url
stringlengths 63
66
| repository_url
stringclasses 1
value | labels_url
stringlengths 77
80
| comments_url
stringlengths 72
75
| events_url
stringlengths 70
73
| html_url
stringlengths 52
56
| id
int64 3.39M
3.04B
| node_id
stringlengths 18
32
| number
int64 1
4.39k
| title
stringlengths 3
367
| user
dict | labels
listlengths 0
5
| state
stringclasses 2
values | locked
bool 2
classes | assignee
dict | assignees
listlengths 0
2
| milestone
null | comments
listlengths 0
30
| created_at
timestamp[s]date 2012-02-26 12:10:46
2025-05-02 18:11:41
| updated_at
timestamp[s]date 2012-02-26 14:52:41
2025-05-02 18:16:27
| closed_at
stringlengths 0
20
| author_association
stringclasses 3
values | type
stringclasses 1
value | active_lock_reason
stringclasses 2
values | sub_issues_summary
dict | body
stringlengths 0
119k
| closed_by
dict | reactions
dict | timeline_url
stringlengths 72
75
| performed_via_github_app
null | state_reason
stringclasses 4
values | is_pull_request
bool 2
classes | draft
bool 2
classes | pull_request
dict |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
https://api.github.com/repos/materialsproject/pymatgen/issues/1801
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1801/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1801/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1801/events
|
https://github.com/materialsproject/pymatgen/issues/1801
| 580,692,526
|
MDU6SXNzdWU1ODA2OTI1MjY=
| 1,801
|
SpaceGroup.from_int_number(hexagonal=False) ineffective
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks. Can you pls implement the fix and submit a PR? Pls include unittests catching the wrong behavior.",
"Do you have a preference on whether I add the fix via option 1 or option 2 that I show above? sg_symbol_from_int_number() is the lower-level function, so it might be better there. But it could also have the biggest implications."
] | 2020-03-13T15:54:18
| 2020-03-16T21:15:13
|
2020-03-16T21:15:13Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When initiating a SpaceGroup class using the SpaceGroup.from_int_number(), the hexagonal option is ineffective for the following spacegroups: 146,148,155,160,161,166,167
NOTE: This is similar to issue #1203. The big difference is that here, it's claimed functionality that is ineffective. While a workaround exists, the functionality of hexagonal=False needs to be corrected.
**To Reproduce**
Below is an example with spacegroup 166 that illustrates the bug:
```
from pymatgen.symmetry.groups import SpaceGroup
SpaceGroup('R-3m') # correctly returns hexagonal info (per hexagonal is set as default)
SpaceGroup('R-3m:H') # correctly returns hexagonal info
SpaceGroup('R-3m:R') # correctly returns rhombohedral info
SpaceGroup.from_int_number(166, hexagonal=True) # correctly returns hexagonal info
SpaceGroup.from_int_number(166, hexagonal=False) # incorrectly returns hexagonal info
```
**Expected behavior**
```
sg = SpaceGroup.from_int_number(166, hexagonal=False)
sg.symbol # should be 'R-3mR', but instead give 'R-3mH' which is the hexagonal form
sg.order # should be 12, but instead gives 36 which is the hexagonal form
```
**Desktop (please complete the following information):**
- OS: Linux
- Version: 2019.12.22 (though code looks to be identical in the up-to-date github version)
**Additional context**
In the from_int_number() code ([located here on github](https://github.com/materialsproject/pymatgen/blob/5a3284fd2dce70ee27e8291e6558e73beaba5164/pymatgen/symmetry/groups.py#L470)), the method really is just calling sg_symbol_from_int_number() and running that result back through SpaceGroup(). The bug exists in sg_symbol_from_int_number(). More specifically, it is a logical error in the following line:
`syms = list(filter(lambda s: not s.endswith("H"), syms))`
This line is run when hexagonal=False. The issue is that this line will return ‘R-3m’, which then calls SpaceGroup(‘R-3m’) – this still gives the hexagonal object! We need that line to return ‘R-3m:R’, not ‘R-3m’. We could add ':R' in multiple places:
OPTION 1: within SpaceGroup.from_int_number():
```
##### original code ######
return SpaceGroup(sg_symbol_from_int_number(int_number,hexagonal=hexagonal))
##### fixed code ######
sym = sg_symbol_from_int_number(int_number,hexagonal=hexagonal)
if not hexagonal and int_number in [146,148,155,160,161,166,167]:
sym += ':R'
return SpaceGroup(sym)
```
OPTION 2: within sg_symbol_from_int_number()
This option is non-ideal because most will want the 'cleaner' symbol 'R-3m' instead of 'R-3m:R'
```
##### original code ######
if hexagonal:
syms = list(filter(lambda s: s.endswith("H"), syms))
else:
syms = list(filter(lambda s: not s.endswith("H"), syms))
##### fixed code ######
if hexagonal:
syms = list(filter(lambda s: s.endswith("H"), syms))
else:
syms = list(filter(lambda s: not s.endswith("H"), syms))
syms = [sym + ':R' for sym in syms]
```
OPTION 3: change the default return of SpaceGroup
I haven't gone through enough to see where this fix would be. I do think this would be the cleanest/best option though. In concept, here's the change in functionality though:
```
### original ###
SpaceGroup('R-3m') # will return hexagonal info
### edit ###
SpaceGroup('R-3m') # will return rhomb info
```
The whole point of the hexagonal=False option is for the spacegroups that have the option between trigonal and hexagonal lattices that we pick the correct one. In the symm_ops.yaml file, the choice between rhombohedral/hexagonal only exists for 146,148,155,160,161,166,167.
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1801/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1801/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1802
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1802/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1802/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1802/events
|
https://github.com/materialsproject/pymatgen/issues/1802
| 580,759,791
|
MDU6SXNzdWU1ODA3NTk3OTE=
| 1,802
|
Structure.from_spacegroup() inaccurate for trigonal crystal systems
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@blondegeek would you be willing to comment here, since this would re-open your #454 issue and possibly break some existing code/tests?",
"I might have had the conventional assignment of spacegroups backwards. This would make the original code correct...\r\nWhen looking at the AFlow library ([here](http://aflowlib.org/CrystalDatabase/trigonal_lattice.html))...\r\nHexagonal lattices:\r\n\r\n> 143, 144, 145, 147, 149, 150, 151, 152,153, 154, 156, 157, 158, 159, 162, 163,164, 165\r\n\r\n Rhombohedral lattices:\r\n\r\n> 146,148,155,160,161,166,167\r\n\r\nI was led to believe the opposite when using another crystallography server, so I apologize for all of the confusion."
] | 2020-03-13T18:02:25
| 2020-03-16T17:10:35
|
2020-03-16T17:10:35Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I apologize in advance for the long post, but I think this is a confusing topic. I wanted to give a full justification for the changes I'm suggesting!
**Summary:**
Structure.from_spacegroup() will inaccurately produce structures for the following spacegroups:
143, 144, 145, 147, 149, 150, 151, 152,153, 154, 156, 157, 158, 159, 162, 163,164, 165.
This occurs because SpaceGroup.is_compatible() requires these spacegroups to be hexagonal lattices, when the [symm_op.yaml](https://github.com/materialsproject/pymatgen/blob/master/pymatgen/symmetry/symm_ops.yaml) only provides operations for the trigonal system. This is a bug that is introduced by issue #454, where discussion is misled by rhombohedral/trigonal/hexagonal confusion.
**Issue #454 inaccuracies:**
In the posted issue (to summarize), [blondegeek](https://github.com/blondegeek) argues that trigonal spacegroups (specifically spacegroup 165) should have hexagonal lattices. Also, [wenhaosun](https://github.com/wenhaosun) states that spacegroup 165 is trigonal. [blondegeek](https://github.com/blondegeek) then makes the final statement that spacegroup 165 is indeed tigonal crystal system but possesses a hexagonal lattice system. The result of this discussion is that all trigonal spacegroups (see list in summary above) are checked to be hexagonal in the [SpaceGroup.is_compatible()](https://github.com/materialsproject/pymatgen/blob/5a3284fd2dce70ee27e8291e6558e73beaba5164/pymatgen/symmetry/groups.py#L362) code.
Their discussion comes close, but it is still incorrect -- leading to a logical error in the code. The confusion can actually be summarized with a quote from wikipedia:
> Crystal systems, crystal families and lattice systems are similar but slightly different, and there is widespread confusion between them: in particular the trigonal crystal system is often confused with the rhombohedral lattice system, and the term "crystal system" is sometimes used to mean "lattice system" or "crystal family".
Space group 165 does NOT have the hexagonal lattice conventionally - it has a **rhombohedral** lattice. But since you can convert between the two, you can have crystal structures that have a hexagonal lattice with space group 165. Because you can have multiple forms of a lattice, this means the is_compatible() function should take that into account - but more that later. To state things clearly, spacegroups like 165 (and the others listed above) belong to the **trigonal crystal system** ([source](https://en.wikipedia.org/wiki/Space_group)) and the **hexagonal crystal family** ([source](https://en.wikipedia.org/wiki/Crystal_system)), but they possess a lattice that belongs to the **rhombohedral lattice system** ([source](https://en.wikipedia.org/wiki/Hexagonal_crystal_family)).
You'll see that in [this page](https://en.wikipedia.org/wiki/Hexagonal_crystal_family), the trigonal crystal system can have either a rhombohedral or hexagonal lattice system. The trigonal space groups (143-167) are a mixture of these lattice systems. Here is how the spacegroup lattices are assigned conventionally...
Rhombohedral lattices:
> 143, 144, 145, 147, 149, 150, 151, 152,153, 154, 156, 157, 158, 159, 162, 163,164, 165
Hexagonal lattices:
> 146,148,155,160,161,166,167
**Bugs & Errors in pymatgen**
Based on the discussion above, I believe the following errors in pymatgen exist:
In pymatgen.symmetry.groups module, the SpaceGroup.is_compatible() ([link](https://github.com/materialsproject/pymatgen/blob/5a3284fd2dce70ee27e8291e6558e73beaba5164/pymatgen/symmetry/groups.py#L362)) function is incorrect. The follow code should be changed:
```
### CURRENT CODE ###
elif crys_system == "hexagonal" or (
crys_system == "trigonal" and (
self.symbol.endswith("H") or
self.int_number in [143, 144, 145, 147, 149, 150, 151, 152,
153, 154, 156, 157, 158, 159, 162, 163,
164, 165])):
### FIXED CODE ###
elif crys_system == "hexagonal" or (
crys_system == "trigonal" and self.symbol.endswith("H")):
```
This is pretty much undoing the changes made in issue #454, so that issue would be reopened. What [blondegeek](https://github.com/blondegeek) should have been told is that her hexagonal lattice is not compatible with the **conventional** lattice of that spacegroup. She needs to first convert her crystal structure to the primitive cell (should give the rhombohedral) which is then compatible. A separate function can be made that does a thorough compatibility check, but as-is the is_compatible() function should only check via convention. Maybe we can add a warning to the doc-string of that function that warns the user of this issue.
Thus far, it's really just a matter of convention, but the really major error comes from the pymatgen.core.structure module. Specifically, the SpaceGroup.from_spacegroup() method will be inaccurate for trigonal crystal systems that are based on the rhombohedral lattice. This because the symmetry operations related to the spacegroups (located in [symm_ops.yaml](https://github.com/materialsproject/pymatgen/blob/5a3284fd2dce70ee27e8291e6558e73beaba5164/pymatgen/symmetry/symm_ops.yaml)) are based on the rhombohedral lattice. If you took a hexagonal lattice and applied the rhombohedral symmetry operations to it, you won't get the correct crystal structure!!!
If someone wants to use the Structure.from_spacegroup() method, where they can choose between a rhombohedral and hexagonal lattice, then the hexagonal=True/False option needs to be added (note there is currently another bug with that option. see issue #1801). This option is only useful for spacegroups 146,148,155,160,161,166, and 167, because only those spacegroups have both the rhomb and hexagonal options available in symm_ops.yaml.
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1802/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1802/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1803
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1803/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1803/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1803/events
|
https://github.com/materialsproject/pymatgen/pull/1803
| 580,779,764
|
MDExOlB1bGxSZXF1ZXN0Mzg3OTI1MzUz
| 1,803
|
Fixing up where #1799 (Fix Q-Chem API) left off
|
{
"login": "espottesmith",
"id": 17201238,
"node_id": "MDQ6VXNlcjE3MjAxMjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/17201238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/espottesmith",
"html_url": "https://github.com/espottesmith",
"followers_url": "https://api.github.com/users/espottesmith/followers",
"following_url": "https://api.github.com/users/espottesmith/following{/other_user}",
"gists_url": "https://api.github.com/users/espottesmith/gists{/gist_id}",
"starred_url": "https://api.github.com/users/espottesmith/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/espottesmith/subscriptions",
"organizations_url": "https://api.github.com/users/espottesmith/orgs",
"repos_url": "https://api.github.com/users/espottesmith/repos",
"events_url": "https://api.github.com/users/espottesmith/events{/privacy}",
"received_events_url": "https://api.github.com/users/espottesmith/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @espottesmith ",
"Test failures: TestQCOutput.test_structural_change",
"I see it. As usual, it's an OpenBabel problem."
] | 2020-03-13T18:46:14
| 2022-01-31T21:04:28
|
2020-03-13T20:25:03Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This copies the changes from #1799 and adds tests for the relevant functions in pymatgen.io.qchem.outputs.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1803/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1803/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1803",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1803",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1803.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1803.patch",
"merged_at": "2020-03-13T20:25:03Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1804
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1804/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1804/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1804/events
|
https://github.com/materialsproject/pymatgen/pull/1804
| 581,361,275
|
MDExOlB1bGxSZXF1ZXN0Mzg4NDMwMDAz
| 1,804
|
[WIP] CP2K Module
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @nwinner, this looks great.\r\n\r\nRegarding the Bader analysis, could you explore the idea of merging implementations -- I'd prefer not to have two separate classes each for Gaussian Cube / VASP Chgcar given that they're basically equivalent. This might also be good time to add `user_bader_settings` as an option to the Bader caller, e.g. for vacuum cut-off and other optional parameters that it takes.\r\n\r\nRegarding xyz parsing and large runs, I just wanted to highlight the `Trajectory` class in `pymatgen.core.trajectory` which is a more light-weight version of storing outputs from large runs: it stores it as a single Structure object + a list of displacements."
] | 2020-03-14T20:33:15
| 2020-03-17T23:13:09
|
2020-03-17T23:13:09Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Work in progress for integration of CP2K in pymatgen. Currently in working condition, but outputs.py and sets.py need to be fleshed out with more features. Also some small changes to existing files have been made that could be useful for this integration.
**Features**
1. Creation of pymatgen.io.cp2k module with the following contents:
- inputs.py: support for the the input file structure of cp2k executable
- sets.py: input sets for running default cp2k calculations using predefined input structure
- outputs.py: cp2k output file parsing
- utils.py: a handful of functions to help with input sets, more to be inserted
- test_files: just a few test files, not exhaustive
2. Added skip_step to the xyz parser. For large MD runs, this can be hugely beneficial. While reading in the raw xyz text as frames is very fast, creating a pymatgen structure object takes a while, so this makes dealing with large xyz files easier.
3. In bader_caller.py in the command line module, it assumed a VASP structure. For Cp2k, the more general .cube files are used. These require a simpler version of the bader caller which I'm currently working on.
**Fixes**
1. pymatgen.io.xyz had a tiny issue in how it was playing with cp2k. It seemed that zopen was not using the 'rt' argument, so when gzipped .xyz files were read in, they couldn't be manipulated like strings.
**TODO**
1. Make the output parsing a little more robust. At the moment it makes some assumptions that will be fine if you know how its written, but for general use it should be a lot better at dealing with variables in the output file structure.
2. Create more input sets.
3. Fix the broken symmetry initialization of the input sets using a util that maps between structure objects and the &BS section of the cp2k input file.
4. Bader caller needs to integrate the "charge transfer" that the VASP one does.
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1804/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1804/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1804",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1804",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1804.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1804.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1805
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1805/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1805/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1805/events
|
https://github.com/materialsproject/pymatgen/pull/1805
| 582,411,989
|
MDExOlB1bGxSZXF1ZXN0Mzg5MzM1NTEx
| 1,805
|
Hexagonal Sym Issues
|
{
"login": "jacksund",
"id": 47992949,
"node_id": "MDQ6VXNlcjQ3OTkyOTQ5",
"avatar_url": "https://avatars.githubusercontent.com/u/47992949?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jacksund",
"html_url": "https://github.com/jacksund",
"followers_url": "https://api.github.com/users/jacksund/followers",
"following_url": "https://api.github.com/users/jacksund/following{/other_user}",
"gists_url": "https://api.github.com/users/jacksund/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jacksund/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jacksund/subscriptions",
"organizations_url": "https://api.github.com/users/jacksund/orgs",
"repos_url": "https://api.github.com/users/jacksund/repos",
"events_url": "https://api.github.com/users/jacksund/events{/privacy}",
"received_events_url": "https://api.github.com/users/jacksund/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I limited the merge request to only fix #1801, as I found a mistake in my post for #1802.",
"Thanks"
] | 2020-03-16T15:59:25
| 2020-03-16T21:14:22
|
2020-03-16T21:14:16Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This merge request addresses issues #1801 and #1802. I decided to add the #1801 fix to the SpaceGroup.from_int_number() method, which is not the lowest function with the hexagonal=True input. The respective unittests for each fix were also added.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1805/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1805/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1805",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1805",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1805.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1805.patch",
"merged_at": "2020-03-16T21:14:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1806
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1806/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1806/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1806/events
|
https://github.com/materialsproject/pymatgen/pull/1806
| 582,667,619
|
MDExOlB1bGxSZXF1ZXN0Mzg5NTUyOTIx
| 1,806
|
[WIP] Add xTB IO to pymatgen
|
{
"login": "espottesmith",
"id": 17201238,
"node_id": "MDQ6VXNlcjE3MjAxMjM4",
"avatar_url": "https://avatars.githubusercontent.com/u/17201238?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/espottesmith",
"html_url": "https://github.com/espottesmith",
"followers_url": "https://api.github.com/users/espottesmith/followers",
"following_url": "https://api.github.com/users/espottesmith/following{/other_user}",
"gists_url": "https://api.github.com/users/espottesmith/gists{/gist_id}",
"starred_url": "https://api.github.com/users/espottesmith/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/espottesmith/subscriptions",
"organizations_url": "https://api.github.com/users/espottesmith/orgs",
"repos_url": "https://api.github.com/users/espottesmith/repos",
"events_url": "https://api.github.com/users/espottesmith/events{/privacy}",
"received_events_url": "https://api.github.com/users/espottesmith/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@espottesmith do you have an update on this PR?",
"I unfortunately have not been able to work towards this recently. However, I know that @arepstein (Alex Epstein) has been working towards this goal, as well as potentially adding xTB/CREST workflows for atomate. I can check in with him about the status there. It might be best to close this PR and let him open another.",
"Ok, will close it for now. Would be nice to see this functionality if someone wants to pick it up though."
] | 2020-03-17T00:05:50
| 2022-09-28T16:12:17
|
2020-12-01T17:07:48Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR will eventually add support in pymatgen for parsing input and output files for Grimme's Extended Tight-Binding (xTB) code (DOI: 10.1021/acs.jctc.7b00118, DOI: 10.1021/acs.jctc.8b01176, DOI: 10.26434/chemrxiv.8326202.v1).
This is not at all finished or tested, and additional work will be needed to get this code in working order.
Note that this code does not actually interface with xTB directly (such functionality is intended to be added to downstream codes such as custodian and atomate).
## TODO (if any)
* Write tests for input class
* Finish main output class
* Write tests for output class
* (Possibly) move QChem IO utils, because it seems like I will be using them for these xTB IO classes.
## Checklist
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1806/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1806/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1806",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1806",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1806.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1806.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1807
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1807/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1807/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1807/events
|
https://github.com/materialsproject/pymatgen/pull/1807
| 582,835,330
|
MDExOlB1bGxSZXF1ZXN0Mzg5NjkwMjIz
| 1,807
|
Reorder check for metal
|
{
"login": "mfherbst",
"id": 11157893,
"node_id": "MDQ6VXNlcjExMTU3ODkz",
"avatar_url": "https://avatars.githubusercontent.com/u/11157893?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mfherbst",
"html_url": "https://github.com/mfherbst",
"followers_url": "https://api.github.com/users/mfherbst/followers",
"following_url": "https://api.github.com/users/mfherbst/following{/other_user}",
"gists_url": "https://api.github.com/users/mfherbst/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mfherbst/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mfherbst/subscriptions",
"organizations_url": "https://api.github.com/users/mfherbst/orgs",
"repos_url": "https://api.github.com/users/mfherbst/repos",
"events_url": "https://api.github.com/users/mfherbst/events{/privacy}",
"received_events_url": "https://api.github.com/users/mfherbst/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2020-03-17T08:18:04
| 2020-03-17T12:20:24
|
2020-03-17T12:01:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Reorder logic in BSPlotter, such that check for metal is only executed when needed.
* The current implementation causes trouble in my downstream usage of pymatgen, so I'd appreciate a merge.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1807/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1807/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1807",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1807",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1807.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1807.patch",
"merged_at": "2020-03-17T12:01:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1808
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1808/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1808/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1808/events
|
https://github.com/materialsproject/pymatgen/pull/1808
| 583,709,005
|
MDExOlB1bGxSZXF1ZXN0MzkwNDE5OTM5
| 1,808
|
[Needs tests] New methods of Completedos in dos.py
|
{
"login": "Nokimann",
"id": 31014779,
"node_id": "MDQ6VXNlcjMxMDE0Nzc5",
"avatar_url": "https://avatars.githubusercontent.com/u/31014779?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Nokimann",
"html_url": "https://github.com/Nokimann",
"followers_url": "https://api.github.com/users/Nokimann/followers",
"following_url": "https://api.github.com/users/Nokimann/following{/other_user}",
"gists_url": "https://api.github.com/users/Nokimann/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Nokimann/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Nokimann/subscriptions",
"organizations_url": "https://api.github.com/users/Nokimann/orgs",
"repos_url": "https://api.github.com/users/Nokimann/repos",
"events_url": "https://api.github.com/users/Nokimann/events{/privacy}",
"received_events_url": "https://api.github.com/users/Nokimann/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"id": 2414169365,
"node_id": "MDU6TGFiZWwyNDE0MTY5MzY1",
"url": "https://api.github.com/repos/materialsproject/pymatgen/labels/needs%20testing",
"name": "needs testing",
"color": "66ddab",
"default": false,
"description": "PRs that are not ready to merge due to lacking tests"
}
] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Many thanks @Nokimann, this will need tests too.",
"Hi @Nokimann, I was wondering if you had any updates on this PR? I'm happy to merge this if it has a test added.",
"@mkhorton Can I request this PR in the newest version of pymatgen? \r\nFirst, I will close this PR section, and then request it again.\r\nI did finish the same thing for the bandstructure.py in indirect ways (not to touch the script of plotting the bandstructure)\r\n"
] | 2020-03-18T12:56:47
| 2020-12-24T08:31:49
|
2020-12-24T08:24:02Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Three methods: `get_orbital_dos`, `get_element_orbital_dos`, `get_bundle_pdos` in `class CompleteDos` are added
`get_orbital_dos` gives projected DOS of s, px, py, pz, dxy, dyz, dz2, dxz, dx2, ...
`get_element_orbital_dos` gives projected DOS of s, px, py, pz, dxy, dyz, dz2, dxz, dx2, ... at a particular element
The previous `get_*spd*_dos` didn't give the above sub-projected dos
`get_bundle_pdos` gives the sum of multiple projected DOS. It uses most of `get_***_dos` functions.
And the input argument can be many general and straightforward formats. But, I'm not sure that the object format like previous input of `get_***_dos` is better or not.
```
site:int, element:str, orbital:str, orbital:str, ortibtal_type:str, (site:int, orbital:str),
(site:int, orbital_type:str), (element:str, orbital:str), (element:str, orbital_type:str)
```
Examples of arguments:
site - 0, 1, 2, ... / element - 'H', 'He', ... / orbital_type - 's', 'p', 'd', 'f' / orbital - 's', 'px', 'py', ...
## TODO (if any)
I will massage and add some functions in a similar way to `bandstructure.py`
|
{
"login": "Nokimann",
"id": 31014779,
"node_id": "MDQ6VXNlcjMxMDE0Nzc5",
"avatar_url": "https://avatars.githubusercontent.com/u/31014779?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Nokimann",
"html_url": "https://github.com/Nokimann",
"followers_url": "https://api.github.com/users/Nokimann/followers",
"following_url": "https://api.github.com/users/Nokimann/following{/other_user}",
"gists_url": "https://api.github.com/users/Nokimann/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Nokimann/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Nokimann/subscriptions",
"organizations_url": "https://api.github.com/users/Nokimann/orgs",
"repos_url": "https://api.github.com/users/Nokimann/repos",
"events_url": "https://api.github.com/users/Nokimann/events{/privacy}",
"received_events_url": "https://api.github.com/users/Nokimann/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1808/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1808/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1808",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1808",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1808.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1808.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1809
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1809/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1809/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1809/events
|
https://github.com/materialsproject/pymatgen/pull/1809
| 588,175,994
|
MDExOlB1bGxSZXF1ZXN0MzkzOTczMDAw
| 1,809
|
Rewrite EconNN implementation
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I think the tests are a bit too rudimentary. Diamond, NaCl and CsCl are hardly difficult structures. Can we include at least one more difficult structure?",
"I added LiFePO4 and tidied the tests a little.\r\n\r\nI didn't add the LiFePO4 test for MinimumVIRENN as it is very slow. ",
"> I didn't add the LiFePO4 test for MinimumVIRENN as it is very slow.\r\n\r\nIs still slow if you run it on a structure that already has oxidation states annotated?",
"Adding oxidation states beforehand didn't change the timing. However, the additional MinimumVIRENN only takes 4 seconds so I added it in.",
"These are so slow :(\r\n\r\nCutoffDictNNs have some obvious optimizations, I really need to re-visit at some point. The NN time is going to be a big factor in driving the visualizations on the web too.",
"<img src=\"https://user-images.githubusercontent.com/1330638/77689245-9673c580-6f5e-11ea-884c-08e1ddedbefc.png\">\r\n\r\nEconNN score will change with this latest update.",
"@sivonxay I remember we discussed optimizations to JmolNN (and other CutoffDictNNs), I remember you had a suggestion but that there was a roadblock with the implementation, could you remind me what the status was with this?\r\n\r\n@utf given that JmolNN is already on the Pareto front, it's probably not urgent that it's improved for this analysis, but I'm going to take a quick look at it",
"@mkhorton Ahhh yes. I had initially thought the CutoffDictNN was poorly optimized, since it was taking a while to run. After a bit of digging, I found out that the slowdown was not due to the neighbor finding code, but due to the NearestNeighbor._get_image() function. With my structures of > 100 atoms, this was a large bottleneck.",
"@utf with the additional LiFePO4 test added, is this ready to merge?\r\n\r\n@sivonxay thanks for the reminder! I'll take a look at optimizing that, will bring discussion of `_get_image` to a new issue/PR",
"Ok, once tests pass this should be ready to merge",
"I also ran into the bottleneck of `get_image` for some large structures. I think a major reason is that frac_coords is continually called and also get_all_neighbors is called many times.\r\n\r\nI made an optimised version of MinimumDistanceNN by only calculating frac_coords once and moving the get_image function inside get_nn_info. I also precompute get_all_neighbors once and reuse that information. This isn't the best modular implementation but might be useful if you are doing some optimisation:\r\n\r\n```python\r\nfrom pymatgen.analysis.local_env import MinimumDistanceNN\r\n\r\nclass FastMinimumDistanceNN(MinimumDistanceNN):\r\n \"\"\"\r\n Optimized version of MinimumDistanceNN\r\n \"\"\"\r\n \r\n def get_all_nn_info(self, structure):\r\n \"\"\"Get a listing of all neighbors for all sites in a structure\r\n\r\n Args:\r\n structure (Structure): Input structure\r\n Return:\r\n List of NN site information for each site in the structure. Each\r\n entry has the same format as `get_nn_info`\r\n \"\"\"\r\n all_neighbors = structure.get_all_neighbors(self.cutoff)\r\n \r\n return [self.get_nn_info(structure, n, all_neighbors=all_neighbors)\r\n for n in range(len(structure))]\r\n\r\n def get_nn_info(self, structure, n, all_neighbors=None):\r\n \"\"\"\r\n Get all near-neighbor sites as well as the associated image locations\r\n and weights of the site with index n using the closest neighbor\r\n distance-based method.\r\n\r\n Args:\r\n structure (Structure): input structure.\r\n n (integer): index of site for which to determine near\r\n neighbors.\r\n\r\n Returns:\r\n siw (list of tuples (Site, array, float)): tuples, each one\r\n of which represents a neighbor site, its image location,\r\n and its weight.\r\n \"\"\"\r\n\r\n site = structure[n]\r\n if not all_neighbors:\r\n neighs_dists = structure.get_neighbors(site, self.cutoff)\r\n else:\r\n neighs_dists = all_neighbors[n]\r\n \r\n from pymatgen.util.coord import pbc_diff\r\n frac_coords = structure.frac_coords\r\n\r\n def get_original_site(site):\r\n \"\"\"Private convenience method for get_nn_info,\r\n gives original site index from ProvidedPeriodicSite.\"\"\"\r\n diff = pbc_diff(site.frac_coords, frac_coords)\r\n matches = np.where(np.all(np.abs(diff < 1e-8), axis=-1))[0]\r\n if len(matches) > 0:\r\n return matches[0]\r\n else:\r\n raise Exception('Site not found!')\r\n\r\n def get_image(site):\r\n original_site_n = get_original_site(site)\r\n original_site = structure[original_site_n]\r\n image = np.around(np.subtract(site.frac_coords, original_site.frac_coords))\r\n image = tuple(image.astype(int))\r\n return original_site_n, image\r\n\r\n siw = []\r\n if self.get_all_sites:\r\n for nn in neighs_dists:\r\n w = nn.nn_distance\r\n original_site, image = get_image(site)\r\n siw.append({'site': nn,\r\n 'image': image,\r\n 'weight': w,\r\n 'site_index': original_site})\r\n else:\r\n min_dist = min([nn.nn_distance for nn in neighs_dists])\r\n for nn in neighs_dists:\r\n dist = nn.nn_distance\r\n if dist < (1.0 + self.tol) * min_dist:\r\n w = min_dist / dist\r\n original_site, image = get_image(site)\r\n siw.append({'site': nn,\r\n 'image': image,\r\n 'weight': w,\r\n 'site_index': original_site})\r\n return siw\r\n```"
] | 2020-03-26T05:57:09
| 2020-03-27T02:25:46
|
2020-03-27T02:25:46Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The EconNN class was rewritten to properly reproduce the algorithm outlined in:
https://www.degruyter.com/view/j/zkri.1979.150.issue-1-4/zkri.1979.150.14.23/zkri.1979.150.14.23.xml
This improves the performance of the method, particularly when only considering bonding between sites of opposing charge (controlled using the new `cation_anion` option).
The method has a tolerance parameter that determines when to consider a site bonded. After trialling several values, it seems like the results are rather insensitive in the range 0.1–0.6. I've set it to the optimal value of 0.2 that I found for my [test set](https://github.com/hackingmaterials/materialscoord).
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1809/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1809/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1809",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1809",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1809.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1809.patch",
"merged_at": "2020-03-27T02:25:46Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1810
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1810/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1810/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1810/events
|
https://github.com/materialsproject/pymatgen/pull/1810
| 588,704,856
|
MDExOlB1bGxSZXF1ZXN0Mzk0NDAyNjE4
| 1,810
|
Improvements to mcsqs integration
|
{
"login": "rwoodsrobinson",
"id": 7270689,
"node_id": "MDQ6VXNlcjcyNzA2ODk=",
"avatar_url": "https://avatars.githubusercontent.com/u/7270689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rwoodsrobinson",
"html_url": "https://github.com/rwoodsrobinson",
"followers_url": "https://api.github.com/users/rwoodsrobinson/followers",
"following_url": "https://api.github.com/users/rwoodsrobinson/following{/other_user}",
"gists_url": "https://api.github.com/users/rwoodsrobinson/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rwoodsrobinson/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rwoodsrobinson/subscriptions",
"organizations_url": "https://api.github.com/users/rwoodsrobinson/orgs",
"repos_url": "https://api.github.com/users/rwoodsrobinson/repos",
"events_url": "https://api.github.com/users/rwoodsrobinson/events{/privacy}",
"received_events_url": "https://api.github.com/users/rwoodsrobinson/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rwoodsrobinson, will merge once tests pass\r\n\r\n@dongsenfo be aware that the keyword args for `run_mcsqs` have changed here to merge `supercell` and `numatoms` options into a single `scaling` option (since both cannot be used simultaneously); I think the improvement is worthwhile to make the change, but since you're the primary user of this code I wanted to make sure you were notified",
"@mkhorton That sounds good to me. \r\n\r\nThanks @rwoodsrobinson!",
"@dongsenfo there was also an issue with the old caller returning structures that were parsed incorrectly (which we discussed in the office), this caller now returns guaranteed correct structures however the underlying parsing bug still has not been fixed yet. Just flagging this in case you've encountered this in your own work. Maybe @rwoodsrobinson can comment further or supply examples",
"You're welcome @dongsenfo, and thanks @mkhorton for the assistance with these changes.\r\n\r\nTo elaborate, I was looking at cif files for some structure outputs from the old code for what should have been a 128 atom sqs derived from mp-1014244 (ZnZrN2 in the \"wurtsalt\" structure) and they looked like this:\r\n<img width=\"505\" alt=\"ZZN-wurtsalt-old\" src=\"https://user-images.githubusercontent.com/7270689/77697688-8a8f0000-6f6c-11ea-832b-cf0cdf4d0525.png\">\r\n\r\nWith the updated code using the \"str2cif\" function in atat, it looks more reasonable:\r\n<img width=\"865\" alt=\"ZZN-wurtsalt-new\" src=\"https://user-images.githubusercontent.com/7270689/77697699-91b60e00-6f6c-11ea-85ff-25d84927ec1b.png\">\r\n\r\nI also added the objective function to the output of run_mcsqs so we can test convergence for future calculations, though this is a work in progress (I plan to add T, w_p, w_n for test). Happy to discuss other changes."
] | 2020-03-26T20:25:23
| 2020-03-27T02:25:58
|
2020-03-27T02:25:58Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Improved tests
* Return objective function
* Simplified method signature
* Improved docs
* Fixed issue with `run_mcsqs` output
## TODO (if any)
* Further improvements in a future pull request (e.g. include MC temperature)
## Checklist
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1810/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1810/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1810",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1810",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1810.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1810.patch",
"merged_at": "2020-03-27T02:25:58Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1811
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1811/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1811/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1811/events
|
https://github.com/materialsproject/pymatgen/issues/1811
| 588,778,346
|
MDU6SXNzdWU1ODg3NzgzNDY=
| 1,811
|
StructureGraph produced a multi-directed graph
|
{
"login": "tawe141",
"id": 13124532,
"node_id": "MDQ6VXNlcjEzMTI0NTMy",
"avatar_url": "https://avatars.githubusercontent.com/u/13124532?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/tawe141",
"html_url": "https://github.com/tawe141",
"followers_url": "https://api.github.com/users/tawe141/followers",
"following_url": "https://api.github.com/users/tawe141/following{/other_user}",
"gists_url": "https://api.github.com/users/tawe141/gists{/gist_id}",
"starred_url": "https://api.github.com/users/tawe141/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/tawe141/subscriptions",
"organizations_url": "https://api.github.com/users/tawe141/orgs",
"repos_url": "https://api.github.com/users/tawe141/repos",
"events_url": "https://api.github.com/users/tawe141/events{/privacy}",
"received_events_url": "https://api.github.com/users/tawe141/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is not a bug. We have to define bonds for _periodic_ solids and this means each bond does not just have a `from_atom` and a `to_atom` but also a `to_image` which encodes which periodic image the bond is pointing towards.\r\n\r\nConsider an example of a simple cubic structure with a single atom whose single atom is bonded to itself -- this now has 6 bonds, the `from_atom` and `to_atom` are identical for all 6 bonds, only the `to_image` changes.\r\n\r\nSince the nodes define the `from_atom` and `to_atom` the edges are between the same two nodes for all 6 bonds, the difference is in the edges which are encoded with the `to_image` metadata.\r\n\r\nTherefore a MultiDiGraph is required if defining graphs in this manner.",
"And just to clarify why it's a directional graph: the directionality is required so that you know which node corresponds to the `to_image` is referring to.\r\n\r\nThere are of course other ways to represent periodic solids as graphs, they variously have different advantages and disadvantages. The situation for molecules is much simpler and a normal `Graph` is fine."
] | 2020-03-26T22:59:39
| 2020-03-26T23:17:28
|
2020-03-26T23:16:12Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Not sure if this is a bug or something that's undocumented, but `StructureGraph.with_local_env_strategy(struct, CrystalNN())` produces a MultiDiGraph. This doesn't make much sense to me, since bonds do not inherently have directionality. This also inhibits intuitive use of the underlying `networkx` object.
**To Reproduce**
Steps to reproduce the behavior:
Using [this cif file](https://www.dropbox.com/s/p6pv1o98x7zwvoh/pn_Mg.cif?dl=0), run the following:
```
struct = CifParser('pn_Mg.cif').get_structures(primitive=False)[0]
g = StructureGraph.with_local_env_strategy(struct, CrystalNN(distance_cutoffs=None, x_diff_weight=0.0))
type(g.graph)
# last line returns "networkx.classes.multidigraph.MultiDiGraph"
```
**Expected behavior**
I would expect a Graph or MultiGraph object.
**Desktop (please complete the following information):**
- OS: Ubuntu 18.04
- Version: 2020.3.13
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1811/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1811/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1812
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1812/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1812/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1812/events
|
https://github.com/materialsproject/pymatgen/issues/1812
| 589,613,063
|
MDU6SXNzdWU1ODk2MTMwNjM=
| 1,812
|
Unexpected structure.get_sorted_structure in structure to lammps data
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Similar issues are with \r\n\r\nhttps://github.com/materialsproject/pymatgen/blob/561b4cfed1f74ede87074fae3903885b9d5c7868/pymatgen/io/lammps/data.py#L806-L807\r\n\r\nHere the structures are rotated without the end users knowing.. This causes unexpected mismatch between force vectors calculated with lammps and the original structures' force vectors when the cell is triclinic. It costed me sometime on this before as well, but the only solution here seems to require the users know this and rotate their structures to consistent direction.",
"Structure sorting can be definitely bypassed since it is only for the purpose of sanity, i.e., grouping atoms of the same element. \r\n\r\nHowever, rotation is inevitable when dealing with structures with non-orthogonal lattice. There are certain [alignment requirements](https://lammps.sandia.gov/doc/Howto_triclinic.html) when the simulation box is triclinic. And unfortunately it only takes care of conversion from structure and never converts the axis back to the original structure. \r\n\r\n\r\n\r\n",
"@adengz good that you are still around. \r\n\r\nFor the lattice, I understand that lammps takes a specific representation of triclinic lattice, but do we really need to rotate all atoms in the cell? Line 807 specifically. \r\n\r\nWe can just redefine the lattice but keep the coordinates right? This way the direction of force vector does not change. It will change the fractional coordinates of course..",
"@chc273 I think the fractional coordinates shall remain unchanged to keep the structure integrated. By constructing the rotated structure from fractional coordinates, there is no need to know the rotation. \r\n\r\nHowever, if (site) velocities are attached to structures, they need to be rotated accordingly under xyz axis. And I think this also applies to atomic forces as well.. ",
"@adengz thanks. My previous assumption of keeping coordinates but only change lattice is wrong. "
] | 2020-03-28T16:42:43
| 2020-03-30T13:47:23
|
2020-03-30T13:47:23Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
https://github.com/materialsproject/pymatgen/blob/47c8223dfe2ef4af6ebc273c950a72ccf038c5c6/pymatgen/io/lammps/data.py#L805
I found this behavior unexpected for some applications.
If I want to calculate the site descriptors for a given site index in a structure using lammps, I can get wrong results since this line of code reordered the sites in the structure before outputting to lammps data.
For example, if my structure has two sites ['S', 'Mo'], and I want to calculate local environment descriptors using lammps on the first site (S), I would always get the properties of the 'Mo' site due to the structure sort.
I had to sort my structures before using this code every time to avoid error and I wonder if this behavior can be changed.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1812/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1812/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1813
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1813/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1813/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1813/events
|
https://github.com/materialsproject/pymatgen/issues/1813
| 589,720,464
|
MDU6SXNzdWU1ODk3MjA0NjQ=
| 1,813
|
POTCAR setup fails
|
{
"login": "paulfons",
"id": 6129100,
"node_id": "MDQ6VXNlcjYxMjkxMDA=",
"avatar_url": "https://avatars.githubusercontent.com/u/6129100?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/paulfons",
"html_url": "https://github.com/paulfons",
"followers_url": "https://api.github.com/users/paulfons/followers",
"following_url": "https://api.github.com/users/paulfons/following{/other_user}",
"gists_url": "https://api.github.com/users/paulfons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/paulfons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/paulfons/subscriptions",
"organizations_url": "https://api.github.com/users/paulfons/orgs",
"repos_url": "https://api.github.com/users/paulfons/repos",
"events_url": "https://api.github.com/users/paulfons/events{/privacy}",
"received_events_url": "https://api.github.com/users/paulfons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-03-29T05:22:41
| 2023-08-10T15:01:42
|
2023-08-10T15:01:42Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
On a new machine (a Macintosh in this case), following the instructions from the pymatgen homepage (using omg config -p <expanded> <pymatgen_potcar> results in a setup that does leads to errors in POTCAR file generation.
**To Reproduce**
I have encountered some problems with the use of pymatgen and the generation of POTCAR VASP files. I thought I would
review the installation procedure in an attempt to determine what is going wrong.
There are several different versions of PAW pseudopotentials on the VASP web site. In principle, it would seem that pymatgen
should work with any of them, but there are problems. At the very least, the documentation on PAW installation using
pmg needs to be updated to avoid the errors I have been encountering.
PAW PS on the VASP site
1. PAW POTCAR files: LDA version 54 (recommend)
2. PAW POTCAR files: PBE version 54 (recommend)
3. PAW POTCAR files: LDA & PBE, 5.2 & 5.4 (original unvie release version)
4. PAW POTCAR files: LDA version 52
5. PAW POTCAR files: PBE version 52
other files are labeled as outdated
The online text states that the distributions 1 and 2 yield identical results to distribution 3, but also
states that the TITEL has been changed and a hash key has been added of the newest distributions 1 and 2.
The above distributions correspond to the following tar images from the www.vasp.at website:
1. 79M Mar 27 21:45 potpaw_PBE.54.tar
2. 71M Mar 27 21:45 potpaw_LDA.54.tar
3. 280M Mar 27 21:45 potpaw_LDA_PBE_52_54_orig.tar
4. 62M Mar 27 22:14 potpaw_LDA.52.tar
5. 70M Mar 27 22:14 potpaw_PBE.52.tar
By way of demonstration, I initially unpacked distribution (3) (PAW POTCAR files: LDA & PBE, 5.2 & 5.4 (original unvie release version))
into a new directory to yield. I was under the impression that this was the older version used by pymatgen (although the
installation instructions on the pymatgen website are fuzzy)
<extracted>
potpaw.52 potpaw.54 potpaw_PBE.52 potpaw_PBE.54
I then ran the pmg config command to generate the pymatgen flavored output
pmg config -p extracted pymatgen_pot
and set the environment variables to reflect the new directory
pmg config --add PMG_VASP_PSP_DIR '/Users/paulfons/Google Drive/Vasp/pymatgen_pot'
Loading a ipython and attempting to generate VASP input fails as can be seen below:
Python 3.7.6 (default, Feb 28 2020, 08:55:06)
Type 'copyright', 'credits' or 'license' for more information
IPython 7.13.0 -- An enhanced Interactive Python. Type '?' for help.
In [2]: import pymatgen as mg
In [3]: structure = mg.Structure.from_file('POSCAR')
In [4]: structure.composition
Out[4]: Comp: Cr24 Ge24 Te72
In [5]: from pymatgen.io.vasp.sets import MPStaticSet
In [7]: static = MPStaticSet(structure)
In [8]: static.write_input(output_dir='crgete')
---------------------------------------------------------------------------
OSError Traceback (most recent call last)
<ipython-input-8-14ad379d82b3> in <module>
----> 1 static.write_input(output_dir='crgete')
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py in write_input(self, output_dir, make_dir_if_not_present, include_cif, potcar_spec, zip_output)
720 include_cif=include_cif,
721 potcar_spec=potcar_spec,
--> 722 zip_output=zip_output
723 )
724 for k, v in self.files_to_transfer.items():
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py in write_input(self, output_dir, make_dir_if_not_present, include_cif, potcar_spec, zip_output)
210 f.write(v.__str__())
211 else:
--> 212 vinput = self.get_vasp_input()
213 vinput.write_input(output_dir, make_dir_if_not_present=make_dir_if_not_present)
214
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py in get_vasp_input(self)
172 kpoints=self.kpoints,
173 poscar=self.poscar,
--> 174 potcar=self.potcar,
175 )
176
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py in potcar(self)
127 Potcar object.
128 """
--> 129 potcar = Potcar(self.potcar_symbols, functional=self.potcar_functional)
130
131 # warn if the selected POTCARs do not correspond to the chosen
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py in __init__(self, symbols, functional, sym_potcar_map)
2211 self.functional = functional
2212 if symbols is not None:
-> 2213 self.set_symbols(symbols, functional, sym_potcar_map)
2214
2215 def as_dict(self):
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py in set_symbols(self, symbols, functional, sym_potcar_map)
2318 else:
2319 for el in symbols:
-> 2320 p = PotcarSingle.from_symbol_and_functional(el, functional)
2321 self.append(p)
2322
~/.pyenv/versions/3.7.6/envs/pymatgen/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py in from_symbol_and_functional(symbol, functional)
1951 raise IOError(
1952 "You do not have the right POTCAR with functional "
-> 1953 + "{} and label {} in your VASP_PSP_DIR".format(functional, symbol)
1954 )
1955
OSError: You do not have the right POTCAR with functional PBE and label Cr_pv in your VASP_PSP_DIR
Any suggestions as to what I can try out to fix this?
**Expected behavior**
It is expected that a functional POTCAR will be generated for a Vast POTCAR setup configured using the instructions provided on the pymatgen homepage.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: [e.g. Mac,Windows,Linux]
- Version [e.g. 2019.9.16]
**Additional context**
Add any other context about the problem here.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1813/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1813/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1814
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1814/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1814/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1814/events
|
https://github.com/materialsproject/pymatgen/pull/1814
| 589,837,352
|
MDExOlB1bGxSZXF1ZXN0Mzk1Mjg0OTc3
| 1,814
|
Lammpsdata fix
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls add a test to ensure this does not occur again. I would also add a new kwarg to allow people to sort if they want.",
"@shyuep Sure. ",
"Thanks."
] | 2020-03-29T17:07:50
| 2020-03-30T13:47:24
|
2020-03-30T13:47:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR fixes #1812
Previously Structure is reordered before writing to lammps data file. This could result in unexpected site index mismatch between structure and the final outcome from lammps.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1814/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1814/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1814",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1814",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1814.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1814.patch",
"merged_at": "2020-03-30T13:47:23Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1815
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1815/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1815/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1815/events
|
https://github.com/materialsproject/pymatgen/issues/1815
| 590,105,404
|
MDU6SXNzdWU1OTAxMDU0MDQ=
| 1,815
|
Boltztrap2.py unit problem
|
{
"login": "adam5353",
"id": 62876892,
"node_id": "MDQ6VXNlcjYyODc2ODky",
"avatar_url": "https://avatars.githubusercontent.com/u/62876892?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/adam5353",
"html_url": "https://github.com/adam5353",
"followers_url": "https://api.github.com/users/adam5353/followers",
"following_url": "https://api.github.com/users/adam5353/following{/other_user}",
"gists_url": "https://api.github.com/users/adam5353/gists{/gist_id}",
"starred_url": "https://api.github.com/users/adam5353/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/adam5353/subscriptions",
"organizations_url": "https://api.github.com/users/adam5353/orgs",
"repos_url": "https://api.github.com/users/adam5353/repos",
"events_url": "https://api.github.com/users/adam5353/events{/privacy}",
"received_events_url": "https://api.github.com/users/adam5353/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Let me check that.\r\nI have a couple of edits done but not pushed yet.\r\nThank for reporting. Do not hesitate if you find something else.",
"It is great to hear that. Looking forward to your edits. Thanks.",
"I pushed a new version. \r\nPlease have a look.\r\nAny suggestion is welcome.\r\n"
] | 2020-03-30T08:46:35
| 2023-08-13T16:33:40
|
2023-08-13T16:33:40Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I was using boltztrap2 module in pymatgen. I discoverd unit problem in electrical conductivity. When I checked the source, I have not found unit conversion of kS/m. I think the unit should be S/m.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1815/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1815/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1816
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1816/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1816/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1816/events
|
https://github.com/materialsproject/pymatgen/issues/1816
| 590,708,848
|
MDU6SXNzdWU1OTA3MDg4NDg=
| 1,816
|
import error in pymatgen
|
{
"login": "srajpand87",
"id": 62912413,
"node_id": "MDQ6VXNlcjYyOTEyNDEz",
"avatar_url": "https://avatars.githubusercontent.com/u/62912413?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/srajpand87",
"html_url": "https://github.com/srajpand87",
"followers_url": "https://api.github.com/users/srajpand87/followers",
"following_url": "https://api.github.com/users/srajpand87/following{/other_user}",
"gists_url": "https://api.github.com/users/srajpand87/gists{/gist_id}",
"starred_url": "https://api.github.com/users/srajpand87/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/srajpand87/subscriptions",
"organizations_url": "https://api.github.com/users/srajpand87/orgs",
"repos_url": "https://api.github.com/users/srajpand87/repos",
"events_url": "https://api.github.com/users/srajpand87/events{/privacy}",
"received_events_url": "https://api.github.com/users/srajpand87/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @srajpand87, I don't think this is a pymatgen issue -- it's likely you're trying to use an old script.",
"Thanks for the reply.\r\nActually i was trying whole day to sort out the issue but I couldn’t.\r\nI am afraid to say do you mean old script is the pymatgen from the above link and I need to use the latest pymatgen 2020 3 13 not the one from the link?",
"I meant that the GaAs.py script might be old (and/or the OPTpy package), it might help if you shared this?",
"```\r\nfrom OPTpy import OPTflow,Structure\r\n\r\nkpt_l = [[4,4,4]] # for optical only, bad practice ther are two places where this is defined :(\r\n\r\n\r\nmodules=''\r\n\r\ntiniba_path='/global/u1/s/srajpand/panday_back_python2/tiniba-v3.2'\r\n\r\nncalc = len(kpt_l)\r\n\r\n# Common arguments for the OPTpy flows.\r\nkwargs = dict(\r\n modules=modules,\r\n dirname = './',\r\n\r\n # Executables:\r\n ABINIT='/global/u2/b/bmunozf2/abinit_cori_haswell/bin/abinit',\r\n RPMNS='/global/u1/s/srajpand/panday_back_python2/tiniba-v3.2/matrix_elements/rpmns',\r\n SET_INPUT_ALL='/global/u1/s/srajpand/panday_back_python2/tiniba-v3.2/latm/set_input_all',\r\n TETRA_METHOD_ALL='/global/u1/s/srajpand/panday_back_python2/tiniba-v3.2/latm/tetra_method_all',\r\n RKRAMER='/global/u1/s/srajpand/panday_back_python2/tiniba-v3.2/kk/rkramer',\r\n\r\n\r\n# Common variables:\r\n prefix = 'gaas', # Root name for files required by Tiniba\r\n# Structure from file:\r\n structure=Structure.from_file('GaAs.json'),\r\n\r\n# Pseudopotentials:\r\n pseudo_dir = './',\r\n pseudos = ['Ga.psp8', 'As.psp8'], # order has to coinside with .json file, e.g.,: SSGeGe or GeGeSS?\r\n\r\n ecut = 50.0, # Wavefunctions cutoff energy, 60 Ha = 120 Ry= what I used in convergence in QE\r\n nspinor = 2, # Numbers of spinorial components\r\n\r\n# Variables for density:\r\n ngkpt = [8,8,8], # k-point grid for density\r\n# nstep = 120, # it ignores this anyways\r\n# nshiftk=3,\r\n kshift = [[.5,.5,.5],[.5,0,0],[0,.5,0],[0,0,.5]], # k-point shift for density, no shifts of kpoint for GeS,ok?\r\n\r\n# Variables for momentum matrix elements and responses:\r\n nband=70, # Number of bands\r\n nbdbuf=2, # Bands in buffer, skip these as are difficult to converge (see Abinit documentation).\r\n nval_total=28, # 20 valence bands (needs to set to all valence bands). \r\n\r\n# Variables for responses:\r\n kgrid_response=[4,4,4],\r\n# Bands to include for transitions (can be less than for RPMNS)\r\n ncond=38,\r\n nval=28, #Ge val=4, S val=4 total =10x2 =20 works\r\n# Response to calculate, see Doc. in responses.py\r\n response=21, #46 for sigma, 21 for SHG\r\n components=[\"xyz\"],\r\n\r\n # These are the default parameters for the MPI runner.\r\n # Please adapt them to your needs.\r\n nproc = 32,\r\n nproc_per_node = '',\r\n mpirun = 'srun -c 2 --cpu_bind=cores',\r\n nproc_flag = '',\r\n nproc_per_node_flag = '',\r\n mpi_extra_vars='',\r\n )\r\n\r\ndirnames = list()\r\n\r\nfor icalc in range(ncalc):\r\n\r\n kgrid_response = kpt_l[icalc]\r\n\r\n dirname = 'k{:0=2}'.format(icalc+1)\r\n dirnames.append(dirname)\r\n\r\n kwargs.update(\r\n dirname = dirname,\r\n kgrid_response=kgrid_response,\r\n )\r\n\r\n\r\n flow = OPTflow(**kwargs)\r\n flow.write()\r\n```\r\n\r\nThis is the content of the file.\r\nThank you\r\n\r\n\r\n\r\n\r\n",
"Thanks for sharing @srajpand87, I would advise you talk to whoever develops `OPTflow`, I don't think this is a pymatgen issue.",
"Yes, it looks like the code was written against pymatgen around three years ago: https://github.com/trangel/OPTpy\r\n\r\nYou could try installing an old version of pymatgen (e.g. `4.0`) but I would advise against this since many bugs have been fixed since then. It would be better for the original script to be updated.\r\n",
"Thanks for the suggestion.\r\nOkay I will contact Tonatiuh Rangel to update this script.",
"Hi mkhorton,\r\nI made an inquiry with the one who wrote OPTpy and above file.\r\nHe suggested me to use an old version of pymatgen like you said to me.\r\nI am wondering how can I get v2018.x pymatgen as i am using python 2.7 and this python only take into account v2018.x pymatgen.\r\nPreviously i was using git clone https://github.com/materialsproject/pymatgen.git which only upload 2020 3 13 version of pymatgen.\r\nI want to get something like git clone similar link above to upload 2018 version of pymatgen.\r\nThank you\r\nSuman",
"Pls refer to standard python installation instructions using pip to get a specific version. Sorry, but we cannot be providing this kind of support for basic python installation questions.\nOn Apr 1, 2020, 12:39 PM -0700, srajpand87 <notifications@github.com>, wrote:\n> Hi mkhorton,\n> I made an inquiry with the one who wrote OPTpy and above file.\n> He suggested me to use an old version of pymatgen like you said to me.\n> I am wondering how can I get v2018.x pymatgen as i am using python 2.7 and this python only take into account v2018.x pymatgen.\n> Previously i was using git clone https://github.com/materialsproject/pymatgen.git which only upload 2020 3 13 version of pymatgen.\n> I want to get something like git clone similar link above to upload 2018 version of pymatgen.\n> Thank you\n> Suman\n> —\n> You are receiving this because you are subscribed to this thread.\n> Reply to this email directly, view it on GitHub, or unsubscribe.\n",
"Really apologized for such a naive question.\r\nI used following commands to install conda and pip.\r\nmodule load python/2.7-anaconda-4.4\r\nconda create -n myenv\r\nsource activate myenv\r\nconda uninstall numpy\r\npip install numpy==1.16.4\r\nNo error using above commands\r\nBut while trying to install pymatgen using command\r\npip install pymatgen==2018.2.13 gave me an error\r\nBuilding wheel for pymatgen (setup.py) ... error\r\n ERROR: Command errored out with exit status 1:\r\n command: /global/homes/s/srajpand/.conda/envs/myenv/bin/python -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '\"'\"'/tmp/pip-install-1gjmt1nc/pymatgen/setup.py'\"'\"'; __file__='\"'\"'/tmp/pip-install-1gjmt1nc/pymatgen/setup.py'\"'\"';f=getattr(tokenize, '\"'\"'open'\"'\"', open)(__file__);code=f.read().replace('\"'\"'\\r\\n'\"'\"', '\"'\"'\\n'\"'\"');f.close();exec(compile(code, __file__, '\"'\"'exec'\"'\"'))' bdist_wheel -d /tmp/pip-wheel-_mbrjwet\r\n cwd: /tmp/pip-install-1gjmt1nc/pymatgen/\r\n\r\nDoes it mean not allowing me to install pymatgen 2018.2.13?",
"Hi @srajpand87, as @shyuep said we really can't help with this kind of issue -- I'd advise reading more about Python and Python packaging in general. Historical versions of pymatgen are available on [PyPI](https://pypi.org/project/pymatgen/#history) and via git using `git checkout`.\r\n\r\nFor this specific issue I couldn't say--you're using an old version of Python, `2.7`, which we don't support anymore since the Python community has almost exclusively moved to 3+.\r\n\r\nMore broadly, whatever calculation you're trying to run, you are likely to have some more success if you use a more recent version of whatever code you're trying to use, and study more closely the specifics of the calculation you're trying to run. Good luck!",
"Okay I will follow your directions and thank you."
] | 2020-03-31T01:13:40
| 2020-04-01T22:27:23
|
2020-04-01T19:43:55Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
I installed pymatgen from github (https://github.com/materialsproject/pymatgen.git) .
When i was trying to run some python file name GaAs.py i got an error which is as follows
File "/global/homes/s/srajpand/.conda/envs/myenv/lib/python3.8/site-packages/OPTpy-4.2.2-py3.8.egg/OPTpy/utils/response.py", line 205
self.runscript.append("cp {0} {1}".format(origin,dest))
ImportError: cannot import name 'units' from 'pymatgen' (/global/homes/s/srajpand/.conda/envs/myenv/lib/python3.8/site-packages/pymatgen-2020.3.13-py3.8-linux-x86_64.egg/pymatgen/__init__.py)
Could you please let me know how to sort out this issue?
Thank you
Suman Raj Panday
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1816/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1816/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1817
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1817/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1817/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1817/events
|
https://github.com/materialsproject/pymatgen/pull/1817
| 592,168,100
|
MDExOlB1bGxSZXF1ZXN0Mzk3MjAyNTkw
| 1,817
|
Tem Calculator
|
{
"login": "welltemperedpaprika",
"id": 38155558,
"node_id": "MDQ6VXNlcjM4MTU1NTU4",
"avatar_url": "https://avatars.githubusercontent.com/u/38155558?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/welltemperedpaprika",
"html_url": "https://github.com/welltemperedpaprika",
"followers_url": "https://api.github.com/users/welltemperedpaprika/followers",
"following_url": "https://api.github.com/users/welltemperedpaprika/following{/other_user}",
"gists_url": "https://api.github.com/users/welltemperedpaprika/gists{/gist_id}",
"starred_url": "https://api.github.com/users/welltemperedpaprika/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/welltemperedpaprika/subscriptions",
"organizations_url": "https://api.github.com/users/welltemperedpaprika/orgs",
"repos_url": "https://api.github.com/users/welltemperedpaprika/repos",
"events_url": "https://api.github.com/users/welltemperedpaprika/events{/privacy}",
"received_events_url": "https://api.github.com/users/welltemperedpaprika/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the quick turnaround on this :-)\r\n\r\nBefore merging, have you also tested a few random structures with random directions? I'm wondering what the test coverage is like here -- I only tested myself along some high symmetry directions with some common structures.",
"> Thanks for the quick turnaround on this :-)\r\n> \r\n> Before merging, have you also tested a few random structures with random directions? I'm wondering what the test coverage is like here -- I only tested myself along some high symmetry directions with some common structures.\r\n\r\nThank you!\r\nCalculations are based on the following indexing procedure:\r\nhttps://www.doitpoms.ac.uk/tlplib/diffraction-patterns/maths.php\r\nI have tested individual formulas of calculation according to the international table of crystallography. I have also generated 600 + patterns with random beam directions of different iron oxides obtained from the materials project, and all of them was able to generate a pattern. There are some directions where the pattern only appears to be a line of dots, however. \r\nI would assume the results for high symmetry system and directions are most accurate (which I was able to compare to some reference pattern online). ",
"Woo, doitpoms 🎉 I worked on that website myself as an undergrad many years ago.\r\n\r\n> I have also generated 600 + patterns with random beam directions of different iron oxides obtained from the materials project, and all of them was able to generate a pattern. \r\n\r\nGreat!\r\n\r\n> There are some directions where the pattern only appears to be a line of dots, however.\r\n\r\nThis would be expected too.\r\n\r\nThere's commercial software out there that's very good called SingleCrystal from the makers of CrystalMaker, if you wanted to do further testing I think they offer free trials, otherwise this sounds great."
] | 2020-04-01T19:58:39
| 2020-04-02T18:52:48
|
2020-04-02T18:52:48Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Include a summary of major changes in bullet points:
Edits made to Frank Wan's previous TEM calculator, a project under Shyam's direction:
- Changed method get_interplanar_spacing to call lattice.d_hkl rather than separately calculating for different crystal systems.
- Vectorized method generate_points to be using numpy arrays rather than nested for loops.
- Vectorized methods to reduce for loops when looping over dictionaries, if possible.
- Changed original implementation of prettytable to pandas dataframe.
- Removed TEM Dot object to a more lightweight named tuple.
- Updated structural factors. Matches international table of crystallography.
---------------------------------------
Updated March 2020:
- Fixed a bug where beam direction of 110 and -1-10 are not generating any patterns.
- Added a new method to more accurately calculate interplanar angles, accounting for different crystal systems.
- Recalculated diffraction pattern using Moore-Penrose inverse.
Request submitted by Jason Liang, student working with Shyam.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1817/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1817/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1817",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1817",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1817.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1817.patch",
"merged_at": "2020-04-02T18:52:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1818
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1818/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1818/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1818/events
|
https://github.com/materialsproject/pymatgen/pull/1818
| 592,171,052
|
MDExOlB1bGxSZXF1ZXN0Mzk3MjA1MDAz
| 1,818
|
Optimize CubicSupercellTransformation for diagonal transformations
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @utf.\r\n\r\nOut of interest, is there an easy way to modify Cubic to give you a square slab instead? e.g. square in x, y but a small z -- this would be useful for some folks doing (S)TEM image simulation, square slabs are useful as multi slice inputs",
"I think that should be possible. I wonder if @rees-c (original author) has any thoughts on how to do that?",
"Once tests pass this should be ready to merge.",
"Thanks @utf and @rees-c",
"Ok, thanks @rees-c, no worries about the 45deg angles etc, this will\nalready be useful as is.\n\nOn Mon, Apr 20, 2020 at 20:56, rees-c <notifications@github.com> wrote:\n\n> *@rees-c* commented on this pull request.\n> ------------------------------\n>\n> In pymatgen/transformations/advanced_transformations.py\n> <https://github.com/materialsproject/pymatgen/pull/1818#discussion_r411849536>\n> :\n>\n> > while sc_not_found:\n> - if self.force_diagonal:\n> - # Update trans_mat (with diagonal constraint)\n> - trans_mat_diagonal += trans_mat_diagonal_update\n> - self.transformation_matrix = np.diag(trans_mat_diagonal)\n> -\n> - else:\n> - # Update trans_mat (without diagonal constraint)\n> - target_sc_lat_vecs = np.eye(3, 3) * target_sc_size\n> -\n> - self.transformation_matrix = (\n> - np.linalg.inv(lat_vecs) @ target_sc_lat_vecs\n> - )\n> + target_sc_lat_vecs = np.eye(3, 3) * target_sc_size\n>\n> Hi @mkhorton <https://github.com/mkhorton>! I see now that my code\n> rounded every entry of each zero column instead of picking a random\n> entry... if there was a good reason for why I would've done that, it has\n> since escaped my brain haha.\n>\n> With regard to the angles, it's my understanding that's it's not always\n> possible to get perfect 90 degree angles (due to the requirement for all\n> entries of the transformation to be integers for preserving periodicity).\n> Not sure if that's why you're getting a 45 degree angle but I'll take a\n> closer look at your test implementation and let you know if I spot anything.\n>\n> —\n> You are receiving this because you were mentioned.\n>\n>\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/pull/1818#discussion_r411849536>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAWWWRFM23KYUZGLYLIC2HTRNUKNVANCNFSM4LZQMMZA>\n> .\n>\n"
] | 2020-04-01T20:04:17
| 2020-04-21T04:26:54
|
2020-04-15T19:31:57Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
`CubicSupercellTransformation` had some unnecessary complexity for diagonal only transformations. This PR implements a more optimised method.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1818/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1818/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1818",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1818",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1818.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1818.patch",
"merged_at": "2020-04-15T19:31:57Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1819
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1819/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1819/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1819/events
|
https://github.com/materialsproject/pymatgen/pull/1819
| 592,173,942
|
MDExOlB1bGxSZXF1ZXN0Mzk3MjA3NDAx
| 1,819
|
Fix BrunnerNN methods
|
{
"login": "utf",
"id": 1330638,
"node_id": "MDQ6VXNlcjEzMzA2Mzg=",
"avatar_url": "https://avatars.githubusercontent.com/u/1330638?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/utf",
"html_url": "https://github.com/utf",
"followers_url": "https://api.github.com/users/utf/followers",
"following_url": "https://api.github.com/users/utf/following{/other_user}",
"gists_url": "https://api.github.com/users/utf/gists{/gist_id}",
"starred_url": "https://api.github.com/users/utf/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/utf/subscriptions",
"organizations_url": "https://api.github.com/users/utf/orgs",
"repos_url": "https://api.github.com/users/utf/repos",
"events_url": "https://api.github.com/users/utf/events{/privacy}",
"received_events_url": "https://api.github.com/users/utf/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @utf!"
] | 2020-04-01T20:09:54
| 2020-04-02T18:52:05
|
2020-04-02T18:51:55Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Two of the BrunnerNN methods were implemented incorrectly leading to poor performance. This PR fixes their implementation to be in line with: https://scripts.iucr.org/cgi-bin/paper?S0567739477000461
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1819/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1819/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1819",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1819",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1819.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1819.patch",
"merged_at": "2020-04-02T18:51:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1820
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1820/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1820/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1820/events
|
https://github.com/materialsproject/pymatgen/pull/1820
| 593,064,148
|
MDExOlB1bGxSZXF1ZXN0Mzk3OTI2MDAz
| 1,820
|
VolumetricData speedup
|
{
"login": "mturiansky",
"id": 8866816,
"node_id": "MDQ6VXNlcjg4NjY4MTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/8866816?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mturiansky",
"html_url": "https://github.com/mturiansky",
"followers_url": "https://api.github.com/users/mturiansky/followers",
"following_url": "https://api.github.com/users/mturiansky/following{/other_user}",
"gists_url": "https://api.github.com/users/mturiansky/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mturiansky/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mturiansky/subscriptions",
"organizations_url": "https://api.github.com/users/mturiansky/orgs",
"repos_url": "https://api.github.com/users/mturiansky/repos",
"events_url": "https://api.github.com/users/mturiansky/events{/privacy}",
"received_events_url": "https://api.github.com/users/mturiansky/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"This is great, thanks @mturiansky! Optimizations always very welcome :-)\r\n\r\nNow there's the question of why large arrays of floats are stored in ascii to start with ... perhaps time to kindly ask the VASP devs for a native hdf5 output.",
"> Now there's the question of why large arrays of floats are stored in ascii to start with ...\r\n\r\nbut ascii has _\\~compatibility\\~_...\r\n\r\n/s\r\n\r\n",
"And also the question of why the ELFCAR and AECCAR are apparently printed in a lower precision. Perhaps there's a good technical reason for this, I'm not sure."
] | 2020-04-03T04:16:13
| 2020-04-03T23:17:36
|
2020-04-03T05:23:53Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Got frustrated while reading a 1.5 GB file, so I tried to speed it up. Changed a couple lines in VolumetricData.parse_file, and here's what I got on my computer:
* test_files/CHGCAR.Fe3O4: 655 ms -> 368 ms (1.78x speedup)
* test_files/CHGCAR.NiO_SOC.gz: 779 ms -> 495 ms (1.57x)
* test_files/LOCPOT: 28.8 ms -> 16.1 ms (1.79x)
* test_files/ELFCAR.gz: 40.6 ms -> 22.7 ms (1.79x)
* 384 MB CHGCAR: 19.8 s -> 10.9 s (1.82x)
* 1.5 GB AECCAR: 81 s -> 48.3 s (1.68x)
<img width=400 alt="It ain't much, but it's honest work." src="https://i.kym-cdn.com/entries/icons/original/000/028/021/work.jpg"/>
## Additional dependencies introduced (if any)
None.
## TODO (if any)
None.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1820/reactions",
"total_count": 1,
"+1": 0,
"-1": 0,
"laugh": 1,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1820/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1820",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1820",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1820.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1820.patch",
"merged_at": "2020-04-03T05:23:53Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1821
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1821/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1821/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1821/events
|
https://github.com/materialsproject/pymatgen/issues/1821
| 593,132,857
|
MDU6SXNzdWU1OTMxMzI4NTc=
| 1,821
|
Connection problem with MPRester
|
{
"login": "liuyuhang599",
"id": 29992375,
"node_id": "MDQ6VXNlcjI5OTkyMzc1",
"avatar_url": "https://avatars.githubusercontent.com/u/29992375?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/liuyuhang599",
"html_url": "https://github.com/liuyuhang599",
"followers_url": "https://api.github.com/users/liuyuhang599/followers",
"following_url": "https://api.github.com/users/liuyuhang599/following{/other_user}",
"gists_url": "https://api.github.com/users/liuyuhang599/gists{/gist_id}",
"starred_url": "https://api.github.com/users/liuyuhang599/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/liuyuhang599/subscriptions",
"organizations_url": "https://api.github.com/users/liuyuhang599/orgs",
"repos_url": "https://api.github.com/users/liuyuhang599/repos",
"events_url": "https://api.github.com/users/liuyuhang599/events{/privacy}",
"received_events_url": "https://api.github.com/users/liuyuhang599/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The likelihood is that the Materials PRoject is down. You will need to contact the Materials Project people.",
"Yes, an error like that is usually a connection issue, not a pymatgen issue.\r\n\r\nWe haven't registered any connection issues with Materials Project since March 6th (35 minutes), though I would not be surprised if we might likely have some issues in the future due to the shelter-in-place. So far, we have mostly been able to work as normal however.\r\n\r\nI notice this is a Windows error, if it persists, perhaps try seeing if the `requests` library is working at all in your installation and/or try an alternative installation (Google Colab is a great resource to try pymatgen, you can `!pip install pymatgen` in the cloud using any Google account)."
] | 2020-04-03T06:32:10
| 2020-04-03T20:05:45
|
2020-04-03T13:02:21Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I got this error message while downloading data from the Materials Project using MPRester.
MPRestError: ('Connection aborted.', OSError("(10051, 'WSAENETUNREACH')"))
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1821/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1821/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1822
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1822/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1822/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1822/events
|
https://github.com/materialsproject/pymatgen/pull/1822
| 596,662,578
|
MDExOlB1bGxSZXF1ZXN0NDAwOTE3NTM2
| 1,822
|
Gau
|
{
"login": "gVallverdu",
"id": 5417018,
"node_id": "MDQ6VXNlcjU0MTcwMTg=",
"avatar_url": "https://avatars.githubusercontent.com/u/5417018?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/gVallverdu",
"html_url": "https://github.com/gVallverdu",
"followers_url": "https://api.github.com/users/gVallverdu/followers",
"following_url": "https://api.github.com/users/gVallverdu/following{/other_user}",
"gists_url": "https://api.github.com/users/gVallverdu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/gVallverdu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/gVallverdu/subscriptions",
"organizations_url": "https://api.github.com/users/gVallverdu/orgs",
"repos_url": "https://api.github.com/users/gVallverdu/repos",
"events_url": "https://api.github.com/users/gVallverdu/events{/privacy}",
"received_events_url": "https://api.github.com/users/gVallverdu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"It seems there is an error because of a whitespace after an open `[` .\r\nLet me know if you want me to fix this.",
"Yes pls. All linting has to pass as well as the tests.",
"Many thanks."
] | 2020-04-08T15:24:12
| 2020-04-08T21:33:55
|
2020-04-08T21:33:50Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Improve parsing of structures if multiple optimization, for example in the case of relaxed scan.
* In case of scan the attribute `opt_structures` store optimized structure contrary to the old `structures` attribute who store all structures
* Fix a bug in read_scan() function linked to a change between g09 and g16.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1822/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1822/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1822",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1822",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1822.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1822.patch",
"merged_at": "2020-04-08T21:33:50Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1823
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1823/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1823/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1823/events
|
https://github.com/materialsproject/pymatgen/issues/1823
| 597,606,899
|
MDU6SXNzdWU1OTc2MDY4OTk=
| 1,823
|
Pourbaix diagrams: AqueousCorrection unexpected behavior re: hydrogen energy
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@JosephMontoya-TRI do you see any problem with just setting `compatible_only=True` as a temporary fix?",
"@JosephMontoya-TRI as part of the longer-term solution, we are thinking of moving the `AqueousCompatibility` scheme to a standalone `Compatibility` class that doesn't require an underlying `Correction` class. That way we could start with already-corrected `ComputedEntry` (e.g., from `MaterialsProjectCompatibility`) and apply the aqueous adjustments directly in the `Compatibility` class, without having to go through the `process_entry` -> `get_correction` -> `get_corrections_dict` flow that causes this bug. See https://github.com/awvio/pymatgen/compare/master...rkingsbury:abstractcompat?expand=1. Do you have any thoughts on this approach?",
"That sounds reasonable to me, my understanding is that writing a custom process_entries method will allow you to implement the sequential logic in a more transparent way?",
"> That sounds reasonable to me, my understanding is that writing a custom process_entries method will allow you to implement the sequential logic in a more transparent way?\r\n\r\nYes, exactly. Also, what do you think about the `compatible_only=True` vs. `False` setting in `get_pourbaix_entries` ?",
"Also the temporary fix sounds fine to me assuming it works. It won't mess up O2 will it?",
"> Also the temporary fix sounds fine to me assuming it works. It won't mess up O2 will it?\r\n\r\nI will verify, but it shouldn't.",
"Update: while #1826 will fix this bug by preventing double-counting of corrections, the pymatgen Pourbaix diagram for Na will in fact not show a hydride. This is a result of a more accurate way of fitting the hydrogen energy that we introduced with the new `MaterialsProjectAqueousCompatibility` class.",
"@rkingsbury sorry why is the hydride no longer present?",
"> \r\n> \r\n> @rkingsbury sorry why is the hydride no longer present?\r\n\r\nAs part of the new `MaterialsProjectAqueousCompatibility` class, we recalculated the chemical potential of H2 (previously -3.60 eV/H and stored in a .yaml file). Due to an updated DFT energy of water in our database and correcting an error in the way the value was fitted before, the new chemical potential is -4.05 eV/H. This more negative hydrogen chemical potential cases most hydrides not to be on the hull. The effect is similar to what was happening previously because of this double-counting bug.\r\n\r\nI'll note that some systems like Li, Ca, Sr, and Ba do gain hydrides after fixing this bug. It's just that Na does not."
] | 2020-04-09T23:17:48
| 2020-05-05T22:45:43
|
2020-05-05T22:45:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
@JosephMontoya-TRI and I recently discovered a bug in the `AqueousCorrection` class (used by `MaterialsProjectAqueousCompatibility` and `MITProjectAqueousCompatibility`) that causes unexpected behavior when correcting the energy of aqueous H2. As a result, the hydrogen energy may vary depending on whether or not the `ComputedEntry` for H2 already has a correction applied (e.g., when downloading from MPRester).
**To Reproduce**
Below are three different ways of generating entries for Pourbaix diagram construction. These should all result in the same energy for H2, but they do not. Note that H2 should always be assigned an energy of -3.6019 eV/atom (the value in the `MPCompatibility.yaml` file.
**Method 1: Manually process MPRester entries**
```
compat = MaterialsProjectAqueousCompatibility()
with MPRester() as m:
ion_ref_entries = m.get_entries_in_chemsys(["O", "H"])
ion_ref_entries = compat.process_entries(ion_ref_entries)
```
(print statements omitted for clarity)
```
Entry ID Correction E_per_atom Corrections
mp-632291 -0.2112 -3.6019 {'MP Gas Correction': 0.30189724724000033}
mp-1188177 -0.2798 -3.6019 {'MP Gas Correction': 1.3179495979200055}
mp-850274 -0.2139 -3.6019 {'MP Gas Correction': 0.2964079872400003}
```
**Method 2: Remove corrections, then manually reprocess MPRester entries**
```
compat = MaterialsProjectAqueousCompatibility()
with MPRester() as m:
ion_ref_entries = m.get_entries_in_chemsys(["O", "H"])
for e in ion_ref_entries:
e.correction=0
ion_ref_entries = compat.process_entries(ion_ref_entries)
```
```
Entry ID Correction E_per_atom Corrections
mp-632291 -0.0603 -3.4509 {'MP Gas Correction': 0.30189724724000033, 'MP Aqueous Correction': -0.30189724724000033}
mp-1188177 -0.1974 -3.5195 {'MP Gas Correction': 1.3179495979200055, 'MP Aqueous Correction': -1.3179495979200055}
mp-850274 -0.0657 -3.4537 {'MP Gas Correction': 0.2964079872400003, 'MP Aqueous Correction': -0.2964079872400003}
```
**Method 3: compatible_only=False, then manually reprocess MPRester entries**
```
compat = MaterialsProjectAqueousCompatibility()
with MPRester() as m:
ion_ref_entries = m.get_entries_in_chemsys(["O", "H"], compatible_only=False)
for e in ion_ref_entries:
e.correction=0
ion_ref_entries = compat.process_entries(ion_ref_entries)
```
```
Entry ID Correction E_per_atom Corrections
mp-632291 -0.0603 -3.4509 {'MP Gas Correction': 0.30189724724000033, 'MP Aqueous Correction': -0.30189724724000033}
mp-1188177 -0.1974 -3.5195 {'MP Gas Correction': 1.3179495979200055, 'MP Aqueous Correction': -1.3179495979200055}
mp-850274 -0.0657 -3.4537 {'MP Gas Correction': 0.2964079872400003, 'MP Aqueous Correction': -0.2964079872400003}
```
Method 1 produces the correct energy, while Method 2 and 3 fail. Method 3 (which parallels the implementation in `get_pourbaix_entries` is equivalent to Method 2.
The reason Method 1 works is that the `ComputedEntry` returned by MPRester already has a non-zero energy correction. `AqueousCorrection` requires the energy of H2 to be already corrected in order to compute the correct final energy, but this is not guaranteed if you process entries "from scratch", as shown in Method 2 or set `compatible_only=False` as in Method 3.
Because of this bug, Pourbaix diagrams generated via `.get_pourbaix_entries` slightly understabilize hydrogen, which can result in hydrides disappearing from the diagrams. For example:
Method 2 /3:
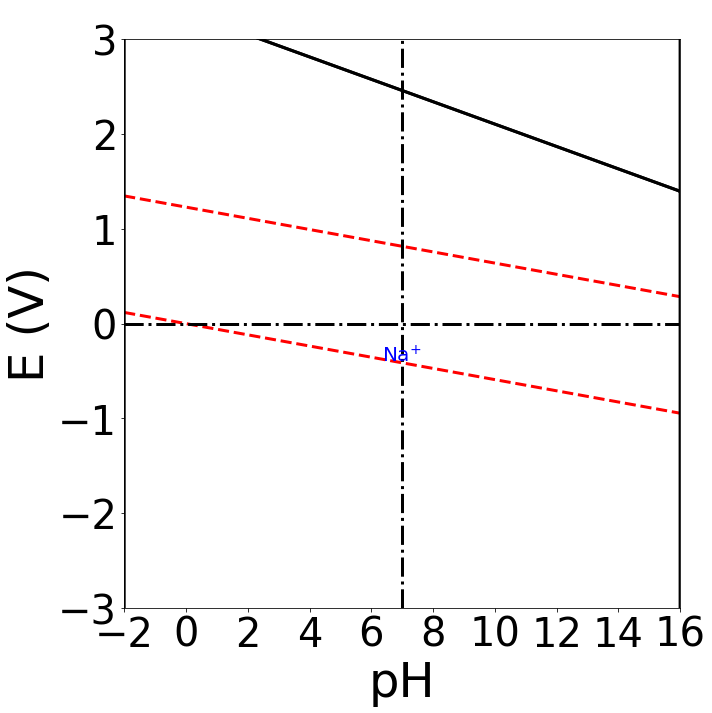
vs. the more correct Method 1:
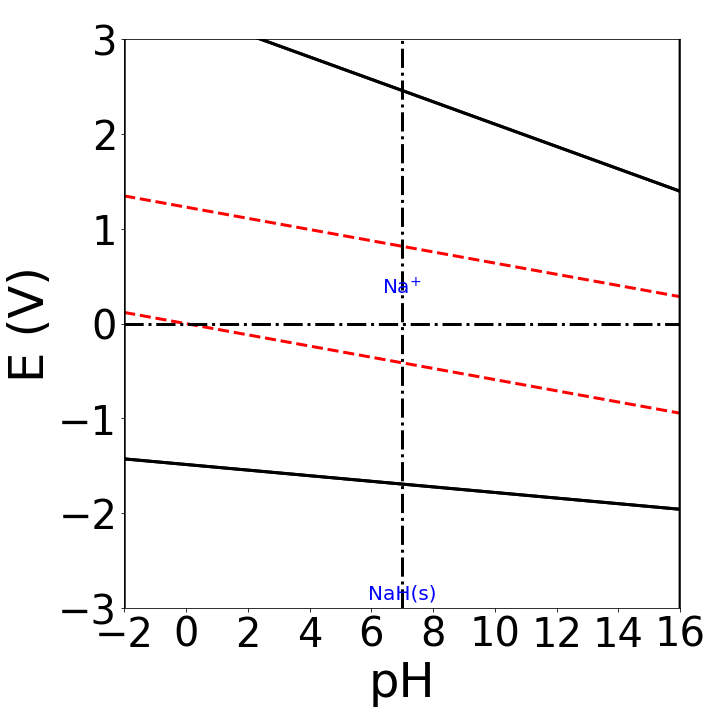
**Additional context**
We are developing a fix to correct this and also to ensure that `MaterialsProjectAqueousCompatibility` is fully compatible with a forthcoming new set of energy corrections (See #1558 ).
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1823/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1823/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1824
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1824/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1824/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1824/events
|
https://github.com/materialsproject/pymatgen/issues/1824
| 598,370,384
|
MDU6SXNzdWU1OTgzNzAzODQ=
| 1,824
|
Pip installation error help!
|
{
"login": "LaboratoryofDexter",
"id": 41708721,
"node_id": "MDQ6VXNlcjQxNzA4NzIx",
"avatar_url": "https://avatars.githubusercontent.com/u/41708721?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/LaboratoryofDexter",
"html_url": "https://github.com/LaboratoryofDexter",
"followers_url": "https://api.github.com/users/LaboratoryofDexter/followers",
"following_url": "https://api.github.com/users/LaboratoryofDexter/following{/other_user}",
"gists_url": "https://api.github.com/users/LaboratoryofDexter/gists{/gist_id}",
"starred_url": "https://api.github.com/users/LaboratoryofDexter/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/LaboratoryofDexter/subscriptions",
"organizations_url": "https://api.github.com/users/LaboratoryofDexter/orgs",
"repos_url": "https://api.github.com/users/LaboratoryofDexter/repos",
"events_url": "https://api.github.com/users/LaboratoryofDexter/events{/privacy}",
"received_events_url": "https://api.github.com/users/LaboratoryofDexter/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @LaboratoryofDexter, the error is this:\r\n\r\n```\r\n error: Microsoft Visual C++ 14.0 is required. Get it with \"Build Tools for Visual Studio\": https://visualstudio.microsoft.com/downloads/\r\n```\r\n\r\nMany Python codes (including pymatgen) have C extensions as an optimization to make certain code perform faster.\r\n\r\nI would recommend you use `conda` instead of `pip` to install pymatgen:\r\n\r\n```\r\nconda install --channel conda-forge pymatgen\r\n```\r\n\r\nThis side-steps the issue since `conda` should perform the compilation for you so you don't need to install your own compiler.",
"Thanks so much for your help, mkhorton.\r\nI installed Anaconda now, and installed the Pymatgen in its CMD.exe prompt.\r\nFor common Python, the Visual Studio installation takes some time and I will wait to see whether it also works or not.\r\nThanks again for your sincerely help!",
"No problem. For the next release of pymatgen we will start including pre-built binaries for all platforms on PyPI/pip, so hopefully in future this will be a bit easier even if not using conda!"
] | 2020-04-11T23:15:21
| 2020-04-22T21:22:44
|
2020-04-11T23:39:34Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I just installed the Python 3.7.7 version and when I used "pip install pymatgen" to install pymatgen, but the terminal gave out error information as:
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Collecting pymatgen
Using cached pymatgen-2020.4.2.tar.gz (2.7 MB)
Requirement already satisfied: numpy>=1.14.3 in c:\program files\python37\lib\site-packages (from pymatgen) (1.18.2)
Requirement already satisfied: requests in c:\program files\python37\lib\site-packages (from pymatgen) (2.23.0)
Requirement already satisfied: ruamel.yaml>=0.15.6 in c:\program files\python37\lib\site-packages (from pymatgen) (0.16.10)
Requirement already satisfied: monty>=3.0.2 in c:\program files\python37\lib\site-packages (from pymatgen) (3.0.2)
Requirement already satisfied: scipy>=1.0.1 in c:\program files\python37\lib\site-packages (from pymatgen) (1.4.1)
Requirement already satisfied: pydispatcher>=2.0.5 in c:\program files\python37\lib\site-packages (from pymatgen) (2.0.5)
Requirement already satisfied: tabulate in c:\program files\python37\lib\site-packages (from pymatgen) (0.8.7)
Collecting spglib>=1.9.9.44
Using cached spglib-1.14.1.post0.tar.gz (694 kB)
Collecting networkx>=2.2
Using cached networkx-2.4-py3-none-any.whl (1.6 MB)
Collecting matplotlib>=1.5
Using cached matplotlib-3.2.1-cp37-cp37m-win_amd64.whl (9.2 MB)
Collecting palettable>=3.1.1
Using cached palettable-3.3.0-py2.py3-none-any.whl (111 kB)
Collecting sympy
Using cached sympy-1.5.1-py2.py3-none-any.whl (5.6 MB)
Collecting pandas
Using cached pandas-1.0.3-cp37-cp37m-win_amd64.whl (8.7 MB)
Collecting plotly>=4.5.0
Using cached plotly-4.6.0-py2.py3-none-any.whl (7.1 MB)
Requirement already satisfied: certifi>=2017.4.17 in c:\program files\python37\lib\site-packages (from requests->pymatgen) (2020.4.5.1)
Requirement already satisfied: chardet<4,>=3.0.2 in c:\program files\python37\lib\site-packages (from requests->pymatgen) (3.0.4)
Requirement already satisfied: idna<3,>=2.5 in c:\program files\python37\lib\site-packages (from requests->pymatgen) (2.9)
Requirement already satisfied: urllib3!=1.25.0,!=1.25.1,<1.26,>=1.21.1 in c:\program files\python37\lib\site-packages (from requests->pymatgen) (1.25.8)
Requirement already satisfied: ruamel.yaml.clib>=0.1.2; platform_python_implementation == "CPython" and python_version < "3.9" in c:\program files\python37\lib\site-packages (from ruamel.yaml>=0.15.6->pymatgen) (0.2.0)
Collecting decorator>=4.3.0
Using cached decorator-4.4.2-py2.py3-none-any.whl (9.2 kB)
Collecting python-dateutil>=2.1
Using cached python_dateutil-2.8.1-py2.py3-none-any.whl (227 kB)
Collecting pyparsing!=2.0.4,!=2.1.2,!=2.1.6,>=2.0.1
Using cached pyparsing-2.4.7-py2.py3-none-any.whl (67 kB)
Collecting cycler>=0.10
Using cached cycler-0.10.0-py2.py3-none-any.whl (6.5 kB)
Collecting kiwisolver>=1.0.1
Using cached kiwisolver-1.2.0-cp37-none-win_amd64.whl (57 kB)
Processing c:\users\dexter\appdata\local\pip\cache\wheels\e2\46\78\e78f76c356bca9277368f1f97a31b37a8cb937176d9511af31\mpmath-1.1.0-py3-none-any.whl
Collecting pytz>=2017.2
Using cached pytz-2019.3-py2.py3-none-any.whl (509 kB)
Collecting six
Using cached six-1.14.0-py2.py3-none-any.whl (10 kB)
Processing c:\users\dexter\appdata\local\pip\cache\wheels\f9\8d\8d\f6af3f7f9eea3553bc2fe6d53e4b287dad18b06a861ac56ddf\retrying-1.3.3-py3-none-any.whl
Building wheels for collected packages: pymatgen, spglib
Building wheel for pymatgen (setup.py) ... error
ERROR: Command errored out with exit status 1:
command: 'c:\program files\python37\python.exe' -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\pymatgen\\setup.py'"'"'; __file__='"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\pymatgen\\setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' bdist_wheel -d 'C:\Users\Dexter\AppData\Local\Temp\pip-wheel-a62s67jp'
cwd: C:\Users\Dexter\AppData\Local\Temp\pip-install-q2sl7tjc\pymatgen\
Complete output (443 lines):
running bdist_wheel
running build
running build_py
creating build
creating build\lib.win-amd64-3.7
creating build\lib.win-amd64-3.7\pymatgen
copying pymatgen\dao.py -> build\lib.win-amd64-3.7\pymatgen
copying pymatgen\__init__.py -> build\lib.win-amd64-3.7\pymatgen
creating build\lib.win-amd64-3.7\pymatgen\alchemy
copying pymatgen\alchemy\filters.py -> build\lib.win-amd64-3.7\pymatgen\alchemy
copying pymatgen\alchemy\materials.py -> build\lib.win-amd64-3.7\pymatgen\alchemy
copying pymatgen\alchemy\transmuters.py -> build\lib.win-amd64-3.7\pymatgen\alchemy
copying pymatgen\alchemy\__init__.py -> build\lib.win-amd64-3.7\pymatgen\alchemy
creating build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\adsorption.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\aflow_prototypes.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\bond_dissociation.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\bond_valence.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\cost.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\diffusion_analyzer.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\dimensionality.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\energy_models.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\eos.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\ewald.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\excitation.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\find_dimension.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\fragmenter.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\functional_groups.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\graphs.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\hhi.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\interface.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\interface_reactions.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\local_env.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\molecule_matcher.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\molecule_structure_comparator.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\nmr.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\path_finder.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\phase_diagram.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\piezo.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\piezo_sensitivity.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\pourbaix_diagram.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\quasiharmonic.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\reaction_calculator.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\structure_analyzer.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\structure_matcher.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\substrate_analyzer.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\surface_analysis.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\thermochemistry.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\transition_state.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\wulff.py -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis
creating build\lib.win-amd64-3.7\pymatgen\apps
copying pymatgen\apps\__init__.py -> build\lib.win-amd64-3.7\pymatgen\apps
creating build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\feff_input_generation.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\feff_plot_cross_section.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\feff_plot_dos.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\gaussian_analyzer.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\get_environment.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_analyze.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_config.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_plot.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_potcar.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_query.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\pmg_structure.py -> build\lib.win-amd64-3.7\pymatgen\cli
copying pymatgen\cli\__init__.py -> build\lib.win-amd64-3.7\pymatgen\cli
creating build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\aconvasp_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\bader_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\critic2_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\enumlib_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\gulp_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\mcsqs_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\vampire_caller.py -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\command_line\__init__.py -> build\lib.win-amd64-3.7\pymatgen\command_line
creating build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\bonds.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\composition.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\ion.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\lattice.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\libxcfunc.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\molecular_orbitals.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\operations.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\periodic_table.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\sites.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\spectrum.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\structure.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\surface.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\tensors.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\trajectory.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\units.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\xcfunc.py -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\__init__.py -> build\lib.win-amd64-3.7\pymatgen\core
creating build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\bandstructure.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\boltztrap.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\boltztrap2.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\cohp.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\core.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\dos.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\plotter.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
copying pymatgen\electronic_structure\__init__.py -> build\lib.win-amd64-3.7\pymatgen\electronic_structure
creating build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\compatibility.py -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\computed_entries.py -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\entry_tools.py -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\exp_entries.py -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\__init__.py -> build\lib.win-amd64-3.7\pymatgen\entries
creating build\lib.win-amd64-3.7\pymatgen\ext
copying pymatgen\ext\cod.py -> build\lib.win-amd64-3.7\pymatgen\ext
copying pymatgen\ext\crystalsai.py -> build\lib.win-amd64-3.7\pymatgen\ext
copying pymatgen\ext\jhu.py -> build\lib.win-amd64-3.7\pymatgen\ext
copying pymatgen\ext\matproj.py -> build\lib.win-amd64-3.7\pymatgen\ext
copying pymatgen\ext\__init__.py -> build\lib.win-amd64-3.7\pymatgen\ext
creating build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\adf.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\aiida.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\ase.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\atat.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\babel.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\cif.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\cssr.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\fiesta.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\gaussian.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\lmto.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\lobster.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\nwchem.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\phonopy.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\prismatic.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\pwscf.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\shengbte.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\xcrysden.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\xr.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\xyz.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\zeopp.py -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\io\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io
creating build\lib.win-amd64-3.7\pymatgen\optimization
copying pymatgen\optimization\linear_assignment_numpy.py -> build\lib.win-amd64-3.7\pymatgen\optimization
copying pymatgen\optimization\__init__.py -> build\lib.win-amd64-3.7\pymatgen\optimization
creating build\lib.win-amd64-3.7\pymatgen\phonon
copying pymatgen\phonon\bandstructure.py -> build\lib.win-amd64-3.7\pymatgen\phonon
copying pymatgen\phonon\dos.py -> build\lib.win-amd64-3.7\pymatgen\phonon
copying pymatgen\phonon\ir_spectra.py -> build\lib.win-amd64-3.7\pymatgen\phonon
copying pymatgen\phonon\plotter.py -> build\lib.win-amd64-3.7\pymatgen\phonon
copying pymatgen\phonon\__init__.py -> build\lib.win-amd64-3.7\pymatgen\phonon
creating build\lib.win-amd64-3.7\pymatgen\plugins
copying pymatgen\plugins\__init__.py -> build\lib.win-amd64-3.7\pymatgen\plugins
creating build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\analyzer.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\bandstructure.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\groups.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\kpath.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\maggroups.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\settings.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\site_symmetries.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\structure.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\__init__.py -> build\lib.win-amd64-3.7\pymatgen\symmetry
creating build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\advanced_transformations.py -> build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\defect_transformations.py -> build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\site_transformations.py -> build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\standard_transformations.py -> build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\transformation_abc.py -> build\lib.win-amd64-3.7\pymatgen\transformations
copying pymatgen\transformations\__init__.py -> build\lib.win-amd64-3.7\pymatgen\transformations
creating build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\convergence.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\coord.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\io_utils.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\num.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\plotting.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\provenance.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\sequence.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\serialization.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\string.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\testing.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\typing.py -> build\lib.win-amd64-3.7\pymatgen\util
copying pymatgen\util\__init__.py -> build\lib.win-amd64-3.7\pymatgen\util
creating build\lib.win-amd64-3.7\pymatgen\vis
copying pymatgen\vis\plotters.py -> build\lib.win-amd64-3.7\pymatgen\vis
copying pymatgen\vis\structure_chemview.py -> build\lib.win-amd64-3.7\pymatgen\vis
copying pymatgen\vis\structure_vtk.py -> build\lib.win-amd64-3.7\pymatgen\vis
copying pymatgen\vis\__init__.py -> build\lib.win-amd64-3.7\pymatgen\vis
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv
copying pymatgen\analysis\chemenv\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv
creating build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\core.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\corrections.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\defect_compatibility.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\dilute_solution_model.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\generators.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\thermodynamics.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
copying pymatgen\analysis\defects\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\defects
creating build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\core.py -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\neutron.py -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\tem.py -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\xrd.py -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
creating build\lib.win-amd64-3.7\pymatgen\analysis\elasticity
copying pymatgen\analysis\elasticity\elastic.py -> build\lib.win-amd64-3.7\pymatgen\analysis\elasticity
copying pymatgen\analysis\elasticity\strain.py -> build\lib.win-amd64-3.7\pymatgen\analysis\elasticity
copying pymatgen\analysis\elasticity\stress.py -> build\lib.win-amd64-3.7\pymatgen\analysis\elasticity
copying pymatgen\analysis\elasticity\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\elasticity
creating build\lib.win-amd64-3.7\pymatgen\analysis\ferroelectricity
copying pymatgen\analysis\ferroelectricity\polarization.py -> build\lib.win-amd64-3.7\pymatgen\analysis\ferroelectricity
copying pymatgen\analysis\ferroelectricity\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\ferroelectricity
creating build\lib.win-amd64-3.7\pymatgen\analysis\gb
copying pymatgen\analysis\gb\grain.py -> build\lib.win-amd64-3.7\pymatgen\analysis\gb
copying pymatgen\analysis\gb\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\gb
creating build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
copying pymatgen\analysis\magnetism\analyzer.py -> build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
copying pymatgen\analysis\magnetism\heisenberg.py -> build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
copying pymatgen\analysis\magnetism\jahnteller.py -> build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
copying pymatgen\analysis\magnetism\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
creating build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\structure_prediction\dopant_predictor.py -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\structure_prediction\substitution_probability.py -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\structure_prediction\substitutor.py -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\structure_prediction\volume_predictor.py -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\structure_prediction\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
creating build\lib.win-amd64-3.7\pymatgen\analysis\xas
copying pymatgen\analysis\xas\spectrum.py -> build\lib.win-amd64-3.7\pymatgen\analysis\xas
copying pymatgen\analysis\xas\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\xas
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
copying pymatgen\analysis\chemenv\connectivity\connected_components.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
copying pymatgen\analysis\chemenv\connectivity\connectivity_finder.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
copying pymatgen\analysis\chemenv\connectivity\environment_nodes.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
copying pymatgen\analysis\chemenv\connectivity\structure_connectivity.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
copying pymatgen\analysis\chemenv\connectivity\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\connectivity
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\chemenv_strategies.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometry_finder.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\structure_environments.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\voronoi.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
copying pymatgen\analysis\chemenv\coordination_environments\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\chemenv_config.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\chemenv_errors.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\coordination_geometry_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\defs_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\func_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\graph_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\math_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\scripts_utils.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
copying pymatgen\analysis\chemenv\utils\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\utils
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
creating build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\strategy_files
copying pymatgen\analysis\chemenv\coordination_environments\strategy_files\__init__.py -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\strategy_files
creating build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\analyzer.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\battery_abc.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\conversion_battery.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\insertion_battery.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\plotter.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
copying pymatgen\apps\battery\__init__.py -> build\lib.win-amd64-3.7\pymatgen\apps\battery
creating build\lib.win-amd64-3.7\pymatgen\apps\borg
copying pymatgen\apps\borg\hive.py -> build\lib.win-amd64-3.7\pymatgen\apps\borg
copying pymatgen\apps\borg\queen.py -> build\lib.win-amd64-3.7\pymatgen\apps\borg
copying pymatgen\apps\borg\__init__.py -> build\lib.win-amd64-3.7\pymatgen\apps\borg
creating build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\abiinspect.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\abiobjects.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\abitimer.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\helpers.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\netcdf.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\pseudos.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\variable.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
copying pymatgen\io\abinit\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\abinit
creating build\lib.win-amd64-3.7\pymatgen\io\exciting
copying pymatgen\io\exciting\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\exciting
copying pymatgen\io\exciting\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\exciting
creating build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\outputs.py -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\sets.py -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\feff
creating build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\lammps\data.py -> build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\lammps\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\lammps\outputs.py -> build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\lammps\utils.py -> build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\lammps\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\lammps
creating build\lib.win-amd64-3.7\pymatgen\io\qchem
copying pymatgen\io\qchem\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\qchem
copying pymatgen\io\qchem\outputs.py -> build\lib.win-amd64-3.7\pymatgen\io\qchem
copying pymatgen\io\qchem\sets.py -> build\lib.win-amd64-3.7\pymatgen\io\qchem
copying pymatgen\io\qchem\utils.py -> build\lib.win-amd64-3.7\pymatgen\io\qchem
copying pymatgen\io\qchem\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\qchem
creating build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\help.py -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\inputs.py -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\outputs.py -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\sets.py -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\__init__.py -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\analysis\bonds_jmol_ob.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\bvparam_1991.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\cn_opt_params.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\icsd_bv.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\op_params.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\vesta_cutoffs.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\aflow_prototypes.json -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\ionic_radii.json -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\costdb_elements.csv -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\analysis\hhi_data.csv -> build\lib.win-amd64-3.7\pymatgen\analysis
copying pymatgen\command_line\OxideTersoffPotentials -> build\lib.win-amd64-3.7\pymatgen\command_line
copying pymatgen\core\bond_lengths.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\func_groups.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\libxc_docs.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\periodic_table.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\quad_data.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\reconstructions_archive.json -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\core\py.typed -> build\lib.win-amd64-3.7\pymatgen\core
copying pymatgen\entries\MITCompatibility.yaml -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\entries\MPCompatibility.yaml -> build\lib.win-amd64-3.7\pymatgen\entries
copying pymatgen\io\BASIS_PBE_54.yaml -> build\lib.win-amd64-3.7\pymatgen\io
copying pymatgen\symmetry\symm_data.yaml -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\symm_ops.yaml -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\symm_data.json -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\symm_ops.json -> build\lib.win-amd64-3.7\pymatgen\symmetry
copying pymatgen\symmetry\symm_data_magnetic.sqlite -> build\lib.win-amd64-3.7\pymatgen\symmetry
creating build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\BaNiO3.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\CsCl.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Graphite.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\He_BCC.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\K2O2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Li10GeP2S12.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Li2O.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Li2O2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Li3V2(PO4)3.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\LiFePO4.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\NaFePO4.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Pb2TiZrO6.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Si.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\SiO2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\Sn.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\SrTiO3.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\TiO2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\TlBiSe2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\util\structures\VO2.json -> build\lib.win-amd64-3.7\pymatgen\util\structures
copying pymatgen\vis\ElementColorSchemes.yaml -> build\lib.win-amd64-3.7\pymatgen\vis
copying pymatgen\analysis\diffraction\atomic_scattering_params.json -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\diffraction\neutron_scattering_length.json -> build\lib.win-amd64-3.7\pymatgen\analysis\diffraction
copying pymatgen\analysis\magnetism\default_magmoms.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis\magnetism
creating build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction\data
copying pymatgen\analysis\structure_prediction\data\lambda.json -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction\data
copying pymatgen\analysis\structure_prediction\data\pair_correlation.json -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction\data
copying pymatgen\analysis\structure_prediction\DLS_bond_params.yaml -> build\lib.win-amd64-3.7\pymatgen\analysis\structure_prediction
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\allcg.txt -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\A#2.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\AC#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\BO_1#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\BO_2#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\BO_3#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\BS_1#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\BS_2#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\C#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\C#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\CO#11.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\DD#20.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\DD#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\DDPN#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\DI#11.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\ET#7.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\FO#7.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\H#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\H#11.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\HA#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\HB#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\HD#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\HP#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\I#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\L#2.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\MI#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\O#6.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\O#6_explicit.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PA#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PB#7.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PBP#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PCPA#11.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PP#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PP#5.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\PP#6.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\S#1.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\S#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\S#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\S#4.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\S#5.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SA#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SBSA#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SBT#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SC#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SH#11.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SH#13.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SMA#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SS#4.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SS#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\ST#7.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\SY#4.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\T#4.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\T#5.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\T#6.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TBSA#10.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TBT#8.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TC#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TI#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TL#3.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TO_1#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TO_2#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TO_3#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TS#3.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TT#12.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TT_1#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TT_2#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TT_3#9.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files\TY#3.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\coordination_geometries_files
copying pymatgen\analysis\chemenv\coordination_environments\strategy_files\ImprovedConfidenceCutoffDefaultParameters.json -> build\lib.win-amd64-3.7\pymatgen\analysis\chemenv\coordination_environments\strategy_files
copying pymatgen\io\feff\MPELNESSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\MPEXAFSSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\MPEXELFSSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\feff
copying pymatgen\io\feff\MPXANESSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\feff
creating build\lib.win-amd64-3.7\pymatgen\io\lammps\templates
copying pymatgen\io\lammps\templates\md.txt -> build\lib.win-amd64-3.7\pymatgen\io\lammps\templates
copying pymatgen\io\lammps\CoeffsDataType.yaml -> build\lib.win-amd64-3.7\pymatgen\io\lammps
copying pymatgen\io\vasp\MITRelaxSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\MPHSERelaxSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\MPRelaxSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\MPSCANRelaxSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\MVLGWSet.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\MVLRelax52Set.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\VASPIncarBase.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\vdW_parameters.yaml -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\incar_parameters.json -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\vasp_potcar_file_hashes.json -> build\lib.win-amd64-3.7\pymatgen\io\vasp
copying pymatgen\io\vasp\vasp_potcar_pymatgen_hashes.json -> build\lib.win-amd64-3.7\pymatgen\io\vasp
running build_ext
building 'pymatgen.optimization.linear_assignment' extension
error: Microsoft Visual C++ 14.0 is required. Get it with "Build Tools for Visual Studio": https://visualstudio.microsoft.com/downloads/
----------------------------------------
ERROR: Failed building wheel for pymatgen
Running setup.py clean for pymatgen
Building wheel for spglib (setup.py) ... error
ERROR: Command errored out with exit status 1:
command: 'c:\program files\python37\python.exe' -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"'; __file__='"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' bdist_wheel -d 'C:\Users\Dexter\AppData\Local\Temp\pip-wheel-hpmr8d0m'
cwd: C:\Users\Dexter\AppData\Local\Temp\pip-install-q2sl7tjc\spglib\
Complete output (14 lines):
setuptools is used.
c:\program files\python37\lib\site-packages\setuptools\dist.py:454: UserWarning: Normalizing '1.14.1-0' to '1.14.1.post0'
warnings.warn(tmpl.format(**locals()))
running bdist_wheel
running build
running build_py
creating build
creating build\lib.win-amd64-3.7
creating build\lib.win-amd64-3.7\spglib
copying spglib\spglib.py -> build\lib.win-amd64-3.7\spglib
copying spglib\__init__.py -> build\lib.win-amd64-3.7\spglib
running build_ext
building 'spglib._spglib' extension
error: Microsoft Visual C++ 14.0 is required. Get it with "Build Tools for Visual Studio": https://visualstudio.microsoft.com/downloads/
----------------------------------------
ERROR: Failed building wheel for spglib
Running setup.py clean for spglib
Failed to build pymatgen spglib
Installing collected packages: spglib, decorator, networkx, six, python-dateutil, pyparsing, cycler, kiwisolver, matplotlib, palettable, mpmath, sympy, pytz, pandas, retrying, plotly, pymatgen
Running setup.py install for spglib ... error
ERROR: Command errored out with exit status 1:
command: 'c:\program files\python37\python.exe' -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"'; __file__='"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' install --record 'C:\Users\Dexter\AppData\Local\Temp\pip-record-uyj9jvv_\install-record.txt' --single-version-externally-managed --compile --install-headers 'c:\program files\python37\Include\spglib'
cwd: C:\Users\Dexter\AppData\Local\Temp\pip-install-q2sl7tjc\spglib\
Complete output (14 lines):
setuptools is used.
c:\program files\python37\lib\site-packages\setuptools\dist.py:454: UserWarning: Normalizing '1.14.1-0' to '1.14.1.post0'
warnings.warn(tmpl.format(**locals()))
running install
running build
running build_py
creating build
creating build\lib.win-amd64-3.7
creating build\lib.win-amd64-3.7\spglib
copying spglib\spglib.py -> build\lib.win-amd64-3.7\spglib
copying spglib\__init__.py -> build\lib.win-amd64-3.7\spglib
running build_ext
building 'spglib._spglib' extension
error: Microsoft Visual C++ 14.0 is required. Get it with "Build Tools for Visual Studio": https://visualstudio.microsoft.com/downloads/
----------------------------------------
ERROR: Command errored out with exit status 1: 'c:\program files\python37\python.exe' -u -c 'import sys, setuptools, tokenize; sys.argv[0] = '"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"'; __file__='"'"'C:\\Users\\Dexter\\AppData\\Local\\Temp\\pip-install-q2sl7tjc\\spglib\\setup.py'"'"';f=getattr(tokenize, '"'"'open'"'"', open)(__file__);code=f.read().replace('"'"'\r\n'"'"', '"'"'\n'"'"');f.close();exec(compile(code, __file__, '"'"'exec'"'"'))' install --record 'C:\Users\Dexter\AppData\Local\Temp\pip-record-uyj9jvv_\install-record.txt' --single-version-externally-managed --compile --install-headers 'c:\program files\python37\Include\spglib' Check the logs for full command output.
~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
Can anyone tell me how to solve this problem?
Thanks so much for your sincerely help!
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1824/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1824/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1825
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1825/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1825/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1825/events
|
https://github.com/materialsproject/pymatgen/issues/1825
| 598,489,554
|
MDU6SXNzdWU1OTg0ODk1NTQ=
| 1,825
|
VoronoiNN._extract_nn_info() ValueError when calculating max_weight
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Hi @CompRhys , `VoronoiNN` takes a `cutoff` parameter as an optimization, but it does need to be increased with some structures. Try `VoronoiNN(cutoff=15)` and you should find it works.\r\n\r\nI'm not ruling out an underlying improvement that could be made here, if you're inclined you could take a look at the behavior of `VoronoiNN().get_all_voronoi_polyhedra(crystal)[14]` for your example vs `VoronoiNN().get_voronoi_polyhedra(crystal, 14)`. The latter works, the former does not, suggesting that the former might be overly aggressive in its optimizations (I did not write this function, so this is just speculation from me). If you find a fix, please feel welcome to open a PR, thanks for reporting.",
"Okay great I will look at the code and see if I can resolve the issue. \r\n\r\nAs a stop gap would a simple PR with a descriptive error message suggesting that users increase the cutoff if they get this exact error at this point be likely to be merged?",
"Yes, PRs always welcome even for things like improved warnings or docstrings – though actually in this case the code need not fail at all. Where the max() statement is, if this is an empty list it could just report no neighbours for that site rather than raising an error, so that change could be made too.\n\nI haven’t made that change myself because my understanding of VoronoiNN (with tolerance 0) is that all sites should at least have some neighbors, which makes me suspect an underlying bug in the optimization when all sites are requested.",
"This is perhaps more complicated, in the minimal code example increasing the cutoff doesn't fix the issue. \r\n\r\nI will open a PR with the assertion to stop if it doesn't find neighbours but only to facilitate fixing the deeper issue",
"That's interesting, the increased tolerance seemed to solve the issue on my machine.",
"So I can call both `local_env.VoronoiNN().get_voronoi_polyhedra(crystal, 14)`\r\n`local_env.VoronoiNN().get_all_voronoi_polyhedra(crystal)[14]` without errors but I get the error if I try `StructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN())`",
"@mkhorton I am currently looking for more edge cases and it appears to have something to do with the fact that I am converting the structures to primitive structures first (using pymatgen to do this). All of the last three edge cases I have found show this behaviour (*-p-* are the primitive structures)\r\n\r\nMaybe the best thing to do in the meantime is just merge a more helpful error message while I continue to look for edge cases?\r\n\r\nexamples:\r\n[breaking-oqmd-62712.txt](https://github.com/materialsproject/pymatgen/files/4482021/breaking-oqmd-62712.txt)\r\n[breaking-oqmd-688003.txt](https://github.com/materialsproject/pymatgen/files/4482022/breaking-oqmd-688003.txt)\r\n[breaking-oqmd-1018337.txt](https://github.com/materialsproject/pymatgen/files/4482023/breaking-oqmd-1018337.txt)\r\n[breaking-p-oqmd-62712.txt](https://github.com/materialsproject/pymatgen/files/4482024/breaking-p-oqmd-62712.txt)\r\n[breaking-p-oqmd-688003.txt](https://github.com/materialsproject/pymatgen/files/4482025/breaking-p-oqmd-688003.txt)\r\n[breaking-p-oqmd-1018337.txt](https://github.com/materialsproject/pymatgen/files/4482026/breaking-p-oqmd-1018337.txt)\r\n",
"I think I am having a similar problem - I had older code using VoronoiNN.get_nn_info that I tried to use today, and it is giving an illegal instruction error. Minimal code is attached.\r\n\r\nDo you know what I can try to do to fix this?\r\n\r\n```\r\nfrom pymatgen.ext.matproj import MPRester\r\n\r\nfrom pymatgen.core.structure import Structure, Composition\r\nfrom pymatgen.analysis.local_env import VoronoiNN\r\n\r\na = MPRester(\"ZbTstiE2olAnyU64\")\r\n\r\nentry = a.get_entry_by_material_id('mp-104',inc_structure='final')\r\n\r\ns = entry.structure\r\n\r\nvcf = VoronoiNN()\r\n\r\nvs = vcf.get_nn_info(s, 0)\r\n```"
] | 2020-04-12T13:51:47
| 2023-08-13T16:33:41
|
2023-08-13T16:33:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Trying to generate a Structure Graph from the Voronoi Neighbours gives `ValueError: max() arg is an empty sequence` for some structures when trying to calculate max_weight in `VoronoiNN._extract_nn_info()` .
**To Reproduce**
Minimal Code:
```
from pymatgen.core.structure import Structure
from pymatgen.analysis import local_env
from pymatgen.analysis.graphs import StructureGraph
with open("breaking-oqmd-4866.txt", 'r') as file:
str_ = file.read()
crystal = Structure.from_str(str_, fmt="json")
print(crystal)
graph = StructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN())
```
breaking example: attatched
[breaking-oqmd-4866.txt](https://github.com/materialsproject/pymatgen/files/4466354/breaking-oqmd-4866.txt)
**Expected behavior**
The code should return a structure graph where each nodes neighbours are its Voronoi neighbours
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1825/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1825/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1826
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1826/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1826/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1826/events
|
https://github.com/materialsproject/pymatgen/pull/1826
| 599,245,055
|
MDExOlB1bGxSZXF1ZXN0NDAyOTI5NDE3
| 1,826
|
Abstract Compatibility interface + self-contained Aqueous Compatibility class
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @rkingsbury, can you talk a little bit about the logic here?\r\n\r\nThe current Compatability (now named MITMPCompatibility in this PR) class is almost an abstract base class itself, it just takes a list of `corrections` to apply, e.g. its `process_entry`, `get_explanation_dict` etc. methods are pretty general.\r\n\r\nExplicitly making this an abstract base class seems fine, but default implementations for these methods seem sensible, and including e.g. default implementations for `process_entry` in the abstract class also seem sensible.",
"@mkhorton the idea here is to define the high-level interface for all `Compatibility` classes. The `MITMPCompatibility` works well as a base class for our existing approach to corrections, which assumes that we want to apply corrections all at once and that they can be computed independently. In that case, it makes sense to have default functionality like a list of corrections and a `get_corrections_dict` method in the base class. \r\n\r\nBut the existing approach is cumbersome and error-prone when we want to apply energy adjustments sequentially (e.g., take corrected solid DFT energies and convert them to Gibbs free energies for Pourbaix diagrams). What I'm trying to do with this PR is facilitate a way to write Compatibility classes that don't have an underlying list of `Correction` classes. They will still have `process_entries`, `explain`, and `process_entry`, but the logic within the class is more flexible - it does not have to depend on a list of `Correction` classes. I should mention that @shyamd was consulted on the initial implementation as well. \r\n\r\nAs I went about implementing one such `Compatibility` class to fix #1823 , I added some extra structure to this abstract implementation (see latest commits). The main change is to populate `Entry.data[\"Energy Adjustments\"]` with the contents of the `corrections_dict` (or equivalent). This allows us to tell by inspection whether \r\n\r\n1. an Entry has already been corrected\r\n2. what `Compatibility` classes have processed it, and \r\n3. whether the documentation in `Entry.data` accounts for all the corrections in `Entry.correction`.\r\n\r\nKnowing these things is important for future energy referencing schemes in which we want to process `ComputedEntry` that already have energy corrections applied (as is the case in the AqueousCorrection / Pourbaix scheme).\r\n\r\nI also added a `validate_corrections` method to ensure that all corrections are documented in `Entry.data` (and raise a warning if this is not the case). Finally, I now provide a new default `explain` method that performs this validation and lists corrections in both eV and eV/atom units.\r\n\r\nI've included default implementations for `process_entries` and `process_entry` in `Compatibility`. However, the `process_entry` implementation is still marked as an abstractclass because it is intended to be overridden. The default code is just an example to illustrate that `.process_entry` needs to both validate the corrections before doing anything and also populate `Entry.data[\"Energy Adjustments\"] before exiting.\r\n\r\nThis all works without changing anything about how the existing `Compatibility` subclasses behave. The only difference is that `ComputedEntry` will have their `.data` attribute populated with a dict documenting all the corrections that are applied.\r\n\r\nBased on my experience the last few months, I believe these changes will lay a foundation for making our energy corrections code substantially more transparent and robust. It will also facilitate schemes like Pourbaix that require knowledge of previously-applied corrections\r\n\r\nLet me know what you think. If this seems like the right direction to go, I'll add some tests for the new `Compatibility` class.",
"Abstract methods shouldn't have an implementation. ",
"> Abstract methods shouldn't have an implementation.\r\n\r\n~Not true... Yes, abstract classes certainly can have default methods, it is only the methods specifically marked as abstract that must be overridden.~\r\n\r\nEdit: sorry @shyamd, I realize you're saying the same thing I said below 🤦🏻♂️",
"> I've included default implementations for process_entries and process_entry in Compatibility. However, the process_entry implementation is still marked as an abstractclass because it is intended to be overridden. [...] The default code is just an example to illustrate that .process_entry needs to both validate the corrections before doing anything and also populate `Entry.data[\"Energy Adjustments\"]` before exiting.\r\n\r\nIf it's marked as abstract it's required to be overridden and should not have a default implementation.\r\n\r\nIf, however, you want to require validation in `process_entry`, that has to be enforced by the base class itself somehow.\r\n\r\nAlso, regarding naming, from your suggested changes it seems like the existing Compatability class is more a CompatabilityList class or similar, there seems to be nothing MP/MIT-specific in it that suggests it should be named `MITMPCompatibility`.\r\n\r\nIn general, I think improving the compatibility provenance is essential, and structural improvements are welcome, I just want to make sure this makes sense.",
"An example using a concrete implementation of `Compatibility` (not part of the PR):\r\n\r\nThe entry returned by `MPRester` has already been processed by `MaterialsProjectCompatibility`\r\n```\r\n>>> with MPRester() as m:\r\n my_entry = m.get_entry_by_material_id(\"mp-697111\", compatible_only=True)\r\n>>> my_entry.data[\"Energy Adjustments\"]\r\n{'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916}}\r\n```\r\n\r\nI now use a new class to apply Aqueous corrections on top of the existing ones:\r\n\r\n```\r\n>>> compat = MaterialsProjectAqueousCompatibility2020()\r\n>>> compat.process_entries([my_entry])\r\n[ComputedEntry mp-697111 - H8 O4\r\n Energy = -59.5408\r\n Correction = -6.3096\r\n Parameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['H', 'O'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE H', 'PBE O']\r\n oxide_type = hydroxide\r\n Data:\r\n oxide_type = hydroxide\r\n correction_uncertainty = nan\r\n Energy Adjustments = {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916}, 'MaterialsProjectAqueousCompatibility2020': {'Aqueous free energy correction': -3.500397110000003}}]\r\n```\r\n```\r\n>>> compat.explain(my_entry)\r\nThe uncorrected energy of H8 O4 is -59.541 eV (-4.962 eV/atom).\r\nThe following energy adjustments have been applied to this entry:\r\n\tMaterialsProjectCompatibility:\r\n\t\tMP Anion Correction: -2.809 eV (-0.234 eV/atom)\r\n\tMaterialsProjectAqueousCompatibility2020:\r\n\t\tAqueous free energy correction: -3.500 eV (-0.292 eV/atom)\r\nThe final energy after adjustments is -59.541 eV (-4.962 eV/atom).\r\n\r\n```\r\n\r\n\r\n\r\nThis new Aqueous class currently has a bug in which reprocessing the entries will keep applying the correction on top of the old ones. This is where the `validate_corrections` method is valuable. If I process `my_entry` again, we get a warning:\r\n\r\n```\r\n>>> compat.process_entries([my_entry])\r\n/Users/ryan/pymatgen/pymatgen/entries/compatibility.py:719: UserWarning: The provenance of the energy correction of -9.810 eV (-0.817 eV/atom) for entry mp-697111 is unknown! The contents of entry.data['Energy Adjustments'] contain total corrections of -6.310 eV (-0.526 eV/atom). Consider re-applying all energy corrections and proceed with caution.\r\n warnings.warn(\"The provenance of the energy correction of {:.3f} \"\r\n\r\n[ComputedEntry mp-697111 - H8 O4\r\n Energy = -59.5408\r\n Correction = -13.3104\r\n Parameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['H', 'O'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE H', 'PBE O']\r\n oxide_type = hydroxide\r\n Data:\r\n oxide_type = hydroxide\r\n correction_uncertainty = nan\r\n Energy Adjustments = {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916}, 'MaterialsProjectAqueousCompatibility2020': {'Aqueous free energy correction': -3.5003971099999998}}]\r\n```",
"For your new class, can you share the `.as_dict()` representation of the corrected ComputedEntry?",
"> For your new class, can you share the `.as_dict()` representation of the corrected ComputedEntry?\r\n\r\n```\r\n>>> my_entry.as_dict()\r\n{'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'ComputedEntry',\r\n 'energy': -59.54084689,\r\n 'composition': defaultdict(float, {'H': 8.0, 'O': 4.0}),\r\n 'parameters': {'run_type': 'GGA',\r\n 'is_hubbard': False,\r\n 'pseudo_potential': {'functional': 'PBE',\r\n 'labels': ['H', 'O'],\r\n 'pot_type': 'paw'},\r\n 'hubbards': {},\r\n 'potcar_symbols': ['PBE H', 'PBE O'],\r\n 'oxide_type': 'hydroxide'},\r\n 'data': {'oxide_type': 'hydroxide',\r\n 'correction_uncertainty': nan,\r\n 'Energy Adjustments': {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916},\r\n 'MaterialsProjectAqueousCompatibility2020': {'Aqueous free energy correction': -3.500397110000003}}},\r\n 'entry_id': 'mp-697111',\r\n 'correction': -6.309557110000003}\r\n```",
"I suggest rather that \"Energy Adjustments\" and \"correction_uncertainty\" become root level keys and that rather than putting the list of corrections within the Compatability scope, they become root level\r\n`'Energy Adjustments': {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916}`\r\nbecomes\r\n`'Energy Adjustments': {'MP Anion Correction': -2.80916}`",
"Why not dump the compatibility class as MSON? There's not enough information in the dict otherwise. Also `energy_adjustments` not `Energy Adjustments`. This is essentially constructing something similar to a transformation pipeline, so you want a list of the actual compatibility classes themselves (e.g. what if they were run with non-default args).",
"> Why not dump the compatibility class as MSON? There's not enough information in the dict otherwise. Also `energy_adjustments` not `Energy Adjustments`. This is essentially constructing something similar to a transformation pipeline, so you want a list of the actual compatibility classes themselves (e.g. what if they were run with non-default args).\r\n\r\nMy main concern here is for the corrections to be human-readable and transparent about what corrections have been applied, rather than to create a complete tranformation pipeline. Wouldn't dumping the whole class be kind of messy? I'm not opposed to adding it to a separate .data dict key though",
"Regarding the `abstractmethod`, the only way I can think of to enforce the `validate_correction` and populating the dict is to separate the actual correction into a separate method (e.g., a class-level `get_correction`. I can do that, but I was trying to avoid it for the sake of simplicity. One of the things that I find challenging about the existing implementation is tracing the logic (and bugs!) through `process_entries` -> `process_entry` -> `get_corrections_dict` -> `get_correction`. \r\n\r\nCan y'all think of any other ways to enforce the overall structure of `process_entry` without specifying the logic for calculating `correction`?",
"> I suggest rather that \"Energy Adjustments\" and \"correction_uncertainty\" become root level keys and that rather than putting the list of corrections within the Compatability scope, they become root level\r\n> `'Energy Adjustments': {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916}`\r\n> becomes\r\n> `'Energy Adjustments': {'MP Anion Correction': -2.80916}`\r\n\r\n@shyamd can I ask why?",
"@mkhorton since this is growing in scope, do you want to see a concrete implementation (e.g, the new AqueousCompatibility in my example) as part of this PR, or in a separate one?",
"> My main concern here is for the corrections to be human-readable and transparent about what corrections have been applied, rather than to create a complete tranformation pipeline\r\n\r\nYou can choose how this is printed using `__str__`, the dict format is not intended to be human readable. If you want to improve provenance it needs to contain necessary information to re-run the compatibility logic.\r\n\r\n> since this is growing in scope, do you want to see a concrete implementation (e.g, the new AqueousCompatibility in my example) as part of this PR, or in a separate one?\r\n\r\nLet's keep it in this PR since they affect each other and it will be easier to prototype.",
"@rkingsbury we're no longer treating them as arbitrary data that comes along with the `Entry` but rather actual pieces of data that are critical to the definition of the `Entry`. The data dict is really a place for us to throw things that we want to bring along but nothing in Compatability / PhaseDiagrams directly interacts with it. ",
"I'm a bit opposed to dumping the Correction class into the Entry. For the transformation pipeline, the dump of the transformation serves two purposes: documentation of provenance and a means of reproducing that structure in a data minimal way. Our alternative would be to store each structure along the way with some minimal documentation provenance, which would quickly blow up in terms of data structure size.\r\n\r\nFor corrections, storing the name, maybe date, and the value is sufficient provenance. We can always take off the corrections and return back to the original energy. By breaking them out into which correction contributes to what we have reproducibility of each step. This doesn't guarantee reproducibility of the correction value, but neither does a dump of the Correction class. It might also make sense to make `correction` a dynamic property of `Entry` that relies on the `energy_adjustments` lists to make the dependence on the list of corrections a stronger. ",
"> I'm a bit opposed to dumping the Correction class into the Entry. For the transformation pipeline, the dump of the transformation serves two purposes: documentation of provenance and a means of reproducing that structure in a data minimal way. Our alternative would be to store each structure along the way with some minimal documentation provenance, which would quickly blow up in terms of data structure size.\r\n> \r\n> For corrections, storing the name, maybe date, and the value is sufficient provenance. We can always take off the corrections and return back to the original energy. By breaking them out into which correction contributes to what we have reproducibility of each step. This doesn't guarantee reproducibility of the correction value, but neither does a dump of the Correction class. It might also make sense to make `correction` a dynamic property of `Entry` that relies on the `energy_adjustments` lists to make the dependence on the list of corrections a stronger.\r\n\r\nI like the idea of dynamically calculating `correction` from the dict; that will simplify the code quite a bit and improve robustness.\r\n\r\nFor the corrections dict, @shyamd am I understanding that what you're proposing would be \r\n\r\n`Entry.energy_adjustments = {{\"MP Anion Correction: -4.2}, {\"MP Aqueous Corrections: -3}}`\r\n\r\netc.?",
"Yeah, that is definitely the right idea, although a set of Dicts isn't allowed. Rather just have it as Dict[str,float]:\r\n```\r\nEntry.energy_adjustments = {\"MP Anion Correction: -4.2, \"MP Aqueous Corrections: -3}\r\n```\r\nI don't see a reason why a correction should ever be applied twice, which the Dict will enforce with only one entry per key. \r\n\r\nWe could add the PMG version but that maybe overkill. It's up to each correction class as to how it stores the individual components. The `CorrectionList` could just store one for each correction it has, while the Pourbaix correction might lump it all into one. Either way the point is to be able to separate out the components of the correction value. \r\n\r\nAs for what is shown in `__repr__`, that is up to you. There is nothing wrong with being detailed and having both the corrected and uncorrected values. There is a diff between `__str__` and `__repr__`. I just forget what.",
"> ```\r\n> Entry.energy_adjustments = {\"MP Anion Correction: -4.2, \"MP Aqueous Corrections: -3}\r\n> ```\r\n\r\n👍 makes sense\r\n\r\n> We could add the PMG version but that maybe overkill. It's up to each correction class as to how it stores the individual components. The `CorrectionList` could just store one for each correction it has, while the Pourbaix correction might lump it all into one. Either way the point is to be able to separate out the components of the correction value.\r\n\r\nSo, if a set of dicts isn't allowed, how would a class store additional data in this structure? Wouldn't each class need its own dict if we wanted to store pmg version, etc. in addition to the correction itself? e.g.\r\n\r\n{'MP Anion Correction': {\"value\": -4.2, \"pmg_version\": 2020-xx-xx, ...}}\r\n\r\n",
"Separate question - for writing new tests for this class, should I:\r\n\r\n1. Try to directly test the abstract class using something like `mock` OR\r\n2. Test a subclass, e.g., `MaterialsProjectCompatibilty` OR\r\n2. write a dummy subclass in the test file ?\r\n\r\nand \r\n\r\n1. use `unittests' OR\r\n2. use 'pytest' ?\r\n\r\nI'm inclined toward No. 2 in both cases, but let me know what you would like to see.",
"> > ```\r\n> > Entry.energy_adjustments = {\"MP Anion Correction: -4.2, \"MP Aqueous Corrections: -3}\r\n> > ```\r\n> \r\n> 👍 makes sense\r\n> \r\n> > We could add the PMG version but that maybe overkill. It's up to each correction class as to how it stores the individual components. The `CorrectionList` could just store one for each correction it has, while the Pourbaix correction might lump it all into one. Either way the point is to be able to separate out the components of the correction value.\r\n> \r\n> So, if a set of dicts isn't allowed, how would a class store additional data in this structure? Wouldn't each class need its own dict if we wanted to store pmg version, etc. in addition to the correction itself? e.g.\r\n> \r\n> {'MP Anion Correction': {\"value\": -4.2, \"pmg_version\": 2020-xx-xx, ...}}\r\n\r\nIt's also occurred to me that I think it would be good to have a record of whether an Entry has been processed by a specific class, even if the resulting correction is zero. I guess we can accomplish that by populating the dict with zero values when appropriate?",
"You're asking if there should be unit tests or integrations tests.\r\nThere should be unit tests that ensure a dummy class with specific functionality creates certain responses for the built-in methods in the Abstract class.\r\nThere should be integration tests that ensure these methods also return the right answer when in actual use in the MPCompatability class.\r\n\r\nI will leave it up to @mkhorton as per pytest vs unittest. I find it easier to write pytest tests.\r\n\r\nIf the entry is processed by a class but it is not a valid correction for that entry, it should probably be set to None. If the correction is valid and its just 0, then it should get set to 0. We want to differentiate between an actual 0, vs an effective 0. ",
"Catching up on the above ...\r\n\r\n> For corrections, storing the name, maybe date, and the value is sufficient provenance. We can always take off the corrections and return back to the original energy. By breaking them out into which correction contributes to what we have reproducibility of each step. This doesn't guarantee reproducibility of the correction value, but neither does a dump of the Correction class. \r\n\r\nI agree there's definitely an argument to be made that the numbers alone are sufficient. The MSON dump does include `@version` though, so it makes it very easy to find out exactly what bit of code generated that correction number, and that's what I'm seeing as most valuable by dumping the class (the `@version` at the root level is presumably sufficient for this).\r\n\r\nI see the argument that it makes the dict a bit heavy, I think an argument can be made either way.\r\n\r\n> I guess we can accomplish that by populating the dict with zero values when appropriate?\r\n\r\nThis would also be the benefit of explicitly storing the list of corrections as dicts, even if the value is zero.\r\n\r\n> I will leave it up to @mkhorton as per pytest vs unittest. I find it easier to write pytest tests.\r\n\r\nI prefer pytest, I stick to unittest in pymatgen for historical consistency, but perhaps it's time we switch. No strong opinions.",
"Yes, I'm going to call it -- let's see a prototype with `energy_adjustments` as a list of the compatibility classes themselves (so that, when serialized, it's going to store these as their dict representations).\r\n\r\n> 'correction_uncertainty': nan,\r\n\r\nSeparately, how is uncertainty going to be propagated when multiple corrections are applied? (and why `nan` not `None`?)",
"That breaks the definition of compatibility which is not defined on a per-entry basis. ",
"> That breaks the definition of compatibility which is not defined on a per-entry basis.\r\n\r\nTo be more specific, what I'm talking about here is e.g. the dict pasted above:\r\n\r\n```\r\n{'@module': 'pymatgen.entries.computed_entries',\r\n '@class': 'ComputedEntry',\r\n 'energy': -59.54084689,\r\n 'composition': defaultdict(float, {'H': 8.0, 'O': 4.0}),\r\n 'parameters': {'run_type': 'GGA',\r\n 'is_hubbard': False,\r\n 'pseudo_potential': {'functional': 'PBE',\r\n 'labels': ['H', 'O'],\r\n 'pot_type': 'paw'},\r\n 'hubbards': {},\r\n 'potcar_symbols': ['PBE H', 'PBE O'],\r\n 'oxide_type': 'hydroxide'},\r\n 'data': {'oxide_type': 'hydroxide',\r\n 'correction_uncertainty': nan,\r\n 'Energy Adjustments': {'MaterialsProjectCompatibility': {'MP Anion Correction': -2.80916},\r\n 'MaterialsProjectAqueousCompatibility2020': {'Aqueous free energy correction': -3.500397110000003}}},\r\n 'entry_id': 'mp-697111',\r\n 'correction': -6.309557110000003}\r\n```\r\n\r\nKeys like `MaterialsProjectAqueousCompatibility2020` are not sufficient to point to a class, dumping the `.as_dict()` gives us pointers (`@module`, `@class`, any other args/kwargs).\r\n\r\nThis is all I'm asking for -- this might not be sufficient to re-produce the number but it at least gives us a pointer to what was applied rather than just a bare string.",
"The other advantage of this is that in a construction of a `PhaseDiagram` it'll make it easier to check that a consistent set of corrections were applied to all `entries` used.",
"Honestly, the provenance of the corrections has never been issue. How complicated they are has been one, but never which is applied, since we can just reply what ever we want to enforce. I don't want to solve the provenance problem and make them even more complicated.\r\n\r\nI also think you're mixing `Compatability` with `Correction`. All of the parameters you want to track are in the `Compatability` classes which get set from code. These are predefined, so there is no need for Args or Kwargs. In the case of the Scan referencing scheme which will require the other entries to parametrized, we simply can't store all of this in one entry. So again, there is not much of a reason to store the `Compatability` class. \r\n\r\nI do agree we should store the corrections and there is a much better way of doing this. I'd rather see `Correction` be converted into something that is just to store that info and not as a means of computing it. For now, there would be two different kinds of correction classes: `CompositionCorrection` and `ConstantCorrection` that store the correction value or multiplier and a description. We can make the entry `correction_list` just those and when dumped via MSONable they serialize. This is far more descriptive and actually provide some traceability of the critical parameters. The `Compatability` classes will build `Correction` objects for each entry and store them. ",
"> I do agree we should store the corrections and there is a much better way of doing this. I'd rather see `Correction` be converted into something that is just to store that info and not as a means of computing it. For now, there would be two different kinds of correction classes: `CompositionCorrection` and `ConstantCorrection` that store the correction value or multiplier and a description. We can make the entry `correction_list` just those and when dumped via MSONable they serialize. This is far more descriptive and actually provide some traceability of the critical parameters. The `Compatability` classes will build `Correction` objects for each entry and store them.\r\n\r\nI like this idea. I didn't want to cause confusion by duplicating the name of the `Corrections` class, so I made a lightweight `EnergyAdjustment` class that stores a name, value, class name, and pymatgen version. So now the `repr` for an entry shows\r\n\r\n```\r\nComputedEntry mp-632291 - H2\r\nEnergy = -6.7814\r\nCorrection = -0.6171\r\nEnergy Adjustments:\r\n\tMP Gas Correction: 0.302 eV (0.151 eV/atom)\r\n\tFit H2 and H2O energy to experiment: -0.919 eV (-0.459 eV/atom)\r\n\tCompound entropy at room temperature: 0.000 eV (0.000 eV/atom)\r\n\tHydrate energy adjustment: 0.000 eV (0.000 eV/atom)\r\nParameters:\r\n\trun_type = GGA\r\n\tis_hubbard = False\r\n\tpseudo_potential = {'functional': 'PBE', 'labels': ['H'], 'pot_type': 'paw'}\r\n\thubbards = {}\r\n\tpotcar_symbols = ['PBE H']\r\n\toxide_type = None\r\nData:\r\n\toxide_type = None\r\n```\r\n"
] | 2020-04-14T02:08:50
| 2020-07-27T16:58:52
|
2020-05-05T22:45:44Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Abstract `Compatibility` interface
This pull request refactors `Compatibility` to define a common interface for pymatgen energy correction and referencing schemes. The goal of doing this is to enable more flexible correction schemes that do not necessarily require a list of `Correction` classes, and also to increase the robustness and transparency of the `Compatibility` classes by preventing double-counting and by documenting energy adjustments within the `ComputedEntry` instead of relying on `get_corrections_dict` or `explain`.
This functionality is important for future enhancements and bugfixes to the Pourbaix scheme (see #1823 ), and to ongoing work with other energy referencing classes.
### Major changes
* `Compatibility` is now an abstract base class, with required methods `process_entries` and `get_adjustments`.
* `process_entries` gains a `clean` kwarg that optionally removes all previously-applied corrections / adjustment before executing
* The old / existing Compatibility base class, which relies on an internal list of `Correction` classes, is renamed `CorrectionsList` and now inherits from the abstract class.
* `ComputedEntry` gains an attribute `energy_adjustments` that stores information about all energy adjustments / corrections that have been applied to the `Entry` by means of lightweight `EnegyAdjustment` objects.
* `ComputedEntry.correction` is now calculated dynamically from the contents of `energy_adjustments`, enforcing transparency regarding the provenance of energy adjustments.
* Manual setting of `ComputedEntry.correction` is still supported, and causes a `ManualEnergyAdjustment` object to replace the contents of `parameters.energy_adjustments`.
* `ComputedEntry` `repr` method has been updated to list uncorrected energy, total corrections, and final energy in both eV and eV/atom. Each `EnergyAdjustment` is also listed.
## Self-contained Aqueous Compatibility class
This PR also replaces `MaterialsProjectAqueousCompatibility` with a new implementation to serve as an example, concrete implementation of the new `Compatibility`. This new class fixes several bugs (see #1823 ) and allows the energy referencing scheme to be applied to any set of DFT entries, regardless of the underlying solid DFT energy corrections.
### Major changes / features
* `MaterialsProjectAqueousCompatibility` is now a standalone `Compatibility` whose sole purpose is to adjust DFT energies to free energies. It does NOT apply any of the solid energy corrections that are applied by `MaterialsProjectCompatibility`. Hence, it is meant to be used after the entries have been processed by another solid DFT energy correction scheme. Another `Compatibility` class can be passed as a kwarg to perform this pre-processing automatically.
* No longer uses `AqueousCorrection`. However, this is still used by `MITAqueousCompatibility`
* Energy adjustments are computed on the fly without reference to hard-coded values in `MPCompatibility.yaml`. To do this, the class requires three DFT energies (the ground-state energies of O2, H2O, and the total energy adjustments / corrections applied to H2O). These can be passed as kwargs to `__init__` or inferred from the entries passed to `process_entries`.
* As a result of recalculating / refitting the energy adjustments, the H2 free energy has changed from a previous value of -3.6019 eV/atom to -4.05 eV/atom
* Fitted DFT H2O free energy (not used by Pourbaix) changed from a previous value of -4.972 eV/atom to -5.26 eV/atom
* The energy correction applied to hydrate compounds is now calculated in a more accurate way. This may slightly change the free energies of some compounds containing OH.
* Added unittests for the complete `MaterialsProjectAqueousCompatibility` class. Previous tests only covered the underlying `AqueousCorrection` class and not the whole compatibility class.
* Fixes #1823 .
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1826/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1826/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1826",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1826",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1826.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1826.patch",
"merged_at": "2020-05-05T22:45:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1827
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1827/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1827/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1827/events
|
https://github.com/materialsproject/pymatgen/pull/1827
| 599,401,787
|
MDExOlB1bGxSZXF1ZXN0NDAzMDUyMDc4
| 1,827
|
[WIP] Edge Case of VoronoiNN() not finding neighbours when called by StructureGraph.with_local_env_strategy()
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"### Fails\r\n```\r\nStructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN(cutoff=8.0778))\r\nprint(\"baz - finds all polyhedra from StructureGraph\")\r\n```\r\n\r\n### \"Works\"\r\n```\r\nStructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN(cutoff=8.0777))\r\nprint(\"baz - finds all polyhedra from StructureGraph\")\r\n```\r\n\r\nwith the caveat that when I try and see whether it's returned the correct graph by running \r\n```\r\nfoo = StructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN(cutoff=8.0777))\r\nfoo.draw_graph_to_file(\"test.png\")\r\n```\r\nI get the following error:\r\n```\r\nTraceback (most recent call last):\r\n File \"/home/reag2/PhD/second-year/pymatgen/break-test.py\", line 19, in <module>\r\n foo.draw_graph_to_file(\"test.png\")\r\n File \"/home/reag2/PhD/second-year/pymatgen/pymatgen/analysis/graphs.py\", line 913, in draw_graph_to_file\r\n write_dot(g, basename + \".dot\")\r\n File \"/home/reag2/anaconda3/lib/python3.7/site-packages/networkx/drawing/nx_agraph.py\", line 194, in write_dot\r\n A.clear()\r\n File \"/home/reag2/anaconda3/lib/python3.7/site-packages/pygraphviz/agraph.py\", line 951, in clear\r\n self.remove_edges_from(self.edges())\r\n File \"/home/reag2/anaconda3/lib/python3.7/site-packages/pygraphviz/agraph.py\", line 573, in edges\r\n return list(self.edges_iter(nbunch=nbunch, keys=keys))\r\n File \"/home/reag2/anaconda3/lib/python3.7/site-packages/pygraphviz/agraph.py\", line 734, in edges_iter\r\n for e in self.out_edges_iter(keys=keys):\r\nRuntimeError: generator raised StopIteration\r\n```\r\n\r\nThe cut-off distance of 8.0777 doesn't appear to have any obvious significance for this structure - the closest reasonable conceptually relevant number is half the body diagonal of the crystal which is ~8.05. I haven't found an upper bound for the cut-off that works",
"The above can be traced to calls to \r\n\r\n```\r\nfoo = local_env.VoronoiNN(cutoff=8.0778).get_all_voronoi_polyhedra(crystal)\r\nbar = local_env.VoronoiNN(cutoff=8.0777).get_all_voronoi_polyhedra(crystal)\r\n```\r\n\r\nThe first returns empty neighbour dictionaries for the 14/15th entries whilst the second provides a full (not confirmed correct) output",
"The issue appears to stem from converting to primitive structures before calling `VoronoiNN()`. As such it is not apparent as of yet what exactly is going wrong. Despite not fixing the issue I suggest that adding this ValueError checking if no Voronoi neighbours are found for a given site is much more transparent as to what the issue is than `ValueError: max() arg is an empty sequence` which was the error previously raised.",
"Thanks @CompRhys for the tip about primitive cells, I'll leave the issue open. Next time we have someone using Voronoi extensively I'll ask them to help track down the underlying issue",
"I opened an issue on Scipy about implementing a periodic Voronoi so if that goes ahead it would be worth baring in mind for updating the `get_all_voronoi_polyhedra()` function",
"That'd be cool! I think they use Qhull under the hood, so it might depend on what Qhull itself supports. Perhaps they'd be willing to add a wrapper though."
] | 2020-04-14T08:47:48
| 2020-04-15T17:15:55
|
2020-04-15T16:35:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR is related to #1825 and aims to track down the source of the bug.
## Minimal Test for Edge Case
### Code
```
from pymatgen.core.structure import Structure
from pymatgen.analysis import local_env
from pymatgen.analysis.graphs import StructureGraph
with open("breaking-oqmd-4866.txt", 'r') as file:
str_ = file.read()
crystal = Structure.from_str(str_, fmt="json")
print(crystal)
StructureGraph.with_local_env_strategy(crystal, local_env.VoronoiNN())
```
### Input File
[breaking-oqmd-4866.txt](https://github.com/materialsproject/pymatgen/files/4474164/breaking-oqmd-4866.txt)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1827/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1827/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1827",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1827",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1827.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1827.patch",
"merged_at": "2020-04-15T16:35:36Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1828
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1828/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1828/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1828/events
|
https://github.com/materialsproject/pymatgen/pull/1828
| 600,535,545
|
MDExOlB1bGxSZXF1ZXN0NDAzOTQ2NTQ0
| 1,828
|
Units k cal
|
{
"login": "mamachra",
"id": 19979746,
"node_id": "MDQ6VXNlcjE5OTc5NzQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/19979746?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mamachra",
"html_url": "https://github.com/mamachra",
"followers_url": "https://api.github.com/users/mamachra/followers",
"following_url": "https://api.github.com/users/mamachra/following{/other_user}",
"gists_url": "https://api.github.com/users/mamachra/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mamachra/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mamachra/subscriptions",
"organizations_url": "https://api.github.com/users/mamachra/orgs",
"repos_url": "https://api.github.com/users/mamachra/repos",
"events_url": "https://api.github.com/users/mamachra/events{/privacy}",
"received_events_url": "https://api.github.com/users/mamachra/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @mamachra, could you add a test to https://github.com/materialsproject/pymatgen/blob/master/pymatgen/core/tests/test_units.py?",
"Thanks!"
] | 2020-04-15T19:28:11
| 2020-04-27T20:42:01
|
2020-04-21T21:09:13Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
kCal units added to units.py
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1828/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1828/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1828",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1828",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1828.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1828.patch",
"merged_at": "2020-04-21T21:09:13Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1829
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1829/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1829/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1829/events
|
https://github.com/materialsproject/pymatgen/issues/1829
| 601,164,797
|
MDU6SXNzdWU2MDExNjQ3OTc=
| 1,829
|
ComputedEntry repr / as_dict display uncorrected energy
|
{
"login": "rkingsbury",
"id": 1908695,
"node_id": "MDQ6VXNlcjE5MDg2OTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/1908695?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rkingsbury",
"html_url": "https://github.com/rkingsbury",
"followers_url": "https://api.github.com/users/rkingsbury/followers",
"following_url": "https://api.github.com/users/rkingsbury/following{/other_user}",
"gists_url": "https://api.github.com/users/rkingsbury/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rkingsbury/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rkingsbury/subscriptions",
"organizations_url": "https://api.github.com/users/rkingsbury/orgs",
"repos_url": "https://api.github.com/users/rkingsbury/repos",
"events_url": "https://api.github.com/users/rkingsbury/events{/privacy}",
"received_events_url": "https://api.github.com/users/rkingsbury/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"@ayushsgupta can you investigate?",
"@mkhorton @shyamd intended behavior would be to show the corrected energy under the 'Energy' key at least in the `repr`, right?",
"I suppose it makes sense to have uncorrected energy in `as_dict`, because that's what one would use to initialize a new entry with `from_dict`, but in`ComputedEntry.as_dict()` we should probably explicitly call it `uncorrected_energy` for clarity. In the `repr`, I think it would be nice to list both `energy` and `uncorrected_energy` for clarity. Thoughts?",
"I agree that it's not obvious if it's corrected or uncorrected energy currently, `uncorrected_energy` is better, but would break backwards compatibility. Adding a `corrected_energy` key would be nice but would be including duplicate information which is not ideal.\r\n\r\nSo, verdict: keep `as_dict()` as it is, but improve `__repr__()` to make it less ambiguous (the only job of `__repr__` is to show something useful about the underlying data, it's really `__str__()` we should be concerned about, but nevertheless people do print the object in Jupyter like in your screenshot so having an unambiguous `__repr__` would be a good thing imo).",
"Here is what I am thinking for the `repr`. This will be part of #1826 .\r\n\r\n```\r\nComputedEntry mp-13 - Fe1\r\nUncorrected Energy = -8.4598\r\nCorrection = 0.0000\r\nFinal Energy = -8.4598\r\nEnergy Adjustments:\r\n None\r\nParameters:\r\n run_type = GGA\r\n is_hubbard = False\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Fe_pv'], 'pot_type': 'paw'}\r\n hubbards = {}\r\n potcar_symbols = ['PBE Fe_pv']\r\n oxide_type = None\r\nData:\r\n oxide_type = None\r\n```",
"and an example with corrections:\r\n\r\n```\r\nComputedEntry mp-626570 - Fe16 H16 O32\r\nUncorrected Energy = -375.8532\r\nCorrection = -66.2013\r\nFinal Energy = -442.0545\r\nEnergy Adjustments:\r\n MP Anion Correction: -22.473 eV (-0.351 eV/atom)\r\n MP Advanced Correction: -43.728 eV (-0.683 eV/atom)\r\nParameters:\r\n run_type = GGA+U\r\n is_hubbard = True\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Fe_pv', 'H', 'O'], 'pot_type': 'paw'}\r\n hubbards = {'Fe': 5.3, 'H': 0.0, 'O': 0.0}\r\n potcar_symbols = ['PBE Fe_pv', 'PBE H', 'PBE O']\r\n oxide_type = hydroxide\r\nData:\r\n oxide_type = hydroxide\r\n```",
"This looks a lot more legible to me :-)",
"added a little extra alignment, reduced composition, and energies in both eV and eV/atom\r\n\r\n```\r\nComputedEntry mp-626570 - Fe16 H16 O32 (FeHO2)\r\nUncorrected Energy = -375.8532 eV (-5.8727 eV/atom)\r\nCorrection = -66.2013 eV (-1.0344 eV/atom)\r\nFinal Energy = -442.0545 eV (-6.9071 eV/atom)\r\nEnergy Adjustments:\r\n MP Anion Correction : -22.4733 eV (-0.3511 eV/atom)\r\n MP Advanced Correction : -43.7280 eV (-0.6833 eV/atom)\r\nParameters:\r\n run_type = GGA+U\r\n is_hubbard = True\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Fe_pv', 'H', 'O'], 'pot_type': 'paw'}\r\n hubbards = {'Fe': 5.3, 'H': 0.0, 'O': 0.0}\r\n potcar_symbols = ['PBE Fe_pv', 'PBE H', 'PBE O']\r\n oxide_type = hydroxide\r\nData:\r\n oxide_type = hydroxide\r\n```",
"Let me weigh in here. Having repr to return the uncorrected energy is fine. But not changing the dict representation. One has to remember that \"corrections\" or any concept therefore, is fundamentally an artificial construction of MP and its correction scheme. Hypothetically, one need not even store it as a correction and simply set it as PDEntry.energy = PDEntry.energy + correction with compatibility. The only reason why we store the correction within COmputedEntry is for reasons of debugging.\r\n\r\nSo the functional API of ComputedEntry is inherited from PDEntry - the most basic and fundamental concept of a Entry that does not require any correction whatsoever. The only reason why the *internal* representation uses `uncorrected_energy` is to enable the energy to be a property computed on the fly. That way, ComputedEntry and PDEntries can be mixed and transparently used. \r\n\r\nThe above repr looks ok to me. But instead of \"Uncorrected Energy' on the second line, I would suggest Energy (Uncorrected).",
"Thanks for the input, @shyuep . How about this?\r\n\r\n```\r\nComputedEntry mp-626570 - Fe16 H16 O32 (FeHO2)\r\nEnergy (Uncorrected) = -375.8532 eV (-5.8727 eV/atom)\r\nCorrection = -66.2013 eV (-1.0344 eV/atom)\r\nEnergy (Final) = -442.0545 eV (-6.9071 eV/atom)\r\nEnergy Adjustments:\r\n MP Anion Correction : -22.4733 eV (-0.3511 eV/atom)\r\n MP Advanced Correction : -43.7280 eV (-0.6833 eV/atom)\r\nParameters:\r\n run_type = GGA+U\r\n is_hubbard = True\r\n pseudo_potential = {'functional': 'PBE', 'labels': ['Fe_pv', 'H', 'O'], 'pot_type': 'paw'}\r\n hubbards = {'Fe': 5.3, 'H': 0.0, 'O': 0.0}\r\n potcar_symbols = ['PBE Fe_pv', 'PBE H', 'PBE O']\r\n oxide_type = hydroxide\r\nData:\r\n oxide_type = hydroxide\r\n```",
"This is fine with me. Though you can probably dump the Parameters and Data (which are both dicts) in YAML like form to make it slightly easier to read. But this is just me nit-picking."
] | 2020-04-16T15:27:24
| 2020-04-30T14:19:33
|
2020-04-30T14:19:33Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The `repr` and `as_dict` methods of `ComputedEntry` display the `uncorrected_energy` attribute under the `energy` key instead of the `energy` attribute (uncorrected energy plus corrections). Energies are still calculated correctly when reconstructing entries using `as_dict()` / `from_dict`, so this bug simply affects presentation.
**To Reproduce**
See steps in the screenshot below.
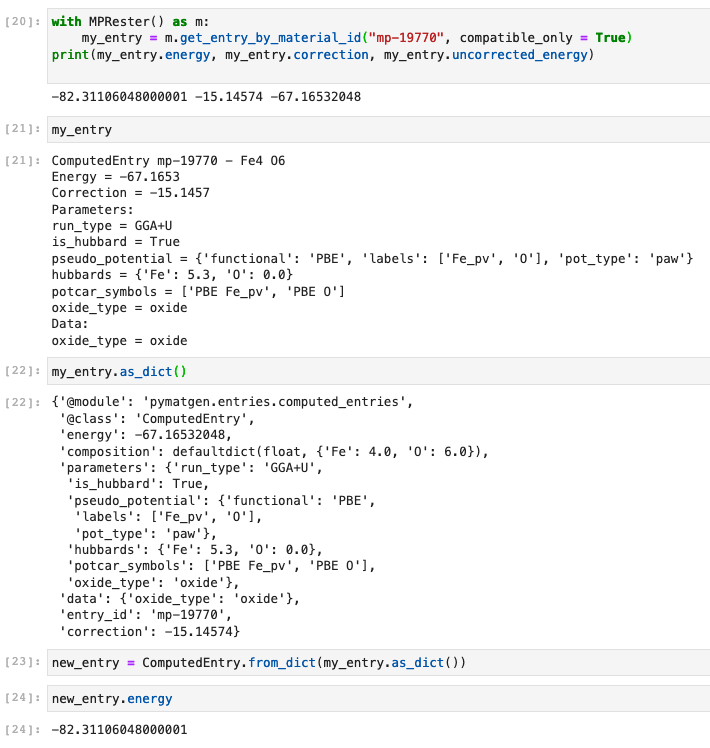
**Expected behavior**
Either display the `energy` attribute with the energy key or rename the key to `uncorrected_energy`.
**Additional context**
I have a suspicion that this bug was introduced when we refactored the `Entry` class (https://github.com/materialsproject/pymatgen/pull/1780), though I could be wrong and it may be intended behavior. If we do not already have one, I think we should add a test to enforce the intended behavior here.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1829/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1829/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1830
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1830/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1830/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1830/events
|
https://github.com/materialsproject/pymatgen/issues/1830
| 602,161,216
|
MDU6SXNzdWU2MDIxNjEyMTY=
| 1,830
|
Cif Parser improvement
|
{
"login": "ramfrey",
"id": 63871671,
"node_id": "MDQ6VXNlcjYzODcxNjcx",
"avatar_url": "https://avatars.githubusercontent.com/u/63871671?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ramfrey",
"html_url": "https://github.com/ramfrey",
"followers_url": "https://api.github.com/users/ramfrey/followers",
"following_url": "https://api.github.com/users/ramfrey/following{/other_user}",
"gists_url": "https://api.github.com/users/ramfrey/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ramfrey/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ramfrey/subscriptions",
"organizations_url": "https://api.github.com/users/ramfrey/orgs",
"repos_url": "https://api.github.com/users/ramfrey/repos",
"events_url": "https://api.github.com/users/ramfrey/events{/privacy}",
"received_events_url": "https://api.github.com/users/ramfrey/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I added an error message to detail unparseable structures. In this case, 18th structure in the file was problematic because there were essentially no atoms defined. You can check that for yourself by opening the CIF file and eyeballing it. \r\n\r\nPymatgen's CIF parser is reasonably robust against a lot of errors. But it cannot guess structures when no atoms are defined.",
"Thanks for the quick response. This already helps. A step further, it would be nice to have an option to read in from such a \"multiple\" cif file and directly resave the individual cif files (even the one with errors). ",
"To split a CIF, I don’t think you need pymatgen for that. You can easily use a Unix tool to do it or a simple few liner python script. "
] | 2020-04-17T18:37:01
| 2020-04-18T13:35:51
|
2020-04-17T22:47:17Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
I'm working with text files that contain multiple cif files (always 20) and would like to parse them with the Cif Parser, see example file. [1.txt](https://github.com/materialsproject/pymatgen/files/4494649/1.txt)
However, reading them into a dict or getting the structures yields different results:
```
for_struc=mg.io.cif.CifParser(r'1.cif',)
for_dict=mg.io.cif.CifParser(r'1.cif').as_dict()
struc=for_struc.get_structures(primitive=False)
print(len(struc))
print(len(list(for_dict.values())))
```
result:
19
20
also if I get the structures directly, I lose information e.g. the keys of the dict are the identifier numbers for each cif file, which i can't correlate directly to the structure anymore because the numbers are different.
**Describe the solution you'd like**
Either more precise error messages, saying which part of the (multiple)cif file was faulty, or a function that allows to create structures from "cif-dictionaries" as obtained like so: `for_dict=mg.io.cif.CifParser(r'1.cif').as_dict() `
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1830/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1830/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1831
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1831/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1831/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1831/events
|
https://github.com/materialsproject/pymatgen/pull/1831
| 602,250,546
|
MDExOlB1bGxSZXF1ZXN0NDA1MzU0NjQ1
| 1,831
|
SpacegroupOperations Description Fix
|
{
"login": "acrutt",
"id": 42151899,
"node_id": "MDQ6VXNlcjQyMTUxODk5",
"avatar_url": "https://avatars.githubusercontent.com/u/42151899?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/acrutt",
"html_url": "https://github.com/acrutt",
"followers_url": "https://api.github.com/users/acrutt/followers",
"following_url": "https://api.github.com/users/acrutt/following{/other_user}",
"gists_url": "https://api.github.com/users/acrutt/gists{/gist_id}",
"starred_url": "https://api.github.com/users/acrutt/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/acrutt/subscriptions",
"organizations_url": "https://api.github.com/users/acrutt/orgs",
"repos_url": "https://api.github.com/users/acrutt/repos",
"events_url": "https://api.github.com/users/acrutt/events{/privacy}",
"received_events_url": "https://api.github.com/users/acrutt/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks,"
] | 2020-04-17T21:45:00
| 2020-04-17T22:36:57
|
2020-04-17T22:36:56Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Fixed description for SpacegroupOperations.are_symmetrically_equivalent(). Input set of sites requires a list of PeriodicSite objects (not a Site objects).
## Additional dependencies introduced (if any)
* None
## TODO (if any)
* n/a
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1831/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1831/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1831",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1831",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1831.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1831.patch",
"merged_at": "2020-04-17T22:36:56Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1832
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1832/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1832/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1832/events
|
https://github.com/materialsproject/pymatgen/pull/1832
| 603,562,415
|
MDExOlB1bGxSZXF1ZXN0NDA2MzI0NDcy
| 1,832
|
Get neighbors save memory
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @chc273 !"
] | 2020-04-20T21:58:27
| 2020-04-20T22:50:28
|
2020-04-20T22:50:21Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This is a minor change to the previous `optimization.neighbors.find_points_in_sphere` cython method to save memory.
The critical step for finding neighbors is the generation of candidate atoms that are potentially within the cutoff radius of some atoms. This is achieved by repeating the unit cell in 3d and generating image cells.
In the previous approach, the memory of the maximum number of atoms `(n_atom_in_unit_cell * n_cell_images)` is allocated first and then those atoms that are beyond the cutoff are filtered out. This introduces a potentially large memory consumption in the beginning.
In the revised methods, I looped through the lattice offset vectors and gradually increase the number of candidate atoms and hence the memory of the calculation is smaller.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1832/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1832/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1832",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1832",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1832.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1832.patch",
"merged_at": "2020-04-20T22:50:21Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1833
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1833/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1833/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1833/events
|
https://github.com/materialsproject/pymatgen/issues/1833
| 603,813,833
|
MDU6SXNzdWU2MDM4MTM4MzM=
| 1,833
|
nullability-completeness related pymatgen build errors on Mac OS X
|
{
"login": "paulfons",
"id": 6129100,
"node_id": "MDQ6VXNlcjYxMjkxMDA=",
"avatar_url": "https://avatars.githubusercontent.com/u/6129100?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/paulfons",
"html_url": "https://github.com/paulfons",
"followers_url": "https://api.github.com/users/paulfons/followers",
"following_url": "https://api.github.com/users/paulfons/following{/other_user}",
"gists_url": "https://api.github.com/users/paulfons/gists{/gist_id}",
"starred_url": "https://api.github.com/users/paulfons/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/paulfons/subscriptions",
"organizations_url": "https://api.github.com/users/paulfons/orgs",
"repos_url": "https://api.github.com/users/paulfons/repos",
"events_url": "https://api.github.com/users/paulfons/events{/privacy}",
"received_events_url": "https://api.github.com/users/paulfons/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Please use conda to install pymatgen. ",
"Hi @paulfons, I can't reproduce -- I'm also on latest macOS (10.15.4) and Xcode (11.4), however I'll keep an eye on this.\r\n\r\n@shyuep is correct that the conda install is more reliable, since there are many more pre-built binary wheels available across platforms."
] | 2020-04-21T08:34:19
| 2020-04-21T16:56:06
|
2020-04-21T08:50:23Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
pymatgen update fails on OS X 10.15.4 and more importantly Xcode version 11.4.1 (11E503a)
**To Reproduce**
Steps to reproduce the behavior:
pip3 install --upgrade pymatgen
The full output is listed below:
[error.txt](https://github.com/materialsproject/pymatgen/files/4508677/error.txt)
The error seems to relate to the flag -Wnullability-completeness from what I can gather from google-based searches.
/Library/Developer/CommandLineTools/SDKs/MacOSX.sdk/usr/include/_stdio.h:93:16: warning: pointer is missing a nullability type specifier (_Nonnull, _Nullabl
e, or _Null_unspecified) [-Wnullability-completeness]
**Expected behavior**
An error free install update.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Desktop (please complete the following information):**
- OS: Mac (Catalina) 10.15.4 note Xcode version is 11.4.1 (11E503a)
- Version [e.g. 2019.9.16]
**Additional context**
Add any other context about the problem here.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1833/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1833/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1834
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1834/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1834/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1834/events
|
https://github.com/materialsproject/pymatgen/pull/1834
| 604,215,810
|
MDExOlB1bGxSZXF1ZXN0NDA2ODUxMjQ3
| 1,834
|
KPathLM labels bug fix
|
{
"login": "munrojm",
"id": 6877334,
"node_id": "MDQ6VXNlcjY4NzczMzQ=",
"avatar_url": "https://avatars.githubusercontent.com/u/6877334?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/munrojm",
"html_url": "https://github.com/munrojm",
"followers_url": "https://api.github.com/users/munrojm/followers",
"following_url": "https://api.github.com/users/munrojm/following{/other_user}",
"gists_url": "https://api.github.com/users/munrojm/gists{/gist_id}",
"starred_url": "https://api.github.com/users/munrojm/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/munrojm/subscriptions",
"organizations_url": "https://api.github.com/users/munrojm/orgs",
"repos_url": "https://api.github.com/users/munrojm/repos",
"events_url": "https://api.github.com/users/munrojm/events{/privacy}",
"received_events_url": "https://api.github.com/users/munrojm/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @munrojm. Is there an existing KPath labels test this case could be added to?",
"> Thanks @munrojm. Is there an existing KPath labels test this case could be added to?\r\n\r\nGood point! I will make one and update the existing test. ",
"Should be fixed now if you merge latest changes from master. Seems to be a _crash_ in pylint for something that seems correct to me -- will revisit in a future version of pylint assuming it's a bug, for now have excluded this specific version from our CI. Thanks for your patience with this.",
"> Should be fixed now if you merge latest changes from master. Seems to be a _crash_ in pylint for something that seems correct to me -- will revisit in a future version of pylint assuming it's a bug, for now have excluded this specific version from our CI. Thanks for your patience with this.\r\n\r\nNot a problem, that sounds good. Thanks @mkhorton!",
"Thanks @munrojm !"
] | 2020-04-21T19:06:48
| 2020-04-28T00:52:07
|
2020-04-28T00:52:07Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR fixes a small bug in the `KPathLM` class where the code would hang when assigning labels to each of the points in a k-path for some complex cases.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1834/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1834/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1834",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1834",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1834.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1834.patch",
"merged_at": "2020-04-28T00:52:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1835
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1835/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1835/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1835/events
|
https://github.com/materialsproject/pymatgen/pull/1835
| 605,205,656
|
MDExOlB1bGxSZXF1ZXN0NDA3NjUzNjY5
| 1,835
|
Sqs caller, io, and transformation changes
|
{
"login": "rwoodsrobinson",
"id": 7270689,
"node_id": "MDQ6VXNlcjcyNzA2ODk=",
"avatar_url": "https://avatars.githubusercontent.com/u/7270689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rwoodsrobinson",
"html_url": "https://github.com/rwoodsrobinson",
"followers_url": "https://api.github.com/users/rwoodsrobinson/followers",
"following_url": "https://api.github.com/users/rwoodsrobinson/following{/other_user}",
"gists_url": "https://api.github.com/users/rwoodsrobinson/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rwoodsrobinson/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rwoodsrobinson/subscriptions",
"organizations_url": "https://api.github.com/users/rwoodsrobinson/orgs",
"repos_url": "https://api.github.com/users/rwoodsrobinson/repos",
"events_url": "https://api.github.com/users/rwoodsrobinson/events{/privacy}",
"received_events_url": "https://api.github.com/users/rwoodsrobinson/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @rwoodsrobinson, this looks great. Will merge once tests pass.",
"Thanks!"
] | 2020-04-23T03:11:10
| 2020-04-24T03:25:57
|
2020-04-24T03:25:50Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Added parallel functionality and directory
* Added Monte Carlo kwargs
* Added new test files for mcsqs_caller, SQSTransformation and atat.io
* Added cluster generation based on structure
* Added file parsing to SQSTransformation
* Added sqs 'Perfect match' handling capabilities
* Updated docstrings
* Updated and improved SQSTransformation
* Fixed parallelization and runtime errors
* Fixed serious input bug on atat.io
* Fixed cif errors
## Checklist
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1835/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1835/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1835",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1835",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1835.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1835.patch",
"merged_at": "2020-04-24T03:25:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1836
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1836/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1836/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1836/events
|
https://github.com/materialsproject/pymatgen/pull/1836
| 606,809,801
|
MDExOlB1bGxSZXF1ZXN0NDA4OTM3MDU3
| 1,836
|
patch to allow scientific notation dielectric function, with possibil…
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can we have a unitest of this. No need for a full OUTCAR. Just testing the regex alone would be sufficient.",
"The regex is within the ```read_freq_dielectric()``` function in the OUTCAR class - which means I can't run the regex without first parsing an entire OUTCAR. As far as I can tell, my options are:\r\n\r\n1. Moving the regex pattern to somewhere universally accessible, e.g., out of read_freq_dielectric and at the top level of the outputs.py module, and then writing a test on that pattern (since now I can access the regex pattern without calling read_freq_dielectric(). This is fine but the code would look a little weirder\r\n\r\n2. Move the function to parse the scientific notation rows to its own static method in Outcar, and then test that static function. Again possible, but again the code would look weirder since now there is a whole other function to do essentially a one-line check.",
"1. is fine. But I would think splitting a sequence of scientific notation numbers is useful in more than one place. So 2 would be useful. I really don't think this is the only place with this particular problem.",
"If you make a regex that can handle both regular notation and scientific, we can just slowly patch that into the various functions. Note that python `float` interprets scientific notation just fine. ",
"Ok I separated the function out and added a unit test\r\n\r\nNote that I think it would be fairly straightforward to try to parse both types of numbers, assuming that if there is no exponent that there is at least a space between values (just change the exponent in the regex to be either the exponent or a set of whitespaces). But, I didn't attempt that here.",
"thanks."
] | 2020-04-25T16:48:22
| 2020-04-28T18:20:08
|
2020-04-28T18:20:05Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Some versions of VASP (notably, run with MedeA) have a different format to dielectric function tensor output in the OUTCAR file. Notably, the format:
- uses scientific notation
- may not have any spaces or delimiter character in between adjacent values on a line
For example, the OUTCAR may contain a section that looks like:
```
frequency dependent IMAGINARY DIELECTRIC FUNCTION (independent particle, no local field effects) density-density
E(ev) X Y Z XY YZ ZX
--------------------------------------------------------------------------------------------------------------
0.00000E+00 0.00000E+00 0.00000E+00 0.00000E+00 0.00000E+00 0.00000E+00 0.00000E+00
0.62963E+00 0.15467E+02 0.15467E+02 0.15467E+02-0.30654E-16-0.91612E-16 0.52388E-16
```
Note the use of scientific notation throughout, as well as the lack of delimiter between some of the values in the second line.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1836/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1836/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1836",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1836",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1836.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1836.patch",
"merged_at": "2020-04-28T18:20:05Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1837
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1837/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1837/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1837/events
|
https://github.com/materialsproject/pymatgen/pull/1837
| 606,900,960
|
MDExOlB1bGxSZXF1ZXN0NDA5MDAzNjUy
| 1,837
|
Add site_weighted_spectrum for XAS
|
{
"login": "yimingchen-eng",
"id": 27871258,
"node_id": "MDQ6VXNlcjI3ODcxMjU4",
"avatar_url": "https://avatars.githubusercontent.com/u/27871258?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yimingchen-eng",
"html_url": "https://github.com/yimingchen-eng",
"followers_url": "https://api.github.com/users/yimingchen-eng/followers",
"following_url": "https://api.github.com/users/yimingchen-eng/following{/other_user}",
"gists_url": "https://api.github.com/users/yimingchen-eng/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yimingchen-eng/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yimingchen-eng/subscriptions",
"organizations_url": "https://api.github.com/users/yimingchen-eng/orgs",
"repos_url": "https://api.github.com/users/yimingchen-eng/repos",
"events_url": "https://api.github.com/users/yimingchen-eng/events{/privacy}",
"received_events_url": "https://api.github.com/users/yimingchen-eng/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-04-26T02:32:54
| 2020-04-26T18:20:08
|
2020-04-26T18:20:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Enable generation of site-weighted XAS object based on site multiplicity for each absorbing index and its corresponding site-wise spectrum.
* Add `site_weighted_spectrum` for generation of site-weighted spectrum from site-wise spectra
* Add tests
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1837/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1837/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1837",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1837",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1837.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1837.patch",
"merged_at": "2020-04-26T18:20:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1838
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1838/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1838/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1838/events
|
https://github.com/materialsproject/pymatgen/pull/1838
| 608,310,054
|
MDExOlB1bGxSZXF1ZXN0NDEwMTExNTA4
| 1,838
|
Requirement typing_extensions
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Just upgrade to Py37 and it will be fine. ",
"Please have a look at the link as well (https://circleci.com/gh/hackingmaterials/atomate/2517?utm_campaign=vcs-integration-link&utm_medium=referral&utm_source=github-build-link): the failing atomate tests are in a python 3.7 environment and it the error is clearly coming from pymatgen. Unfortunately, upgrading does not help. ",
"I am also happy with any other solution that avoids the failing tests with python 3.7.",
"I do not think this is used in pymatgen at all. I do not see typing extensions anywhere in pymatgen master. Is atomate even running the latest pymatgen?\r\n\r\nIn any case, Py37 and above only pls. We will not backport to Py36.\r\n",
"Well, I do see it here:\r\nhttps://github.com/materialsproject/pymatgen/blob/master/pymatgen/command_line/mcsqs_caller.py",
"This was added by someone else recently. It isn't part of the released code I think. @mkhorton, we shouldn't allow people to add requirements unless explicitly approved by us? Literal is already part of typing. I don't see what the typing_extensions is supposed to do.",
"Hi all -- `Literal` is an official type hint as of Python 3.8, `typing-extensions` is the official Python-provided backport for Python <= 3.7.\r\n\r\nWe can add it to requirements.txt in the same way we add the dataclasses backport, so it will only be installed for older Python versions.\r\n\r\nI merged the above commit in. We have two options, either using:\r\n\r\n```\r\ntry:\r\n from typing import Literal\r\nexcept ImportError:\r\n from typing_extensions import Literal\r\n```\r\n\r\nOr just not use the `Literal` type at all, which is actually quite a useful hint. So I think we should do the former, though it's a little unfortunate.",
"that is, `typing-extensions==3.7.4.2; python_version < '3.8'`",
"I think we remove it for now. Or just downgrade Literal to str. It achieves most of the aims.",
"Your call -- I do like `Literal` as a really convenient way of not having to create an `Enum` while still getting most of the benefits for the type checker, it's very useful for settings strings (e.g. `LLL` or `niggli`, for a core example). `str` can still allow bad code to be written that isn't caught by the type checker.",
"Let's do str first. There are also costs to adding requirements. We have far too many as it is...",
"As a side note, I did a quick audit of pymatgen requirements recently:\r\n\r\n```\r\npymatgen==2020.4.2\r\n - matplotlib [required: >=1.5, installed: 3.1.0]\r\n - cycler [required: >=0.10, installed: 0.10.0]\r\n - six [required: Any, installed: 1.14.0]\r\n - kiwisolver [required: >=1.0.1, installed: 1.1.0]\r\n - setuptools [required: Any, installed: 41.0.1]\r\n - numpy [required: >=1.11, installed: 1.18.1]\r\n - pyparsing [required: >=2.0.1,!=2.1.6,!=2.1.2,!=2.0.4, installed: 2.4.6]\r\n - python-dateutil [required: >=2.1, installed: 2.8.1]\r\n - six [required: >=1.5, installed: 1.14.0]\r\n - monty [required: >=3.0.2, installed: 3.0.2]\r\n - networkx [required: >=2.2, installed: 2.3]\r\n - decorator [required: >=4.3.0, installed: 4.4.2]\r\n - numpy [required: >=1.14.3, installed: 1.18.1]\r\n - palettable [required: >=3.1.1, installed: 3.1.1]\r\n - pandas [required: Any, installed: 1.0.1]\r\n - numpy [required: >=1.13.3, installed: 1.18.1]\r\n - python-dateutil [required: >=2.6.1, installed: 2.8.1]\r\n - six [required: >=1.5, installed: 1.14.0]\r\n - pytz [required: >=2017.2, installed: 2019.3]\r\n - plotly [required: >=4.5.0, installed: 4.5.1]\r\n - retrying [required: >=1.3.3, installed: 1.3.3]\r\n - six [required: >=1.7.0, installed: 1.14.0]\r\n - six [required: Any, installed: 1.14.0]\r\n - pydispatcher [required: >=2.0.5, installed: 2.0.5]\r\n - requests [required: Any, installed: 2.22.0]\r\n - certifi [required: >=2017.4.17, installed: 2019.3.9]\r\n - chardet [required: >=3.0.2,<3.1.0, installed: 3.0.4]\r\n - idna [required: >=2.5,<2.9, installed: 2.8]\r\n - urllib3 [required: >=1.21.1,<1.26,!=1.25.1,!=1.25.0, installed: 1.25.3]\r\n - ruamel.yaml [required: >=0.15.6, installed: 0.16.10]\r\n - ruamel.yaml.clib [required: >=0.1.2, installed: 0.2.0]\r\n - scipy [required: >=1.0.1, installed: 1.3.0]\r\n - numpy [required: >=1.13.3, installed: 1.18.1]\r\n - spglib [required: >=1.9.9.44, installed: 1.12.2.post0]\r\n - numpy [required: Any, installed: 1.18.1]\r\n - sympy [required: Any, installed: 1.4]\r\n - mpmath [required: >=0.19, installed: 1.1.0]\r\n - tabulate [required: Any, installed: 0.8.3]\r\n```\r\n\r\nWe have standard science packages (scipy, numpy, pandas, networkx, sympy).\r\n\r\n`Matplotlib` is a very heavy dep (one of the reasons I'm favoring `plotly`). It was also a key stopping point in having pymatgen work with PyPy.\r\n\r\n`palettable` and `tabulate` are very lightweight utilities, pure Python.\r\n\r\n`requests`, `monty`, `ruamel.yaml`, `spglib` obviously essential\r\n\r\n`dataclasses`, similarly to `typing_extensions`, just a backport so we can use newer Python features without having to drop support for older Python\r\n\r\nThere's also `pydispatcher` and `beautifulsoup4` -- I'm not sure why either of these are in core requirements.",
"If plotly can do everything as per matplotlib, I am more than in favor of switching. My experience is that while plot.ly wins in terms of interactivity, the overall quality of achievable quality with minimal coding is still matplotlib. Further, many packages build on top of matplotlib, e.g., seaborn. If you do data science, seaborn is much easier to use.",
"As for pydispatcher, it is only used in the ABINIT package. beautifulsoup is a recent addition to support extraction of VASP help. I think this can be a general feature. E.g., a \"man page' of sorts for VASP and other codes.",
"> As for pydispatcher, it is only used in the ABINIT package.\r\n\r\nFeel free to remove pydispatcher from requirements.txt.\r\nThe package was used to implement the workflow machinery but now this part has been moved to AbiPy. "
] | 2020-04-28T13:16:31
| 2020-04-28T16:07:26
|
2020-04-28T14:41:19Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Hi,
we are currently working on including a workflow in atomate. Unfortunately, some of the tests fail with the current development version of pymatgen: https://circleci.com/gh/hackingmaterials/atomate/2517?utm_campaign=vcs-integration-link&utm_medium=referral&utm_source=github-build-link
It looks like the requirement "type-extensions" is missing.
@gpetretto did a short test. He gets the same error in "pymatgen/command_line/tests/test_mcsqs_caller.py" after installing pymatgen in a clean python3.6 environment. The error vanishes when he installs the optional dependencies.
Best,
JG
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1838/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1838/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1838",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1838",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1838.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1838.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1839
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1839/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1839/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1839/events
|
https://github.com/materialsproject/pymatgen/issues/1839
| 609,485,167
|
MDU6SXNzdWU2MDk0ODUxNjc=
| 1,839
|
CifParser distorts structure
|
{
"login": "peterschindler",
"id": 45672907,
"node_id": "MDQ6VXNlcjQ1NjcyOTA3",
"avatar_url": "https://avatars.githubusercontent.com/u/45672907?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/peterschindler",
"html_url": "https://github.com/peterschindler",
"followers_url": "https://api.github.com/users/peterschindler/followers",
"following_url": "https://api.github.com/users/peterschindler/following{/other_user}",
"gists_url": "https://api.github.com/users/peterschindler/gists{/gist_id}",
"starred_url": "https://api.github.com/users/peterschindler/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/peterschindler/subscriptions",
"organizations_url": "https://api.github.com/users/peterschindler/orgs",
"repos_url": "https://api.github.com/users/peterschindler/repos",
"events_url": "https://api.github.com/users/peterschindler/events{/privacy}",
"received_events_url": "https://api.github.com/users/peterschindler/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"In the call to get_structures, set primitive=False. When primitive is True (the default), the code performs a search for primitive lattice vectors and that can change the setting.",
"Oh, I wasn't aware of that tag. Thank you for the quick solution @shyuep !",
"Have verified this seems to be a bug in CifParser, though not in get_primitive ... potentially serious, am investigating.",
"Actually, I take this back, it seems fine, just unusual given the setting change. We _are_ trying to track down a potentially serious symmetrization/structure matching bug though, and thought this might have been a candidate, but evidently not."
] | 2020-04-30T01:17:52
| 2020-06-30T17:11:05
|
2020-06-30T17:11:04Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Recently came across a strange bug when parsing a primitive cell (cif downloaded from MP). Parsing the cif with CifParser and then immediately exporting it again as a new cif (without any other operations in between) completely changed/distorted the unit cell and atomic positions.
```Python
from pymatgen.io.cif import CifParser
file = "InI_mp-23202_primitive.cif"
structure = CifParser(file).get_structures()[0]
structure.to(filename="structure.cif")
```
See attached input and output cif files. Done with most recent Pymatgen version.
Not sure what could cause this, as the input cif looks normal to me and it's not a specifically weird structure. The lattice matrix of the imported structure seems already strange because the second-row vector `structure.lattice.matrix[1]` appears to be aligned with the z-direction (which normally should be the third-row vector).
Thank you for looking into it!
[cifs.zip](https://github.com/materialsproject/pymatgen/files/4555612/cifs.zip)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1839/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1839/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1840
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1840/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1840/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1840/events
|
https://github.com/materialsproject/pymatgen/issues/1840
| 609,667,107
|
MDU6SXNzdWU2MDk2NjcxMDc=
| 1,840
|
Getting all HKL with their intensities
|
{
"login": "bokertof",
"id": 43165697,
"node_id": "MDQ6VXNlcjQzMTY1Njk3",
"avatar_url": "https://avatars.githubusercontent.com/u/43165697?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bokertof",
"html_url": "https://github.com/bokertof",
"followers_url": "https://api.github.com/users/bokertof/followers",
"following_url": "https://api.github.com/users/bokertof/following{/other_user}",
"gists_url": "https://api.github.com/users/bokertof/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bokertof/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bokertof/subscriptions",
"organizations_url": "https://api.github.com/users/bokertof/orgs",
"repos_url": "https://api.github.com/users/bokertof/repos",
"events_url": "https://api.github.com/users/bokertof/events{/privacy}",
"received_events_url": "https://api.github.com/users/bokertof/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"There is an XRDCalculator in pymatgen.analysis.diffraction.xrd.\r\nIn future, please use https://discuss.matsci.org/ for queries on usage. Github Issues is meant for reporting bugs and requesting new features."
] | 2020-04-30T07:30:11
| 2020-04-30T14:19:16
|
2020-04-30T14:19:16Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Is there possibility to get theoretical HKL file from cif.file? I know that Platon software has such feature.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1840/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1840/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1841
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1841/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1841/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1841/events
|
https://github.com/materialsproject/pymatgen/issues/1841
| 611,257,238
|
MDU6SXNzdWU2MTEyNTcyMzg=
| 1,841
|
Comment in KPOINTS generator
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi Andrew; agreed, I think this could be better phrased. A pull request is welcome if you'd like to change it!\r\n\r\nAs a general comment, a lot of the default MP sets are kept as they are for historical, backwards compatibility reasons, including how we determine k-points. For our newer sets (e.g. SCAN) we're looking at KSPACING or one of the generalized schemes (e.g. the Mueller/JHU scheme) anyway. Regardless, the comments for the kppa scheme should be accurate and unambiguous, and I agree the current comment string is misleading.",
"Alright, I'll make a PR. I just wanted to make sure I wasn't misunderstanding it!\r\n\r\nThanks for the comment about the k-point generation. That's useful to know. I had been using `automatic_density()` for a while now, mainly drawing parallels with the original MP sets but have been meaning to switch over to the Mueller/JHU scheme myself. I just have to do some convergence checks before I feel comfortable porting my workflows over, which I keep putting off...",
"Yes, I've also been running convergence tests on the Mueller scheme, not 100% comfortable yet – I had some surprising results in the first batch, which makes me suspect an error on my part. There's a similar code from Gus Hart et al too https://github.com/msg-byu/kgridGen, but we don't have an interface to that. Opinions on these matters definitely welcome!"
] | 2020-05-02T19:24:07
| 2020-05-09T15:22:08
|
2020-05-09T15:22:08Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When using [`pymatgen.io.inputs.vasp.Kpoints.automatic_density()`](https://github.com/materialsproject/pymatgen/blob/v2020.4.29/pymatgen/io/vasp/inputs.py#L1237) (and likely related functions), the [comment line](https://github.com/materialsproject/pymatgen/blob/v2020.4.29/pymatgen/io/vasp/inputs.py#L1256) is
```python
comment = "pymatgen v%s with grid density = %.0f / atom" % (__version__, kppa)
```
I am having some self-doubt on whether this is _strictly speaking_ the best way to write this, specifically the "/ atom". For instance, if I set `kppa=1000`, it writes out "... grid density = 1000 / atom". Is that better described 1000/number of atoms, as written out in the [Materials Project documentation](https://materialsproject.org/docs/calculations)? Should it instead be written as 1000 per reciprocal atom, as in [the JHU module](https://pymatgen.org/pymatgen.ext.jhu.html)? 1000/per atom always confuses me, but I see that kind of phrasing in [the VASP documentation](https://cms.mpi.univie.ac.at/vasp/vasp/Number_k_points_method_smearing.html). Regardless of the best way to do this, my question really boils down to; "is 1000/atom" the ideal way to phrase this, and is it strictly accurate? A user may see that and think "so, if I increase the number of atoms, the number of k-points goes up," which of course is the opposite.
I know this is me just being pedantic and isn't really a bug, but I thought it'd be worth bringing up here.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1841/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1841/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1842
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1842/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1842/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1842/events
|
https://github.com/materialsproject/pymatgen/pull/1842
| 611,286,983
|
MDExOlB1bGxSZXF1ZXN0NDEyNTE2NTk1
| 1,842
|
Clearer description in generated KPOINTS comment line
|
{
"login": "Andrew-S-Rosen",
"id": 8674072,
"node_id": "MDQ6VXNlcjg2NzQwNzI=",
"avatar_url": "https://avatars.githubusercontent.com/u/8674072?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Andrew-S-Rosen",
"html_url": "https://github.com/Andrew-S-Rosen",
"followers_url": "https://api.github.com/users/Andrew-S-Rosen/followers",
"following_url": "https://api.github.com/users/Andrew-S-Rosen/following{/other_user}",
"gists_url": "https://api.github.com/users/Andrew-S-Rosen/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Andrew-S-Rosen/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Andrew-S-Rosen/subscriptions",
"organizations_url": "https://api.github.com/users/Andrew-S-Rosen/orgs",
"repos_url": "https://api.github.com/users/Andrew-S-Rosen/repos",
"events_url": "https://api.github.com/users/Andrew-S-Rosen/events{/privacy}",
"received_events_url": "https://api.github.com/users/Andrew-S-Rosen/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2020-05-02T22:25:41
| 2020-05-04T23:56:54
|
2020-05-04T23:56:54Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Change k-points / atom to k-points / number of atoms
## Summary
As mentioned in https://github.com/materialsproject/pymatgen/issues/1841, the comment line in the Pymatgen-generated VASP KPOINTS file is somewhat misleading. This PR changes it from being # kpoints/atom to # kpoints/number of atoms in the comment line, which is also consistent with what's on the [Materials Project documentation](https://materialsproject.org/docs/calculations). This should make it clearer that the number of k-points would be higher for smaller systems.
Perhaps worth noting: there is still the very minor ambiguity in that [pymatgen.ext.jhu](https://pymatgen.org/pymatgen.ext.jhu.html) takes `kppra` as an input argument, whereas [`pymatgen.io.vasp.inputs.Kpoints.automatic_density()`](https://github.com/materialsproject/pymatgen/blob/v2020.4.29/pymatgen/io/vasp/inputs.py#L1237) takes `kppa` as an input argument, but obviously the keywords should not be adjusted, and I think the documentation for the latter (which simply states `kppa` = grid density) is accurate without being confusing. Therefore, I did not make any changes to the documentation.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1842/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1842/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1842",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1842",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1842.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1842.patch",
"merged_at": "2020-05-04T23:56:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1843
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1843/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1843/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1843/events
|
https://github.com/materialsproject/pymatgen/issues/1843
| 611,391,656
|
MDU6SXNzdWU2MTEzOTE2NTY=
| 1,843
|
Reference doesn't exist
|
{
"login": "JabinHao",
"id": 47690236,
"node_id": "MDQ6VXNlcjQ3NjkwMjM2",
"avatar_url": "https://avatars.githubusercontent.com/u/47690236?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JabinHao",
"html_url": "https://github.com/JabinHao",
"followers_url": "https://api.github.com/users/JabinHao/followers",
"following_url": "https://api.github.com/users/JabinHao/following{/other_user}",
"gists_url": "https://api.github.com/users/JabinHao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JabinHao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JabinHao/subscriptions",
"organizations_url": "https://api.github.com/users/JabinHao/orgs",
"repos_url": "https://api.github.com/users/JabinHao/repos",
"events_url": "https://api.github.com/users/JabinHao/events{/privacy}",
"received_events_url": "https://api.github.com/users/JabinHao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @JabinHao, there are many textbook references available on Ewald summation. The specific one linked from our documentation was:\r\n\r\n*Ewald summation techniques in perspective: a survey* by Abdulnour Y. Toukmaji and John A. Board Jr., doi: 10.1016/0010-4655(96)00016-1\r\n\r\nA pdf seems to be available here: https://math.la.asu.edu/~jtaylor/teaching/Fall2010/APM530/papers/Toukmaji96.pdf\r\n\r\nI'll update the docstring to reference this paper specifically, thanks for letting us know the URL was broken!"
] | 2020-05-03T11:34:34
| 2020-05-03T19:24:40
|
2020-05-03T19:24:40Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I want to know more about the Ewald electrostatic energy, but when I try to open the reference in
```python
pymatgen.analysis.ewald module
class EwaldSummation(structure, real_space_cut=None, recip_space_cut=None, eta=None, acc_factor=12.0, w=0.7071067811865475, compute_forces=False)
Bases: object
Calculates the electrostatic energy of a periodic array of charges using the Ewald technique. Ref: http://www.ee.duke.edu/~ayt/ewaldpaper/ewaldpaper.html
This matrix can be used to do fast calculations of Ewald sums after species removal.
E = E_recip + E_real + E_point
Atomic units used in the code, then converted to eV.
```
the URL
>Ref: http://www.ee.duke.edu/~ayt/ewaldpaper/ewaldpaper.html
doesn't exist. I guess they have changed their site to other adress, Could you please provide the correct URL?
Thanks a lot.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1843/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1843/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1844
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1844/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1844/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1844/events
|
https://github.com/materialsproject/pymatgen/issues/1844
| 613,343,639
|
MDU6SXNzdWU2MTMzNDM2Mzk=
| 1,844
|
Suggest upgrading to spglib-1.15.1 for solving the pymatgen installation freezing problem.
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-05-06T14:00:33
| 2020-05-10T00:38:48
|
2020-05-09T14:31:16Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
I meet the following problem when installing the git master pymatgen: it freezes for ever during the installation progress. Then I use ` pip install -v ` to locate the problem and find that this is caused by spglib package. Simply to say, spglib needs numpy. If the numpy doesn't exist in the virtualenv where we try to install pymatgen, spglib's setup.py cannot successfully install the numpy and as a result, the installation freezes there for ever. And finally, the pymatgen's installation will also freeze there. Recently, the spglib's author has upgraded the spglib which including the fix for this problem and other bug fixes / enhancements. So I suggest upgrade the spglib's version in requirements.txt of pymatgen for solving this problem.
Regards,
HY
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1844/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1844/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1845
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1845/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1845/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1845/events
|
https://github.com/materialsproject/pymatgen/issues/1845
| 614,786,806
|
MDU6SXNzdWU2MTQ3ODY4MDY=
| 1,845
|
pymatgen.analysis.interface_reactions InterfacialReactivity issue
|
{
"login": "roterojama",
"id": 65028338,
"node_id": "MDQ6VXNlcjY1MDI4MzM4",
"avatar_url": "https://avatars.githubusercontent.com/u/65028338?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/roterojama",
"html_url": "https://github.com/roterojama",
"followers_url": "https://api.github.com/users/roterojama/followers",
"following_url": "https://api.github.com/users/roterojama/following{/other_user}",
"gists_url": "https://api.github.com/users/roterojama/gists{/gist_id}",
"starred_url": "https://api.github.com/users/roterojama/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/roterojama/subscriptions",
"organizations_url": "https://api.github.com/users/roterojama/orgs",
"repos_url": "https://api.github.com/users/roterojama/repos",
"events_url": "https://api.github.com/users/roterojama/events{/privacy}",
"received_events_url": "https://api.github.com/users/roterojama/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the report @roterojama. Can you paste in the two separate error messages you see?",
"@mkhorton I reinstalled the newest version of pymatgen to reproduce the error messages and just couldn't. In other words. Everything seems to be working fine. Im very sorry about this. I can't explain what changed. Thus I guess the issue is resolved. Good day to you.",
"No problem!"
] | 2020-05-08T14:43:35
| 2020-05-11T16:57:31
|
2020-05-11T16:57:31Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
When using the newest version of pymatgen (v2020.4.29) the InterfacialReactivity() function produces an error. Two seperate errors can be generated depeding on whether the argument "include_no_mixing_energy=" is set to True or False. However this was not the case before the newest version of pymatgen. I changed the package version to e.g. pymatgen=2019.2.28 and everything works fine.
**To Reproduce**
Steps to reproduce the behavior:
Just run the example code from the binder hub. Its the "2019-03-11-Interface Reactions.ipynb" notebook. Here you have the code as a txt file: [Example-file.txt](https://github.com/materialsproject/pymatgen/files/4599647/Example-file.txt) (I changed it from a .py file. I hope that didnt introduce any Errors).
**Expected behavior**
Can be seen when running the notebook on the binder hub (binder hub: http://matgenb.materialsvirtuallab.org/). There it works fine.
**Desktop (please complete the following information):**
- OS: Windows 10 Home, OS build: 18362.778
- Version 1903
- Im running the Spyder 4.1.2 IDE with the Anaconda Navigator for the environement.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1845/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1845/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1846
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1846/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1846/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1846/events
|
https://github.com/materialsproject/pymatgen/issues/1846
| 615,180,274
|
MDU6SXNzdWU2MTUxODAyNzQ=
| 1,846
|
Fail to run pymatgen package from within ipython.
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-05-09T13:48:17
| 2020-05-09T15:21:57
|
2020-05-09T15:21:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
Ubuntun 19.10 + git master version pymatgen and ipython. Fail to run some simple tests.
See [here](https://github.com/ipython/ipython/issues/12300) for more info.
Regards
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1846/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1846/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1847
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1847/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1847/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1847/events
|
https://github.com/materialsproject/pymatgen/pull/1847
| 616,474,192
|
MDExOlB1bGxSZXF1ZXN0NDE2NTc3ODI2
| 1,847
|
File update
|
{
"login": "rajeshsharma98",
"id": 36369426,
"node_id": "MDQ6VXNlcjM2MzY5NDI2",
"avatar_url": "https://avatars.githubusercontent.com/u/36369426?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/rajeshsharma98",
"html_url": "https://github.com/rajeshsharma98",
"followers_url": "https://api.github.com/users/rajeshsharma98/followers",
"following_url": "https://api.github.com/users/rajeshsharma98/following{/other_user}",
"gists_url": "https://api.github.com/users/rajeshsharma98/gists{/gist_id}",
"starred_url": "https://api.github.com/users/rajeshsharma98/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/rajeshsharma98/subscriptions",
"organizations_url": "https://api.github.com/users/rajeshsharma98/orgs",
"repos_url": "https://api.github.com/users/rajeshsharma98/repos",
"events_url": "https://api.github.com/users/rajeshsharma98/events{/privacy}",
"received_events_url": "https://api.github.com/users/rajeshsharma98/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I don't understand the difference between this and Element.Si.electronic_structure?",
"It looks like `Element.Si.electronic_structure` contains HTML tags, so is not as versatile?",
"Yes @mkhorton @shyuep, this was the only reason I created this function.\r\nAs for 'Fe'; _Element.Si.electronic_structure_ output contains HTML tags whereas my function(noblegas_electronic_structure) gives output: ['Ar', (3, 'd', 6), (4, 's', 2)].\r\n\r\n**Drawbacks** of using - _Element.Si.electronic_structure_\r\n1. its output is not easily understandable because of tags.\r\n2. its output is of string type that means it needs some time and few lines of code to fetch a particular part of the output.\r\n3. output format is not useful.\r\n\r\n**Improvements I made** with the development of noblegas_electronic_structure function:\r\n1. one can easily understand the output.\r\n2. output formatting of full_electronic_structure function and my function is the same.\r\n3. output is of list type, which makes its implementation easier.\r\n4. its(noblegas_electronic_structure) name also suggests that it gives a similar or comparable output as compared to _full_electronic_structure_ function.\r\n\r\n\r\n\r\n",
"We are not creating a function just to remove html tags. By all means remove the html tags and make the the output of electronic structure. ",
"Yes, I'd agree -- I think it'd be easier to address this just by changing the underlying data in the periodic_table.json file, e.g. see https://github.com/materialsproject/pymatgen/blob/4e138b9f0d1caf040aabc7799875809a4596a05a/pymatgen/core/periodic_table.json#L36",
"I thought creating a function would be an easy task compared to changing the values, as function is just manipulating the output of full electronic structure function. Means it’s implementation and use will be easy. \r\nThis was the reason I created the function , I am also working on a project that requires noble gas configuration, mainly the electrons that are not valanced.",
"Actually, modifying the file is not necessary at all. What you want can be achieved with a single line of code:\r\n```python\r\nre.sub(\"</*sup>\", \"\", Element.Si.electronic_structure)\r\n```\r\ngives\r\n```\r\n'[Ne].3s2.3p2'\r\n```"
] | 2020-05-12T08:43:37
| 2020-05-13T12:16:34
|
2020-05-13T12:14:49Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
### Summary
Depending on the element, the electronic configuration may be very long. Because of this, scientists have developed a shorthand notation that involves using a noble gas to represent electrons that are not valence electrons. This function does the same, it simplifies the electronic configuration for better understanding.
Function Name: nobelgas_electronic_structure()
Example- for 'Fe' this function will output: ['Ar', (3, 'd', 6), (4, 's', 2)]
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1847/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1847/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1847",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1847",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1847.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1847.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1848
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1848/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1848/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1848/events
|
https://github.com/materialsproject/pymatgen/issues/1848
| 620,503,154
|
MDU6SXNzdWU2MjA1MDMxNTQ=
| 1,848
|
Can't instantiate numpy array of Composition objects
|
{
"login": "KirbyBroderick",
"id": 57768514,
"node_id": "MDQ6VXNlcjU3NzY4NTE0",
"avatar_url": "https://avatars.githubusercontent.com/u/57768514?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/KirbyBroderick",
"html_url": "https://github.com/KirbyBroderick",
"followers_url": "https://api.github.com/users/KirbyBroderick/followers",
"following_url": "https://api.github.com/users/KirbyBroderick/following{/other_user}",
"gists_url": "https://api.github.com/users/KirbyBroderick/gists{/gist_id}",
"starred_url": "https://api.github.com/users/KirbyBroderick/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/KirbyBroderick/subscriptions",
"organizations_url": "https://api.github.com/users/KirbyBroderick/orgs",
"repos_url": "https://api.github.com/users/KirbyBroderick/repos",
"events_url": "https://api.github.com/users/KirbyBroderick/events{/privacy}",
"received_events_url": "https://api.github.com/users/KirbyBroderick/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I don't think a numpy array is a good container for a Composition object. The Composition object is in fact a dict-like object, which is iterable.\r\nYou can use lists if you wish, or pandas DataFrames, which are supposed to work with heterogenous objects works too. In general, numpy arrays are for numeric types."
] | 2020-05-18T21:00:35
| 2020-05-18T21:07:15
|
2020-05-18T21:07:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I'm on pymatgen version 2020.1.28 in JupyterLab, and I'm trying to instantiate a numpy array of Composition objects.
`import numpy as np
np.array([pymatgen.core.Composition('Ru')])` and several variants thereof result in an error
`TypeError: Invalid key 0, <class 'int'> for Composition
ValueError exception:
No element with this atomic number 0`
I can, however, get this array by going through a pandas DataFrame:
`pd.DataFrame([[pymatgen.core.Composition('Ru')]]).values[0]`
I was hoping the direct way would work.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1848/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1848/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1849
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1849/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1849/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1849/events
|
https://github.com/materialsproject/pymatgen/issues/1849
| 622,141,356
|
MDU6SXNzdWU2MjIxNDEzNTY=
| 1,849
|
Negative electrostatic energy for charged cells
|
{
"login": "lbluque",
"id": 9788715,
"node_id": "MDQ6VXNlcjk3ODg3MTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9788715?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lbluque",
"html_url": "https://github.com/lbluque",
"followers_url": "https://api.github.com/users/lbluque/followers",
"following_url": "https://api.github.com/users/lbluque/following{/other_user}",
"gists_url": "https://api.github.com/users/lbluque/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lbluque/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lbluque/subscriptions",
"organizations_url": "https://api.github.com/users/lbluque/orgs",
"repos_url": "https://api.github.com/users/lbluque/repos",
"events_url": "https://api.github.com/users/lbluque/events{/privacy}",
"received_events_url": "https://api.github.com/users/lbluque/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"I don't think that is correct. A charged cell has a background charge of the opposite charge. That results in a negative electrostatic energy.",
"Ah, I see now. It is the energy of the structure + the compensating background. Thanks for the clarification @shyuep. \r\n\r\nI think the docstrings could be made more explicit then. Stating that the result for charged systems. Additionally there is a URL in there that does not work anymore, and I believe the actual paper it refers does not treat charged systems at all. I would suggest this as an additional reference:\r\n\r\n- R. M. Martin, Ed., in Electronic Structure: Basic Theory and Practical Methods (Cambridge University Press, Cambridge, 2004; (https://www.cambridge.org/core/books/electronic-structure/coulomb-interactions-in-extended-systems/51B0E2F27A831B78BCEFA1444299E92B) pp. 499–511.\r\n",
"I updated the reference in the code to the original source for the old URL.",
"Yup that's the one I also assumed the old URL referred to."
] | 2020-05-20T23:15:10
| 2020-05-28T15:59:43
|
2020-05-28T15:59:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I'm getting nonphysical negative values for the electrostatic energy of charged cells using `EwaldSummation`.
Seems the `_charged_cell_energy` needs to be adjusted by a minus sign?
https://github.com/materialsproject/pymatgen/blob/72d336d2d17ac140058fdcb74eb3336125bb12b7/pymatgen/analysis/ewald.py#L114
|
{
"login": "lbluque",
"id": 9788715,
"node_id": "MDQ6VXNlcjk3ODg3MTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9788715?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lbluque",
"html_url": "https://github.com/lbluque",
"followers_url": "https://api.github.com/users/lbluque/followers",
"following_url": "https://api.github.com/users/lbluque/following{/other_user}",
"gists_url": "https://api.github.com/users/lbluque/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lbluque/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lbluque/subscriptions",
"organizations_url": "https://api.github.com/users/lbluque/orgs",
"repos_url": "https://api.github.com/users/lbluque/repos",
"events_url": "https://api.github.com/users/lbluque/events{/privacy}",
"received_events_url": "https://api.github.com/users/lbluque/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1849/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1849/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1850
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1850/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1850/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1850/events
|
https://github.com/materialsproject/pymatgen/pull/1850
| 623,705,113
|
MDExOlB1bGxSZXF1ZXN0NDIyMzAzMTg1
| 1,850
|
Minor fix to as_dict and from_dict in StructureMatcher
|
{
"login": "lbluque",
"id": 9788715,
"node_id": "MDQ6VXNlcjk3ODg3MTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9788715?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lbluque",
"html_url": "https://github.com/lbluque",
"followers_url": "https://api.github.com/users/lbluque/followers",
"following_url": "https://api.github.com/users/lbluque/following{/other_user}",
"gists_url": "https://api.github.com/users/lbluque/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lbluque/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lbluque/subscriptions",
"organizations_url": "https://api.github.com/users/lbluque/orgs",
"repos_url": "https://api.github.com/users/lbluque/repos",
"events_url": "https://api.github.com/users/lbluque/events{/privacy}",
"received_events_url": "https://api.github.com/users/lbluque/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks."
] | 2020-05-23T17:28:15
| 2020-05-25T15:07:43
|
2020-05-25T15:07:43Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Saves missing attributes in `StructureMatcher.as_dict` so that `StuctureMatcher.from_dict` can faithfully recreate the original object.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [X] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [X] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [X] Tests have been added for any new functionality or bug fixes.
- [X] All existing tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1850/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1850/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1850",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1850",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1850.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1850.patch",
"merged_at": "2020-05-25T15:07:43Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1851
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1851/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1851/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1851/events
|
https://github.com/materialsproject/pymatgen/issues/1851
| 624,439,154
|
MDU6SXNzdWU2MjQ0MzkxNTQ=
| 1,851
|
any function to get site-specific xas from MP?
|
{
"login": "davidcn21",
"id": 7539330,
"node_id": "MDQ6VXNlcjc1MzkzMzA=",
"avatar_url": "https://avatars.githubusercontent.com/u/7539330?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/davidcn21",
"html_url": "https://github.com/davidcn21",
"followers_url": "https://api.github.com/users/davidcn21/followers",
"following_url": "https://api.github.com/users/davidcn21/following{/other_user}",
"gists_url": "https://api.github.com/users/davidcn21/gists{/gist_id}",
"starred_url": "https://api.github.com/users/davidcn21/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/davidcn21/subscriptions",
"organizations_url": "https://api.github.com/users/davidcn21/orgs",
"repos_url": "https://api.github.com/users/davidcn21/repos",
"events_url": "https://api.github.com/users/davidcn21/events{/privacy}",
"received_events_url": "https://api.github.com/users/davidcn21/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Hi @davidcn21, we're currently developing a new API, and intend to add a site-specific API route for this. I'll update this thread when we have a public beta available. If you send us your email address, we may be able to share private access earlier if this is important to your research. My email is mkhorton@lbl.gov, or you can contact feedback@materialsproject.org."
] | 2020-05-25T18:42:30
| 2023-08-13T16:33:41
|
2023-08-13T16:33:41Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I've been using "get_xas_data" to read the site-average XAS for a given adsorption element. Just curious if there is a function to read the site-specific XAS data? thanks!
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1851/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1851/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1852
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1852/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1852/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1852/events
|
https://github.com/materialsproject/pymatgen/issues/1852
| 624,523,719
|
MDU6SXNzdWU2MjQ1MjM3MTk=
| 1,852
|
Option of plotter class to remove the grid lines on the background.
|
{
"login": "hongyi-zhao",
"id": 11155854,
"node_id": "MDQ6VXNlcjExMTU1ODU0",
"avatar_url": "https://avatars.githubusercontent.com/u/11155854?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hongyi-zhao",
"html_url": "https://github.com/hongyi-zhao",
"followers_url": "https://api.github.com/users/hongyi-zhao/followers",
"following_url": "https://api.github.com/users/hongyi-zhao/following{/other_user}",
"gists_url": "https://api.github.com/users/hongyi-zhao/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hongyi-zhao/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hongyi-zhao/subscriptions",
"organizations_url": "https://api.github.com/users/hongyi-zhao/orgs",
"repos_url": "https://api.github.com/users/hongyi-zhao/repos",
"events_url": "https://api.github.com/users/hongyi-zhao/events{/privacy}",
"received_events_url": "https://api.github.com/users/hongyi-zhao/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"No there is no option. You can just adjust the grid yourself using the code given by @gVallverdu . In general, there are an infinite variety of customizations for plots. There is no way we can support all of them. ",
"Hi\r\nIn case somebody look for an answer, hereafter is the way to remove the grid. You need first to get back the figure object.\r\n\r\n```py\r\napi = MPRester(api_key=\"XXX\")\r\nbs = api.get_bandstructure_by_material_id('mp-XYZT')\r\ndos = api.get_dos_by_material_id('mp-XYZT')\r\nbsdosplot = BSDOSPlotter(\r\n bs_projection=\"elements\",\r\n dos_projection=\"elements\",\r\n vb_energy_range=11,\r\n cb_energy_range=10,\r\n)\r\nplt = bsdosplot.get_plot(bs, dos=dos)\r\nplt.subplots_adjust(wspace = 0)\r\nfig = plt.gcf()\r\nall_axes = fig.get_axes()\r\nfor ax in all_axes:\r\n ax.grid(False)\r\n```\r\n\r\nFrom the `all_axes` array you can show/hide grids on one specific axes or do whatever you want on that axes.\r\n\r\n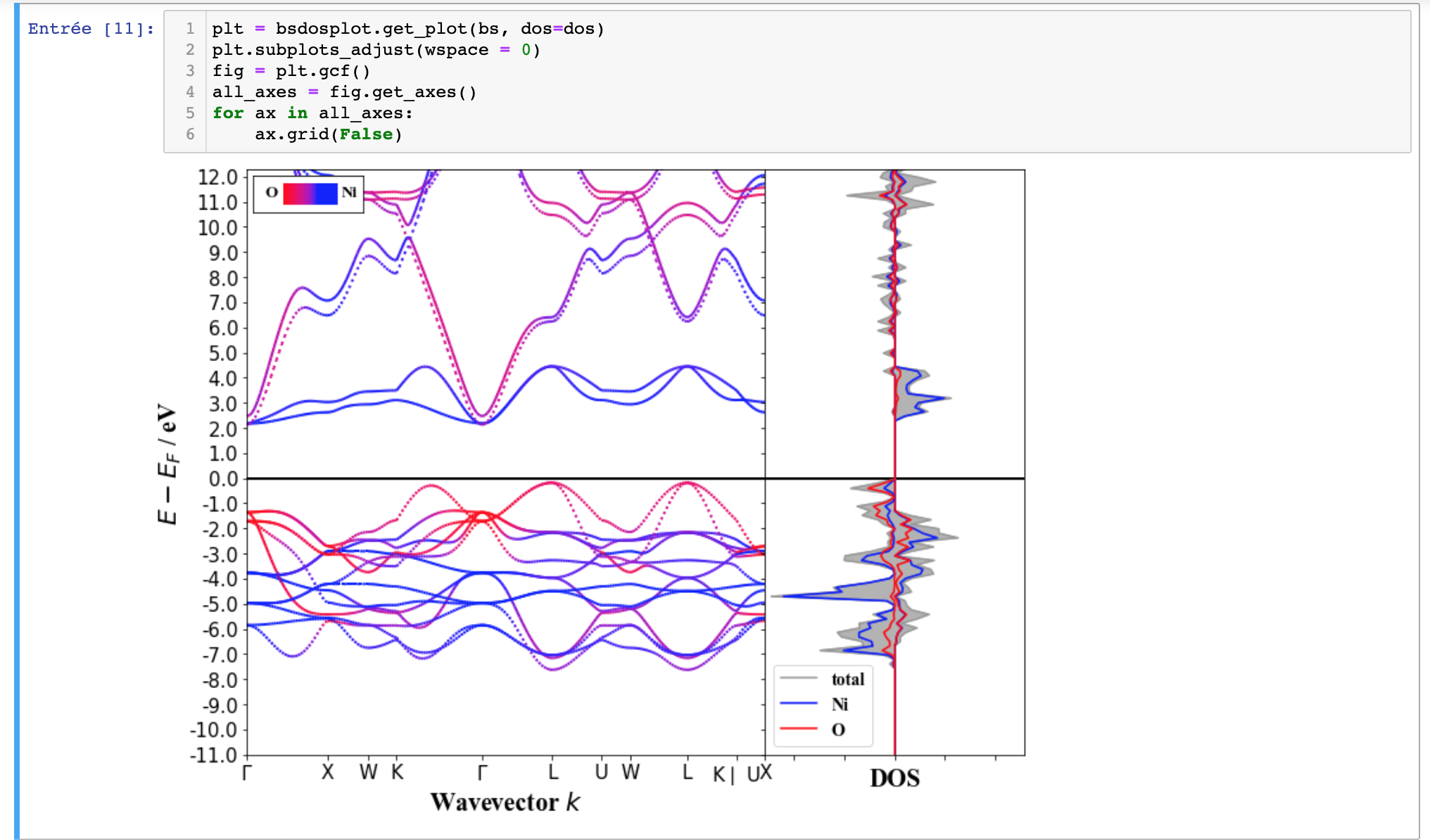\r\n",
"@gVallverdu Thanks for your notes. \r\nAnother issue: How to add the vertical lines on the plot at each high symmetry points?\r\n\r\nRegards"
] | 2020-05-25T23:49:26
| 2020-05-26T08:39:35
|
2020-05-26T02:40:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi,
The pymatgen plotter class uses grid lines on the background in default. I try to remove the grid lines, say, for the example below:
from matplotlib import pyplot as plt
from pymatgen.electronic_structure.plotter import BSDOSPlotter
from pymatgen.ext.matproj import MPRester
```
a = MPRester()
bs = a.get_bandstructure_by_material_id('mp-19009')
dos = a.get_dos_by_material_id('mp-19009')
bsdosplot = BSDOSPlotter(
bs_projection="elements",
dos_projection="elements",
vb_energy_range=11,
cb_energy_range=10,
# egrid_interval=2
)
plt = bsdosplot.get_plot(bs, dos=dos)
plt.subplots_adjust(wspace=0)
fig = plt.gcf()
```
Germain Salvato Vallverdu < germain.vallverdu @ univ - pau.fr > told me the following codes to do the trick by adding them at the end of the above example:
```
all_axes = fig.get_axes()
for ax in all_axes:
ax.grid(False)
```
But, I want to know whether there is an option of this pymatgen class for me to achieve the same aim?
Regards,
HY
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1852/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1852/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1853
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1853/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1853/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1853/events
|
https://github.com/materialsproject/pymatgen/issues/1853
| 625,013,493
|
MDU6SXNzdWU2MjUwMTM0OTM=
| 1,853
|
module 'pymatgen' has no attribute 'DeformedStructureSet'
|
{
"login": "glwhart",
"id": 7816856,
"node_id": "MDQ6VXNlcjc4MTY4NTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/7816856?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/glwhart",
"html_url": "https://github.com/glwhart",
"followers_url": "https://api.github.com/users/glwhart/followers",
"following_url": "https://api.github.com/users/glwhart/following{/other_user}",
"gists_url": "https://api.github.com/users/glwhart/gists{/gist_id}",
"starred_url": "https://api.github.com/users/glwhart/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/glwhart/subscriptions",
"organizations_url": "https://api.github.com/users/glwhart/orgs",
"repos_url": "https://api.github.com/users/glwhart/repos",
"events_url": "https://api.github.com/users/glwhart/events{/privacy}",
"received_events_url": "https://api.github.com/users/glwhart/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi Gus, welcome :-)\r\n\r\n`DeformedStructureSet` lives in `pymatgen.analysis.elasticity.strain' [here](https://pymatgen.org/pymatgen.analysis.elasticity.strain.html#pymatgen.analysis.elasticity.strain.DeformedStructureSet).\r\n\r\nRather than needing to include a long method call, it's usually easier to import explicitly at the top of your script, so you could write as follows:\r\n\r\n```\r\nimport pymatgen as mg\r\nfrom pymatgen.analysis.elasticity.strain import DeformedStructureSet\r\n\r\nhcp = mg.Lattice.hexagonal(3.23, 5.14)\r\nstruct = mg.Structure(hcp, [\"Zr\", \"Zr\"], [[0, 0, 0], [1/3, 1/3, 1/2]])\r\ndfm = DeformedStructureSet(struct, [0.1])\r\n```\r\n\r\nAnd then you access the deformed structures using `def.deformed_structures`.\r\n\r\nFinally, just a note to be careful with naming -- if you call your structure `str`, you're going to over-write the Python built-in \"string\" type.",
"Matthew, thank you for the quick and helpful response. I suspected it was something simple. "
] | 2020-05-26T16:23:51
| 2020-05-26T16:45:53
|
2020-05-26T16:45:04Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Not sure if this is a bug or just newbie user error---my first time trying to use pymatgen
When I call the `DeformedStructureSet` function, I get the message shown in the title
```
hcp = mg.Lattice.hexagonal(3.23,5.14)
str = mg.Structure(hcp,["Zr","Zr"],[[0,0,0],[1/3,1/3,1/2]])
dfm = mg.DeformedStructureSet(str,[0.1])
```
|
{
"login": "glwhart",
"id": 7816856,
"node_id": "MDQ6VXNlcjc4MTY4NTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/7816856?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/glwhart",
"html_url": "https://github.com/glwhart",
"followers_url": "https://api.github.com/users/glwhart/followers",
"following_url": "https://api.github.com/users/glwhart/following{/other_user}",
"gists_url": "https://api.github.com/users/glwhart/gists{/gist_id}",
"starred_url": "https://api.github.com/users/glwhart/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/glwhart/subscriptions",
"organizations_url": "https://api.github.com/users/glwhart/orgs",
"repos_url": "https://api.github.com/users/glwhart/repos",
"events_url": "https://api.github.com/users/glwhart/events{/privacy}",
"received_events_url": "https://api.github.com/users/glwhart/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1853/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1853/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1854
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1854/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1854/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1854/events
|
https://github.com/materialsproject/pymatgen/pull/1854
| 625,079,715
|
MDExOlB1bGxSZXF1ZXN0NDIzMzY2NTcx
| 1,854
|
[WIP] Adding interpolation functionality to VolumetricData class
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @ayushsgupta , thanks for this, this looks useful.\r\n\r\nDo you think you could add a method to take a slice of a plane, returning a 2D numpy array? This could make use of the functionality in `Lattice` to provide a plane as a `(h, k, l)` tuple. If this feature would not be useful for your work however, please disregard this request.",
"Hi @mkhorton, I think that would be a very useful feature. I did do some plane slicing, however it was limited to getting data on the `(x, 0, 0)`, `(0, x, 0)`, and `(0, 0, x)` planes only, not something like `(1, 1, 1)` or `(0, 2, 1)`. The first case is easier to add, but not quite sure how to do the second, as it would produce a possibly non-rectangular 2d array (?). I can look into it some more - what methods in `Lattice` would be helpful? I saw `get_miller_index_from_coords`, is there an inverse?",
"Also the CircleCI tests are failing, but it seems to be unrelated to the code I edited. The problem comes from some imports from sympy to `pymatgen.alchemy` and `pymatgen.analysis`",
"Hi @ayushsgupta, apologies for letting this PR sit here, everything looks great, thanks!",
"Can I confirm from you that this is no longer a [WIP] before I merge?",
"Hi @mkhorton no worries. I want to hold off for just a sec because I know @rkingsbury had some suggestions (change method name) and I want to make sure I covered it all. Also if you think that no warning needs to be raised when data is interpolated then I can combine two methods.",
"I wouldn't raise a warning, I think that's fine and clear from the docstring that interpolation is used.",
"@mkhorton ok, I've made all changes that I can think of. Build errors seem to come from the `Ion` class in `pymatgen.core`",
"Thanks Ayush, looks good! The IonEntry failures are known.",
"Could we have a test here too?",
"Ok. There don't seem to be any current tests for the VolumetricData class, so do you want me to create one or test the interpolation through a different class (elfcar, chgcar)?",
"Testing on one of the inherited classes is fine, thanks",
"@mkhorton done!",
"Thanks @ayushsgupta!"
] | 2020-05-26T18:03:37
| 2020-06-22T22:01:46
|
2020-06-21T23:39:59Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Adds the ability to interpolate between discrete data points
* Adds the `val_at` method, which accepts three coordinates and returns the data value at the associated point (possibly interpolated)
* Adds the `linear_slice` method, which gives a linear slice of the volumetric data between two specified points, with a specified resolution
## Additional dependencies introduced
* Uses RegularGridInterpolator from scipy.interpolate
## TODO
* Should there be some mechanism to raise a warning whenever a data point is interpolated as opposed to directly taken from data? There's a little bit of that (3222-3224, 3241) but not sure about the best implementation
* Is there a better way of checking list length than the assert statements on lines 3258-3259?
* If no warnings are necessary, then `val_at` and `_interpolate` can be combined
* Need to add testing!
* Add type checking for method arguments, specify return types
* Remove extra comments once everything's done
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1854/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1854/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1854",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1854",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1854.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1854.patch",
"merged_at": "2020-06-21T23:39:59Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1855
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1855/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1855/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1855/events
|
https://github.com/materialsproject/pymatgen/issues/1855
| 625,855,446
|
MDU6SXNzdWU2MjU4NTU0NDY=
| 1,855
|
symmetry_reduce: AttributeError: 'Structure' object has no attribute 'transform'
|
{
"login": "glwhart",
"id": 7816856,
"node_id": "MDQ6VXNlcjc4MTY4NTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/7816856?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/glwhart",
"html_url": "https://github.com/glwhart",
"followers_url": "https://api.github.com/users/glwhart/followers",
"following_url": "https://api.github.com/users/glwhart/following{/other_user}",
"gists_url": "https://api.github.com/users/glwhart/gists{/gist_id}",
"starred_url": "https://api.github.com/users/glwhart/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/glwhart/subscriptions",
"organizations_url": "https://api.github.com/users/glwhart/orgs",
"repos_url": "https://api.github.com/users/glwhart/repos",
"events_url": "https://api.github.com/users/glwhart/events{/privacy}",
"received_events_url": "https://api.github.com/users/glwhart/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The symmetry_reduce method is meant to operate on symmetry operations rather than a list of structures.\r\n\r\nFor the purposes of this method, you can simply initialize the DeformedStructureSet with the kwarg symmtry=True. E.g.,\r\n\r\n```\r\ndfm = DeformedStructureSet(structure, symmetry=True)\r\n```\r\n\r\nThis will perform symmetry reduction.\r\nWe have a Discourse forum at https://matsci.org/c/pymatgen which is meant for answering all help questions regarding pymatgen. In general, matsci.org is a collection of forums for various software packages used by materials scientists.",
"Thanks Shyue. Quite helpful.",
"Just a quick note, we've added redirects for each project to make the url easier to remember, so you can use just https://matsci.org/pymatgen",
"We're also welcoming other projects to host their discussions on that website for their own codes, an open offer !"
] | 2020-05-27T16:43:56
| 2020-05-27T18:00:22
|
2020-05-27T16:50:48Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
This is probably my misuse of pymatgen or my weak python skills but...
I created an hcp structure:
```
>>> hcpZr
Structure Summary
Lattice
abc : 3.23 3.2300000000000004 5.14
angles : 90.0 90.0 119.99999999999999
volume : 46.440704076633324
A : 3.23 0.0 1.9778045806229754e-16
B : -1.6149999999999993 2.7972620542237374 1.9778045806229754e-16
C : 0.0 0.0 5.14
PeriodicSite: Zr (0.0000, 0.0000, 0.0000) [0.0000, 0.0000, 0.0000]
PeriodicSite: Zr (0.5383, 0.9324, 2.5700) [0.3333, 0.3333, 0.5000]
```
then generated 24 distortions using `DeformedStructureSet`. Then I tried to reduce them using `symmetry_reduce`. The deformed set of structures are in the list `dfm`. I get the following error:
```
>>> uqdfm = symmetry_reduce(dfm,hcpZr)
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
File "/Users/hart/.virtualenvs/p3/lib/python3.7/site-packages/pymatgen/core/tensors.py", line 1039, in symmetry_reduce
if np.allclose(unique_tensor.transform(symmop), tensor, atol=tol):
AttributeError: 'Structure' object has no attribute 'transform'
```
Is there a forum where I should be asking for this kind of help? I'm not sure I should be raising github issues for these kinds of questions...
Thanks so much.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1855/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1855/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1856
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1856/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1856/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1856/events
|
https://github.com/materialsproject/pymatgen/issues/1856
| 626,749,243
|
MDU6SXNzdWU2MjY3NDkyNDM=
| 1,856
|
Illegal Instruction on neighbors.cpython-38-x86_64-linux-gnu.so for pymatgen==2020.4.29
|
{
"login": "hibagus",
"id": 9344035,
"node_id": "MDQ6VXNlcjkzNDQwMzU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9344035?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hibagus",
"html_url": "https://github.com/hibagus",
"followers_url": "https://api.github.com/users/hibagus/followers",
"following_url": "https://api.github.com/users/hibagus/following{/other_user}",
"gists_url": "https://api.github.com/users/hibagus/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hibagus/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hibagus/subscriptions",
"organizations_url": "https://api.github.com/users/hibagus/orgs",
"repos_url": "https://api.github.com/users/hibagus/repos",
"events_url": "https://api.github.com/users/hibagus/events{/privacy}",
"received_events_url": "https://api.github.com/users/hibagus/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @hibagus, thanks for the report -- we'll need more information to be able to fix this though, specifically a small/reproducible example of a specific call to the neighbor finding code. The CGCNN repo is not maintained by us, so we can't track down how exactly it's using pymatgen.\r\n\r\n@chc273 wrote many of these optimizations, it might be related to this pull request: https://github.com/materialsproject/pymatgen/pull/1832 ",
"Try doing a pip install pymatgen --upgrade to see if it solves the problem. But yes, in general, we do not maintain packages outside our domain....",
"You can also try a pip install directly from source, i.e. compiling the optimizations yourself. It could be that for some reason the pre-build wheel is bad on your system.",
"Hi @shyuep and @mkhorton, \r\nThanks for all of your suggestions.\r\n\r\nHere is what we have done based on your suggestions.\r\n1. We install the pymatgen using conda as suggested by the cgcnn as follows.\r\n`conda create --prefix /work/06156/bagus/maverick2/cgcnn python=3 scikit-learn pytorch torchvision pymatgen -c pytorch -c conda-forge`\r\n\r\nThis will install pymatgen 2020.4.29. The pymatgen installation gives us the Illegal Instruction problem.\r\n\r\n2. We uninstall the pymatgen and re-install the pymatgen using pip.\r\n```\r\npip uninstall pymatgen\r\npip install pymatgen\r\n```\r\nThis will also install pymatgen 2020.4.29. But now, there is no more illegal instruction problem. It seems that the build provided with conda-forge has the problem.\r\n\r\nThanks all.\r\n",
"Yes, I have no idea how conda-forge manages the install for Linux systems. They generally work well for Windows and Macs. For Linux, pip install is the most reliable given that you should have all the necessary gcc libs..."
] | 2020-05-28T19:17:51
| 2020-05-28T19:47:39
|
2020-05-28T19:47:39Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
We encounter illegal instruction when we are running the regression sample of [https://github.com/txie-93/cgcnn](url) that uses pymatgen.
We use the latest pymatgen version (2020.4.29).
Our gdb output is as follows.
```
(gdb) run main.py --train-ratio 0.6 --val-ratio 0.2 --test-ratio 0.2 data/sample-regression
The program being debugged has been started already.
Start it from the beginning? (y or n) y
Starting program: /work/06156/bagus/maverick2/cgcnn/bin/python main.py --train-ratio 0.6 --val-ratio 0.2 --test-ratio 0.2 data/sample-regression
[Thread debugging using libthread_db enabled]
Using host libthread_db library "/lib64/libthread_db.so.1".
Detaching after fork from child process 2240.
main.py:117: UserWarning: Dataset has less than 500 data points. Lower accuracy is expected.
warnings.warn('Dataset has less than 500 data points.
Program received signal SIGILL, Illegal instruction.
0x00002aab207b9ff2 in __pyx_pymod_exec_neighbors(_object*) () from /work/06156/bagus/maverick2/cgcnn/lib/python3.8/site-packages/pymatgen/optimization/neighbors.cpython-38-x86_64-linux-gnu.so
(gdb) quit
A debugging session is active.
Inferior 1 [process 2239] will be killed.
Quit anyway? (y or n) y
```
We have tried to run on two different machines. The first machine has Intel Core i5-8600 CPU running Ubuntu 18.04. The second machine has Intel Xeon E5-2620 v4 CPU running CentOS 7.4.1708.
**Downgrading the pymatgen to previous release 2020.4.2 solves the problem.**
Below is the installed packages in our environment when the problem arises.
```
package | build
---------------------------|-----------------
_libgcc_mutex-0.1 | conda_forge 3 KB conda-forge
_openmp_mutex-4.5 | 1_llvm 5 KB conda-forge
apscheduler-3.6.3 | py38h32f6830_1 81 KB conda-forge
ase-3.19.1 | py_0 1.5 MB conda-forge
beautifulsoup4-4.9.1 | py38h32f6830_0 163 KB conda-forge
blas-2.15 | mkl 10 KB conda-forge
brotlipy-0.7.0 |py38h1e0a361_1000 346 KB conda-forge
bzip2-1.0.8 | h516909a_2 396 KB conda-forge
ca-certificates-2020.4.5.1 | hecc5488_0 146 KB conda-forge
certifi-2020.4.5.1 | py38h32f6830_0 151 KB conda-forge
cffi-1.14.0 | py38hd463f26_0 223 KB conda-forge
cftime-1.1.3 | py38h8790de6_0 335 KB conda-forge
chardet-3.0.4 |py38h32f6830_1006 170 KB conda-forge
click-7.1.2 | pyh9f0ad1d_0 64 KB conda-forge
cryptography-2.9.2 | py38h766eaa4_0 620 KB conda-forge
cudatoolkit-10.2.89 | hfd86e86_1 365.1 MB
curl-7.69.1 | h33f0ec9_0 137 KB conda-forge
cycler-0.10.0 | py_2 9 KB conda-forge
decorator-4.4.2 | py_0 11 KB conda-forge
expat-2.2.9 | he1b5a44_2 191 KB conda-forge
fastcache-1.1.0 | py38h1e0a361_1 29 KB conda-forge
flask-1.1.2 | pyh9f0ad1d_0 70 KB conda-forge
freetype-2.10.2 | he06d7ca_0 905 KB conda-forge
future-0.18.2 | py38h32f6830_1 715 KB conda-forge
gmp-6.2.0 | he1b5a44_2 811 KB conda-forge
gmpy2-2.1.0b1 | py38h04dde30_0 212 KB conda-forge
hdf4-4.2.13 | hf30be14_1003 964 KB conda-forge
hdf5-1.10.6 |nompi_h3c11f04_100 3.0 MB conda-forge
icu-64.2 | he1b5a44_1 12.6 MB conda-forge
idna-2.9 | py_1 52 KB conda-forge
itsdangerous-1.1.0 | py_0 16 KB conda-forge
jinja2-2.11.2 | pyh9f0ad1d_0 93 KB conda-forge
joblib-0.15.1 | py_0 202 KB conda-forge
jpeg-9c | h14c3975_1001 251 KB conda-forge
jsoncpp-1.8.4 | hc9558a2_1002 162 KB conda-forge
kiwisolver-1.2.0 | py38hbf85e49_0 87 KB conda-forge
krb5-1.17.1 | h2fd8d38_0 1.5 MB conda-forge
latexcodec-2.0.0 | py_0 17 KB conda-forge
ld_impl_linux-64-2.34 | h53a641e_4 616 KB conda-forge
libblas-3.8.0 | 15_mkl 10 KB conda-forge
libcblas-3.8.0 | 15_mkl 10 KB conda-forge
libcurl-7.69.1 | hf7181ac_0 573 KB conda-forge
libedit-3.1.20170329 | hf8c457e_1001 172 KB conda-forge
libffi-3.2.1 | he1b5a44_1007 47 KB conda-forge
libgcc-ng-9.2.0 | h24d8f2e_2 8.2 MB conda-forge
libgfortran-ng-7.5.0 | hdf63c60_6 1.7 MB conda-forge
libiconv-1.15 | h516909a_1006 2.0 MB conda-forge
liblapack-3.8.0 | 15_mkl 10 KB conda-forge
liblapacke-3.8.0 | 15_mkl 10 KB conda-forge
libnetcdf-4.7.4 |nompi_h84807e1_104 1.3 MB conda-forge
libpng-1.6.37 | hed695b0_1 308 KB conda-forge
libssh2-1.9.0 | hab1572f_2 298 KB conda-forge
libstdcxx-ng-9.2.0 | hdf63c60_2 4.5 MB conda-forge
libtiff-4.1.0 | hc7e4089_6 668 KB conda-forge
libuuid-2.32.1 | h14c3975_1000 26 KB conda-forge
libwebp-base-1.1.0 | h516909a_3 845 KB conda-forge
libxcb-1.13 | h14c3975_1002 396 KB conda-forge
libxml2-2.9.10 | hee79883_0 1.3 MB conda-forge
llvm-openmp-10.0.0 | hc9558a2_0 2.8 MB conda-forge
lz4-c-1.9.2 | he1b5a44_1 226 KB conda-forge
markupsafe-1.1.1 | py38h1e0a361_1 26 KB conda-forge
matplotlib-base-3.2.1 | py38h2af1d28_0 7.1 MB conda-forge
mkl-2020.1 | 219 200.8 MB conda-forge
monty-3.0.2 | py_0 35 KB conda-forge
mpc-1.1.0 | h04dde30_1007 151 KB conda-forge
mpfr-4.0.2 | he80fd80_1 648 KB conda-forge
mpmath-1.1.0 | py_0 432 KB conda-forge
ncurses-6.1 | hf484d3e_1002 1.3 MB conda-forge
netcdf4-1.5.3 |nompi_py38hfd55d45_105 578 KB conda-forge
networkx-2.4 | py_1 1.2 MB conda-forge
ninja-1.10.0 | hc9558a2_0 1.9 MB conda-forge
numpy-1.18.4 | py38h8854b6b_0 5.3 MB conda-forge
olefile-0.46 | py_0 31 KB conda-forge
openssl-1.1.1g | h516909a_0 2.1 MB conda-forge
palettable-3.3.0 | py_0 86 KB conda-forge
pandas-1.0.3 | py38hcb8c335_1 11.5 MB conda-forge
pillow-7.1.2 | py38h9776b28_0 654 KB conda-forge
pip-20.1.1 | py_1 1.1 MB conda-forge
plotly-4.8.0 | pyh9f0ad1d_0 4.7 MB conda-forge
pthread-stubs-0.4 | h14c3975_1001 5 KB conda-forge
pybtex-0.22.2 | py38h32f6830_1 304 KB conda-forge
pycparser-2.20 | py_0 89 KB conda-forge
pymatgen-2020.4.29 | py38hbf85e49_0 3.3 MB conda-forge
pyopenssl-19.1.0 | py_1 47 KB conda-forge
pyparsing-2.4.7 | pyh9f0ad1d_0 60 KB conda-forge
pysocks-1.7.1 | py38h32f6830_1 27 KB conda-forge
python-3.8.2 |he5300dc_7_cpython 70.8 MB conda-forge
python-dateutil-2.8.1 | py_0 220 KB conda-forge
python_abi-3.8 | 1_cp38 4 KB conda-forge
pytorch-1.5.0 |py3.8_cuda10.2.89_cudnn7.6.5_0 424.7 MB pytorch
pytz-2020.1 | pyh9f0ad1d_0 227 KB conda-forge
pyyaml-5.3.1 | py38h1e0a361_0 190 KB conda-forge
readline-8.0 | hf8c457e_0 441 KB conda-forge
requests-2.23.0 | pyh8c360ce_2 47 KB conda-forge
retrying-1.3.3 | py_2 11 KB conda-forge
ruamel.yaml-0.16.6 | py38h1e0a361_1 167 KB conda-forge
ruamel.yaml.clib-0.2.0 | py38h1e0a361_1 160 KB conda-forge
scikit-learn-0.23.1 | py38h3a94b23_0 7.0 MB conda-forge
scipy-1.4.1 | py38h18bccfc_3 19.0 MB conda-forge
setuptools-47.1.0 | py38h32f6830_0 652 KB conda-forge
six-1.15.0 | pyh9f0ad1d_0 14 KB conda-forge
soupsieve-2.0.1 | py38h32f6830_0 57 KB conda-forge
spglib-1.15.1 | py38h8790de6_0 575 KB conda-forge
sqlite-3.30.1 | hcee41ef_0 2.0 MB conda-forge
sympy-1.6 | py38h32f6830_0 10.6 MB conda-forge
tabulate-0.8.7 | pyh9f0ad1d_0 24 KB conda-forge
tbb-2020.1 | hc9558a2_0 1.4 MB conda-forge
threadpoolctl-2.0.0 | pyh5ca1d4c_0 14 KB conda-forge
tk-8.6.10 | hed695b0_0 3.2 MB conda-forge
torchvision-0.6.0 | py38_cu102 11.7 MB pytorch
tornado-6.0.4 | py38h1e0a361_1 642 KB conda-forge
tqdm-4.46.0 | pyh9f0ad1d_0 50 KB conda-forge
tzlocal-2.1 | pyh9f0ad1d_0 18 KB conda-forge
urllib3-1.25.9 | py_0 92 KB conda-forge
vtk-8.2.0 | py38hf2e56f5_217 39.3 MB conda-forge
werkzeug-1.0.1 | pyh9f0ad1d_0 239 KB conda-forge
wheel-0.34.2 | py_1 24 KB conda-forge
xorg-kbproto-1.0.7 | h14c3975_1002 26 KB conda-forge
xorg-libice-1.0.10 | h516909a_0 57 KB conda-forge
xorg-libsm-1.2.3 | h84519dc_1000 25 KB conda-forge
xorg-libx11-1.6.9 | h516909a_0 918 KB conda-forge
xorg-libxau-1.0.9 | h14c3975_0 13 KB conda-forge
xorg-libxdmcp-1.1.3 | h516909a_0 18 KB conda-forge
xorg-libxt-1.2.0 | h516909a_0 374 KB conda-forge
xorg-xproto-7.0.31 | h14c3975_1007 72 KB conda-forge
xz-5.2.5 | h516909a_0 430 KB conda-forge
yaml-0.2.4 | h516909a_0 82 KB conda-forge
zlib-1.2.11 | h516909a_1006 105 KB conda-forge
zstd-1.4.4 | h6597ccf_3 991 KB conda-forge
------------------------------------------------------------
Total: 1.23 GB
```
|
{
"login": "hibagus",
"id": 9344035,
"node_id": "MDQ6VXNlcjkzNDQwMzU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9344035?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/hibagus",
"html_url": "https://github.com/hibagus",
"followers_url": "https://api.github.com/users/hibagus/followers",
"following_url": "https://api.github.com/users/hibagus/following{/other_user}",
"gists_url": "https://api.github.com/users/hibagus/gists{/gist_id}",
"starred_url": "https://api.github.com/users/hibagus/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/hibagus/subscriptions",
"organizations_url": "https://api.github.com/users/hibagus/orgs",
"repos_url": "https://api.github.com/users/hibagus/repos",
"events_url": "https://api.github.com/users/hibagus/events{/privacy}",
"received_events_url": "https://api.github.com/users/hibagus/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1856/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1856/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1857
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1857/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1857/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1857/events
|
https://github.com/materialsproject/pymatgen/pull/1857
| 627,273,293
|
MDExOlB1bGxSZXF1ZXN0NDI1MTA3MTI4
| 1,857
|
Fix labels in plot_thermodynamic_properties
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for noticing and submitting the fix @ab5424!"
] | 2020-05-29T13:35:06
| 2020-05-29T18:08:34
|
2020-05-29T18:08:19Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The labels seem to be incorrect (see https://github.com/materialsproject/pymatgen/blob/72d336d2d17ac140058fdcb74eb3336125bb12b7/pymatgen/phonon/dos.py#L229-L236 and https://github.com/materialsproject/pymatgen/blob/72d336d2d17ac140058fdcb74eb3336125bb12b7/pymatgen/phonon/dos.py#L263-L270 respecitvely)
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1857/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1857/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1857",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1857",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1857.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1857.patch",
"merged_at": "2020-05-29T18:08:19Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1858
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1858/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1858/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1858/events
|
https://github.com/materialsproject/pymatgen/issues/1858
| 627,527,743
|
MDU6SXNzdWU2Mjc1Mjc3NDM=
| 1,858
|
Elfcar.as_dict() from pymatgen.io.vasp.outputs not working
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-05-29T20:43:17
| 2020-06-01T19:37:31
|
2020-06-01T19:37:31Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I could be doing something wrong, but the `as_dict()` method of the `Elfcar` class doesn't appear to work. `Elfcar` ultimately inherits `as_dict()` from `monty.MSONable`, which requires that every argument passed to the constructor be an attribute of that class. However, one argument for the `Elfcar` constructor is `poscar`, but in the super call, `Elfcar` passes up `poscar.structure` to the `VolumetricData` constructor and does nothing further with `poscar`, so there isn't any `poscar` attribute associated with `Elfcar`. So `as_dict()` tries looking for a `poscar` attribute that doesn't exist, and a `NotImplementedError` is [raised](https://github.com/materialsvirtuallab/monty/blob/952ea44164d22767809374d04e8b9d6c41e13cd7/monty/json.py#L142). A simple fix could be to include `self.poscar = poscar` in the `Elfcar` constructor.
**To Reproduce**
`any_elfcar_object.as_dict()`
In my case, I created the object by calling `Elfcar.from_file()` on a gz file.
**Expected behavior**
Expected that a dictionary object would be returned.
**Screenshots**

**Desktop (please complete the following information):**
- Windows 10
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1858/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1858/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1859
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1859/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1859/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1859/events
|
https://github.com/materialsproject/pymatgen/pull/1859
| 627,579,642
|
MDExOlB1bGxSZXF1ZXN0NDI1MzYxMjU1
| 1,859
|
Add poscar attribute to Elfcar modeled after Chgcar
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for identifying the bug and for the fix, @ayushsgupta ! Can you add a test to verify that `elfcar = elfcar.from_dict(elfcar.as_dict())` ?\r\n",
"@rkingsbury ok I added tests & updated the docstring. Did you want the new tests in a different method or are they fine as is?",
"> \r\n> \r\n> @rkingsbury ok I added tests & updated the docstring. Did you want the new tests in a different method or are they fine as is?\r\n\r\nHi @ayushsgupta , I think the test is great where it is. For the docstring, we typically include all types in parentheses (unless we're using type hinting, which we are not in this module, yet). So the docstring should read\r\n\r\n```\r\nposcar (Poscar or Structure): Object containing structure.\r\n```\r\n ",
"Other than that, I think this is ready for review by @mkhorton . You might want to do another git pull in case that resolves the issues with the CircleCI test failure (not sure if that is fixed yet in master). Then you can go ahead and remove the [WIP]. \r\n ",
"Thanks @ayushsgupta and @rkingsbury!\r\n\r\nCircleCI has been failing recently, going to look into it today."
] | 2020-05-29T22:32:28
| 2020-06-22T22:01:57
|
2020-06-01T18:49:34Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* Add a `poscar` attribute to the `Elfcar` class patterned after the `Chgcar` constructor
* Fixes bug described in #1858
@rkingsbury @mkhorton
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1859/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1859/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1859",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1859",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1859.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1859.patch",
"merged_at": "2020-06-01T18:49:34Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1860
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1860/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1860/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1860/events
|
https://github.com/materialsproject/pymatgen/issues/1860
| 628,937,996
|
MDU6SXNzdWU2Mjg5Mzc5OTY=
| 1,860
|
Getting different INCAR files for 'mp-22590' using MPRester.query() within pymatgen and from materials-project website
|
{
"login": "kgmat",
"id": 66295557,
"node_id": "MDQ6VXNlcjY2Mjk1NTU3",
"avatar_url": "https://avatars.githubusercontent.com/u/66295557?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kgmat",
"html_url": "https://github.com/kgmat",
"followers_url": "https://api.github.com/users/kgmat/followers",
"following_url": "https://api.github.com/users/kgmat/following{/other_user}",
"gists_url": "https://api.github.com/users/kgmat/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kgmat/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kgmat/subscriptions",
"organizations_url": "https://api.github.com/users/kgmat/orgs",
"repos_url": "https://api.github.com/users/kgmat/repos",
"events_url": "https://api.github.com/users/kgmat/events{/privacy}",
"received_events_url": "https://api.github.com/users/kgmat/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-06-02T06:33:03
| 2023-08-13T16:33:42
|
2023-08-13T16:33:41Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hello,
I am getting different INCAR files for ID 'mp-22590' using **MPRester.query()** within pymatgen and from the **materials-project website**.
**using MPRester.query():**
mpr = MPRester('My_API_key')
data=mpr.query('mp-22590',['input.incar'])[0]
INCAR that i get is:
ALGO = Normal
AMIX = 0.2
AMIX_MAG = 0.8
BMIX = 0.001
BMIX_MAG = 0.001
ENCUT = 520
**IBRION = 1**
ICHARG = 1
ISIF = 3
**ISMEAR = 1**
ISPIN = 2
LDAU = True
LDAUJ = 0 0 0
LDAUL = 0 2 0
LDAUTYPE = 2
LDAUU = 0 5.3 0
LREAL = Auto
LWAVE = True
MAGMOM = 12 * 0.6 4 * 5.0 4 * 0.6
NELM = 100
NELMDL = 6
NELMIN = 3
NPAR = 1
**NSW = 200**
PREC = Accurate
**SIGMA = 0.2**
SYSTEM = Rubyvaspy :: o fe la
and accesing VASP input files for 'mp-22590' from the materials-project website, INCAR is:
ALGO = Fast
EDIFF = 0.001
ENCUT = 520
**IBRION = 2**
ICHARG = 1
ISIF = 3
**ISMEAR = -5**
ISPIN = 2
LDAU = True
LDAUJ = 0 0 0
LDAUL = 0 2 0
LDAUPRINT = 1
LDAUTYPE = 2
LDAUU = 0 5.3 0
LMAXMIX = 6
LORBIT = 11
LREAL = Auto
LWAVE = False
MAGMOM = 4 * 0.6 4 * 5.0 12 * 0.6
NELM = 100
**NSW = 99**
PREC = Accurate
**SIGMA = 0.05**
Am i doing something wrong?
I would appreciate any help on this.
Thanks
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1860/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1860/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1861
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1861/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1861/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1861/events
|
https://github.com/materialsproject/pymatgen/pull/1861
| 629,731,306
|
MDExOlB1bGxSZXF1ZXN0NDI2OTkyMjk0
| 1,861
|
SOC WAVECARs and general cleanup
|
{
"login": "mturiansky",
"id": 8866816,
"node_id": "MDQ6VXNlcjg4NjY4MTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/8866816?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mturiansky",
"html_url": "https://github.com/mturiansky",
"followers_url": "https://api.github.com/users/mturiansky/followers",
"following_url": "https://api.github.com/users/mturiansky/following{/other_user}",
"gists_url": "https://api.github.com/users/mturiansky/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mturiansky/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mturiansky/subscriptions",
"organizations_url": "https://api.github.com/users/mturiansky/orgs",
"repos_url": "https://api.github.com/users/mturiansky/repos",
"events_url": "https://api.github.com/users/mturiansky/events{/privacy}",
"received_events_url": "https://api.github.com/users/mturiansky/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The tests are failing because of sympy (not related to my work so far). Looks like check_assumptions is no longer in sympy.core.assumptions.",
"Thanks @mturiansky. As far as I can see it _is_ still in sympy.core.assumptions, been meaning to take a look at this. Nevertheless not related to this PR, is this ready to merge except for this test?",
"I still want to add the WAVECAR to UNK conversion. I'll mark it \"ready for review\" to turn it from a draft to a PR when it's ready.",
"Great, thanks.",
"By the way, #1585 can probably be closed now, it was addressed previously but will definitely be addressed now.\r\n\r\nAlso, heads up @bernstei, I removed the \"gamma\" argument and replaced it with the \"vasp_type\" argument. If you don't specify \"gamma\" it will work fine, but if you specified \"gamma\" somewhere, then you will get a unexpected keyword TypeError.",
"This PR is ready to merge. The failing circleci is not my fault.\r\n\r\nThat being said, I think I know how to fix it. I played around with your docker image and was able to fix the sympy error. The version of sympy that is installed by conda seems to be bad. My guess is that it was generated with an older python version and something with the module loading has changed. If you install sympy with pip instead of conda, everything works. However, I found that using \"conda uninstall sympy\" didn't uninstall things completely/correctly. Here is the sequence of commands I used:\r\n```bash\r\n$ docker pull materialsvirtuallab/circle-ci-pmg-py3:3.7.3\r\n$ docker run --name test -it materialsvirtuallab/circle-ci-pmg-py3:3.7.3 bash\r\n```\r\nand then inside the docker container:\r\n```bash\r\n$ export PATH=$HOME/miniconda3/bin:$PATH\r\n$ conda config --system --add channels conda-forge\r\n$ # NOTE: no sympy in next command\r\n$ conda create -q -n test-environment python=3.7 numpy scipy matplotlib cython pandas networkx pytest pip cmake h5py openbabel python-igraph\r\n$ source activate test-environment\r\n$ git clone https://github.com/materialsproject/pymatgen.git && cd pymatgen/\r\n$ pip install --ignore-installed -r requirements.txt -r requirements-optional.txt -r requirements-dev.txt\r\n$ pip install -e .\r\n$ # now, check that it worked with one of the tests that failed before\r\n$ pytest pymatgen/analysis/tests/test_surface_analysis.py\r\n```\r\nI hope that fixes things, @shyuep @mkhorton . You may need to clear the caches on circleci (is that a thing you can do? I haven't used circleci myself).",
"See #1864 ",
"Can I get a release once this is merged, please?",
"Sure thing, and thanks for investigating the failed test. CircleCI calculates a hash to determine if it needs to clear its cache, I'll check the logic there.",
"This PR is excellent, thanks for all your work on it.",
"Thanks! I noticed that CirceCI calculates a hash, but it only does this for the requirements*.txt files. When you change the CircleCI config file, which has the conda installation scripts, the cache isn't regenerated, so the changes don't propagate. I addressed this in my other pull request.",
"Release pushed. Trying out a new way of building wheels for PyPI for this release, let me know if there are any issues.",
"Thanks, again! Seems to work well on my machine so far.",
"We are having an issue during building for Windows: https://dev.azure.com/conda-forge/feedstock-builds/_build/results?buildId=172210&view=logs&j=171a126d-c574-5c8c-1269-ff3b989e923d&t=1183ba29-a0b5-5324-8463-2a49ace9e213&l=2044\r\n\r\n`ImportError: cannot import name 'FortranEOFError'`\r\n\r\nDoes this depend on a specific scipy version?",
"Oh boy, I hadn't even thought of that. The fortran reader was added as early as v0.13, but the FortranEORError wasn't added until v1.4.0 as far as I can tell.",
"No worries, I can push a supplemental release if necessary.",
"Is there documentation on this? Or an example on how to use it? :)",
"Hi @mangerij , you can read the api documentation for the Wavecar [here](https://pymatgen.org/pymatgen.io.vasp.outputs.html#pymatgen.io.vasp.outputs.Wavecar). There's a few different ways to use the code:\r\n- use the convenience functions like get_parchg or write_unks\r\n- use fft_mesh to get the pseudowavefunctions on a FFT grid\r\n- use the Wavecar.coeffs variable directly, which stores the pseudowavefunction coefficients on the reduced grid in reciprocal space (the grid points are in Wavecar.Gpoints)\r\nIt really depends on what you're trying to do with it.",
"Thanks Mark. Everything seems to work pretty nicely. I'm interested in write_unk for the SOC WAVECARs to try to feed to wannier90.\n\nHowever, my WAVECAR is pretty large... 5GB, so it seems to be pretty slow. Any tips to speed up?\n\nI am wondering if we could add some gpu Accelerate library lines to some of these classes as an option to speed up these member functions if needed. Might play around with that\n________________________________\nFrom: Mark Turiansky <notifications@github.com>\nSent: Monday, June 22, 2020 10:23 PM\nTo: materialsproject/pymatgen <pymatgen@noreply.github.com>\nCc: Mangeri, John <john.mangeri@uconn.edu>; Mention <mention@noreply.github.com>\nSubject: Re: [materialsproject/pymatgen] SOC WAVECARs and general cleanup (#1861)\n\n\n*Message sent from a system outside of UConn.*\n\n\nHi @mangerij<https://nam10.safelinks.protection.outlook.com/?url=https%3A%2F%2Fgithub.com%2Fmangerij&data=02%7C01%7Cjohn.mangeri%40uconn.edu%7Cc14e52fefa2447dcdd5808d816ea32b2%7C17f1a87e2a254eaab9df9d439034b080%7C0%7C0%7C637284542420499309&sdata=UZVld7UWFQKsi6ZzJx6KNQcJlGEyAqaHMTQtbP8J%2BSA%3D&reserved=0> , you can read the api documentation for the Wavecar here<https://nam10.safelinks.protection.outlook.com/?url=https%3A%2F%2Fpymatgen.org%2Fpymatgen.io.vasp.outputs.html%23pymatgen.io.vasp.outputs.Wavecar&data=02%7C01%7Cjohn.mangeri%40uconn.edu%7Cc14e52fefa2447dcdd5808d816ea32b2%7C17f1a87e2a254eaab9df9d439034b080%7C0%7C0%7C637284542420509304&sdata=kJQevRycXdbTAtGpU2jvq2M71B7duMsC0xzTWjY18hE%3D&reserved=0>. There's a few different ways to use the code:\n\n * use the convenience functions like get_parchg or write_unks\n * use fft_mesh to get the pseudowavefunctions on a FFT grid\n * use the Wavecar.coeffs variable directly, which stores the pseudowavefunction coefficients on the reduced grid in reciprocal space (the grid points are in Wavecar.Gpoints)\nIt really depends on what you're trying to do with it.\n\n—\nYou are receiving this because you were mentioned.\nReply to this email directly, view it on GitHub<https://nam10.safelinks.protection.outlook.com/?url=https%3A%2F%2Fgithub.com%2Fmaterialsproject%2Fpymatgen%2Fpull%2F1861%23issuecomment-647750717&data=02%7C01%7Cjohn.mangeri%40uconn.edu%7Cc14e52fefa2447dcdd5808d816ea32b2%7C17f1a87e2a254eaab9df9d439034b080%7C0%7C0%7C637284542420509304&sdata=a6yx9cFXAti3IUygm6NQqwx4Jip2cyQ9i5hgGLfmrj4%3D&reserved=0>, or unsubscribe<https://nam10.safelinks.protection.outlook.com/?url=https%3A%2F%2Fgithub.com%2Fnotifications%2Funsubscribe-auth%2FABZ65FECXNG5JVLLXT2LNKDRX64V7ANCNFSM4NRLWKVA&data=02%7C01%7Cjohn.mangeri%40uconn.edu%7Cc14e52fefa2447dcdd5808d816ea32b2%7C17f1a87e2a254eaab9df9d439034b080%7C0%7C0%7C637284542420519294&sdata=S7VkZtyA9GqQZEQWAzJMiETF1ewdHQE8dtiqR%2FqI%2FvY%3D&reserved=0>.\n",
"I believe most of the code is IO limited. Definitely in reading in the WAVECAR, you're limited by IO. A few of the computations can be sped up using something like numba or cython in principle (for example, generating the Gpoints), but the speed up you'll gain is essentially negligible for large files like this.\r\n\r\nFor writing the UNK files, I think you're again mostly IO limited, but generating the FFT mesh could probably be sped up. Right now, it has a python loop that could probably be sped up with numba. If I have some time, I'll do some profiling and see if that would make a noticeable difference."
] | 2020-06-03T06:33:36
| 2020-06-23T19:33:28
|
2020-06-08T19:53:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* add ability to read SOC WAVECARs
* seemingly robust ability to determine if WAVECAR is from std, gam, or ncl binary
* add conversion from WAVECAR to UNK files for wannier90 (will add needed support for UNKs from ncl calculations)
## Additional dependencies introduced (if any)
None.
## TODO (if any)
- [x] update evaluate_wavefunc and fft_mesh functions to handle ncl WAVECARs
- [x] update get_parchg for ncl WAVECARs
- [x] update old tests
- [x] add new tests for ncl WAVECAR
- [x] add UNK class + tests
- [x] add function to generate UNKs from WAVECAR
- [x] type annotations
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1861/reactions",
"total_count": 2,
"+1": 2,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1861/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1861",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1861",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1861.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1861.patch",
"merged_at": "2020-06-08T19:53:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1862
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1862/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1862/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1862/events
|
https://github.com/materialsproject/pymatgen/issues/1862
| 630,096,127
|
MDU6SXNzdWU2MzAwOTYxMjc=
| 1,862
|
Restore structures from their pymatgen __str__ representation
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"While we can certainly implement a parser for the `Structure.__str__`, the general preference is to use one of the other formats for storing structures, e.g., CIF, POSCAR or json. These are well-established representations that people have been using widely. The structure string is more for the purposes of readability and a quick glance at the basic information. I am not sure that we'd want to introduce a new representation for these purposes.\r\n\r\n@mkhorton Views?\r\n\r\nIf we are really going to introduce a new format, I'd rather it be something simple and structured. E.g., we use YAML for the `__str__` and rely on the yaml parser to get it back. This will have the added feature that we can easily transform to json if needed. I do not really want to be responsible for maintaining a new format for structures....",
"One issue to highlight is that the co-ordinates are truncated to 3dp in this str as opposed to 6dp in a CIF. This alone would make this a bad way to store structures but the functionality may still be helpful?",
"Yes, the number of significant figures can be changed of course....",
"Yes, much better to write using the JSON representation (to_json / as_dict / from_dict), these are also string representations.\r\n\r\n> The need for such functionality arises when restoring a dataframe of structures that has been saved in a csv format\r\n\r\nRegarding this need specifically, I know pandas offers extension types ... is there a way we can hook into pandas so that when written to csv, the `to_json` method is used instead of `str`?",
"Note that the `str` representation is a little lossy too, it may not show site properties sufficiently etc. and will be very difficult to write a general parser for since site properties can take many compound data types."
] | 2020-06-03T15:32:17
| 2020-06-03T16:56:50
|
2020-06-03T16:56:50Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
`Structure` has a str representation defined below that can be written to a file however I can't find an easy way to restore a `Structure` from that string, is there one? it doesn't appear to be an option when using `Stucture.from_str()`
The need for such functionality arises when restoring a dataframe of structures that has been saved in a csv format
### Example of str
> Reduced Formula: CrNi3
abc : 3.501092 3.501092 4.348644
angles: 113.737682 113.737682 90.000000
Sites (4)
\# SP a b c magmom
\--- ---- ---- ---- ---- --------
0 Cr 0 0 0 1.484
1 Ni 0.5 0.5 -0 -0.008
2 Ni 0.25 0.75 0.5 -0.057
3 Ni 0.75 0.25 0.5 -0.057
https://github.com/materialsproject/pymatgen/blob/72d336d2d17ac140058fdcb74eb3336125bb12b7/pymatgen/core/structure.py#L2014
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1862/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1862/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1863
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1863/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1863/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1863/events
|
https://github.com/materialsproject/pymatgen/issues/1863
| 631,158,327
|
MDU6SXNzdWU2MzExNTgzMjc=
| 1,863
|
Bug: Finding nn distances in the pymatgen_diffusion package
|
{
"login": "chemist29",
"id": 45610381,
"node_id": "MDQ6VXNlcjQ1NjEwMzgx",
"avatar_url": "https://avatars.githubusercontent.com/u/45610381?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chemist29",
"html_url": "https://github.com/chemist29",
"followers_url": "https://api.github.com/users/chemist29/followers",
"following_url": "https://api.github.com/users/chemist29/following{/other_user}",
"gists_url": "https://api.github.com/users/chemist29/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chemist29/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chemist29/subscriptions",
"organizations_url": "https://api.github.com/users/chemist29/orgs",
"repos_url": "https://api.github.com/users/chemist29/repos",
"events_url": "https://api.github.com/users/chemist29/events{/privacy}",
"received_events_url": "https://api.github.com/users/chemist29/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Pls make sure ou have the latest pymatgen and pymatgen-diffusion packages. Also, for pymatgen-diffusion, pls post on the Github issues of that package. "
] | 2020-06-04T21:25:28
| 2020-06-05T15:28:23
|
2020-06-05T15:28:23Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
Hello Pymatgen Dev,
There appears to be an error in the pymatgen_diffusion package. Exactly it is finding nn distances with the pymatgen_diffusion package. That has not been updated in a long time and it appears to be broken.
**To Reproduce**
Steps to reproduce the behavior:
Unzip file
1. Command is python neb_end_member_setup Na2O.cif
2. In diffusing species, enter Na
3. You should see error, attached is my output; might have to make some paths work as per your environment.
[neb_end_member_setup.zip](https://github.com/materialsproject/pymatgen/files/4732787/neb_end_member_setup.zip)
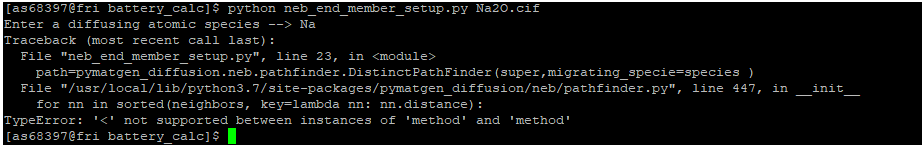
**Expected behavior**
The line,
cubic=cube(min_length=10,min_atoms=50, max_atoms=150)
Should make a cubic supercell that is at least 10 Angstrom wide with between 50 and 150 atoms
**Desktop (please complete the following information):**
- OS: Linux
- Version: 4.19.0-6-amd64
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1863/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1863/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1864
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1864/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1864/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1864/events
|
https://github.com/materialsproject/pymatgen/pull/1864
| 632,204,566
|
MDExOlB1bGxSZXF1ZXN0NDI4OTc4ODgw
| 1,864
|
pip sympy instead of conda sympy
|
{
"login": "mturiansky",
"id": 8866816,
"node_id": "MDQ6VXNlcjg4NjY4MTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/8866816?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mturiansky",
"html_url": "https://github.com/mturiansky",
"followers_url": "https://api.github.com/users/mturiansky/followers",
"following_url": "https://api.github.com/users/mturiansky/following{/other_user}",
"gists_url": "https://api.github.com/users/mturiansky/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mturiansky/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mturiansky/subscriptions",
"organizations_url": "https://api.github.com/users/mturiansky/orgs",
"repos_url": "https://api.github.com/users/mturiansky/repos",
"events_url": "https://api.github.com/users/mturiansky/events{/privacy}",
"received_events_url": "https://api.github.com/users/mturiansky/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2020-06-06T04:48:45
| 2020-06-08T20:05:46
|
2020-06-08T20:00:51Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Attempt to fix circleci.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1864/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1864/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1864",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1864",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1864.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1864.patch",
"merged_at": "2020-06-08T20:00:51Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1865
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1865/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1865/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1865/events
|
https://github.com/materialsproject/pymatgen/issues/1865
| 634,712,173
|
MDU6SXNzdWU2MzQ3MTIxNzM=
| 1,865
|
Illegal instruction error when running VoronoiNN.get_nn_info()
|
{
"login": "mtdunstan",
"id": 7153579,
"node_id": "MDQ6VXNlcjcxNTM1Nzk=",
"avatar_url": "https://avatars.githubusercontent.com/u/7153579?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mtdunstan",
"html_url": "https://github.com/mtdunstan",
"followers_url": "https://api.github.com/users/mtdunstan/followers",
"following_url": "https://api.github.com/users/mtdunstan/following{/other_user}",
"gists_url": "https://api.github.com/users/mtdunstan/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mtdunstan/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mtdunstan/subscriptions",
"organizations_url": "https://api.github.com/users/mtdunstan/orgs",
"repos_url": "https://api.github.com/users/mtdunstan/repos",
"events_url": "https://api.github.com/users/mtdunstan/events{/privacy}",
"received_events_url": "https://api.github.com/users/mtdunstan/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Please install using pip install the latest version of pymatgen.",
"Hi @mtdunstan , I can't reproduce this -- note that VoronoiNN relies on qhull integrated into scipy, try upgrading your scipy and/or your Python version also."
] | 2020-06-08T15:18:23
| 2020-06-08T16:46:50
|
2020-06-08T15:19:07Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I had older code using VoronoiNN.get_nn_info on structures that I tried to use today, and it is giving an illegal instruction error. Minimal code is below. Normally it should produce substructural information for each site in the structure.
Do you know what I can try to do to fix this?
```
from pymatgen.ext.matproj import MPRester
from pymatgen.core.structure import Structure, Composition
from pymatgen.analysis.local_env import VoronoiNN
a = MPRester("ZbTstiE2olAnyU64")
entry = a.get_entry_by_material_id('mp-104',inc_structure='final')
s = entry.structure
vcf = VoronoiNN()
vs = vcf.get_nn_info(s, 0)
```
Desktop:
- OS: Mac 10.12.6 (but running on University cluster)
- Version 2020.4.29
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1865/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1865/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1866
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1866/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1866/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1866/events
|
https://github.com/materialsproject/pymatgen/pull/1866
| 634,991,814
|
MDExOlB1bGxSZXF1ZXN0NDMxNDQzMTM2
| 1,866
|
Fix appveyor
|
{
"login": "mturiansky",
"id": 8866816,
"node_id": "MDQ6VXNlcjg4NjY4MTY=",
"avatar_url": "https://avatars.githubusercontent.com/u/8866816?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mturiansky",
"html_url": "https://github.com/mturiansky",
"followers_url": "https://api.github.com/users/mturiansky/followers",
"following_url": "https://api.github.com/users/mturiansky/following{/other_user}",
"gists_url": "https://api.github.com/users/mturiansky/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mturiansky/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mturiansky/subscriptions",
"organizations_url": "https://api.github.com/users/mturiansky/orgs",
"repos_url": "https://api.github.com/users/mturiansky/repos",
"events_url": "https://api.github.com/users/mturiansky/events{/privacy}",
"received_events_url": "https://api.github.com/users/mturiansky/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"We're probably overdue removing AppVeyor now anyway, we used it because it offered Windows testing, but we can do that within Github Actions now and reduce our CI overload.",
"Yeah, probably worth it. I was just annoyed that the appveyor badge said the build was failing :-)",
"Haha, I admire your diligence"
] | 2020-06-08T23:09:25
| 2020-06-08T23:20:10
|
2020-06-08T23:15:54Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I assume it has the same problem as CircleCI with conda sympy.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1866/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1866/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1866",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1866",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1866.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1866.patch",
"merged_at": "2020-06-08T23:15:54Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1867
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1867/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1867/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1867/events
|
https://github.com/materialsproject/pymatgen/issues/1867
| 635,284,072
|
MDU6SXNzdWU2MzUyODQwNzI=
| 1,867
|
Fail to read CIF file from the CSD database
|
{
"login": "danieleongari",
"id": 25899092,
"node_id": "MDQ6VXNlcjI1ODk5MDky",
"avatar_url": "https://avatars.githubusercontent.com/u/25899092?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/danieleongari",
"html_url": "https://github.com/danieleongari",
"followers_url": "https://api.github.com/users/danieleongari/followers",
"following_url": "https://api.github.com/users/danieleongari/following{/other_user}",
"gists_url": "https://api.github.com/users/danieleongari/gists{/gist_id}",
"starred_url": "https://api.github.com/users/danieleongari/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/danieleongari/subscriptions",
"organizations_url": "https://api.github.com/users/danieleongari/orgs",
"repos_url": "https://api.github.com/users/danieleongari/repos",
"events_url": "https://api.github.com/users/danieleongari/events{/privacy}",
"received_events_url": "https://api.github.com/users/danieleongari/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"After further testing, using the previous version `pymatgen-2020.4.29`, the script as:\r\n```\r\nparser = CifParser(\"FIQCEN.cif\", occupancy_tolerance=100)\r\n```\r\nworks fine.\r\nPlease check if the is a bug in the latest version and document the need of `occupancy_tolerance=somehighnumber` in https://pymatgen.org/usage.html\r\n",
"Hi @danieleongari ,\r\n\r\nThese are not errors, they are warnings:\r\n\r\n```\r\nUserWarning: Issues encountered while parsing CIF:\r\nUserWarning: Some occupancies ([10, 19, 6, 6, 6, 6]) sum to > 1! If they are within the tolerance, they will be rescaled.\r\nUserWarning: Species occupancies sum to more than 1!\r\n```\r\n\r\n(removed the file paths so it can be read more easily)\r\n\r\nThe warning is to tell you that some aspects of your CIF are unusual: it's claiming that there are more than one atom on a given site, so some manual attention is suggested to make sure this is what you intend.\r\n\r\n> which results in core dumped.\r\n\r\nThis is typically an installation issue, it could be that your `spglib` needs to be updated? I'll test also to see if 2020.6.8 has a specific issue, thanks for reporting.",
"Hi @danieleongari , just to update on this we did find an issue with the Linux wheels for 2020.6.8 so a re-install might help.",
"Thanks @mkhorton for letting me know.\r\nAside from the \"core dumped\" problem,\r\nI would still suggest to make the error more verbose and suggesting how to fix the warning, because the definition of \"occupancy_tolerance\" is not a trivial concept and might confuse the user: I had to spend some time diving into the code to catch it!\r\n\r\nSomething like:\r\n```\r\nUserWarning: Species occupancies sum to more than 1! Increase occupancy_tolerance to get the structure.\r\n```",
"That's a good suggestion, thanks! Will make a change to improve the phrasing.\r\n\r\nFor some context (or for anyone else reading this for which it might be useful), occupancies on an individual site should ideally sum to 1.0. What this means is that if the site is disordered (it can contain more than one element on average) then statistically we know what elements are present, and what the probability is that the site contains a given element.\r\n\r\nA sum > 1.0 doesn't make physical sense because then the probabilities are no longer well defined. However, in some large databases, this does happen sometimes -- often because occupation probabilities for individual elements are measured independently and so the small amount of error means they round to just about 1.0 and that's where \"occupancy_tolerance\" is most useful. The other scenario is that some databases don't follow the CIF specification appropriately, which means they're using occupancies that sum up to 100 instead of 1. Technically, this is an error, but if you know to look for it you can just scale occupancies down by a factor of 100 to get the correct result."
] | 2020-06-09T09:51:38
| 2020-06-11T16:07:09
|
2020-06-11T08:43:56Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When running
```
from pymatgen.io.cif import CifParser
parser = CifParser("FIQCEN.cif")
structure = parser.get_structures()[0]
```
with FIQCEN being a structure from the CSD database (https://www.ccdc.cam.ac.uk/solutions/csd-system/components/csd/), the script returns the error:
```
/home/daniele/anaconda3/envs/csd/lib/python3.7/site-packages/pymatgen/io/cif.py:1106: UserWarning: Issues encountered while parsing CIF:
warnings.warn("Issues encountered while parsing CIF:")
/home/daniele/anaconda3/envs/csd/lib/python3.7/site-packages/pymatgen/io/cif.py:1108: UserWarning: Some occupancies ([10, 19, 6, 6, 6, 6]) sum to > 1! If they are within the tolerance, they will be rescaled.
warnings.warn(error)
/home/daniele/anaconda3/envs/csd/lib/python3.7/site-packages/pymatgen/io/cif.py:1108: UserWarning: Species occupancies sum to more than 1!
```
Note that the file is correctly visualized by software such as VESTA:

[FIQCEN.cif.log](https://github.com/materialsproject/pymatgen/files/4751327/FIQCEN.cif.log)
I tried also
```
parser = CifParser("FIQCEN.cif", occupancy_tolerance=100)
```
which results in `core dumped`.
Thanks for your support!
Environment:
- OS: Linux
- Version: 2020.6.8
|
{
"login": "danieleongari",
"id": 25899092,
"node_id": "MDQ6VXNlcjI1ODk5MDky",
"avatar_url": "https://avatars.githubusercontent.com/u/25899092?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/danieleongari",
"html_url": "https://github.com/danieleongari",
"followers_url": "https://api.github.com/users/danieleongari/followers",
"following_url": "https://api.github.com/users/danieleongari/following{/other_user}",
"gists_url": "https://api.github.com/users/danieleongari/gists{/gist_id}",
"starred_url": "https://api.github.com/users/danieleongari/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/danieleongari/subscriptions",
"organizations_url": "https://api.github.com/users/danieleongari/orgs",
"repos_url": "https://api.github.com/users/danieleongari/repos",
"events_url": "https://api.github.com/users/danieleongari/events{/privacy}",
"received_events_url": "https://api.github.com/users/danieleongari/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1867/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1867/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1868
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1868/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1868/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1868/events
|
https://github.com/materialsproject/pymatgen/issues/1868
| 636,109,762
|
MDU6SXNzdWU2MzYxMDk3NjI=
| 1,868
|
StructureMatcher.get_rms_dist illegal instruction on release pymatgen==2020.6.8.
|
{
"login": "mortengjerding",
"id": 27740483,
"node_id": "MDQ6VXNlcjI3NzQwNDgz",
"avatar_url": "https://avatars.githubusercontent.com/u/27740483?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mortengjerding",
"html_url": "https://github.com/mortengjerding",
"followers_url": "https://api.github.com/users/mortengjerding/followers",
"following_url": "https://api.github.com/users/mortengjerding/following{/other_user}",
"gists_url": "https://api.github.com/users/mortengjerding/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mortengjerding/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mortengjerding/subscriptions",
"organizations_url": "https://api.github.com/users/mortengjerding/orgs",
"repos_url": "https://api.github.com/users/mortengjerding/repos",
"events_url": "https://api.github.com/users/mortengjerding/events{/privacy}",
"received_events_url": "https://api.github.com/users/mortengjerding/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"If you installed pymatgen via conda, pls try to do pip install --upgrade pymatgen. This error is usually a symptom that something is wrong with the compilation of the C extensions and happens quite frequently on Linux based systems.",
"I have not installed pymatgen through conda. I have simply pip installed it. This is also making my pipelines fail on gitlab so it's not just my laptap. I just ran the above script in a virtual environment and it still fails. I set the venv up like in the following:\r\n\r\n```\r\n$ python3 -m venv venv\r\n$ . venv/bin/activate\r\n$ pip install pymatgen\r\n$ python test_script.py\r\nIllegal instruction (core dumped)\r\n```\r\nSo I don't see that it should be something with my setup. Since you closed the issue I was hoping you could help me figure out what is wrong. As I wrote: It worked in the previous release.",
"Sorry I forgot that I also had to install ASE in the venv to make the script work. The magic incantation to set up the environment is:\r\n\r\n```\r\n$ python3 -m venv venv\r\n$ . venv/bin/activate\r\n$ pip install pymatgen ase\r\n$ python test_script.py\r\nIllegal instruction (core dumped)\r\n```",
"Does it fail for all scripts, or is it just the specific code you showed that is an issue? As far as I know, nothing has changed since v2020.4 that would cause this. There were some issues with sympy...",
"I'm sorry I don't know whether anything has broken since I am essentially only using the StructureMatcher functionality in my scripts. Could it be that something went wrong in the building of the wheels (I am no expert on this)? Just to make absolute sure that everything works in the previous release I checked this in a venv:\r\n\r\n```\r\n$ python3 -m venv venv\r\n$ . venv/bin/activate\r\n$ pip install pymatgen==2020.4.29 ase\r\n$ python test_script.py\r\n$ # No error\r\n```\r\nCan the issue be reproduced on your pc?",
"I do not have such an error on my Mac. It would help if you can run other scripts to check if this is something wrong in the core of pymatgen or whether it is just in StructureMatcher. Otherwise, I have no way of debugging this.",
"Ok so I have found that the function \"find_points_in_spheres\" is causing the crash. I have made a minimal example here which crashed on my computer:\r\n\r\n```\r\nimport numpy as np \r\nfrom pymatgen.optimization.neighbors import find_points_in_spheres \r\n \r\nall_coords = np.array([[0., 0., 0.]], dtype=float) \r\ncenter_coords = np.array([[0., 0., 0.]], dtype=float) \r\nr = 2.863855528636639 \r\npbc = np.array([1, 1, 1]) \r\nlattice = np.array([[0., 2.025, 2.025], \r\n [2.025, 0., 2.025], \r\n [2.025, 2.025, 0.]], dtype=float) \r\n \r\nfind_points_in_spheres(all_coords=all_coords, \r\n center_coords=center_coords, \r\n r=r, \r\n pbc=pbc, \r\n lattice=lattice, \r\n tol=1e-8)\r\n```\r\nThis function is being called when the structurematcher tries to Niggli reduce the input structures. Has something changed in this function?",
"I hope this helps, otherwise let me know.",
"We're also encountering this issue in our CI, see the https://github.com/aiidateam/aiida-core/pull/4111\r\n\r\nMy suspicion is that this is caused by the wheel being built against a CPU instruction set that is not fully supported on the machines where it crashes.\r\n\r\n@mortengjerding what's the output of `cat /proc/cpuinfo` for you?\r\n\r\nLooking at the wheel itself (`pymatgen-2020.6.8-cp38-cp38-manylinux2010_x86_64.whl` in our case) it seems it's using AVX2 instructions. Grepping from the root of the unpacked wheel:\r\n\r\n```bash\r\n$ grep -inR AVX2 .\r\nBinary file ./pymatgen/optimization/neighbors.cpython-38-x86_64-linux-gnu.so matches\r\n```\r\n\r\nIf that's the case, the immediate workaround would be installing it without the wheel, i.e.\r\n```bash\r\npip install --no-binary pymatgen pymatgen\r\n```",
"Ah thank you. Indeed, that is likely the issue. The strange thing is, I never upload wheels for Linux precisely because of this reason. @mkhorton Do you know what happened with the previous release? Usually I just upload the Windows and Mac wheels and for Linux, I just assume that it can be compiled from the tar.gz easily. Linux is very fussy about compilers etc.",
"@greschd That would also be my understanding. The output of cpuinfo indicates that my processor supports avx2 instructions:\r\n\r\n```\r\n$ cat /proc/cpuinfo\r\nprocessor : 0 \r\nvendor_id : GenuineIntel \r\ncpu family : 6 \r\nmodel : 142 \r\nmodel name : Intel(R) Core(TM) i5-8250U CPU @ 1.60GHz \r\nstepping : 10 \r\nmicrocode : 0xca \r\ncpu MHz : 761.181 \r\ncache size : 6144 KB \r\nphysical id : 0 \r\nsiblings : 8 \r\ncore id : 0 \r\ncpu cores : 4 \r\napicid : 0 \r\ninitial apicid : 0 \r\nfpu : yes \r\nfpu_exception : yes \r\ncpuid level : 22 \r\nwp : yes \r\nflags : fpu vme de pse tsc msr pae mce cx8 apic sep mtrr pge mca cmov pat pse36 clflush dts acpi mmx fxsr sse sse2 ss ht tm pbe syscall nx pdpe1gb rdtscp lm\\\r\n constant_tsc art arch_perfmon pebs bts rep_good nopl xtopology nonstop_tsc cpuid aperfmperf tsc_known_freq pni pclmulqdq dtes64 monitor ds_cpl vmx est tm2 ssse3 sdbg\\\r\n fma cx16 xtpr pdcm pcid sse4_1 sse4_2 x2apic movbe popcnt tsc_deadline_timer aes xsave avx f16c rdrand lahf_lm abm 3dnowprefetch cpuid_fault epb invpcid_single pti s\\\r\nsbd ibrs ibpb stibp tpr_shadow vnmi flexpriority ept vpid fsgsbase tsc_adjust bmi1 avx2 smep bmi2 erms invpcid mpx rdseed adx smap clflushopt intel_pt xsaveopt xsavec\\\r\n xgetbv1 xsaves dtherm ida arat pln pts hwp hwp_notify hwp_act_window hwp_epp md_clear flush_l1d \r\nbugs : cpu_meltdown spectre_v1 spectre_v2 spec_store_bypass l1tf mds swapgs itlb_multihit \r\nbogomips : 3600.00 \r\nclflush size : 64 \r\ncache_alignment : 64 \r\naddress sizes : 39 bits physical, 48 bits virtual \r\npower management:\r\n```",
"@mortengjerding Can you try pip install --no-binary pymatgen and see if that causes the problem to disappear? We will remove the Linux wheels in future releases.",
"Yeah, it could well be a different incompatibility in which instructions are supported..\r\n\r\n> \r\n> \r\n> @mortengjerding Can you try pip install --no-binary pymatgen and see if that causes the problem to disappear? We will remove the Linux wheels in future releases.\r\n\r\nNote that you have to pass `pymatgen` _twice_ to pip - once to tell it which package to ignore binaries for, and once to tell it which package to install.",
"I have just tried and can confirm that installing with `pip install --no-binary pymatgen pymatgen` makes the problem disappear. Thanks a lot for the help!",
"Thanks for the debug. We will handle the rest from our end.",
"@shyuep maybe you could _remove_ the Linux wheels for the current release? Not sure what your policy on removing already-published wheels is, but in this case it's likely to fix more issues than it causes.",
"I already did. Removing wheels is fine. For Linux users, they will default to the tar.gz anyway.",
"Fantastic, thanks!",
"> Do you know what happened with the previous release?\r\n\r\nYes, this was on me -- I was trying to improve our cross-platform support with the latest release for pip users. Wheels should have been compliant with manylinux2010 (see [PEP 571](https://www.python.org/dev/peps/pep-0571/)) and seemed to work. However, evidently this is not the case, so thanks for the report @mortengjerding and @greschd and my apologies for the inconvenience. Back to the drawing board on this!",
"Ok. @mkhorton Let's not distribute wheels for Linux in future. The benefit of wheels on Linux platforms is marginal, since there are seldom compilation issues on those platforms. Wheels are useful mainly for Windows and Macs where the compilers may sometimes not be available."
] | 2020-06-10T09:54:34
| 2020-06-10T15:44:16
|
2020-06-10T15:44:16Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
On the most recent release of pymatgen (2020.6.8) I am now getting "Illegal instruction" when running the `pymatgen.structure_matcher.StructureMatcher.get_rms_dist`. This did not happen on the previous release (2020.4.29).
**To Reproduce**
The following script
```
from ase.build import bulk
from pymatgen.io.ase import AseAtomsAdaptor
from pymatgen.analysis.structure_matcher import StructureMatcher
al = bulk('Al')
adaptor = AseAtomsAdaptor()
structure = adaptor.get_structure(al)
StructureMatcher().get_rms_dist(structure, structure)
```
results in "Illegal instruction (core dumped)".
**Expected behavior**
The result of the get_rms_dist should be zero.
**Desktop (please complete the following information):**
```
morten@hp ~ $ python3
Python 3.6.9 (default, Apr 18 2020, 01:56:04)
[GCC 8.4.0] on linux
>>> import pymatgen
>>> pymatgen.__version__
'2020.6.8'
```
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1868/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1868/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1869
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1869/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1869/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1869/events
|
https://github.com/materialsproject/pymatgen/issues/1869
| 636,432,533
|
MDU6SXNzdWU2MzY0MzI1MzM=
| 1,869
|
Invalid entries in MPRester
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
},
{
"login": "shyamd",
"id": 16827130,
"node_id": "MDQ6VXNlcjE2ODI3MTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/16827130?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyamd",
"html_url": "https://github.com/shyamd",
"followers_url": "https://api.github.com/users/shyamd/followers",
"following_url": "https://api.github.com/users/shyamd/following{/other_user}",
"gists_url": "https://api.github.com/users/shyamd/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyamd/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyamd/subscriptions",
"organizations_url": "https://api.github.com/users/shyamd/orgs",
"repos_url": "https://api.github.com/users/shyamd/repos",
"events_url": "https://api.github.com/users/shyamd/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyamd/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[] | 2020-06-10T17:42:04
| 2023-08-13T16:33:42
|
2023-08-13T16:33:42Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
When querying for entries in MPRester, e.g., for phase diagram construction, I keep getting lots of "Entry mp-*** has invalid run type None or LDA+rVV10. Discarding.". I have no idea why there are so many runs with invalid run types. But shouldn't a better way be to exclude these from the query, e.g., by setting the criteria, when the compatibly_only=True is set, rather than querying for all entries and then removing them? I would assume not getting them from MP in the first place would be much more efficient from a bandwidth and speed perspective.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1869/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1869/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1870
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1870/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1870/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1870/events
|
https://github.com/materialsproject/pymatgen/issues/1870
| 637,151,485
|
MDU6SXNzdWU2MzcxNTE0ODU=
| 1,870
|
pymatgen.Composition, Invalid formula when atom fraction is < 1e-4
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-06-11T16:15:08
| 2020-06-11T23:27:53
|
2020-06-11T23:27:53Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The `Compostion.to_data_dict` property will report `CompositionError: e-05 is an invalid formula!` error when any atom fraction is less than 1e-4.
**To Reproduce**
Steps to reproduce the behavior:
```
from pymatgen import Composition
comp = Composition('Fe0.00009Ni0.99991')
print(comp.to_data_dict)
```
However, `comp._data` shows the correct atom fraction without problems.
**Expected behavior**
I would not expect such an error.
**Desktop (please complete the following information):**
- OS: MacOS
- Version 10.15.4
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1870/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1870/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1871
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1871/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1871/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1871/events
|
https://github.com/materialsproject/pymatgen/pull/1871
| 637,200,663
|
MDExOlB1bGxSZXF1ZXN0NDMzMjMyNzgz
| 1,871
|
Add "unit" to plot_compare in PhononBSPlotter
|
{
"login": "ab5424",
"id": 57258530,
"node_id": "MDQ6VXNlcjU3MjU4NTMw",
"avatar_url": "https://avatars.githubusercontent.com/u/57258530?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ab5424",
"html_url": "https://github.com/ab5424",
"followers_url": "https://api.github.com/users/ab5424/followers",
"following_url": "https://api.github.com/users/ab5424/following{/other_user}",
"gists_url": "https://api.github.com/users/ab5424/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ab5424/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ab5424/subscriptions",
"organizations_url": "https://api.github.com/users/ab5424/orgs",
"repos_url": "https://api.github.com/users/ab5424/repos",
"events_url": "https://api.github.com/users/ab5424/events{/privacy}",
"received_events_url": "https://api.github.com/users/ab5424/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hello, \r\n\r\nis there anything else we would need to do before this can be merged? I think, only py3test is currently failing but this has nothing to do with this change.\r\n\r\nBest, \r\nJG",
"Thanks @ab5424! And apologies both @ab5424 and @JaGeo, been a little slow reviewing PRs this week. We currently have a test failing on master that needs to be addressed, it is unrelated to this PR.",
"Thank you very much for merging, @mkhorton !"
] | 2020-06-11T17:28:06
| 2020-06-20T20:28:54
|
2020-06-20T17:24:07Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
The frequency unit can only be changed for a single bandstructure plot: https://github.com/materialsproject/pymatgen/blob/604d59c93f21c4a180663bdc6cd64bf1329c5239/pymatgen/phonon/plotter.py#L329 This is missing for plot_compare. I also provided a test (although it is not very useful to compare a bandstructure object with itself).
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1871/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1871/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1871",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1871",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1871.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1871.patch",
"merged_at": "2020-06-20T17:24:07Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1872
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1872/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1872/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1872/events
|
https://github.com/materialsproject/pymatgen/pull/1872
| 638,209,804
|
MDExOlB1bGxSZXF1ZXN0NDM0MDQwNzM3
| 1,872
|
Adding the option to plot other side of slab
|
{
"login": "mhsiron",
"id": 49957831,
"node_id": "MDQ6VXNlcjQ5OTU3ODMx",
"avatar_url": "https://avatars.githubusercontent.com/u/49957831?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mhsiron",
"html_url": "https://github.com/mhsiron",
"followers_url": "https://api.github.com/users/mhsiron/followers",
"following_url": "https://api.github.com/users/mhsiron/following{/other_user}",
"gists_url": "https://api.github.com/users/mhsiron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mhsiron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mhsiron/subscriptions",
"organizations_url": "https://api.github.com/users/mhsiron/orgs",
"repos_url": "https://api.github.com/users/mhsiron/repos",
"events_url": "https://api.github.com/users/mhsiron/events{/privacy}",
"received_events_url": "https://api.github.com/users/mhsiron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @mhsiron, some linting failures here otherwise looks useful",
"Hi @mhsiron, any updates on this?",
"Thanks @mhsiron !"
] | 2020-06-13T18:34:07
| 2020-09-26T02:51:10
|
2020-09-26T02:51:04Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Adding the option to plot other side of slab, that way the user has the option to plot either surface of the slab, especially if the two surfaces are different.
## Additional dependencies introduced (if any)
None
Before a pull request can be merged, the following items must be checked:
- [ x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ x] Tests have been added for any new functionality or bug fixes.
- [ x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1872/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1872/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1872",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1872",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1872.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1872.patch",
"merged_at": "2020-09-26T02:51:04Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1873
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1873/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1873/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1873/events
|
https://github.com/materialsproject/pymatgen/pull/1873
| 642,920,719
|
MDExOlB1bGxSZXF1ZXN0NDM3ODE3NzE1
| 1,873
|
Add new functionality to Lobsterin class and some fixes
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi!\r\nI don't think the failing tests are from my changes.\r\n\r\nBest, \r\nJG",
"We've had an IonEntry bug in master for the past week, I think a PR from @knc6 just fixed it. Please try merging in from master and hopefully tests should pass, otherwise this looks great.\r\n\r\nWhat's the distinction between the min/max PBE_54 bases?",
"I hope it is working now.\r\n\r\nmin: all occupied valence orbitals according to the POTCAR\r\nmax: all valence orbitals according to the POTCAR\r\nBoth files have an additional condition: basis functions have to be present in Lobster.",
"Sorry, my fault. Will fix the test - I have moved one file.",
"Thanks @JaGeo !"
] | 2020-06-22T09:45:34
| 2020-06-23T00:55:50
|
2020-06-23T00:55:40Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
-I have introduced a new static method to the Lobsterin class that can return a list of possible basis sets for Lobster runs. And, I also added files that help creating theses basis sets.
-To restructure the code, I have also created a lobster folder and moved tests and classes there. I have also made sure that the code is backwards compatible.
-I have fixed some tests for Lobster and also added a new one for the new functionalities.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1873/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1873/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1873",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1873",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1873.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1873.patch",
"merged_at": "2020-06-23T00:55:40Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1874
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1874/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1874/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1874/events
|
https://github.com/materialsproject/pymatgen/issues/1874
| 643,384,489
|
MDU6SXNzdWU2NDMzODQ0ODk=
| 1,874
|
Link to Bader analysis code no longer working
|
{
"login": "ayushsgupta",
"id": 16586331,
"node_id": "MDQ6VXNlcjE2NTg2MzMx",
"avatar_url": "https://avatars.githubusercontent.com/u/16586331?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/ayushsgupta",
"html_url": "https://github.com/ayushsgupta",
"followers_url": "https://api.github.com/users/ayushsgupta/followers",
"following_url": "https://api.github.com/users/ayushsgupta/following{/other_user}",
"gists_url": "https://api.github.com/users/ayushsgupta/gists{/gist_id}",
"starred_url": "https://api.github.com/users/ayushsgupta/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/ayushsgupta/subscriptions",
"organizations_url": "https://api.github.com/users/ayushsgupta/orgs",
"repos_url": "https://api.github.com/users/ayushsgupta/repos",
"events_url": "https://api.github.com/users/ayushsgupta/events{/privacy}",
"received_events_url": "https://api.github.com/users/ayushsgupta/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Edit to docs_rst, thanks!"
] | 2020-06-22T21:54:36
| 2022-04-20T03:25:26
|
2022-04-20T03:25:26Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
For the [Bader caller module](https://pymatgen.org/pymatgen.command_line.bader_caller.html) in the docs, the link to download the original fortran code for the Bader analysis is broken. The link should be http://theory.cm.utexas.edu/henkelman/code/bader/. Not sure how to fix (edit docs or docs_rst?).
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1874/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1874/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1875
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1875/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1875/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1875/events
|
https://github.com/materialsproject/pymatgen/issues/1875
| 643,394,586
|
MDU6SXNzdWU2NDMzOTQ1ODY=
| 1,875
|
XYZ File Parser can incorrectly handle comment string, miscounting the number of sites
|
{
"login": "stevetorr",
"id": 12805012,
"node_id": "MDQ6VXNlcjEyODA1MDEy",
"avatar_url": "https://avatars.githubusercontent.com/u/12805012?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/stevetorr",
"html_url": "https://github.com/stevetorr",
"followers_url": "https://api.github.com/users/stevetorr/followers",
"following_url": "https://api.github.com/users/stevetorr/following{/other_user}",
"gists_url": "https://api.github.com/users/stevetorr/gists{/gist_id}",
"starred_url": "https://api.github.com/users/stevetorr/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/stevetorr/subscriptions",
"organizations_url": "https://api.github.com/users/stevetorr/orgs",
"repos_url": "https://api.github.com/users/stevetorr/repos",
"events_url": "https://api.github.com/users/stevetorr/events{/privacy}",
"received_events_url": "https://api.github.com/users/stevetorr/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Update: I have discovered that removing the bottom lines does not solve the problem, nor did removing the comment lines, **but removing the scientific notation from the _first_ atomic coordinate does**. When the same coordinate is supplied to a different atom, the parsing appears to complete correctly; so the issue may be when the scientific notation occurs on the very first coordinate line.",
"OK, I got it. **The issue arises from the fact that the asterisk and ^ notation for the coordinate values is unsupported, so no bug in XYZ.** The index error occurs because the regular expression parser that detects when the coord. lines begin doesn't look for the asterisk or carat characters. The parsing can complete after that, but it will not correctly handle the *^ notation. I'll look at adding this functionality in."
] | 2020-06-22T22:17:27
| 2020-06-22T22:51:59
|
2020-06-22T22:51:36Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
For some XYZ files, the parser will incorrectly use the end of the comment line as the number of sites present on the molecule, which can result in an `IndexError` if this exceeds the number of sites on the Molecule. (This could presumably also result in fewer atoms being loaded in then should be, but the 'too many sites' case is how I caught the bug).
**To Reproduce**
A minimal example is as follows (I included a case which works, if it helps with debugging):
```
xyz_string_good = """17
gdb 73000 2.52453 1.59492 1.2866 1.942 70.04 -0.2468 -0.0322 0.2146 1026.4232 0.133427 -458.930556 -458.92199 -458.921046 -458.963286 32.347
C 0.0518622525 1.4677402164 0.1677182445 -0.508727
C 0.149931291 -0.0198512527 0.1549183945 0.387141
O -1.0420748387 -0.7755524938 -0.1098982654 -0.333377
C -0.6310443779 -1.7822217642 0.8717895672 -0.126641
C 0.6995752769 -1.0823398662 1.0716407643 -0.218953
C 1.3176925774 -0.8886070534 -0.3223080253 0.261566
O 1.126586626 -1.8073146011 -1.3481691631 -0.482546
C 2.6622014749 -0.2647569257 -0.3439137992 0.098299
O 3.1033556604 0.4506871538 0.5272840719 -0.280588
H 1.022706895 1.903540259 0.4101533166 0.164412
H -0.2702279642 1.8391291673 -0.8111459282 0.133684
H -0.6788048518 1.7939332661 0.9141342159 0.133712
H -1.290366036 -1.7421645256 1.743319629 0.126989
H -0.6147407007 -2.7905570041 0.4474086287 0.123682
H 1.2938641569 -0.9969950588 1.9709215363 0.126881
H 0.2057168389 -1.7109885321 -1.6301219799 0.297897
H 3.2419713095 -0.515832035 -1.256903708 0.096568
126.6464 179.5404 202.2048 207.8956 237.9939 281.042 297.7535 342.195 453.9626 488.255 552.616 632.3338 681.2005 700.0387 753.8057 906.3622 916.4433 947.4678 966.6321 981.1379 1039.9824 1069.6445 1114.022 1136.5327 1161.4481 1178.0761 1207.8283 1245.4708 1326.0224 1366.8841 1396.7 1423.2805 1443.7657 1482.1065 1496.0189 1507.6194 1797.0553 2919.6036 3040.9772 3050.7498 3103.7278 3106.9445 3145.2569 3215.8312 3736.7032
CC12OCC1C2(O)C=O C[C@]12OC[C@H]1[C@]2(O)C=O
InChI=1S/C6H8O3/c1-5-4(2-9-5)6(5,8)3-7/h3-4,8H,2H2,1H3 InChI=1S/C6H8O3/c1-5-4(2-9-5)6(5,8)3-7/h3-4,8H,2H2,1H3/t4-,5+,6-/m1/s1
"""
xyz_string_bad = """21
gdb 73686 2.29238 1.75978 1.36114 1.4885 76.6 -0.2457 0.0799 0.3256 1046.4543 0.185139 -424.239329 -424.231407 -424.230463 -424.271698 31.284
C 8.3963*^-6 1.5082640218 0.0003678516 -0.475108
C -0.0079983479 -0.0204373839 -0.0071456046 0.318224
C -1.4203683358 -0.6309054519 0.0819864887 -0.14791
O -1.2692539395 -1.8561456788 0.7815062589 -0.247008
C -0.3553538009 -1.5796789091 1.8355022746 -0.106377
C 0.7198523887 -0.6676543053 1.2216354779 -0.080206
C 1.9191247691 -1.4078050364 0.5876323806 -0.096694
O 2.1134029805 -0.823343344 -0.6902996964 -0.241434
C 0.8087264769 -0.5914550703 -1.1926885825 -0.137307
H 1.023353157 1.8877808556 -0.0816471023 0.129439
H -0.5750176226 1.9127699899 -0.8407064652 0.115221
H -0.4363110498 1.9050689906 0.9234010836 0.113272
H -2.0994717752 0.0473780136 0.6260414552 0.082664
H -1.8654921365 -0.8530151745 -0.8927588239 0.107815
H -0.8699247922 -1.0714572655 2.6672231337 0.087199
H 0.0305838701 -2.5351905239 2.2031794572 0.108945
H 1.0602691131 0.0888559672 1.9323936092 0.075195
H 2.8500581495 -1.303628955 1.1507960083 0.106419
H 1.690607141 -2.4820838452 0.4874835885 0.091712
H 0.3581226539 -1.5342926331 -1.5436952327 0.092353
H 0.8901048443 0.0967787983 -2.0385808704 0.103587
73.5764 196.484 253.7954 257.9211 270.4176 343.3211 382.1862 403.1166 549.8545 601.6342 647.5136 727.6591 789.9801 866.5038 917.2463 942.4403 960.8136 967.8604 978.0593 996.6799 1012.2977 1029.2324 1102.3525 1108.503 1138.8784 1176.679 1188.3401 1203.2474 1235.4067 1250.0353 1282.1394 1290.1216 1305.3509 1334.4425 1359.7115 1370.0445 1387.0336 1404.1107 1422.9901 1495.1326 1497.5941 1500.3616 1506.3038 1516.5771 1518.5447 2959.1115 2962.6023 2971.3962 2971.876 3025.3569 3074.6102 3079.041 3081.7835 3086.3615 3091.1404 3096.6227 3104.1382
CC12COCC1COC2 C[C@]12COC[C@H]1COC2
InChI=1S/C7H12O2/c1-7-4-8-2-6(7)3-9-5-7/h6H,2-5H2,1H3 InChI=1S/C7H12O2/c1-7-4-8-2-6(7)3-9-5-7/h6H,2-5H2,1H3/t6-,7+
"""
from pymatgen.io.xyz import XYZ
correct_parse = XYZ.from_string(xyz_string_good)
bad_parse = XYZ.from_string(xyz_string_bad)
```
The error message collected by the bad_parse is an `IndexError` which results when the `num_sites` variable within the `_from_frame_string` method is incorrectly set to 284, which are the final three digits of the comment line. The index error then occurs when the parser attempts to parse a line that is not there.
**Expected behavior**
The parsing is from a file from the QM9 database, and the XYZ parser works great for most of them; There are only a small handful of files which exhibit this error, one of which is included above.
**Desktop (please complete the following information):**
- OS: Mac OSX 10.15.5 (19F101)
- Version 2020.3.13 (which has the most recent edits on [the XYZ file](https://github.com/materialsproject/pymatgen/blob/4ec6ba1ee328fdfbec0e83b9401fe8b92120c2df/pymatgen/io/xyz.py#L15) ).
**Screenshots** I could glean no difference in the strings that would give a clear explanation of why the parsing fails, but I have included two screenshots of the first two lines as they are loaded into the `XYZ` parser from the files themselves (however, in the version I copy+pasted above, the tabs have been converted to spaces-- the error still occurs either way).
The string which works:

The string which doesn't:

I hope this helps-- I tried to debug it by comparing the strings themselves but couldn't make progress in the time I tried.
|
{
"login": "stevetorr",
"id": 12805012,
"node_id": "MDQ6VXNlcjEyODA1MDEy",
"avatar_url": "https://avatars.githubusercontent.com/u/12805012?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/stevetorr",
"html_url": "https://github.com/stevetorr",
"followers_url": "https://api.github.com/users/stevetorr/followers",
"following_url": "https://api.github.com/users/stevetorr/following{/other_user}",
"gists_url": "https://api.github.com/users/stevetorr/gists{/gist_id}",
"starred_url": "https://api.github.com/users/stevetorr/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/stevetorr/subscriptions",
"organizations_url": "https://api.github.com/users/stevetorr/orgs",
"repos_url": "https://api.github.com/users/stevetorr/repos",
"events_url": "https://api.github.com/users/stevetorr/events{/privacy}",
"received_events_url": "https://api.github.com/users/stevetorr/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1875/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1875/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1876
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1876/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1876/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1876/events
|
https://github.com/materialsproject/pymatgen/pull/1876
| 643,441,997
|
MDExOlB1bGxSZXF1ZXN0NDM4MjQ0Nzcz
| 1,876
|
Add support for *^ scientific notation for XYZ parser (Useful for QM9 Database)
|
{
"login": "stevetorr",
"id": 12805012,
"node_id": "MDQ6VXNlcjEyODA1MDEy",
"avatar_url": "https://avatars.githubusercontent.com/u/12805012?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/stevetorr",
"html_url": "https://github.com/stevetorr",
"followers_url": "https://api.github.com/users/stevetorr/followers",
"following_url": "https://api.github.com/users/stevetorr/following{/other_user}",
"gists_url": "https://api.github.com/users/stevetorr/gists{/gist_id}",
"starred_url": "https://api.github.com/users/stevetorr/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/stevetorr/subscriptions",
"organizations_url": "https://api.github.com/users/stevetorr/orgs",
"repos_url": "https://api.github.com/users/stevetorr/repos",
"events_url": "https://api.github.com/users/stevetorr/events{/privacy}",
"received_events_url": "https://api.github.com/users/stevetorr/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @stevetorr!"
] | 2020-06-23T00:30:58
| 2020-06-23T01:25:56
|
2020-06-23T01:25:55Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
The QM9 database uses a special format for scientific notation in their .xyz files, wherein e.g. `3.14e-6` would be represented `3.14*^-6`. This addresses issue #1875 (which was not a bug per se, but simply brushing up against unimplemented behavior).
- The XYZ file parser can now handle this formatting for coordinates.
* This means changing the regular expressions which detect coordinate lines, as well as a substitution from the '*^' notation to 'e' notation.
- This capability is unit tested.
## Checklist
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [N/A] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [N/A] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [X] Tests have been added for any new functionality or bug fixes.
- [X] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1876/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1876/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1876",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1876",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1876.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1876.patch",
"merged_at": "2020-06-23T01:25:55Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1877
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1877/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1877/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1877/events
|
https://github.com/materialsproject/pymatgen/pull/1877
| 643,713,779
|
MDExOlB1bGxSZXF1ZXN0NDM4NDY3NzA5
| 1,877
|
change setup.py
|
{
"login": "JaGeo",
"id": 22094846,
"node_id": "MDQ6VXNlcjIyMDk0ODQ2",
"avatar_url": "https://avatars.githubusercontent.com/u/22094846?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/JaGeo",
"html_url": "https://github.com/JaGeo",
"followers_url": "https://api.github.com/users/JaGeo/followers",
"following_url": "https://api.github.com/users/JaGeo/following{/other_user}",
"gists_url": "https://api.github.com/users/JaGeo/gists{/gist_id}",
"starred_url": "https://api.github.com/users/JaGeo/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/JaGeo/subscriptions",
"organizations_url": "https://api.github.com/users/JaGeo/orgs",
"repos_url": "https://api.github.com/users/JaGeo/repos",
"events_url": "https://api.github.com/users/JaGeo/events{/privacy}",
"received_events_url": "https://api.github.com/users/JaGeo/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks -- the only way to really test this would be a test that does a pip install + then runs the test suite, which is perhaps not a bad idea. The other thing is that we could change the setup.py to just include all .json/.yaml it finds, which might also be sensible (rather than the current method which just includes those in specific submodules).",
"Both are fine with me. It's just that I forgot to include the yaml file in the setup.py the last time I added one... ",
"Let me know what you decide (I am mainly making these changes to finally include the Lobster workflow in atomate)",
"I've merged this now to get it working, I think we should do both: modify the setup.py to include all `.json` and `.yaml`, and run the test suite on pip installs, I'll add it to the list of things to do :) If you want to modify the setup.py in a future PR, that would also be welcome."
] | 2020-06-23T10:12:13
| 2020-06-23T18:15:16
|
2020-06-23T18:14:04Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Hi,
I forget to change the setup.py to reflect the correct folder structure of the new lobster module. Currently, the basis functions are not found when pymatgen is installed... (Is it maybe possible to check for this within an automatic test in the future?)
Best,
JG
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1877/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1877/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1877",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1877",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1877.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1877.patch",
"merged_at": "2020-06-23T18:14:04Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1878
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1878/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1878/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1878/events
|
https://github.com/materialsproject/pymatgen/issues/1878
| 643,898,804
|
MDU6SXNzdWU2NDM4OTg4MDQ=
| 1,878
|
ValueError: No POTCAR for C with functional PBE found
|
{
"login": "Jaydu1996",
"id": 66191420,
"node_id": "MDQ6VXNlcjY2MTkxNDIw",
"avatar_url": "https://avatars.githubusercontent.com/u/66191420?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/Jaydu1996",
"html_url": "https://github.com/Jaydu1996",
"followers_url": "https://api.github.com/users/Jaydu1996/followers",
"following_url": "https://api.github.com/users/Jaydu1996/following{/other_user}",
"gists_url": "https://api.github.com/users/Jaydu1996/gists{/gist_id}",
"starred_url": "https://api.github.com/users/Jaydu1996/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/Jaydu1996/subscriptions",
"organizations_url": "https://api.github.com/users/Jaydu1996/orgs",
"repos_url": "https://api.github.com/users/Jaydu1996/repos",
"events_url": "https://api.github.com/users/Jaydu1996/events{/privacy}",
"received_events_url": "https://api.github.com/users/Jaydu1996/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @Jaydu1996, what does your directory structure look like? Your `pseudopotentials` folder should have a `POT_GGA_PAW_PBE_52` folder inside it, and that has the POTCARs gzipped, e.g. `POTCAR.Ac.gz`",
"The directory structure is below:\r\n```\r\n[user@master POT_GGA_PAW_PBE_52]$ pwd\r\n/home/user/pseudopotentials/POT_GGA_PAW_PBE_52\r\n[user@master POT_GGA_PAW_PBE_52]$ ls\r\nPOTCAR.C.gz\r\n[user@master POT_GGA_PAW_PBE_52]$\r\n```\r\nThe test code is according to the [#1813 ](https://github.com/materialsproject/pymatgen/issues/1813) and the directory stucture is set according to the [website](https://matsci.org/t/hash-failure-for-potcar/3461)",
"Hm, perhaps the instructions are out of date, try:\r\n\r\n```\r\nstatic = MPStaticSet(structure, user_potcar_functional='PBE_52')\r\n```\r\n\r\nIf this works, I think it may be an error [here](https://github.com/materialsproject/pymatgen/blob/c65a6a32f35b345be1a76f6f9bad6f94e1e2cae7/pymatgen/io/vasp/sets.py#L411) with a missing \"PMG\" prefix.",
"Hi @mkhorton, I have a try, but it doesn't work. But do you have the experience of using atomate. If I use 'rlaunch' to run the job, it will appear the error `ValueError: No POTCAR for C with functional PBE found.`. And if I use 'qlaunch' to run the job, it will appear the error `ModuleNotFoundError: No module named 'atomate'`, and this error is described [here](https://stackoverflow.com/questions/62410345/modulenotfounderror-no-module-named-atomate-but-i-have-install-it) in detail. But I don't know why. \r\n```\r\n>>> import pymatgen as mg\r\n>>> structure = mg.Structure.from_file('POSCAR')\r\n>>> structure.composition\r\nComp: C2\r\n>>> from pymatgen.io.vasp.sets import MPStaticSet\r\n>>> static = MPStaticSet(structure, user_potcar_functional='PBE_52')\r\n/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py:437: BadInputSetWarning: Overriding the POTCAR functional is generally not recommended as it significantly affect the results of calculations and compatibility with other calculations done with the same input set. Note that some POTCAR symbols specified in the configuration file may not be available in the selected functional.\r\n BadInputSetWarning,\r\n>>> static.write_input(output_dir='crgete')\r\nTraceback (most recent call last):\r\n File \"<stdin>\", line 1, in <module>\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py\", line 722, in write_input\r\n zip_output=zip_output\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py\", line 212, in write_input\r\n vinput = self.get_vasp_input()\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py\", line 174, in get_vasp_input\r\n potcar=self.potcar,\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py\", line 129, in potcar\r\n potcar = Potcar(self.potcar_symbols, functional=self.potcar_functional)\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py\", line 2213, in __init__\r\n self.set_symbols(symbols, functional, sym_potcar_map)\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py\", line 2320, in set_symbols\r\n p = PotcarSingle.from_symbol_and_functional(el, functional)\r\n File \"/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py\", line 1939, in from_symbol_and_functional\r\n \"are using newer psps from VASP.\" % (symbol, functional)\r\nValueError: No POTCAR for C with functional PBE_52 found. Please set the PMG_VASP_PSP_DIR environment in .pmgrc.yaml, or you may need to set PMG_DEFAULT_FUNCTIONAL to PBE_52 or PBE_54 if you are using newer psps from VASP.\r\n>>> \r\n```",
"`ModuleNotFoundError: No module named 'atomate'`\r\n\r\nThis suggests an issue with your Python installation, and not related to pymatgen. Try creating a dedicated conda environment, and make sure this environment is _active_ when you try to run qlaunch. http://matsci.org/atomate is a place for atomate questions",
"Ah, I see you posted in the atomate forum too, please don't cross-post the same question in multiple places, it just creates work for the moderators, thanks!"
] | 2020-06-23T14:39:05
| 2020-06-25T19:14:25
|
2020-06-25T19:13:29Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
The pymatgen can't recognize the POTCAR(PBE_52), although I think I have set the correct pmgrc.yaml file.
The bug is
```
Traceback (most recent call last):
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/fireworks/core/rocket.py", line 262, in run
m_action = t.run_task(my_spec)
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/atomate/vasp/firetasks/write_inputs.py", line 85, in run_task
vis.write_input(".", potcar_spec=potcar_spec)
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py", line 722, in write_input
zip_output=zip_output
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py", line 212, in write_input
vinput = self.get_vasp_input()
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py", line 174, in get_vasp_input
potcar=self.potcar,
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/sets.py", line 129, in potcar
potcar = Potcar(self.potcar_symbols, functional=self.potcar_functional)
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py", line 2213, in __init__
self.set_symbols(symbols, functional, sym_potcar_map)
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py", line 2320, in set_symbols
p = PotcarSingle.from_symbol_and_functional(el, functional)
File "/home/user/app/anaconda3/envs/firework/lib/python3.7/site-packages/pymatgen/io/vasp/inputs.py", line 1939, in from_symbol_and_functional
"are using newer psps from VASP." % (symbol, functional)
ValueError: No POTCAR for C with functional PBE found. Please set the PMG_VASP_PSP_DIR environment in .pmgrc.yaml, or you may need to set PMG_DEFAULT_FUNCTIONAL to PBE_52 or PBE_54 if you are using newer psps from VASP.
```
And my .pmgrc.ymal 's content is below:
```
PMG_DEFAULT_FUNCTIONAL: PBE_52
PMG_MAPI_KEY: ****************
PMG_VASP_PSR_DIR: /home/user/pseudopotentials
```
what's the problem with it? How can I solve it
Provide any example files that are needed to reproduce the error,
especially if the bug pertains to parsing of a file.
```
In [2]: import pymatgen as mg
In [3]: structure = mg.Structure.from_file(‘POSCAR’)
In [4]: structure.composition
Out[4]: Comp: C2
In [5]: from pymatgen.io.vasp.sets import MPStaticSet
In [7]: static = MPStaticSet(structure)
In [8]: static.write_input(output_dir=‘crgete’)
```
**Desktop (please complete the following information):**
- OS: [Linux]
- Version [2020.4.29]
- python=3.7.7
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1878/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1878/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1879
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1879/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1879/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1879/events
|
https://github.com/materialsproject/pymatgen/pull/1879
| 644,422,331
|
MDExOlB1bGxSZXF1ZXN0NDM5MDQ2Mzgw
| 1,879
|
Boltztrap2 spin
|
{
"login": "fraricci",
"id": 17427517,
"node_id": "MDQ6VXNlcjE3NDI3NTE3",
"avatar_url": "https://avatars.githubusercontent.com/u/17427517?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/fraricci",
"html_url": "https://github.com/fraricci",
"followers_url": "https://api.github.com/users/fraricci/followers",
"following_url": "https://api.github.com/users/fraricci/following{/other_user}",
"gists_url": "https://api.github.com/users/fraricci/gists{/gist_id}",
"starred_url": "https://api.github.com/users/fraricci/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/fraricci/subscriptions",
"organizations_url": "https://api.github.com/users/fraricci/orgs",
"repos_url": "https://api.github.com/users/fraricci/repos",
"events_url": "https://api.github.com/users/fraricci/events{/privacy}",
"received_events_url": "https://api.github.com/users/fraricci/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2020-06-24T08:26:57
| 2020-06-25T13:41:02
|
2020-06-25T13:40:57Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Here an upgrade of the boltztrap2 interface.
Major points are:
- transport properties, dos, and bands can be computed for spin polarized DFT computations
- PMGLoader should be used to load a vasprun/bandstructure pymatgen object, instead of the two previous loaders.
- load and save methods of BztInterpolator and BztTransportProperties allows to store/read in/from file the data they contain
- it includes the fix to the typo in the plotter raised in issue #1815
- bands can be interpolated along a custom path
- carrier concentration for different doping levels and temperatures is now available
I did some tests. But those of you that use it, please let me know any issue. Thanks.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1879/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1879/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1879",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1879",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1879.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1879.patch",
"merged_at": "2020-06-25T13:40:57Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1880
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1880/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1880/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1880/events
|
https://github.com/materialsproject/pymatgen/pull/1880
| 644,711,684
|
MDExOlB1bGxSZXF1ZXN0NDM5Mjg2NDc0
| 1,880
|
Fix suggest_larger_supercells bug
|
{
"login": "kavanase",
"id": 51478689,
"node_id": "MDQ6VXNlcjUxNDc4Njg5",
"avatar_url": "https://avatars.githubusercontent.com/u/51478689?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/kavanase",
"html_url": "https://github.com/kavanase",
"followers_url": "https://api.github.com/users/kavanase/followers",
"following_url": "https://api.github.com/users/kavanase/following{/other_user}",
"gists_url": "https://api.github.com/users/kavanase/gists{/gist_id}",
"starred_url": "https://api.github.com/users/kavanase/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/kavanase/subscriptions",
"organizations_url": "https://api.github.com/users/kavanase/orgs",
"repos_url": "https://api.github.com/users/kavanase/repos",
"events_url": "https://api.github.com/users/kavanase/events{/privacy}",
"received_events_url": "https://api.github.com/users/kavanase/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks for the bug fix @kavanase, my apologies for letting this sit for a while. I've merged this in now as-is since it's a fairly minimal change, but for future bug fixes including a minimal extra test that demonstrates the buggy behavior is always appreciated.\r\n\r\nPlease make sure to include your details in [this form](https://forms.gle/JnisFb38QDR8QTFTA) so that you can be credited appropriately in the [pymatgen documentation](https://pymatgen.org/team.html) too.",
"Great, thanks @mkhorton !!\r\nI'll make sure to include extra tests for future bug fixes!\r\n\r\nI've added my details to the developer attribution form now."
] | 2020-06-24T15:32:46
| 2020-10-04T14:45:35
|
2020-10-03T03:07:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
There's a small bug in the `DefectPhaseDiagram.suggest_larger_supercells()` method (line 390 in `pymatgen/analysis/defects/thermodynamics.py`, where the current code is:
```python
for tl, chgset in self.transition_level_map.items():
```
which gives the wrong variables of Defect-Name, {Transition-Level: Charge-set}. It should be:
```python
for tl, chgset in self.transition_level_map[def_type].items():
```
(it's within the `for def_type in self.defect_types:` loop), which gives the correct variables of which gives the wrong variables of Transition-Level, Charge-set.
See below for a quick demonstration of the bug, and the result when fixed:
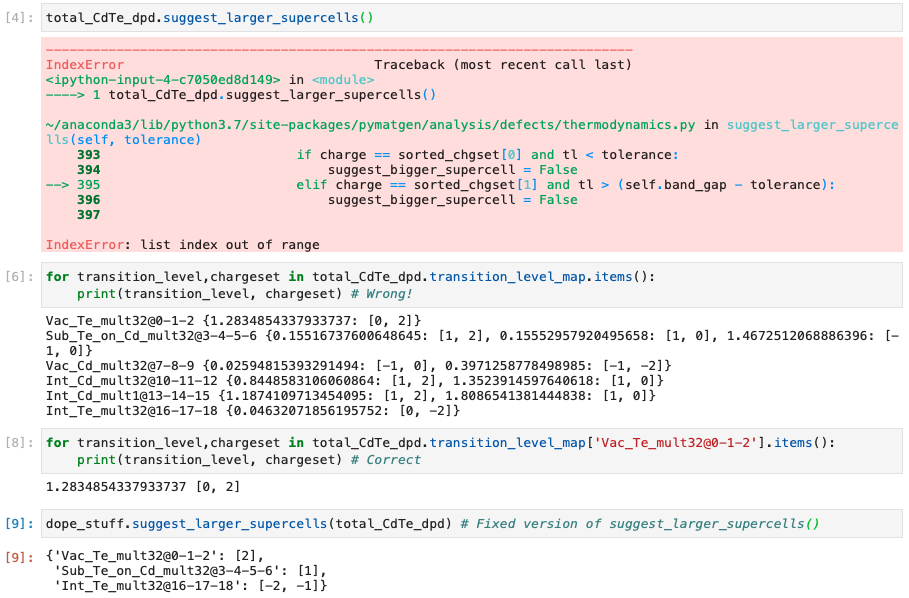
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1880/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1880/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1880",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1880",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1880.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1880.patch",
"merged_at": "2020-10-03T03:07:49Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1881
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1881/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1881/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1881/events
|
https://github.com/materialsproject/pymatgen/pull/1881
| 645,723,830
|
MDExOlB1bGxSZXF1ZXN0NDQwMTUyMzMw
| 1,881
|
[WIP] Add F* diagrams
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-06-25T17:11:34
| 2020-06-25T17:22:41
|
2020-06-25T17:22:41Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Added folder to pymatgen/analysis called Fstar
contains script for generating f* diagrams, an example notebook, and scattering factor files
## TODO (if any)
add additional examples to show off more features.
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1881/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1881/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1881",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1881",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1881.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1881.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1882
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1882/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1882/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1882/events
|
https://github.com/materialsproject/pymatgen/pull/1882
| 645,732,407
|
MDExOlB1bGxSZXF1ZXN0NDQwMTU5NDUw
| 1,882
|
[WIP] Add F* diagram generator
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-06-25T17:26:58
| 2020-06-25T17:43:09
|
2020-06-25T17:43:09Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
* generator code
* example
* scattering factors
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
* need more examples of features and workarounds
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [ ] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [ ] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [ ] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [ ] Tests have been added for any new functionality or bug fixes.
- [ ] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1882/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1882/timeline
| null | true
| true
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1882",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1882",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1882.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1882.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1883
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1883/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1883/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1883/events
|
https://github.com/materialsproject/pymatgen/pull/1883
| 645,743,124
|
MDExOlB1bGxSZXF1ZXN0NDQwMTY4MzAy
| 1,883
|
[WIP]Add F* diagram generator
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @jonathanjdenney, nice to see this PR. I noticed you were uploading these files manually. You may find [GitHub Desktop](https://desktop.github.com) useful to commit your changes locally.",
"Hi @mkhorton I will check that out. In the meantime I'm getting the error \"D100: Missing docstring in public module\" from the pydocstyle check and I'm not sure how to fix it. ",
"Every module file should have a docstring at the top to say what it does, for example see [here](https://github.com/materialsproject/pymatgen/blob/7327a260259e6a0fa32b719582b68720554a400a/pymatgen/core/structure.py#L5) at the top of structure.py.",
"To be more specific, I see your file has a module-level docstring, but it's *below* the import statements, so it's not at the top of the file. The error message contains this information: `pymatgen/analysis/Fstar/f_star_diagram_generator.py:1 at module level:` meaning, it found the error in `f_star_diagram_generator.py` and the :1 refers to line 1.",
"Ok, I can see the linters pass, which is good. This will also need a test or two before it can be merged. A good place to put this would be in analysis/fstar/tests/test_fstar.py (I think renaming the folder to `fstar` lowercase and the diagram to `fstar.py` instead of `fstar_diagram_generator.py` would be good).\r\n\r\nRegarding the example notebook, that's definitely useful, but would be better put into the [`matgenb` notebook repository](https://matgenb.materialsvirtuallab.org) after the functionality has been merged into pymatgen.",
"Hi @mkhorton , It's been a while since I have had time to work on this. I've added a test folder and made the name changes you suggested. What kind of testing needs done to get this merged? ",
"If you look inside the `pymatgen/analysis/tests` directory, you'll see tests for other modules which can give you an idea. Basically, any user-facing feature or function has to have a corresponding test that verifies it generates the output you expect.\r\n\r\nThe purpose of the tests is so that if a future update, e.g. to Python, to a library, or if someone comes and modifies your code (yourself or a future collaborator), they won't inadvertently either break the module or simply unknowingly change historical behavior that someone might be depending on for their own research."
] | 2020-06-25T17:46:38
| 2020-10-20T17:33:46
|
2020-10-20T17:33:46Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Script to generate f* diagrams
## TODO (if any)
If this is a work-in-progress, write something about what else needs
to be done
* add more examples
## Checklist
Work-in-progress pull requests are encouraged, but please put [WIP]
in the pull request title.
Before a pull request can be merged, the following items must be checked:
- [x] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [x] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [x] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [x] Tests have been added for any new functionality or bug fixes.
- [x] All existing tests pass.
Note that the CI system will run all the above checks. But it will be much more
efficient if you already fix most errors prior to submitting the PR. It is
highly recommended that you use the pre-commit hook provided in the pymatgen
repository. Simply `cp pre-commit .git/hooks` and a check will be run prior to
allowing commits.
|
{
"login": "jonathanjdenney",
"id": 48103676,
"node_id": "MDQ6VXNlcjQ4MTAzNjc2",
"avatar_url": "https://avatars.githubusercontent.com/u/48103676?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/jonathanjdenney",
"html_url": "https://github.com/jonathanjdenney",
"followers_url": "https://api.github.com/users/jonathanjdenney/followers",
"following_url": "https://api.github.com/users/jonathanjdenney/following{/other_user}",
"gists_url": "https://api.github.com/users/jonathanjdenney/gists{/gist_id}",
"starred_url": "https://api.github.com/users/jonathanjdenney/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/jonathanjdenney/subscriptions",
"organizations_url": "https://api.github.com/users/jonathanjdenney/orgs",
"repos_url": "https://api.github.com/users/jonathanjdenney/repos",
"events_url": "https://api.github.com/users/jonathanjdenney/events{/privacy}",
"received_events_url": "https://api.github.com/users/jonathanjdenney/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1883/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1883/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1883",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1883",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1883.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1883.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1884
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1884/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1884/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1884/events
|
https://github.com/materialsproject/pymatgen/issues/1884
| 646,559,289
|
MDU6SXNzdWU2NDY1NTkyODk=
| 1,884
|
Possible Potcar hash changes in new pymatgen versions
|
{
"login": "chc273",
"id": 22353204,
"node_id": "MDQ6VXNlcjIyMzUzMjA0",
"avatar_url": "https://avatars.githubusercontent.com/u/22353204?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/chc273",
"html_url": "https://github.com/chc273",
"followers_url": "https://api.github.com/users/chc273/followers",
"following_url": "https://api.github.com/users/chc273/following{/other_user}",
"gists_url": "https://api.github.com/users/chc273/gists{/gist_id}",
"starred_url": "https://api.github.com/users/chc273/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/chc273/subscriptions",
"organizations_url": "https://api.github.com/users/chc273/orgs",
"repos_url": "https://api.github.com/users/chc273/repos",
"events_url": "https://api.github.com/users/chc273/events{/privacy}",
"received_events_url": "https://api.github.com/users/chc273/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi Chi. I agree that it's unlikely it's a \"bad\" POTCAR -- in fact, I've recently changed this warning in master to be `UnknownPotcarWarning` instead, so as not to alarm people. I suspect the database of potcar hashes distributed with pymatgen is simply incomplete, we did use the latest potcars to generate them but perhaps VASP has released an update, I'll check.",
"If interested, here is the current list of [file hashes (checksums of the file itself)](https://github.com/materialsproject/pymatgen/blob/master/pymatgen/io/vasp/vasp_potcar_file_hashes.json) and [\"pymatgen hashes\"( hashes of the content of the potcar)](https://github.com/materialsproject/pymatgen/blob/master/pymatgen/io/vasp/vasp_potcar_pymatgen_hashes.json), so you could check if your Na_pv hash is already included in one of those, it could be a logic bug.\r\n",
"Thanks Matt. In fact, I have never updated my potcars. I assume that for PBE functional used in MP, the potcars should not changed right? maybe i am wrong.. It leads me to think that If somehow the potcar changes slightly, for example adding extra spaces, that will also change the hashes? I wonder if there are alternative ways to test the equivalencies of pseudo potentials..",
"Yes, that's why there are two hashes, the \"file hash\" (which will change even if a title or a space changes) and a \"pymatgen hash\" (which is meant to be more robust against small changes). VASP have released multiple versions of the same POTCARs sometimes, because of small typos.\r\n\r\nWe have the hashing system because some people have spent many CPU hours on incompatible calculations because they accidentally use the wrong POTCAR, so the hashing is meant to prevent that, however I think it's too strict right now.",
"Thanks Matt. Yes, hash can be too sensitive to very small changes and the files that you used to generate the hashes may be slightly different from what our group is using.\r\n\r\nIt would be nice to have the option to turn off the warnings if I am certain that the POTCAR I am using is the correct one, judging from some meta information (e.g., VASP PBE) and the potcar symbol (e.g., Na_pv).\r\n",
"The \"pymatgen hash\" is intended to be robust against small changes, could you share this hash for your Na_pv in this issue so we can diagnose?\r\n\r\n```\r\np = Potcar(['Na'])\r\np[0].file_hash # gives me `5a81584fb022b82fbb3658e6decf92de`\r\np[0].hash # the \"pymatgen hash\" gives me `1a89e79f7e21d99e8cf5788979f6a987`\r\n```\r\n\r\nNote that the meta information is _not_ always correct in the file, some pseudopotentials distributed by VASP have (unfortunately) sometimes had incorrect metadata, e.g. they had a release where the title string gave PAW instead of PAW_PBE.",
"I confirm that these two hashes are the same as yours for `Na`. The one problematic is `Na_pv` (MPRelaxSet uses this one).\r\n\r\n```\r\np_na_pv = Potcar(['Na_pv'])[0]\r\n```\r\nWill give me warning \r\n```BadPotcarWarning: POTCAR data with symbol Na_pv does not match any VASP POTCAR known to pymatgen. We advise verifying the integrity of your POTCAR files.\r\n BadPotcarWarning)\r\n```\r\nThe hashes are\r\n\r\n```\r\np_na_pv.hash # gives me 'c71d0ed99a871c91fccae9347860d8ba'\r\np_na_pv.file_hash # gives me 'c2bf3a84744e11107e8c18128fe55175'\r\n```\r\npymatgen version is `2020.4.2`. \r\n",
"Thanks! I'll follow-up.",
"Thanks Matt! ",
"> If interested, here is the current list of [file hashes (checksums of the file itself)](https://github.com/materialsproject/pymatgen/blob/master/pymatgen/io/vasp/vasp_potcar_file_hashes.json) and [\"pymatgen hashes\"( hashes of the content of the potcar)](https://github.com/materialsproject/pymatgen/blob/master/pymatgen/io/vasp/vasp_potcar_pymatgen_hashes.json), so you could check if your Na_pv hash is already included in one of those, it could be a logic bug.\r\n\r\nIn these two hash files, it seems they only cover `\"potpaw_PBE.54\", \"unvie_potpaw_PBE.54\", \"potpaw_PBE.52\", \"unvie_potpaw_PBE.52\"`. I think Chi used the oldest VASP PSP (`\"potpaw_PBE\" or \"POT_GGA_PAW_PBE\"`) and the MP did the same. \r\n\r\n According to the VASP release notes, this oldest VASP PSP is outdated. The developers recommend PBE54 (`potpaw_PBE.54`)\r\nThe original release of PBE52 and PBE54 are now called \"unvie PBE52\" and \"unvie PBE54\", respectively. (@chc273 I think you also used these old ones.) In VASP portal, they have the lastest PBE52 and PBE54. ",
"Thanks Zhenbin @zbwang In your experience, will the calculation results change if I switch from the older PBE to PBE54? What are the differences? My concern is about the result consistency with MP if I update to the latest PBE. It may affect things like ehull calculations etc.\r\n\r\n",
"Sorry, I am not sure. But based on these two hash files, it seems the MP is going to discard the oldest VASP PSP but use the newest ones. Matt may know how MP is going to deal with this. (some potcar correction?)",
"@zbwang the hash database does include the oldest VASP pseudopotentials. Regarding the issue, no POTCAR correction is planned, we are using the oldest pseudopotentials for MP compatibility. We are switching to PBE54 for newer calculations with different functionals, but our standard PBE calcs are staying on the old pseudos for now.",
"ah. my bad, I just realized that the hash keys for the oldest PSP are at the bottom of the files you shared. Then, it is strange that each time when generating VASP inputs, pymatgen complains the warnings. ",
"`pymatgen/io/vasp/inputs.py:1856: UnknownPotcarWarning: POTCAR with symbol Pr has metadata that does not match any VASP POTCAR known to pymatgen. The data in this POTCAR is known to match the following functionals: ['PBE_52', 'PBE_54', 'unvie_PBE_52']`\r\n.\r\nWhen post-processing results from a **TbO2** relaxation, I get the same UnknownPotcarWarning and pymatgen interprets the Tb ions as Ce when printing output structure data, plotting DOS, etc.\r\n\r\nHow should this behaviour be handled since the post-processing of VASP results is being affected in the case of TbO2.\r\n\r\nCan results generated by pymatgen modules on raw VASP output data be interpreted as the same when this warning is not present? Does this affect the integrity of the results in a general sense?\r\n",
"@darnoceloc I have the same problem as you.Have you solved this problem?\r\n`/pymatgen/io/vasp/inputs.py:1820: UnknownPotcarWarning: POTCAR with symbol Zr_sv has metadata that does not match any VASP POTCAR known to pymatgen. The data in this POTCAR is known to match the following functionals: ['unvie_PBE_52', 'PBE_54', 'PBE_52']`",
"> When post-processing results from a TbO2 relaxation, I get the same UnknownPotcarWarning and pymatgen interprets the Tb ions as Ce when printing output structure data, plotting DOS, etc.\r\n\r\nMore information would be required to diagnose this issue; this sounds unusual and potentially troublesome.\r\n\r\nThe warning itself is not necessarily a problem (it is just a warning). Mis-parsing one ion as another ion, however, would be a significant bug if it could be reproduced and was not an error in usage. "
] | 2020-06-26T23:28:58
| 2023-08-13T16:33:42
|
2023-08-13T16:33:42Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Describe the bug**
I was trying to generate vasp input sets for a Na-containing system using io.vasp.sets.MPRelaxSet. However, it seems that the `Na_pv` potcar failed the potcar check.
I then go to my vasp potcar directory, `VASP_PSP/POT_GGA_PAW_PBE`, and run the following codes
```
from pymatgen.io.vasp.inputs import PotcarSingle
from glob import glob
for pot_file in glob('*.gz'):
p = PotcarSingle.from_file(i)
```
It gives me a long list of warnings like the following.
```
pymatgen/pymatgen/io/vasp/inputs.py:1853: BadPotcarWarning: POTCAR data with symbol Na_pv does not match any VASP POTCAR known to pymatgen. We advise verifying the integrity of your POTCAR files.
BadPotcarWarning)
```
If I change pymatgen version to an earlier version, for example, `pymatgen-2019.5.8`. There is no such problem. I wonder if this is expected. I cannot see why the potcar is wrong, since it is from VASP distribution..
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1884/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1884/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1885
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1885/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1885/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1885/events
|
https://github.com/materialsproject/pymatgen/pull/1885
| 646,714,976
|
MDExOlB1bGxSZXF1ZXN0NDQwOTM1MDUy
| 1,885
|
parse scientific notation in OUTCAR (possibly without spaces in betwe…
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Closing for now, since two features got intermingled in this PR"
] | 2020-06-27T16:47:53
| 2020-06-27T17:18:34
|
2020-06-27T17:18:34Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR is very similar to #1836 , except it now supports the dielectric tensor section of the OUTCAR as well (instead of just plasma frequencies)
The unit test from #1836 should also cover this one
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1885/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1885/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1885",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1885",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1885.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1885.patch",
"merged_at": null
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1886
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1886/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1886/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1886/events
|
https://github.com/materialsproject/pymatgen/pull/1886
| 646,720,208
|
MDExOlB1bGxSZXF1ZXN0NDQwOTM4Nzg1
| 1,886
|
two changes: parse scientific notation dielectric data, update cost information
|
{
"login": "computron",
"id": 986759,
"node_id": "MDQ6VXNlcjk4Njc1OQ==",
"avatar_url": "https://avatars.githubusercontent.com/u/986759?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/computron",
"html_url": "https://github.com/computron",
"followers_url": "https://api.github.com/users/computron/followers",
"following_url": "https://api.github.com/users/computron/following{/other_user}",
"gists_url": "https://api.github.com/users/computron/gists{/gist_id}",
"starred_url": "https://api.github.com/users/computron/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/computron/subscriptions",
"organizations_url": "https://api.github.com/users/computron/orgs",
"repos_url": "https://api.github.com/users/computron/repos",
"events_url": "https://api.github.com/users/computron/events{/privacy}",
"received_events_url": "https://api.github.com/users/computron/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks @computron :) "
] | 2020-06-27T17:22:42
| 2020-06-27T22:04:02
|
2020-06-27T22:03:25Z
|
MEMBER
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR includes two different changes (poor style, I know - but the Github UI makes it difficult to separate them into different PRs after the fact):
### Change 1:
This update for scientific notation is very similar to #1836 , except it now supports the dielectric tensor section of the OUTCAR as well (instead of just plasma frequencies)
The unit test from #1836 should also cover this one
### Change 2:
Added cost data to CSV file for He,H,Ar,Ne,Kr,Tc; remove duplicate O row.
There is no new code here, just a data file update.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1886/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1886/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1886",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1886",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1886.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1886.patch",
"merged_at": "2020-06-27T22:03:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1887
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1887/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1887/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1887/events
|
https://github.com/materialsproject/pymatgen/pull/1887
| 647,187,544
|
MDExOlB1bGxSZXF1ZXN0NDQxMjU2ODQ2
| 1,887
|
Fix regular expression of read_onsite_density_matrices
|
{
"login": "yuuukuma",
"id": 4986887,
"node_id": "MDQ6VXNlcjQ5ODY4ODc=",
"avatar_url": "https://avatars.githubusercontent.com/u/4986887?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yuuukuma",
"html_url": "https://github.com/yuuukuma",
"followers_url": "https://api.github.com/users/yuuukuma/followers",
"following_url": "https://api.github.com/users/yuuukuma/following{/other_user}",
"gists_url": "https://api.github.com/users/yuuukuma/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yuuukuma/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yuuukuma/subscriptions",
"organizations_url": "https://api.github.com/users/yuuukuma/orgs",
"repos_url": "https://api.github.com/users/yuuukuma/repos",
"events_url": "https://api.github.com/users/yuuukuma/events{/privacy}",
"received_events_url": "https://api.github.com/users/yuuukuma/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks.",
"This had a linter failure, nothing major but the other linters didn't have chance to run.",
"I know. But the linter failure is in the test file."
] | 2020-06-29T07:35:27
| 2020-06-29T20:12:20
|
2020-06-29T20:05:22Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Include a summary of major changes in bullet points:
* Fixed regular expression to parse density matrix
At Line 1476 -- 1490 in OUTCAR_merged_numbers2 file in test_files
-------------------------------------------------------
spin component 1
6.1320 -7.9985 9.9720 5.1366 -1.5481
-7.9985 11.2215-13.7153 -6.9893 2.1083 **(Parse this line was okay.)**
9.9720-13.7153 17.1305 8.6130 -2.6035 **(Parse this line was failed.)**
5.1366 -6.9893 8.6130 4.6441 -1.3773
-1.5481 2.1083 -2.6035 -1.3773 0.5186
spin component 2
0.2072 0.0820 -0.0441 0.0155 0.0005
0.0820 0.1381 0.0032 0.0204 -0.0103
-0.0441 0.0032 0.1180 -0.0734 0.0235
0.0155 0.0204 -0.0734 0.1874 -0.0333
0.0005 -0.0103 0.0235 -0.0333 0.0992
-------------------------------------------------------
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1887/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1887/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1887",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1887",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1887.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1887.patch",
"merged_at": "2020-06-29T20:05:22Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1888
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1888/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1888/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1888/events
|
https://github.com/materialsproject/pymatgen/pull/1888
| 647,584,802
|
MDExOlB1bGxSZXF1ZXN0NDQxNTgzNjcx
| 1,888
|
selling_vector property
|
{
"login": "bwjustus",
"id": 45983292,
"node_id": "MDQ6VXNlcjQ1OTgzMjky",
"avatar_url": "https://avatars.githubusercontent.com/u/45983292?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/bwjustus",
"html_url": "https://github.com/bwjustus",
"followers_url": "https://api.github.com/users/bwjustus/followers",
"following_url": "https://api.github.com/users/bwjustus/following{/other_user}",
"gists_url": "https://api.github.com/users/bwjustus/gists{/gist_id}",
"starred_url": "https://api.github.com/users/bwjustus/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/bwjustus/subscriptions",
"organizations_url": "https://api.github.com/users/bwjustus/orgs",
"repos_url": "https://api.github.com/users/bwjustus/repos",
"events_url": "https://api.github.com/users/bwjustus/events{/privacy}",
"received_events_url": "https://api.github.com/users/bwjustus/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"Hi @bwjustus, nice to see this :-)\r\n\r\nIn terms of implementation, would a method to return all 24 Selling lattices be sensible?\r\n\r\nUltimately, the benefit of the Selling reduction is to calculate a distance between lattices `A` and `B`, which I believe takes the form of `min([dist(any_selling_lattice_A, a_selling_lattice_B) for a_selling_lattice_B in all_selling_lattices_for_B])`, and for that we'd need an implementation of the `dist` function too (see section 4 in https://journals.iucr.org/a/issues/2019/03/00/ae5061/ae5061.pdf).\r\n\r\nI haven't thought too deeply about implementation, so let me know your thoughts.",
"Hi @mkhorton, I haven't actually seen this paper yet, I had only seen two others on the same topic, but I will try to make an implementation of the distance function. I can also see if returning all 24 lattices would be reasonable in terms of time cost as compared to Niggli reduction.",
"I did my best to follow the paper and write an implementation for distance using the virtual Cartesian Point method they described. I had a bit of trouble following it so I'm not sure if it's functioning exactly how it should, but it seems correct. The main thing I noticed is that computing all 24 reflections is pretty costly considering all 24 are computed for each of the six virtual cartesian points as well as the original point, so there ends up being 168 vectors calculated. Would it be helpful for me to add the code to this pull request so you can take a look? Currently it returns the selling vector of the reflection that gives the minimum distance.",
"Thanks, yes adding the code would be helpful. It might be possible to vectorize this code, and calculate all 24 reflections in one with numpy, which may make it less costly. As for verification, I'll see if I can find a way to independently verify.",
"Alright, I just pushed it. Most of the body of the function is just the relevant matrices, so let me know if there's a better way to represent these so that they don't take up so much space. I'm not sure I'm familiar with vectorizing code, but if there's a certain NumPy function that would make this process faster, let me know what it is and I can try to change the implementation.",
"From a numpy perspective, the basic principle is removing for loops where possible and trying to incorporate multiple operations into a single operation, for example there's `numpy.linalg.multi_dot` which might help, functions like `tensordot` (across an appropriate axis) and `einsum` can be helpful.\r\n\r\nIt's not clear the optimization will be necessary however, how long does this take typically? (If you're in Jupyter, you can use `%timeit` to get some statistics).",
"Just calculating the selling vectors takes about 60-90 microseconds, while the whole distance function takes about 1.5-2 milliseconds. I'm familiar with einsum but I'll check out the others too to see if I can speed it up. I'm not sure if this is an issue we want to consider yet, but I was also wondering how to convert a reduced selling vector back to lattice vectors so I can return a Selling-reduced lattice. Shyam and I had looked at [this paper](https://journals.iucr.org/a/issues/2020/01/00/ae5074/) but the basis for E3x3 given there in Table 1 only has components in the x-direction, which doesn't seem to make sense for creating a 3D cell. Let me know if you have any ideas on how to do this.",
"Another possible issue that I noticed is that all 6 virtual cartesian points that I calculated are all the same, and are all just the original, unreduced selling vector multipled by -1. Maybe I misunderstood the wording of the paper, so let me know if it seems like there's something that I'm misinterpreting. Also, I've been able to eliminate some of the for loops by using multi_dot. It doesn't seem like it's changing the runtime in any significant way right now but I'll keep getting rid of for loops where I can.",
"Ah, yes, that sounds suspect. In terms of profiling, there are line profilers available which can tell you where most of the time is being spent. Two milliseconds is in general very good, but could still be prohibitive if we wanted to do a search across the entire MP database.\r\n\r\nDrop me an email at mkhorton@lbl.gov and I'll see if I can put you in touch with someone who could help explain the VCPs.",
"I tried replacing all the for loops with just a few lines that accomplish the same thing, and the time did not really change. I tried some code profiling and it seems as though more than half the time comes from using numpy.linalg.norm to find the distances between all 168 vcp reflections and the point representing the other lattice. I'm not sure if there's a way around this because it is part of the algorithm, but hopefully if we do get in touch with Nicholas Sauter he might have some advice.",
"Hi @bwjustus, just following up on that status of this PR, what are the next steps? ",
"The distance function is implemented, so Shyam has been helping me work on finding different ways to test it and try to understand the behavior a bit better. It seems to behave as expected (more dissimilar lattices have a greater selling distance, the distance between two identical lattices is 0, etc.) but I’m still working on writing a lot of different tests to try to get a better feel for what a selling distance value actually means so we can use it effectively. I can push the new selling distance function if you’d like to take a look. I won’t have access to my computer again until Sunday afternoon but I can push it then.",
"Sounds great! Just wanted to check in that there wasn't anything that was stuck, would be happy to take a look when it's ready.",
"Sorry, took me a long time to get back to this. The one thing to modify is to make it so the selling vector is a 1-D array rather than 2d. Right now, we reshape it to (1,6). Let's keep it as (6,) . Can you update the @bwjustus and then the PR will be ready to merge.",
"Sure, I'll take care of that today.",
"Hi @bwjustus, nice PR! Can I clarify whether the test cases have been independently verified? e.g. did you use a lattice that was given in a paper where the Selling vectors/distances were known?",
"Hi @mkhorton! For reference, this is the main paper I referenced when working on this function: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6492488/. They did not give any test cases/examples for selling distance, but in the 2nd section of that paper where it discusses S6 space, they have one example of a calculated selling vector for an orthorhombic structure. The parameters for the structure were (10, 12, 20, 90, 90, 90) and the selling vector was [0, 0, 0, -100, -144, -400]. I didn't use this exact example, but in general what I found was that for orthorhombic lattices, the selling vector generally has the form [0, 0, 0, -a^2, -b^2, -c^2]. One of the tests I wrote has this exact same form, just different lengths. The other two selling vector tests I wrote are for a cubic structure and a tetragonal structure, which follow the same rules. I did not have any reference for the selling distances, so those tests were used to verify the assumptions that the selling distance between two identical structures is 0, and that as they become more dissimilar, the distance increases. Additionally, I also did some tests where I incrementally increased/decreased lattice lengths and angles to make sure the distance also increased/decreased monotonically. I didn't include these tests in the PR, but if you'd like me to add them in I can do that. If there's anything else that needs to be added / changed let me know and I'll take care of it!",
"Ok, that's perfect, thanks! I just wanted some more context on how the code was verified. No need to add additional tests for the monotonic case since I imagine that might be slow, but perhaps add a code comment to the test file just to mention that that verification was performed.",
"Sounds good. Should I say that verification was performed in the sense that I referenced it against that paper?",
"Sure, or you can link the comment you posted above. I just want the additional context to be easy to find if someone else is examining this code at a later date.",
"Alright that makes sense, I'll add that in right now.",
"Merged, thanks @bwjustus!"
] | 2020-06-29T18:40:25
| 2021-04-14T20:37:36
|
2021-04-14T20:37:27Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Adding a property in pymatgen/core/lattice.py to calculate the (1, 6) array containing the six selling scalars.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1888/reactions",
"total_count": 1,
"+1": 1,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1888/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1888",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1888",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1888.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1888.patch",
"merged_at": "2021-04-14T20:37:27Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1889
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1889/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1889/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1889/events
|
https://github.com/materialsproject/pymatgen/issues/1889
| 647,609,062
|
MDU6SXNzdWU2NDc2MDkwNjI=
| 1,889
|
Jupyter notebook bug.
|
{
"login": "keduemml",
"id": 67605140,
"node_id": "MDQ6VXNlcjY3NjA1MTQw",
"avatar_url": "https://avatars.githubusercontent.com/u/67605140?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/keduemml",
"html_url": "https://github.com/keduemml",
"followers_url": "https://api.github.com/users/keduemml/followers",
"following_url": "https://api.github.com/users/keduemml/following{/other_user}",
"gists_url": "https://api.github.com/users/keduemml/gists{/gist_id}",
"starred_url": "https://api.github.com/users/keduemml/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/keduemml/subscriptions",
"organizations_url": "https://api.github.com/users/keduemml/orgs",
"repos_url": "https://api.github.com/users/keduemml/repos",
"events_url": "https://api.github.com/users/keduemml/events{/privacy}",
"received_events_url": "https://api.github.com/users/keduemml/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Which vasprun.xml are you using?",
"The vasprun.xml file that is generated from running VASP. Im not sure how else to explain it, are there different vasprun.xml files.",
"The reason I ask is that usually this kind of error is the result of a mal-formed vasprun.xml file, e.g. one in which VASP didn't complete writing the full file. You could try again with one of the example vasprun.xml files in our [test_files folder](https://github.com/materialsproject/pymatgen/tree/master/test_files), to diagnose whether the issue is with the specific file you're using or not.\r\n\r\nIf this isn't a bug, a good place to ask questions is in our user forum at matsci.org/pymatgen.",
"I will check with the forum page, I was unaware that there was one. I have compared my vasprun.xml file to the one in the folder you linked and there are significant differences mine ends much earlier and explains why I was getting the error I was. Which mean this isn't a bug in pymatgen.\r\nThank you for your help mkhorton.",
"No problem, glad it was helpful. And yes, we should advertise the forum more loudly on our documentation page, I'll have to make that change."
] | 2020-06-29T19:25:00
| 2020-06-29T19:47:03
|
2020-06-29T19:47:02Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
I am going through the Jupiter notebook tutorials and am trying to reproduce the 2013-01-01-Bandstructure of NiO tutorial. The issue I am running into is with the run = BSVasp("vasprun.xml", parse_projected_eigen=True) command. The error is ParseError: no element found: line 486, column 0.
That is the last line in the vasprun.xml file and when I try the same line of code for a different vasprun.xml file it always has the same error on the last line in the vasprun.xml file.
I am running pymatgen 2020.4.2 on windows.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1889/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1889/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1890
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1890/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1890/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1890/events
|
https://github.com/materialsproject/pymatgen/issues/1890
| 648,025,298
|
MDU6SXNzdWU2NDgwMjUyOTg=
| 1,890
|
Error of Critic2Caller
|
{
"login": "yuuukuma",
"id": 4986887,
"node_id": "MDQ6VXNlcjQ5ODY4ODc=",
"avatar_url": "https://avatars.githubusercontent.com/u/4986887?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yuuukuma",
"html_url": "https://github.com/yuuukuma",
"followers_url": "https://api.github.com/users/yuuukuma/followers",
"following_url": "https://api.github.com/users/yuuukuma/following{/other_user}",
"gists_url": "https://api.github.com/users/yuuukuma/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yuuukuma/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yuuukuma/subscriptions",
"organizations_url": "https://api.github.com/users/yuuukuma/orgs",
"repos_url": "https://api.github.com/users/yuuukuma/repos",
"events_url": "https://api.github.com/users/yuuukuma/events{/privacy}",
"received_events_url": "https://api.github.com/users/yuuukuma/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Hi @yuuukuma, you need to compile a newer version of Critic2, the zpsp version isn't in older versions of the code.",
"Dear @mkhorton,\r\nthanks for your reply. \r\nI can only use the stable version of critic2 due to the compiler issue, so try to use the older version of Pymatgen.",
"Ok, you can also check with the critic2 developer to see if you can get a newer compilation to work, it usually works quite well.",
"Dear @mkhorton \r\n\r\nThe latest version of critic2 downloaded from the GitHub page works quit well with the latest Pymatgen.\r\nHowever, I think there is a bug in Critic2Caller\r\n\r\nPlease see line 300 in critic2_caller.py\r\n\"symbol\" should be \"element\"; otherwise, it does not work for e.g. \"Ca_pv\" PAW potential.\r\n\r\n```\r\n if not zpsp:\r\n\r\n potcar_path = _get_filepath(\r\n \"POTCAR\",\r\n \"Could not find POTCAR, will not be able to calculate charge transfer.\",\r\n )\r\n\r\n if potcar_path:\r\n potcar = Potcar.from_file(potcar_path)\r\n zpsp = {p.symbol: p.zval for p in potcar}\r\n```",
"Yes, I agree, that looks like a bug. Would you be willing to submit a PR to fix?",
"Okay, I will submit a PR.\r\nDo you think I need to prepare a unittest for this?",
"No, I think this is fine for now, thanks."
] | 2020-06-30T09:36:04
| 2020-07-02T17:40:15
|
2020-07-02T17:40:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
[test.zip](https://github.com/materialsproject/pymatgen/files/4850861/test.zip)
**Describe the bug**
When using the critic2 with Critic2Caller, the result shows
```
%% load int.CHGCAR id chg_int zpsp Al 3 As 5 O 6
!error! id chg_int zpsp Al 3 As 5 O 6
ERROR(setfield): Unknown extra keyword
ERROR(setfield): Unknown extra keyword
```
and fails.
This result does not depend on the system.
However, when using Pymatgen ver.2020.1.28, Critic2Caller works.
So, the update between ver.2020.1.28 and ver.2020.6.8 may break the code related to Critic2Caller.
The version of critic2 is the latest stable version (+ critic2 (stable), commit (config) 3081176) downloaded last Saturday.
**To Reproduce**
Steps to reproduce the behavior:
1. Copy the attached files to a directory.
2. Run the following command with the attached files. First argument is the path to the directory.
```
from pymatgen.command_line.critic2_caller import Critic2Caller
def create_critic2_output(path: str):
critic2_caller = Critic2Caller.from_path(path)
if __name__ == "__main__":
create_critic2_output(sys.argv[1])
```
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1890/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1890/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1891
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1891/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1891/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1891/events
|
https://github.com/materialsproject/pymatgen/pull/1891
| 649,737,904
|
MDExOlB1bGxSZXF1ZXN0NDQzMzU3NzA1
| 1,891
|
Fix zpsp to have element names as its keys
|
{
"login": "yuuukuma",
"id": 4986887,
"node_id": "MDQ6VXNlcjQ5ODY4ODc=",
"avatar_url": "https://avatars.githubusercontent.com/u/4986887?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/yuuukuma",
"html_url": "https://github.com/yuuukuma",
"followers_url": "https://api.github.com/users/yuuukuma/followers",
"following_url": "https://api.github.com/users/yuuukuma/following{/other_user}",
"gists_url": "https://api.github.com/users/yuuukuma/gists{/gist_id}",
"starred_url": "https://api.github.com/users/yuuukuma/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/yuuukuma/subscriptions",
"organizations_url": "https://api.github.com/users/yuuukuma/orgs",
"repos_url": "https://api.github.com/users/yuuukuma/repos",
"events_url": "https://api.github.com/users/yuuukuma/events{/privacy}",
"received_events_url": "https://api.github.com/users/yuuukuma/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Thanks!"
] | 2020-07-02T08:24:09
| 2020-07-02T17:40:15
|
2020-07-02T17:40:15Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This fix is needed as symbol shows the POTCAR label e.g. "Ca_pv" while element its name "Ca",
Fix #1890
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1891/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1891/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1891",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1891",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1891.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1891.patch",
"merged_at": "2020-07-02T17:40:15Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1892
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1892/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1892/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1892/events
|
https://github.com/materialsproject/pymatgen/issues/1892
| 649,768,961
|
MDU6SXNzdWU2NDk3Njg5NjE=
| 1,892
|
One should be able to put pymatgen.core.surface.Slab.as_dict() into a MongoDB database using pymongo.
|
{
"login": "MichaelWolloch",
"id": 66372013,
"node_id": "MDQ6VXNlcjY2MzcyMDEz",
"avatar_url": "https://avatars.githubusercontent.com/u/66372013?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/MichaelWolloch",
"html_url": "https://github.com/MichaelWolloch",
"followers_url": "https://api.github.com/users/MichaelWolloch/followers",
"following_url": "https://api.github.com/users/MichaelWolloch/following{/other_user}",
"gists_url": "https://api.github.com/users/MichaelWolloch/gists{/gist_id}",
"starred_url": "https://api.github.com/users/MichaelWolloch/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/MichaelWolloch/subscriptions",
"organizations_url": "https://api.github.com/users/MichaelWolloch/orgs",
"repos_url": "https://api.github.com/users/MichaelWolloch/repos",
"events_url": "https://api.github.com/users/MichaelWolloch/events{/privacy}",
"received_events_url": "https://api.github.com/users/MichaelWolloch/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Check out `jsanitize` in `monty.json` and see if that helps. There's a subtle distinction between `to_dict()` and `to_json()` methods, to insert into Mongo it needs to be JSON-compatible, you might try JSON with `MontyEncoder` too. Both these methods should just work without manual code tweaks required. If this doesn't work, let me know.",
"Thanks a lot for the quick reply and the options to solve my problem. Both `monty.json.jsanitize(Slab.as_dict(), allow_bson=True) ` and `json.dumps(Slab, cls=monty.json.MontyEncoder)` can be inserted into MongoDB without problems.\r\n\r\nI think I will to stick to `jsanitize`, as I can use the same code to load `Structures` and `Slabs` from the DB without rewriting stuff to feature `json.dumps` and `json.loads`.\r\nAlso, while `json.loads(json_slab, cls=monty.json.MontyDecoder)` produces a valid `pymatgen.core.surface.Slab` object, I am not getting the expected results. E.g. for the `Slab` object from `json.loads` I get a `dict` for `Slab.scale_factor`:\r\n~~~\r\n{'@module': 'numpy',\r\n '@class': 'array',\r\n 'dtype': 'int64',\r\n 'data': [[0, 0, -1], [1, -1, 0], [0, -1, 0]]}\r\n~~~\r\nwhile I get a list of lists with the `jsanitize` option. I probably are making a mistake again, somewhere with json, but this is not too important for now and I guess a `list of lists` is closer to the original `numpy.ndarray` than a `dict`, even if the `dict` contains all the information of the original array.\r\n\r\nThanks again for the help!",
"I pushed a fix for this.\r\n"
] | 2020-07-02T09:09:23
| 2020-07-03T15:12:10
|
2020-07-03T09:12:08Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
**Is your feature request related to a problem? Please describe.**
When I try to put a `Slab.as_dict()` into a MongoDB database within a FireWorks workflow it fails. The `as_dict()` method has to be MSON serializable according to the documentation, but apparently for pymongo this is not enough.
The problematic entries for me where:
1. `site['properties']['bulk_equivalent']` for site in` Slab.as_dict()['sites']` and for site in `Slab.as_dict()['oriented_unit_cell']['sites']`. These are of type` numpy.int32` which pymongo can not encode.
2. `Slab.as_dict()['scale_factor']` which has type `numpy.ndarray`
**Describe the solution you'd like**
I would appreciate if the `as_dict()` method of `pymatgen.core.surface.Slab` would result in a dictionary that has only data types encodable by pymongo. This would mean to change some types from `numpy.int32` to `int` and `numpy.ndarray` to `list`.
If needed, the `from_dict()` method should also be adapted to convert the types back.
**Describe alternatives you've considered**
I have written a small function to change the problematic types myself before database insertion, and I can confirm that I can create a Slab object again with the from_dict() method. However, I have not yet tested if there might be a problem when using the slabs (e.g. for the creation of an interface) later on, which could be true especially for the scale_factor which remains a list of lists. Of course another small function to change the types back would be sufficient to solve this problem, but overall this is not very elegant and I think to do it inside the `as_dict() `and `to_dict()` methods would be better and cleaner.
|
{
"login": "MichaelWolloch",
"id": 66372013,
"node_id": "MDQ6VXNlcjY2MzcyMDEz",
"avatar_url": "https://avatars.githubusercontent.com/u/66372013?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/MichaelWolloch",
"html_url": "https://github.com/MichaelWolloch",
"followers_url": "https://api.github.com/users/MichaelWolloch/followers",
"following_url": "https://api.github.com/users/MichaelWolloch/following{/other_user}",
"gists_url": "https://api.github.com/users/MichaelWolloch/gists{/gist_id}",
"starred_url": "https://api.github.com/users/MichaelWolloch/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/MichaelWolloch/subscriptions",
"organizations_url": "https://api.github.com/users/MichaelWolloch/orgs",
"repos_url": "https://api.github.com/users/MichaelWolloch/repos",
"events_url": "https://api.github.com/users/MichaelWolloch/events{/privacy}",
"received_events_url": "https://api.github.com/users/MichaelWolloch/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1892/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1892/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1893
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1893/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1893/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1893/events
|
https://github.com/materialsproject/pymatgen/pull/1893
| 650,436,879
|
MDExOlB1bGxSZXF1ZXN0NDQzOTI3NjQ3
| 1,893
|
Avoid division by zero if nb == 0
|
{
"login": "mfherbst",
"id": 11157893,
"node_id": "MDQ6VXNlcjExMTU3ODkz",
"avatar_url": "https://avatars.githubusercontent.com/u/11157893?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mfherbst",
"html_url": "https://github.com/mfherbst",
"followers_url": "https://api.github.com/users/mfherbst/followers",
"following_url": "https://api.github.com/users/mfherbst/following{/other_user}",
"gists_url": "https://api.github.com/users/mfherbst/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mfherbst/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mfherbst/subscriptions",
"organizations_url": "https://api.github.com/users/mfherbst/orgs",
"repos_url": "https://api.github.com/users/mfherbst/repos",
"events_url": "https://api.github.com/users/mfherbst/events{/privacy}",
"received_events_url": "https://api.github.com/users/mfherbst/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[] | 2020-07-03T08:31:59
| 2020-07-03T15:18:47
|
2020-07-03T15:17:25Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Correct bugfix if `nb` happens to come out as zero for a particular branch.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1893/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1893/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1893",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1893",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1893.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1893.patch",
"merged_at": "2020-07-03T15:17:25Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1894
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1894/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1894/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1894/events
|
https://github.com/materialsproject/pymatgen/issues/1894
| 650,482,228
|
MDU6SXNzdWU2NTA0ODIyMjg=
| 1,894
|
SlabGenerator missed some unique surfaces
|
{
"login": "lixinyuu",
"id": 33710988,
"node_id": "MDQ6VXNlcjMzNzEwOTg4",
"avatar_url": "https://avatars.githubusercontent.com/u/33710988?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lixinyuu",
"html_url": "https://github.com/lixinyuu",
"followers_url": "https://api.github.com/users/lixinyuu/followers",
"following_url": "https://api.github.com/users/lixinyuu/following{/other_user}",
"gists_url": "https://api.github.com/users/lixinyuu/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lixinyuu/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lixinyuu/subscriptions",
"organizations_url": "https://api.github.com/users/lixinyuu/orgs",
"repos_url": "https://api.github.com/users/lixinyuu/repos",
"events_url": "https://api.github.com/users/lixinyuu/events{/privacy}",
"received_events_url": "https://api.github.com/users/lixinyuu/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"The code is correct. In your first example, there are TWO surfaces - one is the FePt terminated and the other is CuPt terminated. In your second figure, that is not even stoichiometric FeCuPt2 anymore. If you want to make sure the surfaces are symmetrized, you need the Tasker 2 reconstructed slabs.",
"Thanks Professor. I understand it now."
] | 2020-07-03T09:48:29
| 2020-07-03T15:33:09
|
2020-07-03T14:50:14Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Hi Pymatgen Developers,
I am using Pymatgen to generate the (1, 0, 0) surfaces of mp-3702 FeCuPt2 crystal, and I get confused why only one CuPt terminated surface is generated. Theoretically, there are two unique surfaces (terminated at (1, 0, 0) and (0.5, 0, 0) respectively)
```
from pymatgen.ext.matproj import MPRester
from pymatgen.core.surface import SlabGenerator
with MPRester(api_key='******') as rester:
structure = rester.get_structure_by_material_id('mp-3702')
slab_gen = SlabGenerator(initial_structure=structure,
miller_index=(1, 0, 0),
min_slab_size=6.,
min_vacuum_size=12.,)
slabs = slab_gen.get_slabs()
len(slabs)
```
We can see that the geometry of the generated slab is
**Expected behavior**
However, a slab which is terminated by FePt should be generated as well. I tried to use (2, 0, 0) as miller index, but it doesn't work as well.
Could anyone give me some suggestions here? Thanks very much.
- OS: Linux
- Version 2019.11.11
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1894/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1894/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1895
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1895/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1895/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1895/events
|
https://github.com/materialsproject/pymatgen/issues/1895
| 650,675,374
|
MDU6SXNzdWU2NTA2NzUzNzQ=
| 1,895
|
'decomposition' enthalpy for stable compounds as an alternative to 'e_above_hull' in Phase Diagrams
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"What you call \"decomposition\" enthalpy is simply some formation energy. This can be obtained from the Reaction Calculator quite easily. But if you want the functions that are reported by the paper, you can ask @computron ",
"Yep it's just a formation energy and for anything that isn't a vertex of the convex hull it's the same as `e_above_hull`. From an epistemological viewpoint, it's interesting as it doesn't preference points on the \"known\" convex hull more than points inside the \"known\" convex hull. However, for vertices of the convex hull it's non-trivial to work out what the reaction is. The authors above solved it as a linear programming problem to minimize the formation energy of a set of compounds in the relevant phase diagram that satisfy conservation of mass with the vertex being removed. \r\n\r\nThe question is if I added translated the functionality of the original work into `pymatgen` would it be taken as a useful addition? For all I know once I start looking at it there may well be design clashes which mean it can't practically be implemented in `pymatgen` even if it might be a reasonable fit but if it's not a good fit then it's best to know that earlier rather than later.\r\n\r\n",
"Hi @CompRhys, if I understand the definition correctly, for entries on the hull, this is the difference in formation energy at that composition for the convex hull without the entry present vs the formation energy of that entry. I agree this is a useful measure (it's useful in testing for database errors too, in fact, since spurious data can manifest as very high decomposition energies).\r\n\r\nTo directly answer your question:\r\n\r\n> The question is if I added translated the functionality of the original work into pymatgen would it be taken as a useful addition? \r\n\r\nYes, I think this would be a good addition! Especially if implemented in a clean, efficient way. Perhaps add it to `PhaseDiagram` and just call it `get_decomposition_energy` or similar, rather than confusing with `e_above_hull`. There should be no design clashes.",
"Hi Rhys,\n\nWhat you are describing sounds a lot like\n\"get_equilibrium_reaction_energy\", which is also in PhaseDiagram. You can\nthink of this as the energy 'below the hull', in other words, if the stable\nphase was removed, what is the reaction energy from the components where\nthe hull *would* have been.\n\nBest,\nWenhao Sun\n\nOn Tue, Jul 7, 2020 at 9:49 PM Matthew Horton <notifications@github.com>\nwrote:\n\n> Hi @CompRhys <https://github.com/CompRhys>, if I understand the\n> definition correctly, for entries on the hull, this is the difference in\n> formation energy at that composition for the convex hull without the entry\n> present vs the formation energy of that entry. I agree this is a useful\n> measure (it's useful in testing for database errors too, in fact, since\n> spurious data can manifest as very high decomposition energies).\n>\n> To directly answer your question:\n>\n> The question is if I added translated the functionality of the original\n> work into pymatgen would it be taken as a useful addition?\n>\n> Yes, I think this would be a good addition! Especially if implemented in a\n> clean, efficient way. Perhaps add it to PhaseDiagram and just call it\n> get_decomposition_energy or similar, rather than confusing with\n> e_above_hull. There should be no design clashes.\n>\n> —\n> You are receiving this because you are subscribed to this thread.\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/issues/1895#issuecomment-655231769>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAN4NI4MUQNJ3KUFXBKF6UDR2PGCPANCNFSM4OP5M2SQ>\n> .\n>\n\n\n-- \nAssistant Professor\nMaterials Science and Engineering\nUniversity of Michigan\nhttp://mse.engin.umich.edu/people/fac/whsun\nhttp://wenhaosun.github.io\n",
"Thanks @wenhaosun :)\r\n\r\nYes, that's it, I didn't realize it was already implemented. It looks like the way this is currently implemented is the literal way as described in my previous message; that is, it calculates the convex hull with that entry absent. This seems a bit inefficient, especially for large phase diagrams, since presumably it can be calculated much more easily just by consideration of the two neighboring stable entries(?) So perhaps room for improvement here.",
"@mkhorton I don't think there is an efficient algo for this. But feel free to think through it. For binaries, it is clear that it should be the two nearest neighbor entries. That is not true in high dimensions.\r\nPersonally, I never like this value as a descriptor. There is no hard evidence this value means anything in terms of how synthesizable a compound is.",
"Ah, true, that sounds plausible. I was only thinking about the binary diagram.",
"One tricky aspect of this situation:\n\nConsider you have the following convex hull, and you are evaluating the\n'inverse hull energy' of AB2. Suppose there is a metastable AB phase.\n\nShould the function return the red line, or the green line? If you are just\nremoving the AB2 phase, it will return the green line. But perhaps it ought\nto return the red line. It depends on what kind of question you are trying\nto ask.\n\nJust something to consider!\n\n[image: image.png]\n\n\n\n\n\n\n\nOn Tue, Jul 7, 2020 at 10:46 PM Shyue Ping Ong <notifications@github.com>\nwrote:\n\n> @mkhorton <https://github.com/mkhorton> I don't think there is an\n> efficient algo for this. But feel free to think through it. For binaries,\n> it is clear that it should be the two nearest neighbor entries. That is not\n> true in high dimensions.\n> Personally, I never like this value as a descriptor. There is no hard\n> evidence this value means anything in terms of how synthesizable a compound\n> is.\n>\n> —\n> You are receiving this because you were mentioned.\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/issues/1895#issuecomment-655248349>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAN4NI6FOUMSELH2PJBIZMTR2PMWTANCNFSM4OP5M2SQ>\n> .\n>\n\n\n-- \nAssistant Professor\nMaterials Science and Engineering\nUniversity of Michigan\nhttp://mse.engin.umich.edu/people/fac/whsun\nhttp://wenhaosun.github.io\n",
"@wenhaosun I think your attachment got removed",
"Oops here's a link to the picture.\nhttps://drive.google.com/file/d/1gN_1hfdZrT2QSEswjUZ1EnphEWqHiWYs/view?usp=sharing\n\n\n\nOn Tue, Jul 7, 2020 at 11:20 PM Matthew Horton <notifications@github.com>\nwrote:\n\n> @wenhaosun <https://github.com/wenhaosun> I think your attachment got\n> removed\n>\n> —\n> You are receiving this because you were mentioned.\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/issues/1895#issuecomment-655258299>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAN4NIZLVYNJNN5VPNMHU63R2PQX7ANCNFSM4OP5M2SQ>\n> .\n>\n\n\n-- \nAssistant Professor\nMaterials Science and Engineering\nUniversity of Michigan\nhttp://mse.engin.umich.edu/people/fac/whsun\nhttp://wenhaosun.github.io\n",
"> Hi Rhys, What you are describing sounds a lot like \"get_equilibrium_reaction_energy\", which is also in PhaseDiagram. You can think of this as the energy 'below the hull', in other words, if the stable phase was removed, what is the reaction energy from the components where the hull *would* have been. Best, Wenhao Sun\r\n> -- Assistant Professor Materials Science and Engineering University of Michigan http://mse.engin.umich.edu/people/fac/whsun http://wenhaosun.github.io\r\n\r\nThank you for pointing this function out. Having checked the function more carefully I think it gives the distance to the red line in your plot as it only constructs the new hull from entries that are stable. I am interested in the distance to the green line which I think requires you to consider all the lowest energy polymorphs left in the space after you remove the compound in question.\r\n\r\nI have been reading up around ways to compute this value more efficiently than by reconstructing additional hulls (I think this would be prohibitively expensive to calculate for all of materials project even if factorising the phase diagrams). From what I've found because we do not need the simplices of the perturbed hull for other things I think the constrained optimisation problem over other materials in the (factorised) phase diagram used in the Bartel et al. paper is indeed the cheapest approach. \r\n\r\nIf you had the Delaunay triangulation of the space then I think you might be able to restrict the points considered by the optimisation further to only include points that are on a simplex including the stable compound in question. However, jumping straight to that might be attempting to run before being able to walk. "
] | 2020-07-03T15:58:19
| 2020-10-03T03:28:48
|
2020-10-03T03:28:48Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
`e_above_hull` is zero for everything on the hull. Whereas the decomposition enthalpy provides a distribution of values for compounds on the hull, therefore, making it potentially more useful for some workflows.
**Describe the solution you'd like**
add a function similar to `get_decomp_and_e_above_hull` that calculates the competing compounds and the decomposition enthalpy. It may be able to nest these functions as the decomposition enthalpy is equivalent to `e_above_hull` for unstable systems (as far as I understand).
**Describe alternatives you've considered & Additional context**
This issue is inspired by see https://arxiv.org/pdf/2001.10591.pdf. The authors release code to replicate their work here https://github.com/CJBartel/TestStabilityML.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1895/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1895/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1896
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1896/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1896/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1896/events
|
https://github.com/materialsproject/pymatgen/pull/1896
| 652,625,468
|
MDExOlB1bGxSZXF1ZXN0NDQ1NjYzNDY0
| 1,896
|
Integration of CP2K into Pymatgen IO
|
{
"login": "nwinner",
"id": 8825901,
"node_id": "MDQ6VXNlcjg4MjU5MDE=",
"avatar_url": "https://avatars.githubusercontent.com/u/8825901?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/nwinner",
"html_url": "https://github.com/nwinner",
"followers_url": "https://api.github.com/users/nwinner/followers",
"following_url": "https://api.github.com/users/nwinner/following{/other_user}",
"gists_url": "https://api.github.com/users/nwinner/gists{/gist_id}",
"starred_url": "https://api.github.com/users/nwinner/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/nwinner/subscriptions",
"organizations_url": "https://api.github.com/users/nwinner/orgs",
"repos_url": "https://api.github.com/users/nwinner/repos",
"events_url": "https://api.github.com/users/nwinner/events{/privacy}",
"received_events_url": "https://api.github.com/users/nwinner/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Just adding a note that I'll have some requests on the Bader portion of this code, seems like there's a lot of duplicated code from the current Bader caller. It seems like it'd be neater just to tidy up the code to factor out the common code (eg ACF.dat parsing) and allow it to initialize from either from Chgcar or Cube.\r\n\r\nWill defer to @shyamd on all matters CP2K in terms of code review but this is looking great.",
"Hi @nwinner, is this still a WIP or ready for review? I saw the title changed but also some test failures",
"Hey Matt, I need i pushed and changed the title, assuming there wouldn't be\nany test failures. Not sure what is going on, ill take a look.\n\nOn Fri, Oct 9, 2020, 7:21 PM Matthew Horton <notifications@github.com>\nwrote:\n\n> Hi @nwinner <https://github.com/nwinner>, is this still a WIP or ready\n> for review? I saw the title changed but also some test failures\n>\n> —\n> You are receiving this because you were mentioned.\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/pull/1896#issuecomment-706470463>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/ACDKYLIRIVSCA7EXZIL5AULSJ7AKNANCNFSM4OTSJTOQ>\n> .\n>\n",
"> Hi @nwinner, is this still a WIP or ready for review? I saw the title changed but also some test failures\r\n\r\nIts ready for review now. Shyam helped discover some errors in referencing the test files that didn't show up on my machine.",
"Thanks @nwinner and @shyamd for the review, this is a very substantial PR, looks great and happy to be merging it"
] | 2020-07-07T20:39:56
| 2020-10-19T19:52:34
|
2020-10-19T19:52:30Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Re-opening a work in progress PR for my CP2K module. Main feature is pymatgen.io.cp2k, which includes modules for reading/writing cp2k input files, input sets, and output parsing.
1. **pymatgen.io.cp2k.inputs**: Handles the CP2K input file structure by mainly defining a "Section" and a "Keyword". Keywords are arguments passed to CP2K with generic format. Sections are how different parts of CP2K are activated. Example, "DFT" section must be included to start a DFT calculation. Section is very similar to a dict, but has been instantiated as a standalone class because it needs to allow for a combination of differences including the need to repeat certain types of sections.
2. **pymatgen.io.cp2k.sets**: Creates CP2K input sets as child classes of the Section objects. Currently includes basics like Static, geometry optimization, hybrid calculation, etc. Is a little different from how this is done in VASP.
3. **pymatgen.io.cp2k.outputs**: one "master" cp2k output parser written here to parse structures, energies, DOS, etc.
4. **pymatgen.io.cp2k.utils**: a few utils for the above three modules.
## TODO (if any)
1. Main thing is that CP2K has very diverse input and output, and so we can *add* to the existing module in the future. The Section/Cp2kInput objects are robust enough that they should have no backwards compatibility issues as we do this. The output parser can only handle standard DFT type outputs right now, not the more exotic functionality of cp2k.
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1896/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1896/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1896",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1896",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1896.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1896.patch",
"merged_at": "2020-10-19T19:52:30Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1897
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1897/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1897/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1897/events
|
https://github.com/materialsproject/pymatgen/pull/1897
| 652,761,041
|
MDExOlB1bGxSZXF1ZXN0NDQ1Nzg0MzQz
| 1,897
|
Make EwaldSummation MSONable
|
{
"login": "lbluque",
"id": 9788715,
"node_id": "MDQ6VXNlcjk3ODg3MTU=",
"avatar_url": "https://avatars.githubusercontent.com/u/9788715?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/lbluque",
"html_url": "https://github.com/lbluque",
"followers_url": "https://api.github.com/users/lbluque/followers",
"following_url": "https://api.github.com/users/lbluque/following{/other_user}",
"gists_url": "https://api.github.com/users/lbluque/gists{/gist_id}",
"starred_url": "https://api.github.com/users/lbluque/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/lbluque/subscriptions",
"organizations_url": "https://api.github.com/users/lbluque/orgs",
"repos_url": "https://api.github.com/users/lbluque/repos",
"events_url": "https://api.github.com/users/lbluque/events{/privacy}",
"received_events_url": "https://api.github.com/users/lbluque/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"If anything in the implementation looks fishy please comment...",
"Thanks but I am unclear about the value of this. The EwaldSummation is an analysis. Typically people execute it once for a particular structure. They may want to store the final results of it in some form (son or otherwise) but I fail to see when I would want to reinitialize a EwaldSummation object. There is no additional analysis that can be done?",
"We are using the ewald matrices as part of Monte Carlo sampling. Most often just the total energy matrix, but having the separate parts allows finer grained control in sampling, having access to the properties defined in the class is quite useful for this.\r\n\r\nFor this use case I could definitely just save all the matrices separately, and then write functions to get the total matrix, energies, etc, but this would essentially be re-implementing the class properties in the EwaldSummation. I guess it comes down to this value extending further than our particular use case...",
"Thanks for the explanation. But I guess my question is how you are using the ewald matrices. You are using them unchanged right? What you really need is an output method. as_dict can be implemented. What I don't see is why we need a from_dict method since the dict (which can be json serialized) can be read in with a two-line code and used quite easily.",
"Thanks for the quick reply!\r\n\r\nYes, we use them unchanged. The reason for the from_dict is to recreate without needing to recompute a previously constructed set of matrices, for future sampling runs.",
"Ah ok, I see what you mean. I put it in there to handle the two cases where the matrices have been and have not been computed.\r\n\r\nHere, I am out of my understanding comfort zone. Can this be automatically handled by proper serialization?\r\n\r\n(not having the matrices computed is just a side-effect of the lazy implementation I used to be able to create instances without calling the computing methods, may not be the best design)",
"I see. Ok. I will merge.",
"Thanks @shyuep \r\nI tried using `functools.cached_property` to clean this up, but since some of the properties (i.e. real & point) are computed at the same time, I could not find a way around it."
] | 2020-07-08T00:06:48
| 2020-07-14T16:53:48
|
2020-07-14T15:06:24Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Computing the Ewald matrices for large structures can be very time consuming. By making the `EwaldSummation` class `MSONable` we can at least save time by reloading instances without recreating the matrices.
* `as_dict` method
* `from_dict` method
* To implement the above I changed the constructor to initialize private ewald parts as None and lazy compute when requested.
## Checklist
- [X] Code is in the [standard Python style](https://www.python.org/dev/peps/pep-0008/).
Run [pycodestyle](https://pycodestyle.readthedocs.io/en/latest/) and [flake8](http://flake8.pycqa.org/en/latest/)
on your local machine.
- [X] Docstrings have been added in the [Google docstring format](https://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html).
Run [pydocstyle](http://www.pydocstyle.org/en/2.1.1/index.html) on your code.
- [X] Type annotations are **highly** encouraged. Run [mypy](http://mypy-lang.org/)
to type check your code.
- [X] Tests have been added for any new functionality or bug fixes.
- [X] All existing tests pass.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1897/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1897/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1897",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1897",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1897.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1897.patch",
"merged_at": "2020-07-14T15:06:24Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1898
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1898/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1898/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1898/events
|
https://github.com/materialsproject/pymatgen/issues/1898
| 653,013,713
|
MDU6SXNzdWU2NTMwMTM3MTM=
| 1,898
|
Simple way to test if a structure is disordered?
|
{
"login": "mm04926412",
"id": 21348397,
"node_id": "MDQ6VXNlcjIxMzQ4Mzk3",
"avatar_url": "https://avatars.githubusercontent.com/u/21348397?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mm04926412",
"html_url": "https://github.com/mm04926412",
"followers_url": "https://api.github.com/users/mm04926412/followers",
"following_url": "https://api.github.com/users/mm04926412/following{/other_user}",
"gists_url": "https://api.github.com/users/mm04926412/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mm04926412/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mm04926412/subscriptions",
"organizations_url": "https://api.github.com/users/mm04926412/orgs",
"repos_url": "https://api.github.com/users/mm04926412/repos",
"events_url": "https://api.github.com/users/mm04926412/events{/privacy}",
"received_events_url": "https://api.github.com/users/mm04926412/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"What format are your structure files? Structure.is_ordered is the property method you are looking for.\r\nFor questions regarding usage, pls use https://matsci.org/c/pymatgen/9"
] | 2020-07-08T06:19:42
| 2020-07-08T14:44:51
|
2020-07-08T14:44:50Z
|
NONE
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
Apologies if github is the wrong place to ask operational questions.
I have a large list of structures and I need to quickly identify the ones with partial occupancies and parse them / clean up. Is there a quick test for if a crystal contains partial occupancies?
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1898/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1898/timeline
| null |
completed
| false
| false
|
{
"url": "",
"html_url": "",
"diff_url": "",
"patch_url": "",
"merged_at": ""
}
|
||
https://api.github.com/repos/materialsproject/pymatgen/issues/1899
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1899/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1899/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1899/events
|
https://github.com/materialsproject/pymatgen/pull/1899
| 653,557,972
|
MDExOlB1bGxSZXF1ZXN0NDQ2NDUzNzIx
| 1,899
|
[WIP] Adding decomposition energy function to Phase Diagram
|
{
"login": "CompRhys",
"id": 26601751,
"node_id": "MDQ6VXNlcjI2NjAxNzUx",
"avatar_url": "https://avatars.githubusercontent.com/u/26601751?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/CompRhys",
"html_url": "https://github.com/CompRhys",
"followers_url": "https://api.github.com/users/CompRhys/followers",
"following_url": "https://api.github.com/users/CompRhys/following{/other_user}",
"gists_url": "https://api.github.com/users/CompRhys/gists{/gist_id}",
"starred_url": "https://api.github.com/users/CompRhys/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/CompRhys/subscriptions",
"organizations_url": "https://api.github.com/users/CompRhys/orgs",
"repos_url": "https://api.github.com/users/CompRhys/repos",
"events_url": "https://api.github.com/users/CompRhys/events{/privacy}",
"received_events_url": "https://api.github.com/users/CompRhys/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
] | null |
[
"```\r\nPDEntry : N4 Zn12 Zr12 with energy = -21.6244\r\n\r\nDecomposition reaction:\r\n(PDEntry : N1 Zr1 with energy = -3.5118, 1.8295715595655295)\r\n(PDEntry : N2 Zn2 Zr6 with energy = -9.2150, 1.0852142202171597)\r\n(PDEntry : Zn4 Zr2 with energy = -1.8542, 0.5739288989128745)\r\n(PDEntry : Zn6 Zr2 with energy = -2.3478, 1.255642660652196)\r\n\r\nEquilibrium reaction:\r\n(PDEntry : N2 Zn2 Zr6 with energy = -9.2150, 0.23809523809523814)\r\n(PDEntry : Zn4 Zr2 with energy = -1.8542, 0.5714285714285714)\r\n(PDEntry : N1 Zr1 with energy = -3.5118, 0.19047619047619047)\r\n```\r\n\r\n@mkhorton While exploring the options I have here I was benchmarking against `get_equilibrium_reaction_energy()`. If I restrict to just using the points on the hull as the competing points my `get_decomposition_energy()` function should be equivalent to `get_equilibrium_reaction_energy()` for points on the hull, however, I found several examples where they give different results (see one above). More importantly the decomposition from `get_equilibrium_reaction_energy()` doesn't have the correct stoichiometry which suggests that `get_decomp_and_e_above_hull(..., allow_negative=True)` is not robust. \r\n\r\nGiven the naming confusion with calling this the decomposition energy I could potentially replace the equilibrium reaction function with my function and add a kwarg to switch between the two behaviours therefore eliminating the robustness issue. The one issue would be that if such a change was excepted I would ideally want the decomposition case to be the default. This would introduce a breaking change requiring a code update for users currently using the other function.",
"For reference the issue with speed I am seeing is not likely to be a problem for materials project, in materials project I believe the largest space is ~ 100 materials for \"Li-Mn-Rh-O\" whilst the dataset I was using has 4000 entries in the \"Zn-Zr-N\" space (all bar ~15 of these being unstable) ",
"Thanks @CompRhys, having a busy week but let me know when you think this is ready for review and I'll take a look.\r\n\r\nIn terms of large spaces, you can check out Si-O (though that's lower dimensionality). Interesting you're looking at Zr-Zn-N, we have someone in the group looking at this system, can put you in touch if you're interested.",
"@mkhorton I think it's ready for review, there are a few comments I left in the code which I will try to address based on feedback. The two big things are what to name the function `get_decomposition_energy` is potentially confusing and then this issue with the decomposition being wrong for some examples with `get_reaction_equilibrium energy` (minimal data to reproduce this\r\n[pymatgen-fail.txt](https://github.com/materialsproject/pymatgen/files/4907034/pymatgen-fail.txt)). I am not sure what to do about that but perhaps a check should be added to `get_decomp_and_e_above_hull` to ensure the decomposition has a valid stoichiometry. \r\n\r\nThe tests I have added are just to ensure that all the different types of entries are treated correctly and give the right sign for the answer.",
"Can you make sure you are up to date with the latest master branch first? We will look at the code after that.",
"Having given this another look I think that the stoichiometry for the strange test case I found is actually fine. I think I was confused by the documentation, it's slightly misleading as for `get_decomp_and_e_above_hull(...)` in the dictionary of `{c: amt}` returned `amt` is not the amount of the composition but the amount of the fractional composition as the calculations are done using the per atom energies not the composition energies. I have changed the docstrings to make this clearer.\r\n\r\nIn light of this, I think that `get_reaction_equilibrium energy` does function as intended and the discrepancy arises due to some geometrical quirk I haven't quite worked my head around yet. The effect can be somewhat understood by comparing the phase diagrams with and without Zn3Zr3N.\r\n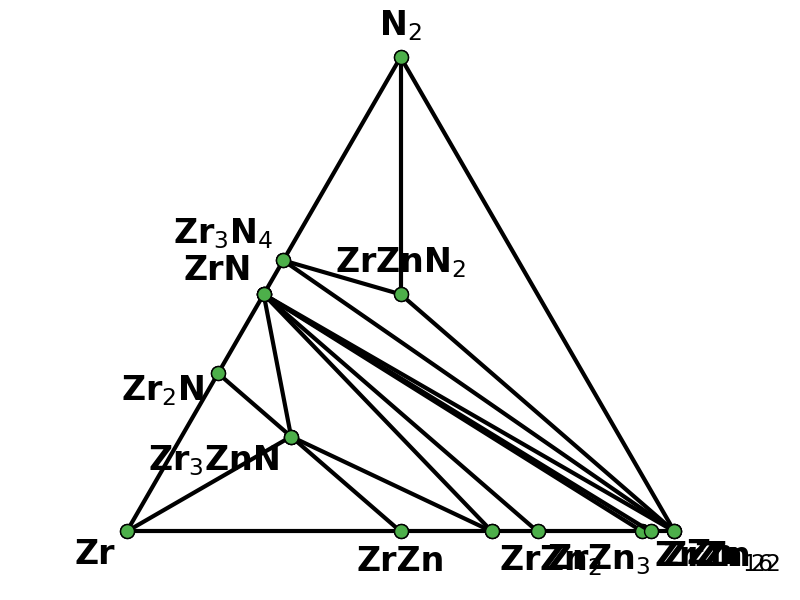\r\n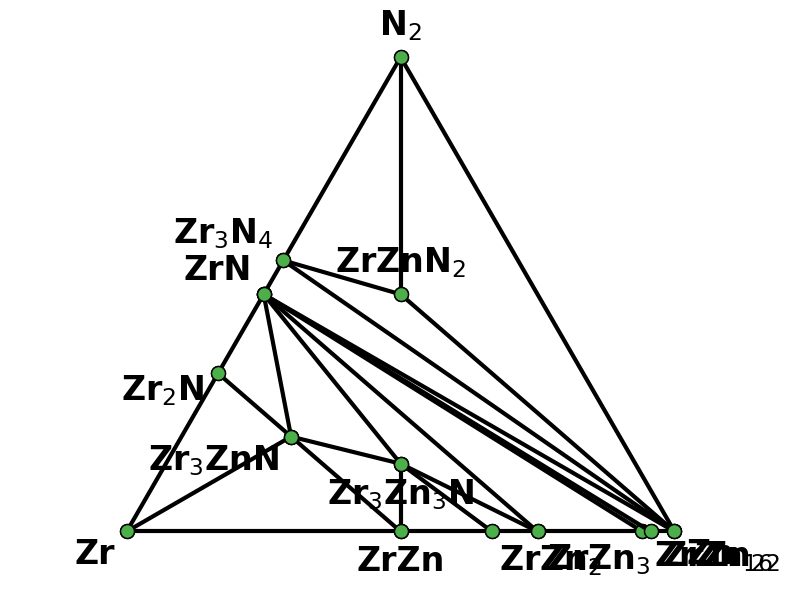\r\n\r\n",
"I have a project going on that uses the get_equilibrium_reaction_energy\nquite extensively.\n\nBefore this gets merged onto the Master branch, Rhys could you give a\ndescription of what the objective of your contribution is? It is not\nentirely clear what you're trying to accomplish, based on what I've seen in\nthese pymatgen discussion threads.\n\n\nBest,\nWenhao Sun\n\nOn Wed, Jul 22, 2020 at 7:19 PM Matthew Horton <notifications@github.com>\nwrote:\n\n> *@mkhorton* commented on this pull request.\n>\n> Apologies for the slowness to review, I think this looks about ready to\n> merge though. How do you want to reconcile with\n> get_equilibrium_reaction_energy? Given their similarity, could roll these\n> into one function with an algo kwarg to let user choose the method?\n> Unless there are good reasons to keep them separate.\n>\n> —\n> You are receiving this because you are subscribed to this thread.\n> Reply to this email directly, view it on GitHub\n> <https://github.com/materialsproject/pymatgen/pull/1899#pullrequestreview-453748529>,\n> or unsubscribe\n> <https://github.com/notifications/unsubscribe-auth/AAN4NI45MCKJI5I3CC6H2ADR45XWTANCNFSM4OU432AQ>\n> .\n>\n\n\n-- \nAssistant Professor\nMaterials Science and Engineering\nUniversity of Michigan\nhttp://mse.engin.umich.edu/people/fac/whsun\nhttp://wenhaosun.github.io\n",
"@wenhaosun I want to attempt to address limitations of my model raised with regard to this energy in the referenced paper. Therefore I need the energy referenced in the paper. Why this energy over `reaction equilibrium energy` is a matter of (the original authors) taste.\r\n\r\nBoth energies seem somewhat fictitious as @shyuep raised. I am not sure the context in which you're using the energy but I don't yet know of physical reason to have that constraint on the number of linked materials. Could you point me to your work using it for interest? \r\n\r\n@mkhorton If I can pin down the constraint I could add this additional constraint via a kwarg and merge the methods but only if that's something viewed as beneficial from @wenhaosun and the rest of the maintainers team. Slsqp should be faster than computing the second hull but there could be edge cases.",
"@mkhorton @wenhaosun After another attempt to get to grips with this observed discrepancy it turns out it's not a geometric effect after all it's actually an artefact of poor optimisation (This in part explains why I couldn't get my head around it before). \r\n\r\nThe problem was that the tolerance values I took from the original paper's implementation (`1e-4`) turn out to be too loose. If I reduce the tolerance down to `1e-10` the two methods give the same energies for the \"problematic\" Zn3Zr3N entry. If I profile the code the slsqp method with this tighter tolerance it still turns out to be 4-5x faster on the testing set I am using. \r\n\r\nAssuming that the only issue was the loose tolerance the two methods still differ with regard to what @wenhaosun raised in #1895 which is that this method gives the green energy and `get_equilibrium_reaction_energy` gives the red energy. It should be pretty easy to add a kwarg to switch between these behaviour patterns.\r\n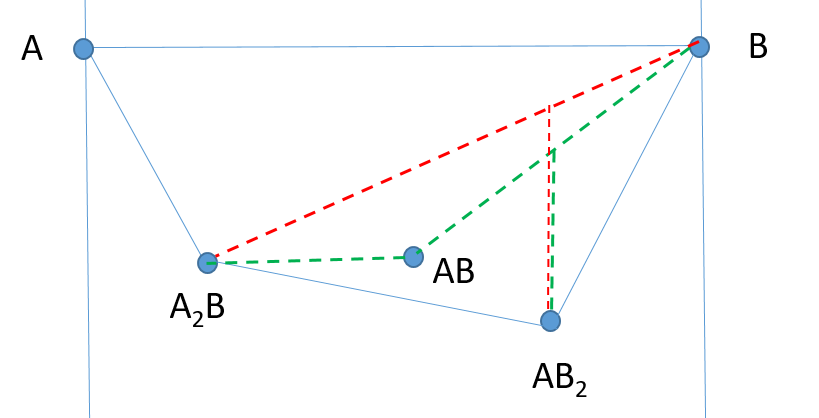\r\n",
"Thanks @CompRhys, @wenhaosun just to clarify, the old behavior will not be going away, this PR adds a new feature.\r\n\r\nThere still remains the question for the name to distinguish these two cases.",
"@mkhorton @shyuep I have amended the name of the function to `get_quasi_e_to_hull` and expanded upon the docstring to make it abundantly clear what the function does. \r\n\r\nThe final thing that occurred to be before asking for this to be merged is that in phase diagrams with duplicated stable entries or when introducing a 'novel' stable duplicate after computation of the phase diagram there are a lot of edge cases that need to be accounted for. I need to introduce tests to treat such edge cases. However, currently it is not clear how duplicates are handled as it doesn't appear to explicit as the choice is made within scipy/qhull. Do you think there would be any merit in handling duplicates explicitly before the call to scipy? This would potentially make it much easier to deal with these edge cases.",
"Thanks, for the duplicates, are you specifically concerned about literal duplicates (same composition, same energy)? I'm not sure duplicates would generally cause any issue with regular phase diagram construction. I would not be against an explicit check during PhaseDiagram construction for duplicates.",
"If we have a literal duplicate it ends up putting one as being stable and another as being unstable. Fortunately, we have a duplicated stable entry already in the unit test file so it's easy to show this\r\n\r\n```python\r\nfrom pymatgen.entries.entry_tools import EntrySet\r\nfrom pymatgen.analysis.phase_diagram import (\r\n PhaseDiagram,\r\n PDEntry,\r\n)\r\n\r\nfrom pymatgen.analysis.tests.test_phase_diagram import module_dir\r\n\r\nentries = EntrySet.from_csv(str(module_dir / \"pdentries_test.csv\"))\r\npd = PhaseDiagram(entries)\r\n\r\ns_lio = [e for e in pd.stable_entries if e.name == \"Li2O\"][0]\r\nun_lio = [e for e in pd.unstable_entries if e.name == \"Li2O\"]\r\n\r\nprint([e == s_lio for e in un_lio])\r\n\r\nduplicate_entry = PDEntry(\"Li2O\", -14.31361175)\r\n\r\nprint(s_lio == duplicate_entry)\r\n\r\n# NOTE added in edit - this is what causes the test to fail\r\n# but given that is it equal to an entry from stable entries I am\r\n# no sure why is it not 'in' it?\r\nprint(duplicate_entry not in pd.stable_entries)\r\n```\r\n```bash\r\n(base) reag2@Laptop:~/PhD/second-year/pymatgen$ python dupl_test.py\r\n[False, False, False, True, False]\r\nTrue\r\nTrue\r\n```\r\nSeeing this behaviour I have written this test where I introduce a novel duplicate of the stable entry and it currently fails but I do not understand why.\r\n\r\n```python\r\nduplicate_entry = PDEntry(\"Li2O\", -14.31361175)\r\nstable_entry = [e for e in self.pd.stable_entries if e.name == \"Li2O\"][0]\r\n\r\nself.assertEqual(\r\n self.pd.get_quasi_e_to_hull(duplicate_entry), self.pd.get_quasi_e_to_hull(stable_entry),\r\n \"Novel duplicates of stable entries should have same decomposition energy!\")\r\n```\r\n\r\n",
"The problem arises due to [this set vs list issue on SO](https://stackoverflow.com/questions/8705378/pythons-in-set-operator/8705406#8705406). \r\n\r\nThere are two ways I can see around this both of which might be breaking changes for some people\r\n\r\n1. change `__hash__` for the `PDEntry` class -- I don't like this because duplicates are still independent things.\r\n1. cast the set `pd._stable_entries` to a list in `pd.stable_entries` -- which would remove the unstable duplicates between `pd.stable_entries` and `pd.unstable_entries`.\r\n\r\nMy preference would be for option 2 but it breaks some of the PDPlotter tests as casting to a list means that `len(pd.stable_entries) + len(pd.unstable_entries) != len(pd.all_entries)` in cases where `pd.all_entries` contains duplicates which is the case for the dataset used to test.\r\n",
"What's the objection to 1? A better hash method might be good, provided that the same hash is generated only for exact equivalence of PDEntries and not almost equivalence. Being able to cast to a set to remove exact duplicates seems like a useful thing.",
"Option 1. just seems as though it was designed to allow the duplicated to be distinguished by the hash as it uses the `id(...)` this makes me think that there would be additional implications. \r\n\r\nIf we go with option 1 of changing the hash there are two sets that are referenced to fixed numbers than need to be changed to account for the 2 duplicated in the unit testing entry set (I was expecting them to be flagged by the CI here as I didn't change them yet but that doesn't appear to be the case?). \r\n\r\nI am also not sure if the grand potential and other phase diagram entry types might need alternative hashes to just the fractional composition and energy per atom",
"> Option 1. just seems as though it was designed to allow the duplicated to be distinguished by the hash as it uses the id(...) this makes me think that there would be additional implications.\r\n\r\nAs far as I know, the default hash is based on an object's id anyway, so I'm not sure why that was added specifically. There's another reason Option 1 may be bad though, and that's that Entries are mutable (e.g. see normalize() method), so going the hash route seems like it might be bug-prone.\r\n\r\n> I am also not sure if the grand potential and other phase diagram entry types might need alternative hashes to just the fractional composition and energy per atom\r\n\r\nThis in and of itself wouldn't have been a big issue though, in that a good hash should be dependent only on the object itself (eg the Entry) and not on what it's used for.\r\n\r\nRegarding this note:\r\n\r\n```\r\n# NOTE added in edit - this is what causes the test to fail\r\n# but given that is it equal to an entry from stable entries I am\r\n# no sure why is it not 'in' it?\r\nprint(duplicate_entry not in pd.stable_entries)\r\n```\r\n\r\nThis isn't working because the `in` is checking based on object identity, e.g. this will work as expected:\r\n\r\n`any([duplicate_entry == e for e in pd.stable_entries])`",
"> As far as I know, the default hash is based on an object's id anyway, so I'm not sure why that was added specifically. There's another reason Option 1 may be bad though, and that's that Entries are mutable (e.g. see normalize() method), so going the hash route seems like it might be bug-prone.\r\n\r\nI tried to avoid potential problems with `Entry.normalise()` by using the `fractional_composition` and the `energy_per_atom` in the hash? These should remain the same even if the `Entry` gets normalised?\r\n\r\n> This isn't working because the in is checking based on object identity, e.g. this will work as expected:\r\n`any([duplicate_entry == e for e in pd.stable_entries])`\r\n\r\nThis is basically what option 2 is as here we are unpacking `pd.stable_entries` as a list but the problem here is that if we only make this change in the functions that I wrote we will still end up with the situation that if duplicates are present one will be in stable entries and the other will be in unstable entries which seems like a more general issue to me and thinking about it more the better solution seems like ensuring that duplicates hash to the same value provided that we can pre-empt the possible bugs.",
"> I tried to avoid potential problems with Entry.normalise() by using the fractional_composition and the energy_per_atom in the hash? These should remain the same even if the Entry gets normalised?\r\n\r\nJust so we're not talking cross-purposes, when I'm talking about the hash I'm specifically referring to the `__hash__` method on the object. In general, if an object is mutable, it's not appropriate to define a hash because it can lead to undefined behavior in Python, since the hash is computed once when adding the object to other data structures and not re-computed when the object changes.\r\n\r\nEven if we don't use the `normalize()` method in this specific code, the fact that the method exists at all means that we shouldn't define a custom hash function on `Entry` based on its contents since it could lead to bugs elsewhere. Therefore, I'd suggest reverting https://github.com/materialsproject/pymatgen/pull/1899/commits/2cf92d84d440c37aa6b3f1efb0ea86b09fad09a1.\r\n\r\nIn the broader sense of a \"hash\" meaning \"how do I group a set of objects, e.g. using `groupby` or similar\", that's a separate discussion.\r\n\r\nMy apologies for being in favour of an alternative hash method previously, I hadn't properly thought it through for this object (I was assuming the Entries were immutable; perhaps they should be, but given that they're not I think that rules out the hash solution).\r\n\r\n---\r\n\r\nI think having a `remove_duplicates` option in the `__init__` might be the cleanest option here, default to False for backwards compatibility, or just handling duplicates specifically in `get_quasi_e_to_hull` if that's the concern.",
"And to add, I understand the point that the hash as-written here would be the same whether or not `normalize` had been called, but the object itself would have changed, so correctly the hash should also change. Even though this would be fine here, I could imagine someone else writing code with Entries where this might cause a bug.",
"> I think having a `remove_duplicates` option in the `__init__` might be the cleanest option here, default to False for backwards compatibility, or just handling duplicates specifically in `get_quasi_e_to_hull` if that's the concern.\r\n\r\nthis would suggest that having duplicated entries in both stable and unstable entries is a feature not a bug? It would involve a fair amount of overhead to handle everything to do with duplicates inside the function but would be possible if this isn't a bug.\r\n\r\nHowever, I do see it as more of a bug as I can't see why anyone would expect this behaviour (my preferred solution here would be going back to making `pd.stable_entries` a list as in 2. so that we test on dictionary equality rather than hash equality and then updating some of the other tests to reflect). I actually think the test for `get_e_above_hull` is failing to test the function properly in this regard.\r\n\r\n```python\r\ndef test_get_e_above_hull(self):\r\n for entry in self.pd.stable_entries:\r\n self.assertLess(self.pd.get_e_above_hull(entry), 1e-11,\r\n \"Stable entries should have e above hull of zero!\")\r\n\r\n for entry in self.pd.all_entries:\r\n if entry not in self.pd.stable_entries:\r\n e_ah = self.pd.get_e_above_hull(entry)\r\n self.assertGreaterEqual(e_ah, 0)\r\n self.assertTrue(isinstance(e_ah, Number))\r\n```\r\n\r\ngiven the first part of the test shouln't the second part be `self.assertGreaterEqual(e_ah, 1e-11)` to be consistent? or maybe`self.assertGreater(e_ah, 0)` based on our understanding that only stable entries should have `e_above_hull == 0`?\r\n\r\nI am sorry for opening what might be a can of worms here",
"@mkhorton I haven't looked at this in a while but took some time today and I think i've managed to resolve all the remaining issues and edge cases without changing functionality elsewhere beyond needing to introduce a non-inplace normalize option for the Entry object (this is causing a linting error somewhere). Can you please give this a review?",
"Hi @CompRhys, I'm sorry for the radio silence from my end on this too, it slipped from my inbox. I'll take a look and review",
"FYI the very final `pylint` lint step has been causing issues lately; this is unrelated to this PR. My apologies for this. As long as there aren't linter failures for the specific files touched in this PR it's not a problem.",
"Many thanks @CompRhys, this looks good! Sorry it took so long to merge in. Regarding the naming, that can be changed if necessary, but no good alternative occurred to me and it seemed silly to wait on the PR for that reason alone. Thanks for the updates to the docstrings too and including a comprehensive docstring for the new functionality.\r\n\r\nPlease do make sure to include your details in [this form](https://forms.gle/JnisFb38QDR8QTFTA) so that you can be credited appropriately in the [pymatgen documentation](https://pymatgen.org/team.html) too."
] | 2020-07-08T19:58:46
| 2020-10-03T03:30:02
|
2020-10-03T03:28:49Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
This PR addresses Issue #1895.
The finalised code should add the functionality to calculate the "decomposition" energy for materials on the convex hull. This is slightly different from the "equilibrium reaction" energy already implemented as it allows for compositions not previously on the hull to be involved in the energy prediction. The solution is obtained by solving a linear programming problem subject to the constraint on the number of moles of each species. The SLSQP method used from SciPy doesn't scale favourably to large competing spaces as the code currently looks at all applicable entries for the optimisation. If the Delaunay triangulation was available we could look at all the vertices in simplices containing the target point however this is very expensive to compute for just this purpose. PR will remain a draft until a reasonable solution is found.
The name decomposition energy may be confused with the `get_decomp_and_e_above_hull()` function and so alternative names are welcome.
## Additional dependencies introduced (if any)
None as of yet, potentially alternative optimizers from `pyOpt` (http://www.pyopt.org/) may scale better than SciPy SLSQP but I would rather solve the current issue by formulating a more physically-constrained problem than by using an alternative optimizer.
## TODO
- [x] Unit Tests need to be added
- [x] Investigate further reductions that can be placed on the space of competing elements
Closes #1895
|
{
"login": "mkhorton",
"id": 2976580,
"node_id": "MDQ6VXNlcjI5NzY1ODA=",
"avatar_url": "https://avatars.githubusercontent.com/u/2976580?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mkhorton",
"html_url": "https://github.com/mkhorton",
"followers_url": "https://api.github.com/users/mkhorton/followers",
"following_url": "https://api.github.com/users/mkhorton/following{/other_user}",
"gists_url": "https://api.github.com/users/mkhorton/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mkhorton/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mkhorton/subscriptions",
"organizations_url": "https://api.github.com/users/mkhorton/orgs",
"repos_url": "https://api.github.com/users/mkhorton/repos",
"events_url": "https://api.github.com/users/mkhorton/events{/privacy}",
"received_events_url": "https://api.github.com/users/mkhorton/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1899/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1899/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1899",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1899",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1899.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1899.patch",
"merged_at": "2020-10-03T03:28:48Z"
}
|
|||
https://api.github.com/repos/materialsproject/pymatgen/issues/1900
|
https://api.github.com/repos/materialsproject/pymatgen
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1900/labels{/name}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1900/comments
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1900/events
|
https://github.com/materialsproject/pymatgen/pull/1900
| 654,088,091
|
MDExOlB1bGxSZXF1ZXN0NDQ2ODgyODcz
| 1,900
|
Changed tolerance for lattice match when creating defect objects
|
{
"login": "mbkumar",
"id": 4041360,
"node_id": "MDQ6VXNlcjQwNDEzNjA=",
"avatar_url": "https://avatars.githubusercontent.com/u/4041360?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/mbkumar",
"html_url": "https://github.com/mbkumar",
"followers_url": "https://api.github.com/users/mbkumar/followers",
"following_url": "https://api.github.com/users/mbkumar/following{/other_user}",
"gists_url": "https://api.github.com/users/mbkumar/gists{/gist_id}",
"starred_url": "https://api.github.com/users/mbkumar/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/mbkumar/subscriptions",
"organizations_url": "https://api.github.com/users/mbkumar/orgs",
"repos_url": "https://api.github.com/users/mbkumar/repos",
"events_url": "https://api.github.com/users/mbkumar/events{/privacy}",
"received_events_url": "https://api.github.com/users/mbkumar/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
[] |
closed
| false
|
{
"login": "",
"id": 0,
"node_id": "",
"avatar_url": "",
"gravatar_id": "",
"url": "",
"html_url": "",
"followers_url": "",
"following_url": "",
"gists_url": "",
"starred_url": "",
"subscriptions_url": "",
"organizations_url": "",
"repos_url": "",
"events_url": "",
"received_events_url": "",
"type": "",
"user_view_type": "",
"site_admin": false
}
|
[] | null |
[
"Can we have a unittest pls, since the claim is that the lattice match is causing functionality failure.",
"Unit test added demonstrating the failure with default lattice match and success with looser absolute tolerance.",
"Thanks."
] | 2020-07-09T14:05:19
| 2020-07-09T16:46:33
|
2020-07-09T16:46:29Z
|
CONTRIBUTOR
|
{
"total": 0,
"completed": 0,
"percent_completed": 0
}
|
## Summary
Stringent atol of 1e-8 used in numpy.allclose function for lattice match is causing problems for users. The tolerance for lattice match check for defect instantiation is now made looser. This fixes many failures experienced by users of pycdt and atomate defect workflows
* Instead of using default lattice match function of pymatgen, direct call to np.allclose function is made. A looser absolute tolerance of 1e-5 is supplied inplace of the default 1e-8 in the call to np.allclose.
|
{
"login": "shyuep",
"id": 577107,
"node_id": "MDQ6VXNlcjU3NzEwNw==",
"avatar_url": "https://avatars.githubusercontent.com/u/577107?v=4",
"gravatar_id": "",
"url": "https://api.github.com/users/shyuep",
"html_url": "https://github.com/shyuep",
"followers_url": "https://api.github.com/users/shyuep/followers",
"following_url": "https://api.github.com/users/shyuep/following{/other_user}",
"gists_url": "https://api.github.com/users/shyuep/gists{/gist_id}",
"starred_url": "https://api.github.com/users/shyuep/starred{/owner}{/repo}",
"subscriptions_url": "https://api.github.com/users/shyuep/subscriptions",
"organizations_url": "https://api.github.com/users/shyuep/orgs",
"repos_url": "https://api.github.com/users/shyuep/repos",
"events_url": "https://api.github.com/users/shyuep/events{/privacy}",
"received_events_url": "https://api.github.com/users/shyuep/received_events",
"type": "User",
"user_view_type": "public",
"site_admin": false
}
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/issues/1900/reactions",
"total_count": 0,
"+1": 0,
"-1": 0,
"laugh": 0,
"hooray": 0,
"confused": 0,
"heart": 0,
"rocket": 0,
"eyes": 0
}
|
https://api.github.com/repos/materialsproject/pymatgen/issues/1900/timeline
| null | true
| false
|
{
"url": "https://api.github.com/repos/materialsproject/pymatgen/pulls/1900",
"html_url": "https://github.com/materialsproject/pymatgen/pull/1900",
"diff_url": "https://github.com/materialsproject/pymatgen/pull/1900.diff",
"patch_url": "https://github.com/materialsproject/pymatgen/pull/1900.patch",
"merged_at": "2020-07-09T16:46:29Z"
}
|
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.