&0&\cdots&0 \\
0&0&1&\cdots&0 \\
\vdots&&&&\vdots \\
0&0&0&\cdots&1 \\
a_0&a_1&a_2&\cdots&a_{n-1}
\end{pmatrix}
\begin{pmatrix}y \\ y' \\ \vdots \\ y^{(n-2)} \\ y^{(n-1)}\end{pmatrix}$$
よって、上の節で述べた方法により初期値問題を解くことができる。
### 具体的な行列に対する計算法
では、具体的な係数行列が与えられたとき、どのようにすれば行列の指数関数が計算できるかを見てみよう。
#### 対角行列の場合
対角行列
$$D=\begin{pmatrix}
c_1&0&\cdots&0 \\
0&c_2&\cdots&0 \\
\vdots&&&\vdots \\
0&0&\cdots&c_n
\end{pmatrix}$$ に対して$e^{xD}$を計算してみよう。
すぐにわかるように、
$$D^k=\begin{pmatrix}
c_1^k&0&\cdots&0 \\
0&c_2^k&\cdots&0 \\
\vdots&&&\vdots \\
0&0&\cdots&c_n^k
\end{pmatrix}$$ である。よって、各成分ごとの計算から
$e^{xD}=\begin{pmatrix}
e^{c_1x}&0&\cdots&0 \\
0&e^{c_2x}&\cdots&0 \\
\vdots&&&\vdots \\
0&0&\cdots&e^{c_nx}
\end{pmatrix}$ である。
#### 対角化可能な行列の場合
行列*A*が$P^{-1}AP=D$と対角化可能な場合も行列の指数関数は容易に計算できる。なぜならば、
$$A^k=(PDP^{-1})^k=PD^kP^{-1}$$ なので、これを代入することで
$$e^{xA}=Pe^{xD}P^{-1}$$
となり、対角行列の指数関数は容易に計算できるからである。
#### 対角化不可能な行列の場合
係数行列が対角化不可能なときは上記のようにはいかず、一般にはJordan標準形を用いることになる。しかし、特殊な場合にはそこまでの計算をする必要はない。たとえば、固有値がすべて等しい場合には次のようにして計算することができる。
$n$ 次正方行列*A*の $n$ 個の固有値がすべて$\lambda$のとき、この行列の固有多項式は$(t-\lambda)^n$なので、Cayley-Hamiltonの定理より
$$(A-\lambda I)^n=O$$ である。このことを用いると、
$$\begin{align}
e^{xA}
&=e^{\lambda xI+x(A-\lambda I)} \\
&=e^{\lambda x}\sum_{k=0}^\infty \frac{x^k(A-\lambda I)^k}{k!} \\
&=e^{\lambda x}\sum_{k=0}^{n-1} \frac{x^k(A-\lambda I)^k}{k!} \\
\end{align}$$ と有限回の計算で指数関数を計算することができる。
# + id="u1rZccHj4jKA"
### 具体例
二階の線型常微分方程式の具体例として、ばねにつながれた物体の運動を記述してみよう。ばねにつながれた物体の時刻 $x$ における変位を $y$ とする。このとき、ばねから物体が受ける力は(負の比例定数で)変位に比例することが知られている。このことを用いて物体の運動方程式を記述すると、
$$y''=\frac{-k}{m}y$$
となる。ただし*k*はばね定数と呼ばれる正の数、*m*は物体の質量である。
この方程式を行列を用いて書き直すと、
$$\frac{d}{dx}\begin{pmatrix}y \\ y'\end{pmatrix}=
\begin{pmatrix}0&1 \\ \frac{-k}{m}&0 \end{pmatrix}
\begin{pmatrix}y \\ y'\end{pmatrix}$$
と表せる。$A=\begin{pmatrix}0&1 \\ \frac{-k}{m}&0 \end{pmatrix}$とする。この行列は対角化できるので、指数関数が計算できて、
$$e^{xA}=
\begin{pmatrix}
\cosh(i\sqrt{\frac{k}{m}}x) & -i\sqrt{\frac{m}{k}}\sinh(i\sqrt{\frac{k}{m}}x) \\
i\sqrt{\frac{k}{m}}\sinh(i\sqrt{\frac{k}{m}}x) & \cosh(i\sqrt{\frac{k}{m}}x) \\
\end{pmatrix}=\begin{pmatrix}
\cos(\sqrt{\frac{k}{m}}x) & \sqrt{\frac{m}{k}}\sin(\sqrt{\frac{k}{m}}x) \\
-\sqrt{\frac{k}{m}}\sin(\sqrt{\frac{k}{m}}x) & \cos(\sqrt{\frac{k}{m}}x) \\
\end{pmatrix}$$ である。初期条件を
$\begin{pmatrix} y(0) \\ y'(0)\end{pmatrix}=\begin{pmatrix} y_0 \\ v_0 \end{pmatrix}$
で定めると、解は
$$\begin{pmatrix} y \\ y'\end{pmatrix}=\begin{pmatrix}
\cos(\sqrt{\frac{k}{m}}x) & \sqrt{\frac{m}{k}}\sin(\sqrt{\frac{k}{m}}x) \\
-\sqrt{\frac{k}{m}}\sin(\sqrt{\frac{k}{m}}x) & \cos(\sqrt{\frac{k}{m}}x) \\
\end{pmatrix}\begin{pmatrix} y_0 \\ v_0 \end{pmatrix}=\begin{pmatrix}y_0\cos(\sqrt{\frac{k}{m}}x)+v_0\sqrt{\frac{m}{k}}\sin(\sqrt{\frac{k}{m}}x) \\
-y_0\sqrt{\frac{k}{m}}\sin(\sqrt{\frac{k}{m}}x)+v_0\cos(\sqrt{\frac{k}{m}}x) \end{pmatrix}$$
と求められた。これがばねによって振動する物体の時刻 $x$ における変位と速度である。
常微分方程式の初期値問題の解の存在と一意性
------------------------------------------
前節まででみたように、いくつかの微分方程式については積分計算によって解を具体的に求めることができるが、一方でそのような求積法の存在しない常微分方程式も多い。だが、そのような方程式についても、ある条件を満たせば解の存在や一意性が保証されることがある。ここではそのような例を見ていこう。
もし解の存在や一意性が保証されるならば、簡単に求積できない微分方程式でも少しは調べやすくなる。一意性が保証されるということは、まぐれやあてずっぽうであっても解をひとつみつけさえすれば、解けたのと同じになるからだ。また、ここで扱う存在と一意性に関する定理は、その解を(ある関数列の極限として)具体的に構成する方法を含んでおり、その意味であてずっぽうではなく解を見つける方法を提供してくれてもいるのである。
本節では、独立変数 $x$ の関数 $y$ についての1階常微分方程式
$$y'=f(x,y) ,\ y(x_0)=y_0$$...(\*)
について考える。関数 $y$ が(\*)を満たすことは、
$$y(x)=y_0+\int_{x_0}^x f(t,y(t)) dt$$...(\*)\'
を満たすことと同値であることも注意しておく。2変数関数$f(x,y)$に対していくつかの仮定を課したときに、この方程式の解がどのように構成されるかを見ていく。
# + id="SfTIofgh4hSV"
### 冪級数による解法
本節では、*f*が次の仮定(H1)を満たすとする。
: (H1)
$f(x,y)$は点$(x_0,y_0)$の近傍で解析的(すなわち冪級数展開可能)であり、$f(x,y)=\sum_{j,l=0}^\infty f_{j,l} (x-x_0)^j (y-y_0)^l$と表される。
このとき、次が成り立つ。
**定理5.1.1**
$f(x,y)$が仮定(H1)を満たすとき、(\*)を満たす$x=x_0$の近傍で解析的な関数 $y$ がただひとつ存在する。
これを証明したい。ただ、冪級数の中心が一般の形だと計算が煩雑になるので、ここでは次の形の定理を証明することにする。
**定理5.1.1**\'
$f(x,y)$が原点の近傍で解析的であり、$f(x,y)=\sum_{j,l=0}^\infty f_{j,l} x^j y^l$と表されるとき、常微分方程式
$$y'=f(x,y) ,\ y(0)=0$$...(☆)
を満たす$x=0$の近傍で解析的な関数 $y$ がただひとつ存在する。
いくつかの補題に分けて証明しよう。
**補題5.1.2**
冪級数$y=\sum_{j=0}^\infty y_j x^j$であって(☆)を満たすものがあるならば、その係数$y_j$は一意に定まる。
(証明)
$y_0=0$である。$j \ge 1$のときは$y=\sum_{j=0}^\infty y_j x^j$を(☆)に代入すると、
$$(lhs)=y_1 +2y_2 x+3y_3 x^2+\cdots$$
$$\begin{align}(rhs)
&=f_{0,0}+f_{0,1}(y_1 x+y_2 x^2+\cdots)+f_{0,2}(y_1 x+y_2 x^2+\cdots)^2+f_{1,0} x+f_{1,1} x(y_1 x+y_2 x^2+\cdots)+f_{2,0} x^2+\cdots \\
&=f_{0,0}+(f_{0,1} y_1+f_{1,0}) x+ (f_{0,1}y_2+f_{0,2}y_1^2+f_{1,1}y_1+f_{2,0}) x^2+\cdots \\
\end{align}$$
であり、次数の低い方から係数を比較することで、係数$y_j$が順に決まっていくことがわかる。//
**補題5.1.3**
$f(x,y)$の冪級数展開の優級数$F(x,y)$があるとき、常微分方程式
$$y'=F(x,y) ,\ y(0)=0$$...(☆)\'
の冪級数解は、補題5.1.2で定まる(☆)の冪級数解の優級数である。
(証明)
$f(x,y)=\sum_{j,l=0}^\infty f_{j,l} x^j y^l,F(x,y)=\sum_{j,l=0}^\infty F_{j,l} x^j y^l$とし、(☆)の解を$y=\sum_{j=0}^\infty y_j x^j$、(☆)\'の解を$Y=\sum_{j=0}^\infty Y_j x^j$とする。
ただし、$y,Y$の冪級数表示は現時点では収束性については何も分かっていない、形式的冪級数である。すべての*j*,*l*について$|f_{j,l}| \le |F_{j,l}|$が成り立つならばすべての*j*について$|y_j| \le |Y_j|$であることを数学的帰納法で証明する。$y_0=Y_0=0,y_1=f_{0,0},Y_1=F_{0,0}$なので、$j=0,1$のときは成り立つ。
$j \le m$なるすべての*j*で成り立つと仮定する。補題5.1.2の証明から、$y_m$は$f_{j,l} (j+l \le m-1),y_{j} (j \le m-1)$に関する多項式の値であり、その係数は非負である。$Y_m$も同様に、同じ非負係数多項式に$F_{j,l} (j+l \le m-1),Y_{j} (j \le m-1)$を代入した値である。よって、帰納法の仮定より、$|y_m| \le |Y_m|$が成り立つ。よって、すべての自然数*j*について$|y_j| \le |Y_j|$が成り立つ。//
**補題5.1.4**
$f(x,y)$が原点の近傍$|x| \le r,|y| \le \rho$において$|f(x,y)| \le M$を満たすとき、$\sum_{j,l=0}^\infty \frac{M}{r^j \rho^l} x^j y^l$は$f(x,y)$の冪級数展開の優級数である。
(証明)
$|f_{j,l}| \le \frac{M}{r^j \rho^l}$を示せばよい。$f(x,y)$の定義域を複素変数に拡張して[w:コーシーの積分公式を用いると](w:コーシーの積分公式 "wikilink")、$|x|<r,|y|<\rho$のとき
$f(x,y)=\int_{|\zeta|=r} \frac{d\zeta}{2i\pi}\int_{|\xi|=\rho} \frac{d\xi}{2i\pi} \frac{f(\zeta,\xi)}{(\zeta-x)(\xi-y)}=-\frac{1}{4\pi^2}\int_{|\zeta|=r} d\zeta \int_{|\xi=\rho} d\xi f(\zeta,\xi)\left(\sum_{j=0}^\infty \frac{x^j}{\zeta^{j+1}}\right)\left(\sum_{l=0}^\infty \frac{y^l}{\xi^{l+1}}\right)=-\sum_{j,l=0}^\infty \frac{1}{4\pi^2} \int_{|\zeta|=r} d\zeta \int_{|\xi|=\rho} d\xi \frac{f(\zeta,\xi)}{\zeta^{j+1}\xi^{l+1}} x^j y^l$であるから、$|f_{j,l}| \le \frac{1}{4\pi^2} \int_{|\zeta|=r} d\zeta \int_{|\xi|=\rho} d\xi \frac{|f(\zeta,\xi)|}{|\zeta|^{j+1}|\xi|^{l+1}} \le \frac{M}{r^j \rho^l}$である。//
**補題5.1.5**
補題5.1.3の微分方程式で$F(x,y)=\sum_{j,l=0}^\infty \frac{M}{r^j \rho^l} x^j y^l$としたものの解は、$x=0$の近傍で解析的な関数であり、収束する冪級数で表される。
(証明)
$$y'=\sum_{j,l=0}^\infty \frac{M}{r^j \rho^l} x^j y^l=\frac{M}{\left(1-\frac{x}{r}\right)\left(1-\frac{y}{\rho}\right)}$$
は変数分離形なので解を求めることができて、
$$\left(1-\frac{y}{\rho}\right)dy=\frac{M}{\left(1-\frac{x}{r}\right)}dx$$
$$-\frac{\rho}{2}\left(1-\frac{y}{\rho}\right)^2=-Mr\log\left(1-\frac{x}{r}\right)+C$$
であり、$y(0)=0$より$C=-\frac{\rho}{2}$であることに注意して整理すると、
$$y=\rho-\sqrt{\rho\left(\rho+2Mr\log\left(1-\frac{x}{r}\right)\right)}$$
である。これは確かに$|x| < r(1-e^{-\frac{\rho}{2Mr}})$で解析的な関数である。//
(定理5.1.1'の証明)
補題5.1.3,5.1.4,5.1.5より、補題5.1.2の冪級数は収束する優級数をもち、したがって自身も収束する。よって、この冪級数の極限として、解が一意的に存在することがわかる。//
# + id="Dmsx5hbO4eNg"
### ピカールの逐次近似法
本節では、*f*が次の仮定(H2)を満たすとする。
: (H2)
$f(x,y)$は点$(x_0,y_0)$の近傍$D=\{|x-x_0| \le r,|y-y_0| \le \rho\}$でリプシッツ連続である、すなわちある*K*に対して$|f(x_1,y_1)-f(x_2,y_2)| \le K \sqrt{(x_1-x_2)^2+(y_1-y_2)^2}$が成り立つ。
このとき、解は次のようにして構成される。
**定義5.2.1**
$f(x,y)$が仮定(H2)を満たすとき、漸化式$y_{j+1}(x)=y_0+\int_{x_0}^x f(t,y_j(t)) dx,\ y_0(x)=y_0$で定まる関数列$y_j$を**ピカールの逐次近似列**という。
**定理5.2.2**
$f(x,y)$が仮定(H2)を満たすとき、$M=\max_{(x,y) \in D} |f(x,y)|,\delta=\min\left\{\frac{\rho}{M},r\right\}$とする。閉区間$[x_0-\delta,x_0+\delta]$において(\*)を満たす関数 $y$ がただひとつ存在し、それはピカールの逐次近似列$y_j$の$j \to \infty$における極限として定義される。
これをいくつかの補題に分けて証明しよう。
**補題5.2.3**
$x_0-\delta \le x \le x_0+\delta$のとき、$\left|\int_{x_0}^x f(t,y(t)) dt\right| \le M|x-x_0| \le \rho$である。
(証明)
$\left|\int_{x_0}^x f(t,y(t)) dt\right| \le M|x-x_0| \le M\delta \le \rho$//
補題5.2.3を帰納的に用いることで、任意の*j*について$y_j$の値域が$|y-y_0| \le \rho$に含まれ、したがって関数列$y_j$がwell-definedであることが従う。
次に、解の一意性を先に示しておこう。
**補題5.2.4**
$f(x,y)$が仮定(H2)を満たすとき、閉区間$[x_0-\delta,x_0+\delta]$において(\*)を満たす関数はただ一つである。
(証明)
$y(x),\tilde{y}(x)$がともに(\*)\'を満たすとすると、*f*がリプシッツ連続であることから
$$|y(x)-\tilde{y}(x)|=\left|\int_{x_0}^x f(t,y(t))-f(t,\tilde{y}(t))dt\right| \le \left|\int_{x_0}^x |f(t,y(t))-f(t,\tilde{y}(t))|dt\right| \le K\left|\int_{x_0}^x |y(t)-\tilde{y}(t)| dt\right|$$
である。一方、補題5.2.3より、
$$|y(x)-\tilde{y}(x)| \le |y(x)-y_0|+|\tilde{y}(x)-y_0|=\left|\int_{x_0}^x f(t,y(t)) dt\right|+\left|\int_{x_0}^x f(t,\tilde{y}(t)) dt\right| \le 2\rho$$
なので、
$$|y(x)-\tilde{y}(x)| \le 2\rho K|x-x_0|$$ であり、よって
$$|y(x)-\tilde{y}(x)| \le \left|\int_{x_0}^x 2\rho K|t-x_0| dt\right|=2\rho \frac{(K|x-x_0|)^2}{2}$$
である。同様に繰り返すことで、任意の自然数*l*に対して
$$|y(x)-\tilde{y}(x)| \le 2\rho \frac{(K|x-x_0|)^l}{l!} \le 2\rho \frac{(K\delta)^l}{l!}$$
であることがわかるが、$\lim_{l \to \infty} \frac{(K\delta)^l}{l!}=0$なので、$y(x)=\tilde{y}(x)$である。//
**補題5.2.6** 関数列$y_j$は一様収束する。
(証明) 補題5.2.3と*f*がリプシッツ連続であることより、
$$|y_1(x)-y_0| \le M|x-x_0|$$
$$|y_2(x)-y_1(x)| \le\left|\int_{x_0}^x |f(t,y_1(t))-f(t,y_0(t))| dt\right| \le K\left|\int_{x_0}^x |y_1(t)-y_0(t)| dt\right| \le K\int_{x_0}^x M|t-x_0| dt \le KM\frac{|x-x_0|^2}{2}$$
以下同様に繰り返して、
$$|y_j(x)-y_{j-1}(x)| \le \frac{M}{K}\frac{(K|x-x_0|)^l}{l!} \le \frac{M}{K}\frac{(K\delta)^l}{l!}$$
である。よって、$j<l$のとき
$$|y_j(x)-y_l(x)| \le \sum_{i=j+1}^l \frac{M}{K}\frac{(K\delta)^i}{i!}$$
であるが、右辺は$j \to \infty$において0に収束するので、$y_j$は一様収束する。//
以上で定理5.2.2を示す準備は整った。
(定理5.2.2の証明)
$y_j$が一様収束することに注意して$y_{j+1}(x)=y_0+\int_{x_0}^x f(t,y_j(t)) dx$の両辺の$j \to \infty$における極限を考えると、
$$\lim_{j \to \infty}y_j(x)=y_0+\int_{x_0}^x f(t,\lim_{j \to \infty}y_j(t)) dx$$
である。これは$\lim_{j \to \infty}y_j(x)$が(\*)\'の解であることを示している。
# + id="k4h7xUEJ4ajw"
### コーシーの折れ線とペアノの定理
本節では、*f*が次の仮定(H3)を満たすとする。
: (H3)
$f(x,y)$は点$(x_0,y_0)$の近傍$D=\{|x-x_0| \le r,|y-y_0| \le \rho\}$で連続である。
このとき、解は次のようにして構成される。
**定義5.3.1**
自然数*j*に対し、$x_0-r \le x \le x_0+r$における関数$y_j$を次のように定める。まず、$y_j(x_0)=y_0$とする。次に、非負整数*i*に対して$x_i=x_0+\frac{ir}{j}$と定め、$x_i< x \le x_{i+1}$のときには
$$y_j(x)=y_0+\frac{r}{j}\sum_{d=0}^{i-1}f(x_d,y_j(x_d))+(x-x_i)f(x_i,y_j(x_i))$$...(\#)
とする。$x<0$についても同様にする。このようにして定まる関数$y_j$のグラフは連続な折れ線になり、これを**コーシーの折れ線**という。
前節までに見た「解析的」や「リプシッツ連続」と比べ、「連続」はとても弱い仮定であり、より多くの関数が満たしている。だがその分本節では解の一意性は失われ、存在しか示すことができない。すなわち、次が成り立つのみである。
**定理5.3.2**
コーシーの折れ線$y_j$は一様収束する部分列$y_{j_l}$を持ち、$\lim_{l \to \infty} y_{j_l}$は方程式(\*)の解である。
まず、次の補題を示す。
**補題5.3.3**
$M=\max_{(x,y) \in D} |f(x,y)|,\delta=\min\left\{\frac{\rho}{M},r\right\}$とする。関数列$y_j$は$x_0-\delta \le x \le x_0+\delta$において一様有界かつ同程度連続である。
(証明) 定義より
$$|y_j(x_1)-y_j(x_2)| \le M|x_1-x_2|$$
なので、同程度連続である。また、この式で$x_2=0$とすると
$$|y_j(x)| \le |y_0|+M\delta \le |y_0|+\rho$$ なので、一様有界である。//
補題5.3.3と[アスコリ=アルツェラの定理により](解析学基礎/関数列の極限#アスコリ=アルツェラの定理 "wikilink")、$y_j$が一様収束する部分列を持つことがわかるので、あとはこの部分列の極限が解になっていることを示せばよい。
(定理5.3.2の証明)
$y(x)=\lim_{l \to \infty} y_{j_l}$が(\*)\'の解になっていることを示したい。(\*)と(\#)を辺々引いた式
$$y(x)-y_{j,l}(x)+\sum_{d=0}^{i-1} \int_{x_d}^{x_{d+1}} (f(x,y(x))-f(x_d,y_j(x_d))) dx+\int_{t_i}^t(f(x,y(x))-f(x_i,y_j(x_i))) dx=0$$
が成り立つことを示せばよい。左辺を$A_{j_l}$とする。$A_{j_l}=0$を示したい。まず、任意の$\varepsilon>0$に対してある $n$ が存在して、$l>N$ならば$|y(x)-y_{j_l}(x)|<\varepsilon$である。次に有界閉集合*D*上の連続関数*f*は一様連続なので、任意の$\varepsilon'>0$に対して、$|x_1-x_2|+|y_1-y_2|<\delta'$ならば$|f(x_1,y_1)-f(x_2,y_2)|<\varepsilon'$となるように、$\delta'>0$をとることができる。この$\delta$に対して$\varepsilon<\frac{\delta'}{2}$を満たすように$\varepsilon>0$をとり、この$\varepsilon$に対して$|y(x)-y_{j_l}(x)|<\varepsilon$かつ$(M+1)\frac{r}{j_l}<\frac{\delta'}{2}$を満たすように*l*をとれば、$x_d< x \le x_{d+1}$のときには
$$|x-x_d|+|y(x)-y_{j_l}(x_d)| \le |x-x_d|+|y(x)-y_{j_l}(x)|+|y_{j_l}(x)-y_{j_l}(x_d)| \le \frac{r}{j_l}+\varepsilon+M\frac{r}{j_l}<\delta'$$
なので、$|f(x,y(x))-f(x_d,y_j(x_d))|<\varepsilon'$である。よって、
$$|A_{j_l}| < \varepsilon+(i+1)\int_{x_d}^{x_{d+1}} \varepsilon'=\varepsilon+\frac{(i+1)r\varepsilon'}{j_l}$$
である。$\varepsilon,\varepsilon'$は任意なので、$A_{j_l}=0$である。
# + id="ZcBz15Rv4Vcr"
陰関数型の1階常微分方程式
-------------------------
陰関数型の1階常微分方程式
$$\displaystyle F \Bigl( k+\ell x+my, \; \; \frac{dy}{dx} \; \Bigr)= 0$$
は求積法で一般解を表示することができる。ここに、$F$
は任意の既知関数であり、$k,\; \ell,\; m$ は任意定数である。
この陰関数型1階常微分方程式の一般解は、次に示す三通りの式で与えられる。
:
: $x = \! \int \! \left\{ \frac{1}{\; \ell + m \psi (t)\;} \cdot \frac{\; d \phi (t) \;}{dt} \; \right\} dt+C,$
<!-- -->
:
: $k + \ell x + my = \phi (t),$
<!-- -->
:
: $F \bigl( \phi (t), \; \psi (t) \bigr) \equiv 0.$
ここに、$t$ は媒介変数であり、$\phi (t)$ と $\psi (t)$ は $t$ の関数で、
$F \bigl( \phi (t),\; \psi (t) \bigr) \equiv 0$ は
$\phi (t),\; \psi (t)$ に関する恒等式である。なお $C$ は積分定数である。
以下で、その解法を示す。
与えられた常微分方程式
$\displaystyle F \Bigl( k+\ell x+my,\; \; \frac{dy}{dx} \; \Bigr)= 0$
に対して、$t$ を媒介変数とする任意関数 $\phi (t), \; \psi (t)$
を導入し、
:
: $k + \ell x + my = \phi (t),$
<!-- -->
:
: $\displaystyle \frac{dy}{dx} = \psi (t)$
と置く。ただし、$m\ne 0$ とする。 上式 $k + \ell x + my =\phi (t)$
の両辺を$x$で微分すると、
$$\displaystyle \ell + m \frac{dy}{dx} = \frac{d \phi (t)}{dt} \cdot \frac{dt}{dx}$$
となる。 ここで、$\frac{dy}{dx} = \psi (t)$ と
$\ell + m \frac{dy}{dx} = \frac{d \phi (t)}{dt} \cdot \frac{dt}{dx}$
から $\frac{dy}{dx}$ を消去すると、
$$\displaystyle \ell + m \psi(t) = \frac{d \phi (t)}{dt} \cdot \frac{dt}{dx}$$
を得る。この式を変形すると、
$$\frac{dx}{dt} = \frac{1}{\; \ell + m \psi (t)\;} \cdot \frac{d \phi (t)}{dt}$$
となる。上式は変数分離形であるから積分すると、
$$x= \! \int \! \left\{ \frac{1}{\; \ell + m \psi (t)\;}
\cdot \frac{\; {d} \phi (t)\; }{dt} \, \right\} dt+C$$
となり、$x$ が $t$ の関数として表示された。 これにより $y$ は
$k + \ell x + my =\phi (t)$ と上式
$x = \int \left\{ \frac{1}{\;\ell+m\psi(t)\;} \cdot \frac{d \phi (t)}{dt} \right\} dt+C$
により $t$ の関数として与えられる。なお $C$ は積分定数である。
# + id="TWhcv9mA4SoQ"
#### 例題 1
陰関数型の関数 $F$ が、
$F \bigl( \phi (t),\; \psi (t) \bigr) = \phi (t) - \psi (t) = 0$
のとき、
:
: $x = \! \int \! \left\{ \frac{1}{\; \ell + m \psi (t)\;} \cdot \frac{\; d \phi (t) \;}{dt} \, \right\} dt+C,$
<!-- -->
:
: $k + \ell x + my = \phi (t)$
から、一般解を求めよ。 解きかたは、$\psi (t)=\phi (t)$ の関係を
$x = \! \int \! \left\{ \frac{1}{\; \ell + m \psi (t)\;} \cdot \frac{\; d \phi (t) \;}{dt} \; \right\} dt+C$
に適用すればよい。
# + id="h6_c-uwA4PZH"
#### 例題 2
陰関数型の関数 $F$ が、
$F \bigl( \phi (t), \; \psi (t) \bigr) = \phi (t) - \psi (t) = 0$
の場合は、解き始めの仮定 $k + \ell x + my = \phi (t)$ と
$\frac{dy}{dx} = \psi (t)$ から、
:
: $k + \ell x + my = \frac{dy}{dx}$
が成り立つ。この式は、1階線型常微分方程式であるから求積法で解ける。
その一般解と、「例題1」の一般解とが一致することを確かめよ。
(注:積分定数は異なる形をしている。)
[しようひふんほうていしき](Category:解析学 "wikilink")
|
math_calc_ordinarydiff.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Application fields
# + hide_input=true
from fastai.gen_doc.nbdoc import *
# + [markdown] hide_input=false
# The fastai library allows you to train a [`Model`](https://pytorch.org/docs/stable/nn.html#torch.nn.Module) on a certain [`DataBunch`](/basic_data.html#DataBunch) very easily by binding them together inside a [`Learner`](/basic_train.html#Learner) object. This module regroups the tools the library provides to help you preprocess and group your data in this format.
#
# ## [`collab`](/collab.html#collab)
#
# This submodule handles the collaborative filtering problems.
#
# ## [`tabular`](/tabular.html#tabular)
#
# This sub-package deals with tabular (or structured) data.
#
# ## [`text`](/text.html#text)
#
# This sub-package contains everything you need for Natural Language Processing.
#
# ## [`vision`](/vision.html#vision)
#
# This sub-package contains the classes that deal with Computer Vision.
#
# ## Module structure
#
# In each case (except for [`collab`](/collab.html#collab)), the module is organized this way:
#
# ### [`transform`](/tabular.transform.html#tabular.transform)
#
# This sub-module deals with the pre-processing (data augmentation for images, cleaning for tabular data, tokenizing and numericalizing for text).
#
# ### [`data`](/tabular.data.html#tabular.data)
#
# This sub-module defines the dataset class(es) to deal with this kind of data.
#
# ### [`models`](/tabular.models.html#tabular.models)
#
# This sub-module defines the specific models used for this kind of data.
#
# ### [`learner`](/text.learner.html#text.learner)
#
# When it exists, this sub-module contains functions will directly bind this data with a suitable model and add the necessary callbacks.
|
docs_src/applications.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "skip"}
# <table>
# <tr align=left><td><img align=left src="./images/CC-BY.png">
# <td>Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved MIT license. (c) <NAME></td>
# </table>
# + slideshow={"slide_type": "skip"}
from __future__ import print_function
from __future__ import absolute_import
# %matplotlib inline
import numpy
import matplotlib.pyplot as plt
# + [markdown] hide_input=true slideshow={"slide_type": "slide"}
# # Numerical Differentiation
#
# **GOAL:** Given a set of $N+1$ points $(x_i, y_i)$ compute the derivative of a given order to a specified accuracy.
#
# **Approaches:**
# * Find the interpolating polynomial $P_N(x)$ and differentiate that.
# * Use Taylor-series expansions and the method of undetermined coefficients to derive finite-difference weights and their error estimates
#
# **Issues:** Order vs accuracy...how to choose
# + [markdown] slideshow={"slide_type": "subslide"}
# # Example 1: how to approximate the derivative $f'(x)$ given a discrete sampling of a function $f(x)$
#
# Here we will consider how to estimate $f'(x_k)$ given a $N$ point sampling of $f(x)=\sin(\pi x) + 1/2 \sin(2\pi x)$ sampled uniformly over the interval $x\in [ 0,1]$
# + hide_input=true slideshow={"slide_type": "-"}
N = 11
x = numpy.linspace(0,1,N)
xfine = numpy.linspace(0,1,101)
f = lambda x: numpy.sin(numpy.pi*x) + 0.5*numpy.sin(4*numpy.pi*x)
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1,1,1)
axes.plot(xfine, f(xfine),'b',label='$f(x)$')
axes.plot(x, f(x), 'ro', markersize=12, label='$f(x_k)$')
axes.grid()
axes.set_xlabel('x')
p = numpy.polyfit(x,f(x),N-1)
axes.plot(xfine,numpy.polyval(p,xfine),'g--',label='$P_{{{N}}}$'.format(N=N-1))
axes.legend(fontsize=15)
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 2: how to approximate derivative $f'(x)$ given a discrete sampling of a function $f(x)$
#
# Here we will consider how to estimate $f'(x_k)$ given a $N$ point sampling of Runge's function sampled uniformly over the interval $x\in [ -1,1]$
# + hide_input=true slideshow={"slide_type": "-"}
N = 11
x = numpy.linspace(-1,1,N)
xfine = numpy.linspace(-1,1,101)
f = lambda x: 1./(1. + 25*x**2)
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1,1,1)
axes.plot(xfine, f(xfine),'b',label='$f(x)$')
axes.plot(x, f(x), 'ro', markersize=12, label='$f(x_k)$')
axes.grid()
axes.set_xlabel('x')
p = numpy.polyfit(x,f(x),N-1)
axes.plot(xfine,numpy.polyval(p,xfine),'g--',label='$P_{{{N}}}$'.format(N=N-1))
axes.legend(fontsize=15)
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The interpolating polynomial: review
#
# From our previous lecture, we showed that we can approximate a function $f(x)$ over some interval in terms of a unique interpolating polynomial through $N+1$ points and a remainder term
#
# $$
# f(x) = P_N(x) + R_N(x)
# $$
#
# Where the Lagrange remainder term is
#
# $$R_N(x) = (x - x_0)(x - x_1)\cdots (x - x_{N})(x - x_{N+1}) \frac{f^{(N+1)}(c)}{(N+1)!}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# While there are multiple ways to represent the interpolating polynomial, both $P_N(x)$ and $R_N(x)$ are polynomials in $x$ and therefore differentiable. Thus we should be able to calculate the first derivative and its error as
#
# $$
# f'(x) = P'_N(x) + R'_N(x)
# $$
#
# and likewise for higher order derivatives up to degree $N$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Derivatives of the Lagrange Polynomials
#
# The Lagrange basis, is a particularly nice basis for calculating numerical differentiation formulas because of their basic interpolating property that
#
# $$
# P_N(x) = \sum_{i=0}^N f(x_i)\ell_i(x)
# $$
#
# where $f(x_i)$ is just the value of our function $f$ at node $x_i$ and all of the $x$ dependence is contained in the Lagrange Polynomials $\ell_i(x)$ (which only depend on the node coordinates $x_i$, $i=0,\ldots,N$). Thus, the interpolating polynomial at any $x$ is simply a linear combination of the values at the nodes $f(x_i)$
# + [markdown] slideshow={"slide_type": "fragment"}
# Likewise its first derivative
# $$
# P'_N(x) = \sum_{i=0}^N f(x_i)\ell'_i(x)
# $$
# is also just a linear combination of the values $f(x_i)$
# + [markdown] hide_input=true slideshow={"slide_type": "slide"}
# ## Examples
#
# Given the potentially, highly oscillatory nature of the interpolating polynomial, in practice we only use a small number of data points around a given point $x_k$ to derive a differentiation formula for the derivative $f'(x_k)$. In the context of differential equations we also often have $f(x)$ so that $f(x_k) = y_k$ and we can approximate the derivative of a known function $f(x)$.
# + hide_input=false slideshow={"slide_type": "-"}
N = 9
f = lambda x: 1./(1. + 25*x**2)
#f = lambda x: numpy.cos(2.*numpy.pi*x)
# + hide_input=true slideshow={"slide_type": "-"}
x = numpy.linspace(-1,1,N)
xfine = numpy.linspace(-1,1,101)
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1,1,1)
axes.plot(xfine, f(xfine),'b',label='$f(x)$')
axes.plot(x, f(x), 'ro', markersize=12, label='$f(x_k)$')
x3 = x[5:8]
x3fine = numpy.linspace(x3[0],x3[-1],20)
p = numpy.polyfit(x3,f(x3),2)
axes.plot(x3,f(x3),'m',label = 'Piecewise $P_1(x)$')
axes.plot(x3fine,numpy.polyval(p,x3fine),'k',label = 'Piecewise $P_2(x)$')
axes.grid()
axes.set_xlabel('x')
p = numpy.polyfit(x,f(x),N-1)
axes.plot(xfine,numpy.polyval(p,xfine),'g--',label='$P_{{{N}}}$'.format(N=N-1))
axes.legend(fontsize=14,loc='best')
plt.show()
# + [markdown] hide_input=false slideshow={"slide_type": "subslide"}
# ### Example: 1st order polynomial through 2 points $x=x_0, x_1$:
#
#
# $$
# P_1(x)=f_0\ell_0(x) + f_1\ell_1(x)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Or written out in full
#
# $$
# P_1(x) = f_0\frac{x-x_1}{x_0-x_1} + f_1\frac{x-x_0}{x_1-x_0}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Thus the first derivative of this polynomial for all $x\in[x_0,x_1]$ is
#
# $$
# P'_1(x) = \frac{f_0}{x_0-x_1} + \frac{f_1}{x_1-x_0} = \frac{f_1 - f_0}{x_1 - x_0} = \frac{f_1 - f_0}{\Delta x}
# $$
#
# Where $\Delta x$ is the width of the interval. This formula is simply the slope of the chord connecting the points $(x_0, f_0)$ and $(x_1,f_1)$. Note also, that the estimate of the first-derivative is constant for all $x\in[x_0,x_1]$.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### "Forward" and "Backward" first derivatives
#
# Even though the first derivative by this method is the same at both $x_0$ and $x_1$, we sometime make a distinction between the "forward Derivative"
#
# $$f'(x_n) \approx D_1^+ = \frac{f(x_{n+1}) - f(x_n)}{\Delta x}$$
#
# and the "backward" finite-difference as
#
# $$f'(x_n) \approx D_1^- = \frac{f(x_n) - f(x_{n-1})}{\Delta x}$$
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Note these approximations should be familiar to use as the limit as $\Delta x \rightarrow 0$ these are no longer approximations but equivalent definitions of the derivative at $x_n$.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: 2nd order polynomial through 3 points $x=x_0, x_1, x_2$:
#
#
# $$
# P_2(x)=f_0\ell_0(x) + f_1\ell_1(x) + f_2\ell_2(x)
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Or written out in full
#
# $$
# P_2(x) = f_0\frac{(x-x_1)(x-x_2)}{(x_0-x_1)(x_0-x_2)} + f_1\frac{(x-x_0)(x-x_2)}{(x_1-x_0)(x_1-x_2)} + f_2\frac{(x-x_0)(x-x_1)}{(x_2-x_0)(x_2-x_1)}
# $$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Thus the first derivative of this polynomial for all $x\in[x_0,x_2]$ is
#
# $$
# P'_2(x) = f_0\frac{(x-x_1)+(x-x_2)}{(x_0-x_1)(x_0-x_2)} + f_1\frac{(x-x_0)+(x-x_2)}{(x_1-x_0)(x_1-x_2)} + f_2\frac{(x-x_0)+(x-x_1)}{(x_2-x_0)(x_2-x_1)}
# $$
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise**: show that the second-derivative $P''_2(x)$ is a constant (find it!) but is also just a linear combination of the function values at the nodes.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Special case of equally spaced nodes $x = [-h, 0, h]$ where $h=\Delta x$ is the grid spacing
#
#
# General Case:
# $$
# P'_2(x) = f_0\frac{(x-x_1)+(x-x_2)}{(x_0-x_1)(x_0-x_2)} + f_1\frac{(x-x_0)+(x-x_2)}{(x_1-x_0)(x_1-x_2)} + f_2\frac{(x-x_0)+(x-x_1)}{(x_2-x_0)(x_2-x_1)}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Becomes:
# $$
# P'_2(x) = f_0\frac{2x-h}{2h^2} + f_1\frac{-2x}{h^2} + f_2\frac{2x+h}{2h^2}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# which if we evaluate at the three nodes $-h,0,h$ yields
#
# $$
# P'_2(-h) = \frac{-3f_0 + 4f_1 -1f_2}{2h}, \quad\quad P'_2(0) = \frac{-f_0 + f_2}{2h}, \quad\quad P'_2(h) = \frac{f_0 -4f_1 + 3f_2}{2h}
# $$
#
# Again, just linear combinations of the values at the nodes $f(x_i)$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Quick Checks
#
# In general, all finite difference formulas can be written as linear combinations of the values of $f(x)$ at the nodes. The formula's can be hard to remember, but they are easy to check.
#
# * The sum of the coefficients must add to zero. Why?
# * The sign of the coefficients can be checked by inserting $f(x_i) = x_i$
# + [markdown] slideshow={"slide_type": "fragment"}
# ##### Example
#
# Given
# $$
# P'_2(-h) =\frac{-3f_0 + 4f_1 -1f_2}{2h}
# $$
#
# What is $P'_2(-h)$ if
#
# * $$f_0=f_1=f_2$$
# * $$f_0 = 0, ~f_1 = 1, ~f_2 = 2$$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Error Analysis
#
# In addition to calculating finite difference formulas, we can also estimate the error
#
# From Lagrange's Theorem, the remainder term looks like
#
# $$R_N(x) = (x - x_0)(x - x_1)\cdots (x - x_{N})(x - x_{N+1}) \frac{f^{(N+1)}(c)}{(N+1)!}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Thus the derivative of the remainder term $R_N(x)$ is
#
# $$R_N'(x) = \left(\sum^{N}_{i=0} \left( \prod^{N}_{j=0,~j\neq i} (x - x_j) \right )\right ) \frac{f^{(N+1)}(c)}{(N+1)!}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# The remainder term contains a sum of $N$'th order polynomials and can be awkward to evaluate, however, if we restrict ourselves to the error at any given node $x_k$, the remainder simplifies to
#
# $$R_N'(x_k) = \left( \prod^{N}_{j=0,~j\neq k} (x_k - x_j) \right) \frac{f^{(N+1)}(c)}{(N+1)!}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# If we let $\Delta x = \max_i |x_k - x_i|$ we then know that the remainder term will be $\mathcal{O}(\Delta x^N)$ as $\Delta x \rightarrow 0$ thus showing that this approach converges and we can find arbitrarily high order approximations (ignoring floating point error).
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Examples
#
# #### First order differences $N=1$
#
# For our first order finite differences, the error term is simply
#
# $$R_1'(x_0) = -\Delta x \frac{f''(c)}{2}$$
# $$R_1'(x_1) = \Delta x \frac{f''(c)}{2}$$
#
# Both of which are $O(\Delta x f'')$
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Second order differences $N=2$
#
#
# For general second order polynomial interpolation, the derivative of the remainder term is
#
# $$\begin{aligned}
# R_2'(x) &= \left(\sum^{2}_{i=0} \left( \prod^{2}_{j=0,~j\neq i} (x - x_j) \right )\right ) \frac{f'''(c)}{3!} \\
# &= \left ( (x - x_{i+1}) (x - x_{i-1}) + (x-x_i) (x-x_{i-1}) + (x-x_i)(x-x_{i+1}) \right ) \frac{f'''(c)}{3!}
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Again evaluating this expression at the center point $x = x_i$ and assuming evenly space points we have
#
# $$R_2'(x_i) = -\Delta x^2 \frac{f'''(c)}{3!}$$
#
# showing that our error is $\mathcal{O}(\Delta x^2)$.
# + [markdown] slideshow={"slide_type": "fragment"}
# ### <font color='red'>Caution</font>
#
# High order does not necessarily imply high-accuracy!
#
# As always, the question remains as to whether the underlying function is well approximated by a high-order polynomial.
#
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Convergence
#
# Nevertheless, we can always check to see if the error reduces as expected as $\Delta x\rightarrow 0$. Here we estimate the 1st and 2nd order first-derivative for evenly spaced points
# + slideshow={"slide_type": "subslide"}
def D1_p(func, x_min, x_max, N):
""" calculate consistent 1st order Forward difference of a function func(x) defined on the interval [x_min,xmax]
and sampled at N evenly spaced points"""
x = numpy.linspace(x_min, x_max, N)
f = func(x)
dx = x[1] - x[0]
f_prime = numpy.zeros(N)
f_prime[0:-1] = (f[1:] - f[0:-1])/dx
# and patch up the end point with a backwards difference
f_prime[-1] = f_prime[-2]
return f_prime
def D1_2(func, x_min, x_max, N):
""" calculate consistent 2nd order first derivative of a function func(x) defined on the interval [x_min,xmax]
and sampled at N evenly spaced points"""
x = numpy.linspace(x_min, x_max, N)
f = func(x)
dx = x[1] - x[0]
f_prime = numpy.zeros(N)
f_prime[0] = f[:3].dot(numpy.array([-3, 4, -1]))/(2*dx)
f_prime[1:-1] = (f[2:N] - f[0:-2])/(2*dx)
f_prime[-1] = f[-3:].dot(numpy.array([1, -4, 3]))/(2*dx)
return f_prime
# + [markdown] slideshow={"slide_type": "fragment"}
# #### Note:
#
# This first derivative operator can also be written as a Matrix $D$ such that $f'(\mathbf{x}) = Df(\mathbf{x})$ where $\mathbf{x}$ is a vector of $x$ coordinates. (exercise left for the homework)
# + slideshow={"slide_type": "subslide"}
N = 11
xmin = 0.
xmax = 1.
func = lambda x: numpy.sin(numpy.pi*x) + 0.5*numpy.sin(4*numpy.pi*x)
func_prime = lambda x: numpy.pi*numpy.cos(numpy.pi*x) + 2.*numpy.pi * numpy.cos(4*numpy.pi*x)
D1f = D1_p(func, xmin, xmax, N)
D2f = D1_2(func, xmin, xmax, N)
# + hide_input=true slideshow={"slide_type": "-"}
xa = numpy.linspace(xmin, xmax, 100)
xi = numpy.linspace(xmin, xmax, N)
fig = plt.figure(figsize=(16, 6))
axes = fig.add_subplot(1, 2, 1)
axes.plot(xa, func(xa), 'b', label="$f(x)$")
axes.plot(xa, func_prime(xa), 'k--', label="$f'(x)$")
axes.plot(xi, func(xi), 'ro')
axes.plot(xi, D1f, 'ko',label='$D^+_1(f)$')
axes.legend(loc='best')
axes.set_title("$f'(x)$")
axes.set_xlabel("x")
axes.set_ylabel("$f'(x)$ and $\hat{f}'(x)$")
axes.grid()
axes = fig.add_subplot(1, 2, 2)
axes.plot(xa, func(xa), 'b', label="$f(x)$")
axes.plot(xa, func_prime(xa), 'k--', label="$f'(x)$")
axes.plot(xi, func(xi), 'ro')
axes.plot(xi, D2f, 'go',label='$D_1^2(f)$')
axes.legend(loc='best')
axes.set_title("$f'(x)$")
axes.set_xlabel("x")
axes.set_ylabel("$f'(x)$ and $\hat{f}'(x)$")
axes.grid()
plt.show()
# + slideshow={"slide_type": "subslide"}
N = 11
xmin = -1
xmax = 1.
func = lambda x: 1./(1 + 25.*x**2)
func_prime = lambda x: -50. * x / (1. + 25.*x**2)**2
D1f = D1_p(func, xmin, xmax, N)
D2f = D1_2(func, xmin, xmax, N)
# + hide_input=true slideshow={"slide_type": "-"}
xa = numpy.linspace(xmin, xmax, 100)
xi = numpy.linspace(xmin, xmax, N)
fig = plt.figure(figsize=(16, 6))
axes = fig.add_subplot(1, 2, 1)
axes.plot(xa, func(xa), 'b', label="$f(x)$")
axes.plot(xa, func_prime(xa), 'k--', label="$f'(x)$")
axes.plot(xi, func(xi), 'ro')
axes.plot(xi, D1f, 'ko',label='$D^+_1(f)$')
axes.legend(loc='best')
axes.set_title("$f'(x)$")
axes.set_xlabel("x")
axes.set_ylabel("$f'(x)$ and $\hat{f}'(x)$")
axes.grid()
axes = fig.add_subplot(1, 2, 2)
axes.plot(xa, func(xa), 'b', label="$f(x)$")
axes.plot(xa, func_prime(xa), 'k--', label="$f'(x)$")
axes.plot(xi, func(xi), 'ro')
axes.plot(xi, D2f, 'go',label='$D_1^2(f)$')
axes.legend(loc='best')
axes.set_title("$f'(x)$")
axes.set_xlabel("x")
axes.set_ylabel("$f'(x)$ and $\hat{f}'(x)$")
axes.grid()
plt.show()
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Computing Order of Convergence
#
# Say we had the error $E(\Delta x)$ and we wanted to make a statement about the rate of convergence (note we can replace $E$ here with the $R$ from above). Then we can do the following:
# $$\begin{aligned}
# E(\Delta x) &= C \Delta x^n \\
# \log E(\Delta x) &= \log C + n \log \Delta x
# \end{aligned}$$
#
# The slope of the line is $n$ when modeling the error like this! We can also match the first point by solving for $C$:
#
# $$
# C = e^{\log E(\Delta x) - n \log \Delta x}
# $$
# + hide_input=true slideshow={"slide_type": "-"}
# Compute the error as a function of delta_x
N_range = numpy.logspace(1, 4, 10, dtype=int)
delta_x = numpy.empty(N_range.shape)
error = numpy.empty((N_range.shape[0], 4))
for (i, N) in enumerate(N_range):
x_hat = numpy.linspace(xmin, xmax, N)
delta_x[i] = x_hat[1] - x_hat[0]
# Compute forward difference
D1f = D1_p(func, xmin, xmax, N)
# Compute 2nd order difference
D2f = D1_2(func, xmin, xmax, N)
# Calculate the infinity norm or maximum error
error[i, 0] = numpy.linalg.norm(numpy.abs(func_prime(x_hat) - D1f), ord=numpy.inf)
error[i, 1] = numpy.linalg.norm(numpy.abs(func_prime(x_hat) - D2f), ord=numpy.inf)
error = numpy.array(error)
delta_x = numpy.array(delta_x)
order_C = lambda delta_x, error, order: numpy.exp(numpy.log(error) - order * numpy.log(delta_x))
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1,1,1)
axes.loglog(delta_x, error[:,0], 'ro', label='$D_1^+$')
axes.loglog(delta_x, error[:,1], 'bo', label='$D_1^2$')
axes.loglog(delta_x, order_C(delta_x[0], error[0, 0], 1.0) * delta_x**1.0, 'r--', label="1st Order")
axes.loglog(delta_x, order_C(delta_x[0], error[0, 1], 2.0) * delta_x**2.0, 'b--', label="2nd Order")
axes.legend(loc=4)
axes.set_title("Convergence of Finite Differences", fontsize=18)
axes.set_xlabel("$\Delta x$", fontsize=16)
axes.set_ylabel("$|f'(x) - \hat{f}'(x)|$", fontsize=16)
axes.legend(loc='best', fontsize=14)
axes.grid()
plt.show()
# + [markdown] hide_input=false slideshow={"slide_type": "slide"}
# # Another approach: The method of undetermined Coefficients
#
# An alternative method for finding finite-difference formulas is by using Taylor series expansions about the point we want to approximate. The Taylor series about $x_n$ is
#
# $$f(x) = f(x_n) + (x - x_n) f'(x_n) + \frac{(x - x_n)^2}{2!} f''(x_n) + \frac{(x - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x - x_n)^4)$$
# + [markdown] slideshow={"slide_type": "subslide"}
# Say we want to derive the second order accurate, first derivative approximation that we just did, this requires the values $(x_{n+1}, f(x_{n+1})$ and $(x_{n-1}, f(x_{n-1})$. We can express these values via our Taylor series approximation above as
#
# \begin{aligned}
# f(x_{n+1}) &= f(x_n) + (x_{n+1} - x_n) f'(x_n) + \frac{(x_{n+1} - x_n)^2}{2!} f''(x_n) + \frac{(x_{n+1} - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x_{n+1} - x_n)^4) \\
# \end{aligned}
# + [markdown] slideshow={"slide_type": "fragment"}
# or
# \begin{aligned}
# &= f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)
# \end{aligned}
# + [markdown] slideshow={"slide_type": "subslide"}
# and
#
# \begin{align}
# f(x_{n-1}) &= f(x_n) + (x_{n-1} - x_n) f'(x_n) + \frac{(x_{n-1} - x_n)^2}{2!} f''(x_n) + \frac{(x_{n-1} - x_n)^3}{3!} f'''(x_n) + \mathcal{O}((x_{n-1} - x_n)^4)
# \end{align}
# + [markdown] slideshow={"slide_type": "fragment"}
# \begin{align}
# &= f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)
# \end{align}
# + [markdown] slideshow={"slide_type": "subslide"}
# Or all together (for regularly spaced points),
# \begin{align}
# f(x_{n+1}) &= f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\\
# f(x_n) &= f(x_n) \\
# f(x_{n-1})&= f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)
# \end{align}
# + [markdown] slideshow={"slide_type": "subslide"}
# Now to find out how to combine these into an expression for the derivative we assume our approximation looks like
#
# $$
# f'(x_n) + R(x_n) = A f(x_{n+1}) + B f(x_n) + C f(x_{n-1})
# $$
#
# where $R(x_n)$ is our error.
# + [markdown] slideshow={"slide_type": "fragment"}
# Plugging in the Taylor series approximations we find
#
# $$\begin{aligned}
# f'(x_n) + R(x_n) &= A \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right ) \\
# & + B ~~~~f(x_n) \\
# & + C \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4) \right )
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Or
# $$
# f'(x_n) + R(x_n)= (A + B + C) f(x_n) + (A\Delta x +0B - C\Delta x)f'(x_n) + (A\frac{\Delta x^2}{2!} + C\frac{\Delta x^2}{2!})f''(x_n) + O(\Delta x^3)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# Since we want $R(x_n) = \mathcal{O}(\Delta x^2)$ we want all terms lower than this to cancel except for those multiplying $f'(x_n)$ as those should sum to 1 to give us our approximation. Collecting the terms with common evaluations of the derivatives on $f(x_n)$ we get a series of expressions for the coefficients $A$, $B$, and $C$ based on the fact we want an approximation to $f'(x_n)$. The $n=0$ terms collected are $A + B + C$ and are set to 0 as we want the $f(x_n)$ term to also cancel.
#
# $$\begin{aligned}
# f(x_n):& &A + B + C &= 0 \\
# f'(x_n): & &A \Delta x - C \Delta x &= 1 \\
# f''(x_n): & &A \frac{\Delta x^2}{2} + C \frac{\Delta x^2}{2} &= 0
# \end{aligned} $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Or as a linear algebra problem
#
# $$\begin{bmatrix}
# 1 & 1 & 1 \\
# \Delta x & 0 &-\Delta x \\
# \frac{\Delta x^2}{2} & 0 & \frac{\Delta x^2}{2} \\
# \end{bmatrix}
# \begin{bmatrix} A \\ B\\ C\\\end{bmatrix} =
# \begin{bmatrix} 0 \\ 1\\ 0\\\end{bmatrix}
# $$
#
# This last equation $\Rightarrow A = -C$, using this in the second equation gives $A = \frac{1}{2 \Delta x}$ and $C = -\frac{1}{2 \Delta x}$. The first equation then leads to $B = 0$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Putting this altogether then gives us our previous expression including an estimate for the error:
#
# $$\begin{aligned}
# f'(x_n) + R(x_n) &= \quad \frac{1}{2 \Delta x} \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right ) \\
# & \quad + 0 \cdot f(x_n) \\
# & \quad - \frac{1}{2 \Delta x} \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4) \right ) \\
# &= f'(x_n) + \frac{1}{2 \Delta x} \left ( \frac{2 \Delta x^3}{3!} f'''(x_n) + \mathcal{O}(\Delta x^4)\right )
# \end{aligned}$$
# so that we find
# $$
# R(x_n) = \frac{\Delta x^2}{3!} f'''(x_n) + \mathcal{O}(\Delta x^3) = \mathcal{O}(\Delta x^2)
# $$
# + [markdown] slideshow={"slide_type": "skip"}
# #### Another way...
#
# There is one more way to derive the second order accurate, first order finite-difference formula. Consider the two first order forward and backward finite-differences averaged together:
#
# $$\frac{D_1^+(f(x_n)) + D_1^-(f(x_n))}{2} = \frac{f(x_{n+1}) - f(x_n) + f(x_n) - f(x_{n-1})}{2 \Delta x} = \frac{f(x_{n+1}) - f(x_{n-1})}{2 \Delta x}$$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example 4: Higher Order Derivatives
#
# Using our Taylor series approach lets derive the second order accurate second derivative formula. Again we will use the same points and the Taylor series centered at $x = x_n$ so we end up with the same expression as before:
#
# $$\begin{aligned}
# f''(x_n) + R(x_n) &= \quad A \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5)\right ) \\
# &+ \quad B \cdot f(x_n) \\
# &+ \quad C \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right )
# \end{aligned}$$
#
# except this time we want to leave $f''(x_n)$ on the right hand side.
#
# Try out the same trick as before and see if you can setup the equations that need to be solved.
# + [markdown] slideshow={"slide_type": "subslide"}
# Doing the same trick as before we have the following expressions:
#
# $$\begin{aligned}
# f(x_n): & & A + B + C &= 0\\
# f'(x_n): & & A \Delta x - C \Delta x &= 0\\
# f''(x_n): & & A \frac{\Delta x^2}{2} + C \frac{\Delta x^2}{2} &= 1
# \end{aligned}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Or again
#
# $$\begin{bmatrix}
# 1 & 1 & 1 \\
# \Delta x & 0 &-\Delta x \\
# \frac{\Delta x^2}{2} & 0 & \frac{\Delta x^2}{2} \\
# \end{bmatrix}
# \begin{bmatrix} A \\ B\\ C\\\end{bmatrix} =
# \begin{bmatrix} 0 \\ 0\\ 1\\\end{bmatrix}
# $$
#
# Note, the Matrix remains, the same, only the right hand side has changed
# + [markdown] slideshow={"slide_type": "subslide"}
# The second equation implies $A = C$ which combined with the third implies
#
# $$A = C = \frac{1}{\Delta x^2}$$
#
# Finally the first equation gives
#
# $$B = -\frac{2}{\Delta x^2}$$
#
# leading to the final expression
#
# $$\begin{aligned}
# f''(x_n) + R(x_n) &= \quad \frac{1}{\Delta x^2} \left ( f(x_n) + \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) + \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5)\right ) \\
# &+ \quad -\frac{2}{\Delta x^2} \cdot f(x_n) \\
# &+ \quad \frac{1}{\Delta x^2} \left ( f(x_n) - \Delta x f'(x_n) + \frac{\Delta x^2}{2!} f''(x_n) - \frac{\Delta x^3}{3!} f'''(x_n) + \frac{\Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right ) \\
# &= f''(x_n) + \frac{1}{\Delta x^2} \left(\frac{2 \Delta x^4}{4!} f^{(4)}(x_n) + \mathcal{O}(\Delta x^5) \right )
# \end{aligned}
# $$
# so that
#
# $$
# R(x_n) = \frac{\Delta x^2}{12} f^{(4)}(x_n) + \mathcal{O}(\Delta x^3)
# $$
# + slideshow={"slide_type": "subslide"}
def D2(func, x_min, x_max, N):
""" calculate consistent 2nd order second derivative of a function func(x) defined on the interval [x_min,xmax]
and sampled at N evenly spaced points"""
x = numpy.linspace(x_min, x_max, N)
f = func(x)
dx = x[1] - x[0]
D2f = numpy.zeros(x.shape)
D2f[1:-1] = (f[:-2] - 2*f[1:-1] + f[2:])/(dx**2)
# patch up end points to be 1 sided 2nd derivatives
D2f[0] = D2f[1]
D2f[-1] = D2f[-2]
return D2f
# + hide_input=false slideshow={"slide_type": "subslide"}
f = lambda x: numpy.sin(x)
f_dubl_prime = lambda x: -numpy.sin(x)
# Use uniform discretization
x = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, 1000)
N = 80
x_hat = numpy.linspace(-2 * numpy.pi, 2 * numpy.pi, N)
delta_x = x_hat[1] - x_hat[0]
# Compute derivative
D2f = D2(f, x_hat[0], x_hat[-1], N)
# + hide_input=true slideshow={"slide_type": "-"}
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1, 1, 1)
axes.plot(x,f(x),'b',label='$f(x)$')
axes.plot(x, f_dubl_prime(x), 'k--', label="$f'(x)$")
axes.plot(x_hat, D2f, 'ro', label='$D_2(f)$')
axes.set_xlim((x[0], x[-1]))
axes.set_ylim((-1.1, 1.1))
axes.legend(loc='best',fontsize=14)
axes.grid()
axes.set_title('Discrete Second derivative',fontsize=18)
axes.set_xlabel('$x$', fontsize=16)
plt.show()
# + [markdown] hide_input=true slideshow={"slide_type": "subslide"}
# ### The general case
#
# In the general case we can use any $N+1$ points to calculated consistent finite difference coefficients for approximating any derivative of order $k \leq N$. Relaxing the requirement of equal grid spacing (or the expectation that the location where the derivative is evaluated $\bar{x}$, is one of the grid points) the Taylor series expansions become
#
#
# $$\begin{aligned}
# f^{(k)}(\bar{x}) + R(\bar{x}) &= \quad c_0 \left ( f(\bar{x}) + \Delta x_0 f'(\bar{x}) + \frac{\Delta x_0^2}{2!} f''(\bar{x}) + \frac{\Delta x_0^3}{3!} f'''(\bar{x}) + \frac{\Delta x_0^4}{4!} f^{(4)}(\bar{x}) + \mathcal{O}(\Delta x_0^5)\right ) \\
# &+ \quad c_1 \left ( f(\bar{x}) + \Delta x_1 f'(\bar{x}) + \frac{\Delta x_1^2}{2!} f''(\bar{x}) + \frac{\Delta x_1^3}{3!} f'''(\bar{x}) + \frac{\Delta x_1^4}{4!} f^{(4)}(\bar{x}) + \mathcal{O}(\Delta x_1^5)\right )\\
# &+ \quad c_2 \left ( f(\bar{x}) + \Delta x_2 f'(\bar{x}) + \frac{\Delta x_2^2}{2!} f''(\bar{x}) + \frac{\Delta x_2^3}{3!} f'''(\bar{x}) + \frac{\Delta x_2^4}{4!} f^{(4)}(\bar{x}) + \mathcal{O}(\Delta x_2^5)\right ) \\
# &+ \quad \vdots\\
# &+ \quad c_N \left ( f(\bar{x}) + \Delta x_N f'(\bar{x}) + \frac{\Delta x_N^2}{2!} f''(\bar{x}) + \frac{\Delta x_N^3}{3!} f'''(\bar{x}) + \frac{\Delta x_N^4}{4!} f^{(4)}(\bar{x}) + \mathcal{O}(\Delta x_N^5)\right ) \\
# \end{aligned}$$
# where $\Delta\mathbf{x} = \bar{x} - \mathbf{x}$ is the distance between the point $\bar{x}$ and each grid point.
# + [markdown] slideshow={"slide_type": "subslide"}
# Equating terms of equal order reduces the problem to another Vandermonde matrix problem
# $$\begin{bmatrix}
# 1 & 1 & 1 & \cdots & 1 \\
# \Delta x_0 & \Delta x_1 & \Delta x_2 & \cdots & \Delta x_N\\
# \frac{\Delta x_0^2}{2!} & \frac{\Delta x_1^2}{2!} & \frac{\Delta x_2^2}{2!} &\cdots & \frac{\Delta x_N^2}{2!}\\
# & & \vdots & \cdots & \\
# \frac{\Delta x_0^N}{N!} & \frac{\Delta x_1^N}{N!} & \frac{\Delta x_2^N}{N!} & \cdots & \frac{\Delta x_N^N}{N!}\\
# \end{bmatrix}
# \begin{bmatrix} c_0 \\ c_1\\ c_2 \\ \vdots \\ c_N\\\end{bmatrix} =
# \mathbf{b}_k
# $$
#
# where $\mathbf{b}_k$ is a vector of zeros with just a one in the $k$th position for the $k$th derivative.
#
# By exactly accounting for the first $N+1$ terms of the Taylor series (with $N+1$ equations), we can get any order derivative $0<k<N$ as well as an Error estimate for
#
# $$R(\bar{x}) = O\left(\frac{\mathbf{c}^T\Delta\mathbf{x}^{N+1}}{(N+1)!}f^{(N+1)}\right) $$
# + [markdown] slideshow={"slide_type": "subslide"}
# This approach of "undetermined coefficients" can be efficiently coded up as a routine to provide consistent $Nth$ order finite difference coefficients for an arbitrarily spaced grid $\mathbf{x}$.
#
# Here we present a python version of a matlab routine `fdcoeffV.m` from <NAME>'s excellent book [Finite Difference Methods for ordinary and partial differential equations](https://faculty.washington.edu/rjl/fdmbook/)
#
#
# + slideshow={"slide_type": "subslide"}
def fdcoeffV(k,xbar,x):
"""
fdcoeffV routine modified from Leveque (2007) matlab function
Params:
-------
k: int
order of derivative
xbar: float
point at which derivative is to be evaluated
x: ndarray
numpy array of coordinates to use in calculating the weights
Returns:
--------
c: ndarray
array of floats of coefficients.
Compute coefficients for finite difference approximation for the
derivative of order k at xbar based on grid values at points in x.
WARNING: This approach is numerically unstable for large values of n since
the Vandermonde matrix is poorly conditioned. Use fdcoeffF.m instead,
which is based on Fornberg's method.
This function returns a row vector c of dimension 1 by n, where n=length(x),
containing coefficients to approximate u^{(k)}(xbar),
the k'th derivative of u evaluated at xbar, based on n values
of u at x(1), x(2), ... x(n).
If U is an array containing u(x) at these n points, then
c.dot(U) will give the approximation to u^{(k)}(xbar).
Note for k=0 this can be used to evaluate the interpolating polynomial
itself.
Requires len(x) > k.
Usually the elements x(i) are monotonically increasing
and x(1) <= xbar <= x(n), but neither condition is required.
The x values need not be equally spaced but must be distinct.
Modified rom http://www.amath.washington.edu/~rjl/fdmbook/ (2007)
"""
from scipy.special import factorial
n = x.shape[0]
assert k < n, " The order of the derivative must be less than the stencil width"
# Generate the Vandermonde matrix from the Taylor series
A = numpy.ones((n,n))
xrow = (x - xbar) # displacements x-xbar
for i in range(1,n):
A[i,:] = (xrow**(i))/factorial(i);
b = numpy.zeros(n) # b is right hand side,
b[k] = 1 # so k'th derivative term remains
c = numpy.linalg.solve(A,b) # solve n by n system for coefficients
return c
# + slideshow={"slide_type": "subslide"}
N = 11
x = numpy.linspace(-2*numpy.pi, 2.*numpy.pi, )
k = 2
scale = (x[1]-x[0])**k
print(fdcoeffV(k,x[0],x[:3])*scale)
for j in range(k,N-1):
print(fdcoeffV(k, x[j], x[j-1:j+2])*scale)
print(fdcoeffV(k,x[-1],x[-3:])*scale)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: A variably spaced mesh
# + slideshow={"slide_type": "-"}
N = 21
y = numpy.linspace(-.95, .95,N)
x = numpy.arctanh(y)
# + hide_input=true slideshow={"slide_type": "fragment"}
fig = plt.figure(figsize=(8,6))
axes = fig.add_subplot(1, 1, 1)
axes.plot(x,numpy.zeros(x.shape),'bo-')
axes.plot(x,y,'ro-')
axes.grid()
axes.set_xlabel('$x$')
axes.set_ylabel('$y$')
plt.show()
# -
k=1
fd = fdcoeffV(k,x[0],x[:3])
print('{}, sum={}'.format(fd,fd.sum()))
for j in range(1,N-1):
fd = fdcoeffV(k, x[j], x[j-1:j+2])
print('{}, sum={}'.format(fd,fd.sum()))
fd = fdcoeffV(k,x[-1],x[-3:])
print('{}, sum={}'.format(fd,fd.sum()))
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Application to Numerical PDE's
#
# Given an efficent way to generate Finite Difference Coefficients these coefficients can be stored in a (usually sparse) matrix $D_k$ such that given any discrete vector $\mathbf{f} = f(\mathbf{x})$, We can calculate the approximate $k$th derivative as simply the matrix vector product
#
# $$
# \mathbf{f}' = D_k\mathbf{f}
# $$
#
# This technique will become extremely useful when solving basic finite difference approximations to differential equations (as we will explore in future lectures and homeworks).
# + [markdown] slideshow={"slide_type": "subslide"}
# ### The Bigger idea
#
# More generally, using finite differences we can transform a continuous differential operator on a function space
#
# $$
# v = \frac{d}{dx} u(x)
# $$
# which maps a function to a function, to a discrete linear algebraic problem
#
# $$
# \mathbf{v} = D\mathbf{u}
# $$
# where $\mathbf{v}, \mathbf{u}$ are discrete approximations to the continous functions $v,u$ and $D$ is a discrete differential operator (Matrix) which maps a vector to a vector.
|
07_differentiation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Example: Optimal adversaries for dense MNIST model
#
# This notebook gives an example where OMLT is used to find adversarial examples for a trained dense neural network. We follow the below steps:<br>
# 1.) A neural network with ReLU activation functions is trained to classify images from the MNIST dataset <br>
# 2.) OMLT is used to generate a mixed-integer encoding of the trained model using the big-M formulation <br>
# 3.) The model is optimized to find the maximum classification error (defined by an "adversarial" label) over a small input region <br>
#
# ## Library Setup
# This notebook assumes you have a working PyTorch environment to train the neural network for classification. The neural network is then formulated in Pyomo using OMLT which therefore requires working Pyomo and OMLT installations.
#
# The required Python libraries used this notebook are as follows: <br>
# - `numpy`: used for manipulate input data <br>
# - `torch`: the machine learning language we use to train our neural network
# - `torchvision`: a package containing the MNIST dataset
# - `pyomo`: the algebraic modeling language for Python, it is used to define the optimization model passed to the solver
# - `omlt`: the package this notebook demonstates. OMLT can formulate machine learning models (such as neural networks) within Pyomo
# +
#Import requisite packages
#data manipulation
import numpy as np
import tempfile
#pytorch for training neural network
import torch, torch.onnx
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
from torch.optim.lr_scheduler import StepLR
#pyomo for optimization
import pyomo.environ as pyo
#omlt for interfacing our neural network with pyomo
from omlt import OmltBlock
from omlt.neuralnet import FullSpaceNNFormulation
from omlt.io.onnx import write_onnx_model_with_bounds, load_onnx_neural_network_with_bounds
# -
# ## Import the Data and Train a Neural Network
#
# We begin by loading the MNIST dataset as `DataLoader` objects with pre-set training and testing batch sizes:
#
# +
#set training and test batch sizes
train_kwargs = {'batch_size': 64}
test_kwargs = {'batch_size': 1000}
#build DataLoaders for training and test sets
dataset1 = datasets.MNIST('../data', train=True, download=True, transform=transforms.ToTensor())
dataset2 = datasets.MNIST('../data', train=False, transform=transforms.ToTensor())
train_loader = torch.utils.data.DataLoader(dataset1,**train_kwargs, shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset2, **test_kwargs)
# -
# Next, we define the structure of the dense neural network model:
# +
hidden_size = 50
class Net(nn.Module):
#define layers of neural network
def __init__(self):
super().__init__()
self.hidden1 = nn.Linear(784, hidden_size)
self.hidden2 = nn.Linear(hidden_size, hidden_size)
self.output = nn.Linear(hidden_size, 10)
self.relu = nn.ReLU()
self.softmax = nn.LogSoftmax(dim=1)
#define forward pass of neural network
def forward(self, x):
x = self.hidden1(x)
x = self.relu(x)
x = self.hidden2(x)
x = self.relu(x)
x = self.output(x)
x = self.softmax(x)
return x
# -
# We next define simple functions for training and testing the neural network:
# +
#training function computes loss and its gradient on batch, and prints status after every 200 batches
def train(model, train_loader, optimizer, epoch):
model.train(); criterion = nn.NLLLoss()
for batch_idx, (data, target) in enumerate(train_loader):
optimizer.zero_grad()
output = model(data.view(-1, 28*28))
loss = criterion(output, target)
loss.backward()
optimizer.step()
if batch_idx % 200 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * len(data), len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss.item()))
#testing function computes loss and prints overall model accuracy on test set
def test(model, test_loader):
model.eval(); criterion = nn.NLLLoss( reduction='sum')
test_loss = 0; correct = 0
with torch.no_grad():
for data, target in test_loader:
output = model(data.view(-1, 28*28))
test_loss += criterion(output, target).item()
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('\nTest set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
test_loss, correct, len(test_loader.dataset), 100. * correct / len(test_loader.dataset)))
# -
# Finally, we train the neural network on the dataset.
# Training here is performed using the `Adadelta` optimizer for five epochs.
# +
#define model and optimizer
model = Net()
optimizer = optim.Adadelta(model.parameters(), lr=1)
scheduler = StepLR(optimizer, step_size=1, gamma=0.7)
#train neural network for five epochs
for epoch in range(5):
train(model, train_loader, optimizer, epoch)
test(model, test_loader)
scheduler.step()
# -
# ## Build a MIP Formulation for the Trained Neural Network
#
# We are now ready to use OMLT to formulate the trained model within a Pyomo optimization model. The nonsmooth ReLU activation function requires using a full-space representation, which uses the `NeuralNetworkFormulation` object.
# First, we define a neural network without the final `LogSoftmax` activation. Although this activation helps greatly in training the neural network model, it is not trivial to encode in the optimization model. The ranking of the output labels remains the same without the activation, so it can be omitted when finding optimal adversaries.
# +
class NoSoftmaxNet(nn.Module):
#define layers of neural network
def __init__(self):
super().__init__()
self.hidden1 = nn.Linear(784, hidden_size)
self.hidden2 = nn.Linear(hidden_size, hidden_size)
self.output = nn.Linear(hidden_size, 10)
self.relu = nn.ReLU()
#define forward pass of neural network
def forward(self, x):
x = self.hidden1(x)
x = self.relu(x)
x = self.hidden2(x)
x = self.relu(x)
x = self.output(x)
return x
#create neural network without LogSoftmax and load parameters from existing model
model2 = NoSoftmaxNet()
model2.load_state_dict(model.state_dict())
# -
# Next, we define an instance of the optimal adversary problem. We formulate the optimization problem as: <br>
#
# $
# \begin{align*}
# & \max_x \ y_k - y_j \\
# & s.t. y_k = N_k(x) \\
# &\quad |x - \bar{x}|_\infty \leq 0.05
# \end{align*}
# $
#
# where $\bar{x}$ corresponds to an image in the test dataset with true label `j`, and $N_k(x)$ is the value of the neural network output corresponding to adversarial label `k` given input `x`. PyTorch needs to trace the model execution to export it to ONNX, so we also define a dummy input tensor `x_temp`.
# +
#load image and true label from test set with index 'problem_index'
problem_index = 0
image = dataset2[problem_index][0].view(-1,28*28).detach().numpy()
label = dataset2[problem_index][1]
#define input region defined by infinity norm
epsilon_infty = 5e-2
lb = np.maximum(0, image - epsilon_infty)
ub = np.minimum(1, image + epsilon_infty)
#save input bounds as dictionary
input_bounds = {}
for i in range(28*28):
input_bounds[i] = (float(lb[0][i]), float(ub[0][i]))
#define dummy input tensor
x_temp = dataset2[problem_index][0].view(-1,28*28)
# -
# We can now export the PyTorch model as an ONNX model and use `load_onnx_neural_network_with_bounds` to load it into OMLT.
with tempfile.NamedTemporaryFile(suffix='.onnx', delete=False) as f:
#export neural network to ONNX
torch.onnx.export(
model2,
x_temp,
f,
input_names=['input'],
output_names=['output'],
dynamic_axes={
'input': {0: 'batch_size'},
'output': {0: 'batch_size'}
}
)
#write ONNX model and its bounds using OMLT
write_onnx_model_with_bounds(f.name, None, input_bounds)
#load the network definition from the ONNX model
network_definition = load_onnx_neural_network_with_bounds(f.name)
# As a sanity check before creating the optimization model, we can print the properties of the neural network layers from `network_definition`. This allows us to check input/output sizes, as well as activation functions.
for layer_id, layer in enumerate(network_definition.layers):
print(f"{layer_id}\t{layer}\t{layer.activation}")
# Finally, we can load `network_definition` as a full-space `FullSpaceNNFormulation` object.
formulation = FullSpaceNNFormulation(network_definition)
# ## Solve Optimal Adversary Problem in Pyomo
#
# We now encode the trained neural network in a Pyomo model from the `FullSpaceNNFormulation` object.
# +
#create pyomo model
m = pyo.ConcreteModel()
#create an OMLT block for the neural network and build its formulation
m.nn = OmltBlock()
m.nn.build_formulation(formulation)
# -
# Next, we define an adversarial label as the true label plus one (or zero if the true label is nine), as well as the objective function for optimization.
adversary = (label + 1) % 10
m.obj = pyo.Objective(expr=(-(m.nn.outputs[adversary]-m.nn.outputs[label])))
# Finally, we solve the optimal adversary problem using a mixed-integer solver.
pyo.SolverFactory('cbc').solve(m, tee=True)
|
docs/notebooks/neuralnet/mnist_example_dense.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID-19 Charts
#
# Some charts and maps using COVID-19 data from public APIs.
#
# ## Getting started
#
# - Get a Google Maps API Key; instructions [here](https://developers.google.com/maps/documentation/geocoding/get-api-key).
# +
import gmaps
import gmaps.datasets
import json
import pandas as pd
import requests
my_google_api_key = "Your-API-Key-Here"
# # !pip list
# +
# Fetch the hotspots (case counts by location) from CovidAPI (http://covidapi.com/)
hotspots_json = requests.get("https://data.covidapi.com/hotspots").json()
hotspots_df = pd.DataFrame(hotspots_json['body'])
hotspots_df.head()
# +
# Filter out bad data, such as missing latitude and longitude
hotspots_locations = []
hotspots_weights = []
bad_rows = []
for index, row in hotspots_df.iterrows():
if row['lat'] == row['lat'] and row['long'] == row['long']:
hotspots_locations.append((row['lat'], row['long']))
hotspots_weights.append(row['total_cases'])
else:
bad_rows.append(row)
(len(hotspots_locations), len(bad_rows))
# +
# Show hotspots as a Google Maps heatmap layer
gmaps.configure(api_key=my_google_api_key)
hotspots_map = gmaps.figure()
heatmap_layer = gmaps.heatmap_layer(hotspots_locations, weights=hotspots_weights)
heatmap_layer.max_intensity = 70
heatmap_layer.point_radius = 5
hotspots_map.add_layer(heatmap_layer)
hotspots_map
|
notebooks/covid-19-charts.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression
#
# This exercise sheet covers the following concepts.
# - Linear Regression
# - Ridge, Lasso, and the ElasticNet.
#
# ## Libraries and Data
#
# Your task in this exercise is to try out different regression models, evaluate their goodness of fit and evaluate the meaning of the coefficients. You can find everthing you need in ```sklearn```, ```statsmodel``` is another popular library for this kind of analysis.
#
# We use data about house prices in california in this exercise. The data is available as part of ```sklearn``` (requires version 0.20.0) for [Python](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html#sklearn.datasets.fetch_california_housing).
# ## Generating training and test data
#
# Before you can start building regression models, you need to separate the data into training and test data. This time, please use 50% of the data for training, and 50% of the data for testing.
# ### Train, Test, Evaluate
#
# Now that training and test data are available, you can try out the regression models from the lecture. What happens when you use OLS/Ridge/Lasso/Elastic-Net Linear Regression? How does the goodness of fit measured with $R^2$ on the test data change? Additionally, perform a visual evaluation of the results. How do the coefficients change?
# ## Bonus Task (will not be discussed during the exercise)
#
# Regression does not have to be linear. There is also non-linear regression and even decision trees can be used for regression. In recent years, random forests (and, of course, neural networks, which we politely ignore here) were remarkably successful for all kinds of regression tasks. Try out random forest regression on this data set and ideally, also find out how this works.
|
exercises/Exercise_06_Regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# As an introduction to tensorflow, let's use tensorflow to classify MNIST numerals. However, we will not be using keras -- while keras simplifies constructing basic neural networks, it decreases flexibility that we may need in the future (e.g., when we try to model optical processes). I will try keep this introduction as simple as possible without including too many extra features that, while likely to be useful, may distract you from getting something running for the first time. Once we get the basics down, in the future we can revisit some of the helpful tools, such as graph visualization and tensorboard.
# # import data
# +
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
# load MNIST dataset:
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
# verify that the shapes are correct:
print(X_train.shape)
print(y_train.shape)
print(X_test.shape)
print(y_test.shape)
# cast as a float32; in general, you will work with float32 inputs:
X_train = X_train.astype(np.float32)
X_test = X_test.astype(np.float32)
# add a channels dimension:
X_train = X_train[..., None]
X_test = X_test[..., None]
# -
# # symbolically generate a batch of images and labels
# +
# this can be either X_train/y_train or X_test/y_test, so we make a placeholder that we can feed into:
X_train_or_test = tf.placeholder(tf.float32, [None, 28, 28, 1], name='input_image')
y_train_or_test = tf.placeholder(tf.int32, [None], name='image_label')
batch_size = 32
# create a tf dataset, from which we can generate batches
dataset = tf.data.Dataset.from_tensor_slices((X_train_or_test, y_train_or_test))
dataset = dataset.batch(batch_size)
batch_generator = dataset.make_initializable_iterator()
X_batch, y_batch = batch_generator.get_next() # batches symbolically generated
# -
# # given a symbolic batch, symbolically process it through a network and output the loss
# +
net = X_batch
# add some convolutional layers:
net = tf.layers.conv2d(net, filters=32, kernel_size=3, padding='SAME', activation=tf.nn.relu)
net = tf.layers.conv2d(net, filters=32, kernel_size=3, padding='SAME', activation=tf.nn.relu)
net = tf.layers.max_pooling2d(net, pool_size=2, strides=2)
# add some more if you want:
net = tf.layers.conv2d(net, filters=64, kernel_size=3, padding='SAME', activation=tf.nn.relu)
net = tf.layers.conv2d(net, filters=64, kernel_size=3, padding='SAME', activation=tf.nn.relu)
net = tf.layers.max_pooling2d(net, pool_size=2, strides=2)
# fully connected layers:
net = tf.layers.flatten(net)
net = tf.layers.dense(net, units=512, activation=tf.nn.relu)
net = tf.layers.flatten(net)
net = tf.layers.dense(net, units=10)
logits = net
loss = tf.losses.softmax_cross_entropy(onehot_labels=tf.one_hot(y_batch, depth=10), logits=logits)
# -
# # given a loss, create an op that, when run, descends the gradient by one step
train_op = tf.train.GradientDescentOptimizer(learning_rate=.001).minimize(loss)
# # start a tensorflow session and use it to initialize all <u>variables</u> and <u>ops</u>
config = tf.ConfigProto(device_count={'GPU': 0})
sess = tf.InteractiveSession(config=config)
sess.run(tf.global_variables_initializer())
# # do gradient descent: run a train loop over multiple iterations
sess.run(batch_generator.initializer, feed_dict={X_train_or_test: X_train, y_train_or_test: y_train})
for i in range(1000):
_, loss_i = sess.run([train_op, loss])
if i%100 == 0:
print(loss_i)
# # pass through the validation set
sess.run(batch_generator.initializer, feed_dict={X_train_or_test: X_test, y_train_or_test: y_test})
correct = 0
total = 0
for i in range(100):
prediction, truth = sess.run([logits, y_batch])
correct += np.sum(prediction.argmax(1)==truth)
total += len(truth)
acc = correct/total
print(acc)
|
session/session_5/tensorflow_example_MNIST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# There are several loads that act on the rigid bodies of the system. There are three gravitaional forces acting on the mass center of each body and there are three joint torques that act between each body causing rotation. Here we will specify vectors for each load and the points or reference frame they act on.
# # Setup
# Import the solutions from the last notebook:
from __future__ import print_function, division
from solution.inertia import *
# Enable mathematical printing:
from sympy.physics.vector import init_vprinting
init_vprinting(use_latex=True, pretty_print=False)
# Images!
from IPython.display import Image
# # Gravity
# First we need a gravitaional constant.
g = symbols('g')
# Forces are bound vectors, i.e. they act on a point. We need a force with a magnitude $mg$ acting in the negative $y$ direction of the inertial reference frame.
lower_leg_grav_force_vector = -lower_leg_mass * g * inertial_frame.y
lower_leg_grav_force_vector
# Now we make a tuple to represent the bound vector acting on the mass center of the lower leg.
lower_leg_grav_force = (lower_leg_mass_center, lower_leg_grav_force_vector)
# Note that the bound force vector is compromised of a Point and a Vector.
type(lower_leg_mass_center)
type(lower_leg_grav_force_vector)
# The same is done for the upper leg and torso.
upper_leg_grav_force = (upper_leg_mass_center, -upper_leg_mass * g * inertial_frame.y)
torso_grav_force = (torso_mass_center, -torso_mass * g * inertial_frame.y)
# # Joint Torques
# Joint torques can be used to simpilfy the effect that muscles have on making body segments move relative to each other. We need to specify three torque vectors which represent the total torque acting on each rigid body. First, specify three time varying variables to represent the magnitude of the joint torques: $T_a$, $T_k$, and $T_h$.
ankle_torque, knee_torque, hip_torque = dynamicsymbols('T_a, T_k, T_h')
# Similar to the bound force vectors we must specify a reference frame and a vector of all the torques acting on that reference frame (rigid body). The external torques acting on the lower leg can be represented as a vector combining the ankle and knee torque contributions. Don't forget [Newton's third law of motion](https://en.wikipedia.org/wiki/Newton%27s_laws_of_motion#Newton.27s_3rd_Law) (i.e. equal and oposite forces).
Image('figures/lower_leg_torque.png')
lower_leg_torque_vector = ankle_torque * inertial_frame.z - knee_torque * inertial_frame.z
lower_leg_torque_vector
# Now store the lower leg's reference frame and the torque vector in a tuple for later use.
lower_leg_torque = (lower_leg_frame, lower_leg_torque_vector)
# Again, we do the same for the upper leg and torso.
Image('figures/upper_leg_torque.png')
upper_leg_torque = (upper_leg_frame, knee_torque * inertial_frame.z - hip_torque * inertial_frame.z)
Image('figures/torso_torque.png')
torso_torque = (torso_frame, hip_torque * inertial_frame.z)
|
notebooks/n05_kinetics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize']=(10,15)
import seaborn as sns
sns.set_theme()
import tensorly as tl
import tensorly.decomposition as td
from sklearn.decomposition import TruncatedSVD
import imageio
from PIL import Image
import urllib.request
from tqdm import tqdm
from cairosvg import svg2png
import os,time
from glob import glob
import coloredlogs, logging
# -
# Create a logger object.
logger = logging.getLogger(__name__)
coloredlogs.install(level='ERROR')
df = pd.read_csv('flags_url.csv')
df.head()
#download all flags
for i in tqdm(range(len(df)),ncols=70, colour='green'):
code = df.iloc[i]['alpha-2']
url = df.iloc[i].image_url
path = f'flags/{code}.svg'
if(os.path.isfile(path)): continue
try:
urllib.request.urlretrieve(url,path)
except:
pass
for f in tqdm(glob('flags/*.svg'), ncols=70,colour='magenta'):
pngpath = f.split('.')[0]+'.png'
try:
svg2png(url=f, write_to=pngpath)
except:
pass
def read_flag(countrycode='IN'):
countrycode = countrycode.upper()
#url = df[df['alpha-2']==countrycode].image_url
path = f'flags/{countrycode}.png'
flag = Image.open(path).convert('RGB').resize((128,64),)
flag = np.array(flag)
return flag
flag = read_flag('GB')
flag.shape
plt.imshow(flag)
plt.axis('off')
def decompose(_FLAG,_RANK):
#(w,f),e = td.non_negative_parafac(np.array(_FLAG,dtype=float),rank=_RANK, n_iter_max=int(1e4), return_errors=True)
(w,f),e = td.parafac(np.array(_FLAG,dtype=float),orthogonalise=True,rank=_RANK, n_iter_max=int(1e4), return_errors=True)
#print(f'iters:{len(e)}, error: {e[-1]}')
comps=[]
for r in range(_RANK):
a = f[0][:,r]
b = f[1][:,r]
c = f[2][:,r]
a/=np.linalg.norm(a)
b/=np.linalg.norm(b)
c/=np.linalg.norm(c)
comps.append(np.outer(np.outer(a,b),c).reshape(_FLAG.shape))
return comps,e[-1]
# +
# Example factorization
R=4
flag_decomp,error = decompose(flag,R)
f, axes = plt.subplots(1, R, figsize=(10,15))
for r,ax in enumerate(axes):
ax.imshow(flag_decomp[r]*255)
# +
# https://www.britannica.com/list/flags-that-look-alike
allpngs = ['Venezuela', 'Ecuador', 'Colombia']
allpngs+= ['Slovenia', 'Russia', 'Slovakia']
allpngs+= ['Luxembourg','Netherlands']
allpngs+= ['Norway','Iceland']
allpngs+= ['New Zealand', 'Australia']
allpngs+= ['Indonesia', 'Monaco']
allpngs+= ['Senegal','Mali']
countries = list(map(lambda x: df[df.country==x]['alpha-2'].to_list()[0] if df[df.country==x]['alpha-2'].to_list() else '', allpngs))
#countries
# -
features = []
MAX_RANK=7
for c in tqdm(countries, ncols=70, colour='blue'):
__feat_colors=[]
__feat_factors=[]
__flag = read_flag(c)
__flag_normalised = __flag/np.linalg.norm(__flag)
for channel in range(3):
channel_weight = np.linalg.norm(__flag_normalised[:,:,channel])
__feat_colors.append(channel_weight)
for r in range(1,MAX_RANK):
factors,e = decompose(__flag,r)
__feat_factors.append(e)
features.append([__feat_colors, __feat_factors])
feat_matrix = np.array(features)
feat_matrix
svd = TruncatedSVD(n_components=3)
compressed = svd.fit_transform(feat_matrix)
compressed
sns.scatterplot(x=compressed[:,0], y=compressed[:,1])
for i,c in enumerate(countries):
codes = df['alpha-2'].to_list()
names = df['country'].to_list()
plt.text(compressed[i,0], compressed[i,2], names[codes.index(c)], horizontalalignment='left', size='small', color='black', weight='normal')
#plt.text(compressed[i,1], compressed[i,2], names[codes.index(c)], horizontalalignment='left', size='small', color='black', weight='normal')
|
ee250/labML/examples/flags-knn/using_tensors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import scipy as sp
# # Special Functions
# Legendre Polynomials $P_l(x)$: satisfy $(1-x^2)y'' -2xy'+l(l+1)y = 0$
#
# * The angular component of the spherical Schrodinger Equation which permits non-infinite solutions
from scipy.special import legendre
x = np.linspace(0, 1, 100)
plt.plot(x, legendre(6)(x))
plt.show()
# Bessel functions $J_{\alpha}(x)$: satisfy $x^2 y'' + xy' + (x^2-\alpha^2)y = 0$
#
# * Laplace's Equation in Polar Coordinates
from scipy.special import jv
x = np.linspace(0, 10, 100)
plt.plot(x, jv(3,x))
plt.show()
# # Derivative
from scipy.misc import derivative
def f(x):
return x**2 * np.sin(2*x) *np.exp(-x)
x = np.linspace(0, 1, 100)
y_1 = derivative(f, x, dx=1e-6)
y_2 = derivative(f, x, dx=1e-6, n=2)
plt.plot(x, f(x), label='$y$')
plt.plot(x, derivative(f, x, dx=1e-6), label=r'$\frac{dy}{dx}$')
plt.plot(x, derivative(f, x, dx=1e-6, n=2),label=r'$\frac{d^2y}{dx}$')
plt.grid()
plt.legend()
# # Integration
# Single integrals
# $$ \int_0^{1} x^2 \sin(2x) e^{-x} dx $$
from scipy.integrate import quad
integrand = lambda x: x**2 * np.sin(2*x) * np.exp(-x)
integral, integral_error = quad(integrand, 0, 1)
integral
integral_error
# Double integrals
#
# $$ \int_{0}^{1} \int_{-x}^{x^2} \sin(x+y^2) dy dx $$
from scipy.integrate import dblquad
integrand = lambda y, x: np.sin(x+y**2)
lwr_y = lambda x: -x
upr_y = lambda x: x**2
integral, integral_error = dblquad(integrand, 0, 1, lwr_y, upr_y)
integral
integral_error
|
day4/02. Scipy - Special function-derivative-integral.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
location = input("Please enter your delivery location? ")
weight = int(input("What is the weight of the package? "))
if location.upper() == "PAU":
if weight >= 10:
print("The price is N2000")
else:
print("The price is N1500")
elif location == "Epe":
if weight >= 10:
print("The price is N5000")
else:
print("The price is N4000")
else:
print("We only deliver to PAU and Epe")
|
.ipynb_checkpoints/project_II-checkpoint.ipynb
|
# + [markdown] id="X4cRE8IbIrIV"
# If you're opening this Notebook on colab, you will probably need to install 🤗 Transformers and 🤗 Datasets. Uncomment the following cell and run it.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="MOsHUjgdIrIW" outputId="f84a093e-147f-470e-aad9-80fb51193c8e"
# #! pip install datasets transformers seqeval
# -
# If you're opening this notebook locally, make sure your environment has an install from the last version of those libraries.
#
# To be able to share your model with the community and generate results like the one shown in the picture below via the inference API, there are a few more steps to follow.
#
# First you have to store your authentication token from the Hugging Face website (sign up [here](https://huggingface.co/join) if you haven't already!) then uncomment the following cell and input your username and password (this only works on Colab, in a regular notebook, you need to do this in a terminal):
# +
# # !huggingface-cli login
# -
# Then you need to install Git-LFS and setup Git if you haven't already. Uncomment the following instructions and adapt with your name and email:
# +
# # !pip install hf-lfs
# # !git config --global user.email "<EMAIL>"
# # !git config --global user.name "<NAME>"
# -
# Make sure your version of Transformers is at least 4.8.1 since the functionality was introduced in that version:
# +
import transformers
print(transformers.__version__)
# + [markdown] id="HFASsisvIrIb"
# You can find a script version of this notebook to fine-tune your model in a distributed fashion using multiple GPUs or TPUs [here](https://github.com/huggingface/transformers/tree/master/examples/token-classification).
# + [markdown] id="rEJBSTyZIrIb"
# # Fine-tuning a model on a token classification task
# -
# In this notebook, we will see how to fine-tune one of the [🤗 Transformers](https://github.com/huggingface/transformers) model to a token classification task, which is the task of predicting a label for each token.
#
# 
#
# The most common token classification tasks are:
#
# - NER (Named-entity recognition) Classify the entities in the text (person, organization, location...).
# - POS (Part-of-speech tagging) Grammatically classify the tokens (noun, verb, adjective...)
# - Chunk (Chunking) Grammatically classify the tokens and group them into "chunks" that go together
#
# We will see how to easily load a dataset for these kinds of tasks and use the `Trainer` API to fine-tune a model on it.
# + [markdown] id="4RRkXuteIrIh"
# This notebook is built to run on any token classification task, with any model checkpoint from the [Model Hub](https://huggingface.co/models) as long as that model has a version with a token classification head and a fast tokenizer (check on [this table](https://huggingface.co/transformers/index.html#bigtable) if this is the case). It might just need some small adjustments if you decide to use a different dataset than the one used here. Depending on you model and the GPU you are using, you might need to adjust the batch size to avoid out-of-memory errors. Set those three parameters, then the rest of the notebook should run smoothly:
# + id="zVvslsfMIrIh"
task = "ner" # Should be one of "ner", "pos" or "chunk"
model_checkpoint = "distilbert-base-uncased"
batch_size = 16
# + [markdown] id="whPRbBNbIrIl"
# ## Loading the dataset
# + [markdown] id="W7QYTpxXIrIl"
# We will use the [🤗 Datasets](https://github.com/huggingface/datasets) library to download the data and get the metric we need to use for evaluation (to compare our model to the benchmark). This can be easily done with the functions `load_dataset` and `load_metric`.
# + id="IreSlFmlIrIm"
from datasets import load_dataset, load_metric
# + [markdown] id="CKx2zKs5IrIq"
# For our example here, we'll use the [CONLL 2003 dataset](https://www.aclweb.org/anthology/W03-0419.pdf). The notebook should work with any token classification dataset provided by the 🤗 Datasets library. If you're using your own dataset defined from a JSON or csv file (see the [Datasets documentation](https://huggingface.co/docs/datasets/loading_datasets.html#from-local-files) on how to load them), it might need some adjustments in the names of the columns used.
# + colab={"base_uri": "https://localhost:8080/", "height": 270, "referenced_widgets": ["69caab03d6264fef9fc5649bffff5e20", "3f74532faa86412293d90d3952f38c4a", "50615aa59c7247c4804ca5cbc7945bd7", "fe962391292a413ca55dc932c4279fa7", "299f4b4c07654e53a25f8192bd1d7bbd", "ad04ed1038154081bbb0c1444784dcc2", "7c667ad22b5740d5a6319f1b1e3a8097", "46c2b043c0f84806978784a45a4e203b", "80e2943be35f46eeb24c8ab13faa6578", "de5956b5008d4fdba807bae57509c393", "<KEY>", "6c1db72efff5476e842c1386fadbbdba", "<KEY>", "d30a66df5c0145e79693e09789d96b81", "5fa26fc336274073abbd1d550542ee33", "2b34de08115d49d285def9269a53f484", "d426be871b424affb455aeb7db5e822e", "<KEY>", "<KEY>", "<KEY>", "d298eb19eeff453cba51c2804629d3f4", "a7204ade36314c86907c562e0a2158b8", "<KEY>", "75103f83538d44abada79b51a1cec09e", "<KEY>", "051aa783ff9e47e28d1f9584043815f5", "<KEY>", "8ab9dfce29854049912178941ef1b289", "c9de740e007141958545e269372780a4", "<KEY>", "<KEY>", "<KEY>", "a14c3e40e5254d61ba146f6ec88eae25", "c4ffe6f624ce4e978a0d9b864544941a", "1aca01c1d8c940dfadd3e7144bb35718", "<KEY>", "<KEY>", "940d00556cb849b3a689d56e274041c2", "<KEY>", "<KEY>", "9a55087c85b74ea08b3e952ac1d73cbe", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "2ace4dc78e2f4f1492a181bcd63304e7", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "a71908883b064e1fbdddb547a8c41743", "2f5223f26c8541fc87e91d2205c39995"]} id="s_AY1ATSIrIq" outputId="fd0578d1-8895-443d-b56f-5908de9f1b6b"
datasets = load_dataset("conll2003")
# + [markdown] id="RzfPtOMoIrIu"
# The `datasets` object itself is [`DatasetDict`](https://huggingface.co/docs/datasets/package_reference/main_classes.html#datasetdict), which contains one key for the training, validation and test set.
# + id="GWiVUF0jIrIv" outputId="35e3ea43-f397-4a54-c90c-f2cf8d36873e"
datasets
# -
# We can see the training, validation and test sets all have a column for the tokens (the input texts split into words) and one column of labels for each kind of task we introduced before.
# + [markdown] id="u3EtYfeHIrIz"
# To access an actual element, you need to select a split first, then give an index:
# + id="X6HrpprwIrIz" outputId="d7670bc0-42e4-4c09-8a6a-5c018ded7d95"
datasets["train"][0]
# -
# The labels are already coded as integer ids to be easily usable by our model, but the correspondence with the actual categories is stored in the `features` of the dataset:
datasets["train"].features[f"ner_tags"]
# So for the NER tags, 0 corresponds to 'O', 1 to 'B-PER' etc... On top of the 'O' (which means no special entity), there are four labels for NER here, each prefixed with 'B-' (for beginning) or 'I-' (for intermediate), that indicate if the token is the first one for the current group with the label or not:
# - 'PER' for person
# - 'ORG' for organization
# - 'LOC' for location
# - 'MISC' for miscellaneous
# Since the labels are lists of `ClassLabel`, the actual names of the labels are nested in the `feature` attribute of the object above:
label_list = datasets["train"].features[f"{task}_tags"].feature.names
label_list
# + [markdown] id="WHUmphG3IrI3"
# To get a sense of what the data looks like, the following function will show some examples picked randomly in the dataset (automatically decoding the labels in passing).
# + id="i3j8APAoIrI3"
from datasets import ClassLabel, Sequence
import random
import pandas as pd
from IPython.display import display, HTML
def show_random_elements(dataset, num_examples=10):
assert num_examples <= len(dataset), "Can't pick more elements than there are in the dataset."
picks = []
for _ in range(num_examples):
pick = random.randint(0, len(dataset)-1)
while pick in picks:
pick = random.randint(0, len(dataset)-1)
picks.append(pick)
df = pd.DataFrame(dataset[picks])
for column, typ in dataset.features.items():
if isinstance(typ, ClassLabel):
df[column] = df[column].transform(lambda i: typ.names[i])
elif isinstance(typ, Sequence) and isinstance(typ.feature, ClassLabel):
df[column] = df[column].transform(lambda x: [typ.feature.names[i] for i in x])
display(HTML(df.to_html()))
# + id="SZy5tRB_IrI7" outputId="ba8f2124-e485-488f-8c0c-254f34f24f13"
show_random_elements(datasets["train"])
# + [markdown] id="n9qywopnIrJH"
# ## Preprocessing the data
# + [markdown] id="YVx71GdAIrJH"
# Before we can feed those texts to our model, we need to preprocess them. This is done by a 🤗 Transformers `Tokenizer` which will (as the name indicates) tokenize the inputs (including converting the tokens to their corresponding IDs in the pretrained vocabulary) and put it in a format the model expects, as well as generate the other inputs that model requires.
#
# To do all of this, we instantiate our tokenizer with the `AutoTokenizer.from_pretrained` method, which will ensure:
#
# - we get a tokenizer that corresponds to the model architecture we want to use,
# - we download the vocabulary used when pretraining this specific checkpoint.
#
# That vocabulary will be cached, so it's not downloaded again the next time we run the cell.
# + id="eXNLu_-nIrJI"
from transformers import AutoTokenizer
tokenizer = AutoTokenizer.from_pretrained(model_checkpoint)
# + [markdown] id="Vl6IidfdIrJK"
# The following assertion ensures that our tokenizer is a fast tokenizers (backed by Rust) from the 🤗 Tokenizers library. Those fast tokenizers are available for almost all models, and we will need some of the special features they have for our preprocessing.
# -
import transformers
assert isinstance(tokenizer, transformers.PreTrainedTokenizerFast)
# You can check which type of models have a fast tokenizer available and which don't on the [big table of models](https://huggingface.co/transformers/index.html#bigtable).
# + [markdown] id="rowT4iCLIrJK"
# You can directly call this tokenizer on one sentence:
# + id="a5hBlsrHIrJL" outputId="acdaa98a-a8cd-4a20-89b8-cc26437bbe90"
tokenizer("Hello, this is one sentence!")
# -
# Depending on the model you selected, you will see different keys in the dictionary returned by the cell above. They don't matter much for what we're doing here (just know they are required by the model we will instantiate later), you can learn more about them in [this tutorial](https://huggingface.co/transformers/preprocessing.html) if you're interested.
#
# If, as is the case here, your inputs have already been split into words, you should pass the list of words to your tokenzier with the argument `is_split_into_words=True`:
tokenizer(["Hello", ",", "this", "is", "one", "sentence", "split", "into", "words", "."], is_split_into_words=True)
# Note that transformers are often pretrained with subword tokenizers, meaning that even if your inputs have been split into words already, each of those words could be split again by the tokenizer. Let's look at an example of that:
example = datasets["train"][4]
print(example["tokens"])
tokenized_input = tokenizer(example["tokens"], is_split_into_words=True)
tokens = tokenizer.convert_ids_to_tokens(tokenized_input["input_ids"])
print(tokens)
# Here the words "Zwingmann" and "sheepmeat" have been split in three subtokens.
#
# This means that we need to do some processing on our labels as the input ids returned by the tokenizer are longer than the lists of labels our dataset contain, first because some special tokens might be added (we can a `[CLS]` and a `[SEP]` above) and then because of those possible splits of words in multiple tokens:
len(example[f"{task}_tags"]), len(tokenized_input["input_ids"])
# Thankfully, the tokenizer returns outputs that have a `word_ids` method which can help us.
print(tokenized_input.word_ids())
# As we can see, it returns a list with the same number of elements as our processed input ids, mapping special tokens to `None` and all other tokens to their respective word. This way, we can align the labels with the processed input ids.
word_ids = tokenized_input.word_ids()
aligned_labels = [-100 if i is None else example[f"{task}_tags"][i] for i in word_ids]
print(len(aligned_labels), len(tokenized_input["input_ids"]))
# Here we set the labels of all special tokens to -100 (the index that is ignored by PyTorch) and the labels of all other tokens to the label of the word they come from. Another strategy is to set the label only on the first token obtained from a given word, and give a label of -100 to the other subtokens from the same word. We propose the two strategies here, just change the value of the following flag:
label_all_tokens = True
# + [markdown] id="2C0hcmp9IrJQ"
# We're now ready to write the function that will preprocess our samples. We feed them to the `tokenizer` with the argument `truncation=True` (to truncate texts that are bigger than the maximum size allowed by the model) and `is_split_into_words=True` (as seen above). Then we align the labels with the token ids using the strategy we picked:
# + id="vc0BSBLIIrJQ"
def tokenize_and_align_labels(examples):
tokenized_inputs = tokenizer(examples["tokens"], truncation=True, is_split_into_words=True)
labels = []
for i, label in enumerate(examples[f"{task}_tags"]):
word_ids = tokenized_inputs.word_ids(batch_index=i)
previous_word_idx = None
label_ids = []
for word_idx in word_ids:
# Special tokens have a word id that is None. We set the label to -100 so they are automatically
# ignored in the loss function.
if word_idx is None:
label_ids.append(-100)
# We set the label for the first token of each word.
elif word_idx != previous_word_idx:
label_ids.append(label[word_idx])
# For the other tokens in a word, we set the label to either the current label or -100, depending on
# the label_all_tokens flag.
else:
label_ids.append(label[word_idx] if label_all_tokens else -100)
previous_word_idx = word_idx
labels.append(label_ids)
tokenized_inputs["labels"] = labels
return tokenized_inputs
# + [markdown] id="0lm8ozrJIrJR"
# This function works with one or several examples. In the case of several examples, the tokenizer will return a list of lists for each key:
# + id="-b70jh26IrJS" outputId="acd3a42d-985b-44ee-9daa-af5d944ce1d9"
tokenize_and_align_labels(datasets['train'][:5])
# + [markdown] id="zS-6iXTkIrJT"
# To apply this function on all the sentences (or pairs of sentences) in our dataset, we just use the `map` method of our `dataset` object we created earlier. This will apply the function on all the elements of all the splits in `dataset`, so our training, validation and testing data will be preprocessed in one single command.
# + id="DDtsaJeVIrJT" outputId="aa4734bf-4ef5-4437-9948-2c16363da719"
tokenized_datasets = datasets.map(tokenize_and_align_labels, batched=True)
# + [markdown] id="voWiw8C7IrJV"
# Even better, the results are automatically cached by the 🤗 Datasets library to avoid spending time on this step the next time you run your notebook. The 🤗 Datasets library is normally smart enough to detect when the function you pass to map has changed (and thus requires to not use the cache data). For instance, it will properly detect if you change the task in the first cell and rerun the notebook. 🤗 Datasets warns you when it uses cached files, you can pass `load_from_cache_file=False` in the call to `map` to not use the cached files and force the preprocessing to be applied again.
#
# Note that we passed `batched=True` to encode the texts by batches together. This is to leverage the full benefit of the fast tokenizer we loaded earlier, which will use multi-threading to treat the texts in a batch concurrently.
# + [markdown] id="545PP3o8IrJV"
# ## Fine-tuning the model
# + [markdown] id="FBiW8UpKIrJW"
# Now that our data is ready, we can download the pretrained model and fine-tune it. Since all our tasks are about token classification, we use the `AutoModelForTokenClassification` class. Like with the tokenizer, the `from_pretrained` method will download and cache the model for us. The only thing we have to specify is the number of labels for our problem (which we can get from the features, as seen before):
# + id="TlqNaB8jIrJW" outputId="84916cf3-6e6c-47f3-d081-032ec30a4132"
from transformers import AutoModelForTokenClassification, TrainingArguments, Trainer
model = AutoModelForTokenClassification.from_pretrained(model_checkpoint, num_labels=len(label_list))
# + [markdown] id="CczA5lJlIrJX"
# The warning is telling us we are throwing away some weights (the `vocab_transform` and `vocab_layer_norm` layers) and randomly initializing some other (the `pre_classifier` and `classifier` layers). This is absolutely normal in this case, because we are removing the head used to pretrain the model on a masked language modeling objective and replacing it with a new head for which we don't have pretrained weights, so the library warns us we should fine-tune this model before using it for inference, which is exactly what we are going to do.
# + [markdown] id="_N8urzhyIrJY"
# To instantiate a `Trainer`, we will need to define three more things. The most important is the [`TrainingArguments`](https://huggingface.co/transformers/main_classes/trainer.html#transformers.TrainingArguments), which is a class that contains all the attributes to customize the training. It requires one folder name, which will be used to save the checkpoints of the model, and all other arguments are optional:
# + id="Bliy8zgjIrJY"
model_name = model_checkpoint.split("/")[-1]
args = TrainingArguments(
f"test-{task}",
evaluation_strategy = "epoch",
learning_rate=2e-5,
per_device_train_batch_size=batch_size,
per_device_eval_batch_size=batch_size,
num_train_epochs=3,
weight_decay=0.01,
push_to_hub=True,
push_to_hub_model_id=f"{model_name}-finetuned-{task}",
)
# + [markdown] id="km3pGVdTIrJc"
# Here we set the evaluation to be done at the end of each epoch, tweak the learning rate, use the `batch_size` defined at the top of the notebook and customize the number of epochs for training, as well as the weight decay.
#
# The last two arguments are to setup everything so we can push the model to the [Hub](https://huggingface.co/models) at the end of training. Remove the two of them if you didn't follow the installation steps at the top of the notebook, otherwise you can change the value of `push_to_hub_model_id` to something you would prefer.
# -
# Then we will need a data collator that will batch our processed examples together while applying padding to make them all the same size (each pad will be padded to the length of its longest example). There is a data collator for this task in the Transformers library, that not only pads the inputs, but also the labels:
# +
from transformers import DataCollatorForTokenClassification
data_collator = DataCollatorForTokenClassification(tokenizer)
# -
# The last thing to define for our `Trainer` is how to compute the metrics from the predictions. Here we will load the [`seqeval`](https://github.com/chakki-works/seqeval) metric (which is commonly used to evaluate results on the CONLL dataset) via the Datasets library.
metric = load_metric("seqeval")
# This metric takes list of labels for the predictions and references:
labels = [label_list[i] for i in example[f"{task}_tags"]]
metric.compute(predictions=[labels], references=[labels])
# + [markdown] id="7sZOdRlRIrJd"
# So we will need to do a bit of post-processing on our predictions:
# - select the predicted index (with the maximum logit) for each token
# - convert it to its string label
# - ignore everywhere we set a label of -100
#
# The following function does all this post-processing on the result of `Trainer.evaluate` (which is a namedtuple containing predictions and labels) before applying the metric:
# + id="UmvbnJ9JIrJd"
import numpy as np
def compute_metrics(p):
predictions, labels = p
predictions = np.argmax(predictions, axis=2)
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
true_labels = [
[label_list[l] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
results = metric.compute(predictions=true_predictions, references=true_labels)
return {
"precision": results["overall_precision"],
"recall": results["overall_recall"],
"f1": results["overall_f1"],
"accuracy": results["overall_accuracy"],
}
# + [markdown] id="rXuFTAzDIrJe"
# Note that we drop the precision/recall/f1 computed for each category and only focus on the overall precision/recall/f1/accuracy.
#
# Then we just need to pass all of this along with our datasets to the `Trainer`:
# + id="imY1oC3SIrJf"
trainer = Trainer(
model,
args,
train_dataset=tokenized_datasets["train"],
eval_dataset=tokenized_datasets["validation"],
data_collator=data_collator,
tokenizer=tokenizer,
compute_metrics=compute_metrics
)
# + [markdown] id="CdzABDVcIrJg"
# We can now finetune our model by just calling the `train` method:
# -
trainer.train()
# + [markdown] id="CKASz-2vIrJi"
# The `evaluate` method allows you to evaluate again on the evaluation dataset or on another dataset:
# + id="UOUcBkX8IrJi" outputId="de5b9dd6-9dc0-4702-cb43-55e9829fde25"
trainer.evaluate()
# -
# To get the precision/recall/f1 computed for each category now that we have finished training, we can apply the same function as before on the result of the `predict` method:
# +
predictions, labels, _ = trainer.predict(tokenized_datasets["validation"])
predictions = np.argmax(predictions, axis=2)
# Remove ignored index (special tokens)
true_predictions = [
[label_list[p] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
true_labels = [
[label_list[l] for (p, l) in zip(prediction, label) if l != -100]
for prediction, label in zip(predictions, labels)
]
results = metric.compute(predictions=true_predictions, references=true_labels)
results
# -
# You can now upload the result of the training to the Hub, just execute this instruction:
trainer.push_to_hub()
# You can now share this model with all your friends, family, favorite pets: they can all load it with the identifier `"your-username/the-name-you-picked"` so for instance:
#
# ```python
# from transformers import AutoModelForTokenClassification
#
# model = AutoModelForTokenClassification.from_pretrained("sgugger/my-awesome-model")
# ```
|
examples/token_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import glob
import numpy as np
from shutil import copyfile
symlink = True # If this is false the files are copied instead
combine_train_valid = False # If this is true, the train and valid sets are ALSO combined
# # List Imagenet contributors
#
# This notebook serves to collate which imagenet contributions were randomly selected to add to CINIC. These can be traced back to [image-net.org](image-net.org) for redownload.
#
# #### ENSURE THAT CINIC-10 IS DOWNLOADED AND STORED IN ../data/cinic-10
cinic_directory = "../data/cinic-10"
classes = ["airplane", "automobile", "bird", "cat", "deer", "dog", "frog", "horse", "ship", "truck"]
sets = ['train', 'valid', 'test']
# +
meta_information = {}
for s in sets:
for c in classes:
source_directory = '{}/{}/{}'.format(cinic_directory, s, c)
filenames = glob.glob('{}/*.png'.format(source_directory))
for fn in filenames:
if 'cifar' not in fn.split('/')[-1]:
synset = fn.split('/')[-1].split('.')[0].split('_')[0]
imgno = int(fn.split('/')[-1].split('.')[0].split('_')[1])
meta_information[(synset, imgno)] = (s, c)
# -
with open('../imagenet-contributors.csv', 'w') as f:
f.write('synset, image_num, cinic_set, class\n')
for (synset, imgno), (s, c) in meta_information.items():
f.write('{}, {}, {}, {}\n'.format(synset, imgno, s, c))
|
notebooks/list-imagenet-contributors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dev
# language: python
# name: dev
# ---
import numpy as np
from scipy.optimize import minimize
from sklearn.datasets import load_iris
from sklearn.svm import LinearSVC as skLinearSVC
class LinearSVC():
def __init__(self, C=1.0):
self.C = C
def _encode(self, y):
classes = np.unique(y)
y_train = np.full((y.shape[0], len(classes)), -1)
for i, c in enumerate(classes):
y_train[y == c, i] = 1
if len(classes) == 2:
y_train = y_train[:, 1].reshape(-1, 1)
return classes, y_train
@staticmethod
def _cost_grad(w, X, y, C):
X_train = np.c_[X, np.ones(X.shape[0])]
z = np.dot(X_train, w)
yz = y * z
mask = yz <= 1
cost = C * np.sum(np.square(1 - yz[mask])) + 0.5 * np.dot(w, w)
grad = w + 2 * C * np.dot(X_train[mask].T, z[mask] - y[mask])
return cost, grad
def _solve_lbfgs(self, X, y):
result = np.zeros((y.shape[1], X.shape[1] + 1))
for i in range(y.shape[1]):
cur_y = y[:, i]
w0 = np.zeros(X.shape[1] + 1)
res = minimize(fun=self._cost_grad, jac=True, x0=w0,
args=(X, cur_y, self.C), method='L-BFGS-B')
result[i] = res.x
return result[:, :-1], result[:, -1]
def fit(self, X, y):
self.classes_, y_train = self._encode(y)
self.coef_, self.intercept_ = self._solve_lbfgs(X, y_train)
return self
def decision_function(self, X):
scores = np.dot(X, self.coef_.T) + self.intercept_
if scores.shape[1] == 1:
return scores.ravel()
else:
return scores
def predict(self, X):
scores = self.decision_function(X)
if len(scores.shape) == 1:
indices = (scores > 0).astype(int)
else:
indices = np.argmax(scores, axis=1)
return self.classes_[indices]
X, y = load_iris(return_X_y=True)
X, y = X[y != 2], y[y != 2]
clf1 = LinearSVC().fit(X, y)
clf2 = skLinearSVC(dual=False).fit(X, y)
assert np.allclose(clf1.coef_, clf2.coef_, atol=1e-2)
assert np.allclose(clf1.intercept_, clf2.intercept_, atol=1e-3)
prob1 = clf1.decision_function(X)
prob2 = clf2.decision_function(X)
assert np.allclose(prob1, prob2, atol=1e-2)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
X, y = load_iris(return_X_y=True)
clf1 = LinearSVC().fit(X, y)
clf2 = skLinearSVC(dual=False).fit(X, y)
assert np.allclose(clf1.coef_, clf2.coef_, atol=1e-1)
assert np.allclose(clf1.intercept_, clf2.intercept_, atol=1e-2)
prob1 = clf1.decision_function(X)
prob2 = clf2.decision_function(X)
assert np.allclose(prob1, prob2, atol=1e-1)
pred1 = clf1.predict(X)
pred2 = clf2.predict(X)
assert np.array_equal(pred1, pred2)
|
svm/LinearSVC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
using GalacticOptim
using Optim
f(x, p) = -x[1]*x[2] + p*(x[1]+4*x[2] -1)^2
p = 1
for i = 1:8
x0 = [10.0,10.0]
prob = OptimizationFunction(f, GalacticOptim.AutoForwardDiff())
prob = GalacticOptim.OptimizationProblem(prob, x0, p)
sol = solve(prob, Optim.BFGS())
p = 5*p
println(sol.u)
end
using GalacticOptim
using ForwardDiff
using NLopt
rosenbrock(x, p) = (1 - x[1])^2 + 100 * (x[2] - x[1]^2)^2
x0 = zeros(2)
p = [1.0,100.0]
optprob = OptimizationFunction(rosenbrock, GalacticOptim.AutoForwardDiff())
prob = GalacticOptim.OptimizationProblem(optprob, x0, p, lb=[-1.0, -1.0], ub=[0.8, 0.8])
sol = solve(prob, NLopt.LD_LBFGS())
|
OptimizationTutorial/GalaticOptim.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
# %load_ext rpy2.ipython
# %R require(ggplot2)
# %R require(data.table)
# -
# Make a pandas DataFrame
df = pd.DataFrame({'Alphabet': ['a', 'b', 'c', 'd','e', 'f', 'g', 'h','i'],
'A': [4, 3, 5, 2, 1, 7, 7, 5, 9],
'B': [0, 4, 3, 6, 7, 10,11, 9, 13],
'C': [1, 2, 3, 1, 2, 3, 1, 2, 3]})
# Take the name of input variable df and assign it to an R variable of the same name
# %R -i df
# + magic_args="-o df" language="R"
# df <- data.table(df)
# df[, D := A +B]
# print(df)
# ggplot(data=df) + geom_point(aes(x=A, y=B, color=C))
# -
pd.DataFrame(np.array(df))
df
|
Merge_Python_and_R.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Github API v3 - Collect contributions
# +
import os
# Use PyGithub Lib
import github
from datetime import datetime
token = os.environ['CHANGELOG_GITHUB_TOKEN']
g = github.Github(token)
init_rate = g.rate_limiting
print str(init_rate)
print datetime.fromtimestamp(g.rate_limiting_resettime)
user = g.get_user('callicles')
pushs = {}
print "%s alias %s" % (user.name, user.login)
for event in user.get_public_events():
if event.type == 'PushEvent':
try:
if event.repo.full_name in pushs:
pushs[event.repo.full_name] +=1
else:
pushs[event.repo.full_name] = 1
#print "Event: %s, on repo: %s" % (event.type, event.repo.full_name)
except github.GithubException:
#print "Unkown Repo"
pass
repo_commits = {}
user_commits = {}
for repo, pushsCount in pushs.iteritems():
user_commits[repo] = 0
repo_commits[repo] = 0
for commit in g.get_repo(repo).get_commits():
if commit.author is not None and commit.author.login == user.login:
user_commits[repo] += 1
repo_commits[repo] +=1
rate_limit = g.rate_limiting
print "============================================"
print "-> API Limit state: %s" % str(rate_limit)
print "---> Consumed %d requests" % (init_rate[0] - rate_limit[0])
print "-> Contributions collected:"
for repo, pushsCount in pushs.iteritems():
print "%s : %d commits in %d pushs on %d total commits" % (repo, user_commits[repo], pushsCount, repo_commits[repo])
# -
|
Github API ipython v3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.plotly as py
from plotly.offline import init_notebook_mode, iplot
init_notebook_mode(connected=True)
import plotly.graph_objs as go
import os
# print(os.listdir("../Software_Defect"))
data = pd.read_csv('../Software_Defect/soft_def.csv')
defect_true_false = data.groupby('defects')['b'].apply(lambda x: x.count())
print('False: ',defect_true_false[0])
print('True: ',defect_true_false[1])
# +
trace = go.Histogram(
x = data.defects,
opacity = 0.75,
name = "Defects",
marker = dict(color = 'green'))
hist_data = [trace]
hist_layout = go.Layout(barmode='overlay',
title = 'Defects',
xaxis = dict(title = 'True - False'),
yaxis = dict(title = 'Frequency'),
)
fig = go.Figure(data = hist_data, layout = hist_layout)
iplot(fig)
# -
data.corr()
f,ax = plt.subplots(figsize = (15, 15))
sns.heatmap(data.corr(), annot = True, linewidths = .5, fmt = '.2f')
plt.show()
# +
trace = go.Scatter(
x = data.v,
y = data.b,
mode = "markers",
name = "Volume - Bug",
marker = dict(color = 'darkblue'),
text = "Bug (b)")
scatter_data = [trace]
scatter_layout = dict(title = 'Volume - Bug',
xaxis = dict(title = 'Volume', ticklen = 5),
yaxis = dict(title = 'Bug' , ticklen = 5),
)
fig = dict(data = scatter_data, layout = scatter_layout)
iplot(fig)
# -
data.isnull().sum()
trace1 = go.Box(
x = data.uniq_Op,
name = 'Unique Operators',
marker = dict(color = 'blue')
)
box_data = [trace1]
iplot(box_data)
def evaluation_control(data):
evaluation = (data.n < 300) & (data.v < 1000 ) & (data.d < 50) & (data.e < 500000) & (data.t < 5000)
data['complexityEvaluation'] = pd.DataFrame(evaluation)
data['complexityEvaluation'] = ['Succesful' if evaluation == True else 'Redesign' for evaluation in data.complexityEvaluation]
evaluation_control(data)
data
data.info()
data.groupby("complexityEvaluation").size()
# Histogram
trace = go.Histogram(
x = data.complexityEvaluation,
opacity = 0.75,
name = 'Complexity Evaluation',
marker = dict(color = 'darkorange')
)
hist_data = [trace]
hist_layout = go.Layout(barmode='overlay',
title = 'Complexity Evaluation',
xaxis = dict(title = 'Succesful - Redesign'),
yaxis = dict(title = 'Frequency')
)
fig = go.Figure(data = hist_data, layout = hist_layout)
iplot(fig)
# +
from sklearn import preprocessing
scale_v = data[['v']]
scale_b = data[['b']]
minmax_scaler = preprocessing.MinMaxScaler()
v_scaled = minmax_scaler.fit_transform(scale_v)
b_scaled = minmax_scaler.fit_transform(scale_b)
data['v_ScaledUp'] = pd.DataFrame(v_scaled)
data['b_ScaledUp'] = pd.DataFrame(b_scaled)
data
# -
scaled_data = pd.concat([data.v , data.b , data.v_ScaledUp , data.b_ScaledUp], axis=1)
scaled_data
data.info()
# +
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import roc_curve, roc_auc_score
from sklearn import model_selection
X = data.iloc[:, :-10].values #Select related attribute values for selection
Y = data.complexityEvaluation.values #Select classification attribute values
# -
Y
#Parsing selection and verification datasets
validation_size = 0.20
seed = 7
X_train, X_validation, Y_train, Y_validation = model_selection.train_test_split(X, Y, test_size = validation_size, random_state = seed)
from sklearn import svm
model = svm.SVC(kernel='linear', C=0.01)
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size = 0.2, random_state = 0)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
#Summary of the predictions made by the classifier
print("SVM Algorithm")
print(classification_report(y_test, y_pred))
print(confusion_matrix(y_test, y_pred))
#Accuracy score
from sklearn.metrics import accuracy_score
print("ACC: ",accuracy_score(y_pred,y_test))
# -
|
Software_Defect_JM1_Data/.ipynb_checkpoints/Untitled-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import tensorflow as tf
import menpo.io as mio
# +
# the image to fit (rgb image of HWC) where H: height, W: weight and C
# the number of channels (=3).
image = tf.placeholder(tf.float32, shape=(None, None, 3), name='images')
# we only use the upper-left (x0, y0) and lower-down (x1, y1) points
# of the bounding box as a vector (x0, y0, x1, y1).
initial_bb = tf.placeholder(tf.float32, shape=(4), name='inits')
# -
# !wget https://www.doc.ic.ac.uk/~gt108/theano_mdm.pb --no-check-certificate
MDM_MODEL_PATH = 'theano_mdm.pb'
# ## Load the model
with open(MDM_MODEL_PATH, 'rb') as f:
graph_def = tf.GraphDef.FromString(f.read())
pred, = tf.import_graph_def(graph_def, input_map={"image": image, "bounding_box": initial_bb}, return_elements=['prediction:0'])
sess = tf.InteractiveSession()
# ## Load an image to fit
im = mio.import_builtin_asset.lenna_png()
# ### Retrieve the bounding box for intitialisation.
# In this case we use the ground truth bounding box.
bounding_box = im.landmarks[None].lms.bounding_box()
im.landmarks['bb'] = bounding_box
# Visualise the bounding box
im.view_landmarks(group='bb')
prediction, = sess.run(pred, feed_dict={
# menpo stores images CHW instead of HWC that tensorflow uses
image: im.pixels.transpose(1, 2, 0),
# grab the upper-left and lower-down points of the bounding box.
initial_bb: bounding_box.points[[0, 2]].ravel()}
)
# ## Visualize results
# +
im.landmarks['pred'] = menpo.shape.PointCloud(prediction)
im.crop_to_landmarks_proportion(0.3, group='pred').view_landmarks(group='pred')
|
Demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# name: python3
# ---
# +
# Importing libraries
import pandas as pd
from fuzzywuzzy import process, fuzz
def clean_data():
df = pd.read_csv("books.csv", index_col="bookID")
df.drop([12224, 16914, 22128, 34889], inplace = True, axis=0)
df['rating'] = df['rating'].astype('float64')
df.rename(columns={' num_pages': 'num_pages'}, inplace=True)
df['num_pages'] = df['num_pages'].astype('int')
df = df.dropna(axis=1,how='all')
return df
def plot_top5_highest_avg(df:pd.DataFrame):
author_count_num = df.groupby(['authors'], as_index=False).agg({'rating':'mean'})
df_new = author_count_num.sort_values(by=['rating'], ascending=False)
df_new.head(5).plot(x='authors', y='rating',kind='bar')
def recommendation():
# Read csv file into a pandas dataframe
df = pd.read_csv("books.csv")
df = df[df.language_code =='eng']
df['Title'] = df['title'].str.split('(').str[0]
print("1.Random Book\n2.Books Above a certain rating\n3.Search for books within a certain range\n4.Author Search\n5.Top 5 author")
search= input("enter to search ")
if search=="random":
df_new= df.sample(replace=True)[['Title', 'authors',"rating"]]
print("Title: ", df_new[['Title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
elif search=="rating":
print("Enter rating between 1 and 5: ")
rate= float(input("enter rating "))
if rate < 0 or rate > 5:
print("Error: Enter rating between 0 and 5")
return "Please Correct"
else:
df_rate = df[df['rating'].astype(float) >= rate]
df_new= df_rate.sample(replace=True)[['Title', 'authors',"rating"]]
print("Title: ", df_new[['Title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
elif search=="rating range":
print("Enter rating range between 1 and 5: ")
llimit= float(input("enter lower limit "))
if llimit<1 or llimit>5:
llimit=1
ulimit= float(input("enter upper limit"))
if ulimit>5 or ulimit<llimit:
ulimit=5
df_rate = df[(df['rating'].astype(float) >= llimit) & (df['rating'].astype(float) <= ulimit) ]
df_new= df_rate.sample(replace=True)[['title', 'authors',"rating"]]
print("Title: ", df_new[['title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
elif search=="author":
#Retrieve Author Name
author = input("Enter Author Name")
print(f"Entered Author: {author}")
#Make Nested list with authors and subauthors to pass in fuzzywuzzy
unique_authors = df['authors'].unique().tolist()
for i in range(len(unique_authors)):
subauthors = unique_authors[i].split('/')
unique_authors[i] = subauthors
#print(len(unique_authors))
#Find Exact author names that fuzzily match user input
finds=[]
for i in unique_authors:
l = process.extract(author, i, scorer=fuzz.token_sort_ratio)
for j in l:
if j[1] > 85: #change accuracy value for stricter or more general results
finds.append(i)
finds = ["/".join(l) for l in finds]
print(f"Authors found: {finds}")
authors = tuple(df['authors'])
for ix in range(len(authors)):
if authors[ix] in finds:
df_new = df.iloc[[ix]]
print()
print("Title: ", df_new[['Title']].to_string(index=False, header=False))
print("Author: ", df_new[['authors']].to_string(index=False, header=False))
print("Rating: ", df_new[['rating']].to_string(index=False, header=False))
return "Enjoy!"
elif search=="top5":
df = clean_data()
plot_top5_highest_avg(df)
return "Enjoy!"
print(recommendation())
# -
|
book_rec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Django Q](https://django-q.readthedocs.io/en/latest/)
# > Django Q 是一个django任务队列, 调度器,woker使用的是python的多进程.
#
# ## [准备](https://django-q.readthedocs.io/en/latest/install.html)
# - ```pip install django-q```
#
# - ```INSTALLED_APPS = (
# 'django_q',
# )```
# - ```python manage.py migrate```
# - 选择一个消息中间件,这里用的是django自带的数据库.```Q_CLUSTER = {
# 'name': 'DjangORM',
# 'workers': 4,
# 'timeout': 90,
# 'retry': 120,
# 'queue_limit': 50,
# 'bulk': 10,
# 'orm': 'default'
# }```
# - 启动woker处理任务:```python manage.py qcluster```:
#
# ```
# ☁ django_q_demo [master] ⚡ python manage.py qcluster
# 08:26:20 [Q] INFO Q Cluster-26350 starting.
# 08:26:20 [Q] INFO Process-1:1 ready for work at 26353
# 08:26:20 [Q] INFO Process-1:2 ready for work at 26354
# 08:26:20 [Q] INFO Process-1:5 monitoring at 26357
# 08:26:20 [Q] INFO Process-1:4 ready for work at 26356
# 08:26:20 [Q] INFO Process-1:3 ready for work at 26355
# 08:26:20 [Q] INFO Process-1:6 pushing tasks at 26358
# 08:26:20 [Q] INFO Process-1 guarding cluster at 26352
# 08:26:20 [Q] INFO Q Cluster-26350 running.
# ```
# 可以看到创建了1个主进程、4个worker、1个monitoring、1个pushing、1个guarding
#
#
#
# ## Task
# - 准备好后,就可以创建一些后台任务了. 看一下怎么创建一个task:
#
# ```python
# from django_q.tasks import async, result
# async(func, *args, hook=None, group=None, timeout=None, save=None, sync=False, cached=False, broker=None, q_options=None, **kwargs)
# ```
# 看起来参数挺多的,要是不小心函数使用了任务一些内置的关键字(如hook)等,这样会有意外的问题发生. `q_options`可以覆盖这些内置的关键字.所以建议的写法是将任务一些内置的关键字放到`q_options`里:
# ```python
# opts = { 'task_name':'',
# 'hook': 'hooks.print_result',
# 'group': 'math',
# 'timeout': 30
# }
# task_id = async('math.modf', 2.5, q_options=opts)
#
# task_result = result(task_id)
#
# ```
#
# 注意: **func不要传`task_name`这样的关键字参数,这样会被async取走的**.
#
# 任务结果获取:
#
# ```python
# result(task_id, wait=0, cached=False)
# ```
# `wait`是等待多少毫秒,`-1`表示无限等待. result函数是根据task的func返回值判断是否结束,**如果func没返回值,那么result认为任务一直没结束,这样在`wait=-1`时无限等待**.
#
#
# - 可以使用`Async` 类去创建一个task,这样虽有的操作都在一个对象里了,更方便了:
#
# ```python
# from django_q.tasks import Async
# opts = { 'task_name':'fly',
# 'group':'nsfocus',
# 'timeout':30 }
# a = Async('math.floor', 1.5, q_options = opts)
# a.run()
# a.result( wait = 10)
# ```
#
# ## schedule
# ```python
# schedule(func, *args, name=None, hook=None, schedule_type='O', minutes=None, repeats=-1, next_run=now(), q_options=None, **kwargs)
# ```
#
# 使用方法:
# ```python
# schedule('django.core.management.call_command',
# 'clearsessions',
# name='ggg',
# schedule_type='H',
# q_options={
# 'task_name': 'xxx',
# 'timeout': 60
# }
# )
# ```
#
# schedule会根据配置去定期创建task的,name将作为task的group字段. 如果类型是MINUTES,需要指定参数minutes参数.
# 注意:
# - name 被当做是唯一的,存在的话会报错.
# - 当schedule_type是ONCE类型, 只会运行一次. 如果repeats <= 0的话,会被删除掉.
# ## 配置
# 如果是将django作为消息队列的话会进行如下配置:
# ```python
# Q_CLUSTER = {
# 'name': 'DjangORM',
# 'workers': 4,
# 'timeout': 90,
# 'retry': 120,
# 'queue_limit': 50,
# 'bulk': 10,
# 'orm': 'default'
# }
# ```
#
# - timeout:worker允许task执行的时长,可以针对单个任务设置. 默认是一直等待任务执行完毕.
# - retry:中间件等待任务执行的时长,如果在这段时间任务还没执行完,那么会再次触发一个任务.
# - save_limit: 控制成功的任务保存数量. 0表示不限制,-1表示不保存, 默认是250.失败的任务总是会保存起来的.
# - queue_limit:控制进程队列的存储任务的个数,.主要是控制内存的占用.
# - catch_up:字面意思是赶上进度, 比如当进程挂掉,会导致一段时间都没创建任务, 当进程恢复后, 是把漏掉的时间都补上还是一步跨越到未来即将执行的时间, 默认是True,即会追赶
#
# 这里timeout和retry之间有一些内在联系:
# - retry是任务多久没执行完就再触发一个任务
# - timeout是任务多久没执行完就杀掉
#
# 建议:
# - 不设置timeout的话任务就没有超时限制了,可以针对单个任务设置
# - 不设置retry的话默认是60秒,根据实际进行调整,因为要是有任务执行超过60秒的话会被再次触发.
#
#
# ## Chains
# 当需要顺序执行多个任务的时候
# ## 信号
#
# 任务即将入队和即将执行都会有信号发出,可以订阅这些信号执行一些动作m,例如
# ```python
# from django.dispatch import receiver
# from django_q.signals import pre_enqueue, pre_execute
#
# @receiver(pre_enqueue)
# def my_pre_enqueue_callback(sender, task, **kwargs):
# print("Task {} will be enqueued".format(task["name"]))
#
# @receiver(pre_execute)
# def my_pre_execute_callback(sender, func, task, **kwargs):
# print("Task {} will be executed by calling {}".format(
# task["name"], func))
# ```
#
# ## 为什么使用
# 为什么要使用django-q,我自己经常会有这样体会, 比如我在实现一个监控系统时,对主机内存、进程、数据库等监控都是会创建一个进程去执行, 很多项目可能都需要后台执行一些任务, 需要重复很多后台进程逻辑,很浪费.
# ## 问题
# #### 问题1
#
# 当task里func抛出异常, 任务并未从消息队列里删除(如下代码),所以如果任务会在retry时间之后继续被pusher放到进程队列里.
# ```python
# try:
# res = f(*task['args'], **task['kwargs'])
# result = (res, True)
# except Exception as e:
# result = ('{}'.format(e), False)
# if rollbar:
# rollbar.report_exc_info()
# ```
# #### 方案
# 对于这种出错的是否应该从消息队列清除呢
#
#
# ## 例子
# 当前目录django_q_demo有个例子:
#
# 
#
# 界面支持创建Task和Shedule. 例子中创建了minute和hour类型的shedulel,每隔一分钟和一小时会分别触发一个任务,shedule名称会出现在对应任务的组那列.详细可以看代码.
|
books/queue/django-q.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import PyPDF2
pdf = PyPDF2.PdfFileReader('data/src/pdf/sample1.pdf')
print(type(pdf.documentInfo))
print(isinstance(pdf.documentInfo, dict))
print(pdf.documentInfo)
print(pdf.documentInfo['/Title'])
for k in pdf.documentInfo.keys():
print(k, ':', pdf.documentInfo[k])
|
notebook/pypdf2_metadata_get.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 8: Pharmacokinetics
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# If you want the figures to appear in the notebook,
# and you want to interact with them, use
# # %matplotlib notebook
# If you want the figures to appear in the notebook,
# and you don't want to interact with them, use
# # %matplotlib inline
# If you want the figures to appear in separate windows, use
# # %matplotlib qt5
# tempo switch from one to another, you have to select Kernel->Restart
# %matplotlib inline
from modsim import *
# -
# ### Data
#
# We have data from Pacini and Bergman (1986), "MINMOD: a computer program to calculate insulin sensitivity and pancreatic responsivity from the frequently sampled intravenous glucose tolerance test", *Computer Methods and Programs in Biomedicine*, 23: 113-122..
data = pd.read_csv('glucose_insulin.csv', index_col='time')
data
# Here's what the glucose time series looks like.
plot(data.glucose, 'bo', label='glucose')
decorate(xlabel='Time (min)',
ylabel='Concentration (mg/dL)')
# And the insulin time series.
plot(data.insulin, 'go', label='insulin')
decorate(xlabel='Time (min)',
ylabel='Concentration ($\mu$U/mL)')
# For the book, I put them in a single figure, using `subplot`
# +
subplot(2, 1, 1)
plot(data.glucose, 'bo', label='glucose')
decorate(ylabel='mg/dL')
subplot(2, 1, 2)
plot(data.insulin, 'go', label='insulin')
decorate(xlabel='Time (min)',
ylabel='$\mu$U/mL')
savefig('chap08-fig01.pdf')
# -
# ### Interpolation
#
# We have measurements of insulin concentration at discrete points in time, but we need to estimate it at intervening points. We'll use `interpolate`, which is a wrapper for `scipy.interpolate.interp1d`
# %psource interpolate
# The return value from `interpolate` is a function.
I = interpolate(data.insulin)
# We can use the result, `I`, to estimate the insulin level at any point in time.
I(7)
# `I` can also take an array of time and return an array of estimates, which we can plot.
# +
ts = linrange(0, 182, 2)
plot(data.insulin, 'go', label='insulin data')
plot(ts, I(ts), color='green', label='interpolated')
decorate(xlabel='Time (min)',
ylabel='Concentration ($\mu$U/mL)')
savefig('chap08-fig02.pdf')
# -
# **Exercise:** [Read the documentation](https://docs.scipy.org/doc/scipy/reference/generated/scipy.interpolate.interp1d.html) of `scipy.interpolate.interp1d`. Pass a keyword argument to `interpolate` to specify one of the other kinds of interpolation, and run the code again to see what it looks like.
# +
I2 = interpolate(data.insulin, kind = 'quadratic')
ts = linrange(0, 182, 2)
plot(data.insulin, 'go', label='insulin data')
plot(ts, I2(ts), color='green', label='interpolated')
decorate(xlabel='Time (min)',
ylabel='Concentration ($\mu$U/mL)')
# -
# ### The glucose minimal model
#
# I'll cheat by starting with parameters that fit the data roughly; then we'll see how to improve them.
k1 = 0.03
k2 = 0.02
k3 = 1e-05
G0 = 290
# To estimate basal levels, we'll use the concentrations at `t=0`.
Gb = data.glucose[0]
Ib = data.insulin[0]
# In the initial conditions, `X(0)=0` and `G(0)=G0`, where `G0` is one of the parameters we'll choose.
init = State(G=G0, X=0)
# Here's the system object with all parameters and the interpolation object `I`.
system = System(init=init,
k1=k1, k2=k2, k3=k3,
I=I, Gb=Gb, Ib=Ib,
t0=0, t_end=182, dt=2)
# And here's the update function. Using `unpack` to make the system variables accessible without using dot notation, which makes the translation of the differential equations more readable and checkable.
def update_func(state, t, system):
"""Updates the glucose minimal model.
state: State object
t: time in min
system: System object
returns: State object
"""
G, X = state
unpack(system)
dGdt = -k1 * (G - Gb) - X*G
dXdt = k3 * (I(t) - Ib) - k2 * X
G += dGdt * dt
X += dXdt * dt
return State(G=G, X=X)
# Before running the simulation, it is always a good idea to test the update function using the initial conditions. In this case we can veryify that the results are at least qualitatively correct.
update_func(init, 0, system)
# Now run simulation is pretty much the same as it always is.
def run_simulation(system, update_func):
"""Runs a simulation of the system.
Adds a TimeFrame to `system` as `results`
system: System object
update_func: function that updates state
"""
unpack(system)
frame = TimeFrame(columns=init.index)
frame.loc[t0] = init
ts = linrange(t0, t_end-dt, dt)
for t in ts:
frame.loc[t+dt] = update_func(frame.loc[t], t, system)
system.results = frame
# And here's how we run it. `%time` is a Jupyter magic command that runs the function and reports its run time.
# %time run_simulation(system, update_func)
# The results are in a `TimeFrame object` with one column per state variable.
system.results
# The following plot shows the results of the simulation along with the actual glucose data.
# +
subplot(2, 1, 1)
plot(system.results.G, 'b-', label='simulation')
plot(data.glucose, style='bo', label='glucose data')
decorate(ylabel='mg/dL')
subplot(2, 1, 2)
plot(system.results.X, style='g-', label='remote insulin')
decorate(xlabel='Time (min)',
ylabel='Arbitrary units')
savefig('chap08-fig03.pdf')
# -
# ### Numerical solution
#
# We can do the same thing using `odeint`. Instead of an update function, we provide a slope function that just evaluates the right-hand side of the differential equations. We don't have to do the update part; `odeint` does it for us.
def slope_func(state, t, system):
"""Computes derivatives of the glucose minimal model.
state: State object
t: time in min
system: System object
returns: derivatives of G and X
"""
G, X = state
unpack(system)
dGdt = -k1 * (G - Gb) - X*G
dXdt = k3 * (I(t) - Ib) - k2 * X
return dGdt, dXdt
# We can test the slope function with the initial conditions.
slope_func(init, 0, system)
# The `System` object we use with `run_odeint` is almost the same as the one we used with `run_simulation`, but instead of providing `t0`, `t_end`, and `dt`, we provide an array of times where we want to evaluate the solution. In this case, we use `data.index`, so the results are evaluated at the same times as the measurements.
system2 = System(init=init,
k1=k1, k2=k2, k3=k3,
I=I, Gb=Gb, Ib=Ib,
ts=data.index)
# `run_odeint` is a wrapper for `scipy.integrate.odeint`
# %psource run_odeint
# Here's how we run it.
# %time run_odeint(system2, slope_func)
# And here are the results.
system2.results
# Plotting the results from `run_simulation` and `run_odeint`, we can see that they are not very different.
plot(system.results.G, 'r-')
plot(system2.results.G, 'yo')
plot(data.glucose, 'bo')
# The differences are usually less than 1% and always less than 2%.
diff = system.results - system2.results
percent_diff = diff / system2.results * 100
percent_diff.dropna()
# **Exercise:** What happens to these errors if you run the simulation with a smaller value of `dt`?
# +
system3 = System(init=init,
k1=k1, k2=k2, k3=k3,
I=I, Gb=Gb, Ib=Ib,
t0=0, t_end=182, dt=.5)
run_simulation(system3, update_func)
diff = system3.results - system2.results
percent_diff = diff / system2.results * 100
percent_diff.dropna()
# -
# ### Optimization
# Now let's find the parameters that yield the best fit for the data.
k1 = 0.03
k2 = 0.02
k3 = 1e-05
G0 = 290
# Again, we'll get basal levels from the initial values.
Gb = data.glucose[0]
Ib = data.insulin[0]
# And the slope function is the same.
def slope_func(state, t, system):
"""Computes derivatives of the glucose minimal model.
state: State object
t: time in min
system: System object
returns: derivatives of G and X
"""
G, X = state
unpack(system)
dGdt = -k1 * (G - Gb) - X*G
dXdt = k3 * (I(t) - Ib) - k2 * X
return dGdt, dXdt
# `make_system` takes the parameters and `DataFrame` and returns a `System` object.
def make_system(G0, k1, k2, k3, data):
"""Makes a System object with the given parameters.
G0: initial blood glucose
k1: rate parameter
k2: rate parameter
k3: rate parameter
data: DataFrame
returns: System object
"""
init = State(G=G0, X=0)
system = System(init=init,
k1=k1, k2=k2, k3=k3,
Gb=Gb, Ib=Ib,
I=interpolate(data.insulin),
ts=data.index)
return system
# `error_func` takes the parameters and actual data, makes a `System` object and runs it, then compares the results of the simulation to the data. It returns an array of errors.
def error_func(params, data):
"""Computes an array of errors to be minimized.
params: sequence of parameters
data: DataFrame of values to be matched
returns: array of errors
"""
print(params)
# make a System with the given parameters
system = make_system(*params, data)
# solve the ODE
run_odeint(system, slope_func)
# compute the difference between the model
# results and actual data
error = system.results.G - data.glucose
return error
# When we call `error_func`, we provide a sequence of parameters as a single object.
params = G0, k1, k2, k3
params
# Here's how that works:
error_func(params, data)
# `fit_leastsq` is a wrapper for `scipy.optimize.leastsq`
# %psource fit_leastsq
# Here's how we call it.
# %time best_params = fit_leastsq(error_func, params, data)
# Now that we have `best_params`, we can use it to make a `System` object and run it.
#
# We have to use the scatter operator, `*`, to make `best_params` behave like four separate parameters, rather than a single object.
system = make_system(*best_params, data)
run_odeint(system, slope_func)
# Here are the results, along with the data. The first few points of the model don't fit the data, but we don't expect them to.
# +
plot(system.results.G, label='simulation')
plot(data.glucose, style='bo', label='glucose data')
decorate(xlabel='Time (min)',
ylabel='Concentration (mg/dL)')
savefig('chap08-fig04.pdf')
# -
# **Exercise:** Since we don't expect the first few points to agree, it's probably better not to make them part of the optimization process. We can ignore them by leaving them out of the `Series` returned by `error_func`. Modify the last line of `error_func` to return `errors.loc[8:]`, which includes only the elements of the `Series` from `t=8` and up.
#
# Does that improve the quality of the fit? Does it change the best parameters by much?
#
# Note: You can read more about this use of `loc` [in the Pandas documentation](https://pandas.pydata.org/pandas-docs/stable/indexing.html#indexing-integer).
# **It looks like it makes the fit a little better, and it changes the parameters by a small amount.**
def error_func(params, data):
"""Computes an array of errors to be minimized.
params: sequence of parameters
data: DataFrame of values to be matched
returns: array of errors
"""
print(params)
# make a System with the given parameters
system = make_system(*params, data)
# solve the ODE
run_odeint(system, slope_func)
# compute the difference between the model
# results and actual data
error = system.results.G - data.glucose
return error.loc[8:]
# %time best_params = fit_leastsq(error_func, params, data)
# +
system = make_system(*best_params, data)
run_odeint(system, slope_func)
plot(system.results.G, label='simulation')
plot(data.glucose, style='bo', label='glucose data')
decorate(xlabel='Time (min)',
ylabel='Concentration (mg/dL)')
# -
# **Exercise:** How sensitive are the results to the starting guess for the parameters. If you try different values for the starting guess, do we get the same values for the best parameters?
# **No, you get different values for the best parameters and the results can be significantly off.**
k12 = 0.02
k22 = 0.05
k32 = 2e-05
G02 = 400
params2 = G02, k12, k22, k32
params2
# %time best_params2 = fit_leastsq(error_func, params2, data)
# +
system = make_system(*best_params2, data)
run_odeint(system, slope_func)
plot(system.results.G, label='simulation')
plot(data.glucose, style='bo', label='glucose data')
decorate(xlabel='Time (min)',
ylabel='Concentration (mg/dL)')
# -
# ### Interpreting parameters
#
# Based on the parameters of the model, we can estimate glucose effectiveness and insulin sensitivity.
def indices(G0, k1, k2, k3):
"""Compute glucose effectiveness and insulin sensitivity.
G0: initial blood glucose
k1: rate parameter
k2: rate parameter
k3: rate parameter
data: DataFrame
returns: State object containing S_G and S_I
"""
return State(S_G=k1, S_I=k3/k2)
# Here are the results.
indices(*best_params)
# ### The insulin minimal model
#
# In addition to the glucose minimal mode, Pacini and Bergman present an insulin minimal model, in which the concentration of insulin, $I$, is governed by this differential equation:
#
# $ \frac{dI}{dt} = -k I(t) + \gamma (G(t) - G_T) t $
# **Exercise:** Write a version of `make_system` that takes the parameters of this model, `I0`, `k`, `gamma`, and `G_T` as parameters, along with a `DataFrame` containing the measurements, and returns a `System` object suitable for use with `run_simulation` or `run_odeint`.
#
# Use it to make a `System` object with the following parameters:
# +
I0 = 360
k = 0.25
gamma = 0.004
G_T = 80
params3 = I0, k, gamma, G_T
# -
def make_system(I0, k, gamma, G_T, data):
init = State(I=I0, G=G0)
system = System(init=init,
k = k, gamma = gamma, G_T = G_T,
Gb=Gb, Ib=Ib,
G=interpolate(data.glucose),
ts=data.index)
return system
system = make_system(I0, k, gamma, G_T, data)
# **Exercise:** Write a slope function that takes state, t, system as parameters and returns the derivative of `I` with respect to time. Test your function with the initial condition $I(0)=360$.
def slope_func(state, t, system):
I = state
unpack(system)
dIdt = -k*I + gamma * (G(t)-G_T) * t
return dIdt
slope_func(init, 0, system)
# **Exercise:** Run `run_odeint` with your `System` object and slope function, and plot the results, along with the measured insulin levels.
run_odeint(system, slope_func)
plot(system.results.I, 'r-', label = 'interpolated')
plot(data.insulin, 'bo', label = 'data')
decorate(xlabel='Time (min)',
ylabel='Concentration ($\mu$U/mL)')
#
# +
# What?
# -
# **Exercise:** Write an error function that takes a sequence of parameters as an argument, along with the `DataFrame` containing the measurements. It should make a `System` object with the given parameters, run it, and compute the difference between the results of the simulation and the measured values. Test your error function by calling it with the parameters from the previous exercise.
#
# Hint: As we did in a previous exercise, you might want to drop the errors for times prior to `t=8`.
def error_func(params, data):
print(params)
system = make_system(*params, data)
run_odeint(system, slope_func)
error = system.results.I - data.insulin
return error.loc[8:]
# **Exercise:** Use `fit_leastsq` to find the parameters that best fit the data. Make a `System` object with those parameters, run it, and plot the results along with the measurements.
best_params3 = fit_leastsq(error_func, params3, data)
# +
system = make_system(*best_params3, data)
run_odeint(system, slope_func)
plot(system.results.I, label='simulation')
plot(data.insulin, style='bo', label='insulin data')
decorate(xlabel='Time (min)',
ylabel='Concentration($\mu$U/mL)')
# -
# **Exercise:** Using the best parameters, estimate the sensitivity to glucose of the first and second phase pancreatic responsivity:
#
# $ \phi_1 = \frac{I_{max} - I_b}{k (G_0 - G_b)} $
#
# $ \phi_2 = \gamma \times 10^4 $
I_max = max(system.results.I)
I_max
first_phase = (I_max - Ib)/(k*(G0-Gb))
first_phase
second_phase = gamma * 10**4
second_phase
|
code/chap08-JLS.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Activity 4: Creating a TensorFlow Model using Keras
# In this notebook we design and compile a deep learning model using Keras as an interface to TensorFlow. We will continue to modify this model in our next lessons and activities by experimenting with different optimization techniques. However, the essential components of the model are entirely designed in this notebook.
from keras.models import Sequential
from keras.layers.recurrent import LSTM
from keras.layers.core import Dense, Activation
# ### Building a Model
# Our dataset contains daily observations and each observation influences a future observation. Also, we are interested in predicting a week--that is, seven days--of Bitcoin prices in the future. For those reasons, we chose the parameters `period_length` and `number_of_observations` as follows:
#
# * `period_length`: the size of the period to use as training input. Our periods are organized in distinct weeks. We will be using a 7-day period to predict a week in the future.
# * `number_of_observations`: how many distinct periods does our dataset has? We hvae 77 weeks available in our dataset, given that we will be using the very last week to test the LSTM network on every epoch, we will use 77 - 1 = 76 periods for training it.
period_length = 7
number_of_periods = 76
# We now build our LSTM model.
def build_model(period_length, number_of_periods, batch_size=1):
"""
Builds an LSTM model using Keras. This function
works as a simple wrapper for a manually created
model.
Parameters
----------
period_length: int
The size of each observation used as input.
number_of_periods: int
The number of periods available in the
dataset.
batch_size: int
The size of the batch used in each training
period.
Returns
-------
model: Keras model
Compiled Keras model that can be trained
and stored in disk.
"""
model = Sequential()
model.add(LSTM(
units=period_length,
batch_input_shape=(batch_size, number_of_periods, period_length),
input_shape=(number_of_periods, period_length),
return_sequences=False, stateful=False))
model.add(Dense(units=period_length))
model.add(Activation("linear"))
model.compile(loss="mse", optimizer="rmsprop")
return model
# ### Saving Model
# We can use the function `build_model()` as a starting point for building our model. That function will be refactored when building our Flask application for making it easier to train the network and use it for predictions. For now, let's store the model output on disk.
model = build_model(period_length=period_length, number_of_periods=number_of_periods)
model.save('bitcoin_lstm_v0.h5')
# The steps above compile the LSTM model as TensorFlow computation graph. We can now train that model using our train and evaluate its results with our test set.
|
Lesson-2/activity_4/Activity_4_Creating_a_TensorFlow_Model_Using_Keras.ipynb
|